WO2025007148A1 - Polymer lipid nanoparticle compositions for delivering circular polynucleotides - Google Patents
Polymer lipid nanoparticle compositions for delivering circular polynucleotides Download PDFInfo
- Publication number
- WO2025007148A1 WO2025007148A1 PCT/US2024/036442 US2024036442W WO2025007148A1 WO 2025007148 A1 WO2025007148 A1 WO 2025007148A1 US 2024036442 W US2024036442 W US 2024036442W WO 2025007148 A1 WO2025007148 A1 WO 2025007148A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- composition
- polymer
- virus
- lipid
- monomer
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K9/00—Medicinal preparations characterised by special physical form
- A61K9/10—Dispersions; Emulsions
- A61K9/127—Synthetic bilayered vehicles, e.g. liposomes or liposomes with cholesterol as the only non-phosphatidyl surfactant
- A61K9/1271—Non-conventional liposomes, e.g. PEGylated liposomes or liposomes coated or grafted with polymers
- A61K9/1272—Non-conventional liposomes, e.g. PEGylated liposomes or liposomes coated or grafted with polymers comprising non-phosphatidyl surfactants as bilayer-forming substances, e.g. cationic lipids or non-phosphatidyl liposomes coated or grafted with polymers
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K9/00—Medicinal preparations characterised by special physical form
- A61K9/0012—Galenical forms characterised by the site of application
- A61K9/0019—Injectable compositions; Intramuscular, intravenous, arterial, subcutaneous administration; Compositions to be administered through the skin in an invasive manner
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K9/00—Medicinal preparations characterised by special physical form
- A61K9/48—Preparations in capsules, e.g. of gelatin, of chocolate
- A61K9/50—Microcapsules having a gas, liquid or semi-solid filling; Solid microparticles or pellets surrounded by a distinct coating layer, e.g. coated microspheres, coated drug crystals
- A61K9/51—Nanocapsules; Nanoparticles
- A61K9/5107—Excipients; Inactive ingredients
- A61K9/5123—Organic compounds, e.g. fats, sugars
Definitions
- the present disclosure generally relates to polymeric nanoparticle compositions for encapsulating therapeutic agents, such as nucleic acids (e.g., circular polynucleotides), for timely release of the nucleic acid cargo.
- therapeutic agents such as nucleic acids (e.g., circular polynucleotides)
- Nucleic acid therapies available include, but are not limited to, the use of DNA or viral vectors for insertion of desired genetic information into the host cell, and/or RNA constructed to encode for a therapeutic protein.
- DNA and viral vector deliveries carry their own setbacks and challenges that make them less favorable to RNA therapeutics.
- the introduced DNA in some cases may be unintentionally inserted into an intact gene and result in a mutation that impedes or even wholly eliminates the function of the endogenous gene leading to an elimination or deleteriously reduced production of an essential enzyme or interruption of a gene critical for the regulating cell growth.
- RNA is substantially safer and more effective gene therapy agent due to its ability to encode for the protein outside of the nucleus to perform its function. With this, the RNA does not involve the risk of being stably integrated into the genome of the transfected cell.
- RNA therapeutics conventionally has consisted of engineering linear messenger RNAs (mRNA). Although more effective than DNA or viral vectors, linear mRNAs have their own set of challenges regarding the stability, immunogenicity, translation efficiency, and delivery. Some of these challenges may lead to size restraints and/or destruction of the linear mRNA due to the challenges present with linear mRNAs’ caps. Partly to overcome these limitations, circular polynucleotides or circular RNAs are increasingly being studied. Due to being covalently closed continuous loops, circular RNAs are useful in the design and production of stable forms of RNA.
- the present application provides, compositions comprising polymeric lipid nanoparticles, and methods for preparing the same.
- the polymeric lipid nanoparticles can comprise RNA polynucleotides, such as circular RNA polynucleotides (aka circRNA or oRNATM), ionizable lipids (lipid) and amphiphilic polymers, thereby forming a polymeric lipid nanoparticle composition.
- RNA polynucleotides such as circular RNA polynucleotides (aka circRNA or oRNATM), ionizable lipids (lipid) and amphiphilic polymers, thereby forming a polymeric lipid nanoparticle composition.
- the lipid forms a complex with the circRNA (circRNA-lipid complex) that is encapsulated in the nanoparticles.
- the circRNA-lipid complex is encapsulated in the core of the nanoparticles and the amphiphilic polymer provides a uniform shell around the circRNA-lipid complex.
- the present application also provides emulsions comprising circRNA, ionizable lipid (lipid) and an amphiphilic polymer useful for preparing the polymeric lipid nanoparticle compositions. Methods of preparing the subject emulsions and compositions are also provided.
- composition comprising a plurality of polymeric lipid nanoparticles, each nanoparticle comprising: a RNA polynucleotide, such as a circular RNA polynucleotide; an ionizable lipid; and an amphiphilic polymer.
- an emulsion comprising an aqueous continuous phase and an organic dispersed phase, wherein the organic dispersed phase comprises droplets that contain: a RNA polynucleotide, such as a circular RNA polynucleotide; an ionizable lipid; and an amphiphilic polymer.
- a RNA polynucleotide such as a circular RNA polynucleotide
- an ionizable lipid such as an amphiphilic polymer
- FIG. 1 is a schematic showing a general flow of a process for the manufacture of subject polymer lipid nanoparticle compositions using an emulsion-based method.
- FIG. 2 is a schematic illustrating the quench step of an emulsion-based method for the manufacture of subject polymer lipid nanoparticle compositions (i.e., adding a subject emulsion to an aqueous bath).
- FIG. 3 is a graphic representation of the core shell of the polymeric lipid nanoparticle containing the circRNA-CFL pair at the core of the nanoparticle and a polymer shell.
- FIG. 4A illustrates RNA integrity data for both, mRNA (mfLuc) and circRNA (ofLuc) after being processed through the high shear process.
- FIG.4B illustrates RNA integrity data for a complex of circRNA (ofLuc) and an ionizable lipid (ethyl lauroyl arginate (EL A)) (ofLuc: EL A) at various ratios.
- FIG. 5B is a graph illustrating the encapsulation efficiency (EE%) for various polymer lipid nanoparticle compositions prepared using the emulsion-based process of Example 2 with various ratios of ionizable lipid (e.g., ELA) to amphiphilic polymer (P) (ELA:P of 2:1, 4:1, and 6:1).
- ELA ionizable lipid
- P amphiphilic polymer
- FIG. 6A is a cryogenic transmission electron microscopy (Cryo-TEM) image of a polymer lipid nanoparticle composition prepared using the emulsion-based process of Example 2 with an ELA : amphiphilic polymer ratio (ELA:P) of 4:1, including the core-shell morphology.
- cryo-TEM cryogenic transmission electron microscopy
- FIG. 6B is a Cryo-TEM image of a composition of ELA and circRNA complex without the inclusion of an amphiphilic polymer.
- FIG. 7 is a graph showing the in vitro release kinetics of the circRNA from polymer lipid nanoparticles performed at 37 °C in PBS over a span of 0 to 6 days.
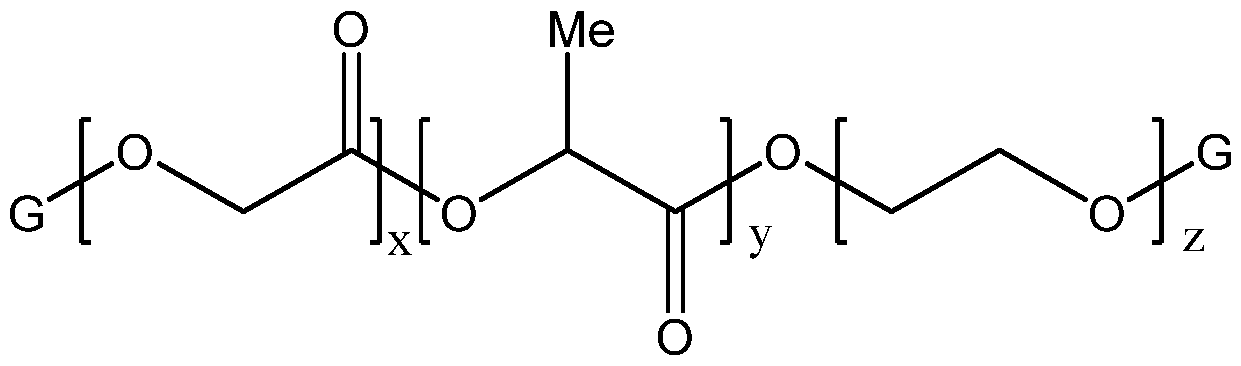
- the polymer lipid nanoparticle composition is made with a 10k-5k polylactic acid-polyethylene glycol polymer (i.e., di-block polymer composed of 10k PLA and 5k PEG) and ELA as the ionizable lipid.
- FIG. 10k-5k polylactic acid-polyethylene glycol polymer i.e., di-block polymer composed of 10k PLA and 5k PEG
- ELA the ionizable lipid
- FISH fluorescence in situ hybridization
- FIG. 10 depicts body weight of the mice injected intravenously with either PLNP GalNAc3, an LNP comprising an ionizable lipid (positive control) (indicated as “Ionizable LNP 1”), or a 20% sucrose (negative control) (indicated as “control”) and circular RNA encoding Flue.
- FIG. 12 is a graphic representation of the glutathione mediated disruption of a polymeric lipid nanoparticle comprising a shell of cross-linked amphiphilic polymers having disulfide linkages.
- FIG. 14 is a graphic representation of polymeric lipid nanoparticles prepared using tri-block amphiphilic polymers.
- FIG. 16 depicts particle size distribution (Z-average, nm), polydispersity index (PDI), encapsulation efficiencies (in %) for PLNPs comprising an ionizable lipid (i.e., ethyl lauroyl arginate (ELA), ionizable lipid 2, or ionizable lipid 3) at either post-quench, post-TFF purification, or postparticle concentration step (indicated as “PQ,” “pTFF,” and “PC” respectively).
- ionizable lipid i.e., ethyl lauroyl arginate (ELA), ionizable lipid 2, or ionizable lipid 3
- FIG. 17 depicts article size distribution (Z-average, nm), polydispersity index (PDI), encapsulation efficiencies (in %) for PLNPs comprising an ionizable lipid (i.e., ethyl lauroyl arginate or ionizable lipid 2).
- ionizable lipid i.e., ethyl lauroyl arginate or ionizable lipid 2.
- FIG. 18 is a circular RNA release curve providing the percentage of circular RNA released in vitro over the span of 8 days for PLNPs comprising either ionizable lipid ethyl lauroyl arginate or ionizable lipid 2.
- FIG. 19A is an ion pair reverse phase (IPRP) high performance liquid chromatograph (HPLC) showing circular RNA integrity of circular RNAs encoding firefly luciferase formulated in a sodium phosphate buffer solution. Intact circular RNAs are labeled as “circular.” Non-intact circular RNA products are labeled as “nicked.”
- FIG. 20 depicts RNA integrity over the span of 7 days of a circular RNA formulated in a sodium phosphate buffer solution (control, “unformulated circRNA”) or a PLNP comprising either ionizable lipid ethyl lauroyl arginate or ionizable lipid 2.
- RNA integrity was calculated from the intact circular RNA peak area under the curve (AUC) percentage present in the IPRP-HPLC chromatographs depicted in FIGs. 19A-19C.
- the dotted line in FIG. 20 indicates the RNA integrity of the circRNA prior to formulation.
- FIG. 21 depicts total ex vivo liver flux in Balb/C mice dosed with circular RNA formulated in a 20% sucrose solution or PLNP comprising either an ionizable lipid ethyl lauroyl arginate, ionizable lipid 2, or ionizable lipid 3 at dose of 4 mpk.
- FIG. 22B is a graph illustrating mean fluorescence signal calculated from FISH imaging of FIG. 22A for circular RNA-PLNP solutions (e.g., circular RNAs formulated with PLNP comprising ethyl lauroyl arginate or ionizable lipid 2 and dosed at 4 mpk or 8mpk) or circular RNA formulated with 20% sucrose solutions (control) at 6 hours post injection.
- circular RNA-PLNP solutions e.g., circular RNAs formulated with PLNP comprising ethyl lauroyl arginate or ionizable lipid 2 and dosed at 4 mpk or 8mpk
- 20% sucrose solutions control
- FIG. 23 is a western blot image of asialoglycoprotein receptor 1 (ASGPR1) expression in Hepa-lClC7, primary human hepatocytes (PHHs), and human skeletal muscle cells (HSKMs) 24 hours post transfection of a circular RNA encoding ASGPR1 (indicated as “1C1C7 Treated,” “PHH Treated,” and “HSKM Treated,” respectively). Endogenous ASGPR1 levels (control) for each of the cell types are indicated as “1C1C7,” “PHH,” and “HSKM.”
- ASGPR1 asialoglycoprotein receptor 1
- FIG. 25 illustrates in vitro luciferase expression in primary human hepatocytes treated with: (1) 50 ng of circular RNA encoding ASGPR (indicated as “oASGPR”); and/ or (2) PLNP unconjugated with GalNAc3 (indicated as “Unconj.”), PLNP conjugated with GalNAc3 (indicated as “GNAc”), or no PLNP transfer vehicle (indicated as “control”).
- PLNPs unconjugated or conjugated with GalNAc3 were formulated with either 0.5 pg, 1 pg or 1.5 p of circular RNA expressing firefly luciferase (fLuc).
- fLuc expression was determined 48 hours post treatment of circular RNA encoding fLuc.
- PHH cells were treated with oASGPR occurred for 6 hours (indicated as “6h txn”) or 24 hours (indicated as “24h txn”). Background signaling is shown as a dotted line.
- FIG. 27 depicts PLNP Z-average (in nm), polydispersity index (indicated as “PDI”) and encapsulation efficiency (indicated as a percentage) of PLNPs having alterative polymer shell compositions (i.e., Shell 1, Shell 2, Shell 3, and/or Shell 4) formulated with circular RNAs encoding firefly luciferase.
- Shell 1 comprised a 4:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer.
- Shell 2 comprised a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer.
- Shell 3 was comprised entirely of a 15K-5K PLGA-PEG block copolymer (the PLGA portion being a random copolymer of 75% lactic acid and 25% glycolic acid).
- Shell 4 comprised a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer, supplemented with 1% by mass 16K-5K PLA-PEG block copolymer.
- Control PLNP comprises ethyl lauroyl arginate (indicated as “control”).
- FIG. 29A is an RNA release curve showing circular RNA released in vitro expressed as a percentage of encapsulated RNA released over the span of 5 days for PLNPs comprising either Shell 1 , Shell 2, Shell 3, or Shell 4 (wherein Shells 1-4 are the same as provided in FIG. 27) and formulated with either 250 ng or 500 ng of circular RNA expressing firefly luciferase (fLuc).
- Control PLNP comprises ethyl lauroyl arginate (indicated as “control”).
- the functional half-life is determined by an in vivo assay, wherein levels of a protein encoded by the expression sequence of the circular RNA polynucleotide are measured in patient serum or tissue samples every 1, 2, 6, 12, or 24 hours over 1, 2, 3, 4, 5, 6, 7, or 14 days.
- the pre-determined threshold value is the functional half-life of a reference linear RNA polynucleotide comprising the same expression sequence as the circular RNA polynucleotide.
- the circular RNA provided herein may have a higher magnitude of expression than equivalent linear mRNA, e.g., a higher magnitude of expression 24 hours after administration of RNA to cells.
- the circular RNA provided herein has a higher magnitude of expression than mRNA comprising the same expression sequence, 5moU modifications, an optimized UTR, a cap, and/or a polyA tail.
- the circular RNA provided herein may be less immunogenic than an equivalent mRNA when exposed to an immune system of an organism or a certain type of immune cell.
- the circular RNA provided herein is associated with modulated production of cytokines when exposed to an immune system of an organism or a certain type of immune cell.
- the circular RNA provided herein is associated with reduced production of IFN-pi, RIG-I, IL-2, IL-6, IFNy, and/or TNFa when exposed to an immune system of an organism or a certain type of immune cell as compared to mRNA comprising the same expression sequence.
- the DNA template, precursor linear RNA polynucleotide and circular RNA provided herein comprise a first (5’) and/or a second (3’) spacer.
- the DNA template or precursor linear RNA polynucleotide comprises one or more spacers in the enhanced intron elements.
- the DNA template, precursor linear RNA polynucleotide comprises one or more spacers in the enhanced exon elements.
- the DNA template or linear RNA polynucleotide comprises a spacer in the 3’ enhanced intron fragment and a spacer in the 5’ enhanced intron fragment.
- DNA template, precursor linear RNA polynucleotide, or circular RNA comprises a spacer in the 3’ enhanced exon fragment and another spacer in the 5’ enhanced exon fragment to aid with circularization or protein expression due to symmetry created in the overall sequence.
- this additional spacer prevents the structured regions of the IRES or aptamer of a TIE from interfering with the folding of the 3’ group I intron fragment or reduces the extent to which this occurs.
- the 5’ spacer sequence is at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25 or 30 nucleotides in length. In some embodiments, the 5’ spacer sequence is no more than 100, 90, 80, 70, 60, 50, 45, 40, 35 or 30 nucleotides in length. In some embodiments the 5’ spacer sequence is between 5 and 50, 10 and 50, 20 and 50, 20 and 40, and/or 25 and 35 nucleotides in length.
- the 5’ spacer sequence is 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49 or 50 nucleotides in length.
- the 5’ spacer sequence is a polyA sequence.
- the 5’ spacer sequence is a poly AC sequence.
- a spacer comprises 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% poly AC content.
- a spacer comprises 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% polypyrimidine (C/T or C/U) content.
- the DNA template and precursor linear RNA polynucleotides and circular RNA polynucleotide provided herein comprise a first (5’) duplex region and a second (3’) duplex region.
- the DNA template and precursor linear RNA polynucleotide comprises a 5’ external duplex region located within the 3’ enhanced intron fragment and a 3’ external duplex region located within the 5’ enhanced intron fragment.
- the DNA template, precursor linear RNA polynucleotide and circular RNA polynucleotide comprise a 5’ internal duplex region located within the 3’ enhanced exon fragment and a 3’ internal duplex region located within the 5’ enhanced exon fragment.
- the DNA polynucleotide and precursor linear RNA polynucleotide comprises a 5’ external duplex region, 5’ internal duplex region, a 3’ internal duplex region, and a 3’ external duplex region.
- duplex regions on the ends of the precursor RNA strand, and adjacent or very close to the group I intron fragment bring the group I intron fragments in close proximity to each other, increasing splicing efficiency.
- the duplex regions are 3 to 100 nucleotides in length (e.g., 3-75 nucleotides in length, 3-50 nucleotides in length, 20-50 nucleotides in length, 35-50 nucleotides in length, 5-25 nucleotides in length, 9-19 nucleotides in length).
- the duplex regions are 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, or 50 nucleotides in length.
- the duplex regions have a length of 9 to 50 nucleotides.
- the duplex regions have a length of 9 to 19 nucleotides.
- the duplex regions have a length of 20 to 40 nucleotides.
- the duplex regions have a length of 30 nucleotides.
- the DNA template, precursor linear RNA polynucleotide, or circular RNA polynucleotide does not comprise of any duplex regions to optimize translation or circularization.
- the DNA template or precursor linear RNA polynucleotide may comprise an affinity tag.
- the affinity tag is located in the 3’ enhanced intron element.
- the affinity tag is located in the 5’ enhanced intron element.
- both (3’ and 5’) enhanced intron elements each comprise an affinity tag.
- an affinity tag of the 3’ enhanced intron element is the length as an affinity tag in the 5’ enhanced intron element.
- an affinity tag of the 3’ enhanced intron element is the same sequence as an affinity tag in the 5’ enhanced intron element.
- the affinity sequence is placed to optimize oligo-dT purification.
- an affinity tag comprises a polyA region.
- the polyA region is at least 15, 30, or 60 nucleotides long.
- one or both polyA regions is 15-50 nucleotides long.
- one or both polyA regions is 20-25 nucleotides long.
- the polyA sequence is removed upon circularization.
- an oligonucleotide hybridizing with the polyA sequence such as a deoxythymine oligonucleotide (oligo(dT)) conjugated to a solid surface (e.g., a resin), can be used to separate circular RNA from its precursor RNA.
- the 3’ enhanced intron element comprises a leading untranslated sequence.
- the leading untranslated sequence is a the 5’ end of the 3’ enhanced intron fragment.
- the leading untranslated sequence comprises of the last nucleotide of a transcription start site (TSS).
- TSS transcription start site
- the TSS is chosen from a viral, bacterial, or eukaryotic DNA template.
- the leading untranslated sequence comprise the last nucleotide of a TSS and 0 to 100 additional nucleotides.
- the TSS is a terminal spacer.
- the leading untranslated sequence contains a guanosine at the 5’ end upon translation of an RNA T7 polymerase.
- the 5’ enhanced intron element comprises a trailing untranslated sequence.
- the 5’ trailing untranslated sequence is located at the 3’ end of the 5’ enhanced intron element.
- the trailing untranslated sequence is a partial restriction digest sequence.
- the trailing untranslated sequence is in whole or in part a restriction digest site used to linearize the DNA template.
- the restriction digest site is in whole or in part from a natural viral, bacterial or eukaryotic DNA template.
- the trailing untranslated sequence is a terminal restriction site fragment.
- the 3’ enhanced intron element and 5’ enhanced intron element each comprise an intron fragment.
- a 3’ intron fragment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to a 3’ proximal fragment of a natural group I or II intron including the 3’ splice site dinucleotide.
- the 3’ intron fragment includes the first and second nucleotides of a 3’ group I or II intron fragment splice site dinucleotide; and the 5’ intron fragment includes the first and second nucleotides of a 3’ group I or II intron fragment dinucleotide.
- the 3’ enhanced intron element and 5’ enhanced intron element comprises a synthetic intron fragment.
- the DNA template, linear precursor RNA polynucleotide, and circular RNA polynucleotide each comprise an enhanced exon fragment.
- the 3’ enhanced exon element is located upstream to core functional element.
- the 5’ enhanced intron element is located downstream to the core functional element.
- the 3’ enhanced exon element and 5’ enhanced exon element each comprise an exon fragment.
- the 3’ enhanced exon element comprises a 3’ exon fragment.
- the 5’ enhanced exon element comprises a 5’ exon fragment.
- the 3’ exon fragment and 5’ exon fragment each comprises a group I or II intron fragment and 1 to 100 nucleotides of an exon sequence.
- a 3’ intron fragment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to a 3’ proximal fragment of a natural group I or II intron including the 3’ splice site dinucleotide.
- a 5’ group I or II intron fragment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to a 5’ proximal fragment of a natural group I or II intron including the 5’ splice site dinucleotide.
- the 3’ exon fragment comprises a second nucleotide of a 3’ group I or II intron splice site dinucleotide and 1 to 100 nucleotides of an exon sequence.
- the exon fragments located within the 5’ enhanced exon element and 3’ enhanced exon element does not comprise of a group I or II splice site dinucleotide.
- a 3’ enhanced intron element comprises in the following 5’ to 3’ order: a leading untranslated sequence, a 5’ affinity tag, an optional 5’ external duplex region, a 5’ external spacer, and a 3’ intron fragment.
- the 3’ enhanced exon element comprises in the following 5’ to 3’ order: a 3’ exon fragment, an optional 5’ internal duplex region, an optional 5’ internal duplex region, and a 5’ internal spacer.
- the 5’ enhanced exon element comprises in the following 5’ to 3’ order: a 3’ internal spacer, an optional 3’ internal duplex region, and a 5’ exon fragment.
- the 3’ enhanced intron element comprises in the following 5’ to 3’ order: a 5’ intron fragment, a 3’ external spacer, an optional 3’ external duplex region, a 3’ affinity tag, and a trailing untranslated sequence.
- the accessory element comprises an IRES transacting factor (ITAF) region.
- IRES transacting factor region modulates the initiation of translation through binding to PCBP1 - PCBP4 (polyC binding protein), PABP1 (polyA binding protein), PTB (polyprimidine tract binding), Argonaute protein family, HNRNPK (Heterogeneous nuclear ribonucleoprotein K protein), or La protein.
- the IRES transacting factor region comprises a polyA, polyC, poly AC, or polyprimidine track.
- the miRNA binding site is located within the spacer within the intron element or exon element. In certain embodiments, the miRNA binding site comprises the entire spacer regions. [0109] In some embodiments, the 5’ intron element and 3’ intron elements each comprise identical miRNA binding sites. In another embodiment, the miRNA binding site of the 5’ intron element comprises a different, in length or nucleotides, miRNA binding site than the 3’ intron element. In one embodiment, the 5’ exon element and 3’ exon element comprise identical miRNA binding sites. In other embodiments, the 5’ exon element and 3’ exon element comprises different, in length or nucleotides, miRNA binding sites.
- the modified nucleoside is m’A (1 -methyladenosine); m 2 A (2-methyladenosine); Am (2’-O-methyladenosine); ms 2 m 6 A (2-methylthio-N 6 -methyladenosine); i 6 A (N 6 -isopentenyladenosine); ms 2 i6A (2-methylthio-N 6 isopentenyladenosine); io 6 A (N 6 -(cis-hydroxyisopentenyl)adenosine); ms 2 io 6 A (2-methylthio-N 6 -(cis-hydroxyisopentenyl)adenosine); g 6 A (N 6 - glycinylcarbamoyladenosine); t 6 A (N 6 -threonylcarbamoyladenosine); ms 2 t 6 A (2-methylthio-N 6 - threon
- polynucleotides may be codon-optimized.
- a codon optimized sequence may be one in which codons in a polynucleotide encoding a polypeptide have been substituted in order to increase the expression, stability and/or activity of the polypeptide.
- the expression sequence encodes a therapeutic protein.
- the expression sequence encodes a cytokine, e.g., IL-12p70, IL-15, IL-2, IL-18, IL-21, IFN-a, IFN- P, IL-10, TGF-beta, IL-4, or IL-35, or a functional fragment thereof.
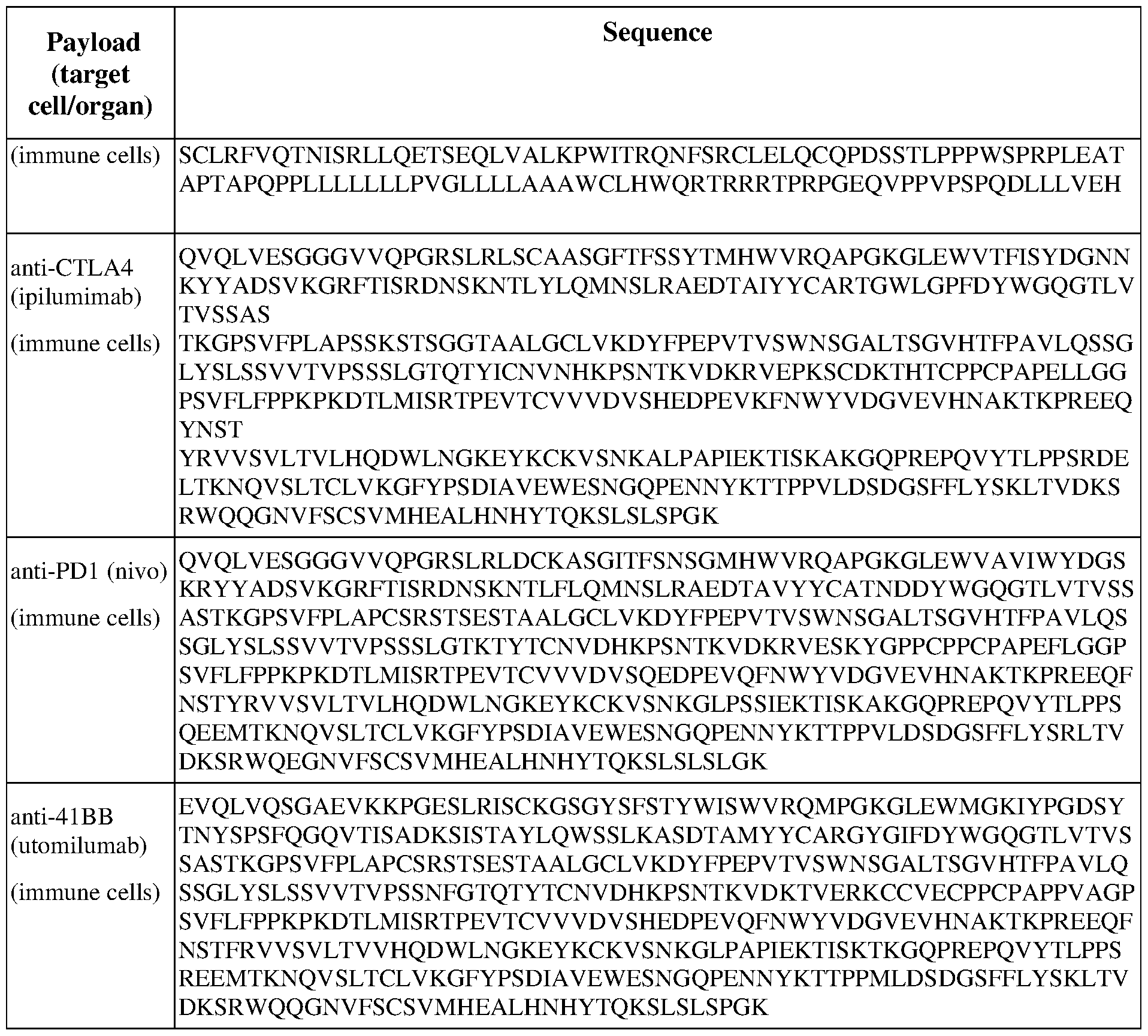
- the expression sequence encodes an immune checkpoint inhibitor.
- the expression sequence encodes an agonist (e.g., a TNFR family member such as CD137L, OX40L,
- the expression sequence encodes a secreted T cell or immune cell engager (e.g., a bispecific antibody such as BiTE, targeting, e.g., CD3, CD137, or CD28 and a tumor-expressed protein e.g., CD19, CD20, or BCMA etc.).
- a transcription factor e.g., FOXP3, HELIOS, TOX1, or T0X2.
- the expression sequence encodes an immunosuppressive enzyme (e.g., IDO or CD39/CD73).
- the expression sequence encodes a GvHD (e.g., anti-HLA- A2 CAR-Tregs).
- the antigen is selected from or derived from the group consisting of rotavirus, foot and mouth disease virus, influenza A virus, influenza B virus, influenza C virus, H1N1, H2N2, H3N2, H5N1, H7N7, H1N2, H9N2, H7N2, H7N3, H10N7, human parainfluenza type 2, herpes simplex virus, Epstein-Barr virus, varicella virus, porcine herpesvirus 1, cytomegalovirus, lyssavirus, Bacillus anthracis, anthrax PA and derivatives, poliovirus, Hepatitis A, Hepatitis B, Hepatitis C, Hepatitis E, distemper virus, Venezuelan equine encephalomyelitis, feline leukemia virus, reovirus, respiratory syncytial virus, Lassa fever virus, polyoma tumor virus, canine parvovirus, papilloma virus, tick borne encephalitis virus
- the antigenic polypeptide is a viral polypeptide from an adenovirus; Herpes simplex, type 1; Herpes simplex, type 2; encephalitis virus; papillomavirus; Varicella-zoster virus; Epstein-barr virus; Human cytomegalovirus; Human herpes virus, type 8; Human papillomavirus; BK virus; JC virus; Smallpox; polio virus; Hepatitis B virus; Human bocavirus; Parvovirus B 19; Human astro virus; Norwalk virus; coxsackievirus; hepatitis A virus; poliovirus; rhinovirus; Severe acute respiratory syndrome virus; Hepatitis C virus; Yellow Fever virus; Dengue virus; West Nile virus; Rubella virus; Hepatitis E virus; Human Immunodeficiency virus (HIV); Influenza virus; Guanarito virus; Junin virus; Lassa virus; Machupo virus; Sabia virus; Crime
- a polynucleotide encodes a protein that is made up of subunits that are encoded by more than one gene.
- the protein may be a heterodimer, wherein each chain or subunit of the protein is encoded by a separate gene. It is possible that more than one circRNA molecule is delivered in the transfer vehicle and each circRNA encodes a separate subunit of the protein. Alternatively, a single circRNA may be engineered to encode more than one subunit. In certain embodiments, separate circRNA molecules encoding the individual subunits may be administered in separate transfer vehicles.
- Additional polynucleotides, including expression sequences, and lipids are in WO2019236673; WO2020237227; WO2021113777; WO2021226597; WO2021189059; WO2021236855;
- Chimeric antigen receptors are genetically-engineered receptors. These engineered receptors may be inserted into and expressed by immune cells, including T cells via circular RNA as described herein. With a CAR, a single receptor may be programmed to both recognize a specific antigen and, when bound to that antigen, activate the immune cell to attack and destroy the cell bearing that antigen. When these antigens exist on tumor cells, an immune cell that expresses the CAR may target and kill the tumor cell.
- the CAR encoded by the polynucleotide comprises (i) an antigen-binding molecule that specifically binds to a target antigen, (ii) a hinge domain, a transmembrane domain, and an intracellular domain, and (iii) an activating domain.
- an orientation of the CARs in accordance with the disclosure comprises an antigen binding domain (such as an scFv) in tandem with a costimulatory domain and an activating domain.
- the costimulatory domain may comprise one or more of an extracellular portion, a transmembrane portion, and an intracellular portion. In other embodiments, multiple costimulatory domains may be utilized in tandem.
- CARs may be engineered to bind to an antigen (such as a cell-surface antigen) by incorporating an antigen binding molecule that interacts with that targeted antigen.
- the antigen binding molecule is an antibody fragment thereof, e.g., one or more single chain antibody fragment (scFv).
- scFv is a single chain antibody fragment having the variable regions of the heavy and light chains of an antibody linked together. See U.S. Patent Nos. 7,741,465, and 6,319,494 as well as Eshhar et al., Cancer Immunol Immunotherapy (1997) 45: 131-136.
- An scFv retains the parent antibody's ability to specifically interact with target antigen.
- scFvs are useful in chimeric antigen receptors because they may be engineered to be expressed as part of a single chain along with the other CAR components. Id. See also Krause et al., J. Exp. Med., Volume 188, No. 4, 1998 (619-626); Finney et al., Journal of Immunology, 1998, 161 : 2791-2797. It will be appreciated that the antigen binding molecule is typically contained within the extracellular portion of the CAR such that it is capable of recognizing and binding to the antigen of interest. Bispecific and multispecific CARs are contemplated within the scope of the present disclosure, with specificity to more than one target of interest.
- the antigen binding molecule comprises a nanobody. In some embodiments, the antigen binding molecule comprises a DARPin. In some embodiments, the antigen binding molecule comprises an anticalin or other synthetic protein capable of specific binding to target protein.
- the CAR comprises an antigen binding domain specific for an antigen selected from CD19, CD123, CD22, CD30, CD171, CS-1, C-type lectin-like molecule-1, CD33, epidermal growth factor receptor variant III (EGFRvIII), ganglioside G2 (GD2), ganglioside GD3, TNF receptor family member B cell maturation (BCMA), Tn antigen ((Tn Ag) or (GalNAca-Ser/Thr)), prostate-specific membrane antigen (PSMA), Receptor tyrosine kinase-like orphan receptor 1 (ROR1), Fms-Like Tyrosine Kinase 3 (FLT3), Tumor-associated glycoprotein 72 (TAG72), CD38, CD44v6, Carcinoembryonic antigen (CEA), Epithelial cell adhesion molecule (EPCAM), B7H3 (CD276), KIT (CD117), Interleukin- 13 receptor subunit alpha-2, me
- an antigen binding domain comprises SEQ ID NO: 321 and/or 322 of WO2023081526.
- a CAR comprises a hinge or spacer domain.
- the hinge/spacer domain may comprise a truncated hinge/spacer domain (THD) the THD domain is a truncated version of a complete hinge/spacer domain (“CHD”).
- an extracellular domain is from or derived from (e.g., comprises all or a fragment of) ErbB2, glycophorin A (GpA), CD2, CD3 delta, CD3 epsilon, CD3 gamma, CD4, CD7, CD8a, CD8[T CD1 la (IT GAL), CD1 lb (IT GAM), CD1 1c (ITGAX), CD1 Id (IT GAD), CD18 (ITGB2), CD19 (B4), CD27 (TNFRSF7), CD28, CD28T, CD29 (ITGB1), CD30 (TNFRSF8), CD40 (TNFRSF5), CD48 (SLAMF2), CD49a (ITGA1), CD49d (ITGA4), CD49f (ITGA6), CD66a (CEACAM1), CD66b (CEACAM8), CD66c (CEACAM6), CD66d (CEACAM3), CD66e (CEACAM5), CD69 (CLEC2), CD79A (B-cell
- a hinge or spacer domain is positioned between an antigen binding molecule (e.g., an scFv) and a transmembrane domain. In this orientation, the hinge/spacer domain provides distance between the antigen binding molecule and the surface of a cell membrane on which the CAR is expressed.
- a hinge or spacer domain is from or derived from an immunoglobulin.
- a hinge or spacer domain is selected from the hinge/spacer regions of IgGl, IgG2, IgG3, IgG4, IgA, IgD, IgE, IgM, or a fragment thereof.
- a hinge or spacer domain comprises, is from, or is derived from the hinge/spacer region of CD8 alpha. In some embodiments, a hinge or spacer domain comprises, is from, or is derived from the hinge/spacer region of CD28. In some embodiments, a hinge or spacer domain comprises a fragment of the hinge/spacer region of CD8 alpha or a fragment of the hinge/spacer region of CD28, wherein the fragment is anything less than the whole hinge/spacer region.
- the fragment of the CD8 alpha hinge/spacer region or the fragment of the CD28 hinge/spacer region comprises an amino acid sequence that excludes at least 1, at least 2, at least 3, at least 4, at least 5, at least 6, at least 7, at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 amino acids at the N-terminus or C-Terminus, or both, of the CD8 alpha hinge/spacer region, or of the CD28 hinge/spacer region.
- the CAR may further comprise a transmembrane domain and/or an intracellular signaling domain.
- the transmembrane domain may be designed to be fused to the extracellular domain of the CAR. It may similarly be fused to the intracellular domain of the CAR.
- the transmembrane domain that naturally is associated with one of the domains in a CAR is used.
- the transmembrane domain may be selected or modified (e.g., by an amino acid substitution) to avoid binding of such domains to the transmembrane domains of the same or different surface membrane proteins to minimize interactions with other members of the receptor complex.
- the transmembrane domain may be derived either from a natural or from a synthetic source. Where the source is natural, the domain may be derived from any membrane-bound or transmembrane protein.
- Transmembrane regions may be derived from (i.e., comprise) a receptor tyrosine kinase (e.g., ErbB2), glycophorin A (GpA), 4-1BB/CD137, activating NK cell receptors, an immunoglobulin protein, B7-H3, BAFFR, BFAME (SEAMF8), BTEA, CD100 (SEMA4D), CD103, CD160 (BY55), CD18, CD19, CD19a, CD2, CD247, CD27, CD276 (B7-H3), CD28, CD29, CD3 delta, CD3 epsilon, CD3 gamma, CD30, CD4, CD40, CD49a, CD49D, CD49f, CD69, CD7, CD84, CD8alpha, CD8beta, CD96 (Tactile), CD1 la, CD1 lb, CD1 1c, CD1 Id, CDS, CEACAM1, CRT AM, cytokine receptor
- suitable intracellular signaling domain include, but are not limited to, activating Macrophage/Myeloid cell receptors CSFR1, MYD88, CD14, TIE2, TLR4, CR3, CD64, TREM2, DAP10, DAP12, CD169, DECTIN1, CD206, CD47, CD163, CD36, MARCO, TIM4, MERTK, F4/80, CD91, C1QR, LOX-1, CD68, SRA, BAI-1, ABCA7, CD36, CD31, Lactoferrin, or a fragment, truncation, or combination thereof.
- a receptor tyrosine kinase may be derived from (e.g., comprise) Insulin receptor (InsR), Insulin-like growth factor I receptor (IGF1R), Insulin receptor-related receptor (IRR), platelet derived growth factor receptor alpha (PDGFRa), platelet derived growth factor receptor beta (PDGFRfi).
- Insulin receptor Insulin receptor
- IGF1R Insulin-like growth factor I receptor
- IRR Insulin receptor-related receptor
- PDGFRa platelet derived growth factor receptor alpha
- PDGFRfi platelet derived growth factor receptor beta
- KIT proto-oncogene receptor tyrosine kinase Kit
- colony stimulating factor 1 receptor CSFR
- fms related tyrosine kinase 3 FLT3
- fms related tyrosine kinase 1 VFGFR-1
- kinase insert domain receptor VAGFR-2
- fms related tyrosine kinase 4 VGFR-3
- FGFR1 fibroblast growth factor receptor 1
- FGFR2 fibroblast growth factor receptor 2
- FGFR3 fibroblast growth factor receptor 4
- FGFR4 protein tyrosine kinase 7
- trkA neurotrophic receptor tyrosine kinase 1
- trkB neurotrophic receptor tyrosine kinase 2
- trkC neurotrophic receptor tyrosine kinase like orphan receptor
- the CAR comprises a costimulatory domain.
- the costimulatory domain comprises 4-1BB (CD137), CD28, or both, and/or an intracellular T cell signaling domain.
- the costimulatory domain is human CD28, human 4- IBB, or both, and the intracellular T cell signaling domain is human CD3 zeta (Q. 4-1BB, CD28, CD3 zeta may comprise less than the whole 4-1BB, CD28 or CD3 zeta, respectively.
- Chimeric antigen receptors may incorporate costimulatory (signaling) domains to increase their potency. See U.S. Patent Nos. 7,741,465, and 6,319,494, as well as Krause etal.
- a costimulatory domain comprises the amino acid sequence of SEQ ID NO: 318 or 320 of WO2023081526.
- the intracellular (signaling) domain of the engineered T cells disclosed herein may provide signaling to an activating domain, which then activates at least one of the normal effector functions of the immune cell. Effector function of a T cell, for example, may be cytolytic activity or helper activity including the secretion of cytokines.
- suitable intracellular signaling domain include (e.g., comprise), but are not limited to 4-1BB/CD137, activating NK cell receptors, an Immunoglobulin protein, B7-H3, BAFFR, BEAME (SEAMF8), BTLA, CD100 (SEMA4D), CD103, CD160 (BY55), CD18, CD19, CD 19a, CD2, CD247, CD27, CD276 (B7-H3), CD28, CD29, CD3 delta, CD3 epsilon, CD3 gamma, CD30, CD4, CD40, CD49a, CD49D, CD49f, CD69, CD7, CD84, CD8alpha, CD8beta, CD96 (Tactile), CD1 la, CD1 lb, CD1 1c, CD1 Id, CDS, CEACAM1, CRT AM, cytokine receptor, DAP-10, DNAM1 (CD226), Fc gamma receptor, GADS, GITR
- CD3 is an element of the T cell receptor on native T cells, and has been shown to be an important intracellular activating element in CARs.
- the CD3 is CD3 zeta.
- the activating domain comprises an amino acid sequence of at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to the polypeptide sequence of SEQ ID NO: 319 of WO2023081526.
- the sequence encoding the CAR comprises a sequence from Table 2.
- TCR T-Cell Receptors
- TCRs are described using the International Immunogenetics (IMGT) TCR nomenclature, and links to the IMGT public database of TCR sequences.
- Native alpha-beta heterodimeric TCRs have an alpha chain and a beta chain.
- each chain may comprise variable, joining and constant regions, and the beta chain also usually contains a short diversity region between the variable and joining regions, but this diversity region is often considered as part of the joining region.
- Each variable region may comprise three CDRs (Complementarity Determining Regions) embedded in a framework sequence, one being the hypervariable region named CDR3.
- Va alpha chain variable
- VP beta chain variable
- the Va types are referred to in IMGT nomenclature by a unique TRAV number.
- TRAV21 defines a TCR Va region having unique framework and CDR1 and CDR2 sequences, and a CDR3 sequence which is partly defined by an amino acid sequence which is preserved from TCR to TCR but which also includes an amino acid sequence which varies from TCR to TCR.
- TRBV5-1 defines a TCR VP region having unique framework and CDR1 and CDR2 sequences, but with only a partly defined CDR3 sequence.
- the joining regions of the TCR are similarly defined by the unique IMGT TRAJ and TRBJ nomenclature, and the constant regions by the IMGT TRAC and TRBC nomenclature.
- the beta chain diversity region is referred to in IMGT nomenclature by the abbreviation TRBD, and, as mentioned, the concatenated TRBD/TRBJ regions are often considered together as the joining region.
- TCRs exist in heterodimeric aP or y5 forms. However, recombinant TCRs consisting of aa or PP homodimers have previously been shown to bind to peptide MHC molecules. Therefore, the TCR of the present disclosure may be a heterodimeric aP TCR or may be an aa or PP homodimeric TCR.
- an aP heterodimeric TCR may, for example, be transfected as full length chains having both cytoplasmic and transmembrane domains.
- TCRs of the present disclosure may have an introduced disulfide bond between residues of the respective constant domains, as described, for example, in WO 2006/000830.
- TCRs of the present disclosure may comprise an alpha chain TRAC constant domain sequence and/or a beta chain TRBC1 or TRBC2 constant domain sequence.
- the alpha and beta chain constant domain sequences may be modified by truncation or substitution to delete the native disulfide bond between Cys4 of exon 2 of TRAC and Cys2 of exon 2 of TRBC 1 or TRBC2.
- the alpha and/or beta chain constant domain sequence(s) may also be modified by substitution of cysteine residues for Thr 48 of TRAC and Ser 57 of TRBC1 or TRBC2, the said cysteines forming a disulfide bond between the alpha and beta constant domains of the TCR.
- Binding affinity (inversely proportional to the equilibrium constant KD) and binding half-life (expressed as P/2) can be determined by any appropriate method. It will be appreciated that doubling the affinity of a TCR results in halving the KD- P/2 is calculated as In 2 divided by the off-rate (koff). So doubling of P/2 results in a halving in koff. KD and koff values for TCRs are usually measured for soluble forms of the TCR, i.e., those forms which are truncated to remove cytoplasmic and transmembrane domain residues.
- a given TCR has an improved binding affinity for, and/or a binding half-life for the parental TCR if a soluble form of that TCR has the said characteristics.
- the binding affinity or binding half-life of a given TCR is measured several times, for example 3 or more times, using the same assay protocol, and an average of the results is taken.
- the present disclosure includes a non-naturally occurring and/or purified and/or or engineered cell, especially a T-cell, presenting a TCR of the present disclosure.
- nucleic acid such as DNA, cDNA or RNA
- T cells expressing the TCRs of the present disclosure will be suitable for use in adoptive therapy-based treatment of cancers such as those of the pancreas and liver.
- suitable methods by which adoptive therapy can be carried out see for example Rosenberg et al. , (2008) Nat Rev Cancer 8(4): 299-308).
- TCRs of the present disclosure may be subject to post-translational modifications when expressed by transfected cells.
- Glycosylation is one such modification, which may comprise the covalent attachment of oligosaccharide moieties to defined amino acids in the TCR chain.
- asparagine residues, or serine/threonine residues are well-known locations for oligosaccharide attachment.
- the glycosylation status of a particular protein depends on a number of factors, including protein sequence, protein conformation and the availability of certain enzymes. Furthermore, glycosylation status (i.e., oligosaccharide type, covalent linkage and total number of attachments) can influence protein function.
- Glycosylation of transfected TCRs may be controlled by mutations of the transfected gene (Kuball J et al. (2009), J Exp Med 206(2):463-475). Such mutations are also encompassed in this disclosure.
- a TCR may be specific for an antigen in the group MAGE-A1, MAGE-A2, MAGE- A3, MAGE-A4, MAGE-A5, MAGE-A6, MAGE-A7, MAGE-A8, MAGE-A9, MAGE-A10, MAGE-A11, MAGE-A12, MAGE-A13, GAGE-1, GAGE-2, GAGE-3, GAGE-4, GAGE-5, GAGE-6, GAGE-7, GAGE-8, BAGE-1, RAGE-1, LB33/MUM-1, PRAME, NAG, MAGE-Xp2 (MAGE-B2), MAGE-Xp3 (MAGE-B3), MAGE-Xp4 (AGE-B4), tyrosinase, brain glycogen phosphorylase, Melan-A, MAGE-CI, MAGE-C2, NY-ESO-1, LAGE-1, SSX-1, SSX-2(HOM-MEL-40), SSX-1, SSX-4, SSX-5, S
- BCR B-Cell Receptors
- B-cell receptors or B-cell antigen receptors are immunoglobulin molecules that form a type I transmembrane protein on the surface of a B cell.
- a BCR is capable of transmitting activatory signal into a B cell following recognition of a specific antigen. Prior to binding of a B cell to an antigen, the BCR will remain in an unstimulated or “resting” stage. Binding of an antigen to a BCR leads to signaling that initiates a humoral immune response.
- a BCR is expressed by mature B cells. These B cells work with immunoglobulins (Igs) in recognizing and tagging pathogens.
- the typical BCR comprises a membrane-bound immunoglobulin (e.g., mlgA, mlgD, mlgE, mlgG, and mlgM), along with associated and Iga/IgP (CD79a/CD79b) heterodimers (a/p).
- membrane-bound immunoglobulins are tetramers consisting of two identical heavy and two light chains.
- the membrane bound immunoglobulin is capable of responding to antigen binding by signal transmission across the plasma membrane leading to B cell activation and consequently clonal expansion and specific antibody production (Friess M et al. (2016), Front. Immunol. 2947(9)).
- the Iga/IgP heterodimer is responsible for transducing signals to the cell interior.
- IT AMs immunoreceptor tyrosine-based activation motifs located on each of the cytosolic tails of the heterodimers.
- IT AMs comprise two tyrosine residues separated by 9-12 amino acids (e.g., tyrosine, leucine, and/or valine).
- tyrosine of the BCR’s IT AMs become phosphorylated by Src-family tyrosine kinases Blk, Fyn, or Lyn (Janeway C etal., Immunobiology: The Immune System in Health and Disease (Garland Science, 5th ed. 2001)).
- the circular RNA polynucleotide may encode for a various number of other chimeric proteins available in the art.
- the chimeric proteins may include recombinant fusion proteins, chimeric mutant protein, or other fusion proteins.
- the circular RNA polynucleotide encodes for an immune modulatory ligand.
- the immune modulatory ligand may be immunostimulatory; while in other embodiments, the immune modulatory ligand may be immunosuppressive.
- the circular RNA polynucleotide encodes for a cytokine.
- the cytokine comprises a chemokine, interferon, interleukin, lymphokine, and tumor necrosis factor.
- Chemokines are chemotactic cytokine produced by a variety of cell types in acute and chronic inflammation that mobilizes and activates white blood cells.
- An interferon comprises a family of secreted a-helical cytokines induced in response to specific extracellular molecules through stimulation of TLRs (Borden, Molecular Basis of Cancer (Fourth Edition) 2015).
- Interleukins are cytokines expressed by leukocytes.
- the circular RNA polynucleotide may encode for a transcription factor.
- Regulatory T cells are important in maintaining homeostasis, controlling the magnitude and duration of the inflammatory response, and in preventing autoimmune and allergic responses.
- Tregs are thought to be mainly involved in suppressing immune responses, functioning in part as a “self-check” for the immune system to prevent excessive reactions.
- Tregs are involved in maintaining tolerance to self-antigens, harmless agents such as pollen or food, and abrogating autoimmune disease.
- Tregs are found throughout the body including, without limitation, the gut, skin, lung, and liver. Additionally, Treg cells may also be found in certain compartments of the body that are not directly exposed to the external environment such as the spleen, lymph nodes, and even adipose tissue. Each of these Treg cell populations is known or suspected to have one or more unique features and additional information may be found in Lehtimaki and Lahesmaa, Regulatory T cells control immune responses through their non-redundant tissue specific features, 2013, FRONTIERS IN IMMUNOL., 4(294): 1- 10, the disclosure of which is hereby incorporated in its entirety.
- Tregs are known to require TGF-P and IL-2 for proper activation and development.
- Tregs expressing abundant amounts of the IL-2 receptor (IL-2R), are reliant on IL-2 produced by activated T cells.
- Tregs are known to produce both IL-10 and TGF-P, both potent immune suppressive cytokines.
- Tregs are known to inhibit the ability of antigen presenting cells (APCs) to stimulate T cells.
- APCs antigen presenting cells
- CTLA-4 may bind to B7 molecules on APCs and either block these molecules or remove them by causing internalization resulting in reduced availability of B7 and an inability to provide adequate co-stimulation for immune responses. Additional discussion regarding the origin, differentiation and function of Tregs may be found in Dhamne et al. , Peripheral and thymic Foxp3+ regulatory T cells in search of origin, distinction, and function, 2013, Frontiers in Immunol., 4 (253): 1-11, the disclosure of which is hereby incorporated in its entirety.
- the coding element of the circular RNA polynucleotide encodes for one or more checkpoint inhibitors or agonists.
- the immune checkpoint inhibitor is an inhibitor of Programmed Death- Ligand 1 (PD-L1, also known as B7-H1, CD274), Programmed Death 1 (PD-1), CTLA-4, PD-L2 (B7- DC, CD273), LAG3, TIM3, 2B4, A2aR, B7H1, B7H3, B7H4, BTLA, CD2, CD27, CD28, CD30, CD40, CD70, CD80, CD86, CD137, CD160, CD226, CD276, DR3, GAL9, GITR, HAVCR2, HVEM, IDO1, IDO2, ICOS (inducible T cell costimulator), KIR, LAIR1, LIGHT, MARCO (macrophage receptor with collageneous structure), PS (phosphatidylserine), OX-40, SLAM, TIGHT, VISTA, VTCN1, or any combinations thereof.
- PD-L1 Programmed Death- Ligand 1
- PD-1 Programmed Death 1
- CTLA-4 PD
- the immune checkpoint inhibitor is an inhibitor of IDO1, CTLA4, PD-1, LAG3, PD-L1, TIM3, or combinations thereof. In some embodiments, the immune checkpoint inhibitor is an inhibitor of PD-L1. In some embodiments, the immune checkpoint inhibitor is an inhibitor of PD-1. In some embodiments, the immune checkpoint inhibitor is an inhibitor of CTLA-4. In some embodiments, the immune checkpoint inhibitor is an inhibitor of LAG3. In some embodiments, the immune checkpoint inhibitor is an inhibitor of TIM3. In some embodiments, the immune checkpoint inhibitor is an inhibitor of IDOL
- Such antagonists of CTLA-4, PD-1, PDL-1, LAG-3, and TIM-3 can include antibodies which specifically bind to CTLA-4, PD-1, PDL-1, LAG-3, and TIM-3, respectively and inhibit and/or block biological activity and function.
- the pay load encoded within one or more of the coding elements is a hormone, FC fusion protein, anticoagulant, blood clotting factor, protein associated with deficiencies and genetic disease, a chaperone protein, an antimicrobial protein, an enzyme (e.g., metabolic enzyme), a structural protein (e.g., a channel or nuclear pore protein), protein variant, small molecule, antibody, nanobody, an engineered non-body antibody, or a combination thereof.
- an ionizable lipid comprises one or more cleavable functional groups (e.g., a disulfide) that allow, for example, a hydrophilic functional head-group to dissociate from a lipophilic functional tail-group of the compound (e.g., upon exposure to oxidative, reducing or acidic conditions).
- cleavable functional groups e.g., a disulfide
- the ionizable lipid has a pKa from 6 to 12. In some embodiments, the ionizable lipid has a pKa from 7 to 9. In some embodiments, the ionizable lipid has a pKa of 6.0, 6.1, 6.2, 6.3, 6.4, 6.5, 6.6, 6.7, 6.8, 6.9, 7.0, 7.1, 7.2, 7.3, 7.4, 7.5, 7.6, 7.7, 7.8, 7.9, 8.0, 8.1, 8.2, 8.3, 8.4, 8.5, 8.6, 8.7, 8.8, 8.9, or 9.0 or any ranges created by these.
- the ionizable lipid comprises an amino group.
- the ionizable lipid comprises a divalent headgroup and one or more straight hydrocarbon lipid tails.
- the straight hydrocarbon lipid tails are from 3- 25 carbon atoms in length, such as 5 to 25, 5 to 20, 5 to 15, 5 to 10, 10 to 15, 10 to 20, or 10 to 25 carbon atoms in length.
- the ionizable lipid comprises a divalent headgroup and one or more branched hydrocarbon lipid tails.
- the branched hydrocarbon lipid tails are from 3-25 carbon atoms in length, such as 5 to 25, 5 to 20, 5 to 15, 5 to 10, 10 to 15, 10 to 20, or 10 to 25 carbon atoms in length.
- the divalent headgroup is selected from guanidine and squaramide.
- the squaramide headgroup is of the following formula: wherein RA and RB are each independently a C1-C6 alkyl group or H; and represents the point of attachment of the headgroup to a straight or branched hydrocarbon lipid tail.
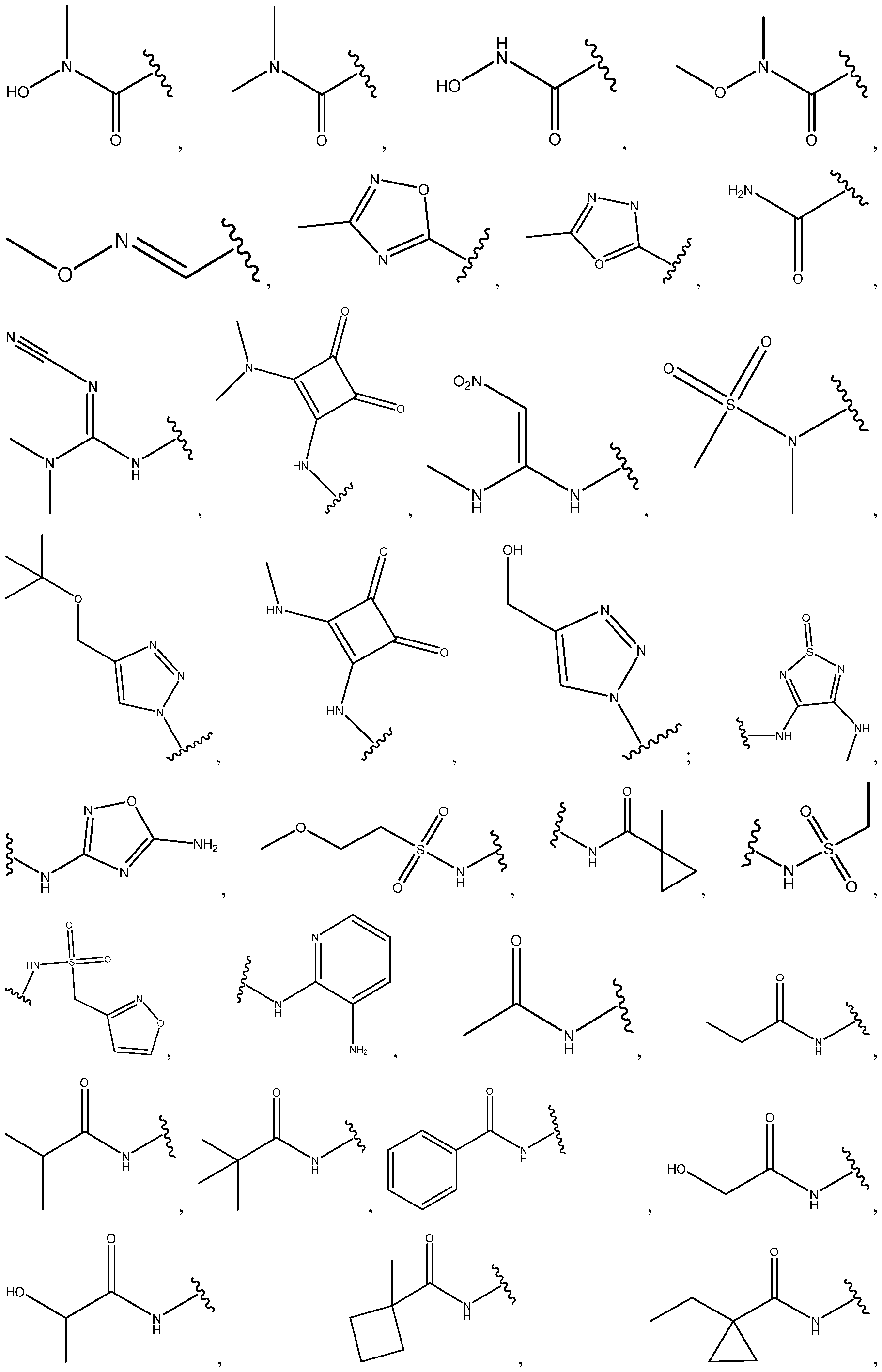
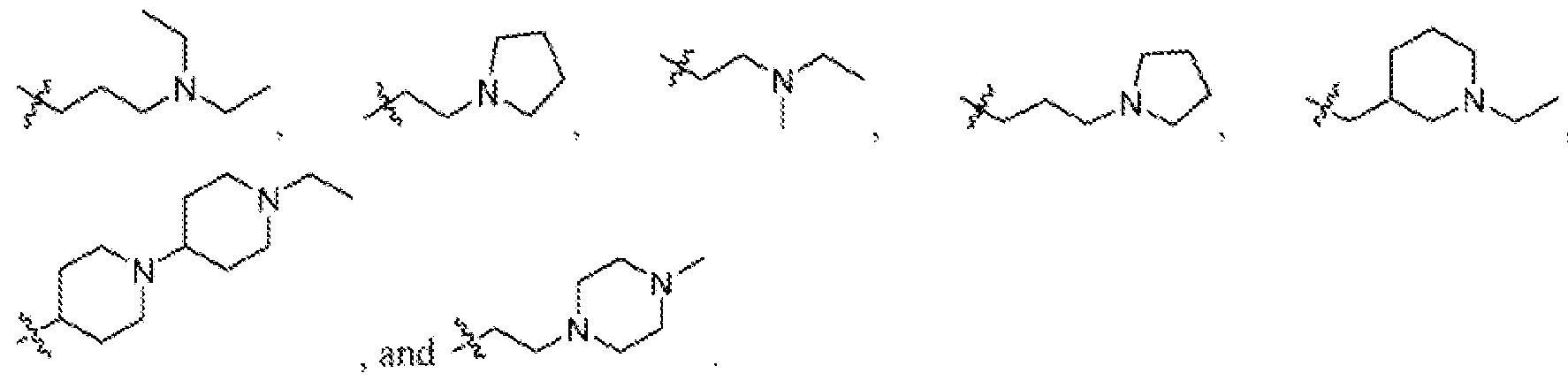
- the ionizable lipid comprises a head group selected from:
- the ionizable lipid comprises a head group selected from: wherein represents the point of attachment of the headgroup to a hydrocarbon lipid tail (e.g., straight or branched).
- the ionizable lipid comprises a hydrophilic headgroup as disclosed in Jayaraman et al. Angew. Chem. Int. Ed. (2012), 51, 8529-8533.
- the ionizable lipid is ionizable lipid 2, wherein the ionizable lipid 2 [0188] In some embodiments, the ionizable lipid is ionizable lipid 3, wherein the ionizable lipid 3 comprises:
- the ionizable lipid is endosomal escape agent 1, wherein the endosomal escape agent comprises:
- the one or more of the cationic or ionizable lipids are represented by Formula (LI):
- n is an integer between 1 and 4;
- R a is hydrogen or hydroxyl
- Ri and R2 are each independently a linear or branched Co-C id alkyl, Co-C id alkenyl, or Co-C id heteroalkyl, optionally substituted by one or more substituents selected from a group consisting of oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonyla
- Ra is hydrogen. In some embodiments, Ra is hydroxyl.
- the ionizable lipid is represented by Formula (Lla -1), Formula (LIa-2), or Formula (LIa-3):
- the ionizable lipid is represented by Formula (LIb-1), Formula (LIb-2), or Formula (LIb-3):
- the ionizable lipid is represented by Formula (LIb-4), Formula (LIb-5), Formula (LIb-6), Formula (LIb-7), Formula (LIb-8), or Formula (LIb-9): Formula (LIb-4) Formula (LIb-5) Formula (LIb-6)
- the one or more of the cationic or ionizable lipids are represented by Formula (LI), wherein Ri and R2 are each independently selected from: [0196] In some embodiments, Ri and R2 are the same. In some embodiments, Ri and R2 are different.
- the one or more of the cationic or ionizable lipids are represented by Formula (LI*):
- n* is an integer between 1 to 7
- R a is hydrogen or hydroxyl
- R b is hydrogen or Ci-Ce alkyl
- Ri and R2 are each independently a linear or branched C1-C30 alkyl, C2-C30 alkenyl, or C1-C30 heteroalkyl, optionally substituted by one or more substituents selected from oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino, hydroxycarbonyl, al
- alkylaminoalkyl (alkyl)aminocarbonyl, alkylaminoalkylcarbonyl, dialkylaminoalkylcarbonyl, heterocyclylcarbonyl, alkenylcarbonyl, alkynylcarbonyl, alkylsulfoxide, alkylsulfoxidealkyl, alkylsulfonyl, and alkylsulfonealkyl.
- the one or more of the cationic or ionizable lipids are represented by Formula (LII):
- n is independently an integer from 2-15;
- Li and L3 are each independently -0C(0)-* or -C(O)O-*, wherein indicates the attachment point to Ri or R3;
- Ri and R3 are each independently a linear or branched C9-C20 alkyl or C9-C20 alkenyl, optionally substituted by one or more substituents selected from a group consisting of oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino, hydroxycarbonyl, alkyloxycarbon
- R2 is selected from a group consisting of:
- the ionizable lipid is selected from an ionizable lipid of Formula LII, wherein Ri and R3 are each independently selected from a group consisting of:
- Ri and R3 are the same. In some embodiments, Ri and R3 are different. [0201] In some embodiments, the one or more of the cationic or ionizable lipids are represented by
- the ionizable lipid is selected from an ionizable lipid of WO 2015/095340. In some embodiments, the ionizable lipid is selected from an ionizable lipid of WO 2021/021634, WO 2020/237227, or WO 2019/236673. In some embodiments, the ionizable lipid is selected from an ionizable lipid of WO 2021/226597 and WO 2021/113777. In some embodiments, the ionizable lipid is selected from an ionizable lipid of WO 2023/056033. In some embodiments, the ionizable lipid is selected from an ionizable lipid of WO 2023/081526.
- the one or more of the cationic or ionizable lipids are represented by Formula (LIII):
- L 1 is C2-C11 alkylene, Cr-Cio-alkenylene, or Cr-Cio-alkynylene;
- X 1 is OR 1 , SR 1 , or N(R’)2, where R 1 is independently H or unsubstituted Ci-Ce alkyl;
- R 2 and R 3 are each independently Ce-Cso-alkyl, Ce-Cso-alkenyl, or Ce-Cso-alkynyl.
- the one or more of the cationic or ionizable lipids are represented by Formula (LIII*):
- L 1 is C2-C11 alkylene, Cr-Cio-alkenylene, or Cr-Cio-alkynylene;
- X 1 is OR 1 , SR 1 , or N(R’)2, where R 1 is independently H or unsubstituted Ci-Ce alkyl;
- R2 and R3 are each independently a linear or branched C1-C30 alkyl, C2-C30 alkenyl, or C1-C30 heteroalkyl, optionally substituted by one or more substituents selected from oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino, hydroxycarbonyl,
- alkylaminoalkyl (alkyl)aminocarbonyl, alkylaminoalkylcarbonyl, dialkylaminoalkylcarbonyl, heterocyclylcarbonyl, alkenylcarbonyl, alkynylcarbonyl, alkylsulfoxide, alkylsulfoxidealkyl, alkylsulfonyl, and alkylsulfonealkyl.
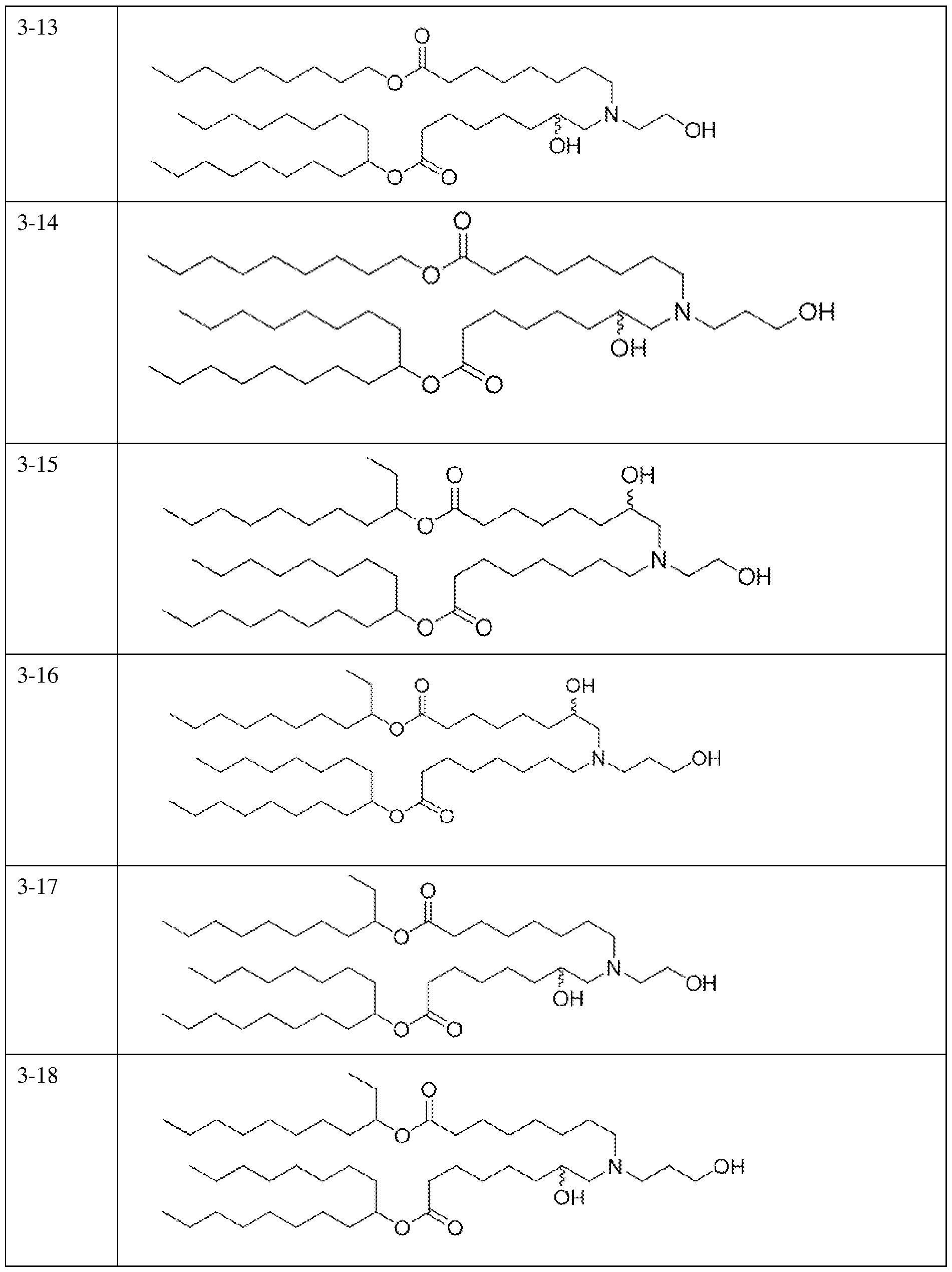
- an ionizable lipid is selected from Table 4.
- an LNP of the present disclosure comprises an ionizable lipid disclosed in one of US 2023/0053437; US 2019/0240354; US 2010/0130588; US 2021/0087135; WO 2021/204179; US 2021/0128488; US 2020/0121809; US 2017/0119904; US 2013/0108685; US 2013/0195920; US 2015/0005363; US 2014/0308304; US 2013/0053572; WO 2019/232095; WO 2021/077067; WO 2019/152557; US 2017/0210697; or WO 2019/089828, each of which is incorporated by reference herein in their entirety.
- an LNP described herein comprises a lipid, e.g., an ionizable lipid, disclosed in US Application publication US 2017/0119904, which is incorporated by reference herein, in its entirety.
- an LNP described herein comprises a lipid, e.g., an ionizable lipid, disclosed in PCT Application publication WO 2021/204179, which is incorporated by reference herein, in its entirety.
- an LNP described herein comprises a lipid, e.g., an ionizable lipid, disclosed in PCT Application WO 2022/251665, which is incorporated by reference herein, in its entirety.
- an LNP described herein comprises an ionizable lipid of Table 5: Table 5: Exemplary Ionizable Lipid Structures
- the ionizable lipid is MC3.
- an ionizable lipid is a compound of Formula (LIV):
- R a is hydrogen or hydroxyl
- R h is hydrogen or Ci-Ce alkyl
- R 1 is C1-C30 alkyl or R’* ;
- R 2 is C1-C30 alkyl or R 2 *;
- R 1 * and R 2 * are independently selected from: -(CH 2 ) q C(O)O(CH 2 ) r C(R 8 )(R 9 )(R 10 ),
- R 8 is H or R"
- R 9 , R 10 , and R” are each independently Ci-C 2 o alkyl or C 2 -C 2 o-alkenyl; and wherein (i) R 1 is R 1 *, (ii) R 2 is R 2 *, or (iii) R 1 is R 1 * and R 2 is R 2 *.
- an ionizable lipid is selected from Table 6.
- an ionizable lipid of the present disclosure is represented by Formula (LV):
- Formula (LV) or is a pharmaceutically acceptable salt thereof, wherein:
- R a is hydrogen or hydroxyl
- R 1 is C1-C30 alkyl or R’* ;
- R 2 is C1-C30 alkyl or R 2 *; R 1 * and R 2 * are independently selected from:
- R 4 is hydrogen or R 7 ;
- R 5 , R 6 , and R 7 are each independently Ci-C 2 o alkyl or C 2 -C 2 o-alkenyl; wherein (i) R 1 is R 1 *, (ii) R 2 is R 2 *, or (iii) R 1 is R 1 * and R 2 is R 2 *; and
- R 3 is L-R’, wherein L is linear or branched Ci-Cio alkylene, and R’ is (i) mono- or bicyclic heterocyclyl or heteroaryl, such as imidazolyl, pyrazolyl, 1 ,2,4-triazolyl, or benzimidazolyl, each optionally substituted at one or more available carbon and nitrogen by Ci-Ce alkyl, or (ii) R A , R B , or R c , wherein R B is selected from:
- the ionizable lipid is selected from an ionizable lipid described or disclosed in any one of PCT Publications WO 2023/044343, WO 2023/044333, WO 2023/122752, WO 2024/044728 and WO 2023/196931 and PCT Application PCT/US2024/019990, or any combination thereof, each of which is incorporated by reference herein in its entirety.
- an ionizable lipid is selected from Table 7.
- amphiphilic polymers In certain embodiments disclosed herein are amphiphilic polymers.
- the subject amphiphilic polymers may be used as a component of a composition to facilitate encapsulation and release of circular RNA to one or more target cells (e.g., by forming a shell around the circular RNA core).
- Any suitable amphiphilic polymer can be used in the disclosed nanoparticles.
- Polymers can be natural or unnatural (synthetic) polymers.
- Polymers can be copolymers comprising two or more monomers. In terms of sequence, copolymers can be random, block, or comprise a combination of random and block sequences.
- polymer as used herein, is given its ordinary meaning as used in the art, i.e., a molecular structure comprising one or more repeat units (monomers), connected by covalent bonds.
- the repeat units may all be identical, or in some embodiments, there may be more than one type of repeat unit present within the polymer. If more than one type of repeat unit is present within the polymer, then the polymer is said to be a “copolymer.” It is to be understood that in any embodiment employing a polymer, the polymer being employed may be a copolymer in some embodiments. The repeat units forming the copolymer may be arranged in any fashion.
- the repeat units may be arranged in a random order, in an alternating order, or as a block copolymer, i.e., comprising one or more regions each comprising a first repeat unit (e.g., a first block), and one or more regions each comprising a second repeat unit (e.g., a second block), etc.
- Block copolymers may have two (a di-block copolymer), three (a tri-block copolymer), or more numbers of distinct blocks.
- Disclosed nanoparticles can include copolymers, which, in some embodiments, describes two or more polymers (such as those described herein) that have been associated with each other, usually by covalent bonding of the two or more polymers together.
- a copolymer may comprise a first polymer and a second polymer, which have been conjugated together to form a block copolymer where the first polymer can be a first block of the block copolymer and the second polymer can be a second block of the block copolymer.
- a block copolymer may, in some embodiments, contain multiple blocks of polymer, and a “block copolymer,” as used herein, is not limited to only block copolymers having only a single first block and a single second block.
- a block copolymer may comprise a first block comprising a first polymer, a second block comprising a second polymer, and a third block comprising a third polymer or the first polymer, etc.
- block copolymers can contain any number of first blocks of a first polymer and second blocks of a second polymer (and in certain embodiments, third blocks, fourth blocks, etc.).
- block copolymers can also be formed, in some instances, from other block copolymers.
- a first block copolymer may be conjugated to another polymer (which may be a homopolymer, a biopolymer, another block copolymer, etc.), to form a new block copolymer containing multiple types of blocks, and/or to other moieties (e.g., to non-polymeric moieties).
- the polymer e.g., copolymer, e.g., block copolymer
- amphiphilic i.e., having a hydrophilic portion and a hydrophobic portion, or a relatively hydrophilic portion and a relatively hydrophobic portion.
- a hydrophilic polymer can be one generally that attracts water, and a hydrophobic polymer can be one that generally repels water.
- the amphiphilic polymer is a block copolymer comprising a hydrophilic block comprising a hydrophilic polymer; and a hydrophobic block comprising a hydrophobic polymer.
- the amphiphilic polymer is a biocompatible polymer.
- the amphiphilic polymer is a biocompatible polymer that can be biodegradable, i.e., the polymer is able to degrade, chemically and/or biologically, within a physiological environment, such as within the body.
- biodegradable polymers are those that, when introduced into cells, are broken down by the cellular machinery (biologically degradable) and/or by a chemical process, such as hydrolysis, (chemically degradable) into components that the cells can either reuse or dispose of without significant toxic effect on the cells.
- the biodegradable polymer and their degradation byproducts can be biocompatible.
- the subject amphiphilic polymer may be one that hydrolyzes spontaneously upon exposure to water (e.g., within a subject) or degrades upon exposure to heat (e.g., at temperatures of about 37° C). Degradation of a subject amphiphilic polymer may occur at varying rates, depending on the polymer or copolymer used. For example, the half-life of the polymer (the time at which 50% of the polymer can be degraded into monomers and/or other nonpolymeric moieties) may be on the order of days, weeks, months, or years, depending on the polymer.
- the polymers may be biologically degraded, e.g., by enzymatic activity or cellular machinery, in some embodiments, for example, through exposure to a lysozyme (e.g., having relatively low pH).
- the polymers may be broken down into monomers and/or other nonpolymeric moieties that cells can either reuse or dispose of without significant toxic effect on the cells (for example, a polylactide polymer may be hydrolyzed to form lactic acid, a polyglycolide polymer may be hydrolyzed to form glycolic acid, etc.).
- the amphiphilic polymer further comprises a cleavable linker (L).
- the cleavable linker can render the amphiphilic polymer biodegradable (i.e., as described herein above). Any convenient cleavable linker can find use in the subject amphiphilic polymers.
- the amphiphilic polymer comprises a cleavable linker that is cleaved by exposure to a stimulus.
- a non-exhaustive list of stimulus includes pH, temperature, light, redox change, overexpressed enzymes, hypoxia, sound, magnetic force, electrical energy, and any combination thereof.
- the cleavable linker L comprises a group selected from disulfide, hydrazone, vinyl ether, imine, ortho ester, borate ester, amide, a peptide, an azo, and any combination thereof.
- the cleavable linker L comprises a disulfide.
- the linker comprises a disulfide and can be cleaved by exposure to a redox change.
- the cleavage of the disulfide linker is mediated by glutathione (GSH).
- GSH glutathione
- Disulfide bonds can be easily broken down by reducing glutathione (GSH) into sulfhydryl groups, which causes the degradation of carriers and facilitates the release of cargoes (e.g.,, circRNA cargo).
- Disulfide bonds are often used in delivery systems as linkers, which can degrade rapidly to release cargoes in the reducing environment of GSH in tumor cells.
- Glutathione is the most abundant thiol species in the cytoplasm, functioning as a natural oxidant scavenger and the major reducing agent in biochemical processes.
- the intracellular GSH concentration (2-10 mM) is substantially higher than extracellular levels (2 pM in plasma), which provides opportunities for intracellular delivery of therapeutic agents by cleavable disulfide linked carriers.
- the cleavable linker L is a pH sensitive linker.
- pH is a commonly used internal stimulus in pathological sites such as tumors and inflammatory tissues, as well as in a physiological environment such as acidic organelles (e.g., endosomes). Endosomes (pH 5-6) and lysosomes (pH 4-5) in mammalian cells appear slightly acidic, while the cytoplasm and endoplasmic reticulum have a neutral pH (e.g., approximately 7.2), the Golgi complex has a pH in the range of 6.0- 6.7, and the mitochondrion a pH of approximately 8.0.
- polymeric lipid nanoparticles After endocytosis, polymeric lipid nanoparticles are entrapped in the early endosome (pH approximately 5.5), which mature into the late endosome. The late endosome fuses with the lysosome (pH less than 5) and these nano-drug delivery systems in the lysosome are subjected to degradation. Polymeric lipid nanoparticles carrying circRNA must avoid endosomal degradation and successfully release the circRNA into the cytoplasm for it to perform its desired therapeutic effect.
- pH-sensitive polymeric lipid nanoparticles can be designed such that upon being exposed to acidic microenvironment in the endosome, the polymeric shell of the polymeric lipid nanoparticles rapidly disintegrates to release the circRNA and ionizable lipid (lipid). The lipid can then drive the escape of the circRNA from the endosome into the cytoplasm.
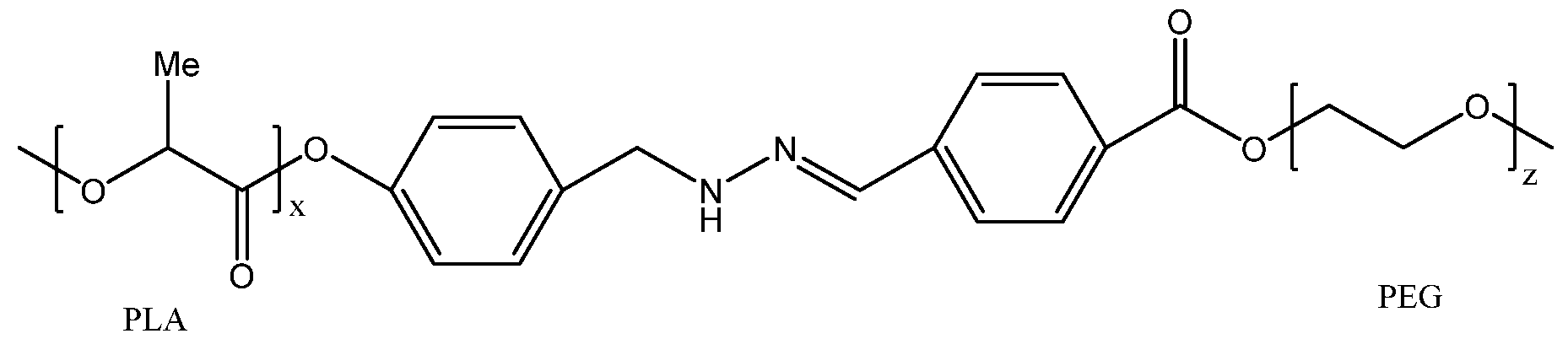
- the cleavable linker L comprises a hydrazone.
- the linker comprises a hydrazone that can be cleaved by exposure to an acidic pH.
- the linker comprises a hydrazone and can be cleaved by exposure to a pH of 6.5 or less, such as a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 or less, a pH of 4 or less, or even less.
- the cleavable linker comprises a vinyl ether (see e.g., Shin, et al. Molecular Pharmaceutics 2012, 9(11), 3266-3276).
- the linker comprises a vinyl ether that can be cleaved by exposure to an acidic pH.
- the linker comprises a vinyl ether and can be cleaved by exposure to a pH of 6.5 or less, a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 of less, a pH of 4 or less, or even less.
- the cleavable linker comprises one or more of an imine, an ortho ester, a borate ester, or an amide.
- the linker comprises one or more of an imine, an ortho ester, a borate ester, or an amide that can be cleaved by exposure to an acidic pH (see e.g., Ding et al. Journal of Controlled Release 2022, 348, 206-238).
- the linker comprises one or more of an imine, an ortho ester, a borate ester, or an amide and can be cleaved by exposure to a pH of 6.5 or less, a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 of less, a pH of 4 or less, or even less.
- the cleavable linker comprises an octapeptide.
- the octapeptide is of the sequence GPLGIAGQ.
- the octapeptide is of the sequence GPLGVRGC.
- the linker comprising an octapeptide is cleaved by exposure to over-expressed enzymes.
- the over-expressed enzyme is matrix metalloproteinase 2 (MMP2) (see e.g., Zhu et al. PNAS 2013, 110(42), 17047-17052).
- MMP2 matrix metalloproteinase 2
- the cleavable linker comprises an azo group.
- the linker comprises an azo group that can be cleaved by exposure to hypoxia (see e.g., Joshi et al. International Journal of Pharmaceutics 2020, 590, 119915).
- the hydrophobic block of the amphiphilic polymer comprises the cleavable linker.
- the cleavable linker covalently connects the hydrophilic block to the hydrophobic block of the amphiphilic polymer.
- the cleavable linker covalently connects two or more amphiphilic polymers, such as three or more, four or more, five or more, or even more amphiphilic polymers. In some embodiments, the cleavable linker covalently connects two amphiphilic polymers. In some embodiments, the cleavable linker covalently connects three amphiphilic polymers. In some embodiments, the cleavable linker covalently connects four amphiphilic polymers. In some embodiments, the cleavable linker covalently connects five amphiphilic polymers. In some embodiments, the cleavable linker covalently connects five or more amphiphilic polymers.
- the amphiphilic polymer is covalently bound to a cationic moiety (e.g., a small molecule cationic moiety such as an ionizable lipid, non-lipid small molecule etc.). In some embodiments, the amphiphilic polymer is covalently bound to the ionizable lipid. In some embodiments, the amphiphilic polymer is not covalently bound to the ionizable lipid.
- a cationic moiety e.g., a small molecule cationic moiety such as an ionizable lipid, non-lipid small molecule etc.
- the molecular weight (or e.g., the ratio of molecular weights of, e.g., different blocks of a copolymer) of the amphiphilic polymers can be optimized for effective treatment of a specific disease or disorder.
- the molecular weight of a polymer may influence particle degradation rate (such as when the molecular weight of a biodegradable polymer can be adjusted), solubility, water uptake, and drug release kinetics.
- the molecular weight of the polymer (or the ratio of molecular weights of, e.g., different blocks of a copolymer) can be adjusted such that the particle biodegrades in the subject being treated within a reasonable period of time (ranging from a few hours to 1-2 weeks, 3-4 weeks, 5-6 weeks, 7-8 weeks, etc.).
- the amphiphilic polymer has a molecular weight from 10k to 100k, such as 10k to 90k, 10k to 80k, 10k to 70k, or 10k to 50k. In some embodiments, the amphiphilic polymer has a molecular weight from 10k to 70k, such as 10k to 65k, 10k to 60k, 10k to 55k, 10k to 50k, 20k to 70k, 25k to 70k, 30k to 70k, or 35k to 70k.
- the amphiphilic polymer has a molecular weight from 10k to 50k, such as 10k to 45k, 10k to 40k, 10k to 35k, 10k to 30k, 20k to 50k, 25k to 50k, 30k to 50k, or 35k to 50k. In some embodiments, the amphiphilic polymer has a molecular weight of 10k, 15k, 20k, 25k, 30k, 35k, 40k, 45k, 50k, 55k, 60k, 65k, or 70k +/- 10%. [0240] In some embodiments, the amphiphilic polymer is a di-block copolymer.
- amphiphilic polymer comprises a block copolymer of Formula I: X-Y (I) wherein:
- X is a hydrophobic block comprising a hydrophobic polymer
- Y is a hydrophilic block comprising a hydrophilic polymer, wherein the amphiphilic polymer optionally further comprises one or more cleavable linkers.
- the amphiphilic polymer does not include any cleavable linkers.
- the amphiphilic polymer includes one or more cleavable linkers (as described herein). In some embodiments of Formula I, the amphiphilic polymer includes one or more cleavable linkers within the hydrophobic block (X). In some embodiments of Formula I, the amphiphilic polymer includes one or more cleavable linkers within the hydrophilic block (Y). In some embodiments of Formula I, the amphiphilic polymer includes one or more cleavable linkers between the hydrophobic block (X) and the hydrophilic block (Y).
- amphiphilic polymer includes a cleavable linker (as described herein), and the amphiphilic polymer comprises a block copolymer of Formula IA:
- X is a hydrophobic block comprising a hydrophobic polymer
- Y is a hydrophilic block comprising a hydrophilic polymer
- L is a cleavable linker
- the amphiphilic polymer comprises a block copolymer of Formula IB: X A -(L-X B ) n -Y (IB) wherein: each of X A and X B is independently a hydrophobic block comprising a hydrophobic polymer;
- Y is a hydrophilic block comprising a hydrophilic polymer
- L is a cleavable linker
- n is an integer from 1 to 5.
- n is 5. In some embodiments, n is 4. In some embodiments of Formula IB, n is an integer from 1 to 3. [0247] In some embodiments of Formula IB, n is 1 such that the amphiphilic polymer comprises a block copolymer of Formula IB-1 :
- n is 2 such that the amphiphilic polymer comprises a block copolymer of Formula IB -2:
- n 3 such that the amphiphilic polymer comprises a block copolymer of Formula IB -3:
- the amphiphilic polymer comprises a block copolymer of Formula IC:
- each of X A and X B is independently a hydrophobic block comprising a hydrophobic polymer; each of Y A and Y B is independently a hydrophilic block comprising a hydrophilic polymer;
- L is a cleavable linker; and each T is independently a trivalent connecting group (e.g., lysine or cysteine).
- the trivalent connecting group T is an amino acid. In some embodiments, the connecting group T is lysine. In some embodiments, the connecting group T is cysteine.
- amphiphilic polymer comprises a block copolymer of Formula ID: R-X-Y (ID) wherein:
- R is a cationic moiety (e.g., an ionizable lipid);
- X is a hydrophobic block comprising a hydrophobic polymer
- Y is a hydrophilic block comprising a hydrophilic polymer; and the amphiphilic polymer optionally further comprises one or more cleavable linkers.
- the amphiphilic polymer does not comprise a cleavable linker.
- the amphiphilic polymer further comprises a cleavable linker (as described herein). In some embodiments of Formula ID, the amphiphilic polymer includes one or more cleavable linkers within the hydrophobic block (X). In some embodiments of Formula ID, the amphiphilic polymer includes one or more cleavable linkers within the hydrophilic block (Y).
- the amphiphilic polymer further comprises a cleavable linker, and is of Formula ID-1 or ID-2:
- the cationic moiety R comprises a lipid, a polymer, or a non-lipid small molecule.
- the cationic moiety R comprises an ionizable lipid (as described herein).
- the ionizable lipid has a pKa from 6-12, such as 7- 9.
- the ionizable lipid comprises an ionizable amino group.
- the ionizable lipid comprises a divalent headgroup and one or more lipid tails (e.g., straight or branched hydrocarbon lipid tails).
- the divalent headgroup is selected from guanidine and squaramide.
- the ionizable lipid comprises a headgroup as described herein.
- the ionizable lipid includes one or more (e.g., 2) straight or branched hydrocarbon lipid tails from 3-25 carbon atoms in length.
- the cationic moiety R is ethyl lauroyl arginate (EL A).
- the cationic moiety R comprises a polymer (i.e., a cationic polymer).
- cationic polymers are able to condense and/or protect negatively charged strands of nucleic acids (e.g., DNA, RNA, or derivatives thereof).
- the cationic moiety R comprises a polymer selected from poly(lysine), polyethylene imine (PEI), poly (amidoamine), poly (histidine), poly (arginines), and poly amine resins.
- the amine-containing compound is selected from choline, betaine, N,N’- dibenzylethylenediamine, diethylamine, 2-diethylaminoethanol, 2-methylaminoethanol, glucosamine, glucamine, ethanolamine, ethylenediamine, hydrabamine, isopropyl amine, methylglucamine, procaine, triethylamine, trimethylamine, tripropylamine, and tromethamine.
- the cationic moiety R comprises a non-lipid small molecule that is an amino acid.
- the amino acid is selected from arginine, histidine, and lysine.
- the cationic moiety R comprises a non-lipid small molecule that is a heterocycle-containing compound, or a heteroaryl-containing compound.
- that cationic moiety R comprises a non-lipid small molecule selected from caffeine, N- ethylmorpholine, N-ethylpiperidine, morpholine, piperazine, piperidine, purines, and theobromine.
- the amphiphilic polymer is a tri-block copolymer.
- the amphiphilic polymer comprises a block copolymer of Formula II A: X A -Y-X B (IIA) wherein: each of X A and X B is a hydrophobic block comprising a hydrophobic polymer;
- Y is a hydrophilic block comprising a hydrophilic polymer; and the amphiphilic polymer optionally further comprises one or more cleavable linkers.
- the amphiphilic polymer does not comprise a cleavable linker.
- the amphiphilic polymer further comprises a cleavable linker (as described herein).
- the amphiphilic polymer includes one or more cleavable linkers within one or both of the hydrophobic blocks (X A and/or X B ).
- the amphiphilic polymer includes one or more cleavable linkers within the hydrophilic block (Y).
- the amphiphilic polymer comprises a cleavable linker, and is of any one of Formulae IIA-1 to IIA-3:
- the amphiphilic polymer comprises a block copolymer of Formula IIB: Y A -X-Y B (IIB) wherein: each of Y A and Y B is a hydrophilic block comprising a hydrophilic polymer;
- X is a hydrophobic block comprising a hydrophobic polymer; and the amphiphilic polymer optionally further comprises one or more cleavable linkers.
- the amphiphilic polymer does not comprise a cleavable linker.
- the amphiphilic polymer further comprises a cleavable linker (as described herein).
- the amphiphilic polymer includes one or more cleavable linkers within one or both of the hydrophilic blocks (Y A and/or Y B ).
- the amphiphilic polymer includes one or more cleavable linkers within the hydrophobic block (X).
- the amphiphilic polymer further comprises a cleavable linker, and is of any one of Formulae IIB-1 to IIB -3:
- X, X A , or X B comprises a polymer selected from a polyester polymer, a polyorthoester (POE) polymer, a polyanhydride polymer, a polyamide polymer, a poly(ester amide) polymer, a poly(phosphoester) polymer, a poly(alkyl cyanoacrylate) (PACA) polymer, a polysaccharide polymer, and any combination thereof.
- any one of formulae I, IA-ID, or IIA-IIB comprises a polyester polymer. In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, X A , or X B comprises a polyester polymer that is also a polycationic polymer.
- X, X A , or X B comprises a polyester polymer selected from a polylactide (PLA) polymer, a polyglycolide (PGA) polymer, a polycaprolactone (PCL) polymer, a polydioxanone (PDO) polymer, a polyhydroxyalkanoate (PHA) polymer, a poly(glycerol sebacate) polymer, a poly(lactic-co-glycolic acid) (PLGA) polymer, a poly([5- amino ester) (PB AE) polymer, a poly(amine-co-ester) (PACE) polymer, a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) polymer, and any combination thereof.
- PLA polylactide
- PGA polyglycolide
- PCL polycaprolactone
- PDO polydioxanone
- PHA polyhydroxyalkanoate
- PACE poly(glyce
- X, X A , or X B comprises a polylactide (PLA) polymer, where poly-L-lactic acid, poly-D-lactic acid, poly-D,L-lactic acid, poly-L- lactide, poly-D-lactide, and poly-D,L-lactide are all encompassed by the term “PLA.”
- PLA polylactide
- X, X A , or X B comprises a polyglycolide (PGA) polymer.
- any one of formulae I, IA-ID, or IIA-IIB, X, X A , or X B comprises a polydioxanone (PDO) polymer.
- PDO polydioxanone
- X, X A , or X B comprises a polyhydroxyalkanoate (PHA) polymer.
- PHA polyhydroxyalkanoate
- the polyhydroxyalkanoate is a polyhydroxybutyrate polymer.
- the polyhydroxyalkanoate is a polyhydroxy valerate .
- X, X A , or X B comprises a poly(glycerol sebacate) polymer.
- X, X A , or X B comprises a poly(lactic-co-glycolic acid) (PLGA) polymer.
- PLGA is a biocompatible and biodegradable copolymer of lactic acid and glycolic acid, and various forms of PLGA can be characterized by the ratio of lactic acid:glycolic acid.
- Lactic acid can be L-lactic acid, D-lactic acid, or D,L-lactic acid.
- the degradation rate of PLGA can be adjusted by altering the lactic acid-glycolic acid ratio.
- PLGA can be characterized by a lactic acid:glycolic acid ratio of approximately 85:15, approximately 75:25, approximately 60:40, approximately 50:50, approximately 40:60, approximately 25:75, or approximately 15:85.
- the ratio of lactic acid to glycolic acid monomers in the polymer of the particle may be selected to optimize for various parameters such as water uptake, therapeutic agent release and/or polymer degradation kinetics can be optimized.
- X, X A , or X B comprises apo poly(P-amino ester) (PBAE) polymer.
- X, X A , or X B comprises a poly(amine-co-ester) (PACE) polymer.
- PACE poly(amine-co-ester)
- X, X A , or X B comprises a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) polymer.
- PDMAEMA poly(2-(dimethylamino)ethyl methacrylate)
- X, X A , or X B comprises a polyorthoester (POE) polymer.
- POE polyorthoester
- X, X A , or X B comprises a polyanhydride polymer.
- any one of formulae I, IA-ID, or IIA-IIB comprises a polyamide polymer.
- X, X A , or X B comprises a poly(ester amide) polymer.
- X, X A , or X B comprises a poly(alkyl cyanoacrylate) (PACA) polymer.
- X, X A , or X B comprises a poly(phosphoester) polymer.
- X, X A , or X B comprises a polysaccharide polymer.
- the polysaccharide polymer is selected from a chitosan polymer, and a hyaluronic acid (HA) polymer, or a combination thereof.
- Y, Y A , or Y B comprises a polymer selected from a polyethylene glycol (PEG) polymer, a polyethylene oxide (PEG) polymer, a polyglutamic acid (PGA) polymer, a poly[N-(2-hydroxypropyl) methacrylamide] (HPMA) polymer, a poly(vinylpyrrolidone) (PVP) polymer, a poly(2-methyl-2-oxazoline) (PMOX) polymer, a poly(N,N- dimethyl acrylamide) (PDMA) polymer, a poly(N-acryloyl morpholine) (PAcM) polymer, and any combination thereof.
- PEG polyethylene glycol
- PEG polyethylene oxide
- PGA polyglutamic acid
- HPMA poly[N-(2-hydroxypropyl) methacrylamide]
- HPMA poly(vinylpyrrolidone)
- PMOX poly(2-methyl-2-oxazoline)
- PDMA poly(
- Y, Y A , or Y B comprises a polyethylene glycol PEG polymer.
- PEG may be terminated and include an end group, for example, when PEG is not conjugated to another polymer.
- PEG may terminate in a hydroxyl, a methoxy or other alkoxyl group, a methyl or other alkyl group, an aryl group, a carboxylic acid, an amine, an amide, an acetyl group, a guanidino group, or an imidazole.
- Other contemplated end groups include azide, alkyne, maleimide, aldehyde, hydrazide, hydroxylamine, alkoxyamine, or thiol moieties.
- Y, Y A , or Y B comprises a polyethylene glycol (PEG) polymer.
- PEG polyethylene glycol
- Y, Y A , or Y B comprises a polyglutamic acid (PGA) polymer.
- PGA polyglutamic acid
- Y, Y A , or Y B comprises a polyethylene oxide (PEO) polymer.
- PEO polyethylene oxide
- any one of formulae I, IA-ID, or IIA-IIB, Y, Y A , or Y B comprises a poly[N-(2-hydroxypropyl) methacrylamide] (HPMA) polymer.
- HPMA poly(2-hydroxypropyl) methacrylamide]
- PVP poly(vinylpyrrolidone)
- any one of formulae I, IA-ID, or IIA-IIB,Y, Y A , or Y B comprises a poly(2-methyl-2-oxazoline) (PMOX) polymer.
- any one of formulae I, IA-ID, or IIA-IIB,Y, Y A , or Y B comprises a poly(N,N-dimethyl acrylamide) (PDMA) polymer.
- PDMA poly(N,N-dimethyl acrylamide)
- any one of formulae I, IA-ID, or IIA-IIB,Y, Y A , or Y B comprises a poly(N-acryloyl morpholine) (PAcM) polymer.
- PAcM poly(N-acryloyl morpholine)
- X, X A , or X B comprises a polyester polymer
- Y, Y A , or Y B comprises a PEG polymer.
- the molar ratio of Y, Y A , and/or Y B collectively to X, X A , and/or X B collectively is 1:2 to 1:4, such as 1:2 to 1:3 or 1:3 to 1:4.
- the molar ratio of Y, Y A , and/or Y B collectively to X, X A , and/or X B collectively is 1:2 +/- 10%, 1:2.5 +/- 10%, 1:3 +/- 10%, 1:3.5 +/- 10%, or 1:4 +/- 10%.
- the polyester polymer when X, X A , or X B comprises a polyester polymer (as described herein), the polyester polymer has a molecular weight from 5k to 20k, such as 5k to 15k, 5k to 10k, 10k to 15k, or 10k to 20k. In some embodiments, the polyester polymer has a molecular weight of from 5k to 10k. In some embodiments, the polyester has a molecular weight of 5k +/- 10%. In some embodiments, the polyester has a molecular weight of 6k +/- 10%. In some embodiments, the polyester has a molecular weight of 7k +/- 10%.
- the polyester has a molecular weight of 8k +/- 10%. In some embodiments, the polyester has a molecular weight of 9k +/- 10%. In some embodiments, the polyester has a molecular weight of 10k +/- 10%.
- the PEG polymer when Y, Y A , or Y B comprises a PEG polymer, the PEG polymer has a molecular weight from 5k to 10k, such as 5k to 9k, 5k to 8k, 5k to 7k, or 5k to 6k. In some embodiments, the PEG polymer has a molecular weight of 5k +/- 10%. In some embodiments, the PEG polymer has a molecular weight of 6k +/- 10%. In some embodiments, the PEG polymer has a molecular weight of 7k +/- 10%.
- the PEG has a molecular weight of 8k +/- 10%. In some embodiments, the PEG has a molecular weight of 9k +/- 10%. In some embodiments, the PEG has a molecular weight of 10k +/- 10%.
- the amphiphilic polymer comprises a block copolymer of Formula III: [A] v -[B] w -[C] x -[D] y -[E] z (III) wherein:
- A is a polyester monomer or a polyethylene glycol (PEG) monomer
- B is a polyester monomer
- C is a polyester monomer
- D is a polyester monomer
- E is a PEG monomer; v is an integer from 0 to 200 w is an integer from 0 to 200; x is an integer from 0 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150, wherein the amphiphilic polymer optionally further comprises one or more intervening cleavable linkers (L) between any of A-E.
- each of v, w and x are integers greater than 0. In some embodiments of Formula (III), each of v, w and x are integers from 10 to 200.
- each of v, w and x are 0, and the amphiphilic polymer comprises a block copolymer of Formula III A:
- D is a polyester monomer
- E is a PEG monomer; y is an integer from 10 to 200; and z is an integer from 10 to 150 wherein the amphiphilic polymer optionally further comprises one or more intervening cleavable linkers (L) between D and E.
- each of v, and w are 0, and the amphiphilic polymer comprises a block copolymer of Formula IIIB:
- C is a polyester monomer
- D is a polyester monomer
- E is a PEG monomer; x is an integer from 10 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150 wherein the amphiphilic polymer optionally further comprises one or more intervening cleavable linkers (L) between any of C-E.
- the amphiphilic polymer further comprises at least one cleavable linker (L) (as described herein).
- the amphiphilic polymer comprises a cleavable linker between one or more of monomers A and B; monomers B and C; monomers C and D; or monomers D and E.
- the amphiphilic polymer comprises a cleavable linker between monomers D and E.
- the amphiphilic polymer comprises a cleavable linker between one or more of monomers C and D; or D and E.
- the amphiphilic polymer comprises one cleavable linker (E), and is of the Formula IIIC:
- C is a polyester monomer
- L is a cleavable linker
- D is a polyester monomer
- E is a PEG monomer
- x is an integer from 10 to 200
- y is an integer from 10 to 200
- z is an integer from 10 to 150.
- the amphiphilic polymer comprises one cleavable linker (L), and is of the Formula IIID:
- A is a polyester monomer
- B is a polyester monomer
- L is a cleavable linker
- C is a polyester monomer
- D is a polyester monomer
- E is a PEG monomer
- v is an integer from 10 to 200
- w is an integer from 10 to 200
- x is an integer from 10 to 200
- y is an integer from 10 to 200
- z is an integer from 10 to 150.
- C is a polyester monomer
- D is a polyester monomer
- L is a cleavable linker
- E is a PEG monomer
- x is an integer from 0 to 200
- y is an integer from 10 to 200
- z is an integer from 10 to 150.
- x is 0, such that the amphiphilic polymer does not include polyester monomer C. In some embodiments of Formula IIIE, x is 10 to 200, such that the amphiphilic polymer does include polyester monomer C.
- v is 0 and each of w and x are integers from 10 to 200, such that the amphiphilic polymer comprises a block copolymer of Formula IIIF:
- amphiphilic polymer is tri-block copolymer of Formula IV: [A] v -[B] w -[E] z -[C] x -[D] y (IV) wherein:
- A is a polyester monomer
- B is a polyester monomer
- E is a PEG monomer
- C is a polyester monomer
- D is a polyester monomer; v is an integer from 10 to 200 w is an integer from 0 to 200; x is an integer from 0 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150, wherein the amphiphilic polymer optionally further comprises one or more intervening cleavable linkers (L) between any of A-E.
- the amphiphilic polymer does not include a cleavable linker (L).
- the amphiphilic polymer further comprises at least one cleavable linker (E) (as described herein).
- the amphiphilic polymer comprises a cleavable linker between one or more of monomers A and B ; monomers A and E (when w is 0); monomers B and E; monomers E and C; monomers E and D (when x is 0); or monomers C and D.
- w is 0, such that the amphiphilic polymer does not include polyester monomer B. In some embodiments of Formula IV, w is 10 to 200, such that the amphiphilic polymer does include polyester monomer B. In some embodiments of Formula IV, x is 0, such that the amphiphilic polymer does not include polyester monomer C. In some embodiments of Formula IV, x is 10 to 200, such that the amphiphilic polymer does include polyester monomer C.
- each of w and x are 0, such that the amphiphilic polymer comprises a block copolymer of Formula IVA:
- the amphiphilic polymer does not include a cleavable linker (L). In some embodiments of Formula IVA, the amphiphilic polymer further comprises at least one cleavable linker (L) (as described herein). In some embodiments of Formula IVA, the amphiphilic polymer comprises a cleavable linker between one or more of monomers A and E; or monomers E and D.
- each of w and x are 10 to 200, such that the amphiphilic polymer comprises a block copolymer of Formula IVB:
- the amphiphilic polymer does not include a cleavable linker (L). In some embodiments of Formula IVB, the amphiphilic polymer further comprises at least one cleavable linker (L) (as described herein). In some embodiments of Formula IVB, the amphiphilic polymer comprises a cleavable linker between one or more of monomers A and B ; or monomers E and
- amphiphilic polymer is a cross-linked polymer comprising a block copolymer of Formula V : wherein:
- E and F are each independently a PEG monomer
- A, B, C, and D are each independently a polyester monomer; v is an integer from 10 to 200 w is an integer from 0 to 200; x is an integer from 0 to 200; y is an integer from 10 to 200; and z and u are each an integer from 10 to 150.
- w is 0, such that the amphiphilic polymer does not include polyester monomer B. In some embodiments of Formula V, w is 10 to 200, such that the amphiphilic polymer does include polyester monomer B. In some embodiments of Formula V, x is 0, such that the amphiphilic polymer does not include polyester monomer C. In some embodiments of Formula V, x is 10 to 200, such that the amphiphilic polymer does include polyester monomer C.
- each of w and x are integers from 10 to 200, such that the amphiphilic polymer comprises all of polyester monomers A, B, C, and D.
- each of w and x are 0, such that the amphiphilic polymer is a block copolymer of Formula VA:
- the trivalent connecting group (T) is a small molecule having a molecular weight of 1000 Daltons (Da) or less, such as 900 Da or less, 850 Da or less, 800 Da or less, 750 Da or less, 700 Da or less, 650 Da or less, 600 Da or less, 550 Da or less, 500 Da or less, 450 Da or less, 400 Da or less, 350 Da or less, 300 Da or less, 250 Da or less, 200 Da or less, or even less.
- Da Daltons
- the trivalent connecting group (T) is a peptide.
- the peptide comprises cysteine.
- the peptide comprises lysine.
- L is cleaved by exposure to a stimulus.
- the stimulus is selected from pH, temperature, light, redox change, over-expressed enzymes, hypoxia, sound, magnetic force, electrical energy, and any combination thereof.
- the cleavable linker L is a pH-sensitive linker.
- the cleavable linker L is stable at physiological pH (approximately pH 7.4), but is cleaved by exposure to an acidic pH, e.g., in acidic regions of cells, such as lysosomes (approximately pH 4.8).
- L is cleaved by exposure to a specific temperature.
- L is cleaved by exposure to light.
- L is cleaved by exposure to a redox change.
- L is cleaved by exposure to over-expressed enzymes.
- L is cleaved by exposure to hypoxia.
- L is cleaved by exposure to a specific frequency of sound.
- L is cleaved by exposure to a specific magnetic force.
- L is cleaved by exposure to electrical energy.
- the cleavable linker L comprises a group selected from disulfide, hydrazone, vinyl ether, imine, ortho ester, borate ester, amide, a peptide, and azo.
- the cleavable linker L comprises a disulfide.
- the linker comprises a disulfide and can be cleaved by exposure to a redox change.
- the cleavable linker L comprises a hydrazone.
- the linker comprises a hydrazone that can be cleaved by exposure to an acidic pH.
- the linker comprises a hydrazone and can be cleaved by exposure to a pH of 6.5 or less, such as a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 or less, a pH of 4 or less, or even less.
- the cleavable linker comprises a vinyl ether (see e.g., Shin, et al. Molecular Pharmaceutics 2012, 9(11), 3266-3276).
- the linker comprises a vinyl ether that can be cleaved by exposure to an acidic pH.
- the linker comprises a vinyl ether and can be cleaved by exposure to a pH of 6.5 or less, a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 of less, a pH of 4 or less, or even less.
- the cleavable linker comprises an imine, an ortho ester, a borate east, or an amide.
- the linker comprises one or more of an imine, an ortho ester, a borate ester, or an amide that can be cleaved by exposure to an acidic pH (see, e.g., Ding et al. Journal of Controlled Release 2022, 348, 206-238).
- the linker comprises one or more of an imine, an ortho ester, a borate ester, or an amide and can be cleaved by exposure to a pH of 6.5 or less, a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 of less, a pH of 4 or less, or even less.
- the cleavable linker comprises an octapeptide.
- the octapeptide is of the sequence GPLGIAGQ.
- the octapeptide is of the sequence GPLGVRGC.
- the linker comprising an octapeptide is cleaved by exposure to over-expressed enzymes.
- the overexpressed enzyme is matrix metalloproteinase 2 (MMP2) (see e.g., Zhu et al. PNAS 2013, 110(42), 17047-17052).
- the cleavable linker comprises an azo group.
- the linker comprises an azo group that can be cleaved by exposure to hypoxia (see e.g., Joshi et al. International Journal of Pharmaceutics 2020, 590, 119915).
- the amphiphilic polymer comprises a block copolymer of Formula VI: R-[C] x -[D] y -[E] z (VI) wherein:
- R is a cationic moiety
- C is a polyester monomer
- D is a polyester monomer
- E is a PEG monomer
- x is an integer from 0 to 200
- y is an integer from 10 to 200
- z is an integer from 10 to 150.
- x is 0, such that the amphiphilic polymer does not include polyester monomer C.
- w is 10 to 200, such that the amphiphilic polymer does include polyester monomer C.
- the cationic moiety R comprises an ionizable lipid (as described herein).
- the cationic moiety R comprises polyethylene imine (PEI).
- the cationic moiety R comprises poly (amidoamine).
- the cationic moiety R comprises poly (histidine).
- the cationic moiety R comprises poly (arginines).
- the cationic moiety R comprises polyamine resins.
- the cationic moiety R comprises a non-lipid small molecule.
- the non-lipid small molecule is selected from an amine-containing compound, an amino acid, a heterocycle-containing compound, and a heteroaryl-containing compound.
- the cationic moiety R comprises a non-lipid small molecule that is an amine-containing compound.
- the amine-containing compound is selected from choline, betaine, N,N’ -dibenzylethylenediamine, diethylamine, 2- diethylaminoethanol, 2-methylaminoethanol, glucosamine, glucamine, ethanolamine, ethylenediamine, hydrabamine, isopropyl amine, methylglucamine, procaine, triethylamine, trimethylamine, tripropylamine, and tromethamine.
- the cationic moiety R is a non-lipid small molecule that is an amino acid.
- the amino acid is arginine.
- the amino acid is histidine.
- the amino acid is lysine.
- the non-lipid small molecule is a heterocycle-containing compound or a heteroaryl-containing compound.
- the non-lipid small molecule is selected from caffeine, N-ethylmorpholine, N-ethylpiperidine, morpholine, piperazine, piperidine, purines, and theobromine.
- polyester monomers A, B, C and D each independently include a polyester monomer selected from a polylactide (PLA) monomer, a polyglycolide (PGA) monomer, a polycaprolactone (PCL) monomer, a polydioxanone (PDO) monomer, a polyhydroxyalkanoate monomer, a poly(glycerol sebacate) monomer, a poly(P-amino ester) (PBAE) monomer, a poly(2- (dimethylamino)ethyl methacrylate) (PDMAEMA) monomer and any combination thereof.
- PLA polylactide
- PGA polyglycolide
- PCL polycaprolactone
- PDO polydioxanone
- PBAE poly(P-amino ester)
- PDMAEMA poly(2- (dimethylamino)ethyl methacrylate)
- one or more of the polyester monomers A, B, C or D comprises a polylactide (PLA) monomer.
- A comprises a polylactide (PLA) monomer.
- B comprises a polylactide (PLA) monomer.
- C comprises a polylactide (PLA) monomer.
- D comprises a polylactide (PLA) monomer.
- each of A, B, C and D each comprise a polylactide (PLA) monomer.
- one or more of the polyester monomers A, B, C or D comprises a polyglycolide (PGA) monomer.
- A comprises a polyglycolide (PGA) monomer.
- B comprises a polyglycolide (PGA) monomer.
- C comprises a polyglycolide (PGA) monomer.
- D comprises a polyglycolide (PGA) monomer.
- one or more of the polyester monomers A, B, C or D comprises a polycaprolactone (PCL) monomer.
- A comprises a polycaprolactone (PCL) monomer.
- B comprises a polycaprolactone (PCL) monomer.
- C comprises a polycaprolactone (PCL) monomer.
- D comprises a polycaprolactone (PCL) monomer.
- B comprises a polyhydroxyalkanoate monomer.
- C comprises a polyhydroxy alkanoate monomer.
- D comprises a polyhydroxy alkanoate monomer.
- the polyhydroxy alkanolate monomer is a polyhydroxybutyrate monomer.
- the polyhydroxylalkanolate monomer is a polyhydroxy valerate .
- one or more of the polyester monomers A, B, C or D comprises a poly(glycerol sebacate) monomer.
- A comprises a poly(glycerol sebacate) monomer.
- B comprises a poly(glycerol sebacate) monomer.
- C comprises poly(glycerol sebacate) monomer.
- D comprises a poly(glycerol sebacate) monomer.
- one or more of the polyester monomers A, B, C or D comprises a poly([5- amino ester) (PBAE) monomer.
- A comprises a poly(P-amino ester) (PBAE) monomer.
- B comprises a poly(P-amino ester) (PBAE) monomer.
- C comprises a poly(P-amino ester) (PBAE) monomer.
- D comprises a poly(P-amino ester) (PBAE) monomer.
- one or more of the polyester monomers A, B, C or D comprises a poly(2- (dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
- A comprises a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
- B comprises a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
- C comprises a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
- D comprises a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
- [A]v and [B]w together form a copolymer comprising any combination of monomers selected from a polylactide (PLA) monomer, a polyglycolide (PGA) monomer, a polycaprolactone (PCL) monomer, a polydioxanone (PDO) monomer, a polyhydroxyalkanoate (e.g., polyhydroxybutyrate) monomer, a poly(glycerol sebacate) monomer a poly(P-amino ester) (PBAE) monomer, and a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
- PLA polylactide
- PGA polyglycolide
- PCL polycaprolactone
- PDO polydioxanone
- PDO polyhydroxyalkanoate
- PBAE poly(P-amino ester)
- PDMAEMA poly(2-(dimethylamino)ethyl methacryl
- polyester monomers [A] v and [B] w together form a poly(lactic-co- glycolic acid) (PLGA) copolymer.
- the molar ratio of lactic acid monomer units to glycolic acid monomer units in the PLGA copolymer is from 1:1 to 9:1, such as 1:1, 2:1, 3:1, 4:1, 5:1, 6:1, 7:1, 8:1, or 9:1.
- [C] x and [D] y together form a copolymer comprising any combination of monomers selected from a polylactide (PLA) monomer, a polyglycolide (PGA) monomer, a polycaprolactone (PCL) monomer, a polydioxanone (PDO) monomer, a polyhydroxyalkanoate (e.g., polyhydroxybutyrate) monomer, a poly(glycerol sebacate) monomer a poly(P-amino ester) (PBAE) monomer, and a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
- PLA polylactide
- PGA polyglycolide
- PCL polycaprolactone
- PDO polydioxanone
- PDO polyhydroxyalkanoate
- PBAE poly(P-amino ester)
- PDMAEMA poly(2-(dimethylamino)ethyl me
- polyester monomers [C] x and [D] y together form a poly(lactic-co- glycolic acid) (PLGA) copolymer.
- the molar ratio of lactic acid monomer units to glycolic acid monomer units in the PLGA copolymer is from 1:1 to 9:1, such as 1:1, 2:1, 3:1, 4:1, 5:1, 6:1, 7:1, 8:1, or 9:1.
- the subject amphiphilic polymer further comprises one or more terminal groups selected from carboxyl, hydroxyl, amino, amido and alkoxy.
- one or more terminal groups is a carboxyl group.
- one or more terminal groups is a hydroxyl group.
- one or more terminal groups is an amino group.
- one or more terminal groups is an amido group.
- one or more terminal groups is an alkoxy group.
- the terminal group is an alkoxy group selected from n-butoxy (-O-(CH 2 ) 3 )-CH 3 ) and tert-butoxy (-O-(C(CH 3 ) 3 ).
- the subject amphiphilic polymer comprises a di-block copolymer of polyester PLA and a PEG polymer.
- the molecular weight of the PLA polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k.
- the molecular weight of the PEG polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k.
- the PLA and PEG blocks have different molecular weights.
- the PLA block has a molecular weight of 10k, and the PEG block has a molecular weight of 5k (also referred to herein as 10k-5k PLA-PEG). In some embodiments, the PLA block has a molecular weight of 9k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has a molecular weight of 8k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has a molecular weight of 7k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has a molecular weight of 6k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has the same molecular weight as the PEG block. In some embodiments, the PLA block has a molecular weight of 5k, and the PEG block has a molecular weight of 5k.
- the subject amphiphilic polymer is of Formula IIIA, and comprises a diblock copolymer of PLA and PEG polymers represented by the following structure:
- PLA PEG wherein n is 10 to 200 and m is 10 to 150.
- the subject amphiphilic polymer is of Formula IIIA, and comprises a diblock copolymer of PLA and PEG polymers represented by the following structure:
- PLA PEG wherein n is 10 to 200, m is 10 to 150, and each G, independently, can be H, CH3, NH2 or COOH.
- the subject amphiphilic polymer comprises a di-block copolymer of polyester PLGA and a PEG polymer.
- the molecular weight of the PLGA polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k.
- the molecular weight of the PEG polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k.
- the PLGA and PEG blocks have different molecular weights.
- the PLGA block has a molecular weight of 10k, and the PEG block has a molecular weight of 5k (also referred to herein as 10k-5k PLGA-PEG). In some embodiments, the PLGA block has a molecular weight of 9k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 8k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 7k, and the PEG block has a molecular weight of 5k.
- the PLGA block has a molecular weight of 6k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has the same molecular weight as the PEG block. In some embodiments, the PLGA block has a molecular weight of 5k, and the PEG block has a molecular weight of 5k.
- the subject amphiphilic polymer is of Formula III, and comprises a diblock copolymer of PLGA and PEG polymers represented by the following structure:
- PLGA PEG wherein x and y are each 10 to 200, and z is 10 to 150.
- the subject amphiphilic polymer is of Formula III, and comprises a di- block copolymer of PLGA and PEG polymers represented by the following structure:
- PLGA PEG wherein x and y are each 10 to 200, z is 10 to 150, and each G, independently, can be H, CH3, NH2 or COOH.
- the amphiphilic polymer is a PLA-PEG, or a PLGA-PEG copolymer as described in U.S. Pat. Pub. No. US2021/0259982, which is incorporated herein by reference in its entirety.
- the amphiphilic polymer includes a cleavable pH sensitive linker (L).
- the amphiphilic polymer includes a pH sensitive cleavable linker (L) that is a hydrazone.
- the hydrazone can be more stable than an ester, ethylene ether or and imine under a physiological condition of pH 7.4.
- the hydrazone hydrolyzes at a pH of about 5-6, producing ketones and hydrazine. Such a bond can be inserted in some of the formulae described herein.
- PLA-hydrazone-PLA-PEG PLA-hydrazone-PLA-hydrazone-PLA-PEG, and PLA-hydrazone-PEG.
- a hydrazone linker can be inserted into a di-block copolymer of Formula IIID, comprising polyester PLGA (PLA and PGA monomers) and a PEG polymer, to provide the following constructs:
- the subject amphiphilic polymer is of Formula IIIA, and comprises a diblock copolymer of PLA and PEG polymers including a cleavable hydrazone linker represented by the following structure: wherein x is 10 to 200, and z is 10 to 150.
- the subject amphiphilic polymer is of Formula IIIA, and comprises a diblock copolymer of PLA and PEG polymers including a cleavable hydrazone linker represented by the following structure: wherein x is 10 to 200, z is 10 to 150, and each G, independently, can be H, CH3, NH2 or COOH.
- the amphiphilic polymer includes a cleavable linker (L), and the polymer is configured as a cross-linked polymer (see, e.g., Formulae IC, V and VA) to ensure the cleavable linker groups (L) are exposed closer to the surface of the nanoparticle.
- the cleavable linker group (L) is a disulfide, and two copolymers are cross-linked via a disulfide bond using cysteine connecting groups.
- the cross-linked copolymers are PLA-PEG.
- the cross-linked copolymers are PLGA-PEG.
- the subject amphiphilic polymer is of the Formula VA, and comprises PLA-PEG copolymers linked via a disulfide bond and cysteine connecting groups: wherein x is 10 to 200, and z is 10 to 150.
- the subject amphiphilic polymer is of the Formula VA, and comprises
- PLA-PEG copolymers linked via a disulfide bond and cysteine connecting groups wherein x is 10 to 200, z is 10 to 150, and each G, independently, can be H, CH3, NH2 or COOH.
- the cleavable linker group (L) is a disulfide, and two copolymers are cross-linked via a hydrazone bond using lysine connecting groups.
- the crosslinked copolymers are PLA-PEG.
- the cross-linked copolymers are PLGA-PEG.
- the subject amphiphilic polymer is of the Formula VA, and comprises PLA-PEG copolymers linked via a cleavable linker comprising hydrazone groups and lysine connecting groups: wherein x is 10 to 200, and z is 10 to 150.
- the subject amphiphilic polymer is of the Formula VA, and comprises PLA-PEG copolymers linked via a cleavable linker comprising hydrazone groups and lysine connecting groups:
- x is 10 to 200
- z is 10 to 150
- each G independently, can independently be H, CH3, NH2 or COOH.
- FIG. 13 is a graphic representation of the pH mediated disruption of a polymeric lipid nanoparticle comprising a shell of cross-linked amphiphilic polymers having hydrazone linkages.
- the subject amphiphilic polymer comprises a tri-block copolymer of polyester PL A and a PEG polymer.
- the molecular weight of the PL A polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k.
- the molecular weight of the PEG polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k.
- the PL A and PEG blocks have different molecular weights.
- the PL A block has a molecular weight of 10k, and the PEG block has a molecular weight of 5k. In some embodiments, the PL A block has a molecular weight of 9k, and the PEG block has a molecular weight of 5k. In some embodiments, the PL A block has a molecular weight of 8k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has a molecular weight of 7k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has a molecular weight of 6k, and the PEG block has a molecular weight of 5k.
- the PLA block has the same molecular weight as the PEG block. In some embodiments, the PLA block has a molecular weight of 5k, and the PEG block has a molecular weight of 5k.
- the subject amphiphilic polymer is of Formula IVA, and comprises a tri-block copolymer of PL A and PEG polymers represented by the following structure: wherein v and y independently are 10 to 200, and z is 10 to 150.
- the subject amphiphilic polymer is of Formula IVA, and comprises a tri- block copolymer of PL A and PEG polymers represented by the following structure: wherein v and y independently are 10 to 200, z is 10 to 150, and each G can independently be H, CH3,
- the subject amphiphilic polymer comprises a tri-block copolymer of polyester PLGA and a PEG polymer.
- the molecular weight of the PLGA polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k.
- the molecular weight of the PEG polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k.
- the PLGA and PEG blocks have different molecular weights.
- the PLGA block has a molecular weight of 10k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 9k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 8k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 7k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 6k, and the PEG block has a molecular weight of 5k.
- the PLGA block has the same molecular weight as the PEG block. In some embodiments, the PLGA block has a molecular weight of 5k, and the PEG block has a molecular weight of 5k.
- the subject amphiphilic polymer is of Formula IVB, and comprises a triblock copolymer of PLGA and PEG polymers represented by the following structure:
- PLGA PEG PLGA wherein each x and y independently are independently 10 to 200, and z is 10 to 150.
- the subject amphiphilic polymer is of Formula IVB, and comprises a tri- block copolymer of PLGA and PEG polymers represented by the following structure:
- PLGA PEG PLGA wherein each x and y independently are independently 10 to 200, z is 10 to 150, and each G can independently be H, CH3, NH2 or COOH.
- polymeric lipid nanoparticles using the triblock polymers shown above, (i.e., PLA-PEG-PLA, and PLGA-PEG-PLGA), the two hydrophobic polymer blocks (PLA or PLGA) can be embedded in the nanoparticle shell, whereas the PEG polymer block can form a loop on the outside of the particle.
- the PEG would not have any end groups and may lead to reduced anti-PEG IgM formation and reduced accelerated blood clearage (ABC) of polymeric lipid nanoparticles after subsequent doses.
- FIG. 14 The graphical representation of polymeric lipid nanoparticles prepared using such tri-block polymer is shown in FIG. 14.
- the subject amphiphilic polymer is of Formula VI and comprises an ionizable lipid as the cationic moiety conjugated to the polymer, such that the ionizable lipid can anchor the core to the shell of the particle.
- a lipid such as ethyl lauroyl arginate (ELA) can be conjugated to a PLA-PEG polymer, forming PEG-PLA-ELA.
- ELA ethyl lauroyl arginate
- PEG-PLA-ELA ethyl lauroyl arginate
- the subject amphiphilic polymer is of Formula IV, and comprises a copolymer of PLA and PEG polymers conjugated to ionizable lipid R, represented by the following structure:
- PLA PEG wherein x is 10 to 200, z is 10 to 150, and R is an ionizable lipid (as described herein).
- the subject amphiphilic polymer is of Formula IV, and comprises a copolymer of PLA and PEG polymers conjugated to ionizable lipid R, represented by the following structure:
- PLA PEG wherein x is 10 to 200, z is 10 to 150, R is an ionizable lipid (as described herein), and G can be H,
- the subject amphiphilic polymer is of Formula IV, and comprises a copolymer of PLGA and PEG polymers conjugated to ionizable lipid R, represented by the following structure: wherein x and y are each independently 10 to 200, z is 10 to 150, and R is an ionizable lipid (as described herein).
- the subject amphiphilic polymer is of Formula IV, and comprises a copolymer of PLGA and PEG polymers conjugated to ionizable lipid R, represented by the following structure:
- PLGA PEG wherein x and y are each independently 10 to 200, z is 10 to 150, R is an ionizable lipid (as described herein), and G can be H, CH3, NH2 or COOH.
- the present disclosure provides emulsions comprising polymeric lipid nanoparticles. Also provided are methods of preparing the subject polymeric lipid nanoparticles.
- the polypeptide is detectable at a therapeutic level in patient tissue (e.g., liver or lung).
- the level of detectable polypeptide is from continuous expression from the circRNA composition over periods of time of more than one, more than four, more than six, more than 12, more than 24, more than 48, or more than 72 hours after administration.
- a protein encoded by circRNA is produced at levels above normal physiological levels.
- the level of protein may be increased as compared to a control.
- the control is the baseline physiological level of the polypeptide in a normal individual or in a population of normal individuals.
- the control is the baseline physiological level of the polypeptide in an individual having a deficiency in the relevant protein or polypeptide or in a population of individuals having a deficiency in the relevant protein or polypeptide.
- the control can be the normal level of the relevant protein or polypeptide in the individual to whom the composition is administered.
- the control is the expression level of the polypeptide upon other therapeutic intervention, e.g., upon direct injection of the corresponding polypeptide, at one or more comparable time points.
- the levels of a protein encoded by a circRNA are detectable at 3 days, 4 days, 5 days, or 1 week or more after administration. Increased levels of protein may be observed in a tissue (e.g., liver or lung).
- Delivery systems also include non-polymer systems that are lipids including sterols such as cholesterol, cholesterol esters, and fatty acids or neutral fats such as mono-di-and tri-glycerides; hydrogel release systems; sylastic systems; peptide based systems: wax coatings; compressed tablets using conventional binders and excipients; partially fused implants; and the like.
- Specific examples include but are not limited to: (a) erosional systems in which the active composition is contained in a form within a matrix such as those described in U.S. Patents 4,452,775, 4,667,014, 4,748,034, and 5,239,660 and (b) diffusional systems in which an active component permeates at a controlled rate from a polymer such as described in U.S. Patents 3,832,253 and 3,854,480.
- pump-based hardware delivery systems can be used, some of which are adapted for implantation.
- the therapeutic agent can be conjugated either directly or indirectly through a linking moiety to a targeting moiety.
- Methods for conjugating therapeutic agents to targeting moieties is known in the art. See, for instance, Wadwa et al., J, Drug Targeting 3:111 (1995) and U.S. Patent 5,087,616.
- the therapeutic agents provided herein are formulated into a depot form, such that the manner in which the therapeutic agent is released into the body to which it is administered is controlled with respect to time and location within the body (see, for example, U.S. Patent 4,450,150).
- Depot forms of therapeutic agents can be, for example, an implantable composition comprising the therapeutic agents and a porous or non-porous material, such as a polymer, wherein the therapeutic agents are encapsulated by or diffused throughout the material and/or degradation of the non-porous material.
- the depot is then implanted into the desired location within the body and the therapeutic agents are released from the implant at a predetermined rate.
- the present disclosure also contemplates the discriminatory targeting of target cells and tissues by both passive and active targeting means.
- the phenomenon of passive targeting exploits the natural distributions patterns of a transfer vehicle in vivo without relying upon the use of additional excipients or means to enhance recognition of the transfer vehicle by target cells.
- transfer vehicles which are subject to phagocytosis by the cells of the reticulo-endothelial system are likely to accumulate in the liver or spleen, and accordingly may provide a means to passively direct the delivery of the subject compositions to such target cells.
- the present disclosure contemplates active targeting, which involves the use of targeting moieties that may be bound (either covalently or non-covalently) to the polymeric lipid nanoparticle to encourage localization of such at certain target cells or target tissues.
- targeting may be mediated by the inclusion of one or more endogenous targeting moieties in or on the polymeric lipid nanoparticle to encourage distribution to the target cells or tissues.
- the composition can comprise a moiety capable of enhancing affinity of the composition to the target cell.
- Targeting moieties may be linked to the outer polymeric layer of the polymeric lipid nanoparticle during formulation or post-formulation.
- compositions of the present disclosure demonstrate improved transfection efficacies, and/or demonstrate enhanced selectivity towards target cells or tissues of interest.
- compositions which comprise one or more moieties (e.g., peptides, aptamers, oligonucleotides, vitamins or other molecules) that are capable of enhancing the affinity of the compositions and their nucleic acid contents for the target cells or tissues.
- moieties may optionally be bound or linked to the surface of the nanoparticle.
- the targeting moiety may span the surface of a nanoparticle or be encapsulated within the nanoparticle.
- Suitable moieties and are selected based upon their physical, chemical or biological properties (e.g., selective affinity and/or recognition of target cell surface markers or features). Cell-specific target sites and their corresponding targeting ligand can vary widely.
- compositions of the present disclosure may include surface markers (e.g., apolipoprotein-B (APOB) or apolipoprotein-E (APOE)) that selectively enhance recognition of, or affinity to hepatocytes (e.g., by receptor-mediated recognition of and binding to such surface markers).
- surface markers e.g., apolipoprotein-B (APOB) or apolipoprotein-E (APOE)
- APOB apolipoprotein-B
- APOE apolipoprotein-E
- the use of galactose as a targeting moiety would be expected to direct the compositions of the present disclosure to parenchymal hepatocytes, or alternatively the use of mannose containing sugar residues as a targeting ligand would be expected to direct the compositions of the present disclosure to liver endothelial cells (e.g., mannose containing sugar residues that may bind preferentially to the asialoglycoprotein receptor present in hepatocytes).
- liver endothelial cells e.g., mannose containing sugar residues that may bind preferentially to the asialoglycoprotein receptor present in hepatocytes.
- targeting moieties that have been conjugated to moieties present in the polymeric lipid nanoparticle composition therefore facilitate recognition and uptake of the compositions of the present disclosure in target cells and tissues.
- suitable targeting moieties include one or more peptides, proteins, aptamers, vitamins and oligonucleotides .
- a polymeric lipid nanoparticle composition comprises a targeting moiety.
- the targeting moiety mediates receptor-mediated endocytosis selectively into a specific population of cells.
- the targeting moiety is operably connected, or linked, to the transfer vehicle.
- the targeting moiety is capable of binding to an immune cell antigen.
- the targeting moiety is capable of binding to a T cell antigen.
- T cell antigens include, but are not limited to, CD2, CD3, CD5, CD7, CD8, CD4, beta7 integrin, beta2ingetrin, and ClqR.
- the targeting moiety is capable of binding to a NK, NKT, or macrophage antigen. In some embodiments, the targeting moiety is capable of binding to a protein selected from CD3, CD4, CD8, PD-1, 4-1BB, and CD2. In some embodiments, the targeting moiety is a single chain Fv (scFv) fragment, nanobody, peptide, peptide-based macrocycle, minibody, heavy chain variable region, light chain variable region or fragment thereof.
- scFv single chain Fv
- the targeting moiety is selected from T-cell receptor motif antibodies, T-cell a chain antibodies, T-cell P chain antibodies, T-cell y chain antibodies, T-cell 5 chain antibodies, CCR7 antibodies, CD3 antibodies, CD4 antibodies, CD5 antibodies, CD7 antibodies, CD8 antibodies, CDl lb antibodies, CDl lc antibodies, CD16 antibodies, CD19 antibodies, CD20 antibodies, CD21 antibodies, CD22 antibodies, CD25 antibodies, CD28 antibodies, CD34 antibodies, CD35 antibodies, CD40 antibodies, CD45RA antibodies, CD45RO antibodies, CD52 antibodies, CD56 antibodies, CD62L antibodies, CD68 antibodies, CD80 antibodies, CD95 antibodies, CD117 antibodies, CD127 antibodies, CD133 antibodies, CD137 (4-1BB) antibodies, CD163 antibodies, F4/80 antibodies, IL-4Ra antibodies, Sca-1 antibodies, CTLA-4 antibodies, GITR antibodies GARP antibodies, LAP antibodies, granzyme B antibodies, LFA-1 antibodies, transferrin receptor antibodies, and fragments thereof.
- the targeting moiety is a small molecule binder of an ectoenzyme on lymphocytes.
- Small molecule binders of ectoenzymes include A2A inhibitors CD73 inhibitors, CD39 or adesines receptors A2aR and A2bR.
- Potential small molecules include AB928.
- the immune cell represents the target cell.
- the compositions of the present disclosure transfect the target cells on a discriminatory basis (i.e., do not transfect non-target cells).
- the compositions of the present disclosure may also be prepared to preferentially target a variety of target cells, which include, but are not limited to, T cells, B cells, macrophages, and dendritic cells.
- the target cells are deficient in a protein or enzyme of interest.
- the hepatocyte represents the target cell.
- the compositions of the present disclosure transfect the target cells on a discriminatory basis (i.e., do not transfect non-target cells).
- compositions of the present disclosure may also be prepared to preferentially target a variety of target cells, which include, but are not limited to, hepatocytes, epithelial cells, hematopoietic cells, epithelial cells, endothelial cells, lung cells, bone cells, stem cells, mesenchymal cells, neural cells (e.g., meninges, astrocytes, motor neurons, cells of the dorsal root ganglia and anterior horn motor neurons), photoreceptor cells (e.g., rods and cones), retinal pigmented epithelial cells, secretory cells, cardiac cells, adipocytes, vascular smooth muscle cells, cardiomyocytes, skeletal muscle cells, beta cells, pituitary cells, synovial lining cells, ovarian cells, testicular cells, fibroblasts, B cells, T cells, reticulocytes, leukocytes, granulocytes and tumor cells.
- target cells include, but are not limited to, hepatocytes,
- compositions of the present disclosure may be prepared to preferentially distribute to target cells such as in the heart, lungs, kidneys, liver, and spleen.
- the compositions of the present disclosure distribute into the cells of the liver or spleen to facilitate the delivery and the subsequent expression of the circRNA comprised therein by the cells of the liver (e.g., hepatocytes) or the cells of spleen (e.g., immune cells).
- the targeted cells may function as a biological “reservoir” or “depot” capable of producing, and systemically excreting a functional protein or enzyme.
- the transfer vehicle may target hepatocytes or immune cells and/or preferentially distribute to the cells of the liver or spleen upon delivery.
- the circRNA loaded in the nanoparticle are translated and a functional protein product is produced, excreted and systemically distributed.
- cells other than hepatocytes e.g., lung, spleen, heart, ocular, or cells of the central nervous system
- the compositions of the present disclosure facilitate a subject's endogenous production of one or more functional proteins and/or enzymes.
- the polymeric lipid nanoparticles comprise circRNA which encode a deficient protein or enzyme.
- the exogenous circRNA loaded into the nanoparticle may be translated in vivo to produce a functional protein or enzyme encoded by the exogenously administered circRNA (e.g., a protein or enzyme in which the subject is deficient).
- the compositions of the present disclosure exploit a subject's ability to translate exogenously- or recombinantly-prepared circRNA to produce an endogenously-translated protein or enzyme, and thereby produce (and where applicable excrete) a functional protein or enzyme.
- the expressed or translated proteins or enzymes may also be characterized by the in vivo inclusion of native post-translational modifications which may often be absent in recombinantly-prepared proteins or enzymes, thereby further reducing the immunogenicity of the translated protein or enzyme.
- circRNA encoding a deficient protein or enzyme avoids the need to deliver the nucleic acids to specific organelles within a target cell. Rather, upon transfection of a target cell and delivery of the nucleic acids to the cytoplasm of the target cell, the circRNA contents of a transfer vehicle may be translated and a functional protein or enzyme expressed.
- a circular RNA comprises one or more miRNA binding sites.
- a circular RNA comprises one or more miRNA binding sites recognized by miRNA present in one or more non-target cells or non-target cell types (e.g., Kupffer cells or hepatic cells) and not present in one or more target cells or target cell types (e.g., hepatocytes or T cells).
- a circular RNA comprises one or more miRNA binding sites recognized by miRNA present in an increased concentration in one or more non-target cells or non-target cell types (e.g., Kupffer cells or hepatic cells) compared to one or more target cells or target cell types (e.g., hepatocytes or T cells). miRNAs are thought to function by pairing with complementary sequences within RNA molecules, resulting in gene silencing.
- the compositions of the present disclosure transfect or distribute to target cells on a discriminatory basis (i.e., do not transfect non-target cells).
- the compositions of the present disclosure may also be prepared to preferentially target a variety of target cells, which include, but are not limited to, hepatocytes, epithelial cells, hematopoietic cells, epithelial cells, endothelial cells, lung cells, bone cells, stem cells, mesenchymal cells, neural cells (e.g., meninges, astrocytes, motor neurons, cells of the dorsal root ganglia and anterior horn motor neurons), photoreceptor cells (e.g., rods and cones), retinal pigmented epithelial cells, secretory cells, cardiac cells, adipocytes, vascular smooth muscle cells, cardiomyocytes, skeletal muscle cells, beta cells, pituitary cells, synovial lining cells, ovarian cells, testicular cells, fibroblasts,
- target cells include,
- provided herein is a method of producing a protein of interest in a subject in need thereof by introducing or administering an effective amount of a pharmaceutical composition described herein comprising at least one polymeric lipid nanoparticle described herein.
- provided herein is a method of treating and/or preventing a condition comprising administering an effective amount of a pharmaceutical composition described herein comprising at least one polymeric lipid nanoparticle described herein.
- the pharmaceutical compositions described herein are co-administered with one or more additional therapeutic agents (e.g., in the same pharmaceutical composition or in separate pharmaceutical compositions).
- the pharmaceutical compositions provided herein can be administered first and the one or more additional therapeutic agents can be administered second, or vice versa.
- the pharmaceutical compositions provided herein and the one or more additional therapeutic agents can be administered simultaneously.
- the subject is a mammal.
- the mammal referred to herein can be any mammal, including, but not limited to, mammals of the order Rodentia, such as mice and hamsters, or mammals of the order Logomorpha, such as rabbits.
- the mammals may be from the order Carnivora, including Felines (cats) and Canines (dogs).
- the mammals may be from the order Artiodactyla, including Bovines (cows) and Swines (pigs), or of the order Perssodactyla, including Equines (horses).
- the mammals may be of the order Primates, Ceboids, or Simoids (monkeys) or of the order Anthropoids (humans and apes).
- the mammal is a human.
- provided herein is a method of vaccinating a subject by introducing or administering an effective amount of a pharmaceutical composition described herein comprising at least one polymeric lipid nanoparticle described herein.
- the method of vaccinating comprises administering an effective amount of an antigen comprising a viral polypeptide from an adenovirus; Herpes simplex, type 1; Herpes simplex, type 2; encephalitis virus; papillomavirus; Varicella-zoster virus; Epstein-barr virus; Human cytomegalovirus; Human herpes virus, type 8; Human papillomavirus; BK virus; JC virus; Smallpox; polio virus; Hepatitis B virus; Human bocavirus; Parvovirus B19; Human astrovirus; Norwalk virus; coxsackievirus; hepatitis A virus; poliovirus; rhinovirus; Severe acute respiratory syndrome virus; Hepatitis C virus; Yellow Fever virus; Dengue virus; West Nile virus; Rubella virus; Hepatitis E virus; Human Immunodeficiency virus (HIV); Influenza virus; Guanarito virus; Junin virus;
- provided herein is a method of treating an autoimmune disorder in a subject by introducing or administering an effective amount of a pharmaceutical composition described herein comprising at least one polymeric lipid nanoparticle described herein.
- provided herein is a method of treating cancer in a subject by introducing or administering an effective amount of a pharmaceutical composition described herein comprising at least one polymeric lipid nanoparticle described herein.
- the circular RNA construct encodes a CAR
- the CARs have biological activity, e.g., ability to recognize an antigen, e.g., CD19, HER2, or BCMA, such that the CAR, when expressed by a cell, is able to mediate an immune response against the cell expressing the antigen, e.g., CD19, HER2, or BCMA, for which the CAR is specific.
- CAR-T chimeric antigen receptor
- CRS cytokine release syndrome
- CRES CAR-T cell-related encephalopathy syndrome
- CRS is the most common and well-described toxicity associated with CAR-T therapy, occurring in over 90% of patients at any grade and is characterized by high fever, hypotension, hypoxia and/ or multiple organ toxicity and can lead to death.
- Neurotoxicity is characterized by damage to nervous tissue that can cause tremors, encephalopathy, dizziness or seizures.
- lymphodepletion is known to increase CAR- T cell expansion and enhanced efficacy of infused CAR-T cells by, for example, altering the tumor phenotype and microenvironment.
- lymphodepletion agents often cause side effects to the patients.
- lymphodepletion can cause neutropenia, anemia, thrombocytopenia, and immunosuppression, leading to a greater risk of infection, along with other toxicities.
- CAR-T therapies require an assortment of protocols to isolate, genetically modify, and selectively expand the redirected cells before infusing them back into the patient.
- the subject has a cancer selected from the group consisting of: acute myeloid leukemia (AML); alveolar rhabdomyosarcoma; B cell malignancies; bladder cancer (e.g., bladder carcinoma); bone cancer; brain cancer (e.g., medulloblastoma and glioblastoma multiforme); breast cancer; cancer of the anus, anal canal, or anorectum; cancer of the eye; cancer of the intrahepatic bile duct; cancer of the joints; cancer of the neck; gallbladder cancer; cancer of the pleura; cancer of the nose, nasal cavity, or middle ear; cancer of the oral cavity; cancer of the vulva; chronic lymphocytic leukemia; chronic myeloid cancer; colon cancer; esophageal cancer, cervical cancer; fibrosarcoma; gastrointestinal carcinoid tumor; head and neck cancer (e.g., head and neck squamous cell carcinoma); Hodgkin lymphom
- AML acute myeloid
- the subject has an autoimmune disorder selected from scleroderma, Grave's disease, Crohn's disease, Sjogren's disease, multiple sclerosis, Hashimoto's disease, psoriasis, myasthenia gravis, autoimmune polyendocrinopathy syndromes, Type I diabetes mellitus (TIDM), autoimmune gastritis, autoimmune uveoretinitis, polymyositis, colitis, thyroiditis, and the generalized autoimmune diseases typified by human Lupus.
- TIDM Type I diabetes mellitus
- RNA refers to a single-stranded RNA polynucleotide wherein the 3’ and 5’ ends that are normally present in a linear RNA polynucleotide have been joined together.
- splice site refers to a dinucleotide that is partially or fully included in a group I intron and between which a phosphodiester bond is cleaved during RNA circularization.
- splice site refers to the dinucleotide or dinucleotides between which cleavage of the phosphodiester bond occurs during a splicing reaction.
- a “5’ splice site” refers to the natural 5’ dinucleotide of the intron e.g., group I intron, while a “3’ splice site” refers to the natural 3’ dinucleotide of the intron).
- a “spacer” refers to a region of a polynucleotide sequence ranging from 1 nucleotide to hundreds or thousands of nucleotides separating two other elements along a polynucleotide sequence.
- the sequences can be defined or can be random.
- a spacer is typically noncoding. In some embodiments, spacers include duplex regions.
- Linear nucleic acid molecules are said to have a “5 ’-terminus” (5’ end) and a “3 ’-terminus” (3’ end) because nucleic acid phosphodiester linkages occur at the 5’ carbon and 3’ carbon of the sugar moieties of the substituent mononucleotides.
- the end nucleotide of a polynucleotide at which a new linkage would be to a 5’ carbon is its 5’ terminal nucleotide.
- the end nucleotide of a polynucleotide at which a new linkage would be to a 3’ carbon is its 3’ terminal nucleotide.
- a terminal nucleotide, as used herein, is the nucleotide at the end position of the 3’- or 5 ’-terminus.
- Transformation means the formation of a polypeptide molecule by a ribosome based upon an RNA template.
- the term “encode” refers broadly to any process whereby the information in a polymeric macromolecule is used to direct the production of a second molecule that is different from the first.
- the second molecule may have a chemical structure that is different from the chemical nature of the first molecule.
- co-administering is meant administering a therapeutic agent provided herein in conjunction with one or more additional therapeutic agents sufficiently close in time such that the therapeutic agent provided herein can enhance the effect of the one or more additional therapeutic agents, or vice versa.
- an “internal ribosome entry site” or “IRES” refers to an RNA sequence or structural element ranging in size from 10 nt to 1000 nt or more, capable of initiating translation of a polypeptide in the absence of a typical RNA cap structure.
- An IRES is typically 500 nt to 700 nt in length.
- aptamer refers in general to either an oligonucleotide of a single defined sequence or a mixture of said nucleotides, wherein the mixture retains the properties of binding specifically to the target molecule (e.g., eukaryotic initiation factor, 40S ribosome, polyC binding protein, polyA binding protein, polypyrimidine tract-binding protein, argonaute protein family, Heterogeneous nuclear ribonucleoprotein K and La and related RNA-binding protein).
- target molecule e.g., eukaryotic initiation factor, 40S ribosome, polyC binding protein, polyA binding protein, polypyrimidine tract-binding protein, argonaute protein family, Heterogeneous nuclear ribonucleoprotein K and La and related RNA-binding protein.
- aptamer is meant to refer to a single- or double-stranded nucleic acid which is capable of binding to a protein or other molecule.
- aptamers preferably comprise 10 to 100 nucleotides, preferably 15 to 40 nucleotides, more preferably 20 to 40 nucleotides, in that oligonucleotides of a length that falls within these ranges are readily prepared by conventional techniques.
- aptamers can further comprise a minimum of approximately 6 nucleotides, preferably 10, and more preferably 14 or 15 nucleotides, that are necessary to effect specific binding.
- an “eukaryotic initiation factor” or “elF” refers to a protein or protein complex used in assembling an initiator tRNA, 40S and 60S ribosomal subunits required for initiating eukaryotic translation.
- an “internal ribosome entry site” or “IRES” refers to an RNA sequence or structural element ranging in size from 10 nt to 1000 nt or more, capable of initiating translation of a polypeptide in the absence of a typical RNA cap structure.
- An IRES is typically 500 nt to 700 nt in length.
- a “miRNA site” refers to a stretch of nucleotides within a polynucleotide that is capable of forming a duplex with at least 8 nucleotides of a natural miRNA sequence.
- an “endonuclease site” refers to a stretch of nucleotides within a polynucleotide that is capable of being recognized and cleaved by an endonuclease protein.
- bicistronic RNA refers to a polynucleotide that includes two expression sequences coding for two distinct proteins. These expression sequences can be separated by a nucleotide sequence encoding a cleavable peptide such as a protease cleavage site. They can also be separated by a ribosomal skipping element.
- ribosomal skipping element refers to a nucleotide sequence encoding a short peptide sequence capable of causing generation of two peptide chains from translation of one RNA molecule. While not wishing to be bound by theory, it is hypothesized that ribosomal skipping elements function by (1) terminating translation of the first peptide chain and re-initiating translation of the second peptide chain; or (2) cleavage of a peptide bond in the peptide sequence encoded by the ribosomal skipping element by an intrinsic protease activity of the encoded peptide, or by another protease in the environment (e.g., cytosol).
- the term “co-formulate” refers to a nanoparticle formulation comprising two or more nucleic acids or a nucleic acid and other active drug substance. Typically, the ratios are equimolar or defined in the ratiometric amount of the two or more nucleic acids or the nucleic acid and other active drug substance.
- the phrase “ionizable lipid” refers to any of a number of lipid species that carry a net positive charge at a selected pH, such as physiological pH 4 and a neutral charge at other pHs such as physiological pH 7.
- an ionizable lipid or polymer e.g., an amphiphilic polymer, disclosed herein comprises one or more cleavable groups.
- cleave and cleavable are used herein to mean that one or more chemical bonds (e.g., one or more of covalent bonds, hydrogen-bonds, van der Waals' forces and/or ionic interactions) between atoms in or adjacent to the subject functional group are broken (e.g., hydrolyzed) or are capable of being broken upon exposure to selected conditions (e.g., upon exposure to enzymatic conditions).
- the cleavable group is a disulfide functional group, and in particular embodiments is a disulfide group that is capable of being cleaved upon exposure to selected biological conditions (e.g., intracellular conditions).
- the cleavable group is an ester functional group that is capable of being cleaved upon exposure to selected biological conditions.
- the disulfide groups may be cleaved enzymatically or by a hydrolysis, oxidation or reduction reaction. Upon cleavage of such disulfide functional group, the one or more functional moieties or groups (e.g., one or more of a head-group and/or a tail-group) that are bound thereto may be liberated.
- a cleavable group is bound (e.g., bound by one or more of hydrogen-bonds, van der Waals' forces, ionic interactions and covalent bonds) to one or more functional moieties or groups.
- Compound described herein may also comprise one or more isotopic substitutions.
- H may be in any isotopic form, including 1H, 2H (D or deuterium), and 3H (T or tritium);
- C may be in any isotopic form, including 12C, 13C, and 14C;
- 0 may be in any isotopic form, including 160 and 180;
- F may be in any isotopic form, including 18F and 19F; and the like.
- Cl-6 alkyl is intended to encompass, Cl, C2, C3, C4, C5, C6, Cl-6, Cl-5, Cl-4, Cl-3, Cl-2, C2-6, C2-5, C2-4, C2-3, C3-6, C3-5, C3-4, C4-6, C4-5, and C5-6 alkyl.
- head-group as used describe the compounds of the present invention, and in particular functional groups that comprise such compounds, are used for ease of reference to describe the orientation of one or more functional groups relative to other functional groups.
- a hydrophilic head-group e.g., an amino group
- a hydrophobic tail-group e.g., cholesterol
- the present invention is intended to encompass the compounds disclosed herein, and the pharmaceutically acceptable salts, pharmaceutically acceptable esters, tautomeric forms, polymorphs, and prodrugs of such compounds.
- the present invention includes a pharmaceutically acceptable addition salt, a pharmaceutically acceptable ester, a solvate (e.g., hydrate) of an addition salt, a tautomeric form, a polymorph, an enantiomer, a mixture of enantiomers, a stereoisomer or mixture of stereoisomers (pure or as a racemic or non-racemic mixture) of a compound described herein.
- Compounds described herein can comprise one or more asymmetric centers, and thus can exist in various isomeric forms, e.g., enantiomers and/or diastereomers.
- the compounds described herein can be in the form of an individual enantiomer, diastereomer or geometric isomer, or can be in the form of a mixture of stereoisomers, including racemic mixtures and mixtures enriched in one or more stereoisomer.
- Isomers can be isolated from mixtures by methods known to those skilled in the art, including chiral high pressure liquid chromatography (HPLC) and the formation and crystallization of chiral salts; or preferred isomers can be prepared by asymmetric syntheses.
- HPLC high pressure liquid chromatography
- the subject compositions exhibit an enhanced (e.g., increased) ability to transfect one or more target cells.
- methods of transfecting one or more target cells generally comprise the step of contacting the one or more target cells with the compositions disclosed herein such that the one or more target cells are transfected with the circular RNA encapsulated therein.
- transfect or “transfection” refer to the intracellular introduction of one or more encapsulated materials (e.g., nucleic acids and/or polynucleotides) into a cell, or preferably into a target cell.
- transfection efficiency refers to the relative amount of such encapsulated material (e.g., polynucleotides) up-taken by, introduced into and/or expressed by the target cell which is subject to transfection.
- nucleotide sequences disclosed herein can represent an RNA sequence or a corresponding DNA sequence. It is understood that deoxythymidine (dT or T) in a DNA is transcribed into a uridine (U) in an RNA. As such, “T” and “U” are used interchangeably herein in nucleotide sequences.
- sequence identity or, for example, comprising a “sequence 50% identical to,” as used herein, refer to the extent that sequences are identical on a nucleotide-by-nucleotide basis or an amino acid-by-amino acid basis over a window of comparison.
- a “percentage of sequence identity” may be calculated by comparing two optimally aligned sequences over the window of comparison, determining the number of positions at which the identical nucleic acid base (e.g., A, T, C, G, I) or the identical amino acid residue (e.g., Ala, Pro, Ser, Thr, Gly, Vai, Leu, He, Phe, Tyr, Trp, Lys, Arg, His, Asp, Glu, Asn, Gin, Cys and Met) occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison (i.e., the window size), and multiplying the result by 100 to yield the percentage of sequence identity.
- the identical nucleic acid base e.g., A, T, C, G, I
- the identical amino acid residue e.g., Ala, Pro, Ser, Thr, Gly, Vai, Leu, He, Phe, Tyr, Trp, Lys,
- nucleotides and polypeptides having at least 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity to any of the reference sequences described herein, typically where the polypeptide variant maintains at least one biological activity of the reference polypeptide.
- the expression sequences in the polynucleotide construct may be separated by a “cleavage site” sequence which enables polypeptides encoded by the expression sequences, once translated, to be expressed separately by the cell.
- a “self-cleaving peptide” refers to a peptide which is translated without a peptide bond between two adjacent amino acids, or functions such that when the polypeptide comprising the proteins and the self-cleaving peptide is produced, it is immediately cleaved or separated into distinct and discrete first and second polypeptides without the need for any external cleavage activity.
- TCR alpha variable domain therefore refers to the concatenation of TRAV and TRAJ regions
- TCR alpha constant domain refers to the extracellular TRAC region, or to a C-terminal truncated TRAC sequence.
- TCR beta variable domain refers to the concatenation of TRBV and TRBD/TRBJ regions
- TCR beta constant domain refers to the extracellular TRBC region, or to a C-terminal truncated TRBC sequence.
- duplexed double-stranded
- hybridized refers to nucleic acids formed by hybridization of two single strands of nucleic acids containing complementary sequences. In most cases, genomic DNA is double-stranded. Sequences can be fully complementary or partially complementary.
- a “vaccine” refers to a composition for generating immunity for the prophylaxis and/or treatment of diseases. Accordingly, vaccines are medicaments which comprise antigens and are intended to be used in humans or animals for generating specific defense and protective substances upon administration to the human or animal.
- Circular RNA used in polymer lipid nanoparticle (PLNP) compositions as described herein can be prepared and purified according to procedures in Wesselhoeft et al. (PCT/US2020/034418, filed May 22, 2020, published as WO2020237227), and Goodman et al. (PCT/US2021/031629, filed May 10, 2021, published as WO2021226597), the contents of which are hereby incorporated by reference in their entirety for all purposes.
- the subject polymer lipid nanoparticle composition are manufactured using an emulsion-based method. A general flow of the process is shown in FIG. 1 and FIG. 2.
- FIG. 4A shows RNA integrity data for both mRNA (mfLuc) and circRNA (ofLuc) after being processed through the high shear process. RNA degradation was observed for both mRNA and circRNA, however, circRNA demonstrated significantly less degradation compared to mRNA. This observation was certainly unexpected and unpredictable.
- PLNPs were prepared according to the procedure set out in Example 2 using 10K-5K molecular weighted polymer (P) (e.g., 10K weighted PLA and 5K weighted PEG) and ionizable lipid ethyl lauroyl arginate (ELA) at ELA:P ratios of 2: 1 , 4: 1 , and 6:1.
- P 10K-5K molecular weighted polymer
- ELA ionizable lipid ethyl lauroyl arginate
- the PLNPs were loaded with circRNAs comprising a 3’ exon fragment, an internal ribosome entry site (IRES), a coding region (e.g., firefly luciferase), and a 5’ exon fragment.
- IRS internal ribosome entry site
- coding region e.g., firefly luciferase
- FIG. 7 shows vitro release (IVR) data in PBS at 37°C for a PLNP of two different particle sizes made with a 10K-5K PLA-PEG. Both PLNPs demonstrated a burst release of approximately 20- 25% and complete RNA release over 5 to 6 days.
- Circular RNAs comprising a 3’ exon, an internal ribosome entry site (IRES), a coding region encoding firefly luciferase, and a 5’ exon, were formulated with delivery vehicle comprising either a PLNP (non-targeted), a PLNP comprising GalNAc3, an exemplary LNP comprising an ionizable lipid (positive control), and a diluent.
- the formulated circular RNA and delivery vehicle were dosed at 4 mpk into Balb/C mice intravenously. Mice were sacrificed at 6 hours, 1 day, 2 days, 5 days post intravenous injection of the circular RNAs. Protein expression was evaluated using Ex vivo IVIS. Liver tissue was characterized for circRNA distribution into hepatocytes and Kupffer cells (within the Liver tissue) using FISH. Mice body weights were collected throughout the study to assess tolerability of the nanoparticle formulations.
- FIG. 8 provides the resulting FISH imaging.
- the top row of images shows liver tissue samples characterized at 6 hours post IV injection using Fluorescence in situ hybridization (FISH) that were imaged at lx magnification. Only the yellow fluorescence channel (corresponding to circular RNA encoding firefly) had been turned on for FIG. 8.
- FISH Fluorescence in situ hybridization
- the diluent image negative control
- no firefly luciferase signal was seen.
- the ionizable LNP 1 image positive control
- the firefly signal was observed suggesting delivery of circular RNA encoding firefly luciferase to liver tissue.
- the firefly luciferase signal was present but more distributed towards the surface of the tissue.
- the firefly luciferase fluorescence was significantly more brighter and spread throughout the liver tissue, suggesting a much more effective and uniform delivery of the circular RNA encoding flue to the target tissue as compared to the PLNP (non-targeted) and LNP groups.
- the bottom row of images correspond to the same 6 hour Liver tissue samples characterized using FISH and being imaged at 20x magnification and had all of the fluorescence channels turned on. As marked in the Diluent image of FIG.
- the hepatocytes appeared red in color, and most Kupffer cells demonstrated yellow overlap suggesting the circular RNA encoding firefly luciferase distribution mainly to Kupffer cells, rather than hepatocyte.
- the hepatocytes appeared yellow-orange in color indicating that a large amount of the circular RNAs encoding firefly luciferase was delivered to hepatocytes.
- most Kupffer cells appeared completely green in color suggesting that GalNAc3 conjugated PLNPs demonstrated preferential uptake by hepatocyte rather than the Kupffer cells.
- the liver organs of the same Balb/C mice were harvested and analyzed for Flue expression using Ex vivo imaging (provided in FIG. 9).
- the ionizable LNP 1 treated mice demonstrated high protein expression at the 6 hour timepoint but the protein levels start to drop rapidly for subsequent timepoints.
- the PLNP GalNAc mice demonstrated lower (compared to the ionizable LNP 1 mice) but sustained expression of protein (for three consecutive timepoints) up to 2 days.
- the non-targeted PLNP (shown as PLNP in FIG. 9) demonstrated an inconsistent expression profile.
- mice body weights were also collected for the circular RNAs formulated with either PLNPs comprising GalNAc3, ionizable LNP 1 or 20% sucrose. Body weights shown in FIG. 10 demonstrated little to no change in mice body weights over the duration of the study, suggesting that all the dosing groups, including PLNP groups, were well tolerated.
- PLNPs were prepared according to the emulsion-based formulation procedure described in Example 2 using 10K-5K molecular weighted PLA-PEG copolymer (e.g., 10K weighted PLA and 5K weighted PEG, acquired from Advanced Polymer Materials, Inc.) and one of three different ionizable lipids.
- the three ionizable lipids used comprised either ethyl lauroyl arginate, ionizable lipid 2, and ionizable lipid 3.
- PLNPs comprising either ethyl lauroyl arginate and/or ionizable lipid 2 were formulated at a lipid to phosphate (RNA backbone) ratio of 3:1; and PLNPs comprising ionizable lipid 3 were formulated at a lipid to phosphate ratio of 4:1. All three PLNPs were loaded with circRNAs comprising a 3’ exon fragment, an internal ribosome entry site (IRES), a coding region (e.g., expressing firefly luciferase), and a 5’ exon fragment.
- IRS internal ribosome entry site
- the PLNPs were characterized throughout the formulation process using dynamic light scattering (DLS) and Quant-iTTM RiboGreenTM RNA encapsulation assay as shown in FIG. 16. DLS measurements were taken following the emulsion quench step (PQ), post-TFF purification (pTFF), and after an additional particle concentration step using AmiconTM centrifugal filter columns (PC). All three PLNPs showed similar and consistent DLS measurements throughout the process, with final particle size around 80 nm with PDI less than 0.1 (illustrated in FIG. 16).
- DLS dynamic light scattering
- PC AmiconTM centrifugal filter columns
- Encapsulation efficiency was determined using RiboGreenTM at the PC step, with encapsulation efficiency of 73%, 80%, and 66% for the ethyl lauroyl arginate, ionizable lipid 2, and ionizable lipid 3 PLNPs, respectively (also illustrated in FIG. 16).
- PLNPs were prepared according to the emulsion-based formulation procedure described in Example 2 using 10K-5K molecular weight PLA-PEG copolymer (e.g., 10K weighted PLA and 5K weighted PEG, acquired from Advanced Polymer Materials, inc.) and one of two different ionizable lipids.
- 10K-5K molecular weight PLA-PEG copolymer e.g., 10K weighted PLA and 5K weighted PEG, acquired from Advanced Polymer Materials, inc.
- PLNPs comprising ionizable lipids ethyl lauroyl arginate or ionizable lipid 2 were formulated at a lipid to phosphate (RNA backbone) ratio of 3: 1 and were loaded with circRNAs comprising a 3’ exon fragment, an internal ribosome entry site (IRES), a coding region (e.g., firefly luciferase), and a 5’ exon fragment.
- the two PLNP formulations had similar post-processing characterization as assessed by dynamic light scattering (DLS) and Quant-iTTM RiboGreenTM RNA encapsulation assay as shown in FIG. 17.
- the PLNPs comprising ethyl lauroyl arginate or ionizable lipid 2 had similar release curves, with about 20% circRNA being released in the first 6 hours of incubation followed by steady release up to 80-90% after 7 days (as depicted in FIG. 18).
- RNA encapsulated in PLNP was isolated and/or released from unencapsulated RNA in the surrounding solution using an ultracentrifugation method. Briefly, 2 mL of each PLNP or control sample was pipetted onto 10 mL of distilled water. The PLNP samples were loaded into a SW 41 Ti rotor in a Beckman Coulter floor and centrifuged at 288,000 g for 3 hours. The supernatant containing RNA in solution was aspirated and discarded. The pellet of PLNPs was reconstituted in distilled water and transferred to a microcentrifuge tube.
- RNA was extracted from the reconstituted pellet using the New England Biolabs Total RNA Miniprep Kit, following the protocol for RNA extraction from animal cells.
- the RNA control sample was not ultracentrifuged but was processed using the RNA extraction method.
- the concentration of the extracted RNA from all samples was measured using a NanoDrop Microvolume UV-Vis Spectrophotometer, diluted to 100 pg/mL with distilled water, and stored at 4 °C until analysis.
- FIGS. 19A, B, C provide exemplary chromatograms for the unformulated circRNA (depicted in FIG. 19A) and circRNA extracted from PLNPs comprising either ethyl lauroyl arginate or ionizable lipid 2 at 0 minutes of the IVR assay (depicted in FIGs. 19B and 19C respectively).
- RNA integrity was calculated as the area under the curve of the intact circular peak as a percentage of all RNA peaks.
- FIG. 20 provides the calculated RNA integrity of the circRNA remaining in the PLNPs comprising ethyl lauroyl arginate and ionizable lipid 2 at timepoints before and throughout the IVR assay, as compared with the RNA integrity of the unformulated RNA control over time.
- CircRNA comprising: a 3’ exon, an internal ribosome entry site (IRES), a coding region encoding firefly luciferase, and a 5’ exon, was formulated with a PLNP delivery vehicle comprising ethyl lauroyl arginate or ionizable lipid 2 as the ionizable lipid. All three PLNP formulations were appended with GalNAc3.
- circRNA PLNPs were dosed at 8 mpk and/or 4 mpk into Balb/C mice intravenously. Dosing was defined as mass of circRNA per kilogram of bodyweight. Mice were euthanized at either 6 hours, 2 days, and 5 days post-intravenous injection. Protein fLuc expression from the mice was evaluated using ex vivo IVIS as depicted FIG. 21.
- Liver tissue was analyzed for circRNA distribution in hepatocytes and Kupffer cells (e.g., within the liver tissue) using FISH. All formulations were well tolerated as determined from clinical observation of the mice using techniques available known in art.
- Fluorescent in situ hybridization is an effective imaging technique for visualizing RNA intracellular deposition and distribution with a tissue. Following harvest and imaging, the livers of the mice were preserved using 4% paraformaldehyde in phosphate buffered solution (PBS) and sent for FISH imaging. A probe for the circRNA coding for firefly luciferase was used to visualize the delivered RNA, while probes for albumin RNA, Adgrel RNA, and DNA were used to label the hepatocytes, Kupffer cells, and cell nuclei, respectively.
- FIG. 22A shows the mean fLuc RNA signal, calculated using ImageJ, across dosed groups at 6 hours post-injection.
- FIG. 22B shows exemplary FISH images for the two PLNPs 4 mpk groups at 6 hours and 5 days post-dose timepoints.
- the PLNP comprising ionizable lipid 2 showed noticeably more RNA signal remaining in the liver tissue compared to PLNPs comprising ethyl lauroyl arginate.
- PLNP in vitro firefly luciferase expression assay was used for PLNP to facilitate the evaluation of various formulation conditions.
- a targeted receptor-mediated uptake method e.g., using GalNac3
- GalNac3 interaction on PLNPs with ASGPR receptors on hepatocytes was used to facilitate and analyze intracellular uptake in vivo.
- PLNPs comprised ethyl lauroyl arginate.
- Three cell types i.e., 1C1C7 cells, primary human hepatocytes (PHH), and human skeletal muscle cells (HSKM) were assayed for endogenous ASGPR expression via western blot.
- circRNA encoding ASGPR was synthesized, and the same cell types were transfected with the circRNA using MessengerMaxTM, a commercially available membrane permeating reagent for either 6 hours or 24 hours. At 24 hours post-transfection, the cells were also assayed for ASGPR expression via western blot as illustrated in FIG. 23. Both 1C1C7S and primary human hepatocytes (PHH) showed endogenous ASGPR expression that was significantly enhanced by transfection of ASGPR circRNA. Human skeletal muscle cells were used as a negative control for endogenous expression. As expected, the human skeletal muscle cells did not show detectable levels of endogenous ASGPR expression and only showed expression upon transfection of ASGPR protein. PHH cells were further transfected with ASGPR circRNA as the basis for an in vitro expression assay.
- Luciferase expression was then measured 24 hours post-ASGPR circRNA transfection after PLNP transfection 6 hours post-ASGPR circRNA transfection as depicted in FIG. 26. Significant luciferase expression over unconjugated PLNP controls was also observed at this timepoint. In all studies, Promega Bright-GloTM Luciferase Assay System was used to evaluate luciferase expression. EXAMPLE 10
- PLNP formulatability, in vitro RNA release profile, and in vitro firefly luciferase expression in primary human hepatocytes were evaluated in PLNPs formulated with different polymer shell constitutions.
- PLNPs were prepared according to the emulsion-based formulation procedure described in Example 2 (e.g., using ethyl lauroyl arginate as the complex forming ionizable lipid at a lipid to phosphate ratio of 3:1 and circRNAs comprising a 3’ exon fragment, an internal ribosome entry site (IRES), an expression sequence encoding firefly luciferase, and a 5’ exon fragment).
- the PLNPs were developed with different polymer components.
- the control PLNP was formulated with a 10K-5K molecular weight PLA-PEG block copolymer.
- Shell 1 comprised a 4:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer.
- Shell 2 comprised a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer.
- Shell 3 was comprised entirely of a 15K-5K PLGA-PEG block copolymer (i.e., the PLGA portion comprised a random copolymer of 75% lactic acid and 25% glycolic acid).
- Shell 4 comprised a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K- 2K molecular weight PLA-PEG block copolymer, supplemented with 1% by mass 16K-5K PLA-PEG block copolymer.
- the PLNPs were characterized using dynamic light scattering (DLS) and Quant-iTTM RiboGreenTM RNA encapsulation assay (illustrated in FIG. 27).
- FIG. 28A shows the percent of encapsulated circRNA released from each PLNP formulation over the course of 5 days;
- FIG. 28B shows the first 24 hours post administration of FIG. 28A.
- the formulations of FIGs. 28A and 28B show a variety of release curves.
- three of the four alternative shell formulations had released more RNA than the control formulation, with Shell 2 showing near 100% RNA release.
- the alternative shell PLNP (i.e., Shells 1-4) formulations were also evaluated for luciferase expression in primary human hepatocytes, using methods described in Example 9. Two circRNA doses were assessed with expression timepoints at 24 and 48 hours post-ASGPR circRNA transfection (shown in FIGs. 29A and 29B). In general, the alternative PLNP shells affected the kinetics of luciferase expression. At 24 hours, three of the four alternative shell formulations showed significantly higher expression (p ⁇ 0.05) than the control formulations, consistent with the RNA release profile at 24 hours in the IVR assay (illustrated in FIG. 29A). At 48 hours, the control formulation shows higher expression than the alternative shell PLNPs (illustrated in FIG. 29B). EXAMPLE 11
- PLNPs were prepared according to the emulsion-based formulation procedure described in Example 2 using 10K-5K molecular weight PLA-PEG block copolymer and ethyl lauroyl arginate as the complex forming ionizable lipid at a lipid to phosphate ratio of 3:1.
- the PLNPs were loaded with circRNA comprising a 3’ exon fragment, an internal ribosome entry site (IRES), an expression sequence encoding firefly luciferase, and a 5’ exon fragment.
- IRS internal ribosome entry site
- the PLNPs were diluted in a lyophilization protectant buffer. Aliquots were lyophilized using a Labconco FreeZone benchtop lyophilizer. The aliquots were stored at either 4 °C or -80 °C and reconstituted at one-week intervals over 4 weeks. Characterization showed minimal changes in PLNP particle size distribution, encapsulation efficiency, and RNA integrity following lyophilization and storage for up to 4 weeks (illustrated in FIGs. 30A, 30B, and 30C respectively).
- PLNP formulatability, in vitro RNA release profile, and in vitro firefly luciferase expression in primary human hepatocytes were evaluated in PLNPs formulated with different polymer shell constitutions.
- PLNPs were prepared according to the emulsion-based formulation procedure described in Example 2 (e.g., using ethyl lauroyl arginate as the complex forming ionizable lipid at a lipid to phosphate ratio of 3:1 and circRNAs comprising a 3’ exon fragment, an internal ribosome entry site (IRES), an expression sequence encoding firefly luciferase, and a 5’ exon fragment).
- the PLNPs were developed with different polymer components.
- the control PLNP was formulated with a 10K-5K molecular weight PLA-PEG block copolymer.
- Shell 2 comprised a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer.
- Shell 3 was comprised entirely of a 15K-5K PLGA-PEG block copolymer (i.e., the PLGA portion comprised a random copolymer of 75% lactic acid and 25% glycolic acid).
- PLNPs comprised core forming lipids, ELA or ionizable lipid 2.
- the endosomal escape agents (e.g., doxepin or endosomal escape agent 1) were applied as separate solutions either concurrently with the circular RNA-PLNP or 1 or 2 hours afterwards (e.g., at a pre-dose, concurrent dose, or post-dose timepoint relative to PLNP dosing).
- doxepin hydrochloride concentration e.g., from Millipore- Sigma
- endosomal escape agent 1 For PLNPs formulated with endosomal escape agent 1, 7.17 pM of endosomal escape agent 1 solution was used either as a pre-dose 1 hour prior to dosing the circular RNA-PLNP solutions into cells, concurrently as a direct dose during circular RNA-PLNP dosing, or 1 or 2 hours as a post dose.Luciferase expression was then measured 48 hours post circRNA transfection as shown in FIG. 31. In all studies, Promega Bright-GloTM Luciferase Assay System was used to evaluate luciferase expression.
- PCT/US2019/035531 PCT/US2020/034418, PCT/US2020/063494, PCT/US2021/031629, PCT/US2021/023540, PCT/US2021/033276, PCT/US2022/033091, PCT/US2022/045408, PCT/US2022/049313,
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Veterinary Medicine (AREA)
- Pharmacology & Pharmacy (AREA)
- Epidemiology (AREA)
- Medicinal Chemistry (AREA)
- Animal Behavior & Ethology (AREA)
- General Health & Medical Sciences (AREA)
- Public Health (AREA)
- Engineering & Computer Science (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Physics & Mathematics (AREA)
- Dermatology (AREA)
- Biomedical Technology (AREA)
- Nanotechnology (AREA)
- Optics & Photonics (AREA)
- Biophysics (AREA)
- Molecular Biology (AREA)
- Dispersion Chemistry (AREA)
- Pharmaceuticals Containing Other Organic And Inorganic Compounds (AREA)
Abstract
Disclosed herein are compositions comprising polymeric lipid nanoparticles, and methods for preparing the same. The polymeric lipid nanoparticles can comprise RNA polynucleotides, such as circular RNA polynucleotides (circRNA), ionizable lipids (lipid) and amphiphilic polymers, thereby forming a polymeric lipid nanoparticle composition. In some embodiments, the lipid forms a complex with the circRNA (circRNA-lipid complex) that is encapsulated in the nanoparticles. In some embodiments, the circRNA-lipid complex is encapsulated in the core of the nanoparticles and the amphiphilic polymer provides a uniform shell around the circRNA-lipid complex. The present application also provides emulsions comprising the polymeric lipid nanoparticles.
Description
POLYMER LIPID NANOPARTICLE COMPOSITIONS FOR DELIVERING CIRCULAR POLYNUCLEOTIDES
FIELD
[0001] The present disclosure generally relates to polymeric nanoparticle compositions for encapsulating therapeutic agents, such as nucleic acids (e.g., circular polynucleotides), for timely release of the nucleic acid cargo.
BACKGROUND
[0002] In the past few decades, the field of nucleic acid therapeutics has rapidly expanded and has become the basis for treating a wide variety of diseases. Nucleic acid therapies available include, but are not limited to, the use of DNA or viral vectors for insertion of desired genetic information into the host cell, and/or RNA constructed to encode for a therapeutic protein. DNA and viral vector deliveries carry their own setbacks and challenges that make them less favorable to RNA therapeutics. For example, the introduced DNA in some cases may be unintentionally inserted into an intact gene and result in a mutation that impedes or even wholly eliminates the function of the endogenous gene leading to an elimination or deleteriously reduced production of an essential enzyme or interruption of a gene critical for the regulating cell growth. Viral vector-based therapies can result in an adverse immune response. Compared to DNA or viral vectors, RNA is substantially safer and more effective gene therapy agent due to its ability to encode for the protein outside of the nucleus to perform its function. With this, the RNA does not involve the risk of being stably integrated into the genome of the transfected cell.
[0003] The field of RNA therapeutics conventionally has consisted of engineering linear messenger RNAs (mRNA). Although more effective than DNA or viral vectors, linear mRNAs have their own set of challenges regarding the stability, immunogenicity, translation efficiency, and delivery. Some of these challenges may lead to size restraints and/or destruction of the linear mRNA due to the challenges present with linear mRNAs’ caps. Partly to overcome these limitations, circular polynucleotides or circular RNAs are increasingly being studied. Due to being covalently closed continuous loops, circular RNAs are useful in the design and production of stable forms of RNA. The circularization of an RNA molecule provides an advantage to the study of RNA structure and function, especially in the case of molecules that are prone to folding in an inactive conformation (Wang and Ruffner, 1998). Circular RNA can also be particularly interesting and useful for in vivo applications, especially in the research area of RNA-based control of gene expression and therapeutics, including protein replacement therapy and vaccination.
[0004] Further, to promote effective delivery of these RNA polynucleotides, polymer lipid nanoparticle delivery systems can be used. This disclosure provides polymeric lipid nanoparticle compositions to encapsule engineered polynucleotides to allow timely release of the polynucleotide cargo for therapeutic purposes.
SUMMARY
[0005] The present application provides, compositions comprising polymeric lipid nanoparticles, and methods for preparing the same. The polymeric lipid nanoparticles can comprise RNA polynucleotides, such as circular RNA polynucleotides (aka circRNA or oRNA™), ionizable lipids (lipid) and amphiphilic polymers, thereby forming a polymeric lipid nanoparticle composition. In some embodiments, the lipid forms a complex with the circRNA (circRNA-lipid complex) that is encapsulated in the nanoparticles. In some embodiments, the circRNA-lipid complex is encapsulated in the core of the nanoparticles and the amphiphilic polymer provides a uniform shell around the circRNA-lipid complex. The present application also provides emulsions comprising circRNA, ionizable lipid (lipid) and an amphiphilic polymer useful for preparing the polymeric lipid nanoparticle compositions. Methods of preparing the subject emulsions and compositions are also provided.
[0006] In one aspect, provided herein is a composition comprising a plurality of polymeric lipid nanoparticles, each nanoparticle comprising: a RNA polynucleotide, such as a circular RNA polynucleotide; an ionizable lipid; and an amphiphilic polymer.
[0007] In another aspect, provided herein is an emulsion comprising an aqueous continuous phase and an organic dispersed phase, wherein the organic dispersed phase comprises droplets that contain: a RNA polynucleotide, such as a circular RNA polynucleotide; an ionizable lipid; and an amphiphilic polymer.
[0008] In another aspect, there is provided a method of preparing a composition comprising a plurality of polymeric lipid nanoparticles, the method comprising: a) providing a mixture comprising: an ionizable lipid; and an amphiphilic polymer, both dissolved in an organic phase; b) combining the mixture with a RNA polynucleotide, such as a circular RNA polynucleotide, dissolved in an aqueous phase to obtain an emulsion; and
c) adding the emulsion to an aqueous bath to form polymeric lipid nanoparticles comprising the ionizable lipid, the amphiphilic polymer, and the RNA polynucleotide.
BRIEF DESCRIPTION OF THE DRAWINGS
[0009] FIG. 1 is a schematic showing a general flow of a process for the manufacture of subject polymer lipid nanoparticle compositions using an emulsion-based method.
[0010] FIG. 2 is a schematic illustrating the quench step of an emulsion-based method for the manufacture of subject polymer lipid nanoparticle compositions (i.e., adding a subject emulsion to an aqueous bath).
[0011] FIG. 3 is a graphic representation of the core shell of the polymeric lipid nanoparticle containing the circRNA-CFL pair at the core of the nanoparticle and a polymer shell.
[0012] FIG. 4A illustrates RNA integrity data for both, mRNA (mfLuc) and circRNA (ofLuc) after being processed through the high shear process.
[0013] FIG.4B illustrates RNA integrity data for a complex of circRNA (ofLuc) and an ionizable lipid (ethyl lauroyl arginate (EL A)) (ofLuc: EL A) at various ratios.
[0014] FIG. 5A is a graph illustrating the particle size distribution (Z-average, nm) and polydispersity index (PDI) for various polymer lipid nanoparticle compositions prepared using the emulsion-based process of Example 2 with various ratios of ionizable lipid (e.g., ELA) to amphiphilic polymer (P) (ELAT of 2:1, 4:1, and 6:1).
[0015] FIG. 5B is a graph illustrating the encapsulation efficiency (EE%) for various polymer lipid nanoparticle compositions prepared using the emulsion-based process of Example 2 with various ratios of ionizable lipid (e.g., ELA) to amphiphilic polymer (P) (ELA:P of 2:1, 4:1, and 6:1).
[0016] FIG. 6A is a cryogenic transmission electron microscopy (Cryo-TEM) image of a polymer lipid nanoparticle composition prepared using the emulsion-based process of Example 2 with an ELA : amphiphilic polymer ratio (ELA:P) of 4:1, including the core-shell morphology.
[0017] FIG. 6B is a Cryo-TEM image of a composition of ELA and circRNA complex without the inclusion of an amphiphilic polymer.
[0018] FIG. 7 is a graph showing the in vitro release kinetics of the circRNA from polymer lipid nanoparticles performed at 37 °C in PBS over a span of 0 to 6 days. The polymer lipid nanoparticle composition is made with a 10k-5k polylactic acid-polyethylene glycol polymer (i.e., di-block polymer composed of 10k PLA and 5k PEG) and ELA as the ionizable lipid.
[0019] FIG. 8 depicts liver tissue of mice samples characterized using fluorescence in situ hybridization (FISH) being imaged at 6 hours at lx magnification (top) or at 20x magnification with fluorescent channels turned on (bottom) for mice dosed intravenously with either a PLNP (nontargeted), PLNP GalNAc3, an LNP comprising an ionizable lipid (positive control) (indicated as “Ionizable LNP 1”), or a diluent (negative control) and circular RNA.
[0020] FIG. 9 depicts firefly luciferase (Flue) expression using ex vivo imaging of the liver of mice over the span of 5 days post intravenous injection of either a PLNP (non-targeted), PLNP GalNAc3, an LNP comprising an ionizable lipid (positive control) (indicated as “Ionizable LNP 1”), or a diluent (negative control) (indicated as “control”) and circular RNA encoding Flue.
[0021] FIG. 10 depicts body weight of the mice injected intravenously with either PLNP GalNAc3, an LNP comprising an ionizable lipid (positive control) (indicated as “Ionizable LNP 1”), or a 20% sucrose (negative control) (indicated as “control”) and circular RNA encoding Flue.
[0022] FIG. 11 is a graphic representation of the glutathione mediated disruption of a polymeric lipid nanoparticle comprising a shell of amphiphilic polymers having disulfide linkages.
[0023] FIG. 12 is a graphic representation of the glutathione mediated disruption of a polymeric lipid nanoparticle comprising a shell of cross-linked amphiphilic polymers having disulfide linkages.
[0024] FIG. 13 is a graphic representation of the acidic pH mediated disruption of a polymeric lipid nanoparticle comprising a shell of cross-linked amphiphilic polymers having hydrazone linkages.
[0025] FIG. 14 is a graphic representation of polymeric lipid nanoparticles prepared using tri-block amphiphilic polymers.
[0026] FIG. 15 is a graphic representation of polymeric lipid nanoparticles prepared using amphiphilic polymers conjugated to an ionizable lipid (an amphiphilic polymer-cation complex).
[0027] FIG. 16 depicts particle size distribution (Z-average, nm), polydispersity index (PDI), encapsulation efficiencies (in %) for PLNPs comprising an ionizable lipid (i.e., ethyl lauroyl arginate (ELA), ionizable lipid 2, or ionizable lipid 3) at either post-quench, post-TFF purification, or postparticle concentration step (indicated as “PQ,” “pTFF,” and “PC” respectively).
[0028] FIG. 17 depicts article size distribution (Z-average, nm), polydispersity index (PDI), encapsulation efficiencies (in %) for PLNPs comprising an ionizable lipid (i.e., ethyl lauroyl arginate or ionizable lipid 2).
[0029] FIG. 18 is a circular RNA release curve providing the percentage of circular RNA released in vitro over the span of 8 days for PLNPs comprising either ionizable lipid ethyl lauroyl arginate or ionizable lipid 2.
[0030] FIG. 19A is an ion pair reverse phase (IPRP) high performance liquid chromatograph (HPLC) showing circular RNA integrity of circular RNAs encoding firefly luciferase formulated in a sodium phosphate buffer solution. Intact circular RNAs are labeled as “circular.” Non-intact circular RNA products are labeled as “nicked.”
[0031] FIG. 19B is an ion pair reverse phase (IPRP) high performance liquid chromatograph (HPLC) showing circular RNA integrity of circular RNAs encoding firefly luciferase extracted from PLNP comprising an ionizable lipid, ethyl lauroyl arginate. Intact circular RNAs are labeled as “circular.” Non-intact circular RNA products are labeled as “nicked.”
[0032] FIG. 19C is an ion pair reverse phase (IPRP) high performance liquid chromatograph (HPLC) showing circular RNA integrity of circular RNAs encoding firefly luciferase extracted from PLNP comprising an ionizable lipid (ionizable lipid 2). Intact circular RNAs are labeled as “circular.” Nonintact circular RNA products are labeled as “nicked.”
[0033] FIG. 20 depicts RNA integrity over the span of 7 days of a circular RNA formulated in a sodium phosphate buffer solution (control, “unformulated circRNA”) or a PLNP comprising either ionizable lipid ethyl lauroyl arginate or ionizable lipid 2. RNA integrity was calculated from the intact circular RNA peak area under the curve (AUC) percentage present in the IPRP-HPLC chromatographs depicted in FIGs. 19A-19C. The dotted line in FIG. 20 indicates the RNA integrity of the circRNA prior to formulation.
[0034] FIG. 21 depicts total ex vivo liver flux in Balb/C mice dosed with circular RNA formulated in a 20% sucrose solution or PLNP comprising either an ionizable lipid ethyl lauroyl arginate, ionizable lipid 2, or ionizable lipid 3 at dose of 4 mpk.
[0035] FIG. 22A depicts liver tissue of mice sample characterized using fluorescence in situ hybridization (FISH) being imaged at 6 hours (left) or 120 hours (right) at 20x magnification with fluorescent channels turned on (bottom) for mice dosed intravenously with either a PLNP comprising either ionizable lipid 2 (top) or ethyl lauroyl arginate (bottom) formulated with circular RNA.
[0036] FIG. 22B is a graph illustrating mean fluorescence signal calculated from FISH imaging of FIG. 22A for circular RNA-PLNP solutions (e.g., circular RNAs formulated with PLNP comprising ethyl lauroyl arginate or ionizable lipid 2 and dosed at 4 mpk or 8mpk) or circular RNA formulated with 20% sucrose solutions (control) at 6 hours post injection.
[0037] FIG. 23 is a western blot image of asialoglycoprotein receptor 1 (ASGPR1) expression in Hepa-lClC7, primary human hepatocytes (PHHs), and human skeletal muscle cells (HSKMs) 24 hours post transfection of a circular RNA encoding ASGPR1 (indicated as “1C1C7 Treated,” “PHH Treated,”
and “HSKM Treated,” respectively). Endogenous ASGPR1 levels (control) for each of the cell types are indicated as “1C1C7,” “PHH,” and “HSKM.”
[0038] FIG. 24 illustrates in vitro luciferase expression in primary human hepatocytes post treatment with: (1) 50 ng circular RNA encoding ASGPR (indicated as “oASGPR”); and/ or (2) PLNP unconjugated GalNAc3 (indicated as “Unconj.”), PLNP conjugated GalNAc3 (indicated as “GNAc”), or no PLNP transfer vehicle (indicated as “control”). PLNPs were formulated with Ip of circular RNA encoding firefly luciferase (fLuc). fLuc expression was determined 48 hours post treatment of circular RNA encoding fLuc. Background signaling is shown as a dotted line.
[0039] FIG. 25 illustrates in vitro luciferase expression in primary human hepatocytes treated with: (1) 50 ng of circular RNA encoding ASGPR (indicated as “oASGPR”); and/ or (2) PLNP unconjugated with GalNAc3 (indicated as “Unconj.”), PLNP conjugated with GalNAc3 (indicated as “GNAc”), or no PLNP transfer vehicle (indicated as “control”). PLNPs unconjugated or conjugated with GalNAc3 were formulated with either 0.5 pg, 1 pg or 1.5 p of circular RNA expressing firefly luciferase (fLuc). fLuc expression was determined 48 hours post treatment of circular RNA encoding fLuc. PHH cells were treated with oASGPR occurred for 6 hours (indicated as “6h txn”) or 24 hours (indicated as “24h txn”). Background signaling is shown as a dotted line.
[0040] FIG. 26 illustrates in vitro luciferase expression in primary human hepatocytes treated with: (1) 50 ng of circular RNA encoding ASGPR (indicated as “oASGPR”); and/ or (2) PLNP unconjugated with GalNAc3 (indicated as “Unconj.”), PLNP conjugated with GalNAc3 (indicated as “GNAc”), or no PLNP transfer vehicle (indicated as “control”). PLNPs unconjugated or conjugated with GalNAc3 were formulated with 0.5 pg of circular RNA expressing firefly luciferase (fLuc). fLuc expression was determined 24 hours post treatment of circular RNA encoding fLuc. Background signaling is shown as a dotted line.
[0041] FIG. 27 depicts PLNP Z-average (in nm), polydispersity index (indicated as “PDI”) and encapsulation efficiency (indicated as a percentage) of PLNPs having alterative polymer shell compositions (i.e., Shell 1, Shell 2, Shell 3, and/or Shell 4) formulated with circular RNAs encoding firefly luciferase. Shell 1 comprised a 4:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer. Shell 2 comprised a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer. Shell 3 was comprised entirely of a 15K-5K PLGA-PEG block copolymer (the PLGA portion being a random copolymer of 75% lactic acid and 25% glycolic acid). Shell 4 comprised a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer, supplemented with 1% by mass 16K-5K PLA-PEG block copolymer. Control PLNP comprises ethyl lauroyl arginate (indicated as “control”).
[0042] FIG. 28A depicts luciferase expression in primary human hepatocytes treated with GalNAc3- appended PLNP formulated with circular RNA encoding firefly luciferase. PLNPs were formulated with 4 different polymer shell compositions (i.e., Shell 1, Shell 2, Shell 3, and Shell 4) at 24 hours post- ASGPR circular RNA administration. Control PLNP comprises ethyl lauroyl arginate (indicated as “control”).
[0043] FIG. 28B depicts luciferase expression in primary human hepatocytes treated with GalNAc3- appended PLNP formulated with circular RNA encoding firefly luciferase. PLNPs were formulated with 4 different polymer shell compositions (i.e., Shell 1, Shell 2, Shell 3, and Shell 4) at 48 hours post- ASGPR circular RNA administration. Control PLNP comprises ethyl lauroyl arginate (indicated as “control”).
[0044] FIG. 29A is an RNA release curve showing circular RNA released in vitro expressed as a percentage of encapsulated RNA released over the span of 5 days for PLNPs comprising either Shell 1 , Shell 2, Shell 3, or Shell 4 (wherein Shells 1-4 are the same as provided in FIG. 27) and formulated with either 250 ng or 500 ng of circular RNA expressing firefly luciferase (fLuc). Control PLNP comprises ethyl lauroyl arginate (indicated as “control”).
[0045] FIG. 29B is an RNA release curve depicting the firefly luciferase expression 48 hours after administration of the PLNP comprising formulated circular RNAs and/or controls present in FIG. 28A.
[0046] FIG. 30A illustrates the particle size distribution of reconstituted lyophilized PLNP aliquots comprising circular RNA stored at 4 °C and 20 °C over 4 weeks.
[0047] FIG. 30B illustrates the encapsulation efficiency of reconstituted lyophilized PLNP aliquots comprising circular RNA stored at 4 °C and 20 °C over 4 weeks.
[0048] FIG. 30C illustrates the circular RNA integrity of reconstituted lyophilized PLNP aliquots comprising circular RNA stored at 4 °C and 20 °C over 3 weeks.
[0049] FIG. 31 illustrates firefly luciferase expression 48 hours after transfection of PLNPs comprising a core forming lipid, wherein the core forming lipid comprised of either ELA or ionizable lipid 1 and treated with doxepin (indicated as “Dox”) or endosomal escape agent 1 into primary human hepatocytes (PHH) cells. Poly (lactic-co-glycolic acid) (PLGA) nanoparticles were used as a control. As depicted in FIG. 31, “1:1” indicates PLNPs comprising a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer.
DETAILED DESCRIPTION
[0050] The present application provides, among other things, compositions comprising polymeric lipid nanoparticles, and methods for preparing the same. The polymeric lipid nanoparticles can comprise
RNA polynucleotides, particularly circular RNA polynucleotides (aka circRNA or oRNA™), ionizable lipids (lipid) and amphiphilic polymers, thereby forming a polymeric lipid nanoparticle composition. In some embodiments, the lipid forms a complex with the circRNA (circRNA-lipid complex) that is encapsulated in the nanoparticles. In some embodiments, the circRNA-lipid complex is encapsulated in the core of the nanoparticles and the amphiphilic polymer provides a uniform shell around the circRNA-lipid complex.
[0051] Also disclosed herein is RNA therapy, along with associated compositions and methods. In some embodiments, the RNA therapy allows for increased RNA stability, expression, and prolonged half-life, among other things.
[0052] In some embodiments, provided herein are methods comprising administration of circular RNA polynucleotides provided herein into cells for therapy or production of useful proteins. In some embodiments, the method is advantageous in providing the production of a desired polypeptide inside eukaryotic cells with a longer half-life than linear RNA, due to the resistance of the circular RNA to ribonucleases.
[0053] Circular RNA polynucleotides lack the free ends necessary for exonuclease-mediated degradation, causing them to be resistant to several mechanisms of RNA degradation and granting extended half-lives when compared to an equivalent linear RNA. Circularization may allow for the stabilization of RNA polynucleotides that generally suffer from short half-lives and may improve the overall efficacy of exogenous mRNA in a variety of applications. In an embodiment, the functional half-life of the circular RNA polynucleotides provided herein in eukaryotic cells (e.g., mammalian cells, such as human cells) as assessed by protein synthesis is at least 20 hours (e.g., at least 40, 60, or 80 hours).
[0054] Various aspects of the disclosure are described in detail in the following sections. The use of sections is not meant to limit the disclosure. Each section can apply to any aspect of the disclosure. In this application, the use of “or” means “and/or” unless stated otherwise.
1. POLYMERIC LIPID NANOPARTICLE COMPOSITIONS
[0055] As summarized herein, the present disclosure provides polymeric lipid nanoparticle compositions, including a RNA polynucleotide, particularly a circular RNA polynucleotide, an ionizable lipid, and an amphiphilic polymer.
[0056] In certain embodiments, as described herein, the polymeric lipid nanoparticle composition is composed of an oily core surrounded by a polymeric shell. In some embodiments, the circular RNA is contained within the core and the polymeric shell controls the release of the circular RNA. In some
embodiments, the circular RNA forms a complex with the ionizable lipid (circRNA-lipid complex). In some embodiments, the circRNA-lipid complex is encapsulated in the core of the nanoparticles and the amphiphilic polymer provides a uniform shell around the circRNA-lipid complex.
[0057] By “uniform shell” it is meant that the shell can be substantially uniform in thickness around the entire outer surface of the polymeric lipid nanoparticle. For instance, the thickness of the shell can be substantially uniform in thickness by not varying in thickness by more than 20% or more, such as by not varying more than 10% in overall thickness around the entire circumference of the shell.
[0058] In some embodiments, the ionizable lipid is configured to be conjugated to the amphiphilic polymer.
[0059] In some embodiments, the molar ratio of ionizable lipid to amphiphilic polymer is from 2:1 to 6:1. In some embodiments, the molar ratio of ionizable lipid to amphiphilic polymer is 2:1, 3:1, 4:1, 5 : 1 , or 6: 1. In some embodiments, the molar ratio of ionizable lipid to amphiphilic polymer is from 2: 1 to 5:1, such as 2:1 to 4:1, 2:1 to 3:1, 3:1 to 4:1, 4:1 to 5:1, or 5:1 to 6:1. In some embodiments, the molar ratio of ionizable lipid to amphiphilic polymer is 2:1, 2.2:1, 2.5:1, 3:1, 3.2:1, 3.5:1, 4:1, 4.2:1, 4.5:1, 5:1, 5.2:1, 5.5:1, or 6:1. In some embodiments, the molar ratio of ionizable lipid to amphiphilic polymer is 3.5:1 to 4.5:1. In some embodiments, the molar ratio of ionizable lipid to amphiphilic polymer is 3.5:1, 3.6:1, 3.7:1, 3.8:1, 3.9:1, 4:1, 4.1:1, 4.2:1, 4.3:1, 4.4:1, or 4.5:1.
[0060] In some embodiments of the polymeric lipid nanoparticle composition, the nanoparticle mean diameter is from 50 to 200 nm, such as 50 to 180 nm, 50 to 150 nm, 50 to 120 nm, 50 to 100 nm, 50 to 80 nm, 80 to 200 nm, 80 to 180 nm, 80 to 150 nm, 80 to 120 nm, or 80 to 100 nm.
[0061] In one embodiment of the polymeric lipid nanoparticle composition, the nanoparticles may have a diameter from 10 to 100 nm such as, but not limited to, 10 to 20 nm, 10 to 30 nm, 10 to 40 nm, 10 to 50 nm, 10 to 60 nm, 10 to 70 nm, 10 to 80 nm, 10 to 90 nm, 20 to 30 nm, 20 to 40 nm, 20 to 50 nm, 20 to 60 nm, 20 to a 70 nm, 20 to 80 nm, 20 to 90 nm, 20 to 100 nm, 30 to 40 nm, 30 to 50 nm, 30 to 60 nm, 30 to 70 nm, 30 to 80 nm, 30 to 90 nm, 30 to 100 nm, 40 to 50 nm, 40 to 60 nm, 40 to 70 nm, 40 to 80 nm, 40 to 90 nm, 40 to 100 nm, 50 to 60 nm, 50 to 70 nm, 50 to 80 nm, 50 to 90 nm, 50 to 100 nm, 60 to 70 nm, 60 to 80 nm, 60 to 90 nm, 60 to 100 nm, 70 to 80 nm, 70 to 90 nm, 70 to 100 nm, 80 to 90 nm, 80 to 100 nm and/or 90 to 100 nm. In one embodiment, the lipid nanoparticles may have a diameter from 10 to 500 nm. In one embodiment, the lipid nanoparticle may have a diameter greater than 100 nm, greater than 150 nm, greater than 200 nm, greater than 250 nm, greater than 300 nm, greater than 350 nm, greater than 400 nm, greater than 450 nm, greater than 500 nm, greater than 550 nm, greater than 600 nm, greater than 650 nm, greater than 700 nm, greater than 750 nm, greater than
800 rnn, greater than 850 nm, greater than 900 nm, greater than 950 nm or greater than 1000 nm. Each possibility represents a separate embodiment of the present disclosure.
[0062] In some embodiments, a nanoparticle (e.g., a polymeric lipid nanoparticle) has a mean diameter of 10-500 nm, 20-400 nm, 30-300 nm, or 40-200 nm. In some embodiments, a nanoparticle (e.g., a lipid nanoparticle) has a mean diameter of 50-150 nm, 50-200 nm, 80-100 nm, or 80-200 nm.
[0063] In some embodiments, the polymeric lipid nanoparticles described herein can have a diameter from below 0.1 pm to up to 1 mm such as, but not limited to, less than 0.1 pm, less than 1.0 pm, less than 5 pm, less than 10 pm, less than 15 pm, less than 20 pm, less than 25 pm, less than 30 pm, less than 35 pm, less than 40 pm, less than 50 pm, less than 55 pm, less than 60 pm, less than 65 pm, less than 70 pm, less than 75 pm, less than 80 pm, less than 85 pm, less than 90 pm, less than 95 pm, less than 100 pm, less than 125 pm, less than 150 pm, less than 175 pm, less than 200 pm, less than 225 pm, less than 250 pm, less than 275 pm, less than 300 pm, less than 325 pm, less than 350 pm, less than 375 pm, less than 400 pm, less than 425 pm, less than 450 pm, less than 475 pm, less than 500 pm, less than 525 pm, less than 550 pm, less than 575 pm, less than 600 pm, less than 625 pm, less than 650 pm, less than 675 pm, less than 700 pm, less than 725 pm, less than 750 pm, less than 775 pm, less than 800 pm, less than 825 pm, less than 850 pm, less than 875 pm, less than 900 pm, less than 925 pm, less than 950 pm, less than 975 pm.
[0064] In some embodiments, a nanoparticle (e.g., a polymeric lipid nanoparticle) may have a diameter from 1 nm to 100 nm, from 1 nm to 10 nm, 1 nm to 20 nm, from 1 nm to 30 nm, from 1 nm to 40 nm, from 1 nm to 50 nm, from 1 nm to 60 nm, from 1 nm to 70 nm, from 1 nm to 80 nm, from 1 nm to 90 nm, from 5 nm to from 100 nm, from 5 nm to 10 nm, 5 nm to 20 nm, from 5 nm to 30 nm, from 5 nm to 40 nm, from 5 nm to 50 nm, from 5 nm to 60 nm, from 5 nm to 70 nm, from 5 nm to 80 nm, from 5 nm to 90 nm, 10 to 50 nm, from 20 to 50 nm, from 30 to 50 nm, from 40 to 50 nm, from 20 to 60 nm, from 30 to 60 nm, from 40 to 60 nm, from 20 to 70 nm, from 30 to 70 nm, from 40 to 70 nm, from 50 to 70 nm, from 60 to 70 nm, from 20 to 80 nm, from 30 to 80 nm, from 40 to 80 nm, from 50 to 80 nm, from 60 to 80 nm, from 20 to 90 nm, from 30 to 90 nm, from 40 to 90 nm, from 50 to 90 nm, from 60 to 90 nm and/or from 70 to 90 nm. Each possibility represents a separate embodiment of the present disclosure.
[0065] A polymeric lipid nanoparticle composition may be relatively homogenous. A poly dispersity index (PDI) may be used to indicate the homogeneity of a polymeric lipid nanoparticle composition, e.g., the particle size distribution of the nanoparticle compositions. A small (e.g., less than 0.3) polydispersity index generally indicates a narrow particle size distribution. In some embodiments, the polydispersity index of the polymeric lipid nanoparticles is 0.3 or less. A nanoparticle composition may have a polydispersity index from 0 to 0.3, such as 0.01, 0.02, 0.03, 0.04, 0.05, 0.06, 0.07, 0.08,
0.09, 0.10, 0.1 1, 0.12, 0.13, 0.14, 0.15, 0.16, 0.17, 0.18, 0.19, 0.20, 0.21, 0.22, 0.23, 0.24, 0.25, 0.26, 0.27, 0.28, 0.29, or 0.30, or any ranges created by these. In some embodiments, the polydispersity index of a nanoparticle composition may be from 0.10 to 0.20. In some embodiments, the polydispersity index of the nanoparticle composition is 0.10, 0.11, 0.12, 0.13, 0.14, or 0.15. In some embodiments, the polydispersity index of the nanoparticle composition is 0.16, 0.17, 0.18, 0.19, or 0.20. In some embodiments, the polydispersity index of the nanoparticle composition is 0.21, 0.22, 0.23, 0.24 or 0.25. In some embodiments, the polydispersity index of the nanoparticle composition is 0.26, 0.27, 0.28, 0.29 or 0.30. Each possibility represents a separate embodiment of the present disclosure.
[0066] The zeta potential of a nanoparticle composition may be used to indicate the electrokinetic potential of the composition. For example, the zeta potential may describe the surface charge of a nanoparticle composition. Nanoparticle compositions with relatively low charges, positive or negative, are generally desirable, as more highly charged species may interact undesirably with cells, tissues, and other elements in the body. In some embodiments, the zeta potential of a polymeric lipid nanoparticle composition may be from -20 mV to +20 mV, from -20 mV to +15 mV, from -20 mV to +10 mV, from -20 mV to +5 mV, from -20 mV to 0 mV, from -20 mV to -5 mV, from -20 mV to -10 mV, from -20 mV to -15 mV from -20 mV to +20 mV, from -20 mV to +15 mV, from -20 mV to +10 mV, from -20 mV to +5 mV, from -20 mV to 0 mV, from 0 mV to +20 mV, from 0 mV to +15 mV, from 0 mV to +10 mV, from 0 mV to +5 mV, from +5 mV to +20 mV, from +5 mV to +15 mV, or from +5 mV to +10 mV. Each possibility represents a separate embodiment of the present disclosure.
[0067] The efficiency of encapsulation of a therapeutic agent describes the amount of therapeutic agent that is encapsulated or otherwise associated with a polymeric lipid nanoparticle composition after preparation, relative to the initial amount provided. The encapsulation efficiency is desirably high (e.g., close to 100%). The encapsulation efficiency may be measured, for example, by comparing the amount of therapeutic agent in a solution containing the polymeric lipid nanoparticle composition before and after breaking up the nanoparticle composition with one or more organic solvents or detergents. Fluorescence may be used to measure the amount of free therapeutic agent (e.g., nucleic acids) in a solution. For the nanoparticle compositions described herein, the encapsulation efficiency of a therapeutic agent may be at least 50%, for example 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100%. In some embodiments, the encapsulation efficiency may be at least 70%. In some embodiments, the encapsulation efficiency may be at least 80%. In certain embodiments, the encapsulation efficiency may be at least 90%. Each possibility represents a separate embodiment of the present disclosure.
[0068] In some embodiments, the polymeric lipid nanoparticle has a polydispersity index of less than 0.3. In some embodiments, the polymeric lipid nanoparticle has a net neutral charge at a neutral pH. In some embodiments, the polymeric lipid nanoparticle has a mean diameter of 50-200 nm.
[0069] In some embodiments, the polymeric lipid nanoparticle has a polydispersity index of less than 0.3, a mean diameter of 50-200 nm, and a net neutral charge at a neutral pH. In some embodiments, the polymeric lipid nanoparticle has a polydispersity index of less than 0.2, a mean diameter of 50-200 nm, and a net neutral charge at a neutral pH. In some embodiments, the polymeric lipid nanoparticle has a polydispersity index of less than 0.1 , a mean diameter of 50-200 nm, and a net neutral charge at a neutral pH. In some embodiments, the polymeric lipid nanoparticle has a polydispersity index of less than 0.3, a mean diameter of 50-100 nm, and a net neutral charge at a neutral pH. In some embodiments, the polymeric lipid nanoparticle has a poly dispersity index of less than 0.2, a mean diameter of 50-100 nm, and a net neutral charge at a neutral pH. In some embodiments, the polymeric lipid nanoparticle has a poly dispersity index of less than 0.1, a mean diameter of 50-100 nm, and a net neutral charge at a neutral pH.
[0070] The properties of a subject polymeric lipid nanoparticle formulation may be influenced by factors including, but not limited to, the selection of the ionizable lipid component, the selection of the amphiphilic polymer, the ratio of the ionizable lipid to amphiphilic polymer components and biophysical parameters such as size.
A. POLYNUCLEOTIDES
[0071] In certain embodiments, the polymeric lipid nanoparticle compositions described herein comprise a polynucleotide. In some embodiments, the polynucleotide is DNA. In some embodiments, the polynucleotide is RNA. In some embodiments, the polynucleotide is linear RNA. In certain embodiments, the polynucleotide is circular RNA.
[0072] Transcription of a DNA template (e.g., comprising a 3’ intron element, 3’ exon element, a core functional element, a 5’ exon element, and a 5’ intron element) results in formation of a precursor linear RNA polynucleotide capable of circularizing. In some embodiments, this DNA template comprises a vector, PCR product, plasmid, minicircle DNA, cosmid, artificial chromosome, complementary DNA (cDNA), extrachromosomal DNA (ecDNA), or a fragment therein. In certain embodiments, the minicircle DNA may be linearized or non-linearized. In certain embodiments, the plasmid may be linearized or non-linearized. In some embodiments, the DNA template may be single-stranded. In other embodiments, the DNA template may be double-stranded. In some embodiments, the DNA template comprises in whole or in part from a viral, bacterial, or eukaryotic vector.
[0073] The present disclosure, as provided herein, comprises a DNA template that shares the same sequence as the precursor linear RNA polynucleotide prior to splicing of the precursor linear RNA polynucleotide (e.g., a 3’ intron element, a 3’ exon element, a core functional element, and a 5’ exon element, a 5’ intron element). In some embodiments, said linear precursor RNA polynucleotide undergoes splicing leading to the removal of the 3’ intron element and 5’ intron element during the process of circularization. In some embodiments, the resulting circular RNA polynucleotide lacks a 3’ intron fragment and a 5’ intron fragment, but maintains a 3’ exon fragment, a core functional element, and a 5’ exon element.
[0074] In some embodiments, the precursor linear RNA polynucleotide circularizes when incubated in the presence of one or more guanosine nucleotides or nucleoside (e.g., GTP) and a divalent cation (e.g., Mg2+). In some embodiments, the 3’ exon element, 5’ exon element, and/or core functional element in whole or in part promotes the circularization of the precursor linear RNA polynucleotide to form the circular RNA polynucleotide provided herein.
[0075] In certain embodiments circular RNA provided herein is produced inside a cell. In some embodiments, precursor RNA is transcribed using a DNA template (e.g., in some embodiments, using a vector provided herein) in the cytoplasm by a bacteriophage RNA polymerase, or in the nucleus by host RNA polymerase II and then circularized.
[0076] In certain embodiments, the circular RNA provided herein is injected into an animal (e.g., a human), such that a polypeptide encoded by the circular RNA molecule is expressed inside the animal.
[0077] In some embodiments, the DNA template (e.g., vector), linear RNA (e.g., precursor RNA), and/or circular RNA polynucleotide provided herein is between 300 and 10000, 400 and 9000, 500 and 8000, 600 and 7000, 700 and 6000, 800 and 5000, 900 and 5000, 1000 and 5000, 1100 and 5000, 1200 and 5000, 1300 and 5000, 1400 and 5000, and/or 1500 and 5000 nucleotides (nt) in length. In some embodiments, the polynucleotide is at least 300 nt, 400 nt, 500 nt, 600 nt, 700 nt, 800 nt, 900 nt, 1000 nt, 1100 nt, 1200 nt, 1300 nt, 1400 nt, 1500 nt, 2000 nt, 2500 nt, 3000 nt, 3500 nt, 4000 nt, 4500 nt, or 5000 nt in length. In some embodiments, the polynucleotide is no more than 3000 nt, 3500 nt, 4000 nt, 4500 nt, 5000 nt, 6000 nt, 7000 nt, 8000 nt, 9000 nt, or 10000 nt in length. In some embodiments, the length of a DNA, linear RNA, and/or circular RNA polynucleotide provided herein is 300 nt, 400 nt, 500 nt, 600 nt, 700 nt, 800 nt, 900 nt, 1000 nt, 1100 nt, 1200 nt, 1300 nt, 1400 nt, 1500 nt, 2000 nt, 2500 nt, 3000 nt, 3500 nt, 4000 nt, 4500 nt, 5000 nt, 6000 nt, 7000 nt, 8000 nt, 9000 nt, or 10000 nt.
[0078] In some embodiments, the circular RNA provided herein has higher functional stability than mRNA comprising the same expression sequence. In some embodiments, the circular RNA provided herein has higher functional stability than mRNA comprising the same expression sequence, 5moU modifications, an optimized UTR, a cap, and/or a polyA tail.
[0079] In some embodiments, the circular RNA polynucleotide provided herein has a functional halflife of at least 5 hours, 10 hours, 15 hours, 20 hours. 30 hours, 40 hours, 50 hours, 60 hours, 70 hours or 80 hours. In some embodiments, the circular RNA polynucleotide provided herein has a functional half-life of 5-80, 10-70, 15-60, or 20-50 hours. In some embodiments, the circular RNA polynucleotide provided herein has a functional half-life greater than (e.g., at least 1.5-fold greater than, at least 2-fold greater than) that of an equivalent linear RNA polynucleotide encoding the same protein. In some embodiments, functional half-life can be assessed through the detection of functional protein synthesis.
[0080] In some embodiments, the circular RNA polynucleotide, or pharmaceutical composition thereof, has a functional half-life in a human cell greater than or equal to that of a pre-determined threshold value. In some embodiments, the functional half-life is determined by a functional protein assay. For example, in some embodiments, the functional half-life is determined by an in vitro luciferase assay, wherein the activity of Gaussia luciferase (GLuc) is measured in the media of human cells (e.g., HepG2) expressing the circular RNA polynucleotide every 1, 2, 6, 12, or 24 hours over 1, 2, 3, 4, 5, 6, 7, or 14 days. In other embodiments, the functional half-life is determined by an in vivo assay, wherein levels of a protein encoded by the expression sequence of the circular RNA polynucleotide are measured in patient serum or tissue samples every 1, 2, 6, 12, or 24 hours over 1, 2, 3, 4, 5, 6, 7, or 14 days. In some embodiments, the pre-determined threshold value is the functional half-life of a reference linear RNA polynucleotide comprising the same expression sequence as the circular RNA polynucleotide.
[0081] In some embodiments, the circular RNA provided herein may have a higher magnitude of expression than equivalent linear mRNA, e.g., a higher magnitude of expression 24 hours after administration of RNA to cells. In some embodiments, the circular RNA provided herein has a higher magnitude of expression than mRNA comprising the same expression sequence, 5moU modifications, an optimized UTR, a cap, and/or a polyA tail.
[0082] In some embodiments, the circular RNA provided herein may be less immunogenic than an equivalent mRNA when exposed to an immune system of an organism or a certain type of immune cell. In some embodiments, the circular RNA provided herein is associated with modulated production of cytokines when exposed to an immune system of an organism or a certain type of immune cell. For example, in some embodiments, the circular RNA provided herein is associated with reduced production of IFN-pi, RIG-I, IL-2, IL-6, IFNy, and/or TNFa when exposed to an immune system of an organism or a certain type of immune cell as compared to mRNA comprising the same expression sequence. In some embodiments, the circular RNA provided herein is associated with less IFN-pi, RIG-I, IL-2, IL-6, IFNy, and/or TNFa transcript induction when exposed to an immune system of an organism or a certain type of immune cell as compared to mRNA comprising the same expression sequence. In some embodiments, the circular RNA provided herein is less immunogenic than mRNA
comprising the same expression sequence. In some embodiments, the circular RNA provided herein is less immunogenic than mRNA comprising the same expression sequence, 5moU modifications, an optimized UTR, a cap, and/or a polyA tail.
[0083] In certain embodiments, the circular RNA provided herein can be transfected into a cell as is or can be transfected in DNA vector form and transcribed in the cell. Transcription of circular RNA from a transfected DNA vector can be via added polymerases or polymerases encoded by nucleic acids transfected into the cell, or preferably via endogenous polymerases. i. Enhanced Intron Elements & Enhanced Exon Elements
[0084] In some embodiments, the DNA template (e.g., vector) or linear RNA (e.g., precursor RNA) comprises an enhanced intron element and/or enhanced exon element. The enhanced intron elements and enhanced exon elements may comprise spacers, duplex regions, affinity sequences, intron fragments, exon fragments and various untranslated elements. These sequences within the enhanced intron elements or enhanced exon elements are arranged to optimize circularization or protein expression.
[0085] In certain embodiments, the DNA template, precursor linear RNA polynucleotide and circular RNA provided herein comprise a first (5’) and/or a second (3’) spacer. In some embodiments, the DNA template or precursor linear RNA polynucleotide comprises one or more spacers in the enhanced intron elements. In some embodiments, the DNA template, precursor linear RNA polynucleotide comprises one or more spacers in the enhanced exon elements. In certain embodiments, the DNA template or linear RNA polynucleotide comprises a spacer in the 3’ enhanced intron fragment and a spacer in the 5’ enhanced intron fragment. In certain embodiments, DNA template, precursor linear RNA polynucleotide, or circular RNA comprises a spacer in the 3’ enhanced exon fragment and another spacer in the 5’ enhanced exon fragment to aid with circularization or protein expression due to symmetry created in the overall sequence.
[0086] In some embodiments, including a spacer between the 3’ group I intron fragment and the core functional element may conserve secondary structures in those regions by preventing them from interacting, thus increasing splicing efficiency. In some embodiments, the first (between 3’ group I intron fragment and core functional element) and second (between the two expression sequences and core functional element) spacers comprise additional base pairing regions that are predicted to base pair with each other and not to the first and second duplex regions. In other embodiments, the first (between 3’ group I intron fragment and core functional element) and second (between the one of the core functional element and 5’ group I intron fragment) spacers comprise additional base pairing regions that are predicted to base pair with each other and not to the first and second duplex regions. In some embodiments, such spacer base pairing brings the group I intron fragments in close proximity to each
other, further increasing splicing efficiency. Additionally, in some embodiments, the combination of base pairing between the first and second duplex regions, and separately, base pairing between the first and second spacers, promotes the formation of a splicing bubble containing the group I intron fragments flanked by adjacent regions of base pairing. Typical spacers are contiguous sequences with one or more of the following qualities: 1) predicted to avoid interfering with proximal structures, for example, the IRES, expression sequence, aptamer, or intron; 2) is at least 7 nt long and no longer than 100 nt; 3) is located after and adjacent to the 3’ intron fragment and/or before and adjacent to the 5’ intron fragment; and 4) contains one or more of the following: a) an unstructured region at least 5 nt long, b) a region of base pairing at least 5 nt long to a distal sequence, including another spacer, and c) a structured region at least 7 nt long limited in scope to the sequence of the spacer. Spacers may have several regions, including an unstructured region, a base pairing region, a hairpin/structured region, and combinations thereof. In an embodiment, the spacer has a structured region with high GC content. In an embodiment, a region within a spacer base pairs with another region within the same spacer. In an embodiment, a region within a spacer base pairs with a region within another spacer. In an embodiment, a spacer comprises one or more hairpin structures. In an embodiment, a spacer comprises one or more hairpin structures with a stem of 4 to 12 nucleotides and a loop of 2 to 10 nucleotides. In an embodiment, there is an additional spacer between the 3’ group I intron fragment and the core functional element. In an embodiment, this additional spacer prevents the structured regions of the IRES or aptamer of a TIE from interfering with the folding of the 3’ group I intron fragment or reduces the extent to which this occurs. In some embodiments, the 5’ spacer sequence is at least 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25 or 30 nucleotides in length. In some embodiments, the 5’ spacer sequence is no more than 100, 90, 80, 70, 60, 50, 45, 40, 35 or 30 nucleotides in length. In some embodiments the 5’ spacer sequence is between 5 and 50, 10 and 50, 20 and 50, 20 and 40, and/or 25 and 35 nucleotides in length. In certain embodiments, the 5’ spacer sequence is 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49 or 50 nucleotides in length. In one embodiment, the 5’ spacer sequence is a polyA sequence. In another embodiment, the 5’ spacer sequence is a poly AC sequence. In one embodiment, a spacer comprises 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% poly AC content. In one embodiment, a spacer comprises 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% polypyrimidine (C/T or C/U) content.
[0087] In some embodiments, the DNA template and precursor linear RNA polynucleotides and circular RNA polynucleotide provided herein comprise a first (5’) duplex region and a second (3’) duplex region. In certain embodiments, the DNA template and precursor linear RNA polynucleotide comprises a 5’ external duplex region located within the 3’ enhanced intron fragment and a 3’ external duplex region located within the 5’ enhanced intron fragment. In some embodiments, the DNA template, precursor linear RNA polynucleotide and circular RNA polynucleotide comprise a 5’ internal
duplex region located within the 3’ enhanced exon fragment and a 3’ internal duplex region located within the 5’ enhanced exon fragment. In some embodiments, the DNA polynucleotide and precursor linear RNA polynucleotide comprises a 5’ external duplex region, 5’ internal duplex region, a 3’ internal duplex region, and a 3’ external duplex region.
[0088] In certain embodiments, the first and second duplex regions may form perfect or imperfect duplexes. Thus, in certain embodiments at least 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% of the first and second duplex regions may be base paired with one another. In some embodiments, the duplex regions are predicted to have less than 50% (e.g., less than 45%, less than 40%, less than 35%, less than 30%, less than 25%) base pairing with unintended sequences in the RNA (e.g., non-duplex region sequences). In some embodiments, including such duplex regions on the ends of the precursor RNA strand, and adjacent or very close to the group I intron fragment, bring the group I intron fragments in close proximity to each other, increasing splicing efficiency. In some embodiments, the duplex regions are 3 to 100 nucleotides in length (e.g., 3-75 nucleotides in length, 3-50 nucleotides in length, 20-50 nucleotides in length, 35-50 nucleotides in length, 5-25 nucleotides in length, 9-19 nucleotides in length). In some embodiments, the duplex regions are 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, or 50 nucleotides in length. In some embodiments, the duplex regions have a length of 9 to 50 nucleotides. In one embodiment, the duplex regions have a length of 9 to 19 nucleotides. In some embodiments, the duplex regions have a length of 20 to 40 nucleotides. In certain embodiments, the duplex regions have a length of 30 nucleotides.
[0089] In other embodiments, the DNA template, precursor linear RNA polynucleotide, or circular RNA polynucleotide does not comprise of any duplex regions to optimize translation or circularization.
[0090] As provided herein, the DNA template or precursor linear RNA polynucleotide may comprise an affinity tag. In some embodiments, the affinity tag is located in the 3’ enhanced intron element. In some embodiments, the affinity tag is located in the 5’ enhanced intron element. In some embodiments, both (3’ and 5’) enhanced intron elements each comprise an affinity tag. In one embodiment, an affinity tag of the 3’ enhanced intron element is the length as an affinity tag in the 5’ enhanced intron element. In some embodiments, an affinity tag of the 3’ enhanced intron element is the same sequence as an affinity tag in the 5’ enhanced intron element. In some embodiments, the affinity sequence is placed to optimize oligo-dT purification.
[0091] In some embodiments, an affinity tag comprises a polyA region. In some embodiments the polyA region is at least 15, 30, or 60 nucleotides long. In some embodiments, one or both polyA regions is 15-50 nucleotides long. In some embodiments, one or both polyA regions is 20-25 nucleotides long. The polyA sequence is removed upon circularization. Thus, an oligonucleotide hybridizing with the
polyA sequence, such as a deoxythymine oligonucleotide (oligo(dT)) conjugated to a solid surface (e.g., a resin), can be used to separate circular RNA from its precursor RNA.
[0092] In certain embodiments, the 3’ enhanced intron element comprises a leading untranslated sequence. In some embodiments, the leading untranslated sequence is a the 5’ end of the 3’ enhanced intron fragment. In some embodiments, the leading untranslated sequence comprises of the last nucleotide of a transcription start site (TSS). In some embodiments, the TSS is chosen from a viral, bacterial, or eukaryotic DNA template. In one embodiment, the leading untranslated sequence comprise the last nucleotide of a TSS and 0 to 100 additional nucleotides. In some embodiments, the TSS is a terminal spacer. In one embodiment, the leading untranslated sequence contains a guanosine at the 5’ end upon translation of an RNA T7 polymerase.
[0093] In certain embodiments, the 5’ enhanced intron element comprises a trailing untranslated sequence. In some embodiments, the 5’ trailing untranslated sequence is located at the 3’ end of the 5’ enhanced intron element. In some embodiments, the trailing untranslated sequence is a partial restriction digest sequence. In one embodiment, the trailing untranslated sequence is in whole or in part a restriction digest site used to linearize the DNA template. In some embodiments, the restriction digest site is in whole or in part from a natural viral, bacterial or eukaryotic DNA template. In some embodiments, the trailing untranslated sequence is a terminal restriction site fragment. a. Enhanced Intron Fragments
[0094] In some embodiments, the 3’ enhanced intron element and 5’ enhanced intron element each comprise an intron fragment. In certain embodiments, a 3’ intron fragment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to a 3’ proximal fragment of a natural group I or II intron including the 3’ splice site dinucleotide. Typically, a 5’ intron fragment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to a 5’ proximal fragment of a natural group I or II intron including the 5’ splice site dinucleotide. In some embodiments, the 3’ intron fragment includes the first nucleotide of a 3’ group I or II splice site dinucleotide. In some embodiments, the 5’ intron fragment includes the first nucleotide of a 5’ group I or II splice site dinucleotide. In other embodiments, the 3’ intron fragment includes the first and second nucleotides of a 3’ group I or II intron fragment splice site dinucleotide; and the 5’ intron fragment includes the first and second nucleotides of a 3’ group I or II intron fragment dinucleotide. In some embodiments the 3’ enhanced intron element and 5’ enhanced intron element comprises a synthetic intron fragment.
b. Enhanced Exon Fragments
[0095] In certain embodiments, as provided herein, the DNA template, linear precursor RNA polynucleotide, and circular RNA polynucleotide each comprise an enhanced exon fragment. In some embodiments, following a 5’ to 3’ order, the 3’ enhanced exon element is located upstream to core functional element. In some embodiments, following a 5’ to 3’ order, the 5’ enhanced intron element is located downstream to the core functional element.
[0096] In some embodiments, the 3’ enhanced exon element and 5’ enhanced exon element each comprise an exon fragment. In some embodiments, the 3’ enhanced exon element comprises a 3’ exon fragment. In some embodiments, the 5’ enhanced exon element comprises a 5’ exon fragment. In certain embodiments, as provided herein, the 3’ exon fragment and 5’ exon fragment each comprises a group I or II intron fragment and 1 to 100 nucleotides of an exon sequence. In certain embodiments, a 3’ intron fragment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to a 3’ proximal fragment of a natural group I or II intron including the 3’ splice site dinucleotide. Typically, a 5’ group I or II intron fragment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to a 5’ proximal fragment of a natural group I or II intron including the 5’ splice site dinucleotide. In some embodiments, the 3’ exon fragment comprises a second nucleotide of a 3’ group I or II intron splice site dinucleotide and 1 to 100 nucleotides of an exon sequence. In some embodiments, the 5’ exon fragment comprises the first nucleotide of a 5’ group I or II intron splice site dinucleotide and 1 to 100 nucleotides of an exon sequence. In some embodiments, the exon sequence comprises in part or in whole from a naturally occurring exon sequence from a virus, bacterium or eukaryotic DNA vector. In other embodiments, the exon sequence further comprises a synthetic, genetically modified (e.g., containing modified nucleotide), or other engineered exon sequence.
[0097] In one embodiment, where the 3’ intron fragment comprises both nucleotides of a 3’ group I or II splice site dinucleotide and the 5’ intron fragment comprises both nucleotides of a 5’ group I or II splice site dinucleotide, the exon fragments located within the 5’ enhanced exon element and 3’ enhanced exon element does not comprise of a group I or II splice site dinucleotide.
[0098] For means of example and not intended to be limiting, in some embodiment, a 3’ enhanced intron element comprises in the following 5’ to 3’ order: a leading untranslated sequence, a 5’ affinity tag, an optional 5’ external duplex region, a 5’ external spacer, and a 3’ intron fragment. In same embodiments, the 3’ enhanced exon element comprises in the following 5’ to 3’ order: a 3’ exon fragment, an optional 5’ internal duplex region, an optional 5’ internal duplex region, and a 5’ internal spacer. In the same embodiments, the 5’ enhanced exon element comprises in the following 5’ to 3’ order: a 3’ internal spacer, an optional 3’ internal duplex region, and a 5’ exon fragment. In still the
same embodiments, the 3’ enhanced intron element comprises in the following 5’ to 3’ order: a 5’ intron fragment, a 3’ external spacer, an optional 3’ external duplex region, a 3’ affinity tag, and a trailing untranslated sequence. ii. Core Functional Element
[0099] In some embodiments, the DNA template, linear precursor RNA polynucleotide, and circular RNA polynucleotide comprise a core functional element. In some embodiments, the core functional element comprises a coding or noncoding element. In certain embodiments, the core functional element may contain both a coding and noncoding element. In some embodiments, the core functional element further comprises translation initiation element (TIE) upstream to the coding or noncoding element. In some embodiments, the core functional element comprises a termination element. In some embodiments, the termination element is located downstream to the TIE and coding element. In some embodiments, the termination element is located downstream to the coding element but upstream to the TIE. In certain embodiments, where the coding element comprises a noncoding region, a core functional element lacks a TIE and/or a termination element.
Hi. Coding or Noncoding Element
[0100] In some embodiments, the polynucleotides herein comprise a coding element, a noncoding element, or a combination of both. In some embodiments, the coding element comprises an expression sequence. In some embodiments, the coding element encodes at least one therapeutic protein.
[0101] In some embodiments, the circular RNA encodes two or more polypeptides. In some embodiments, the circular RNA is a bicistronic RNA. The sequences encoding the two or more polypeptides can be separated by a ribosomal skipping element or a nucleotide sequence encoding a protease cleavage site. In certain embodiments, the ribosomal skipping element encodes thosea-asigna virus 2A peptide (T2A), porcine teschovirus-1 2 A peptide (P2A), foot-and-mouth disease virus 2 A peptide (F2A), equine rhinitis A vims 2 A peptide (E2A), cytoplasmic polyhedrosis vims 2 A peptide (BmCPV 2A), or flacherie vims of B. mori 2A peptide (BmIFV 2A).
Hi. Translation Initiation Element (TIE)
[0102] As provided herein in some embodiments, the core functional element comprises at least one translation initiation element (TIE). TIEs are designed to allow translation efficiency of an encoded protein. Thus, optimal core functional elements comprising only of noncoding elements lack any TIEs. In some embodiments, core functional elements comprising one or more coding element will further comprise one or more TIEs.
[0103] In some embodiments, a TIE comprises an untranslated region (UTR). In certain embodiments, the TIE provided herein comprise an internal ribosome entry site (IRES). Inclusion of an IRES permits the translation of one or more open reading frames from a circular RNA (e.g., open reading frames that
form the expression sequences). The IRES element attracts a eukaryotic ribosomal translation initiation complex and promotes translation initiation. See, e.g., Kaufman etal., Nuc. Acids Res. (1991) 19:4485- 4490; Gurtu et al., Biochem. Biophys. Res. Comm. (1996) 229:295-298; Rees et al., BioTechniques (1996) 20: 102-110; Kobayashi et al., BioTechniques (1996) 21 :399-402; and Mosser et al., BioTechniques 1997 22 150-161. In some embodiments, the IRES element is selected from those disclosed in international publication WO/2022/261490, the contents of which are hereby incorporated in their entireties. iv. Additional Accessory Elements (Sequence Elements)
[0104] As described in this disclosure, the circular RNA polynucleotide, linear RNA polynucleotide, and/or DNA template may further comprise of accessory elements. In certain embodiments, these accessory elements may be included within the sequences of the circular RNA, linear RNA polynucleotide and/or DNA template for enhancing circularization, translation or both. Accessory elements are sequences, in certain embodiments that are located with specificity between or within the enhanced intron elements, enhanced exon elements, or core functional element of the respective polynucleotide. As an example, but not intended to be limiting, an accessory element includes, a IRES transacting factor region, a miRNA binding site, a restriction site, an RNA editing region, a structural or sequence element, a granule site, a zip code element, an RNA trafficking element or another specialized sequence as found in the art that enhances promotes circularization and/or translation of the protein encoded within the circular RNA polynucleotide.
[0105] In certain embodiments, the accessory element comprises an IRES transacting factor (ITAF) region. In some embodiments, the IRES transacting factor region modulates the initiation of translation through binding to PCBP1 - PCBP4 (polyC binding protein), PABP1 (polyA binding protein), PTB (polyprimidine tract binding), Argonaute protein family, HNRNPK (Heterogeneous nuclear ribonucleoprotein K protein), or La protein. In some embodiments, the IRES transacting factor region comprises a polyA, polyC, poly AC, or polyprimidine track.
[0106] In some embodiments, the ITAF region is located within the core functional element. In some embodiments, the ITAF region is located within the TIE.
[0107] In certain embodiments, the accessory element comprises a miRNA binding site. In some embodiments the miRNA binding site is located within the 5’ intron element, 5’ exon element, core functional element, 3’ exon element, and/or 3’ intron element.
[0108] In some embodiments, wherein the miRNA binding site is located within the spacer within the intron element or exon element. In certain embodiments, the miRNA binding site comprises the entire spacer regions.
[0109] In some embodiments, the 5’ intron element and 3’ intron elements each comprise identical miRNA binding sites. In another embodiment, the miRNA binding site of the 5’ intron element comprises a different, in length or nucleotides, miRNA binding site than the 3’ intron element. In one embodiment, the 5’ exon element and 3’ exon element comprise identical miRNA binding sites. In other embodiments, the 5’ exon element and 3’ exon element comprises different, in length or nucleotides, miRNA binding sites.
[0110] In some embodiments, the miRNA binding sites are located adjacent to each other within the circular RNA polynucleotide, linear RNA polynucleotide precursor, and/or DNA template. In certain embodiments, the first nucleotide of one of the miRNA binding sites follows the first nucleotide last nucleotide of the second miRNA binding site.
[0111] In some embodiments, the miRNA binding site is located within a translation initiation element (TIE) of a core functional element. In one embodiment, the miRNA binding site is located before, trailing or within an internal ribosome entry site (IRES). In another embodiment, the miRNA binding site is located before, trailing, or within an aptamer complex.
[112] The unique sequences defined by the miRNA nomenclature are widely known and accessible to those working in the micrcircRNA field. For example, they can be found in the miRDB public database. v. Natural Ties: Viral & Eukaryotic/Cellular Internal Ribosome Entry Sites (IRES)
[0113] A multitude of IRES sequences are available and include sequences derived from a wide variety of viruses, such as from leader sequences of piccircRNAviruses such as the encephalomyocarditis virus (EMCV) UTR (Jang et al., J. Virol. (1989) 63: 1651-1660), the polio leader sequence, the hepatitis A virus leader, the hepatitis C virus IRES, human rhinovirus type 2 IRES (Dobrikova et al., Proc. Natl. Acad. Sci. (2003) 100(25): 15125- 15130), an IRES element from the foot and mouth disease virus (Ramesh et al., Nucl. Acid Res. (1996) 24:2697-2700), a giardiavirus IRES (Garlapati et al., J. Biol. Chem. (2004) 279(5):3389-3397), and the like.
[0114] For driving protein expression, the circular RNA comprises an IRES operably linked to a protein coding sequence. Modifications of IRES and accessory sequences are disclosed herein to increase or reduce IRES activities, for example, by truncating the 5’ and/or 3’ ends of the IRES, adding a spacer 5’ to the IRES, modifying the 6 nucleotides 5’ to the translation initiation site (Kozak sequence), modification of alternative translation initiation sites, and creating chimeric/hybrid IRES sequences. In some embodiments, the IRES sequence in the circular RNA disclosed herein comprises one or more of these modifications relative to a native IRES.
[0115] In some embodiments, the IRES is an IRES sequence of Taura syndrome virus, Triatoma virus, Theiler's encephalomyelitis virus, Simian Virus 40, Solenopsis invicta virus 1, Rhopalosiphum padi virus, Reticuloendotheliosis virus, Human poliovirus 1, Plautia stali intestine virus, Kashmir bee virus, Human rhinovirus 2, Homalodisca coagulata virus- 1, Human Immunodeficiency Virus type 1, Himetobi P virus, Hepatitis C virus, Hepatitis A virus, Hepatitis GB virus, Foot and mouth disease virus, Human enterovirus 71, Equine rhinitis virus, Ectropis obliqua piccircRNA-like virus, Encephalomyocarditis virus, Drosophila C Virus, Human coxsackievirus B3, Crucifer tobamovirus, Cricket paralysis virus, Bovine viral diarrhea virus 1, Black Queen Cell Virus, Aphid lethal paralysis virus, Avian encephalomyelitis virus, Acute bee paralysis virus, Hibiscus chlorotic ringspot virus, Classical swine fever virus, Human FGF2, Human SFTPA1, Human AML1/RUNX1, Drosophila antennapedia, Human AQP4, Human AT1R, Human BAG-1, Human BCL2, Human BiP, Human c- IAP1, Human c-myc, Human eIF4G, Mouse NDST4L, Human LEF1, Mouse HIF1 alpha, Human n.myc, Mouse Gtx, Human p27kipl, Human PDGF2/c-sis, Human p53, Human Pim-1, Mouse Rbm3, Drosophila reaper, Canine Scamper, Drosophila Ubx, Human UNR, Mouse UtrA, Human VEGF-A, Human XIAP, Drosophila hairless, S. cerevisiae TFIID, S. cerevisiae YAP1, tobacco etch virus, turnip crinkle virus, EMCV-A, EMCV-B, EMCV-Bf, EMCV-Cf, EMCV pEC9, Picobirnavirus, HCV QC64, Human Cosavirus E/D, Human Cosavirus F, Human Cosavirus JMY, Rhino virus NAT001, HRV14, HRV89, HRVC-02, HRV-A21, Salivirus A SHI, Salivirus FHB, Salivirus NG-J1, Human Parechovirus 1, Crohivirus B, Yc-3, Rosavirus M-7, Shanbavirus A, Pasivirus A, Pasivirus A 2, Echovirus E14, Human Parechovirus 5, Aichi Virus, Hepatitis A Virus HA16, Phopivirus, CVA10, Enterovirus C, Enterovirus D, Enterovirus J, Human Pegivirus 2, GBV-C GT110, GBV-C K1737, GBV-C Iowa, Pegivirus A 1220, Pasivirus A 3, Sapelovirus, Rosavirus B, Bakunsa Virus, Tremovirus A, Swine Pasivirus 1, PLV-CHN, Pasivirus A, Sicinivirus, Hepacivirus K, Hepacivirus A, BVDV1, Border Disease Virus, BVDV2, CSFV-PK15C, SF573 Dicistro virus, Hubei PiccircRNA-like Virus, CRPV, Salivirus A BN5, Salivirus A BN2, Salivirus A 02394, Salivirus A GUT, Salivirus A CH, Salivirus A SZ1, Salivirus FHB, CVB3, CVB1, Echovirus 7, CVB5, EVA71, CVA3, CVA12, EV24 or an aptamer to eIF4G.
[0116] In some embodiments, the IRES comprises in whole or in part from a eukaryotic or cellular IRES. In certain embodiments, the IRES is from a human gene, where the human gene is ABCF1, ABCG1, ACAD10, ACOT7, ACSS3, ACTG2, ADCYAP1, ADK, AGTR1, AHCYL2, AHI1, AKAP8L, AKR1A1, ALDH3A1, ALDOA, ALG13, AMMECR1L, ANGPTL4, ANK3, AOC3, AP4B1, AP4E1, APAF1, APBB1, APC, APH1A, APOBEC3D, APOM, APP, AQP4, ARHGAP36, ARL13B, ARMC8, ARMCX6, ARPC1A, ARPC2, ARRDC3, ASAP1, ASB3, ASB5, ASCL1, ASMTL, ATF2, ATF3, ATG4A, ATP5B, ATP6V0A1, ATXN3, AURKA, AURKA, AURKA, AURKA, B3GALNT1, B3GNTL1, B4GALT3, BAAT, BAG1, BAIAP2, BAIAP2L2, BAZ2A, BBX, BCAR1, BCL2, BCS1L, BET1, BID, BIRC2, BPGM, BPIFA2, BRINP2, BSG, BTN3A2, C12orf43,
C14orf93, C17orf62, Clorf226, C21orf62, C2orfl5, C4BPB, C4orf22, C9orf84, CACNA1A, CALC0C02, CAPN11, CASP12, CASP8AP2, CAV1, CBX5, CCDC120, CCDC17, CCDC186, CCDC51, CCN1, CCND1, CCNT1, CD2BP2, CD9, CDC25C, CDC42, CDC7, CDCA7L, CDIP1, CDK1, CDK11A, CDKN1B, CEACAM7, CEP295NL, CFLAR, CHCHD7, CHIA, CHICI, CHMP2A, CHRNA2, CLCN3, CLEC12A, CLEC7A, CLECL1, CLRN1, CMSS1, CNIH1, CNR1, CNTN5, C0G4, C0MMD1, COMMD5, CPEB1, CPS1, CRACR2B, CRBN, CREM, CRYBG1, CSDE1, CSF2RA, CSNK2A1, CSTF3, CTCFL, CTH, CTNNA3, CTNNB1, CTNNB1, CTNND1, CTSL, CUTA, CXCR5, CYB5R3, CYP24A1, CYP3A5, DAG1, DAP3, DAP5, DAXX, DCAF4, DCAF7, DCLRE1A, DCP1A, DCTN1, DCTN2, DDX19B, DDX46, DEFB123, DGKA, DGKD, DHRS4, DHX15, DIO3, DLG1, DLL4, DMD UTR, DMD ex5, DMKN, DNAH6, DNAL4, DUSP13, DUSP19, DYNC1I2, DYNLRB2, DYRK1A, ECI2, ECT2, EIF1AD, EIF2B4, EIF4G1, EIF4G2, EIF4G3, ELANE, ELOVL6, ELP5, EMCN, ENO1, EPB41, ERMN, ERVV-1, ESRRG, ETFB, ETFBKMT, ETV1, ETV4, EXD1, EXT1, EZH2, FAM111B, FAM157A, FAM213A, FBXO25, FBXO9, FBXW7, FCMR, FGF1, FGF1, FGF1A, FGF2, FGF2, FGF-9, FHL5, FMRI, FN1, FOXP1, FTH1, FUBP1, G3BP1, GABBR1, GALC, GART, GAS7, gastrin, GATA1, GATA4, GFM2, GHR, GJB2, GLI1, GLRA2, GMNN, GPAT3, GPATCH3, GPR137, GPR34, GPR55, GPR89A, GPRASP1, GRAP2, GSDMB, GSTO2, GTF2B, GTF2H4, GUCY1B2, HAX1, HCST, HIGD1A, HIGD1B, HIPK1, HIST1H1C, HIST1H3H, HK1, HLA-DRB4, HMBS, HMGA1, HNRNPC, HOPX, HOXA2, HOXA3, HPCAL1, HR, HSP90AB1, HSPA1A, HSPA4L, HSPA5, HYPK, IFFO1, IFT74, IFT81, IGF1, IGF1R, IGF1R, IGF2, IL11, IL17RE, IL1RL1, IL1RN, IL32, IL6, ILF2, ILVBL, INSR, INTS13, IP6K1, ITGA4, ITGAE, KCNE4, KERA, KIAA0355, KIAA0895L, KIAA1324, KIAA1522, KIAA1683, KIF2C, KIZ, KLHL31, KLK7, KRR1, KRT14, KRT17, KRT33A, KRT6A, KRTAP10-2, KRTAP13- 3, KRTAP13-4, KRTAP5-11, KRTCAP2, LACRT, LAMB1, LAMB3, LANCL1, LBX2, LCAT, LDHA, LDHAL6A, LEF1, LINC-PINT, LM03, LRRC4C, LRRC7, LRTOMT, LSM5, LTB4R, LYRM1, LYRM2, MAGEA11, MAGEA8, MAGEB1, MAGEB16, MAGEB3, MAPT, MARS, MC1R, MCCC1, METTL12, METTL7A, MGC 16025, MGC 16025, MIA2, MIA2, MITF, MKLN1, MNT, MORF4L2, MPD6, MRFAP1, MRPL21, MRPS12, MSI2, MSLN, MSN, MT2A, MTFR1L, MTMR2, MTRR, MTUS1, MYB, MYC, MYCL, MYCN, MYL10, MYL3, MYLK, MYO1A, MYT2, MZB1, NAP1L1, NAVI, NBAS, NCF2, NDRG1, NDST2, NDUFA7, NDUFB11, NDUFC1, NDUFS1, NEDD4L, NFAT5, NFE2L2, NFE2L2, NFIA, NHEJ1, NHP2, NITI, NKRF, NME1-NME2, NPAT, NR3C1, NRBF2, NRF1, NTRK2, NUDCD1, NXF2, NXT2, ODC1, ODF2, OPTN, OR10R2, OR11L1, OR2M2, OR2M3, OR2M5, OR2T10, OR4C15, OR4F17, OR4F5, OR5H1, OR5K1, OR6C3, OR6C75, OR6N1, OR7G2, p53, P2RY4, PAN2, PAQR6, PARP4, PARP9, PC, PCBP4, PCDHGC3, PCLAF, PDGFB, PDZRN4, PELO, PEMT, PEX2, PFKM, PGBD4, PGLYRP3, PHLDA2, PHTF1, PI4KB, PIGC, PIM1, PKD2L1, PKM, PLCB4, PLD3, PLEKHA1, PLEKHB1, PLS3, PML, PNMA5, PNN, POC1A, POC1B, POLD2, POLD4, POU5F1, PPIG, PQBP1, PRAME, PRPF4, PRR11, PRRT1,
PRSS8, PSMA2, PSMA3, PSMA4, PSMD11, PSMD4, PSMD6, PSME3, PSMG3, PTBP3, PTCHI, PTHLH, PTPRD, PUS7L, PVRIG, QPRT, RAB27A, RAB7B, RABGGTB, RAET1E, RALGDS, RALYL, RARB, RCVRN, REG3G, RFC5, RGL4, RGS19, RGS3, RHD, RINL, RIP0R2, RITA1, RMDN2, RNASE1, RNASE4, RNF4, RPA2, RPL17, RPL21, RPL26L1, RPL28, RPL29, RPL41, RPL9, RPS11, RPS13, RPS14, RRBP1, RSU1, RTP2, RUNX1, RUNX1T1, RUNX1T1, RUNX2, RUSC1, RXRG, S100A13, S100A4, SAT1, SCHIP1, SCMH1, SEC14L1, SEMA4A, SERPINA1, SERPINB4, SERTAD3, SFTPD, SH3D19, SHC1, SHMT1, SHPRH, SIM1, SIRT5, SLC11A2, SLC12A4, SLC16A1, SLC25A3, SLC26A9, SLC5A11, SLC6A12, SLC6A19, SLC7A1, SLFN11, SLIRP, SMAD5, SMARCAD1, SMN1, SNCA, SNRNP200, SNRPB2, SNX12, S0D1, SOX13, SOX5, SP8, SPARCL1, SPATA12, SPATA31C2, SPN, SPOP, SQSTM1, SRBD1, SRC, SREBF1, SRPK2, SSB, SSB, SSBP1, ST3GAL6, STAB1, STAMBP, STAU1, STAU1, STAU1, STAU1, STAU1, STK16, STK24, STK38, STMN1, STX7, SULT2B1, SYK, SYNPR, TAF1C, TAGLN, TANK, TAS2R40, TBC1D15, TBXAS1, TCF4, TDGF1, TDP2, TDRD3, TDRD5, TESK2, THAP6, THBD, THTPA, TIAM2, TKFC, TKTL1, TLR10, TM9SF2, TMC6, TMCO2, TMED10, TMEM116, TMEM126A, TMEM159, TMEM208, TMEM230, TMEM67, TMPRSS13, TMUB2, TNFSF4, TNIP3, TP53, TP53, TP73, TRAF1, TRAK1, TRIM31, TRIM6, TRMT1, TRMT2B, TRPM7, TRPM8, TSPEAR, TTC39B, TTLL11, TUBB6, TXLNB, TXNIP, TXNL1, TXNRD1, TYROBP, U2AF1, UBA1, UBE2D3, UBE2I, UBE2L3, UBE2V1, UBE2V2, UMPS, UNG, UPP2, USMG5, USP18, UTP14A, UTRN, UTS2, VDR, VEGFA, VEGFA, VEPH1, VIPAS39, VPS29, VSIG10L, WDHD1, WDR12, WDR4, WDR45, WDYHV1, WRAP53, XIAP, XPNPEP3, YAP1, YWHAZ, YY1AP1, ZBTB32, ZNF146, ZNF250, ZNF385A, ZNF408, ZNF410, ZNF423, ZNF43, ZNF502, ZNF512, ZNF513, ZNF580, ZNF609, ZNF707, or ZNRDE vi. Synthetic Ties: Aptamer Complexes, Modified Nucleotides, IRES Variants & Other Engineered Ties
[0117] As contemplated herein, in some embodiments, a translation initiation element (TIE) comprises a synthetic TIE. In some embodiments, a synthetic TIE comprises aptamer complexes, synthetic IRES or other engineered TIES capable of initiating translation of a linear RNA or circular RNA polynucleotide.
[0118] In some embodiments, one or more aptamer sequences is capable of binding to a component of a eukaryotic initiation factor to either enhance or initiate translation. In some embodiments, aptamer may be used to enhance translation in vivo and in vitro by promoting specific eukaryotic initiation factors (elF) (e.g., aptamer in WO2019081383 is capable of binding to eukaryotic initiation factor 4F (eIF4F). In some embodiments, the aptamer or a complex of aptamers may be capable of binding to EIF4G, EIF4E, EIF4A, EIF4B, EIF3, EIF2, EIF5, EIF1, EIF1A, 40S ribosome, PCBP1 (polyC binding protein), PCBP2, PCBP3, PCBP4, PABP1 (polyA binding protein), PTB, Argonaute protein family, HNRNPK (heterogeneous nuclear ribonucleoprotein K), or La protein.
vii. Termination Sequence
[0119] In some embodiments, the core functional element comprises a termination sequence. In some embodiments, the termination sequence comprises a stop codon. In one embodiment, the termination sequence comprises a stop cassette. In some embodiments, the stop cassette comprises at least 2 stop codons. In some embodiments, the stop cassette comprises at least 2 frames of stop codons. In the same embodiment, the frames of the stop codons in a stop cassette each comprise 1, 2 or more stop codons. In some embodiments, the stop cassette comprises a LoxP or a RoxStopRox, or frt-flanked stop cassette. In the same embodiment, the stop cassette comprises a lox-stop-lox stop cassette. viii. Variants
[0120] In some embodiments, a circular RNA polynucleotide provided herein comprises modified RNA nucleotides and/or modified nucleosides. In some embodiments, the modified nucleoside is m5C (5-methylcytidine). In one embodiment, the modified nucleoside is m5U (5 -methyluridine). In another embodiment, the modified nucleoside is m6A (N6-methyladenosine). In another embodiment, the modified nucleoside is s2U (2-thiouridine). In another embodiment, the modified nucleoside is T (pseudouridine). In another embodiment, the modified nucleoside is Um (2'-O-methyluridine). In other embodiments, the modified nucleoside is m’A (1 -methyladenosine); m2A (2-methyladenosine); Am (2’-O-methyladenosine); ms2m6A (2-methylthio-N6-methyladenosine); i6A (N6-isopentenyladenosine); ms2i6A (2-methylthio-N6isopentenyladenosine); io6A (N6-(cis-hydroxyisopentenyl)adenosine); ms2io6A (2-methylthio-N6-(cis-hydroxyisopentenyl)adenosine); g6A (N6- glycinylcarbamoyladenosine); t6A (N6-threonylcarbamoyladenosine); ms2t6A (2-methylthio-N6- threonyl carbamoyladenosine); m6t6A (N6-methyl-N6-threonylcarbamoyladenosine); hn6A(N6- hydroxynorvalylcarbamoyladenosine) ; ms2hn6A (2-methylthio-N6-hydroxynorvalyl carbamoyladenosine); Ar(p) (2’ -O-ribosyladenosine (phosphate)); I (inosine); m’l (1 -methylinosine); m’lm (l,2’-O-dimethylinosine); m3C (3-methylcytidine); Cm (2’-O-methylcytidine); s2C (2- thiocytidine); ac4C (N4-acetylcytidine); UC (5 -formylcytidine); m5Cm (5,2'-O-dimethylcytidine); ac4Cm (N4-acetyl-2’-O-methylcytidine); k2C (lysidine); m’G (1 -methylguanosine); m2G (N2- methylguanosine); m7G (7-methylguanosine); Gm (2'-O-methylguanosine); m2 2G (N2,N2- dimethylguanosine); m2Gm (N2,2’-O-dimethylguanosine); m2 2Gm (N2,N2,2’-O-trimethylguanosine); Gr(p) (2’-O-ribosylguanosine(phosphate)); yW (wybutosine); O2yW (peroxy wybutosine); OHyW (hydroxywybutosine); OHyW* (undermodified hydroxy wybutosine); imG (wyosine); mimG (methylwyosine); Q (queuosine); oQ (epoxy queuosine); galQ (galactosyl-queuosine); manQ (mannosyl-queuosine); preQo (7-cyano-7-deazaguanosine); preQi (7-aminomethyl-7-deazaguanosine); G+ (archaeosine); D (dihydrouridine); m5Um (5,2’-O-dimethyluridine); s4U (4-thiouridine); m5s2U (5- methyl-2-thiouridine); s2Um (2-thio-2’-O-methyluridine); acp3U (3-(3-amino-3- carboxypropyl)uridine); ho5U (5 -hydroxyuridine); mo5U (5 -methoxyuridine); cmo5U (uridine 5- oxy acetic acid); mcmo5U (uridine 5 -oxy ace tic acid methyl ester); chm5U (5-
(carboxyhydroxymethyl)uridine)); mchm5U (5-(carboxyhydroxymethyl)uridine methyl ester); mcm5U (5-methoxycarbonylmethyluridine) ; mcm5Um (5-methoxycarbonylmethyl-2’ -O-methyluridine) ; mcm5s2U (5-methoxycarbonylmethyl-2-thiouridine); nm5S2U (5-aminomethyl-2-thiouridine); mnm5U (5 -methylaminomethyluridine); mnm5s2U (5-methylaminomethyl-2-thiouridine); mnm5se2U (5- methylaminomethyl-2-selenouridine); ncm5U (5 -carbamoylmethyluridine); ncm5Um (5- carbamoylmethyl-2'-O-methyluridine); cmnm5U (5-carboxymethylaminomethyluridine); cmnm5Um (5-carboxymethylaminomethyl-2'-O-methyluridine); cmnm5s2U (5-carboxymethylaminomethyl-2- thiouridine); m6 2A (N6,N6-dimethyladenosine); Im (2’-O-methylinosine); m4C (N4-methylcytidine); m4Cm (N4,2’-O-dimethylcytidine); hm5C (5 -hydroxymethylcytidine); m3U (3 -methyluridine); cm5U (5-carboxymethyluridine); m6Am (N6,2’-O-dimethyladenosine); m6 2Am (N6,N6,O-2’- trimethyladenosine); m2,7G (N2,7-dimethylguanosine); m2,2,7G (N2,N2,7-trimethylguanosine); m3Um (3,2’-O-dimethyluridine); m5D (5 -methyldihydrouridine); FCm (5-formyl-2’-O-methylcytidine); m’Gm (l,2’-O-dimethylguanosine); m’Am (l,2’-O-dimethyladenosine); rm 5U (5- taurinomethyluridine); rm5s2U (5-taurinomethyl-2-thiouridine)); imG-14 (4-demethylwyosine); imG2 (isowyosine); or ac6A (N6-acetyladenosine).
[0121] In some embodiments, the modified nucleoside may include a compound selected from: pyridin-4-one ribonucleoside, 5-aza-uridine, 2-thio-5-aza-uridine, 2-thiouridine, 4-thio-pseudouridine, 2-thio-pseudouridine, 5-hydroxyuridine, 3 -methyluridine, 5-carboxymethyl-uridine, 1 -carboxymethylpseudouridine, 5-propynyl-uridine, 1-propynyl-pseudouridine, 5-taurinomethyluridine, 1- taurinomethyl-pseudouridine, 5-taurinomethyl-2-thio-uridine, l-taurinomethyl-4-thio-uridine, 5- methyl-uridine , 1 -methyl-pseudouridine , 4-thio- 1 -methyl-pseudouridine , 2-thio- 1 -methyl- pseudouridine, 1 -methyl- 1 -deaza-pseudouridine, 2-thio- 1 -methyl- 1 -deaza-pseudouridine, dihydrouridine, dihydropseudouridine, 2-thio-dihydrouridine, 2-thio-dihydropseudouridine, 2- methoxyuridine, 2-methoxy-4-thio-uridine, 4-methoxy-pseudouridine, 4-m ethoxy-2-thio- pseudouridine, 5-aza-cytidine, pseudoisocytidine, 3-methyl-cytidine, N4-acetylcytidine, 5- formylcytidine, N4-methylcytidine, 5-hydroxymethylcytidine, 1 -methyl-pseudoisocytidine, pyrrolo- cytidine, pyrrolo-pseudoisocytidine, 2-thio-cytidine, 2-thio-5-methyl-cytidine, 4-thio- pseudoisocytidine, 4-thio- 1 -methyl-pseudoisocytidine, 4-thio- 1 -methyl- 1 -deaza-pseudoisocytidine, 1 - methyl- 1-deaza-pseudoisocytidine, zebularine, 5-aza-zebularine, 5-methyl-zebularine, 5-aza-2-thio- zebularine, 2-thio-zebularine, 2-methoxy-cytidine, 2-methoxy-5-methyl-cytidine, 4-methoxy- pseudoisocytidine, 4-methoxy-l -methyl-pseudoisocytidine, 2-aminopurine, 2, 6-diaminopurine, 7- deaza-adenine, 7-deaza-8-aza-adenine, 7-deaza-2-aminopurine, 7-deaza-8-aza-2-aminopurine, 7- deaza-2, 6-diaminopurine, 7-deaza-8-aza-2, 6-diaminopurine, 1 -methyladenosine, N6-methyladenosine, N6-isopentenyladenosine, N6-(cis-hydroxyisopentenyl)adenosine, 2-methylthio-N6-(cis- hydroxyisopentenyl) adenosine, N6-glycinylcarbamoyladenosine, N6-threonylcarbamoyladenosine, 2- methylthio-N6-threonyl carbamoyladenosine, N6,N6-dimethyladenosine, 7-methyladenine, 2-
methylthio-adenine, 2-methoxy-adenine, inosine, 1-methyl-inosine, wyosine, wybutosine, 7-deaza- guanosine, 7-deaza-8-aza-guanosine, 6-thio-guanosine, 6-thio-7-deaza-guanosine, 6-thio-7-deaza-8- aza-guanosine, 7-methyl-guanosine, 6-thio-7-methyl-guanosine, 7-methylinosine, 6-methoxy- guanosine, 1 -methylguanosine, N2-methylguanosine, N2,N2-dimethylguanosine, 8-oxo-guanosine, 7- methyl- 8 -oxo-guanosine, l-methyl-6-thio-guanosine, N2-methyl-6-thio-guanosine, and N2,N2- dimethyl-6-thio-guanosine. In another embodiment, the modifications are independently selected from 5-methylcytosine, pseudouridine and 1 -methylpseudouridine.
[0122] In some embodiments, the modified ribonucleosides include 5 -methylcytidine, 5- methoxyuridine, 1-methyl-pseudouridine, N6-methyladenosine, and/or pseudouridine. In some embodiments, such modified nucleosides provide additional stability and resistance to immune activation.
[0123] In particular embodiments, polynucleotides may be codon-optimized. A codon optimized sequence may be one in which codons in a polynucleotide encoding a polypeptide have been substituted in order to increase the expression, stability and/or activity of the polypeptide. Factors that influence codon optimization include, but are not limited to one or more of: (i) variation of codon biases between two or more organisms or genes or synthetically constructed bias tables, (ii) variation in the degree of codon bias within an organism, gene, or set of genes, (iii) systematic variation of codons including context, (iv) variation of codons according to their decoding tRNAs, (v) variation of codons according to GC %, either overall or in one position of the triplet, (vi) variation in degree of similarity to a reference sequence for example a naturally occurring sequence, (vii) variation in the codon frequency cutoff, (viii) structural properties of mRNAs transcribed from the DNA sequence, (ix) prior knowledge the function of the DNA sequences upon which design of the codon substitution set is to be based, and/or (x) systematic variation of codon sets for each amino acid. In some embodiments, a codon optimized polynucleotide may minimize ribozyme collisions and/or limit structural interference between the expression sequence and the core functional element. ix. Payloads
[0124] In some embodiments, the polynucleotide (e.g., circRNA) expression sequence encodes a therapeutic protein. In some embodiments, the therapeutic protein is selected from the proteins listed in Table 1.
[0125] In some embodiments, the expression sequence encodes a therapeutic protein. In some embodiments, the expression sequence encodes a cytokine, e.g., IL-12p70, IL-15, IL-2, IL-18, IL-21, IFN-a, IFN- P, IL-10, TGF-beta, IL-4, or IL-35, or a functional fragment thereof. In some embodiments, the expression sequence encodes an immune checkpoint inhibitor. In some embodiments, the expression sequence encodes an agonist (e.g., a TNFR family member such as CD137L, OX40L,
ICOSL, LIGHT, or CD70). In some embodiments, the expression sequence encodes a chimeric antigen receptor. In some embodiments, the expression sequence encodes an inhibitory receptor agonist (e.g., PDL1, PDL2, Galectin-9, VISTA, B7H4, or MHCII) or inhibitory receptor (e.g., PD1, CTLA4, TIGIT, LAG3, or TIM3). In some embodiments, the expression sequence encodes an inhibitory receptor antagonist. In some embodiments, the expression sequence encodes one or more TCR chains (alpha and beta chains or gamma and delta chains). In some embodiments, the expression sequence encodes a secreted T cell or immune cell engager (e.g., a bispecific antibody such as BiTE, targeting, e.g., CD3, CD137, or CD28 and a tumor-expressed protein e.g., CD19, CD20, or BCMA etc.). In some embodiments, the expression sequence encodes a transcription factor (e.g., FOXP3, HELIOS, TOX1, or
T0X2). In some embodiments, the expression sequence encodes an immunosuppressive enzyme (e.g., IDO or CD39/CD73). In some embodiments, the expression sequence encodes a GvHD (e.g., anti-HLA- A2 CAR-Tregs).
[0126] In some embodiments, the precursor RNA polynucleotide and circular RNA constructs comprise at least one expression sequence encoding an antigen, adjuvant, or adjuvant-like protein, e.g., from an infectious agent. In these embodiments, the circular RNA construct may be used as a vaccine. In some embodiments, the one or more circular RNA polynucleotide encodes an antigen or adjuvant derived from an infectious agent. In some embodiments the infectious agent from which the antigen or adjuvant is derived or engineered includes, but is not limited to a virus, bacterium, fungus, protozoan, and/or parasite. In some embodiments, the antigen is a viral antigen or viral antigenic polypeptide.
[0127] In an embodiment, the antigen is selected from or derived from the group consisting of rotavirus, foot and mouth disease virus, influenza A virus, influenza B virus, influenza C virus, H1N1, H2N2, H3N2, H5N1, H7N7, H1N2, H9N2, H7N2, H7N3, H10N7, human parainfluenza type 2, herpes simplex virus, Epstein-Barr virus, varicella virus, porcine herpesvirus 1, cytomegalovirus, lyssavirus, Bacillus anthracis, anthrax PA and derivatives, poliovirus, Hepatitis A, Hepatitis B, Hepatitis C, Hepatitis E, distemper virus, Venezuelan equine encephalomyelitis, feline leukemia virus, reovirus, respiratory syncytial virus, Lassa fever virus, polyoma tumor virus, canine parvovirus, papilloma virus, tick borne encephalitis virus, rinderpest virus, human rhinovirus species, Enterovirus species, Mengo virus, paramyxovirus, avian infectious bronchitis virus, human T-cell leukemia-lymphoma virus 1, human immunodeficiency virus- 1, human immunodeficiency virus-2, lymphocytic choriomeningitis virus, parvovirus B19, adenovirus, rubella virus, yellow fever virus, dengue virus, bovine respiratory syncitial virus, corona virus, Bordetella pertussis, Bordetella bronchiseptica, Bordetella parapertussis, Brucella abortis, Brucella melitensis, Brucella suis, Brucella ovis, Brucella species, Escherichia coli, Salmonella species, Salmonella typhi, Streptococci, Vibrio cholera, Vibrio parahaemolyticus, Shigella, Pseudomonas, tuberculosis, avium, Bacille Calmette Guerin, Mycobacterium leprae, Pneumococci, Staphlylococci, Enterobacter species, Rochalimaia henselae, Pasteurella haemolytica, Pasteurella multocida, Chlamydia trachomatis, Chlamydia psittaci, Lymphogranuloma venereum, Treponema pallidum, Haemophilus species, Mycoplasma bovigenitalium, Mycoplasma pulmonis, Mycoplasma species, Borrelia burgdorferi, Legionalla pneumophila, Colstridium botulinum, Corynebacterium diphtheriae, Yersinia entercolitica, Rickettsia rickettsii, Rickettsia typhi, Rickettsia prowsaekii, Ehrlichia chaffeensis, Anaplasma phagocytophilum, Plasmodium falciparum, Plasmodium vivax, Plasmodium malariae, Schistosomes, trypanosomes, Leishmania species, Filarial nematodes, trichomoniasis, sarcosporidiasis, Taenia saginata, Taenia solium, Leishmania, Toxoplasma gondii, Trichinella spiralis, coccidiosis, Eimeria tenella, Cryptococcus neoformans, Candida albican, Aspergillus fumigatus, coccidioidomycosis, Neisseria gonorrhoeae, malaria circumsporozoite protein,
malaria merozoite protein, trypanosome surface antigen protein, pertussis, alphaviruses, adenovirus, diphtheria toxoid, tetanus toxoid, meningococcal outer membrane protein, streptococcal M protein, Influenza hemagglutinin, cancer antigen, tumor antigens, toxins, Clostridium perfringens epsilon toxin, ricin toxin, pseudomonas exotoxin, exotoxins, neurotoxins, cytokines, cytokine receptors, monokines, monokine receptors, plant pollens, animal dander, and dust mites.
[0128] In some embodiments, the antigenic polypeptide is a viral polypeptide from an adenovirus; Herpes simplex, type 1; Herpes simplex, type 2; encephalitis virus; papillomavirus; Varicella-zoster virus; Epstein-barr virus; Human cytomegalovirus; Human herpes virus, type 8; Human papillomavirus; BK virus; JC virus; Smallpox; polio virus; Hepatitis B virus; Human bocavirus; Parvovirus B 19; Human astro virus; Norwalk virus; coxsackievirus; hepatitis A virus; poliovirus; rhinovirus; Severe acute respiratory syndrome virus; Hepatitis C virus; Yellow Fever virus; Dengue virus; West Nile virus; Rubella virus; Hepatitis E virus; Human Immunodeficiency virus (HIV); Influenza virus; Guanarito virus; Junin virus; Lassa virus; Machupo virus; Sabia virus; Crimean-Congo hemorrhagic fever virus; Ebola virus; Marburg virus; Measles virus; Mumps virus; Parainfluenza virus; Respiratory syncytial virus; Human metapneumo virus; Hendra virus; Nipah virus; Rabies virus; Hepatitis D; Rotavirus; Orbivirus; Coltivirus; Banna virus; Human Enterovirus; Hanta virus; West Nile virus; Middle East Respiratory Syndrome Corona Virus; Japanese encephalitis virus; Vesicular exanthernavirus; SARS- CoV-2; Eastern equine encephalitis, or a combination of any two or more of the foregoing.
[0129] In some embodiments, the adjuvant is selected from or derived from the group consisting of BCSP31, MOMP, FomA, MymA, ESAT6, PorB, PVL, Porin, OmpA, PepO, OmpU, Lumazine synthase, 0mpl6, 0mpl9, CobT, RpfE, Rv0652, HBHA, NhhA, DnaJ, Pneumolysin, Falgellin, IFN- alpha, IFN-gamma, IL-2, IL-12, IL-15, IL-18, IL-21, GM-CSF, IL-lb, IL-6, TNF-a, IL-7, IL-17, IL- IBeta, anti-CTLA4, anti-PDl, anti-41BB, PD-L1, Tim-3, Lag-3, TIGIT, GITR, and andti-CD3.
[0130] In some embodiments, a polynucleotide encodes a protein that is made up of subunits that are encoded by more than one gene. For example, the protein may be a heterodimer, wherein each chain or subunit of the protein is encoded by a separate gene. It is possible that more than one circRNA molecule is delivered in the transfer vehicle and each circRNA encodes a separate subunit of the protein. Alternatively, a single circRNA may be engineered to encode more than one subunit. In certain embodiments, separate circRNA molecules encoding the individual subunits may be administered in separate transfer vehicles.
[0131] Additional polynucleotides, including expression sequences, and lipids are in WO2019236673; WO2020237227; WO2021113777; WO2021226597; WO2021189059; WO2021236855;
W 02022261490; W02023056033; WO2023081526; the contents of which are hereby incorporated by reference in their entireties.
(1) Chimeric Antigen Receptors ( CARS)
[0132] Chimeric antigen receptors (CARs or CAR-Ts) are genetically-engineered receptors. These engineered receptors may be inserted into and expressed by immune cells, including T cells via circular RNA as described herein. With a CAR, a single receptor may be programmed to both recognize a specific antigen and, when bound to that antigen, activate the immune cell to attack and destroy the cell bearing that antigen. When these antigens exist on tumor cells, an immune cell that expresses the CAR may target and kill the tumor cell. In some embodiments, the CAR encoded by the polynucleotide comprises (i) an antigen-binding molecule that specifically binds to a target antigen, (ii) a hinge domain, a transmembrane domain, and an intracellular domain, and (iii) an activating domain.
[0133] In some embodiments, an orientation of the CARs in accordance with the disclosure comprises an antigen binding domain (such as an scFv) in tandem with a costimulatory domain and an activating domain. The costimulatory domain may comprise one or more of an extracellular portion, a transmembrane portion, and an intracellular portion. In other embodiments, multiple costimulatory domains may be utilized in tandem.
[0134] CARs may be engineered to bind to an antigen (such as a cell-surface antigen) by incorporating an antigen binding molecule that interacts with that targeted antigen. In some embodiments, the antigen binding molecule is an antibody fragment thereof, e.g., one or more single chain antibody fragment (scFv). An scFv is a single chain antibody fragment having the variable regions of the heavy and light chains of an antibody linked together. See U.S. Patent Nos. 7,741,465, and 6,319,494 as well as Eshhar et al., Cancer Immunol Immunotherapy (1997) 45: 131-136. An scFv retains the parent antibody's ability to specifically interact with target antigen. scFvs are useful in chimeric antigen receptors because they may be engineered to be expressed as part of a single chain along with the other CAR components. Id. See also Krause et al., J. Exp. Med., Volume 188, No. 4, 1998 (619-626); Finney et al., Journal of Immunology, 1998, 161 : 2791-2797. It will be appreciated that the antigen binding molecule is typically contained within the extracellular portion of the CAR such that it is capable of recognizing and binding to the antigen of interest. Bispecific and multispecific CARs are contemplated within the scope of the present disclosure, with specificity to more than one target of interest.
[0135] In some embodiments, the antigen binding molecule comprises a single chain, wherein the heavy chain variable region and the light chain variable region are connected by a linker. In some embodiments, the VH is located at the N terminus of the linker and the VL is located at the C terminus of the linker. In other embodiments, the VL is located at the N terminus of the linker and the VH is located at the C terminus of the linker. In some embodiments, the linker comprises at least 5, at least 8, at least 10, at least 13, at least 15, at least 18, at least 20, at least 25, at least 30, at least 35, at least 40, at least 45, at least 50, at least 60, at least 70, at least 80, at least 90, or at least 100 amino acids.
[0136] In some embodiments, the antigen binding molecule comprises a nanobody. In some embodiments, the antigen binding molecule comprises a DARPin. In some embodiments, the antigen binding molecule comprises an anticalin or other synthetic protein capable of specific binding to target protein.
[0137] In some embodiments, the CAR comprises an antigen binding domain specific for an antigen selected from CD19, CD123, CD22, CD30, CD171, CS-1, C-type lectin-like molecule-1, CD33, epidermal growth factor receptor variant III (EGFRvIII), ganglioside G2 (GD2), ganglioside GD3, TNF receptor family member B cell maturation (BCMA), Tn antigen ((Tn Ag) or (GalNAca-Ser/Thr)), prostate-specific membrane antigen (PSMA), Receptor tyrosine kinase-like orphan receptor 1 (ROR1), Fms-Like Tyrosine Kinase 3 (FLT3), Tumor-associated glycoprotein 72 (TAG72), CD38, CD44v6, Carcinoembryonic antigen (CEA), Epithelial cell adhesion molecule (EPCAM), B7H3 (CD276), KIT (CD117), Interleukin- 13 receptor subunit alpha-2, mesothelin, Interleukin 11 receptor alpha (IL-1 IRa), prostate stem cell antigen (PSCA), Protease Serine 21, vascular endothelial growth factor receptor 2 (VEGFR2), Lewis(Y) antigen, CD24, Platelet-derived growth factor receptor beta (PDGFR-beta), Stage-specific embryonic antigen-4 (SSEA-4), CD20, Folate receptor alpha, HER2, HER3, Mucin 1, cell surface associated (MUC1), epidermal growth factor receptor (EGFR), neural cell adhesion molecule (NCAM), Prostase, prostatic acid phosphatase (PAP), elongation factor 2 mutated (ELF2M), Ephrin B2, fibroblast activation protein alpha (FAP), insulin-like growth factor 1 receptor (IGF-I receptor), carbonic anhydrase IX (CAIX), Proteasome (Prosome, Macropain) Subunit, Beta Type, 9 (LMP2), glycoprotein 100 (gplOO), oncogene fusion protein consisting of breakpoint cluster region (BCR) and Abelson murine leukemia viral oncogene homolog 1 (Abl) (bcr-abl), tyrosinase, ephrin type- A receptor 2 (EphA2), Fucosyl GM1, sialyl Lewis adhesion molecule (sLe), ganglioside GM3, transglutaminase 5 (TGS5), high molecular weight-melanoma-associated antigen (HMWMAA), o- acetyl-GD2 ganglioside (OAcGD2), Folate receptor beta, tumor endothelial marker 1 (TEM1/CD248), tumor endothelial marker 7-related (TEM7R), claudin 6 (CLDN6), thyroid stimulating hormone receptor (TSHR), G protein-coupled receptor class C group 5, member D (GPRC5D), chromosome X open reading frame 61 (CXORF61), CD97, CD179a, anaplastic lymphoma kinase (ALK), Polysialic acid, placenta-specific 1 (PLAC1), hexasaccharide portion of globoH glycoceramide (GloboH), mammary gland differentiation antigen (NY-BR-1), uroplakin 2 (UPK2), Hepatitis A virus cellular receptor 1 (HAVCR1), adrenoceptor beta 3 (ADRB3), pannexin 3 (PANX3), G protein-coupled receptor 20 (GPR20), lymphocyte antigen 6 complex, locus K 9 (LY6K), Olfactory receptor 51E2 (OR51E2), TCR Gamma Alternate Reading Frame Protein (TARP), Wilms tumor protein (WT1), Cancer/testis antigen 1 (NY-ESO-1), Cancer/testis antigen 2 (LAGE-la), MAGE family members (including MAGE-A1, MAGE- A3 and MAGE-A4), ETS translocation-variant gene 6, located on chromosome 12p (ETV6-AML), sperm protein 17 (SPA17), X Antigen Family, Member 1A (XAGE1), angiopoietin-binding cell surface receptor 2 (Tie 2), melanoma cancer testis antigen- 1 (MAD-CT-1),
melanoma cancer testis antigen-2 (MAD-CT-2), Fos-related antigen 1, tumor protein p53 (p53), p53 mutant, prostein, surviving, telomerase, prostate carcinoma tumor antigen- 1, melanoma antigen recognized by T cells 1, Rat sarcoma (Ras) mutant, human Telomerase reverse transcriptase (hTERT), sarcoma translocation breakpoints, melanoma inhibitor of apoptosis (ML-IAP), ERG (transmembrane protease, serine 2 (TMPRSS2) ETS fusion gene), N-Acetyl glucosaminyl-transferase V (NA17), paired box protein Pax-3 (PAX3), Androgen receptor, Cyclin Bl, v-myc avian myelocytomatosis viral oncogene neuroblastoma derived homolog (MYCN), Ras Homolog Family Member C (RhoC), Tyrosinase-related protein 2 (TRP-2), Cytochrome P450 1B1 (CYP1B1), CCCTC-Binding Factor (Zinc Finger Protein)-Like, Squamous Cell Carcinoma Antigen Recognized By T Cells 3 (SART3), Paired box protein Pax-5 (PAX5), proacrosin binding protein sp32 (OY-TES1), lymphocyte-specific protein tyrosine kinase (LCK), A kinase anchor protein 4 (AKAP-4), synovial sarcoma, X breakpoint 2 (SSX2), Receptor for Advanced Glycation Endproducts (RAGE-1), renal ubiquitous 1 (RU1), renal ubiquitous 2 (RU2), legumain, human papilloma virus E6 (HPV E6), human papilloma virus E7 (HPV E7), intestinal carboxyl esterase, heat shock protein 70-2 mutated (mut hsp70-2), CD79a, CD79b, CD72, Leukocyte-associated immunoglobulin-like receptor 1 (LAIR1), Fc fragment of IgA receptor (FCAR or CD89), Leukocyte immunoglobulin-like receptor subfamily A member 2 (LILRA2), CD300 molecule-like family member f (CD300LF), C-type lectin domain family 12 member A (CLEC12A), bone marrow stromal cell antigen 2 (BST2), EGF-like module-containing mucin-like hormone receptorlike 2 (EMR2), lymphocyte antigen 75 (LY75), Glypican-3 (GPC3), Fc receptor-like 5 (FCRL5), MUC16, 5T4, 8H9, avP0 integrin, avP6 integrin, alphafetoprotein (AFP), B7-H6, ca-125, CA9, CD44, CD44v7/8, CD52, E-cadherin, EMA (epithelial membrane antigen), epithelial glycoprotein-2 (EGP-2), epithelial glycoprotein-40 (EGP-40), ErbB4, epithelial tumor antigen (ETA), folate binding protein (FBP), kinase insert domain receptor (KDR), k-light chain, LI cell adhesion molecule, MUC18, NKG2D, oncofetal antigen (h5T4), tumor/testis-antigen IB, GAGE, GAGE-1, BAGE, SCP-1, CTZ9, SAGE, CAGE, CT10, MART-1, immunoglobulin lambda-like polypeptide 1 (IGLL1), Hepatitis B Surface Antigen Binding Protein (HBsAg), viral capsid antigen (VCA), early antigen (EA), EBV nuclear antigen (EBNA), HHV-6 p41 early antigen, HHV-6B U94 latent antigen, HHV-6B p98 late antigen , cytomegalovirus (CMV) antigen, large T antigen, small T antigen, adenovirus antigen, respiratory syncytial virus (RSV) antigen, haemagglutinin (HA), neuraminidase (NA), parainfluenza type 1 antigen, parainfluenza type 2 antigen, parainfluenza type 3 antigen, parainfluenza type 4 antigen, Human Metapneumovirus (HMPV) antigen, hepatitis C virus (HCV) core antigen, HIV p24 antigen, human T-cell lympotrophic virus (HTLV-1) antigen, Merkel cell polyoma virus small T antigen, Merkel cell polyoma virus large T antigen, Kaposi sarcoma-associated herpesvirus (KSHV) lytic nuclear antigen and KSHV latent nuclear antigen. In some embodiments, an antigen binding domain comprises SEQ ID NO: 321 and/or 322 of WO2023081526.
[0138] In some embodiments, a CAR comprises a hinge or spacer domain. In some embodiments, the hinge/spacer domain may comprise a truncated hinge/spacer domain (THD) the THD domain is a truncated version of a complete hinge/spacer domain (“CHD”). In some embodiments, an extracellular domain is from or derived from (e.g., comprises all or a fragment of) ErbB2, glycophorin A (GpA), CD2, CD3 delta, CD3 epsilon, CD3 gamma, CD4, CD7, CD8a, CD8[T CD1 la (IT GAL), CD1 lb (IT GAM), CD1 1c (ITGAX), CD1 Id (IT GAD), CD18 (ITGB2), CD19 (B4), CD27 (TNFRSF7), CD28, CD28T, CD29 (ITGB1), CD30 (TNFRSF8), CD40 (TNFRSF5), CD48 (SLAMF2), CD49a (ITGA1), CD49d (ITGA4), CD49f (ITGA6), CD66a (CEACAM1), CD66b (CEACAM8), CD66c (CEACAM6), CD66d (CEACAM3), CD66e (CEACAM5), CD69 (CLEC2), CD79A (B-cell antigen receptor complex-associated alpha chain), CD79B (B-cell antigen receptor complex-associated beta chain), CD84 (SLAMF5), CD96 (Tactile), CD100 (SEMA4D), CD103 (ITGAE), CD134 (0X40), CD137 (4- 1BB), CD150 (SLAMF1), CD158A (KIR2DL1), CD158B1 (KIR2DL2), CD158B2 (KIR2DL3), CD158C (KIR3DP1), CD158D (KIRDL4), CD158F1 (KIR2DL5A), CD158F2 (KIR2DL5B), CD158K (KIR3DL2), CD160 (BY55), CD162 (SELPLG), CD226 (DNAM1), CD229 (SLAMF3), CD244 (SLAMF4), CD247 (CD3-zeta), CD258 (LIGHT), CD268 (BAFFR), CD270 (TNFSF14), CD272 (BTLA), CD276 (B7-H3), CD279 (PD-1), CD314 (NKG2D), CD319 (SLAMF7), CD335 (NK-p46), CD336 (NK-p44), CD337 (NK-p30), CD352 (SLAMF6), CD353 (SLAMF8), CD355 (CRT AM), CD357 (TNFRSF18), inducible T cell co-stimulator (ICOS), LFA-1 (CD1 la/CD18), NKG2C, DAP-10, ICAM-1, NKp80 (KLRF1), IL-2R beta, IL-2R gamma, IL-7R alpha, LFA-1, SLAMF9, LAT, GADS (GrpL), SLP-76 (LCP2), PAG1/CBP, a CD83 ligand, Fc gamma receptor, MHC class 1 molecule, MHC class 2 molecule, a TNF receptor protein, an immunoglobulin protein, a cytokine receptor, an integrin, activating NK cell receptors, a Toll ligand receptor, and fragments or combinations thereof. A hinge or spacer domain may be derived either from a natural or from a synthetic source.
[0139] In some embodiments, a hinge or spacer domain is positioned between an antigen binding molecule (e.g., an scFv) and a transmembrane domain. In this orientation, the hinge/spacer domain provides distance between the antigen binding molecule and the surface of a cell membrane on which the CAR is expressed. In some embodiments, a hinge or spacer domain is from or derived from an immunoglobulin. In some embodiments, a hinge or spacer domain is selected from the hinge/spacer regions of IgGl, IgG2, IgG3, IgG4, IgA, IgD, IgE, IgM, or a fragment thereof. In some embodiments, a hinge or spacer domain comprises, is from, or is derived from the hinge/spacer region of CD8 alpha. In some embodiments, a hinge or spacer domain comprises, is from, or is derived from the hinge/spacer region of CD28. In some embodiments, a hinge or spacer domain comprises a fragment of the hinge/spacer region of CD8 alpha or a fragment of the hinge/spacer region of CD28, wherein the fragment is anything less than the whole hinge/spacer region. In some embodiments, the fragment of the CD8 alpha hinge/spacer region or the fragment of the CD28 hinge/spacer region comprises an amino acid sequence that excludes at least 1, at least 2, at least 3, at least 4, at least 5, at least 6, at least 7, at
least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 amino acids at the N-terminus or C-Terminus, or both, of the CD8 alpha hinge/spacer region, or of the CD28 hinge/spacer region.
[0140] The CAR may further comprise a transmembrane domain and/or an intracellular signaling domain. The transmembrane domain may be designed to be fused to the extracellular domain of the CAR. It may similarly be fused to the intracellular domain of the CAR. In some embodiments, the transmembrane domain that naturally is associated with one of the domains in a CAR is used. In some instances, the transmembrane domain may be selected or modified (e.g., by an amino acid substitution) to avoid binding of such domains to the transmembrane domains of the same or different surface membrane proteins to minimize interactions with other members of the receptor complex. The transmembrane domain may be derived either from a natural or from a synthetic source. Where the source is natural, the domain may be derived from any membrane-bound or transmembrane protein.
[0141] Transmembrane regions may be derived from (i.e., comprise) a receptor tyrosine kinase (e.g., ErbB2), glycophorin A (GpA), 4-1BB/CD137, activating NK cell receptors, an immunoglobulin protein, B7-H3, BAFFR, BFAME (SEAMF8), BTEA, CD100 (SEMA4D), CD103, CD160 (BY55), CD18, CD19, CD19a, CD2, CD247, CD27, CD276 (B7-H3), CD28, CD29, CD3 delta, CD3 epsilon, CD3 gamma, CD30, CD4, CD40, CD49a, CD49D, CD49f, CD69, CD7, CD84, CD8alpha, CD8beta, CD96 (Tactile), CD1 la, CD1 lb, CD1 1c, CD1 Id, CDS, CEACAM1, CRT AM, cytokine receptor, DAP-10, DNAM1 (CD226), Fc gamma receptor, GADS, GITR, HVEM (EIGHTR), IA4, ICAM-1, ICAM-1, Ig alpha (CD79a), IE-2R beta, IE-2R gamma, IE-7R alpha, inducible T cell costimulator (ICOS), integrins, ITGA4, ITGA4, ITGA6, IT GAD, ITGAE, ITGAE, IT GAM, ITGAX, ITGB2, ITGB7, ITGB1, KIRDS2, EAT, LFA-1, LFA-1, a ligand that specifically binds with CD83, LIGHT, LIGHT, LTBR, Ly9 (CD229), lymphocyte function-associated antigen-1 (LFA-1; CDl-la/CD18), MHC class 1 molecule, NKG2C, NKG2D, NKp30, NKp44, NKp46, NKp80 (KLRF1), OX-40, PAG/Cbp, programmed death- 1 (PD-1), PSGL1, SELPLG (CD 162), Signaling Lymphocytic Activation Molecules (SLAM proteins), SLAM (SLAMF1; CD150; IPO-3), SLAMF4 (CD244; 2B4), SLAMF6 (NTB-A; Lyl08), SLAMF7, SLP-76, TNF receptor proteins, TNFR2, TNFSF14, a Toll ligand receptor, TRANCE/RANKL, VLA1, or VLA-6, or a fragment, truncation, or a combination thereof.
[0142] In some embodiments, suitable intracellular signaling domain include, but are not limited to, activating Macrophage/Myeloid cell receptors CSFR1, MYD88, CD14, TIE2, TLR4, CR3, CD64, TREM2, DAP10, DAP12, CD169, DECTIN1, CD206, CD47, CD163, CD36, MARCO, TIM4, MERTK, F4/80, CD91, C1QR, LOX-1, CD68, SRA, BAI-1, ABCA7, CD36, CD31, Lactoferrin, or a fragment, truncation, or combination thereof.
[0143] In some embodiments, a receptor tyrosine kinase may be derived from (e.g., comprise) Insulin receptor (InsR), Insulin-like growth factor I receptor (IGF1R), Insulin receptor-related receptor (IRR), platelet derived growth factor receptor alpha (PDGFRa), platelet derived growth factor receptor beta (PDGFRfi). KIT proto-oncogene receptor tyrosine kinase (Kit), colony stimulating factor 1 receptor (CSFR), fms related tyrosine kinase 3 (FLT3), fms related tyrosine kinase 1 (VEGFR-1), kinase insert domain receptor (VEGFR-2), fms related tyrosine kinase 4 (VEGFR-3), fibroblast growth factor receptor 1 (FGFR1), fibroblast growth factor receptor 2 (FGFR2), fibroblast growth factor receptor 3 (FGFR3), fibroblast growth factor receptor 4 (FGFR4), protein tyrosine kinase 7 (CCK4), neurotrophic receptor tyrosine kinase 1 (trkA), neurotrophic receptor tyrosine kinase 2 (trkB), neurotrophic receptor tyrosine kinase 3 (trkC), receptor tyrosine kinase like orphan receptor 1 (ROR1), receptor tyrosine kinase like orphan receptor 2 (ROR2), muscle associated receptor tyrosine kinase (MuSK), MET protooncogene, receptor tyrosine kinase (MET), macrophage stimulating 1 receptor (Ron), AXL receptor tyrosine kinase (Axl), TYR03 protein tyrosine kinase (Tyro3), MER proto-oncogene, tyrosine kinase (Mer), tyrosine kinase with immunoglobulin like and EGF like domains 1 (TIE1), TEK receptor tyrosine kinase (TIE2), EPH receptor Al (EphAl), EPH receptor A2 (EphA2), (EPH receptor A3) EphA3, EPH receptor A4 (EphA4), EPH receptor A5 (EphA5), EPH receptor A6 (EphA6), EPH receptor A7 (EphA7), EPH receptor A8 (EphA8), EPH receptor A10 (EphAlO), EPH receptor Bl (EphBl), EPH receptor B2 (EphB2), EPH receptor B3 (EphB3), EPH receptor B4 (EphB4), EPH receptor B6 (EphB6), ret proto oncogene (Ret), receptor-like tyrosine kinase (RYK), discoidin domain receptor tyrosine kinase 1 (DDR1), discoidin domain receptor tyrosine kinase 2 (DDR2), c-ros oncogene 1, receptor tyrosine kinase (ROS), apoptosis associated tyrosine kinase (Lmrl), lemur tyrosine kinase 2 (Lmr2), lemur tyrosine kinase 3 (Lmr3), leukocyte receptor tyrosine kinase (LTK), ALK receptor tyrosine kinase (ALK), or serine/threonine/tyrosine kinase 1 (STYK1).
[0144] In some embodiments, the CAR comprises a costimulatory domain. In some embodiments, the costimulatory domain comprises 4-1BB (CD137), CD28, or both, and/or an intracellular T cell signaling domain. In one embodiment, the costimulatory domain is human CD28, human 4- IBB, or both, and the intracellular T cell signaling domain is human CD3 zeta (Q. 4-1BB, CD28, CD3 zeta may comprise less than the whole 4-1BB, CD28 or CD3 zeta, respectively. Chimeric antigen receptors may incorporate costimulatory (signaling) domains to increase their potency. See U.S. Patent Nos. 7,741,465, and 6,319,494, as well as Krause etal. and Finney et al. (supra), Song et al., Blood 119:696- 706 (2012); Kalos etal., Sci Transl. Med. 3:95 (2011); Porter etal., N. Engl. J. Med. 365:725-33 (2011), and Gross et al., Amur. Rev. Pharmacol. Toxicol. 56:59-83 (2016).
[0145] In some embodiments, a costimulatory domain comprises the amino acid sequence of SEQ ID NO: 318 or 320 of WO2023081526.
[0146] The intracellular (signaling) domain of the engineered T cells disclosed herein may provide signaling to an activating domain, which then activates at least one of the normal effector functions of the immune cell. Effector function of a T cell, for example, may be cytolytic activity or helper activity including the secretion of cytokines.
[0147] In some embodiments, suitable intracellular signaling domain include (e.g., comprise), but are not limited to 4-1BB/CD137, activating NK cell receptors, an Immunoglobulin protein, B7-H3, BAFFR, BEAME (SEAMF8), BTLA, CD100 (SEMA4D), CD103, CD160 (BY55), CD18, CD19, CD 19a, CD2, CD247, CD27, CD276 (B7-H3), CD28, CD29, CD3 delta, CD3 epsilon, CD3 gamma, CD30, CD4, CD40, CD49a, CD49D, CD49f, CD69, CD7, CD84, CD8alpha, CD8beta, CD96 (Tactile), CD1 la, CD1 lb, CD1 1c, CD1 Id, CDS, CEACAM1, CRT AM, cytokine receptor, DAP-10, DNAM1 (CD226), Fc gamma receptor, GADS, GITR, HVEM (LIGHTR), IA4, ICAM-1, Ig alpha (CD79a), IL- 2R beta, IL-2R gamma, IL-7R alpha, inducible T cell costimulator (ICOS), integrins, ITGA4, ITGA6, ITGAD, ITGAE, ITGAL, ITGAM, ITGAX, ITGB2, ITGB7, ITGB1, KIRDS2, LAT, LFA-1, ligand that specifically binds with CD83, LIGHT, LTBR, Ly9 (CD229), Lyl08, lymphocyte function- associated antigen- 1 (LFA-1; CDl-la/CD18), MHC class 1 molecule, NKG2C, NKG2D, NKp30, NKp44, NKp46, NKp80 (KLRF1), OX-40, PAG/Cbp, programmed death-1 (PD-1), PSGL1, SELPLG (CD162), Signaling Lymphocytic Activation Molecules (SLAM proteins), SLAM (SLAMF1; CD150; IPO-3), SLAMF4 (CD244; 2B4), SLAMF6 (NTB-A), SLAMF7, SLP-76, TNF receptor proteins, TNFR2, TNFSF14, a Toll ligand receptor, TRANCE/RANKL, VLA1, or VLA-6, or a fragment, truncation, or a combination thereof.
[0148] CD3 is an element of the T cell receptor on native T cells, and has been shown to be an important intracellular activating element in CARs. In some embodiments, the CD3 is CD3 zeta. In some embodiments, the activating domain comprises an amino acid sequence of at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to the polypeptide sequence of SEQ ID NO: 319 of WO2023081526.
[0149] In some embodiments, the sequence encoding the CAR comprises a sequence from Table 2.
(2) T-Cell Receptors (TCR)
[0150] TCRs are described using the International Immunogenetics (IMGT) TCR nomenclature, and links to the IMGT public database of TCR sequences. Native alpha-beta heterodimeric TCRs have an alpha chain and a beta chain. Broadly, each chain may comprise variable, joining and constant regions, and the beta chain also usually contains a short diversity region between the variable and joining regions, but this diversity region is often considered as part of the joining region. Each variable region may comprise three CDRs (Complementarity Determining Regions) embedded in a framework sequence, one being the hypervariable region named CDR3. There are several types of alpha chain variable (Va) regions and several types of beta chain variable (VP) regions distinguished by their
framework, CDR1 and CDR2 sequences, and by a partly defined CDR3 sequence. The Va types are referred to in IMGT nomenclature by a unique TRAV number. Thus “TRAV21” defines a TCR Va region having unique framework and CDR1 and CDR2 sequences, and a CDR3 sequence which is partly defined by an amino acid sequence which is preserved from TCR to TCR but which also includes an amino acid sequence which varies from TCR to TCR. In the same way, “TRBV5-1” defines a TCR VP region having unique framework and CDR1 and CDR2 sequences, but with only a partly defined CDR3 sequence.
[0151] The joining regions of the TCR are similarly defined by the unique IMGT TRAJ and TRBJ nomenclature, and the constant regions by the IMGT TRAC and TRBC nomenclature.
[0152] The beta chain diversity region is referred to in IMGT nomenclature by the abbreviation TRBD, and, as mentioned, the concatenated TRBD/TRBJ regions are often considered together as the joining region.
[0153] The unique sequences defined by the IMGT nomenclature are widely known and accessible to those working in the TCR field. For example, they can be found in the IMGT public database. The “T cell Receptor Factsbook,” (2001) LeFranc and LeFranc, Academic Press, ISBN 0-12-441352-8 also discloses sequences defined by the IMGT nomenclature, but because of its publication date and consequent time-lag, the information therein sometimes needs to be confirmed by reference to the IMGT database.
[0154] Native TCRs exist in heterodimeric aP or y5 forms. However, recombinant TCRs consisting of aa or PP homodimers have previously been shown to bind to peptide MHC molecules. Therefore, the TCR of the present disclosure may be a heterodimeric aP TCR or may be an aa or PP homodimeric TCR.
[0155] For use in adoptive therapy, an aP heterodimeric TCR may, for example, be transfected as full length chains having both cytoplasmic and transmembrane domains. In certain embodiments TCRs of the present disclosure may have an introduced disulfide bond between residues of the respective constant domains, as described, for example, in WO 2006/000830.
[0156] TCRs of the present disclosure, particularly alpha-beta heterodimeric TCRs, may comprise an alpha chain TRAC constant domain sequence and/or a beta chain TRBC1 or TRBC2 constant domain sequence. The alpha and beta chain constant domain sequences may be modified by truncation or substitution to delete the native disulfide bond between Cys4 of exon 2 of TRAC and Cys2 of exon 2 of TRBC 1 or TRBC2. The alpha and/or beta chain constant domain sequence(s) may also be modified by substitution of cysteine residues for Thr 48 of TRAC and Ser 57 of TRBC1 or TRBC2, the said cysteines forming a disulfide bond between the alpha and beta constant domains of the TCR.
[0157] Binding affinity (inversely proportional to the equilibrium constant KD) and binding half-life (expressed as P/2) can be determined by any appropriate method. It will be appreciated that doubling the affinity of a TCR results in halving the KD- P/2 is calculated as In 2 divided by the off-rate (koff). So doubling of P/2 results in a halving in koff. KD and koff values for TCRs are usually measured for soluble forms of the TCR, i.e., those forms which are truncated to remove cytoplasmic and transmembrane domain residues. Therefore, it is to be understood that a given TCR has an improved binding affinity for, and/or a binding half-life for the parental TCR if a soluble form of that TCR has the said characteristics. Preferably the binding affinity or binding half-life of a given TCR is measured several times, for example 3 or more times, using the same assay protocol, and an average of the results is taken.
[0158] Since the TCRs of the present disclosure have utility in adoptive therapy, the present disclosure includes a non-naturally occurring and/or purified and/or or engineered cell, especially a T-cell, presenting a TCR of the present disclosure. There are a number of methods suitable for the transfection of T cells with nucleic acid (such as DNA, cDNA or RNA) encoding the TCRs of the present disclosure (see for example Robbins et al., (2008) J Immunol. 180: 6116-6131). T cells expressing the TCRs of the present disclosure will be suitable for use in adoptive therapy-based treatment of cancers such as those of the pancreas and liver. As will be known to those skilled in the art, there are a number of suitable methods by which adoptive therapy can be carried out (see for example Rosenberg et al. , (2008) Nat Rev Cancer 8(4): 299-308).
[0159] As is well-known in the art TCRs of the present disclosure may be subject to post-translational modifications when expressed by transfected cells. Glycosylation is one such modification, which may comprise the covalent attachment of oligosaccharide moieties to defined amino acids in the TCR chain. For example, asparagine residues, or serine/threonine residues are well-known locations for oligosaccharide attachment. The glycosylation status of a particular protein depends on a number of factors, including protein sequence, protein conformation and the availability of certain enzymes. Furthermore, glycosylation status (i.e., oligosaccharide type, covalent linkage and total number of attachments) can influence protein function. Therefore, when producing recombinant proteins, controlling glycosylation is often desirable. Glycosylation of transfected TCRs may be controlled by mutations of the transfected gene (Kuball J et al. (2009), J Exp Med 206(2):463-475). Such mutations are also encompassed in this disclosure.
[0160] A TCR may be specific for an antigen in the group MAGE-A1, MAGE-A2, MAGE- A3, MAGE-A4, MAGE-A5, MAGE-A6, MAGE-A7, MAGE-A8, MAGE-A9, MAGE-A10, MAGE-A11, MAGE-A12, MAGE-A13, GAGE-1, GAGE-2, GAGE-3, GAGE-4, GAGE-5, GAGE-6, GAGE-7, GAGE-8, BAGE-1, RAGE-1, LB33/MUM-1, PRAME, NAG, MAGE-Xp2 (MAGE-B2), MAGE-Xp3 (MAGE-B3), MAGE-Xp4 (AGE-B4), tyrosinase, brain glycogen phosphorylase, Melan-A, MAGE-CI,
MAGE-C2, NY-ESO-1, LAGE-1, SSX-1, SSX-2(HOM-MEL-40), SSX-1, SSX-4, SSX-5, SCP-1, CT- 7, alpha-actinin-4, Bcr-Abl fusion protein, Casp-8, beta-catenin, cdc27, cdk4, cdkn2a, coa-1, dek-can fusion protein, EF2, ETV6-AML1 fusion protein, LDLR-fucosyltransferaseAS fusion protein, HLA- A2, HLA-A11, hsp70-2, KIAAO205, Mart2, Mum-2, and 3, neo-PAP, myosin class I, OS-9, pml-RARa fusion protein, PTPRK, K-ras, N-ras, Triosephosphate isomeras, GnTV, Herv-K-mel, Lage-1, Mage- 02, NA-88, Lage-2, SP17, and TRP2-Int2, (MART-I), gplOO (Pmel 17), TRP-1, TRP-2, MAGE-1, MAGE-3, pl5(58), CEA, NY-ESO (LAGE), SCP-1, Hom/Mel-40, p53, H-Ras, HER-2/neu, BCR- ABL, E2A-PRL, H4-RET, IGH-IGK, MYL-RAR, Epstein Barr virus antigens, EBNA, human papillomavirus (HPV) antigens E6 and E7, TSP-180, MAGE-4, MAGE-5, MAGE-6, pl85erbB2, pl80erbB-3, c-met, nm-23Hl, PSA, TAG-72-4, CA 19-9, CA 72-4, CAM 17.1, NuMa, K-ras, beta- catenin, CDK4, Mum-1, pl6, TAGE, PSMA, PSCA, CT7, telomerase, 43-9F, 5T4, 791Tgp72, a- fetoprotein, 13HCG, BCA225, BTAA, CA 125, CA 15-3 (CA 27.29\BCAA), CA 195, CA 242, CA- 50, CAM43, CD68\KP1, CO-029, FGF-5, G250, Ga733 (EpCAM), HTgp-175, M344, MA-50, MG7- Ag, M0V18, NBM70K, NY-CO-1, RCAS1, SDCCAG16, TA-90 (Mac-2 binding protein\cyclophilin C-associated protein), TAAL6, TAG72, TLP, and TPS.
(3) B-Cell Receptors (BCR)
[0161] B-cell receptors (BCRs) or B-cell antigen receptors are immunoglobulin molecules that form a type I transmembrane protein on the surface of a B cell. A BCR is capable of transmitting activatory signal into a B cell following recognition of a specific antigen. Prior to binding of a B cell to an antigen, the BCR will remain in an unstimulated or “resting” stage. Binding of an antigen to a BCR leads to signaling that initiates a humoral immune response.
[0162] A BCR is expressed by mature B cells. These B cells work with immunoglobulins (Igs) in recognizing and tagging pathogens. The typical BCR comprises a membrane-bound immunoglobulin (e.g., mlgA, mlgD, mlgE, mlgG, and mlgM), along with associated and Iga/IgP (CD79a/CD79b) heterodimers (a/p). These membrane-bound immunoglobulins are tetramers consisting of two identical heavy and two light chains. Within the BCR, the membrane bound immunoglobulin is capable of responding to antigen binding by signal transmission across the plasma membrane leading to B cell activation and consequently clonal expansion and specific antibody production (Friess M et al. (2018), Front. Immunol. 2947(9)). The Iga/IgP heterodimer is responsible for transducing signals to the cell interior.
[0163] A Iga/IgP heterodimer signaling relies on the presence of immunoreceptor tyrosine-based activation motifs (IT AMs) located on each of the cytosolic tails of the heterodimers. IT AMs comprise two tyrosine residues separated by 9-12 amino acids (e.g., tyrosine, leucine, and/or valine). Upon binding of an antigen, the tyrosine of the BCR’s IT AMs become phosphorylated by Src-family tyrosine
kinases Blk, Fyn, or Lyn (Janeway C etal., Immunobiology: The Immune System in Health and Disease (Garland Science, 5th ed. 2001)).
(4) Other Chimeric Proteins
[0164] In addition to the chimeric proteins provided above, the circular RNA polynucleotide may encode for a various number of other chimeric proteins available in the art. The chimeric proteins may include recombinant fusion proteins, chimeric mutant protein, or other fusion proteins.
[0165] In some embodiments, the circular RNA polynucleotide encodes for an immune modulatory ligand. In certain embodiments, the immune modulatory ligand may be immunostimulatory; while in other embodiments, the immune modulatory ligand may be immunosuppressive.
[0166] In some embodiments, the circular RNA polynucleotide encodes for a cytokine. In some embodiments, the cytokine comprises a chemokine, interferon, interleukin, lymphokine, and tumor necrosis factor. Chemokines are chemotactic cytokine produced by a variety of cell types in acute and chronic inflammation that mobilizes and activates white blood cells. An interferon comprises a family of secreted a-helical cytokines induced in response to specific extracellular molecules through stimulation of TLRs (Borden, Molecular Basis of Cancer (Fourth Edition) 2015). Interleukins are cytokines expressed by leukocytes.
[0167] Descriptions and/or amino acid sequences of IL-2, IL-7, IL-10, IL-12, IL-15, IL-18, IL-27P, IFNy, and/or TGFpi are provided herein and at the www.uniprot.org database at accession numbers: P60568 (IL-2), P29459 (IL-12A), P29460 (IL-12B), P13232 (IL-7), P22301 (IL-10), P40933 (IL-15), Q14116 (IL-18), Q14213 (IL-27P), P01579 (IFNy), and/or P01137 (TGFpi).
[0168] In some embodiments, the circular RNA polynucleotide may encode for a transcription factor. Regulatory T cells (Treg) are important in maintaining homeostasis, controlling the magnitude and duration of the inflammatory response, and in preventing autoimmune and allergic responses.
[0169] In general, Tregs are thought to be mainly involved in suppressing immune responses, functioning in part as a “self-check” for the immune system to prevent excessive reactions. In particular, Tregs are involved in maintaining tolerance to self-antigens, harmless agents such as pollen or food, and abrogating autoimmune disease.
[0170] Tregs are found throughout the body including, without limitation, the gut, skin, lung, and liver. Additionally, Treg cells may also be found in certain compartments of the body that are not directly exposed to the external environment such as the spleen, lymph nodes, and even adipose tissue. Each of these Treg cell populations is known or suspected to have one or more unique features and additional information may be found in Lehtimaki and Lahesmaa, Regulatory T cells control immune responses
through their non-redundant tissue specific features, 2013, FRONTIERS IN IMMUNOL., 4(294): 1- 10, the disclosure of which is hereby incorporated in its entirety.
[0171] Typically, Tregs are known to require TGF-P and IL-2 for proper activation and development. Tregs, expressing abundant amounts of the IL-2 receptor (IL-2R), are reliant on IL-2 produced by activated T cells. Tregs are known to produce both IL-10 and TGF-P, both potent immune suppressive cytokines. Additionally, Tregs are known to inhibit the ability of antigen presenting cells (APCs) to stimulate T cells. One proposed mechanism for APC inhibition is via CTLA-4, which is expressed by Foxp3+ Tregs. It is thought that CTLA-4 may bind to B7 molecules on APCs and either block these molecules or remove them by causing internalization resulting in reduced availability of B7 and an inability to provide adequate co-stimulation for immune responses. Additional discussion regarding the origin, differentiation and function of Tregs may be found in Dhamne et al. , Peripheral and thymic Foxp3+ regulatory T cells in search of origin, distinction, and function, 2013, Frontiers in Immunol., 4 (253): 1-11, the disclosure of which is hereby incorporated in its entirety.
[0172] As provided herein, in certain embodiments, the coding element of the circular RNA polynucleotide encodes for one or more checkpoint inhibitors or agonists.
[0173] In some embodiments, the immune checkpoint inhibitor is an inhibitor of Programmed Death- Ligand 1 (PD-L1, also known as B7-H1, CD274), Programmed Death 1 (PD-1), CTLA-4, PD-L2 (B7- DC, CD273), LAG3, TIM3, 2B4, A2aR, B7H1, B7H3, B7H4, BTLA, CD2, CD27, CD28, CD30, CD40, CD70, CD80, CD86, CD137, CD160, CD226, CD276, DR3, GAL9, GITR, HAVCR2, HVEM, IDO1, IDO2, ICOS (inducible T cell costimulator), KIR, LAIR1, LIGHT, MARCO (macrophage receptor with collageneous structure), PS (phosphatidylserine), OX-40, SLAM, TIGHT, VISTA, VTCN1, or any combinations thereof. In some embodiments, the immune checkpoint inhibitor is an inhibitor of IDO1, CTLA4, PD-1, LAG3, PD-L1, TIM3, or combinations thereof. In some embodiments, the immune checkpoint inhibitor is an inhibitor of PD-L1. In some embodiments, the immune checkpoint inhibitor is an inhibitor of PD-1. In some embodiments, the immune checkpoint inhibitor is an inhibitor of CTLA-4. In some embodiments, the immune checkpoint inhibitor is an inhibitor of LAG3. In some embodiments, the immune checkpoint inhibitor is an inhibitor of TIM3. In some embodiments, the immune checkpoint inhibitor is an inhibitor of IDOL
[0174] As described herein, at least in one aspect, the present disclosure encompasses the use of immune checkpoint antagonists. Such immune checkpoint antagonists include antagonists of immune checkpoint molecules such as Cytotoxic T-Lymphocyte Antigen 4 (CTLA-4), Programmed Cell Death Protein 1 (PD-1), Programmed Death-Ligand 1 (PDL-1), Lymphocyte- activation gene 3 (LAG-3), and T-cell immunoglobulin and mucin domain 3 (TIM-3). An antagonist of CTLA-4, PD-1, PDL-1, LAG-3, or TIM-3 interferes with CTLA-4, PD-1, PDL-1, LAG-3, or TIM-3 function,
respectively. Such antagonists of CTLA-4, PD-1, PDL-1, LAG-3, and TIM-3 can include antibodies which specifically bind to CTLA-4, PD-1, PDL-1, LAG-3, and TIM-3, respectively and inhibit and/or block biological activity and function.
[0175] In some embodiments, the pay load encoded within one or more of the coding elements is a hormone, FC fusion protein, anticoagulant, blood clotting factor, protein associated with deficiencies and genetic disease, a chaperone protein, an antimicrobial protein, an enzyme (e.g., metabolic enzyme), a structural protein (e.g., a channel or nuclear pore protein), protein variant, small molecule, antibody, nanobody, an engineered non-body antibody, or a combination thereof.
B. IONIZABLE LIPIDS
[0176] In certain embodiments disclosed herein are ionizable lipids. The subject ionizable lipids may be used as a component of a composition to facilitate encapsulation and release of circular RNA to one or more target cells (e.g., by forming an ion pair/complex with the circular RNA within the core of the subject polymeric lipid nanoparticle composition). In certain embodiments, an ionizable lipid comprises one or more cleavable functional groups (e.g., a disulfide) that allow, for example, a hydrophilic functional head-group to dissociate from a lipophilic functional tail-group of the compound (e.g., upon exposure to oxidative, reducing or acidic conditions).
[0177] In some embodiments, the ionizable lipid has a pKa from 6 to 12. In some embodiments, the ionizable lipid has a pKa from 7 to 9. In some embodiments, the ionizable lipid has a pKa of 6.0, 6.1, 6.2, 6.3, 6.4, 6.5, 6.6, 6.7, 6.8, 6.9, 7.0, 7.1, 7.2, 7.3, 7.4, 7.5, 7.6, 7.7, 7.8, 7.9, 8.0, 8.1, 8.2, 8.3, 8.4, 8.5, 8.6, 8.7, 8.8, 8.9, or 9.0 or any ranges created by these.
[0178] In some embodiments, the ionizable lipid comprises an amino group.
[0179] In some embodiments, the ionizable lipid comprises a divalent headgroup and one or more straight hydrocarbon lipid tails. In some embodiments, the straight hydrocarbon lipid tails are from 3- 25 carbon atoms in length, such as 5 to 25, 5 to 20, 5 to 15, 5 to 10, 10 to 15, 10 to 20, or 10 to 25 carbon atoms in length.
[0180] In some embodiments, the ionizable lipid comprises a divalent headgroup and one or more branched hydrocarbon lipid tails. In some embodiments, the branched hydrocarbon lipid tails are from 3-25 carbon atoms in length, such as 5 to 25, 5 to 20, 5 to 15, 5 to 10, 10 to 15, 10 to 20, or 10 to 25 carbon atoms in length.
[0181] In some embodiments, the divalent headgroup is selected from guanidine and squaramide.
[0182] In some embodiments, the squaramide headgroup is of the following formula:
wherein RA and RB are each independently a C1-C6 alkyl group or H; and represents the point of attachment of the headgroup to a straight or branched hydrocarbon lipid tail.
wherein represents the point of attachment of the headgroup to a straight or branched hydrocarbon lipid tail. [0184] In some embodiments, the ionizable lipid comprises a head group selected from:
wherein represents the point of attachment of the headgroup to a hydrocarbon lipid tail (e.g., straight or branched).
[0185] In some embodiments, the ionizable lipid comprises a hydrophilic headgroup as disclosed in Jayaraman et al. Angew. Chem. Int. Ed. (2012), 51, 8529-8533.
[0186] In some embodiments, the ionizable lipid is ethyl lauroyl arginate (ELA). In some embodiments, the ionizable lipid is ionizable lipid 1, wherein ionizable lipid 1 comprises:
[0187] In some embodiments, the ionizable lipid is ionizable lipid 2, wherein the ionizable lipid 2
[0188] In some embodiments, the ionizable lipid is ionizable lipid 3, wherein the ionizable lipid 3 comprises:
[0189] In some embodiments, the ionizable lipid is endosomal escape agent 1, wherein the endosomal escape agent comprises:
[0190] In some embodiments, the one or more of the cationic or ionizable lipids are represented by Formula (LI):
Formula (LI) wherein: n is an integer between 1 and 4;
Ra is hydrogen or hydroxyl; and
Ri and R2 are each independently a linear or branched Co-C id alkyl, Co-C id alkenyl, or Co-C id heteroalkyl, optionally substituted by one or more substituents selected from a group consisting of oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino, hydroxycarbonyl, alkyloxycarbonyl, aminocarbonyl, aminoalkylaminocarbonyl, alkylaminoalkylaminocarbonyl, dialkylaminoalkylaminocarbonyl, heterocyclylalkylaminocarbonyl, (alkylaminoalkyl)(alkyl)aminocarbonyl, alkylaminoalkylcarbonyl,
dialkylaminoalkylcarbonyl, heterocyclylcarbonyl, alkenylcarbonyl, alkynylcarbonyl, alkylsulfoxide, alkylsulfoxidealkyl, alkylsulfonyl, and alkylsulfonealkyl.
[0191] In some embodiments, Ra is hydrogen. In some embodiments, Ra is hydroxyl.
[0192] In some embodiments, the ionizable lipid is represented by Formula (Lla -1), Formula (LIa-2), or Formula (LIa-3):
Formula (LIa-1) Formula (Lla -2) Formula (LIa-3)
[0193] In some embodiments, the ionizable lipid is represented by Formula (LIb-1), Formula (LIb-2), or Formula (LIb-3):
Formula (LIb-1) Formula (LIb-2) Formula (LIb-3)
[0194] In some embodiments, the ionizable lipid is represented by Formula (LIb-4), Formula (LIb-5), Formula (LIb-6), Formula (LIb-7), Formula (LIb-8), or Formula (LIb-9):
Formula (LIb-4) Formula (LIb-5) Formula (LIb-6)
Formula (LIb-7) Formula (LIb-8) Formula (LIb-9)
[0195] In some embodiments, the one or more of the cationic or ionizable lipids are represented by Formula (LI), wherein Ri and R2 are each independently selected from:
[0196] In some embodiments, Ri and R2 are the same. In some embodiments, Ri and R2 are different.
[0197] In various embodiments, the one or more of the cationic or ionizable lipids are represented by Formula (LI*):
Formula (LI*) wherein: n* is an integer between 1 to 7,
Ra is hydrogen or hydroxyl,
Rb is hydrogen or Ci-Ce alkyl,
Ri and R2 are each independently a linear or branched C1-C30 alkyl, C2-C30 alkenyl, or C1-C30 heteroalkyl, optionally substituted by one or more substituents selected from oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino, hydroxycarbonyl, alkyloxycarbonyl, alkylcarbonyloxy, alkylcarbonate, alkenyloxycarbonyl, alkenylcarbonyloxy, alkenylcarbonate, alkynyloxycarbonyl, alkynylcarbonyloxy, alkynylcarbonate, aminocarbonyl, aminoalkylaminocarbonyl, alkylaminoalkylaminocarbonyl, dialkylaminoalkylaminocarbonyl, heterocyclylalkylaminocarbonyl,
(alkylaminoalkyl)(alkyl)aminocarbonyl, alkylaminoalkylcarbonyl, dialkylaminoalkylcarbonyl, heterocyclylcarbonyl, alkenylcarbonyl, alkynylcarbonyl, alkylsulfoxide, alkylsulfoxidealkyl, alkylsulfonyl, and alkylsulfonealkyl.
[0198] In some embodiments, the one or more of the cationic or ionizable lipids are represented by Formula (LII):
Formula (LII) wherein: each n is independently an integer from 2-15;
Li and L3 are each independently -0C(0)-* or -C(O)O-*, wherein indicates the attachment point to Ri or R3;
Ri and R3 are each independently a linear or branched C9-C20 alkyl or C9-C20 alkenyl, optionally substituted by one or more substituents selected from a group consisting of oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino, hydroxycarbonyl, alkyloxycarbonyl, aminocarbonyl, aminoalkylaminocarbonyl, alkylaminoalkylaminocarbonyl, dialkylaminoalkylaminocarbonyl, heterocyclylalkylaminocarbonyl, (alkylaminoalkyl)(alkyl)aminocarbonyl, alkylaminoalkylcarbonyl, dialkylaminoalkylcarbonyl, heterocyclylcarbonyl, alkenylcarbonyl, alkynylcarbonyl, alkylsulfoxide, alkylsulfoxidealkyl, alkylsulfonyl, and alkylsulfonealkyl; and
[0199] In some embodiments, the ionizable lipid is selected from an ionizable lipid of Formula LII, wherein Ri and R3 are each independently selected from a group consisting of:
[0200] In some embodiments, Ri and R3 are the same. In some embodiments, Ri and R3 are different. [0201] In some embodiments, the one or more of the cationic or ionizable lipids are represented by
Formula (LII-2).
[0202] In some embodiments, the ionizable lipid is selected from an ionizable lipid of WO 2015/095340. In some embodiments, the ionizable lipid is selected from an ionizable lipid of WO 2021/021634, WO 2020/237227, or WO 2019/236673. In some embodiments, the ionizable lipid is selected from an ionizable lipid of WO 2021/226597 and WO 2021/113777. In some embodiments, the ionizable lipid is selected from an ionizable lipid of WO 2023/056033. In some embodiments, the ionizable lipid is selected from an ionizable lipid of WO 2023/081526.
[0203] In some embodiments, the one or more of the cationic or ionizable lipids are represented by Formula (LIII):
Formula (LIU) or a pharmaceutically acceptable salt thereof, wherein
L1 is C2-C11 alkylene, Cr-Cio-alkenylene, or Cr-Cio-alkynylene;
X1 is OR1, SR1, or N(R’)2, where R1 is independently H or unsubstituted Ci-Ce alkyl; and
R2 and R3 are each independently Ce-Cso-alkyl, Ce-Cso-alkenyl, or Ce-Cso-alkynyl.
[0204] In some embodiments, the one or more of the cationic or ionizable lipids are represented by Formula (LIII*):
Formula (LUI*) or a pharmaceutically acceptable salt thereof, wherein
L1 is C2-C11 alkylene, Cr-Cio-alkenylene, or Cr-Cio-alkynylene;
X1 is OR1, SR1, or N(R’)2, where R1 is independently H or unsubstituted Ci-Ce alkyl; and
R2 and R3 are each independently a linear or branched C1-C30 alkyl, C2-C30 alkenyl, or C1-C30 heteroalkyl, optionally substituted by one or more substituents selected from oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino, hydroxycarbonyl, alkyloxycarbonyl, alkylcarbonyloxy, alkylcarbonate, alkenyloxycarbonyl, alkenylcarbonyloxy, alkenylcarbonate, alkynyloxycarbonyl, alkynylcarbonyloxy, alkynylcarbonate, aminocarbonyl, aminoalkylaminocarbonyl, alkylaminoalkylaminocarbonyl, dialkylaminoalkylaminocarbonyl, heterocyclylalkylaminocarbonyl,
(alkylaminoalkyl)(alkyl)aminocarbonyl, alkylaminoalkylcarbonyl, dialkylaminoalkylcarbonyl,
heterocyclylcarbonyl, alkenylcarbonyl, alkynylcarbonyl, alkylsulfoxide, alkylsulfoxidealkyl, alkylsulfonyl, and alkylsulfonealkyl.
[0205] In some embodiments, an ionizable lipid is selected from Table 4.
[0206] In some embodiments, an LNP of the present disclosure comprises an ionizable lipid disclosed in one of US 2023/0053437; US 2019/0240354; US 2010/0130588; US 2021/0087135; WO 2021/204179; US 2021/0128488; US 2020/0121809; US 2017/0119904; US 2013/0108685; US 2013/0195920; US 2015/0005363; US 2014/0308304; US 2013/0053572; WO 2019/232095; WO 2021/077067; WO 2019/152557; US 2017/0210697; or WO 2019/089828, each of which is incorporated by reference herein in their entirety.
[0207] In some embodiments, an LNP described herein comprises a lipid, e.g., an ionizable lipid, disclosed in US Application publication US 2017/0119904, which is incorporated by reference herein, in its entirety. [0208] In some embodiments, an LNP described herein comprises a lipid, e.g., an ionizable lipid, disclosed in PCT Application publication WO 2021/204179, which is incorporated by reference herein, in its entirety.
[0209] In some embodiments, an LNP described herein comprises a lipid, e.g., an ionizable lipid, disclosed in PCT Application WO 2022/251665, which is incorporated by reference herein, in its entirety.
[0210] In some embodiments, an LNP described herein comprises an ionizable lipid of Table 5:
Table 5: Exemplary Ionizable Lipid Structures
[0211] In some embodiments, the ionizable lipid is MC3.
Formula (LIV) or is a pharmaceutically acceptable salt thereof, wherein: n* is an integer from 1 to 7 ;
Ra is hydrogen or hydroxyl;
Rh is hydrogen or Ci-Ce alkyl;
R1 is C1-C30 alkyl or R’*;
R2 is C1-C30 alkyl or R2*;
R1* and R2* are independently selected from: -(CH2)qC(O)O(CH2)rC(R8)(R9)(R10),
-(CH2)qOC(O)(CH2)rC(R8)(R9)(R10), and
-(CH2)qOC(O)O(CH2)rC(R8)(R9)(R10); wherein: q is an integer from 0 to 12, r is an integer from 0 to 6, wherein at least one occurrence of r is not 0;
R8 is H or R";
R9, R10, and R” are each independently Ci-C2o alkyl or C2-C2o-alkenyl; and wherein (i) R1 is R1*, (ii) R2 is R2*, or (iii) R1 is R1* and R2 is R2*.
[213] In some embodiments, an ionizable lipid is selected from Table 6.
[0214] In some embodiments, an ionizable lipid of the present disclosure is represented by Formula (LV):
Formula (LV) or is a pharmaceutically acceptable salt thereof, wherein:
Ra is hydrogen or hydroxyl;
R1 is C1-C30 alkyl or R’*;
R2 is C1-C30 alkyl or R2*;
R1* and R2* are independently selected from:
-(CH2)qC(O)O(CH2)rC(R4)(R5)(R6),
-(CH2)qOC(O)(CH2)rC(R4)(R5)(R6), and
-(CH2)qOC(O)O(CH2)rC(R4)(R5)(R6); wherein: q is an integer from 0 to 12, r is an integer from 0 to 6, wherein at least one occurrence of r is not 0;
R4 is hydrogen or R7;
R5, R6, and R7 are each independently Ci-C2o alkyl or C2-C2o-alkenyl; wherein (i) R1 is R1*, (ii) R2 is R2*, or (iii) R1 is R1* and R2 is R2*; and
R3 is L-R’, wherein L is linear or branched Ci-Cio alkylene, and R’ is (i) mono- or bicyclic heterocyclyl or heteroaryl, such as imidazolyl, pyrazolyl, 1 ,2,4-triazolyl, or benzimidazolyl, each optionally substituted at one or more available carbon and nitrogen by Ci-Ce alkyl, or (ii) RA, RB, or Rc, wherein
RB is selected from:
[0215] In some embodiments, the ionizable lipid is selected from an ionizable lipid described or disclosed in any one of PCT Publications WO 2023/044343, WO 2023/044333, WO 2023/122752, WO 2024/044728 and WO 2023/196931 and PCT Application PCT/US2024/019990, or any combination thereof, each of which is incorporated by reference herein in its entirety.
[0216] In some embodiments, an ionizable lipid is selected from Table 7.
C. AMPHIPHILIC POLYMER
[0217] In certain embodiments disclosed herein are amphiphilic polymers. The subject amphiphilic polymers may be used as a component of a composition to facilitate encapsulation and release of circular RNA to one or more target cells (e.g., by forming a shell around the circular RNA core). [0218] Any suitable amphiphilic polymer can be used in the disclosed nanoparticles. Polymers can be natural or unnatural (synthetic) polymers. Polymers can be copolymers comprising two or more monomers. In terms of sequence, copolymers can be random, block, or comprise a combination of random and block sequences.
[0219] The term “polymer,” as used herein, is given its ordinary meaning as used in the art, i.e., a molecular structure comprising one or more repeat units (monomers), connected by covalent bonds.
The repeat units may all be identical, or in some embodiments, there may be more than one type of repeat unit present within the polymer. If more than one type of repeat unit is present within the polymer, then the polymer is said to be a “copolymer.” It is to be understood that in any embodiment employing a polymer, the polymer being employed may be a copolymer in some embodiments. The repeat units forming the copolymer may be arranged in any fashion. For example, the repeat units may be arranged in a random order, in an alternating order, or as a block copolymer, i.e., comprising one or more regions each comprising a first repeat unit (e.g., a first block), and one or more regions each comprising a second repeat unit (e.g., a second block), etc. Block copolymers may have two (a di-block copolymer), three (a tri-block copolymer), or more numbers of distinct blocks.
[0220] Disclosed nanoparticles can include copolymers, which, in some embodiments, describes two or more polymers (such as those described herein) that have been associated with each other, usually by covalent bonding of the two or more polymers together. Thus, a copolymer may comprise a first polymer and a second polymer, which have been conjugated together to form a block copolymer where the first polymer can be a first block of the block copolymer and the second polymer can be a second block of the block copolymer. A block copolymer may, in some embodiments, contain multiple blocks of polymer, and a “block copolymer,” as used herein, is not limited to only block copolymers having only a single first block and a single second block. For instance, a block copolymer may comprise a first block comprising a first polymer, a second block comprising a second polymer, and a third block comprising a third polymer or the first polymer, etc. In some embodiments, block copolymers can contain any number of first blocks of a first polymer and second blocks of a second polymer (and in certain embodiments, third blocks, fourth blocks, etc.). In addition, it should be noted that block copolymers can also be formed, in some instances, from other block copolymers. For example, a first block copolymer may be conjugated to another polymer (which may be a homopolymer, a biopolymer, another block copolymer, etc.), to form a new block copolymer containing multiple types of blocks, and/or to other moieties (e.g., to non-polymeric moieties).
[0221] As disclosed herein, the polymer (e.g., copolymer, e.g., block copolymer) is amphiphilic, i.e., having a hydrophilic portion and a hydrophobic portion, or a relatively hydrophilic portion and a relatively hydrophobic portion. A hydrophilic polymer can be one generally that attracts water, and a hydrophobic polymer can be one that generally repels water.
[0222] In some embodiments, the amphiphilic polymer is a block copolymer comprising a hydrophilic block comprising a hydrophilic polymer; and a hydrophobic block comprising a hydrophobic polymer.
[0223] In some embodiments, the amphiphilic polymer is a biocompatible polymer. In some embodiments, the amphiphilic polymer is a biocompatible polymer that can be biodegradable, i.e., the polymer is able to degrade, chemically and/or biologically, within a physiological environment, such as within the body. As used herein, “biodegradable” polymers are those that, when introduced into cells, are broken down by the cellular machinery (biologically degradable) and/or by a chemical process, such as hydrolysis, (chemically degradable) into components that the cells can either reuse or dispose of without significant toxic effect on the cells. In one embodiment, the biodegradable polymer and their degradation byproducts can be biocompatible.
[0224] In certain embodiments, the subject amphiphilic polymer may be one that hydrolyzes spontaneously upon exposure to water (e.g., within a subject) or degrades upon exposure to heat (e.g., at temperatures of about 37° C). Degradation of a subject amphiphilic polymer may occur at varying rates, depending on the polymer or copolymer used. For example, the half-life of the polymer (the time
at which 50% of the polymer can be degraded into monomers and/or other nonpolymeric moieties) may be on the order of days, weeks, months, or years, depending on the polymer. The polymers may be biologically degraded, e.g., by enzymatic activity or cellular machinery, in some embodiments, for example, through exposure to a lysozyme (e.g., having relatively low pH). In some embodiments, the polymers may be broken down into monomers and/or other nonpolymeric moieties that cells can either reuse or dispose of without significant toxic effect on the cells (for example, a polylactide polymer may be hydrolyzed to form lactic acid, a polyglycolide polymer may be hydrolyzed to form glycolic acid, etc.).
[0225] In some embodiments, the amphiphilic polymer further comprises a cleavable linker (L). The cleavable linker can render the amphiphilic polymer biodegradable (i.e., as described herein above). Any convenient cleavable linker can find use in the subject amphiphilic polymers. In some embodiments, the amphiphilic polymer comprises a cleavable linker that is cleaved by exposure to a stimulus. A non-exhaustive list of stimulus includes pH, temperature, light, redox change, overexpressed enzymes, hypoxia, sound, magnetic force, electrical energy, and any combination thereof.
[0226] In some embodiments, the cleavable linker L comprises a group selected from disulfide, hydrazone, vinyl ether, imine, ortho ester, borate ester, amide, a peptide, an azo, and any combination thereof.
[0227] In some embodiments, the cleavable linker L comprises a disulfide. In some embodiments, the linker comprises a disulfide and can be cleaved by exposure to a redox change. In some embodiments, the cleavage of the disulfide linker is mediated by glutathione (GSH). Disulfide bonds can be easily broken down by reducing glutathione (GSH) into sulfhydryl groups, which causes the degradation of carriers and facilitates the release of cargoes (e.g.,, circRNA cargo). Disulfide bonds are often used in delivery systems as linkers, which can degrade rapidly to release cargoes in the reducing environment of GSH in tumor cells.
[0228] More generally, it is noted that Glutathione (GSH) is the most abundant thiol species in the cytoplasm, functioning as a natural oxidant scavenger and the major reducing agent in biochemical processes. The intracellular GSH concentration (2-10 mM) is substantially higher than extracellular levels (2 pM in plasma), which provides opportunities for intracellular delivery of therapeutic agents by cleavable disulfide linked carriers.
[0229] In some embodiments, the cleavable linker L is a pH sensitive linker. pH is a commonly used internal stimulus in pathological sites such as tumors and inflammatory tissues, as well as in a physiological environment such as acidic organelles (e.g., endosomes). Endosomes (pH 5-6) and lysosomes (pH 4-5) in mammalian cells appear slightly acidic, while the cytoplasm and endoplasmic
reticulum have a neutral pH (e.g., approximately 7.2), the Golgi complex has a pH in the range of 6.0- 6.7, and the mitochondrion a pH of approximately 8.0. After endocytosis, polymeric lipid nanoparticles are entrapped in the early endosome (pH approximately 5.5), which mature into the late endosome. The late endosome fuses with the lysosome (pH less than 5) and these nano-drug delivery systems in the lysosome are subjected to degradation. Polymeric lipid nanoparticles carrying circRNA must avoid endosomal degradation and successfully release the circRNA into the cytoplasm for it to perform its desired therapeutic effect. Accordingly, pH-sensitive polymeric lipid nanoparticles can be designed such that upon being exposed to acidic microenvironment in the endosome, the polymeric shell of the polymeric lipid nanoparticles rapidly disintegrates to release the circRNA and ionizable lipid (lipid). The lipid can then drive the escape of the circRNA from the endosome into the cytoplasm.
[0230] In some embodiments, the cleavable linker L comprises a hydrazone. In some embodiments, the linker comprises a hydrazone that can be cleaved by exposure to an acidic pH. In some embodiments, the linker comprises a hydrazone and can be cleaved by exposure to a pH of 6.5 or less, such as a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 or less, a pH of 4 or less, or even less.
[0231] In some embodiments, the cleavable linker comprises a vinyl ether (see e.g., Shin, et al. Molecular Pharmaceutics 2012, 9(11), 3266-3276). In some embodiments, the linker comprises a vinyl ether that can be cleaved by exposure to an acidic pH. In some embodiments, the linker comprises a vinyl ether and can be cleaved by exposure to a pH of 6.5 or less, a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 of less, a pH of 4 or less, or even less.
[0232] In some embodiments, the cleavable linker comprises one or more of an imine, an ortho ester, a borate ester, or an amide. In some embodiments, the linker comprises one or more of an imine, an ortho ester, a borate ester, or an amide that can be cleaved by exposure to an acidic pH (see e.g., Ding et al. Journal of Controlled Release 2022, 348, 206-238). In some embodiments, the linker comprises one or more of an imine, an ortho ester, a borate ester, or an amide and can be cleaved by exposure to a pH of 6.5 or less, a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 of less, a pH of 4 or less, or even less.
[0233] In some embodiments, the cleavable linker comprises an octapeptide. In some embodiments, the octapeptide is of the sequence GPLGIAGQ. In some embodiments, the octapeptide is of the sequence GPLGVRGC. In some embodiments, the linker comprising an octapeptide is cleaved by exposure to over-expressed enzymes. In some embodiments, the over-expressed enzyme is matrix metalloproteinase 2 (MMP2) (see e.g., Zhu et al. PNAS 2013, 110(42), 17047-17052).
[0234] In some embodiments, the cleavable linker comprises an azo group. In some embodiments, the linker comprises an azo group that can be cleaved by exposure to hypoxia (see e.g., Joshi et al. International Journal of Pharmaceutics 2020, 590, 119915).
[0235] In some embodiments, the hydrophobic block of the amphiphilic polymer comprises the cleavable linker. In some embodiments, the cleavable linker covalently connects the hydrophilic block to the hydrophobic block of the amphiphilic polymer.
[0236] In some embodiments, the cleavable linker covalently connects two or more amphiphilic polymers, such as three or more, four or more, five or more, or even more amphiphilic polymers. In some embodiments, the cleavable linker covalently connects two amphiphilic polymers. In some embodiments, the cleavable linker covalently connects three amphiphilic polymers. In some embodiments, the cleavable linker covalently connects four amphiphilic polymers. In some embodiments, the cleavable linker covalently connects five amphiphilic polymers. In some embodiments, the cleavable linker covalently connects five or more amphiphilic polymers.
[0237] In some embodiments, the amphiphilic polymer is covalently bound to a cationic moiety (e.g., a small molecule cationic moiety such as an ionizable lipid, non-lipid small molecule etc.). In some embodiments, the amphiphilic polymer is covalently bound to the ionizable lipid. In some embodiments, the amphiphilic polymer is not covalently bound to the ionizable lipid.
[0238] In some embodiments, the molecular weight (or e.g., the ratio of molecular weights of, e.g., different blocks of a copolymer) of the amphiphilic polymers can be optimized for effective treatment of a specific disease or disorder. For example, the molecular weight of a polymer may influence particle degradation rate (such as when the molecular weight of a biodegradable polymer can be adjusted), solubility, water uptake, and drug release kinetics. For example, the molecular weight of the polymer (or the ratio of molecular weights of, e.g., different blocks of a copolymer) can be adjusted such that the particle biodegrades in the subject being treated within a reasonable period of time (ranging from a few hours to 1-2 weeks, 3-4 weeks, 5-6 weeks, 7-8 weeks, etc.).
[0239] In some embodiments, the amphiphilic polymer has a molecular weight from 10k to 100k, such as 10k to 90k, 10k to 80k, 10k to 70k, or 10k to 50k. In some embodiments, the amphiphilic polymer has a molecular weight from 10k to 70k, such as 10k to 65k, 10k to 60k, 10k to 55k, 10k to 50k, 20k to 70k, 25k to 70k, 30k to 70k, or 35k to 70k. In some embodiments, the amphiphilic polymer has a molecular weight from 10k to 50k, such as 10k to 45k, 10k to 40k, 10k to 35k, 10k to 30k, 20k to 50k, 25k to 50k, 30k to 50k, or 35k to 50k. In some embodiments, the amphiphilic polymer has a molecular weight of 10k, 15k, 20k, 25k, 30k, 35k, 40k, 45k, 50k, 55k, 60k, 65k, or 70k +/- 10%.
[0240] In some embodiments, the amphiphilic polymer is a di-block copolymer.
[0241] In some embodiments, the amphiphilic polymer comprises a block copolymer of Formula I: X-Y (I) wherein:
X is a hydrophobic block comprising a hydrophobic polymer; and
Y is a hydrophilic block comprising a hydrophilic polymer, wherein the amphiphilic polymer optionally further comprises one or more cleavable linkers.
[0242] In some embodiments of Formula I, the amphiphilic polymer does not include any cleavable linkers.
[0243] In some embodiments of Formula I, the amphiphilic polymer includes one or more cleavable linkers (as described herein). In some embodiments of Formula I, the amphiphilic polymer includes one or more cleavable linkers within the hydrophobic block (X). In some embodiments of Formula I, the amphiphilic polymer includes one or more cleavable linkers within the hydrophilic block (Y). In some embodiments of Formula I, the amphiphilic polymer includes one or more cleavable linkers between the hydrophobic block (X) and the hydrophilic block (Y).
[0244] In some embodiments the amphiphilic polymer includes a cleavable linker (as described herein), and the amphiphilic polymer comprises a block copolymer of Formula IA:
X-L-Y (IA) wherein:
X is a hydrophobic block comprising a hydrophobic polymer;
Y is a hydrophilic block comprising a hydrophilic polymer; and
L is a cleavable linker.
[0245] In some embodiments, the amphiphilic polymer comprises a block copolymer of Formula IB: XA-(L-XB)n-Y (IB) wherein: each of XA and XB is independently a hydrophobic block comprising a hydrophobic polymer;
Y is a hydrophilic block comprising a hydrophilic polymer;
L is a cleavable linker; and n is an integer from 1 to 5.
[0246] In some embodiments of Formula IB, n is 5. In some embodiments, n is 4. In some embodiments of Formula IB, n is an integer from 1 to 3.
[0247] In some embodiments of Formula IB, n is 1 such that the amphiphilic polymer comprises a block copolymer of Formula IB-1 :
XA-L-XB-Y (IB-1)
[0248] In some embodiments of Formula IB, n is 2 such that the amphiphilic polymer comprises a block copolymer of Formula IB -2:
XA-L-XB-L-XB-Y (IB-2)
[0249] In some embodiments of Formula IB, n is 3 such that the amphiphilic polymer comprises a block copolymer of Formula IB -3:
XA-L-XB-L-XB-L-XB-Y (IB-3)
[0250] In some embodiments, the amphiphilic polymer comprises a block copolymer of Formula IC:
XA - T - L - T - XB
Y IA Y IB (IC) wherein: each of XA and XB is independently a hydrophobic block comprising a hydrophobic polymer; each of YA and YB is independently a hydrophilic block comprising a hydrophilic polymer;
L is a cleavable linker; and each T is independently a trivalent connecting group (e.g., lysine or cysteine).
[0251] In some embodiments of Formula IC, the trivalent connecting group T is an amino acid. In some embodiments, the connecting group T is lysine. In some embodiments, the connecting group T is cysteine.
[0252] In some embodiments, the amphiphilic polymer comprises a block copolymer of Formula ID: R-X-Y (ID) wherein:
R is a cationic moiety (e.g., an ionizable lipid);
X is a hydrophobic block comprising a hydrophobic polymer;
Y is a hydrophilic block comprising a hydrophilic polymer; and the amphiphilic polymer optionally further comprises one or more cleavable linkers.
[0253] In some embodiments of Formula ID, the amphiphilic polymer does not comprise a cleavable linker.
[0254] In some embodiments of Formula ID, the amphiphilic polymer further comprises a cleavable linker (as described herein). In some embodiments of Formula ID, the amphiphilic polymer includes
one or more cleavable linkers within the hydrophobic block (X). In some embodiments of Formula ID, the amphiphilic polymer includes one or more cleavable linkers within the hydrophilic block (Y).
[0255] In some embodiments of Formula ID, the amphiphilic polymer further comprises a cleavable linker, and is of Formula ID-1 or ID-2:
R-L-X-Y (ID-1);
R-X-L-Y (ID-2).
[0256] In some embodiments of Formula ID-ID-2, the cationic moiety R comprises a lipid, a polymer, or a non-lipid small molecule. In some embodiments, the cationic moiety R comprises an ionizable lipid (as described herein). In some embodiments, the ionizable lipid has a pKa from 6-12, such as 7- 9. In some embodiments, the ionizable lipid comprises an ionizable amino group. In some embodiments, the ionizable lipid comprises a divalent headgroup and one or more lipid tails (e.g., straight or branched hydrocarbon lipid tails). In some embodiments, the divalent headgroup is selected from guanidine and squaramide. In some embodiments, the ionizable lipid comprises a headgroup as described herein. In some embodiments, the ionizable lipid includes one or more (e.g., 2) straight or branched hydrocarbon lipid tails from 3-25 carbon atoms in length.
[0257] In some embodiments, the cationic moiety R is ethyl lauroyl arginate (EL A).
[0258] In some embodiments of Formula ID-ID-2, the cationic moiety R comprises a polymer (i.e., a cationic polymer). In general, cationic polymers are able to condense and/or protect negatively charged strands of nucleic acids (e.g., DNA, RNA, or derivatives thereof). In some embodiments, the cationic moiety R comprises a polymer selected from poly(lysine), polyethylene imine (PEI), poly (amidoamine), poly (histidine), poly (arginines), and poly amine resins.
[0259] In some embodiments of Formula ID-ID-2, the cationic moiety R comprises a non-lipid small molecule. In some embodiments, the non-lipid small molecule is selected from an amine-containing compound, an amino acid, a heterocycle-containing compound, and a heteroaryl-containing compound. In some embodiments, the cationic moiety R comprises an amine-containing compound. In some embodiments, the amine-containing compound is selected from choline, betaine, N,N’- dibenzylethylenediamine, diethylamine, 2-diethylaminoethanol, 2-methylaminoethanol, glucosamine, glucamine, ethanolamine, ethylenediamine, hydrabamine, isopropyl amine, methylglucamine, procaine, triethylamine, trimethylamine, tripropylamine, and tromethamine.
[0260] In some embodiments of Formula ID-ID-2, the cationic moiety R comprises a non-lipid small molecule that is an amino acid. In some embodiments, the amino acid is selected from arginine, histidine, and lysine.
[0261] In some embodiments of Formula ID-ID-2, the cationic moiety R comprises a non-lipid small molecule that is a heterocycle-containing compound, or a heteroaryl-containing compound. In some embodiments, that cationic moiety R comprises a non-lipid small molecule selected from caffeine, N- ethylmorpholine, N-ethylpiperidine, morpholine, piperazine, piperidine, purines, and theobromine.
[0262] In some embodiments, the amphiphilic polymer is a tri-block copolymer.
[0263] In some embodiments, the amphiphilic polymer comprises a block copolymer of Formula II A: XA-Y-XB (IIA) wherein: each of XA and XB is a hydrophobic block comprising a hydrophobic polymer;
Y is a hydrophilic block comprising a hydrophilic polymer; and the amphiphilic polymer optionally further comprises one or more cleavable linkers.
[0264] In some embodiments of Formula IIA, the amphiphilic polymer does not comprise a cleavable linker.
[0265] In some embodiments of Formula IIA, the amphiphilic polymer further comprises a cleavable linker (as described herein). In some embodiments of Formula IIA, the amphiphilic polymer includes one or more cleavable linkers within one or both of the hydrophobic blocks (XA and/or XB). In some embodiments of Formula IIA, the amphiphilic polymer includes one or more cleavable linkers within the hydrophilic block (Y).
[0266] In some embodiments of Formula IIA, the amphiphilic polymer comprises a cleavable linker, and is of any one of Formulae IIA-1 to IIA-3:
XA-L-Y-XB (IIA-1);
XA-Y-L-XB (IIA-2); or XA-L-Y-L-XB (IIA-3).
[0267] In some embodiments, the amphiphilic polymer comprises a block copolymer of Formula IIB: YA-X-YB (IIB) wherein: each of YA and YB is a hydrophilic block comprising a hydrophilic polymer;
X is a hydrophobic block comprising a hydrophobic polymer; and the amphiphilic polymer optionally further comprises one or more cleavable linkers.
[0268] In some embodiments of Formula IIB, the amphiphilic polymer does not comprise a cleavable linker.
[0269] In some embodiments of Formula IIB, the amphiphilic polymer further comprises a cleavable linker (as described herein). In some embodiments of Formula IIB, the amphiphilic polymer includes one or more cleavable linkers within one or both of the hydrophilic blocks (YA and/or YB). In some embodiments of Formula IIB, the amphiphilic polymer includes one or more cleavable linkers within the hydrophobic block (X).
[0270] In some embodiments of Formula IIB, the amphiphilic polymer further comprises a cleavable linker, and is of any one of Formulae IIB-1 to IIB -3:
YA-L-X-YB (IIB-1);
YA-X-L-YB (IIB-2); or YA-L-X-L-YB (IIB -3).
[0271] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polymer selected from a polyester polymer, a polyorthoester (POE) polymer, a polyanhydride polymer, a polyamide polymer, a poly(ester amide) polymer, a poly(phosphoester) polymer, a poly(alkyl cyanoacrylate) (PACA) polymer, a polysaccharide polymer, and any combination thereof.
[0272] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polyester polymer. In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polyester polymer that is also a polycationic polymer.
[0273] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polyester polymer selected from a polylactide (PLA) polymer, a polyglycolide (PGA) polymer, a polycaprolactone (PCL) polymer, a polydioxanone (PDO) polymer, a polyhydroxyalkanoate (PHA) polymer, a poly(glycerol sebacate) polymer, a poly(lactic-co-glycolic acid) (PLGA) polymer, a poly([5- amino ester) (PB AE) polymer, a poly(amine-co-ester) (PACE) polymer, a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) polymer, and any combination thereof.
[0274] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polylactide (PLA) polymer, where poly-L-lactic acid, poly-D-lactic acid, poly-D,L-lactic acid, poly-L- lactide, poly-D-lactide, and poly-D,L-lactide are all encompassed by the term “PLA.”
[0275] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polyglycolide (PGA) polymer.
[0276] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polycaprolactone (PCL) polymer.
[0277] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polydioxanone (PDO) polymer.
[0278] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polyhydroxyalkanoate (PHA) polymer. In some embodiments, the polyhydroxyalkanoate is a polyhydroxybutyrate polymer. In some embodiments, the polyhydroxyalkanoate is a polyhydroxy valerate .
[0279] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a poly(glycerol sebacate) polymer.
[0280] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a poly(lactic-co-glycolic acid) (PLGA) polymer. PLGA is a biocompatible and biodegradable copolymer of lactic acid and glycolic acid, and various forms of PLGA can be characterized by the ratio of lactic acid:glycolic acid. Lactic acid can be L-lactic acid, D-lactic acid, or D,L-lactic acid. The degradation rate of PLGA can be adjusted by altering the lactic acid-glycolic acid ratio. In some embodiments, PLGA can be characterized by a lactic acid:glycolic acid ratio of approximately 85:15, approximately 75:25, approximately 60:40, approximately 50:50, approximately 40:60, approximately 25:75, or approximately 15:85. In some embodiments, the ratio of lactic acid to glycolic acid monomers in the polymer of the particle (e.g., a PLGA block copolymer or a PLGA-PEG block copolymer), may be selected to optimize for various parameters such as water uptake, therapeutic agent release and/or polymer degradation kinetics can be optimized.
[0281] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises apo poly(P-amino ester) (PBAE) polymer.
[0282] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a poly(amine-co-ester) (PACE) polymer.
[0283] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) polymer.
[0284] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polyorthoester (POE) polymer.
[0285] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polyanhydride polymer.
[0286] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polyamide polymer.
[0287] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a poly(ester amide) polymer.
[0288] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a poly(alkyl cyanoacrylate) (PACA) polymer.
[0289] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a poly(phosphoester) polymer.
[0290] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, X, XA, or XB comprises a polysaccharide polymer. In some embodiments, the polysaccharide polymer is selected from a chitosan polymer, and a hyaluronic acid (HA) polymer, or a combination thereof.
[0291] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, Y, YA, or YB comprises a polymer selected from a polyethylene glycol (PEG) polymer, a polyethylene oxide (PEG) polymer, a polyglutamic acid (PGA) polymer, a poly[N-(2-hydroxypropyl) methacrylamide] (HPMA) polymer, a poly(vinylpyrrolidone) (PVP) polymer, a poly(2-methyl-2-oxazoline) (PMOX) polymer, a poly(N,N- dimethyl acrylamide) (PDMA) polymer, a poly(N-acryloyl morpholine) (PAcM) polymer, and any combination thereof.
[0292] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, Y, YA, or YB comprises a polyethylene glycol PEG polymer. In some embodiments (where chemically feasible), PEG may be terminated and include an end group, for example, when PEG is not conjugated to another polymer. For example, PEG may terminate in a hydroxyl, a methoxy or other alkoxyl group, a methyl or other alkyl group, an aryl group, a carboxylic acid, an amine, an amide, an acetyl group, a guanidino group, or an imidazole. Other contemplated end groups include azide, alkyne, maleimide, aldehyde, hydrazide, hydroxylamine, alkoxyamine, or thiol moieties.
[0293] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, Y, YA, or YB comprises a polyethylene glycol (PEG) polymer.
[0294] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, Y, YA, or YB comprises a polyglutamic acid (PGA) polymer.
[0295] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, Y, YA, or YB comprises a polyethylene oxide (PEO) polymer.
[0296] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, Y, YA, or YB comprises a poly[N-(2-hydroxypropyl) methacrylamide] (HPMA) polymer.
[0297] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB,Y, YA, or YB comprises a poly(vinylpyrrolidone) (PVP) polymer.
[0298] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB,Y, YA, or YB comprises a poly(2-methyl-2-oxazoline) (PMOX) polymer.
[0299] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB,Y, YA, or YB comprises a poly(N,N-dimethyl acrylamide) (PDMA) polymer.
[0300] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB,Y, YA, or YB comprises a poly(N-acryloyl morpholine) (PAcM) polymer.
[0301] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB:
X, XA, or XB comprises a polyester polymer; and
Y, YA, or YB comprises a PEG polymer.
[0302] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, the molar ratio of Y, YA, and/or YB collectively to X, XA, and/or XB collectively is 1:2 to 1:4, such as 1:2 to 1:3 or 1:3 to 1:4. In some embodiments, the molar ratio of Y, YA, and/or YB collectively to X, XA, and/or XB collectively is 1:2 +/- 10%, 1:2.5 +/- 10%, 1:3 +/- 10%, 1:3.5 +/- 10%, or 1:4 +/- 10%.
[0303] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, when X, XA, or XB comprises a polyester polymer (as described herein), the polyester polymer has a molecular weight from 5k to 20k, such as 5k to 15k, 5k to 10k, 10k to 15k, or 10k to 20k. In some embodiments, the polyester polymer has a molecular weight of from 5k to 10k. In some embodiments, the polyester has a molecular weight of 5k +/- 10%. In some embodiments, the polyester has a molecular weight of 6k +/- 10%. In some embodiments, the polyester has a molecular weight of 7k +/- 10%. In some embodiments, the polyester has a molecular weight of 8k +/- 10%. In some embodiments, the polyester has a molecular weight of 9k +/- 10%. In some embodiments, the polyester has a molecular weight of 10k +/- 10%.
[0304] In some embodiments of any one of formulae I, IA-ID, or IIA-IIB, when Y, YA, or YB comprises a PEG polymer, the PEG polymer has a molecular weight from 5k to 10k, such as 5k to 9k, 5k to 8k, 5k to 7k, or 5k to 6k. In some embodiments, the PEG polymer has a molecular weight of 5k +/- 10%. In some embodiments, the PEG polymer has a molecular weight of 6k +/- 10%. In some embodiments, the PEG polymer has a molecular weight of 7k +/- 10%. In some embodiments, the PEG has a molecular weight of 8k +/- 10%. In some embodiments, the PEG has a molecular weight of 9k +/- 10%. In some embodiments, the PEG has a molecular weight of 10k +/- 10%.
[0305] In some embodiments, the amphiphilic polymer comprises a block copolymer of Formula III:
[A]v-[B]w-[C]x-[D]y-[E]z (III) wherein:
A is a polyester monomer or a polyethylene glycol (PEG) monomer;
B is a polyester monomer;
C is a polyester monomer;
D is a polyester monomer;
E is a PEG monomer; v is an integer from 0 to 200 w is an integer from 0 to 200; x is an integer from 0 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150, wherein the amphiphilic polymer optionally further comprises one or more intervening cleavable linkers (L) between any of A-E.
[0306] In some embodiments of Formula (III), each of v, w and x are integers greater than 0. In some embodiments of Formula (III), each of v, w and x are integers from 10 to 200.
[0307] In some embodiments of Formula (III), each of v, w and x are 0, and the amphiphilic polymer comprises a block copolymer of Formula III A:
[D]y-[E]Z (IIIA) wherein:
D is a polyester monomer;
E is a PEG monomer; y is an integer from 10 to 200; and z is an integer from 10 to 150 wherein the amphiphilic polymer optionally further comprises one or more intervening cleavable linkers (L) between D and E.
[0308] In some embodiments of Formula (III), each of v, and w are 0, and the amphiphilic polymer comprises a block copolymer of Formula IIIB:
[C]x-[D]y-[E]z (IIIB) wherein:
C is a polyester monomer;
D is a polyester monomer;
E is a PEG monomer; x is an integer from 10 to 200;
y is an integer from 10 to 200; and z is an integer from 10 to 150 wherein the amphiphilic polymer optionally further comprises one or more intervening cleavable linkers (L) between any of C-E.
[0309] In some embodiments of any of Formula III-IIIB, the amphiphilic polymer further comprises at least one cleavable linker (L) (as described herein). In some embodiments of Formula III, the amphiphilic polymer comprises a cleavable linker between one or more of monomers A and B; monomers B and C; monomers C and D; or monomers D and E. In some embodiments of Formula IIIA, the amphiphilic polymer comprises a cleavable linker between monomers D and E. In some embodiments of Formula IIIB, the amphiphilic polymer comprises a cleavable linker between one or more of monomers C and D; or D and E.
[0310] In some embodiments, the amphiphilic polymer comprises one cleavable linker (E), and is of the Formula IIIC:
[C]x-E-[D]y-[E]z (IIIC) wherein:
C is a polyester monomer;
L is a cleavable linker;
D is a polyester monomer;
E is a PEG monomer; x is an integer from 10 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150.
[0311] In some embodiments, the amphiphilic polymer comprises one cleavable linker (L), and is of the Formula IIID:
[A]v-[B]w-L-[C]x-[D]y-[E]z (IIID) wherein:
A is a polyester monomer;
B is a polyester monomer;
L is a cleavable linker;
C is a polyester monomer;
D is a polyester monomer;
E is a PEG monomer; v is an integer from 10 to 200; w is an integer from 10 to 200;
x is an integer from 10 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150.
[0312] In some embodiments, the amphiphilic polymer comprises one cleavable linker (L), and is of the Formula IIIE:
[C]x-[D]y-L-[E]z (IIIE) wherein:
C is a polyester monomer;
D is a polyester monomer;
L is a cleavable linker;
E is a PEG monomer; x is an integer from 0 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150.
[0313] In some embodiments of Formula IIIE, x is 0, such that the amphiphilic polymer does not include polyester monomer C. In some embodiments of Formula IIIE, x is 10 to 200, such that the amphiphilic polymer does include polyester monomer C.
[0314] In some embodiments of Formula III, v is 0 and each of w and x are integers from 10 to 200, such that the amphiphilic polymer comprises a block copolymer of Formula IIIF:
[B]w-[C]x-[D]y-[E]z (IIIF).
[0315] In some embodiments, the amphiphilic polymer is tri-block copolymer of Formula IV: [A]v-[B]w-[E]z-[C]x-[D]y (IV) wherein:
A is a polyester monomer;
B is a polyester monomer;
E is a PEG monomer;
C is a polyester monomer;
D is a polyester monomer; v is an integer from 10 to 200 w is an integer from 0 to 200; x is an integer from 0 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150,
wherein the amphiphilic polymer optionally further comprises one or more intervening cleavable linkers (L) between any of A-E.
[0316] In some embodiments of Formula IV, the amphiphilic polymer does not include a cleavable linker (L).
[0317] In some embodiments of Formula IV, the amphiphilic polymer further comprises at least one cleavable linker (E) (as described herein). In some embodiments of Formula IV, the amphiphilic polymer comprises a cleavable linker between one or more of monomers A and B ; monomers A and E (when w is 0); monomers B and E; monomers E and C; monomers E and D (when x is 0); or monomers C and D.
[0318] In some embodiments of Formula IV, w is 0, such that the amphiphilic polymer does not include polyester monomer B. In some embodiments of Formula IV, w is 10 to 200, such that the amphiphilic polymer does include polyester monomer B. In some embodiments of Formula IV, x is 0, such that the amphiphilic polymer does not include polyester monomer C. In some embodiments of Formula IV, x is 10 to 200, such that the amphiphilic polymer does include polyester monomer C.
[0319] In some embodiments of Formula IV, each of w and x are 0, such that the amphiphilic polymer comprises a block copolymer of Formula IVA:
[A]v-[E]z-[D]y (IVA).
[0320] In some embodiments of Formula IVA, the amphiphilic polymer does not include a cleavable linker (L). In some embodiments of Formula IVA, the amphiphilic polymer further comprises at least one cleavable linker (L) (as described herein). In some embodiments of Formula IVA, the amphiphilic polymer comprises a cleavable linker between one or more of monomers A and E; or monomers E and D.
[0321] In some embodiments of Formula IV, each of w and x are 10 to 200, such that the amphiphilic polymer comprises a block copolymer of Formula IVB:
[A]v-[B]w-[E]z-[C]x-[D]y (IVB) wherein: v, w, x, and y are each independently an integer from 10 to 200; and z is an integer from 10 to 150.
[0322] In some embodiments of Formula IVB, the amphiphilic polymer does not include a cleavable linker (L). In some embodiments of Formula IVB, the amphiphilic polymer further comprises at least one cleavable linker (L) (as described herein). In some embodiments of Formula IVB, the amphiphilic
polymer comprises a cleavable linker between one or more of monomers A and B ; or monomers E and
D.
[0323] In some embodiments, the amphiphilic polymer is a cross-linked polymer comprising a block copolymer of Formula V :
wherein:
L is a cleavable linker; each T is independently a trivalent connecting group;
E and F are each independently a PEG monomer;
A, B, C, and D are each independently a polyester monomer; v is an integer from 10 to 200 w is an integer from 0 to 200; x is an integer from 0 to 200; y is an integer from 10 to 200; and z and u are each an integer from 10 to 150.
[0324] In some embodiments of Formula V, w is 0, such that the amphiphilic polymer does not include polyester monomer B. In some embodiments of Formula V, w is 10 to 200, such that the amphiphilic polymer does include polyester monomer B. In some embodiments of Formula V, x is 0, such that the amphiphilic polymer does not include polyester monomer C. In some embodiments of Formula V, x is 10 to 200, such that the amphiphilic polymer does include polyester monomer C.
[0325] In some embodiments of Formula V, each of w and x are integers from 10 to 200, such that the amphiphilic polymer comprises all of polyester monomers A, B, C, and D.
[0326] In some embodiments of Formula V, each of w and x are 0, such that the amphiphilic polymer is a block copolymer of Formula VA:
[0327] In some embodiments of Formula V or VA, the trivalent connecting group (T) is a small molecule having a molecular weight of 1000 Daltons (Da) or less, such as 900 Da or less, 850 Da or less, 800 Da or less, 750 Da or less, 700 Da or less, 650 Da or less, 600 Da or less, 550 Da or less, 500
Da or less, 450 Da or less, 400 Da or less, 350 Da or less, 300 Da or less, 250 Da or less, 200 Da or less, or even less.
[0328] In some embodiments of Formula V or VA, the trivalent connecting group (T) is a peptide. In some embodiments, the peptide comprises cysteine. In some embodiments, the peptide comprises lysine.
[0329] In some embodiments of Formula V or VA, L is cleaved by exposure to a stimulus. In some embodiments, the stimulus is selected from pH, temperature, light, redox change, over-expressed enzymes, hypoxia, sound, magnetic force, electrical energy, and any combination thereof.
[0330] In some embodiments of Formula V or VA, the cleavable linker L is a pH-sensitive linker. In some embodiments, the cleavable linker L is stable at physiological pH (approximately pH 7.4), but is cleaved by exposure to an acidic pH, e.g., in acidic regions of cells, such as lysosomes (approximately pH 4.8).
[0331] In some embodiments, L is cleaved by exposure to a specific temperature.
[0332] In some embodiments, L is cleaved by exposure to light.
[0333] In some embodiments, L is cleaved by exposure to a redox change.
[0334] In some embodiments, L is cleaved by exposure to over-expressed enzymes.
[0335] In some embodiments, L is cleaved by exposure to hypoxia.
[0336] In some embodiments, L is cleaved by exposure to a specific frequency of sound.
[0337] In some embodiments, L is cleaved by exposure to a specific magnetic force.
[0338] In some embodiments, L is cleaved by exposure to electrical energy.
[0339] In some embodiments of Formula V or VA, the cleavable linker L comprises a group selected from disulfide, hydrazone, vinyl ether, imine, ortho ester, borate ester, amide, a peptide, and azo.
[0340] In some embodiments of Formula V or VA, the cleavable linker L comprises a disulfide. In some embodiments, the linker comprises a disulfide and can be cleaved by exposure to a redox change.
[0341] In some embodiments of Formula V or VA, the cleavable linker L comprises a hydrazone. In some embodiments, the linker comprises a hydrazone that can be cleaved by exposure to an acidic pH. In some embodiments, the linker comprises a hydrazone and can be cleaved by exposure to a pH of 6.5
or less, such as a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 or less, a pH of 4 or less, or even less.
[0342] In some embodiments of Formula V or VA, the cleavable linker comprises a vinyl ether (see e.g., Shin, et al. Molecular Pharmaceutics 2012, 9(11), 3266-3276). In some embodiments, the linker comprises a vinyl ether that can be cleaved by exposure to an acidic pH. In some embodiments, the linker comprises a vinyl ether and can be cleaved by exposure to a pH of 6.5 or less, a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 of less, a pH of 4 or less, or even less.
[0343] In some embodiments of Formula V or VA, the cleavable linker comprises an imine, an ortho ester, a borate east, or an amide. In some embodiments, the linker comprises one or more of an imine, an ortho ester, a borate ester, or an amide that can be cleaved by exposure to an acidic pH (see, e.g., Ding et al. Journal of Controlled Release 2022, 348, 206-238). In some embodiments, the linker comprises one or more of an imine, an ortho ester, a borate ester, or an amide and can be cleaved by exposure to a pH of 6.5 or less, a pH of 6 or less, a pH of 5.5 or less, a pH of 5 or less, a pH of 4.5 of less, a pH of 4 or less, or even less.
[0344] In some embodiments of Formula V or VA, the cleavable linker comprises an octapeptide. In some embodiments, the octapeptide is of the sequence GPLGIAGQ. In some embodiments, the octapeptide is of the sequence GPLGVRGC. In some embodiments, the linker comprising an octapeptide is cleaved by exposure to over-expressed enzymes. In some embodiments, the overexpressed enzyme is matrix metalloproteinase 2 (MMP2) (see e.g., Zhu et al. PNAS 2013, 110(42), 17047-17052).
[0345] In some embodiments of Formula V or VA, the cleavable linker comprises an azo group. In some embodiments, the linker comprises an azo group that can be cleaved by exposure to hypoxia (see e.g., Joshi et al. International Journal of Pharmaceutics 2020, 590, 119915).
[0346] In some embodiments, the amphiphilic polymer comprises a block copolymer of Formula VI: R-[C]x-[D]y-[E]z (VI) wherein:
R is a cationic moiety;
C is a polyester monomer;
D is a polyester monomer;
E is a PEG monomer; x is an integer from 0 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150.
[0347] In some embodiments of Formula VI, x is 0, such that the amphiphilic polymer does not include polyester monomer C. In some embodiments of Formula x, w is 10 to 200, such that the amphiphilic polymer does include polyester monomer C.
[0348] In some embodiments of Formula VI, the cationic moiety R comprises a lipid, a polymer, or a non-lipid small molecule.
[0349] In some embodiments of Formula VI, the cationic moiety R comprises an ionizable lipid (as described herein).
[0350] In some embodiments of Formula VI, the cationic moiety R comprises a polymer (as described herein). In some embodiments, the polymer is selected from poly(lysine), polyethylene imine (PEI), poly (amidoamine), poly (histidine), poly (arginines), and poly amine resins.
[0351] In some embodiments of Formula VI, the cationic moiety R comprises poly(lysine).
[0352] In some embodiments of Formula VI, the cationic moiety R comprises polyethylene imine (PEI).
[0353] In some embodiments of Formula VI, the cationic moiety R comprises poly (amidoamine).
[0354] In some embodiments of Formula VI, the cationic moiety R comprises poly (histidine).
[0355] In some embodiments of Formula VI, the cationic moiety R comprises poly (arginines).
[0356] In some embodiments of Formula VI, the cationic moiety R comprises polyamine resins.
[0357] In some embodiments of Formula VI, the cationic moiety R comprises a non-lipid small molecule. In some embodiments, the non-lipid small molecule is selected from an amine-containing compound, an amino acid, a heterocycle-containing compound, and a heteroaryl-containing compound.
[0358] In some embodiments of Formula VI, the cationic moiety R comprises a non-lipid small molecule that is an amine-containing compound. In some embodiments, the amine-containing compound is selected from choline, betaine, N,N’ -dibenzylethylenediamine, diethylamine, 2- diethylaminoethanol, 2-methylaminoethanol, glucosamine, glucamine, ethanolamine, ethylenediamine, hydrabamine, isopropyl amine, methylglucamine, procaine, triethylamine, trimethylamine, tripropylamine, and tromethamine.
[0359] In some embodiments of Formula VI, the cationic moiety R is a non-lipid small molecule that is an amino acid. In some embodiments, the amino acid is arginine. In some embodiments, the amino acid is histidine. In some embodiments, the amino acid is lysine.
[0360] In some embodiments of Formula VI, the non-lipid small molecule is a heterocycle-containing compound or a heteroaryl-containing compound.
[0361] In some embodiments of Formula VI, the non-lipid small molecule is selected from caffeine, N-ethylmorpholine, N-ethylpiperidine, morpholine, piperazine, piperidine, purines, and theobromine.
[0362] In some embodiments, polyester monomers A, B, C and D each independently include a polyester monomer selected from a polylactide (PLA) monomer, a polyglycolide (PGA) monomer, a polycaprolactone (PCL) monomer, a polydioxanone (PDO) monomer, a polyhydroxyalkanoate monomer, a poly(glycerol sebacate) monomer, a poly(P-amino ester) (PBAE) monomer, a poly(2- (dimethylamino)ethyl methacrylate) (PDMAEMA) monomer and any combination thereof.
[0363] In some embodiments, one or more of the polyester monomers A, B, C or D comprises a polylactide (PLA) monomer. In some embodiments, A comprises a polylactide (PLA) monomer. In some embodiments, B comprises a polylactide (PLA) monomer. In some embodiments, C comprises a polylactide (PLA) monomer. In some embodiments, D comprises a polylactide (PLA) monomer. In some embodiments, each of A, B, C and D each comprise a polylactide (PLA) monomer.
[0364] In some embodiments, one or more of the polyester monomers A, B, C or D comprises a polyglycolide (PGA) monomer. In some embodiments, A comprises a polyglycolide (PGA) monomer. In some embodiments, B comprises a polyglycolide (PGA) monomer. In some embodiments, C comprises a polyglycolide (PGA) monomer. In some embodiments, D comprises a polyglycolide (PGA) monomer.
[0365] In some embodiments, one or more of the polyester monomers A, B, C or D comprises a polycaprolactone (PCL) monomer. In some embodiments, A comprises a polycaprolactone (PCL) monomer. In some embodiments, B comprises a polycaprolactone (PCL) monomer. In some embodiments, C comprises a polycaprolactone (PCL) monomer. In some embodiments, D comprises a polycaprolactone (PCL) monomer.
[0366] In some embodiments, one or more of the polyester monomers A, B, C or D comprises a polydioxanone (PDO) monomer. In some embodiments, A comprises a polydioxanone (PDO) monomer. In some embodiments, B comprises a polydioxanone (PDO) monomer. In some embodiments, C comprises a polydioxanone (PDO) monomer. In some embodiments, D comprises a polydioxanone (PDO) monomer.
[0367] In some embodiments, one or more of the polyester monomers A, B, C or D comprises a polyhydroxyalkanoate monomer. In some embodiments, A comprises a polyhydroxyalkanoate monomer. In some embodiments, B comprises a polyhydroxyalkanoate monomer. In some embodiments, C comprises a polyhydroxy alkanoate monomer. In some embodiments, D comprises a polyhydroxy alkanoate monomer. In some embodiments, the polyhydroxy alkanolate monomer is a polyhydroxybutyrate monomer. In some embodiments, the polyhydroxylalkanolate monomer is a polyhydroxy valerate .
[0368] In some embodiments, one or more of the polyester monomers A, B, C or D comprises a poly(glycerol sebacate) monomer. In some embodiments, A comprises a poly(glycerol sebacate) monomer. In some embodiments, B comprises a poly(glycerol sebacate) monomer. In some embodiments, C comprises poly(glycerol sebacate) monomer. In some embodiments, D comprises a poly(glycerol sebacate) monomer.
[0369] In some embodiments, one or more of the polyester monomers A, B, C or D comprises a poly([5- amino ester) (PBAE) monomer. In some embodiments, A comprises a poly(P-amino ester) (PBAE) monomer. In some embodiments, B comprises a poly(P-amino ester) (PBAE) monomer. In some embodiments, C comprises a poly(P-amino ester) (PBAE) monomer. In some embodiments, D comprises a poly(P-amino ester) (PBAE) monomer.
[0370] In some embodiments, one or more of the polyester monomers A, B, C or D comprises a poly(2- (dimethylamino)ethyl methacrylate) (PDMAEMA) monomer. In some embodiments, A comprises a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer. In some embodiments, B comprises a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer. In some embodiments, C comprises a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer. In some embodiments, D comprises a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
[0371] In some embodiments, [A]v and [B]w together form a copolymer comprising any combination of monomers selected from a polylactide (PLA) monomer, a polyglycolide (PGA) monomer, a polycaprolactone (PCL) monomer, a polydioxanone (PDO) monomer, a polyhydroxyalkanoate (e.g., polyhydroxybutyrate) monomer, a poly(glycerol sebacate) monomer a poly(P-amino ester) (PBAE) monomer, and a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
[0372] In some embodiments, polyester monomers [A]v and [B]w together form a poly(lactic-co- glycolic acid) (PLGA) copolymer. In some embodiments, the molar ratio of lactic acid monomer units to glycolic acid monomer units in the PLGA copolymer is from 1:1 to 9:1, such as 1:1, 2:1, 3:1, 4:1, 5:1, 6:1, 7:1, 8:1, or 9:1.
[0373] In some embodiments, [C]x and [D]y together form a copolymer comprising any combination of monomers selected from a polylactide (PLA) monomer, a polyglycolide (PGA) monomer, a polycaprolactone (PCL) monomer, a polydioxanone (PDO) monomer, a polyhydroxyalkanoate (e.g., polyhydroxybutyrate) monomer, a poly(glycerol sebacate) monomer a poly(P-amino ester) (PBAE) monomer, and a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
[0374] In some embodiments, polyester monomers [C]x and [D]y together form a poly(lactic-co- glycolic acid) (PLGA) copolymer. In some embodiments, the molar ratio of lactic acid monomer units to glycolic acid monomer units in the PLGA copolymer is from 1:1 to 9:1, such as 1:1, 2:1, 3:1, 4:1, 5:1, 6:1, 7:1, 8:1, or 9:1.
[0375] In some embodiments, the subject amphiphilic polymer further comprises one or more terminal groups selected from carboxyl, hydroxyl, amino, amido and alkoxy. In some embodiments, one or more terminal groups is a carboxyl group. In some embodiments, one or more terminal groups is a hydroxyl group. In some embodiments one or more terminal groups is an amino group. In some embodiments, one or more terminal groups is an amido group. In some embodiments, one or more terminal groups is an alkoxy group. In some embodiments, the terminal group is an alkoxy group selected from n-butoxy (-O-(CH2)3)-CH3) and tert-butoxy (-O-(C(CH3)3).
[0376] In some embodiments, the subject amphiphilic polymer comprises a di-block copolymer of polyester PLA and a PEG polymer. In some embodiments, the molecular weight of the PLA polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k. In some embodiments, the molecular weight of the PEG polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k. In some embodiments, the PLA and PEG blocks have different molecular weights. In some embodiments, the PLA block has a molecular weight of 10k, and the PEG block has a molecular weight of 5k (also referred to herein as 10k-5k PLA-PEG). In some embodiments, the PLA block has a molecular weight of 9k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has a molecular weight of 8k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has a molecular weight of 7k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has a molecular weight of 6k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has the same molecular weight as the PEG block. In some embodiments, the PLA block has a molecular weight of 5k, and the PEG block has a molecular weight of 5k.
[0377] In some embodiments the subject amphiphilic polymer is of Formula IIIA, and comprises a diblock copolymer of PLA and PEG polymers represented by the following structure:
PLA PEG wherein n is 10 to 200 and m is 10 to 150.
[0378] In some embodiments the subject amphiphilic polymer is of Formula IIIA, and comprises a diblock copolymer of PLA and PEG polymers represented by the following structure:
PLA PEG wherein n is 10 to 200, m is 10 to 150, and each G, independently, can be H, CH3, NH2 or COOH.
[0379] In some embodiments, the subject amphiphilic polymer comprises a di-block copolymer of polyester PLGA and a PEG polymer. In some embodiments, the molecular weight of the PLGA polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k. In some embodiments, the molecular weight of the PEG polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k. In some embodiments, the PLGA and PEG blocks have different molecular weights. In some embodiments, the PLGA block has a molecular weight of 10k, and the PEG block has a molecular weight of 5k (also referred to herein as 10k-5k PLGA-PEG). In some embodiments, the PLGA block has a molecular weight of 9k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 8k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 7k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 6k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has the same molecular weight as the PEG block. In some embodiments, the PLGA block has a molecular weight of 5k, and the PEG block has a molecular weight of 5k.
[0380] In some embodiments, the subject amphiphilic polymer is of Formula III, and comprises a diblock copolymer of PLGA and PEG polymers represented by the following structure:
PLGA PEG wherein x and y are each 10 to 200, and z is 10 to 150.
[0381] In some embodiments the subject amphiphilic polymer is of Formula III, and comprises a di- block copolymer of PLGA and PEG polymers represented by the following structure:
PLGA PEG wherein x and y are each 10 to 200, z is 10 to 150, and each G, independently, can be H, CH3, NH2 or COOH.
[0382] In some embodiments, the amphiphilic polymer is a PLA-PEG, or a PLGA-PEG copolymer as described in U.S. Pat. Pub. No. US2021/0259982, which is incorporated herein by reference in its entirety.
[0383] In some embodiments, the amphiphilic polymer includes a cleavable linker (L). In some embodiments, the linker is a disulfide. A disulfide bond can be incorporated into compound of Formula III, such as the PLA-PEG or PLGA-PEG backbone as a linker to help facilitate the breakdown of the polymeric lipid nanoparticle shell once the particles are exposed to higher glutathione concentrations in the cell cytoplasm.
[0384] For example, a disulfide linker can be inserted into a di-block copolymer of polyester PL A and a PEG polymer, where the combined molecular weight of PLA polymer is 16k and the molecular weight of PEG polymer is 5k. For example, an 8k block of PLA can be connected to a copolymer of PLA (8k) and PEG (5k) using a disulfide bond as a linker (PLAsk-SS-PLAsk-PEGsk). The PLAsk-SS-PLAsk-PEGsk polymer can form a stable shell around the circRNA-lipid complex cargo until the particle reaches the cytoplasm, where the PLAsk-SS-PLAsk-PEGsk can break into PLAxk and PLAsk-PEGsk, thus breaking the shell of the particle and releasing the cargo inside the cytoplasm of the cell. FIG. 12 is a graphic
representation of the glutathione mediated disruption of a polymeric lipid nanoparticle comprising a shell of amphiphilic polymers having disulfide linkages. See also FIG. 11.
[0385] In some embodiments the subject amphiphilic polymer is of Formula IIIC, and comprises a diblock copolymer of PLA and PEG polymers including a cleavable disulfide linker represented by the following structure:
wherein x and y are 10 to 200, z is 10 to 150, and m and m’ independently are 1 to 20.
[0386] In some embodiments the subject amphiphilic polymer is of Formula IIIC, and comprises a diblock copolymer of PLA and PEG polymers including a cleavable disulfide linker represented by the following structure:
wherein x and y are 10 to 200, z is 10 to 150, m and m’ independently are 1 to 20, and each G, independently, can be H, CH3, NH2 or COOH.
[0387] In some embodiments, the amphiphilic polymer includes a cleavable pH sensitive linker (L). In some embodiments, the amphiphilic polymer includes a pH sensitive cleavable linker (L) that is a hydrazone. Hydrazone (C=NNHR) is a compound produced by condensation of hydrazine and aldehyde or ketone. It contains a C=N bond, and the double bond breaks upon the attack of protons under acidic conditions. Acid hydrolysis of the component is rapid. The hydrazone can be more stable than an ester, ethylene ether or and imine under a physiological condition of pH 7.4. In addition, the hydrazone hydrolyzes at a pH of about 5-6, producing ketones and hydrazine. Such a bond can be inserted in some of the formulae described herein.
[0388] For example, a hydrazone linker can be inserted into a di-block copolymer of Formula IIIC, comprising polyester PLA and a PEG polymer, to provide the following constructs:
PLA-hydrazone-PLA-PEG,
PLA-hydrazone-PLA-hydrazone-PLA-PEG, and PLA-hydrazone-PEG.
[0389] In another example, a hydrazone linker can be inserted into a di-block copolymer of Formula IIID, comprising polyester PLGA (PLA and PGA monomers) and a PEG polymer, to provide the following constructs:
PLGA-hydrazone-PLGA-PEG,
PLGA-hydrazone-PLGA-hydrazone-PLGA-PEG, and PLGA-hydrazone-PEG.
[0390] In some embodiments, the subject amphiphilic polymer is of Formula IIIA, and comprises a diblock copolymer of PLA and PEG polymers including a cleavable hydrazone linker represented by the following structure:
wherein x is 10 to 200, and z is 10 to 150.
[0391] In some embodiments, the subject amphiphilic polymer is of Formula IIIA, and comprises a diblock copolymer of PLA and PEG polymers including a cleavable hydrazone linker represented by the following structure:
wherein x is 10 to 200, z is 10 to 150, and each G, independently, can be H, CH3, NH2 or COOH.
[0392] In some embodiments, the amphiphilic polymer includes a cleavable linker (L), and the polymer is configured as a cross-linked polymer (see, e.g., Formulae IC, V and VA) to ensure the cleavable linker groups (L) are exposed closer to the surface of the nanoparticle. In some embodiments the cleavable linker group (L) is a disulfide, and two copolymers are cross-linked via a disulfide bond using cysteine connecting groups. In some embodiments, the cross-linked copolymers are PLA-PEG. In some embodiments, the cross-linked copolymers are PLGA-PEG.
[0393] In some embodiments the subject amphiphilic polymer is of the Formula VA, and comprises PLA-PEG copolymers linked via a disulfide bond and cysteine connecting groups:
wherein x is 10 to 200, and z is 10 to 150. [0394] In some embodiments, the subject amphiphilic polymer is of the Formula VA, and comprises
PLA-PEG copolymers linked via a disulfide bond and cysteine connecting groups:
wherein x is 10 to 200, z is 10 to 150, and each G, independently, can be H, CH3, NH2 or COOH.
[0395] FIG. 12 is a graphic representation of the glutathione mediated disruption of a polymeric lipid nanoparticle comprising a shell of cross-linked amphiphilic polymers having disulfide linkages.
[0396] In some embodiments the cleavable linker group (L) is a disulfide, and two copolymers are cross-linked via a hydrazone bond using lysine connecting groups. In some embodiments, the crosslinked copolymers are PLA-PEG. In some embodiments, the cross-linked copolymers are PLGA-PEG.
[0397] In some embodiments the subject amphiphilic polymer is of the Formula VA, and comprises PLA-PEG copolymers linked via a cleavable linker comprising hydrazone groups and lysine connecting groups:
wherein x is 10 to 200, and z is 10 to 150.
[0398] In some embodiments the subject amphiphilic polymer is of the Formula VA, and comprises PLA-PEG copolymers linked via a cleavable linker comprising hydrazone groups and lysine connecting groups:
wherein x is 10 to 200, z is 10 to 150, and each G, independently, can independently be H, CH3, NH2 or COOH.
[0399] FIG. 13 is a graphic representation of the pH mediated disruption of a polymeric lipid nanoparticle comprising a shell of cross-linked amphiphilic polymers having hydrazone linkages.
[0400] In some embodiments, the subject amphiphilic polymer comprises a tri-block copolymer of polyester PL A and a PEG polymer. In some embodiments, the molecular weight of the PL A polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k. In some embodiments, the molecular weight of the PEG polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k. In some embodiments, the PL A and PEG blocks have different molecular weights. In some embodiments, the PL A block has a molecular weight of 10k, and the PEG block has a molecular weight of 5k. In some embodiments, the PL A block has a molecular weight of 9k, and the PEG block has a molecular weight of 5k. In some embodiments, the PL A block has a molecular weight of 8k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has a molecular weight of 7k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has a molecular weight of 6k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLA block has the same molecular weight as the PEG block. In some embodiments, the PLA block has a molecular weight of 5k, and the PEG block has a molecular weight of 5k.
[0401] In some embodiments, the subject amphiphilic polymer is of Formula IVA, and comprises a tri-block copolymer of PL A and PEG polymers represented by the following structure:
wherein v and y independently are 10 to 200, and z is 10 to 150.
[0402] In some embodiments the subject amphiphilic polymer is of Formula IVA, and comprises a tri- block copolymer of PL A and PEG polymers represented by the following structure:
wherein v and y independently are 10 to 200, z is 10 to 150, and each G can independently be H, CH3,
NH2 or COOH.
[0403] In some embodiments, the subject amphiphilic polymer comprises a tri-block copolymer of polyester PLGA and a PEG polymer. In some embodiments, the molecular weight of the PLGA polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k. In some embodiments, the molecular weight of the PEG polymer block can be less than 10k, such as 10k to 9k, 9k to 8k, 8k to 7k, 7k to 6k or 6k to 5k. In some embodiments, the PLGA and PEG blocks have different molecular weights. In some embodiments, the PLGA block has a molecular weight of 10k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 9k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 8k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 7k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has a molecular weight of 6k, and the PEG block has a molecular weight of 5k. In some embodiments, the PLGA block has the same molecular weight as the PEG block. In some embodiments, the PLGA block has a molecular weight of 5k, and the PEG block has a molecular weight of 5k.
[0404] In some embodiments the subject amphiphilic polymer is of Formula IVB, and comprises a triblock copolymer of PLGA and PEG polymers represented by the following structure:
PLGA PEG PLGA wherein each x and y independently are independently 10 to 200, and z is 10 to 150.
[0405] In some embodiments the subject amphiphilic polymer is of Formula IVB, and comprises a tri- block copolymer of PLGA and PEG polymers represented by the following structure:
PLGA PEG PLGA wherein each x and y independently are independently 10 to 200, z is 10 to 150, and each G can independently be H, CH3, NH2 or COOH.
[0406] When forming polymeric lipid nanoparticles using the triblock polymers shown above, (i.e., PLA-PEG-PLA, and PLGA-PEG-PLGA), the two hydrophobic polymer blocks (PLA or PLGA) can be embedded in the nanoparticle shell, whereas the PEG polymer block can form a loop on the outside of the particle. In this configuration, the PEG would not have any end groups and may lead to reduced anti-PEG IgM formation and reduced accelerated blood clearage (ABC) of polymeric lipid nanoparticles after subsequent doses. The graphical representation of polymeric lipid nanoparticles prepared using such tri-block polymer is shown in FIG. 14.
[0407] In some embodiments, the subject amphiphilic polymer is of Formula VI and comprises an ionizable lipid as the cationic moiety conjugated to the polymer, such that the ionizable lipid can anchor the core to the shell of the particle. For example, a lipid such as ethyl lauroyl arginate (ELA) can be conjugated to a PLA-PEG polymer, forming PEG-PLA-ELA. This polymer-lipid complex, when incorporated within the polymeric lipid nanoparticle, can recruit the circRNA from the aqueous phase and form a stable core of the polymeric lipid nanoparticle, while still being connected the polymeric shell of the nanoparticle. Such an assembly of a polymeric lipid nanoparticle is illustrated in FIG. 15.
[0408] In some embodiments the subject amphiphilic polymer is of Formula IV, and comprises a copolymer of PLA and PEG polymers conjugated to ionizable lipid R, represented by the following structure:
PLA PEG wherein x is 10 to 200, z is 10 to 150, and R is an ionizable lipid (as described herein).
[0409] In some embodiments the subject amphiphilic polymer is of Formula IV, and comprises a copolymer of PLA and PEG polymers conjugated to ionizable lipid R, represented by the following structure:
PLA PEG wherein x is 10 to 200, z is 10 to 150, R is an ionizable lipid (as described herein), and G can be H,
CH3, NH2 or COOH.
[0410] In some embodiments the subject amphiphilic polymer is of Formula IV, and comprises a copolymer of PLGA and PEG polymers conjugated to ionizable lipid R, represented by the following structure:
wherein x and y are each independently 10 to 200, z is 10 to 150, and R is an ionizable lipid (as described herein).
[0411] In some embodiments the subject amphiphilic polymer is of Formula IV, and comprises a copolymer of PLGA and PEG polymers conjugated to ionizable lipid R, represented by the following structure:
PLGA PEG wherein x and y are each independently 10 to 200, z is 10 to 150, R is an ionizable lipid (as described herein), and G can be H, CH3, NH2 or COOH.
[0412] In certain embodiments, the lipid nanoparticle further comprises one or more additional amphiphilic PEG lipids, other than and in addition to those described above. In certain embodiments, the lipid nanoparticle further comprises a PEG lipid disclosed or described in PCT Publication WO 2024/044728, which is incorporated by reference herein, in its entirety. In certain embodiments, the lipid nanoparticle further comprises a PEG lipid selected from PEG-c-DOMG, PEG-DMG, PEG- DLPE, PEG-DMPE, PEG-DPPC, PEG-DSPE, PEG-DAG, PEG-S-DAG, PEG-PE, PEG-S-DMG, PEG-CER, PEG-dialkoxypropylcarbamate, PEG-OR, PEG-OH, PEG-c-DOMG, and PEG-1.
2. EMULSIONS AND METHODS OF PREPARATION
[0413] As summarized herein, the present disclosure provides emulsions comprising polymeric lipid nanoparticles. Also provided are methods of preparing the subject polymeric lipid nanoparticles.
A. EMULSIONS
[0414] As summarized herein, the present disclosure provides emulsion comprising an aqueous continuous phase and an organic dispersed phase, wherein the organic dispersed phase comprises a circular RNA polynucleotide (circRNA), an ionizable lipid, and an amphiphilic polymer.
[0415] In some embodiments of the emulsion, the circRNA, ionizable lipid and amphiphilic polymer components are as described herein above.
[0416] In some embodiments, the emulsion is an intermediate in the process for forming the subject polymeric lipid nanoparticles.
[0417] In some embodiments of the emulsion, the ratio of the organic dispersed phase to the aqueous continuous phase is from 1:3 to 1.7, such as 1:3 to 1:5, 1:4 to 1:6, and 1:5 to 1.7. In some embodiments
of the emulsion, the ratio of the organic phase to the aqueous continuous phase is 1:3, 1:3.5, 1:4, 1:4.5, 1:5, 1:5.5, 1:6, 1:6.5, or 1:7.
[0418] In some embodiments of the emulsion, the organic dispersed phase is composed of a mixture of organic solvents. In some embodiments, the organic solvents are at least partially water miscible.
[0419] As used herein the term “miscible” is intended to mean that two or more components form a single -phase solution. The term “water miscible” or “water soluble” is intended to mean that a component is miscible in water over an extensive concentration range to form a single homogeneous solution. The term “partially miscible” with respect to two liquid phases is intended to mean that the two liquid phases are separated into two liquid phases after mixing and each liquid phase is in a dissolved state at ambient temperature and pressure. For example, the “partially water miscible organic solvent” is partially, for example, 10% (v/v) partially miscible solvent is fully miscible up to 10% (v/v) and immiscible over 10% over v/v at ambient temperature and pressure.
[0420] In some embodiments, the partially water miscible organic solvent is selected from one or more of benzyl alcohol, ethyl acetate, toluene, methyl ethyl ketone, acetonitrile, tetrahydrofuran (THF), isopropyl alcohol, isopropyl acetate, dimethylformamide (DMF), dichloromethane, chloroform, acetone, dimethylsulfoxide (DMSO), N-methylpyrrolidinone (NMP), 2-methyltetrahydrofuran, sulfolane, 1,1, 3,3-tetramethylurea (TMU), N,N'-dimethyl-N,N'-trimethyleneurea (1,3-dimethyl- 3,4,5,6-tetrahydro-2-pyrimidinone (DMPU)), 1.3 -dimethyl-2-imidazolidinone (DMI), 1,4-dioxane, 1,3-dioxolane, lactate esters, polyethers, anisole, benzonitrile, 1 -butanol, 2-butanol, tert-butanol, 2 - butoxyethanol, 2-butoxyethyl acetate, 2-(2-butoxyethoxy)ethyl acetate, tert-butyl acetoacetate, carbon disulfide, diethoxy ethane, 1 ,2-dimethoxyethane, N,N-dimethylacetamide, ethanol, 2-ethoxyethanol, ethyl formate, ethylene glycol, ethylene glycol diethyl ether, 2-ethyl-l -hexanol, formamide, glycerol, 1 -heptanol, 2-heptanone, 1 -hexanol, 2-hexanol, 2-methoxyethanol, 2-methoxyethyl acetate, 1 - methoxy-2-propanol, 3-methyl-l -butanol, methyl butyl ketone a, methyl isobutyl ketone, methyl formate, 5-methyl-2-hexanone, 4-methyl-2-pentanol, 2-methyl-l -propanol, nitrobenzene, nitromethane, 1-cctanol, 1 -pentanol, 2-pentanol, 3-pentanol, 2-pentanone, 3-pentanone, 1 -propanol, 2- propanol, propionaldehyde, 2-propoxyethanol, tetralin, triethyl orthoformate, and 3,5,5-trimethyl-l- hexanol.
[0421] In some embodiments, the partially water-miscible organic solvent is selected from benzyl alcohol, ethyl acetate, and a combination thereof.
[0422] In some embodiments, the emulsion further comprises a surfactant. In some embodiments, the surfactant comprises sodium cholate, sodium, polysorbate 20, polysorbate 80, sorbitan monooleate,
polyoxyethylene (100) stearyl ether (e.g., BRIJ® S100), Polyoxyethylene (23) Lauryl Ether (e.g., BRIJ® 35), Polyethylene glycol hexadecyl ether (e.g., BRIJ® 52), or any combination thereof.
B. PRODUCTION OF POLYNUCLEOTIDES
[0423] DNA templates can be made using standard techniques of molecular biology. For example, the various elements of the vectors provided herein can be obtained using recombinant methods, such as by screening cDNA and genomic libraries from cells, or by deriving the polynucleotides from a DNA template known to include the same.
[0424] The various elements of the DNA template can also be produced synthetically, rather than cloned, based on the known sequences. The complete sequence can be assembled from overlapping oligonucleotides prepared by standard methods and assembled into the complete sequence. See, e.g., Edge, Nature (1981) 292:756; Nambair et al., Science (1984) 223: 1299; and Jay et al., J. Biol. Chem. (1984) 259:631 1.
[0425] Thus, particular nucleotide sequences can be obtained from DNA template harboring the desired sequences or synthesized completely, or in part, using various oligonucleotide synthesis techniques known in the art, such as site-directed mutagenesis and polymerase chain reaction (PCR) techniques where appropriate. One method of obtaining nucleotide sequences encoding the desired DNA template elements is by annealing complementary sets of overlapping synthetic oligonucleotides produced in a conventional, automated polynucleotide synthesizer, followed by ligation with an appropriate DNA ligase and amplification of the ligated nucleotide sequence via PCR. See, e.g., Jayaraman et al., Proc. Natl. Acad. Sci. USA (1991) 88:4084-4088. Additionally, oligonucleotide- directed synthesis (Jones et al., Nature (1986) 54:75-82), oligonucleotide directed mutagenesis of preexisting nucleotide regions (Riechmann et al., Nature (1988) 332:323-327 and Verhoeyen et al., Science (1988) 239: 1534-1536), and enzymatic filling-in of gapped oligonucleotides using T4 DNA polymerase (Queen et al., Proc. Natl. Acad. Sci. USA (1989) 86: 10029-10033) can be used.
[0426] The precursor RNA can be generated by incubating a DNA template under conditions permissive of transcription of the precursor RNA encoded by the DNA template. For example, in some embodiments a precursor RNA is synthesized by incubating a DNA template provided herein that comprises an RNA polymerase promoter upstream of its 5' duplex sequence and/or expression sequences with a compatible RNA polymerase enzyme under conditions permissive of in vitro transcription. In some embodiments, the DNA template is incubated inside of a cell by a bacteriophage RNA polymerase or in the nucleus of a cell by host RNA polymerase II.
C. PRODUCTION OF POLYMER LIPID NANQPARTICLES
[0427] Provided herein are methods for preparing a composition comprising the subject polymeric lipid nanoparticles. In one embodiment, the polymeric lipid nanoparticles are formed by providing an organic solution comprising one or more amphiphilic polymers and one or more ionizable lipids and contacting the solution with a polymer nonsolvent (e.g., water) to produce the polymeric lipid nanoparticles. The organic solution may be partially miscible or immiscible with the polymer nonsolvent. For example, a partially water miscible organic solution may contain the polymers/ionizable lipids, and polymeric lipid nanoparticles are formed as the partially water miscible solution is contacted with water, a polymer nonsolvent, e.g., by pouring the partially water miscible solution into the water at a controlled rate. The polymer contained within the organic solution, upon contact with the polymer nonsolvent, may then precipitate to form particles such as nanoparticles. Typically, an organic solution (e.g., dichloromethane, acetonitrile, chloroform, tetrahydrofuran, acetone, formamide, dimethylformamide, pyridines, dioxane, dimethylsulfoxide, etc.) and an aqueous liquid (e.g., water, or water containing dissolved salts or other species, cell or biological media, ethanol, etc.) are immiscible or only partially miscible with respect to each other. In some embodiments, the rate of introduction of the organic solution into the aqueous solution is carefully controlled and kept at a relatively slow rate.
[0428] Properties such as surface functionality, surface charge, size, zeta ( £) potential, hydrophobicity, ability to control immunogenicity, and the like, may be highly controlled using the disclosed process. For instance, a library of polymeric lipid nanoparticles may be synthesized, and screened to identify the particles having a particular ratio of polymers that allows the particles to have a specific density of moieties present on the surface of the particle. This allows particles having one or more specific properties to be prepared, for example, a specific size and a specific surface density of moieties, without an undue degree of effort. Accordingly, certain embodiments are directed to screening techniques using such libraries, as well as any particles identified using such libraries. In addition, identification may occur by any suitable method. For instance, the identification may be direct or indirect, or proceed quantitatively or qualitatively.
[0429] In some embodiments a circRNA, one or more ionizable lipids, and one or more amphiphilic polymers, may be combined with an organic solution to form a first organic phase. In other processes, the circRNA is added to the aqueous phase, and the ionizable lipid and amphiphilic polymer are combined in an organic phase. The organic phase may be combined with the aqueous solution to form a second phase. The organic solution can include, for example, toluene, methyl ethyl ketone, acetonitrile, tetrahydrofuran, ethyl acetate, isopropyl alcohol, isopropyl acetate, dimethylformamide, dichloromethane, chloroform, acetone, benzyl alcohol, or the like, and combinations thereof. In one
embodiment, the organic phase may include benzyl alcohol, ethyl acetate, and combinations thereof. The aqueous solution can be water, optionally in combination with one or more of sodium cholate, polysorbate 80, sorbitan monooleate ethyl acetate, polyvinyl acetate and benzyl alcohol. In some embodiments, the pH of the aqueous phase may be selected and adjusted based on the pKa of the circRNA.
[0430] In one embodiment, provided herein is a method of preparing a composition comprising a plurality of polymeric lipid nanoparticles, the method comprising: a) providing a mixture comprising: an ionizable lipid; and an amphiphilic polymer, both dissolved in an organic phase; b) combining the mixture with a circular RNA polynucleotide dissolved in an aqueous phase to obtain an emulsion; and c) adding the emulsion to an aqueous bath (e.g., 8x-10x) to form polymeric lipid nanoparticles comprising the ionizable lipid, the amphiphilic polymer, and the circular RNA polynucleotide.
[0431] In some embodiments of the method of preparing a subject composition, the aqueous phase in step b) includes one or more surfactants. In some embodiments of the method of preparing a subject composition, the aqueous phase includes one or more dissolved organic solvents, such that the aqueous phase is saturated with the organic solvents. In some embodiments, the dissolved organic solvents are selected from benzyl alcohol, ethyl acetate, and combinations thereof.
[0432] In some embodiments, the organic phase may use a solvent that is only partially miscible with the aqueous phase (nonsolvent). Therefore, when mixed at a low enough ratio and/or when using water pre-saturated with the organic solvents, the oil phase remains liquid. The oil phase may be emulsified into an aqueous solution and, as liquid droplets, sheared into nanoparticles using, for example, high energy dispersion systems, such as homogenizers or sonicators. The aqueous portion (aqueous phase) of the emulsion may be a surfactant solution consisting of a surfactant (e.g., sodium cholate, polysorbate 80, or sorbitan monooleate) and pre-saturated with partially water miscible organic solvents (e.g., ethyl acetate and benzyl alcohol). In some embodiments, the organic phase may include the circRNA. In some embodiments, the circRNA, ionizable lipid and amphiphilic polymer, may be dissolved in the organic phase. In some embodiments, the circRNA may be dissolved in the aqueous phase.
[0433] Emulsifying the aqueous phase to form an emulsion phase may be performed in one or two emulsification steps. For example, a primary coarse emulsion may be prepared, and then emulsified in a secondary emulsification step to form a fine emulsion. The primary emulsion can be formed, for
example, using simple mixing, a high-pressure homogenizer, probe sonicator, stir bar, or a rotor stator homogenizer. The primary emulsion may be formed into a fine emulsion through the use of e.g., probe sonicator or a high-pressure homogenizer, e.g., by using 1, 2, 3, or more passes through a homogenizer. For example, when a high pressure homogenizer is used, the pressure used may be 30 to 60 psi, 40 to 50 psi, 1000 to 8000 psi, 2000 to 4000 psi, 4000 to 8000 psi, 4000 to 5000 psi, 9,000 to 10,000 psi, and up to 30,000 psi, e.g., 2000 psi, 2500 psi, 4000 psi, or 5000 psi.
[0434] In some embodiments of the method of preparing a subject composition, in step b) the ionizable lipid forms a complex with the circular RNA polynucleotide (circRNA-lipid complex).
[0435] In some embodiments, fine emulsion conditions, which can be characterized by a very high surface to volume ratio of the droplets in the emulsion, can be chosen to maximize the solubility of the circRNA-lipid complex. In certain embodiments, under fine emulsion conditions, equilibration of dissolved components can occur very quickly, i.e., faster than solidification of the nanoparticles.
[0436] In some embodiments, the ionizable lipid forms a complex with the circRNA (circRNA-lipid complex) and the ionizable lipid during the primary coarse emulsion step. In some embodiments, the circRNA-lipid complex is formed during a second emulsion step to form a fine emulsion.
[0437] In some embodiments of the method of preparing a subject composition, in step c) the fine emulsion is quenched by adding to a larger quantity of water in an aqueous bath to from solid polymeric lipid nanoparticles comprising the circRNA-lipid complex encapsulated in an amphiphilic polymer matrix. In some embodiments, the aqueous bath is 10 times or more the volume of the fine emulsion, such as 5 times or more, 6 times or more, 7 times or more, 8 times or more, 9 times or more, or 10 times or more. In some embodiments, the aqueous bath is 8-10 times the volume of the fine emulsion, such as 8 times, 9 times or 10 times the volume of the fine emulsion.
[0438] In some embodiments, the emulsion can be diluted into cold water to a concentration sufficient to dissolve all of the organic solvent to form the quenched phase. In some embodiments, quenching may be performed at least partially at a temperature of about 5 °C. or less. For example, water used in the quenching may be at a temperature that is less that room temperature (e.g., 0 to 10°C, or 0 to 5 °C). In certain embodiments, the quench may be chosen having a pH that is advantageous for quenching the emulsion phase, e.g., by improving the properties of the polymeric lipid nanoparticles, such as the release profile, or improving a nanoparticle parameter, such as the drug loading. The pH of the quench may be adjusted by acid or base titration, for example, or by appropriate selection of a buffer. In some embodiments, the pH of the quench may be selected based on the pKa of the circRNA or the ionizable lipid.
[0439] In some embodiments, the method of preparing a subject composition further comprises separating the polymeric lipid nanoparticles from the aqueous bath.
[0440] In some embodiments, not all of the circRNA is encapsulated in the particles at the quenching stage, and a drug solubilizer is optionally added to form a solubilized phase. The drug solubilizer may be for example, polysorbate 80, polysorbate 20, polyvinyl pyrrolidone, cyclodextran, sodium dodecyl sulfate, sodium cholate, diethylnitrosamine, sodium acetate, urea, glycerin, propylene glycol, glycofurol, poly(ethylene)glycol, bris(polyoxyethyleneglycolddodecyl ether, sodium benzoate, sodium salicylate, or combinations thereof. For example, polysorbate 80 may be added to the quenched nanoparticle suspension to solubilize the free drug and prevent the formation of drug crystals.
[0441] The solubilized phase may be filtered to recover the solid polymeric lipid nanoparticles. For example, ultrafiltration membranes may be used to concentrate the polymeric lipid nanoparticle suspension and substantially eliminate organic solvent, free drug (i.e., unencapsulated circRNA), drug solubilizer, and other processing aids (surfactants). Exemplary filtration may be performed using a tangential flow filtration system.
[0442] After concentrating the nanoparticle suspension, the polymeric lipid nanoparticles can be collected and stored frozen in a sucrose solution. In some embodiments, the sucrose solution is a 10- 30 wt% sucrose solution, such as 10-15 wt%, 15-20 wt%, 20-25 wt%, or a 25-30 wt% sucrose solution.
[0443] It will be appreciated that the amounts of amphiphilic polymer, circRNA, and ionizable lipid that are used in the preparation of the formulation may differ from a final formulation. For example, some of the circRNA may not become completely incorporated in a polymeric lipid nanoparticle and such free circRNA can be removed e.g., by filtering away.
[0444] In some embodiments of the method of preparing a subject composition, in step c) the circRNA- lipid complex is encapsulated in the nanoparticles with an encapsulation efficiency of at least 70%, such as at least 75%, at least 80%, at least 85%, at least 90% or at least 95%.
[0445] In some embodiments of the method of preparing a subject composition, in step c) the circRNA- lipid complex is encapsulated in the nanoparticles with an encapsulation efficiency of 70-90%, such as 75-90%, 80-90%, or 85-90%.
[0446] A polymeric lipid nanoparticle composition may optionally comprise one or more coatings. For example, a nanoparticle composition may be formulated in a capsule, film, or tablet having a coating. A capsule, film, or tablet including a composition described herein may have any useful size, tensile strength, hardness, or density.
[0447] In some embodiments, the polymeric lipid nanoparticles described herein may be synthesized using methods comprising microfluidic mixers and/or high-pressure homogenizers (e.g., microfluidizer). Exemplary microfluidic mixers may include, but are not limited to, a slit interdigitial micromixer including, but not limited to, those manufactured by Precision Nanosystems (Vancouver, BC, Canada), Microinnova (Allerheiligen bei Wildon, Austria) and/or a staggered herringbone micromixer (SHM) (Zhigaltsev, I.V. et al. (2012) Langmuir. 28:3633-40; Belliveau, N.M. et al. Mol. Ther. Nucleic. Acids. (2012) l:e37; Chen, D. et al. J. Am. Chem. Soc. (2012) 134(16):6948-51; each of which is herein incorporated by reference in its entirety).
[0448] In some embodiments, methods of polymeric lipid nanoparticle generation comprising SHM, further comprise the mixing of at least two input streams wherein mixing occurs by microstructure- induced chaotic advection (MICA). According to this method, fluid streams flow through channels present in a herringbone pattern causing rotational flow and folding the fluids around each other. This method may also comprise a surface for fluid mixing wherein the surface changes orientations during fluid cycling.
[0449] In one embodiment, the polymeric lipid nanoparticles may be formulated using a micromixer such as, but not limited to, a Slit Interdigital Microstructured Mixer (SIMM-V2) or a Standard Slit Interdigital Micro Mixer (SSIMM) or Caterpillar (CPMM) or Impinging-jet (IJMM)from the Institut fur Mikrotechnik Mainz GmbH, Mainz Germany). In one embodiment, the lipid nanoparticles are created using microfluidic technology (see, Whitesides (2006) Nature. 442: 368-373; and Abraham et al. (2002) Science. 295: 647-651; each of which is herein incorporated by reference in its entirety). As a non-limiting example, controlled microfluidic formulation includes a passive method for mixing streams of steady pressure-driven flows in micro channels at a low Reynolds number (see, e.g., Abraham et al. (2002) Science. 295: 647651, which is herein incorporated by reference in its entirety).
[0450] In one embodiment, the circRNA of the present invention may be formulated in polymeric lipid nanoparticles created using a micromixer chip such as, but not limited to, those from Harvard Apparatus (Holliston, MA), Dolomite Microfluidics (Royston, UK), or Precision Nanosystems (Van Couver, BC, Canada). A micromixer chip can be used for rapid mixing of two or more fluid streams with a split and recombine mechanism.
[0451] In certain embodiments, the polymeric lipid nanoparticle may be formulated with one or more triantennary N-acetyl galactosamine (i.e., GalNAc3). In some embodiments, the GalNAc3 is conjugated, appended and/or affixed to the polymeric lipid nanoparticle. In some embodiments, the PLNP has a GalNAc3 attached to its surface. In certain embodiments, GalNAc3 comprises a PEGylated linker and/or diarylcyclooctyne (DBCO) functional group. In some embodiments, the DBCO functional group of the GalNAc3 may chemically react with a polylactide-block-PEG (PLA-PEG) (e.g., via
copper-free azide-alkyne click chemistry). In some embodiments, the PLNP comprises a PLA-PEG lipid comprising one or more azide group (N3). In some embodiments, the PLA-PEG of the PLNP reacts with GalNAc3 to conjugate, append and/or become affixed to the surface of the PLNP. In some embodiments, GalNAc3 interacts with the ASGPR receptor on hepatocytes to facilitate intracellular uptake in vivo. In some embodiments, the PLNP is unconjugated (i.e., the PLNP does not have a GalNAc3 affixed to its surface and/or bonded to an attached PEG).
3. PHARMACEUTICAL COMPOSITIONS
[0452] In certain embodiments, provided herein are compositions (e.g., pharmaceutical compositions) comprising a therapeutic agent provided herein. In some embodiments, the therapeutic agent is a circular RNA polynucleotide. In some embodiments the therapeutic agent is a vector. In some embodiments, the therapeutic agent is a cell comprising a circular RNA or vector (e.g., a human cell, such as a human T cell). In certain embodiments, the composition further comprises a pharmaceutically acceptable carrier. In some embodiments, the compositions provided herein comprise a therapeutic agent provided herein in combination with other pharmaceutically active agents or drugs, such as antiinflammatory drugs or antibodies capable of targeting B cell antigens, e.g., anti-CD20 antibodies, e.g., rituximab.
[0453] With respect to pharmaceutical compositions, the pharmaceutically acceptable carrier can be any of those conventionally used and is limited only by chemical-physical considerations, such as solubility and lack of reactivity with the active agent(s), and by the route of administration. The pharmaceutically acceptable carriers described herein, for example, vehicles, adjuvants, excipients, and diluents, are well-known to those skilled in the art and are readily available to the public. It is preferred that the pharmaceutically acceptable carrier be one which is chemically inert to the therapeutic agent(s) and one which has no detrimental side effects or toxicity under the conditions of use.
[0454] The choice of carrier will be determined in part by the particular therapeutic agent, as well as by the particular method used to administer the therapeutic agent. Accordingly, there are a variety of suitable formulations of the pharmaceutical compositions provided herein.
[0455] In certain embodiments, the pharmaceutical composition comprises a preservative. In certain embodiments, suitable preservatives may include, for example, methylparaben, propylparaben, sodium benzoate, and benzalkonium chloride. Optionally, a mixture of two or more preservatives may be used. The preservative or mixtures thereof are typically present in an amount of 0.0001% to 2% by weight of the total composition.
[0456] In some embodiments, the pharmaceutical composition comprises a buffering agent. In some embodiments, suitable buffering agents may include, for example, citric acid, sodium citrate,
phosphoric acid, potassium phosphate, and various other acids and salts. A mixture of two or more buffering agents optionally may be used. The buffering agent or mixtures thereof are typically present in an amount of 0.001% to 4% by weight of the total composition.
[0457] In some embodiments, the concentration of therapeutic agent in the pharmaceutical composition can vary, e.g., less than 1%, or at least 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9% 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, or 50% or more by weight, and can be selected primarily by fluid volumes, and viscosities, in accordance with the particular mode of administration selected.
[0458] The following formulations for oral, aerosol, parenteral (e.g., subcutaneous, intravenous, intraarterial, intramuscular, intradermal, intraperitoneal, and intrathecal), and topical administration are merely exemplary and are in no way limiting. More than one route can be used to administer the therapeutic agents provided herein, and in certain instances, a particular route can provide a more immediate and more effective response than another route.
[0459] Formulations suitable for oral administration can comprise or consist of (a) liquid solutions, such as an effective amount of the therapeutic agent dissolved in diluents, such as water, saline, or orange juice; (b) capsules, sachets, tablets, lozenges, and troches, each containing a predetermined amount of the active ingredient, as solids or granules; (c) powders; (d) suspensions in an appropriate liquid; and (e) suitable emulsions. Liquid formulations may include diluents, such as water and alcohols, for example, ethanol, benzyl alcohol and the polyethylene alcohols, either with or without the addition of a pharmaceutically acceptable surfactant. Capsule forms can be of the ordinary hard- or soft-shelled gelatin type containing, for example, surfactants, lubricants, and inert fillers, such as lactose, sucrose, calcium phosphate, and corn starch. Tablet forms can include one or more of lactose, sucrose, mannitol, corn starch, potato starch, alginic acid, microcrystalline cellulose, acacia, gelatin, guar gum, colloidal silicon dioxide, croscarmellose sodium, talc, magnesium stearate, calcium stearate, zinc stearate, stearic acid, and other excipients, colorants, diluents, buffering agents, disintegrating agents, moistening agents, preservatives, flavoring agents, and other pharmacologically compatible excipients. Lozenge forms can comprise the therapeutic agent with a flavorant, usually sucrose, acacia or tragacanth. Pastilles can comprise the therapeutic agent with an inert base, such as gelatin and glycerin, or sucrose and acacia, emulsions, gels, and the like containing, in addition to, such excipients as are known in the art.
[0460] Formulations suitable for parenteral administration include aqueous and nonaqueous isotonic sterile injection solutions, which can contain antioxidants, buffers, bacteriostats, and solutes that render the formulation isotonic with the blood of the intended recipient, and aqueous and nonaqueous sterile suspensions that can include suspending agents, solubilizers, thickening agents, stabilizers, and preservatives. In some embodiments, the therapeutic agents provided herein can be administered in a
physiologically acceptable diluent in a pharmaceutical carrier, such as a sterile liquid or mixture of liquids including water, saline, aqueous dextrose and related sugar solutions, an alcohol such as ethanol or hexadecyl alcohol, a glycol such as propylene glycol or polyethylene glycol, dimethylsulfoxide, glycerol, ketals such as 2,2-dimethyl-l,3-dioxolane-4-methanol, ethers, poly(ethyleneglycol) 400, oils, fatty acids, fatty acid esters or glycerides, or acetylated fatty acid glycerides with or without the addition of a pharmaceutically acceptable surfactant such as a soap or a detergent, suspending agent such as pectin, carbomers, methylcellulose, hydroxypropylmethylcellulose, or carboxymethylcellulose, or emulsifying agents and other pharmaceutical adjuvants.
[0461] Oils, which can be used in parenteral formulations in some embodiments, include petroleum, animal oils, vegetable oils, or synthetic oils. Specific examples of oils include peanut, soybean, sesame, cottonseed, corn, olive, petrolatum, and mineral oil. Suitable fatty acids for use in parenteral formulations include oleic acid, stearic acid, and isostearic acid. Ethyl oleate and isopropyl myristate are examples of suitable fatty acid esters.
[0462] Suitable soaps for use in certain embodiments of parenteral formulations include fatty alkali metal, ammonium, and triethanolamine salts, and suitable detergents include (a) cationic detergents such as, for example, dimethyl dialkyl ammonium halides, and alkyl pyridinium halides, (b) anionic detergents such as, for example, alkyl, aryl, and olefin sulfonates, alkyl, olefin, ether, and monoglyceride sulfates, and sulfosuccinates, (c) nonionic detergents such as, for example, fatty amine oxides, fatty acid alkanolamides, and polyoxyethylenepolypropylene copolymers, (d) amphoteric detergents such as, for example, alkyl-P-aminopropionates, and 2-alkyl-imidazoline quaternary ammonium salts, and (e) mixtures thereof.
[0463] In some embodiments, the parenteral formulations will contain, for example, from 0.5% to 25% by weight of the therapeutic agent in solution. Preservatives and buffers may be used. In order to minimize or eliminate irritation at the site of injection, such compositions may contain one or more nonionic surfactants having, for example, a hydrophile-lipophile balance (HLB) of from 12 to 17. The quantity of surfactant in such formulations will typically range, for example, from 5% to 15% by weight. Suitable surfactants include polyethylene glycol, sorbitan, fatty acid esters such as sorbitan monooleate, and high molecular weight adducts of ethylene oxide with a hydrophobic base formed by the condensation of propylene oxide with propylene glycol. The parenteral formulations can be presented in unit-dose or multi-dose sealed containers, such as ampoules or vials, and can be stored in a freeze- dried (lyophilized) condition requiring only the addition of a sterile liquid excipient, for example, water for injections, immediately prior to use. Extemporaneous injection solutions and suspensions can be prepared from sterile powders, granules, and tablets of the kind previously described.
[0464] In certain embodiments, injectable formulations are provided herein. The requirements for effective pharmaceutical carriers for injectable compositions are well-known to those of ordinary skill in the art (see, e.g., Pharmaceutics and Pharmacy Practice, J.B. Lippincott Company, Philadelphia, PA, Banker and Chalmers, eds., pages 238-250 (1982), and ASHP Handbook on Injectable Drugs, Toissel, 4th ed, pages 622-630 (1986)).
[0465] In some embodiments, the therapeutic agents provided herein are formulated in time -released, delayed release, or sustained release delivery systems such that the delivery of the composition occurs prior to, and with sufficient time to, cause sensitization of the site to be treated. Such systems can avoid repeated administrations of the therapeutic agent, thereby increasing convenience to the subject and the physician, and may be particularly suitable for certain composition embodiments provided herein. In one embodiment, the compositions of the invention are formulated such that they are suitable for extended release of the circRNA contained therein. Such extended-release compositions may be conveniently administered to a subject at extended dosing intervals. For example, in one embodiment, the compositions of the present invention are administered to a subject twice a day, daily or every other day. In an embodiment, the compositions of the present invention are administered to a subject twice a week, once a week, every ten days, every two weeks, every three weeks, every four weeks, once a month, every six weeks, every eight weeks, every three months, every four months, every six months, every eight months, every nine months or annually.
[0466] In some embodiments, a protein encoded by a circRNA is produced by a target cell for sustained amounts of time. For example, the protein may be produced for more than one hour, more than four, more than six, more than 12, more than 24, more than 48 hours, or more than 72 hours after administration. In some embodiments the polypeptide is expressed at a peak level six hours after administration. In some embodiments the expression of the polypeptide is sustained at least at a therapeutic level. In some embodiments, the polypeptide is expressed at least at a therapeutic level for more than one, more than four, more than six, more than 12, more than 24, more than 48, or more than 72 hours after administration. In some embodiments, the polypeptide is detectable at a therapeutic level in patient tissue (e.g., liver or lung). In some embodiments, the level of detectable polypeptide is from continuous expression from the circRNA composition over periods of time of more than one, more than four, more than six, more than 12, more than 24, more than 48, or more than 72 hours after administration.
[0467] In certain embodiments, a protein encoded by circRNA is produced at levels above normal physiological levels. The level of protein may be increased as compared to a control. In some embodiments, the control is the baseline physiological level of the polypeptide in a normal individual or in a population of normal individuals. In other embodiments, the control is the baseline physiological
level of the polypeptide in an individual having a deficiency in the relevant protein or polypeptide or in a population of individuals having a deficiency in the relevant protein or polypeptide. In some embodiments, the control can be the normal level of the relevant protein or polypeptide in the individual to whom the composition is administered. In other embodiments, the control is the expression level of the polypeptide upon other therapeutic intervention, e.g., upon direct injection of the corresponding polypeptide, at one or more comparable time points.
[0468] In certain embodiments, the levels of a protein encoded by a circRNA are detectable at 3 days, 4 days, 5 days, or 1 week or more after administration. Increased levels of protein may be observed in a tissue (e.g., liver or lung).
[0469] In some embodiments, the method yields a sustained circulation half-life of a protein encoded by a circRNA. For example, the protein may be detected for hours or days longer than the half-life observed via subcutaneous injection of the protein or mRNA encoding the protein. In some embodiments, the half-life of the protein is 1 day, 2 days, 3 days, 4 days, 5 days, or 1 week or more.
[0470] Many types of release delivery systems are available and known to those of ordinary skill in the art. They include polymer-based systems such as poly(lactide-glycolide), copoly oxalates, polycaprolactones, polyesteramides, poly orthoesters, polyhydroxybutyric acid, and polyanhydrides. Microcapsules of the foregoing polymers containing drugs are described in, for example, U.S. Patent 5,075,109. Delivery systems also include non-polymer systems that are lipids including sterols such as cholesterol, cholesterol esters, and fatty acids or neutral fats such as mono-di-and tri-glycerides; hydrogel release systems; sylastic systems; peptide based systems: wax coatings; compressed tablets using conventional binders and excipients; partially fused implants; and the like. Specific examples include but are not limited to: (a) erosional systems in which the active composition is contained in a form within a matrix such as those described in U.S. Patents 4,452,775, 4,667,014, 4,748,034, and 5,239,660 and (b) diffusional systems in which an active component permeates at a controlled rate from a polymer such as described in U.S. Patents 3,832,253 and 3,854,480. In addition, pump-based hardware delivery systems can be used, some of which are adapted for implantation.
[0471] In some embodiments, the therapeutic agent can be conjugated either directly or indirectly through a linking moiety to a targeting moiety. Methods for conjugating therapeutic agents to targeting moieties is known in the art. See, for instance, Wadwa et al., J, Drug Targeting 3:111 (1995) and U.S. Patent 5,087,616.
[0472] In some embodiments, the therapeutic agents provided herein are formulated into a depot form, such that the manner in which the therapeutic agent is released into the body to which it is administered is controlled with respect to time and location within the body (see, for example, U.S. Patent 4,450,150).
Depot forms of therapeutic agents can be, for example, an implantable composition comprising the therapeutic agents and a porous or non-porous material, such as a polymer, wherein the therapeutic agents are encapsulated by or diffused throughout the material and/or degradation of the non-porous material. The depot is then implanted into the desired location within the body and the therapeutic agents are released from the implant at a predetermined rate.
[473] The present disclosure also contemplates the discriminatory targeting of target cells and tissues by both passive and active targeting means. The phenomenon of passive targeting exploits the natural distributions patterns of a transfer vehicle in vivo without relying upon the use of additional excipients or means to enhance recognition of the transfer vehicle by target cells. For example, transfer vehicles which are subject to phagocytosis by the cells of the reticulo-endothelial system are likely to accumulate in the liver or spleen, and accordingly may provide a means to passively direct the delivery of the subject compositions to such target cells.
[474] Alternatively, the present disclosure contemplates active targeting, which involves the use of targeting moieties that may be bound (either covalently or non-covalently) to the polymeric lipid nanoparticle to encourage localization of such at certain target cells or target tissues. For example, targeting may be mediated by the inclusion of one or more endogenous targeting moieties in or on the polymeric lipid nanoparticle to encourage distribution to the target cells or tissues. Recognition of the targeting moiety by the target tissues actively facilitates tissue distribution and cellular uptake of the polymeric lipid nanoparticle and/or its contents in the target cells and tissues (e.g., the inclusion of an apolipoprotein-E targeting ligand encourages recognition and binding of the polymeric lipid nanoparticle o endogenous low density lipoprotein receptors expressed by hepatocytes). As provided herein, the composition can comprise a moiety capable of enhancing affinity of the composition to the target cell. Targeting moieties may be linked to the outer polymeric layer of the polymeric lipid nanoparticle during formulation or post-formulation. In addition, some polymeric lipid nanoparticle formulations may employ fusogenic polymers such as PEAA, hemagluttinin, other lipopeptides (see U.S. patent application Ser. No. 08/835,281, and 60/083,294, which are incorporated herein by reference) and other features useful for in vivo and/or intracellular delivery. In some embodiments, the compositions of the present disclosure demonstrate improved transfection efficacies, and/or demonstrate enhanced selectivity towards target cells or tissues of interest. Contemplated therefore are compositions which comprise one or more moieties (e.g., peptides, aptamers, oligonucleotides, vitamins or other molecules) that are capable of enhancing the affinity of the compositions and their nucleic acid contents for the target cells or tissues. Suitable moieties may optionally be bound or linked to the surface of the nanoparticle. In some embodiments, the targeting moiety may span the surface of a nanoparticle or be encapsulated within the nanoparticle. Suitable moieties and are selected based upon their physical, chemical or biological properties (e.g., selective affinity and/or recognition of target cell surface
markers or features). Cell-specific target sites and their corresponding targeting ligand can vary widely. Suitable targeting moieties are selected such that the unique characteristics of a target cell are exploited, thus allowing the composition to discriminate between target and non-target cells. For example, compositions of the present disclosure may include surface markers (e.g., apolipoprotein-B (APOB) or apolipoprotein-E (APOE)) that selectively enhance recognition of, or affinity to hepatocytes (e.g., by receptor-mediated recognition of and binding to such surface markers). As an example, the use of galactose as a targeting moiety would be expected to direct the compositions of the present disclosure to parenchymal hepatocytes, or alternatively the use of mannose containing sugar residues as a targeting ligand would be expected to direct the compositions of the present disclosure to liver endothelial cells (e.g., mannose containing sugar residues that may bind preferentially to the asialoglycoprotein receptor present in hepatocytes). (See Hillery A M, et al. “Drug Delivery and Targeting: For Pharmacists and Pharmaceutical Scientists” (2002) Taylor & Francis, Inc.) The presentation of such targeting moieties that have been conjugated to moieties present in the polymeric lipid nanoparticle composition therefore facilitate recognition and uptake of the compositions of the present disclosure in target cells and tissues. Examples of suitable targeting moieties include one or more peptides, proteins, aptamers, vitamins and oligonucleotides .
[475] In particular embodiments, a polymeric lipid nanoparticle composition comprises a targeting moiety. In some embodiments, the targeting moiety mediates receptor-mediated endocytosis selectively into a specific population of cells. In some embodiments, the targeting moiety is operably connected, or linked, to the transfer vehicle. In some embodiments, the targeting moiety is capable of binding to an immune cell antigen. In some embodiments, the targeting moiety is capable of binding to a T cell antigen. Exemplary T cell antigens include, but are not limited to, CD2, CD3, CD5, CD7, CD8, CD4, beta7 integrin, beta2ingetrin, and ClqR. In some embodiments, the targeting moiety is capable of binding to a NK, NKT, or macrophage antigen. In some embodiments, the targeting moiety is capable of binding to a protein selected from CD3, CD4, CD8, PD-1, 4-1BB, and CD2. In some embodiments, the targeting moiety is a single chain Fv (scFv) fragment, nanobody, peptide, peptide-based macrocycle, minibody, heavy chain variable region, light chain variable region or fragment thereof. In some embodiments, the targeting moiety is selected from T-cell receptor motif antibodies, T-cell a chain antibodies, T-cell P chain antibodies, T-cell y chain antibodies, T-cell 5 chain antibodies, CCR7 antibodies, CD3 antibodies, CD4 antibodies, CD5 antibodies, CD7 antibodies, CD8 antibodies, CDl lb antibodies, CDl lc antibodies, CD16 antibodies, CD19 antibodies, CD20 antibodies, CD21 antibodies, CD22 antibodies, CD25 antibodies, CD28 antibodies, CD34 antibodies, CD35 antibodies, CD40 antibodies, CD45RA antibodies, CD45RO antibodies, CD52 antibodies, CD56 antibodies, CD62L antibodies, CD68 antibodies, CD80 antibodies, CD95 antibodies, CD117 antibodies, CD127 antibodies, CD133 antibodies, CD137 (4-1BB) antibodies, CD163 antibodies, F4/80 antibodies, IL-4Ra antibodies,
Sca-1 antibodies, CTLA-4 antibodies, GITR antibodies GARP antibodies, LAP antibodies, granzyme B antibodies, LFA-1 antibodies, transferrin receptor antibodies, and fragments thereof. In some embodiments, the targeting moiety is a small molecule binder of an ectoenzyme on lymphocytes. Small molecule binders of ectoenzymes include A2A inhibitors CD73 inhibitors, CD39 or adesines receptors A2aR and A2bR. Potential small molecules include AB928.
[476] Where it is desired to deliver a nucleic acid to an immune cell, the immune cell represents the target cell. In some embodiments, the compositions of the present disclosure transfect the target cells on a discriminatory basis (i.e., do not transfect non-target cells). The compositions of the present disclosure may also be prepared to preferentially target a variety of target cells, which include, but are not limited to, T cells, B cells, macrophages, and dendritic cells.
[477] In some embodiments, the target cells are deficient in a protein or enzyme of interest. For example, where it is desired to deliver a nucleic acid to a hepatocyte, the hepatocyte represents the target cell. In some embodiments, the compositions of the present disclosure transfect the target cells on a discriminatory basis (i.e., do not transfect non-target cells). The compositions of the present disclosure may also be prepared to preferentially target a variety of target cells, which include, but are not limited to, hepatocytes, epithelial cells, hematopoietic cells, epithelial cells, endothelial cells, lung cells, bone cells, stem cells, mesenchymal cells, neural cells (e.g., meninges, astrocytes, motor neurons, cells of the dorsal root ganglia and anterior horn motor neurons), photoreceptor cells (e.g., rods and cones), retinal pigmented epithelial cells, secretory cells, cardiac cells, adipocytes, vascular smooth muscle cells, cardiomyocytes, skeletal muscle cells, beta cells, pituitary cells, synovial lining cells, ovarian cells, testicular cells, fibroblasts, B cells, T cells, reticulocytes, leukocytes, granulocytes and tumor cells.
[478] The compositions of the present disclosure may be prepared to preferentially distribute to target cells such as in the heart, lungs, kidneys, liver, and spleen. In some embodiments, the compositions of the present disclosure distribute into the cells of the liver or spleen to facilitate the delivery and the subsequent expression of the circRNA comprised therein by the cells of the liver (e.g., hepatocytes) or the cells of spleen (e.g., immune cells). The targeted cells may function as a biological “reservoir” or “depot” capable of producing, and systemically excreting a functional protein or enzyme. Accordingly, in one embodiment of the present disclosure the transfer vehicle may target hepatocytes or immune cells and/or preferentially distribute to the cells of the liver or spleen upon delivery. In an embodiment, following transfection of the target hepatocytes or immune cells, the circRNA loaded in the nanoparticle are translated and a functional protein product is produced, excreted and systemically distributed. In other embodiments, cells other than hepatocytes (e.g., lung, spleen, heart, ocular, or cells of the central nervous system) can serve as a depot location for protein production.
[479] In one embodiment, the compositions of the present disclosure facilitate a subject's endogenous production of one or more functional proteins and/or enzymes. In an embodiment of the present disclosure, the polymeric lipid nanoparticles comprise circRNA which encode a deficient protein or enzyme. Upon distribution of such compositions to the target tissues and the subsequent transfection of such target cells, the exogenous circRNA loaded into the nanoparticle may be translated in vivo to produce a functional protein or enzyme encoded by the exogenously administered circRNA (e.g., a protein or enzyme in which the subject is deficient). Accordingly, the compositions of the present disclosure exploit a subject's ability to translate exogenously- or recombinantly-prepared circRNA to produce an endogenously-translated protein or enzyme, and thereby produce (and where applicable excrete) a functional protein or enzyme. The expressed or translated proteins or enzymes may also be characterized by the in vivo inclusion of native post-translational modifications which may often be absent in recombinantly-prepared proteins or enzymes, thereby further reducing the immunogenicity of the translated protein or enzyme.
[480] The administration of circRNA encoding a deficient protein or enzyme avoids the need to deliver the nucleic acids to specific organelles within a target cell. Rather, upon transfection of a target cell and delivery of the nucleic acids to the cytoplasm of the target cell, the circRNA contents of a transfer vehicle may be translated and a functional protein or enzyme expressed.
[481] In some embodiments, a circular RNA comprises one or more miRNA binding sites. In some embodiments, a circular RNA comprises one or more miRNA binding sites recognized by miRNA present in one or more non-target cells or non-target cell types (e.g., Kupffer cells or hepatic cells) and not present in one or more target cells or target cell types (e.g., hepatocytes or T cells). In some embodiments, a circular RNA comprises one or more miRNA binding sites recognized by miRNA present in an increased concentration in one or more non-target cells or non-target cell types (e.g., Kupffer cells or hepatic cells) compared to one or more target cells or target cell types (e.g., hepatocytes or T cells). miRNAs are thought to function by pairing with complementary sequences within RNA molecules, resulting in gene silencing.
[482] In some embodiments, the compositions of the present disclosure transfect or distribute to target cells on a discriminatory basis (i.e., do not transfect non-target cells). The compositions of the present disclosure may also be prepared to preferentially target a variety of target cells, which include, but are not limited to, hepatocytes, epithelial cells, hematopoietic cells, epithelial cells, endothelial cells, lung cells, bone cells, stem cells, mesenchymal cells, neural cells (e.g., meninges, astrocytes, motor neurons, cells of the dorsal root ganglia and anterior horn motor neurons), photoreceptor cells (e.g., rods and cones), retinal pigmented epithelial cells, secretory cells, cardiac cells, adipocytes, vascular smooth muscle cells, cardiomyocytes, skeletal muscle cells, beta cells, pituitary cells, synovial lining cells,
ovarian cells, testicular cells, fibroblasts, B cells, T cells, reticulocytes, leukocytes, granulocytes and tumor cells.
4. THERAPEUTIC METHODS
[0483] In certain aspects, provided herein is a method of producing a protein of interest in a subject in need thereof by introducing or administering an effective amount of a pharmaceutical composition described herein comprising at least one polymeric lipid nanoparticle described herein.
[0484] In certain aspects, provided herein is a method of treating and/or preventing a condition comprising administering an effective amount of a pharmaceutical composition described herein comprising at least one polymeric lipid nanoparticle described herein.
[0485] In certain embodiments, the pharmaceutical compositions described herein are co-administered with one or more additional therapeutic agents (e.g., in the same pharmaceutical composition or in separate pharmaceutical compositions). In some embodiments, the pharmaceutical compositions provided herein can be administered first and the one or more additional therapeutic agents can be administered second, or vice versa. Alternatively, the pharmaceutical compositions provided herein and the one or more additional therapeutic agents can be administered simultaneously.
[0486] In some embodiments, the subject is a mammal. In some embodiments, the mammal referred to herein can be any mammal, including, but not limited to, mammals of the order Rodentia, such as mice and hamsters, or mammals of the order Logomorpha, such as rabbits. The mammals may be from the order Carnivora, including Felines (cats) and Canines (dogs). The mammals may be from the order Artiodactyla, including Bovines (cows) and Swines (pigs), or of the order Perssodactyla, including Equines (horses). The mammals may be of the order Primates, Ceboids, or Simoids (monkeys) or of the order Anthropoids (humans and apes). In some embodiments, the mammal is a human.
[0487] In some embodiments, provided herein is a method of vaccinating a subject by introducing or administering an effective amount of a pharmaceutical composition described herein comprising at least one polymeric lipid nanoparticle described herein.
[0488] In some embodiments, provided herein the method of vaccinating comprises administering an effective amount of an antigen comprising a viral polypeptide from an adenovirus; Herpes simplex, type 1; Herpes simplex, type 2; encephalitis virus; papillomavirus; Varicella-zoster virus; Epstein-barr virus; Human cytomegalovirus; Human herpes virus, type 8; Human papillomavirus; BK virus; JC virus; Smallpox; polio virus; Hepatitis B virus; Human bocavirus; Parvovirus B19; Human astrovirus; Norwalk virus; coxsackievirus; hepatitis A virus; poliovirus; rhinovirus; Severe acute respiratory syndrome virus; Hepatitis C virus; Yellow Fever virus; Dengue virus; West Nile virus; Rubella virus;
Hepatitis E virus; Human Immunodeficiency virus (HIV); Influenza virus; Guanarito virus; Junin virus; Lassa virus; Machupo virus; Sabia virus; Crimean-Congo hemorrhagic fever virus; Ebola virus; Marburg virus; Measles virus; Mumps virus; Parainfluenza virus; Respiratory syncytial virus; Human metapneumo virus; Hendra virus; Nipah virus; Rabies virus; Hepatitis D; Rotavirus; Orbivirus; Coltivirus; Banna virus; Human Enterovirus; Hanta virus; West Nile virus; Middle East Respiratory Syndrome Corona Virus; Japanese encephalitis virus; Vesicular exanthema virus; SARS-CoV-2; Eastern equine encephalitis, or a combination of any two or more of the foregoing.
[0489] In some embodiments, provided herein is a method of treating an autoimmune disorder in a subject by introducing or administering an effective amount of a pharmaceutical composition described herein comprising at least one polymeric lipid nanoparticle described herein.
[0490] In some embodiments, provided herein is a method of treating cancer in a subject by introducing or administering an effective amount of a pharmaceutical composition described herein comprising at least one polymeric lipid nanoparticle described herein.
[0491] In some embodiments, the circular RNA construct encodes a CAR, the CARs have biological activity, e.g., ability to recognize an antigen, e.g., CD19, HER2, or BCMA, such that the CAR, when expressed by a cell, is able to mediate an immune response against the cell expressing the antigen, e.g., CD19, HER2, or BCMA, for which the CAR is specific.
[0492] Adoptive T-cell immunotherapy is a rapidly growing field, in particular in cancer treatments. In general, chimeric antigen receptor (CAR) T cell or “CAR-T” engagement of CD19-expressing cancer cells results in T-cell activation, proliferation and secretion of inflammatory cytokines and chemokines resulting in tumor cell lysis. However, while CAR-T therapies have become an important tool in cancer treatments, they have toxic side effects and involve complex procedures. Treatment with CAR-T can lead to a large and rapid release of cytokines into the blood and can cause cytokine release syndrome (CRS) or CAR-T cell-related encephalopathy syndrome (CRES), also referred to as neurotoxicity associated with CAR-T. CRS is the most common and well-described toxicity associated with CAR-T therapy, occurring in over 90% of patients at any grade and is characterized by high fever, hypotension, hypoxia and/ or multiple organ toxicity and can lead to death. Neurotoxicity is characterized by damage to nervous tissue that can cause tremors, encephalopathy, dizziness or seizures. Additionally, prior to infusion, the patients generally undergo lymphodepletion. Lymphodepletion is known to increase CAR- T cell expansion and enhanced efficacy of infused CAR-T cells by, for example, altering the tumor phenotype and microenvironment. However, lymphodepletion agents often cause side effects to the patients. For example, lymphodepletion can cause neutropenia, anemia, thrombocytopenia, and immunosuppression, leading to a greater risk of infection, along with other toxicities. In addition to the toxicities associated with targeted CAR-T therapies, there are procedures, specialized equipment, and
costs involved in producing the modified lymphocytes. CAR-T therapies require an assortment of protocols to isolate, genetically modify, and selectively expand the redirected cells before infusing them back into the patient.
[0493] In some embodiments, the subject has a cancer selected from the group consisting of: acute myeloid leukemia (AML); alveolar rhabdomyosarcoma; B cell malignancies; bladder cancer (e.g., bladder carcinoma); bone cancer; brain cancer (e.g., medulloblastoma and glioblastoma multiforme); breast cancer; cancer of the anus, anal canal, or anorectum; cancer of the eye; cancer of the intrahepatic bile duct; cancer of the joints; cancer of the neck; gallbladder cancer; cancer of the pleura; cancer of the nose, nasal cavity, or middle ear; cancer of the oral cavity; cancer of the vulva; chronic lymphocytic leukemia; chronic myeloid cancer; colon cancer; esophageal cancer, cervical cancer; fibrosarcoma; gastrointestinal carcinoid tumor; head and neck cancer (e.g., head and neck squamous cell carcinoma); Hodgkin lymphoma; hypopharynx cancer; kidney cancer; larynx cancer; leukemia; liquid tumors; lipoma; liver cancer; lung cancer (e.g., non-small cell lung carcinoma, lung adenocarcinoma, and small cell lung carcinoma); lymphoma; mesothelioma; mastocytoma; melanoma; multiple myeloma; nasopharynx cancer; non-Hodgkin lymphoma; B -chronic lymphocytic leukemia; hairy cell leukemia; Burkitt's lymphoma; ovarian cancer; pancreatic cancer; cancer of the peritoneum; cancer of the omentum; mesentery cancer; pharynx cancer; prostate cancer; rectal cancer; renal cancer; skin cancer; small intestine cancer; soft tissue cancer; solid tumors; synovial sarcoma; gastric cancer; teratoma; testicular cancer; thyroid cancer; and ureter cancer.
[0494] In some embodiments, the subject has an autoimmune disorder selected from scleroderma, Grave's disease, Crohn's disease, Sjogren's disease, multiple sclerosis, Hashimoto's disease, psoriasis, myasthenia gravis, autoimmune polyendocrinopathy syndromes, Type I diabetes mellitus (TIDM), autoimmune gastritis, autoimmune uveoretinitis, polymyositis, colitis, thyroiditis, and the generalized autoimmune diseases typified by human Lupus.
5. DEFINITIONS
[0495] As used herein, the terms “circRNA” or “circular polyribonucleotide” or “circular RNA” or “oRNA” are used interchangeably and refers to a single-stranded RNA polynucleotide wherein the 3’ and 5’ ends that are normally present in a linear RNA polynucleotide have been joined together.
[0496] As used herein, the term “DNA template” refers to a DNA sequence capable of transcribing a linear RNA polynucleotide. For example, but not intending to be limiting, a DNA template may include a DNA vector, PCR product or plasmid.
[0497] As used herein, the term “3’ group I intron fragment” refers to a sequence with 75% or higher similarity to the 3 ’-proximal end of a natural group I intron including the splice site dinucleotide and
optionally a stretch of natural exon sequence. As used herein, the term “3’ group II intron fragment” refers to a sequence with 75% or higher similarity to the 3 ’-proximal end of a natural group II intron including the splice site dinucleotide and optionally a stretch of natural exon sequence.
[0498] As used herein, the term “5’ group I intron fragment” refers to a sequence with 75% or higher similarity to the 5 ’-proximal end of a natural group I intron including the splice site dinucleotide and optionally a stretch of natural exon sequence. As used herein, the term “5’ group II intron fragment” refers to a sequence with 75% or higher similarity to the 5 ’-proximal end of a natural group II intron including the splice site dinucleotide and optionally a stretch of natural exon sequence.
[0499] In some embodiments, provided herein are circular RNA polynucleotides comprising a post splicing 3’ group I or II intron fragment (e.g., a stretch of exon sequence), optionally a first spacer, an IRES, an expression sequence, optionally a second spacer, and a post splicing 5’ group I or II intron fragment (e.g., a stretch of exon sequence).
[0500] As used herein, the term “permutation site” refers to the site in a group I or II intron where a cut is made prior to permutation of the intron. This cut generates 3’ and 5’ group I or II intron fragments that are permuted to be on either side of a stretch of precursor RNA to be circularized.
[0501] As used herein, the term “splice site” refers to a dinucleotide that is partially or fully included in a group I intron and between which a phosphodiester bond is cleaved during RNA circularization. (As used herein, “splice site” refers to the dinucleotide or dinucleotides between which cleavage of the phosphodiester bond occurs during a splicing reaction. A “5’ splice site” refers to the natural 5’ dinucleotide of the intron e.g., group I intron, while a “3’ splice site” refers to the natural 3’ dinucleotide of the intron).
[0502] As used herein, the term “expression sequence” refers to a nucleic acid sequence that encodes a product, e.g., a peptide or polypeptide, regulatory nucleic acid, or non-coding nucleic acid. An exemplary expression sequence that codes for a peptide or polypeptide can comprise a plurality of nucleotide triads, each of which can code for an amino acid and is termed as a “codon.”
[0503] As used herein, “coding element” or “coding region” is region located within the expression sequence and encodings for one or more proteins or polypeptides (e.g., therapeutic protein).
[0504] As used herein, a “noncoding element” or “non-coding nucleic acid” is a region located within the expression sequence. This sequence, but itself does not encode for a protein or polypeptide, but may have other regulatory functions, including but not limited, allow the overall polynucleotide to act as a biomarker or adjuvant to a specific cell.
[0505] As used herein, the term “therapeutic protein” refers to any protein that, when administered to a subject directly or indirectly in the form of a translated nucleic acid, has a therapeutic, diagnostic, and/or prophylactic effect and/or elicits a desired biological and/or pharmacological effect.
[0506] As used herein, the term “immunogenic” refers to a potential to induce an immune response to a substance. An immune response may be induced when an immune system of an organism or a certain type of immune cells is exposed to an immunogenic substance. The term “non-immunogenic” refers to a lack of or absence of an immune response above a detectable threshold to a substance. No immune response is detected when an immune system of an organism or a certain type of immune cells is exposed to a non-immunogenic substance. In some embodiments, a non-immunogenic circular polyribonucleotide as provided herein, does not induce an immune response above a pre-determined threshold when measured by an immunogenicity assay. In some embodiments, no innate immune response is detected when an immune system of an organism or a certain type of immune cells is exposed to a non-immunogenic circular polyribonucleotide as provided herein. In some embodiments, no adaptive immune response is detected when an immune system of an organism or a certain type of immune cell is exposed to a non-immunogenic circular polyribonucleotide as provided herein.
[0507] As used herein, the term “circularization efficiency” refers to a measurement of resultant circular polyribonucleotide as compared to its linear starting material.
[0508] As used herein, the term “translation efficiency” refers to a rate or amount of protein or peptide production from a ribonucleotide transcript. In some embodiments, translation efficiency can be expressed as amount of protein or peptide produced per given amount of transcript that codes for the protein or peptide.
[0509] The term “nucleotide” refers to a ribonucleotide, a deoxyribonucleotide, a modified form thereof, or an analog thereof. Nucleotides include species that comprise purines, e.g., adenine, hypoxanthine, guanine, and their derivatives and analogs, as well as pyrimidines, e.g., cytosine, uracil, thymine, and their derivatives and analogs. Nucleotide analogs include nucleotides having modifications in the chemical structure of the base, sugar and/or phosphate, including, but not limited to, 5’ -position pyrimidine modifications, 8’ -position purine modifications, modifications at cytosine exocyclic amines, and substitution of 5-bromo-uracil; and 2’ -position sugar modifications, including but not limited to, sugar-modified ribonucleotides in which the 2’ -OH is replaced by a group such as an H, OR, R, halo, SH, SR, NH2, NHR, NR2, or CN, wherein R is an alkyl moiety as defined herein. Nucleotide analogs are also meant to include nucleotides with bases such as inosine, queuosine, xanthine; sugars such as 2’ -methyl ribose; non-natural phosphodiester linkages such as methylphosphonate, phosphorothioate and peptide linkages. Nucleotide analogs include 5- methoxyuridine, 1 -methylpseudouridine, and 6-methyladenosine.
[0510] The term “nucleic acid” and “polynucleotide” are used interchangeably herein to describe a polymer of any length, e.g., greater than 2 bases, greater than 10 bases, greater than 100 bases, greater than 500 bases, greater than 1000 bases, or up to 10,000 or more bases, composed of nucleotides, e.g., deoxyribonucleotides or ribonucleotides, and may be produced enzymatically or synthetically (e.g., as described in U.S. Pat. No. 5,948,902 and the references cited therein), which can hybridize with naturally occurring nucleic acids in a sequence specific manner analogous to that of two naturally occurring nucleic acids, e.g., can participate in Watson-Crick base pairing interactions. Naturally occurring nucleic acids are comprised of nucleotides including guanine, cytosine, adenine, thymine, and uracil (G, C, A, T, and U respectively).
[0511] The terms “ribonucleic acid” and “RNA” as used herein mean a polymer composed of ribonucleotides.
[0512] The terms “deoxyribonucleic acid” and “DNA” as used herein mean a polymer composed of deoxyribonucleotides .
[0513] ‘ Isolated” or “purified” generally refers to isolation of a substance (for example, in some embodiments, a compound, a polynucleotide, a protein, a polypeptide, a polynucleotide composition, or a polypeptide composition) such that the substance comprises a significant percent (e.g., greater than 1%, greater than 2%, greater than 5%, greater than 10%, greater than 20%, greater than 50%, or more, usually up to 90%-100%) of the sample in which it resides. In certain embodiments, a substantially purified component comprises at least 50%, 80%-85%, or 90%-95% of the sample. Techniques for purifying polynucleotides and polypeptides of interest are well-known in the art and include, for example, ion-exchange chromatography, affinity chromatography and sedimentation according to density. Generally, a substance is purified when it exists in a sample in an amount, relative to other components of the sample, that is more than as it is found naturally.
[0514] The terms “duplexed,” “double-stranded,” or “hybridized” as used herein refer to nucleic acids formed by hybridization of two single strands of nucleic acids containing complementary sequences. In most cases, genomic DNA is double-stranded. Sequences can be fully complementary or partially complementary.
[0515] As used herein, “unstructured” with regard to RNA refers to an RNA sequence that is not predicted by the RNAFold software or similar predictive tools to form a structure (e.g., a hairpin loop) with itself or other sequences in the same RNA molecule. In some embodiments, unstructured RNA can be functionally characterized using nuclease protection assays.
[0516] As used herein, “structured” with regard to RNA refers to an RNA sequence that is predicted by the RNAFold software or similar predictive tools to form a structure (e.g., a hairpin loop) with itself or other sequences in the same RNA molecule.
[0517] As used herein, two “duplex sequences,” “duplex region,” “duplex regions,” “homology arms,” or “homology regions” may be any two regions that are thermodynamically favored to cross-pair in a sequence specific interaction. In some embodiments, two duplex sequences, duplex regions, homology arms, or homology regions, share a sufficient level of sequence identity to one another’s reverse complement to act as substrates for a hybridization reaction. As used herein polynucleotide sequences have “homology” when they are either identical or share sequence identity to a reverse complement or “complementary” sequence. The percent sequence identity between a homology region and a counterpart homology region’s reverse complement can be any percent of sequence identity that allows for hybridization to occur. In some embodiments, an internal duplex region of an inventive polynucleotide is capable of forming a duplex with another internal duplex region and does not form a duplex with an external duplex region.
[0518] As used herein, an “affinity sequence” or “affinity tag” is a region of polynucleotide sequences polynucleotide sequence ranging from 1 nucleotide to hundreds or thousands of nucleotides containing a repeated set of nucleotides for the purposes of aiding purification of a polynucleotide sequence. For example, an affinity sequence may comprise, but is not limited to, a polyA or poly AC sequence.
[0519] As used herein, a “spacer” refers to a region of a polynucleotide sequence ranging from 1 nucleotide to hundreds or thousands of nucleotides separating two other elements along a polynucleotide sequence. The sequences can be defined or can be random. A spacer is typically noncoding. In some embodiments, spacers include duplex regions.
[0520] Linear nucleic acid molecules are said to have a “5 ’-terminus” (5’ end) and a “3 ’-terminus” (3’ end) because nucleic acid phosphodiester linkages occur at the 5’ carbon and 3’ carbon of the sugar moieties of the substituent mononucleotides. The end nucleotide of a polynucleotide at which a new linkage would be to a 5’ carbon is its 5’ terminal nucleotide. The end nucleotide of a polynucleotide at which a new linkage would be to a 3’ carbon is its 3’ terminal nucleotide. A terminal nucleotide, as used herein, is the nucleotide at the end position of the 3’- or 5 ’-terminus.
[0521] As used herein, a “leading untranslated sequence” is a region of polynucleotide sequences ranging from 1 nucleotide to hundreds of nucleotides located at the upmost 5' end of a polynucleotide sequence. The sequences can be defined or can be random. A leading untranslated sequence is noncoding.
[0522] As used herein, a “leading untranslated sequence” is a region of polynucleotide sequences ranging from 1 nucleotide to hundreds of nucleotides located at the downmost 3' end of a polynucleotide sequence. The sequences can be defined or can be random. A leading untranslated sequence is noncoding.
[0523] “Transcription” means the formation or synthesis of an RNA molecule by an RNA polymerase using a DNA molecule as a template. The invention is not limited with respect to the RNA polymerase that is used for transcription. For example, in some embodiments, a T7-type RNA polymerase can be used.
[0524] ‘ ‘Translation” means the formation of a polypeptide molecule by a ribosome based upon an RNA template.
[0525] It is to be understood that the terminology used herein is for the purpose of describing particular embodiments only and is not intended to be limiting. As used in this specification and the appended claims, the singular forms “a,” “an,” and “the” include plural referents unless the content clearly dictates otherwise. Thus, for example, reference to “a cell” includes combinations of two or more cells, or entire cultures of cells; reference to “a polynucleotide” includes, as a practical matter, many copies of that polynucleotide. Unless specifically stated or obvious from context, as used herein, the term “or” is understood to be inclusive. Unless defined herein and below in the reminder of the specification, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which the invention pertains.
[0526] As used herein, the term “encode” refers broadly to any process whereby the information in a polymeric macromolecule is used to direct the production of a second molecule that is different from the first. The second molecule may have a chemical structure that is different from the chemical nature of the first molecule.
[0527] By “co-administering” is meant administering a therapeutic agent provided herein in conjunction with one or more additional therapeutic agents sufficiently close in time such that the therapeutic agent provided herein can enhance the effect of the one or more additional therapeutic agents, or vice versa.
[0528] The terms “treat” and “prevent” as well as words stemming therefrom, as used herein, do not necessarily imply 100% or complete treatment or prevention. Rather, there are varying degrees of treatment or prevention of which one of ordinary skill in the art recognizes as having a potential benefit or therapeutic effect. The treatment or prevention provided by the method disclosed herein can include treatment or prevention of one or more conditions or symptoms of the disease. Also, for purposes
herein, “prevention” can encompass delaying the onset of the disease, or a symptom or condition thereof.
[0529] As used herein, an “internal ribosome entry site” or “IRES” refers to an RNA sequence or structural element ranging in size from 10 nt to 1000 nt or more, capable of initiating translation of a polypeptide in the absence of a typical RNA cap structure. An IRES is typically 500 nt to 700 nt in length.
[0530] As used herein, “aptamer” refers in general to either an oligonucleotide of a single defined sequence or a mixture of said nucleotides, wherein the mixture retains the properties of binding specifically to the target molecule (e.g., eukaryotic initiation factor, 40S ribosome, polyC binding protein, polyA binding protein, polypyrimidine tract-binding protein, argonaute protein family, Heterogeneous nuclear ribonucleoprotein K and La and related RNA-binding protein). Thus, as used herein “aptamer” denotes both singular and plural sequences of nucleotides, as defined hereinabove. The term “aptamer” is meant to refer to a single- or double-stranded nucleic acid which is capable of binding to a protein or other molecule. In general, aptamers preferably comprise 10 to 100 nucleotides, preferably 15 to 40 nucleotides, more preferably 20 to 40 nucleotides, in that oligonucleotides of a length that falls within these ranges are readily prepared by conventional techniques. Optionally, aptamers can further comprise a minimum of approximately 6 nucleotides, preferably 10, and more preferably 14 or 15 nucleotides, that are necessary to effect specific binding.
[0531] An “eukaryotic initiation factor” or “elF” refers to a protein or protein complex used in assembling an initiator tRNA, 40S and 60S ribosomal subunits required for initiating eukaryotic translation.
[0532] As used herein, an “internal ribosome entry site” or “IRES” refers to an RNA sequence or structural element ranging in size from 10 nt to 1000 nt or more, capable of initiating translation of a polypeptide in the absence of a typical RNA cap structure. An IRES is typically 500 nt to 700 nt in length.
[0533] As used herein, a “miRNA site” refers to a stretch of nucleotides within a polynucleotide that is capable of forming a duplex with at least 8 nucleotides of a natural miRNA sequence.
[0534] As used herein, an “endonuclease site” refers to a stretch of nucleotides within a polynucleotide that is capable of being recognized and cleaved by an endonuclease protein.
[0535] As used herein, “bicistronic RNA” refers to a polynucleotide that includes two expression sequences coding for two distinct proteins. These expression sequences can be separated by a
nucleotide sequence encoding a cleavable peptide such as a protease cleavage site. They can also be separated by a ribosomal skipping element.
[0536] As used herein, the term “ribosomal skipping element” refers to a nucleotide sequence encoding a short peptide sequence capable of causing generation of two peptide chains from translation of one RNA molecule. While not wishing to be bound by theory, it is hypothesized that ribosomal skipping elements function by (1) terminating translation of the first peptide chain and re-initiating translation of the second peptide chain; or (2) cleavage of a peptide bond in the peptide sequence encoded by the ribosomal skipping element by an intrinsic protease activity of the encoded peptide, or by another protease in the environment (e.g., cytosol).
[0537] As used herein, the term “co-formulate” refers to a nanoparticle formulation comprising two or more nucleic acids or a nucleic acid and other active drug substance. Typically, the ratios are equimolar or defined in the ratiometric amount of the two or more nucleic acids or the nucleic acid and other active drug substance.
[0538] As used herein, the phrase “ionizable lipid” refers to any of a number of lipid species that carry a net positive charge at a selected pH, such as physiological pH 4 and a neutral charge at other pHs such as physiological pH 7.
[0539] In some embodiments, an ionizable lipid or polymer, e.g., an amphiphilic polymer, disclosed herein comprises one or more cleavable groups. The terms “cleave” and “cleavable” are used herein to mean that one or more chemical bonds (e.g., one or more of covalent bonds, hydrogen-bonds, van der Waals' forces and/or ionic interactions) between atoms in or adjacent to the subject functional group are broken (e.g., hydrolyzed) or are capable of being broken upon exposure to selected conditions (e.g., upon exposure to enzymatic conditions). In certain embodiments, the cleavable group is a disulfide functional group, and in particular embodiments is a disulfide group that is capable of being cleaved upon exposure to selected biological conditions (e.g., intracellular conditions). In certain embodiments, the cleavable group is an ester functional group that is capable of being cleaved upon exposure to selected biological conditions. For example, the disulfide groups may be cleaved enzymatically or by a hydrolysis, oxidation or reduction reaction. Upon cleavage of such disulfide functional group, the one or more functional moieties or groups (e.g., one or more of a head-group and/or a tail-group) that are bound thereto may be liberated. In some embodiments, a cleavable group is bound (e.g., bound by one or more of hydrogen-bonds, van der Waals' forces, ionic interactions and covalent bonds) to one or more functional moieties or groups.
[0540] Compound described herein may also comprise one or more isotopic substitutions. For example, H may be in any isotopic form, including 1H, 2H (D or deuterium), and 3H (T or tritium); C
may be in any isotopic form, including 12C, 13C, and 14C; 0 may be in any isotopic form, including 160 and 180; F may be in any isotopic form, including 18F and 19F; and the like.
[0541] When describing the invention, which may include compounds and pharmaceutically acceptable salts thereof, pharmaceutical compositions containing such compounds and methods of using such compounds and compositions, the following terms, if present, have the following meanings unless otherwise indicated. It should also be understood that when described herein any of the moieties defined forth below may be substituted with a variety of substituents, and that the respective definitions are intended to include such substituted moieties within their scope as set out below. Unless otherwise stated, the term “substituted” is to be defined as set out below. It should be further understood that the terms “groups” and “radicals” can be considered interchangeable when used herein.
[0542] When a range of values is listed, it is intended to encompass each value and sub-range within the range. For example, “Cl-6 alkyl” is intended to encompass, Cl, C2, C3, C4, C5, C6, Cl-6, Cl-5, Cl-4, Cl-3, Cl-2, C2-6, C2-5, C2-4, C2-3, C3-6, C3-5, C3-4, C4-6, C4-5, and C5-6 alkyl.
[0543] It should be noted that the term “head-group” as used describe the compounds of the present invention, and in particular functional groups that comprise such compounds, are used for ease of reference to describe the orientation of one or more functional groups relative to other functional groups. For example, in certain embodiments a hydrophilic head-group (e.g., an amino group) is bound (e.g., by one or more of hydrogen-bonds, van der Waals' forces, ionic interactions and covalent bonds) to a hydrophobic tail-group (e.g., cholesterol).
[0544] In typical embodiments, the present invention is intended to encompass the compounds disclosed herein, and the pharmaceutically acceptable salts, pharmaceutically acceptable esters, tautomeric forms, polymorphs, and prodrugs of such compounds. In some embodiments, the present invention includes a pharmaceutically acceptable addition salt, a pharmaceutically acceptable ester, a solvate (e.g., hydrate) of an addition salt, a tautomeric form, a polymorph, an enantiomer, a mixture of enantiomers, a stereoisomer or mixture of stereoisomers (pure or as a racemic or non-racemic mixture) of a compound described herein.
[0545] Compounds described herein can comprise one or more asymmetric centers, and thus can exist in various isomeric forms, e.g., enantiomers and/or diastereomers. For example, the compounds described herein can be in the form of an individual enantiomer, diastereomer or geometric isomer, or can be in the form of a mixture of stereoisomers, including racemic mixtures and mixtures enriched in one or more stereoisomer. Isomers can be isolated from mixtures by methods known to those skilled in the art, including chiral high pressure liquid chromatography (HPLC) and the formation and crystallization of chiral salts; or preferred isomers can be prepared by asymmetric syntheses. See, for
example, Jacques et al., Enantiomers, Racemates and Resolutions (Wiley Interscience, New York, 1981); Wilen et al., Tetrahedron 33:2725 (1977); Eliel, Stereochemistry of Carbon Compounds (McGraw-Hill, NY, 1962); and Wilen, Tables of Resolving Agents and Optical Resolutions p. 268 (E.L. Eliel, Ed., Univ, of Notre Dame Press, Notre Dame, IN 1972). The invention additionally encompasses compounds described herein as individual isomers substantially free of other isomers, and alternatively, as mixtures of various isomers.
[0546] In certain embodiments the subject compositions (e.g., polymeric lipid nanoparticles) exhibit an enhanced (e.g., increased) ability to transfect one or more target cells. Accordingly, also provided herein are methods of transfecting one or more target cells. Such methods generally comprise the step of contacting the one or more target cells with the compositions disclosed herein such that the one or more target cells are transfected with the circular RNA encapsulated therein. As used herein, the terms “transfect” or “transfection” refer to the intracellular introduction of one or more encapsulated materials (e.g., nucleic acids and/or polynucleotides) into a cell, or preferably into a target cell. The term “transfection efficiency” refers to the relative amount of such encapsulated material (e.g., polynucleotides) up-taken by, introduced into and/or expressed by the target cell which is subject to transfection.
[0547] All nucleotide sequences disclosed herein can represent an RNA sequence or a corresponding DNA sequence. It is understood that deoxythymidine (dT or T) in a DNA is transcribed into a uridine (U) in an RNA. As such, “T” and “U” are used interchangeably herein in nucleotide sequences.
[0548] The recitations “sequence identity” or, for example, comprising a “sequence 50% identical to,” as used herein, refer to the extent that sequences are identical on a nucleotide-by-nucleotide basis or an amino acid-by-amino acid basis over a window of comparison. Thus, a “percentage of sequence identity” may be calculated by comparing two optimally aligned sequences over the window of comparison, determining the number of positions at which the identical nucleic acid base (e.g., A, T, C, G, I) or the identical amino acid residue (e.g., Ala, Pro, Ser, Thr, Gly, Vai, Leu, He, Phe, Tyr, Trp, Lys, Arg, His, Asp, Glu, Asn, Gin, Cys and Met) occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison (i.e., the window size), and multiplying the result by 100 to yield the percentage of sequence identity. Included are nucleotides and polypeptides having at least 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity to any of the reference sequences described herein, typically where the polypeptide variant maintains at least one biological activity of the reference polypeptide.
[0549] The expression sequences in the polynucleotide construct may be separated by a “cleavage site” sequence which enables polypeptides encoded by the expression sequences, once translated, to be expressed separately by the cell.
[0550] A “self-cleaving peptide” refers to a peptide which is translated without a peptide bond between two adjacent amino acids, or functions such that when the polypeptide comprising the proteins and the self-cleaving peptide is produced, it is immediately cleaved or separated into distinct and discrete first and second polypeptides without the need for any external cleavage activity.
[0551] The a and P chains of aP TCR's are generally regarded as each having two domains or regions, namely variable and constant domains/regions. The variable domain consists of a concatenation of variable regions and joining regions. In the present specification and claims, the term “TCR alpha variable domain” therefore refers to the concatenation of TRAV and TRAJ regions, and the term TCR alpha constant domain refers to the extracellular TRAC region, or to a C-terminal truncated TRAC sequence. Likewise, the term “TCR beta variable domain” refers to the concatenation of TRBV and TRBD/TRBJ regions, and the term TCR beta constant domain refers to the extracellular TRBC region, or to a C-terminal truncated TRBC sequence.
[0552] The terms “duplexed,” “double-stranded,” or “hybridized” as used herein refer to nucleic acids formed by hybridization of two single strands of nucleic acids containing complementary sequences. In most cases, genomic DNA is double-stranded. Sequences can be fully complementary or partially complementary.
[0553] As used herein, a “vaccine” refers to a composition for generating immunity for the prophylaxis and/or treatment of diseases. Accordingly, vaccines are medicaments which comprise antigens and are intended to be used in humans or animals for generating specific defense and protective substances upon administration to the human or animal.
EXAMPLES
[554] Wesselhoeft et al., (2019) RNA Circularization Diminishes Immunogenicity and Can Extend Translation Duration In vivo. Molecular Cell. 74(3), 508-520 and Wesselhoeft et al., (2018) Engineering circular RNA for Potent and Stable Translation in Eukaryotic Cells. Nature Communications. 9, 2629 are incorporated by reference in their entirety.
[555] The present disclosure further includes the following examples that provide those of ordinary skill in the art with a description of how to make and use the various embodiments of the present disclosure. These examples are not intended to limit the scope of what is regarded as the claimed invention.
EXAMPLE 1
[0556] Circular RNA used in polymer lipid nanoparticle (PLNP) compositions as described herein can be prepared and purified according to procedures in Wesselhoeft et al. (PCT/US2020/034418, filed May 22, 2020, published as WO2020237227), and Goodman et al. (PCT/US2021/031629, filed May 10, 2021, published as WO2021226597), the contents of which are hereby incorporated by reference in their entirety for all purposes. Additional polynucleotides, including expression sequences, and lipids are disclosed in WO2019236673; WO2020237227; WO2021113777; WO2021226597; WO2021189059; WO2021236855; WO2022261490; W02023056033; WO2023081526; the contents of which are hereby incorporated by reference in their entireties.
EXAMPLE 2
General procedure for polymer lipid nanoparticle (PLNP) composition.
[0557] The subject polymer lipid nanoparticle composition are manufactured using an emulsion-based method. A general flow of the process is shown in FIG. 1 and FIG. 2.
[0558] A polylactic acid-polyethylene glycol (PLA-PEG) copolymer, along with the ionizable lipid (also referred to herein as a complex forming lipid, CFL, or simply “lipid”) was dissolved in the organic phase. The organic phase was composed of a mixture of solvents (e.g., benzyl alcohol and ethyl acetate). The organic phase was then mixed (using a probe homogenizer) with the aqueous phase at a ratio of 1:3 to 1:7 to achieve a coarse emulsion. The aqueous phase consisted of circRNA, surfactant and some dissolved solvents to saturate the aqueous phase with organic solvents. The coarse emulsion was then passed through a high shear homogenizer. The fine emulsion droplets particle size was controlled using the surfactant concentration and pressure on the homogenizer. The fine emulsion was then quenched into 8x-10x larger amount of water. The unencapsulated circRNA, ionizable lipid (CFL), surfactant and the processing solvents were then removed using tangential flow filtration (TFF). To determine whether the circRNA can withstand the high shear process and maintain its integrity, naked RNA (without polymer and ionizable lipid) was processed through the whole process and then the samples were characterized using an automated electrophoresis system (e.g., Tapestation, Agilent).
[0559] During the quench step, as the solvents are starting to leave the oil droplet and as the polymeric lipid nanoparticles (PLNPs) are starting to solidify (from the emulsion state), the circRNA-CFL gets trapped at the core of PLNPs whereas the polymer forms a shell around the encapsulated circRNA-CFL pair. Without the CFL, a large hydrophilic molecule such as the circRNA could not get trapped within the core. Shown in FIG. 3 is a graphic representation of the core shell PLNP containing the circRNA- CFL pair at the core of the PLNP and a polymer shell.
EXAMPLE 3
Testing integrity of circular RN A polynucleotide (circRNA) in the absence of polymer.
[0560] To determine whether the circRNA can withstand the high shear process and maintain its integrity, naked RNA (ofLuc and mfLuc, without amphiphilic polymer and ionizable lipid) was processed through the whole process and then the samples were characterized using automated electrophoresis (TapeStation). FIG. 4A shows RNA integrity data for both mRNA (mfLuc) and circRNA (ofLuc) after being processed through the high shear process. RNA degradation was observed for both mRNA and circRNA, however, circRNA demonstrated significantly less degradation compared to mRNA. This observation was certainly unexpected and unpredictable.
[0561] In a follow up experiment, complex of circRNA (ofLuc) and ionizable lipid ethyl lauroyl arginate (ELA) at various ratios (no amphiphilic polymer) were processed through the high shear process and characterized for RNA integrity using automated electrophoresis (TapeStation). FIG. 4B shows that when the circRNA is complexed with ELA, no RNA degradation was observed for any circRNA:ELA complex. This data supported using the emulsion-based process of Example 2 for making circRNA loaded PLNPs.
EXAMPLE 4
In vitro release of a PLNP.
[0562] PLNPs were prepared according to the procedure set out in Example 2 using 10K-5K molecular weighted polymer (P) (e.g., 10K weighted PLA and 5K weighted PEG) and ionizable lipid ethyl lauroyl arginate (ELA) at ELA:P ratios of 2: 1 , 4: 1 , and 6:1. The PLNPs were loaded with circRNAs comprising a 3’ exon fragment, an internal ribosome entry site (IRES), a coding region (e.g., firefly luciferase), and a 5’ exon fragment. circRNA loaded particles with a particle size of 77nm and 85nm and a PDI below 0.2 were prepared using the emulsion based process provided in Example 2. Encapsulation efficiency of 70-90% were achieved for all the ELA:P ratio as shown in FIGs. 5A and 5B. The nanoparticle made with ELA:P ratio of 4:1 were analyzed for morphology using Cryo-TEM shown in FIG. 6A. For comparison purposes, samples from a “no-polymer” formulation were tested concurrently (shown in FIG. 6B). FIG. 7 shows vitro release (IVR) data in PBS at 37°C for a PLNP of two different particle sizes made with a 10K-5K PLA-PEG. Both PLNPs demonstrated a burst release of approximately 20- 25% and complete RNA release over 5 to 6 days.
EXAMPLE 5
PLNP in vivo delivery of circRNA.
[0563] Circular RNAs, comprising a 3’ exon, an internal ribosome entry site (IRES), a coding region encoding firefly luciferase, and a 5’ exon, were formulated with delivery vehicle comprising either a
PLNP (non-targeted), a PLNP comprising GalNAc3, an exemplary LNP comprising an ionizable lipid (positive control), and a diluent. The formulated circular RNA and delivery vehicle were dosed at 4 mpk into Balb/C mice intravenously. Mice were sacrificed at 6 hours, 1 day, 2 days, 5 days post intravenous injection of the circular RNAs. Protein expression was evaluated using Ex vivo IVIS. Liver tissue was characterized for circRNA distribution into hepatocytes and Kupffer cells (within the Liver tissue) using FISH. Mice body weights were collected throughout the study to assess tolerability of the nanoparticle formulations.
[0564] FIG. 8 provides the resulting FISH imaging. In FIG. 8, the top row of images shows liver tissue samples characterized at 6 hours post IV injection using Fluorescence in situ hybridization (FISH) that were imaged at lx magnification. Only the yellow fluorescence channel (corresponding to circular RNA encoding firefly) had been turned on for FIG. 8. For the diluent image (negative control), no firefly luciferase signal was seen. For the ionizable LNP 1 image (positive control), the firefly signal was observed suggesting delivery of circular RNA encoding firefly luciferase to liver tissue. For the PLNP group (non-targeted) the firefly luciferase signal was present but more distributed towards the surface of the tissue. Finally, in the PLNP-GalNAc3 image, the firefly luciferase fluorescence was significantly more brighter and spread throughout the liver tissue, suggesting a much more effective and uniform delivery of the circular RNA encoding flue to the target tissue as compared to the PLNP (non-targeted) and LNP groups. In FIG. 8, the bottom row of images correspond to the same 6 hour Liver tissue samples characterized using FISH and being imaged at 20x magnification and had all of the fluorescence channels turned on. As marked in the Diluent image of FIG. 8 blue corresponds to DAPI (nuclei of the hepatocytes), red corresponds to hepatocytes, green corresponds to Kupffer cells and yellow corresponds to circular RNAs encoding firefly luciferase. In the Diluent image, no signal for firefly luciferase was observed. In the ionizable LNP 1 image, the hepatocytes appeared slightly orange in color and some of the Kupffer cells demonstrate yellow overlap suggesting a mixed distribution in hepatocyte and Kupffer cells. For the PLNP group (non-targeted), the hepatocytes appeared red in color, and most Kupffer cells demonstrated yellow overlap suggesting the circular RNA encoding firefly luciferase distribution mainly to Kupffer cells, rather than hepatocyte. For PLNP-GalNAc3 group, the hepatocytes appeared yellow-orange in color indicating that a large amount of the circular RNAs encoding firefly luciferase was delivered to hepatocytes. Also, most Kupffer cells appeared completely green in color suggesting that GalNAc3 conjugated PLNPs demonstrated preferential uptake by hepatocyte rather than the Kupffer cells.
[0565] The liver organs of the same Balb/C mice were harvested and analyzed for Flue expression using Ex vivo imaging (provided in FIG. 9). The ionizable LNP 1 treated mice demonstrated high protein expression at the 6 hour timepoint but the protein levels start to drop rapidly for subsequent timepoints. Comparatively, the PLNP GalNAc mice demonstrated lower (compared to the ionizable
LNP 1 mice) but sustained expression of protein (for three consecutive timepoints) up to 2 days. The non-targeted PLNP (shown as PLNP in FIG. 9) demonstrated an inconsistent expression profile.
[0566] Mice body weights were also collected for the circular RNAs formulated with either PLNPs comprising GalNAc3, ionizable LNP 1 or 20% sucrose. Body weights shown in FIG. 10 demonstrated little to no change in mice body weights over the duration of the study, suggesting that all the dosing groups, including PLNP groups, were well tolerated.
EXAMPLE 6
Formulation of PLNP with different ionizable lipids.
[0567] PLNPs were prepared according to the emulsion-based formulation procedure described in Example 2 using 10K-5K molecular weighted PLA-PEG copolymer (e.g., 10K weighted PLA and 5K weighted PEG, acquired from Advanced Polymer Materials, Inc.) and one of three different ionizable lipids. The three ionizable lipids used comprised either ethyl lauroyl arginate, ionizable lipid 2, and ionizable lipid 3. PLNPs comprising either ethyl lauroyl arginate and/or ionizable lipid 2 were formulated at a lipid to phosphate (RNA backbone) ratio of 3:1; and PLNPs comprising ionizable lipid 3 were formulated at a lipid to phosphate ratio of 4:1. All three PLNPs were loaded with circRNAs comprising a 3’ exon fragment, an internal ribosome entry site (IRES), a coding region (e.g., expressing firefly luciferase), and a 5’ exon fragment.
[0568] The PLNPs were characterized throughout the formulation process using dynamic light scattering (DLS) and Quant-iT™ RiboGreen™ RNA encapsulation assay as shown in FIG. 16. DLS measurements were taken following the emulsion quench step (PQ), post-TFF purification (pTFF), and after an additional particle concentration step using Amicon™ centrifugal filter columns (PC). All three PLNPs showed similar and consistent DLS measurements throughout the process, with final particle size around 80 nm with PDI less than 0.1 (illustrated in FIG. 16). Encapsulation efficiency was determined using RiboGreen™ at the PC step, with encapsulation efficiency of 73%, 80%, and 66% for the ethyl lauroyl arginate, ionizable lipid 2, and ionizable lipid 3 PLNPs, respectively (also illustrated in FIG. 16).
EXAMPLE 7
In vitro release and RNA integrity of PLNP with different ionizable lipids.
[0569] The rate of RNA released into solution over time in physiological buffer conditions and the integrity of the unreleased RNA was evaluated in PLNPs.
[0570] PLNPs were prepared according to the emulsion-based formulation procedure described in Example 2 using 10K-5K molecular weight PLA-PEG copolymer (e.g., 10K weighted PLA and 5K
weighted PEG, acquired from Advanced Polymer Materials, inc.) and one of two different ionizable lipids. PLNPs comprising ionizable lipids ethyl lauroyl arginate or ionizable lipid 2 were formulated at a lipid to phosphate (RNA backbone) ratio of 3: 1 and were loaded with circRNAs comprising a 3’ exon fragment, an internal ribosome entry site (IRES), a coding region (e.g., firefly luciferase), and a 5’ exon fragment. The two PLNP formulations had similar post-processing characterization as assessed by dynamic light scattering (DLS) and Quant-iT™ RiboGreen™ RNA encapsulation assay as shown in FIG. 17.
[0571] In vitro release (IVR) assays were conducted, wherein PLNPs comprising ethyl lauroyl arginate or ionizable lipid 2 formulations were each diluted into 10 mM sodium phosphate buffer (pH 7.4) and incubated with gentle shaking at 37 °C. Aliquots were taken at t=0 and over the course of 7 days. An unformulated RNA control comprised circRNA that had been formulated into the PLNP diluted directly into the sodium phosphate buffer. These aliquots were first assessed for extent of circRNA released from the PLNPs into solution using the RiboGreen™ RNA encapsulation assay. When plotted as percentage of total circRNA, the PLNPs comprising ethyl lauroyl arginate or ionizable lipid 2 had similar release curves, with about 20% circRNA being released in the first 6 hours of incubation followed by steady release up to 80-90% after 7 days (as depicted in FIG. 18).
[0572] To evaluate the integrity of the circRNA remaining in the PLNPs at given timepoints throughout the IVR assay, RNA encapsulated in PLNP was isolated and/or released from unencapsulated RNA in the surrounding solution using an ultracentrifugation method. Briefly, 2 mL of each PLNP or control sample was pipetted onto 10 mL of distilled water. The PLNP samples were loaded into a SW 41 Ti rotor in a Beckman Coulter floor and centrifuged at 288,000 g for 3 hours. The supernatant containing RNA in solution was aspirated and discarded. The pellet of PLNPs was reconstituted in distilled water and transferred to a microcentrifuge tube. The RNA was extracted from the reconstituted pellet using the New England Biolabs Total RNA Miniprep Kit, following the protocol for RNA extraction from animal cells. The RNA control sample was not ultracentrifuged but was processed using the RNA extraction method. The concentration of the extracted RNA from all samples was measured using a NanoDrop Microvolume UV-Vis Spectrophotometer, diluted to 100 pg/mL with distilled water, and stored at 4 °C until analysis.
[0573] Samples were analyzed using an RP-IP (reverse phase ion pairing) HPLC method to assess RNA integrity (e.g., samples were separated and measured for relative quantities of intact circRNA (“circular”) against partially degraded circRNA (“nicked”) and other degradation products). FIGS. 19A, B, C provide exemplary chromatograms for the unformulated circRNA (depicted in FIG. 19A) and circRNA extracted from PLNPs comprising either ethyl lauroyl arginate or ionizable lipid 2 at 0 minutes of the IVR assay (depicted in FIGs. 19B and 19C respectively). RNA integrity was calculated as the area under the curve of the intact circular peak as a percentage of all RNA peaks. FIG. 20 provides
the calculated RNA integrity of the circRNA remaining in the PLNPs comprising ethyl lauroyl arginate and ionizable lipid 2 at timepoints before and throughout the IVR assay, as compared with the RNA integrity of the unformulated RNA control over time.
EXAMPLE 8
In vivo delivery of circRNA formulated into PLNP with different ionizable lipids.
[0574] CircRNA comprising: a 3’ exon, an internal ribosome entry site (IRES), a coding region encoding firefly luciferase, and a 5’ exon, was formulated with a PLNP delivery vehicle comprising ethyl lauroyl arginate or ionizable lipid 2 as the ionizable lipid. All three PLNP formulations were appended with GalNAc3. These formulated circRNA PLNPs, along with an exemplary circRNA LNP comprising an ionizable lipid, ethyl lauroyl arginate (positive control) and a diluent, 20% sucrose (negative control), were dosed at 8 mpk and/or 4 mpk into Balb/C mice intravenously. Dosing was defined as mass of circRNA per kilogram of bodyweight. Mice were euthanized at either 6 hours, 2 days, and 5 days post-intravenous injection. Protein fLuc expression from the mice was evaluated using ex vivo IVIS as depicted FIG. 21. Liver tissue was analyzed for circRNA distribution in hepatocytes and Kupffer cells (e.g., within the liver tissue) using FISH. All formulations were well tolerated as determined from clinical observation of the mice using techniques available known in art. The PLNP comprising ethyl lauroyl arginate or ionizable lipid 2 both showed fLuc expression above baseline for up to 2 days post-injection in FIG. 21.
[0575] Fluorescent in situ hybridization (FISH) is an effective imaging technique for visualizing RNA intracellular deposition and distribution with a tissue. Following harvest and imaging, the livers of the mice were preserved using 4% paraformaldehyde in phosphate buffered solution (PBS) and sent for FISH imaging. A probe for the circRNA coding for firefly luciferase was used to visualize the delivered RNA, while probes for albumin RNA, Adgrel RNA, and DNA were used to label the hepatocytes, Kupffer cells, and cell nuclei, respectively. FIG. 22A shows the mean fLuc RNA signal, calculated using ImageJ, across dosed groups at 6 hours post-injection. PLNPs comprising either ionizable lipid ethyl lauroyl arginate or ionizable lipid 2 resulted in significant RNA signal over baseline, with the 4 mpk and 8 mpk doses resulting in similar total RNA delivered in the liver. FIG. 22B shows exemplary FISH images for the two PLNPs 4 mpk groups at 6 hours and 5 days post-dose timepoints. The PLNP comprising ionizable lipid 2 showed noticeably more RNA signal remaining in the liver tissue compared to PLNPs comprising ethyl lauroyl arginate.
EXAMPLE 9
In vitro delivery of circRNA- PLNP.
[0576] An in vitro firefly luciferase expression assay was used for PLNP to facilitate the evaluation of various formulation conditions. To emulate in vivo conditions, a targeted receptor-mediated uptake method (e.g., using GalNac3) was designed into the assay. GalNac3 interaction on PLNPs with ASGPR receptors on hepatocytes was used to facilitate and analyze intracellular uptake in vivo. PLNPs comprised ethyl lauroyl arginate. Three cell types (i.e., 1C1C7 cells, primary human hepatocytes (PHH), and human skeletal muscle cells (HSKM)) were assayed for endogenous ASGPR expression via western blot. Additionally, to explore the option of supplementing endogenous expression, circRNA encoding ASGPR was synthesized, and the same cell types were transfected with the circRNA using MessengerMax™, a commercially available membrane permeating reagent for either 6 hours or 24 hours. At 24 hours post-transfection, the cells were also assayed for ASGPR expression via western blot as illustrated in FIG. 23. Both 1C1C7S and primary human hepatocytes (PHH) showed endogenous ASGPR expression that was significantly enhanced by transfection of ASGPR circRNA. Human skeletal muscle cells were used as a negative control for endogenous expression. As expected, the human skeletal muscle cells did not show detectable levels of endogenous ASGPR expression and only showed expression upon transfection of ASGPR protein. PHH cells were further transfected with ASGPR circRNA as the basis for an in vitro expression assay.
[0577] Multiple timepoints were evaluated for the PLNP transfection and luciferase expression in PHH cells. After ASGPR circRNA transfection using MessengerMax™, a cell membrane permeating reagent, the PHH cells were allowed to recover before nanoparticle transfection to avoid additional nonspecific particle uptake as shown in FIG. 24, wherein transfection with unconjugated PLNP resulted in significant luciferase expression above baseline in cells concurrently treated with ASGPR circRNA. CircRNA transfection via the PLNPs at 6 hours and 24 hours post- ASGPR transfection timepoints; luciferase expression was analyzed at 48 hours post-ASGPR circRNA transfection (as shown in FIG. 25). At both timepoints of ASGPR transfection (e.g., at 6 hours and 24 hours) and for all doses of firefly luciferase circRNA assessed, the unconjugated PLNPs did not indicate significant non-specific uptake and the GalNAc3-appended PLNP showed significant increases in luciferase expression over unconjugated PLNP controls in cells transfected with exogenous ASGPR circRNA.
[0578] Luciferase expression was then measured 24 hours post-ASGPR circRNA transfection after PLNP transfection 6 hours post-ASGPR circRNA transfection as depicted in FIG. 26. Significant luciferase expression over unconjugated PLNP controls was also observed at this timepoint. In all studies, Promega Bright-Glo™ Luciferase Assay System was used to evaluate luciferase expression.
EXAMPLE 10
Assessment of PLNPs with alternative polymer shell formulations.
[0579] PLNP formulatability, in vitro RNA release profile, and in vitro firefly luciferase expression in primary human hepatocytes were evaluated in PLNPs formulated with different polymer shell constitutions.
[0580] PLNPs were prepared according to the emulsion-based formulation procedure described in Example 2 (e.g., using ethyl lauroyl arginate as the complex forming ionizable lipid at a lipid to phosphate ratio of 3:1 and circRNAs comprising a 3’ exon fragment, an internal ribosome entry site (IRES), an expression sequence encoding firefly luciferase, and a 5’ exon fragment). The PLNPs were developed with different polymer components. The control PLNP was formulated with a 10K-5K molecular weight PLA-PEG block copolymer. Shell 1 comprised a 4:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer. Shell 2 comprised a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer. Shell 3 was comprised entirely of a 15K-5K PLGA-PEG block copolymer (i.e., the PLGA portion comprised a random copolymer of 75% lactic acid and 25% glycolic acid). Shell 4 comprised a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K- 2K molecular weight PLA-PEG block copolymer, supplemented with 1% by mass 16K-5K PLA-PEG block copolymer. The PLNPs were characterized using dynamic light scattering (DLS) and Quant-iT™ RiboGreen™ RNA encapsulation assay (illustrated in FIG. 27).
[0581] An in vitro RNA release assay as described in Example 7 was conducted with the alternative shell PLNP formulations (i.e., Shells 1-4) and a circRNA control. FIG. 28A shows the percent of encapsulated circRNA released from each PLNP formulation over the course of 5 days; FIG. 28B shows the first 24 hours post administration of FIG. 28A. The formulations of FIGs. 28A and 28B show a variety of release curves. By 5 days, three of the four alternative shell formulations had released more RNA than the control formulation, with Shell 2 showing near 100% RNA release.
[0582] The alternative shell PLNP (i.e., Shells 1-4) formulations were also evaluated for luciferase expression in primary human hepatocytes, using methods described in Example 9. Two circRNA doses were assessed with expression timepoints at 24 and 48 hours post-ASGPR circRNA transfection (shown in FIGs. 29A and 29B). In general, the alternative PLNP shells affected the kinetics of luciferase expression. At 24 hours, three of the four alternative shell formulations showed significantly higher expression (p < 0.05) than the control formulations, consistent with the RNA release profile at 24 hours in the IVR assay (illustrated in FIG. 29A). At 48 hours, the control formulation shows higher expression than the alternative shell PLNPs (illustrated in FIG. 29B).
EXAMPLE 11
Lyophilization stability of PLNP formulations.
[0583] Lyophilized storage stability of circRNA PLNP formulations was assessed using dynamic light scattering (DLS), Quant-iT™ RiboGreen™ RNA encapsulation assay, and IPRP-HPLC RNA integrity assay on reconstituted samples.
[0584] PLNPs were prepared according to the emulsion-based formulation procedure described in Example 2 using 10K-5K molecular weight PLA-PEG block copolymer and ethyl lauroyl arginate as the complex forming ionizable lipid at a lipid to phosphate ratio of 3:1. The PLNPs were loaded with circRNA comprising a 3’ exon fragment, an internal ribosome entry site (IRES), an expression sequence encoding firefly luciferase, and a 5’ exon fragment.
[0585] After TFF purification, the PLNPs were diluted in a lyophilization protectant buffer. Aliquots were lyophilized using a Labconco FreeZone benchtop lyophilizer. The aliquots were stored at either 4 °C or -80 °C and reconstituted at one-week intervals over 4 weeks. Characterization showed minimal changes in PLNP particle size distribution, encapsulation efficiency, and RNA integrity following lyophilization and storage for up to 4 weeks (illustrated in FIGs. 30A, 30B, and 30C respectively).
EXAMPLE 12
PLNPs formulated with endosomal escape agents.
[0586] PLNP formulatability, in vitro RNA release profile, and in vitro firefly luciferase expression in primary human hepatocytes were evaluated in PLNPs formulated with different polymer shell constitutions.
[0587] PLNPs were prepared according to the emulsion-based formulation procedure described in Example 2 (e.g., using ethyl lauroyl arginate as the complex forming ionizable lipid at a lipid to phosphate ratio of 3:1 and circRNAs comprising a 3’ exon fragment, an internal ribosome entry site (IRES), an expression sequence encoding firefly luciferase, and a 5’ exon fragment). The PLNPs were developed with different polymer components. The control PLNP was formulated with a 10K-5K molecular weight PLA-PEG block copolymer. Shell 2 comprised a 1:1 ratio of 10K-5K molecular weight PLA-PEG block copolymer and 5K-2K molecular weight PLA-PEG block copolymer. Shell 3 was comprised entirely of a 15K-5K PLGA-PEG block copolymer (i.e., the PLGA portion comprised a random copolymer of 75% lactic acid and 25% glycolic acid). PLNPs comprised core forming lipids, ELA or ionizable lipid 2. The endosomal escape agents (e.g., doxepin or endosomal escape agent 1) were applied as separate solutions either concurrently with the circular RNA-PLNP or 1 or 2 hours afterwards (e.g., at a pre-dose, concurrent dose, or post-dose timepoint relative to PLNP dosing). For PLNPs formulated with doxepin, 50 pM of doxepin hydrochloride concentration (e.g., from Millipore-
Sigma) was used either as a pre-dose 1 hour prior to dosing the circular RNA-PLNP solutions into cells, concurrently as a direct dose during circular RNA-PLNP dosing, or 1 or 2 hours as a post dose. For PLNPs formulated with endosomal escape agent 1, 7.17 pM of endosomal escape agent 1 solution was used either as a pre-dose 1 hour prior to dosing the circular RNA-PLNP solutions into cells, concurrently as a direct dose during circular RNA-PLNP dosing, or 1 or 2 hours as a post dose.Luciferase expression was then measured 48 hours post circRNA transfection as shown in FIG. 31. In all studies, Promega Bright-Glo™ Luciferase Assay System was used to evaluate luciferase expression.
INCORPORATION BY REFERENCE
[0588] All publications, patents, and patent applications mentioned in this specification are herein incorporated by reference to the same extent as if each individual publication, patent, or patent application were specifically and individually indicated as being incorporated by reference herein, including, for example, U.S. provisional patent application nos. 63/511,574 (filed June 30, 2023), 63/492,988, and 63/492,974; International patent application nos. PCT/US2019/035531, PCT/US2020/034418, PCT/US2020/063494, PCT/US2021/031629, PCT/US2021/023540, PCT/US2021/033276, PCT/US2022/033091, PCT/US2022/045408, PCT/US2022/049313,
PCT/US2023/78946, PCT/US2023/78950, and PCT/US2024/019990; and PCT Publications WO 2023/044343, WO 2023/044333, WO 2023/122752, WO 2024/044728 and WO 2023/196931.
Claims
1. A composition comprising a plurality of polymeric lipid nanoparticles, each nanoparticle comprising: a circular RNA polynucleotide; an ionizable lipid; and an amphiphilic polymer.
2. The composition of claim 1, wherein the amphiphilic polymer is a block copolymer comprising: a hydrophilic block comprising a hydrophilic polymer; and a hydrophobic block comprising a hydrophobic polymer.
3. The composition of claim 2, wherein the amphiphilic polymer further comprises a cleavable linker.
4. The composition of claim 3, wherein the hydrophobic block of the amphiphilic polymer comprises the cleavable linker.
5. The composition of claim 3, wherein the cleavable linker covalently connects the hydrophilic block to the hydrophobic block of the amphiphilic polymer.
6. The composition of claim 3, wherein the cleavable linker covalently connects two or more amphiphilic polymers.
7. The composition of any one of claims 2 to 6, wherein the amphiphilic polymer is covalently bound to a cationic moiety.
8. The composition of claim 7, wherein the amphiphilic polymer is covalently bound to the ionizable lipid.
9. The composition of any one of claims 2 to 6, wherein the amphiphilic polymer is not covalently bound to the ionizable lipid.
10. The composition of any one of claims 1 to 9, wherein the amphiphilic polymer has a molecular weight from 10k to 70k.
11. The composition according to any one of claims 2 to 10, wherein the amphiphilic polymer is a di-block copolymer.
12. The composition according to any one of claims 2 to 10, wherein the amphiphilic polymer is a tri-block copolymer.
13. The composition of claim 11, wherein the amphiphilic polymer comprises a block copolymer of Formula I:
X-Y (I) wherein:
X is a hydrophobic block comprising a hydrophobic polymer; and
Y is a hydrophilic block comprising a hydrophilic polymer, wherein the amphiphilic polymer optionally further comprises one or more cleavable linkers.
14. The composition of claim 13, wherein the amphiphilic polymer comprises a block copolymer of Formula IA:
X-L-Y (IA) wherein:
X is a hydrophobic block comprising a hydrophobic polymer;
Y is a hydrophilic block comprising a hydrophilic polymer; and
L is a cleavable linker.
15. The composition of claim 13, wherein the amphiphilic polymer comprises a block copolymer of Formula IB:
XA-(L-XB)n-Y (IB) wherein: each of XA and XB is independently a hydrophobic block comprising a hydrophobic polymer;
Y is a hydrophilic block comprising a hydrophilic polymer;
L is a cleavable linker; and n is an integer from 1 to 5.
16. The composition of claim 15, wherein n is an integer from 1 to 3.
17. The composition of claim 2, wherein the amphiphilic polymer comprises a block copolymer of Formula IC:
XA - T - L - T - XB
Y IA Y IB (IC) wherein: each of XA and XB is independently a hydrophobic block comprising a hydrophobic polymer; each of YA and YB is independently a hydrophilic block comprising a hydrophilic polymer;
L is a cleavable linker; and each T is independently a trivalent connecting group.
18. The composition of claim 2, wherein the amphiphilic polymer comprises a block copolymer of Formula ID:
R-X-Y (ID) wherein:
R is a cationic moiety;
X is a hydrophobic block comprising a hydrophobic polymer;
Y is a hydrophilic block comprising a hydrophilic polymer; and the amphiphilic polymer optionally further comprises one or more cleavable linkers.
19. The composition of claim 12, wherein the amphiphilic polymer comprises a block copolymer of Formula II A:
XA-Y-XB (IIA) wherein: each of XA and XB is independently a hydrophobic block comprising a hydrophobic polymer;
Y is a hydrophilic block comprising a hydrophilic polymer; and the amphiphilic polymer optionally further comprises one or more cleavable linkers.
20. The composition of claim 12, wherein the amphiphilic polymer comprises a block copolymer of Formula IIB:
YA-X-YB (IIB)
wherein: each of YA and YB is independently a hydrophilic block comprising a hydrophilic polymer;
X is a hydrophobic block comprising a hydrophobic polymer; and the amphiphilic polymer optionally further comprises one or more cleavable linkers.
21. The composition of any one of claims 13 to 20, wherein X, XA, or XB comprises a polymer selected from a polyester polymer, a polyorthoester (POE) polymer, a polyanhydride polymer, a polyamide polymer, a poly(ester amide) polymer, a poly(phosphoester) polymer, a poly(alkyl cyanoacrylate) (PACA) polymer, a polysaccharide polymer, and any combination thereof.
22. The composition of claim 21, wherein X, XA, or XB comprises a polyester polymer.
23. The composition of claim 21, where X, XA, or XB comprises a polysaccharide polymer and the polysaccharide polymer is selected from a chitosan polymer, and a hyaluronic acid (HA) polymer, or a combination thereof.
24. The composition of any one of claims 13 to 23, wherein Y, YA, or YB comprises a polymer selected from a polyethylene glycol (PEG) polymer, a polyethylene oxide (PEG) polymer, a polyglutamic acid (PGA) polymer, a poly[N-(2-hydroxypropyl) methacrylamide] (HPMA) polymer, a poly(vinylpyrrolidone) (PVP) polymer, a poly(2-methyl-2-oxazoline) (PMOX) polymer, a poly(N,N- dimethyl acrylamide) (PDMA) polymer, a poly(N-acryloyl morpholine) (PAcM) polymer, and any combination thereof.
25. The composition of claim 24, wherein Y, YA, or YB comprises a PEG polymer.
26. The composition of any one of claims 13 to 20 wherein:
X, XA, or XB comprises a polyester polymer; and
Y, YA, or YB comprises a PEG polymer.
27. The composition of claim 26, wherein the molar ratio of Y, YA, and/or YB collectively to X, XA, and/or XB collectively is 1:2 to 1:4.
28. The composition of claim 26 or 27, wherein the polyester polymer has a molecular weight from 5k to 20k.
29. The composition of claim 28, wherein the polyester polymer has a molecular weight of from
5k to 10k.
30. The composition of any one of claims 25 to 29, wherein the PEG polymer has a molecular weight of from 5k to 10k.
31. The composition of claim 30, wherein the PEG polymer has a molecular weight of 5k +/- 10%.
32. The composition of any one of claims 21 to 31, wherein the polyester polymer is selected from a polylactide (PLA) polymer, a polyglycolide (PGA) polymer, a polycaprolactone (PCL) polymer, a polydioxanone (PDO) polymer, a polyhydroxyalkanoate polymer, a poly(glycerol sebacate) polymer, a poly(lactic-co-glycolic acid) (PLGA) polymer, a poly(P-amino ester) (PBAE), a poly(amine-co-ester) (PACE), a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA), and any combination thereof.
33. The composition of claim 32, wherein the polyester polymer comprises polylactide (PLA).
34. The composition of claim 32, wherein the polyester polymer comprises poly(lactic-co- gly colic acid) (PLGA).
35. The composition of claim 2, wherein the amphiphilic polymer comprises a block copolymer of Formula III:
[A] V-[B] w-[C]x-[D]y-[E]z (III) wherein:
A is a polyester monomer or a polyethylene glycol (PEG) monomer;
B is a polyester monomer;
C is a polyester monomer;
D is a polyester monomer;
E is a PEG monomer; v is an integer from 0 to 200; w is an integer from 0 to 200; x is an integer from 0 to 200; y is an integer 10 to 200; and z is an integer from 10 to 150, wherein the amphiphilic polymer optionally further comprises one or more intervening cleavable linkers (L) between any of A-E.
36. The composition of claim 13 or 35, wherein the amphiphilic polymer comprises a block copolymer of Formula IIIA:
[D]y-[E]Z (IIIA) wherein:
D is a polyester monomer;
E is a PEG monomer; y is an integer from 10 to 200; and z is an integer from 10 to 150.
37. The composition of claim 13 or 35, wherein the amphiphilic polymer comprises a block copolymer of Formula IIIB :
[C]x-[D]y-[E]z (IIIB) wherein:
C is a polyester monomer;
D is a polyester monomer;
E is a PEG monomer; x is an integer from 10 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150.
38. The composition of claim 13 or 35, wherein the amphiphilic polymer further comprises at least one cleavable linker (L).
39. The composition of claim 15 or 38, wherein the amphiphilic polymer comprises a block copolymer of Formula IIIC:
[C]x-L-[D]y-[E]z (IIIC) wherein:
C is a polyester monomer;
L is a cleavable linker;
D is a polyester monomer;
E is a PEG monomer;
x is an integer from 10 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150.
40. The composition of claim 15 or 38, wherein the amphiphilic polymer comprises a block copolymer of Formula IIID:
[A]v-[B]w-L-[C]x-[D]y-[E]z (IIID) wherein:
A is a polyester monomer;
B is a polyester monomer;
L is a cleavable linker;
C is a polyester monomer;
D is a polyester monomer;
E is a PEG monomer; v is 10 to 200; w is an integer from 10 to 200; x is an integer from 10 to 200; y is an integer from 10 to 200; and z is an integer from 10 to 150.
41. The composition of claim 14 or 38, wherein the amphiphilic polymer comprises a block copolymer of Formula IIIE:
[C]x-[D]y-L-[E]z (IIIE) wherein:
C is a polyester monomer;
D is a polyester monomer;
L is a cleavable linker;
E is a PEG monomer; x is an integer from 0 to 200;
y is an integer from 10 to 200; and z is an integer from 10 to 150.
42. The composition of claim 2 or 19, wherein the amphiphilic polymer comprises a block copolymer of Formula IV :
[A]v-[B]w-[E]z-[C]x-[D]y (IV) wherein:
A is a polyester monomer;
B is a polyester monomer;
E is a PEG monomer;
C is a polyester monomer;
D is a polyester monomer; v is an integer from 10 to 200; w is an integer from 0 to 200; x is an integer from 0 to 200; y is an integer from 10 to 200; z is an integer from 10 to 150, wherein the amphiphilic polymer optionally further comprises one or more intervening cleavable linkers (L) between any of A-E.
43. The composition of claim 42, wherein the amphiphilic polymer comprises a block copolymer of Formula IVA:
[A]v-[E]z-[D]y (IVA).
44. The composition of claim 42, wherein the amphiphilic polymer comprises a block copolymer of Formula IVB:
[A]v-[B]w-[E]z-[C]x-[D]y (IVB) wherein: w is an integer from 10 to 200; and x is an integer from 10 to 200.
45. The composition of claim 2 or 17, wherein the amphiphilic polymer comprises a block copolymer of Formula V :
wherein:
L is a cleavable linker; each T is independently a trivalent connecting group;
E and F are each independently a PEG monomer;
A, B, C, and D are each independently a polyester monomer; v is an integer from 10 to 200 w is an integer from 0 to 200; x is an integer from 0 to 200; y is an integer from 10 to 200; and z and u are each an integer from 10 to 150.
47. The composition of claim 45 or 46, wherein T is a small molecule having a molecular weight of 1000 Daltons or less.
48. The composition of claim 47, wherein the small molecule is a peptide.
49. The composition of claim 48, wherein the peptide comprises cysteine or lysine.
50. The composition of any one of claims 35 to 49, wherein L is cleaved by exposure to a stimulus.
51. The composition of claim 50, wherein the stimulus is selected from pH, temperature, light, redox change, over-expressed enzymes, hypoxia, sound, magnetic force, electrical energy, and any combination thereof.
52. The composition of claim 50 or 51 , wherein L comprises a group selected from disulfide, hydrazone, vinyl ether, imine, ortho ester, borate ester, amide, a peptide, and azo.
53. The composition of claim 52, wherein L comprises a disulfide.
54. The composition of claim 52, wherein L comprises a hydrazone.
55. The composition of claim 52, wherein L comprises a peptide and the peptide is a cleavable octapeptide.
56. The composition of claim 2 or 18, wherein the amphiphilic polymer comprises a block copolymer of Formula VI:
R-[C]x-[D]y-[E]z (VI) wherein:
R is a cationic moiety;
C is a polyester monomer;
D is a polyester monomer;
E is a PEG monomer; x is an integer from 0 to 200; y is an integer from 10 to 200; z is an integer from 10 to 150.
57. The composition of claim 56, wherein R comprises a lipid, a polymer, or a non-lipid small molecule.
58. The composition of claim 57, wherein R comprises an ionizable lipid.
59. The composition of claim 58, wherein the ionizable lipid has a pKa from 6 to 9.
60. The composition of claim 59, wherein the ionizable lipid has a pKa from 7 to 9.
61. The composition of any one of claims 58-60, wherein the ionizable lipid comprises an ionizable amino group.
62. The composition of claim 61, wherein the ionizable lipid comprises a divalent headgroup and one or more hydrocarbon lipid tails (e.g., straight or branched hydrocarbon lipid tails).
63. The composition of claim 62, wherein the straight or branched hydrocarbon lipid tails are from 3-25 carbon atoms in length.
64. The composition of claim 62 or 63, wherein the divalent headgroup is selected from guanidine and squaramide.
65. The composition of claim 64, wherein the ionizable lipid is ethyl lauroyl arginate (ELA).
66. The composition of claim 57, wherein R comprises a polymer.
67. The composition of claim 66, wherein the polymer is selected from poly(lysine), polyethylene imine (PEI), poly (amidoamine), poly (histidine), poly( arginines), and polyamine resins.
68. The composition of claim 57, wherein R comprises a non-lipid small molecule.
69. The composition of claim 68, wherein the non-lipid small molecule is selected from an amine-containing compound, an amino acid, a heterocycle-containing compound, and a heteroaryl- containing compound.
70. The composition of claim 69, wherein the non-lipid small molecule is an amine-containing compound.
71. The composition of claim 70, wherein the amine-containing compound is selected from choline, betaine, N,N’ -dibenzylethylenediamine, diethylamine, 2-diethylaminoethanol, 2- methylaminoethanol, glucosamine, glucamine, ethanolamine, ethylenediamine, hydrabamine, isopropyl amine, methylglucamine, procaine, triethylamine, trimethylamine, tripropylamine, and tromethamine.
72. The composition of claim 69, wherein the non-lipid small molecule is an amino acid.
73. The composition of claim 72, wherein the amino acid is selected from arginine, histidine, and lysine.
74. The composition of claim 69, wherein the non-lipid small molecule is a heterocyclecontaining compound, or a heteroaryl-containing compound.
75. The composition of claim 74, wherein the non-lipid small molecule is selected from caffeine, N-ethylmorpholine, N-ethylpiperidine, morpholine, piperazine, piperidine, purines, and theobromine.
76. The composition of any one of claims 35 to 75, wherein A, B, C and D each independently comprise a monomer selected from a polylactide (PLA) monomer, a polyglycolide (PGA) monomer, a
polycaprolactone (PCL) monomer, a polydioxanone (PDO) monomer, a polyhydroxyalkanoate (e.g., polyhydroxybutyrate) monomer, a poly(glycerol sebacate) monomer, a poly(P-amino ester) (PBAE) monomer, a poly(2-(dimethylamino)ethyl methacrylate) (PDMAEMA) monomer, and any combination thereof.
77. The composition of claim 76, wherein A, B, C or D each comprise a polylactide (PLA) monomer.
78. The composition of any one of claims 35 to 75, wherein [A] v and [B] „ together and/or [C]x and [D]y together form a copolymer comprising any combination of monomers selected from a polylactide (PLA) monomer, a polyglycolide (PGA) monomer, a polycaprolactone (PCL) monomer, a polydioxanone (PDO) monomer, a polyhydroxyalkanoate (e.g., polyhydroxybutyrate) monomer, a poly(glycerol sebacate) monomer, a poly(P-amino ester) (PBAE) monomer, and a poly(2- (dimethylamino)ethyl methacrylate) (PDMAEMA) monomer.
79. The composition of claim 78, wherein [ A] v and [B] „ together and/or [C]x and [D]y together forms poly(lactic-co-glycolic acid) (PLGA).
80. The composition of claim 79, wherein the molar ratio of lactic acid monomer units to glycolic acid monomer units in the PLGA copolymer is from 1:1 to 9:1.
81. The composition of any one of claims 35 to 80, wherein the amphiphilic polymer further comprises one or more terminal groups selected from carboxyl, hydroxyl, amino, amido, and alkoxy.
82. The composition of claim 81, wherein the terminal group is an alkoxy group selected from n- butoxy (-O-(CH2)3)-CH3) and tert-butoxy (-O-(C(CH3)3).
83. The composition of any one of claims 1 to 82, wherein the ionizable lipid has a pKa from 6 to 9.
84. The composition of claim 83, wherein the ionizable lipid has a pKa from 7 to 9.
85. The composition of any one of claims 1 to 84, wherein the ionizable lipid comprises an ionizable amino group.
86. The composition of claim 85, wherein the ionizable lipid comprises a di-valent headgroup and one or more straight or branched hydrocarbon lipid tails.
87. The composition of claim 86, wherein the straight or branched hydrocarbon lipid tails are from 3-25 carbon atoms in length.
88. The composition of claim 86 or 87, wherein the di-valent headgroup is selected from guanidine and squaramide.
89. The composition of any one of claims 1-58, wherein the ionizable lipid is selected from ethyl lauroyl arginate (ELA), ionizable lipid 2, ionizable lipid 3, endosomal escape agent 1, and any of the lipids in Tables 3-7, preferably wherein the ionizable lipid is ELA.
90. The composition of any one of claims 1 to 89, wherein the molar ratio of ionizable lipid to amphiphilic polymer is from 2:1 to 6:1.
91. The composition of claim 90, wherein the molar ratio of ionizable lipid to amphiphilic polymer is 4:1.
92. The composition of any one of claims 1 to 91, wherein the ionizable lipid forms a complex with the circular RNA polynucleotide (circRNA-lipid complex).
93. The composition of claim 92, wherein the circRNA-lipid complex is encapsulated in the core of the nanoparticles and the amphiphilic polymer provides a uniform shell around the circRNA-lipid complex.
94. The composition of any one of claims 1 to 93, wherein the nanoparticle has a mean diameter from 50-200 nm.
95. The composition of any one of claims 1 to 94, wherein the poly dispersity index of the nanoparticles is 0.3 or less.
96. The composition of any one of claims 1 to 95, wherein the circular RNA comprises a first expression sequence.
97. The composition of claim 96, wherein the first expression sequence encodes a therapeutic protein.
98. The composition of claim 96, wherein the first expression sequence encodes a cytokine or a functional fragment thereof.
99. The composition of claim 96, wherein the first expression sequence encodes a transcription factor.
100. The composition of claim 96, wherein the first expression sequence encodes an immune checkpoint inhibitor.
101. The composition of claim 96, wherein the first expression sequence encodes a chimeric antigen receptor (CAR).
102. The composition of any one of claims 96 to 101 wherein the circular RNA polynucleotide further comprises a second expression sequence.
103. The composition of claim 102, wherein the circular RNA polynucleotide further comprises an internal ribosome entry site (IRES).
104. The composition of claim 102 or 103, wherein the first and second expression sequences are separated by a ribosomal skipping element or a nucleotide sequence encoding a protease cleavage site.
105. The composition of any one of claims 102 to 104, wherein the first expression sequence encodes a first T-cell receptor (TCR) chain, and the second expression sequence encodes a second TCR chain.
106. The composition of any one of claims 1 to 105, wherein the circular RNA polynucleotide comprises one or more micrcircRNA binding sites.
107. The composition of claim 106, wherein the micrcircRNA binding site is recognized by a micrcircRNA expressed in the liver.
108. The composition of claim 105 or 106, wherein the micrcircRNA binding site is recognized by miR-122.
109. The composition of any one of claims 1 to 108, wherein the circular RNA polynucleotide comprises a first IRES associated with greater protein expression in a human immune cell than in a reference human cell.
110. The composition of claim 109, wherein the human immune cell is a T cell, an NK cell, an NKT cell, a macrophage, or a neutrophil.
111. The composition of claim 109 or 110, wherein the reference human cell is a hepatic cell.
112. The composition of any one of claims 1 to 111, wherein the circular RNA polynucleotide comprises, in the following order: a. a 3’ exon fragment, b. a core functional element, and c. a 5’ exon fragment.
113. The composition of any one of claim 112, wherein the circular RNA polynucleotide further comprises a post-splicing intron fragment.
114. The composition of claim 112 or 113, wherein the circular RNA comprises a 5’ internal duplex region located downstream to the 3’ exon fragment.
115. The composition of any one of claims 112 to 114, wherein the circular RNA comprises a 5’ internal spacer located downstream to the 3’ exon fragment.
116. The composition of claim 115, wherein the 5’ internal spacer has a length of 10 to 60 nucleotides.
117. The composition of claim 115 or 116, wherein the 5’ internal spacer comprises a polyA or polyA-C sequence.
118. The composition of claim 117, wherein the polyA or polyA-C sequence comprises a length of 10-50 nucleotides.
119. The composition of any one of claims 112 to 118, wherein the core functional element comprises a translation initiation element (TIE).
120. The composition of claim 119, wherein the translation initiation element (TIE) comprises an untranslated region (UTR) or fragment thereof.
121. The composition of claim 120, wherein the UTR or fragment thereof comprises a viral internal ribosome entry site (IRES) or eukaryotic IRES.
122. The composition of claim 121, wherein the IRES has a sequence in whole or in part from a Taura syndrome virus, Triatoma virus, Theiler's encephalomyelitis virus, Simian Virus 40, Solenopsis invicta virus 1, Rhopalosiphum padi virus, Reticuloendotheliosis virus, Human poliovirus 1, Plautia stali intestine virus, Kashmir bee virus, Human rhinovirus 2, Homalodisca coagulata virus- 1, Human Immunodeficiency Virus type 1, Homalodisca coagulata virus- 1, Himetobi P virus, Hepatitis C virus, Hepatitis A virus, Hepatitis GB virus , Foot and mouth disease virus, Human enterovirus 71, Equine rhinitis virus, Ectropis obliqua piccircRNA-like virus, Encephalomyocarditis virus, Drosophila C Virus, Human coxsackievirus B3, Crucifer tobamovirus, Cricket paralysis virus, Bovine viral diarrhea virus 1, Black Queen Cell Virus, Aphid lethal paralysis virus, Avian encephalomyelitis virus, Acute bee paralysis virus, Hibiscus chlorotic ringspot virus, Classical swine fever virus, Human FGF2, Human SFTPA1, Human AML1/RUNX1, Drosophila antennapedia, Human AQP4, Human AT1R, Human BAG-1, Human BCL2, Human BiP, Human c-IAPl, Human c-myc, Human eIF4G, Mouse
NDST4L, Human LEF1, Mouse HIF1 alpha, Human n.myc, Mouse Gtx, Human p27kipl, Human PDGF2/c-sis, Human p53, Human Pim-1, Mouse Rbm3, Drosophila reaper, Canine Scamper, Drosophila Ubx, Human UNR, Mouse UtrA, Human VEGF-A, Human XIAP, Drosophila hairless, S. cerevisiae TFIID, S. cerevisiae YAP1, tobacco etch virus, turnip crinkle virus, EMCV-A, EMCV-B, EMCV-Bf, EMCV-Cf, EMCV pEC9, Picobirna virus, HCV QC64, Human Cosavirus E/D, Human Cosavirus F, Human Cosavirus JMY, Rhinovirus NAT001, HRV14, HRV89, HRVC-02, HRV-A21, Salivirus A SHI, Sali virus FHB, Sali virus NG-J1, Human Parecho virus 1, Crohi virus B, Yc-3, Rosavirus M-7, Shanbavirus A, Pasivirus A, Pasivirus A 2, Echovirus E14, Human Parechovirus 5, Aichi Virus, Hepatitis A Virus HA16, Phopivirus, CVA10, Enterovirus C, Enterovirus D, Enterovirus J, Human Pegivirus 2, GBV-C GT110, GBV-C K1737, GBV-C Iowa, Pegivirus A 1220, Pasivirus A 3, Sapelovirus, Rosavirus B, Bakunsa Virus, Tremovirus A, Swine Pasivirus 1, PLV-CHN, Pasivirus A, Sicinivirus, Hepacivirus K, Hepacivirus A, BVDV1, Border Disease Virus, BVDV2, CSFV- PK15C, SF573 Dicistro virus, Hubei PiccircRNA-like Virus, CRPV, Apodemus Agrarius PiccircRNAvirus, Caprine Kobuvirus, Parabovirus, Salivirus A BN5, Salivirus A BN2, Salivirus A 02394, Salivirus A GUT, Salivirus A CH, Salivirus A SZ1, Salivirus FHB, CVB3, CVB1, Echovirus 7, CVB5, EVA71, CVA3, CVA12, EV24, or an aptamer to eIF4G.
123. The composition of any one of claims 119 to 122, wherein the translation initiation element (TIE) comprises an aptamer complex.
124. The composition of claim 123, wherein the aptamer complex comprises at least two aptamers.
125. The composition of any one of claims 119 to 124, wherein the core functional element comprises a coding region.
126. The composition of claim 125, wherein the coding region encodes for a therapeutic protein.
127. The composition of claim 126, wherein the therapeutic protein is a chimeric antigen receptor (CAR), a cytokine, a transcription factor, a T cell receptor (TCR), B-cell receptor (BCR), ligand, immune cell activation or inhibitory receptor, recombinant fusion protein, chimeric mutant protein, or fusion protein or a functional fragment thereof.
128. The composition of claim 127, wherein the therapeutic protein is an antigen.
129. The composition of claim 128, wherein the antigen is a viral polypeptide from an adenovirus; Herpes simplex, type 1; Herpes simplex, type 2; encephalitis virus; papillomavirus; Varicella-zoster virus; Epstein-barr virus; Human cytomegalovirus; Human herpes virus, type 8; Human papillomavirus; BK virus; JC virus; Smallpox; polio virus; Hepatitis B virus; Human bocavirus; Parvovirus B19; Human astrovirus; Norwalk virus; coxsackievirus; hepatitis A virus; poliovirus;
rhinovirus; Severe acute respiratory syndrome virus; Hepatitis C virus; Yellow Fever virus; Dengue virus; West Nile virus; Rubella virus; Hepatitis E virus; Human Immunodeficiency virus (HIV); Influenza virus; Guanarito virus; Junin virus; Lassa virus; Machupo virus; Sabia virus; Crimean- Congo hemorrhagic fever virus; Ebola virus; Marburg virus; Measles virus; Mumps virus; Parainfluenza virus; Respiratory syncytial virus; Human metapneumo virus; Hendra virus; Nipah virus; Rabies virus; Hepatitis D; Rotavirus; Orbi virus; Colti virus; Banna virus; Human Enterovirus; Hanta virus; West Nile virus; Middle East Respiratory Syndrome Corona Virus; Japanese encephalitis virus; Vesicular exanthernavirus; SARS-CoV-2; Eastern equine encephalitis, or a combination of any two or more of the foregoing.
130. The composition of any one of claims 119 to 129, wherein the core functional element comprises a stop codon or a stop cassette.
131. The composition of any one of claims 119 to 129, wherein the core functional element comprises a noncoding region.
132. The composition of any one of claims 119 to 129, wherein the core functional element comprises an accessory or modulatory element.
133. The composition of claim 132, wherein the accessory or modulatory element comprises a miRNA binding site or a fragment thereof, a restriction site or a fragment thereof, an RNA editing motif or a fragment thereof, a zip code element or a fragment thereof, an RNA trafficking element or fragment thereof, or a combination thereof.
134. The composition of claim 132, wherein the accessory or modulatory element comprises a binding domain to an IRES transacting factor (ITAF).
135. The composition of claim 134, wherein the circular RNA comprises a 3’ internal spacer located upstream to the 5’ exon fragment.
136. The composition of claim 135, wherein the 3’ internal spacer is a polyA or polyA-C sequence.
137. The composition of claim 135 or 136, wherein the 3’ internal spacer has a length of 10 to 60 nucleotides.
138. The composition of claim 135 to 137, wherein the circular RNA comprises a 3’ internal duplex element located upstream to the 5’ exon fragment.
139. The composition of any one of claims 1 to 138, wherein the circular RNA polynucleotide is made via circularization of a RNA polynucleotide comprising, in the following order: a 3’ intron fragment, b. a 3’ exon fragment, c. a core functional element, d. a 5’ exon fragment, and e. a 5’ intron fragment.
140. The composition of claim 139, wherein the 3’ intron fragment comprises a first or a first and second nucleotide of a 3’ group I intron splice site dinucleotide.
141. The composition of claim 139 or 140, wherein the RNA polynucleotide comprises a 5’ affinity tag located upstream to the 3’ intron fragment.
142. The composition of any one of claims 139 to 141, wherein the 5’ RNA polynucleotide comprises a 5’ external spacer located upstream to the 3’ intron fragment.
143. The composition of any one of claims 139 to 142 wherein the 5’ RNA polynucleotide comprises a leading untranslated sequence located at the 5’ end of said 5’ enhanced intron element.
144. The composition of any one of claims 139 to 143, wherein the RNA polynucleotide comprises a 3’ external spacer located downstream to the 5’ intron fragment.
145. The composition of any one of claims 139 to 144, wherein the RNA polynucleotide comprises a 3’ affinity tag located downstream to the 5’ intron fragment.
146. The composition of any one of claims 139 to 145, wherein the RNA polynucleotide comprises a 3’ terminal untranslated sequence at the 3’ end of the said RNA polynucleotide.
147. The composition of any one of claims 138 to 146, wherein the RNA polynucleotide comprises a 5’ external duplex region upstream to the 3’ intron fragment, and a 3’ external duplex region downstream to the 5’ intron fragment.
148. The composition of claim 147, wherein the 5’ external duplex region and the 3’ external duplex region are the same.
149. The composition of claim 147, wherein the 5’ external duplex region and the 3’ external duplex region are different.
150. The composition of any one of claims 139 to 149, wherein the group I intron comprises in part or in whole from a bacterial phage, viral vector, organelle genome, or a nuclear rDNA gene.
151. The composition of claim 150, wherein the nuclear rDNA gene comprises a nuclear rDNA gene derived from a fungi, plant, or algae, or a fragment thereof.
152. The composition of any one of claims 1 to 151, wherein the circular RNA polynucleotide contains at least 80% naturally occurring nucleotides.
153. The composition of claim 151 or 152, wherein the expression sequence is codon optimized.
154. The composition of any one of claims 1 to 153, wherein the circular RNA polynucleotide is optimized to lack at least one micrcircRNA binding site present in an equivalent pre-optimized polynucleotide.
155. The composition of any one of claims 1 to 154, wherein the circular RNA polynucleotide is optimized to lack at least one micrcircRNA binding site capable of binding to a micrcircRNA present in a cell within which the RNA polynucleotide is expressed.
156. The composition of any one of claims 1 to 155, wherein the circular RNA polynucleotide is optimized to lack at least one endonuclease susceptible site present in an equivalent pre-optimized polynucleotide.
157. The composition of any one of claims 1 to 156, wherein the circular RNA polynucleotide is optimized to lack at least one endonuclease susceptible site capable of being cleaved by an endonuclease present in a cell within which the endonuclease is expressed.
158. The composition of any one of claims 1 to 157, wherein the circular RNA polynucleotide is optimized to lack at least one RNA editing susceptible site present in an equivalent pre-optimized polynucleotide.
159. The composition of any one of claims 1 to 158, wherein the circular RNA polynucleotide is from lOOnt to 15,000nt in length.
160. The composition of claim 159, wherein the RNA polynucleotide is from lOOnt to 10,000nt in length.
161. An emulsion comprising an aqueous continuous phase and an organic dispersed phase, wherein the organic dispersed phase comprises droplets that contain: a circular RNA polynucleotide;
an ionizable lipid; and an amphiphilic polymer.
162. A method of preparing a composition comprising a plurality of polymeric lipid nanoparticles, the method comprising: a) providing a mixture comprising: an ionizable lipid; and an amphiphilic polymer, both dissolved in an organic phase; b) combining the mixture with a circular RNA polynucleotide dissolved in an aqueous phase to obtain an emulsion; and c) adding the emulsion to an aqueous bath to form polymeric lipid nanoparticles comprising the ionizable lipid, the amphiphilic polymer, and the circular RNA polynucleotide.
163. The method of claim 162, wherein the method further comprises: d) separating the polymeric lipid nanoparticles from the aqueous bath.
164. The method of claim 162, wherein in step b) the ionizable lipid forms a complex with the circular RNA polynucleotide (circRNA-lipid complex).
165. The method of claim 162, wherein in step c) the circRNA-lipid complex is encapsulated in the nanoparticles with an encapsulation efficiency of at least 70%.
166. The method of claim 165, wherein in step c) the circRNA-lipid complex is encapsulated in the nanoparticles with an encapsulation efficiency of 70-90%.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363511574P | 2023-06-30 | 2023-06-30 | |
| US63/511,574 | 2023-06-30 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025007148A1 true WO2025007148A1 (en) | 2025-01-02 |
Family
ID=92108292
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/036442 Pending WO2025007148A1 (en) | 2023-06-30 | 2024-07-01 | Polymer lipid nanoparticle compositions for delivering circular polynucleotides |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025007148A1 (en) |
Citations (51)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US3832253A (en) | 1973-03-21 | 1974-08-27 | Baxter Laboratories Inc | Method of making an inflatable balloon catheter |
| US3854480A (en) | 1969-04-01 | 1974-12-17 | Alza Corp | Drug-delivery system |
| US4450150A (en) | 1973-05-17 | 1984-05-22 | Arthur D. Little, Inc. | Biodegradable, implantable drug delivery depots, and method for preparing and using the same |
| US4452775A (en) | 1982-12-03 | 1984-06-05 | Syntex (U.S.A.) Inc. | Cholesterol matrix delivery system for sustained release of macromolecules |
| US4667014A (en) | 1983-03-07 | 1987-05-19 | Syntex (U.S.A.) Inc. | Nonapeptide and decapeptide analogs of LHRH, useful as LHRH antagonists |
| US4748034A (en) | 1983-05-13 | 1988-05-31 | Nestec S.A. | Preparing a heat stable aqueous solution of whey proteins |
| US5075109A (en) | 1986-10-24 | 1991-12-24 | Southern Research Institute | Method of potentiating an immune response |
| US5087616A (en) | 1986-08-07 | 1992-02-11 | Battelle Memorial Institute | Cytotoxic drug conjugates and their delivery to tumor cells |
| US5239660A (en) | 1990-10-31 | 1993-08-24 | Nec Corporation | Vector processor which can be formed by an integrated circuit of a small size |
| US5948902A (en) | 1997-11-20 | 1999-09-07 | South Alabama Medical Science Foundation | Antisense oligonucleotides to human serine/threonine protein phosphatase genes |
| US6319494B1 (en) | 1990-12-14 | 2001-11-20 | Cell Genesys, Inc. | Chimeric chains for receptor-associated signal transduction pathways |
| WO2006000830A2 (en) | 2004-06-29 | 2006-01-05 | Avidex Ltd | Cells expressing a modified t cell receptor |
| US20100130588A1 (en) | 2008-04-15 | 2010-05-27 | Protiva Biotherapeutics, Inc. | Novel lipid formulations for nucleic acid delivery |
| US7741465B1 (en) | 1992-03-18 | 2010-06-22 | Zelig Eshhar | Chimeric receptor genes and cells transformed therewith |
| US20130053572A1 (en) | 2010-01-22 | 2013-02-28 | Steven L. Colletti | Novel Cationic Lipids for Oligonucleotide Delivery |
| US20130108685A1 (en) | 2010-04-28 | 2013-05-02 | Takeshi Kuboyama | Cationic lipid |
| US20130195920A1 (en) | 2011-12-07 | 2013-08-01 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| US20140308304A1 (en) | 2011-12-07 | 2014-10-16 | Alnylam Pharmaceuticals, Inc. | Lipids for the delivery of active agents |
| US20150005363A1 (en) | 2011-12-07 | 2015-01-01 | Alnylam Pharmaceuticals, Inc. | Branched Alkyl And Cycloalkyl Terminated Biodegradable Lipids For The Delivery Of Active Agents |
| WO2015095340A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| US20170119904A1 (en) | 2015-10-28 | 2017-05-04 | Acuitas Therapeutics, Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| US20170204422A1 (en) * | 2014-07-16 | 2017-07-20 | Moderna Therapeutics, Inc. | Circular polynucleotides |
| US20170210697A1 (en) | 2015-09-17 | 2017-07-27 | Modernatx, Inc. | Compounds and compositions for intracellular delivery of therapeutic agents |
| WO2019089828A1 (en) | 2017-10-31 | 2019-05-09 | Acuitas Therapeutics, Inc. | Lamellar lipid nanoparticles |
| WO2019110067A1 (en) * | 2017-12-07 | 2019-06-13 | Aarhus Universitet | Hybrid nanoparticle |
| US20190240354A1 (en) | 2016-06-30 | 2019-08-08 | Arbutus Biopharma Corporation | Compositions and methods for delivering messenger rna |
| WO2019152557A1 (en) | 2018-01-30 | 2019-08-08 | Modernatx, Inc. | Compositions and methods for delivery of agents to immune cells |
| WO2019232095A1 (en) | 2018-05-30 | 2019-12-05 | Translate Bio, Inc. | Vitamin cationic lipids |
| WO2019236673A1 (en) | 2018-06-06 | 2019-12-12 | Massachusetts Institute Of Technology | Circular rna for translation in eukaryotic cells |
| WO2020237227A1 (en) | 2019-05-22 | 2020-11-26 | Massachusetts Institute Of Technology | Circular rna compositions and methods |
| WO2021021634A1 (en) | 2019-07-29 | 2021-02-04 | Georgia Tech Research Corporation | Nanomaterials containing constrained lipids and uses thereof |
| US20210087135A1 (en) | 2019-09-19 | 2021-03-25 | Modernatx, Inc. | Branched tail lipid compounds and compositions for intracellular delivery of therapeutic agents |
| WO2021077067A1 (en) | 2019-10-18 | 2021-04-22 | The Trustees Of The University Of Pennsylvania | Lipid nanoparticles and formulations thereof for car mrna delivery |
| US20210128488A1 (en) | 2017-08-16 | 2021-05-06 | Acuitas Therapeutics, Inc. | Lipids for use in lipid nanoparticle formulations |
| WO2021113777A2 (en) | 2019-12-04 | 2021-06-10 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| US20210259982A1 (en) | 2015-08-21 | 2021-08-26 | Pfizer Inc. | Therapeutic Nanoparticles Comprising A Therapeutic Agent And Methods of Making And Using Same |
| WO2021189059A2 (en) | 2020-03-20 | 2021-09-23 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021204179A1 (en) | 2020-04-09 | 2021-10-14 | Suzhou Abogen Biosciences Co., Ltd. | Nucleic acid vaccines for coronavirus |
| WO2021226597A2 (en) | 2020-05-08 | 2021-11-11 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021236855A1 (en) | 2020-05-19 | 2021-11-25 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| EP4023249A1 (en) * | 2014-04-23 | 2022-07-06 | ModernaTX, Inc. | Nucleic acid vaccines |
| WO2022251665A1 (en) | 2021-05-28 | 2022-12-01 | Renagade Therapeutics Management Inc. | Lipid nanoparticles and methods of use thereof |
| WO2022261490A2 (en) | 2021-06-10 | 2022-12-15 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| US20230053437A1 (en) | 2020-08-20 | 2023-02-23 | Suzhou Abogen Biosciences Co., Ltd. | Lipid compounds and lipid nanoparticle compositions |
| WO2023044343A1 (en) | 2021-09-14 | 2023-03-23 | Renagade Therapeutics Management Inc. | Acyclic lipids and methods of use thereof |
| WO2023044333A1 (en) | 2021-09-14 | 2023-03-23 | Renagade Therapeutics Management Inc. | Cyclic lipids and methods of use thereof |
| WO2023056033A1 (en) | 2021-09-30 | 2023-04-06 | Orna Therapeutics, Inc. | Lipid nanoparticle compositions for delivering circular polynucleotides |
| WO2023081526A1 (en) | 2021-11-08 | 2023-05-11 | Orna Therapeutics, Inc. | Lipid nanoparticle compositions for delivering circular polynucleotides |
| WO2023122752A1 (en) | 2021-12-23 | 2023-06-29 | Renagade Therapeutics Management Inc. | Constrained lipids and methods of use thereof |
| WO2023196931A1 (en) | 2022-04-07 | 2023-10-12 | Renagade Therapeutics Management Inc. | Cyclic lipids and lipid nanoparticles (lnp) for the delivery of nucleic acids or peptides for use in vaccinating against infectious agents |
| WO2024044728A1 (en) | 2022-08-26 | 2024-02-29 | Renagade Therapeutics Management Inc. | Pegylated lipid compounds and methods of use thereof |
-
2024
- 2024-07-01 WO PCT/US2024/036442 patent/WO2025007148A1/en active Pending
Patent Citations (52)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US3854480A (en) | 1969-04-01 | 1974-12-17 | Alza Corp | Drug-delivery system |
| US3832253A (en) | 1973-03-21 | 1974-08-27 | Baxter Laboratories Inc | Method of making an inflatable balloon catheter |
| US4450150A (en) | 1973-05-17 | 1984-05-22 | Arthur D. Little, Inc. | Biodegradable, implantable drug delivery depots, and method for preparing and using the same |
| US4452775A (en) | 1982-12-03 | 1984-06-05 | Syntex (U.S.A.) Inc. | Cholesterol matrix delivery system for sustained release of macromolecules |
| US4667014A (en) | 1983-03-07 | 1987-05-19 | Syntex (U.S.A.) Inc. | Nonapeptide and decapeptide analogs of LHRH, useful as LHRH antagonists |
| US4748034A (en) | 1983-05-13 | 1988-05-31 | Nestec S.A. | Preparing a heat stable aqueous solution of whey proteins |
| US5087616A (en) | 1986-08-07 | 1992-02-11 | Battelle Memorial Institute | Cytotoxic drug conjugates and their delivery to tumor cells |
| US5075109A (en) | 1986-10-24 | 1991-12-24 | Southern Research Institute | Method of potentiating an immune response |
| US5239660A (en) | 1990-10-31 | 1993-08-24 | Nec Corporation | Vector processor which can be formed by an integrated circuit of a small size |
| US6319494B1 (en) | 1990-12-14 | 2001-11-20 | Cell Genesys, Inc. | Chimeric chains for receptor-associated signal transduction pathways |
| US7741465B1 (en) | 1992-03-18 | 2010-06-22 | Zelig Eshhar | Chimeric receptor genes and cells transformed therewith |
| US5948902A (en) | 1997-11-20 | 1999-09-07 | South Alabama Medical Science Foundation | Antisense oligonucleotides to human serine/threonine protein phosphatase genes |
| WO2006000830A2 (en) | 2004-06-29 | 2006-01-05 | Avidex Ltd | Cells expressing a modified t cell receptor |
| US20100130588A1 (en) | 2008-04-15 | 2010-05-27 | Protiva Biotherapeutics, Inc. | Novel lipid formulations for nucleic acid delivery |
| US20130053572A1 (en) | 2010-01-22 | 2013-02-28 | Steven L. Colletti | Novel Cationic Lipids for Oligonucleotide Delivery |
| US20130108685A1 (en) | 2010-04-28 | 2013-05-02 | Takeshi Kuboyama | Cationic lipid |
| US20130195920A1 (en) | 2011-12-07 | 2013-08-01 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| US20140308304A1 (en) | 2011-12-07 | 2014-10-16 | Alnylam Pharmaceuticals, Inc. | Lipids for the delivery of active agents |
| US20150005363A1 (en) | 2011-12-07 | 2015-01-01 | Alnylam Pharmaceuticals, Inc. | Branched Alkyl And Cycloalkyl Terminated Biodegradable Lipids For The Delivery Of Active Agents |
| WO2015095340A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| EP4023249A1 (en) * | 2014-04-23 | 2022-07-06 | ModernaTX, Inc. | Nucleic acid vaccines |
| US20170204422A1 (en) * | 2014-07-16 | 2017-07-20 | Moderna Therapeutics, Inc. | Circular polynucleotides |
| US20210259982A1 (en) | 2015-08-21 | 2021-08-26 | Pfizer Inc. | Therapeutic Nanoparticles Comprising A Therapeutic Agent And Methods of Making And Using Same |
| US20170210697A1 (en) | 2015-09-17 | 2017-07-27 | Modernatx, Inc. | Compounds and compositions for intracellular delivery of therapeutic agents |
| US20170119904A1 (en) | 2015-10-28 | 2017-05-04 | Acuitas Therapeutics, Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| US20200121809A1 (en) | 2015-10-28 | 2020-04-23 | Erikc A. HARWOOD | Lipid nanoparticle formulations |
| US20190240354A1 (en) | 2016-06-30 | 2019-08-08 | Arbutus Biopharma Corporation | Compositions and methods for delivering messenger rna |
| US20210128488A1 (en) | 2017-08-16 | 2021-05-06 | Acuitas Therapeutics, Inc. | Lipids for use in lipid nanoparticle formulations |
| WO2019089828A1 (en) | 2017-10-31 | 2019-05-09 | Acuitas Therapeutics, Inc. | Lamellar lipid nanoparticles |
| WO2019110067A1 (en) * | 2017-12-07 | 2019-06-13 | Aarhus Universitet | Hybrid nanoparticle |
| WO2019152557A1 (en) | 2018-01-30 | 2019-08-08 | Modernatx, Inc. | Compositions and methods for delivery of agents to immune cells |
| WO2019232095A1 (en) | 2018-05-30 | 2019-12-05 | Translate Bio, Inc. | Vitamin cationic lipids |
| WO2019236673A1 (en) | 2018-06-06 | 2019-12-12 | Massachusetts Institute Of Technology | Circular rna for translation in eukaryotic cells |
| WO2020237227A1 (en) | 2019-05-22 | 2020-11-26 | Massachusetts Institute Of Technology | Circular rna compositions and methods |
| WO2021021634A1 (en) | 2019-07-29 | 2021-02-04 | Georgia Tech Research Corporation | Nanomaterials containing constrained lipids and uses thereof |
| US20210087135A1 (en) | 2019-09-19 | 2021-03-25 | Modernatx, Inc. | Branched tail lipid compounds and compositions for intracellular delivery of therapeutic agents |
| WO2021077067A1 (en) | 2019-10-18 | 2021-04-22 | The Trustees Of The University Of Pennsylvania | Lipid nanoparticles and formulations thereof for car mrna delivery |
| WO2021113777A2 (en) | 2019-12-04 | 2021-06-10 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021189059A2 (en) | 2020-03-20 | 2021-09-23 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021204179A1 (en) | 2020-04-09 | 2021-10-14 | Suzhou Abogen Biosciences Co., Ltd. | Nucleic acid vaccines for coronavirus |
| WO2021226597A2 (en) | 2020-05-08 | 2021-11-11 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021236855A1 (en) | 2020-05-19 | 2021-11-25 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| US20230053437A1 (en) | 2020-08-20 | 2023-02-23 | Suzhou Abogen Biosciences Co., Ltd. | Lipid compounds and lipid nanoparticle compositions |
| WO2022251665A1 (en) | 2021-05-28 | 2022-12-01 | Renagade Therapeutics Management Inc. | Lipid nanoparticles and methods of use thereof |
| WO2022261490A2 (en) | 2021-06-10 | 2022-12-15 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2023044343A1 (en) | 2021-09-14 | 2023-03-23 | Renagade Therapeutics Management Inc. | Acyclic lipids and methods of use thereof |
| WO2023044333A1 (en) | 2021-09-14 | 2023-03-23 | Renagade Therapeutics Management Inc. | Cyclic lipids and methods of use thereof |
| WO2023056033A1 (en) | 2021-09-30 | 2023-04-06 | Orna Therapeutics, Inc. | Lipid nanoparticle compositions for delivering circular polynucleotides |
| WO2023081526A1 (en) | 2021-11-08 | 2023-05-11 | Orna Therapeutics, Inc. | Lipid nanoparticle compositions for delivering circular polynucleotides |
| WO2023122752A1 (en) | 2021-12-23 | 2023-06-29 | Renagade Therapeutics Management Inc. | Constrained lipids and methods of use thereof |
| WO2023196931A1 (en) | 2022-04-07 | 2023-10-12 | Renagade Therapeutics Management Inc. | Cyclic lipids and lipid nanoparticles (lnp) for the delivery of nucleic acids or peptides for use in vaccinating against infectious agents |
| WO2024044728A1 (en) | 2022-08-26 | 2024-02-29 | Renagade Therapeutics Management Inc. | Pegylated lipid compounds and methods of use thereof |
Non-Patent Citations (51)
| Title |
|---|
| "ASHP Handbook on Injectable Drugs", 1986, pages: 622 - 630 |
| "Pharmaceutics and Pharmacy Practice", 1982, J.B. LIPPINCOTT COMPANY, pages: 238 - 250 |
| ABRAHAM ET AL., SCIENCE, vol. 295, 2002, pages 647651 - 651 |
| BELLIVEAU, N.M. ET AL., MOL. THER. NUCLEIC. ACIDS., vol. 1, 2012, pages 37 |
| BORDEN: "Molecular Basis of Cancer", 2015 |
| CHEN, D. ET AL., J. AM. CHEM. SOC., vol. 134, no. 16, 2012, pages 6948 - 51 |
| DHAMNE ET AL.: "Peripheral and thymic Foxp3+ regulatory T cells in search of origin, distinction, and function", FRONTIERS IN IMMUNOL, vol. 4, no. 253, 2013, pages 1 - 11 |
| DING ET AL., JOURNAL OF CONTROLLED RELEASE, vol. 348, 2022, pages 206 - 238 |
| DOBRIKOVA ET AL., PROC. NATL. ACAD. SCI., vol. 100, no. 25, 2003, pages 15125 - 15130 |
| EDGE, NATURE, vol. 292, 1981, pages 756 |
| ELIEL: "Stereochemistry of Carbon Compounds", 1962, MCGRAW-HILL |
| ESHHAR ET AL., CANCER IMMUNOL IMMUNOTHERAPY, vol. 45, 1997, pages 131 - 136 |
| FINNEY ET AL., JOURNAL OF IMMUNOLOGY, vol. 161, 1998, pages 2791 - 2797 |
| FRIESS M ET AL., FRONT. IMMUNOL., vol. 2947, no. 9, 2018 |
| GARLAPATI ET AL., J. BIOL. CHEM., vol. 279, no. 5, 2004, pages 3389 - 3397 |
| GROSS ET AL., AMUR. REV. PHARMACOL. TOXICOL., vol. 56, 2016, pages 59 - 83 |
| GURTU ET AL., BIOCHEM. BIOPHYS. RES. COMM., vol. 229, 1996, pages 295 - 298 |
| HILLERY A M ET AL.: "Drug Delivery and Targeting: For Pharmacists and Pharmaceutical Scientists", 2002, TAYLOR & FRANCIS, INC. |
| JACQUES ET AL.: "Enantiomers, Racemates and Resolutions", 1981, WILEY INTERSCIENCE |
| JANEWAY C ET AL.: "Immunobiology: The Immune System in Health and Disease", 2001, GARLAND SCIENCE |
| JANG ET AL., J. VIROL., vol. 63, 1989, pages 1651 - 1660 |
| JAY ET AL., J. BIOL. CHEM., vol. 259, 1984, pages 631 - 1 |
| JAYARAMAN ET AL., ANGEW. CHEM. INT. ED., vol. 51, 2012, pages 8529 - 8533 |
| JAYARAMAN ET AL., PROC. NATL. ACAD. SCI. USA, vol. 88, 1991, pages 4084 - 4088 |
| JONES ET AL., NATURE, vol. 54, 1986, pages 75 - 82 |
| JOSHI ET AL., INTERNATIONAL JOURNAL OF PHARMACEUTICS, vol. 590, 2020, pages 119915 |
| KALOS ET AL., SCI TRANSL. MED., vol. 3, 2011, pages 95 |
| KAUFMAN ET AL., NUC. ACIDS RES., vol. 19, 1991, pages 4485 - 4490 |
| KOBAYASHI ET AL., BIOTECHNIQUES, vol. 21, 1996, pages 399 - 402 |
| KRAUSE ET AL., J. EXP. MED., vol. 188, no. 4, 1998, pages 619 - 626 |
| KUBALL J ET AL., J EXP MED, vol. 206, no. 2, 2009, pages 463 - 475 |
| LEHTIMAKILAHESMAA: "Regulatory T cells control immune responses through their non-redundant tissue specific features", FRONTIERS IN IMMUNOL, vol. 4, no. 294, 2013, pages 1 - 10 |
| MOSSER ET AL., BIOTECHNIQUES, vol. 22, 1997, pages 150 - 161 |
| NAMBAIR ET AL., SCIENCE, vol. 223, 1984, pages 1299 |
| PORTER ET AL., N. ENGL. J. MED., vol. 365, 2011, pages 725 - 33 |
| QUEEN ET AL., PROC. NATL. ACAD. SCI. USA, vol. 86, 1989, pages 10029 - 10033 |
| RAMESH ET AL., NUCL. ACID RES., vol. 24, 1996, pages 2697 - 2700 |
| RIECHMANN ET AL., NATURE, vol. 332, 1988, pages 323 - 327 |
| ROBBINS ET AL., J IMMUNOL., vol. 180, 2008, pages 6116 - 6131 |
| ROSENBERG ET AL., NAT REV CANCER, vol. 8, no. 4, 2008, pages 299 - 308 |
| SHIN ET AL., MOLECULAR PHARMACEUTICS, vol. 9, no. 11, 2012, pages 3266 - 3276 |
| SONG ET AL., BLOOD, vol. 119, 2012, pages 696 - 706 |
| VERHOEYEN ET AL., SCIENCE, vol. 239, 1988, pages 1534 - 1536 |
| WADWA ET AL., J, DRUG TARGETING, vol. 3, 1995, pages 111 |
| WESSELHOEFT ET AL.: "Engineering circular RNA for Potent and Stable Translation in Eukaryotic Cells", NATURE COMMUNICATIONS, vol. 9, 2018, pages 2629, XP055622155, DOI: 10.1038/s41467-018-05096-6 |
| WESSELHOEFT ET AL.: "RNA Circularization Diminishes Immunogenicity and Can Extend Translation Duration In vivo", MOLECULAR CELL, vol. 74, no. 3, 2019, pages 508 - 520 |
| WHITESIDES, NATURE, vol. 442, 2006, pages 368 - 373 |
| WILEN ET AL.: "Tetrahedron", vol. 33, 1977, pages: 2725 |
| WILEN: "Tables of Resolving Agents and Optical Resolutions", 1972, UNIV. OF NOTRE DAME PRESS, pages: 268 |
| ZHIGALTSEV, I.V. ET AL., LANGMUIR, vol. 28, 2012, pages 3633 - 40 |
| ZHU ET AL., PNAS, vol. 110, no. 42, 2013, pages 17047 - 17052 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US12415776B2 (en) | Lipid nanoparticle compositions for delivering circular polynucleotides | |
| US20250283073A1 (en) | Circular RNA Compositions and Methods | |
| EP3920976B1 (en) | Circular rna compositions and methods | |
| WO2023056033A1 (en) | Lipid nanoparticle compositions for delivering circular polynucleotides | |
| KR20230069042A (en) | Circular RNA compositions and methods | |
| US12297285B2 (en) | Circular RNA encoding chimeric antigen receptors targeting BCMA | |
| US20250304994A1 (en) | Circular RNA Compositions and Methods | |
| WO2024102762A1 (en) | Lipids and lipid nanoparticle compositions for delivering polynucleotides | |
| WO2024233308A2 (en) | Circular rna compositions and methods | |
| WO2024205657A2 (en) | Lipids and lipid nanoparticle compositions for delivering polynucleotides | |
| WO2025007148A1 (en) | Polymer lipid nanoparticle compositions for delivering circular polynucleotides | |
| WO2025117969A1 (en) | Process for manufacturing lipid nanoparticles | |
| WO2025049690A1 (en) | Circular polyethylene glycol lipids | |
| WO2026006593A1 (en) | Synthetic internal ribosome entry sites | |
| WO2025166238A1 (en) | Fast-shedding polyethylene glycol lipids | |
| TW202425959A (en) | Lipids and nanoparticle compositions for delivering polynucleotides | |
| HK40065368B (en) | Circular rna compositions and methods | |
| HK40065368A (en) | Circular rna compositions and methods |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24748731 Country of ref document: EP Kind code of ref document: A1 |