WO2024233308A2 - Circular rna compositions and methods - Google Patents
Circular rna compositions and methods Download PDFInfo
- Publication number
- WO2024233308A2 WO2024233308A2 PCT/US2024/027627 US2024027627W WO2024233308A2 WO 2024233308 A2 WO2024233308 A2 WO 2024233308A2 US 2024027627 W US2024027627 W US 2024027627W WO 2024233308 A2 WO2024233308 A2 WO 2024233308A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- sequence
- exon
- intron
- segment
- rna
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/85—Vectors or expression systems specially adapted for eukaryotic hosts for animal cells
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K39/39—Medicinal preparations containing antigens or antibodies characterised by the immunostimulating additives, e.g. chemical adjuvants
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K40/00—Cellular immunotherapy
- A61K40/30—Cellular immunotherapy characterised by the recombinant expression of specific molecules in the cells of the immune system
- A61K40/31—Chimeric antigen receptors [CAR]
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/555—Medicinal preparations containing antigens or antibodies characterised by a specific combination antigen/adjuvant
- A61K2039/55511—Organic adjuvants
- A61K2039/55561—CpG containing adjuvants; Oligonucleotide containing adjuvants
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2830/00—Vector systems having a special element relevant for transcription
- C12N2830/42—Vector systems having a special element relevant for transcription being an intron or intervening sequence for splicing and/or stability of RNA
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2830/00—Vector systems having a special element relevant for transcription
- C12N2830/52—Vector systems having a special element relevant for transcription encoding ribozyme for self-inactivation
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2840/00—Vectors comprising a special translation-regulating system
- C12N2840/20—Vectors comprising a special translation-regulating system translation of more than one cistron
- C12N2840/203—Vectors comprising a special translation-regulating system translation of more than one cistron having an IRES
Definitions
- gene therapy with DNA may result in the impairment of a vital genetic function in the treated host, such as e.g., elimination or deleteriously reduced production of an essential enzyme or interruption of a gene critical for the regulation of cell growth, resulting in unregulated or cancerous cell proliferation.
- a vital genetic function such as e.g., elimination or deleteriously reduced production of an essential enzyme or interruption of a gene critical for the regulation of cell growth, resulting in unregulated or cancerous cell proliferation.
- it is necessary for effective expression of the desired gene product to include a strong promoter sequence which again may lead to undesirable changes in the regulation of normal gene expression in the cell.
- the DNA based genetic material will result in the induction of undesired anti-DNA antibodies, which in turn, may trigger a possibly fatal immune response.
- Gene therapy approaches using viral vectors can also result in an adverse immune response. In some circumstances, the viral vector may even integrate into the host genome.
- RNA does not involve the risk of being stably integrated into the genome of the transfected cell, thus eliminating the concern that the introduced genetic material will disrupt the normal functioning of an essential gene, or cause a mutation that results in deleterious or oncogenic effects, and extraneous promoter sequences are not required for effective translation of the encoded protein, again avoiding possible deleterious side effects.
- RNA it is not necessary for RNA to enter the nucleus to perform its function, while DNA must overcome this major barrier.
- Circular RNA (circRNA or oRNA®) is a stable form of RNA that provides an advantage compared to linear RNA in structure and function, especially in the case of molecules that are prone to folding in an inactive conformation (Wang and Ruffner, 1998).
- Circular RNA polynucleotides lack the free ends necessary for exonuclease-mediated degradation, causing them to be resistant to several mechanisms of RNA degradation and granting extended half-lives when compared to an equivalent linear RNA. Circularization may allow for the stabilization of RNA polynucleotides that generally suffer from short half-lives and may improve the overall efficacy of exogenous mRNA in a variety of applications.
- Circular RNA can also be particularly interesting and useful for in vivo applications, especially in the research area of RNA-based control of gene expression and therapeutics, including protein replacement therapy and vaccination.
- RNA ligase-mediated method Three main techniques for making circularized RNA in vitro are the splint-mediated method, the permuted intron-exon method, and the RNA ligase-mediated method.
- existing methodologies may be limited by the size of RNA that can be circularized, thus limiting their therapeutic application.
- the present disclosure addresses this need by providing methods and compositions for the manufacture and preparation of circularized RNAs via engineering of the sequences for the DNA template, precursor linear RNA, and ultimately the circular RNA, along with methods of manufacturing and preparing the circular RNA, and methods of treating a subject in need using said circular RNA.
- circular RNA polynucleotides also referred to herein as “circular RNA” comprising, in the following order, a 3’ self-spliced exon segment, an intervening region, and a 5’ self-spliced exon segment.
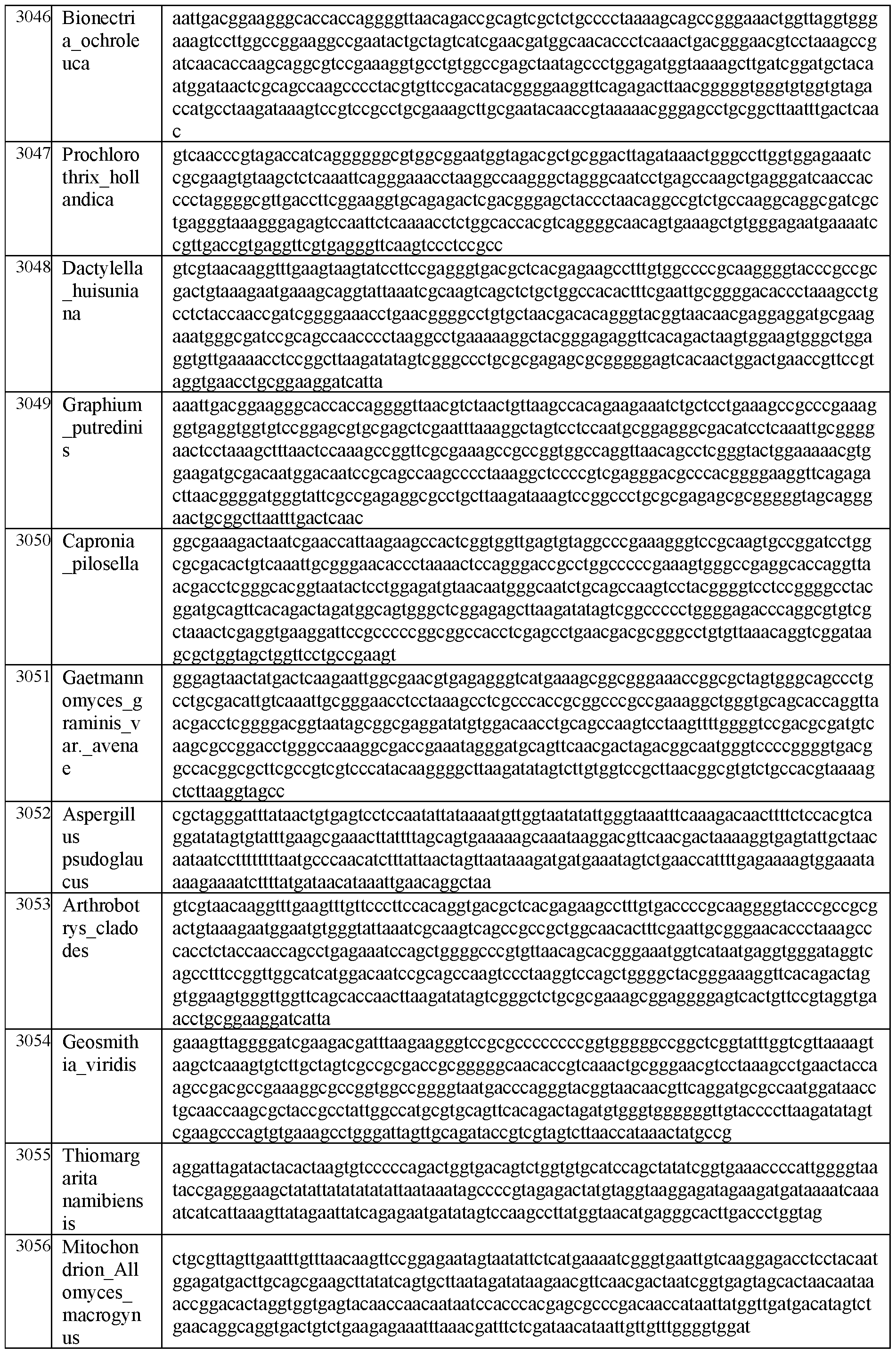
- the 3’ self-spliced exon segment and/or the 5’ self-spliced exon segment is selected from an exon segment disclosed herein, e.g., in Table A or Table B, or SEQ ID NOs: 2990-3668, 25573, and 25574.
- RNA polynucleotides comprising, in the following order, i) a 5’ combined accessory element comprising a 3’ self- spliced exon segment; ii) an intervening region; and iii) a 3’ combined accessory element comprising a 5’ self-spliced exon segment.
- the 3’ self-spliced exon segment and/or the 5’ self-spliced exon segment is selected from an exon segment disclosed herein, e.g., in Table A or Table B, or SEQ ID NOs: 2990-3668, 25573, and 25574.
- RNA polynucleotides comprising, in the following order, i) a 5’ combined accessory element comprising a 3’ self- spliced exon segment, wherein the 3’ self-spliced exon segment comprises an exon segment; ii) an intervening region; and iii) a 3’ combined accessory element comprising a 5’ self-spliced exon segment, wherein the 5’ self-spliced exon segment comprises an exon segment.
- the 3’ self-spliced exon segment and/or the 5’ self-spliced exon segment is selected from an exon segment disclosed herein, e.g., in Table A or Table B, or SEQ ID NOs: 2990-3668, 25573, and 25574.
- RNA polynucleotides comprising, in the following order, i) a 5’ combined accessory element comprising a 3’ self- spliced exon segment, wherein the 3’ self-spliced exon segment comprises an exon segment and a 3’ nucleotide of a 3’ splice site dinucleotide; ii) an intervening region; and iii) a 3’ combined accessory element comprising a 5’ self-spliced exon segment, wherein the 5’ self- spliced exon segment comprises an exon segment and a 5’ nucleotide of a 5’ splice site dinucleotide.
- the 3’ self-spliced exon segment and/or the 5’ self-spliced exon segment is selected from an exon segment disclosed herein, e.g., in Table A or Table B, or SEQ ID NOs: 2990-3668, 25573, and 25574.
- RNA polynucleotide comprising, in the following order, a 3’ self-spliced exon segment, an intervening region, and a 5’ self-spliced exon segment, wherein at least one of the 3’ or 5’ self-spliced exon segments is selected from an exon segment comprising a sequence selected from SEQ ID NOs: 2990- 3668, 25573, and 25574.
- precursor linear RNA polynucleotides that are capable of autocatalytically self-splicing and forming the circular RNA described herein, retaining the intervening region and a 3’ exon segment and a 5’ exon segment.
- precursor linear RNA comprising, for example, a 5 ' combined accessory element comprising a 3 ' permuted intron segment; an intervening region; and a 3 ' combined accessory element comprising a 5 ' permuted intron segment.
- precursor linear RNA comprising, for example, a 5 ' combined accessory element comprising a 3 ' permuted intron segment and a 3 ' permuted exon segment; an intervening region; and a 3 ' combined accessory element comprising a 5 ' permuted intron segment and a 5 ' permuted exon segment.
- precursor linear RNA comprising a monotron, an intervening region, and a terminal sequence; or a terminal sequence, an intervening region, and a monotron.
- precursor linear RNA comprising at least one modified A, C, G, or U/T nucleotide or nucleoside.
- RNA produced from these precursors results in improved circularization efficiency and/or splicing efficiency as compared to a control RNA polynucleotide comprising a native intronic sequence.
- FIG. 1A depicts a size exclusion-high-performance liquid chromatography (SEC- HPLC) analysis of a post-IVT reaction solution. Milli-absorbance units (mAU) were measured over the course of the retention time. The IVT reaction was performed on a DNA template comprising Anabaena intron and exon segments. The largest peak in the figure (beginning approximately at 9.25 min) provides the circular RNA collected post-IVT of the DNA template. The two peaks to the right of the largest peak (beginning approximately at 11.5 and 13 minutes) correspond to the spliced-out 5’ and 3’ intron fragments produced post-IVT of the DNA template.
- FIG. 1A depicts a size exclusion-high-performance liquid chromatography (SEC- HPLC) analysis of a post-IVT reaction solution. Milli-absorbance units (mAU) were measured over the course of the retention time. The IVT reaction was performed on a DNA template comprising Anabaena intron and exon segments. The largest peak in the
- FIG. 1B illustrates a permuted intron construct design schematic used to develop the exemplary DNA templates of FIG. la.
- FIG. 1C shows the general placement of the splice site dinucleotides in the exemplary DNA templates.
- FIG. ID provides an illustration of the splicing reaction of a linear precursor wherein the permutation occurs in a 5 ’ orientation created post IVT of the exemplary DNA template and resulting circular product.
- FIG. IE provides an illustration of the splicing reaction of a linear precursor wherein the permutation occurs in a 3’ orientation created post IVT of the exemplary DNA template and resulting circular product.
- FIG. 2 depicts fragment analyzer analysis of a post-IVT reaction solution.
- the IVT reaction was performed on a DNA template comprising Anabaena intron and exon segments. Relative fluorescent units (RFU) were measured for each of the peaks.
- the largest peak in the figure (2530 nucleotides) contains the circular RNA, precursor RNAs, or nicked circular RNAs produced post-IVT of the DNA template.
- the two peaks to the left of the largest peak (265 and 357) correspond to the spliced-out 5 ' and 3 ' intron fragments produced post-IVT of the DNA template.
- LM in the figure indicates the lowest marker.
- FIG. 3 depicts estimated percent circulation measured for various post-IVT reaction solutions using a fragment analyzer (“FA”) or a SEC-HPLC (“SEC”).
- IntronPeak and circPeak in the figure correspond to whether the intron peaks (“IntronPeak”) or the circular RNA peaks (“circPeak”) in the fragment analyzer and/or SEC-HPLC results were used to measure the estimated percent circularization.
- Post-IVT reaction solutions were formed from IVT reactions on 12 different DNA templates comprising a 5’ and 3 ’ Anabaena intron segment.
- FIG. 4A and FIG. 4B show intronic activity for permuted introns of several different species origins (e.g., from Azoarcus, Twort, Nostoc, Nodularia, S795, large subunit ribosomal RNA (LSU), Pleurocapsa, and/or Planktothrix).
- FIG. 4A Depicted in FIG. 4A are SEC- HPLC chromatographs of post-IVT DNA template solutions comprising introns for the species.
- FIG. 4B provides the circular to precursor RNA present post IVT fraction of each of the DNA templates used in FIG. 4a.
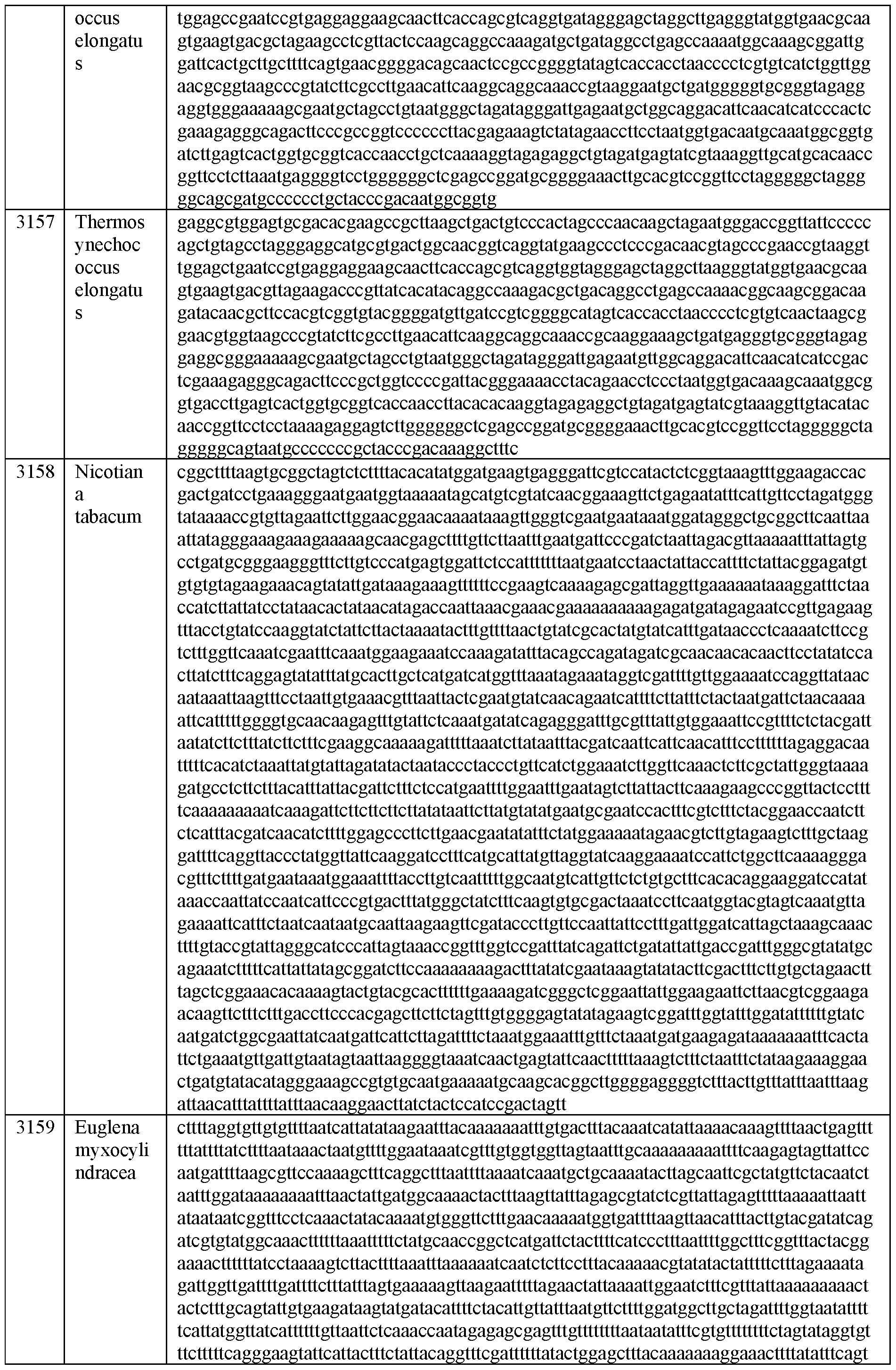
- FIG. 5A and FIG. 5B depict percent estimated excised introns for Group I introns (FIG. 5A) and Group II introns (FIG. 5B) from various species origins. % estimated intron in the figures represent the percent of intronic fragments present post IVT of DNA templates comprising said Group I or Group II introns. Intron and exon sequences for FIGs. 5a and 5b are present in SEQ ID NOS: 2990-3130 (Group I) and 3131-3187 (Group II), respectively.
- FIG. 6A, FIG. 6B, and FIG. 6C illustrate the permutation screening process used to preliminarily determine whether Group I or Group II intron generated maintained splicing activity post permutation.
- FIG. 6A represents the general intron screening process.
- FIG. 6B provides an example of 3 permutations - indicated by the arrows in the figure - that were made on a Hypocrea pallida sequence.
- FIG. 6C illustrates an exemplary linear RNA sequence (i.e., precursor RNA polynucleotide) schematic used to test the splicing activity of the permuted introns from FIG. 6A.
- RNA sequence i.e., precursor RNA polynucleotide
- FIG. 7 shows percent circularization for 6 naturally occurring introns (e.g. , Coxiella burnetii, Geosmithia argillacea, Agrobacterium tumefaciens, Hypocrea pallida, bulbithecium hyalosporum, and Myocarachis inversa at three different permutation sites. Percent circularization was measured from the IVT reaction of a DNA template comprising the permuted introns.
- DNA templates in FIG. 7 comprise sequences from SEQ ID NOS: 2991- 2993, 2997, 3054 and 2998.
- FIG. 8A and FIG. 8B depict permutation heat maps of regions of an exemplary group I intron (e.g., Anabaena (FIG. 8A) and Azoarcus (FIG. 8B)).
- Each location in FIG. 8A corresponds with a permuted intron sequence from SEQ ID NOS: 3222-3483.
- Each number in each figure indicates a tested permutation site.
- the splice junctions are indicated by the triangles.
- FIG. 9A provides percent circularization (e.g., estimated percent splicing) of Coxiella burnetti and Hypocrea pallida introns at 8 permutation sites. Percent circularization was measured from the IVT reaction of a DNA template comprising the permuted introns.
- FIG. 9B depicts the 8 permutation sites of Coxiella burnetti.
- FIG. 9C depicts the 8 permutation sites of Hypocrea pallida.
- each of the numbers in the key to the left of the figure are represented by the arrows along the sequence and correspond to a specific permutation site.
- FIG. 10A and FIG. 10B depict estimated percent splicing for RNA constructs with incrementally minimized exon segments.
- Estimated percent splicing for both FIG. 10a and 10b was collected from constructs comprising Anabaena introns segments, exon segments, a CVB3 internal ribosome entry site (IRES), and firefly luciferase coding regions.
- FIG. 10A depicts results from constructs with 5 ’-terminal exon deletions from naturally occurring Anabaena 5’ exons.
- Original (51nt) in FIG. 10a pertains to the RNA construct control comprising full-length 51 nucleotide (e.g., non-minimized) exon segments derived from SEQ ID NO: 3188.
- Constructs comprising minimized exon sequences used in FIG. 10a comprise a sequence from SEQ ID NOS: 3189-3197.
- FIG. 10B depicts results from constructs with 3’ terminal deletions from naturally occurring Anabaena 3’ exons.
- Original (15nt) in FIG. 10a pertains to the RNA construct control comprising full-length 15 nucleotide (e.g., nonminimized) exon segments.
- Constructs comprising minimized exon sequences used in FIG. 10b comprise a sequence from SEQ ID NOS: 3198-3205.
- 10C illustrates the direction of deletion for the 3’ and/or 5’ exon segments (ie., left arrow shows the incremental deletion from the 3’ end of the 3’ exon segment; the right arrow shows the incremental deletion from the 5’ end of the 5’ exon segment).
- FIG. 11A and FIG. 11B depict estimated percent splicing for RNA constructs with incrementally minimized exon segments.
- Estimated percent splicing for FIG. 11A and FIG. 11B was collected from constructs comprising Anabaena (FIG. 11 A) or Coxiella burnetti (FIG. 11B) introns segments and exon segments, a Caprine kobuvirus internal ribosome entry site (IRES), and firefly luciferase coding regions.
- FIG. 11A depicts results from constructs with 5’ or 3 ’-terminal exon deletions from a permuted Anabaena 5’ or 3’ exons. Constructs comprising minimized exon sequences in FIG.
- FIG. 11A comprise a sequence from SEQ ID NOS: 3579-3596.
- FIG. 11B depicts results from constructs with 5’ or 3 ’-terminal exon deletions from Coxiella burnetti 5’ or 3’ exons. Constructs comprising minimized exon sequences in FIG. 11B comprise a sequence from SEQ ID NOS: 3642-3664. 3’ exons (Pl) were deleted from the 3’ terminal end of the 3’ exon; 5’ exons (P2) were deleted from the 5’ terminal end of the 5’ exon in both FIG. 11A and FIG. 11B. Std refers to a non-minimized pair of exons in both FIG. 11A and FIG. 11B
- FIG. 12A and FIG. 12B depict estimated percent splicing of permuted intron-exon (PIE) constructs with a single nucleotide swap within the splice junctions of either the 3’ intron and/or sequences from Anabaena (FIG. 12A) or Coxiella burnettii (FIG. 12B).
- PIE constructs were derived from naturally occurring Anabaena intron and exon sequences and comprise of SEQ ID NOS: 3572-3578.
- FIG. 12A depicts nucleotide swaps from natural intron and exon sequences.
- FIG. 12B depicts nucleotide swaps for spacer constructs and Coxiella burnetti exon sequences.
- PIE constructs were derived from naturally occurring Anabaena intron and exon sequences. DNA templates for FIG. 12B comprised of SEQ ID NOS: 3635-3641.
- FIG. 13A and FIG. 13B depict percent estimated splicing for constructs comprising Anabaena (FIG. 13A) or Coxiella burnetti (FIG. 13B) permuted introns and exons with one or more nucleotide swaps in the exon segments. Nucleotide swaps were reverse complements and/or a random scramble of one or more nucleotides in either the 3 ' exon (Pl) or 5 ' exon (P2).
- FIG. 13A constructs comprised SEQ ID NOS: 3622 and 3624-3627.
- FIG. 13B constructs comprised SEQ ID NOS: 3665-3668.
- FIG. 14 provides a schematic of an intron deletion in one or more exemplary DNA templates.
- FIG. 14 depicts the deletion of 3’ intron segment including the naturally occurring nucleotide of the splice site dinucleotide.
- the internal and external accessory sequences indicated in the figure may comprise a spacer and/or homology arm.
- FIG. 15 depicts estimated percent splicing of DNA templates with a deleted 3" intron (Pl) (e.g., DNA templates comprised in the following 5 ' to 3 ' order: a 3 ' exon segment, an internal ribosome entry site (IRES), an expression sequence, a 5 ' exon segment and a 5 ' intron segment).
- a base pair of one of the splice junctions had also been swapped to one of the other three base pairs e.g., indicated in the figure as initial nucleotide > swapped nucleotide, e.g., C to G).
- Intron and exon segments were derived from Anabaena DNA plasmids.
- IRESes were derived from CVB3 IRESes and the expression sequence encodes firefly luciferase.
- DNA templates were comprised of sequences in whole or in part from SEQ ID NOS: 3597-3603.
- FIG. 16 depicts estimated percent splicing of constructs lacking a 3 ' intron segment.
- DNA templates comprised a 3 ' exon segment, an internal ribosome entry site (IRES), a firefly luciferase coding region, a 5 ' exon segment and a 5 ' intron segment. Each of the DNA templates also received exon minimization incrementally in the 5 ' and or 3 ' exon segments.
- DNA templates were comprised of sequences in whole or in part from SEQ ID NOS: 3597, and 3604-3621.
- FIG. 17A, FIG. 17B, and FIG. 17C depict a construct comprising a 5 ' terminal sequence and a 3 ' monotron sequence.
- FIG. 17A illustrates an exemplary DNA template comprising the 5 ' terminal sequence and 3 ' monotron sequence along with the placement of the splice site nucleotides.
- FIG. 17B provides an illustration of splicing and circularization process of a linear precursor of the DNA template in FIG. 17A.
- FIG. 17C depicts a size exclusion-high-performance liquid chromatography (SEC-HPLC) analysis of a post-IVT reaction solution of the DNA template in FIG. 17A. Milli-absorbance units (mAU) were measured over the course of the retention time.
- the IVT reaction was performed on a DNA template comprising Anabaena intron and exon segments. The largest peak in the figure (beginning approximately at 10 min) provides the circular RNA collected post-IVT of the DNA template.
- FIG. 18A, FIG. 18B, and FIG. 18C depicts a construct comprising a 3 ' terminal sequence and a 5 ' monotron sequence.
- FIG. 18A illustrates an exemplary DNA template comprising the 3 ' terminal sequence and 5 ' monotron sequence along with the placement of the splice site nucleotides.
- FIG. 18B provides an illustration of splicing and circularization process of a linear precursor of the DNA template in FIG. 18A.
- FIG. 18C depicts a size exclusion-high-performance liquid chromatography (SEC-HPLC) analysis of a post-IVT reaction solution of the DNA template in FIG. 18A. Milli-absorbance units (mAU) were measured over the course of the retention time.
- SEC-HPLC size exclusion-high-performance liquid chromatography
- FIG. 19A and FIG. 19B depict percent circular RNAs produced in RNA constructs developed with either Anabaena position 189 or position 230 permutation site in FIG. 8A.
- the constructs of the figures were designed to include accessory sequences (e.g., internal or external spacers and/or homology arms) or not include any accessory sequences (indicated by NA). Percent circular RNA produced was measured post SEC-HPLC analysis.
- FIG. 20 provides percent splicing for initial DNA templates comprising accessory elements, including internal homology arms (IH), external homology arms (EH), internal spacers (IS), and/or external spacers (ES) of different lengths, as compared to a control lacking accessory elements.
- the DNA templates were comprised of sequences from SEQ ID NOS: 3484-3571.
- FIG. 21 depicts estimated percent splicing for a construct with a 3 ' monotron element (P2) lacking internal homology arms (IH).
- the standard construct (“Std”) comprises two permuted intron exon elements and no monotron or terminal elements.
- DNA template comprised a sequence from SEQ ID NO: 3628 or 3633-3634.
- FIG. 22A and FIG. 22B depict estimated percent splicing for RNA constructs that were allowed to circularize co-transcriptionally and optionally allowed to refold (refolded constructs are indicated by "_R” in the figures).
- FIG. 22A illustrates results from two RNA constructs with different intron permutation sites. Each of the constructs in FIG. 22A were allowed to undergo co-transcription and the constructs indicated with "_R” were allowed an additional refold step.
- FIG. 22B shows results from RNA constructs with various 5 ' -terminal exon deletions (e.g., 10, 20, 40, 42, 44 nucleotide deletions). "Original” indicates the constructs containing non-minimized Anabaena exon structures with 51 nucleotides.
- FIG. 23A provides a schematic showing the incorporation of m6A modifications in linear RNA constructs to form circular RNA constructs comprising m6A modifications.
- FIG. 23B depicts a gel of a post IVT reaction of various RNA samples. RNA samples comprised RNA comprising either 0%, 1%, 5%, 10% or 100% fed m6A modifications, a CBV3 internal ribosome entry site (IRES), firefly luciferase coding region, and Anabaena permuted intron-exon (PIE) segments.
- IRS CBV3 internal ribosome entry site
- PIE Anabaena permuted intron-exon
- FIG. 24A illustrates naturally occurring DNA plasmids comprising exon and Group I or Group II introns sequences used to form linear precursor RNA with selective modification regions for FIG. 24B.
- FIG. 24B, FIG. 24C, and FIG. 24D illustrate exemplary depiction of the two linear precursor RNA used to form a single construct with certain regions lacking modifications (e.g., introns).
- one of the strands (top) comprises no modified nucleotides or nucleosides; the other of the two strands (bottom) comprises one or more modified nucleotides or nucleosides (indicated by the stars).
- FIG. 24A illustrates naturally occurring DNA plasmids comprising exon and Group I or Group II introns sequences used to form linear precursor RNA with selective modification regions for FIG. 24B.
- FIG. 24B, FIG. 24C, and FIG. 24D illustrate exemplary depiction of the two linear precursor RNA used to form a single construct with certain regions lacking modifications (e.g.
- FIG. 24B depicts two linear precursor RNAs, wherein each linear precursor RNA comprises two introns and two exon segments.
- FIG. 24C depicts two linear precursor RNAs, wherein each linear precursor RNA comprises a monotron intron and two exon segments.
- the dotted and non-dotted reactions in steps 2 and 3 may occur simultaneously or independently of each other (e.g., the dotted reactions could occur first for steps 2 and 3 then be followed by the non-dotted reactions in steps 2 and 3 (not depicted in FIG. 24B) or the dotted and non-reaction of step 2 occurs at the same time and then is followed by the dotted and non-dotted reactions of step 3 (depicted in FIG.
- FIG. 24D depicts two linear precursor RNAs, wherein one linear RNA precursor comprises an intron and two exon segments, while the other comprises a monotron intron and (non-monotron) intron segment along with two exons segments.
- Strand 1 may have a transesterification reaction first.
- Strand 2 may have a transesterification reaction first.
- FIG. 25 depicts a size exclusion-high-performance liquid chromatography (SEC- HPLC) analysis of a post-IVT reaction solution of the DNA template comprising an intron developed from an Azoarcus position 11 permutation site in FIG. 8B. Milli-absorbance units (mAU) were measured over the course of the retention time.
- the IVT reaction was performed on a DNA template comprising Azoarcus intron and exon segments at low magnesium levels of 12.75 mM and treated either with or without exonuclease digestion.
- the arrows in the figure indicate the circular RNA and linear RNA collected post-IVT of the DNA template.
- FIG. 26 provides percent circular RNAs produced in RNA constructs developed with either Anabaena position 230 permutation site (i.e., "L9a5" as labeled in FIG. 26) in FIG. 8A or Azoarcus position 11 permutation site in FIG. 8B.
- the constructs of the figures were designed to include accessory sequences (e.g., internal or external spacers and/or homology arms).
- Percent circular RNA produced was measured post SEC-HPLC analysis. Circular RNAs were produced from IVT reactions of DNA templates at either low magnesium levels (i.e., 12.75 mM) or standard reaction levels (i.e., 34 mM).
- FIG. 27 depicts a permutation heat map of regions of an exemplary group I intron Tetrahyema. Each location in FIG. 27 corresponds with a permuted intron sequence from SEQ ID NO: 25573. Each number in each figure indicates a tested permutation site.
- FIG. 28 depicts a permutation heat map of regions of an exemplary group I intron T4 td. Each location in FIG. 28 corresponds with a permuted intron sequence from SEQ ID NO: 25574. Each number in each figure indicates a tested permutation site.
- FIG. 29 depicts a permutation heat map of regions of an exemplary group I intron Staphylococcus phage Twort. Each location in FIG. 29 corresponds with a permuted intron sequence from SEQ ID NO: 3006. Each number in each figure indicates a tested permutation site.
- FIG. 30 depicts a permutation heat map of regions of an exemplary group I intron Coxiella Burnetii . Each location in FIG. 30 corresponds with a permuted intron sequence from SEQ ID NO: 2997. Each number in each figure indicates a tested permutation site.
- FIG. 31A and FIG. 31B illustrate percent m6A modification incorporation (i.e., "% M6A Peak Area") into IVT reactions of DNA templates comprising introns developed with either Anabaena position 230 permutation site in FIG. 8A (FIG. 31A) or Azoarcus position 12 permutation site in FIG. 8B (FIG. 31B).
- Amount of m6A modified nucleotide introduced into the IVT reaction was either at 0%, 1% 5%, 10%, or 50% (i.e., "% Fed M6A").
- FIG. 32 depicts estimated percent circular RNAs produced in RNA constructs developed with either m6A or mlV modifications post IVT reaction of DNA template comprising a caprine kobuvirus internal ribosome entry site (IRES) and Anabaena intron permuted at position of 230 in FIG. 8 A. Percent circular RNA produced was measured post SEC-HPLC analysis. Amount of m6A or mlV modified nucleotide introduced into the IVT reaction was either at 0%, 1% 5%, 10%, or 50% (i.e., "% Fed Base Modification").
- FIG. 33 depicts estimated percent circular RNAs produced in RNA constructs developed with either m6A or mlV modifications post IVT reaction of DNA template comprising a caprine kobuvirus internal ribosome entry site (IRES) and Anabaena intron permuted at position of 230 in FIG. 8A and subsequently purified using oligo-dT purification methods. Percent circular RNA produced was measured post SEC-HPLC analysis. Amount of m6A or mlV modified nucleotide introduced into the IVT reaction was either at 0%, 1% 5%, 10%, or 50% (i.e., "% Fed Base Modification").
- IVT caprine kobuvirus internal ribosome entry site
- FIG. 34A provides luminescence of circular RNAs encoding firefly luciferase in relative light units (“RLU") (i.e., "Flue Activity”) and percent circularization (i.e., "circ, %") post IVT reaction of three DNA templates (e.g., "control”, “Anabaena” and “Azoarcus” as depicted in FIG. 34A) with transfection of either 0%, 1%, 5% or 10% m6A or mlV base modifications.
- FIG. 34B provides IFNp secretion levels of constructs provided in FIG. 34A. In FIGs.
- FIG. 35A-35C depicts IFN[3 secretion (FIG. 35 A), IFN[3 (FIG. 35B), IL-6 fold induction (FIG. 35C) of circular RNAs formed from DNA templates undergoing IVT reaction with either 0% or 5% m6A or mlV base modifications.
- DNA templates comprised: (1) a T7 polymerase promoter, (2) 5 ' and 3 ' permuted intron segments formed from the permuted site(s), (4) 5 ' and 3 ' exon segments, (5) Caprine kobuvirus internal ribosome entry site (IRES), (6) Flue coding sequence and an Xbal restriction site.
- IRS Caprine kobuvirus internal ribosome entry site
- FIG. 36A depicts loss of circularization post IVT reaction of DNA templates with either 0%, 1%, 5%, 10%, and 50% m6A base modifications as compared to 0% m6A base modifications.
- DNA templates comprised introns of either Anabaena positions 8 or 230 permutation sites in FIG. 8A or Azoarcus positions 12 or 119 permutation sites in FIG. 8B.
- FIG. 36B depicts estimated percent circularization of constructs present in FIG. 36A as determined using SEC-HPLC.
- FIG. 37 depicts a gel of a post IVT reaction of various RNA samples.
- RNA samples comprised RNA comprising a CBV3 internal ribosome entry site (IRES), firefly luciferase coding region, and Anabaena permuted intron-exon (PIE) segments.
- IVT CBV3 internal ribosome entry site
- PIE Anabaena permuted intron-exon
- FIG. 38A depicts SEC-HPLC analysis of a two linear precursor ("Strand 1" and "Strand 2").
- Strand 1 comprises a monotron sequence and a (non-monotron) intron segment.
- Strand 2 comprises two (non-monotron) intron segments.
- “dT+Exonuclease” are the circular RNA product formed from Strand 1 and Strand 2 after oligo-dT and exonuclease purification.
- FIG. 38B depicts the circular RNA product present in FIG. 38A after purification using oligo- dT only purification (“dT Purified”), and after purification using both oligo-dT and an exonuclease solution.
- FIG. 38C depicts an exonuclease control post IVT reaction of a DNA template capable of self-circularization and comprising Anabaena 5 ' and 3 ' intron segments.
- FIG. 39A and FIG. 39B show circular RNA generated using in vitro transcription followed by purification reducing reactogenicity in a Balb/c mouse model.
- FIG. 40A and FIG. 40B show circular RNA generated using in vitro transcription followed by purification reducing reactogenicity in a BLaERl model.
- the present disclosure provides, among other things, precursor RNAs for producing circular RNAs and the produced circular RNAs.
- such produced circular RNAs have improved properties, such as improved circularization efficiency.
- the precursor RNAs comprise Group I or Group II exon and/or intron segments.
- the precursor RNAs and/or circular RNAs comprise one or more modified nucleotides or nucleosides.
- compositions e.g., template DNAs or lipid nanoparticles.
- methods for the selection, design, preparation, manufacture, formulation, and/or use of RNA preparations, such as precursor RNAs or circular RNAs are also provided herein.
- linear nucleic acid molecules are said to have a “5’-terminus” (or “5’ end”) and a “3’-terminus” (or “3’ end”) because nucleic acid phosphodiester linkages occur at the 5’ carbon and 3’ carbon of the sugar moi eties of the substituent mononucleotides.
- the end nucleotide of a polynucleotide at which a new linkage would be to a 5’ carbon is its 5’ terminal nucleotide.
- the end nucleotide of a polynucleotide at which a new linkage would be to a 3’ carbon is its 3’ terminal nucleotide.
- a “terminal nucleotide,” as used herein, is the nucleotide at the end position of the 3’ - or 5 ’-terminus.
- the term “3’ intron segment” refers to a sequence with at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% similarity to the 3’-proximal end of a natural intron (e.g., a group I or group II intron).
- the 3’ intron segment includes the 5’ nucleotide of the splice site dinucleotide.
- 3’ exon segment refers to a sequence with at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% similarity to the 5 ’-proximal end of an exon adjacent to a “3’ intron segment” as described herein.
- the 3’ exon segment includes the 3’ nucleotide of the splice site dinucleotide.
- the term “5’ intron segment” refers to a sequence with at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or higher 100% similarity to the 5’-proximal end of a natural intron (e.g., a group I or group II intron).
- the 5’ intron segment includes the 3’ nucleotide of the splice site dinucleotide.
- “5’ exon segment” refers to a sequence with at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or higher 100% similarity to the 3’-proximal end of an exon adjacent to a “5’ intron segment” as described herein.
- the 5’ exon segment includes the 5’ nucleotide of the splice site dinucleotide.
- the 3 ’ intron segment and the 3 ’ exon segment together form a first portion of an autocatalytic or self-splicing intron-exon sequence.
- the 5’ intron segment and the 5’ exon segment together form the remainder (i.e., second portion) of the autocatalytic or self-splicing intron-exon sequence.
- a linear nucleic acid molecule e.g., RNA, comprising the 3’ intron segment and the 3’ exon segment at the 5’ end of the linear nucleic acid molecule and further the 5’ intron segment and the 5’ exon segment at the 3’ end the linear nucleic acid molecule, is capable of autocatalytically self-splicing and thereby capable of forming a circular nucleic acid molecule, e.g., circular RNA.
- the 3’ intron segment and the 5’ intron segments are excised from the circular nucleic acid molecule, e.g., circular RNA, and the 3’ exon segment and the 5’ exon segment are retained in the circular nucleic acid molecule, e.g., circular RNA.
- Each retained post-splicing exon segment may be referred to as a self-splicing or self-spliced exon segment, e.g., a 3’ self-splicing or self-spliced exon segment and a 5’ self-splicing or selfspliced exon segment.
- the intron segment is a “Group I intron” and the corresponding exon segment may be referred to as a “Group I exon” or “Group 1 self-splicing exon” or “Group I self-spliced exon segment” or the like.
- the intron segment is a “Group II intron” and the corresponding exon segment may be referred to as a “Group II exon” or “Group II self-splicing exon” or “Group II self-spliced exon segment” or the like.
- the retained, post-splicing, self-splicing 3’ or 5’ exon segment is a non-coding sequence in the circular nucleic acid molecule, e.g., circular RNA.
- the circular nucleic acid molecule, e.g., circular RNA further comprises a desired coding sequence
- the retained, post-splicing, self-splicing 3’ or 5’ exon segment is (e.g., designed) to be a portion of the desired expression sequence, contiguous with the desired coding sequence, and/or in frame with the desired coding sequence.
- a circular nucleic acid molecule e.g., derived from a linear nucleic acid precursor, and comprising a coding sequence
- the 5’ to 3’ orientation of the coding sequence may be used to inform whether other sequences within the circular nucleic acid are 5’ and/or 3’, e.g., for example, 5’ is nearer to the 5’ of the coding sequence, and the 3’ end is downstream of the coding sequence.
- reference to a “5”’ or “3”’ portion of the molecule may correspond to the orientation of the sequence within the linear nucleic acid precursor.
- splice site refers to the junction consisting of a dinucleotide between an exon and an intron in an unspliced RNA.
- splice site refers to a dinucleotide that is partially or fully included in a group I or group II intron and/or exon and between which a phosphodiester bond is cleaved during RNA circularization.
- a “splice site dinucleotide” refers two nucleotides: a 5’ splice site nucleotide and the 3’ splice site nucleotide.
- a “5’ splice site” refers to the natural 5’ dinucleotide of the intron and/or exon e.g., group I or group II intron and/or exon, while a “3’ splice site” refers to the natural 3’ dinucleotide of the intron and/or exon. Exemplary splice site dinucleotides are shown in the table below.
- permutation site refers to a site in an intron and/or exon (e.g., a group I or II intron and/or exon) where a cut is made prior to permutation of the intron/or exon.
- a cut generates an intron sequence comprising a 3’ intron segment and a sequence comprising a 5’ intron segment (e.g., group I or group II intron fragments) that are permuted to be on either side of a stretch of precursor RNA to be circularized.
- permuted intron segments are thereby called “3’ permuted intron segments” or “3’ permuted elements” and “5’ permuted intron segments” or “5’ permuted elements” in the context of said precursor RNA.
- permuted intron segment and “permuted intron element” are used interchangeably.
- the permutation site consists of a dinucleotide.
- the terms “about,” or “approximately” are understood as within a range of normal tolerance in the art, for example within 2 standard deviations of the mean. “About” can be understood as within 10%, 9%, 8%, 7%, 6%, 5%, 4%, 3%, 2%, 1%, 0.9%, 0.8%, 0.7%, 0.6%, 0.5%, 0.4%, 0.3%, 0.2%, 0.1%, 0.09%, 0.08%, 0.07%, 0.06%, 0.05%, 0.04%, 0.03%, 0.02%, or 0.01% of the stated value.
- a “combined accessory element” or “combined accessory sequences” comprises the accessory element and further comprises an intron and/or exon segment. In some embodiments, the accessory element increases circularization efficiency and/or translation efficiency in a circular RNA as compared to a control circular RNA without the accessory sequences.
- an “affinity sequence” or “affinity tag” is a region of a polynucleotide sequence ranging from one (1) nucleotide to hundreds or thousands of nucleotides containing a repeated set of nucleotides for the purposes of aiding purification of a polynucleotide sequence.
- an affinity sequence may comprise, but is not limited to, a polyA or poly AC sequence.
- affinity tags are used in purification methods, referred to herein as “affinity-purification,” in which selective binding of a binding agent to molecules comprising an affinity tag facilitates separation from molecules that do not comprise an affinity tag.
- an affinity-purification method is a “negative selection” purification method, in which unwanted species, such as linear RNA, are selectively bound and removed and wanted species, such as circular RNA, are eluted and separated from unwanted species.
- an “antigen” refers to any molecule that provokes an immune response or is capable of being bound by an antibody or an antigen binding molecule.
- the immune response may involve either antibody production, or the activation of specific immunologically - competent cells, or both.
- An antigen may be endogenously expressed, i.e. expressed by genomic DNA, or may be recombinantly expressed.
- An antigen may be specific to a certain tissue, such as a cancer cell, or it may be broadly expressed.
- fragments of larger molecules may act as antigens.
- antigens are tumor antigens.
- an antigen binding molecule refers to any molecule that specifically binds to a desired antigen.
- an antigen binding molecule comprises the antigen binding parts (e.g., CDRs) of an antibody or antibody-like molecule.
- An antigen binding molecule may include the antigenic complementarity determining regions (CDRs).
- antibody fragments include, but are not limited to, Fab, Fab', F(ab')2, Fv fragments, dAb, linear antibodies, scFv antibodies, and multispecific antibodies formed from antigen binding molecules.
- Peptibodies i.e., Fc fusion molecules comprising peptide binding domains
- suitable antigen binding molecules are another example of suitable antigen binding molecules.
- the antigen binding molecule binds to an antigen on a tumor cell. In some embodiments, the antigen binding molecule binds to an antigen on a cell involved in a hyperproliferative disease or to a viral or bacterial antigen. In further embodiments, the antigen binding molecule is an antibody fragment, including one or more of the complementarity determining regions (CDRs) thereof, that specifically binds to the antigen. In further embodiments, the antigen binding molecule is a single chain variable fragment (scFv). In some embodiments, the antigen binding molecule comprises or consists of avimers. [76] The term “antibody” (Ab) includes, without limitation, a glycoprotein immunoglobulin which binds specifically to an antigen.
- an antibody may comprise at least two heavy (H) chains and two light (L) chains interconnected by disulfide bonds, or an antigen-binding molecule thereof.
- Each H chain may comprise a heavy chain variable region (abbreviated herein as VH) and a heavy chain constant region.
- the heavy chain constant region can comprise three constant domains, CHI, CH2 and CH3.
- Each light chain can comprise a light chain variable region (abbreviated herein as VL) and a light chain constant region.
- the light chain constant region can comprise one constant domain, CL.
- the VH and VL regions may be further subdivided into regions of hypervariability, termed complementarity determining regions (CDRs), interspersed with regions that are more conserved, termed framework regions (FR).
- CDRs complementarity determining regions
- CDRs may be described by numbering known in the art, for example, Kabat numbering, Chothia numbering, AbM numbering, or contact numbering.
- Each VH and VL may comprise three CDRs and four FRs, arranged from amino-terminus to carboxyterminus in the following order: FR1, CDR1, FR2, CDR2, FR3, CDR3, and FR4.
- the variable regions of the heavy and light chains contain a binding domain that interacts with an antigen.
- the constant regions of the Abs may mediate the binding of the immunoglobulin to host tissues or factors, including various cells of the immune system (e.g., effector cells) and the first component of the classical complement system.
- Antibodies may include, for example, monoclonal antibodies, recombinantly produced antibodies, monospecific antibodies, multispecific antibodies (including bispecific antibodies), human antibodies, engineered antibodies, humanized antibodies, chimeric antibodies, immunoglobulins, synthetic antibodies, tetrameric antibodies comprising two heavy chain and two light chain molecules, an antibody light chain monomer, an antibody heavy chain monomer, an antibody light chain dimer, an antibody heavy chain dimer, an antibody light chain- antibody heavy chain pair, intrabodies, antibody fusions (sometimes referred to herein as “antibody conjugates”), heteroconjugate antibodies, single domain antibodies, monovalent antibodies, single chain antibodies or singlechain variable fragments (scFv), camelized antibodies, affybodies, Fab fragments, F(ab’)2 fragments, disulfide-linked variable fragments (sdFv), anti -idiotypic (anti-id) antibodies (including, e.g., anti-anti-Id antibodies), minibodies, domain antibodies, synthetic antibodies (sometimes referred to
- An immunoglobulin may derive from any of the commonly known isotypes, including but not limited to IgA, secretory IgA, IgG and IgM.
- IgG subclasses are also well known to those in the art and include but are not limited to human IgGl, IgG2, IgG3 and IgG4.
- “Isotype” refers to the Ab class or subclass (e.g., IgM or IgGl) that is encoded by the heavy chain constant region genes.
- antibody includes, by way of example, both naturally occurring and non-naturally occurring Abs; monoclonal and polyclonal Abs; chimeric and humanized Abs; human or nonhuman Abs; wholly synthetic Abs; and single chain Abs.
- a nonhuman Ab may be humanized by recombinant methods to reduce its immunogenicity in humans.
- the term “antibody” also includes an antigen-binding fragment or an antigen-binding portion of any of the aforementioned immunoglobulins, and includes a monovalent and a divalent fragment or portion, and a single chain Ab.
- variable region typically refers to a portion of an antibody, generally, a portion of a light or heavy chain, typically about the amino-terminal 110 to 120 amino acids in the mature heavy chain and about 90 to 115 amino acids in the mature light chain, which differ extensively in sequence among antibodies and are used in the binding and specificity of a particular antibody for its particular antigen.
- CDRs complementarity determining regions
- FR framework regions
- variable region is a human variable region.
- variable region comprises rodent or murine CDRs and human framework regions (FRs).
- FRs human framework regions
- the variable region is a primate (e.g., non-human primate) variable region.
- the variable region comprises rodent or murine CDRs and primate (e.g., non-human primate) framework regions (FRs).
- VL and VL domain are used interchangeably to refer to the light chain variable region of an antibody or an antigen-binding molecule thereof.
- VH and “VH domain” are used interchangeably to refer to the heavy chain variable region of an antibody or an antigen-binding molecule thereof.
- constant region and “constant domain” are interchangeable and have a meaning common in the art.
- the constant region is an antibody portion, e.g., a carboxyl terminal portion of a light and/or heavy chain which is not directly involved in binding of an antibody to antigen but which may exhibit various effector functions, such as interaction with the Fc receptor.
- the constant region of an immunoglobulin molecule generally has a more conserved amino acid sequence relative to an immunoglobulin variable domain.
- aptamer refers in general to either an oligonucleotide of a single defined sequence or a mixture of said nucleotides, wherein the mixture retains the properties of binding specifically to the target molecule e.g., eukaryotic initiation factor, 40S ribosome, polyC binding protein, polyA binding protein, polypyrimidine tract-binding protein, argonaute protein family, Heterogeneous nuclear ribonucleoprotein K and La and related RNA-binding protein).
- target molecule e.g., eukaryotic initiation factor, 40S ribosome, polyC binding protein, polyA binding protein, polypyrimidine tract-binding protein, argonaute protein family, Heterogeneous nuclear ribonucleoprotein K and La and related RNA-binding protein.
- aptamer is meant to refer to a single- or double-stranded nucleic acid which is capable of binding to a protein or other molecule.
- aptamers preferably comprise about 10 to about 100 nucleotides, preferably about 15 to about 40 nucleotides, more preferably about 20 to about 40 nucleotides, in that oligonucleotides of a length that falls within these ranges are readily prepared by conventional techniques.
- aptamers can further comprise a minimum of approximately 6 nucleotides, preferably 10, and more preferably 14 or 15 nucleotides, that are necessary to effect specific binding.
- autoimmunity is defined as persistent and progressive immune reactions to non-infectious self-antigens, as distinct from infectious non self-antigens from bacterial, viral, fungal, or parasitic organisms which invade and persist within mammals and humans.
- Autoimmune conditions include scleroderma, Grave's disease, Crohn's disease, Sjorgen's disease, multiple sclerosis, Hashimoto's disease, psoriasis, myasthenia gravis, autoimmune polyendocrinopathy syndromes, Type I diabetes mellitus (TIDM), autoimmune gastritis, autoimmune uveoretinitis, polymyositis, colitis, and thyroiditis, as well as in the generalized autoimmune diseases typified by human Lupus.
- TIDM Type I diabetes mellitus
- Autoantigen or “self-antigen” as used herein refers to an antigen or epitope which is native to the mammal and which is immunogenic in said mammal.
- autologous refers to any material derived from the same individual to which it is later to be re-introduced.
- eACTTM engineered autologous cell therapy
- Binding affinity generally refers to the strength of the sum total of non-covalent interactions between a single binding site of a molecule (e.g., an antibody) and its binding partner (e.g., an antigen). Unless indicated otherwise, as used herein, “binding affinity” refers to intrinsic binding affinity which reflects a 1 : 1 interaction between members of a binding pair (e.g., antibody and antigen).
- the affinity of a molecule X for its partner Y may generally be represented by the dissociation constant (KD or Ka). Affinity may be measured and/or expressed in a number of ways known in the art, including, but not limited to, equilibrium dissociation constant (KD), and equilibrium association constant (KA or K a ).
- the KD is calculated from the quotient of koir/kon, whereas KA is calculated from the quotient of k 0 n/k 0 ff.
- kon refers to the association rate constant of, e.g., an antibody to an antigen
- koir refers to the dissociation of, e.g., an antibody to an antigen.
- the k on and koir may be determined by techniques known to one of ordinary skill in the art, such as BIACORE® or KinExA.
- the term “specifically binds,” refers to molecules that bind to an antigen (e.g., epitope or immune complex) as such binding is understood by one skilled in the art.
- a molecule that specifically binds to an antigen may bind to other peptides or polypeptides, generally with lower affinity as determined by, e.g., immunoassays, BIACORE®, KinExA 3000 instrument (Sapidyne Instruments, Boise, ID), or other assays known in the art.
- molecules that specifically bind to an antigen bind to the antigen with a KA that is at least 2 logs, 2.5 logs, 3 logs, 4 logs or greater than the KA when the molecules bind to another antigen.
- bicistronic RNA refers to a polynucleotide that includes two expression sequences coding for two distinct proteins. These expression sequences can be separated by a nucleotide sequence encoding a cleavable peptide such as a protease cleavage site. They can also be separated by a ribosomal skipping element.
- a “cancer” refers to a broad group of various diseases characterized by the uncontrolled growth of abnormal cells in the body. Unregulated cell division and growth results in the formation of malignant tumors that invade neighboring tissues and may also metastasize to distant parts of the body through the lymphatic system or bloodstream.
- a “cancer” or “cancer tissue” may include a tumor. Examples of cancers that may be treated by the methods disclosed herein include, but are not limited to, cancers of the immune system including lymphoma, leukemia, myeloma, and other leukocyte malignancies.
- the methods disclosed herein may be used to reduce the tumor size of a tumor derived from, for example, bone cancer, pancreatic cancer, skin cancer, cancer of the head or neck, cutaneous or intraocular malignant melanoma, uterine cancer, ovarian cancer, rectal cancer, cancer of the anal region, stomach cancer, testicular cancer, uterine cancer, multiple myeloma, Hodgkin's Disease, nonHodgkin's lymphoma (NHL), primary mediastinal large B cell lymphoma (PMBC), diffuse large B cell lymphoma (DLBCL), follicular lymphoma (FL), transformed follicular lymphoma, splenic marginal zone lymphoma (SMZL), cancer of the esophagus, cancer of the small intestine, cancer of the endocrine system, cancer of the thyroid gland, cancer of the parathyroid gland, cancer of the adrenal gland, cancer of the urethra, cancer of the penis, chronic or
- the methods disclosed herein may be used to reduce the tumor size of a tumor derived from, for example, sarcomas and carcinomas, fibrosarcoma, myxosarcoma, liposarcoma, chondrosarcoma, osteogenic sarcoma, Kaposi's sarcoma, sarcoma of soft tissue, and other sarcomas, synovioma, mesothelioma, Ewing's tumor, leiomyosarcoma, rhabdomyosarcoma, colon carcinoma, pancreatic cancer, breast cancer, ovarian cancer, prostate cancer, hepatocellular carcinomna, lung cancer, colorectal cancer, squamous cell carcinoma, basal cell carcinoma, adenocarcinoma (for example adenocarcinoma of the pancreas, colon, ovary, lung, breast, stomach, prostate, cervix, or esophagus), sweat gland carcinoma,
- the particular cancer may be responsive to chemo- or radiation therapy or the cancer may be refractory.
- a refractory cancer refers to a cancer that is not amenable to surgical intervention and the cancer is either initially unresponsive to chemo- or radiation therapy or the cancer becomes unresponsive over time.
- circRNA As used herein, the terms “circRNA,” “circular polyribonucleotide,” “circular RNA,” “circularized RNA,” “circular RNA polynucleotide” and “oRNA” are used interchangeably and refer to a single-stranded polyribonucleotide wherein the 3’ and 5’ ends that are normally present in a linear RNA polynucleotide have been joined together, e.g., by covalent bonds. As used herein, such terms also include preparations comprising circRNAs.
- circularization efficiency refers to a measurement of the rate of formation of amount of resultant circular polyribonucleotide as compared to its linear starting material.
- the expression sequences in the polynucleotide construct may be separated by a “cleavage site” sequence which enables polypeptides encoded by the expression sequences, once translated, to be expressed separately by the cell, e.g., eukaryotic cell.
- a “self-cleaving peptide” refers to a peptide which is translated without a peptide bond between two adjacent amino acids, or functions such that when the polypeptide comprising the proteins and the selfcleaving peptide is produced, it is immediately cleaved or separated into distinct and discrete first and second polypeptides without the need for any external cleavage activity.
- co-administering refers to administering a therapeutic agent provided herein in conjunction with one or more additional therapeutic agents sufficiently close in time such that the therapeutic agent provided herein can enhance the effect of the one or more additional therapeutic agents, or vice versa.
- coding element As used herein, “coding element,” “coding sequence,” “coding nucleic acid,” or “coding region” is region located within the expression sequence and encodings for one or more proteins or polypeptides e.g., therapeutic protein).
- a “noncoding element,” “noncoding sequence,” “non-coding nucleic acid,” or “noncoding nucleic acid” is a region located within the expression sequence. This sequence by itself does not encode for a protein or polypeptide, but may have other regulatory functions, including but not limited, allow the overall polynucleotide to act as a biomarker or adjuvant to a specific cell.
- a “costimulatory ligand,” as used herein, includes a molecule on an antigen presenting cell that specifically binds a cognate co-stimulatory molecule on a T cell. Binding of the costimulatory ligand provides a signal that mediates a T cell response, including, but not limited to, proliferation, activation, differentiation, and the like. A costimulatory ligand induces a signal that is in addition to the primary signal provided by a stimulatory molecule, for instance, by binding of a T cell receptor (TCR)/CD3 complex with a major histocompatibility complex (MHC) molecule loaded with peptide.
- TCR T cell receptor
- MHC major histocompatibility complex
- a co-stimulatory ligand may include, but is not limited to, 3/TR6, 4-IBB ligand, agonist or antibody that binds Toll-like receptor, B7-1 (CD80), B7-2 (CD86), CD30 ligand, CD40, CD7, CD70, CD83, herpes virus entry mediator (HVEM), human leukocyte antigen G (HLA-G), ILT4, immunoglobulin-like transcript (ILT) 3, inducible costimulatory ligand (ICOS-L), intercellular adhesion molecule (ICAM), ligand that specifically binds with B7-H3, lymphotoxin beta receptor, MHC class I chain-related protein A (MICA), MHC class I chain-related protein B (MICB), 0X40 ligand, PD-L2, or programmed death (PD) LI.
- HVEM herpes virus entry mediator
- HLA-G human leukocyte antigen G
- ILT4 immunoglobulin-like transcript
- ILT immunoglobul
- a co-stimulatory ligand includes, without limitation, an antibody that specifically binds with a co-stimulatory molecule present on a T cell, such as, but not limited to, 4-IBB, B7-H3, CD2, CD27, CD28, CD30, CD40, CD7, ICOS, ligand that specifically binds with CD83, lymphocyte function- associated antigen-1 (LFA-1), natural killer cell receptor C (NKG2C), 0X40, PD-1, or tumor necrosis factor superfamily member 14 (TNFSF14 or LIGHT).
- LFA-1 lymphocyte function- associated antigen-1
- NSG2C natural killer cell receptor C
- 0X40 PD-1
- TNFSF14 or LIGHT tumor necrosis factor superfamily member 14
- costimulatory molecule is a cognate binding partner on a T cell that specifically binds with a costimulatory ligand, thereby mediating a costimulatory response by the T cell, such as, but not limited to, proliferation.
- Costimulatory molecules include, but are not limited to, 4-1BB/CD137, B7-H3, BAFFR, BLAME (SLAMF8), BTLA, CD 33, CD 45, CD100 (SEMA4D), CD103, CD134, CD137, CD154, CD16, CD160 (BY55), CD 18, CD19, CD 19a, CD2, CD22, CD247, CD27, CD276 (B7-H3), CD28, CD29, CD3 (alpha; beta; delta; epsilon; gamma; zeta), CD30, CD37, CD4, CD4, CD40, CD49a, CD49D, CD49f, CD5, CD64, CD69, CD7, CD80, CD83 ligand, CD84, CD86, CD8alpha, CD8beta, CD9, CD96 (Tactile), CD1- la, CDl-lb, CDl-lc, CDl-ld, CDS, CEACAM1, CRT AM, DAP-10, DNAM
- an antigen binding molecule, an antibody, or an antigen binding molecule thereof “cross-competes” with a reference antibody or an antigen binding molecule thereof if the interaction between an antigen and the first binding molecule, an antibody, or an antigen binding molecule thereof blocks, limits, inhibits, or otherwise reduces the ability of the reference binding molecule, reference antibody, or an antigen binding molecule thereof to interact with the antigen.
- Cross competition may be complete, e.g., binding of the binding molecule to the antigen completely blocks the ability of the reference binding molecule to bind the antigen, or it may be partial, e.g., binding of the binding molecule to the antigen reduces the ability of the reference binding molecule to bind the antigen.
- an antigen binding molecule that cross-competes with a reference antigen binding molecule binds the same or an overlapping epitope as the reference antigen binding molecule. In other embodiments, the antigen binding molecule that cross-competes with a reference antigen binding molecule binds a different epitope as the reference antigen binding molecule.
- RIA solid phase direct or indirect radioimmunoassay
- EIA solid phase direct or indirect enzyme immunoassay
- sandwich competition assay Stahli et al., 1983, Methods in Enzymology 9:242-253
- solid phase direct biotin-avidin EIA Karlin et al., 1986, J. Immunol.
- solid phase direct labeled assay solid phase direct labeled sandwich assay (Harlow and Lane, 1988, Antibodies, A Laboratory Manual, Cold Spring Harbor Press); solid phase direct label RIA using 1-125 label (Morel et al., 1988, Molec. Immunol. 25:7-15); solid phase direct biotin-avidin EIA (Cheung, et al., 1990, Virology 176:546-552); and direct labeled RIA (Moldenhauer et al., 1990, Scand. J. Immunol. 32:77-82).
- a “cytokine,” as used herein, refers to a non-antibody protein that is released by one cell in response to contact with a specific antigen, wherein the cytokine interacts with a second cell to mediate a response in the second cell.
- a cytokine may be endogenously expressed by a cell or administered to a subject.
- Cytokines may be released by immune cells, including macrophages, B cells, T cells, neutrophils, dendritic cells, eosinophils and mast cells to propagate an immune response. Cytokines may induce various responses in the recipient cell. Cytokines may include homeostatic cytokines, chemokines, pro- inflammatory cytokines, effectors, and acute-phase proteins.
- homeostatic cytokines including interleukin (IL) 7 and IL- 15, promote immune cell survival and proliferation, and pro- inflammatory cytokines may promote an inflammatory response.
- homeostatic cytokines include, but are not limited to, IL-2, IL-4, IL-5, IL-7, IL-10, IL-12p40, IL-12p70, IL-15, and interferon (IFN) gamma.
- IFN interferon
- pro-inflammatory cytokines include, but are not limited to, IL-la, IL-lb, IL- 6, IL-13, IL-17a, IL-23, IL-27, tumor necrosis factor (TNF)-alpha, TNF-beta, fibroblast growth factor (FGF) 2, granulocyte macrophage colony-stimulating factor (GM- CSF), soluble intercellular adhesion molecule 1 (sICAM-1), soluble vascular adhesion molecule 1 (sVCAM-1), vascular endothelial growth factor (VEGF), VEGF-C, VEGF-D, and placental growth factor (PLGF).
- TNF tumor necrosis factor
- FGF fibroblast growth factor
- GM- CSF granulocyte macrophage colony-stimulating factor
- sICAM-1 soluble intercellular adhesion molecule 1
- sVCAM-1 soluble vascular adhesion molecule 1
- VEGF vascular endothelial growth factor
- effectors include, but are not limited to, granzyme A, granzyme B, soluble Fas ligand (sFasL), TGF-beta, IL-35, and perforin.
- sFasL soluble Fas ligand
- TGF-beta TGF-beta
- IL-35 TGF-beta
- perforin perforin.
- acute phase-proteins include, but are not limited to, C-reactive protein (CRP) and serum amyloid A (SAA).
- CRP C-reactive protein
- SAA serum amyloid A
- deoxyribonucleic acid and “DNA” as used herein mean a polymer composed of deoxyribonucleotides.
- ribonucleic acid and “RNA” as used herein mean a polymer composed of ribonucleotides.
- DNA template refers to a DNA sequence capable of transcribing a linear RNA polynucleotide.
- a DNA template may include a DNA vector, PCR product or plasmid.
- duplexed As used herein, the terms “duplexed,” “double-stranded,” and “hybridized” are used interchangeably and refer to double-stranded nucleic acids formed by hybridization of two single strands of nucleic acids containing complementary sequences. Sequences of the two single-stranded nucleic acids can be fully complementary or partially complementary. In some embodiments, a nucleic acid provided herein may be fully double-stranded or partially doublestranded. In most cases, genomic DNA is double-stranded.
- duplex sequences As used herein, two “duplex sequences,” “duplex forming sequences,” “duplex region,” “duplex forming regions,” “homology arms,” or “homology regions,” complement, or are complementary, fully or partially, to one another when the two regions share a sufficient level of sequence identity to one another’s reverse complement to act as substrates for a hybridization reaction.
- two duplex forming sequences are thermodynamically favored to cross-pair in a sequence specific interaction.
- polynucleotide sequences have “homology” when they are either identical or share sequence identity to a reverse complement or “complementary” sequence.
- an internal duplex forming region of a polynucleotide disclosed herein is capable of forming a duplex with another internal duplex forming region and does not form a duplex with an external duplex forming region.
- a DNA template e.g., a DNA vector
- a precursor RNA polynucleotide e.g., a linear precursor RNA polynucleotide
- a mature RNA polynucleotide e.g., a circular RNA polynucleotide
- endogenous means a substance that is native to, i.e., naturally originated from, a biological system (e.g., an organism, a tissue, or a cell).
- a “endogenous polynucleotide” is normally expressed in a cell or tissue.
- a polynucleotide is still considered endogenous if the control sequences, such as a promoter or enhancer sequences which activate transcription or translation, have been altered through recombinant techniques.
- heterologous means from any source other than naturally occurring sequences.
- an “endonuclease site” refers to a stretch of nucleotides within a polynucleotide that is capable of being recognized and cleaved by an endonuclease protein.
- an “eukaryotic initiation factor” or “elF” refers to a protein or protein complex used in assembling an initiator tRNA, 40S and 60S ribosomal subunits required for initiating eukaryotic translation.
- an “epitope” is a term in the art and refers to a localized region of an antigen to which an antibody may specifically bind.
- An epitope may be, for example, contiguous amino acids of a polypeptide (linear or contiguous epitope) or an epitope can, for example, come together from two or more non-contiguous regions of a polypeptide or polypeptides (conformational, non-linear, discontinuous, or non-contiguous epitope).
- the epitope to which an antibody binds may be determined by, e.g., NMR spectroscopy, X-ray diffraction crystallography studies, ELISA assays, hydrogen/deuterium exchange coupled with mass spectrometry (e.g., liquid chromatography electrospray mass spectrometry), array -based oligo-peptide scanning assays, and/or mutagenesis mapping (e.g., site- directed mutagenesis mapping).
- NMR spectroscopy e.g., NMR spectroscopy, X-ray diffraction crystallography studies, ELISA assays, hydrogen/deuterium exchange coupled with mass spectrometry (e.g., liquid chromatography electrospray mass spectrometry), array -based oligo-peptide scanning assays, and/or mutagenesis mapping (e.g., site- directed mutagenesis mapping).
- crystallization may be accomplished using any of the known methods in the art (e.g., Giege R et al., (1994) Acta Crystallogr D Biol Crystallogr 50(Pt 4): 339-350; McPherson A (1990) Eur J Biochem 189: 1- 23; Chayen NE (1997) Structure 5: 1269- 1274; McPherson A (1976) J Biol Chem 251 : 6300- 6303).
- Antibody antigen crystals may be studied using well known X-ray diffraction techniques and may be refined using computer software such as X- PLOR (Yale University, 1992, distributed by Molecular Simulations, Inc.; see e.g.
- expression sequence refers to a nucleic acid sequence that encodes a product, e.g., a peptide or polypeptide, regulatory nucleic acid, or non-coding nucleic acid.
- An exemplary expression sequence that codes for a peptide or polypeptide can comprise a plurality of nucleotide triads, each of which can code for an amino acid and is termed as a “codon.”
- fusion protein is a protein with at least two domains that are encoded by separate genes that have been joined to transcribe for a single peptide.
- the term “genetically engineered” or “engineered” refers to a method of modifying the genome of a cell, including, but not limited to, deleting a coding or non-coding region or a portion thereof or inserting a coding region or a portion thereof.
- the cell that is modified is a lymphocyte, e.g., a T cell, which may either be obtained from a patient or a donor.
- the cell may be modified to express an exogenous construct, such as, e.g., a chimeric antigen receptor (CAR) or a T cell receptor (TCR), which is incorporated into the cell's genome.
- CAR chimeric antigen receptor
- TCR T cell receptor
- an “immune response” refers to the action of a cell of the immune system (for example, T lymphocytes, B lymphocytes, natural killer (NK) cells, macrophages, eosinophils, mast cells, dendritic cells and neutrophils) and soluble macromolecules produced by any of these cells or the liver (including Abs, cytokines, and complement) that results in selective targeting, binding to, damage to, destruction of, and/or elimination from a vertebrate's body of invading pathogens, cells or tissues infected with pathogens, cancerous or other abnormal cells, or, in cases of autoimmunity or pathological inflammation, normal human cells or tissues.
- a cell of the immune system for example, T lymphocytes, B lymphocytes, natural killer (NK) cells, macrophages, eosinophils, mast cells, dendritic cells and neutrophils
- soluble macromolecules produced by any of these cells or the liver including Abs, cytokines, and complement
- the term “immunogenic” or “immunostimulatory” refers to a potential to induce an immune response to a substance. An immune response may be induced when an immune system of an organism or a certain type of immune cells is exposed to an immunogenic substance.
- the term “non-immunogenic” refers to a lack of or absence of an immune response above a detectable threshold to a substance. No immune response is detected when an immune system of an organism or a certain type of immune cells is exposed to a non- immunogenic substance.
- a non-immunogenic circular polyribonucleotide as provided herein does not induce an immune response above a predetermined threshold when measured by an immunogenicity assay.
- no innate immune response is detected when an immune system of an organism or a certain type of immune cells is exposed to a non-immunogenic circular polyribonucleotide as provided herein.
- no adaptive immune response is detected when an immune system of an organism or a certain type of immune cell is exposed to a non-immunogenic circular polyribonucleotide as provided herein.
- an “internal ribosome entry site” or “IRES” refers to an RNA sequence or structural element ranging in size from 10 nt to 1000 nt or more, capable of initiating translation of a polypeptide in the absence of a typical RNA cap structure.
- An exemplary IRES can be about 500 nt to about 700 nt in length.
- an “intervening region” refers to the portion of an RNA sequence that comprises one or more noncoding or one or more coding elements, or combinations thereof (e.g., translation initiation element, coding element, and/or stop codon) between splice sites.
- the intervening regions are between the 5’ combined accessory element and the 3’ combined accessory element or between the 3’ intron fragment and the 5’ intron fragment in a precursor RNA polynucleotide.
- the intervening region is between the monotron element and terminal element in other precursor RNA polynucleotides.
- isolated or “purified” generally refers to isolation of a substance (for example, in some embodiments, a compound, a polynucleotide, a protein, a polypeptide, a polynucleotide composition, or a polypeptide composition) such that the substance comprises a significant percent (e.g., greater than 1%, greater than 2%, greater than 5%, greater than 10%, greater than 20%, greater than 50%, or more, usually up to about 90%-100%) of the sample in which it resides.
- a substantially purified component comprises at least 50, 60, 70, 75, 80, 85, 90, 95, 96, 97, 98, or 99% of the sample.
- a substantially purified component comprises about, 80%-85%, or 90%-95%, 95-99%, 96-99%, 97-99%, or 95-100% of the sample.
- Techniques for purifying polynucleotides and polypeptides of interest are well-known in the art and include, for example, ion-exchange chromatography, affinity chromatography and sedimentation according to density.
- a substance is purified when it exists in a sample in an amount, relative to other components of the sample, that is more than as it is found naturally.
- a “leading untranslated sequence” is a region of polynucleotide sequences ranging from 1 nucleotide to hundreds of nucleotides located at the upmost 5' end of a polynucleotide sequence. The sequences can be defined or can be random. A leading untranslated sequence is non-coding.
- a “terminal untranslated sequence” is a region of polynucleotide sequences ranging from 1 nucleotide to hundreds of nucleotides located at the downmost 3' end of a polynucleotide sequence. The sequences can be defined or can be random. A terminal untranslated sequence is non-coding.
- terminal sequence or “terminal element” are used interchangeably to refer to an RNA sequence capable of complexing with a monotron sequence or monotron element.
- the terminal sequence comprises a splice site nucleotide from the natural group I or group II intron present in the monotron.
- the terminal sequence further comprises a natural exon or a fragment thereof and/or a synthetic sequence.
- NK cells include natural killer (NK) cells, T cells, or B cells.
- NK cells are a type of cytotoxic (cell toxic) lymphocyte that represent a major component of the innate immune system. NK cells reject tumors and cells infected by viruses. It works through the process of apoptosis or programmed cell death. They were termed “natural killers” because they do not require activation in order to kill cells.
- T cells play a major role in cell-mediated-immunity (no antibody involvement).
- T cell receptors (TCR) differentiate T cells from other lymphocyte types. The thymus, a specialized organ of the immune system, is the primary site for T cell maturation.
- T cells There are numerous types of T cells, including: helper T cells (e.g., CD4+ cells), cytotoxic T cells (also known as TC, cytotoxic T lymphocytes, CTL, T-killer cells, cytolytic T cells, CD8+ T cells or killer T cells), memory T cells ((i) stem memory cells (TSCM), like naive cells, are CD45RO-, CCR7+, CD45RA+, CD62L+ (L- selectin), CD27+, CD28+ and IL-7Ra+, but also express large amounts of CD95, IL-2R, CXCR3, and LFA-1, and show numerous functional attributes distinctive of memory cells); (ii) central memory cells (TCM) express L-selectin and CCR7, they secrete IL-2, but not IFNy or IL-4, and (iii) effector memory cells (TEM), however, do not express L-selectin or CCR7 but produce effector cytokines like IFNY an d IL-4),
- B-cells play a principal role in humoral immunity (with antibody involvement).
- B-cells make antibodies, are capable of acting as antigen- presenting cells (APCs) and turn into memory B-cells and plasma cells, both short-lived and long-lived, after activation by antigen interaction.
- APCs antigen-presenting cells
- B-cells In mammals, immature B-cells are formed in the bone marrow.
- a “miRNA site” refers to a stretch of nucleotides within a polynucleotide that is capable of forming a duplex with at least 8 nucleotides of a natural miRNA sequence.
- the terms “monotron,” “monotron sequence,” or “monotron element” are used interchangeably to refer a segment of a precursor RNA polynucleotide that is located at either the 5’ or 3’ end of the polynucleotide, i.e., either 5’ or 3’ from the intervening region.
- a monotron element refers to a sequence with 70% or higher similarity to a natural group I or group II intron including the splice site dinucleotide. In some embodiments, the monotron is capable of contributing to ribozymatic activity that allows it to enzymatically selfcleave.
- the monotron is capable of forming a phosphodiester bond with a terminal sequence, i.e., a sequence containing a splice site dinucleotide and optionally a natural exon sequence or fragment thereof.
- the terminal sequence is upstream of the monotron in a linear precursor.
- the monotron sequence is upstream of the terminal sequence in a linear precursor.
- the monotron can perform two transesterification reactions, e.g., sequentially, self-cleavage and formation of a phosphodiester bond with the terminal sequence.
- the monotron is capable of interacting with a nucleophile that is capable of cleaving at the splice site dinucleotide at or near the 5’ end of the monotron, and (b) the cleavage product of (a), i.e., the 5’ splice site nucleotide, e.g., having a 3’ hydroxyl group, engages in a transesterification reaction (cleaves) at the splice site nucleotide of the terminal sequence, yielding a circular RNA or oRNA.
- the monotron interacts with the nucleophile (e.g., a guanosine, e.g., a free guanosine that is introduced to the precursor) by forming a binding pocket with the nucleophile, and the linear precursor is capable of adopting a conformation in which the nucleophile is in proximity to and is capable of cleaving at the splice site dinucleotide at or near the 5’ end of the monotron.
- the monotron can also perform two transesterification reactions.
- the monotron is capable of interacting with a nucleophile that is capable of cleaving at the splice site nucleotide of the terminal element, and (b) the cleavage product of (a), i.e., the 5’ splice site nucleotide, e.g., having a 3’ hydroxyl group, engages in a transesterification reaction (cleaves) at the splice site dinucleotide at or near the 3’ end of the monotron, yielding a circular RNA or oRNA.
- the monotron interacts with the nucleophile (e.g., a guanosine, e.g., a free guanosine that is introduced to the precursor) by forming a binding pocket with the nucleophile, and the linear precursor is capable of adopting a conformation in which the nucleophile is in proximity to and is capable of cleaving the splice site nucleotide of the terminal element.
- the nucleophile e.g., a guanosine, e.g., a free guanosine that is introduced to the precursor
- the monotron comprises a 5’ proximal end of a natural group I or group II intron including the splice site dinucleotide and optionally a natural exon sequence or fragment thereof.
- the 5’ end of the monotron refers to nucleotides within the 5’ half of the monotron.
- the 3’ end of the monotron refers to nucleotides within the 3’ half of the monotron.
- at or near the 5’ end of the monotron refers to within the 5’ half of the monotron.
- at or near the 5’ end of the monotron refers to within the first ten 5’ positions in the monotron.
- At the 5’ end of the monotron refers to the first 5’ position(s) in the monotron. In some embodiments, at or near the 3’ end of the monotron refers to within the 3’ half of the monotron. In some embodiments, at or near the 3’ end of the monotron refers to within the last ten 3’ positions in the monotron. In some embodiments, at the 3’ end of the monotron refers to last 3’ position(s) in the monotron.
- nucleophile refers to a nucleophilic nucleotide or nucleoside capable of initiating a nucleophilic attack at a splice site and/or transesterification reaction (cleavage) at a splice site.
- nucleotide and “nucleoside” refer to a ribonucleotide, a deoxyribonucleotide, or an analog thereof.
- Nucleotides include species that comprise purines, e.g., adenine, hypoxanthine, guanine, and their derivatives and analogs, as well as pyrimidines, e.g., cytosine, uracil, thymine, and their derivatives and analogs.
- Nucleosides are similar to nucleotides, e.g., comprising purines and pyrimidines, but without the additional phosphate group.
- Modified nucleotide or nucleosides, or nucleoside or nucleotide “analogs” include nucleotides or nucleoside having modifications in the chemical structure of the base, sugar and/or phosphate, including, but not limited to, 5’-position pyrimidine modifications, 8’- position purine modifications, modifications at cytosine exocyclic amines, and substitution of 5 -bromo-uracil; and 2’ -position sugar modifications, including but not limited to, sugar- modified ribonucleotides in which the 2’ -OH is replaced by a group such as an H, OR, R, halo, SH, SR, NH2, NHR, NR2, or CN, wherein R is an alkyl moiety as defined herein.
- Nucleotide or nucleoside modifications are also meant to include nucleotides or nucleoside with bases such as inosine, queuosine, xanthine; sugars such as 2’ -methyl ribose; non-natural phosphodiester linkages such as methylphosphonate, phosphorothioate and peptide linkages.
- the “modified” nucleotide or nucleoside may be naturally occurring (e.g., pseudouridine) or synthetic.
- Nucleotide or nucleoside modifications include 5-methoxyuridine, 1- methylpseudouridine, and 6-methyladenosine. Exemplary nucleotide or nucleotide modifications are described herein.
- nucleotide or nucleotide modification differ from mutations selected from insertions, deletions, addition, or subtraction of nucleotides, for example, the mutations in a permuted Group I and Group II intron segment.
- a nucleotide or nucleoside “comprising no nucleotide or nucleoside modifications” i.e., comprising 0% modifications
- an unmodified nucleotide or nucleoside in context.
- a modified polynucleotide sequence contains at least one nucleotide or nucleoside having a modification, e.g., between 1% and 100%, 1% and 2%, 1% and 3%, 1% and 4%, 1% and 5%, 5% and 6%, 5% and 7%, 5% and 8%, 5% and 9%, 5% and 10%, 10% and 20%, 20% and 30%, 30% and 40%, 40% and 50%, 50% and 60%, 60% and 70%, 70% and 80%, 80% and 90% or 90% and 100% of the nucleotides or nucleosides are modified.
- a modification e.g., between 1% and 100%, 1% and 2%, 1% and 3%, 1% and 4%, 1% and 5%, 5% and 6%, 5% and 7%, 5% and 8%, 5% and 9%, 5% and 10%, 10% and 20%, 20% and 30%, 30% and 40%, 40% and 50%, 50% and 60%, 60% and 70%, 70% and 80%, 80% and 90% or 90% and 100%
- % modification refers to the level of incorporation within a polynucleotide, i.e., the number of modified nucleotides or nucleosides in a polynucleotide sequence divided by the total number of nucleotides or nucleosides (modified or unmodified) in the polynucleotide sequence. In some embodiments, “% modification” refers to the relative quantity of modified nucleotide or nucleoside used to generate the polynucleotide (e.g., 5% modified adenosine refers to feeding 5 mM modified adenosine and 95 mM unmodified adenosine to generate a polynucleotide sequence).
- nucleotide sequences disclosed herein can represent an RNA sequence or a corresponding DNA sequence. It is understood that deoxythymidine (dT or T) in a DNA is transcribed into a uridine (U) in an RNA. As such, “T” and “U” may be used interchangeably herein in nucleotide sequences.
- nucleic acid refers to any length, e.g., greater than about 2 bases, greater than about 10 bases, greater than about 100 bases, greater than about 500 bases, greater than 1000 bases, or up to about 10,000 or more bases, composed of nucleotides, e.g., deoxyribonucleotides or ribonucleotides, and may be produced enzymatically or synthetically (e.g., as described in U.S. Pat. No.
- nucleic acid “backbone” can be made up of a variety of linkages, including one or more of sugar-phosphodiester linkages, peptide-nucleic acid bonds (“peptide nucleic acids” or PNA; PCT No. WO 95/32305), phosphorothioate linkages, methylphosphonate linkages, or combinations thereof.
- Sugar moieties of a nucleic acid can be ribose, deoxyribose, or similar compounds with substitutions, e.g., 2’ methoxy or 2’ halide substitutions.
- Nitrogenous bases can be conventional bases (A, G, C, T, U), analogs thereof (e.g., modified uridines such as 5- methoxyuridine, pseudouridine, or N1 -methylpseudouridine, or others); inosine; derivatives of purines or pyrimidines (e.g., N4-methyl deoxyguanosine, deaza- or aza-purines, deaza- or azapyrimidines, pyrimidine bases with substituent groups at the 5 or 6 position (e.g., 5- methylcytosine), purine bases with a substituent at the 2, 6, or 8 positions, 2-amino-6- methylaminopurine, O6-methylguanine, 4-thio-pyrimidines, 4-amino-pyrim
- Nucleic acids can include one or more “abasic” residues where the backbone includes no nitrogenous base for position(s) of the polymer (US Pat. No. 5,585,481).
- a nucleic acid can comprise only conventional RNA or DNA sugars, bases and linkages, or can include both conventional components and substitutions (e.g., conventional bases with 2’ methoxy linkages, or polymers containing both conventional bases and one or more base analogs).
- Naturally occurring nucleic acids are comprised of nucleotides, including guanine, cytosine, adenine, thymine, and uracil containing nucleotides (G, C, A, T, and U respectively).
- an “oligonucleotide” is a polynucleotide comprising fewer than 1000 nucleotides, such as a polynucleotide comprising fewer than 500 nucleotides or fewer than 100 nucleotides.
- polyA means a polynucleotide or a portion of a polynucleotide consisting of nucleotides comprising adenine.
- polyT means a polynucleotide or a portion of a polynucleotide consisting of nucleotides comprising thymine.
- polyAC means a polynucleotide or a portion of a polynucleotide consisting of nucleotides comprising adenine or cytosine.
- ribosomal skipping element refers to a nucleotide sequence encoding a short peptide sequence capable of causing generation of two peptide chains from translation of one RNA molecule. While not wishing to be bound by theory, it is hypothesized that ribosomal skipping elements function by (1) terminating translation of the first peptide chain and re-initiating translation of the second peptide chain; or (2) cleavage of a peptide bond in the peptide sequence encoded by the ribosomal skipping element by an intrinsic protease activity of the encoded peptide, or by another protease in the environment (e.g., cytosol).
- sequence identity refers to the extent that sequences are identical on a nucleotide-by-nucleotide basis or an amino acid- by-amino acid basis over a window of comparison.
- a “percentage of sequence identity” may be calculated by comparing two optimally aligned sequences over the window of comparison, determining the number of positions at which the identical nucleic acid base (e.g., A, T, C, G, I) or the identical amino acid residue (e.g., Ala, Pro, Ser, Thr, Gly, Vai, Leu, He, Phe, Tyr, Trp, Lys, Arg, His, Asp, Glu, Asn, Gin, Cys and Met) occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison (i.e., the window size), and multiplying the result by 100 to yield the percentage of sequence identity.
- the identical nucleic acid base e.g., A, T, C, G, I
- the identical amino acid residue e.g., Ala, Pro, Ser, Thr, Gly, Vai, Leu, He, Phe, Tyr, Trp, Lys,
- nucleotides and polypeptides having at least about 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99% or 100% sequence identity to any of the reference sequences described herein, typically where the polypeptide variant maintains at least one biological activity of the reference polypeptide.
- a “spacer” refers to a region of a polynucleotide sequence ranging from 1 nucleotide to hundreds or thousands of nucleotides separating two other elements along a polynucleotide sequence.
- the sequences can be defined or can be random.
- a spacer is typically non-coding. In some embodiments, spacers include duplex regions.
- the term “splicing efficiency” refers to a measurement of the rate of splicing activity (e.g., none, low, or high) in a splicing or self-splicing reaction, for example, in portions of a precursor RNA polynucleotide capable of self-circularization.
- the splicing activity of, e.g., a monotron element or intron segment is affected by the structure and/or sequence of the linear RNA polynucleotide.
- structured with regard to RNA refers to an RNA sequence that is predicted by the RNAFold software or similar predictive tools to form a structure (e.g., a hairpin loop) with itself or other sequences in the same RNA molecule.
- unstructured with regard to RNA refers to an RNA sequence that is not predicted by RNA structure predictive tools to form a structure (e.g., a hairpin loop) with itself or other sequences in the same RNA molecule.
- unstructured RNA can be functionally characterized using nuclease protection assays.
- therapeutic protein refers to any protein that, when administered to a subject directly or indirectly in the form of a translated nucleic acid, has a therapeutic, diagnostic, and/or prophylactic effect and/or elicits a desired biological and/or pharmacological effect.
- translation initiation element refers to a portion of the intervening region comprising a sequence to allow translation efficiency of an encoded protein.
- core functional elements comprising one or more coding elements will further comprise one or more TIEs.
- the TIE can be part of the noncoding element.
- the TIE comprises an internal ribosome entry site (IRES).
- transcription refers to the formation or synthesis of an RNA molecule by an RNA polymerase using a DNA molecule as a template.
- the disclosure is not limited with respect to the RNA polymerase that is used for transcription.
- a T7-type RNA polymerase can be used.
- translation refers to the formation of a polypeptide molecule by a ribosome based upon an RNA template.
- translation efficiency refers to a rate or amount of protein or peptide production from a ribonucleotide transcript. In some embodiments, translation efficiency can be expressed as amount of protein or peptide produced per given amount of transcript that codes for the protein or peptide.
- transfect refers to the intracellular introduction of one or more encapsulated materials (e.g., nucleic acids and/or polynucleotides) into a cell, or preferably into a target cell.
- transfection efficiency refers to the relative amount of such encapsulated material (e.g., polynucleotides) up-taken by, introduced into and/or expressed by the target cell which is subject to transfection. In some embodiments, transfection efficiency may be estimated by the amount of a reporter polynucleotide product produced by the target cells following transfection.
- a transfer vehicle has high transfection efficiency. In some embodiments, a transfer vehicle has at least about 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, or 90% transfection efficiency.
- transfer vehicle includes any of the standard pharmaceutical carriers, diluents, excipients, and the like, which are generally intended for use in connection with the administration of biologically active agents, including nucleic acids.
- the transfer vehicles e.g., lipid nanoparticles
- the transfer vehicles are prepared to encapsulate one or more materials or therapeutic agents (e.g., circRNA).
- the process of incorporating a desired therapeutic agent (e.g., circRNA) into a transfer vehicle is referred to herein as or “loading” or “encapsulating” (Lasic, et al., FEBS Lett., 312: 255-258, 1992).
- the transfer vehicle-loaded or -encapsulated materials may be completely or partially located in the interior space of the transfer vehicle, within a bilayer membrane of the transfer vehicle, or associated with the exterior surface of the transfer vehicle.
- treatment or prevention can include treatment or prevention of one or more conditions or symptoms of the disease. Also, for purposes herein, “prevention” can encompass delaying the onset of the disease, or a symptom or condition thereof, e.g., prophylaxis of disease.
- upstream and downstream refer to relative positions of genetic code, e.g., nucleotides, sequence elements, in polynucleotide sequences.
- upstream is toward the 5’ end of the polynucleotide and downstream is toward the 3’ end.
- upstream is toward the 5’ end of the coding strand for the gene in question and downstream is toward the 3’ end.
- a “vaccine” refers to a composition for generating immunity for the prophylaxis and/or treatment of diseases. Accordingly, vaccines are medicaments which comprise antigens and are intended to be used in humans or animals for generating specific defense and protective substances upon administration to the human or animal.
- biodegradable lipid or “degradable lipid” refers to any of a number of lipid species that are broken down in a host environment on the order of minutes, hours, or days ideally making them less toxic and unlikely to accumulate in a host over time. Common modifications to lipids include ester bonds, and disulfide bonds among others to increase the biodegradability of a lipid.
- biodegradable PEG lipid or “degradable PEG lipid” refers to any of a number of lipid species where the PEG molecules are cleaved from the lipid in a host environment on the order of minutes, hours, or days ideally making them less immunogenic. Common modifications to PEG lipids include ester bonds, and disulfide bonds among others to increase the biodegradability of a lipid.
- cationic lipid or “ionizable lipid” refers to any of a number of lipid species that carry a net positive charge at a selected pH, such as physiological pH 4 and a neutral charge at other pHs such as physiological pH 7.
- PEG means any polyethylene glycol or other polyalkylene ether polymer.
- a “PEG-OH lipid” (also referred to herein as “hydroxy-PEGylated lipid”) is a PEGylated lipid having one or more hydroxyl (-OH) groups on the lipid.
- a “phospholipid” is a lipid that includes a phosphate moiety and one or more carbon chains, such as unsaturated fatty acid chains.
- structural lipid refers to sterols and also to lipids containing sterol moieties.
- sterols are a subgroup of steroids consisting of steroid alcohols.
- head-group and tail-group when used herein to describe the compounds (e.g., lipids) of the present disclosure, and in particular functional groups that are comprised in such compounds, are used for ease of reference to describe the orientation of such compounds or of one or more functional groups relative to other functional groups.
- a hydrophilic head-group e.g., guanidinium
- a cleavable functional group e.g., a disulfide group
- a hydrophobic tail-group e.g., cholesterol
- the compounds disclosed herein comprise, for example, at least one hydrophilic head-group and at least one hydrophobic tail-group, each bound to at least one cleavable group, thereby rendering such compounds amphiphilic.
- the term “amphiphilic” means the ability to dissolve in both polar (e.g., water) and non-polar (e.g., lipid) environments.
- the compounds (e.g., lipids) disclosed herein comprise at least one lipophilic tail-group (e.g., cholesterol or a C6-20 alkyl) and at least one hydrophilic head-group (e.g., imidazole), each bound to a cleavable group (e.g., disulfide).
- hydrophilic is used to indicate in qualitative terms that a functional group is water-preferring, and typically such groups are water-soluble.
- compounds e.g., ionizable lipids
- a cleavable group e.g., a disulfide (S — S) group
- hydrophilic groups e.g., a hydrophilic head-group
- such hydrophilic groups comprise or are selected from the group consisting of imidazole, guanidinium, amino, imine, enamine, an optionally-substituted alkyl amino (e.g., an alkyl amino such as dimethylamino) and pyridyl.
- hydrophobic is used to indicate in qualitative terms that a functional group is water-avoiding, and typically such groups are not water soluble.
- at least one of the functional groups of moieties that comprise the compounds disclosed herein is hydrophobic in nature (e.g., a hydrophobic tail-group comprising a naturally occurring lipid such as cholesterol).
- ionizable lipids that comprise a cleavable functional group (e.g., a disulfide (S — S) group) bound to one or more hydrophobic groups, wherein such hydrophobic groups may comprise, or may be selected from, one or more naturally occurring lipids such as cholesterol, an optionally substituted, variably saturated or unsaturated C6-C20 alkyl, and/or an optionally substituted, variably saturated or unsaturated C6-C20 acyl.
- a cleavable functional group e.g., a disulfide (S — S) group
- hydrophobic groups may comprise, or may be selected from, one or more naturally occurring lipids such as cholesterol, an optionally substituted, variably saturated or unsaturated C6-C20 alkyl, and/or an optionally substituted, variably saturated or unsaturated C6-C20 acyl.
- liposome generally refers to a vesicle composed of lipids (e.g., amphiphilic lipids) arranged in one or more spherical bilayer or bilayers. Such liposomes may be unilamellar or multilamellar vesicles which have a membrane formed from a lipophilic material and an aqueous interior that contains the encapsulated circRNA to be delivered to one or more target cells, tissues and organs.
- lipids e.g., amphiphilic lipids
- Such liposomes may be unilamellar or multilamellar vesicles which have a membrane formed from a lipophilic material and an aqueous interior that contains the encapsulated circRNA to be delivered to one or more target cells, tissues and organs.
- lipid nanoparticle refers to a transfer vehicle comprising one or more cationic or ionizable lipids, stabilizing lipids, structural lipids, and helper lipids.
- compositions described herein comprise one or more liposomes or lipid nanoparticles.
- suitable lipids e.g., ionizable lipids
- suitable lipids include one or more of the compounds disclosed herein (e.g., HGT4001, HGT4002, HGT4003, HGT4004 and/or HGT4005).
- Such liposomes and lipid nanoparticles may also comprise additional ionizable lipids such as C12-200, dLin-KC2-DMA, and/or HGT5001, helper lipids, structural lipids, PEG-modified lipids, MC3, DLinDMA, DLinkC2DMA, cKK-E12, ICE, HGT5000, DODAC, DDAB, DMRIE, DOSPA, DOGS, DODAP, DODMA, DMDMA, DODAC, DLenDMA, DMRIE, CLinDMA, CpLinDMA, DMOBA, DOcarbDAP, DLinDAP, DLincarbDAP, DLinCDAP, kLin-K-DMA, dLin-K-XTC2-DMA, HGT4003, and combinations thereof.
- additional ionizable lipids such as C12-200, dLin-KC2-DMA, and/or HGT5001, helper lipids,
- a lipid e.g., an ionizable lipid, disclosed herein comprises one or more cleavable groups.
- cleave and “cleavable” are used in this regard to mean that one or more chemical bonds (e.g., one or more of covalent bonds, hydrogen-bonds, van der Waals' forces and/or ionic interactions) between atoms in or adjacent to the subject functional group are broken (e.g., hydrolyzed) or are capable of being broken upon exposure to selected conditions (e.g., upon exposure to enzymatic conditions).
- the cleavable group is a disulfide functional group, and in particular embodiments is a disulfide group that is capable of being cleaved upon exposure to selected biological conditions (e.g., intracellular conditions).
- the cleavable group is an ester functional group that is capable of being cleaved upon exposure to selected biological conditions.
- the disulfide groups may be cleaved enzymatically or by a hydrolysis, oxidation or reduction reaction. Upon cleavage of such disulfide functional group, the one or more functional moieties or groups (e.g., one or more of a head-group and/or a tail-group) that are bound thereto may be liberated.
- Exemplary cleavable groups may include, but are not limited to, disulfide groups, ester groups, ether groups, and any derivatives thereof (e.g., alkyl and aryl esters). In certain embodiments, the cleavable group is not an ester group or an ether group. In some embodiments, a cleavable group is bound (e.g., bound by one or more of hydrogen-bonds, van der Waals’ forces, ionic interactions and covalent bonds) to one or more functional moieties or groups (e.g., at least one head-group and at least one tail-group).
- a cleavable group is bound (e.g., bound by one or more of hydrogen-bonds, van der Waals’ forces, ionic interactions and covalent bonds) to one or more functional moieties or groups (e.g., at least one head-group and at least one tail-group).
- At least one of the functional moieties or groups is hydrophilic (e.g., a hydrophilic head-group comprising one or more of imidazole, guanidinium, amino, imine, enamine, optionally-substituted alkyl amino and pyridyl).
- hydrophilic e.g., a hydrophilic head-group comprising one or more of imidazole, guanidinium, amino, imine, enamine, optionally-substituted alkyl amino and pyridyl.
- the disclosure may include compounds and pharmaceutically acceptable salts thereof, pharmaceutical compositions containing such compounds and methods of using such compounds and compositions, and the following terms, if present, have the following meanings unless otherwise indicated. It should also be understood that when described herein any of the moieties defined forth below may be substituted with a variety of substituents, and that the respective definitions are intended to include such substituted moieties within their scope as set out below. Unless otherwise stated, the term “substituted” is to be defined as set out below. It should be further understood that the terms “groups” and “radicals” can be considered interchangeable when used herein.
- H may be in any isotopic form, including 'H, 2 H (D or deuterium), and 3 H (T or tritium);
- C may be in any isotopic form, including 12 C, 13 C, and 14 C;
- O may be in any isotopic form, including 16 O and 18 O;
- F may be in any isotopic form, including 18 F and 19 F; and the like.
- Ci-6 alkyl is intended to encompass, Ci, C2, C3, C4, Cs, c 6 , C1-6, Ci-5, Ci-4, Ci-3, Ci-2, C2-6, C2-5, C2-1, C2-3, C3-6, C3-5, C3-4, C4 6, C4 -5, and C5-6 alkyl.
- alkyl refers to both straight and branched chain C1-40 hydrocarbons (e.g., C6-20 hydrocarbons), and include both saturated and unsaturated hydrocarbons.
- the alkyl may comprise one or more cyclic alkyls and/or one or more heteroatoms such as oxygen, nitrogen, or sulfur and may optionally be substituted with substituents (e.g., one or more of alkyl, halo, alkoxyl, hydroxy, amino, aryl, ether, ester or amide).
- a contemplated alkyl includes (9Z,12Z)-octadeca-9,12- dien.
- C6-20 refers to an alkyl (e.g., straight or branched chain and inclusive of alkenes and alkyls) having the recited range carbon atoms.
- an alkyl group has 1 to 10 carbon atoms (“C1-10 alkyl”).
- an alkyl group has 1 to 9 carbon atoms (“C1-9 alkyl”).
- an alkyl group has 1 to 8 carbon atoms (“C1-8 alkyl”).
- an alkyl group has 1 to 7 carbon atoms (“C1-7 alkyl”).
- an alkyl group has 1 to 6 carbon atoms (“Ci-6 alkyl”).
- an alkyl group has 1 to 5 carbon atoms (“C1-5 alkyl”). In some embodiments, an alkyl group has 1 to 4 carbon atoms (“Ci-4 alkyl”). In some embodiments, an alkyl group has 1 to 3 carbon atoms (“C1-3 alkyl”). In some embodiments, an alkyl group has 1 to 2 carbon atoms (“C1-2 alkyl”). In some embodiments, an alkyl group has 1 carbon atom (“Ci alkyl”). Examples of C1-6 alkyl groups include methyl, ethyl, propyl, isopropyl, butyl, isobutyl, pentyl, hexyl, and the like.
- alkenyl refers to a radical of a straight-chain or branched hydrocarbon group having from 2 to 20 carbon atoms, one or more carbon-carbon double bonds (e.g., 1, 2, 3, or 4 carbon-carbon double bonds), and optionally one or more carboncarbon triple bonds (e.g., 1, 2, 3, or 4 carbon-carbon triple bonds) (“C2-20 alkenyl”). In certain embodiments, alkenyl does not contain any triple bonds. In some embodiments, an alkenyl group has 2 to 10 carbon atoms (“C2-10 alkenyl”). In some embodiments, an alkenyl group has 2 to 9 carbon atoms (“C2-9 alkenyl”).
- an alkenyl group has 2 to 8 carbon atoms (“C2-8 alkenyl”). In some embodiments, an alkenyl group has 2 to 7 carbon atoms (“C2- 7 alkenyl”). In some embodiments, an alkenyl group has 2 to 6 carbon atoms (“C2-6 alkenyl”). In some embodiments, an alkenyl group has 2 to 5 carbon atoms (“C2-5 alkenyl”). In some embodiments, an alkenyl group has 2 to 4 carbon atoms (“C2-4 alkenyl”). In some embodiments, an alkenyl group has 2 to 3 carbon atoms (“C2-3 alkenyl”). In some embodiments, an alkenyl group has 2 carbon atoms (“C2 alkenyl”).
- the one or more carboncarbon double bonds can be internal (such as in 2-butenyl) or terminal (such as in 1-butenyl).
- C2-4 alkenyl groups include ethenyl (C2), 1-propenyl (C3), 2-propenyl (C3), 1- butenyl (C4), 2-butenyl (C4), butadienyl (C4), and the like.
- C2-6 alkenyl groups include the aforementioned C2-4 alkenyl groups as well as pentenyl (Cs), pentadienyl (Cs), hexenyl (Ce), and the like.
- Additional examples of alkenyl include heptenyl (C7), octenyl (Cs), octatrienyl (Cs), and the like.
- alkynyl refers to a radical of a straight-chain or branched hydrocarbon group having from 2 to 20 carbon atoms, one or more carbon-carbon triple bonds (e.g., 1, 2, 3, or 4 carbon-carbon triple bonds), and optionally one or more carbon-carbon double bonds (e.g., 1, 2, 3, or 4 carbon-carbon double bonds) (“C2-20 alkynyl”). In certain embodiments, alkynyl does not contain any double bonds. In some embodiments, an alkynyl group has 2 to 10 carbon atoms (“C2-10 alkynyl”). In some embodiments, an alkynyl group has 2 to 9 carbon atoms (“C2-9 alkynyl”).
- an alkynyl group has 2 to 8 carbon atoms (“C2-8 alkynyl”). In some embodiments, an alkynyl group has 2 to 7 carbon atoms (“C2- 7 alkynyl”). In some embodiments, an alkynyl group has 2 to 6 carbon atoms (“C2-6 alkynyl”). In some embodiments, an alkynyl group has 2 to 5 carbon atoms (“C2-5 alkynyl”). In some embodiments, an alkynyl group has 2 to 4 carbon atoms (“C2-4 alkynyl”). In some embodiments, an alkynyl group has 2 to 3 carbon atoms (“C2-3 alkynyl”).
- an alkynyl group has 2 carbon atoms (“C2 alkynyl”).
- the one or more carboncarbon triple bonds can be internal (such as in 2-butynyl) or terminal (such as in 1-butynyl).
- Examples of C2-4 alkynyl groups include, without limitation, ethynyl (C2), 1-propynyl (C3), 2- propynyl (C3), 1-butynyl (C4), 2-butynyl (C4), and the like.
- Examples of C2-6 alkenyl groups include the aforementioned C2-4 alkynyl groups as well as pentynyl (Cs), hexynyl (Ce), and the like. Additional examples of alkynyl include heptynyl (C7), octynyl (Cs), and the like.
- alkylene As used herein, “alkylene,” “alkenylene,” and “alkynylene,” refer to a divalent radical of an alkyl, alkenyl, and alkynyl group respectively. When a range or number of carbons is provided for a particular “alkylene,” “alkenylene,” or “alkynylene” group, it is understood that the range or number refers to the range or number of carbons in the linear carbon divalent chain. “Alkylene,” “alkenylene,” and “alkynylene” groups may be substituted or unsubstituted with one or more substituents as described herein.
- alkoxy refers to an alkyl group which is attached to another moiety via an oxygen atom (-O(alkyl)).
- Non-limiting examples include e.g., methoxy, ethoxy, propoxy, and butoxy.
- aryl refers to aromatic groups (e.g., monocyclic, bicyclic and tricyclic structures) containing six to ten carbons in the ring portion.
- the aryl groups may be optionally substituted through available carbon atoms and in certain embodiments may include one or more heteroatoms such as oxygen, nitrogen or sulfur.
- an aryl group has six ring carbon atoms (“Ce aryl”; e.g., phenyl).
- an aryl group has ten ring carbon atoms (“C10 aryl”; e.g., naphthyl such as 1-naphthyl and 2-naphthyl).
- cycloalkyl refers to a monovalent saturated cyclic, bicyclic, or bridged cyclic (e.g., adamantyl) hydrocarbon group of 3-12, 3-8, 4-8, or 4-6 carbons, referred to herein, e.g., as "C4-8 cycloalkyl,” derived from a cycloalkane.
- exemplary cycloalkyl groups include, but are not limited to, cyclohexanes, cyclopentanes, cyclobutanes and cyclopropanes.
- cyano refers to -CN.
- heteroaryl refers to a radical of a 5-10 membered monocyclic or bicyclic 4n+2 aromatic ring system (e.g., having 6 or 10 electrons shared in a cyclic array) having ring carbon atoms and 1-4 ring heteroatoms provided in the aromatic ring system, wherein each heteroatom is independently selected from nitrogen, oxygen and sulfur (“5-10 membered heteroaryl”).
- heteroaryl groups that contain one or more nitrogen atoms, the point of attachment can be a carbon or nitrogen atom, as valency permits.
- Heteroaryl bicyclic ring systems can include one or more heteroatoms in one or both rings.
- Heteroaryl includes ring systems wherein the heteroaryl ring, as defined above, is fused with one or more carbocyclyl or heterocyclyl groups wherein the point of attachment is on the heteroaryl ring, and in such instances, the number of ring members continue to designate the number of ring members in the heteroaryl ring system. “Heteroaryl” also includes ring systems wherein the heteroaryl ring, as defined above, is fused with one or more aryl groups wherein the point of attachment is either on the aryl or heteroaryl ring, and in such instances, the number of ring members designates the number of ring members in the fused (aryl/heteroaryl) ring system.
- Bicyclic heteroaryl groups wherein one ring does not contain a heteroatom e.g., indolyl, quinolinyl, carbazolyl, and the like
- the point of attachment can be on either ring, i.e., either the ring bearing a heteroatom (e.g., 2-indolyl) or the ring that does not contain a heteroatom (e.g., 5- indolyl).
- heterocyclyl refers to a radical of a 3- to 10- membered non-aromatic ring system having ring carbon atoms and 1 to 4 ring heteroatoms, wherein each heteroatom is independently selected from nitrogen, oxygen, sulfur, boron, phosphorus, and silicon (“3-10 membered heterocyclyl”).
- the point of attachment can be a carbon or nitrogen atom, as valency permits.
- a heterocyclyl group can either be monocyclic (“monocyclic heterocyclyl”) or a fused, bridged or spiro ring system such as a bicyclic system (“bicyclic heterocyclyl”), and can be saturated or can be partially unsaturated.
- Heterocyclyl bicyclic ring systems can include one or more heteroatoms in one or both rings.
- Heterocyclyl also includes ring systems wherein the heterocyclyl ring, as defined above, is fused with one or more carbocyclyl groups wherein the point of attachment is either on the carbocyclyl or heterocyclyl ring, or ring systems wherein the heterocyclyl ring, as defined above, is fused with one or more aryl or heteroaryl groups, wherein the point of attachment is on the heterocyclyl ring, and in such instances, the number of ring members continue to designate the number of ring members in the heterocyclyl ring system.
- the terms “heterocycle,” “heterocyclyl,” “heterocyclyl ring,” “heterocyclic group,” “heterocyclic moiety,” and “heterocyclic radical,” may be used interchangeably.
- halo and “halogen” as used herein refer to an atom selected from fluorine (fluoro, F), chlorine (chloro, Cl), bromine (bromo, Br), and iodine (iodo, I).
- the halo group is either fluoro or chloro.
- substituted means that at least one hydrogen present on a group (e.g., a carbon or nitrogen atom) is replaced with a permissible substituent, e.g., a substituent which upon substitution results in a stable compound, e.g., a compound which does not spontaneously undergo transformation such as by rearrangement, cyclization, elimination, or other reaction.
- a “substituted” group has a substituent at one or more substitutable positions of the group, and when more than one position in any given structure is substituted, the substituent is either the same or different at each position.
- “pharmaceutically acceptable salt” refers to those salts which are, within the scope of sound medical judgment, suitable for use in contact with the tissues of humans and lower animals without undue toxicity, irritation, allergic response and the like, and are commensurate with a reasonable benefit/risk ratio.
- Pharmaceutically acceptable salts are well known in the art. For example, Berge et al., describes pharmaceutically acceptable salts in detail in J. Pharmaceutical Sciences (1977) 66: 1-19.
- Pharmaceutically acceptable salts include those derived from suitable inorganic and organic acids and bases.
- Examples of pharmaceutically acceptable, nontoxic acid addition salts are salts of an amino group formed with inorganic acids such as hydrochloric acid, hydrobromic acid, phosphoric acid, sulfuric acid and perchloric acid or with organic acids such as acetic acid, oxalic acid, maleic acid, tartaric acid, citric acid, succinic acid or malonic acid or by using other methods used in the art such as ion exchange.
- inorganic acids such as hydrochloric acid, hydrobromic acid, phosphoric acid, sulfuric acid and perchloric acid
- organic acids such as acetic acid, oxalic acid, maleic acid, tartaric acid, citric acid, succinic acid or malonic acid or by using other methods used in the art such as ion exchange.
- salts include adipate, alginate, ascorbate, aspartate, benzenesulfonate, benzoate, bisulfate, borate, butyrate, camphorate, camphorsulfonate, citrate, cyclopentanepropionate, digluconate, dodecyl sulfate, ethanesulfonate, formate, fumarate, glucoheptonate, glycerophosphate, gluconate, hemisulfate, heptanoate, hexanoate, hydroiodide, 2-hydroxy-ethanesulfonate, lactobionate, lactate, laurate, lauryl sulfate, malate, maleate, malonate, methanesulfonate, 2-naphthalenesulfonate, nicotinate, nitrate, oleate, oxalate, palmitate, pamoate, pectinate
- Pharmaceutically acceptable salts derived from appropriate bases include alkali metal, alkaline earth metal, ammonium and N + (Ci-4alkyl)4 salts.
- Representative alkali or alkaline earth metal salts include sodium, lithium, potassium, calcium, magnesium, and the like.
- Further pharmaceutically acceptable salts include, when appropriate, nontoxic ammonium, quaternary ammonium, and amine cations formed using counterions such as halide, hydroxide, carboxylate, sulfate, phosphate, nitrate, lower alkyl sulfonate, and aryl sulfonate.
- the present disclosure is intended to encompass the compounds disclosed herein, and the pharmaceutically acceptable salts, pharmaceutically acceptable esters, tautomeric forms, polymorphs, and prodrugs of such compounds.
- the present disclosure includes a pharmaceutically acceptable addition salt, a pharmaceutically acceptable ester, a solvate (e.g., hydrate) of an addition salt, a tautomeric form, a polymorph, an enantiomer, a mixture of enantiomers, a stereoisomer or mixture of stereoisomers (pure or as a racemic or non-racemic mixture) of a compound described herein.
- Compounds described herein can comprise one or more asymmetric centers, and thus can exist in various isomeric forms, e.g., enantiomers and/or diastereomers.
- the compounds described herein can be in the form of an individual enantiomer, diastereomer or geometric isomer, or can be in the form of a mixture of stereoisomers, including racemic mixtures and mixtures enriched in one or more stereoisomer.
- Isomers can be isolated from mixtures by methods known to those skilled in the art, including chiral high pressure liquid chromatography (HPLC) and the formation and crystallization of chiral salts; or preferred isomers can be prepared by asymmetric syntheses.
- HPLC high pressure liquid chromatography
- the compounds e.g., ionizable lipids
- the transfer vehicles e.g., lipid nanoparticles
- Such methods generally comprise the step of contacting the one or more target cells with the compounds and/or pharmaceutical compositions disclosed herein such that the one or more target cells are transfected with the circular RNA encapsulated therein.
- the present disclosure is based in part on precursor RN As comprising both 5' intron and exon elements and 3' exon and intron elements or comprising only 3' exon and intron elements for producing circular RNAs with enhanced circularization efficiency.
- RNA polynucleotide capable of producing a circular RNA polynucleotide after splicing, wherein the precursor RNA polynucleotide comprises both 5' intron and exon elements and 3’ exon and intron elements (e.g., combined accessory elements). Also provided is a precursor RNA polynucleotide capable of producing a circular RNA polynucleotide after splicing, wherein the precursor RNA polynucleotide comprises only 3' exon and intron elements.
- RNA polynucleotide comprises (i) 3' permuted intron segment comprising a 5' nucleotide of a 3' splice site dinucleotide; and (ii) a 3' exon segment comprising a 3' nucleotide of a 3' splice site dinucleotide.
- Exemplary splice site dinucleotides are provided in the Table set forth herein.
- a provided precursor RNA polynucleotide comprises (i) a 5' exon segment comprising a 5' nucleotide of a 5' splice site dinucleotide; and (ii) a 5' permuted intron segment comprising a 3' nucleotide of a 5' splice site dinucleotide.
- a provided precursor RNA polynucleotide comprises a terminal element comprising (a) an excised terminal segment and a retained terminal segment or (b) a natural exon or a fragment thereof.
- RNA polynucleotide comprises (i) a 5' intron element comprising a 3' permuted intron segment comprising a 5' nucleotide of a 3' splice site dinucleotide; (ii) a 5' exon element comprising a 3' exon segment comprising a 3' nucleotide of a 3' splice site dinucleotide; (iii) a 3' exon element comprising a 5' exon segment comprising a 5' nucleotide of a 5' splice site; and (iv) a 3' intron element comprising a 5' permuted intron segment comprising a 3' nucleotide of a 5' splice site dinucleotide.
- RNA polynucleotide comprises (i) a terminal element comprising (a) an excised terminal segment and/or a retained terminal segment or (b) a natural exon or a fragment thereof; (ii) a 5' intron element comprising a 3' permuted intron segment comprising a 5' nucleotide of a 3' splice site dinucleotide; and (iii) a 5' exon element comprising a 3' exon segment comprising a 3' nucleotide of a 3' splice site dinucleotide.
- a provided precursor RNA polynucleotide comprises a 5’ combined accessory element comprising (i) a 3' permuted intron segment comprising a 5' nucleotide of a 3' splice site dinucleotide; and (ii) a 3' exon segment comprising a 3' nucleotide of a 3' splice site dinucleotide.
- element (ii) is located upstream to the intervening region.
- the 5 ' combined accessory element comprises a 3 ' exon segment comprising a Group I or Group II exon 3 ' nucleotide of a 3 ' splice site dinucleotide.
- a provided precursor RNA polynucleotide comprises a 3' combined accessory element comprising (i) a 5' exon segment comprising a 5' nucleotide of a 5' splice site dinucleotide; and (ii) a 5' permuted intron segment comprising a 3' nucleotide of a 5' splice site dinucleotide.
- element (ii) is located downstream to the intervening region.
- a 3 ' combined accessory element comprises a 5 ' exon segment comprising a Group I or Group II exon 5 ' nucleotide of a 5 ' splice site dinucleotide.
- a provided precursor RNA polynucleotide comprises a 5’ combined accessory element, an intervening region, and a 3' combined accessory element.
- the 5’ combined accessory element comprises (i) a 3' permuted intron segment comprising a 5' nucleotide of a 3' splice site dinucleotide; and (ii) a 3' exon segment comprising a 3' nucleotide of a 3' splice site dinucleotide; and (b) the 3' combined accessory element comprising (i) a 5' exon segment comprising a 5' nucleotide of a 5' splice site dinucleotide; and (ii) a 5' permuted intron segment comprising a 3' nucleotide of a 5' splice site dinucleotide.
- the 5' nucleotide of a 3' splice site dinucleotide, 3' nucleotide of a 3' splice site dinucleotide, 5' nucleotide of a 5' splice site dinucleotide and 3' nucleotide of a 5' splice site dinucleotide are optionally a combination of nucleotides or a portion of a sequence selected from SEQ ID NOS: 2990-3668.
- the 5’ combined accessory element is located 5' to the intervening region; and the intervening region is located is 5' to the 3' combined accessory element.
- a provided precursor RNA polynucleotide comprises a terminal element, an intervening region, and a monotron element.
- the monotron element is located 5’ to the intervening region, which is located 5’ to the terminal element.
- the monotron element is located 3’ to the intervening region, which is located 3’ to the terminal element.
- the terminal element comprises a splice site nucleotide and the monotron element comprises a splice site dinucleotide and a splice site nucleotide.
- the precursor RNA polynucleotide is linear.
- permuted intron-exon splicing results in circularization of the precursor RNA polynucleotide.
- a transesterification reaction can occur at the 5 ’ splice site and a second transesterification reaction can occur at the 3 ’ splice site.
- splicing of the precursor RNA polynucleotide results in the removal of the 3' intron element and the 5' intron element. Accordingly, the circular RNA polynucleotide produced after splicing of the precursor RNA polynucleotide lacks the 3' intron segment and the 5' intron segment, but retains the 3' exon segment and the 5' exon segment.
- the precursor RNA polynucleotide is capable of circularizing when incubated in the presence of one or more guanosine nucleotides or nucleoside (e.g., GTP) and a divalent cation (e.g., Mg 2+ ).
- GTP guanosine nucleotides or nucleoside
- Mg 2+ divalent cation
- the precursor RNA polynucleotide is between 300 and 10000, between 400 and 9000, between 500 and 8000, between 600 and 7000, between 700 and 6000, between 800 and 5000, between 900 and 5000, between 1000 and 5000, between 1100 and 5000, between 1200 and 5000, between 1300 and 5000, between 1400 and 5000, or between 1500 and 5000 nucleotides (nt) in length.
- the precursor RNA polynucleotide is at least 300 nt, at least 400 nt, at least 500 nt, at least 600 nt, at least 700 nt, at least 800 nt, at least 900 nt, at least 1000 nt, at least 1100 nt, at least 1200 nt, at least 1300 nt, at least 1400 nt, at least 1500 nt, at least 2000 nt, at least 2500 nt, at least 3000 nt, at least 3500 nt, at least 4000 nt, at least 4500 nt, or at least 5000 nt in length.
- the precursor RNA polynucleotide is no more than 3000 nt, no more than 3500 nt, no more than 4000 nt, no more than 4500 nt, no more than 5000 nt, no more than 6000 nt, no more than 7000 nt, no more than 8000 nt, no more than 9000 nt, or no more than 10000 nt in length.
- the precursor RNA polynucleotide is about 300 nt, about 400 nt, about 500 nt, about 600 nt, about 700 nt, about 800 nt, about 900 nt, about 1000 nt, about 1100 nt, about 1200 nt, about 1300 nt, about 1400 nt, about 1500 nt, about 2000 nt, about 2500 nt, about 3000 nt, about 3500 nt, about 4000 nt, about 4500 nt, about 5000 nt, about 6000 nt, about 7000 nt, about 8000 nt, about 9000 nt, or about 10000 nt in length.
- DNA templates that transcribe into precursor RNA polynucleotides of the disclosure. Accordingly, provided herein are DNA templates comprising sequences encoding the precursor RNAs of the disclosure.
- the DNA template or polynucleotide of the present disclosure comprises a vector, a PCR product, a plasmid, a minicircle DNA, a cosmid, an artificial chromosome, a complementary DNA (cDNA), an extrachromosomal DNA (ecDNA), a doggybone DNA (dbDNA), a close-ended DNA (ceDNA), a viral polynucleotide, or a fragment thereof.
- the polynucleotide of the present disclosure is selected from a DNA plasmid, a cosmid, a PCR product, dbDNA, close-ended DNA (ceDNA), and a viral polynucleotide.
- the polynucleotide further comprises a promoter segment or sequence.
- the DNA template is linearized. In other embodiments, the DNA template is non-linearized. In some embodiments, the DNA template is single-stranded. In some embodiments, the DNA template is double-stranded. In some embodiments, the DNA template comprises in whole or in part from a viral, bacterial or eukaryotic vector.
- RNA polynucleotide produced by circularization of a precursor RNA polynucleotide described herein.
- the circular RNA polynucleotide is produced inside a cell.
- a provided precursor RNA is transcribed using a DNA template in the cytoplasm (e.g., by a bacteriophage RNA polymerase) or nucleus (e.g., by host RNA polymerase II) and then circularized.
- the circular RNA polynucleotide is between 300 and 10000, between 400 and 9000, between 500 and 8000, between 600 and 7000, between 700 and 6000, between 800 and 5000, between 900 and 5000, between 1000 and 5000, between 1100 and 5000, between 1200 and 5000, between 1300 and 5000, between 1400 and 5000, or between 1500 and 5000 nucleotides (nt) in length.
- the circular RNA polynucleotide is at least 300 nt, at least 400 nt, at least 500 nt, at least 600 nt, at least 700 nt, at least 800 nt, at least 900 nt, at least 1000 nt, at least 1100 nt, at least 1200 nt, at least 1300 nt, at least 1400 nt, at least 1500 nt, at least 2000 nt, at least 2500 nt, at least 3000 nt, at least 3500 nt, at least 4000 nt, at least 4500 nt, or at least 5000 nt in length.
- the circular RNA polynucleotide is no more than 3000 nt, no more than 3500 nt, no more than 4000 nt, no more than 4500 nt, no more than 5000 nt, no more than 6000 nt, no more than 7000 nt, no more than 8000 nt, no more than 9000 nt, or no more than 10000 nt in length.
- the circular RNA polynucleotide is about 300 nt, about 400 nt, about 500 nt, about 600 nt, about 700 nt, about 800 nt, about 900 nt, about 1000 nt, about 1100 nt, about 1200 nt, about 1300 nt, about 1400 nt, about 1500 nt, about 2000 nt, about 2500 nt, about 3000 nt, about 3500 nt, about 4000 nt, about 4500 nt, about 5000 nt, about 6000 nt, about 7000 nt, about 8000 nt, about 9000 nt, or about 10000 nt in length.
- Circular RNA polynucleotides lack the free ends necessary for exonuclease- mediated degradation, causing them to be resistant to several mechanisms of RNA degradation and granting extended half-lives when compared to an equivalent linear RNA. Circularization may allow for the stabilization of RNA polynucleotides that generally suffer from short halflives and may improve the overall efficacy of exogenous mRNA in a variety of applications.
- the circular RNA polynucleotide has a functional half-life of at least 5 hours, 10 hours, 15 hours, 20 hours, 30 hours, 40 hours, 50 hours, 60 hours, 70 hours, or 80 hours. In some embodiments, the circular RNA polynucleotide has a functional half-life of 5-80, 10-70, 15-60, or 20-50 hours. In some embodiments, the circular RNA polynucleotide has a functional half-life greater (e.g., at least 1.5-fold greater or at least 2-fold greater) than that of an equivalent linear RNA polynucleotide comprising the same expression sequence.
- the circular RNA polynucleotide, or a pharmaceutical composition thereof has a functional half-life in a human cell greater than or equal to that of a pre-determined threshold value.
- the functional half-life is determined by a functional protein assay.
- the functional half-life is determined by an in vitro luciferase assay, wherein the activity of Gaussia luciferase (GLuc) is measured in the media of human cells (e.g., HepG2) expressing the circular RNA polynucleotide every 1, 2, 6, 12, or 24 hours over 1, 2, 3, 4, 5, 6, 7, or 14 days.
- the functional half-life is determined by an in vivo assay, wherein levels of a protein encoded by the expression sequence of the circular RNA polynucleotide are measured in patient serum or tissue samples every 1, 2, 6, 12, or 24 hours over 1, 2, 3, 4, 5, 6, 7, or 14 days.
- the pre-determined threshold value is the functional half-life of a reference linear RNA polynucleotide comprising the same expression sequence as the circular RNA polynucleotide.
- the functional half-life of a circular RNA polynucleotides provided herein in eukaryotic cells is at least 20 hours (e.g., at least 80 hours).
- the circular RNA polynucleotide provided herein has higher functional stability than an mRNA comprising the same expression sequence. In some embodiments, the circular RNA provided herein has higher functional stability than an mRNA comprising the same expression sequence, 5moU modifications, optimized UTR, cap, and/or poly A tail.
- a provided circular RNA polynucleotide may have a higher magnitude of expression, e.g., a higher magnitude of expression 24 hours after administration of RNA to cells, than an equivalent linear mRNA. In some embodiments, the circular RNA polynucleotide has a higher magnitude of expression than an mRNA comprising the same expression sequence, 5moU modifications, optimized UTR, cap, and/or polyA tail.
- a provided circular RNA polynucleotide is transfected into a cell.
- the DNA template which transcribes into the precursor RNA polynucleotide from which the circular RNA polynucleotide is produced, is transfected into a cell and subsequently transcribed in the cell. Transcription of the circular RNA from the transfected DNA template may be induced via polymerases.
- the polymerases are endogenous polymerases of the cell.
- the polymerases are added to the cell.
- the polymerases are encoded by one or more nucleic acids transfected into the cell.
- the circular RNA polynucleotide is administered to an animal (e.g., a human) such that a polypeptide (e.g., an adjuvant, an adjuvant-like molecule, or an immunomodulatory molecule) encoded by the circular RNA polynucleotide is expressed inside the animal.
- an animal e.g., a human
- a polypeptide e.g., an adjuvant, an adjuvant-like molecule, or an immunomodulatory molecule
- a provided circular RNA is less immunogenic than an equivalent mRNA when exposed to an immune system of an organism or a certain type of immune cell.
- the circular RNA is associated with modulated production of cytokines when exposed to an immune system of an organism or a certain type of immune cell.
- the circular RNA is associated with reduced production of IFN-pi, RIG-I, IL-2, IL-6, IFNy, and/or TNFa when exposed to an immune system of an organism or a certain type of immune cell as compared to mRNA comprising the same expression sequence.
- the circular RNA is associated with less IFN-pi, RIG-I, IL-2, IL-6, IFNy, and/or TNFa transcript induction when exposed to an immune system of an organism or a certain type of immune cell as compared to mRNA comprising the same expression sequence.
- the circular RNA is less immunogenic than mRNA comprising the same expression sequence.
- the circular RNA is less immunogenic than mRNA comprising the same expression sequence, 5moU modifications, an optimized UTR, a cap, and/or a polyA tail.
- Polynucleotides provided herein may comprise one or more intron elements, exon elements, and/or terminal elements.
- each intron element, exon element, and terminal element may independently comprise one or more spacers, intron segments, exon segments, duplex regions, affinity sequences, and/or untranslated elements. These sequence elements within the intron elements, exon elements, or terminal elements are arranged to optimize circularization and/or protein expression.
- an intron element (e.g., 3’ intron element or 5’ intron element) comprises a permuted intron segment.
- a 3’ permuted intron segment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to a 3’ proximal fragment of a natural intron (e.g., a group I or group II intron) including the 5’ nucleotide of the 3’ splice site dinucleotide.
- a natural intron e.g., a group I or group II intron
- a 5’ permuted intron fragment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to a 5’ proximal fragment of a natural intron (e.g., a group I or group II intron) including the 3’ nucleotide of the 5’ splice site dinucleotide.
- a natural intron e.g., a group I or group II intron
- Exemplary splice site dinucleotides are described in the Table herein.
- an intron element comprises an intron derived from a transsplicing ribozyme.
- the intron element comprises a Group I transsplicing ribozyme (e.g., a Tetrahymena trans-splicing ribozyme) segment.
- the trans-splicing ribozyme segment along with an exon segment that may cleave a target site (e.g., a sequence of interest and/or a coding element) and subsequently ligate cleaved targe site to a 3’ exon to form a circular RNA product.
- a provided polynucleotide (e.g., a DNA template or a precursor RNA polynucleotide) comprises a 5’ exon element located upstream to the intervening region.
- a provided polynucleotide comprises a 3’ intron element located downstream to the intervening region.
- a provided polynucleotide comprises a 3’ exon element located upstream to the intervening region.
- a provided polynucleotide comprises a 3’ intron element located upstream to the intervening region.
- the 3’ exon element and 5’ exon element each comprise an exon segment.
- the 5’ exon element comprises a 3’ exon segment.
- the 3’ exon element comprises a 5’ exon segment.
- the 3’ and/or 5’ exon segment is a self-spliced or self-splicing exon segment.
- the self-spliced and/or self-splicing exon segment comprises in part or in whole a naturally occurring exon sequence from a virus, bacterium or eukaryotic DNA vector.
- the self-spliced and/or self-splicing exon segment comprises in part or in whole a non-naturally occurring sequence.
- a 3’ exon segment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to the 5 ’-proximal end of an exon adjacent a 3’ intron segment as described herein, including the 3’ nucleotide of the splice site dinucleotide.
- a 5’ exon segment is a contiguous sequence at least 75% homologous (e.g., at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% homologous) to the 3 ’-proximal end of an exon adjacent a 5’ intron segment as described herein, including the 5’ nucleotide of the splice site dinucleotide.
- At least one of the exon segments is less than 15 nucleotides in length.
- the 3' exon segment and/or 5' exon segment comprises a Group I exon segment or a Group II exon segment less than 15 nucleotides in length.
- the circular RNA comprises a self-spliced exon segment that is 5 nucleotides, 6 nucleotides, 7 nucleotides, 8 nucleotides, 9 nucleotides, 10 nucleotides, 11 nucleotides, 12 nucleotides, 13 nucleotides, 14 nucleotides, or 15 nucleotides.
- the circular RNA comprises a self-spliced exon segment that is 5 nucleotides, 6 nucleotides, 7 nucleotides, 8 nucleotides, 9 nucleotides, 10 nucleotides, 11 nucleotides, 12 nucleotides, 13 nucleotides, 14 nucleotides, or 15 nucleotides from the exonic sequences of Table A (in which sequences are shown as 15-nucleotide exonic sequence, intronic sequence, 15-nucleotide exonic sequence), e.g., contiguous nucleotides from the 5’ or 3’ end of the exonic sequences of Table A.
- the circular RNA comprises a self-spliced exon segment that is 5 nucleotides, 6 nucleotides, 7 nucleotides, 8 nucleotides, 9 nucleotides, or 10 nucleotides from the exonic sequences of Table B (in which sequences are shown as 10- nucleotide exonic sequence, intronic sequence, 10-nucleotide exonic sequence), e.g., contiguous nucleotides from the 5’ or 3’ end of the exonic sequences of Table B. See also SEQ ID NOs: 2990-3668, 25573, and 25574.
- the intron segment is a Group I intron and the exon segment comprises a Group I self-splicing exon segment. In some embodiments, the intron segment is a Group II intron and the exon segment comprises a Group II self-splicing exon segment.
- the exon element comprises a sequence directed to a native Group I intron-adjacent exon segment sequence or Group II intron-adjacent exon segment sequence, or fragment thereof. In some embodiments, the exon element comprises at least one mutation of a native Group I intron-adjacent exon segment sequence or Group II intron- adjacent exon segment sequence, or fragment thereof. In some embodiments, the exon element comprises at least one deletion of a native Group I intron-adjacent exon segment sequence or Group II intron-adjacent exon segment sequence, or fragment thereof.
- the exon element comprises at least one insertion of a native Group I intron-adjacent exon segment sequence or Group II intron-adjacent exon segment sequence, or fragment thereof.
- the native Group I intron segment or Group II intron segment sequences are selected from a sequence in Table A or Table B, below. b. TERMINAL ELEMENTS
- a provided polynucleotide (e.g., a DNA template or a precursor RNA polynucleotide) comprises a terminal element.
- the terminal element is located upstream to the intervening region.
- the terminal element is non-intronic.
- the terminal element lacks one or both nucleotides of a natural splice site dinucleotide associated with a natural Group I or Group II intron sequence.
- a portion or the entire terminal element is excised after circularization of a precursor RNA polynucleotide comprising said terminal element.
- a polynucleotide comprises a terminal element, an intervening region, and a monotron. In some embodiments, the polynucleotide comprises, in the following order, a terminal element, an intervening region, and a monotron. In some embodiments, the polynucleotide comprises, in the following order, a monotron, an intervening region, and a terminal element. In some embodiments, the terminal element comprises a splice site nucleotide capable of engaging in a transesterification reaction with the monotron.
- the terminal element comprises an excised terminal segment and a retained terminal segment.
- the retained terminal segment is retained after circularization of a precursor RNA polynucleotide comprising such a terminal element.
- the exercised terminal segment is not retained after circularization of a precursor RNA polynucleotide comprising such a terminal element.
- the nucleotide sequence of the terminal element is non-natural or synthetic.
- the terminal element comprises a natural exon or a fragment thereof. In some embodiments, the terminal element is retained after circularization of a precursor RNA polynucleotide comprising said terminal element.
- the terminal element is capable of binding to a 3’ intron element (e.g., the 3’ intron element comprised in the same polynucleotide). In some embodiments, the terminal element is capable of directing or functionalizing the splicing activity of a 3’ intron element (e.g., the 3’ intron element comprised in the same polynucleotide).
- a 3’ intron element e.g., the 3’ intron element comprised in the same polynucleotide.
- a 5’ intron element comprises, in the following 5’ to 3’ order: a 5’ leading sequence, an optional 5’ external duplex, a 5’ affinity tag, a 5’ external spacer, and a 3’ permuted intron segment.
- the 5’ exon element comprises, in the following 5’ to 3’ order: a 3’ exon segment, an optional 5’ internal duplex, and a 5’ internal spacer.
- the 3’ exon element comprises, in the following 5’ to 3’ order: a 3’ internal spacer, an optional 3’ internal duplex, and a 5’ exon segment.
- the 3’ intron element comprises, in the following 5’ to 3’ order: a 5’ permuted intron segment, a 3’ external spacer, an optional 3’ external duplex, a 3’ affinity tag, and a 3’ lagging sequence.
- a terminal element comprises, in the following 5’ to 3’ order: a 5’ leading sequence, a 5’ external spacer, an excised terminal segment, a retained terminal segment, an optional 5’ internal duplex, and a 5’ internal spacer.
- the 3’ exon element comprises, in the following 5’ to 3’ order: a 3’ internal spacer, an optional 3’ internal duplex, and a 5’ exon segment.
- the 3’ intron element comprises, in the following 5’ to 3’ order: a 5’ permuted intron segment, a 3’ external spacer, an optional 3’ external duplex, and a 3’ lagging sequence.
- the terminal element sequence has a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more to an exon fragment of a sequence selected from Tables A or B.
- a 3’ intron element comprises in the following 5’ to 3’ order: a leading untranslated sequence, a 5’ affinity tag, an optional 5’ external duplex region, a 5’ external spacer, and a 3’ intron fragment.
- the 3’ exon element comprises in the following 5’ to 3’ order: a 3’ exon fragment, an optional 5’ internal duplex region, an optional 5’ internal duplex region, and a 5’ internal spacer.
- the 5’ exon element comprises in the following 5’ to 3’ order: a 3’ internal spacer, an optional 3’ internal duplex region, and a 5’ exon fragment.
- the 3’ intron element comprises in the following 5’ to 3’ order: a 5’ intron fragment, a 3’ external spacer, an optional 3’ external duplex region, a 3’ affinity tag, and a trailing untranslated sequence.
- the affinity tag is a polyA affinity tag.
- the 5' intron element is located 5' to the 5' exon element. In some embodiments, the 5' intron element is adjacent to the 5' exon element. In some embodiments, the 3' intron element is located 3' to the 3' exon element. In some embodiments, the 3' intron element is adjacent to the 3' exon element.
- the 5' exon element comprises a 5' internal duplex sequence located 3' to the 3' exon segment. In some embodiments, the 3' exon element comprises a 3' internal duplex sequence located 5' to the 5' exon segment. In some embodiments, the 5' intron element comprises a 5' external duplex sequence located 5' to the 3' permuted intron segment. In some embodiments, the 3' intron element comprises a 3' external duplex sequence located 3' to the 5' permuted intron segment. In some embodiments, the 5' intron element is adjacent to the 5' exon element. In some embodiments, the 3' intron element is located 3' to the 3' exon element. In some embodiments, the 3' intron element is adjacent to the 3' exon element.
- the 5' intron comprises a 5' affinity tag, a 5' external spacer, and the 3' permuted intron segment.
- the 5' exon comprises the 3' exon segment, a 5' internal duplex sequence, and a 5' internal spacer.
- the 5' affinity tag is adjacent to the 5' external spacer.
- the 5' affinity tag is located 5' to the 5' external spacer.
- the 5' internal duplex sequence is adjacent to the 5' internal spacer.
- the 5' internal duplex sequence is located 5' to the 5' internal spacer.
- the 3' exon comprises a 3' internal spacer, 3' internal duplex sequence, and the 5' exon segment.
- the 3' intron comprises the 5' permuted intron segment, a 3' external spacer, and a 3' affinity tag.
- the 3' affinity tag is adjacent to the 3' external spacer.
- the 3' affinity tag is located 3' to the 3' external spacer.
- the 3' internal duplex sequence is adjacent to the 3' internal spacer.
- the 3' internal duplex sequence is located 3' to the 3' internal spacer.
- the affinity tag is a polyA affinity tag.
- the 5' exon comprises a 5' internal duplex sequence located between the 3' exon segment and the intervening region.
- the 3' exon comprises a 3' internal duplex sequence positioned between the intervening region and the 5' exon segment.
- the polynucleotide comprises a 5' internal duplex sequence and a 3' internal duplex sequence.
- the 3' and 5' permuted intron segments each independently comprise a Group I intron segment, a Group II intron segment, a synthetic intron segment, or a variant thereof.
- the 3' permuted intron segment comprises a 3' Group I intron segment or a variant thereof.
- the 5' permuted intron segment comprises a 5' Group I intron segment or a variant thereof.
- the 3' permuted intron segment comprises a 3' Group II intron segment or a variant thereof.
- the 5' permuted intron segment comprises a 5' Group II intron segment or a variant thereof.
- the 3' permuted intron segment or element, 5' permuted intron segment or element, or both the 3' and 5' permuted intron segments or elements are at least 100, at least 90, at least 80, at least 70, at least 60, and/or at least 50 nucleotides in length. In some embodiments, the 3' permuted intron element, 5' permuted intron element, or both the 3' and 5' permuted intron elements are at least 50 nucleotides in length.
- the 3' permuted intron element, 5' permuted intron element, or both the 3' and 5' permuted intron elements have a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or more to a naturally occurring intron.
- the 3' permuted intron element, 5' permuted intron element, or both the 3' and 5' permuted intron elements comprise a native Group I intron segment or Group II intron segment sequence. In some embodiments, the 3' permuted intron element, 5' permuted intron element, or both the 3' and 5' permuted intron elements comprise one or more nucleotide substitutions of a native Group I intron segment or Group II intron segment sequence.
- the 3' permuted intron element, 5' permuted intron element, or both the 3' and 5' permuted intron elements comprise one or more nucleotide insertions of a native Group I intron segment or Group II intron segment sequence. In some embodiments, the 3' permuted intron element, 5' permuted intron element, or both the 3' and 5' permuted intron elements comprise one or more nucleotide deletions of a native Group I intron segment or Group II intron segment sequence.
- the 3' permuted intron element, 5' permuted intron element, or both the 3' and 5' permuted intron elements comprise a nucleotide substitution of one or both the dinucleotide of a native Group I or Group II intron splice site dinucleotide.
- the 3' Group I or Group II intron segment or the 5' Group I or Group II intron segment comprises one, two, three, four, five, six, seven, eight, nine, ten, or more mutations of a native Group I intron or Group II intron sequence.
- the mutations are selected from insertion, deletion, mutation, addition, and subtraction.
- the mutations are deletions of two or more nucleotides of the 3' Group I or Group II intron segment or the 5' Group I or Group II intron segment, or combinations thereof. In some embodiments, the mutations are two or more deletions of the 5' Group I intron segment at the 3' end or two or more deletions of the 3' Group I intron segment at the 5' end.
- the native Group I intron segment or Group II intron segment sequences are selected from a sequence in Table A or Table B, below.
- the 3' and/or 5' permuted intron element comprise a polynucleotide sequence having a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or more to a naturally occurring intron selected from a sequence set forth in Table A or Table B, below, or a fragment or segment thereof.
- the 3' and/or 5' permuted intron element comprise a polynucleotide sequence selected from a sequence set forth in SEQ ID NOS: 2990-3668. In some embodiments, the 3' and/or 5' permuted intron element comprise a polynucleotide sequence selected from a sequence set forth in SEQ ID NOS: 3188-3668.
- the 3' permuted intron segment comprises a 3' Group I or Group II intron segment derived from a gene selected from a genus and/or species selected from column 2 of Tables A or B; and/or the 5' permuted intron segment comprises a 5' Group I or Group II intron segment derived from a gene selected from a genus and/or species selected from column 2 of Tables A or B.
- the 3' Group I or Group II intron segment or the 5' Group I or Group II intron segment are derived from a gene selected from a species selected from: Cyanobacterium Anabaena sp., T4 phage, Hypocrea pallida, Bulbithecium hyalosporum, Myoarachis inversa, Geosmithia argillacea, Coxiella burnetii, Agrobacterium tumefaciens, Azoarcus, Nostoc, Cordyceps capitata, Prochlorothrix hollandica, Tilletiopsis orzyzicola, Tetrahymena therm ophila, and Staphylococcus phage Twort.
- Table B Group II introns (flanked by lOnt exons)
- the 3' or 5’ intron segments and/or 3’ or 5’ exon segments are derived from a gene selected from a species selected from: Cyanobacterium Anabaena sp., T4 phage, Coxiella burnetii, Azoarcus, Tetrahymena thermophila, and Staphylococcus phage Twort.
- the 3’ or 5’ intron segment and/or 3’ or 5’ exon segment are developed from permuting at a position along a Cyanobacterium Anabaena sp., T4 phage, Coxiella burnetii, Azoarcus, Tetrahymena thermophila, and Staphylococcus phage Twort intron and/or exon sequence.
- the 5’ or 3 monotron element are derived from a gene selected from a species selected from: Cyanobacterium Anabaena sp., T4 phage, Coxiella burnetii, Azoarcus, Tetrahymena thermophila, and Staphylococcus phage Twort.
- the 3’ or 5’ intron segment and/or 3’ or 5’ exon segment are developed from permuting at a position along a Cyanobacterium Anabaena sp., T4 phage, Coxiella burnetii, Azoarcus, Tetrahymena thermophila, and Staphylococcus phage Twort intron and/or exon sequence.
- the intron segments and/or exon segments of a provided polynucleotide are derived from a gene from the same species (e.g., a polynucleotide comprises Azoarcus 3’ and 5’ exon segments and Azoarcus 3’ and 5’ intron segments).
- the 3’ or 5’ intron segments or 3’ or 5’ exon segments of a provided polynucleotide are derived from genes of different species (e.g., a polynucleotide comprises an Anabaena intron segment and Staphylococcus phage Twort exon segment).
- the monotron element of a provided polynucleotide is derived from a gene of a different species than the 3’ or 5’ intron segments and/or 3’ or 5’ exon segments (e.g., a polynucleotide comprises a Staphylococcus phage Twort montron element and an Anabaena intron segment).
- a polynucleotide comprises a Staphylococcus phage Twort montron element and an Anabaena intron segment.
- use of genes of one species of an intron segment and/or exon segment may allow for more efficient or effective circularization or self-splicing of one or more polynucleotides as compared to another gene of a different species.
- the gene used of one species develop an intron segment may more efficiently promote the interaction between an intron segment and a nucleophile (e.g., form a more efficient or effective binding pocket that promotes the transesterification reaction of a splice site nucleotide) as compared to an intron segment developed from a gene of a different species.
- the gene of one species from which an intron segment is derived may be more efficient in forming a binding pocket for a nucleophile as compared to a different gene of the same species.
- the species of gene from which the intron segment is derived may be more efficient in forming a binding pocket for a nucleophile as compared to a species of genes comprising the same and/or homologous sequence from a different species.
- a provided polynucleotide comprises an intron segment and/or exon segment derived from permuting at a position along a Group I or Group II gene selected from Table A or Table B. Location or position of the permutation sites may enhance the ability of an intron segment to effectively splice and/or circularize in a provided polynucleotide.
- the Group I or Group II genes are permuted at a position that enhances splicing or circularization activity of an intron segment of a provided polynucleotide as compared to a different permutation site.
- the Group I or II genes are permuted at a position in an intron segment of a provided polynucleotide that enhances the provided polynucleotide’s ability to self-circularize as compared to a different permutation site. In some embodiments, the Group I or II genes are permuted at a position that enhances or promotes the splicing activity of an intron segment to another intron segment, monotron element and/or exon segment. In some embodiments, the Group I or II genes are permuted at a position that allows the intron segment to more efficiently splice or self-splice than an intron segment permuted at a different position.
- a position of a permutation site may promote the interaction between an intron segment and a nucleophile (e.g., form a more efficient or effective binding pocket that promotes the transesterification reaction of a splice site nucleotide).
- permutation sites positions are described to mean that the permutation of the natural or synthetic intron occurs at the junction between the listed amino acid and the adjacent downstream amino acid (e.g., an Anabaena position 189 permutation site corresponds herein to a permutation site between amino acids 189 and 190).
- a provided polynucleotide comprises an intron segment derived from permuting at a position along a gene selected from a species selected from: Cyanobacterium Anabaena sp., T4 phage, Coxiella burnetii, Azoarcus, Tetrahymena thermophila, and Staphylococcus phage Twort.
- a provided polynucleotide comprises an intron segment derived from permuting a Cyanobacterium Anabaena sp. gene.
- the permutation site of the Cyanobacterium Anabaena sp. gene may be downstream relative to amino acid positions 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34,
- a provided polynucleotide comprises an intron segment derived from permuting an Azoarcus gene.
- the permutation site of the Azoarcus gene may be downstream relative to amino acid positions 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16,
- a provided polynucleotide comprises an intron segment derived from permuting an Coxiella burnetii gene.
- the permutation site of the Coxiella burnetii gene may be downstream relative to amino acid positions 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33,
- a provided polynucleotide comprises an intron segment derived from permuting a Tetrahymena thermophila gene.
- the permutation site of the Tetrahymena thermophila gene may be downstream relative to amino acid positions 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20,
- a provided polynucleotide comprises an intron segment derived from permuting an T4 phage (td) gene.
- the permutation site of the T4 phage (td) gene may be downstream relative to amino acid positions 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103, 104, 105, 106, 107, 108,
- a provided polynucleotide comprises an intron segment derived from permuting a Staphylococcus phage Twort gene.
- the permutation site of the Staphylococcus phage Twort gene may be downstream relative to amino acid positions 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22,
- a method comprises:
- the mutated Group I or Group II exon and/or intron element or segment comprises a deletion, insertion or substitution of at least one nucleotide, including but not limited to a nucleotide substitution of one or both the dinucleotides of the 5' and/or 3' Group I splice site dinucleotides.
- the 5' or 3' Group I or Group II intronic sequences, or combinations thereof are sequenced.
- the method further comprises comparing the circularization efficiency of the polynucleotide with a polynucleotide comprising a native intronic sequence, or a parent polynucleotide.
- the method further comprises comparing the circularization efficiency of the polynucleotide with a polynucleotide comprising a native intronic sequence, or a parent polynucleotide.
- a provided polynucleotide (e.g., a DNA template, a precursor RNA polynucleotide, or a circular RNA polynucleotide) comprises one or more spacers.
- the DNA template, precursor linear RNA polynucleotide and circular RNA provided herein comprise a 5’ and/or a 3’ spacer.
- the polynucleotide comprises one or more spacers in the intron elements.
- the polynucleotide comprises one or more spacers in the exon elements.
- the polynucleotide comprises a spacer in the 3’ intron fragment (also referred to as “5’ external spacer”).
- the polynucleotide comprises a spacer in the 5’ intron fragment (also referred to as “3’ external spacer”).
- the polynucleotide comprises a spacer in the 3’ exon fragment (also referred to as “5’ internal spacer”). In some embodiments, the polynucleotide comprises a spacer in the 5’ exon fragment (also referred to as “3’ internal spacer”).
- the polynucleotide comprises a spacer in the 3’ intron fragment and/or a spacer in the 5’ intron fragment.
- the 5' external spacer is located 5' to the 3' permuted intron segment.
- the 5' internal spacer is located 3' to the 3' exon segment.
- the 3' external spacer is located 3' to the 5' permuted intron segment.
- the 3' external spacer is located 5' to the 5' exon segment.
- the polynucleotide comprises a 5' external spacer located between a leading untranslated sequence and the 5' or 3' intron element. In certain embodiments, the polynucleotide comprises a 3' external spacer located between the 3' or 5' intron element and a lagging untranslated sequence.
- the polynucleotide can comprise an internal spacer sequence positioned between the terminal element and the intervening region, and/or between the intervening region and the monotron element.
- the polynucleotide can comprise an external spacer positioned adjacent to the terminal element and/or an external spacer positioned adjacent to the monotron element.
- the spacers aid with circularization or protein expression due to symmetry created in the overall sequence of the precursor RNA polynucleotide.
- including a 5’ internal spacer and/or including a spacer between the 3’ group I intron fragment and the intervening region may conserve secondary structures in those regions by preventing them from interacting, thus increasing splicing efficiency.
- the polynucleotide further comprises an aptamer.
- the aptamer is synthetic.
- the first spacer e.g., between the 3’ group I or II intron fragment and intervening region
- second spacer e.g., between the two expression sequences and intervening region
- the first spacer e.g., between 3’ group I or II intron fragment and intervening region
- second spacer e.g., between the one of the intervening region and 5’ group I or II intron fragment
- the polynucleotide comprises a first (5’) and a second (3’) spacer.
- the polynucleotide comprises a 5’ external spacer and a 3’ external spacer, wherein the spacers comprise additional base pairing regions that are predicted to base pair with each other and not to the first and second duplex regions.
- the polynucleotide comprises a 5’ internal spacer and a 3’ internal spacer, wherein the spacers comprise additional base pairing regions that are predicted to base pair with each other and not to the first and second duplex regions.
- such spacer base pairing brings the permuted intron segments in close proximity to each other, which may increase splicing efficiency.
- the combination of base pairing between the first and second duplex regions, and separately, base pairing between the first and second spacers promotes the formation of a splicing bubble containing the permuted intron segments flanked by adjacent regions of base pairing.
- Typical spacers are contiguous sequences with one or more of the following qualities: (1) predicted to avoid interfering (e.g., forming duplex) with proximal structures, for example, the IRES, expression sequence, aptamer, or intron; (2) is at least 5 nt long and no longer than 100 nt; (3) is located adjacent to the permuted intron segment; and (4) contains one or more of the following: (a) an unstructured region at least 5 nt long, (b) a region of base pairing at least 5 nt long to a distal sequence, such as another spacer, and (c) a structured region at least 5 nt long limited in scope to the sequence of the spacer.
- a spacer is not predicted to form a duplex of more than 8 nucleotides in length with any sequences within 250 nucleotides in either direction. In some embodiments, the spacer is not predicted to form a duplex of more than 8 nucleotides in length with any sequences within 1000 nucleotides in either direction.
- the spacer comprises an unstructured, structured or randomly generated polynucleotide sequence. Spacers may have several regions, including an unstructured region, a base pairing region, a hairpin/structured region, and combinations thereof. In an embodiment, the spacer has a structured region with high GC content. In an embodiment, a region within a spacer base pairs with another region within the same spacer.
- a region within a spacer base pairs with a region within another spacer.
- a spacer comprises one or more hairpin structures.
- a spacer comprises one or more hairpin structures with a stem of 4 to 12 nucleotides and a loop of 2 to 10 nucleotides.
- a spacer sequence is at least 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 25, or 30 nucleotides in length. In some embodiments, a spacer sequence is no more than 100, 90, 80, 70, 60, 50, 45, 40, 35, or 30 nucleotides in length. In some embodiments, a spacer sequence is between 5 and 50, 10 and 50, 20 and 50, 20 and 40, and/or 25 and 35 nucleotides in length.
- a spacer sequence is 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49 or 50 nucleotides in length. In some embodiments, the spacer sequence is at least 5 nucleotides in length, and/or about 5 to about 60 nucleotides in length. [254] In some embodiments, a spacer sequence is a polyA sequence. In some embodiments, a spacer sequence is a polyAC sequence. In some embodiments, a spacer comprises about 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% polyAC content. In some embodiments, a spacer comprises about 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% polypyrimidine (C/T or C/U) content. e. DUPLEX
- a provided polynucleotide (e.g., a DNA template, a precursor RNA polynucleotide, or a circular RNA polynucleotide) comprises one or more duplexes.
- the polynucleotide comprises a 5’ external duplex located within the 3 ’ intron fragment. In some embodiments, the polynucleotide comprises a 3 ’ external duplex located within the 5’ intron fragment. In some embodiments, the polynucleotide comprises a 5' internal duplex sequence and a 3' internal duplex sequence. In some embodiments, the polynucleotide comprises a 5’ internal duplex located within the 3’ exon fragment. In some embodiments, the 5' internal duplex sequence is positioned between the 5' exon element and the intervening region. In some embodiments, the polynucleotide comprises a 3’ internal duplex located within the 5’ exon fragment.
- the 3' internal duplex sequence is positioned between the intervening region and the 3' exon element.
- the polynucleotide comprises a 5’ external duplex located within the 3’ intron fragment and a 3’ external duplex located within the 5’ intron fragment.
- the polynucleotide comprises a 5’ internal duplex located within the 3’ exon fragment and a 3’ internal duplex located within the 5’ exon fragment.
- the polynucleotide comprises a 5’ external duplex, 5’ internal duplex, a 3’ internal duplex region, and a 3’ external duplex.
- the polynucleotide comprises a monotron element, intervening region, and terminal element, and a 5' internal duplex sequence and a 3' internal duplex sequence.
- the terminal element is upstream of the monotron element
- the 5' internal duplex sequence is positioned between the terminal element and the intervening region
- the 3' internal duplex sequence is positioned between the intervening region and the monotron element.
- the monotron element is upstream of the terminal element
- the 5' internal duplex sequence is positioned between monotron and the intervening region
- the 3' internal duplex sequence is positioned between the intervening region and the terminal element.
- the 5’ or 3’ internal duplex is positioned adjacent to a 5’ or 3’ internal spacer.
- the polynucleotide comprises a first (5’) duplex and a second (3’) duplex (e.g., a 5’ external duplex region and a 3’ external duplex region).
- the first and second duplex regions may form perfect or imperfect duplexes.
- at least 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% of the first duplex and second duplex may be base paired with one another.
- the duplexes regions are predicted to have less than 50% (e.g., less than 45%, less than 40%, less than 35%, less than 30%, less than 25%) base pairing with unintended sequences in the RNA (e.g., non-duplex sequences).
- the 5' internal duplex sequence and 3' internal duplex sequence are at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99% or 100% complementary.
- first duplex and second duplex on the 5’ and 3’ ends of the precursor RNA strand, respectively, and adjacent or very close to the permuted intron segment bring the permuted intron segments in close proximity to each other, increasing splicing efficiency.
- a duplex is 3-100 nt in length (e.g., 3-75 nt in length, 3-50 nt in length, 20-50 nt in length, 35-50 nt in length, 5-25 nt in length, 5-20 nt in length, 9-19 nt in length).
- a duplex has a length of about 9 to about 50 nt.
- a duplex has a length of about 9 to about 19 nt.
- a duplex has a length of about 5 to about 20 nt nucleotides in length, inclusive.
- the 5' internal duplex sequence and 3' internal duplex sequence are each independently about 9 to about 50 nt, about 9 to about 19 nt, or about 5 to about 20 nt nucleotides in length, inclusive.
- a duplex has a length of about 20 to about 40 nt.
- a duplex is about 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49 or 50 nt in length.
- a duplex has a length of about 30 nt.
- the 5' and 3' internal duplex sequences are predicted to form a contiguous duplex.
- the contiguous duplex has a length of 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49 or 50 nt in length.
- the contiguous duplex has a length of no longer than 35 nucleotides.
- at least one of the exon segments is less than 15 nucleotides in length.
- the 5' internal duplex sequence and/or 3' internal duplex sequence each have a GC content of at least 10%.
- the polynucleotide does not comprise of any duplex to optimize translation or circularization. f. AFFINITY SEQUENCES
- a provided polynucleotide may comprise an affinity sequence (or affinity tag)
- a precursor RNA polynucleotide comprises at least one affinity tag.
- the affinity tag is located in the 3’ intron element.
- the affinity tag is located in the 5’ intron element.
- both (3’ and 5’) intron elements each comprise an affinity tag.
- the 5' affinity tag is located 5' to the 3' permuted intron segment.
- the 3' affinity tag is located 3' to the 5' permuted intron segment.
- the polynucleotide comprises a monotron element comprising an affinity tag and/or terminal element comprising an affinity tag.
- the terminal element comprises (a) a 5' affinity tag if the terminal element is located upstream of the monotron element, wherein the 5' affinity tag is located 5' to the terminal element; or (b) a 3' affinity tag if the monotron element is located upstream of the terminal element, wherein the 3' affinity tag is located 3' to the terminal element.
- the monotron element comprises (a) a 3' affinity tag if the terminal element is located upstream of the monotron element, wherein the 3' affinity tag is located 3' to the monotron element; or (b) a 5' affinity tag if the monotron element is located upstream of the terminal element, wherein the 5' affinity tag is located 5' to the monotron element.
- the precursor RNA polynucleotide comprises an external spacer
- the 5’ or 3’ affinity tag is positioned adjacent to the external spacer.
- an affinity tag of the 3’ intron element is the length as an affinity tag in the 5’ intron element. In some embodiments, an affinity tag of the 3’ intron element is the same sequence as an affinity tag in the 5’ intron element. In some embodiments, the affinity sequence is placed to optimize oligo-dT purification.
- the one or more affinity tags present in a precursor RNA polynucleotide are removed upon circularization. See, for example, Figures 97A and 97B from WO2022261490, which are incorporated by reference herein in entirety.
- affinity tags are added to remaining linear RNA after circularization of precursor RNA is performed.
- affinity tags are added enzymatically to linear RNA.
- the presence of one or more affinity tags in linear RNA and their absence from circular RNA can facilitate purification of circular RNA.
- such purification is performed using a negative selection or affinity-purification method.
- such purification is performed using a binding agent that preferentially or specifically binds to the affinity tag.
- an affinity sequence such as biotin
- an oligonucleotide comprising an affinity sequence is ligated to linear RNA.
- an oligonucleotide conjugated to an affinity handle is ligated to the linear RNA.
- a solution comprising the linear RNA ligated to the affinity sequence or handle and the circular RNA that does not comprise an affinity sequence or handle are contacted with a binding agent comprising a solid support conjugated to an oligonucleotide complementary to the affinity sequence or to a binding partner of the affinity handle, such that the linear RNA binds to the binding agent, and the circular RNA is eluted or separated from the solid support.
- an affinity tag comprises a polyA sequence or is a polyA affinity tag.
- the polyA sequence is at least 15, 30, or 60 nt in length.
- the affinity tag comprising a polyA sequence is present in two places in a precursor linear RNA.
- one or both polyA sequences are 15-50 nt in length.
- one or both polyA sequences are 20-25 nt in length.
- the polyA sequence(s) is removed upon circularization.
- an oligonucleotide hybridizing with the polyA sequence such as a deoxythymidine oligonucleotide (oligo(dT)) conjugated to a solid surface (e.g., a resin), can be used to separate circular RNA from its precursor RNA.
- a deoxythymidine oligonucleotide oligo(dT) conjugated to a solid surface
- a solid surface e.g., a resin
- any purification method for circular RNA described herein may comprise one or more buffer exchange steps.
- buffer exchange is performed after in vitro transcription (IVT) and before additional purification steps.
- the IVT reaction solution is buffer exchanged into a buffer comprising Tris.
- the IVT reaction solution is buffer exchanged into a buffer comprising greater than 1 mM or greater than 10 mM one or more monovalent salts, such as NaCl or KC1, and optionally comprising EDTA.
- buffer exchange is performed after purification of circular RNA is complete.
- buffer exchange is performed after IVT and after purification of circular RNA.
- the buffer exchange that is performed after purification of circular RNA comprises exchange of the circular RNA into water or storage buffer.
- the storage buffer comprises ImM sodium citrate, pH 6.5. g. LEADING SEQUENCES & LAGGING SEQUENCES
- provided polynucleotide e.g., a DNA template, a precursor linear RNA polynucleotide, or a circular RNA polynucleotide
- a leading untranslated sequence comprises a leading untranslated sequence.
- the leading untranslated sequence is located at the 5’ end in the 3’ intron fragment (also referred to as “5’ leading sequence”).
- the leading untranslated sequence comprises the last nucleotide of a transcription start site (TSS).
- TSS transcription start site
- the TSS is chosen from a viral, bacterial, or eukaryotic DNA template.
- the leading untranslated sequence comprises the last nucleotide of a TSS and 0 to 100 additional nucleotides.
- the TSS is a spacer.
- the leading untranslated sequence contains a guanosine at the 5’ end.
- provided polynucleotide (e.g., a DNA template, a precursor linear RNA polynucleotide, or a circular RNA polynucleotide) comprises a lagging untranslated sequence (also referred to as “trailing sequence”).
- the lagging untranslated sequence is located at the 3’ end.
- the polynucleotide comprises a 3' external spacer located between the 3' intron element and a lagging untranslated sequence.
- the polynucleotide a leading untranslated sequence at the 5' end.
- the polynucleotide comprises a 5' external spacer located between a leading untranslated sequence and the 5' intron element.
- the polynucleotide comprises a monotron element and a leading untranslated sequence. In some embodiments, the polynucleotide comprises a 5' external spacer positioned between a leading untranslated sequence and either the terminal element or monotron element. In some embodiments, the polynucleotide comprises a monotron element and a lagging untranslated sequence. In some embodiments, the polynucleotide comprises a 3’ external spacer positioned between the lagging untranslated sequence and either the monotron element or terminal element.
- the lagging untranslated sequence comprises a restriction site sequence or a fragment thereof.
- the restriction site sequence or fragment thereof is used to linearize the polypeptide (e.g., DNA template).
- the restriction site sequence is derived from a natural viral, bacterial or eukaryotic DNA template.
- RNA polynucleotide comprising a monotron (also called a monotron element or monotron sequence) and a terminal element (also called a terminal sequence).
- the monotron has ribozymatic activity that allows it to enzymatically self-cleave.
- the monotron is capable of forming a phosphodiester bond with a terminal sequence, i.e., a sequence containing a splice site dinucleotide and optionally a natural exon sequence or fragment thereof.
- the precursor RNA polynucleotide comprises a terminal element; an intervening region, and a monotron element.
- the precursor RNA polynucleotide comprises, in the following order, (a) a terminal element; (b) an intervening region, and (c) a monotron element.
- the terminal sequence is upstream of the monotron sequence in the precursor RNA polynucleotide.
- the terminal element comprises a splice site nucleotide
- the monotron element comprises a splice site dinucleotide at or near the 5’ end of the monotron
- the monotron element is capable of interacting with a nucleophile that is capable of cleaving at the splice site dinucleotide at or near the 5’ end of the monotron, where the cleavage product of (iii) comprises a 5’ splice site nucleotide that is capable of cleaving at the splice site nucleotide of the terminal element.
- the nucleophile is a free nucleophile that is introduced to the precursor RNA polynucleotide, e.g., not in cis and/or covalently linked to the precursor RNA polynucleotide.
- the nucleophile is a guanosine that is capable of cleaving at the splice site dinucleotide at or near the 5’ end of the monotron.
- the guanosine is a free guanosine that is introduced to the precursor RNA polynucleotide, e.g., not in cis and/or covalently linked to the precursor RNA polynucleotide.
- the cleavage product of (iii) comprises a 5’ splice site nucleotide having a 3’ hydroxyl group that is capable of cleaving at the splice site nucleotide of the terminal element.
- the precursor RNA polynucleotide comprises, in the following order, (a) a monotron element; (b) an intervening region, and (c) terminal element.
- the monotron sequence is upstream of the terminal sequence in the precursor RNA polynucleotide.
- the monotron element comprises a splice site dinucleotide at or near the 3’ end of the monotron
- the terminal element comprises a splice site nucleotide
- the monotron element is capable of interacting with a nucleophile that is capable of cleaving at the splice site nucleotide of the terminal element, where the cleavage product of (iii) comprises a 5’ splice site nucleotide that is capable of cleaving at the splice site dinucleotide at or near the 3’ end of the monotron.
- the nucleophile is a free nucleophile that is introduced to the precursor RNA polynucleotide, e.g., not in cis and/or covalently linked to the precursor RNA polynucleotide.
- the nucleophile is a guanosine that is capable of cleaving at the splice site nucleotide of the terminal element.
- the guanosine is a free guanosine that is introduced to the precursor RNA polynucleotide, e.g., not in cis and/or covalently linked to the precursor RNA polynucleotide.
- the cleavage product of (iii) comprises a 5’ splice site nucleotide having a 3’ hydroxyl group that is capable of cleaving at the splice site nucleotide of the terminal element.
- the monotron can perform two transesterification reactions.
- the monotron can (a) self-cleave and (b) form a phosphodiester bond with the terminal sequence.
- the reactions (a) and (b) are sequential.
- the monotron is capable of interacting with a nucleophile that is capable of cleaving at the splice site dinucleotide at or near the 5’ end of the monotron, and (b) the cleavage product of (a), i.e., the 5’ splice site nucleotide, e.g., having a 3’ hydroxyl group, engages in a transesterification reaction (cleaves) at the splice site nucleotide of the terminal sequence, yielding a circular RNA or oRNA.
- the monotron interacts with the nucleophile by forming a binding pocket with the nucleophile, and the linear precursor is capable of adopting a conformation in which the nucleophile is in proximity to and is capable of cleaving at the splice site dinucleotide at or near the 5’ end of the monotron.
- the nucleophile can be a guanosine, e.g., a free guanosine that is introduced to the precursor RNA polynucleotide.
- the monotron can also perform two transesterification reactions.
- the monotron is capable of interacting with a nucleophile that is capable of cleaving at the splice site nucleotide of the terminal element, and (b) the cleavage product of (a), i.e., the 5’ splice site nucleotide, e.g., having a 3’ hydroxyl group, engages in a transesterification reaction (cleaves) at the splice site dinucleotide at or near the 3 ’ end of the monotron, yielding a circular RNA or oRNA.
- the monotron interacts with the nucleophile by forming a binding pocket with the nucleophile, and the linear precursor
- I l l is capable of adopting a conformation in which the nucleophile is in proximity to and is capable of cleaving the splice site nucleotide of the terminal element.
- the nucleophile can be a guanosine, e.g., a free guanosine that is introduced to the precursor RNA polynucleotide.
- the monotron comprises a 5’ proximal end of a natural group I or group II intron including the splice site dinucleotide and optionally a natural exon sequence or fragment thereof.
- the 5’ end of the monotron refers to nucleotides within the 5’ half of the monotron.
- the 3’ end of the monotron refers to nucleotides within the 3’ half of the monotron.
- at or near the 5’ end of the monotron refers to within the 5’ half of the monotron.
- at or near the 5’ end of the monotron refers to within the first ten 5’ positions in the monotron.
- At the 5’ end of the monotron refers to the first 5’ position(s) in the monotron. In some embodiments, at or near the 3’ end of the monotron refers to within the 3’ half of the monotron. In some embodiments, at or near the 3’ end of the monotron refers to within the last ten 3’ positions in the monotron. In some embodiments, at the 3’ end of the monotron refers to last 3’ position(s) in the monotron.
- the splice site nucleotide of the terminal element is not a natural splice site dinucleotide associated with a natural Group I or Group II intron sequence.
- the terminal element comprises at least a portion of a natural exon or a fragment of a natural exon.
- the natural exon is a Group I or Group II exon.
- the natural exon or fragment thereof is 10-20 nucleotides in length.
- the terminal element comprises a synthetic derivative of a natural exon or fragment thereof.
- the terminal element comprises an exon or synthetic nucleotides that are longer than the splice site nucleotide that can help with splicing.
- the terminal element sequence has a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more to an exon fragment of a sequence selected from Tables A or B.
- the terminal element sequence comprises an exon fragment comprising one, two, three, four, five, six, seven, eight, nine, ten, or more mutations to a sequence selected from Tables A or B.
- the mutations are, for example, selected from insertions, deletions, mutations, additions, and subtractions.
- the terminal element or exon fragment thereof comprises a polynucleotide sequence selected from a sequence set forth in SEQ ID NOS: 2990-3668.
- the terminal element is less than 500, less than 450, less than 400, less than 350, less than 300, less than 250, less than 200, less than 150, less than 100, less than 90, less than 80, less than 70, less than 60, less than 50, less than 40, less than 30, less than 20, or less than 10 nucleotides in length.
- the terminal element is capable of directing or functionalizing the splicing activity of the monotron element.
- a portion of the terminal segment is retained upon circularization. In some embodiments, a portion of the terminal segment is excised upon circularization. In some embodiments, all or a portion of the terminal element is excised postcircularization. In some embodiments, the terminal element is not excised upon cleavage and is retained post-cleavage.
- the monotron element comprises at least a portion of a Group I or Group II intron.
- Group I or Group II intron is selected from a genus and/or species described in Tables A or B.
- the Group I or Group II intron is from a gene selected from Cyanobacterium Anabaena sp., T4 phage, Hypocrea pallida, Bulbithecium hyalosporum, Myoarachis inversa, Geosmithia argillacea, Coxiella burnetii, Agrobacterium tumefaciens, Azoarcus, Nostoc, Cordyceps capitata, Prochlorothrix hollandica, and Tilletiopsis orzyzicola.
- the monotron element or a fragment thereof has a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more to a sequence selected from Tables A or B.
- the monotron element sequence or fragment thereof comprises one, two, three, four, five, six, seven, eight, nine, ten, or more mutations to a sequence selected from Tables A or B.
- the mutations are, for example, selected from insertions, deletions, additions, and subtractions.
- the monotron element sequence or fragment thereof comprises a polynucleotide sequence having a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or more to a portion of a sequence set forth in SEQ ID NOS: 2990-3668.
- the Group I or Group II intron or introns, or portion thereof are at least 10 nucleotides in length.
- the monotron element comprises at least one mutation of a native Group I intron-adjacent exon sequence or Group II intron-adjacent exon sequence.
- the at least one mutation is at least one substation mutation of a native Group I intron-adjacent exon sequence or Group II intron-adjacent exon sequence.
- the at least one mutation is at least one deletion of a native Group I intron- adjacent exon sequence or Group II intron-adjacent exon sequence.
- at least one of the exon segments is less than 15 nucleotides in length.
- the monotron element comprises a 3' exon segment and/or 5' exon segment, wherein the 3’ or 5’ exon segment comprises a Group I exon segment or a Group II exon segment less than 15 nucleotides in length.
- the 3' exon segments and/or 5' exon segments have a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more to a native Group I intron-adjacent exon sequence or Group II intron-adjacent exon sequence.
- the monotron element is less than 500 nucleotides in length.
- the monotron element is capable of inducing circularization when it interacts with the terminal element. In some embodiments, a portion of the monotron element is excised post-circularization. In some embodiments, the monotron element is fully excised post-circularization. In some embodiments, a portion of the monotron element and a portion of the terminal element are retained and excised post-circularization.
- the precursor RNA polynucleotide comprises at least one affinity tag or affinity sequence. Affinity sequences are described in further detail herein.
- the affinity tag comprises a polyA sequence or is a polyA affinity tag.
- the terminal element comprises an affinity tag. In some embodiments, the terminal element comprises a 5' affinity tag or a 3' affinity tag.
- the monotron element comprises an affinity tag. In some embodiments, the monotron element comprises a 3' affinity tag or a 5' affinity tag.
- the precursor RNA polynucleotide comprises an internal and/or external spacer. Spacers of the present disclosure are described in further detail herein.
- the precursor RNA polynucleotide comprises an internal spacer sequence positioned between the terminal element and the intervening region. In some embodiments, the precursor RNA polynucleotide comprises an internal spacer sequence positioned between the intervening region and the monotron element. In some embodiments, the precursor RNA polynucleotide comprises an external spacer. In some embodiments, the external spacer is positioned adjacent to the terminal element. In some embodiments, the external spacer is positioned adjacent to the monotron element.
- the precursor RNA polynucleotide comprises internal spacers and/or external spacers.
- the internal spacers and external spacers can each comprise an unstructured, structured or randomly generated polynucleotide sequence.
- the internal spacers and external spacers are at least 5 nucleotides in length and can be about 5 - 60 nucleotides in length. In some embodiments, the internal and external spacers are 5 - 60 nucleotides in length, inclusive.
- the precursor RNA polynucleotide comprises one or more duplexes. Duplexes of the present disclosure are described in further detail herein.
- the precursor RNA polynucleotide comprises a 5' internal duplex sequence and a 3' internal duplex sequence. In embodiments where the terminal element is upstream of the monotron element, the 5' internal duplex sequence is positioned between the terminal element and the intervening region, and the 3' internal duplex sequence is positioned between the intervening region and the monotron element.
- the 5' internal duplex sequence is positioned between monotron and the intervening region, and the 3' internal duplex sequence is positioned between the intervening region and the terminal element.
- the 5' internal duplex sequence and 3' internal duplex sequence are at least 80% complementary.
- a duplex is 3-100 nucleotides in length.
- a duplex is 5-20 nucleotides in length, inclusive.
- the 5' and 3' internal duplex sequences are capable of forming, and are predicted to form, a contiguous duplex.
- the continuous duplex has a length of no longer than about 35 nucleotides.
- the 5' internal duplex sequence and/or 3' internal duplex sequence each have a GC content of at least 10%.
- the precursor RNA polynucleotide comprises at least one affinity tag and at least one external spacer. In some embodiments, the precursor RNA polynucleotide comprises at least one internal duplex and at least one internal spacer, for example a 5’ affinity tag and 5’ internal spacer and/or a 3’ affinity tag and 3’ internal spacer. In embodiments where the polynucleotide comprises a 5’ affinity tag, the 5' affinity tag is positioned adjacent to the 5' external spacer, and in certain embodiments is positioned 5' to the 5' external spacer. In some embodiments where the polynucleotide comprises a 3' affinity tag, the 3' affinity tag is positioned adjacent to the 3’ external spacer, and in certain embodiments is positioned 3' to the 3' external spacer.
- the precursor RNA polynucleotide comprises at least one duplex and at least one internal spacer. In some embodiments, the precursor RNA polynucleotide comprises at least one internal duplex and at least one internal spacer, for example a 5’ internal duplex and a 5’ internal spacer and/or a 3’ internal duplex and a 3’ internal spacer. In some embodiments where the polynucleotide comprises a 5' internal duplex, the 5' internal duplex is positioned adjacent to the 5' internal spacer, and in certain embodiments is positioned 5' to the 5' internal spacer.
- the 3' internal duplex is positioned adjacent to the 3' internal spacer, and in certain embodiments, the 3' internal duplex is positioned 3' to the 3' internal spacer.
- the precursor polynucleotide comprises a 3’ and/or 5’ exon segment. In some embodiments, at least a portion of the 3’ and/or 5’ exon segment is codon optimized.
- the precursor RNA polynucleotide described above further comprises a leading untranslated sequence and/or a lagging untranslated sequence.
- the precursor RNA polynucleotide can comprise a 5' external spacer that is positioned between a leading untranslated sequence and the terminal element if the terminal element is upstream of the monotron element; or between a leading untranslated sequence and the monotron element if the monotron element is upstream of the terminal element.
- the precursor RNA polynucleotide comprises a 3' external spacer that is positioned between the monotron element and a lagging untranslated sequence if the terminal element is upstream of the monotron element; or between the terminal element and a lagging untranslated sequence if the monotron element is upstream of the terminal element.
- the intervening region of the precursor RNA polynucleotide can comprise sequences directed to, for example, an aptamer, a coding element, a stop codon or stop cassette, an intervening region comprising an untranslated region, a noncoding element.
- the intervening region can comprise a coding element where the coding element comprises, for example, a sequence encoding a therapeutic protein.
- the intervening region comprises an untranslated region, which can comprise one or more non-coding element, including but not limited to, a natural 5' Untranslated Region (UTR), a natural 3' Untranslated Region (UTR), a synthetic spacer sequence, an aptamer, TIE, a viral or eukaryotic IRES, or sequences selected from, e.g., IncRNA, miRNA, or a miRNA sponge.
- a natural 5' Untranslated Region UTR
- UTR natural 3' Untranslated Region
- synthetic spacer sequence an aptamer
- TIE a viral or eukaryotic IRES
- sequences selected from, e.g., IncRNA, miRNA, or a miRNA sponge e.g., IncRNA, miRNA, or a miRNA sponge.
- the polynucleotide comprises at least one mutation of a native Group I intron-adjacent exon sequence or Group II intron-adjacent exon sequence.
- the mutation can be, for example, at least one mutation of a native Group I or native Group II intron-adjacent exon sequence.
- the mutation can be one substitution, at least one deletion, and/or at least one insertion of a native Group I or Group II intron-adjacent exon sequence.
- at least one of the exon segments is less than 15 nucleotides in length.
- the 3' exon segment and/or 5' exon segment comprises a Group I exon segment or a Group II exon segment.
- the at least one exon segment is less than 15 nucleotides in length. In some embodiments, the at least one exon segment has a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more to a native Group I intron-adjacent exon sequence or Group II intron-adjacent exon sequence. In some embodiments, the at least one segment is selected from a 3' exon segment, 5' exon segment, or a 3' and 5' permuted exon segment. In some embodiments, the exon sequence or fragment is in the terminal element.
- polynucleotides encoding the precursor RNA polynucleotides described above that comprise a monotron and terminal element.
- Polynucleotides for example, DNA templates comprising sequences encoding the precursor RNAs described above, and their uses in related methods are described elsewhere herein.
- Polynucleotides of the present disclosure can comprise, for example, an expression vector, DNA plasmid, a cosmid, a PCR product, dbDNA close-ended DNA (ceDNA), and a viral polynucleotide.
- the polynucleotides can comprises a promoter segment, for example a T7 promoter, SP6 promoter or a fragment thereof.
- RNA polynucleotides produced by the precursor RNAs described above that comprise a monotron and terminal element. Circular RNAs are described in detail elsewhere herein.
- a circular RNA polynucleotide comprises: at least a portion of a terminal element, an intervening region, and at least a portion of a monotron element.
- a circular RNA comprises: (a) at least a portion of a terminal element, (b) a 3' exon segment comprising a 3' nucleotide of a 3' splice site dinucleotide, (c) an intervening region, (d) a 5' exon segment comprising a 5' nucleotide of a 5' splice site dinucleotide, and (e) at least a portion of a monotron element.
- (d) comprises the first nucleotide of a 5' Group I or Group II splice site dinucleotide and a natural exon sequence and (b) comprises the second nucleotide of a 3' Group I or Group II exon splice site dinucleotide and a natural exon sequence.
- the 5' and/or 3' splice site dinucleotides are distinct from the natural splice site dinucleotide(s) associated with a natural Group I or Group II intron sequence.
- the circular RNA polynucleotides comprise additional elements, including but not limited to, a 5' internal duplex and/or 3' internal duplex; a 5' internal spacer and/or 3' internal spacer. In some embodiments, the circular RNA polynucleotide is from about 50 nucleotides to about 15 kilobases in length.
- RNA polynucleotides comprising the precursor RNA polynucleotides, delivery or transfer vehicles, and pharmaceutical compositions thereof are described elsewhere herein in further detail.
- Related methods of producing circularized RNA and related methods of treating a subject in need thereof are also provided herein.
- Also provided herein are methods of identifying a monotron element and terminal element pair that allows production of a circular RNA that is translatable or biologically active inside a eukaryotic cell comprising, for example: (i) inserting a mutated 5' and 3' Group I or Group II intron sequence derived from a database of native intronic sequence to form a monotron element into a precursor RNA polynucleotide described above; (ii) inserting a synthetic polynucleotide sequence to form a terminal element into a precursor RNA polynucleotide described above; (iii) transcribing the polynucleotide into RNA in vitro or allowing the polynucleotide to be transcribed into RNA by a cell; and (iv) determining the circularization efficiency of the RNA produced by the polynucleotide by identifying the amount of circularized RNA, the amount of excised intronic sequences, the amount of precursor RNA remaining after circularization,
- the mutated 5' and 3' Group I or Group II intron sequence comprises at least one deletion, insertion or substitution of at least one nucleotide. In some embodiments, the 5' or 3' Group I or Group II intronic sequences, or combinations thereof are sequenced.
- Also provided herein are methods for determining a polynucleotide sequence that improves RNA circularization efficiency compared to a polynucleotide comprising a native intronic sequence or to a parent polynucleotide with a known sequence comprising modifying a DNA sequence encoding the precursor RNA polynucleotide described above, the modifying comprising: (i) mutating at least one nucleotide and/or altering the length of the terminal element and/or monotron element of the DNA sequence encoding the precursor RNA polynucleotide described above; (ii) altering the length of the 5' and/or 3' internal and/or external spacer sequence of the DNA sequence encoding precursor RNA polynucleotide described above; (iii) altering the length of the 5' and/or 3' internal duplex sequence of the DNA sequence encoding the precursor RNA polynucleotide described above; (iv) altering the length of the 5' and
- a provided polynucleotide (e.g., a DNA template, a linear precursor RNA polynucleotide, or a circular RNA polynucleotide) comprises an intervening region.
- the intervening region and/or core functional element comprises one or more noncoding elements, e.g., microRNA binding site, IRES transacting factor region, restriction site, a RNA editing region, structural or sequence element, a granule site, a zip code element, or an RNA trafficking element.
- the intervening region and/or core functional element comprises one or more coding elements.
- the intervening region and/or core functional element comprises a combination of coding and noncoding elements.
- the coding or non-coding region is a part of the core functional element or intervening region located between the 5’ end and 3’ end of the linear precursor RNA polynucleotide and resultant circular RNA.
- the coding element comprises an expression sequence. In some embodiments, the coding element comprises a sequence encoding at least one therapeutic protein. In some embodiments, the coding element encodes two or more polypeptides. In some embodiments, the sequences encoding the two or more polypeptides are separated by a ribosomal skipping element or a nucleotide sequence encoding a protease cleavage site.
- the ribosomal skipping element encodes thosea-asigna virus 2A peptide (T2A), porcine teschovirus-1 2 A peptide (P2A), foot-and-mouth disease virus 2 A peptide (F2A), equine rhinitis A vims 2A peptide (E2A), cytoplasmic polyhedrosis vims 2A peptide (BmCPV 2 A), or flacherie vims of B. mori 2 A peptide (BmIFV 2 A). Coding elements or regions and payloads are described in further detail elsewhere herein.
- the intervening region comprises at least one translation initiation element (TIE).
- TIEs are designed to allow translation efficiency of an encoded protein.
- core functional elements comprising one or more coding elements will further comprise one or more TIEs.
- a translation initiation element (TIE) comprises a synthetic TIE.
- a synthetic TIE comprises aptamer complexes, synthetic IRES or other engineered TIEs capable of initiating translation of a linear RNA or circular RNA polynucleotide.
- the intervening region comprises one or more noncoding elements.
- the noncoding element comprises an untranslated region (UTR) or fragment thereof.
- the noncoding element is a natural 5' UTR.
- the noncoding element is a natural 3' UTR.
- the noncoding element is a synthetic spacer sequence.
- the noncoding element is an aptamer.
- the noncoding element is or comprises a translation initiation element (TIE).
- TIE translation initiation element
- the noncoding element comprises a IncRNA, miRNA, or a miRNA sponge.
- the intervening region comprises a TIE comprising an untranslated region (UTR) or a fragment thereof, an aptamer complex or a fragment thereof, or a combination thereof.
- the TIE contains modified nucleotides.
- the TIE provided herein comprise an internal ribosome entry site (IRES).
- the TIE provided herein comprise a viral or eukaryotic internal ribosome entry site (IRES) or a fragment or variant thereof.
- the IRES comprises one or more modified nucleotides compared to the wildtype viral IRES or eukaryotic IRES. See, e.g., PCT Application No. US2022/33091, which is incorporated herein by reference in its entirety.
- the noncoding element comprises an untranslated region (UTR). In some embodiments, the noncoding element is a natural 5’ UTR. In some embodiments, the noncoding element is a natural 3’ UTR. In some embodiments, the noncoding element is a synthetic spacer sequence. In some embodiments, the noncoding element is an aptamer or synthetic aptamer. In some embodiments, the noncoding element is or comprises a translation initiation element (TIE). b . TRANSLATION INITIATION ELEMENT
- the DNA template, linear precursor RNA polynucleotide, and circular RNA polynucleotide comprise an intervening region and/or core functional element.
- the intervening region and/or core functional element comprises a coding and/or noncoding element.
- the intervening region and/or core functional element further comprises a translation initiation element (TIE) upstream to the coding or noncoding element, and/or a termination element.
- TIE translation initiation element
- the polynucleotide comprises a translation initiation element (TIE).
- TIE translation initiation element
- the intervening region comprises at least one TIE.
- the TIE is upstream to a coding or noncoding element.
- TIEs are designed to allow translation efficiency of an encoded protein. Accordingly, in some embodiments, an intervening region comprising one or more coding elements further comprises one or more TIEs. In other embodiments, an intervening region comprising only noncoding elements lacks any TIEs.
- a TIE comprises an internal ribosome entry site (IRES).
- the TIE provided herein comprise a viral or eukaryotic internal ribosome entry site (IRES) or a fragment or variant thereof.
- inclusion of an IRES permits the translation of one or more open reading frames from a circular RNA (e.g., open reading frames that form the expression sequences).
- IRES attracts a eukaryotic ribosomal translation initiation complex and promotes translation initiation. See, e.g., PCT Application No.WO202261490, which is incorporated herein by reference in its entirety.
- Natural TIES viral & eukaryotic/cellular IRES
- the payload encoded by the circular RNA polynucleotide may be optimized through use of a specific internal ribosome entry sites (IRES) within the translation initiation element (TIE).
- IRES internal ribosome entry sites
- TIE translation initiation element
- IRES specificity within a circular RNA can significantly enhance expression of specific proteins encoded within the coding element.
- the IRES comprises a viral IRES or eukaryotic IRES.
- IRES sequences are available and include sequences derived from a wide variety of viruses, such as from leader sequences of picornaviruses such as the encephalomyocarditis virus (EMCV) UTR (Jang et al., J. Virol. (1989) 63: 1651-1660), the polio leader sequence, the hepatitis A virus leader, the hepatitis C virus IRES, human rhinovirus type 2 IRES (Dobrikova et al., Proc. Natl. Acad. Sci. (2003) 100(25): 15125- 15130), an IRES element from the foot and mouth disease virus (Ramesh et al., Nucl. Acid Res.
- EMCV encephalomyocarditis virus
- UTR the polio leader sequence
- the hepatitis A virus leader the hepatitis C virus IRES
- human rhinovirus type 2 IRES Dobrikova et al., Proc. Natl. Acad. Sci.
- IRES giardiavirus
- a giardiavirus IRES a giardiavirus IRES
- IRES element attracts a eukaryotic ribosomal translation initiation complex and promotes translation initiation. See, e.g., Kaufman et al., Nuc. Acids Res. (1991) 19:4485-4490; Gurtu et al., Biochem. Biophys. Res. Comm.
- the IRES is capable of facilitating expression of a protein encoded by the precursor RNA in a cell. In some embodiments, the IRES is capable of facilitating expression of the protein, such that the expression level of the protein is comparable to or higher than when a control IRES is used.
- IRES sequences have varying ability to drive protein expression, and the ability of any particular identified or predicted IRES sequence to drive protein expression from linear mRNA or circular RNA constructs is unknown and unpredictable.
- potential IRES sequences can be bioinformatically identified based on sequence positions in viral sequences. However, the activity of such sequences has been previously uncharacterized. As demonstrated herein, such IRES sequences may have differing protein expression capability depending on cell type, for example in T cells, liver cells, or muscle cells.
- the novel IRES sequences described herein may have at least 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 50, or 100 fold increased expression in a particular cell type compared to previously described EMCV IRES sequences.
- a polynucleotide for driving protein expression, comprises an IRES operably linked to a protein coding sequence.
- the IRES comprises a sequence selected from the sequences in Table 1 or a fragment thereof or a sequence selected from SEQ ID NOS: 1-2989 and 4045-25570 (GIRES 0-10762), or a fragment thereof.
- the IRES comprises a sequence at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identical to a sequence selected from the sequences in Table 1 or a sequence selected from SEQ ID NOS: 1-2989 and 4045- 25570 (GIRES 0-10762), or a fragment thereof. See also, e.g., PCT Application No. US2022/33091 (WO202261490), which is incorporated herein by reference in its entirety.
- the IRES is derived from Aalivirus, Ailurivirus, Ampivirus, Anativirus, Aphthovirus, Aquamavirus, Avihepatovirus, Avisivirus, Boosepivirus, Bopivirus, Caecilivirus, Cardiovirus, Cosavirus, Crahelivirus, Crohivirus, Danipivirus, Dicipivirus, Diresapivirus, Enterovirus, Erbovirus, Felipivirus, Fipivirus, Gallivirus, Gruhelivirus, Grusopivirus, Harkavirus, Hemipivirus, Hepatovirus, Hunnivirus, Kobuvirus, Kunsagivirus, Limnipivirus, Livupivirus, Ludopivirus, Malagasivirus, Marsupivirus, Megrivirus, Mischivirus, Mosavirus, Mupivirus, Myrropivirus, Orivirus, Oscivirus, Parabovirus, Parechovirus, Pasivirus, Passerivirus, Pemapivirus
- the IRES is an IRES sequence derived from Taura syndrome virus, Triatoma virus, Theiler's encephalomyelitis virus, Simian Virus 40, Solenopsis invicta virus 1, Rhopalosiphum padi virus, Reticuloendotheliosis virus, Human poliovirus 1, Plautia stali intestine virus, Kashmir bee virus, Human rhinovirus 2, Homalodisca coagulata virus- 1, Human Immunodeficiency Virus type 1, Himetobi P virus, Hepatitis C virus, Hepatitis A virus, Hepatitis GB virus, Foot and mouth disease virus, Human enterovirus 71, Equine rhinitis virus, Ectropis obliqua picorna-like virus, Encephalomyocarditis virus, Drosophila C Virus, Human coxsackievirus B3, Crucifer tobamovirus, Cricket paralysis virus, Bovine viral diarrhea virus 1, Black Queen Cell Virus, Aphid lethal
- the IRES comprises in whole or in part a eukaryotic or cellular IRES.
- the IRES is an IRES sequence derived from a human gene, wherein the human gene is ABCF1, ABCG1, ACAD10, ACOT7, ACSS3, ACTG2, ADCYAP1, ADK, AGTR1, AHCYL2, AHI1, AKAP8L, AKR1A1, ALDH3A1, ALDOA, ALG13, AMMECR1L, ANGPTL4, ANK3, AOC3, AP4B1, AP4E1, APAF1, APBB1, APC, APH1A, APOBEC3D, APOM, APP, AQP4, ARHGAP36, ARL13B, ARMC8, ARMCX6, ARPC1A, ARPC2, ARRDC3, ASAP1, ASB3, ASB5, ASCL1, ASMTL, ATF2, ATF3, ATG4A, ATP5B, ATP6V0A1, ATXN3, A
- the cell is a myotube.
- the IRES is derived from Bopivirus, Oscivirus, Hunnivirus, Passerivirus, Mischivirus, Kobuvirus, Enterovirus, Cardiovirus, Salivirus, Rabovirus, Parechovirus, Gallivirus, or Sicinivirus.
- the IRES is derived from Hunnivirus, Passerivirus, Kobuvirus, Bopivirus, or Enterovirus.
- the IRES is derived from Enterovirus I, Enterovirus F, Enterovirus E, Enterovirus J, Enterovirus C, Enterovirus A, Enterovirus B, Aichivirus B, Parechovirus A, Cardiovirus F, Cardiovirus B, or Cardiovirus E.
- the cell is a hepatocyte.
- the IRES is derived from Enterovirus, Bopivirus, Mischivirus, Gallivirus, Oscivirus, Cardiovirus, Kobuvirus, Rabovirus, Salivirus, Parechovirus, Hunnivirus, Tottorivirus, Passerivirus, Cosavirus, or Sicinivirus.
- the IRES is derived from Enterovirus, Mischivirus, Kobuvirus, Bopivirus, or Gallivirus.
- the IRES is derived from Enterovirus B, Enterovirus A, Enterovirus D, Enterovirus J, Enterovirus C, Rhinovirus B, Enterovirus H, Enterovirus I, Enterovirus E, Enterovirus F, Aichivirus B, Aichivirus A, Parechovirus A, Cardiovirus F, Cardiovirus E, or Cardiovirus B.
- the cell is a T cell.
- the IRES is derived from Passerivirus, Bopivirus, Hunnivirus, Mischivirus, Enterovirus, Kobuvirus, Rabovirus, Tottorivirus, Salivirus, Cardiovirus, Parechovirus, Megrivirus, Allexivirus, Oscivirus, or Shanbavirus.
- the IRES is derived from Passerivirus, Hunnivirus, Mischivirus, Enterovirus, or Kobuvirus.
- the IRES is derived from Enterovirus I, Enterovirus D, Enterovirus C, Enterovirus A, Enterovirus J, Enterovirus H, Aichivirus B, Parechovirus A, or Cardiovirus B.
- a provided circular RNA comprises an IRES operably linked to a protein coding sequence.
- the IRES comprises a sequence selected from SEQ ID NOS: 1-2989 and 4045-25570 (GIRES 0- 10762) or Table 1 below, or SEQ ID NOs: 1-2983 and 3282-3287 of PCT Application No. US2022/33091 (WO202261490) or a fragment thereof.
- the IRES comprises a sequence at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identical to a sequence selected from SEQ ID NOS: 1-2989 and 4045-25570 or Table 1 below, or a sequence selected from SEQ ID NOs: 1-2983 and 3282-3287 of PCT Application No. US2022/33091 (WO202261490).
- the circular RNA disclosed herein comprises an IRES sequence at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identical to a sequence selected from SEQ ID NOS: 1-2989 and 4045-25570 or Table 1 below, or a sequence selected from SEQ ID NOs: 1-2983 and 3282-3287 of PCT Application No. US2022/33091 (WO202261490).
- the circular RNA disclosed herein comprises an IRES sequence selected from SEQ ID NOS: 1-2989 and 4045- 25570 or Table 1 below or a fragment thereof, or SEQ ID NOs: 1-2983 and 3282-3287 of PCT Application No. US2022/33091 (WO202261490) or a fragment thereof.
- RNA polynucleotide, circular RNA constructs and related pharmaceutical compositions disclosed herein comprise an IRES sequence that is at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identical to an IRES sequence in Table 1.
- precursor RNA polynucleotide, circular RNA constructs and related pharmaceutical compositions disclosed herein comprise an IRES sequence in Table 1.
- IRES sequence in the polynucleotide disclosed herein comprises one or more of these modifications relative to a natural or native IRES.
- Synthetic TIEs aptamer complexes, modified nucleotides, IRES variants & other engineered TIEs
- a TIE provided herein is a synthetic TIE.
- a synthetic TIE comprises aptamer complex, synthetic IRES, or other engineered TIE capable of initiating translation of a linear RNA or circular RNA polynucleotide.
- one or more aptamer sequences are capable of binding to a component of a eukaryotic initiation factor to either enhance or initiate translation.
- an aptamer may be used to enhance translation in vivo and in vitro by promoting specific eukaryotic initiation factors (elF) (e.g., certain aptamers disclosed in International Pat. Appl. No. PCTZEP2018/078794 are capable of binding to eukaryotic initiation factor 4F (eIF4F)).
- elF eukaryotic initiation factors
- eIF4F eukaryotic initiation factor 4F
- an aptamer or a complex of aptamers may be capable of binding to EIF4G, EIF4E, EIF4A, EIF4B, EIF3, EIF2, EIF5, EIF1, EIF1A, 40S ribosome, PCBP1 (polyC binding protein), PCBP2, PCBP3, PCBP4, PABP1 (poly A binding protein), PTB, Argonaute protein family, HNRNPK (heterogeneous nuclear ribonucleoprotein K), or La protein.
- the intervening region and/or core functional element comprises a stop codon or stop cassette.
- the sequence is located downstream to a TIE and coding element. In some embodiments, the sequence is located downstream to a coding element and upstream to a TIE.
- the intervening region comprises a stop codon. In one embodiment, the intervening region comprises a stop cassette. In some embodiments, the stop cassette comprises at least 2 stop codons. In some embodiments, the stop cassette comprises at least 2 frames of stop codons. In the same embodiment, the frames of the stop codons in a stop cassette each comprise 1, 2 or more stop codons. In some embodiments, the stop cassette comprises a LoxP or a RoxStopRox, or frt- flanked stop cassette. In the same embodiment, the stop cassette comprises a lox-stop-lox stop cassette.
- a provided polynucleotide e.g., a DNA template, a linear precursor RNA polynucleotide, or a circular RNA polynucleotide
- a provided polynucleotide further comprises one or more elements for enhancing circularization, translation, or both.
- these elements are located with specificity between or within the intron elements, exon elements, or intervening region of the polynucleotide.
- a polynucleotide, a precursor RNA polynucleotide, or circular RNA can comprise an IRES transacting factor region, a miRNA binding site, a restriction site, an RNA editing region, a structural or sequence element, a granule site, a zip code element, and/or an RNA trafficking element or another specialized sequence as found in the art that enhances promotes circularization and/or translation of the protein encoded within the circular RNA polynucleotide.
- the polynucleotide, precursor RNA polynucleotide, or circular RNA comprises an IRES transacting factor (ITAF) region.
- the IRES transacting factor region modulates the initiation of translation through binding to PC- P1 - PCBP4 (polyC binding protein), PABP1 (poly A binding protein), PTB (polyprimidine tract binding), Argonaute protein family, HNRNPK (Heterogeneous nuclear ribonucleoprotein K protein), or La protein.
- the IRES transacting factor region comprises a poly A, polyC, poly AC, or polyprimidine track.
- the ITAF region is located within the intervening region or core functional element. In some embodiments, the ITAF region is located within the TIE.
- the polynucleotide, precursor RNA polynucleotide, or circular RNA comprises a IncRNA, miRNA, or a miRNA sponge.
- at least one miRNA binding site is included.
- the miRNA binding site is located within the 5’ intron element, 5’ exon element, intervening region or core functional element, 3’ exon element, and/or 3’ intron element.
- the miRNA binding site is located within the spacer within the intron element or exon element.
- the miRNA binding site comprises the entire spacer regions.
- the 5’ intron element and 3’ intron elements each comprise identical miRNA binding sites.
- the miRNA binding site of the 5’ intron element comprises a different, in length or nucleotides, miRNA binding site than the 3’ intron element.
- the 5’ exon element and 3’ exon element comprise identical miRNA binding sites.
- the 5’ exon element and 3’ exon element comprise different, in length or nucleotides, miRNA binding sites.
- the miRNA binding sites are located adjacent to each other within the circular RNA construct, linear RNA polynucleotide precursor, and/or DNA template.
- the first nucleotide of one of the miRNA binding sites follows the first nucleotide last nucleotide of the second miRNA binding site.
- the miRNA binding site is located within a translation initiation element (TIE) of an intervening region or core functional element. In one embodiment, the miRNA binding site is located before, trailing or within an internal ribosome entry site (IRES). In another embodiment, the miRNA binding site is located before, trailing, or within an aptamer complex.
- TIE translation initiation element
- IRS internal ribosome entry site
- miRNA binding sequences can permit tissue-specific expression of a coding sequence within an intervening region or core functional element.
- miRNA binding sequences resulting in expression suppression in tissues such as the liver or kidney may be desired.
- Such miRNA binding sequences may be selected based on the cell or tissue expression of miRNAs.
- the unique sequences defined by the miRNA nomenclature are widely known and accessible to those working in the microRNA field. For example, they can be found in the miRDB public database.
- one or more miR-122 target sites can be inserted in the circular RNA.
- the miR-122 site can comprise the following sequence:
- CAAACACCATTGTCACACTCCAA (SEQ ID NO: 4018).
- RNAs in some instances produced by the precursor RNA polynucleotides described herein.
- RNA polynucleotide comprising, in the following order, a 3 ' self-spliced exon segment, an intervening region, and a 5 ' self-spliced exon segment.
- a circular RNA polynucleotide comprising, in the following order, a 3 ' self-spliced exon segment, a coding sequence, and a 5 ' self-spliced exon segment.
- RNA polynucleotide comprising, in the following order, a 3 ' self-spliced exon segment, a translation initiation element (TIE), a coding sequence, and a 5 ’ self-spliced exon segment.
- TIE translation initiation element
- a circular RNA polynucleotide comprising, in the following order, a 3 ’ self-spliced exon segment, a translation initiation element (TIE), a coding sequence with which the TIE is not naturally associated, and a 5 ’ self-spliced exon segment.
- RNA polynucleotide comprising: i) a 5’ combined accessory element; ii) an intervening region; and iii) a 3’ combined accessory element, where the intervening region is between the 5’ combined accessory element and the 3’ combined accessory element.
- the 5 ' combined accessory element comprises a 3’ selfspliced exon segment.
- the 3’ self-spliced exon segment comprises an exon segment or fragment thereof.
- the 3’ self-spliced exon segment comprises a 3 ' nucleotide of a 3 ' splice site dinucleotide.
- the 3’ selfspliced exon segment comprises an exon segment and a 3’ nucleotide of a 3’ splice site dinucleotide.
- the exon segment comprises a natural exon sequence or non-naturally occurring sequence.
- the 3' splice site dinucleotides are distinct from the natural splice site dinucleotide(s) associated with a natural Group I or Group II intron sequence.
- the 3 ' self-spliced exon segment comprises a sequence having a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more to a sequence selected from SEQ ID NOs: 2990-3668, 25573, and 25574.
- the 3’ self-spliced exon segment is selected from an exon segment disclosed herein, e.g., in Table A or Table B, or SEQ ID NOs: 2990-3668, 25573, and 25574. See, e.g., supra.
- the self-spliced exon segment is, e.g., 5 nucleotides, 6 nucleotides, 7 nucleotides, 8 nucleotides, 9 nucleotides, 10 nucleotides, 11 nucleotides, 12 nucleotides, 13 nucleotides, 14 nucleotides, or 15 nucleotides.
- the circular RNA comprises a self-spliced exon segment that is 5 nucleotides, 6 nucleotides, 7 nucleotides, 8 nucleotides, 9 nucleotides, 10 nucleotides, 11 nucleotides, 12 nucleotides, 13 nucleotides, 14 nucleotides, or 15 nucleotides from the exonic sequences of Table A or is e.g., 5 nucleotides, 6 nucleotides, 7 nucleotides, 8 nucleotides, 9 nucleotides, or 10 nucleotides from the exonic sequences of Table B. See also SEQ ID NOs: 2990-3668, 25573, and 25574.
- the 3 ' combined accessory element comprises a 5’ selfspliced exon segment.
- the 5’ self-spliced exon segment comprises an exon segment or fragment thereof.
- the 5’ self-spliced exon segment comprises a 5 ' nucleotide of a 5 ' splice site dinucleotide.
- the 5’ selfspliced exon segment comprises an exon segment and a 5’ nucleotide of a 5’ splice site dinucleotide.
- the exon segment comprises a natural exon sequence or non-naturally occurring sequence.
- the 5 ’ splice site dinucleotides are distinct from the natural splice site dinucleotide(s) associated with a natural Group I or Group II intron sequence.
- the 5 ' self-spliced exon segment comprises a sequence having a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more to a sequence selected from SEQ ID NOs: 2990-3668, 25573, and 25574.
- the 5’ self-spliced exon segment is selected from an exon segment disclosed herein, e.g., in Table A or Table B, or SEQ ID NOs: 2990-3668, 25573, and 25574. See, e.g., supra.
- the self-spliced exon segment is e.g., 5 nucleotides, 6 nucleotides, 7 nucleotides, 8 nucleotides, 9 nucleotides, 10 nucleotides, 11 nucleotides, 12 nucleotides, 13 nucleotides, 14 nucleotides, or 15 nucleotides.
- the circular RNA comprises a self-spliced exon segment that is 5 nucleotides, 6 nucleotides, 7 nucleotides, 8 nucleotides, 9 nucleotides, 10 nucleotides, 11 nucleotides, 12 nucleotides, 13 nucleotides, 14 nucleotides, or 15 nucleotides from the exonic sequences of Table A or is e.g., 5 nucleotides, 6 nucleotides, 7 nucleotides, 8 nucleotides, 9 nucleotides, or 10 nucleotides from the exonic sequences of Table B. See also SEQ ID NOs: 2990-3668, 25573, and 25574.
- the intervening region comprises a noncoding region or a coding region.
- the intervening region comprises at least one translation initiation element (TIE).
- the TIE comprises a viral or eukaryotic internal ribosome entry site (IRES) and a noncoding or coding region.
- the IRES comprises a sequence selected from the sequences in Table 1 or a fragment thereof or a sequence selected from SEQ ID NOS: 1-2989 and 4045-25570. See, e.g., supra.
- the TIE comprises a coding sequence with which the TIE is not naturally associated.
- the intervening region comprises an untranslated region (UTR).
- the UTR comprises one or more noncoding elements.
- the one or more noncoding elements are selected from, e.g., a natural 3 ' Untranslated Region (UTR), a natural 5 ' Untranslated Region (UTR), a synthetic spacer sequence, an aptamer, and IncRNA, miRNA, and a miRNA sponge.
- the noncoding element is or comprises the TIE.
- the intervening region comprises a comprises a coding element or coding region.
- the coding element comprises a sequence encoding at least one therapeutic protein.
- the coding element encodes two or more polypeptides.
- the coding element or coding region comprises a sequence encoding, for example, a therapeutic protein, cytokine, immune checkpoint inhibitor, an agonist, a chimeric antigen receptor, an inhibitory receptor agonist or inhibitory receptor, an inhibitory receptor antagonist, one or more TCR chains, a secreted T cell or immune cell engager, a transcription factor, an immunosuppressive enzyme, or a TvHd, as set forth in detail herein.
- the coding element or coding region comprises one or more expression sequences or portions thereof, e.g., Table 2, infra.
- RNA polynucleotides comprising, in the following order, i) a 5’ combined accessory element comprising a 3’ selfspliced exon segment; ii) an intervening region; and iii) a 3’ combined accessory element comprising a 5’ self-spliced exon segment.
- the 3’ self-spliced exon segment and/or the 5’ self-spliced exon segment is selected from an exon segment disclosed herein, e.g., in Table A or Table B, or SEQ ID NOs: 2990-3668, 25573, and 25574.
- circular RNA polynucleotides comprising, in the following order, i) a 5’ combined accessory element comprising a 3’ selfspliced exon segment, wherein the 3’ self-spliced exon segment comprises an exon segment; ii) an intervening region; and iii) a 3’ combined accessory element comprising a 5’ self-spliced exon segment, wherein the 5’ self-spliced exon segment comprises an exon segment.
- the 3’ self-spliced exon segment and/or the 5’ self-spliced exon segment is selected from an exon segment disclosed herein, e.g., in Table A or Table B, or SEQ ID NOs: 2990-3668, 25573, and 25574.
- RNA polynucleotides comprising, in the following order, i) a 5’ combined accessory element comprising a 3’ selfspliced exon segment, wherein the 3’ self-spliced exon segment comprises an exon segment and a 3’ nucleotide of a 3’ splice site dinucleotide; ii) an intervening region; and iii) a 3’ combined accessory element comprising a 5’ self-spliced exon segment, wherein the 5’ selfspliced exon segment comprises an exon segment and a 5’ nucleotide of a 5’ splice site dinucleotide.
- the 3’ self-spliced exon segment and/or the 5’ self-spliced exon segment is selected from an exon segment disclosed herein, e.g., in Table A or Table B, or SEQ ID NOs: 2990-3668, 25573, and 25574.
- a circular RNA polynucleotide comprising, in the following order, a 3’ self-spliced exon segment, an intervening region, and a 5’ self-spliced exon segment, wherein at least one of the 3’ or 5’ self-spliced exon segments is selected from an exon segment comprising a sequence selected from SEQ ID NOs: 2990-3668, 25573, and 25574.
- RNA polynucleotide comprises the following elements operably connected and arranged in the following sequence:
- a circular RNA polynucleotide comprises a retained portion of a monotron element. See, e.g., supra.
- a circular RNA polynucleotide comprises: a 5’ internal spacer, a 5’ internal duplex, at least a portion of a terminal element (or sequence or segment), at least a portion of a monotron element (or sequence or segment), a 3’ internal duplex, a 3’ internal spacer, a coding or noncoding region, and an intervening region.
- the circular RNA polynucleotide comprises a coding region and the intervening region comprises an IRES.
- the monotron element present in the precursor RNA polynucleotide, of which a portion is retained in the circular RNA polynucleotide comprises a polynucleotide sequence that has a percent sequence identity of about 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or more to a sequence selected from SEQ ID NOs: 2990- 3187, 25573, and 25574.
- the circular RNA polynucleotide comprises the following elements operably connected and arranged in the following sequence:
- the circular RNA polynucleotide comprises the following elements operably connected and arranged in the following sequence:
- RNA polynucleotide comprises the following elements operably connected and arranged in the following sequence:
- element (d) comprises the first nucleotide of a 5 ' Group I or Group II splice site dinucleotide and a natural exon sequence.
- element (b) comprises the second nucleotide of a 3 ' Group I or Group II exon splice site dinucleotide and a natural exon sequence.
- the 5' exon element comprises the second nucleotide of a 3' Group I or Group II exon splice site dinucleotide and a natural exon sequence.
- the 3' exon element fragment comprises the first nucleotide of a 5' Group I or Group II splice site dinucleotide and a natural exon sequence.
- the 5' exon element comprises a 5' internal duplex; and the 3' exon element comprises a 3' internal duplex.
- the 5' exon element comprises a 5' internal spacer.
- the 3' exon element comprises a 3' internal spacer.
- the circular RNA polynucleotide comprises a 5’ internal duplex and a 3’ internal duplex. See, e.g., supra.
- the circular RNA polynucleotide comprises a 5’ internal homology region and/or a 3’ internal homology region. See, e.g., supra.
- the circular RNA polynucleotide comprises internal spacers (IS) of different lengths, e.g., a 5 ' internal spacer and/or a 3 ' internal spacer. See, e.g., supra.
- IS internal spacers
- the circular RNA polynucleotide retains portions of the precursor RNA polynucleotides, described elsewhere herein in detail. In some embodiments, portions of the precursor RNA polynucleotide are removed upon circularization. For example, in some embodiments, the circular RNA polynucleotide does not comprise a 5 ' external spacer and/or a 3 ' external spacer. In some embodiments, the circular RNA polynucleotide does not comprise a 5 ' intron segment and/or 3 ' intron segment. In some embodiments, the circular RNA polynucleotide does not comprise affinity tags. In some embodiments, the circular RNA polynucleotide does not retain a portion of a monotron element. In certain embodiments, the circular RNA polynucleotide does not retain a monotron element.
- the circular RNA polynucleotide comprises modified nucleotides and/or modified nucleosides, namely comprising at least one modified A, C, G, or U/T nucleotide or nucleoside. Exemplary modifications are described in detail elsewhere herein. See, e.g., infra.
- a circular RNA polynucleotide comprises modified nucleotides and/or modified nucleosides where between 1% and 100%, 1% and 2%, 1% and 3%, 1% and 4%, 1% and 5%, 5% and 6%, 5% and 7%, 5% and 8%, 5% and 9%, 5% and 10%, 10% and 20%, 20% and 30%, 30% and 40%, 40% and 50%, 50% and 60%, 60% and 70%, 70% and 80%, 80% and 90%, or 90% and 100% of the nucleotides or nucleosides are modified.
- portions of the polynucleotide comprise between 1% and 10% modification of the nucleotides or nucleosides.
- portions of the circular RNA polynucleotide comprise less than 10% modification. In some embodiments, portions of the polynucleotide or the polynucleotide in its entirety comprise no nucleotide or nucleoside modifications. In some embodiments, a circular RNA polynucleotide may lack modifications, where the linear precursors used to produce the circular RNA polynucleotide contained modifications (e.g., in the introns). See, e.g., Figures 24B, 24C, and 24D.
- incorporation of a nucleotide or nucleoside modification to a precursor RNA polynucleotide hinders or lowers the capacity of the circular RNA to circularize, splice, or express.
- the circular RNA polynucleotide is from about 50 nucleotides to about 15 kilobases in length.
- the circular RNA polynucleotide has an in vivo duration of therapeutic effect in a subject of at least about 10 hours. In some embodiments, the circular RNA polynucleotide has a functional half-life of at least about 10 hours. In some embodiments, the circular RNA polynucleotide has a duration of therapeutic effect in a cell greater than or equal to that of an equivalent linear RNA polynucleotide comprising the same expression sequence. In some embodiments, the circular RNA polynucleotide has a functional half-life in a cell greater than or equal to that of an equivalent linear RNA polynucleotide comprising the same expression sequence.
- the circular RNA polynucleotide has an in vivo duration of therapeutic effect in a subject greater than that of an equivalent linear RNA polynucleotide having the same expression sequence. In some embodiments, the circular RNA polynucleotide has an in vivo functional half-life in a subject greater than that of an equivalent linear RNA polynucleotide having the same expression sequence.
- RNA polynucleotide comprising a translation initiation element (TIE), a coding sequence (e.g., with which the TIE is not naturally associated), and a means for self-splicing. See, e.g., Example 8, demonstrating that self-splicing efficiency and/or circularization efficiency is linked to the structures herein, e.g., at Examples 1, 2, 8.
- TIE translation initiation element
- coding sequence e.g., with which the TIE is not naturally associated
- RNA polynucleotide comprising a translation initiation element (TIE), a coding sequence (e.g., with which the TIE is not naturally associated), and an autocatalytic intron-exon means for self-splicing. See, e.g., id.
- TIE translation initiation element
- coding sequence e.g., with which the TIE is not naturally associated
- autocatalytic intron-exon means for self-splicing See, e.g., id.
- TIE translation initiation element
- TIE translation initiation element
- coding sequence e.g., with which the TIE is not naturally associated
- RNA polynucleotide comprising, in the following order, a 3 ' exon segment means for self-splicing, a translation initiation element, a coding sequence, and a 5 ' exon segment means for self-splicing. See, e.g., id.
- a non-naturally occurring RNA polynucleotide comprising, in the following order, a 3 ' exon segment means for self-circularization, a translation initiation element, a coding sequence, and a 5 ' exon segment means for self-circularization.
- RNA polynucleotide comprising, in the following order, a 3 ' exon segment, a translation initiation element, a coding sequence, and a 5 ' exon segment, wherein the exon segments are means for self-splicing. See, e.g., id.
- a non-naturally occurring RNA polynucleotide comprising, in the following order, a 3 ' exon segment, a translation initiation element, a coding sequence, and a 5 ' exon segment, wherein the exon segments are means for self-circularization.
- RNA polynucleotide comprising, in the following order, a 3 ' exon segment means for self-circularization, a translation initiation element, a coding sequence, and a 5 ' exon segment means for self-circularization. See, e.g., id.
- a circular RNA polynucleotide comprising, in the following order, a 3 ' exon segment, a translation initiation element, a coding sequence, and a 5 ' exon segment, wherein the exon segments are means for self-splicing. See, e.g., id.
- a provided polynucleotide (e.g., a precursor RNA polynucleotide, a circular RNA polynucleotide, or a DNA template) comprises modified nucleotides and/or modified nucleosides, namely comprising at least one modified A, C, G, or U/T nucleotide or nucleoside.
- modified nucleotides and/or modified nucleosides namely comprising at least one modified A, C, G, or U/T nucleotide or nucleoside.
- such modifications differ from mutations selected from insertions, deletions, addition, or subtraction of nucleotides, for example, the mutations in a permuted Group I and Group II intron segment.
- the polynucleotide is a precursor RNA polynucleotide and comprises at least one modified A, C, G, or U nucleotide or nucleoside.
- the precursor RNA polynucleotide is linear.
- the precursor RNA polynucleotide is capable of producing a circular RNA comprising at least one modified nucleotide or nucleoside after splicing.
- the precursor RNA polynucleotide comprising one or more modified nucleotide or nucleoside is capable of circularizing when incubated in the presence of one or more guanosine nucleotides or nucleoside (e.g., GTP) and a divalent cation (e.g., Mg2+).
- the polynucleotide is a circular RNA polynucleotide and comprises at least one modified A, C, G, or U nucleotide or nucleoside modifications.
- modified nucleotides or nucleosides occur throughout a precursor RNA polynucleotide.
- the RNA polynucleotide comprises 5 ' and 3 ' combined accessory elements comprising one or more modified nucleotides.
- the RNA polynucleotide comprises an intron element and/or exon element comprising one or more modified nucleotide or nucleoside.
- portions of the 3 ' and/or 5 ' intron and/or exon segments in a linear precursor RNA polynucleotide of the present disclosure contain modified nucleotides or nucleosides.
- the secondary structures of at least the intron and/or exon segments are preserved.
- the terminal element comprises at least one modified nucleotide or nucleoside.
- the terminal element, intervening region, and/or monotron comprises at least one modified nucleotide or nucleoside.
- the RNA polynucleotide comprises a spacer comprising at least one modified nucleotide or nucleoside.
- the RNA polynucleotide comprises a duplex comprising at least one modified nucleotide or nucleoside. In certain embodiments, the RNA polynucleotide comprises an affinity sequence comprising at least one modified nucleotide or nucleoside. In certain embodiments, the RNA polynucleotide comprises a leading and/or lagging strand comprising at least one modified nucleotide or nucleoside. In some embodiments, the RNA polynucleotide comprises a coding or a noncoding element comprising at least one modified nucleotide or nucleoside.
- the RNA polynucleotide comprises a translation initiation element (TIE) comprising at least one modified nucleotide or nucleoside.
- TIE translation initiation element
- the polynucleotide comprises a stop codon and/or stop cassette comprising one or more modified nucleotide or nucleoside.
- a precursor RNA polynucleotide comprising at least one modified A, C, G, or U nucleotide or nucleoside comprises at least a portion of each of a. a 5’ combined accessory element, comprising: i. a 3’ intron segment, ii. a 3’ exon segment, b. an intervening region comprising an internal ribosome entry site (IRES) and a noncoding or coding region, c. a 3’ combined accessory element, comprising: i. a 5’ exon segment, and ii. a 5’ intron segment.
- a 5’ combined accessory element comprising: i. a 3’ intron segment, ii. a 3’ exon segment, and ii. a 5’ intron segment.
- a circular RNA comprising at least one modified A, C, G, or U nucleotide or nucleoside comprises at least a portion of each of: a. a post-splicing 3’ exon segment, b. optionally a 5’ internal homology region, c. optionally a 5’ spacer, d. an intervening region comprising an internal ribosome entry site (IRES) and a noncoding or coding region, e. optionally a 3’ spacer, f. optionally a 3’ internal homology region, and g. a post-splicing 5’ exon segment.
- a post-splicing 3’ exon segment b. optionally a 5’ internal homology region, c. optionally a 5’ spacer, d. an intervening region comprising an internal ribosome entry site (IRES) and a noncoding or coding region, e. optionally a 3’ spacer, f. optionally a 3’ internal homology region, and
- the modified nucleoside is m 5 C (5-methylcytidine). In another embodiment, the modified nucleoside is m 5 U (5-methyluridine). In another embodiment, the modified nucleoside is m 6 A (N 6 -methyladenosine). In another embodiment, the modified nucleoside is s 2 U (2-thiouridine). In another embodiment, the modified nucleoside is W (pseudouridine). In another embodiment, the modified nucleoside is Um (2 ' - O-methyluridine).
- the modified nucleoside is nfA (1- methyladenosine); m 2 A (2 -methyladenosine); Am (2' -O-methyladenosine); ms 2 m 6 A (2- methylthio-N 6 -methyladenosine); i 6 A (N 6 -isopentenyladenosine); ms 2 i6A (2-methylthio- N 6 isopentenyladenosine); io 6 A (N 6 -(cis-hydroxyisopentenyl)adenosine); ms 2 io 6 A (2- methylthio-N 6 -(cis-hydroxyisopentenyl)adenosine); g 6 A (N 6 -glycinylcarbamoyladenosine); t 6 A (N 6 -threonylcarbamoyladenosine); ms 2 t 6 A (2-methylthio-N 6 -threonyl
- the modified nucleoside may include a compound selected from the group of 146yridine-4-one ribonucleoside, 5-aza-uridine, 2-thio-5-aza-uridine, 2- thiouridine, 4-thio-pseudouridine, 2-thio-pseudouridine, 5-hydroxyuridine, 3 -methyluridine, 5-carboxymethyl-uridine, 1-carboxymethyl-pseudouridine, 5-propynyl-uridine, 1-propynyl- pseudouridine, 5-taurinomethyluridine, 1-taurinomethyl-pseudouridine, 5-taurinomethyl-2- thio-uridine, l-taurinomethyl-4-thio-uridine, 5-methyl-uridine, 1-methyl-pseudouridine, 4- thio- 1 -methyl-pseudouridine, 2-thio- 1 -methyl-pseudouridine, 1
- the modifications are independently selected from 5- methylcytosine, pseudouridine and 1 -methylpseudouridine.
- the modified ribonucleosides include 5 -methylcytidine, 5- methoxyuridine, 1-methyl-pseudouridine, N6-methyladenosine, and/or pseudouridine.
- the modified nucleoside is N1 -methylpseudouridine.
- the modified nucleotide or nucleoside is selected from one or more of 5-propynyluridine, 5-propynylcytidine, 6-methyladenine, 6-methylguanine, N,N,- dimethyladenine, 2-propyladenine, 2-propylguanine, 2-aminoadenine, 1 -methylinosine, 3- methyluridine, 5-methylcytidine, 5-methyluridine, 5-(2-amino)propyl uridine, 5- halocytidine, 5-halouridine, 4-acetylcytidine, 1 -methyladenosine, 2-methyladenosine, 3- methyicytidine, 6- methyluridine, 2-methylguanosine, 7-m ethylguanosine, 2,2- dimethylguanosine, 5- methylaminoethyluridine, 5-methyloxyuridine, 7-deaza-adenosine, 6- azouridine, 6- azo
- Additional modified nucleotides and nucleosides can be selected from clinically validated modified nucleotides described in the art. See, e.g., US20190345503A1 (m 6 A- modified circRNA); US20220288176A1 (m 6 A modification of circRNA); US20220251578A1 (at least one N6-methyladenosine (m 6 A)); WO2022271965A2 (N6-methyladenosine, 2- thiouridine, and 2' -O-methyl cytidine), which are each incorporated by reference in their entireties.
- a first and second precursor polynucleotide are provided, where the first precursor RNA polynucleotide comprises a 3 ' intron fragment of a first intron (Intron 1), a 5 ' intron fragment of a second intron (Intron 2), a translation initiation element, a fragment of a sequence of interest (e.g., coding region), and two exon fragments that correspond with the intron fragments; and the second precursor comprises a 3 ' intron fragment of the second intron (Intron 2) and a 5 ' intron fragment of the first intron (Intron 1), a fragment of the sequence of interest of the first precursor, and exon fragments corresponding to those in the first precursor.
- the first and second linear precursor RNA polynucleotides are capable of forming a circular RNA.
- the first precursor comprises no nucleotide or nucleoside modifications and the second precursor comprises nucleotide or nucleoside modifications.
- the first precursor comprises nucleotide or nucleoside modifications and the second precursor comprises no nucleotide or nucleoside modifications.
- the first precursor and the second precursor comprise no nucleotide or nucleoside modifications.
- the first precursor and the second precursor comprise nucleotide or nucleoside modifications.
- the presence of a modified nucleotide or nucleoside can affect the folding patterns and/or function of an accessory element, translation initiation element (TIE), and/or coding element within the circular RNA or linear precursor.
- Position and composition of a nucleotide or nucleoside modification in a polynucleotide are impacted by the nucleotide or nucleoside composition (i.e., A, C, G, or U nucleotide or nucleoside) of the accessory elements, TIE, or coding elements.
- a provided polynucleotide e.g., a precursor RNA polynucleotide, or a circular RNA polynucleotide, described in more detail elsewhere herein
- a provided polynucleotide e.g., a precursor RNA polynucleotide, or a circular RNA polynucleotide, described in more detail elsewhere herein
- 1% and 100% 1% and 2%, 1% and 3%, 1% and 4%, 1% and 5%, 5% and 6%, 5% and 7%, 5% and 8%, 5% and 9%, 5% and 10%, 10% and 20%, 20% and 30%, 30% and 40%
- 40% and 50%, 50% and 60%, 60% and 70%, 70% and 80%, 80% and 90%, or 90% and 100% of the nucleotides or nucleosides are unmodified.
- a provided polynucleotide (e.g., a precursor RNA polynucleotide, or a circular RNA polynucleotide, described in more detail elsewhere herein) comprises modified nucleotides and/or modified nucleosides where between 1% and 100%, 1% and 2%, 1% and 3%, 1% and 4%, 1% and 5%, 5% and 6%, 5% and 7%, 5% and 8%, 5% and 9%, 5% and 10%, 10% and 20%, 20% and 30%, 30% and 40%, 40% and 50%, 50% and 60%, 60% and 70%, 70% and 80%, 80% and 90%, or 90% and 100% of the nucleotides or nucleosides are modified. In some embodiments, between 1% and 10% of the nucleotides or nucleosides are modified.
- portions of the polynucleotide e.g., a precursor RNA polynucleotide, or a circular RNA polynucleotide, described in more detail elsewhere herein
- the polynucleotide e.g., a precursor RNA polynucleotide, or a circular RNA polynucleotide, described in more detail elsewhere herein
- the secondary structures of at least the intron and/or exon segments are preserved.
- the secondary structure of the coding or noncoding region is preserved.
- the IRES is unmodified or substantially unmodified to preserve secondary structure needed to initiate translation.
- nucleotides or nucleosides are modified in a polynucleotide of the present disclosure (e.g., a precursor RNA polynucleotide, or a circular RNA polynucleotide, described in more detail elsewhere herein).
- portions of the polynucleotide comprise between 1% and 10% modification of the nucleotides or nucleosides.
- between 1% and 10% of the nucleotides or nucleosides in the intervening region are modified.
- between 11% and 10% of the nucleotides or nucleosides in the IRES are modified.
- between 1% and 10% of the nucleotides or nucleosides in the noncoding or coding region are modified. In some embodiments, between 1% and 10% of the nucleotides or nucleosides in the 5 ' intron segment and/or 3 ' intron segment are modified. In some embodiments, between 11% and 10% of the nucleotides or nucleosides in the 5 ' exon segment or post-splicing exon segment and/or 3 ' exon segment or post-splicing exon segment are modified. In some embodiments, between 1% and 10% of the nucleotides or nucleosides in the 5 ' spacer and/or 3 ' spacer are modified. In some embodiments, between 1% and 10% of the nucleotides or nucleosides in the 5 ' homology region and/or 3 ' homology region are modified.
- polynucleotides comprising modified nucleotides and/or modified nucleosides provide additional stability and resistance to immune activation. In some embodiments, polynucleotides comprising modified nucleotides and/or modified nucleosides maintain stability and resistance to immune activation as compared to a corresponding polynucleotide comprising no modified nucleotides and/or modified nucleosides.
- a precursor RNA polynucleotide with modified nucleotides and/or nucleosides improves circularization as compared to a corresponding linear precursor RNA polynucleotide comprising no nucleotide or nucleoside modifications or other appropriate control.
- a precursor RNA polynucleotide with modified nucleotides and/or nucleosides maintains the same circularization as compared to a corresponding precursor RNA polynucleotide comprising no nucleotide or nucleoside modifications or other appropriate control.
- the precursor polynucleotides comprising modified nucleotides and/or modified nucleosides maintain circularization at greater than 50%, greater than 60%, greater than 70%, greater than 80%, greater than 90%, or greater than 100% (i.e., improves circularization), as compared to a corresponding precursor polynucleotide comprising no nucleotide or nucleoside modifications.
- the precursor polynucleotides maintain circularization at greater than 70% as compared to a corresponding precursor polynucleotide comprising no nucleotide or nucleoside modifications.
- the precursor polynucleotides maintain circularization at greater than 80% as compared to a corresponding precursor polynucleotide comprising no nucleotide or nucleoside modifications. In some embodiments, the precursor polynucleotides maintain circularization at greater than 90% as compared to a corresponding precursor polynucleotide comprising no nucleotide or nucleoside modifications. In some embodiments, the precursor polynucleotides exhibit greater than 100% circularization (i.e., improved circularization) as compared to a corresponding precursor polynucleotide comprising no nucleotide or nucleoside modifications.
- a circular RNA with modified nucleotides and/or nucleosides reduces immunogenicity and/or improves translation of the coding region as compared to a corresponding circular RNA comprising no nucleotide or nucleoside modifications.
- a circular RNA polynucleotide with modified nucleotides and/or nucleosides maintains the same immunogenicity and/or translation of the coding region as compared to a corresponding circular RNA comprising no nucleotide or nucleoside modifications.
- the circular RNAs described herein comprising at least one modified A, C, G, or U nucleotide or nucleoside exhibit reduced immunogenicity, without losing circularization and/or translation.
- the circular RNAs described herein exhibit immunogenicity that is reduced by about 10% to about 99%, for example reduced by about 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 99% as compared to a corresponding circular RNA comprising no nucleotide or nucleoside modifications.
- the polynucleotides comprising modified nucleosides provide additional stability and resistance to immune activation.
- portions of the polynucleotide e.g., a precursor RNA polynucleotide, or a circular RNA polynucleotide
- portions of the polynucleotide e.g., a precursor RNA polynucleotide, or a circular RNA polynucleotide, described in more detail elsewhere herein
- the intervening region comprises no nucleotide or nucleoside modifications or less than 10% of the nucleotides or nucleosides therein are modified.
- the IRES comprises no nucleotide or nucleoside modifications or less than 10% of the nucleotides or nucleosides therein are modified.
- the noncoding or coding region comprises no nucleotide or nucleoside modifications or less than 10% of the nucleotides or nucleosides therein are modified.
- the 5 ' intron segment and/or 3 ' intron segment comprises no nucleotide or nucleoside modifications or less than 10% of the nucleotides or nucleosides therein are modified.
- the 5 ' exon segment or post-splicing exon segment and/or 3 ' exon segment or post-splicing exon segment comprises no nucleotide or nucleoside modifications or less than 10% of the nucleotides or nucleosides therein are modified.
- the 5 ' spacer and/or 3 ' spacer comprises no nucleotide or nucleoside modifications or less than 10% of the nucleotides or nucleosides therein are modified.
- the 5 ' homology region, 3 ' homology region comprises no nucleotide or nucleoside modifications or less than 10% of the nucleotides or nucleosides therein are modified.
- the intervening region comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the IRES comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the noncoding or coding region comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the 5 ' intron segment and/or 3 ' intron segment comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the 5 ' exon segment or post-splicing exon segment and/or 3 ' exon segment or post-splicing exon segment comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- modified nucleotides or nucleotides occur throughout a precursor RNA polynucleotide.
- portions of the 3 ' and/or 5 ' intron and/or exon segments in a linear precursor RNA polynucleotide of the present disclosure contain modified nucleotides or nucleosides, but the remaining portions of the linear precursor do not comprise nucleotide or nucleoside modifications.
- portions of the 3 ' and/or 5 ' intron and/or exon segments in a linear precursor RNA polynucleotide of the present disclosure contain modified nucleotides or nucleosides, but the remaining portions of the linear precursor comprise minimal nucleotide or nucleoside modifications.
- portions of the 3 ' and/or 5 ' intron and/or exon segments in a linear precursor RNA polynucleotide of the present disclosure contain modified nucleotides or nucleosides, but the remaining portions of the linear precursor comprise less than 90%, 80%, 70%, 60%, 50%, 40%, 30%, 20%, or 10% modified nucleotides or nucleosides.
- the circular RNA is produced from a linear precursor and where the linear precursor is modified at the 3 ' and/or 5 ' ends only, the circular RNA contains only the modified nucleotide or nucleosides that remain after circularization.
- a circular RNA is prepared by providing modified nucleotides or nucleosides to precursor RNA comprising: a. a 5’ combined accessory element, comprising: i. a 3’ intron segment, ii. a 3’ exon segment, b. an intervening region comprising an internal ribosome entry site (IRES) and a noncoding or coding region, c. a 3’ combined accessory element, comprising: i. a 5’ exon segment, and ii. a 5’ intron segment.
- precursor RNA comprising: a. a 5’ combined accessory element, comprising: i. a 3’ intron segment, ii. a 3’ exon segment, b. an intervening region comprising an internal ribosome entry site (IRES) and a noncoding or coding region, c. a 3’ combined accessory element, comprising: i. a 5’ exon segment, and ii. a 5’ intron segment.
- IRS internal
- a circular RNA is prepared by providing a first and second linear precursor RNA polynucleotide, wherein the first and second linear precursor RNA polynucleotides are capable of forming a circular RNA.
- first precursor or the second precursor but not both precursors comprises at least one modified A, C, G, or U nucleotide or nucleoside.
- the first precursor comprises at least one modified A, C, G, or U nucleotide or nucleoside and the second precursor comprises no modified nucleotides or nucleosides.
- the second precursor comprises least one modified A, C, G, or U nucleotide or nucleoside and the first precursor comprises no modified nucleotides or nucleosides.
- the first precursor comprises a 3 ’ intron fragment of a first intron (Intron 1), a 5 ' intron fragment of a second intron (Intron 2), a translation initiation element, a fragment of a sequence of interest (e.g., coding region), and two exon fragments that correspond with the intron fragments.
- the second precursor comprises a 3 ’ intron fragment of the second intron (Intron 2) and a 5 ' intron fragment of the first intron (Intron 1), a fragment of the sequence of interest of the first precursor, and exon fragments corresponding to those in the first precursor.
- the TIE of the first precursor RNA polynucleotides comprises an IRES. In some embodiments, the first precursor RNA polynucleotide comprises a noncoding or coding region.
- the first and second precursor RNA polynucleotides further comprise spacers and/or homology arms.
- portions of the first linear polynucleotide between 0% and 99%, 1% and 2%, 1% and 3%, 1% and 4%, 1% and 5%, 5% and 6%, 5% and 7%, 5% and 8%, 5% and 9%, 5% and 10%, 10% and 20%, 20% and 30%, 30% and 40%, 40% and 50%, 50% and 60%, 60% and 70%, 70% and 80%, 80% and 90% or 90% and 99% of the nucleotides or nucleosides are modified.
- TIE e.g., IRES
- less than 10% of the nucleotides or nucleosides are modified.
- less than 10% of the nucleotides or nucleosides in the intervening region of the first linear polynucleotide are modified.
- less than 10% of the nucleotides or nucleosides in the TIE (e.g., IRES) of the first linear polynucleotide are modified.
- less than 10% of the nucleotides or nucleosides in the noncoding or coding region of the first linear polynucleotide are modified.
- less than 10% of the nucleotides or nucleosides in the 5 ' and/or 3 ' intron fragment of the first linear polynucleotide are modified. In some embodiments, less than 10% of the nucleotides or nucleosides in the 5 ' and/or 3 ' exon fragment of the first linear polynucleotide are modified. In some embodiments, less than 10% of the nucleotides or nucleosides in the spacer of the first linear polynucleotide are modified. In some embodiments, less than 10% of the nucleotides or nucleosides in the internal and/or external homology region of the first linear polynucleotide are modified.
- the intervening region of the first linear polynucleotide comprises no nucleotide or nucleoside modifications.
- the TIE e.g, IRES
- the noncoding or coding region of the first linear polynucleotide comprises no nucleotide or nucleoside modifications.
- the 5 ' and/or 3 ' intron fragment of the first linear polynucleotide comprises no nucleotide or nucleoside modifications.
- the 5 ' and/or 3 ' exon fragment of the first linear polynucleotide comprises no nucleotide or nucleoside modifications.
- the spacer of the first linear polynucleotide comprises no nucleotide or nucleoside modifications.
- the internal and/or external homology region of the first linear polynucleotide comprises no nucleotide or nucleoside modifications.
- portions of the second linear polynucleotide between 1% and 100%, 1% and 2%, 1% and 3%, 1% and 4%, 1% and 5%, 5% and 6%, 5% and 7%, 5% and 8%, 5% and 9%, 5% and 10%, 10% and 20%, 20% and 30%, 30% and 40%, 40% and 50%, 50% and 60%, 60% and 70%, 70% and 80%, 80% and 90% or 90% and 100% of the nucleotides or nucleosides are modified.
- less than 10% of the nucleotides or nucleosides are modified.
- less than 10% of the nucleotides or nucleosides in the noncoding or coding region of the second linear polynucleotide are modified.
- less than 10% of the nucleotides or nucleosides in the 5 ' and/or 3 ' intron fragment of the second linear polynucleotide are modified.
- less than 10% of the nucleotides or nucleosides in the 5 ' and/or 3 ' exon fragment of the second linear polynucleotide is modified.
- less than 10% of the nucleotides or nucleosides in the spacer of the second linear polynucleotide are modified. In some embodiments, less than 10% of the nucleotides or nucleosides in the internal and/or external homology region of the second linear polynucleotide are modified.
- the noncoding or coding region of the second linear polynucleotide comprises no nucleotide or nucleoside modifications.
- the 5 ' intron fragment and/or 3 ' intron fragment of the second linear polynucleotide comprises no nucleotide or nucleoside modifications.
- the 5 ' exon fragment and/or 3 ' exon fragment of the second linear polynucleotide comprises no nucleotide or nucleoside modifications.
- the spacer of the second linear polynucleotide comprises no nucleotide or nucleoside modifications.
- the internal and/or external homology region of the second linear polynucleotide comprises no nucleotide or nucleoside modifications.
- the intervening region comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the TIE e.g., IRES
- the noncoding or coding region comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the 5 ' and/or 3 ' intron fragment comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the 5 ' and/or 3 ' exon fragment comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the spacer comprises no nucleotide or nucleoside modifications or is less than 10% modified; and/or the internal and/or external homology region comprises no nucleotide or nucleoside modifications or is less than 10% modified.
- the noncoding or coding region comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the 5 ’ intron fragment and/or 3 ’ intron fragment comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the 5 ’ exon fragment and/or 3 ’ exon fragment comprises no nucleotide or nucleoside modifications or is less than 10% modified;
- the spacer comprises no nucleotide or nucleoside modifications or is less than 10% modified; and/or the internal and/or external homology region comprises no nucleotide or nucleoside modifications or is less than 10% modified.
- incorporation of a nucleotide or nucleoside modification to a precursor RNA polynucleotide hinders or lowers the capacity of the circular RNA to circularize, splice, or express.
- the precursor polynucleotide comprising no modified nucleotides and/or nucleosides maintains or improves circularization as compared to a precursor polynucleotide comprising one or more nucleotide or nucleoside modification.
- the precursor polynucleotide comprising no modified nucleotides or nucleosides maintains circularization at greater than 50%, greater than 60%, greater than 70%, greater than 80%, greater than 90%, or greater than 100% (i.e., improves circularization), as compared to a corresponding precursor polynucleotide comprising one or more nucleotide or nucleoside modification.
- the polynucleotides comprising no nucleotide or nucleoside modifications for an example a circular RNA, has comparable or reduced immunogenicity as compared to a polynucleotide comprising one or more nucleotide or nucleoside modification.
- the circular RNAs described herein exhibit immunogenicity that is reduced by about 10% to about 99%, for example reduced by about 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 99% as compared to a corresponding circular RNA comprising one or more nucleotide or nucleoside modifications.
- the polynucleotides comprising no modified nucleotides and/or modified nucleosides maintain or improve translation of a coding region as compared to a corresponding polynucleotide comprising one or more nucleotide or nucleoside modifications.
- the polynucleotides comprising no modified nucleosides provide additional stability and resistance to immune activation.
- the polynucleotide comprising no modified A, C, G, or U nucleotide or nucleoside maintains expression at greater than 50%, greater than 60%, greater than 70%, greater than 80%, greater than 90%, or greater than 100%, as compared to a corresponding precursor polynucleotide comprising one or more nucleotide or nucleoside modifications.
- the non-modified polynucleotides maintain expression at greater than 70% as compared to a corresponding polynucleotide comprising one or more nucleotide or nucleoside modifications.
- the non-modified polynucleotides maintain expression at greater than 80% as compared to a corresponding polynucleotide comprising one or more nucleotide or nucleoside modifications. In some embodiments, the non-modified polynucleotides maintain expression at greater than 90% as compared to a corresponding polynucleotide comprising one or more nucleotide or nucleoside modifications. In some embodiments, the non-modified polynucleotides exhibit greater than 100% expression (i.e., improved expression) as compared to a corresponding polynucleotide comprising one or more nucleotide or nucleoside modifications.
- the non-modified polynucleotides exhibit greater purification efficacy as compared to a corresponding polynucleotide comprising one or more nucleotide or nucleoside modifications.
- the polynucleotide comprising no modified A, C, G, or U nucleotide or nucleoside exhibits greater purification efficacy at greater than 50%, greater than 60%, greater than 70%, greater than 80%, greater than 90%, or greater than 100%, as compared to a corresponding precursor comprising one or more nucleotide or nucleoside modification.
- polynucleotides may be codon-optimized.
- a codon- optimized sequence may be one in which codons in a polynucleotide encoding a polypeptide have been substituted in order to increase the expression, stability and/or activity of the polypeptide.
- Factors that influence codon optimization include, but are not limited to one or more of: (i) variation of codon biases between two or more organisms or genes or synthetically constructed bias tables, (ii) variation in the degree of codon bias within an organism, gene, or set of genes, (iii) systematic variation of codons including context, (iv) variation of codons according to their decoding tRNAs, (v) variation of codons according to GC %, either overall or in one position of the triplet, (vi) variation in degree of similarity to a reference sequence for example a naturally occurring sequence, (vii) variation in the codon frequency cutoff, (viii) structural properties of mRNAs transcribed from the DNA sequence, (ix) prior knowledge about the function of the DNA sequences upon which design of the codon substitution set is to be based, and/or (x) systematic variation of codon sets for each amino acid.
- a codon optimized polynucleotide may minimize ribozyme collisions and/or limit
- a provided polynucleotide (e.g., a DNA template, a linear precursor RNA polynucleotide, or a circular RNA polynucleotide) comprises one or more expression sequences or portions thereof.
- the precursor RNA polynucleotide and circular RNA constructs comprise at least one expression sequence encoding a binding molecule.
- the precursor RNA polynucleotide and the circular RNA constructs comprise at least one expression sequence encoding a therapeutic protein and an IRES, wherein the IRES can facilitate expression of the protein when delivered in vivo.
- the coding (or non-coding region) is a part of the intervening region or core functional element located in between the 5’ end and 3’ end of a linear precursor RNA polynucleotide and resultant circular RNA.
- the precursor RNA polynucleotide and circular RNA may encode for various therapeutic proteins, cytokines, immune checkpoint inhibitors, agonists, chimeric antigen receptors, inhibitory receptor agonists, one or more T-Cell Receptors, and/or B- cell Receptors that are available in the art.
- the chimeric proteins may also include, for example, recombinant fusion proteins, chimeric mutant protein, or other fusion proteins.
- the circular RNA comprises more than 1 expression sequence, e.g., 2, 3,
- the circular RNA is a bicistronic RNA.
- the bicistronic RNA is codon optimized. Exemplary bicistronic circular RNA are described in WO2021/189059A2, which is incorporated by reference herein in its entirety.
- the precursor RNA polynucleotide and circular RNA constructs comprise at least one expression sequence encoding an antigen, adjuvant, or adjuvant-like protein, e.g., from an infectious agent.
- the circular RNA construct may be used as a vaccine.
- the expression sequence encodes a therapeutic protein.
- therapeutic proteins are listed in Table 2.
- the scFv, heavy variable domain, light variable domain, heavy CDR sequences, and/or light CDR sequences of the therapeutic proteins listed in Table 2 may be used.
- the therapeutic protein is selected from a CD19-targted chimeric antigen receptor (CAR), a BCMA-targeted CAR, MAGE-A4 T-cell receptor (TCR), NY-ESO TCR, erythropoietin (EPO), phenylalanine hydroxylase (PAH), carbamoyl phosphate synthetase I (CPS1), Cas9, ADAMTS13, FOXP3, IL-10, or IL-2.
- CAR CD19-targted chimeric antigen receptor
- BCMA-targeted CAR MAGE-A4 T-cell receptor (TCR), NY-ESO TCR
- EPO erythropoietin
- PAH phenylalanine hydroxylase
- CPS1 carbamoyl phosphate synthetase I
- Cas9 Cas9
- ADAMTS13 ADAMTS13
- FOXP3 IL-10
- IL-2 IL-2
- the target amino acid has an amino acid sequence that is identical to or at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identical to a sequence selected from SEQ ID NOS: 4019-4040, or a fragment thereof.
- the expression sequence encodes a therapeutic protein.
- the expression sequence encodes a cytokine, e.g., IL-12p70, IL- 15, IL-2, IL-18, IL-21, IFN-a, IFN- P, IL-10, TGF-beta, IL-4, or IL-35, or a functional fragment thereof.
- the expression sequence encodes an immune checkpoint inhibitor.
- the expression sequence encodes an agonist (e.g., a TNFR family member such as CD137L, OX40L, ICOSL, LIGHT, or CD70).
- the expression sequence encodes a chimeric antigen receptor.
- the expression sequence encodes an inhibitory receptor agonist (e.g., PDL1, PDL2, Galectin-9, VISTA, B7H4, or MHCII) or inhibitory receptor (e g., PD1, CTLA4, TIGIT, LAG3, or TIM3).
- the expression sequence encodes an inhibitory receptor antagonist.
- the expression sequence encodes one or more TCR chains (alpha and beta chains or gamma and delta chains).
- the expression sequence encodes a secreted T cell or immune cell engager (e.g., a bispecific antibody such as BiTE, targeting, e.g., CD3, CD137, or CD28 and a tumor-expressed protein e.g., CD19, CD20, or BCMA etc.).
- the expression sequence encodes a transcription factor (e.g., FOXP3, HELIOS, T0X1, or T0X2).
- the expression sequence encodes an immunosuppressive enzyme (e.g., IDO or CD39/CD73).
- the expression sequence encodes a GvHD (e.g., anti -HL A- A2 CAR-Tregs).
- a provided polynucleotide encodes a protein that is made up of subunits that are encoded by more than one gene.
- the protein may be a heterodimer, wherein each chain or subunit of the protein is encoded by a separate gene.
- more than one provided polynucleotides e.g., circular RNA polynucleotides
- polynucleotides encoding the individual subunits may be administered in separate transfer vehicles.
- a single polynucleotide e.g., circular RNA polynucleotide
- a provided polynucleotide encodes one or more chimeric antigen receptors (CARs).
- CARs are genetically-engineered receptors. These engineered receptors may be inserted into and expressed by immune cells, including T cells via circular RNA as described herein. With a CAR, a single receptor may be programmed to both recognize a specific antigen and, when bound to that antigen, activate the immune cell to attack and destroy the cell bearing that antigen. When these antigens exist on tumor cells, an immune cell that expresses the CAR may target and kill the tumor cell.
- the CAR encoded by the polynucleotide comprises (i) an antigen-binding molecule that specifically binds to a target antigen, (ii) a hinge domain, a transmembrane domain, and an intracellular domain, and (iii) an activating domain.
- an orientation of the CARs in accordance with the disclosure comprises an antigen binding domain (such as an scFv) in tandem with a costimulatory domain and an activating domain.
- the costimulatory domain may comprise one or more of an extracellular portion, a transmembrane portion, and an intracellular portion. In other embodiments, multiple costimulatory domains may be utilized in tandem. i. Antigen binding domain
- CARs may be engineered to bind to an antigen (such as a cell-surface antigen) by incorporating an antigen binding molecule that interacts with that targeted antigen.
- the antigen binding molecule is an antibody fragment thereof, e.g., one or more single chain antibody fragment (scFv).
- scFv is a single chain antibody fragment having the variable regions of the heavy and light chains of an antibody linked together. See U.S. Patent Nos. 7,741,465, and 6,319,494, as well as Eshhar et al., Cancer Immunol Immunotherapy (1997) 45: 131-136.
- An scFv retains the parent antibody's ability to specifically interact with target antigen.
- scFvs are useful in chimeric antigen receptors because they may be engineered to be expressed as part of a single chain along with the other CAR components. Id. See also Krause et al., J. Exp. Med., Volume 188, No. 4, 1998 (619-626); Finney et al., Journal of Immunology, 1998, 161 : 2791-2797. It will be appreciated that the antigen binding molecule is typically contained within the extracellular portion of the CAR such that it is capable of recognizing and binding to the antigen of interest. Bispecific and multispecific CARs are contemplated within the scope of the disclosure, with specificity to more than one target of interest.
- the antigen binding molecule comprises a single chain, wherein the heavy chain variable region and the light chain variable region are connected by a linker.
- the VH is located at the N terminus of the linker and the VL is located at the C terminus of the linker. In other embodiments, the VL is located at the N terminus of the linker and the VH is located at the C terminus of the linker.
- the linker comprises at least about 5, at least about 8, at least about 10, at least about 13, at least about 15, at least about 18, at least about 20, at least about 25, at least about 30, at least about 35, at least about 40, at least about 45, at least about 50, at least about 60, at least about 70, at least about 80, at least about 90, or at least about 100 amino acids.
- the antigen binding molecule comprises a nanobody. In some embodiments, the antigen binding molecule comprises a DARPin. In some embodiments, the antigen binding molecule comprises an anticalin or other synthetic protein capable of specific binding to target protein.
- the CAR comprises an antigen binding domain specific for an antigen selected from the group CD 19, CD 123, CD22, CD30, CD171, CS-1, C-type lectin- like molecule- 1, CD33, epidermal growth factor receptor variant III (EGFRvIII), ganglioside G2 (GD2), ganglioside GD3, TNF receptor family member B cell maturation (BCMA), Tn antigen ((Tn Ag) or (GalNAca-Ser/Thr)), prostate-specific membrane antigen (PSMA), Receptor tyrosine kinase-like orphan receptor 1 (R0R1), Fms-Like Tyrosine Kinase 3 (FLT3), Tumor-associated glycoprotein 72 (TAG72), CD38, CD44v6, Carcinoembryonic antigen (CEA), Epithelial cell adhesion molecule (EPCAM), B7H3 (CD276), KIT (CD 117), Interleukin- 13 receptor subunit al
- melanoma cancer testis antigen-2 MAD-CT-2
- Fos-related antigen tumor protein p53 (p53), p53 mutant, prostein, surviving, telomerase, prostate carcinoma tumor antigen- 1, melanoma antigen recognized by T cells 1, Rat sarcoma (Ras) mutant, human Telomerase reverse transcriptase (hTERT), sarcoma translocation breakpoints, melanoma inhibitor of apoptosis (ML-IAP), ERG (transmembrane protease, serine 2 (TMPRSS2) ETS fusion gene), N-Acetyl glucosaminyl-transferase V (NA17), paired box protein Pax-3 (PAX3), Androgen receptor, Cyclin Bl, v-myc avian myelocytomatosis viral oncogene neuroblastoma derived homolog (MYCN), Ras Homolog Family Member C (Rho
- EMR2 lymphocyte antigen 75
- Glypican-3 Glypican-3
- FCRL5 Fc receptor-like 5
- MUC16 5T4, 8H9, avP0 integrin, avP6 integrin, alphafetoprotein (AFP), B7-H6, ca-125, CA9, CD44, CD44v7/8, CD52, E-cadherin, EMA (epithelial membrane antigen), epithelial glycoprotein-2 (EGP-2), epithelial glycoprotein-40 (EGP-40), ErbB4, epithelial tumor antigen (ETA), folate binding protein (FBP), kinase insert domain receptor (KDR), k-light chain, LI cell adhesion molecule, MUC18, NKG2D, oncofetal antigen (h5T4), tumor/testis-antigen IB, GAGE, GAGE-1, BAGE, SCP-1, CTZ9, SAGE, CAGE, CT 10, MART-1, immunoglobulin lambd
- the circular RNA constructs and related pharmaceutical compositions comprise the expression sequences described in Table 2, above.
- the circular RNA constructs and related pharmaceutical compositions disclosed herein comprise an expression sequence that is at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identical to a sequence in Table 2, wherein the codon sequence produces a protein having the desired sequence. ii. Hinge / spacer domain
- a CAR of the instant disclosure comprises a hinge or spacer domain.
- the hinge/spacer domain may comprise a truncated hinge/spacer domain (THD) the THD domain is a truncated version of a complete hinge/spacer domain (“CHD”).
- THD truncated hinge/spacer domain
- CHD complete hinge/spacer domain
- an extracellular domain is from or derived from (e.g., comprises all or a fragment of) ErbB2, glycophorin A (GpA), CD2, CD3 delta, CD3 epsilon, CD3 gamma, CD4, CD7, CD8a, CD8[T CD1 la (IT GAL), CD1 lb (IT GAM), CD1 1c (ITGAX), CD1 Id (IT GAD), CD 18 (ITGB2), CD 19 (B4), CD27 (TNFRSF7), CD28, CD28T, CD29 (ITGB1), CD30 (TNFRSF8), CD40 (TNFRSF5), CD48 (SLAMF2), CD49a (ITGA1), CD49d (ITGA4), CD49f (ITGA6), CD66a (CEACAM1), CD66b (CEACAM8), CD66c (CEACAM6), CD66d (CEACAM3), CD66e (CEACAM5), CD69 (CLEC2), CD79A (B-cell
- a hinge or spacer domain is positioned between an antigen binding molecule (e.g., an scFv) and a transmembrane domain. In this orientation, the hinge/spacer domain provides distance between the antigen binding molecule and the surface of a cell membrane on which the CAR is expressed.
- a hinge or spacer domain is from or derived from an immunoglobulin.
- a hinge or spacer domain is selected from the hinge/spacer regions of IgGl, IgG2, IgG3, IgG4, IgA, IgD, IgE, IgM, or a fragment thereof.
- a hinge or spacer domain comprises, is from, or is derived from the hinge/spacer region of CD8 alpha. In some embodiments, a hinge or spacer domain comprises, is from, or is derived from the hinge/spacer region of CD28. In some embodiments, a hinge or spacer domain comprises a fragment of the hinge/spacer region of CD8 alpha or a fragment of the hinge/spacer region of CD28, wherein the fragment is anything less than the whole hinge/spacer region.
- the fragment of the CD8 alpha hinge/spacer region or the fragment of the CD28 hinge/spacer region comprises an amino acid sequence that excludes at least 1, at least 2, at least 3, at least 4, at least 5, at least 6, at least 7, at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, at least 19, or at least 20 amino acids at the N- terminus or C-Terminus, or both, of the CD8 alpha hinge/spacer region, or of the CD28 hinge/spacer region.
- the CAR of the present disclosure may further comprise a transmembrane domain and/or an intracellular signaling domain.
- the transmembrane domain may be designed to be fused to the extracellular domain of the CAR. It may similarly be fused to the intracellular domain of the CAR.
- the transmembrane domain that naturally is associated with one of the domains in a CAR is used.
- the transmembrane domain may be selected or modified (e.g., by an amino acid substitution) to avoid binding of such domains to the transmembrane domains of the same or different surface membrane proteins to minimize interactions with other members of the receptor complex.
- the transmembrane domain may be derived either from a natural or from a synthetic source. Where the source is natural, the domain may be derived from any membrane-bound or transmembrane protein.
- Transmembrane regions may be derived from (i.e. comprise) a receptor tyrosine kinase e.g., ErbB2), glycophorin A (GpA), 4-1BB/CD137, activating NK cell receptors, an immunoglobulin protein, B7-H3, BAFFR, BFAME (SEAMF8), BTEA, CD100 (SEMA4D), CD103, CD160 (BY55), CD18, CD19, CD19a, CD2, CD247, CD27, CD276 (B7-H3), CD28, CD29, CD3 delta, CD3 epsilon, CD3 gamma, CD30, CD4, CD40, CD49a, CD49D, CD49f, CD69, CD7, CD84, CD8alpha, CD8beta, CD96 (Tactile), CD1 la, CD1 lb, CD1 1c, CD1 Id, CDS, CEACAM1, CRT AM, cytokine receptor,
- suitable intracellular signaling domain include, but are not limited to, activating Macrophage/Myeloid cell receptors CSFR1, MYD88, CD14, TIE2, TLR4, CR3, CD64, TREM2, DAP10, DAP12, CD169, DECTIN1, CD206, CD47, CD163, CD36, MARCO, TIM4, MERTK, F4/80, CD91, C1QR, LOX-1, CD68, SRA, BALI, ABCA7, CD36, CD31, Lactoferrin, or a fragment, truncation, or combination thereof.
- a receptor tyrosine kinase may be derived from (e.g., comprise) Insulin receptor (InsR), Insulin-like growth factor I receptor (IGF1R), Insulin receptor-related receptor (IRR), platelet derived growth factor receptor alpha (PDGFRa), platelet derived growth factor receptor beta (PDGFRfi).
- Insulin receptor Insulin receptor
- IGF1R Insulin-like growth factor I receptor
- IRR Insulin receptor-related receptor
- PDGFRa platelet derived growth factor receptor alpha
- PDGFRfi platelet derived growth factor receptor beta
- KIT proto-oncogene receptor tyrosine kinase Kit
- colony stimulating factor 1 receptor CSFR
- fms related tyrosine kinase 3 FLT3
- fms related tyrosine kinase 1 VFGFR-1
- kinase insert domain receptor VAGFR-2
- fms related tyrosine kinase 4 VGFR-3
- FGFR1 fibroblast growth factor receptor 1
- FGFR2 fibroblast growth factor receptor 2
- FGFR3 fibroblast growth factor receptor 4
- FGFR4 protein tyrosine kinase 7
- trkA neurotrophic receptor tyrosine kinase 1
- trkB neurotrophic receptor tyrosine kinase 2
- trkC neurotrophic receptor tyrosine kinase like orphan receptor
- the CAR comprises a costimulatory domain.
- the costimulatory domain comprises 4-1BB (CD137), CD28, or both, and/or an intracellular T cell signaling domain.
- the costimulatory domain is human CD28, human 4-1BB, or both, and the intracellular T cell signaling domain is human CD3 zeta (Q. 4-1BB, CD28, CD3 zeta may comprise less than the whole 4-1BB, CD28 or CD3 zeta, respectively.
- Chimeric antigen receptors may incorporate costimulatory (signaling) domains to increase their potency. See U.S. Patent Nos.
- a costimulatory domain comprises the amino acid sequence of SEQ ID NO: 4041 (KRGRKKLLYIFKQPFMRPVQTTQEEDGCSCRFPEEEEGGCEL) or 4042
- the intracellular (signaling) domain of the engineered T cells disclosed herein may provide signaling to an activating domain, which then activates at least one of the normal effector functions of the immune cell.
- Effector function of a T cell for example, may be cytolytic activity or helper activity including the secretion of cytokines.
- suitable intracellular signaling domain comprise, but are not limited to 4-1BB/CD137, activating NK cell receptors, an Immunoglobulin protein, B7-H3, BAFFR, BLAME (SLAMF8), BTLA, CD100 (SEMA4D), CD103, CD160 (BY55), CD18, CD19, CD 19a, CD2, CD247, CD27, CD276 (B7-H3), CD28, CD29, CD3 delta, CD3 epsilon, CD3 gamma, CD30, CD4, CD40, CD49a, CD49D, CD49f, CD69, CD7, CD84, CD8alpha, CD8beta, CD96 (Tactile), CD1 la, CD1 lb, CD1 1c, CD1 Id, CDS, CEACAM1, CRT AM, cytokine receptor, DAP-10, DNAM1 (CD226), Fc gamma receptor, GADS, GITR, HVEM (
- CD3 is an element of the T cell receptor on native T cells, and has been shown to be an important intracellular activating element in CARs.
- the CD3 is CD3 zeta.
- the activating domain comprises an amino acid sequence at least about 60%, at least about 65%, at least about 70%, at least about 75%, at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 96%, at least about 97%, at least about 98%, at least about 99%, or about 100% identical to the sequence of SEQ ID NO: 4043
- TCR T-CELL RECEPTORS
- a provided polynucleotide encodes a T-cell receptor.
- TCRs are described using the International Immunogenetics (IMGT) TCR nomenclature, and links to the IMGT public database of TCR sequences.
- IMGT International Immunogenetics
- Native alpha-beta heterodimeric TCRs have an alpha chain and a beta chain. Broadly, each chain may comprise variable, joining and constant regions, and the beta chain also usually contains a short diversity region between the variable and joining regions, but this diversity region is often considered as part of the joining region.
- Each variable region may comprise three CDRs (Complementarity Determining Regions) embedded in a framework sequence, one being the hypervariable region named CDR3.
- CDRs Complementarity Determining Regions
- the Va types are referred to in IMGT nomenclature by a unique TRAV number.
- TRAV21 defines a TCR Va region having unique framework and CDR1 and CDR2 sequences, and a CDR3 sequence which is partly defined by an amino acid sequence which is preserved from TCR to TCR but which also includes an amino acid sequence which varies from TCR to TCR.
- TRBV5-1 defines a TCR VP region having unique framework and CDR1 and CDR2 sequences, but with only a partly defined CDR3 sequence.
- the joining regions of the TCR are similarly defined by the unique IMGT TRAJ and TRBJ nomenclature, and the constant regions by the IMGT TRAC and TRBC nomenclature.
- the beta chain diversity region is referred to in IMGT nomenclature by the abbreviation TRBD, and, as mentioned, the concatenated TRBD/TRBJ regions are often considered together as the joining region.
- TRBD abbreviation
- TRBJ concatenated TRBD/TRBJ regions are often considered together as the joining region.
- the unique sequences defined by the IMGT nomenclature are widely known and accessible to those working in the TCR field. For example, they can be found in the IMGT public database.
- the “T cell Receptor Factsbook”, (2001) LeFranc and LeFranc, Academic Press, ISBN 0-12-441352-8 also discloses sequences defined by the IMGT nomenclature, but because of its publication date and consequent time-lag, the information therein sometimes needs to be confirmed by reference to the IMGT database.
- TCRs exist in heterodimeric a[3 or y5 forms. However, recombinant TCRs consisting of aa or PP homodimers have previously been shown to bind to peptide MHC molecules. Therefore, the TCR of the present disclosure may be a heterodimeric aP TCR or may be an aa or PP homodimeric TCR.
- an aP heterodimeric TCR may, for example, be transfected as full length chains having both cytoplasmic and transmembrane domains.
- TCRs of the present disclosure may have an introduced disulfide bond between residues of the respective constant domains, as described, for example, in WO 2006/000830.
- TCRs of the present disclosure may comprise an alpha chain TRAC constant domain sequence and/or a beta chain TRBC1 or TRBC2 constant domain sequence.
- the alpha and beta chain constant domain sequences may be mutated by truncation or substitution to delete the native disulfide bond between Cys4 of exon 2 of TRAC and Cys2 of exon 2 of TRBC1 or TRBC2.
- the alpha and/or beta chain constant domain sequence(s) may also be mutated by substitution of cysteine residues for Thr 48 of TRAC and Ser 57 of TRBC1 or TRBC2, the said cysteines forming a disulfide bond between the alpha and beta constant domains of the TCR.
- Binding affinity (inversely proportional to the equilibrium constant KD) and binding half-life (expressed as T’ ) can be determined by any appropriate method. It will be appreciated that doubling the affinity of a TCR results in halving the KD. T’ is calculated as In 2 divided by the off-rate (koff). So doubling of T’ results in a halving in koff. KD and koff values for TCRs are usually measured for soluble forms of the TCR, i.e. those forms which are truncated to remove cytoplasmic and transmembrane domain residues.
- a given TCR has an improved binding affinity for, and/or a binding half-life for the parental TCR if a soluble form of that TCR has the said characteristics.
- the binding affinity or binding half-life of a given TCR is measured several times, for example, 3 or more times, using the same assay protocol, and an average of the results is taken.
- the disclosure includes a non-naturally occurring and/or purified and/or or engineered cell, especially a T-cell, presenting a TCR of the present disclosure.
- T cells expressing the TCRs will be suitable for use in adoptive therapy-based treatment of cancers such as those of the pancreas and liver.
- suitable methods by which adoptive therapy can be carried out see for example Rosenberg et al., (2008) Nat Rev Cancer 8(4): 299-308).
- TCRs of the present disclosure may be subject to post- translational modifications when expressed by transfected cells.
- Glycosylation is one such modification, which may comprise the covalent attachment of oligosaccharide moieties to defined amino acids in the TCR chain.
- asparagine residues, or serine/threonine residues are well-known locations for oligosaccharide attachment.
- the glycosylation status of a particular protein depends on a number of factors, including protein sequence, protein conformation and the availability of certain enzymes. Furthermore, glycosylation status (i.e., oligosaccharide type, covalent linkage and total number of attachments) can influence protein function.
- Glycosylation of transfected TCRs may be controlled by mutations of the transfected gene (Kuball J et al. (2009), J Exp Med 206(2):463-475). Such mutations are also encompassed herein.
- a TCR may be specific for an antigen in the group MAGE-A1 , MAGE-A2, MAGE- A3, MAGE-A4, MAGE-A5, MAGE-A6, MAGE-A7, MAGE-A8, MAGE-A9, MAGE-A10, MAGE-A11, MAGE-A12, MAGE-A13, GAGE-1, GAGE-2, GAGE-3, GAGE-4, GAGE-5, GAGE-6, GAGE-7, GAGE-8, BAGE-1, RAGE-1, LB33/MUM-1, PRAME, NAG, MAGE- Xp2 (MAGE-B2), MAGE-Xp3 (MAGE-B3), MAGE-Xp4 (AGE-B4), tyrosinase, brain glycogen phosphorylase, Melan-A, MAGE-CI, MAGE-C2, NY-ESO-1, LAGE-1, SSX-1, SSX-2(HOM-MEL-40), SSX-1, SSX-4, SSX
- a provided polynucleotide encodes one or more B-cell receptors (BCRs).
- BCRs (or B-cell antigen receptors) are immunoglobulin molecules that form a type I transmembrane protein on the surface of a B cell.
- a BCR is capable of transmitting activatory signal into a B cell following recognition of a specific antigen. Prior to binding of a B cell to an antigen, the BCR will remain in an unstimulated or “resting” stage. Binding of an antigen to a BCR leads to signaling that initiates a humoral immune response.
- a BCR is expressed by mature B cells. These B cells work with immunoglobulins (Igs) in recognizing and tagging pathogens.
- the typical BCR comprises a membrane-bound immunoglobulin (e.g., mlgA, mlgD, mlgE, mlgG, and mlgM), along with associated and Iga/IgP (CD79a/CD79b) heterodimers (a/p).
- membrane-bound immunoglobulins are tetramers consisting of two identical heavy and two light chains.
- the membrane bound immunoglobulins is capable of responding to antigen binding by signal transmission across the plasma membrane leading to B cell activation and consequently clonal expansion and specific antibody production (Friess M et al. (2016), Front. Immunol. 2947(9)).
- the Iga/IgP heterodimers is responsible for transducing signals to the cell interior.
- a Iga/IgP heterodimer signaling relies on the presence of immunoreceptor tyrosinebased activation motifs (ITAMs) located on each of the cytosolic tails of the heterodimers.
- ITAMs comprise two tyrosine residues separated by 9-12 amino acids e.g., tyrosine, leucine, and/or valine).
- tyrosine of the BCR’s ITAMs become phosphorylated by Src-family tyrosine kinases Blk, Fyn, or Lyn (Janeway C et al., Immunobiology: The Immune System in Health and Disease (Garland Science, 5th ed. 2001)).
- a provided polynucleotide e.g., a DNA template, a linear precursor RNA polynucleotide, or a circular RNA polynucleotide
- the chimeric proteins may include recombinant fusion proteins, chimeric mutant protein, or other fusion proteins.
- a provided polynucleotide encodes for an immune modulatory ligand.
- the immune modulatory ligand may be immunostimulatory; while in other embodiments, the immune modulatory ligand may be immunosuppressive .
- the circular RNA polynucleotide encodes for a cytokine or a functional fragment thereof, including but not limited to interferons, chemokines, interleukins, growth factors, and other cytokines known in the art.
- the cytokine comprises a chemokine, interferon, interleukin, lymphokine, and/or tumor necrosis factor.
- Chemokines are chemotactic cytokine produced by a variety of cell types in acute and chronic inflammation that mobilizes and activates white blood cells.
- An interferon comprises a family of secreted a-helical cytokines induced in response to specific extracellular molecules through stimulation of TLRs (Borden, Molecular Basis of Cancer (Fourth Edition) 2015). Interleukins are cytokines expressed by leukocytes.
- Regulatory T cells are important in maintaining homeostasis, controlling the magnitude and duration of the inflammatory response, and in preventing autoimmune and allergic responses.
- Tregs are thought to be mainly involved in suppressing immune responses, functioning in part as a “self-check” for the immune system to prevent excessive reactions.
- Tregs are involved in maintaining tolerance to self-antigens, harmless agents such as pollen or food, and abrogating autoimmune disease.
- Tregs are found throughout the body including, without limitation, the gut, skin, lung, and liver. Additionally, Treg cells may also be found in certain compartments of the body that are not directly exposed to the external environment such as the spleen, lymph nodes, and even adipose tissue. Each of these Treg cell populations is known or suspected to have one or more unique features and additional information may be found in Lehtimaki and Lahesmaa, Regulatory T cells control immune responses through their non-redundant tissue specific features, 2013, FRONTIERS IN IMMUNOL., 4(294): 1-10, the disclosure of which is hereby incorporated in its entirety.
- Tregs are known to require TGF-P and IL-2 for proper activation and development.
- Tregs expressing abundant amounts of the IL-2 receptor (IL-2R), are reliant on IL-2 produced by activated T cells.
- Tregs are known to produce both IL-10 and TGF-P, both potent immune suppressive cytokines.
- Tregs are known to inhibit the ability of antigen presenting cells (APCs) to stimulate T cells.
- APCs antigen presenting cells
- CTLA-4 may bind to B7 molecules on APCs and either block these molecules or remove them by causing internalization resulting in reduced availability of B7 and an inability to provide adequate costimulation for immune responses. Additional discussion regarding the origin, differentiation and function of Tregs may be found in Dhamne et al., Peripheral and thymic Foxp3+ regulatory T cells in search of origin, distinction, and function, 2013, Frontiers in Immunol., 4 (253): 1- 11, the disclosure of which is hereby incorporated in its entirety.
- a provided polynucleotide e.g., a DNA template, a linear precursor RNA polynucleotide, or a circular RNA polynucleotide encodes one or more checkpoint inhibitors or agonists.
- the immune checkpoint inhibitor is an inhibitor of Programmed Death-Ligand 1 (PD-L1, also known as B7-H1, CD274), Programmed Death 1 (PD-1), CTLA-4, PD-L2 (B7-DC, CD273), LAG3, TIM3, 2B4, A2aR, B7H1, B7H3, B7H4, BTLA, CD2, CD27, CD28, CD30, CD40, CD70, CD80, CD86, CD 137, CD 160, CD226, CD276, DR3, GAL9, GITR, HAVCR2, HVEM, IDO1, IDO2, ICOS (inducible T cell costimulator), KIR, LAIR1, LIGHT, MARCO (macrophage receptor with collageneous structure), PS (phosphatidylserine), OX-40, SLAM, TIGHT, VISTA, VTCN1, or any combinations thereof.
- PD-L1 Programmed Death-Ligand 1
- PD-1 Programmed Death 1
- the immune checkpoint inhibitor is an inhibitor of IDO1, CTLA4, PD-1, LAG3, PD-L1, TIM3, or combinations thereof. In some embodiments, the immune checkpoint inhibitor is an inhibitor of PD-L1. In some embodiments, the immune checkpoint inhibitor is an inhibitor of PD-1. In some embodiments, the immune checkpoint inhibitor is an inhibitor of CTLA-4. In some embodiments, the immune checkpoint inhibitor is an inhibitor of LAG3. In some embodiments, the immune checkpoint inhibitor is an inhibitor of TIM3. In some embodiments, the immune checkpoint inhibitor is an inhibitor of IDOL
- the disclosure encompasses the use of immune checkpoint antagonists.
- immune checkpoint antagonists include antagonists of immune checkpoint molecules such as Cytotoxic T-Lymphocyte Antigen 4 (CTLA-4), Programmed Cell Death Protein 1 (PD-1), Programmed Death-Ligand 1 (PDL-1), Lymphocyte- activation gene 3 (LAG-3), and T-cell immunoglobulin and mucin domain 3 (TIM-3).
- CTLA-4 Cytotoxic T-Lymphocyte Antigen 4
- PD-1 Programmed Cell Death Protein 1
- PDL-1 Programmed Death-Ligand 1
- LAG-3 Lymphocyte- activation gene 3
- TIM-3 T-cell immunoglobulin and mucin domain 3
- An antagonist of CTLA-4, PD-1, PDL-1, LAG-3, or TIM-3 interferes with CTLA-4, PD-1, PDL-1, LAG-3, or TIM-3 function, respectively.
- Such antagonists of CTLA-4, PD-1, PDL-1, LAG-3, and TIM-3 can include antibodies which specifically bind to CTLA-4, PD-1, PDL-1, LAG-3, and TIM-3, respectively and inhibit and/or block biological activity and function.
- the payload encoded within one or more of the coding elements is a hormone, FC fusion protein, anticoagulant, blood clotting factor, protein associated with deficiencies and genetic disease, a chaperone protein, an antimicrobial protein, an enzyme (e.g., metabolic enzyme), a structural protein (e.g., a channel or nuclear pore protein), protein variant, small molecule, antibody, nanobody, an engineered non-body antibody, or a combination thereof.
- the DNA templates provided herein can be made using standard techniques of molecular biology.
- the various elements of the vectors provided herein can be obtained using recombinant methods, such as by screening cDNA and genomic libraries from cells, or by deriving the polynucleotides from a DNA template known to include the same.
- the various elements of the DNA template provided herein can also be produced synthetically, rather than cloned, based on the known sequences.
- the complete sequence can be assembled from overlapping oligonucleotides prepared by standard methods and assembled into the complete sequence.
- nucleotide sequences can be obtained from DNA template harboring the desired sequences or synthesized completely, or in part, using various oligonucleotide synthesis techniques known in the art, such as site-directed mutagenesis and polymerase chain reaction (PCR) techniques where appropriate.
- oligonucleotide synthesis techniques known in the art, such as site-directed mutagenesis and polymerase chain reaction (PCR) techniques where appropriate.
- PCR polymerase chain reaction
- One method of obtaining nucleotide sequences encoding the desired DNA template elements is by annealing complementary sets of overlapping synthetic oligonucleotides produced in a conventional, automated polynucleotide synthesizer, followed by ligation with an appropriate DNA ligase and amplification of the ligated nucleotide sequence via PCR. See, e.g., Jayaraman et al., Proc. Natl.
- oligonucleotide-directed synthesis Jones et al., Nature (1986) 54:75-82
- oligonucleotide directed mutagenesis of preexisting nucleotide regions Riechmann et al., Nature (1988) 332:323-327 and Verhoeyen et al., Science (1988) 239: 1534-1536
- enzymatic filling-in of gapped oligonucleotides using T4 DNA polymerase Queen et al., Proc. Natl. Acad. Sci. USA (1989) 86: 10029-10033
- a DNA template e.g., comprising a 3’ intron element, 3’ exon element, an intervening region or core functional element including an IRES and expression sequence, a 5’ exon element, and a 5’ intron element
- this DNA template comprises a vector, PCR product, plasmid, minicircle DNA, cosmid, artificial chromosome, complementary DNA (cDNA), extrachromosomal DNA (ecDNA), or a fragment therein.
- the minicircle DNA may be linearized or non-linearized.
- the plasmid may be linearized or non-linearized.
- the DNA template may be single-stranded. In other embodiments, the DNA template may be double-stranded. In some embodiments, the DNA template comprises in whole or in part from a viral, bacterial or eukaryotic vector. In some embodiments, the polynucleotide of the present disclosure is an expression vector.
- the precursor RNA provided herein can be generated by incubating a DNA template provided herein under conditions permissive of transcription of the precursor RNA encoded by the DNA template.
- a precursor RNA is synthesized by incubating a DNA template provided herein that comprises an RNA polymerase promoter or promoter segment upstream of its 5’ duplex sequence and/or expression sequences with a compatible RNA polymerase enzyme under conditions permissive of in vitro transcription.
- the DNA template is incubated inside of a cell by a bacteriophage RNA polymerase or in the nucleus of a cell by host RNA polymerase II.
- the polynucleotide of the present disclosure is an expression vector, wherein the expression vector comprises a polymerase promoter sequence or segment.
- RNA template provided herein as a template (e.g., a vector provided herein with an RNA polymerase promoter or promoter segment positioned upstream of the 5’ duplex region).
- the DNA template shares the same sequence as the precursor linear RNA polynucleotide prior to splicing of the precursor linear RNA polynucleotide (e.g., a 3’ intron element, a 3’ exon element, an intervening region core functional element, and a 5’ exon element, a 5’ intron element).
- said linear precursor RNA polynucleotide undergoes splicing leading to the removal of the 3’ intron element and 5’ intron element during the process of circularization.
- the resulting circular RNA polynucleotide lacks a 3’ intron fragment and a 5’ intron fragment, but maintains a 3’ exon fragment, an intervening region or a core functional element, and a 5’ exon element.
- the resulting precursor RNA can be used to generate circular RNA (e.g., a circular RNA polynucleotide provided herein) by incubating it in the presence of magnesium ions and guanosine nucleotide or nucleoside at a temperature at which RNA circularization occurs (e.g., between 20 °C and 60 °C).
- circular RNA e.g., a circular RNA polynucleotide provided herein
- a temperature at which RNA circularization occurs e.g., between 20 °C and 60 °C.
- the method comprises synthesizing precursor RNA by transcription (e.g., run-off transcription) using a vector provided herein (e.g., a 5’ intron element, a 5’ exon element, an intervening region or core functional element, a 3’ exon element, and a 3’ intron element) as a template, and incubating the resulting precursor RNA in the presence of divalent cations (e.g., magnesium ions) and GTP such that it circularizes to form circular RNA.
- a vector provided herein e.g., a 5’ intron element, a 5’ exon element, an intervening region or core functional element, a 3’ exon element, and a 3’ intron element
- divalent cations e.g., magnesium ions
- the precursor RNA disclosed herein is capable of circularizing in the absence of magnesium ions and GTP and/or without the step of incubation with magnesium ions and GTP.
- the precursor linear RNA polynucleotide circularizes when incubated in the presence of one or more guanosine nucleotides or nucleoside (e.g., GTP) and a divalent cation (e.g., Mg2+).
- transcription occurs at a Mg2+ concentration of at least 3 mM of magnesium. In certain embodiments, the transcription occurs at a Mg2+ concentration of no more than lOOmM of magnesium. In certain embodiments, transcription occurs at a Mg2+ concentration of or about 1 mM, 2 mM, 3 mM, 4 mM, 5 mM, 6 mM, 7 mM, 8 mM, 9 mM, 10 mM, 11 mM, 12 mM, 13 mM, 14 mM, 15 mM, 16 mM, 17 mM, 18 mM, 19 mM, 20 mM, 21 mM, 22 mM, 23 mM, 24 mM, 25 mM, 26 mM, 27 mM, 28 mM, 29 mM, 30 mM, 31 mM, 32 mM, 33 mM, 34 mM, 35 mM, 36 mM,
- the greater concentration of Mg2+ during transcription of a linear RNA polynucleotide improves circularization and/or splicing as compared to the same linear RNA polynucleotide undergoing transcription at a lower Mg2+ concentration.
- the 3’ exon element, 5’ exon element, and/or core functional element in whole or in part promotes the circularization of the precursor linear RNA polynucleotide to form the circular RNA construct provided herein.
- the method comprises ligation.
- the method comprises chemical ligation.
- the method comprises splint mediated ligation.
- the ligation is performed with a T4 ligase using splint DNA.
- the method of preparing a circular RNA comprises providing modified nucleotides or nucleosides to precursor RNA comprising: a. a 5’ combined accessory element, comprising: i. a 3’ intron segment, ii. a 3’ exon segment, b. an intervening region comprising an internal ribosome entry site (IRES) and a noncoding or coding region, c. a 3’ combined accessory element, comprising: i. a 5’ exon segment, and ii. a 5’ intron segment.
- precursor RNA comprising: a. a 5’ combined accessory element, comprising: i. a 3’ intron segment, ii. a 3’ exon segment, b. an intervening region comprising an internal ribosome entry site (IRES) and a noncoding or coding region, c. a 3’ combined accessory element, comprising: i. a 5’ exon segment, and ii. a 5’ intron segment.
- the method of preparing a circular RNA comprises providing a first and second linear precursor RNA polynucleotide, wherein the first and second linear precursor RNA polynucleotides are capable of forming a circular RNA (e.g., ligation or permuted introns).
- a first precursor and a second precursor may be ligated to form a circular RNA.
- the first precursor and the second precursor each comprise a short adapter sequence at their 5 ' and 3 ' ends.
- the adapter sequences comprise homology arms with splints used for circularization. Splint ligation may be performed in the presence of a DNA splint using a suitable ligase to generate a circular RNA polynucleotide. Ligation methods are known in the art. See, e.g., Wesselhoeft et al., 2019.
- a first precursor and a second precursor may splice to form a circular RNA comprising a sequence of interest, e.g., a coding region.
- Each of the first precursor and the second precursor comprises at least one fragment of the sequence of interest, e.g., the first precursor comprises the 5 ' fragment of the sequence of interest and the second precursor comprises the 3 ' fragment of the sequence of interest.
- the 5 ' fragment of the sequence of interest, the 3 ' fragment of the sequence of interest, and two additional fragments of the sequence of interest together form the sequence of interest.
- sequence of interest consists of, e.g., in 5 ' to 3 ' order, the 5 ' fragment of the sequence of interest, an exonic fragment of the sequence of interest (Exon 2A), an exonic fragment of the sequence of interest (Exon 2B), and the 3 ' fragment of the sequence of interest.
- the first precursor comprises the following:
- a translation initiation element e.g., IRES
- one exon fragment (e.g., Exon 2A) is a part of a sequence of interest, for example in the coding or noncoding region.
- the coding region is scanned for sequences that are homologous to this exon (Exon 2A) fragment, thereby allowing splicing to occur without altering the resulting coding sequence in the circular RNA.
- the second precursor comprises the following:
- the coding sequence is scanned for regions that are homologous to this exon (Exon 2B) fragment, thereby allowing splicing to occur without altering the resulting coding sequence in the circular RNA.
- the first precursor and the second precursor may be incubated together to facilitate splicing between the first precursor and the second precursor in order to generate a circular RNA polynucleotide, which comprises specific modified regions and specific unmodified regions.
- a first precursor comprises an optional first external homology region (Arm 1 A), a first intron fragment (3 ' intron fragment of a first intron (Intron 1)), a first exon fragment (Exon IB), an optional internal homology region, an optional spacer, a translation initiation element (e.g., IRES), the 5 ' fragment of the sequence of interest (e.g., coding region), a second exon fragment (Exon 2A), a second intron fragment (5 ' intron fragment of a second intron (Intron 2)), and an optional second external homology region (Arm 2A).
- Arm 1 A optional first external homology region
- first intron fragment 3 ' intron fragment of a first intron (Intron 1)
- a first exon fragment Exon IB
- an optional internal homology region an optional spacer
- a translation initiation element e.g., IRES
- the 5 ' fragment of the sequence of interest e.g., coding region
- a second exon fragment Exon 2A
- the second precursor comprises an optional first external homology region (Arm 2B), a first intron fragment (3 ' intron fragment of the second intron (Intron 2)), a first exon fragment (Exon 2B corresponding to the 5 ' fragment of the 3 ' fragment of the sequence of interest (e.g., coding region)), the 3 ' fragment of the sequence of interest (e.g., coding region), an optional spacer, an optional internal homology region, a second exon fragment (Exon 1A), a second intron fragment (5' fragment of the first intron (Intron 1)), and an optional second external homology region (Arm IB).
- Arm 2B optional first external homology region
- a first intron fragment (3 ' intron fragment of the second intron (Intron 2)
- a first exon fragment Exon 2B corresponding to the 5 ' fragment of the 3 ' fragment of the sequence of interest (e.g., coding region)
- the 3 ' fragment of the sequence of interest e.g., coding region
- either the first precursor or the second precursor comprises a monotron.
- the first precursor comprises an optional first external homology region (Arm 1 A), a first intron fragment (3 ' intron fragment of a first intron (Intron 1)), a first exon fragment (Exon IB), an optional internal homology region, an optional spacer, a translation initiation element (e.g., IRES), the 5 ' fragment of the sequence of interest (e.g., coding region), a second exon fragment (Exon 2A), a terminal element corresponding to a monotron sequence, and an optional second external homology region (Arm 2A).
- Arm 1 A a first intron fragment (3 ' intron fragment of a first intron (Intron 1)), a first exon fragment (Exon IB), an optional internal homology region, an optional spacer, a translation initiation element (e.g., IRES), the 5 ' fragment of the sequence of interest (e.g., coding region), a second exon fragment (Exon 2A), a terminal element corresponding to a monotron sequence, and an optional second external homo
- the second precursor comprises an optional external homology region (Arm 2B), the monotron sequence via Intron 2, a first exon fragment (Exon 2B, which corresponds to the 5 ' fragment of the 3 ' fragment of the sequence of interest (e.g., coding region)), the 3 ' fragment of the sequence of interest (e.g., coding region), an optional spacer, an optional internal homology region, a second exon fragment (Exon 1A), an intron fragment (5 ' intron fragment of Intron 2), and an optional second external homology region (Arm IB).
- the first precursor comprises an optional first external homology region (Arm 1A), a monotron sequence via Intron 1, a first exon fragment (Exon IB), an optional internal homology region, an optional spacer, a translation initiation element (e.g., IRES), the 5 ' fragment of the sequence of interest (e.g., coding region), a second exon fragment (Exon 2A), an intron fragment (5 ' intron fragment of Intron 2), and an optional second external homology region.
- the second precursor comprises an optional external homology region (Arm 2B), an intron fragment (3 ' intron fragment of Intron
- Example 2B which corresponds to the 5 ’ fragment of the 3 ’ fragment of the sequence of interest (e.g., coding region)), a 3 ’ fragment of the sequence of interest (e.g., coding region), an optional spacer, an optional internal homology region, a second exon fragment (Exon 1A), a terminal element corresponding to the monotron sequence, and an optional second external homology region (Arm IB).
- each of the first precursor and the second precursor comprises a monotron.
- the first precursor comprises an optional first external homology region (Arm 1A), a first monotron sequence via Intron 1, a first exon fragment (Exon IB), an optional internal homology region, an optional spacer, a translation initiation element (e.g., IRES), the 5 ' fragment of the sequence of interest, a second exon fragment (Exon 2A), a terminal element corresponding to a second monotron sequence via Intron 2, and an optional second external homology region (Arm 2A).
- the second precursor comprises an optional external homology region (Arm 2B), a second monotron sequence via Intron 2, a first exon fragment (Exon 2B), the 3 ' fragment of the sequence of interest (e.g., coding region), an optional spacer, an optional internal homology region, a second exon fragment (Exon 1A), a terminal element corresponding to the first monotron sequence via Intron 1, and an optional second external homology region (Arm IB).
- circular RNA that do not comprise modified nucleotides and/or modified nucleosides.
- modified circular RNA i.e., comprising at least one modified nucleotide and/or modified nucleoside
- the modified circular RNA affects immunogenicity, circularization, and/or translation as compared to circular RNA prepared with RNA precursor polynucleotides that comprise no nucleotide or nucleoside modifications.
- RNA has reduced immunogenicity relative to a corresponding mRNA, at least partially because the mRNA contains an immunogenic 5’ cap.
- a DNA vector from certain promoters e.g., a T7 promoter, SP6 promoter, or a fragment thereof
- the 5’ end of the precursor RNA is G.
- an excess of GMP relative to GTP can be provided during transcription such that most transcripts contain a 5’ GMP, which cannot be capped. Therefore, in some embodiments, transcription is carried out in the presence of an excess of GMP.
- transcription is carried out where the ratio of GMP concentration to GTP concentration is within the range of about 3 : 1 to about 15 : 1 , for example, about 3:1 to about 10: 1, about 3: 1 to about 5: 1, about 3: 1, about 4: 1, or about 5: 1.
- Circular RNA may be purified by any known method commonly used in the art, such as column chromatography, gel filtration chromatography, and size exclusion chromatography.
- purification comprises one or more of the following steps: phosphatase treatment, HPLC size exclusion purification, and RNase R digestion.
- purification comprises the following steps in order: RNase R digestion, phosphatase treatment, and HPLC size exclusion purification.
- purification comprises reverse phase HPLC.
- a purified composition contains less double stranded RNA, DNA splints, triphosphorylated RNA, phosphatase proteins, protein ligases, capping enzymes and/or nicked RNA than unpurified RNA.
- purification of circular RNA comprises an affinity-purification or negative selection method described herein.
- purification of circular RNA comprises separation of linear RNA from circular RNA using oligonucleotides that are complementary to a sequence in the linear RNA but are not complementary to a sequence in the circular RNA.
- a purified composition is less immunogenic than an unpurified composition.
- immune cells exposed to a purified composition produce less TNFa, RIG-I, IL-2, IL-6, IFNy, and/or a type 1 interferon, e.g., IFN-pi, than immune cells exposed to an unpurified composition.
- circular RNA is produced by transcribing a DNA polynucleotide sequence that is complementary to a precursory RNA polynucleotide that is described herein.
- circular RNA provided herein is produced in vitro.
- circular RNA provided herein is produced inside a cell.
- the cell selected from, for example, an immune cell, muscle cell, neural cell, epithelial cell and a tumor cell.
- precursor RNA is transcribed using a DNA template (e.g., in some embodiments, using a vector provided herein) in the cytoplasm by a bacteriophage RNA polymerase, or in the nucleus by host RNA polymerase II and then circularized.
- a DNA template e.g., in some embodiments, using a vector provided herein
- WO2020/237227 which is incorporated by reference herein in its entirety.
- PIE permuted intron exon
- a l.lkb sequence containing a full-length encephalomyocarditis virus (EMCV) IRES, a Gaussia luciferase (GLuc) expression sequence, and two short exon fragments of the permuted intron-exon (PIE) construct were inserted between the 3’ and 5’ introns of the permuted group I catalytic intron in the thymidylate synthase (Td) gene of the T4 phage.
- Precursor RNA was synthesized by run-off transcription. Circularization was attempted by heating the precursor RNA in the presence of magnesium ions and GTP, but splicing products were not obtained.
- an ionizable lipid that may be used as a component of a transfer vehicle to facilitate or enhance the delivery and release of circular RNA to one or more target cells (e.g, by permeating or fusing with the lipid membranes of such target cells).
- an ionizable lipid comprises one or more cleavable functional groups (e.g, a disulfide) that allow, for example, a hydrophilic functional head- group to dissociate from a lipophilic functional tail-group of the compound (e.g., upon exposure to oxidative, reducing or acidic conditions), thereby facilitating a phase transition in the lipid bilayer of the one or more target cells.
- cleavable functional groups e.g, a disulfide
- an ionizable lipid is as described in international patent application PCT/US2020/038678.
- an ionizable lipid is a lipid as represented by formula 1 of or as listed in Tables 1 or 2 of US Patent No. 9,708,628, the content of which is herein incorporated by reference in its entirety.
- an ionizable lipid is as described in pages 7-13 of US Patent No. 9,765,022 or as represented by formula 1 of US Patent No. 9,765,022, the content of which is herein incorporated by reference in its entirety.
- an ionizable lipid is described in pages 12-24 of International Patent Application No.
- a lipid or transfer vehicle is a lipid as described in International Patent Application Nos. PCT/US2010/061058, PCT/US2018/058555, PCT/US2018/053569, PCT/US2017/028981, PCT/US2019/025246, PCT/US2019/015913, PCT/US2019/016362, PCT/US2019/016362, US Application Publication Nos.
- Suitable cationic lipids for use in the compositions and methods herein include those described in international patent publication WO 2010/053572 and/or US patent application 15/809,680, e.g., C12-200.
- compositions and methods herein employ an ionizable cationic lipid described in WO2013149140 (incorporated herein by reference), such as, e.g., (15Z,18Z) — N,N-dimethyl-6-(9Z,12Z)-octadeca-9,12-dien-l-yl)tetracosa-15,18-dien- 1 -amine (HGT5000), ( 15Z, 18Z)— N,N-dimethyl-6-((9Z, 12Z)-octadeca-9, 12-dien- 1 - yl)tetracosa-4,15,18-trien-l-amine (HGT5001), and (15Z,18Z) — N,N-dimethyl-6-((9Z,12Z)- octadeca-9,12-dien-l-yl)tetracosa-5,15,18-trien-l-amine (HGT5002).
- the cationic lipid N-[l-(2,3-dioleyloxy)propyl]-N,N,N- trimethylammonium chloride or “DOTMA” is used.
- DOTMA can be formulated alone or can be combined with the neutral lipid, dioleoylphosphatidyl-ethanolamine or “DOPE” or other cationic or noncationic lipids into a transfer vehicle or a lipid nanoparticle, and such liposomes can be used to enhance the delivery of nucleic acids into target cells.
- Suitable cationic lipids include, for example, 5-carboxyspermylglycinedioctadecylamide or “DOGS,” 2,3-dioleyloxy-N- [2(spermine-carboxamido)ethyl]-N,N-dimethyl-l-propanaminium or “DOSPA” (Behr et al. Proc. Nat.'l Acad. Sci. 86, 6982 (1989); U.S. Pat. Nos.
- Contemplated cationic lipids also include l,2-distearyloxy-N,N-dimethyl-3- aminopropane or “DSDMA”, l,2-dioleyloxy-N,N-dimethyl-3 -aminopropane or “DODMA,” l,2-dilinoleyloxy-N,N-dimethyl-3 -aminopropane or “DLinDMA,” l,2-dilinolenyloxy-N,N- dimethyl-3 -aminopropane or “DLenDMA,” N-dioleyl-N,N-dimethylammonium chloride or “DODAC,” N,N-distearyl-N,N-dimethylammonium chloride or “DODAC,” N,N-distearyl-N,N-dimethylammonium chloride or “DODAC,” N,N-distearyl-N,N-dimethylammonium chloride or “DODAC,” N,N
- cholesterol -based cationic lipids to formulate the transfer vehicles (e.g., lipid nanoparticles) is also contemplated herein.
- Such cholesterol-based cationic lipids can be used, either alone or in combination with other cationic or non-cationic lipids.
- Suitable cholesterol-based cationic lipids include, for example, GL67, DC-Chol (N,N-dimethyl-N- ethylcarboxamidocholesterol), l,4-bis(3-N-oleylamino-propyl)piperazine (Gao, et al. Biochem. Biophys. Res. Comm. 179, 280 (1991); Wolf et al. BioTechniques 23, 139 (1997); U.S. Pat. No. 5,744,335), or ICE.
- the one or more of the cationic or ionizable lipids provide increased activity of the nucleic acid and improved tolerability of the compositions in vivo.
- PCT/US2022/033091 (WO 2022/261490) describes representative cationic lipids of any one of the disclosed embodiments and is incorporated by reference herein in its entirety.
- the cationic lipid (or a pharmaceutically acceptable salt, tautomer, prodrug or stereoisomer thereof) has a structure selected from one of the following that are described in detail on pages 113-118 of WO 2022/261490 and page 113-118 of WO 2023/056033, which are incorporated by reference herein in their entireties:
- a cationic lipid of any one of the disclosed embodiments has a structure of Formula I and/or is selected from structure numbers 1-1 through 1-41, set forth at pages 119-130 and in Table 1 of WO 2022/261490; has a structure of Formula II and/or is selected from structure numbers II- 1 through 11-46, set forth at pages 130-146 and in Table 2 of WO 2022/261490; has a structure of Formula III and/or is selected from structure numbers III-l through III-49, set forth at pages 146-157 and in Table 3 of WO 2022/261490; has a structure of Formula IV or V and/or is selected from structure numbers IV- 1 through IV- 3, set forth at pages 157-174 and in Table 4 of WO 2022/261490; has a structure of Formula VI and/or is selected from structure numbers VI-1 through VI-37, set forth at pages 174-188 and in Table 5 of WO 2022/261490; has a structure of Formula I and/or is selected from structure numbers 1-1
- an ionizable lipid is a compound of Formula (1), Formula (1-1), Formula (1-2), Formula (2), Formula (3), Formula (3-1), Formula (3-2), Formula (3-3), Formula (5), or Formula (6), in WO 2022/261490, which is incorporated by reference herein in its entirety.
- WO 2022/261490 provides exemplary reaction schemes that illustrate an exemplary method to make compounds of Formula (1).
- WO 2023/056033 describes similar structures, and is incorporated by reference in its entirety.
- an ionizable lipid of the disclosure is a lipid selected from Table 10a, Table 10b, or Table 10 on pages 235-271- of WO 2022/261490, which is incorporated herein by reference in its entirety.
- the ionizable lipid is Lipid 26, 27, 53, 54, 45, 46, 137, 138, 139, 128, or 130 in Table 10a of WO 2022/261490.
- an ionizable lipid of the disclosure is Lipid 15 from Table 10b of WO 2022/261490.
- the ionizable lipid is described in US patent publication number US20170210697A1. In an embodiment, the ionizable lipid is described in US patent publication number US20170119904A1.
- the ionizable lipid has one of the structures set forth in Table 11 of WO 2022/261490, which is incorporated herein by reference in its entirety, certain of which are described in international patent application PCT/US2010/061058.
- the ionizable lipids may include a lipid selected from Tables 12, 13, 14, or 15a of WO 2022/261490.
- the transfer vehicle comprises Lipid A, Lipid B, Lipid C, and/or Lipid D, described in detail, including methods of synthesis that are known in the art, in WO 2022/261490, WO 2023/056033, and PCT/US2017/028981, which are incorporated herein by reference in their entireties.
- ionizable lipids such as the dialkylamino-based, imidazole- based, and guanidinium-based lipids. See, e.g., PCT/US2010/058457, incorporated herein by reference.
- certain embodiments are directed to a composition comprising one or more imidazole-based ionizable lipids, for example, the imidazole cholesterol ester or “ICE” lipid, (3S, 10R, 13R, 17R)-10, 13 -dimethyl- 17-((R)-6-methylheptan-2-yl)-2, 3, 4, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17-tetradecahydro-lH-cyclopenta[a]phenanthren-3-yl 3-(lH-imidazol- 4-yl)propanoate, as represented by structure (XIII) of WO 2022/261490 and WO 2023/056033, which are incorporated herein by reference in their entireties.
- imidazole cholesterol ester or “ICE” lipid 3S, 10R, 13R, 17R)-10, 13 -dimethyl- 17-((R)-6-methylheptan-2-yl)-2, 3, 4, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17-t
- a transfer vehicle for delivery of circRNA may comprise one or more imidazole-based ionizable lipids, for example, the imidazole cholesterol ester or “ICE” lipid (3S, 10R, 13R, 17R)-10, 13- dimethyl-17-((R)-6-methylheptan-2-yl)-2, 3, 4, 7, 9, 10, 11, 12, 13, 14, 15, 16, 17- tetradecahydro-lH-cyclopenta[a]phenanthren-3-yl 3-(lH-imidazol-4-yl)propanoate, as represented by structure (XIII).
- an ionizable lipid is described by US patent publication number 20190314284.
- the ionizable lipid is described by structure (XIV), structure XVII (referred to herein as “HGT4001”), structure XVIII (referred to herein as “HGT4002”), structure XIX (referred to herein as “HGT4003”), structure XX (referred to herein as “HGT4004”), or structure XXI (referred to herein as “HGT4005”) of WO 2022/261490 and WO 2023/056033, which are incorporated herein by reference in their entireties.
- the ionizable lipid is selected from a lipid with a structure depicted on 390-457 of WO 2022/261490, which is incorporated herein by reference in its entirety.
- WO 2023/056033 also describes representative cationic lipids of any one of the disclosed embodiments and is incorporated by reference herein in its entirety.
- a cationic lipid of any one of the disclosed embodiments has the structure of Formula (7), (7-1), (7-2), (7-3), (8), (8-1), (8-2), (8-3), (8-4), (9), (10), (11), and/or (12) of WO 2023/056033.
- the cationic lipid is selected from a lipid with a structure depicted in any of Tables lOa-lOf, Table 11, Tables 12, 13, 14, or 15a of WO 2023/056033.
- the ionizable lipid is described by structure (XIV) of WO 2023/056033, and pharmaceutical compositions comprising the compound of structure XIV are envisioned.
- the cationic lipid is selected from a lipid with a structure depicted on pages 386-439 of WO 2023/056033.
- the one or more of the cationic or ionizable lipids are represented by Formula (I):
- n is an integer between 1 and 4;
- Ra is hydrogen or hydroxyl
- Ri and R2 are each independently a linear or branched C6-C30 alkyl, C6-C30 alkenyl, or C6-C30 heteroalkyl, optionally substituted by one or more substituents selected from a group consisting of oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino,
- Ra is hydrogen. In some embodiments, Ra is hydroxyl.
- the ionizable lipid is represented by Formula (la -1), Formula (la- 2), or Formula (la-3):
- the ionizable lipid is represented by Formula (Ib-1), Formula (Ib-
- the ionizable lipid is represented by Formula (Ib-4), Formula (Ib-
- Formula (Ib-7) Formula (Ib-8) Formula (Ib-9) [513]
- the one or more of the cationic or ionizable lipids are represented by Formula (I), wherein Ri and R2 are each independently selected from:
- Ri and R2 are the same. In some embodiments, Ri and R2 are different.
- the one or more of the cationic or ionizable lipids are represented by Formula (I*):
- n* is an integer between 1 to 7
- R a is hydrogen or hydroxyl
- R b is hydrogen or Ci-Ce alkyl
- Ri and R2 are each independently a linear or branched C1-C30 alkyl, C2-C30 alkenyl, or C1-C30 heteroalkyl, optionally substituted by one or more substituents selected from oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino, hydroxycarbonyl, al
- the one or more of the cationic or ionizable lipids are represented by Formula (II):
- Li and L3 are each independently -OC(O)-* or -C(O)O-*, wherein indicates the attachment point to Ri or R3;
- Ri and R3 are each independently a linear or branched C9-C20 alkyl or C9-C20 alkenyl, optionally substituted by one or more substituents selected from a group consisting of oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino, hydroxy carbonyl, alkyloxycarbon
- alkylaminoalkyl (alkyl)aminocarbonyl, alkylaminoalkylcarbonyl, dialkylaminoalkylcarbonyl, heterocyclylcarbonyl, alkenylcarbonyl, alkynylcarbonyl, alkylsulfoxide, alkylsulfoxidealkyl, alkylsulfonyl, and alkyl sulfonealkyl;
- R2 is selected from a group consisting of:
- the ionizable lipid is selected from an ionizable lipid of Formula II, wherein Ri and Rs are each independently selected from a group consisting of:
- Ri and Rs are the same. In some embodiments, Ri and Rs are different.
- the one or more of the cationic or ionizable lipids are represented by Formula (II- 1) or Formula (II-2):
- the ionizable lipid is selected from an ionizable lipid of W02015/095340 (lipid number 123 of Table 3). In some embodiments, the ionizable lipid is selected from an ionizable lipid ofWO2021/021634, WO2020/237227, or WO2019/236673 (lipid numbers 124- 127 of Table 3). In some embodiments, the ionizable lipid is selected from an ionizable lipid of WO2021226597 and WO2021113777 (lipid numbers 128-131 ofTable 3).
- the transfer vehicle comprises an ionizable lipid selected from an ionizable lipid represented in Table 3.
- the ionizable lipid is selected from lipid numbers 16, 45, 86, 89, and 90 of Table 3, below.
- the ionizable lipid is selected from lipid numbers 128-131 of Table 3, below.
- the one or more of the cationic or ionizable lipids are represented by Formula (III):
- L 1 is C2-C11 alkylene, C4-Cio-alkenylene, or C4-Cio-alkynylene;
- X 1 is OR 1 , SR 1 , or N(R')2, where R 1 is independently H or unsubstituted Ci-Ce alkyl;
- R 2 and R 3 are each independently Ce-Cso-alkyl, Ce-Cso-alkenyl, or Ce-Cso-alkynyl.
- the one or more of the cationic or ionizable lipids are represented by Formula (III*):
- L 1 is C2-C11 alkylene, C4-Cio-alkenylene, or C4-Cio-alkynylene;
- X 1 is OR 1 , SR 1 , or N(R')2, where R 1 is independently H or unsubstituted Ci-Ce alkyl;
- R2 and R3 are each independently a linear or branched C1-C30 alkyl, C2-C30 alkenyl, or C1-C30 heteroalkyl, optionally substituted by one or more substituents selected from oxo, halo, hydroxy, cyano, alkyl, alkenyl, aldehyde, heterocyclylalkyl, hydroxyalkyl, dihydroxyalkyl, hydroxyalkylaminoalkyl, aminoalkyl, alkylaminoalkyl, dialkylaminoalkyl, (heterocyclyl)(alkyl)aminoalkyl, heterocyclyl, heteroaryl, alkylheteroaryl, alkynyl, alkoxy, amino, dialkylamino, aminoalkylcarbonylamino, aminocarbonylalkylamino, (aminocarbonylalkyl)(alkyl)amino, alkenylcarbonylamino, hydroxycarbonyl,
- an ionizable lipid is a compound of Formula (15):
- n* is an integer from 1 to 7;
- R a is hydrogen or hydroxyl
- R h is hydrogen or Ci-Ce alkyl
- R 1 * and R 2 * are independently selected from:
- R 8 is H or R 11 ;
- R 9 , R 10 , and R 11 are each independently C1-C20 alkyl or C2-C2o-alkenyl; and wherein (i) R 1 is R 1 *, (ii) R 2 is R 2 *, or (iii) R 1 is R 1 * and R 2 is R 2 *.
- R a is hydrogen and the ionizable lipid is of
- the ionizable lipid is of Formula (17):
- n is an integer from 1 to 7; q and q’ are each independently integers from 0 to 12; r and r’ are each independently integers from 0 to 6, wherein at least one of r or r’ is not 0;
- Z A and Z B are each independently selected from A -C(O)O-, A -OC(O), and -OC(O)O-; where denotes the attachment point to -(C Dq- or -(CH2) q and
- R 9A , R 9B , R 1OA and R 10B are each independently C1-C20 alkyl or C2-C20 alkenyl.
- Z A and Z B are A -C(O)O-, and the ionizable lipid is of Formula (17a- 1)
- Z A and Z B are A -OC(O)-, and the ionizable lipid is of Formula ( 17a-2)
- Z A and Z B are -O(C)(O)O-, and the ionizable lipid is represented by Formula (17a-3):
- R a is hydroxyl and the ionizable lipid is of Formula (18):
- n* is an integer from 1 to 7;
- R h is hydrogen or Ci-Ce alkyl
- R 1 * and R 2 * are independently selected from:
- R 8 is hydrogen or R 11 ;
- R 9 , R 10 , and R 11 are each independently Ci-C 2 o alkyl or C 2 -C 2 o-alkenyl; wherein (i) R 1 is R 1 *, (ii) R 2 is R 2 *, or (iii) R 1 is R 1 * and R 2 is R 2 *; and wherein, for (iii), (a) R 1 * and R 2 * are different or (b) R 9 and R 10 have different numbers of carbon atoms for at least one of R 1 * and R 2 * .
- the ionizable lipid of is of Formula (19):
- n is an integer from 1 to 7; q and q’ are each independently integers from 0 to 12; r and r’ are each independently integers from 0 to 6, wherein at least one of r or r’ is not 0;
- Z A and Z B are each independently selected from A -C(O)O-, A -OC(O), and -OC(O)O-; where denotes the attachment point to -(CH 2 ) q - or -(CH 2 ) q ;-and R 9A , R 9B , R 1OA and R 10B are each independently C1-C20 alkyl or C2-C20 alkenyl.
- Z A and Z B are A -C(O)O-, and the ionizable lipid is of Formula (19a- 1):
- Z A and Z B are A -OC(O)-, and the ionizable lipid is of Formula ( 19a-2) :
- Z A and Z B are -O(C)(O)O-, and the ionizable lipid is represented by Formula (19a-3):
- R 1 is C1-C30 alkyl, and the ionizable lipid is of
- R 9A and R 1OA are each independently C1-C20 alkyl or C2-C20 alkenyl; n is an integer from 1 to 7; q is an integer from 0 to 12; and r is an integer from 1 to 6.
- Z A is A -C(O)O-
- the ionizable lipid is of Formula (20a- 1):
- Z A is A -OC(O)-
- the ionizable lipid is of Formula (20a-2):
- Z A is -OC(O)O-, and the ionizable lipid is of
- R 2 is C1-C30 alkyl, and the ionizable lipid is of
- Z B is selected from A -C(O)O-, A -OC(O)-, and -OC(O)O-; where A denotes the attachment point to -(CH 2 ) q -;
- R 9B and R 1OB are each independently C1-C20 alkyl or C2-C20 alkenyl; n is an integer from 1 to 7; q’ is an integer from 0 to 12; and r’ is an integer from 1 to 6.
- Z B is A -C(O)O-, and the ionizable lipid is of
- Z B is A -OC(O)-
- the ionizable lipid is of Formula (2 la-2):
- Z B is -OC(O)O-
- the ionizable lipid is of Formula (2 la-3):
- an ionizable lipid is selected from the table below:
- an ionizable lipid of the present disclosure is represented by Formula (22):
- R a is hydrogen or hydroxyl
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Genetics & Genomics (AREA)
- Engineering & Computer Science (AREA)
- General Health & Medical Sciences (AREA)
- General Engineering & Computer Science (AREA)
- Biomedical Technology (AREA)
- Organic Chemistry (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Microbiology (AREA)
- Veterinary Medicine (AREA)
- Epidemiology (AREA)
- Public Health (AREA)
- Biotechnology (AREA)
- Animal Behavior & Ethology (AREA)
- Mycology (AREA)
- Biophysics (AREA)
- Physics & Mathematics (AREA)
- Pharmacology & Pharmacy (AREA)
- Molecular Biology (AREA)
- Plant Pathology (AREA)
- Medicinal Chemistry (AREA)
- Immunology (AREA)
- General Chemical & Material Sciences (AREA)
- Biochemistry (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Pharmaceuticals Containing Other Organic And Inorganic Compounds (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
Abstract
Description
Claims
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| AU2024269222A AU2024269222A1 (en) | 2023-05-05 | 2024-05-03 | Circular rna compositions and methods |
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363464435P | 2023-05-05 | 2023-05-05 | |
| US63/464,435 | 2023-05-05 | ||
| US202363470064P | 2023-05-31 | 2023-05-31 | |
| US63/470,064 | 2023-05-31 |
Publications (3)
| Publication Number | Publication Date |
|---|---|
| WO2024233308A2 true WO2024233308A2 (en) | 2024-11-14 |
| WO2024233308A3 WO2024233308A3 (en) | 2025-01-16 |
| WO2024233308A9 WO2024233308A9 (en) | 2025-04-10 |
Family
ID=91302109
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/027627 Pending WO2024233308A2 (en) | 2023-05-05 | 2024-05-03 | Circular rna compositions and methods |
Country Status (2)
| Country | Link |
|---|---|
| AU (1) | AU2024269222A1 (en) |
| WO (1) | WO2024233308A2 (en) |
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2025259931A1 (en) * | 2024-06-14 | 2025-12-18 | Orbital Therapeutics, Inc. | Compositions and methods for rna circularization |
Citations (129)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US3832253A (en) | 1973-03-21 | 1974-08-27 | Baxter Laboratories Inc | Method of making an inflatable balloon catheter |
| US3854480A (en) | 1969-04-01 | 1974-12-17 | Alza Corp | Drug-delivery system |
| US4450150A (en) | 1973-05-17 | 1984-05-22 | Arthur D. Little, Inc. | Biodegradable, implantable drug delivery depots, and method for preparing and using the same |
| US4452775A (en) | 1982-12-03 | 1984-06-05 | Syntex (U.S.A.) Inc. | Cholesterol matrix delivery system for sustained release of macromolecules |
| US4667014A (en) | 1983-03-07 | 1987-05-19 | Syntex (U.S.A.) Inc. | Nonapeptide and decapeptide analogs of LHRH, useful as LHRH antagonists |
| US4748034A (en) | 1983-05-13 | 1988-05-31 | Nestec S.A. | Preparing a heat stable aqueous solution of whey proteins |
| US4897355A (en) | 1985-01-07 | 1990-01-30 | Syntex (U.S.A.) Inc. | N[ω,(ω-1)-dialkyloxy]- and N-[ω,(ω-1)-dialkenyloxy]-alk-1-yl-N,N,N-tetrasubstituted ammonium lipids and uses therefor |
| US5075109A (en) | 1986-10-24 | 1991-12-24 | Southern Research Institute | Method of potentiating an immune response |
| US5087616A (en) | 1986-08-07 | 1992-02-11 | Battelle Memorial Institute | Cytotoxic drug conjugates and their delivery to tumor cells |
| US5171678A (en) | 1989-04-17 | 1992-12-15 | Centre National De La Recherche Scientifique | Lipopolyamines, their preparation and their use |
| WO1993013121A1 (en) | 1991-12-24 | 1993-07-08 | Isis Pharmaceuticals, Inc. | Gapped 2' modified oligonucleotides |
| US5239660A (en) | 1990-10-31 | 1993-08-24 | Nec Corporation | Vector processor which can be formed by an integrated circuit of a small size |
| US5334761A (en) | 1992-08-28 | 1994-08-02 | Life Technologies, Inc. | Cationic lipids |
| US5378825A (en) | 1990-07-27 | 1995-01-03 | Isis Pharmaceuticals, Inc. | Backbone modified oligonucleotide analogs |
| WO1995032305A1 (en) | 1994-05-19 | 1995-11-30 | Dako A/S | Pna probes for detection of neisseria gonorrhoeae and chlamydia trachomatis |
| US5585481A (en) | 1987-09-21 | 1996-12-17 | Gen-Probe Incorporated | Linking reagents for nucleotide probes |
| US5744335A (en) | 1995-09-19 | 1998-04-28 | Mirus Corporation | Process of transfecting a cell with a polynucleotide mixed with an amphipathic compound and a DNA-binding protein |
| US5885613A (en) | 1994-09-30 | 1999-03-23 | The University Of British Columbia | Bilayer stabilizing components and their use in forming programmable fusogenic liposomes |
| US5948902A (en) | 1997-11-20 | 1999-09-07 | South Alabama Medical Science Foundation | Antisense oligonucleotides to human serine/threonine protein phosphatase genes |
| US6066626A (en) | 1997-10-29 | 2000-05-23 | Genzyme Corporation | Compositions and method for treating lysosomal storage disease |
| US6319494B1 (en) | 1990-12-14 | 2001-11-20 | Cell Genesys, Inc. | Chimeric chains for receptor-associated signal transduction pathways |
| US20030022649A1 (en) | 2001-05-03 | 2003-01-30 | Mitsubishi Denki Kabushiki Kaisha | Signal reception method and device |
| US20040014194A1 (en) | 2002-03-27 | 2004-01-22 | Schering Corporation | Beta-secretase crystals and methods for preparing and using the same |
| US20040110709A1 (en) | 2000-02-17 | 2004-06-10 | Genzyme Corporation | Genetic modification of the lung as a portal for gene delivery |
| US20050222064A1 (en) | 2002-02-20 | 2005-10-06 | Sirna Therapeutics, Inc. | Polycationic compositions for cellular delivery of polynucleotides |
| WO2005120152A2 (en) | 2004-06-07 | 2005-12-22 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods of use |
| WO2005121348A1 (en) | 2004-06-07 | 2005-12-22 | Protiva Biotherapeutics, Inc. | Lipid encapsulated interfering rna |
| WO2006000830A2 (en) | 2004-06-29 | 2006-01-05 | Avidex Ltd | Cells expressing a modified t cell receptor |
| WO2006007712A1 (en) | 2004-07-19 | 2006-01-26 | Protiva Biotherapeutics, Inc. | Methods comprising polyethylene glycol-lipid conjugates for delivery of therapeutic agents |
| WO2006069782A2 (en) | 2004-12-27 | 2006-07-06 | Silence Therapeutics Ag. | Lipid complexes coated with peg and their use |
| WO2008042973A2 (en) | 2006-10-03 | 2008-04-10 | Alnylam Pharmaceuticals, Inc. | Lipid containing formulations |
| WO2008103276A2 (en) | 2007-02-16 | 2008-08-28 | Merck & Co., Inc. | Compositions and methods for potentiated activity of biologicaly active molecules |
| WO2009086558A1 (en) | 2008-01-02 | 2009-07-09 | Tekmira Pharmaceuticals Corporation | Improved compositions and methods for the delivery of nucleic acids |
| WO2009127060A1 (en) | 2008-04-15 | 2009-10-22 | Protiva Biotherapeutics, Inc. | Novel lipid formulations for nucleic acid delivery |
| WO2009132131A1 (en) | 2008-04-22 | 2009-10-29 | Alnylam Pharmaceuticals, Inc. | Amino lipid based improved lipid formulation |
| US20100061058A1 (en) | 2008-09-11 | 2010-03-11 | Gen Tanabe | Electrical unit cooling structure for motorcycles |
| WO2010042877A1 (en) | 2008-10-09 | 2010-04-15 | Tekmira Pharmaceuticals Corporation | Improved amino lipids and methods for the delivery of nucleic acids |
| WO2010048536A2 (en) | 2008-10-23 | 2010-04-29 | Alnylam Pharmaceuticals, Inc. | Processes for preparing lipids |
| WO2010054384A1 (en) | 2008-11-10 | 2010-05-14 | Alnylam Pharmaceuticals, Inc. | Lipids and compositions for the delivery of therapeutics |
| WO2010054406A1 (en) | 2008-11-10 | 2010-05-14 | Alnylam Pharmaceuticals, Inc. | Novel lipids and compositions for the delivery of therapeutics |
| WO2010053572A2 (en) | 2008-11-07 | 2010-05-14 | Massachusetts Institute Of Technology | Aminoalcohol lipidoids and uses thereof |
| US7741465B1 (en) | 1992-03-18 | 2010-06-22 | Zelig Eshhar | Chimeric receptor genes and cells transformed therewith |
| WO2010088537A2 (en) | 2009-01-29 | 2010-08-05 | Alnylam Pharmaceuticals, Inc. | Improved lipid formulation |
| WO2010129709A1 (en) | 2009-05-05 | 2010-11-11 | Alnylam Pharmaceuticals, Inc. | Lipid compositions |
| WO2010144740A1 (en) | 2009-06-10 | 2010-12-16 | Alnylam Pharmaceuticals, Inc. | Improved lipid formulation |
| WO2011000107A1 (en) | 2009-07-01 | 2011-01-06 | Protiva Biotherapeutics, Inc. | Novel lipid formulations for delivery of therapeutic agents to solid tumors |
| WO2011000106A1 (en) | 2009-07-01 | 2011-01-06 | Protiva Biotherapeutics, Inc. | Improved cationic lipids and methods for the delivery of therapeutic agents |
| WO2011022460A1 (en) | 2009-08-20 | 2011-02-24 | Merck Sharp & Dohme Corp. | Novel cationic lipids with various head groups for oligonucleotide delivery |
| WO2011038160A2 (en) | 2009-09-23 | 2011-03-31 | Protiva Biotherapeutics, Inc. | Compositions and methods for silencing genes expressed in cancer |
| WO2011066651A1 (en) | 2009-12-01 | 2011-06-09 | Protiva Biotherapeutics, Inc. | Snalp formulations containing antioxidants |
| WO2011071860A2 (en) | 2009-12-07 | 2011-06-16 | Alnylam Pharmaceuticals, Inc. | Compositions for nucleic acid delivery |
| WO2011090965A1 (en) | 2010-01-22 | 2011-07-28 | Merck Sharp & Dohme Corp. | Novel cationic lipids for oligonucleotide delivery |
| WO2011127255A1 (en) | 2010-04-08 | 2011-10-13 | Merck Sharp & Dohme Corp. | Preparation of lipid nanoparticles |
| WO2011141705A1 (en) | 2010-05-12 | 2011-11-17 | Protiva Biotherapeutics, Inc. | Novel cationic lipids and methods of use thereof |
| WO2011141704A1 (en) | 2010-05-12 | 2011-11-17 | Protiva Biotherapeutics, Inc | Novel cyclic cationic lipids and methods of use |
| US8071082B2 (en) | 2006-07-21 | 2011-12-06 | Massachusetts Institute Of Technology | End-modified poly(beta-amino esters) and uses thereof |
| WO2011153120A1 (en) | 2010-06-04 | 2011-12-08 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012000104A1 (en) | 2010-06-30 | 2012-01-05 | Protiva Biotherapeutics, Inc. | Non-liposomal systems for nucleic acid delivery |
| WO2012016184A2 (en) | 2010-07-30 | 2012-02-02 | Alnylam Pharmaceuticals, Inc. | Methods and compositions for delivery of active agents |
| WO2012024526A2 (en) | 2010-08-20 | 2012-02-23 | Cerulean Pharma Inc. | Conjugates, particles, compositions, and related methods |
| WO2012031043A1 (en) | 2010-08-31 | 2012-03-08 | Novartis Ag | Pegylated liposomes for delivery of immunogen-encoding rna |
| WO2012040184A2 (en) | 2010-09-20 | 2012-03-29 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012044638A1 (en) | 2010-09-30 | 2012-04-05 | Merck Sharp & Dohme Corp. | Low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012054365A2 (en) | 2010-10-21 | 2012-04-26 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| US20120178702A1 (en) | 1995-01-23 | 2012-07-12 | University Of Pittsburgh | Stable lipid-comprising drug delivery complexes and methods for their production |
| WO2012099755A1 (en) | 2011-01-11 | 2012-07-26 | Alnylam Pharmaceuticals, Inc. | Pegylated lipids and their use for drug delivery |
| WO2012162210A1 (en) | 2011-05-26 | 2012-11-29 | Merck Sharp & Dohme Corp. | Ring constrained cationic lipids for oligonucleotide delivery |
| WO2013006825A1 (en) | 2011-07-06 | 2013-01-10 | Novartis Ag | Liposomes having useful n:p ratio for delivery of rna molecules |
| WO2013016058A1 (en) | 2011-07-22 | 2013-01-31 | Merck Sharp & Dohme Corp. | Novel bis-nitrogen containing cationic lipids for oligonucleotide delivery |
| WO2013033563A1 (en) | 2011-08-31 | 2013-03-07 | Novartis Ag | Pegylated liposomes for delivery of immunogen-encoding rna |
| WO2013049328A1 (en) | 2011-09-27 | 2013-04-04 | Alnylam Pharmaceuticals, Inc. | Di-aliphatic substituted pegylated lipids |
| WO2013086354A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| WO2013086373A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Lipids for the delivery of active agents |
| WO2013086322A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Branched alkyl and cycloalkyl terminated biodegradable lipids for the delivery of active agents |
| US20130156845A1 (en) | 2010-04-29 | 2013-06-20 | Alnylam Pharmaceuticals, Inc. | Lipid formulated single stranded rna |
| WO2013089151A1 (en) | 2011-12-12 | 2013-06-20 | 協和発酵キリン株式会社 | Lipid nanoparticles for drug delivery system containing cationic lipids |
| WO2013093648A2 (en) | 2011-11-04 | 2013-06-27 | Nitto Denko Corporation | Method of producing lipid nanoparticles for drug delivery |
| WO2013116126A1 (en) | 2012-02-01 | 2013-08-08 | Merck Sharp & Dohme Corp. | Novel low molecular weight, biodegradable cationic lipids for oligonucleotide delivery |
| WO2013126803A1 (en) | 2012-02-24 | 2013-08-29 | Protiva Biotherapeutics Inc. | Trialkyl cationic lipids and methods of use thereof |
| WO2013149140A1 (en) | 2012-03-29 | 2013-10-03 | Shire Human Genetic Therapies, Inc. | Ionizable cationic lipids |
| WO2013148541A1 (en) | 2012-03-27 | 2013-10-03 | Merck Sharp & Dohme Corp. | DIETHER BASED BIODEGRADABLE CATIONIC LIPIDS FOR siRNA DELIVERY |
| US8734853B2 (en) | 2008-11-17 | 2014-05-27 | University Of North Texas Health Science Center At Fort Worth | HDL particles for delivery of nucleic acids |
| WO2014136086A1 (en) | 2013-03-08 | 2014-09-12 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2015061467A1 (en) | 2013-10-22 | 2015-04-30 | Shire Human Genetic Therapies, Inc. | Lipid formulations for delivery of messenger rna |
| WO2015074085A1 (en) | 2013-11-18 | 2015-05-21 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid for rna delivery |
| WO2015095340A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2015095346A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| US20150239926A1 (en) | 2013-11-18 | 2015-08-27 | Arcturus Therapeutics, Inc. | Asymmetric ionizable cationic lipid for rna delivery |
| WO2015130584A2 (en) | 2014-02-25 | 2015-09-03 | Merck Sharp & Dohme Corp. | Lipid nanoparticle vaccine adjuvants and antigen delivery systems |
| WO2015199952A1 (en) | 2014-06-25 | 2015-12-30 | Acuitas Therapeutics Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2016081029A1 (en) | 2014-11-18 | 2016-05-26 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid for rna delivery |
| US20160151284A1 (en) | 2013-07-23 | 2016-06-02 | Protiva Biotherapeutics, Inc. | Compositions and methods for delivering messenger rna |
| US20160376224A1 (en) | 2015-06-29 | 2016-12-29 | Acuitas Therapeutics, Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017049245A2 (en) | 2015-09-17 | 2017-03-23 | Modernatx, Inc. | Compounds and compositions for intracellular delivery of therapeutic agents |
| US20170119904A1 (en) | 2015-10-28 | 2017-05-04 | Acuitas Therapeutics, Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017099823A1 (en) | 2015-12-10 | 2017-06-15 | Modernatx, Inc. | Compositions and methods for delivery of therapeutic agents |
| WO2017117528A1 (en) | 2015-12-30 | 2017-07-06 | Acuitas Therapeutics, Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| US20170190661A1 (en) | 2015-12-30 | 2017-07-06 | Arcturus Therapeutics, Inc. | Aromatic Ionizable Cationic Lipid |
| US9708628B2 (en) | 2011-11-18 | 2017-07-18 | Nof Corporation | Cationic lipid having improved intracellular kinetics |
| US9765022B2 (en) | 2013-02-28 | 2017-09-19 | Tufts University | Disulfide compounds for delivery of pharmaceutical agents |
| WO2017173054A1 (en) | 2016-03-30 | 2017-10-05 | Intellia Therapeutics, Inc. | Lipid nanoparticle formulations for crispr/cas components |
| US20180005363A1 (en) | 2015-01-30 | 2018-01-04 | Hitachi High-Technologies Corporation | Pattern Matching Device and Computer Program for Pattern Matching |
| WO2018011633A1 (en) | 2016-07-13 | 2018-01-18 | Alcatel Lucent | Underlying recessed component placement |
| US20180053569A1 (en) | 2015-05-04 | 2018-02-22 | Huawei Technologies Co., Ltd. | DRAM Refresh Method, Apparatus, and System |
| US20190015913A1 (en) | 2017-07-12 | 2019-01-17 | Rexon Industrial Corp., Ltd. | Miter saw with enhanced adjustable miter angle |
| US20190091164A1 (en) | 2015-08-21 | 2019-03-28 | Pfizer Inc. | Therapeutic Nanoparticles Comprising A Therapeutic Agent And Methods of Making And Using Same |
| WO2019131770A1 (en) | 2017-12-27 | 2019-07-04 | 武田薬品工業株式会社 | Nucleic acid-containing lipid nano-particle and use thereof |
| US20190314284A1 (en) | 2011-06-08 | 2019-10-17 | Translate Bio, Inc. | Cleavable Lipids |
| US20190321489A1 (en) | 2011-06-08 | 2019-10-24 | Translate Bio, Inc. | Lipid nanoparticle compositions and methods for mrna delivery |
| US20190345503A1 (en) | 2016-06-20 | 2019-11-14 | The Board Of The Leland Stanford Junior University | Circular rnas and their use in immunomodulation |
| WO2019236673A1 (en) | 2018-06-06 | 2019-12-12 | Massachusetts Institute Of Technology | Circular rna for translation in eukaryotic cells |
| US20200038678A1 (en) | 2017-02-08 | 2020-02-06 | Michelson Diagnostics Limited | Processing optical coherence tomography (oct) scans |
| WO2020237227A1 (en) | 2019-05-22 | 2020-11-26 | Massachusetts Institute Of Technology | Circular rna compositions and methods |
| WO2021021634A1 (en) | 2019-07-29 | 2021-02-04 | Georgia Tech Research Corporation | Nanomaterials containing constrained lipids and uses thereof |
| WO2021041473A1 (en) | 2019-08-27 | 2021-03-04 | The Trustees Of Columbia University In The City Of New York | Engineered exosomes for targeted delivery |
| WO2021113777A2 (en) | 2019-12-04 | 2021-06-10 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021189059A2 (en) | 2020-03-20 | 2021-09-23 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021226597A2 (en) | 2020-05-08 | 2021-11-11 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021236855A1 (en) | 2020-05-19 | 2021-11-25 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| US20220033091A1 (en) | 2020-07-30 | 2022-02-03 | Hamilton Sundstrand Corporation | Aircraft environmental control system |
| WO2022061490A1 (en) | 2020-09-22 | 2022-03-31 | 华为数字能源技术有限公司 | Photovoltaic power generation system, and method and device for detecting ground fault in photovoltaic strings |
| US20220251578A1 (en) | 2020-06-25 | 2022-08-11 | The Board Of Trustees Of The Leland Stanford Junior University | Genetic elements driving circular rna translation and methods of use |
| US20220288176A1 (en) | 2019-08-28 | 2022-09-15 | The Board Of Trustees Of The Leland Stanford Junior University | Circular rna modification and methods of use |
| WO2022261490A2 (en) | 2021-06-10 | 2022-12-15 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2022271965A2 (en) | 2021-06-25 | 2022-12-29 | The Board Of Trustees Of The Leland Stanford Junior University | Compositions and methods for improved protein translation from recombinant circular rnas |
| WO2023056033A1 (en) | 2021-09-30 | 2023-04-06 | Orna Therapeutics, Inc. | Lipid nanoparticle compositions for delivering circular polynucleotides |
| WO2023081526A1 (en) | 2021-11-08 | 2023-05-11 | Orna Therapeutics, Inc. | Lipid nanoparticle compositions for delivering circular polynucleotides |
| WO2023141586A1 (en) | 2022-01-21 | 2023-07-27 | Orna Therapeutics, Inc. | Systemic administration of circular rna polynucleotides encoding muscle proteins or protein complexes |
| WO2023250375A1 (en) | 2022-06-24 | 2023-12-28 | Orna Therapeutics, Inc. | Circular rna encoding chimeric antigen receptors targeting bcma |
Family Cites Families (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN115404240A (en) * | 2021-05-28 | 2022-11-29 | 上海环码生物医药有限公司 | Constructs, methods for making circular RNA and uses thereof |
| EP4271818B1 (en) * | 2021-09-17 | 2024-07-31 | Flagship Pioneering Innovations VI, LLC | Compositions and methods for producing circular polyribonucleotides |
| KR20240082384A (en) * | 2021-09-26 | 2024-06-10 | 센터 포 엑설런스 인 몰레큘러 셀 사이언스, 차이니즈 아카데미 오브 사이언시스 | Circular RNA and method for producing the same |
-
2024
- 2024-05-03 WO PCT/US2024/027627 patent/WO2024233308A2/en active Pending
- 2024-05-03 AU AU2024269222A patent/AU2024269222A1/en active Pending
Patent Citations (181)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US3854480A (en) | 1969-04-01 | 1974-12-17 | Alza Corp | Drug-delivery system |
| US3832253A (en) | 1973-03-21 | 1974-08-27 | Baxter Laboratories Inc | Method of making an inflatable balloon catheter |
| US4450150A (en) | 1973-05-17 | 1984-05-22 | Arthur D. Little, Inc. | Biodegradable, implantable drug delivery depots, and method for preparing and using the same |
| US4452775A (en) | 1982-12-03 | 1984-06-05 | Syntex (U.S.A.) Inc. | Cholesterol matrix delivery system for sustained release of macromolecules |
| US4667014A (en) | 1983-03-07 | 1987-05-19 | Syntex (U.S.A.) Inc. | Nonapeptide and decapeptide analogs of LHRH, useful as LHRH antagonists |
| US4748034A (en) | 1983-05-13 | 1988-05-31 | Nestec S.A. | Preparing a heat stable aqueous solution of whey proteins |
| US4897355A (en) | 1985-01-07 | 1990-01-30 | Syntex (U.S.A.) Inc. | N[ω,(ω-1)-dialkyloxy]- and N-[ω,(ω-1)-dialkenyloxy]-alk-1-yl-N,N,N-tetrasubstituted ammonium lipids and uses therefor |
| US5087616A (en) | 1986-08-07 | 1992-02-11 | Battelle Memorial Institute | Cytotoxic drug conjugates and their delivery to tumor cells |
| US5075109A (en) | 1986-10-24 | 1991-12-24 | Southern Research Institute | Method of potentiating an immune response |
| US5585481A (en) | 1987-09-21 | 1996-12-17 | Gen-Probe Incorporated | Linking reagents for nucleotide probes |
| US5171678A (en) | 1989-04-17 | 1992-12-15 | Centre National De La Recherche Scientifique | Lipopolyamines, their preparation and their use |
| US5378825A (en) | 1990-07-27 | 1995-01-03 | Isis Pharmaceuticals, Inc. | Backbone modified oligonucleotide analogs |
| US5239660A (en) | 1990-10-31 | 1993-08-24 | Nec Corporation | Vector processor which can be formed by an integrated circuit of a small size |
| US6319494B1 (en) | 1990-12-14 | 2001-11-20 | Cell Genesys, Inc. | Chimeric chains for receptor-associated signal transduction pathways |
| WO1993013121A1 (en) | 1991-12-24 | 1993-07-08 | Isis Pharmaceuticals, Inc. | Gapped 2' modified oligonucleotides |
| US7741465B1 (en) | 1992-03-18 | 2010-06-22 | Zelig Eshhar | Chimeric receptor genes and cells transformed therewith |
| US5334761A (en) | 1992-08-28 | 1994-08-02 | Life Technologies, Inc. | Cationic lipids |
| WO1995032305A1 (en) | 1994-05-19 | 1995-11-30 | Dako A/S | Pna probes for detection of neisseria gonorrhoeae and chlamydia trachomatis |
| US5885613A (en) | 1994-09-30 | 1999-03-23 | The University Of British Columbia | Bilayer stabilizing components and their use in forming programmable fusogenic liposomes |
| US20120178702A1 (en) | 1995-01-23 | 2012-07-12 | University Of Pittsburgh | Stable lipid-comprising drug delivery complexes and methods for their production |
| US5744335A (en) | 1995-09-19 | 1998-04-28 | Mirus Corporation | Process of transfecting a cell with a polynucleotide mixed with an amphipathic compound and a DNA-binding protein |
| US6066626A (en) | 1997-10-29 | 2000-05-23 | Genzyme Corporation | Compositions and method for treating lysosomal storage disease |
| US5948902A (en) | 1997-11-20 | 1999-09-07 | South Alabama Medical Science Foundation | Antisense oligonucleotides to human serine/threonine protein phosphatase genes |
| US20040110709A1 (en) | 2000-02-17 | 2004-06-10 | Genzyme Corporation | Genetic modification of the lung as a portal for gene delivery |
| US20030022649A1 (en) | 2001-05-03 | 2003-01-30 | Mitsubishi Denki Kabushiki Kaisha | Signal reception method and device |
| US20050222064A1 (en) | 2002-02-20 | 2005-10-06 | Sirna Therapeutics, Inc. | Polycationic compositions for cellular delivery of polynucleotides |
| US20040014194A1 (en) | 2002-03-27 | 2004-01-22 | Schering Corporation | Beta-secretase crystals and methods for preparing and using the same |
| WO2005121348A1 (en) | 2004-06-07 | 2005-12-22 | Protiva Biotherapeutics, Inc. | Lipid encapsulated interfering rna |
| US20060008910A1 (en) | 2004-06-07 | 2006-01-12 | Protiva Biotherapeuties, Inc. | Lipid encapsulated interfering RNA |
| WO2005120152A2 (en) | 2004-06-07 | 2005-12-22 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods of use |
| US20060083780A1 (en) | 2004-06-07 | 2006-04-20 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods of use |
| WO2006000830A2 (en) | 2004-06-29 | 2006-01-05 | Avidex Ltd | Cells expressing a modified t cell receptor |
| WO2006007712A1 (en) | 2004-07-19 | 2006-01-26 | Protiva Biotherapeutics, Inc. | Methods comprising polyethylene glycol-lipid conjugates for delivery of therapeutic agents |
| US20060051405A1 (en) | 2004-07-19 | 2006-03-09 | Protiva Biotherapeutics, Inc. | Compositions for the delivery of therapeutic agents and uses thereof |
| US20100062967A1 (en) | 2004-12-27 | 2010-03-11 | Silence Therapeutics Ag | Coated lipid complexes and their use |
| WO2006069782A2 (en) | 2004-12-27 | 2006-07-06 | Silence Therapeutics Ag. | Lipid complexes coated with peg and their use |
| US8071082B2 (en) | 2006-07-21 | 2011-12-06 | Massachusetts Institute Of Technology | End-modified poly(beta-amino esters) and uses thereof |
| WO2008042973A2 (en) | 2006-10-03 | 2008-04-10 | Alnylam Pharmaceuticals, Inc. | Lipid containing formulations |
| US20090023673A1 (en) | 2006-10-03 | 2009-01-22 | Muthiah Manoharan | Lipid containing formulations |
| WO2008103276A2 (en) | 2007-02-16 | 2008-08-28 | Merck & Co., Inc. | Compositions and methods for potentiated activity of biologicaly active molecules |
| WO2009086558A1 (en) | 2008-01-02 | 2009-07-09 | Tekmira Pharmaceuticals Corporation | Improved compositions and methods for the delivery of nucleic acids |
| US20110117125A1 (en) | 2008-01-02 | 2011-05-19 | Tekmira Pharmaceuticals Corporation | Compositions and methods for the delivery of nucleic acids |
| WO2009127060A1 (en) | 2008-04-15 | 2009-10-22 | Protiva Biotherapeutics, Inc. | Novel lipid formulations for nucleic acid delivery |
| US8492359B2 (en) | 2008-04-15 | 2013-07-23 | Protiva Biotherapeutics, Inc. | Lipid formulations for nucleic acid delivery |
| US20100130588A1 (en) | 2008-04-15 | 2010-05-27 | Protiva Biotherapeutics, Inc. | Novel lipid formulations for nucleic acid delivery |
| WO2009132131A1 (en) | 2008-04-22 | 2009-10-29 | Alnylam Pharmaceuticals, Inc. | Amino lipid based improved lipid formulation |
| US20100061058A1 (en) | 2008-09-11 | 2010-03-11 | Gen Tanabe | Electrical unit cooling structure for motorcycles |
| US20110256175A1 (en) | 2008-10-09 | 2011-10-20 | The University Of British Columbia | Amino lipids and methods for the delivery of nucleic acids |
| WO2010042877A1 (en) | 2008-10-09 | 2010-04-15 | Tekmira Pharmaceuticals Corporation | Improved amino lipids and methods for the delivery of nucleic acids |
| WO2010048536A2 (en) | 2008-10-23 | 2010-04-29 | Alnylam Pharmaceuticals, Inc. | Processes for preparing lipids |
| WO2010053572A2 (en) | 2008-11-07 | 2010-05-14 | Massachusetts Institute Of Technology | Aminoalcohol lipidoids and uses thereof |
| US20120058144A1 (en) | 2008-11-10 | 2012-03-08 | Alnylam Pharmaceuticals, Inc. | Lipids and compositions for the delivery of therapeutics |
| WO2010054401A1 (en) | 2008-11-10 | 2010-05-14 | Alnylam Pharmaceuticals, Inc. | Novel lipids and compositions for the delivery of therapeutics |
| WO2010054405A1 (en) | 2008-11-10 | 2010-05-14 | Alnylam Pharmaceuticals, Inc. | Novel lipids and compositions for the delivery of therapeutics |
| WO2010054384A1 (en) | 2008-11-10 | 2010-05-14 | Alnylam Pharmaceuticals, Inc. | Lipids and compositions for the delivery of therapeutics |
| US20120027796A1 (en) | 2008-11-10 | 2012-02-02 | Alnylam Pharmaceuticals, Inc. | Novel lipids and compositions for the delivery of therapeutics |
| WO2010054406A1 (en) | 2008-11-10 | 2010-05-14 | Alnylam Pharmaceuticals, Inc. | Novel lipids and compositions for the delivery of therapeutics |
| US8734853B2 (en) | 2008-11-17 | 2014-05-27 | University Of North Texas Health Science Center At Fort Worth | HDL particles for delivery of nucleic acids |
| US20120101148A1 (en) | 2009-01-29 | 2012-04-26 | Alnylam Pharmaceuticals, Inc. | lipid formulation |
| WO2010088537A2 (en) | 2009-01-29 | 2010-08-05 | Alnylam Pharmaceuticals, Inc. | Improved lipid formulation |
| US20120128760A1 (en) | 2009-05-05 | 2012-05-24 | Alnylam Pharmaceuticals, Inc. | Lipid compositions |
| WO2010129709A1 (en) | 2009-05-05 | 2010-11-11 | Alnylam Pharmaceuticals, Inc. | Lipid compositions |
| US8158601B2 (en) | 2009-06-10 | 2012-04-17 | Alnylam Pharmaceuticals, Inc. | Lipid formulation |
| US20100324120A1 (en) | 2009-06-10 | 2010-12-23 | Jianxin Chen | Lipid formulation |
| WO2010144740A1 (en) | 2009-06-10 | 2010-12-16 | Alnylam Pharmaceuticals, Inc. | Improved lipid formulation |
| WO2011000106A1 (en) | 2009-07-01 | 2011-01-06 | Protiva Biotherapeutics, Inc. | Improved cationic lipids and methods for the delivery of therapeutic agents |
| WO2011000107A1 (en) | 2009-07-01 | 2011-01-06 | Protiva Biotherapeutics, Inc. | Novel lipid formulations for delivery of therapeutic agents to solid tumors |
| US20120202871A1 (en) | 2009-07-01 | 2012-08-09 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods for the delivery of therapeutic agents |
| US20110076335A1 (en) | 2009-07-01 | 2011-03-31 | Protiva Biotherapeutics, Inc. | Novel lipid formulations for delivery of therapeutic agents to solid tumors |
| US20120149894A1 (en) | 2009-08-20 | 2012-06-14 | Mark Cameron | Novel cationic lipids with various head groups for oligonucleotide delivery |
| WO2011022460A1 (en) | 2009-08-20 | 2011-02-24 | Merck Sharp & Dohme Corp. | Novel cationic lipids with various head groups for oligonucleotide delivery |
| US20130065939A1 (en) | 2009-09-23 | 2013-03-14 | Protiva Biotherapeutics, Inc. | Compositions and methods for silencing genes expressed in cancer |
| WO2011038160A2 (en) | 2009-09-23 | 2011-03-31 | Protiva Biotherapeutics, Inc. | Compositions and methods for silencing genes expressed in cancer |
| WO2011066651A1 (en) | 2009-12-01 | 2011-06-09 | Protiva Biotherapeutics, Inc. | Snalp formulations containing antioxidants |
| US20130338210A1 (en) | 2009-12-07 | 2013-12-19 | Alnylam Pharmaceuticals, Inc. | Compositions for nucleic acid delivery |
| WO2011071860A2 (en) | 2009-12-07 | 2011-06-16 | Alnylam Pharmaceuticals, Inc. | Compositions for nucleic acid delivery |
| WO2011090965A1 (en) | 2010-01-22 | 2011-07-28 | Merck Sharp & Dohme Corp. | Novel cationic lipids for oligonucleotide delivery |
| WO2011127255A1 (en) | 2010-04-08 | 2011-10-13 | Merck Sharp & Dohme Corp. | Preparation of lipid nanoparticles |
| US20130156845A1 (en) | 2010-04-29 | 2013-06-20 | Alnylam Pharmaceuticals, Inc. | Lipid formulated single stranded rna |
| WO2011141705A1 (en) | 2010-05-12 | 2011-11-17 | Protiva Biotherapeutics, Inc. | Novel cationic lipids and methods of use thereof |
| US20130116307A1 (en) | 2010-05-12 | 2013-05-09 | Protiva Biotherapeutics Inc. | Novel cyclic cationic lipids and methods of use |
| US20130123338A1 (en) | 2010-05-12 | 2013-05-16 | Protiva Biotherapeutics, Inc. | Novel cationic lipids and methods of use thereof |
| WO2011141704A1 (en) | 2010-05-12 | 2011-11-17 | Protiva Biotherapeutics, Inc | Novel cyclic cationic lipids and methods of use |
| US20130090372A1 (en) | 2010-06-04 | 2013-04-11 | Brian W. Budzik | Novel Low Molecular Weight Cationic Lipids for Oligonucleotide Delivery |
| WO2011153120A1 (en) | 2010-06-04 | 2011-12-08 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| US20130303587A1 (en) | 2010-06-30 | 2013-11-14 | Protiva Biotherapeutics, Inc. | Non-liposomal systems for nucleic acid delivery |
| WO2012000104A1 (en) | 2010-06-30 | 2012-01-05 | Protiva Biotherapeutics, Inc. | Non-liposomal systems for nucleic acid delivery |
| US20130323269A1 (en) | 2010-07-30 | 2013-12-05 | Muthiah Manoharan | Methods and compositions for delivery of active agents |
| WO2012016184A2 (en) | 2010-07-30 | 2012-02-02 | Alnylam Pharmaceuticals, Inc. | Methods and compositions for delivery of active agents |
| WO2012024526A2 (en) | 2010-08-20 | 2012-02-23 | Cerulean Pharma Inc. | Conjugates, particles, compositions, and related methods |
| US20130202684A1 (en) | 2010-08-31 | 2013-08-08 | Lichtstrasse | Pegylated liposomes for delivery of immunogen encoding rna |
| WO2012031043A1 (en) | 2010-08-31 | 2012-03-08 | Novartis Ag | Pegylated liposomes for delivery of immunogen-encoding rna |
| WO2012040184A2 (en) | 2010-09-20 | 2012-03-29 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| US20130178541A1 (en) | 2010-09-20 | 2013-07-11 | Matthew G. Stanton | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012044638A1 (en) | 2010-09-30 | 2012-04-05 | Merck Sharp & Dohme Corp. | Low molecular weight cationic lipids for oligonucleotide delivery |
| US20130274523A1 (en) | 2010-09-30 | 2013-10-17 | John A. Bawiec, III | Low molecular weight cationic lipids for oligonucleotide delivery |
| WO2012054365A2 (en) | 2010-10-21 | 2012-04-26 | Merck Sharp & Dohme Corp. | Novel low molecular weight cationic lipids for oligonucleotide delivery |
| US20130274504A1 (en) | 2010-10-21 | 2013-10-17 | Steven L. Colletti | Novel Low Molecular Weight Cationic Lipids For Oligonucleotide Delivery |
| WO2012099755A1 (en) | 2011-01-11 | 2012-07-26 | Alnylam Pharmaceuticals, Inc. | Pegylated lipids and their use for drug delivery |
| US20140200257A1 (en) | 2011-01-11 | 2014-07-17 | Alnylam Pharmaceuticals, Inc. | Pegylated lipids and their use for drug delivery |
| WO2012162210A1 (en) | 2011-05-26 | 2012-11-29 | Merck Sharp & Dohme Corp. | Ring constrained cationic lipids for oligonucleotide delivery |
| US20190321489A1 (en) | 2011-06-08 | 2019-10-24 | Translate Bio, Inc. | Lipid nanoparticle compositions and methods for mrna delivery |
| US20190314284A1 (en) | 2011-06-08 | 2019-10-17 | Translate Bio, Inc. | Cleavable Lipids |
| US20140141070A1 (en) | 2011-07-06 | 2014-05-22 | Andrew Geall | Liposomes having useful n:p ratio for delivery of rna molecules |
| WO2013006825A1 (en) | 2011-07-06 | 2013-01-10 | Novartis Ag | Liposomes having useful n:p ratio for delivery of rna molecules |
| WO2013016058A1 (en) | 2011-07-22 | 2013-01-31 | Merck Sharp & Dohme Corp. | Novel bis-nitrogen containing cationic lipids for oligonucleotide delivery |
| US20140255472A1 (en) | 2011-08-31 | 2014-09-11 | Andrew Geall | Pegylated liposomes for delivery of immunogen-encoding rna |
| WO2013033563A1 (en) | 2011-08-31 | 2013-03-07 | Novartis Ag | Pegylated liposomes for delivery of immunogen-encoding rna |
| US20150203446A1 (en) | 2011-09-27 | 2015-07-23 | Takeda Pharmaceutical Company Limited | Di-aliphatic substituted pegylated lipids |
| WO2013049328A1 (en) | 2011-09-27 | 2013-04-04 | Alnylam Pharmaceuticals, Inc. | Di-aliphatic substituted pegylated lipids |
| WO2013093648A2 (en) | 2011-11-04 | 2013-06-27 | Nitto Denko Corporation | Method of producing lipid nanoparticles for drug delivery |
| US20130164400A1 (en) | 2011-11-04 | 2013-06-27 | Nitto Denko Corporation | Single use system for sterilely producing lipid-nucleic acid particles |
| US9708628B2 (en) | 2011-11-18 | 2017-07-18 | Nof Corporation | Cationic lipid having improved intracellular kinetics |
| WO2013086373A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Lipids for the delivery of active agents |
| US20130195920A1 (en) | 2011-12-07 | 2013-08-01 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| WO2013086354A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| US20140308304A1 (en) | 2011-12-07 | 2014-10-16 | Alnylam Pharmaceuticals, Inc. | Lipids for the delivery of active agents |
| WO2013086322A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Branched alkyl and cycloalkyl terminated biodegradable lipids for the delivery of active agents |
| US20140039032A1 (en) | 2011-12-12 | 2014-02-06 | Kyowa Hakko Kirin Co., Ltd. | Lipid nano particles comprising cationic lipid for drug delivery system |
| WO2013089151A1 (en) | 2011-12-12 | 2013-06-20 | 協和発酵キリン株式会社 | Lipid nanoparticles for drug delivery system containing cationic lipids |
| WO2013116126A1 (en) | 2012-02-01 | 2013-08-08 | Merck Sharp & Dohme Corp. | Novel low molecular weight, biodegradable cationic lipids for oligonucleotide delivery |
| WO2013126803A1 (en) | 2012-02-24 | 2013-08-29 | Protiva Biotherapeutics Inc. | Trialkyl cationic lipids and methods of use thereof |
| US20150064242A1 (en) | 2012-02-24 | 2015-03-05 | Protiva Biotherapeutics, Inc. | Trialkyl cationic lipids and methods of use thereof |
| US20150057373A1 (en) | 2012-03-27 | 2015-02-26 | Sirna Therapeutics, Inc | DIETHER BASED BIODEGRADABLE CATIONIC LIPIDS FOR siRNA DELIVERY |
| WO2013148541A1 (en) | 2012-03-27 | 2013-10-03 | Merck Sharp & Dohme Corp. | DIETHER BASED BIODEGRADABLE CATIONIC LIPIDS FOR siRNA DELIVERY |
| WO2013149140A1 (en) | 2012-03-29 | 2013-10-03 | Shire Human Genetic Therapies, Inc. | Ionizable cationic lipids |
| US9765022B2 (en) | 2013-02-28 | 2017-09-19 | Tufts University | Disulfide compounds for delivery of pharmaceutical agents |
| WO2014136086A1 (en) | 2013-03-08 | 2014-09-12 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| US20160151284A1 (en) | 2013-07-23 | 2016-06-02 | Protiva Biotherapeutics, Inc. | Compositions and methods for delivering messenger rna |
| US20150140070A1 (en) | 2013-10-22 | 2015-05-21 | Shire Human Genetic Therapies, Inc. | Lipid formulations for delivery of messenger rna |
| WO2015061467A1 (en) | 2013-10-22 | 2015-04-30 | Shire Human Genetic Therapies, Inc. | Lipid formulations for delivery of messenger rna |
| US20170114010A1 (en) | 2013-11-18 | 2017-04-27 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid for rna delivery |
| US20150239926A1 (en) | 2013-11-18 | 2015-08-27 | Arcturus Therapeutics, Inc. | Asymmetric ionizable cationic lipid for rna delivery |
| US20150141678A1 (en) | 2013-11-18 | 2015-05-21 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid for rna delivery |
| WO2015074085A1 (en) | 2013-11-18 | 2015-05-21 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid for rna delivery |
| WO2015095346A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2015095340A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| US20160311759A1 (en) | 2013-12-19 | 2016-10-27 | Luis Brito | Lipids and Lipid Compositions for the Delivery of Active Agents |
| US20160317458A1 (en) | 2013-12-19 | 2016-11-03 | Luis Brito | Lipids and Lipid Compositions for the Delivery of Active Agents |
| WO2015130584A2 (en) | 2014-02-25 | 2015-09-03 | Merck Sharp & Dohme Corp. | Lipid nanoparticle vaccine adjuvants and antigen delivery systems |
| WO2015199952A1 (en) | 2014-06-25 | 2015-12-30 | Acuitas Therapeutics Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| US20150376115A1 (en) | 2014-06-25 | 2015-12-31 | Acuitas Therapeutics Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2016081029A1 (en) | 2014-11-18 | 2016-05-26 | Arcturus Therapeutics, Inc. | Ionizable cationic lipid for rna delivery |
| US20180005363A1 (en) | 2015-01-30 | 2018-01-04 | Hitachi High-Technologies Corporation | Pattern Matching Device and Computer Program for Pattern Matching |
| US20180053569A1 (en) | 2015-05-04 | 2018-02-22 | Huawei Technologies Co., Ltd. | DRAM Refresh Method, Apparatus, and System |
| WO2017004143A1 (en) | 2015-06-29 | 2017-01-05 | Acuitas Therapeutics Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| US20160376224A1 (en) | 2015-06-29 | 2016-12-29 | Acuitas Therapeutics, Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| US20190091164A1 (en) | 2015-08-21 | 2019-03-28 | Pfizer Inc. | Therapeutic Nanoparticles Comprising A Therapeutic Agent And Methods of Making And Using Same |
| US20170210697A1 (en) | 2015-09-17 | 2017-07-27 | Modernatx, Inc. | Compounds and compositions for intracellular delivery of therapeutic agents |
| WO2017049245A2 (en) | 2015-09-17 | 2017-03-23 | Modernatx, Inc. | Compounds and compositions for intracellular delivery of therapeutic agents |
| US20170119904A1 (en) | 2015-10-28 | 2017-05-04 | Acuitas Therapeutics, Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017075531A1 (en) | 2015-10-28 | 2017-05-04 | Acuitas Therapeutics, Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| US20190314524A1 (en) | 2015-10-28 | 2019-10-17 | Acuitas Therapeutics, Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| US20180028664A1 (en) | 2015-12-10 | 2018-02-01 | Modernatx, Inc. | Compositions and methods for delivery of agents |
| WO2017099823A1 (en) | 2015-12-10 | 2017-06-15 | Modernatx, Inc. | Compositions and methods for delivery of therapeutic agents |
| US20170190661A1 (en) | 2015-12-30 | 2017-07-06 | Arcturus Therapeutics, Inc. | Aromatic Ionizable Cationic Lipid |
| WO2017117528A1 (en) | 2015-12-30 | 2017-07-06 | Acuitas Therapeutics, Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017173054A1 (en) | 2016-03-30 | 2017-10-05 | Intellia Therapeutics, Inc. | Lipid nanoparticle formulations for crispr/cas components |
| US20190345503A1 (en) | 2016-06-20 | 2019-11-14 | The Board Of The Leland Stanford Junior University | Circular rnas and their use in immunomodulation |
| WO2018011633A1 (en) | 2016-07-13 | 2018-01-18 | Alcatel Lucent | Underlying recessed component placement |
| US20200038678A1 (en) | 2017-02-08 | 2020-02-06 | Michelson Diagnostics Limited | Processing optical coherence tomography (oct) scans |
| US20190015913A1 (en) | 2017-07-12 | 2019-01-17 | Rexon Industrial Corp., Ltd. | Miter saw with enhanced adjustable miter angle |
| WO2019131770A1 (en) | 2017-12-27 | 2019-07-04 | 武田薬品工業株式会社 | Nucleic acid-containing lipid nano-particle and use thereof |
| WO2019236673A1 (en) | 2018-06-06 | 2019-12-12 | Massachusetts Institute Of Technology | Circular rna for translation in eukaryotic cells |
| WO2020237227A1 (en) | 2019-05-22 | 2020-11-26 | Massachusetts Institute Of Technology | Circular rna compositions and methods |
| WO2021021634A1 (en) | 2019-07-29 | 2021-02-04 | Georgia Tech Research Corporation | Nanomaterials containing constrained lipids and uses thereof |
| WO2021041473A1 (en) | 2019-08-27 | 2021-03-04 | The Trustees Of Columbia University In The City Of New York | Engineered exosomes for targeted delivery |
| US20220288176A1 (en) | 2019-08-28 | 2022-09-15 | The Board Of Trustees Of The Leland Stanford Junior University | Circular rna modification and methods of use |
| WO2021113777A2 (en) | 2019-12-04 | 2021-06-10 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021189059A2 (en) | 2020-03-20 | 2021-09-23 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021226597A2 (en) | 2020-05-08 | 2021-11-11 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2021236855A1 (en) | 2020-05-19 | 2021-11-25 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| US20220251578A1 (en) | 2020-06-25 | 2022-08-11 | The Board Of Trustees Of The Leland Stanford Junior University | Genetic elements driving circular rna translation and methods of use |
| US20220033091A1 (en) | 2020-07-30 | 2022-02-03 | Hamilton Sundstrand Corporation | Aircraft environmental control system |
| WO2022061490A1 (en) | 2020-09-22 | 2022-03-31 | 华为数字能源技术有限公司 | Photovoltaic power generation system, and method and device for detecting ground fault in photovoltaic strings |
| WO2022261490A2 (en) | 2021-06-10 | 2022-12-15 | Orna Therapeutics, Inc. | Circular rna compositions and methods |
| WO2022271965A2 (en) | 2021-06-25 | 2022-12-29 | The Board Of Trustees Of The Leland Stanford Junior University | Compositions and methods for improved protein translation from recombinant circular rnas |
| WO2023056033A1 (en) | 2021-09-30 | 2023-04-06 | Orna Therapeutics, Inc. | Lipid nanoparticle compositions for delivering circular polynucleotides |
| WO2023081526A1 (en) | 2021-11-08 | 2023-05-11 | Orna Therapeutics, Inc. | Lipid nanoparticle compositions for delivering circular polynucleotides |
| WO2023141586A1 (en) | 2022-01-21 | 2023-07-27 | Orna Therapeutics, Inc. | Systemic administration of circular rna polynucleotides encoding muscle proteins or protein complexes |
| WO2023250375A1 (en) | 2022-06-24 | 2023-12-28 | Orna Therapeutics, Inc. | Circular rna encoding chimeric antigen receptors targeting bcma |
Non-Patent Citations (80)
| Title |
|---|
| "Encyclopedia of Reagents for Organic Synthesis", 2009, JOHN WILEY AND SONS |
| "Medical Applications of Liposomes.", 1998, ELSEVIER |
| "T cell Receptor Factsbook", 2001, ACADEMIC PRESS, article "LeFranc and LeFranc" |
| ACTA CRYSTALLOGR D BIOL CRYSTALLOGR, vol. 56, no. 10, 2000, pages 1316 - 1323 |
| BEHR ET AL., PROC. NAT.'L ACAD. SCI., vol. 86, 1989, pages 6982 |
| BRICOGNE G, ACTA CRYSTALLOGR D BIOL CRYSTALLOGR, vol. 49, no. 1, 1993, pages 37 - 60 |
| CHAYEN NE, STRUCTURE, vol. 5, 1997, pages 1269 - 1274 |
| CHEN ET AL., NATURE BIOTECHNOLOGY, 2022 |
| CHEUNG ET AL., VIROLOGY, vol. 176, 1990, pages 546 - 552 |
| DHAMNE ET AL.: "Peripheral and thymic Foxp3+ regulatory T cells in search of origin, distinction, and function", FRONTIERS IN IMMUNOL., vol. 4, no. 253, 2013, pages 1 - 11 |
| DOBRIKOVA ET AL., PROC. NATL. ACAD. SCI., vol. 100, no. 25, 2003, pages 15125 - 15130 |
| EDGE, NATURE, vol. 292, 1981, pages 756 |
| ELIEL: "Stereochemistry of Carbon Compounds", 1962, MCGRAW-HILL |
| ESHHAR ET AL., CANCER IMMUNOL IMMUNOTHERAPY, vol. 45, 1997, pages 131 - 136 |
| FELGNER ET AL., PROC. NAT'L ACAD. SCI., vol. 84, 1987, pages 7413 |
| FINNEY ET AL., JOURNAL OF IMMUNOLOGY, vol. 161, 1998, pages 2791 - 2797 |
| FRIESS M ET AL., FRONT. IMMUNOL., vol. 2947, no. 9, 2018 |
| GAO ET AL., BIOCHEM. BIOPHYS. RES. COMM., vol. 179, 1991, pages 280 |
| GARLAPATI ET AL., J. BIOL. CHEM., vol. 279, no. 5, 2004, pages 3389 - 3397 |
| GIEGE R ET AL., ACTA CRYSTALLOGR D BIOL CRYSTALLOGR, vol. 50, no. 4, 1994, pages 339 - 350 |
| GREEN, T. W.P. G. M. WUTZ: "Protective Groups in Organic Synthesis", 1999, VCH PUBLISHERS |
| GROSS ET AL., AMUR. REV. PHARMACOL. TOXICOL., vol. 56, 2016, pages 59 - 83 |
| GURTU ET AL., BIOCHEM. BIOPHYS. RES. COMM., vol. 229, 1996, pages 295 - 298 |
| HEYES, J. ET AL., J CONTROLLED RELEASE, vol. 107, 2005, pages 276 - 287 |
| HILLERY A M ET AL.: "Drug Delivery and Targeting: For Pharmacists and Pharmaceutical Scientists", 2002, TAYLOR & FRANCIS, INC. |
| J CONTROL RELEASE., vol. 207, 2015, pages 18 - 30 |
| J. PHARMACEUTICAL SCIENCES, vol. 66, 1977, pages 1 - 19 |
| JANEWAY C ET AL.: "Garland Science", 2001, article "Immunobiology: The Immune System in Health and Disease" |
| JANG ET AL., J. VIROL., vol. 63, 1989, pages 1651 - 1660 |
| JAY ET AL., J. BIOL. CHEM., vol. 259, 1984, pages 631 1 |
| JAYARAMA ET AL.: "Angew. Chem.", vol. 51, 2012, pages: 8529 - 8533 |
| JAYARAMAN ET AL., PROC. NATL. ACAD. SCI. USA, vol. 88, 1991, pages 4084 - 4088 |
| JONES ET AL., NATURE, vol. 54, 1986, pages 75 - 82 |
| KALOS ET AL., SCI TRANSL. MED., vol. 3, no. 95, 2011 |
| KARIKO ET AL., IMMUNITY, 2005 |
| KARIKO ET AL., MOL THER, 2005 |
| KAUFMAN ET AL., NUC. ACIDS RES., vol. 19, 1991, pages 4485 - 4490 |
| KIRKLAND ET AL., J. IMMUNOL., vol. 137, 1986, pages 3614 - 3619 |
| KLIBANOV ET AL., FEBS LETTERS, vol. 268, no. 1, 1990, pages 235 - 237 |
| KOBAYASHI ET AL., BIOTECHNIQUES, vol. 21, 1996, pages 399 - 402 |
| KOLTOVER ET AL., SCIENCE, vol. 281, 1998, pages 78 - 81 |
| KRAUSE ET AL., J. EXP. MED., vol. 188, no. 4, 1998, pages 619 - 626 |
| KUBALL JET, J EXP MED, vol. 206, no. 2, 2009, pages 463 - 475 |
| L. FIESERM. FIESER: "Fieser and Fieser' s Reagents for Organic Synthesis", 1994, JOHN WILEY AND SONS |
| LASIC ET AL., FEBS LETT., vol. 312, 1992, pages 255 - 258 |
| LEHTIMAKILAHESMAA: "Regulatory T cells control immune responses through their non-redundant tissue specific features", FRONTIERS IN IMMUNOL., vol. 4, no. 294, 2013, pages 1 - 10 |
| LUAN ET AL., ACTA PHARMACOLOGICA SINICA, vol. 38, 2017, pages 754 - 763 |
| MAIER ET AL., MOLECULAR THERAPY, vol. 21, 2013, pages 1570 - 1578 |
| MAIER, M.A. ET AL.: "Biodegradable Lipids Enabling Rapidly Eliminated Lipid Nanoparticles for Systemic Delivery of RNAi Therapeutics", MOL. THER., vol. 21, no. 8, 2013, pages 1570 - 78, XP055237159, DOI: 10.1038/mt.2013.124 |
| MCPHERSON A, EUR J BIOCHEM, vol. 189, 1990, pages 1 - 23 |
| MCPHERSON A, J BIOL CHEM, vol. 251, 1976, pages 6300 - 6303 |
| MENDES L ET AL., MOLECULES, vol. 22, no. 9, 2017, pages 1401 |
| MOLDENHAUER ET AL., J. IMMUNOL., vol. 32, 1990, pages 77 - 82 |
| MOREL ET AL., MOLEC. IMMUNOL., vol. 25, 1988, pages 7 - 15 |
| MORRISSEY, D V. ET AL., NAT. BIOTECHNOL., vol. 23, no. 8, 2005, pages 1003 - 1007 |
| MOSSER ET AL., BIOTECHNIQUES, vol. 114,115, 1997, pages 139 - 161 |
| MUKHERJEE ET AL., IN J. NANOMEDICINE., vol. 14, 2019, pages 1937 - 1952 |
| NAMBAIR ET AL., SCIENCE, vol. 223, 1984, pages 1299 |
| P. G. M. WUTST. W. GREENE: "Greene's Protective Groups in Organic Synthesis", 2007, JOHN WILEY AND SONS |
| PORTER ET AL., N. ENGL. J. MED., vol. 365, 2011, pages 725 - 33 |
| QUEEN ET AL., PROC. NATL. ACAD. SCI. USA, vol. 86, 1989, pages 10029 - 10033 |
| RAMESH ET AL., NUCL. ACID RES., vol. 24, 1996, pages 2697 - 2700 |
| RIECHMANN ET AL., NATURE, vol. 332, 1988, pages 323 - 327 |
| ROBBINS ET AL., J IMMUNOL., vol. 180, pages 6116 - 6131 |
| ROSENBERG ET AL., NAT REV CANCER, vol. 8, no. 4, 2008, pages 299 - 308 |
| S. LI ET AL., GENE THER., vol. 4, 1997, pages 891 |
| SEE, N. J. CAPLEN ET AL., GENE THER., vol. 2, 1995, pages 603 |
| SEMPLE ET AL., NAT. BIOTECHNOL., vol. 28, 2010, pages 172 - 176 |
| SEMPLE ET AL., NATURE BIOTECH., vol. 28, 2010, pages 172 - 176 |
| SHOBAKI NSATO YHARASHIMA H: "Mixing lipids to manipulate the ionization status of lipid nanoparticles for specific tissue targeting", INT J NANOMEDICINE, vol. 13, 2018, pages 8395 - 8410, XP055924551, DOI: 10.2147/IJN.S188016 |
| SON ET AL., NANO LETT., vol. 20, no. 3, 2020, pages 1499 - 1509 |
| SONG ET AL., BLOOD, vol. 119, 2012, pages 696 - 706 |
| STAHLI ET AL., METHODS IN ENZYMOLOGY, vol. 9, 1983, pages 242 - 253 |
| VERHOEYEN ET AL., SCIENCE, vol. 239, 1988, pages 1534 - 1536 |
| WADWA ET AL., J, DRUG TARGETING, vol. 3, 1995, pages 111 |
| WANG ET AL., ACS SYNTHETIC BIOLOGY, vol. 1, 2012, pages 403 - 07 |
| WENDER ET AL., ADV. DRUG DEL. REV., vol. 60, 2008, pages 452 - 472 |
| WESSELHOEFT ET AL., MOL CELL, 2019 |
| WILEN ET AL., TETRAHEDRON, vol. 33, 1977, pages 2725 |
| WILEN: "Tables of Resolving Agents and Optical Resolutions", 1972, UNIV. OF NOTRE DAME PRESS, pages: 268 |
Cited By (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2025259931A1 (en) * | 2024-06-14 | 2025-12-18 | Orbital Therapeutics, Inc. | Compositions and methods for rna circularization |
Also Published As
| Publication number | Publication date |
|---|---|
| WO2024233308A3 (en) | 2025-01-16 |
| AU2024269222A1 (en) | 2025-10-09 |
| WO2024233308A9 (en) | 2025-04-10 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US12384741B2 (en) | Lipid nanoparticle compositions for delivering circular polynucleotides | |
| US20250283073A1 (en) | Circular RNA Compositions and Methods | |
| EP4408825A1 (en) | Lipid nanoparticle compositions for delivering circular polynucleotides | |
| US12297285B2 (en) | Circular RNA encoding chimeric antigen receptors targeting BCMA | |
| AU2023398825A1 (en) | Circular rna compositions and methods | |
| WO2024102762A1 (en) | Lipids and lipid nanoparticle compositions for delivering polynucleotides | |
| WO2024102677A1 (en) | Circular rna compositions | |
| AU2024269222A1 (en) | Circular rna compositions and methods | |
| WO2024205657A2 (en) | Lipids and lipid nanoparticle compositions for delivering polynucleotides | |
| WO2026006593A1 (en) | Synthetic internal ribosome entry sites | |
| WO2025250751A1 (en) | Circular rna compositions and methods | |
| WO2025117969A1 (en) | Process for manufacturing lipid nanoparticles | |
| WO2025101501A1 (en) | Circular rna compositions | |
| CN118103514A (en) | Cyclic RNA compositions and methods | |
| EA049788B1 (en) | CIRCULAR RNA COMPOSITIONS AND METHODS |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24729520 Country of ref document: EP Kind code of ref document: A2 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: AU2024269222 Country of ref document: AU |
|
| ENP | Entry into the national phase |
Ref document number: 2024269222 Country of ref document: AU Date of ref document: 20240503 Kind code of ref document: A |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2024729520 Country of ref document: EP |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2024729520 Country of ref document: EP Effective date: 20251205 |
|
| ENP | Entry into the national phase |
Ref document number: 2024729520 Country of ref document: EP Effective date: 20251205 |