WO2025085810A2 - Antisense oligomers for treatment of centronuclear myopathies - Google Patents
Antisense oligomers for treatment of centronuclear myopathies Download PDFInfo
- Publication number
- WO2025085810A2 WO2025085810A2 PCT/US2024/052063 US2024052063W WO2025085810A2 WO 2025085810 A2 WO2025085810 A2 WO 2025085810A2 US 2024052063 W US2024052063 W US 2024052063W WO 2025085810 A2 WO2025085810 A2 WO 2025085810A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- dnm2
- seq
- antisense oligomer
- target region
- exon
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
- C12N15/1137—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing against enzymes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/11—Antisense
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/32—Chemical structure of the sugar
- C12N2310/323—Chemical structure of the sugar modified ring structure
- C12N2310/3233—Morpholino-type ring
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/35—Nature of the modification
- C12N2310/351—Conjugate
- C12N2310/3513—Protein; Peptide
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2320/00—Applications; Uses
- C12N2320/30—Special therapeutic applications
- C12N2320/33—Alteration of splicing
Definitions
- Dynamin 2 is a ubiquitously expressed protein that belongs to the superfamily of large GTPases.
- Dynamin 2 acts as a mechanochemical scaffolding molecule that deforms biological membranes leading to the release of nascent vesicles.
- DNM2 acts as a mechanochemical scaffolding molecule that deforms biological membranes leading to the release of nascent vesicles.
- DNM2 acts as a mechanochemical scaffolding molecule that deforms biological membranes leading to the release of nascent vesicles.
- DNM2 is involved in clathrin-dependent and clathrin-independent endocytosis. This protein is also involved in the formation of vesicles from endosomes and trans-Golgi network.
- Several studies have highlighted the role of Dynamin 2 as regulator of actin and microtubule cytoskeletons.
- AD-CNM Autosomal dominant centronuclear myopathy
- CMT Charcot-Marie-Tooth disease
- HSP Hereditary spastic paraplegia
- DNM2 small cell lung cancer
- CCM centronuclear myopathy
- an antisense oligomer comprising a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of human dynamin 2 (DNM2) gene represented by SEQ ID NO: 1, wherein the target region is within an exon internal region, an intron internal region, an intron-exon junction, or an exon-intron junction of the human DNM2 gene pre-mRNA.
- DNM2 human dynamin 2
- the target region is within the exon internal region of exon 2 (SEQ ID NO: 2), the intron 1/exon 2 junction, the exon 2/intron 2 junction, or the intron internal region of intron 2 (SEQ ID NO: 11) of the human DNM2 gene pre-mRNA.
- the target region is within the exon internal region of exon 9 (SEQ ID NO: 6), the intron 8/exon 9 junction, or the exon 9/intron 9 junction of the human DNM2 gene pre-mRNA.
- the target region is within the exon internal region of exon 12 (SEQ ID NO: 7), the intron 11/exon 12 junction, or the exon 12/intron 12 junction of the human DNM2 gene pre-mRNA. [0013] In certain embodiments, the target region is within the exon internal region of exon 13 (SEQ ID NO: 8), the intron 12/exon 13 junction, the exon 13/intron 13 junction, or the intron internal region of intron 12 (SEQ ID NO: 12).
- the target region is within the exon internal region of exon 6 (SEQ ID NO: 3), the intron 5/exon 6 junction, the exon 6/intron 6 junction, or the intron internal region of intron 6 (SEQ ID NO: 11).
- the target region is within the exon internal region of exon 7 (SEQ ID NO: 4), the intron 6/exon 7 junction, or the exon 7/intron 7 junction.
- the target region is within the exon internal region of exon 8 (SEQ ID NO: 5), the intron 7/exon 8 junction, or the exon 8/intron 8 junction.
- the target region is within the exon internal region of exon 16 (SEQ ID NO: 9), the intron 15/exon 16 junction, or the exon 16/intron 16 junction. [0018] In certain embodiments, the target region is within the exon internal region of exon 19 (SEQ ID NO: 9), the intron 18/exon 19 junction, or the exon 18/intron 18 junction.
- an antisense oligomer, or a pharmaceutically acceptable salt thereof comprising: a non-natural chemical backbone selected from a peptide nucleic acid, a locked nucleic acid, a phosphorodiamidate morpholino oligomer, a 2′-OMe phosphorothioate oligomer, or a combination thereof; and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of human dynamin 2 (DNM2) gene represented by SEQ ID NO: 1, wherein the target region is within an exon internal region, an intron internal region, an intron-exon junction, or an exon-intron junction of the human DNM2 gene pre-mRNA.
- DNM2 human dynamin 2
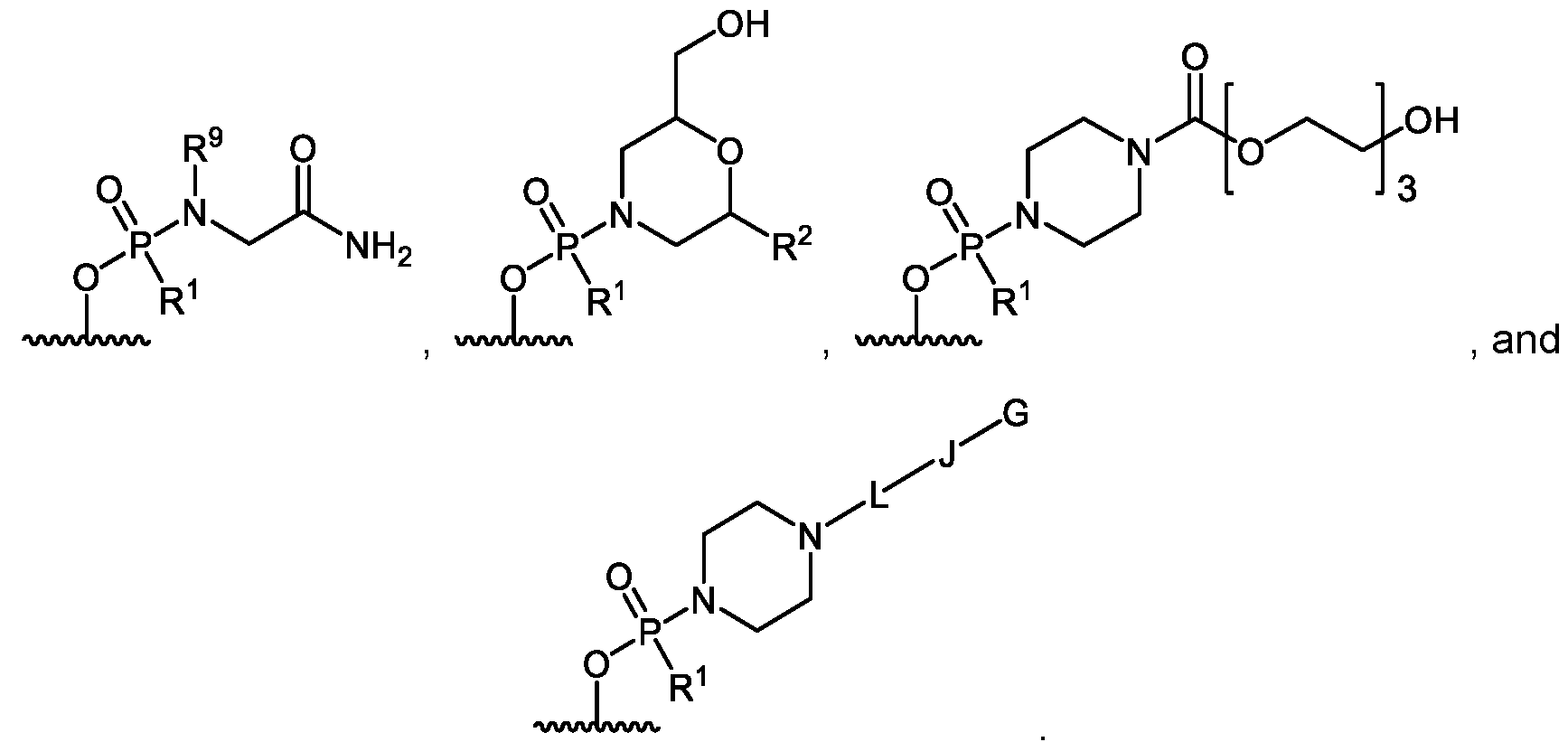
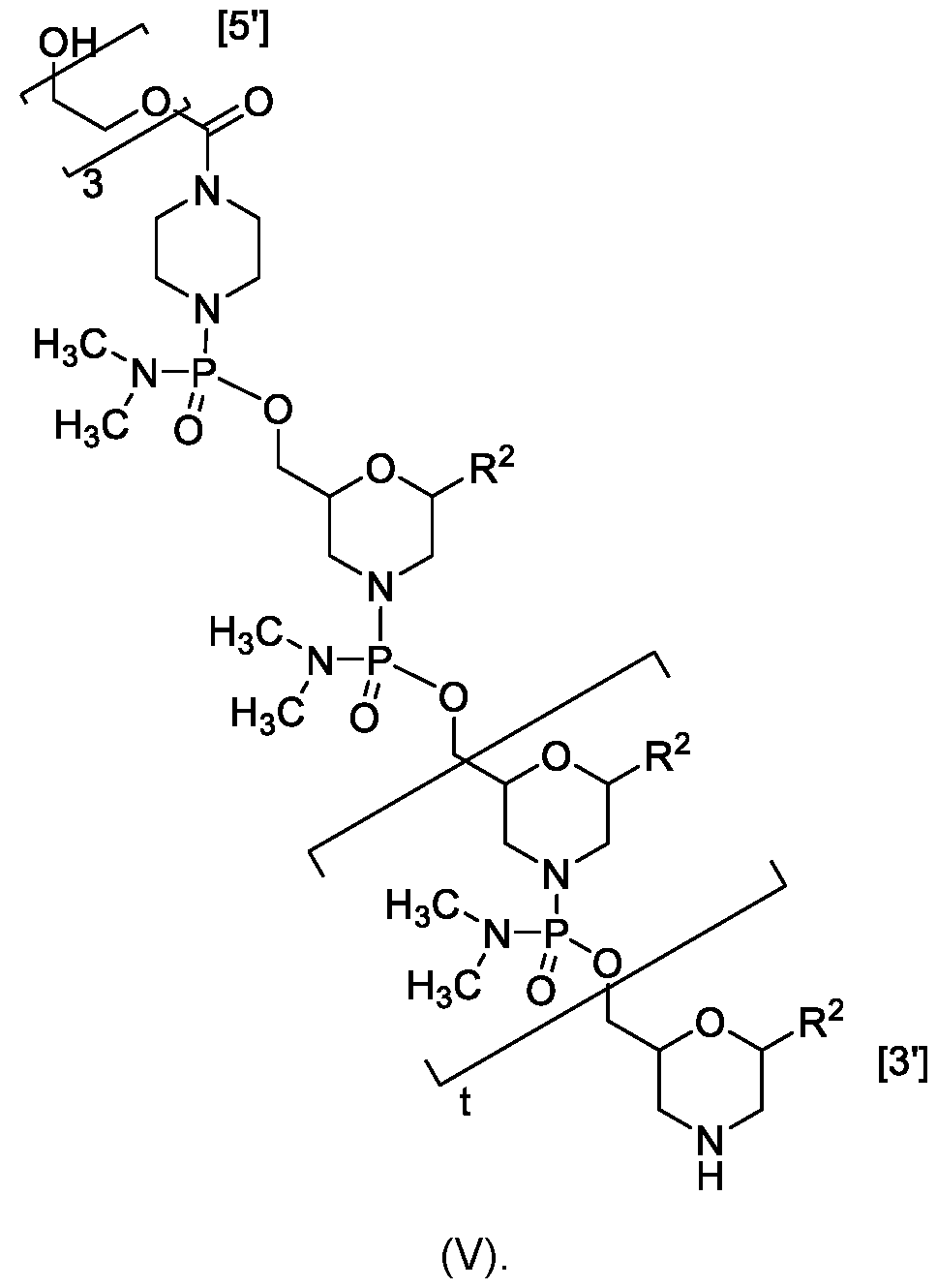
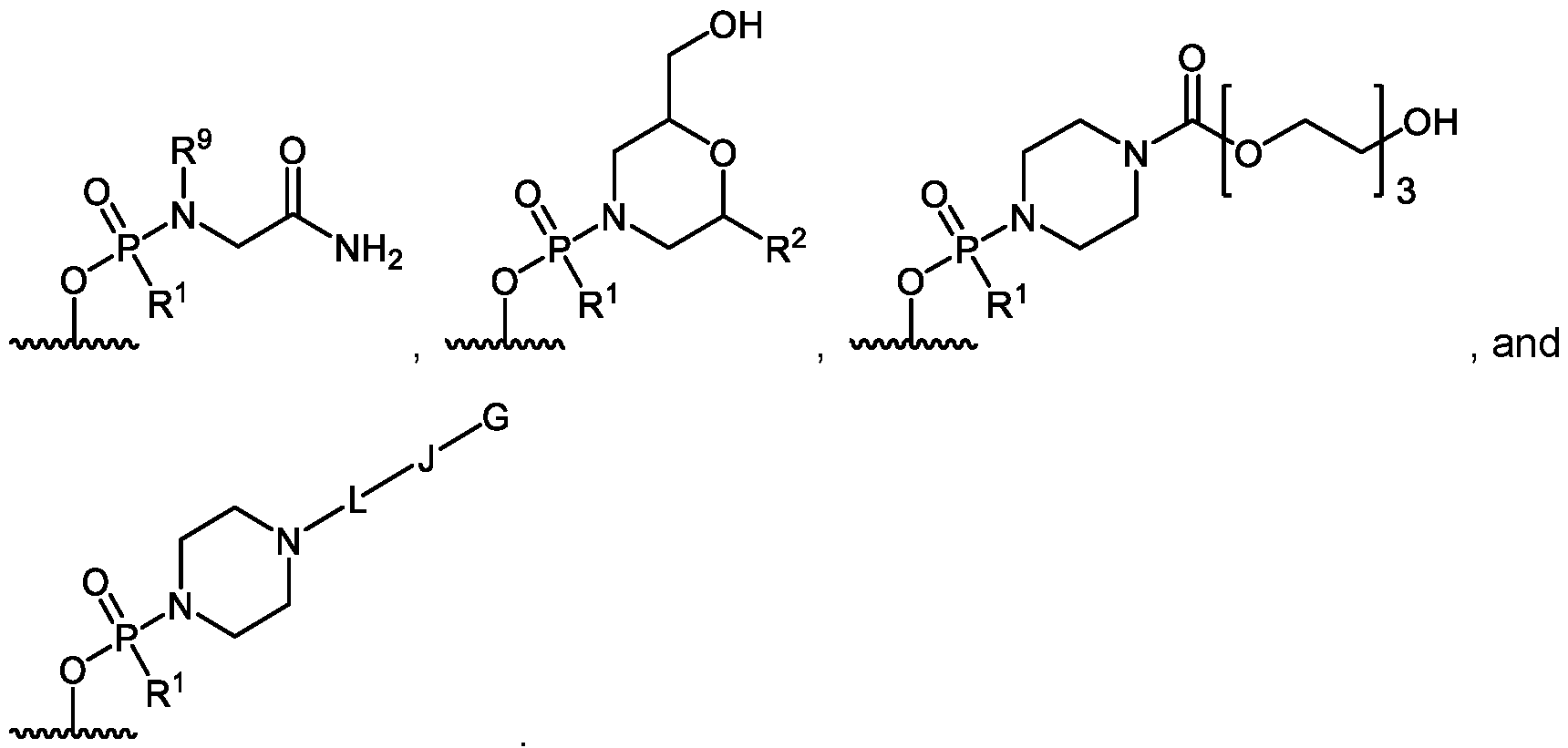
- R 5 is -C(O)(O-alkyl) x -OH, wherein x is 3-10 and each alkyl group is, independently at each occurrence, C 2-6 -alkyl, or R 5 is selected from H, -C(O)C 1-6 -alkyl, trityl, monomethoxytrityl, -(C 1-6 -alkyl)-R 6 , -(C 1-6 - heteroalkyl)-R 6 , -C 6-10 -aryl-R 6 , 5-10 membered heteroaryl-R 6 , -C(O)O-(C 1-6 -alkyl)-R 6 , - C(O)O-aryl-R 6 , -C(O)O-(5-10 membered heteroaryl)-R 6 , and ; R 6 is selected from -OH, -SH, and -NH 2 , or R 6 is O, S, or NH, each of which is
- A′ is a moiety selected from: [0022]
- a pharmaceutical composition comprising: an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formulae (I), (IA), (II), (III), (IV) and/or (V); and a pharmaceutically acceptable carrier.
- a method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein comprising administering to a subject in need thereof a therapeutically effective amount of an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formula (I):
- a method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein comprising administering to a subject in need thereof a therapeutically effective amount of an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formula (III):
- n is 11-28.
- a method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein comprising administering to a subject in need thereof a therapeutically effective amount of an antisense oligomer, or a pharmaceutically acceptable salt thereof, of In an aspect provided herein, is antisense oligomer of Formula (IV): (IV), wherein R 2 and n are as described herein.
- the disease or disorder is a genetic muscle disorder.
- the disease or disorder is a neuromuscular disorder.
- the disease or disorder is a centronuclear myopathy (CNM).
- the disease or disorder is myotubular myopathy (XLMTM or XLCNM), autosomal dominant congenital myopathy (ADCNM), autosomal recessive centronuclear myopathy (ARCNM), or Charcot-Marie- Tooth disease (CMT).
- the CMT is DNM2-implicated CMT.
- the Charcot-Marie-Tooth disease (CMT) is Charcot-Marie-Tooth disease, dominant intermediate B (CMTDIB or DI-CMTB).
- the Charcot- Marie-Tooth disease is Charcot-Marie-Tooth disease, axonal type 2M (CMT2M).
- the disease or disorder is a hereditary spastic paraplegia (HSP).
- HSP hereditary spastic paraplegia
- the antisense oligomers disclosed herein are useful in methods of inducing exon skipping to create premature stop codon in a human DNM2 pre-mRNA, thereby triggering nonsense-mediated decay and reducing DNM2-aggregation in diseased cells. Accordingly, the antisense oligomers described herein are useful for the treatment for various diseases in a subject in need thereof, including, but not limited to, diseases such as a centronuclear myopathy.
- FIGs.1A-1E show normalized DNM2 transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exons 6, 8 and 16.1A-1B: RT-qPCR after treatment with indicated PMO compounds for exon 8.1C: RT-qPCR after treatment with indicated PMO compounds for exon 16.1D-1E: RT-qPCR after treatment with indicated PMO compounds for exon 6.
- FIGs.2A-2F show normalized DNM2 transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exons 2, 7, 9, 12, 13, and 19.2A: RT-qPCR after treatment with indicated PMO compounds for exon 2.2B: RT-qPCR after treatment with indicated PMO compounds for exon 9.2C: RT-qPCR after treatment with indicated PMO compounds for exon 12.2D: RT-qPCR after treatment with indicated PMO compounds for exon 13.2E: RT-qPCR after treatment with indicated PMO compounds for exon 19.2F: RT-qPCR after treatment with indicated PMO compounds for exon 7.
- FIGs.3A-3J show normalized DNM2 transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exons, 2, 9, 12, and 13.3A-3B: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 2.3C-3E: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 9.3F-3G: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 12.3H-3J: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 13.
- FIGs.4A-4G show the dose response of PMOs targeted against exons 2, 9, 12, and 13 as measured by RT-qPCR.4A-4B and G: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 2.4C-4D: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 9.4E: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 12.4F: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 13.
- FIGs.5A-5B show DNM2 exon skipping confirmation after treatment with select compounds targeting exon 2 and exon 9 as measured by RT-PCR.
- 5A photographic depiction of DNA agarose gel electrophoresis of DNM2 exon skipping confirmation after treatment with select compounds targeting exon
- 2.5B photographic depiction of DNA agarose gel electrophoresis of DNM2 exon skipping confirmation after treatment with select compounds targeting exon 9.
- FIGs.6A-6B show treatment with select exon 2 targeting compound results in DNM2 protein reduction.
- 6A photographic depiction of capillary electrophoresis of DNM2 protein reduction in RD Cells after treatment with PPMOs.6B: is a graphic representation of exon 2 targeting compound results in DNM2 protein reduction.
- FIG.7 shows normalized DNM2 transcript levels as measured by RT-qPCR after PPMO treatment in AB1190C16PV myoblast immortalized cell line, differentiation method 1.
- FIG.8 shows normalized DNM2 transcript levels as measured by RT-qPCR after PPMO treatment in AB1190C16PV myoblast immortalized cell line, differentiation method 2.
- FIGs.12A-12G show dose response of normalized DNM2 transcript levels as measured by RT-qPCR with select compounds in GM06813 C1 myotubes.
- 12A dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 2 in GM06813 C1 myotubes.
- 12B dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 3 in GM06813 C1 myotubes.
- 12C dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 4 in GM06813 C1 myotubes.
- 12D dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 5 in GM06813 C1 myotubes.
- 12E dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 6 in GM06813 C1 myotubes.
- 12F dose response of normalized DNM2
- FIGs.13A-13D show DNM2 exon skipping confirmation in myotubes after treatment with indicated PPMO compounds as measured by RT-PCR.
- 13A DNM2 exon skipping confirmation in myotubes after treatment with PPMO 10 as measured by RT-PCR.
- 13B DNM2 exon skipping confirmation in myotubes after treatment with PPMOs 14, 22, and 28 as measured by RT-PCR.
- 13C DNM2 exon skipping confirmation in myotubes after treatment with PPMO 12 as measured by RT-PCR.
- 13D DNM2 exon skipping confirmation in myotubes after treatment with PPMOs 27 and 28 as measured by RT-PCR.
- FIGs.14A-14D show DNM2 exon skipping confirmation in myotubes after treatment with indicated PPMO compounds as measured by RT-PCR.
- 14A DNM2 exon skipping confirmation in myotubes after treatment with PPMO 29 as measured by RT-PCR.
- 14B DNM2 exon skipping confirmation in myotubes after treatment with PPMOs 30, 22, and 28 as measured by RT-PCR.
- 14C DNM2 exon skipping confirmation in myotubes after treatment with PPMOs 1 and 29 as measured by RT-PCR.14D: DNM2 exon skipping confirmation in myotubes after treatment with PPMO 35 as measured by RT-PCR.
- FIGs.15A-15B show DNM2 protein reduction in myotubes after treatment with indicated PPMO compounds.
- 15A photographic depiction of a Western blot of DNM2 protein reduction in myotubes after treatment with PPMOs 10, 27, and 28.
- 15B a graphic representation of DNM2 protein reduction in myotubes after treatment with PPMOs 10, 27, and 28.
- FIGs.16A-16B show DNM2 protein reduction in myotubes after treatment with indicated PPMO compounds.
- 16A photographic depiction of a Western blot of DNM2 protein reduction in myotubes after treatment with PPMOs 1, 29, and 35.
- 16B a graphic representation of DNM2 protein reduction in myotubes after treatment with PPMOs 1, 29, and 35.
- DETAILED DESCRIPTION Provided herein is an antisense oligomer or pharmaceutically acceptable salt thereof, comprising a modified chemical backbone and a targeting sequence of 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of the human DNM2 gene (SEQ ID NO: 1).
- the target region may be an intron/exon junction or an exon internal region of the human DNM2 gene pre-mRNA.
- Such antisense oligomers are useful for the treatment of various diseases in a subject in need thereof, including, but not limited to, centronuclear myopathies (CNM).
- CCM centronuclear myopathies
- Certain embodiments relate to methods of inducing exon skipping to create a premature stop codon on a human DNM2 pre-mRNA in a cell, comprising contacting the cell with an antisense oligomer of sufficient length and complementarity to specifically hybridize to a region within the human DNM2 gene, such that the expression of DNM2 is reduced.
- the cell is in a subject, and the method comprises administering the antisense oligomer to the subject.
- the cell is a diseased cell.
- the antisense oligomer further comprises a cell-penetrating peptide, an antibody, an antibody fragment, or an antigen fragment of an antibody.
- the antisense oligomer comprises a cell penetrating peptide, wherein the cell-penetrating peptide is any of the peptides provided herein or known in the art.
- pharmaceutical compositions and methods for treating diseases or disorders, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein are also provided herein.
- C 1-6 -alkyl moieties include, but are not limited to, methyl, ethyl, propyl, isopropyl, n-butyl, tert-butyl, neopentyl, n-hexyl moieties; and examples of C 1-8 -alkyl moieties include, but are not limited to, methyl, ethyl, propyl, isopropyl, n-butyl, tert-butyl, neopentyl, n- hexyl, heptyl, and octyl moieties.
- the number of carbon atoms in an alkyl substituent can be indicated by the prefix “C x-y ,” or “C x -C y ” where x is the minimum and y is the maximum number of carbon atoms in the substituent.
- a C x chain means an alkyl chain containing x carbon atoms.
- heteroalkyl by itself or in combination with another term means, unless otherwise stated, a stable straight or branched chain alkyl group consisting of the stated number of carbon atoms and up to five heteroatoms selected from O, N, S, and P, wherein the nitrogen, sulfur, and phosphorus atoms may be optionally oxidized, and the nitrogen heteroatom may be optionally quaternized.
- the heteroatom(s) may be placed at any position of the heteroalkyl group, including between the rest of the heteroalkyl group and the fragment to which it is attached, as well as attached to the most distal carbon atom in the heteroalkyl group.
- Up to three heteroatoms may be consecutive, such as, for example, -CH2-NH-OCH3, or -CH2-CH2-S-S-CH3.
- aryl employed alone or in combination with other terms, means, unless otherwise stated, a carbocyclic aromatic system containing one or more rings (typically one, two, or three rings), wherein such rings may be attached together in a pendent manner, such as a biphenyl, or may be fused, such as naphthalene.
- aryl groups include phenyl, anthracyl, and naphthyl.
- examples of an aryl group may include phenyl (i.e., C 6 -aryl) and biphenyl (i.e., C 12 -aryl).
- aryl groups have from six to sixteen carbon atoms (i.e., C 6-16 -aryl). In some embodiments, aryl groups have from six to twelve carbon atoms (i.e., C 6-12 -aryl). In some embodiments, aryl groups have six carbon atoms (i.e., C 6 -aryl). [0060] As used herein, the term “heteroaryl” or “heteroaromatic” refers to a heterocycle having aromatic character. Heteroaryl substituents may be described by the number of carbon atoms, e.g., C 1-9 -heteroaryl indicates the number of carbon atoms contained in the heteroaryl group without including the number of heteroatoms.
- a C 1-9 -heteroaryl will include an additional one to four heteroatoms.
- a polycyclic heteroaryl may include one or more rings that are partially saturated.
- heteroaryls include pyridyl, pyrazinyl, pyrimidinyl (including, e.g., 2- and 4-pyrimidinyl), pyridazinyl, thienyl, furyl, pyrrolyl (including, e.g., 2-pyrrolyl), imidazolyl, thiazolyl, oxazolyl, pyrazolyl (including, e.g., 3- and 5-pyrazolyl), isothiazolyl, 1,2,3-triazolyl, 1,2,4-triazolyl, 1,3,4-triazolyl, tetrazolyl, 1,2,3-thiadiazolyl, 1,2,3-oxadiazolyl, 1,3,4-thiadiazolyl and 1,3,4-oxadiazolyl.
- Non-limiting examples of polycyclic heterocycles and heteroaryls include indolyl (including, e.g., 3-, 4-, 5-, 6- and 7-indolyl), indolinyl, quinolyl, tetrahydroquinolyl, isoquinolyl (including, e.g., 1- and 5-isoquinolyl), 1,2,3,4-tetrahydroisoquinolyl, cinnolinyl, quinoxalinyl (including, e.g., 2- and 5-quinoxalinyl), quinazolinyl, phthalazinyl, 1,8-naphthyridinyl, 1,4-benzodioxanyl, coumarin, dihydrocoumarin, 1,5-naphthyridinyl, benzofuryl (including, e.g., 3-, 4-, 5-, 6- and 7-benzofuryl), 2,3-dihydrobenzofuryl, 1,2-benzisox
- protecting group or “chemical protecting group” refers to chemical moieties that block some or all reactive moieties of a compound and prevent such moieties from participating in chemical reactions until the protective group is removed, for example, those moieties listed and described in T.W. Greene, P.G.M. Wuts, Protective Groups in Organic Synthesis, 3rd ed. John Wiley & Sons (1999). It may be advantageous, where different protecting groups are employed, that each (different) protective group be removable by a different means. Protective groups that are cleaved under totally disparate reaction conditions allow differential removal of such protecting groups. For example, protective groups can be removed by acid, base, and hydrogenolysis.
- Groups such as trityl, monomethoxytrityl, dimethoxytrityl, acetal and tert-butyldimethylsilyl are acid labile and may be used to protect carboxy and hydroxy reactive moieties in the presence of amino groups protected with Cbz groups, which are removable by hydrogenolysis, and Fmoc groups, which are base labile.
- Carboxylic acid moieties may be blocked with base labile groups such as, without limitation, methyl, or ethyl, and hydroxy reactive moieties may be blocked with base labile groups such as acetyl in the presence of amines blocked with acid labile groups such as tert-butyl carbamate or with carbamates that are both acid and base stable but hydrolytically removable.
- Carboxylic acid and hydroxyl reactive moieties may also be blocked with hydrolytically removable protective groups such as the benzyl group, while amine groups may be blocked with base labile groups such as Fmoc.
- a particularly useful amine protecting group for the synthesis of compound of Formula (I) is trifluoroacetamide.
- Carboxylic acid reactive moieties may be blocked with oxidatively-removable protective groups such as 2,4-dimethoxybenzyl, while coexisting amino groups may be blocked with fluoride labile silyl carbamates.
- Allyl blocking groups are useful in the presence of acid- and base-protecting groups since the former are stable and can be subsequently removed by metal or pi-acid catalysts.
- an allyl-blocked carboxylic acid can be deprotected with a palladium(0)-catalyzed reaction in the presence of acid labile t-butyl carbamate or base-labile acetate amine protecting groups.
- Yet another form of protecting group is a resin to which a compound or intermediate may be attached.
- nucleobase refers to the heterocyclic ring portion of a nucleoside, nucleotide, and/or morpholino subunit. Nucleobases may be naturally occurring (e.g., uracil, thymine, adenine, cytosine, and guanine), or may be modified or analogs of these naturally occurring nucleobases, e.g., one or more nitrogen atoms of the nucleobase may be independently at each occurrence replaced by carbon.
- Exemplary analogs include hypoxanthine (the base component of the nucleoside inosine); 2, 6-diaminopurine; 5-methyl cytosine; C5-propynyl-modified pyrimidines; 10-(9- (aminoethoxy)phenoxazinyl) (G-clamp) and the like.
- base pairing moieties include, but are not limited to, uracil, thymine, adenine, cytosine, guanine and hypoxanthine having their respective amino groups protected by acyl protecting groups, 2-fluorouracil, 2-fluorocytosine, 5-bromouracil, 5-iodouracil, 2,6- diaminopurine, azacytosine, pyrimidine analogs such as pseudoisocytosine and pseudouracil and other modified nucleobases such as 8-substituted purines, xanthine, or hypoxanthine (the latter two being the natural degradation products).
- nucleobases disclosed in Chiu and Rana (2003) RNA 9:1034-1048, Limbach et al. (1994) Nucleic Acids Res.22:2183-2196 and Revankar and Rao, Comprehensive Natural Products Chemistry, vol.7, 313, are also contemplated, the contents of which are incorporated herein by reference.
- Further examples of base pairing moieties include, but are not limited to, expanded-size nucleobases in which one or more benzene rings have been added. Nucleic base replacements described in the Glen Research catalog (www.glenresearch.com); Krueger AT et al. (2007) Acc. Chem. Res.40:141-150; Kool ET (2002) Acc. Chem.
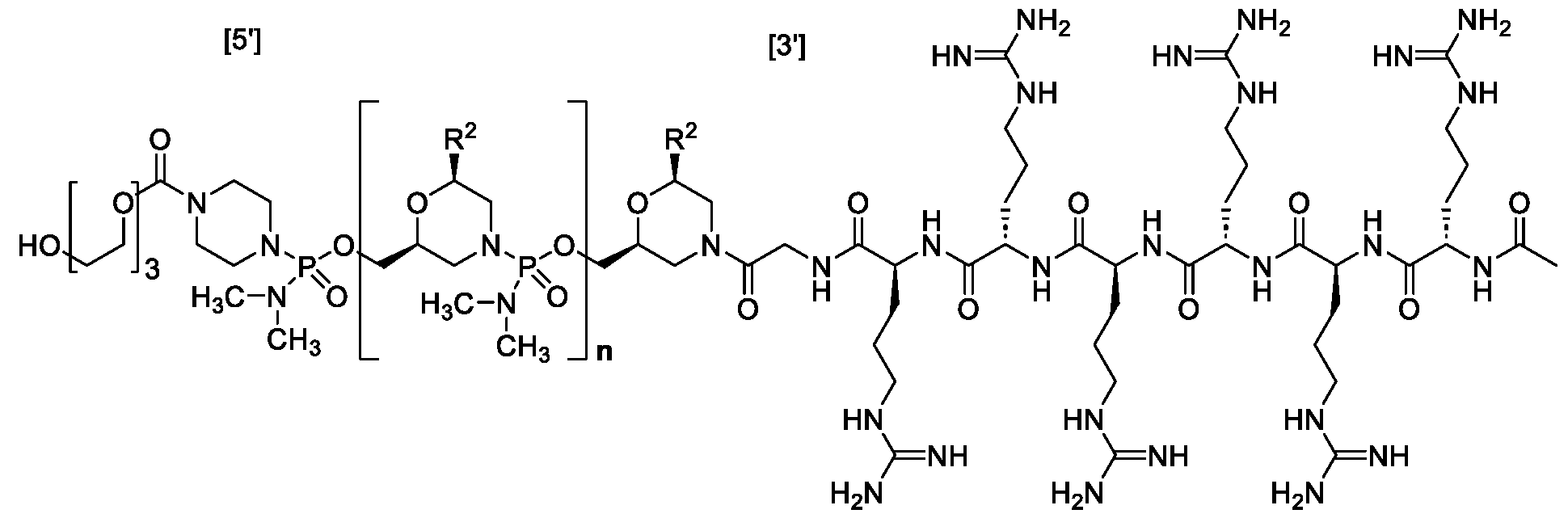
- the arginine residues can have any stereo configuration, for example, the arginine residues can be L-arginine residues, D- arginine residues, or a mixture of D- and L-arginine residues.
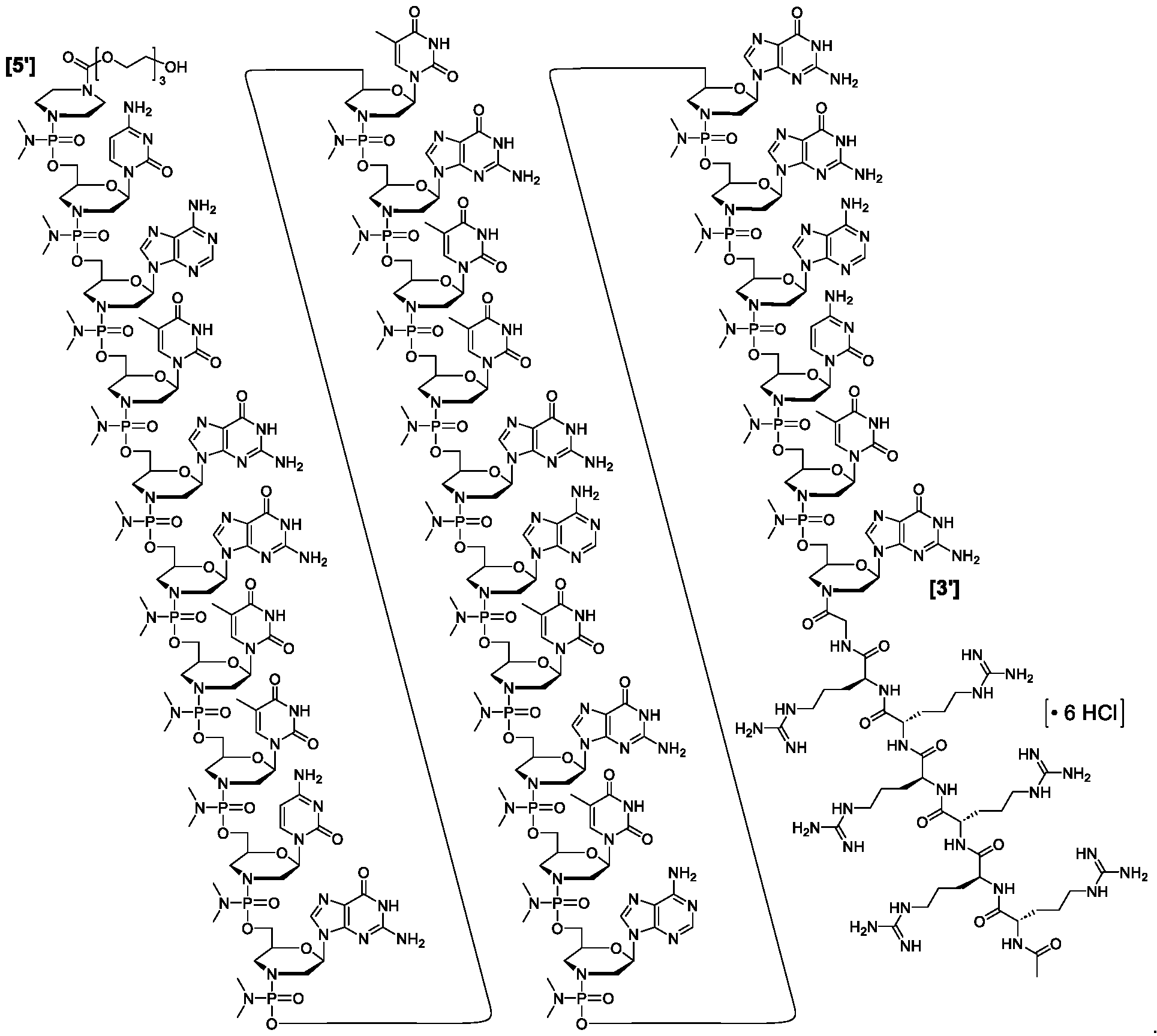
- “-G-R6” or “-G-R 6 -Ac” is conjugated to the morpholine ring nitrogen of the 3′ most morpholino subunit of a PMO antisense oligomer of the disclosure.
- “-G-R 6 ” or “-G-R 6 -Ac” is conjugated to the 3′ end of an antisense oligomer of the disclosure and is of the following formula: .
- oligonucleotide or “oligomer” refer to a compound comprising a plurality of linked nucleosides, nucleotides, or a combination of both nucleosides and nucleotides.
- an oligonucleotide is a morpholino oligonucleotide.
- nucleoside means a compound comprising a nucleobase and a sugar moiety. The nucleobase and sugar moiety are each, independently, unmodified or modified.
- “Modified nucleoside” means a nucleoside comprising a modified nucleobase and/or a modified sugar moiety. Modified nucleosides include abasic nucleosides, which lack a nucleobase.
- the antisense oligonucleotides comprising an abasic subunit have the partial structure:
- modified oligonucleotides comprise one or more nucleosides comprising a modified nucleobase. In certain embodiments, modified oligonucleotides comprise one or more nucleosides that do not comprise a nucleobase, referred to as an abasic nucleoside.
- “subunit” refers to a nucleotide, nucleoside, nucleobase, or sugar, or a modified nucleotide, nucleoside, nucleobase, or sugar as provided herein.
- antisense oligomer or “antisense compound” are used interchangeably and refer to a sequence of subunits, each having a base carried on a backbone subunit composed of ribose or other pentose sugar or morpholino group, and where the backbone groups are linked by intersubunit linkages that allow the bases in the compound to hybridize to a target sequence in a nucleic acid (typically an RNA) by Watson-Crick base pairing, to form a nucleic acid:oligomer heteroduplex within the target sequence.
- the oligomer may have exact sequence complementarity to the target sequence or nearly exact complementarity.
- antisense oligomers are designed to block or inhibit translation of the mRNA containing the target sequence and may be said to be “directed to” a sequence with which it hybridizes.
- antisense oligomers or “antisense compounds” are phosphorothioate-modified oligomers, peptide nucleic acids (PNAs), locked nucleic acids (LNAs), 2′-fluoro-modified oligomers, 2′-O,4′-C-ethylene-bridged nucleic acids (ENAs), tricyclo- DNAs, tricylo-DNA phosphorothioate-modified oligomers, 2′-O-[2-(N-methylcarbamoyl) ethyl] modified oligomers, 2′-O-methyl phosphorothioate modified oligomers, 2′-O-methoxyethyl (2′-O- MOE) modified oligomers, and 2
- An antisense oligomer “specifically hybridizes” to a target polynucleotide if the oligomer hybridizes to the target under physiological conditions, with a Tm greater than 37 °C, greater than 45 °C, at least 50 °C, and typically 60 °C-80 °C or higher.
- the “Tm” of an oligomer is the temperature at which 50% hybridizes to a complementary polynucleotide. Tm is determined under standard conditions in physiological saline, as described, for example, in Miyada et al. (1987) Methods Enzymol.154:94-107.
- complementarity refers to oligonucleotides (i.e., a sequence of nucleotides) related by base-pairing rules.
- sequence “T-G-A (5′- 3′)” is complementary to the sequence “T-C-A (5′-3′).”
- Complementarity may be “partial,” in which only some of the nucleic acids’ bases are matched according to base pairing rules, or there may be “complete,” “total,” or “perfect” (100%) complementarity between the nucleic acids.
- oligonucleotides can include one or more mismatches (e.g., 6, 5, 4, 3, 2, or 1 mismatches) with respect to the target RNA. Such hybridization may occur with “near” or “substantial” complementarity of the antisense oligomer to the target sequence, as well as with exact complementarity.
- an oligomer may hybridize to a target sequence with about 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 99% or 100% complementarity. Variations at any location within the oligomer are included.
- variations in sequence near the termini of an oligomer, if present, are typically within about 6, 5, 4, 3, 2, or 1 nucleotides of the 5′-terminus, 3′-terminus, or both termini.
- the terms “TEG,” “EG3,” or “triethylene glycol tail” refer to triethylene glycol moieties conjugated to the oligomer, e.g., at its 3′- or 5′-end.
- “TEG” includes, for example, wherein A′ of the conjugate of Formula (I) is of the formula: .
- Naturally occurring nucleotide bases include adenine, guanine, cytosine, thymine, and uracil, which have the symbols A, G, C, T, and U, respectively. Nucleotide bases can also encompass analogs of naturally occurring nucleotide bases. Base pairing typically occurs between purine A and pyrimidine T or U, and between purine G and pyrimidine C. [0078] Oligonucleotides may also include nucleobase modifications or substitutions. Oligonucleotides containing a modified or substituted base include oligonucleotides in which one or more purine or pyrimidine bases most found in nucleic acids are replaced with less common or non-natural bases.
- the nucleobase is covalently linked at the N9 atom of the purine base, or at the N1 atom of the pyrimidine base, to the morpholine ring of a nucleotide or nucleoside.
- Purine bases comprise a pyrimidine ring fused to an imidazole ring, as described by the general formula: .
- Adenine and guanine are the two purine nucleobases most found in nucleic acids. These may be substituted with other naturally occurring purines, including, but not limited to, N6- methyladenine, N2-methylguanine, hypoxanthine, and 7-methylguanine.
- Pyrimidine bases comprise a six-membered pyrimidine ring as described by the general formula: .
- Cytosine, uracil, and thymine are the pyrimidine bases most found in nucleic acids. These may be substituted with other naturally occurring pyrimidines, including, but not limited to, 5-methylcytosine, 5-hydroxymethylcytosine, pseudouracil, and 4-thiouracil.
- the oligonucleotides described herein contain thymine bases in place of uracil.
- modified or substituted bases include, but are not limited to, 2,6-diaminopurine, orotic acid, agmatidine, lysidine, 2-thiopyrimidine (e.g.2-thiouracil, 2-thiothymine), G-clamp and its derivatives, 5-substituted pyrimidine (e.g.5-halouracil, 5-propynyluracil, 5-propynylcytosine, 5-aminomethyluracil, 5-hydroxymethyluracil, 5-aminomethylcytosine, 5-hydroxymethylcytosine, Super T), 7-deazaguanine, 7-deazaadenine, 7-aza-2,6-diaminopurine, 8-aza-7-deazaguanine, 8-aza-7-deazaadenine, 8-aza-7-deaza-2,6-diaminopurine, Super G, Super A, and N4- ethylcytosine, or derivatives thereof; N2-cyclopent
- Pseudouracil is a naturally occurring isomerized version of uracil, with a C-glycoside rather than the regular N- glycoside as in uridine.
- modified or substituted nucleobases are useful for facilitating the purification of antisense oligonucleotides.
- antisense oligonucleotides may contain three or more (e.g., 3, 4, 5, 6, or more) consecutive guanine bases. In certain antisense oligonucleotides, a string of three or more consecutive guanine bases can result in aggregation of the oligonucleotides, complicating purification.
- one or more of the consecutive guanines can be substituted with hypoxanthine.
- the substitution of hypoxanthine for one or more guanines in a string of three or more consecutive guanine bases can reduce aggregation of the antisense oligonucleotide, thereby facilitating purification.
- the oligonucleotides provided herein are synthesized and do not include antisense compositions of biological origin.
- nucleic acid analog refers to a non-naturally occurring nucleic acid molecule.
- a nucleic acid is a polymer of nucleotide subunits linked together into a linear structure. Each nucleotide consists of a nitrogen-containing aromatic base attached to a pentose (five-carbon) sugar, which is in turn attached to a phosphate group.
- phosphate groups are linked together through phosphodiester bonds to form the polymer.
- the two common forms of naturally occurring nucleic acids are deoxyribonucleic acid (DNA) and ribonucleic acid (RNA).
- DNA deoxyribonucleic acid
- RNA ribonucleic acid
- One end of the chain carries a free phosphate group attached to the 5′-carbon atom of a sugar moiety; this is called the 5′ end of the molecule.
- the other end has a free hydroxyl (-OH) group at the 3′-carbon of a sugar moiety and is called the 3′ end of the molecule.
- a nucleic acid analog can include one or more non-naturally occurring nucleobases, sugars, and/or internucleotide linkages, for example, a phosphorodiamidate morpholino oligomer (PMO).
- PMO phosphorodiamidate morpholino oligomer
- a “nucleic acid analog” is a PMO
- a “nucleic acid analog” is a positively charged cationic PMO.
- a “morpholino oligomer” or “PMO” refers to a polymeric molecule having a backbone that supports bases capable of hydrogen bonding to typical polynucleotides, wherein the polymer lacks a pentose sugar backbone moiety, and more specifically a ribose backbone linked by phosphodiester bonds which is typical of nucleotides and nucleosides, but instead contains a ring nitrogen with coupling through the ring nitrogen.
- An exemplary “morpholino” oligomer comprises morpholino subunit structures linked together by phosphoramidate or phosphorodiamidate linkages, joining the morpholino nitrogen of one subunit to the 5′ exocyclic carbon of an adjacent subunit, each subunit comprising a purine or pyrimidine base-pairing moiety effective to bind, by base-specific hydrogen bonding, to a base in a polynucleotide.
- Morpholino oligomers are detailed, for example, in U.S. Pat.
- a preferred morpholino oligomer is a phosphorodiamidate-linked morpholino oligomer, referred to herein as a PMO.
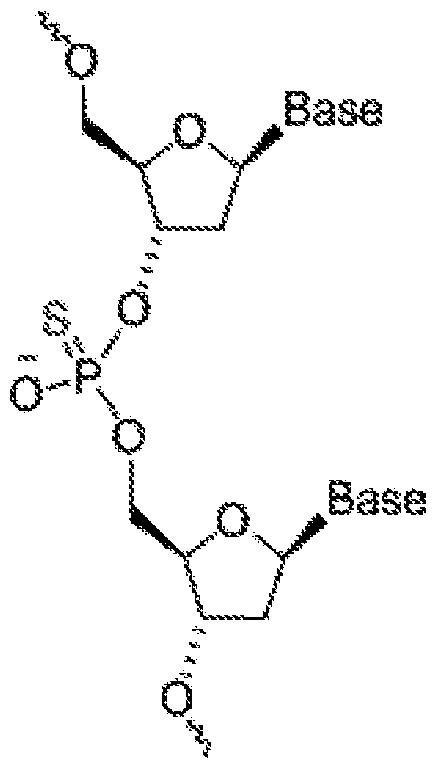
- Such oligomers are composed of morpholino subunit structures such as those shown below: where X is NH2, NHR, or NR2 (where R is lower alkyl, (e.g., methyl)), Y1 is O, and Z is O, and Pi and Pj are purine or pyrimidine base-pairing moieties effective to bind, by base-specific hydrogen bonding, to a base in a polynucleotide. Also preferred are structures having an alternate phosphorodiamidate linkage, where X is lower alkoxy, such as methoxy or ethoxy, Y1 is NH or NR, where R is lower alkyl, and Z is O.
- Representative PMOs include PMOs wherein the intersubunit linkages are linkage (A1). See Table 1. Table 1. Representative Intersubunit Linkages [0090] A “phosphoramidate” group comprises phosphorus having three attached oxygen atoms and one attached nitrogen atom, while a “phosphorodiamidate” group comprises phosphorus having two attached oxygen atoms and two attached nitrogen atoms. A representative phosphorodiamidate example is below:
- each P i is independently selected from H, a nucleobase, and a nucleobase functionalized with a chemical protecting-group, wherein the nucleobase independently at each occurrence comprises a pyridine, a pyrimidine, a triazinane, purine, or a deaza-purine; and n is an integer of 6-38 (i.e., n is 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, or 38).
- the ring nitrogen of the subunit at the 3′ terminus of a PMO may be capped with a capping group such as acetyl or may be uncapped with a free hydrogen.
- PMOs are water-soluble, uncharged, or substantially uncharged antisense molecules that inhibit gene expression by preventing binding or progression of splicing or translational machinery components. PMOs have also been shown to inhibit or block viral replication (Stein, Skilling et al.2001; McCaffrey, Meuse et al.2003), and are highly resistant to enzymatic digestion (Hudziak, Barofsky et al.1996).
- PMOs have demonstrated high antisense specificity and efficacy in vitro in cell-free and cell culture models (Stein, Foster et al.1997; Summerton and Weller 1997), and in vivo in zebrafish, frog, and sea urchin embryos (Heasman, Kofron et al.2000; Nasevicius and Ekker 2000), as well as in adult animal models, such as rats, mice, rabbits, dogs, and pigs (see e.g.
- Antisense PMO oligomers have been shown to be taken up into cells and to be more consistently effective in vivo, with fewer nonspecific effects, than other widely used antisense oligonucleotides (see e.g., P. Iversen, “Phosphoramidite Morpholino Oligomers,” in Antisense Drug Technology, S.T.
- a “cationic PMO” or “PMO+” refers to a phosphorodiamidate morpholino oligomer comprising any number of (1-piperazino)phosphinylideneoxy, (1-(4-( ⁇ -guanidino-alkanoyl))- piperazino)phosphinylideneoxy linkages (A2 and A3; see Table 1) that have been described previously (see e.g., PCT publication WO 2008/036127 which is incorporated herein by reference in its entirety).
- the “backbone” of an antisense oligomer refers to the structure supporting the base-pairing moieties, e.g., for a morpholino oligomer, as described herein, the “backbone” includes morpholino ring structures connected by intersubunit linkages (e.g., phosphorus-containing linkages).
- a “substantially uncharged backbone” refers to the backbone of an oligonucleotide analogue wherein less than 50% of the intersubunit linkages are charged at near-neutral pH.
- a substantially uncharged backbone may comprise less than 50%, less than 40%, less than 30%, less than 20%, less than 10%, less than 5% or even 0% intersubunit linkages which are charged at near neutral pH.
- the substantially uncharged backbone comprises at most one charged (at physiological pH) intersubunit linkage for every four uncharged (at physiological pH) linkages, at most one for every eight or at most one for every sixteen uncharged linkages.
- the nucleic acid analogs described herein are fully uncharged.
- the compounds described herein comprise or consist of an oligonucleotide comprising a targeting base sequence that is complementary to a target nucleic acid.
- the target nucleic acid is an endogenous RNA molecule. In certain embodiments, the target nucleic acid encodes a protein. In certain embodiments, the target nucleic acid is non-coding. In certain such embodiments, the target nucleic acid is selected from an mRNA and a pre-mRNA, including intronic, exonic and untranslated regions. In certain embodiments, the target RNA is an mRNA. In certain embodiments, the target nucleic acid is a pre-mRNA. In certain such embodiments, the target region is entirely within an exon. In certain such embodiments, the target region is entirely within an intron. In certain embodiments, the target region spans an intron/exon junction.
- the target region is at least 50% within an intron.
- targeting base sequence or “targeting sequence” is the (nucleobase) sequence in the antisense oligomer that is complementary or substantially complementary to a target nucleic acid. The entire sequence, or only a portion, of the oligomer compound may be complementary to the target sequence.
- target region refers to a sequence in a nucleic acid that is targeted by, and is therefore complementary, substantially complementary, or partially complementary, to the targeting sequence of the antisense oligomer. The entire sequence, or only a portion, of the target region may be complementary to the targeting sequence of the oligomer compound described herein.
- a plurality of antisense compounds is directed to a single target region.
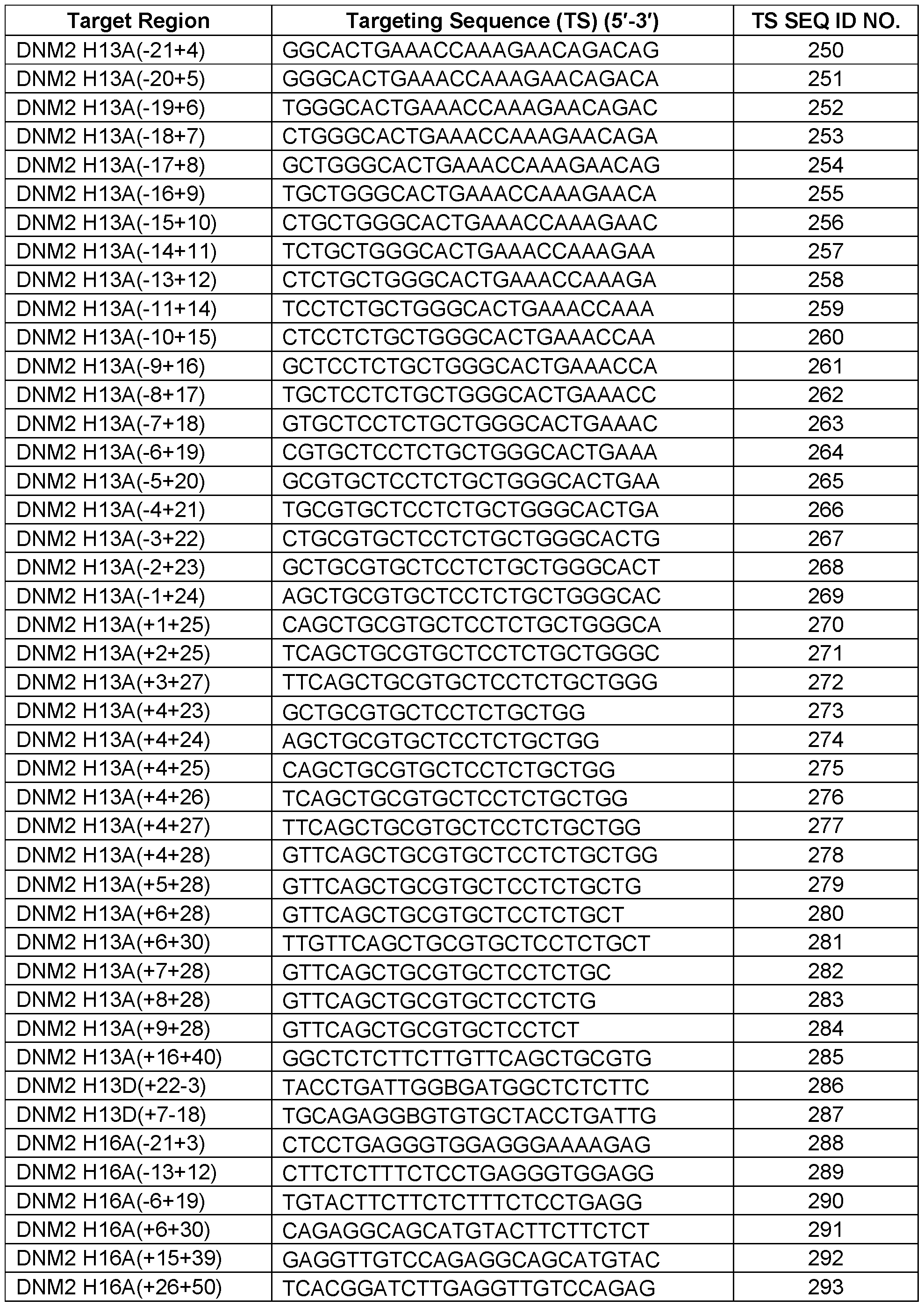
- a nomenclature system according to gene, species, exon number, and annealing coordinates is used herein to identify or indicate a target region or annealing coordinates of a targeted nucleic acid (e.g., a nucleic acid transcript) as follows: G SpZA/D ( ⁇ X ⁇ Y), where G is gene name; Sp is species (e.g., “H” for human, “M” for mouse); Z is exon number; A/D is selected from acceptor (A) and donor (D); The symbol “–“ denotes an intronic position; The symbol “+” denotes an exonic position; X is the position of the first 5′ base relative to A or D site; and Y is the position of the first 3′ base relative to A or D site.
- the nomenclature begins with the name of the transcript (e.g., dynamin 2; DNM2), then the species of the target mRNA (e.g., H: human or M: mouse), followed by the target exon number of the specified transcript and specification of an acceptor (A) or donor (D) site.
- the annealing coordinates are shown in brackets within the pre-mRNA transcript.
- the intronic bases are designated with a negative prefix (-) and the exonic position with a positive (+) symbol.
- the annealing coordinates are the positions of bases relative to the bases relative to the acceptor or donor sites of the reference transcript as denoted by National Center for Biotechnology Information and Ensembl genome browser 96.
- H5A(+76+100) indicates the site within the 76 th and 100 th nucleotides as measured from the 5′ end of an exon 5 (e.g., exon 5 of human DNM2 gene pre-mRNA) (i.e., the region within the 76 th and 100 th nucleotides as measured from the start of exon 5 at the 5′ end).
- exon 5 e.g., exon 5 of human DNM2 gene pre-mRNA
- DNM2 H2D (+13-12) indicates the last 13 exonic bases and the first 12 intronic bases of the target exon donor splice site (i.e., the target region within the 13 th nucleotide of exon 2 measured from the 3′ end of exon 2 to the 12 th nucleotide of intron 2 measured from the 3′ end of exon 2 (e.g., human exon 2 of the human DNM2 gene pre-mRNA).
- DNM2 H13A(-8+17) indicates the last 8 bases of the intron preceding the target exon 2 and the first 17 bases of the target exon 2 (i.e., the target region within the 8 th nucleotide of intron 12 measured from the 5′ end of exon 13 to the 17 th nucleotide of exon 13 measured from the 5′ end of exon 13).
- the closest splice site is the acceptor; therefore, these coordinates are preceded with “A.”
- peptide refers to a compound comprising a plurality of linked amino acids.
- the peptides provided herein are cell-penetrating peptides (CPP).
- CPP cell-penetrating peptide
- carrier peptide is a relatively short peptide capable of promoting the uptake of PMOs by cells, thereby delivering the PMOs to the interior (cytoplasm) of the cells.
- the CPP or carrier peptide typically is about 12 to about 40 amino acids long.
- the length of the carrier peptide is not particularly limited and varies in different embodiments. In some embodiments, the carrier peptide comprises from 4 to 40 amino acid subunits.
- the carrier peptide comprises from 6 to 30, from 6 to 20, from 8 to 25, or from 10 to 20 amino acid subunits.
- the CPPs disclosed herein comprise an arginine-rich peptide as described further below.
- a “peptide-conjugated phosphorodiamidate-linked morpholino oligomer” or “PPMO” refers to a PMO covalently linked to a peptide, such as a cell-penetrating peptide (CPP) or carrier peptide. The cell-penetrating peptide promotes the uptake of the PMO by cells, thereby delivering the PMO to the interior (cytoplasm) of the cells.
- CPP cell-penetrating peptide
- a CPP can be generally effective, or it can be specifically or selectively effective for PMO delivery to a particular type or particular types of cells.
- PMOs and CPPs are typically linked at their ends, e.g., the C-terminal end of the CPP can be linked to the 5′ end of the PMO, or the 3′ end of the PMO can be linked to the N-terminal end of the CPP.
- PPMOs can include uncharged PMOs, charged (e.g., cationic) PMOs, and mixtures thereof.
- the linking moiety of the conjugates described herein may be cleaved to release a PPMO.
- the carrier peptide may be linked to the nucleic acid analog either directly or via an optional linker, e.g., one or more additional naturally occurring amino acids, e.g., cysteine (C), glycine (G), or proline (P), or additional amino acid analogs, e.g., 6-aminohexanoic acid (X), beta-alanine (B), or XB.
- an optional linker e.g., one or more additional naturally occurring amino acids, e.g., cysteine (C), glycine (G), or proline (P), or additional amino acid analogs, e.g., 6-aminohexanoic acid (X), beta-alanine (B), or XB.
- additional amino acid analogs e.g., 6-aminohexanoic acid (X), beta-alanine (B), or XB.
- Other linking moieties known in the art may also be employed.
- amino acid subunit is generally an ⁇ -amino acid residue (–CO-CHR-NH-); but may also be a ⁇ - or other amino acid residue (e.g., -CO-CH 2 CHR-NH-), where R is an amino acid side chain.
- naturally occurring amino acid refers to an amino acid present in proteins found in nature; examples include Alanine (A), Cysteine (C), Aspartic acid (D), Glutamic acid (E), Phenyalanine (F), Glycine (G), Histidine (H), Isoleucine (I), Lysine (K), Leucine (L).
- non-natural amino acids refers to those amino acids not present in proteins found in nature; examples include beta-alanine ( ⁇ -Ala) and 6-aminohexanoic acid (Ahx).
- the agent may be transported, for example, by “active transport,” referring to transport of agents across a mammalian cell membrane by, e.g., an ATP-dependent transport mechanism, or by “facilitated transport,” referring to transport of antisense agents across the cell membrane by a transport mechanism that requires binding of the agent to a transport protein, which then facilitates passage of the bound agent across the membrane.
- active transport referring to transport of agents across a mammalian cell membrane by, e.g., an ATP-dependent transport mechanism
- facilitated transport referring to transport of antisense agents across the cell membrane by a transport mechanism that requires binding of the agent to a transport protein, which then facilitates passage of the bound agent across the membrane.
- an “effective amount” refers to any amount of a substance that is sufficient to achieve a desired biological result.
- a “therapeutically effective amount” refers to any amount of a substance that is sufficient to achieve a desired therapeutic result.
- Peptide-oligonucleotides [0114] Provided herein is an antisense oligomer comprising a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of human dynamin 2 (DNM2) gene represented by SEQ ID NO: 1, wherein the target region is within an exon internal region, an intron internal region, an intron-exon junction, or an exon-intron junction of the human DNM2 gene pre-mRNA.
- DNM2 human dynamin 2
- the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 6 (SEQ ID NO: 3), exon 7 (SEQ ID NO: 4), exon 8 (SEQ ID NO: 5), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), exon 13 (SEQ ID NO: 8), exon 16 (SEQ ID NO: 9), and exon 19 (SEQ ID NO: 10).
- exon 2 SEQ ID NO: 2
- exon 6 SEQ ID NO: 3
- exon 7 SEQ ID NO: 4
- exon 8 SEQ ID NO: 5
- exon 9 SEQ ID NO: 6
- exon 12 SEQ ID NO: 7
- exon 13 SEQ ID NO: 8
- exon 16 SEQ ID NO: 9
- exon 19 SEQ ID NO: 10
- the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), and exon 13 (SEQ ID NO: 8).
- the target region within an exon internal region of the human DNM2 gene pre-mRNA is exon 2 (SEQ ID NO: 2).
- the target region within an exon internal region of the human DNM2 gene pre-mRNA is exon 9 (SEQ ID NO: 6).
- the target region within an exon internal region of the human DNM2 gene pre-mRNA is exon 12 (SEQ ID NO: 7).
- the target region within an exon internal region of the human DNM2 gene pre- mRNA and exon 13 (SEQ ID NO: 8). [0117] In certain embodiments, the target region is within an intron-exon junction or an exon- intron junction region of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 5/exon 6 junction, exon 6/intron 6 junction, intron 6/exon 7 junction, exon 7/intron 7 junction, intron 7/exon 8 junction, exon 8/intron 8 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, exon 13/intron 13 junction, intron 15/exon 16 junction, exon 16/intron 16 junction, intron 18/exon 19 junction, and exon 19/intron 19 junction.
- the target region is within an intron-exon junction or an exon- intron junction of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, and exon 13/intron 13 junction.
- the target region within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA is intron 1/exon 2 junction.
- the target region within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA is exon 2/intron 2 junction. In certain embodiments, the target region within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre- mRNA is intron 8/exon 9 junction. In certain embodiments, the target region within an intron- exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA is exon 9/intron 9 junction. In certain embodiments, the target region within an intron-exon junction or an exon- intron junction of the human DNM2 gene pre-mRNA is intron 11/exon 12 junction.
- the target region within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA is exon 12/intron 12 junction, intron 12/exon 13 junction. In certain embodiments, the target region within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA is exon 13/intron 13 junction. [0120] In certain embodiments, the target region is within an intron internal region of the human DNM2 gene pre-mRNA selected from intron 2 (SEQ ID NO: 11) and intron 12 (SEQ ID NO: 12). In certain embodiments, the target region within an intron internal region of the human DNM2 gene pre-mRNA is intron 2 (SEQ ID NO: 11).
- the target region within an intron internal region of the human DNM2 gene pre-mRNA is intron 12 (SEQ ID NO: 12).
- the antisense oligomer is 13-30 subunits in length.
- the antisense oligonucleotide is 13-30, 14-30, 15-30, 16-30, 17-30, 18-30, 19-30, 20-30, 21-30, 22-30, 23-30, 24-30, 25-30, 26-30, 27-30, 28-30, or 29-30 subunits in length.
- the antisense oligomer is 13-29 subunits in length.
- the antisense oligonucleotide is 13-29, 14-29, 15-29, 16-29, 17-29, 18-29, 19-29, 20-29, 21-29, 22-29, 23-29, 24-29, 25-29, 26-29, 27-29, or 28-29 subunits in length. [0123] In certain embodiments, the antisense oligomer is 13-28 subunits in length.
- the antisense oligonucleotide is 13-28, 14-28, 15-28, 16-28, 17-28, 18-28, 19-28, 20-28, 21-28, 22-28, 23-28, 24-28, 25-28, 26-28, or 27-28 subunits in length. [0124] In certain embodiments, the antisense oligomer is 13-27 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-27, 14-27, 15-27, 16-27, 17-27, 18-27, 19-27, 20-27, 21-27, 22-27, 23-27, 24-27, 25-27, or 26-27 subunits in length.
- the antisense oligomer is 13-26 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-26, 14-26, 15-26, 16-26, 17-26, 18-26, 19-26, 20-26, 21-26, 22-26, 23-26, 24-26, or 25-26 subunits in length. [0126] In certain embodiments, the antisense oligomer is 13-25 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-25, 14-25, 15-25, 16-25, 17-25, 18-25, 19-25, 20-25, 21-25, 22-25, 23-25, or 24-25 subunits in length.
- the antisense oligomer is 13-24 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-24, 14-24, 15-24, 16-24, 17-24, 18-24, 19-24, 20-24, 21-24, 22-24, or 23-24 subunits in length. [0128] In certain embodiments, the antisense oligomer is 13-23 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-23, 14-23, 15-23, 16-23, 17-23, 18-23, 19-23, 20-23, 21-23, or 22-23 subunits in length.
- the antisense oligomer is 13-22 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-22, 14-22, 15-22, 16-22, 17-22, 18-22, 19-22, 20-22, or 21-22 subunits in length. [0130] In certain embodiments, the antisense oligomer is 13-21 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-21, 14-21, 15-21, 16-21, 17-21, 18-21, 19-21, or 20-21 subunits in length. [0131] In certain embodiments, the antisense oligomer is 13-20 subunits in length.
- the antisense oligonucleotide is 13-20, 14-20, 15-20, 16-20, 17-20, 18-20, or 19- 20 subunits in length. [0132] In certain embodiments, the antisense oligomer is 13-19 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-19, 14-19, 15-19, 16-19, 17-19, or 18-19 subunits in length. [0133] In certain embodiments, the antisense oligomer is 13-18 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-18, 14-18, 15-18, 16-18, or 17-18 subunits in length.
- the antisense oligonucleotide is 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 subunits in length. In certain embodiments, the antisense oligonucleotide is 13 subunits in length. In certain embodiments, the antisense oligonucleotide is 14 subunits in length. In certain embodiments, the antisense oligonucleotide is 15 subunits in length. In certain embodiments, the antisense oligonucleotide is 16 subunits in length. In certain embodiments, the antisense oligonucleotide is 17 subunits in length.
- the antisense oligonucleotide is 18 subunits in length. In certain embodiments, the antisense oligonucleotide is 19 subunits in length. In certain embodiments, the antisense oligonucleotide is 20 subunits in length. In certain embodiments, the antisense oligonucleotide is 21 subunits in length. In certain embodiments, the antisense oligonucleotide is 22 subunits in length. In certain embodiments, the antisense oligonucleotide is 23 subunits in length. In certain embodiments, the antisense oligonucleotide is 24 subunits in length.
- the antisense oligonucleotide is 25 subunits in length. In certain embodiments, the antisense oligonucleotide is 26 subunits in length. In certain embodiments, the antisense oligonucleotide is 27 subunits in length. In certain embodiments, the antisense oligonucleotide is 28 subunits in length. In certain embodiments, the antisense oligonucleotide is 29 subunits in length. In certain embodiments, the antisense oligonucleotide is 30 subunits in length.
- the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12- 13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2 H2A (-10+15),
- the target region is DNM2 H2A (-10+15). In certain embodiments, the target region is DNM2 H2A(-7+17). In certain embodiments, the target region is DNM2 H2A(+3+27). In certain embodiments, the target region is DNM2 H2A(+15+37). In certain embodiments, the target region is DNM2 H2A(+21+45). In certain embodiments, the target region is DNM2 H2A(+23+47). In certain embodiments, the target region is DNM2 H2A(+27+51). In certain embodiments, the target region is DNM2 H2A(+37+61). In certain embodiments, the target region is DNM2 H2A(+46+70).
- the target region is DNM2 H2A (+48+72). In certain embodiments, the target region is DNM2 H2D(+20-5). In certain embodiments, the target region is DNM2 H2D (+13-12). In certain embodiments, the target region is DNM2 H2D(+12-13). In certain embodiments, the target region is DNM2 H2D(+11-14). In certain embodiments, the target region is DNM2 H2D(+10-15). In certain embodiments, the target region is DNM2 H2D(+9-16). In certain embodiments, the target region is DNM2 H2D(+8-17). In certain embodiments, the target region is DNM2 H2D(+7-18).
- the target region is DNM2 H2D(+6-19). In certain embodiments, the target region is DNM2 H2D(+5-20). In certain embodiments, the target region is DNM2 H2D(+4-21). In certain embodiments, the target region is DNM2 H2D(+3-22). In certain embodiments, the target region is DNM2 H2D(+2-23) . In certain embodiments, the target region is DNM2 H2D(+1-24). In certain embodiments, the target region is DNM2 H2D(-1-25). In certain embodiments, the target region is DNM2 H2D(-2-26). In certain embodiments, the target region is DNM2 H2D(-3-27).
- the target region is DNM2 H2D(-4-28). In certain embodiments, the target region is DNM2 H2D(-5-29). In certain embodiments, the target region is DNM2 H2D(-6-30). In certain embodiments, the target region is DNM2 H6A(-20+5). In certain embodiments, the target region is DNM2 H6A(-6+19). In certain embodiments, the target region is DNM2 H6A(-2+23). In certain embodiments, the target region is DNM2 H6A(+3+26). In certain embodiments, the target region is DNM2 H6A(+10+3). In certain embodiments, the target region is DNM2 H6A(+13+37).
- the target region is DNM2 H6A(+23+47). In certain embodiments, the target region is DNM2 H6A(+32+56). In certain embodiments, the target region is DNM2 H6A(+35+59). In certain embodiments, the target region is DNM2 H6A(+44+68). In certain embodiments, the target region is DNM2 H6A(+54+78). In certain embodiments, the target region is DNM2 H6A(+60+84). In certain embodiments, the target region is DNM2 H6A(+63+87). In certain embodiments, the target region is DNM2 H6A(+88+112). In certain embodiments, the target region is DNM2 H6A(+96+119).
- the target region is DNM2 H7A(+1+25). In certain embodiments, the target region is DNM2 H7A(+6+30). In certain embodiments, the target region is. In certain embodiments, the target region is DNM2 H7A(+22+46). In certain embodiments, the target region is DNM2 H7A(+29+53). In certain embodiments, the target region is DNM2 H7A(+41+65). In certain embodiments, the target region is DNM2 H7A(+51+75). In certain embodiments, the target region is DNM2 H7A(+57+81). In certain embodiments, the target region is DNM2 H7A(+62+86).
- the target region is DNM2 H7A(+72+96). In certain embodiments, the target region is DNM2 H7A(+74+98). In certain embodiments, the target region is DNM2 H7A(+81+105). In certain embodiments, the target region is DNM2 H7A(+91+115). In certain embodiments, the target region is DNM2 H7A(+110+134). In certain embodiments, the target region is DNM2 H7D(+8-16). In certain embodiments, the target region is DNM2 H7D(+9-16). In certain embodiments, the target region is DNM2 H7D(+21-4). In certain embodiments, the target region is DNM2 H8A(-20+5).
- the target region is DNM2 H8A(-14+11). In certain embodiments, the target region is DNM2 H8A(-7+18). In certain embodiments, the target region is DNM2 H8A(-6+19). In certain embodiments, the target region is DNM2 H8A(-2+23). In certain embodiments, the target region is DNM2 H8A(+5+29). In certain embodiments, the target region is DNM2 H8A(+18+42). In certain embodiments, the target region is DNM2 H8A(+21+45). In certain embodiments, the target region is DNM2 H8A(+27+51). In certain embodiments, the target region is DNM2 H8A(+36+60).
- the target region is DNM2 H8A(+46+70). In certain embodiments, the target region is DNM2 H8A(+51+75). In certain embodiments, the target region is DNM2 H8A(+56+80). In certain embodiments, the target region is DNM2 H8A(+65+88). In certain embodiments, the target region is DNM2 H8A(+76+100). In certain embodiments, the target region is DNM2 H8A(+83+107). In certain embodiments, the target region is DNM2 H8A(+96+119). In certain embodiments, the target region is DNM2 H8A(+105+129). In certain embodiments, the target region is DNM2 H8D(+18-7).
- the target region is DNM2 H8D(+10-15). In certain embodiments, the target region is DNM2 H9A(-12+13). In certain embodiments, the target region is DNM2 H9A(-5+20). In certain embodiments, the target region is DNM2 H9A(+1+25). In certain embodiments, the target region is DNM2 H9A(+6+30). In certain embodiments, the target region is DNM2 H9A(+16+40). In certain embodiments, the target region is DNM2 H9A(+25+49). In certain embodiments, the target region is DNM2 H9A(+36+60). In certain embodiments, the target region is DNM2 H9A(+37+61).
- the target region is DNM2 H9A(+42+66). In certain embodiments, the target region is DNM2 H9A(+43+62). In certain embodiments, the target region is DNM2 H9A(+43+63). In certain embodiments, the target region is DNM2 H9A(+43+64). In certain embodiments, the target region is DNM2 H9A(+43+65). In certain embodiments, the target region is DNM2 H9A(+43+66). In certain embodiments, the target region is DNM2 H9A(+43+67). In certain embodiments, the target region is DNM2 H9A(+44+67). In certain embodiments, the target region is DNM2 H9A(+44+68).
- the target region is DNM2 H9A(+45+67). In certain embodiments, the target region is DNM2 H9A (+46+67). In certain embodiments, the target region is DNM2 H9A(+47+67). In certain embodiments, the target region is DNM2 H9A(+48+67). In certain embodiments, the target region is DNM2 H9D(+24-1). In certain embodiments, the target region is DNM2 H9D(+23-2). In certain embodiments, the target region is DNM2 H9D(+22-3). In certain embodiments, the target region is DNM2 H9D(+21-4). In certain embodiments, the target region is DNM2 H9D(+20-5).
- the target region is DNM2 H9D(+19-6). In certain embodiments, the target region is DNM2 H9D (+18-7). In certain embodiments, the target region is DNM2 H9D(+17-8). In certain embodiments, the target region is DNM2 H9D(+16-9). In certain embodiments, the target region is DNM2 H9D(+15-10). In certain embodiments, the target region is DNM2 H9D(+14-11). In certain embodiments, the target region is DNM2 H9D(+13-12). In certain embodiments, the target region is DNM2 H9D(+12-13). In certain embodiments, the target region is DNM2 H9D(+11-14).
- the target region is DNM2 H9D(+10-15). In certain embodiments, the target region is DNM2 H9D(+9-16). In certain embodiments, the target region is DNM2 H9D(+8-12). In certain embodiments, the target region is DNM2 H9D(+8-13). In certain embodiments, the target region is DNM2 H9D(+8-14). In certain embodiments, the target region is DNM2 H9D(+8-15). In certain embodiments, the target region is DNM2 H9D(+8-17). In certain embodiments, the target region is DNM2 H9D(+7-18). In certain embodiments, the target region is DNM2 H9D(+7-17).
- the target region is DNM2 H9D(+6-19). In certain embodiments, the target region is DNM2 H9D(+6-17). In certain embodiments, the target region is DNM2 H9D(+5-17). In certain embodiments, the target region is DNM2 H9D(+5-20). In certain embodiments, the target region is DNM2 H9D(+4-17). In certain embodiments, the target region is DNM2 H9D(+4-21). In certain embodiments, the target region is DNM2 H9D(+3-17). In certain embodiments, the target region is DNM2 H9D(+3-22). In certain embodiments, the target region is DNM2 H12A(-13+12).
- the target region is DNM2 H12A(-3+22). In certain embodiments, the target region is DNM2 H12A(+4+28). In certain embodiments, the target region is DNM2 H12A(+7+31). In certain embodiments, the target region is DNM2 H12A(+10+34). In certain embodiments, the target region is DNM2 H12A(+11+35). In certain embodiments, the target region is DNM2 H12A(+12+36). In certain embodiments, the target region is DNM2 H12A(+13+37). In certain embodiments, the target region is DNM2 H12A(+14+38). In certain embodiments, the target region is DNM2 H12A(+15+39).
- the target region is DNM2 H12A(+16+40). In certain embodiments, the target region is DNM2 H12A(+17+41). In certain embodiments, the target region is DNM2 H12A(+18+42). In certain embodiments, the target region is DNM2 H12A(+19+43). In certain embodiments, the target region is DNM2 H12A(+20+44). In certain embodiments, the target region is DNM2 H12A(+25+44). In certain embodiments, the target region is DNM2 H12A(+25+45). In certain embodiments, the target region is DNM2 H12A(+25+46). In certain embodiments, the target region is DNM2 H12A(+25+47).
- the target region is DNM2 H12A(+25+48). In certain embodiments, the target region is DNM2 H12A(+25+49). In certain embodiments, the target region is DNM2 H12A(+26+49). In certain embodiments, the target region is DNM2 H12A(+27+49). In certain embodiments, the target region is DNM2 H12A(+28+49). In certain embodiments, the target region is DNM2 H12A(+29+49). In certain embodiments, the target region is DNM2 H12A(+29+53). In certain embodiments, the target region is DNM2 H12A(+30+49). In certain embodiments, the target region is DNM2 H12A(+30+54).
- the target region is DNM2 H12A(+36+60). In certain embodiments, the target region is DNM2 H12A(+37+61). In certain embodiments, the target region is DNM2 H12A(+43+67). In certain embodiments, the target region is DNM2 H12D(+22- 3). In certain embodiments, the target region is DNM2 H12D(+20-5). In certain embodiments, the target region is DNM2 H12D(+15-10). In certain embodiments, the target region is DNM2 H13A(-27-3). In certain embodiments, the target region is DNM2 H13A(-26-2). In certain embodiments, the target region is DNM2 H13A(-25-1).
- the target region is DNM2 H13A(-24+1). In certain embodiments, the target region is DNM2 H13A(-23+2). In certain embodiments, the target region is DNM2 H13A(-22+3). In certain embodiments, the target region is DNM2 H13A(-21+4). In certain embodiments, the target region is DNM2 H13A(- 20+5). In certain embodiments, the target region is DNM2 H13A(-19+6). In certain embodiments, the target region is DNM2 H13A(-18+7). In certain embodiments, the target region is DNM2 H13A(-17+8). In certain embodiments, the target region is DNM2 H13A(-16+9).
- the target region is DNM2 H13A(-15+10). In certain embodiments, the target region is DNM2 H13A(-14+11). In certain embodiments, the target region is DNM2 H13A(- 13+12). In certain embodiments, the target region is DNM2 H13A(-11+14). In certain embodiments, the target region is DNM2 H13A(-10+15). In certain embodiments, the target region is DNM2 H13A(-9+16). In certain embodiments, the target region is DNM2 H13A(-8+17). In certain embodiments, the target region is DNM2 H13A(-7+18). In certain embodiments, the target region is DNM2 H13A(-6+19).
- the target region is DNM2 H13A(- 5+20). In certain embodiments, the target region is DNM2 H13A(-4+21). In certain embodiments, the target region is DNM2 H13A(-3+22). In certain embodiments, the target region is DNM2 H13A(-2+23). In certain embodiments, the target region is DNM2 H13A(-1+24). In certain embodiments, the target region is DNM2 H13A(+1+25). In certain embodiments, the target region is DNM2 H13A(+2+25). In certain embodiments, the target region is DNM2 H13A(+3+27). In certain embodiments, the target region is DNM2 H13A(+4+23).
- the target region is DNM2 H13A(+4+24). In certain embodiments, the target region is DNM2 H13A(+4+25). In certain embodiments, the target region is DNM2 H13A(+4+26). In certain embodiments, the target region is DNM2 H13A(+4+27). In certain embodiments, the target region is DNM2 H13A(+4+28). In certain embodiments, the target region is DNM2 H13A(+5+28). In certain embodiments, the target region is DNM2 H13A(+6+28). In certain embodiments, the target region is DNM2 H13A(+6+30). In certain embodiments, the target region is DNM2 H13A(+7+28).
- the target region is DNM2 H13A(+8+28). In certain embodiments, the target region is DNM2 H13A(+9+28). In certain embodiments, the target region is DNM2 H13A(+16+40). In certain embodiments, the target region is DNM2 H13D(+22-3). In certain embodiments, the target region is DNM2 H13D(+7-18). In certain embodiments, the target region is DNM2 H16A(-21+3). In certain embodiments, the target region is DNM2 H16A(-13+12). In certain embodiments, the target region is DNM2 H16A(- 6+19). In certain embodiments, the target region is DNM2 H16A(+6+30).
- the target region is DNM2 H16A(+15+39). In certain embodiments, the target region is DNM2 H16A(+26+50). In certain embodiments, the target region is DNM2 H16A(+36+60). In certain embodiments, the target region is DNM2 H16A(+45+69). In certain embodiments, the target region is DNM2 H16A(+55+79). In certain embodiments, the target region is DNM2 H16A(+66+90). In certain embodiments, the target region is DNM2 H16A(+75+99). In certain embodiments, the target region is DNM2 H16A(+82+106). In certain embodiments, the target region is DNM2 H16D(+19-6).
- the target region is DNM2 H16D(+11-14). In certain embodiments, the target region is DNM2 H19A(- 11+14). In certain embodiments, the target region is DNM2 H19A(-4+21). In certain embodiments, the target region is DNM2 H19A(+1+25). In certain embodiments, the target region is DNM2 H19A(+7+31). In certain embodiments, the target region is DNM2 H19A(+16+40). In certain embodiments, the target region is DNM2 H19A(+26+50). In certain embodiments, the target region is DNM2 H19A(+37+61). In certain embodiments, the target region is DNM2 H19A(+47+71).
- the target region is DNM2 H19A(+56+80). In certain embodiments, the target region is DNM2 H19A(+66+90). In certain embodiments, the target region is DNM2 H19A(+76+100). In certain embodiments, the target region is DNM2 H19A(+86+110). In certain embodiments, the target region is DNM2 H19A(+96+120). In certain embodiments, the target region is DNM2 H19A(+106+130). In certain embodiments, the target region is DNM2 H19A(+117+141). In certain embodiments, the target region is DNM2 H19A(+127+151). In certain embodiments, the target region is DNM2 H19A(+137+161).
- the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGGCCGC); SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC); SEQ ID NO: 77 (TTTACCTGTTT
- B is an abasic subunit.
- the targeting sequence is SEQ ID NO: 67. In certain embodiments, the targeting sequence is SEQ ID NO: 68. In certain embodiments, the targeting sequence is SEQ ID NO: 69. In certain embodiments, the targeting sequence is SEQ ID NO: 70. [0139] In certain embodiments, the targeting sequence is SEQ ID NO: 71. In certain embodiments, the targeting sequence is SEQ ID NO: 72. In certain embodiments, the targeting sequence is SEQ ID NO: 73. In certain embodiments, the targeting sequence is SEQ ID NO: 74. In certain embodiments, the targeting sequence is SEQ ID NO: 75. In certain embodiments, the targeting sequence is SEQ ID NO: 76.
- the targeting sequence is SEQ ID NO: 77. In certain embodiments, the targeting sequence is SEQ ID NO: 78. In certain embodiments, the targeting sequence is SEQ ID NO: 79. In certain embodiments, the targeting sequence is SEQ ID NO: 80. In certain embodiments, the targeting sequence is SEQ ID NO: 81. In certain embodiments, the targeting sequence is SEQ ID NO: 82. In certain embodiments, the targeting sequence is SEQ ID NO: 83. In certain embodiments, the targeting sequence is SEQ ID NO: 84. In certain embodiments, the targeting sequence is SEQ ID NO: 85. In certain embodiments, the targeting sequence is SEQ ID NO: 86. In certain embodiments, the targeting sequence is SEQ ID NO: 87.
- the targeting sequence is SEQ ID NO: 88. In certain embodiments, the targeting sequence is SEQ ID NO: 89. In certain embodiments, the targeting sequence is SEQ ID NO: 90. In certain embodiments, the targeting sequence is SEQ ID NO: 91. In certain embodiments, the targeting sequence is SEQ ID NO: 92. In certain embodiments, the targeting sequence is SEQ ID NO: 93. In certain embodiments, the targeting sequence is SEQ ID NO: 94. In certain embodiments, the targeting sequence is SEQ ID NO: 95. In certain embodiments, the targeting sequence is SEQ ID NO: 96. In certain embodiments, the targeting sequence is SEQ ID NO: 97. In certain embodiments, the targeting sequence is SEQ ID NO: 98.
- the targeting sequence is SEQ ID NO: 99. In certain embodiments, the targeting sequence is SEQ ID NO: 100. In certain embodiments, the targeting sequence is SEQ ID NO: 101. In certain embodiments, the targeting sequence is SEQ ID NO: 102. In certain embodiments, the targeting sequence is SEQ ID NO: 103. In certain embodiments, the targeting sequence is SEQ ID NO: 104. In certain embodiments, the targeting sequence is SEQ ID NO: 105. In certain embodiments, the targeting sequence is SEQ ID NO: 106. In certain embodiments, the targeting sequence is SEQ ID NO: 107. In certain embodiments, the targeting sequence is SEQ ID NO: 108. In certain embodiments, the targeting sequence is SEQ ID NO: 109.
- the targeting sequence is SEQ ID NO: 110. In certain embodiments, the targeting sequence is SEQ ID NO: 111. In certain embodiments, the targeting sequence is SEQ ID NO: 112. In certain embodiments, the targeting sequence is SEQ ID NO: 113. In certain embodiments, the targeting sequence is SEQ ID NO: 114. In certain embodiments, the targeting sequence is SEQ ID NO: 115. In certain embodiments, the targeting sequence is SEQ ID NO: 116. In certain embodiments, the targeting sequence is SEQ ID NO: 117. In certain embodiments, the targeting sequence is SEQ ID NO: 118. In certain embodiments, the targeting sequence is SEQ ID NO: 119. In certain embodiments, the targeting sequence is SEQ ID NO: 120.
- the targeting sequence is SEQ ID NO: 121. In certain embodiments, the targeting sequence is SEQ ID NO: 122. In certain embodiments, the targeting sequence is SEQ ID NO: 123. In certain embodiments, the targeting sequence is SEQ ID NO: 124. In certain embodiments, the targeting sequence is SEQ ID NO: 125. In certain embodiments, the targeting sequence is SEQ ID NO: 126. In certain embodiments, the targeting sequence is SEQ ID NO: 127. In certain embodiments, the targeting sequence is SEQ ID NO: 128. In certain embodiments, the targeting sequence is SEQ ID NO: 129. In certain embodiments, the targeting sequence is SEQ ID NO: 130. In certain embodiments, the targeting sequence is SEQ ID NO: 131.
- the targeting sequence is SEQ ID NO: 132. In certain embodiments, the targeting sequence is SEQ ID NO: 133. In certain embodiments, the targeting sequence is SEQ ID NO: 134. In certain embodiments, the targeting sequence is SEQ ID NO: 135. In certain embodiments, the targeting sequence is SEQ ID NO: 136. In certain embodiments, the targeting sequence is SEQ ID NO: 137. In certain embodiments, the targeting sequence is SEQ ID NO: 138. In certain embodiments, the targeting sequence is SEQ ID NO: 139. In certain embodiments, the targeting sequence is SEQ ID NO: 140. In certain embodiments, the targeting sequence is SEQ ID NO: 141. In certain embodiments, the targeting sequence is SEQ ID NO: 142.
- the targeting sequence is SEQ ID NO: 154. In certain embodiments, the targeting sequence is SEQ ID NO: 155. In certain embodiments, the targeting sequence is SEQ ID NO: 156. In certain embodiments, the targeting sequence is SEQ ID NO: 157. In certain embodiments, the targeting sequence is SEQ ID NO: 158. In certain embodiments, the targeting sequence is SEQ ID NO: 159. In certain embodiments, the targeting sequence is SEQ ID NO: 160. In certain embodiments, the targeting sequence is SEQ ID NO: 161. In certain embodiments, the targeting sequence is SEQ ID NO: 162. In certain embodiments, the targeting sequence is SEQ ID NO: 163. In certain embodiments, the targeting sequence is SEQ ID NO: 164.
- the targeting sequence is SEQ ID NO: 165. In certain embodiments, the targeting sequence is SEQ ID NO: 166. In certain embodiments, the targeting sequence is SEQ ID NO: 167. In certain embodiments, the targeting sequence is SEQ ID NO: 168. In certain embodiments, the targeting sequence is SEQ ID NO: 169. In certain embodiments, the targeting sequence is SEQ ID NO: 170. In certain embodiments, the targeting sequence is SEQ ID NO: 171. In certain embodiments, the targeting sequence is SEQ ID NO: 172. In certain embodiments, the targeting sequence is SEQ ID NO: 173. In certain embodiments, the targeting sequence is SEQ ID NO: 174. In certain embodiments, the targeting sequence is SEQ ID NO: 175.
- the targeting sequence is SEQ ID NO: 176. In certain embodiments, the targeting sequence is SEQ ID NO: 177. In certain embodiments, the targeting sequence is SEQ ID NO: 178. In certain embodiments, the targeting sequence is SEQ ID NO: 179. In certain embodiments, the targeting sequence is SEQ ID NO: 180. In certain embodiments, the targeting sequence is SEQ ID NO: 181. In certain embodiments, the targeting sequence is SEQ ID NO: 182. In certain embodiments, the targeting sequence is SEQ ID NO: 183. In certain embodiments, the targeting sequence is SEQ ID NO: 184. In certain embodiments, the targeting sequence is SEQ ID NO: 185. In certain embodiments, the targeting sequence is SEQ ID NO: 186.
- the targeting sequence is SEQ ID NO: 187. In certain embodiments, the targeting sequence is SEQ ID NO: 188. In certain embodiments, the targeting sequence is SEQ ID NO: 189. In certain embodiments, the targeting sequence is SEQ ID NO: 190. In certain embodiments, the targeting sequence is SEQ ID NO: 191. In certain embodiments, the targeting sequence is SEQ ID NO: 192. In certain embodiments, the targeting sequence is SEQ ID NO: 193. In certain embodiments, the targeting sequence is SEQ ID NO: 194. In certain embodiments, the targeting sequence is SEQ ID NO: 195. In certain embodiments, the targeting sequence is SEQ ID NO: 196. In certain embodiments, the targeting sequence is SEQ ID NO: 197.
- the targeting sequence is SEQ ID NO: 198. In certain embodiments, the targeting sequence is SEQ ID NO: 199. In certain embodiments, the targeting sequence is SEQ ID NO: 200. In certain embodiments, the targeting sequence is SEQ ID NO: 201. In certain embodiments, the targeting sequence is SEQ ID NO: 202. In certain embodiments, the targeting sequence is SEQ ID NO: 203. In certain embodiments, the targeting sequence is SEQ ID NO: 204. In certain embodiments, the targeting sequence is SEQ ID NO: 205. In certain embodiments, the targeting sequence is SEQ ID NO: 206. In certain embodiments, the targeting sequence is SEQ ID NO: 207. In certain embodiments, the targeting sequence is SEQ ID NO: 208.
- the targeting sequence is SEQ ID NO: 209. In certain embodiments, the targeting sequence is SEQ ID NO: 210. In certain embodiments, the targeting sequence is SEQ ID NO: 211. In certain embodiments, the targeting sequence is SEQ ID NO: 212. In certain embodiments, the targeting sequence is SEQ ID NO: 213. In certain embodiments, the targeting sequence is SEQ ID NO: 214. In certain embodiments, the targeting sequence is SEQ ID NO: 215. In certain embodiments, the targeting sequence is SEQ ID NO: 216. In certain embodiments, the targeting sequence is SEQ ID NO: 217. In certain embodiments, the targeting sequence is SEQ ID NO: 218. In certain embodiments, the targeting sequence is SEQ ID NO: 219.
- the targeting sequence is SEQ ID NO: 220. In certain embodiments, the targeting sequence is SEQ ID NO: 221. In certain embodiments, the targeting sequence is SEQ ID NO: 222. In certain embodiments, the targeting sequence is SEQ ID NO: 223. In certain embodiments, the targeting sequence is SEQ ID NO: 224. In certain embodiments, the targeting sequence is SEQ ID NO: 225. In certain embodiments, the targeting sequence is SEQ ID NO: 226. In certain embodiments, the targeting sequence is SEQ ID NO: 227. In certain embodiments, the targeting sequence is SEQ ID NO: 228. In certain embodiments, the targeting sequence is SEQ ID NO: 229. In certain embodiments, the targeting sequence is SEQ ID NO: 230.
- the targeting sequence is SEQ ID NO: 231. In certain embodiments, the targeting sequence is SEQ ID NO: 232. In certain embodiments, the targeting sequence is SEQ ID NO: 233. In certain embodiments, the targeting sequence is SEQ ID NO: 234. In certain embodiments, the targeting sequence is SEQ ID NO: 235. In certain embodiments, the targeting sequence is SEQ ID NO: 236. In certain embodiments, the targeting sequence is SEQ ID NO: 237. In certain embodiments, the targeting sequence is SEQ ID NO: 238. In certain embodiments, the targeting sequence is SEQ ID NO: 239. In certain embodiments, the targeting sequence is SEQ ID NO: 240. In certain embodiments, the targeting sequence is SEQ ID NO: 241.
- the targeting sequence is SEQ ID NO: 242. In certain embodiments, the targeting sequence is SEQ ID NO: 243. In certain embodiments, the targeting sequence is SEQ ID NO: 244. In certain embodiments, the targeting sequence is SEQ ID NO: 245. In certain embodiments, the targeting sequence is SEQ ID NO: 246. In certain embodiments, the targeting sequence is SEQ ID NO: 247. In certain embodiments, the targeting sequence is SEQ ID NO: 248. In certain embodiments, the targeting sequence is SEQ ID NO: 249. In certain embodiments, the targeting sequence is SEQ ID NO: 250. In certain embodiments, the targeting sequence is SEQ ID NO: 251. In certain embodiments, the targeting sequence is SEQ ID NO: 252.
- the targeting sequence is SEQ ID NO: 253. In certain embodiments, the targeting sequence is SEQ ID NO: 254. In certain embodiments, the targeting sequence is SEQ ID NO: 255. In certain embodiments, the targeting sequence is SEQ ID NO: 256. In certain embodiments, the targeting sequence is SEQ ID NO: 257. In certain embodiments, the targeting sequence is SEQ ID NO: 258. In certain embodiments, the targeting sequence is SEQ ID NO: 259. In certain embodiments, the targeting sequence is SEQ ID NO: 260. In certain embodiments, the targeting sequence is SEQ ID NO: 261. In certain embodiments, the targeting sequence is SEQ ID NO: 262. In certain embodiments, the targeting sequence is SEQ ID NO: 263.
- the targeting sequence is SEQ ID NO: 264. In certain embodiments, the targeting sequence is SEQ ID NO: 265. In certain embodiments, the targeting sequence is SEQ ID NO: 266. In certain embodiments, the targeting sequence is SEQ ID NO: 267. In certain embodiments, the targeting sequence is SEQ ID NO: 268. In certain embodiments, the targeting sequence is SEQ ID NO: 269. In certain embodiments, the targeting sequence is SEQ ID NO: 270. In certain embodiments, the targeting sequence is SEQ ID NO: 271. In certain embodiments, the targeting sequence is SEQ ID NO: 272. In certain embodiments, the targeting sequence is SEQ ID NO: 273. In certain embodiments, the targeting sequence is SEQ ID NO: 274.
- the targeting sequence is SEQ ID NO: 275. In certain embodiments, the targeting sequence is SEQ ID NO: 276. In certain embodiments, the targeting sequence is SEQ ID NO: 277. In certain embodiments, the targeting sequence is SEQ ID NO: 278. In certain embodiments, the targeting sequence is SEQ ID NO: 279. In certain embodiments, the targeting sequence is SEQ ID NO: 280. In certain embodiments, the targeting sequence is SEQ ID NO: 281. In certain embodiments, the targeting sequence is SEQ ID NO: 282. In certain embodiments, the targeting sequence is SEQ ID NO: 283. In certain embodiments, the targeting sequence is SEQ ID NO: 284. In certain embodiments, the targeting sequence is SEQ ID NO: 285.
- the targeting sequence is SEQ ID NO: 286. In certain embodiments, the targeting sequence is SEQ ID NO: 287. In certain embodiments, the targeting sequence is SEQ ID NO: 288. In certain embodiments, the targeting sequence is SEQ ID NO: 289. In certain embodiments, the targeting sequence is SEQ ID NO: 290. In certain embodiments, the targeting sequence is SEQ ID NO: 291. In certain embodiments, the targeting sequence is SEQ ID NO: 292. In certain embodiments, the targeting sequence is SEQ ID NO: 293. In certain embodiments, the targeting sequence is SEQ ID NO: 294. In certain embodiments, the targeting sequence is SEQ ID NO: 295. In certain embodiments, the targeting sequence is SEQ ID NO: 296.
- the targeting sequence is SEQ ID NO: 297. In certain embodiments, the targeting sequence is SEQ ID NO: 298. In certain embodiments, the targeting sequence is SEQ ID NO: 299. In certain embodiments, the targeting sequence is SEQ ID NO: 300. In certain embodiments, the targeting sequence is SEQ ID NO: 301. In certain embodiments, the targeting sequence is SEQ ID NO: 302. In certain embodiments, the targeting sequence is SEQ ID NO: 303. In certain embodiments, the targeting sequence is SEQ ID NO: 304. In certain embodiments, the targeting sequence is SEQ ID NO: 305. In certain embodiments, the targeting sequence is SEQ ID NO: 306. In certain embodiments, the targeting sequence is SEQ ID NO: 307.
- the targeting sequence is SEQ ID NO: 308. In certain embodiments, the targeting sequence is SEQ ID NO: 309. In certain embodiments, the targeting sequence is SEQ ID NO: 310. In certain embodiments, the targeting sequence is SEQ ID NO: 311. In certain embodiments, the targeting sequence is SEQ ID NO: 312. In certain embodiments, the targeting sequence is SEQ ID NO: 313. In certain embodiments, the targeting sequence is SEQ ID NO: 314. In certain embodiments, the targeting sequence is SEQ ID NO: 315. In certain embodiments, the targeting sequence is SEQ ID NO: 316. In certain embodiments, the targeting sequence is SEQ ID NO: 317. In certain embodiments, the targeting sequence is SEQ ID NO: 318.
- the targeting sequence is SEQ ID NO: 319. In certain embodiments, the targeting sequence is SEQ ID NO: 320. In certain embodiments, the targeting sequence is SEQ ID NO: 321. In certain embodiments, the targeting sequence is SEQ ID NO: 322. In certain embodiments, the targeting sequence is SEQ ID NO: 323. In certain embodiments, the targeting sequence is SEQ ID NO: 324. In certain embodiments, the targeting sequence is SEQ ID NO: 325. In certain embodiments, the targeting sequence is SEQ ID NO: 326.
- the target region is within the exon internal region of exon 2 (SEQ ID NO: 2), the intron 1/exon 2 junction, the exon 2/intron 2 junction, or the intron internal region of intron 2 (SEQ ID NO: 11) of the human DNM2 gene pre-mRNA.
- the target region of the human DNM2 gene pre-mRNA is within the 21 st nucleotide to the 47 th nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2).
- the target region is a region within SEQ ID NO: 13.
- the target region is a region within DNM2 H2A(+21+47).
- the target region of the human DNM2 gene pre-mRNA is within the 23 rd nucleotide to the 51 st nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 14. In certain embodiments, the target region is a region within DNM2 H2A(+23+51). [0142] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 46 th nucleotide to the 72 nd nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2).
- the target region is a region within SEQ ID NO: 15. In certain embodiments, the target region is a region within DNM2 H2A(+46+72). [0143] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 13 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 24 th nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 16. In certain embodiments, the target region is a region within DNM2 H2D(+13-24).
- the target region of the human DNM2 gene pre-mRNA is within the 13 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 14 th nucleotide of intron 2 measured from the 3′ end of exon 2.
- the target region is a region within SEQ ID NO: 17.
- the target region is a region within DNM2 H2D(+13-14).
- the target region of the human DNM2 gene pre-mRNA is within the 10 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 19 th nucleotide of intron 2 measured from the 3′ end of exon 2.
- the target region is a region within SEQ ID NO: 18.
- the target region is a region within DNM2 H2D(+10-19).
- the target region of the human DNM2 gene pre-mRNA is within the 9 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 18 th nucleotide of intron 2 measured from the 3′ end of exon 2.
- the target region is a region within SEQ ID NO: 19.
- the target region is a region within DNM2 H2D(+9-18).
- the target region of the human DNM2 gene pre-mRNA is within the 6 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23 rd nucleotide of intron 2 measured from the 3′ end of exon 2.
- the target region is a region within SEQ ID NO: 20.
- the target region is a region within DNM2 H2D(+6-23).
- the target region of the human DNM2 gene pre-mRNA is within the 4 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23 rd nucleotide of intron 2 measured from the 3′ end of exon 2.
- the target region is a region within SEQ ID NO: 21.
- the target region is a region within DNM2 H2D(+4-23).
- the target region of the human DNM2 gene pre-mRNA is within the 1 st nucleotide to the 30 th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 22. In certain embodiments, the target region is a region within DNM2 H2D(-1-30).
- the target region of the human DNM2 gene pre-mRNA is within the 1 st nucleotide to the 29 th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 23. In certain embodiments, the target region is a region within DNM2 H2D(-1-29).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12- 13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2 H2A (-10+15),
- the target region of the human DNM2 gene pre-mRNA is DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9- 16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23) , DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6- 30).
- the target region is DNM2 H2A(+21+45) or DNM2 H2A(+23+47). In certain embodiments, the target region is DNM2 H2A(+23+47) or DNM2 H2A(+27+51). In certain embodiments, the target region is DNM2 H2A(+46+70) or DNM2 H2A(+48+72).
- the target region is DNM2 H2D(+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), or DNM2 H2D(+1-24).
- the target region is DNM2 H2D (+13-12), DNM2 H2D(+12-13), or DNM2 H2D(+11-14). [0155] In certain embodiments, the target region is DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), or DNM2 H2D(+6-19). In certain embodiments, the target region is DNM2 H2D(+9-16), DNM2 H2D(+8-17), or DNM2 H2D(+7-18). In certain embodiments, the target region is DNM2 H2D(+8-17).
- the target region is DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23). In certain embodiments, the target region is DNM2 H2D(+4-21), DNM2 H2D(+3-22), or DNM2 H2D(+2-23). In certain embodiments, the target region is DNM2 H2D(+3-22).
- the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30). In certain embodiments, the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), or DNM2 H2D(-5-29). [0158] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(-7+17). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H2A(+3+27). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+15+37). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+37+61). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2D(+20-5).
- the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGGCCGC); SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC); SEQ ID NO: 77 (TTTACCTGTTT
- B is an abasic subunit.
- the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTGTT);
- the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTC
- the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); and SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG).
- the targeting sequence is selected from: SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); and SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT).
- the targeting sequence is selected from: SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); and SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT). [0168] In certain embodiments, the targeting sequence is SEQ ID NO: 83.
- the targeting sequence is selected from: SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); and SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT).
- the targeting sequence is selected from: SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); and SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT). [0171] In certain embodiments, the targeting sequence is SEQ ID NO: 88.
- the targeting sequence is selected from: SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT); and SEQ ID NO: 96 (CCTGCTGAACCTCAGGCCGCCCCAT).
- the targeting sequence is selected from: SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); and SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT).
- the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGAGGCCGC); and SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC), wherein “B” represents an abasic subunit or guanine. In certain embodiments, B is an abasic subunit.
- the target region is within the exon internal region of exon 9 (SEQ ID NO: 6), the intron 8/exon 9 junction, or the exon 9/intron 9 junction of the human DNM2 gene pre-mRNA.
- the target region of the human DNM2 gene pre-mRNA is within the 36 th nucleotide to the 61 st nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6).
- the target region is a region within SEQ ID NO: 24.
- the target region is a region within DNM2 H9A(+36+61).
- the target region of the human DNM2 gene pre-mRNA is within the 42 nd nucleotide to the 68 th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6). In certain embodiments, the target region is a region within SEQ ID NO: 25. In certain embodiments, the target region is a region within DNM2 H9A(+42+70). [0178] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 43 rd nucleotide to the 68 th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6).
- the target region is a region within SEQ ID NO: 26. In certain embodiments, the target region is a region within DNM2 H9A(+42+68). [0179] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 24 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15 th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 27. In certain embodiments, the target region is a region within DNM2 H9D(+24-15).
- the target region of the human DNM2 gene pre-mRNA is within the 24 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 5 th nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 28.
- the target region is a region within DNM2 H9D(+24-5).
- the target region of the human DNM2 gene pre-mRNA is within the 19 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 10 th nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 29.
- the target region is a region within DNM2 H9D(+19-10).
- the target region of the human DNM2 gene pre-mRNA is within the 9 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22 nd nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 31.
- the target region is a region within DNM2 H9D(+9-22).
- the target region of the human DNM2 gene pre-mRNA is within the 9 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 20 th nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 32.
- the target region is a region within DNM2 H9D(+9-20).
- the target region of the human DNM2 gene pre-mRNA is within the 9 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 18 th nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 33.
- the target region is a region within DNM2 H9D(+9-18).
- the target region of the human DNM2 gene pre-mRNA is within the 8 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 21 st nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 34.
- the target region is a region within DNM2 H9D(+8-21).
- the target region of the human DNM2 gene pre-mRNA is within the 6 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22 nd nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 35.
- the target region is a region within DNM2 H9D(+6-22).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13), DNM2 H9A(-5+20), DNM2 H9A(+1+25),DNM2 H9A(+6+30), DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(N-N-N-(+
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+43+67), DNM2 H9A(+42+66), DNM2 H9A(+44+68), DNM2 H9D(+24-1), DNM2 H9D(+23- 2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+36+60) or DNM2 H9A(+37+61).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), or DNM2 H9A(+48+67).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), or DNM2 H9A(+44+68).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+67).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11- 14), or DNM2 H9D(+10-15).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), or DNM2 H9D(+20-5). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), or DNM2 H9D(+15-10).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), or DNM2 H9D(+10-15).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6- 17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6- 17), DNM2 H9D(+5-17), or DNM2 H9D(+5-20).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), or DNM2 H9D(+7-18).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), or DNM2 H9D(+4-21).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-5+20).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H9A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+6+30). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+25+49).
- the targeting sequence is selected from: SEQ ID NO: 158 (CGTCAAACTCCATCTGAGAAGAGAC); SEQ ID NO: 159 (TCCTTCTCGTCAAACTCCATCTGAG); SEQ ID NO: 160 (GTAAGTCCTTCTCGTCAAACTCCAT); SEQ ID NO: 161 (CCGTCGTAAGTCCTTCTCGTCAAAC); SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG); SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT); SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 (TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (C
- the targeting sequence is selected from: SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG); SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT); SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 181 (TGCCTGACT
- the targeting sequence is selected from: SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); and SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT).
- the targeting sequence is selected from: SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG);
- the targeting sequence is selected from: SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); and SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG).
- the targeting sequence is SEQ ID NO: 172.
- the targeting sequence is selected from: SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT); SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG); SEQ ID NO: 189
- the targeting sequence is selected from: SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); and SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT).
- the targeting sequence is selected from: SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); and SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG).
- the targeting sequence is selected from: SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 (TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); and SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT).
- the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACT
- the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); and SEQ ID NO: 205 (ACTCTCCTCGTGGA
- the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); and SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC).
- the targeting sequence is selected from: SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); and SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA).
- the targeting sequence is SEQ ID NO: 199.
- the targeting sequence is selected from: SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 208 (CTCCTCGTGGAACTTGCCTG); and SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG).
- the targeting sequence is selected from: SEQ ID NO: 158 (CGTCAAACTCCATCTGAGAAGAGAC); SEQ ID NO: 159 (TCCTTCTCGTCAAACTCCATCTGAG); SEQ ID NO: 160 (GTAAGTCCTTCTCGTCAAACTCCAT); SEQ ID NO: 161 (CCGTCGTAAGTCCTTCTCGTCAAAC); SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); and SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG).
- the target region is within the exon internal region of exon 12 (SEQ ID NO: 7), the intron 11/exon 12 junction, or the exon 12/intron 12 junction of the human DNM2 gene pre-mRNA.
- the target region of the human DNM2 gene pre-mRNA is within the 4 th nucleotide to the 31 st nucleotide measured of exon 12 from the 5′ end of exon 12 (SEQ ID NO: 7).
- the target region is a region within SEQ ID NO: 36.
- the target region is a region within DNM2 H12A(+4+31).
- the target region of the human DNM2 gene pre-mRNA is within the 10 th nucleotide to the 44 th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 37. In certain embodiments, the target region is a region within DNM2 H12A(+10+44). [0220] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 10 th nucleotide to the 38 th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
- the target region is a region within SEQ ID NO: 38. In certain embodiments, the target region is a region within DNM2 H12A(+10+38). [0221] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 16 th nucleotide to the 44 th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 39. In certain embodiments, the target region is a region within DNM2 H12A(+16+44).
- the target region of the human DNM2 gene pre-mRNA is within the 14 th nucleotide to the 42 nd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 40. In certain embodiments, the target region is a region within DNM2 H12A(+14+42). [0223] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 25 th nucleotide to the 54 th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
- the target region is a region within SEQ ID NO: 41. In certain embodiments, the target region is a region within DNM2 H12A(+25+54). [0224] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 25 th nucleotide to the 53 rd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 42. In certain embodiments, the target region is a region within DNM2 H12A(+25+53).
- the target region of the human DNM2 gene pre-mRNA is within the 26 th nucleotide to the 54 th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 43. In certain embodiments, the target region is a region within DNM2 H12A(+26+54). [0226] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 36 th nucleotide to the 61 st nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
- the target region is a region within SEQ ID NO: 44. In certain embodiments, the target region is a region within DNM2 H12A(+36+61). [0227] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 22nd nucleotide of exon 12 measured from the 3′ end of exon 12 (SEQ ID NO: 7) to the 5th nucleotide of intron 12 measured from the 3′ end of exon 12. In certain embodiments, the target region is a region within SEQ ID NO: 45. In certain embodiments, the target region is a region within DNM2 H12D(+22-5).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-13+12), DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12
- the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+49), DNM2 H12A(+29+53), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H
- the target region is DNM2 H12A(+4+28) or DNM2 H12A(+7+31).
- the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44).
- the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), or DNM2 H12A(+14+38). [0233] In certain embodiments, the target region is DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), or DNM2 H12A(+20+44).
- the target region is DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), or DNM2 H12A(+18+42).
- the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54).
- the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), or DNM2 H12A(+29+53).
- the target region is DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54).
- the target region is DNM2 H12A(+25+49).
- the target region is DNM2 H12A(+36+60) or DNM2 H12A(+37+61).
- the target region is DNM2 H12D(+22-3) or DNM2 H12D(+20-5).
- the target region is DNM2 H12A(-13+12). In certain embodiments, the target region is DNM2 H12A(-3+22). In certain embodiments, the target region is DNM2 H12A(+43+67). In certain embodiments, the target region is DNM2 H12D(+15-10).
- the targeting sequence is selected from: SEQ ID NO: 210 (CAGCAGAAGAATCTGGBGGAGAGCG); SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG); SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 210 (CAG
- the targeting sequence is selected from: SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG); SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ
- the targeting sequence is selected from: SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); and SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG).
- the targeting sequence is selected from: SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222
- the targeting sequence is selected from: SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); and SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA).
- the targeting sequence is selected from: SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); and SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA).
- the targeting sequence is selected from: SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); and SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG).
- the targeting sequence is selected from: SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAGG);
- the targeting sequence is selected from: SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); and SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG).
- the targeting sequence is selected from: SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); and SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG).
- the targeting sequence is SEQ ID NO: 230
- the targeting sequence is selected from: SEQ ID NO: 238 (GATGAAGTCCTCATGGTTCGTGTTG); and SEQ ID NO: 239 (CGATGAAGTCCTCATGGTTCGTGTT).
- the targeting sequence is selected from: SEQ ID NO: 241 (TACTTGGCAAACCCGATGAAGTCCT); and SEQ ID NO: 242 (CCTACTTGGCAAACCCGATGAAGTC).
- the targeting sequence is selected from: SEQ ID NO: 210 (CAGCAGAAGAATCTGGBGGAGAGCG); SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 240 (CAAACCCGATGAAGTCCTCATGGTT); and SEQ ID NO: 243 (AAGTACCTACTTGGCAAACCCGATG).
- the target region is within the exon internal region of exon 13 (SEQ ID NO: 8), the intron 12/exon 13 junction, the exon 13/intron 13 junction, or the intron internal region of intron 12 (SEQ ID NO: 12).
- the target region of the human DNM2 gene pre-mRNA is within the 27 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12 th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 46. In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within DNM2 H13A(-27+12).
- the target region of the human DNM2 gene pre-mRNA is within the 27 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 2 nd nucleotide of exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 47.
- the target region is a region within DNM2 H13A(-27+2).
- the target region of the human DNM2 gene pre-mRNA is within the 22 nd nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 7 th nucleotide of exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 48.
- the target region is a region within DNM2 H13A(-22+7).
- the target region of the human DNM2 gene pre-mRNA is within the 17 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12 th nucleotide of exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 49.
- the target region is a region within DNM2 H13A(-17+12).
- the target region of the human DNM2 gene pre-mRNA is within the 11 th nucleotide of intron 12 (SEQ ID NO: 12) measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24 th nucleotide of exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 50.
- the target region is a region within DNM2 H13A(-11+24).
- the target region of the human DNM2 gene pre-mRNA is within the 11 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 18 th nucleotide of exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 51.
- the target region is a region within DNM2 H13A(-11+18).
- the target region of the human DNM2 gene pre-mRNA is within the 5 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24 th nucleotide exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 52.
- the target region is a region within DNM2 H13A(-5+24).
- the target region of the human DNM2 gene pre-mRNA is within the 8 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 21 st nucleotide exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 53. In certain embodiments, the target region is a region within DNM2 H13A(-8+21). [0262] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8).
- the target region is a region within SEQ ID NO: 54. In certain embodiments, the target region is a region within DNM2 H13A(+1+30). [0263] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 28th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 55. In certain embodiments, the target region is a region within DNM2 H13A(+1+28).
- the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 56. In certain embodiments, the target region is a region within DNM2 H13A(+6+30). [0265] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 3rd nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8).
- the target region is a region within SEQ ID NO: 57. In certain embodiments, the target region is a region within DNM2 H13A(+3+30). [0266] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15),
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), or DNM2 H13A(- 23+2).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), or DNM2 H13A(- 18+7).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(- 7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(-1+24).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), or DNM2 H13A(-7+18).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(- 1+24).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), or DNM2 H13A(- 4+21).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), or DNM2 H13A(+5+28).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), or DNM2 H13A(+6+30).
- the target region of the human DNM2 gene pre-mRNA DNM2 H13A(+4+28).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+16+40). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H13D(+22-3). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13D(+7-18).
- the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 254 (GCTGG
- B is an abasic subunit.
- the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCT
- the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 254 (GCTGG
- the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); and SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG).
- the targeting sequence is selected from: SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); and SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA).
- the targeting sequence is selected from: SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); and SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA).
- the targeting sequence is selected from: SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); and SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC).
- the targeting sequence is selected from: SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); and SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC).
- the targeting sequence is selected from: SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); and SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC).
- the targeting sequence is selected from: SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); and SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA).
- the targeting sequence is selected from: SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (CAGCTGCGTGCTCC
- the targeting sequence is selected from: SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); and SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG).
- the targeting sequence is selected from: SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 282 (GTTCAGCTGCGTGCTCCTCTGC); SEQ ID NO: 283 (GTTCAGCTGCGTGCTCCTCTG); and SEQ ID NO: 284 (GTTCAGCTGCGTGCTCCTCT).
- the targeting sequence is selected from: SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); and SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT).
- the targeting sequence is selected from: SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 285 (GGCTCTCTTCTTGTTCAGCTGCGTG); SEQ ID NO: 286 (TACCTGATTGGBGATGGCTCTCTTC); and SEQ ID NO: 287 (TGCAGAGGBGTGTGCTACCTGATTG).
- the target region is within the exon internal region of exon 6 (SEQ ID NO: 3), the intron 5/exon 6 junction, the exon 6/intron 6 junction, or the intron internal region of intron 6 (SEQ ID NO: 11).
- the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 37th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 58. In certain embodiments, the target region is a region within DNM2 H6A(+10+37). [0298] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 32nd nucleotide to the 59th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3).
- the target region is a region within SEQ ID NO: 59. In certain embodiments, the target region is a region within DNM2 H6A(+32+59). [0299] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 60th nucleotide to the 87th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 60. In certain embodiments, the target region is a region within DNM2 H6A(+60+87).
- the target region of the human DNM2 gene pre-mRNA is within the 130th nucleotide to the 156th nucleotide measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 61. In certain embodiments, the target region is a region within DNM2 H6A(+130+156). [0301] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 17th nucleotide of exon 6 measured from the 3′ end of exon 6 (SEQ ID NO: 3) to the 10th nucleotide of intron 6 measured from the 3′ end of exon 6.
- the target region is a region within SEQ ID NO: 62. In certain embodiments, the target region is a region within DNM2 H6D(+17-10). [0302] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5), DNM2 H6A(-6+19), DNM2 H6A(-2+23), DNM2 H6A(+3+26), DNM2 H6A(+10+3), DNM2 H6A(+13+37), DNM2 H6A(+23+47), DNM2 H6A(+32+56), DNM2 H6A(+35+59), DNM2 H6A(+44+68), DNM2 H6A(+54+78), DNM2 H6A(+60+84), DNM2 H6A(+63+87), DNM2 H6A(+88+112), DNM2 H6A(+96+119), DNM2 H6A(+105
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+10+3) or DNM2 H6A(+13+37).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+32+56) or DNM2 H6A(+35+59).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+60+84) or DNM2 H6A(+63+87).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+130+154) or DNM2 H6A(+132+156). [0307] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+15-10) or DNM2 H6D (+17-8). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-6+19). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-2+23).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+3+26). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+23+47). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+44+68). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+54+78). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+88+112).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H6A(+96+119). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+105+129). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+121+145). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+6-19).
- the targeting sequence is selected from: SEQ ID NO: 97 (GTAGCCTGTGGGGAGAGGCAGTCAG); SEQ ID NO: 98 (TTCACCACGCCAATGTAGCCTGTGG); SEQ ID NO: 99 (GCGGTTCACCACGCCAATGTAGCCT); SEQ ID NO: 100 (GCTGCGGTTCACCACGCCAATGTA); SEQ ID NO: 101 (TCCTTCTGGCTGCGGTTCACCACGC); SEQ ID NO: 102 (ATATCCTTCTGGCTGCGGTTCACCA); SEQ ID NO: 103 (CTTGCCCTCAATATCCTTCTGGCTG); SEQ ID NO: 104 (GATGTCCTTCTTGCCCTCAATATCC); SEQ ID NO: 105 (ACGGATGTCCTTCTTGCCCTCAATA); SEQ ID NO: 106 (CAGTGCTGCACGGATGTCCTTCTTG); SEQ ID NO: 107 (TCTCAGCTGCCAGTGCTGCACGGAT); SEQ ID NO: 107 (TCTCAGCTG
- the target region is within the exon internal region of exon 7 (SEQ ID NO: 4), the intron 6/exon 7 junction, or the exon 7/intron 7 junction.
- the target region of the human DNM2 gene pre-mRNA is within the 72nd nucleotide to the 98th nucleotide of exon 7 measured from the 5′ end of exon 7 (SEQ ID NO: 4).
- the target region is a region within SEQ ID NO: 63.
- the target region is a region within DNM2 H7A(+72+98).
- the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 7 measured from the 3′ end of exon 7 (SEQ ID NO: 4) to the 16th nucleotide of intron 7 measured from the 3′ end of exon 7.
- the target region is a region within SEQ ID NO: 64.
- the target region is a region within DNM2 H7D(+9-16).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+1+25), DNM2 H7A(+6+30), DNM2 H7A(+11+35), DNM2 H7A(+22+46), DNM2 H7A(+29+53), DNM2 H7A(+41+65), DNM2 H7A(+51+75), DNM2 H7A(+57+81), DNM2 H7A(+62+86), DNM2 H7A(+72+96), DNM2 H7A(+74+98), DNM2 H7A(+81+105), DNM2 H7A(+91+115), or DNM2 H7A(+110+134).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+72+96) or DNM2 H7A(+74+98).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+8-16) or DNM2 H7D(+9-16).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-15+10). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-5+20).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H7A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+6+30). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+11+35). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+22+46). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+29+53). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+41+65).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+51+75). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+57+81). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+62+86). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H7A(+81+105). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+91+115).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+110+134). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+21-4).
- the targeting sequence is selected from: SEQ ID NO: 119 (TGGTCAGTTGCTGCAGGAGAGAGGT); SEQ ID NO: 120 (CGGATGTGGTTGGTCAGTTGCTGCA); SEQ ID NO: 121 (ACTCCCGGATGTGGTTGGTCAGTTG); SEQ ID NO: 122 (CAGCGACTCCCGGATGTGGTTGGTC); SEQ ID NO: 123 (GCCGGCAGCGACTCCCGGATGTGGT); SEQ ID NO: 124 (TGCTACGTAGGGCCGGCAGCGACTC); SEQ ID NO: 125 (TGTAGTTTGCTACGTAGGGCCGGCA); SEQ ID NO: 126 (AGCAGCTGGCTCTGTAGTTTGCTAC); SEQ ID NO: 129 (AGCAGCTGGCTC
- B is an abasic subunit.
- the target region is within the exon internal region of exon 8 (SEQ ID NO: 5), the intron 7/exon 8 junction, or the exon 8/intron 8 junction.
- the target region of the human DNM2 gene pre-mRNA is within the 7th nucleotide of intron 7 measured from the 5′ end of exon 8 (SEQ ID NO: 5) to the 19th nucleotide of exon 8 measured from the 5′ end of exon 8.
- the target region is a region within SEQ ID NO: 65.
- the target region is a region within DNM2 H8A(-7+19).
- the target region of the human DNM2 gene pre-mRNA is within the 18th nucleotide to the 45th nucleotide of exon 8 measured from the 5′ end of exon 8 (SEQ ID NO: 5). In certain embodiments, the target region is a region within SEQ ID NO: 66. In certain embodiments, the target region is a region within DNM2 H8A(+18+45).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-20+5), DNM2 H8A(-14+11), DNM2 H8A(-7+18), DNM2 H8A(-6+19), DNM2 H8A(-2+23), DNM2 H8A(+5+29), DNM2 H8A(+18+42), DNM2 H8A(+21+45), DNM2 H8A(+27+51), DNM2 H8A(+36+60), DNM2 H8A(+46+70), DNM2 H8A(+51+75), DNM2 H8A(+56+80), DNM2 H8A(+65+88), DNM2 H8A(+76+100), DNM2 H8A(+83+107), DNM2 H8A(+96+119), DNM2 H8A(+105+129), DNM2 H8D(+18-7), or DNM2 H8D(+
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-7+18) or DNM2 H8A(-6+19).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+18+42) or DNM2 H8A(+21+45).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-20+5).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-14+11).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H8A(-2+23).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+5+29). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+27+51). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+36+60). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+46+70). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+51+75). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+56+80).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+65+88). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+76+100). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H8A(+83+107). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+96+119). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+105+129). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+18-7).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+10-15).
- the targeting sequence is selected from: SEQ ID NO: 138 (CCATCCTATGAGGAAAAGGAAAAAG); SEQ ID NO: 139 (GCTGGACCATCCTATGAGGAAAAGG); SEQ ID NO: 140 (CCAAACTGCTGGACCATCCTATGAG); SEQ ID NO: 141 (CCCAAACTGCTGGACCATCCTATGA); SEQ ID NO: 142 (CCACCCCAAACTGCTGGACCATCCT); SEQ ID NO: 143 (CAAAATCCACCCCAAACTGCTGGAC); SEQ ID NO: 144 (TCGATCCTCTTCTCAAAATCCACCC); SEQ ID NO: 145 (CCCTCGATCCTCTTCTCAAAATCCA); SEQ ID NO: 146 (CCTGAGCCCTCGATCCTCTTCTCAA); SEQ ID NO: 147 (ACCTGATCTCCTG
- the target region is within the exon internal region of exon 16 (SEQ ID NO: 9), the intron 15/exon 16 junction, or the exon 16/intron 16 junction.
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3), DNM2 H16A(-13+12), DNM2 H16A(-6+19), DNM2 H16A(+6+30), DNM2 H16A(+15+39), DNM2 H16A(+26+50), DNM2 H16A(+36+60), DNM2 H16A(+45+69), DNM2 H16A(+55+79), DNM2 H16A(+66+90), DNM2 H16A(+75+99), DNM2 H16A(+82+106), DNM2 H16D(+19-6), or DNM2 H16D(+11-14).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-13+12).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H16A(-6+19).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+6+30).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+15+39).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+26+50). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+36+60). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+45+69). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+55+79). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+66+90). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+75+99).
- the targeting sequence is selected from: SEQ ID NO: 288 (CTCCTGAGGGTGGAGGGAAAAGAG); SEQ ID NO: 289 (CTTCTCTTTCTCCTGAGGGTGGAGG); SEQ ID NO: 290 (TGTACTTCTTCTCTTTCTCCTGAGG); SEQ ID NO: 291 (CAGAGGCAGCATGTACTTCTTCTCT); SEQ ID NO: 292 (GAGGTTGTCCAGAGGCAGCATGTAC); SEQ ID NO: 293 (TCACGGATCTTGAGGTTGTCCAGAG); SEQ ID NO: 294 (CTTCTCCACATCACGGATCTTGAGG); SEQ ID NO: 295 (CATGAAGCCCTTCTCCACATCACGG); SEQ ID NO: 296 (GCTTGTTGGACATGAAGCCCTTCTC); SEQ ID NO: 297 (GGCGAAGACGTGCTTGTTGGACATG); SEQ ID NO: 298 (GTTGAAGATGGCGAAGACGTGCTTG); SEQ ID NO: 298 (GTTGAAG
- the target region is within the exon internal region of exon 19 (SEQ ID NO: 9), the intron 18/exon 19 junction, or the exon 18/intron 18 junction.
- the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14), DNM2 H19A(-4+21), DNM2 H19A(+1+25), DNM2 H19A(+7+31), DNM2 H19A(+16+40), DNM2 H19A(+26+50), DNM2 H19A(+37+61), DNM2 H19A(+47+71), DNM2 H19A(+56+80), DNM2 H19A(+66+90), DNM2 H19A(+76+100), DNM2 H19A(+86+110), DNM2 H19A(+96+120), DNM2 H19A(+106+130), DNM2 H19A(+117+141), DNM
- the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-4+21). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+7+31). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+16+40).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+26+50). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+37+61). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+47+71). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+56+80). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+66+90). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+76+100).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+86+110). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+96+120). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+106+130). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+117+141). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+127+151).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+137+161). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+147+171). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+157+181). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+166+189). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+191+215).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+200+224). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+23-2). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+17-8). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+10-15).
- the targeting sequence is selected from: SEQ ID NO: 302 (ATGAAGGCCTTCGTCTGCGGGCAGG); SEQ ID NO: 303 (GTGGTGGATGAAGGCCTTCGTCTGC); SEQ ID NO: 304 (GCTCGTGGTGGATGAAGGCCTTCGT); SEQ ID NO: 305 (CCAGCAGCTCGTGGTGGATGAAGGC); SEQ ID NO: 306 (ATAGGTAGGCCAGCAGCTCGTGGTG); SEQ ID NO: 307 (GCCGAGGAGTATAGGTAGGCCAGCA); SEQ ID NO: 308 (TGCTCTGGTCTGCCGAGGAGTATAG); SEQ ID NO: 309 (TCCATGAGGCTGCTCTGGTCTGCCG); SEQ ID NO: 310 (GCCGACTCCTCCATGAGGCTGCTCT); SEQ ID NO: 311 (TGCCTGGTCAGCCGACTCCTCCATG); SEQ ID NO: 312 (CCCGCCGCTGTGCCTGGTCAGCCGA); SEQ ID NO:
- each R 2 , taken together forms a targeting sequence having SEQ ID NO: 83. In certain embodiments, each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 88. In certain embodiments, each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 172. In certain embodiments, R 2 , taken together, forms a targeting sequence having SEQ ID NO: 199. In certain embodiments, each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 230. In certain embodiments, each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 278.
- the targeting sequence is at least 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99 percent complementary to the target region.
- the targeting sequence is at least 84%, at least 88%, or at least 92% complementary to the target region.
- the targeting sequence is at least 90% complementary to the target region.
- the targeting sequence is at least 95% complementary to the target region.
- the targeting sequence is 100% complementary to the target region.
- the antisense oligomer of the present disclosure can have a modified chemical backbone as described herein or known in the art.
- the antisense oligomer may be a peptide nucleic acid (PNA), a locked nucleic acid, a phosphorodiamidate morpholino oligomer, a 2′-OMe phosphorothioate oligomer, or a combination thereof.
- the antisense oligomer is a phosphorodiamidate morpholino oligomer.
- the antisense oligomer is a peptide nucleic acid.
- the antisense oligomer is a locked nucleic acid.
- the antisense oligomer is a 2′-OMe phosphorothioate oligomer. In still another embodiment, the antisense oligomer is a combination of any of the above oligonucleotides having a modified chemical backbone. [0336] In some embodiments, the antisense oligomer is conjugated to, e.g., covalently attached to, a delivery moiety such as, for example, a cell-penetrating peptide, an antibody, a fragment of an antibody, or an antigen fragment of an antibody. In an embodiment, the antisense oligomer is conjugated to a cell-penetrating peptide.
- the antisense oligomer is conjugated to a cell-penetrating peptide, wherein the cell-penetrating peptide is selected from rTAT, Tat, R9F2, R5F2R4, R4, R5, R6, R7, R8, R9, (RXR)4, (RXR)5, (RXRRBR)2, (RAR)4F2, (RGR)4F2, and RBRBYLIQFRBRRBR, wherein A represents alanine, B represents beta-alanine, F represents phenylalanine, G represents glycine, I represents isoleucine, L represents leucine, Q represents glutamine, R represents arginine, X represent 6-aminohexanoic acid, and Y represents tyrosine.
- the cell-penetrating peptide is selected from rTAT, Tat, R9F2, R5F2R4, R4, R5, R6, R7, R8, R9, (RXR)4, (RXR
- the antisense oligomer is conjugated to an antibody, an antibody fragment, or an antigen fragment of an antibody.
- the antisense oligomer of the present disclosure may be attached to the delivery moiety via a linker.
- the linker is a direct bond from the antisense oligomer to the delivery moiety.
- the linker is an amino acid such as, for example, a glycine amino acid, a proline amino acid, or a glutamic amino acid.
- the linker is a glycine amino acid or a proline amino acid.
- an antisense oligomer or a pharmaceutically acceptable salt thereof, having a structure of Formula (I): (I), wherein A′, R 1 , R 2 , t, E′, and G are as described herein.
- t is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28.
- each R 2 taken together forms a targeting sequence of 13-30 bases.
- each R 2 taken together forms a targeting sequence of 11-28 bases.
- each R 2 taken together forms a targeting sequence of 18-27 bases.
- each R 2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases.
- the pharmaceutically acceptable salt is an HCl salt.
- E′ is selected from H, -C1-6-alkyl, -C(O)C1-6-alkyl, benzoyl, stearoyl, trityl, monomethoxytrityl, dimethoxytrityl, trimethoxytrityl, and .
- E′ is selected from -H, -C(O)CH3, benzoyl, stearoyl, trityl, 4- methoxytrityl, and .
- A′ is selected from: and E′ is .
- A′ is [0344]
- E′ is selected from H, -C(O)CH 3 , trityl, 4-methoxytrityl, benzoyl, and stearoyl.
- each R 1 is -N(CH3)2.
- L is glycine, proline, or b-alanine.
- L is glycine. In an embodiment, L is proline. In an embodiment, L is b-alanine. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt. [0345] In some embodiments, provided herein is an antisense oligomer, which is an oligomer of Formula (IA):
- each R 2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases.
- the pharmaceutically acceptable salt is an HCl salt.
- an antisense oligomer which is an oligomer of Formula (II): or a pharmaceutically acceptable salt thereof, wherein t, G, and R 2 are as described herein.
- t is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28.
- each R 2 taken together forms a targeting sequence of 13-30 bases.
- each R 2 taken together forms a targeting sequence of 11-28 bases.
- each R 2 taken together forms a targeting sequence of 18-27 bases.
- each R 2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases.
- the pharmaceutically acceptable salt is an HCl salt.
- an antisense oligomer which is an oligomer of Formula (III): or a pharmaceutically acceptable salt thereof, wherein n is 11-28 and R 2 is as described herein.
- n is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28.
- each R 2 taken together forms a targeting sequence of 13- 30 bases.
- each R 2 taken together forms a targeting sequence of 11-28 bases.
- each R 2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt.
- an antisense oligomer which is an oligomer of Formula (IV): (IV), wherein n is 11-28 and R 2 is as described herein.
- n is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28.
- each R 2 taken together forms a targeting sequence of 13-30 bases.
- each R 2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases.
- J is selected from rTAT, TAT, R 9 F 2 , R 5 F 2 R 4 , R 4 , R 5 , R 6 , R 7 , R 8 , R 9 , (RXR) 4 , (RXR) 5 , (RXRRBR) 2 , (RAR) 4 F 2 , (RGR) 4 F 2 , and RBRBYLIQFRBRRBR, wherein A represents alanine, B represents ⁇ beta-alanine, F represents phenylalanine, G represents glycine, R represents arginine, and X represent 6-aminohexanoic acid.
- G is selected from H, C(O)CH 3 , benzoyl, and stearoyl. In an embodiment, G is H or -C(O)CH 3 . In an embodiment, G is H. In an embodiment, G is -C(O)CH 3 . [0355] In certain embodiments, the antisense oligomer is selected from: ,
- the antisense oligomer has the structure: . [0357] In certain embodiments, the antisense oligomer has the structure:
- the antisense oligomer has the structure:
- the antisense oligomer has the structure:
- the antisense oligomer has the structure:
- the antisense oligomer has the structure: .
- a pharmaceutical composition comprising: an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formulae (I), (IA), (II), (III), (IV) and/or (V); and at least one pharmaceutically acceptable carrier.
- a method of treating a disease in a subject in need thereof comprising administering a therapeutically effective amount of an antisense oligomer, including an oligomer of Formulae (I), (IA), (II), (III), (IV), and/or (V), or a pharmaceutical composition disclosed herein to the subject.
- the disease is associated with aberrant expression of dynamin 2 (DNM2) protein.
- the disease is a centronuclear myopathy (CNM).
- the subject is a human.
- modified antisense oligomers include, without limitation, morpholino oligomers, phosphorothioate modified oligomers, 2′-O-methyl modified oligomers, peptide nucleic acid (PNA), locked nucleic acid (LNA), phosphorothioate oligomers, 2′-O-MOE modified oligomers, 2′-fluoro-modified oligomer, 2′-O,4′-C-ethylene-bridged nucleic acids (ENAs), tricyclo-DNAs, tricyclo-DNA phosphorothioate subunits, 2′-O-[2-(N- methylcarbamoyl)ethyl] modified oligomers, including combinations of any of the foregoing.
- nucleobases of the modified antisense oligomer are linked to morpholino ring structures, wherein the morpholino ring structures are joined by phosphoros- containing intersubunit linkages joining a morpholino nitrogen of one ring structure to a 5′ exocyclic carbon of an adjacent ring structure.
- the nucleobases of the antisense oligomer are linked to a peptide nucleic acid (PNA), wherein the phosphate-sugar polynucleotide backbone is replaced by a flexible pseudo-peptide polymer to which the nucleobases are linked.
- PNA peptide nucleic acid
- at least one of the nucleobases of the antisense oligomer is linked to a locked nucleic acid (LNA), wherein the locked nucleic acid structure is a nucleotide analog that is chemically modified where the ribose moiety has an extra bridge connecting the 2′ oxygen and the 4′ carbon.
- At least one of the nucleobases of the antisense oligomer is linked to a bridged nucleic acid (BNA), wherein the sugar conformation is restricted or locked by the introduction of an additional bridged structure to the furanose skeleton.
- BNA bridged nucleic acid
- at least one of the nucleobases of the antisense oligomer is linked to a 2′-O,4′-C-ethylene- bridged nucleic acid (ENA).
- ENA 2′-O,4′-C-ethylene- bridged nucleic acid
- the modified antisense oligomer may contain unlocked nucleic acid (UNA) subunits.
- UNAs and UNA oligomers are an analogue of RNA in which the C2′-C3′ bond of the subunit has been cleaved.
- the modified antisense oligomer contains one or more phosphorothioates (or S-oligos), in which one of the nonbridging oxygens is replaced by sulfur.
- the modified antisense oligomer contains one or more 2′ O-Methyl, 2′ O-MOE, MCE, and 2′-F in which the 2′-OH of the ribose is substituted with a methyl, methoxy ethyl, 2-(N- methylcarbamoyl)ethyl, or fluoro group, respectively.
- the modified antisense oligomer is a tricyclo-DNA (tc-DNA) which is a constrained DNA analog in which each nucleotide is modified by the introduction of a cyclopropane ring to restrict conformational flexibility of the backbone and to optimize the backbone geometry of the torsion angle g.
- tc-DNA tricyclo-DNA
- at least one of the nucleobases of the antisense oligomer is linked to a bridged nucleic acid (BNA), wherein the sugar conformation is restricted or locked by the introduction of an additional bridged structure to the furanose skeleton.
- BNA bridged nucleic acid
- PNAs Peptide Nucleic Acids
- PNAs Peptide nucleic acids
- PNAs are analogs of DNA in which the backbone is structurally homomorphous with a deoxyribose backbone, consisting of N-(2-aminoethyl) glycine units to which pyrimidine or purine bases are attached.
- PNAs containing natural pyrimidine and purine bases hybridize to complementary oligomers obeying Watson-Crick base-pairing rules, and mimic DNA in terms of base-pair recognition.
- the backbone of PNAs is formed by peptide bonds rather than phosphodiester bonds, making them well-suited for antisense applications (see structure below).
- the backbone is uncharged, resulting in PNA/DNA or PNA/RNA duplexes that exhibit greater than normal thermal stability.
- PNAs are not recognized by nucleases or proteases. A non-limiting example of a PNA is depicted below. [0374] Despite a radical structural change to the natural structure, PNAs are capable of sequence-specific binding in a helix form to DNA or RNA.
- PNAs Characteristics of PNAs include a high binding affinity to complementary DNA or RNA, a destabilizing effect caused by single-base mismatch, resistance to nucleases and proteases, hybridization with DNA or RNA independent of salt concentration, and triplex formation with homopurine DNA.
- PANAGENETM has developed its proprietary Bts PNA monomers (Bts; benzothiazole-2-sulfonyl group) and proprietary oligomerization process.
- Bts benzothiazole-2-sulfonyl group
- the PNA oligomerization using Bts PNA monomers is composed of repetitive cycles of deprotection, coupling, and capping.
- PNAs can be produced synthetically using any technique known in the art. See, e.g., U.S. Pat.
- LNAs Locked Nucleic Acids
- Antisense oligomers may also contain "locked nucleic acid” subunits (LNAs).
- LNAs are a member of a class of modifications called bridged nucleic acid (BNA).
- BNA is characterized by a covalent linkage that locks the conformation of the ribose ring in a C30-endo (northern) sugar pucker.
- the bridge is composed of a methylene between the 2′-O and the 4′-C positions. LNA enhances backbone preorganization and base stacking to increase hybridization and thermal stability.
- the structures of LNAs can be found, for example, in Wengel, et al., Chemical Communications (1998) 455; Koshkin et al., Tetrahedron (1998) 54:3607; Jesper Wengel, Accounts of Chem.
- Antisense oligomers of the disclosure may incorporate one or more LNAs; in some cases, the antisense oligomers may be entirely composed of LNAs. Methods for the synthesis of individual LNA nucleoside subunits and their incorporation into oligomers are described, for example, in U.S.
- intersubunit linkers include phosphodiester and phosphorothioate moieties; alternatively, non- phosphorous-containing linkers may be employed.
- Further embodiments include an LNA containing antisense oligomer where each LNA subunit is separated by a DNA subunit. Certain antisense oligomers are composed of alternating LNA and DNA subunits where the intersubunit linker is phosphorothioate. 3.
- Ethylene-Bridged Nucleic Acids [0378] 2′-O,4′C-ethylene-bridged nucleic acids (ENAs) are another member of the class of BNAs. A non-limiting example is depicted below. [0379] ENA oligomers and their preparation are described in Obika et al., Tetrahedron Lett (1997) 38 (50): 8735, which is hereby incorporated by reference in its entirety. Antisense oligomers of the disclosure may incorporate one or more ENA subunits. 4. Unlocked nucleic acid (UNA) [0380] Antisense oligomers may also contain unlocked nucleic acid (UNA) subunits.
- UNAs and UNA oligomers are an analog of RNA in which the C2′-C3′ bond of the subunit has been cleaved. Whereas LNA is conformationally restricted (relative to DNA and RNA), UNA is very flexible. UNAs are disclosed, for example, in WO 2016/070166. A non-limiting example of an UNA is depicted below. [0381] Typical intersubunit linkers include phosphodiester and phosphorothioate moieties; alternatively, non-phosphorous containing linkers may be employed. 5. Phosphorothioates [0382] "Phosphorothioates" (or S-oligos) are a variant of normal DNA in which one of the nonbridging oxygens is replaced by sulfur.
- a non-limiting example of a phosphorothioate is depicted below.
- the sulfurization of the internucleotide bond reduces the action of endo-and exonucleases including 5′ to 3′ and 3′ to 5′ DNA POL 1 exonuclease, nucleases S1 and P1, RNases, serum nucleases, and snake venom phosphodiesterase.
- Phosphorothioates are made by two principal routes: by the action of a solution of elemental sulfur in carbon disulfide on a hydrogen phosphonate, or by the method of sulfurizing phosphite triesters with either tetraethylthiuram disulfide (TETD) or 3H-1, 2-benzodithiol-3-one 1, 1-dioxide (BDTD) (see, e.g., Iyer et al., J. Org. Chem.55, 4693-4699, 1990, which is hereby incorporated by reference in its entirety).
- TETD tetraethylthiuram disulfide
- BDTD 2-benzodithiol-3-one 1, 1-dioxide
- Tricyclo-DNAs are a class of constrained DNA analogs in which each nucleotide is modified by the introduction of a cyclopropane ring to restrict conformational flexibility of the backbone and to optimize the backbone geometry of the torsion angle ⁇ . Homobasic adenine- and thymine-containing tc-DNAs form extraordinarily stable A-T base pairs with complementary RNAs. Tricyclo-DNAs and their synthesis are described in International Patent Application Publication No.
- Antisense oligomers of the disclosure may incorporate one or more tricycle-DNA subunits; in some cases, the antisense oligomers may be entirely composed of tricycle-DNA subunits.
- Tricyclo-phosphorothioate subunits are tricyclo-DNA subunits with phosphorothioate intersubunit linkages. Tricyclo-phosphorothioate subunits and their synthesis are described in International Patent Application Publication No. WO 2013/053928, which is hereby incorporated by reference in its entirety.
- Antisense oligomers of the disclosure may incorporate one or more tricycle-DNA subunits; in some cases, the antisense oligomers may be entirely composed of tricycle-DNA subunits.
- a non-limiting example of a tricycle-DNA/tricycle- phosphorothioate subunit is depicted below. 7.
- 2′-O-Methyl, 2′-O-MOE, and 2′-F Oligomers [0386] "2′-O-Me oligomer" molecules carry a methyl group at the 2′-OH residue of the ribose molecule.2′-O-Me-RNAs show the same (or similar) behavior as DNA but are protected against nuclease degradation.2′-O-Me-RNAs can also be combined with phosphorothioate oligomers (PTOs) for further stabilization.2′O-Me oligomers (phosphodiester or phosphorothioate) can be synthesized according to routine techniques in the art (see, e.g., Yoo et al., Nucleic Acids Res.
- 2′-O-Me oligomers carry a methoxyethyl group at the 2′-OH residue of the ribose molecule and are discussed in Martin et al., Helv. Chim. Acta, 78, 486-504, 1995, which is hereby incorporated by reference in its entirety.
- 2′-O- MOE subunit is depicted below.
- 2′-Fluoro (2′-F) oligomers have a fluoro radical in at the 2′ position in place of the 2′-OH.
- a non-limiting example of a 2′-F oligomer is depicted below.
- 2′-fluoro oligomers are further described in WO 2004/043977, which is hereby incorporated by reference in its entirety.
- 2′-O-Methyl, 2′-O-MOE, and 2′-F oligomers may also comprise one or more phosphorothioate (PS) linkages as depicted below.
- PS phosphorothioate
- 2′-O-Methyl PS 2′-O-MOE PS 2′-F PS may comprise PS intersubunit linkages throughout the oligomer, for example, as in the 2′-O-methyl PS oligomer drisapersen depicted below.
- 2′-O-Methyl, 2′-O-MOE, and/or 2′-F oligomers may comprise PS linkages at the ends of the oligomer, as depicted below:
- Antisense oligomers of the disclosure may incorporate one or more 2′-O-methyl, 2′-O- MOE, and 2′-F subunits and may utilize any of the intersubunit linkages described here. In some instances, an antisense oligomer of the disclosure may be composed of entirely 2′-O-methyl, 2′- O-MOE, or 2′-F subunits.
- MCEs are another example of 2′-O modified ribonucleosides useful in the antisense oligomers of the disclosure.
- the 2′-OH is derivatized to a 2-(N-methylcarbamoyl)ethyl moiety to increase nuclease resistance.
- a non-limiting example of an MCE oligomer is depicted below.
- Stereo-specific oligomers are those in which the stereo chemistry of each phosphorous- containing linkage is fixed by the method of synthesis such that a substantially stereo-pure oligomer is produced. A non-limiting example of a stereo-specific oligomer is depicted below. [0397] In the above example, each phosphorous of the oligomer has the same stereo configuration.
- LNAs, ENAs, Tricyclo-DNAs, MCEs, 2′-O-Methyl, 2′-O-MOE, 2′-F, and morpholino-based oligomers can be prepared with stereo-specific phosphorous-containing internucleoside linkages such as, for example, phosphorothioate, phosphodiester, phosphoramidate, phosphorodiamidate, or other phosphorous-containing internucleoside linkages.
- Stereo specific oligomers, methods of preparation, chiral controlled synthesis, chiral design, and chiral auxiliaries for use in the preparation of such oligomers are detailed, for example, in WO2017192664, WO2017192679, WO2017062862, WO2017015575, WO2017015555, WO2015107425, WO2015108048, WO2015108046, WO2015108047, WO2012039448, WO2010064146, WO2011034072, WO2014010250, WO2014012081, WO20130127858, and WO2011005761, each of which is hereby incorporated by reference in its entirety.
- Stereo-specific oligomers can have phosphorous-containing internucleoside linkages in an R P or S P configuration. Chiral phosphorous-containing linkages in which the stereo configuration of the linkages is controlled is referred to as "stereopure,” while chiral phosphorous-containing linkages in which the stereo configuration of the linkages is uncontrolled is referred to as "stereorandom.”
- the oligomers of the disclosure comprise a plurality of stereopure and stereorandom linkages, such that the resulting oligomer has stereopure subunits at pre-specified positions of the oligomer.
- stereopure subunits An example of the location of the stereopure subunits is provided in international patent application publication number WO 2017/062862 A2 in Figures 7A and 7B.
- all the chiral phosphorous-containing linkages in an oligomer are stereorandom.
- all the chiral phosphorous-containing linkages in an oligomer are stereopure.
- all n of the chiral phosphorous-containing linkages in the oligomer are stereorandom.
- an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), all n of the chiral phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 10% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 20% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure.
- an oligomer with n chiral phosphorous- containing linkages (where n is an integer of 1 or greater), at least 30% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 40% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure.
- an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 50% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 60% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure.
- an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 70% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 80% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure.
- an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 90% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure.
- the oligomer contains at least 2 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either S P or R P ).
- the oligomer contains at least 3 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either S P or R P ). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 4 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either S P or R P ).
- the oligomer contains at least 5 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either S P or R P ). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 6 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP).
- the oligomer contains at least 7 contiguous stereopure phosphorous- containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 8 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP).
- the oligomer contains at least 9 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 10 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP).
- the oligomer contains at least 11 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either S P or R P ). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 12 contiguous stereopure phosphorous- containing linkages of the same stereo orientation (i.e., either S P or R P ).
- the oligomer contains at least 13 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either S P or R P ). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 14 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either S P or R P ).
- the oligomer contains at least 15 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either S P or R P ). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 16 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either S P or R P ).
- the oligomer contains at least 17 contiguous stereopure phosphorous- containing linkages of the same stereo orientation (i.e., either S P or R P ). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 18 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either S P or R P ).
- the oligomer contains at least 19 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 20 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP).
- the oligomer contains at least 2 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP) and at least 2 contiguous stereopure phosphorous-containing linkages of the other stereo orientation.
- the oligomer can contain at least 2 contiguous stereopure phosphorous-containing linkages of the SP orientation and at least 2 contiguous stereopure phosphorous-containing linkages of the R P orientation.
- the oligomer contains at least 2 contiguous stereopure phosphorous-containing linkages of the same stereo orientation in an alternating pattern.
- the oligomer can contain the following in order: 2 or more R P, 2 or more S P , and 2 or more R P , etc. 10.
- Morpholino Oligomers [0403] Exemplary embodiments of the disclosure relate to phosphorodiamidate morpholino oligomers of the following general structure: and as described in Figure 2 of Summerton, J., et al., Antisense & Nucleic Acid Drug Development, 7: 187-195 (1997).
- Morpholinos as described herein are intended to cover all stereoisomers and tautomers of the foregoing general structure.
- the synthesis, structures, and binding characteristics of morpholino oligomers are detailed in U.S. Patent Nos.: 5,698,685; 5,217,866; 5,142,047; 5,034,506; 5,166,315; 5,521,063; 5,506,337; 8,076,476; and 8,299,206, all of which are incorporated herein by reference.
- a morpholino is conjugated at the 5′ or 3′ end of the oligomer with a "tail" moiety to increase its stability and/or solubility.
- Exemplary tails include: [0405]
- the disclosure provides antisense oligomers according to Formulae (I), (IA), (II), (III), (IV), and/or (V), and pharmaceutically acceptable salts thereof.
- the antisense oligomers disclosed herein including antisense oligomers of Formulae (I), (IA), (II), (III), (IV), and/or (V), comprise a targeting sequence complementary to a target region within a pre-mRNA of the human DNM2 gene represented by SEQ ID NO: 1.
- the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 6 (SEQ ID NO: 3), exon 7 (SEQ ID NO: 4), exon 8 (SEQ ID NO: 5), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), exon 13 (SEQ ID NO: 8), exon 16 (SEQ ID NO: 9), and exon 19 (SEQ ID NO: 10).
- the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), and exon 13 (SEQ ID NO: 8).
- the target region is within an intron-exon junction or an exon-intron junction region of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 5/exon 6 junction, exon 6/intron 6 junction, intron 6/exon 7 junction, exon 7/intron 7 junction, intron 7/exon 8 junction, exon 8/intron 8 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, exon 13/intron 13 junction, intron 15/exon 16 junction, exon 16/intron 16 junction, intron 18/exon 19 junction, and exon 19/intron 19 junction
- the target region is within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, and exon 13/intron 13 junction.
- the target region is within an intron internal region of the human DNM2 gene pre-mRNA selected from intron 2 (SEQ ID NO: 11) and intron 12 (SEQ ID NO: 12).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12- 13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2 H2A (-10+15),
- the target region is DNM2 H2A (-10+15). In certain embodiments, the target region is DNM2 H2A(-7+17). In certain embodiments, the target region is DNM2 H2A(+3+27). In certain embodiments, the target region is DNM2 H2A(+15+37). In certain embodiments, the target region is DNM2 H2A(+21+45). In certain embodiments, the target region is DNM2 H2A(+23+47). In certain embodiments, the target region is DNM2 H2A(+27+51). In certain embodiments, the target region is DNM2 H2A(+37+61). In certain embodiments, the target region is DNM2 H2A(+46+70).
- the target region is DNM2 H2A (+48+72). In certain embodiments, the target region is DNM2 H2D(+20-5). In certain embodiments, the target region is DNM2 H2D (+13-12). In certain embodiments, the target region is DNM2 H2D(+12-13). In certain embodiments, the target region is DNM2 H2D(+11-14). In certain embodiments, the target region is DNM2 H2D(+10-15). In certain embodiments, the target region is DNM2 H2D(+9-16). In certain embodiments, the target region is DNM2 H2D(+8-17). In certain embodiments, the target region is DNM2 H2D(+7-18).
- the target region is DNM2 H2D(+6-19). In certain embodiments, the target region is DNM2 H2D(+5-20). In certain embodiments, the target region is DNM2 H2D(+4-21). In certain embodiments, the target region is DNM2 H2D(+3-22). In certain embodiments, the target region is DNM2 H2D(+2-23) . In certain embodiments, the target region is DNM2 H2D(+1-24). In certain embodiments, the target region is DNM2 H2D(-1-25). In certain embodiments, the target region is DNM2 H2D(-2-26). In certain embodiments, the target region is DNM2 H2D(-3-27).
- the target region is DNM2 H2D(-4-28). In certain embodiments, the target region is DNM2 H2D(-5-29). In certain embodiments, the target region is DNM2 H2D(-6-30). In certain embodiments, the target region is DNM2 H6A(-20+5). In certain embodiments, the target region is DNM2 H6A(-6+19). In certain embodiments, the target region is DNM2 H6A(-2+23). In certain embodiments, the target region is DNM2 H6A(+3+26). In certain embodiments, the target region is DNM2 H6A(+10+3). In certain embodiments, the target region is DNM2 H6A(+13+37).
- the target region is DNM2 H6A(+23+47). In certain embodiments, the target region is DNM2 H6A(+32+56). In certain embodiments, the target region is DNM2 H6A(+35+59). In certain embodiments, the target region is DNM2 H6A(+44+68). In certain embodiments, the target region is DNM2 H6A(+54+78). In certain embodiments, the target region is DNM2 H6A(+60+84). In certain embodiments, the target region is DNM2 H6A(+63+87). In certain embodiments, the target region is DNM2 H6A(+88+112). In certain embodiments, the target region is DNM2 H6A(+96+119).
- the target region is DNM2 H6A(+105+129). In certain embodiments, the target region is DNM2 H6A(+121+145). In certain embodiments, the target region is DNM2 H6A(+130+154). In certain embodiments, the target region is DNM2 H6A(+132+156). In certain embodiments, the target region is DNM2 H6D (+6-19). In certain embodiments, the target region is DNM2 H6D (+15-10). In certain embodiments, the target region is DNM2 H6D (+17-8). In certain embodiments, the target region is DNM2 H7A(-15+10). In certain embodiments, the target region is DNM2 H7A(- 5+20).
- the target region is DNM2 H7A(+1+25). In certain embodiments, the target region is DNM2 H7A(+6+30). In certain embodiments, the target region is. In certain embodiments, the target region is DNM2 H7A(+22+46). In certain embodiments, the target region is DNM2 H7A(+29+53). In certain embodiments, the target region is DNM2 H7A(+41+65). In certain embodiments, the target region is DNM2 H7A(+51+75). In certain embodiments, the target region is DNM2 H7A(+57+81). In certain embodiments, the target region is DNM2 H7A(+62+86).
- the target region is DNM2 H7A(+72+96). In certain embodiments, the target region is DNM2 H7A(+74+98). In certain embodiments, the target region is DNM2 H7A(+81+105). In certain embodiments, the target region is DNM2 H7A(+91+115). In certain embodiments, the target region is DNM2 H7A(+110+134). In certain embodiments, the target region is DNM2 H7D(+8-16). In certain embodiments, the target region is DNM2 H7D(+9-16). In certain embodiments, the target region is DNM2 H7D(+21-4). In certain embodiments, the target region is DNM2 H8A(-20+5).
- the target region is DNM2 H8A(-14+11). In certain embodiments, the target region is DNM2 H8A(-7+18). In certain embodiments, the target region is DNM2 H8A(-6+19). In certain embodiments, the target region is DNM2 H8A(-2+23). In certain embodiments, the target region is DNM2 H8A(+5+29). In certain embodiments, the target region is DNM2 H8A(+18+42). In certain embodiments, the target region is DNM2 H8A(+21+45). In certain embodiments, the target region is DNM2 H8A(+27+51). In certain embodiments, the target region is DNM2 H8A(+36+60).
- the target region is DNM2 H8A(+46+70). In certain embodiments, the target region is DNM2 H8A(+51+75). In certain embodiments, the target region is DNM2 H8A(+56+80). In certain embodiments, the target region is DNM2 H8A(+65+88). In certain embodiments, the target region is DNM2 H8A(+76+100). In certain embodiments, the target region is DNM2 H8A(+83+107). In certain embodiments, the target region is DNM2 H8A(+96+119). In certain embodiments, the target region is DNM2 H8A(+105+129). In certain embodiments, the target region is DNM2 H8D(+18-7).
- the target region is DNM2 H8D(+10-15). In certain embodiments, the target region is DNM2 H9A(-12+13). In certain embodiments, the target region is DNM2 H9A(-5+20). In certain embodiments, the target region is DNM2 H9A(+1+25). In certain embodiments, the target region is DNM2 H9A(+6+30). In certain embodiments, the target region is DNM2 H9A(+16+40). In certain embodiments, the target region is DNM2 H9A(+25+49). In certain embodiments, the target region is DNM2 H9A(+36+60). In certain embodiments, the target region is DNM2 H9A(+37+61).
- the target region is DNM2 H9A(+42+66). In certain embodiments, the target region is DNM2 H9A(+43+62). In certain embodiments, the target region is DNM2 H9A(+43+63). In certain embodiments, the target region is DNM2 H9A(+43+64). In certain embodiments, the target region is DNM2 H9A(+43+65). In certain embodiments, the target region is DNM2 H9A(+43+66). In certain embodiments, the target region is DNM2 H9A(+43+67). In certain embodiments, the target region is DNM2 H9A(+44+67). In certain embodiments, the target region is DNM2 H9A(+44+68).
- the target region is DNM2 H9A(+45+67). In certain embodiments, the target region is DNM2 H9A (+46+67). In certain embodiments, the target region is DNM2 H9A(+47+67). In certain embodiments, the target region is DNM2 H9A(+48+67). In certain embodiments, the target region is DNM2 H9D(+24-1). In certain embodiments, the target region is DNM2 H9D(+23-2). In certain embodiments, the target region is DNM2 H9D(+22-3). In certain embodiments, the target region is DNM2 H9D(+21-4). In certain embodiments, the target region is DNM2 H9D(+20-5).
- the target region is DNM2 H9D(+19-6). In certain embodiments, the target region is DNM2 H9D (+18-7). In certain embodiments, the target region is DNM2 H9D(+17-8). In certain embodiments, the target region is DNM2 H9D(+16-9). In certain embodiments, the target region is DNM2 H9D(+15-10). In certain embodiments, the target region is DNM2 H9D(+14-11). In certain embodiments, the target region is DNM2 H9D(+13-12). In certain embodiments, the target region is DNM2 H9D(+12-13). In certain embodiments, the target region is DNM2 H9D(+11-14).
- the target region is DNM2 H9D(+10-15). In certain embodiments, the target region is DNM2 H9D(+9-16). In certain embodiments, the target region is DNM2 H9D(+8-12). In certain embodiments, the target region is DNM2 H9D(+8-13). In certain embodiments, the target region is DNM2 H9D(+8-14). In certain embodiments, the target region is DNM2 H9D(+8-15). In certain embodiments, the target region is DNM2 H9D(+8-17). In certain embodiments, the target region is DNM2 H9D(+7-18). In certain embodiments, the target region is DNM2 H9D(+7-17).
- the target region is DNM2 H9D(+6-19). In certain embodiments, the target region is DNM2 H9D(+6-17). In certain embodiments, the target region is DNM2 H9D(+5-17). In certain embodiments, the target region is DNM2 H9D(+5-20). In certain embodiments, the target region is DNM2 H9D(+4-17). In certain embodiments, the target region is DNM2 H9D(+4-21). In certain embodiments, the target region is DNM2 H9D(+3-17). In certain embodiments, the target region is DNM2 H9D(+3-22). In certain embodiments, the target region is DNM2 H12A(-13+12).
- the target region is DNM2 H12A(-3+22). In certain embodiments, the target region is DNM2 H12A(+4+28). In certain embodiments, the target region is DNM2 H12A(+7+31). In certain embodiments, the target region is DNM2 H12A(+10+34). In certain embodiments, the target region is DNM2 H12A(+11+35). In certain embodiments, the target region is DNM2 H12A(+12+36). In certain embodiments, the target region is DNM2 H12A(+13+37). In certain embodiments, the target region is DNM2 H12A(+14+38). In certain embodiments, the target region is DNM2 H12A(+15+39).
- the target region is DNM2 H12A(+16+40). In certain embodiments, the target region is DNM2 H12A(+17+41). In certain embodiments, the target region is DNM2 H12A(+18+42). In certain embodiments, the target region is DNM2 H12A(+19+43). In certain embodiments, the target region is DNM2 H12A(+20+44). In certain embodiments, the target region is DNM2 H12A(+25+44). In certain embodiments, the target region is DNM2 H12A(+25+45). In certain embodiments, the target region is DNM2 H12A(+25+46). In certain embodiments, the target region is DNM2 H12A(+25+47).
- the target region is DNM2 H12A(+25+48). In certain embodiments, the target region is DNM2 H12A(+25+49). In certain embodiments, the target region is DNM2 H12A(+26+49). In certain embodiments, the target region is DNM2 H12A(+27+49). In certain embodiments, the target region is DNM2 H12A(+28+49). In certain embodiments, the target region is DNM2 H12A(+29+49). In certain embodiments, the target region is DNM2 H12A(+29+53). In certain embodiments, the target region is DNM2 H12A(+30+49). In certain embodiments, the target region is DNM2 H12A(+30+54).
- the target region is DNM2 H12A(+36+60). In certain embodiments, the target region is DNM2 H12A(+37+61). In certain embodiments, the target region is DNM2 H12A(+43+67). In certain embodiments, the target region is DNM2 H12D(+22- 3). In certain embodiments, the target region is DNM2 H12D(+20-5). In certain embodiments, the target region is DNM2 H12D(+15-10). In certain embodiments, the target region is DNM2 H13A(-27-3). In certain embodiments, the target region is DNM2 H13A(-26-2). In certain embodiments, the target region is DNM2 H13A(-25-1).
- the target region is DNM2 H13A(-24+1). In certain embodiments, the target region is DNM2 H13A(-23+2). In certain embodiments, the target region is DNM2 H13A(-22+3). In certain embodiments, the target region is DNM2 H13A(-21+4). In certain embodiments, the target region is DNM2 H13A(- 20+5). In certain embodiments, the target region is DNM2 H13A(-19+6). In certain embodiments, the target region is DNM2 H13A(-18+7). In certain embodiments, the target region is DNM2 H13A(-17+8). In certain embodiments, the target region is DNM2 H13A(-16+9).
- the target region is DNM2 H13A(-15+10). In certain embodiments, the target region is DNM2 H13A(-14+11). In certain embodiments, the target region is DNM2 H13A(- 13+12). In certain embodiments, the target region is DNM2 H13A(-11+14). In certain embodiments, the target region is DNM2 H13A(-10+15). In certain embodiments, the target region is DNM2 H13A(-9+16). In certain embodiments, the target region is DNM2 H13A(-8+17). In certain embodiments, the target region is DNM2 H13A(-7+18). In certain embodiments, the target region is DNM2 H13A(-6+19).
- the target region is DNM2 H13A(- 5+20). In certain embodiments, the target region is DNM2 H13A(-4+21). In certain embodiments, the target region is DNM2 H13A(-3+22). In certain embodiments, the target region is DNM2 H13A(-2+23). In certain embodiments, the target region is DNM2 H13A(-1+24). In certain embodiments, the target region is DNM2 H13A(+1+25). In certain embodiments, the target region is DNM2 H13A(+2+25). In certain embodiments, the target region is DNM2 H13A(+3+27). In certain embodiments, the target region is DNM2 H13A(+4+23).
- the target region is DNM2 H13A(+4+24). In certain embodiments, the target region is DNM2 H13A(+4+25). In certain embodiments, the target region is DNM2 H13A(+4+26). In certain embodiments, the target region is DNM2 H13A(+4+27). In certain embodiments, the target region is DNM2 H13A(+4+28). In certain embodiments, the target region is DNM2 H13A(+5+28). In certain embodiments, the target region is DNM2 H13A(+6+28). In certain embodiments, the target region is DNM2 H13A(+6+30). In certain embodiments, the target region is DNM2 H13A(+7+28).
- the target region is DNM2 H13A(+8+28). In certain embodiments, the target region is DNM2 H13A(+9+28). In certain embodiments, the target region is DNM2 H13A(+16+40). In certain embodiments, the target region is DNM2 H13D(+22-3). In certain embodiments, the target region is DNM2 H13D(+7-18). In certain embodiments, the target region is DNM2 H16A(-21+3). In certain embodiments, the target region is DNM2 H16A(-13+12). In certain embodiments, the target region is DNM2 H16A(- 6+19). In certain embodiments, the target region is DNM2 H16A(+6+30).
- the target region is DNM2 H16A(+15+39). In certain embodiments, the target region is DNM2 H16A(+26+50). In certain embodiments, the target region is DNM2 H16A(+36+60). In certain embodiments, the target region is DNM2 H16A(+45+69). In certain embodiments, the target region is DNM2 H16A(+55+79). In certain embodiments, the target region is DNM2 H16A(+66+90). In certain embodiments, the target region is DNM2 H16A(+75+99). In certain embodiments, the target region is DNM2 H16A(+82+106). In certain embodiments, the target region is DNM2 H16D(+19-6).
- the target region is DNM2 H16D(+11-14). In certain embodiments, the target region is DNM2 H19A(- 11+14). In certain embodiments, the target region is DNM2 H19A(-4+21). In certain embodiments, the target region is DNM2 H19A(+1+25). In certain embodiments, the target region is DNM2 H19A(+7+31). In certain embodiments, the target region is DNM2 H19A(+16+40). In certain embodiments, the target region is DNM2 H19A(+26+50). In certain embodiments, the target region is DNM2 H19A(+37+61). In certain embodiments, the target region is DNM2 H19A(+47+71).
- the target region is DNM2 H19A(+56+80). In certain embodiments, the target region is DNM2 H19A(+66+90). In certain embodiments, the target region is DNM2 H19A(+76+100). In certain embodiments, the target region is DNM2 H19A(+86+110). In certain embodiments, the target region is DNM2 H19A(+96+120). In certain embodiments, the target region is DNM2 H19A(+106+130). In certain embodiments, the target region is DNM2 H19A(+117+141). In certain embodiments, the target region is DNM2 H19A(+127+151). In certain embodiments, the target region is DNM2 H19A(+137+161).
- the target region is DNM2 H19A(+147+171). In certain embodiments, the target region is DNM2 H19A(+157+181). In certain embodiments, the target region is DNM2 H19A(+166+189). In certain embodiments, the target region is DNM2 H19A(+191+215). In certain embodiments, the target region is DNM2 H19A(+200+224). In certain embodiments, the target region is DNM2 H19D(+23-2). In certain embodiments, the target region is DNM2 H19D(+17-8). In certain embodiments, the target region is DNM2 H19D(+10-15).
- the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGGCCGC); SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC); SEQ ID NO: 77 (TTTACCTGTTT
- the targeting sequence is SEQ ID NO: 67. In certain embodiments, the targeting sequence is SEQ ID NO: 68. In certain embodiments, the targeting sequence is SEQ ID NO: 69. In certain embodiments, the targeting sequence is SEQ ID NO: 70. [0416] In certain embodiments, the targeting sequence is SEQ ID NO: 71. In certain embodiments, the targeting sequence is SEQ ID NO: 72. In certain embodiments, the targeting sequence is SEQ ID NO: 73. In certain embodiments, the targeting sequence is SEQ ID NO: 74. In certain embodiments, the targeting sequence is SEQ ID NO: 75. In certain embodiments, the targeting sequence is SEQ ID NO: 76. In certain embodiments, the targeting sequence is SEQ ID NO: 77.
- the targeting sequence is SEQ ID NO: 78. In certain embodiments, the targeting sequence is SEQ ID NO: 79. In certain embodiments, the targeting sequence is SEQ ID NO: 80. In certain embodiments, the targeting sequence is SEQ ID NO: 81. In certain embodiments, the targeting sequence is SEQ ID NO: 82. In certain embodiments, the targeting sequence is SEQ ID NO: 83. In certain embodiments, the targeting sequence is SEQ ID NO: 84. In certain embodiments, the targeting sequence is SEQ ID NO: 85. In certain embodiments, the targeting sequence is SEQ ID NO: 86. In certain embodiments, the targeting sequence is SEQ ID NO: 87. In certain embodiments, the targeting sequence is SEQ ID NO: 88.
- the targeting sequence is SEQ ID NO: 89. In certain embodiments, the targeting sequence is SEQ ID NO: 90. In certain embodiments, the targeting sequence is SEQ ID NO: 91. In certain embodiments, the targeting sequence is SEQ ID NO: 92. In certain embodiments, the targeting sequence is SEQ ID NO: 93. In certain embodiments, the targeting sequence is SEQ ID NO: 94. In certain embodiments, the targeting sequence is SEQ ID NO: 95. In certain embodiments, the targeting sequence is SEQ ID NO: 96. In certain embodiments, the targeting sequence is SEQ ID NO: 97. In certain embodiments, the targeting sequence is SEQ ID NO: 98. In certain embodiments, the targeting sequence is SEQ ID NO: 99.
- the targeting sequence is SEQ ID NO: 100. In certain embodiments, the targeting sequence is SEQ ID NO: 101. In certain embodiments, the targeting sequence is SEQ ID NO: 102. In certain embodiments, the targeting sequence is SEQ ID NO: 103. In certain embodiments, the targeting sequence is SEQ ID NO: 104. In certain embodiments, the targeting sequence is SEQ ID NO: 105. In certain embodiments, the targeting sequence is SEQ ID NO: 106. In certain embodiments, the targeting sequence is SEQ ID NO: 107. In certain embodiments, the targeting sequence is SEQ ID NO: 108. In certain embodiments, the targeting sequence is SEQ ID NO: 109. In certain embodiments, the targeting sequence is SEQ ID NO: 110.
- the targeting sequence is SEQ ID NO: 111. In certain embodiments, the targeting sequence is SEQ ID NO: 112. In certain embodiments, the targeting sequence is SEQ ID NO: 113. In certain embodiments, the targeting sequence is SEQ ID NO: 114. In certain embodiments, the targeting sequence is SEQ ID NO: 115. In certain embodiments, the targeting sequence is SEQ ID NO: 116. In certain embodiments, the targeting sequence is SEQ ID NO: 117. In certain embodiments, the targeting sequence is SEQ ID NO: 118. In certain embodiments, the targeting sequence is SEQ ID NO: 119. In certain embodiments, the targeting sequence is SEQ ID NO: 120. In certain embodiments, the targeting sequence is SEQ ID NO: 121.
- the targeting sequence is SEQ ID NO: 122. In certain embodiments, the targeting sequence is SEQ ID NO: 123. In certain embodiments, the targeting sequence is SEQ ID NO: 124. In certain embodiments, the targeting sequence is SEQ ID NO: 125. In certain embodiments, the targeting sequence is SEQ ID NO: 126. In certain embodiments, the targeting sequence is SEQ ID NO: 127. In certain embodiments, the targeting sequence is SEQ ID NO: 128. In certain embodiments, the targeting sequence is SEQ ID NO: 129. In certain embodiments, the targeting sequence is SEQ ID NO: 130. In certain embodiments, the targeting sequence is SEQ ID NO: 131. In certain embodiments, the targeting sequence is SEQ ID NO: 132.
- the targeting sequence is SEQ ID NO: 133. In certain embodiments, the targeting sequence is SEQ ID NO: 134. In certain embodiments, the targeting sequence is SEQ ID NO: 135. In certain embodiments, the targeting sequence is SEQ ID NO: 136. In certain embodiments, the targeting sequence is SEQ ID NO: 137. In certain embodiments, the targeting sequence is SEQ ID NO: 138. In certain embodiments, the targeting sequence is SEQ ID NO: 139. In certain embodiments, the targeting sequence is SEQ ID NO: 140. In certain embodiments, the targeting sequence is SEQ ID NO: 141. In certain embodiments, the targeting sequence is SEQ ID NO: 142. In certain embodiments, the targeting sequence is SEQ ID NO: 143.
- the targeting sequence is SEQ ID NO: 144. In certain embodiments, the targeting sequence is SEQ ID NO: 145. In certain embodiments, the targeting sequence is SEQ ID NO: 146. In certain embodiments, the targeting sequence is SEQ ID NO: 147. In certain embodiments, the targeting sequence is SEQ ID NO: 148. In certain embodiments, the targeting sequence is SEQ ID NO: 149. In certain embodiments, the targeting sequence is SEQ ID NO: 150. In certain embodiments, the targeting sequence is SEQ ID NO: 151. In certain embodiments, the targeting sequence is SEQ ID NO: 152. In certain embodiments, the targeting sequence is SEQ ID NO: 153. In certain embodiments, the targeting sequence is SEQ ID NO: 154.
- the targeting sequence is SEQ ID NO: 155. In certain embodiments, the targeting sequence is SEQ ID NO: 156. In certain embodiments, the targeting sequence is SEQ ID NO: 157. In certain embodiments, the targeting sequence is SEQ ID NO: 158. In certain embodiments, the targeting sequence is SEQ ID NO: 159. In certain embodiments, the targeting sequence is SEQ ID NO: 160. In certain embodiments, the targeting sequence is SEQ ID NO: 161. In certain embodiments, the targeting sequence is SEQ ID NO: 162. In certain embodiments, the targeting sequence is SEQ ID NO: 163. In certain embodiments, the targeting sequence is SEQ ID NO: 164. In certain embodiments, the targeting sequence is SEQ ID NO: 165.
- the targeting sequence is SEQ ID NO: 166. In certain embodiments, the targeting sequence is SEQ ID NO: 167. In certain embodiments, the targeting sequence is SEQ ID NO: 168. In certain embodiments, the targeting sequence is SEQ ID NO: 169. In certain embodiments, the targeting sequence is SEQ ID NO: 170. In certain embodiments, the targeting sequence is SEQ ID NO: 171. In certain embodiments, the targeting sequence is SEQ ID NO: 172. In certain embodiments, the targeting sequence is SEQ ID NO: 173. In certain embodiments, the targeting sequence is SEQ ID NO: 174. In certain embodiments, the targeting sequence is SEQ ID NO: 175. In certain embodiments, the targeting sequence is SEQ ID NO: 176.
- the targeting sequence is SEQ ID NO: 177. In certain embodiments, the targeting sequence is SEQ ID NO: 178. In certain embodiments, the targeting sequence is SEQ ID NO: 179. In certain embodiments, the targeting sequence is SEQ ID NO: 180. In certain embodiments, the targeting sequence is SEQ ID NO: 181. In certain embodiments, the targeting sequence is SEQ ID NO: 182. In certain embodiments, the targeting sequence is SEQ ID NO: 183. In certain embodiments, the targeting sequence is SEQ ID NO: 184. In certain embodiments, the targeting sequence is SEQ ID NO: 185. In certain embodiments, the targeting sequence is SEQ ID NO: 186. In certain embodiments, the targeting sequence is SEQ ID NO: 187.
- the targeting sequence is SEQ ID NO: 188. In certain embodiments, the targeting sequence is SEQ ID NO: 189. In certain embodiments, the targeting sequence is SEQ ID NO: 190. In certain embodiments, the targeting sequence is SEQ ID NO: 191. In certain embodiments, the targeting sequence is SEQ ID NO: 192. In certain embodiments, the targeting sequence is SEQ ID NO: 193. In certain embodiments, the targeting sequence is SEQ ID NO: 194. In certain embodiments, the targeting sequence is SEQ ID NO: 195. In certain embodiments, the targeting sequence is SEQ ID NO: 196. In certain embodiments, the targeting sequence is SEQ ID NO: 197. In certain embodiments, the targeting sequence is SEQ ID NO: 198.
- the targeting sequence is SEQ ID NO: 199. In certain embodiments, the targeting sequence is SEQ ID NO: 200. In certain embodiments, the targeting sequence is SEQ ID NO: 201. In certain embodiments, the targeting sequence is SEQ ID NO: 202. In certain embodiments, the targeting sequence is SEQ ID NO: 203. In certain embodiments, the targeting sequence is SEQ ID NO: 204. In certain embodiments, the targeting sequence is SEQ ID NO: 205. In certain embodiments, the targeting sequence is SEQ ID NO: 206. In certain embodiments, the targeting sequence is SEQ ID NO: 207. In certain embodiments, the targeting sequence is SEQ ID NO: 208. In certain embodiments, the targeting sequence is SEQ ID NO: 209.
- the targeting sequence is SEQ ID NO: 210. In certain embodiments, the targeting sequence is SEQ ID NO: 211. In certain embodiments, the targeting sequence is SEQ ID NO: 212. In certain embodiments, the targeting sequence is SEQ ID NO: 213. In certain embodiments, the targeting sequence is SEQ ID NO: 214. In certain embodiments, the targeting sequence is SEQ ID NO: 215. In certain embodiments, the targeting sequence is SEQ ID NO: 216. In certain embodiments, the targeting sequence is SEQ ID NO: 217. In certain embodiments, the targeting sequence is SEQ ID NO: 218. In certain embodiments, the targeting sequence is SEQ ID NO: 219. In certain embodiments, the targeting sequence is SEQ ID NO: 220.
- the targeting sequence is SEQ ID NO: 221. In certain embodiments, the targeting sequence is SEQ ID NO: 222. In certain embodiments, the targeting sequence is SEQ ID NO: 223. In certain embodiments, the targeting sequence is SEQ ID NO: 224. In certain embodiments, the targeting sequence is SEQ ID NO: 225. In certain embodiments, the targeting sequence is SEQ ID NO: 226. In certain embodiments, the targeting sequence is SEQ ID NO: 227. In certain embodiments, the targeting sequence is SEQ ID NO: 228. In certain embodiments, the targeting sequence is SEQ ID NO: 229. In certain embodiments, the targeting sequence is SEQ ID NO: 230. In certain embodiments, the targeting sequence is SEQ ID NO: 231.
- the targeting sequence is SEQ ID NO: 232. In certain embodiments, the targeting sequence is SEQ ID NO: 233. In certain embodiments, the targeting sequence is SEQ ID NO: 234. In certain embodiments, the targeting sequence is SEQ ID NO: 235. In certain embodiments, the targeting sequence is SEQ ID NO: 236. In certain embodiments, the targeting sequence is SEQ ID NO: 237. In certain embodiments, the targeting sequence is SEQ ID NO: 238. In certain embodiments, the targeting sequence is SEQ ID NO: 239. In certain embodiments, the targeting sequence is SEQ ID NO: 240. In certain embodiments, the targeting sequence is SEQ ID NO: 241. In certain embodiments, the targeting sequence is SEQ ID NO: 242.
- the targeting sequence is SEQ ID NO: 243. In certain embodiments, the targeting sequence is SEQ ID NO: 244. In certain embodiments, the targeting sequence is SEQ ID NO: 245. In certain embodiments, the targeting sequence is SEQ ID NO: 246. In certain embodiments, the targeting sequence is SEQ ID NO: 247. In certain embodiments, the targeting sequence is SEQ ID NO: 248. In certain embodiments, the targeting sequence is SEQ ID NO: 249. In certain embodiments, the targeting sequence is SEQ ID NO: 250. In certain embodiments, the targeting sequence is SEQ ID NO: 251. In certain embodiments, the targeting sequence is SEQ ID NO: 252. In certain embodiments, the targeting sequence is SEQ ID NO: 253.
- the targeting sequence is SEQ ID NO: 254. In certain embodiments, the targeting sequence is SEQ ID NO: 255. In certain embodiments, the targeting sequence is SEQ ID NO: 256. In certain embodiments, the targeting sequence is SEQ ID NO: 257. In certain embodiments, the targeting sequence is SEQ ID NO: 258. In certain embodiments, the targeting sequence is SEQ ID NO: 259. In certain embodiments, the targeting sequence is SEQ ID NO: 260. In certain embodiments, the targeting sequence is SEQ ID NO: 261. In certain embodiments, the targeting sequence is SEQ ID NO: 262. In certain embodiments, the targeting sequence is SEQ ID NO: 263. In certain embodiments, the targeting sequence is SEQ ID NO: 264.
- the targeting sequence is SEQ ID NO: 276. In certain embodiments, the targeting sequence is SEQ ID NO: 277. In certain embodiments, the targeting sequence is SEQ ID NO: 278. In certain embodiments, the targeting sequence is SEQ ID NO: 279. In certain embodiments, the targeting sequence is SEQ ID NO: 280. In certain embodiments, the targeting sequence is SEQ ID NO: 281. In certain embodiments, the targeting sequence is SEQ ID NO: 282. In certain embodiments, the targeting sequence is SEQ ID NO: 283. In certain embodiments, the targeting sequence is SEQ ID NO: 284. In certain embodiments, the targeting sequence is SEQ ID NO: 285. In certain embodiments, the targeting sequence is SEQ ID NO: 286.
- the targeting sequence is SEQ ID NO: 287. In certain embodiments, the targeting sequence is SEQ ID NO: 288. In certain embodiments, the targeting sequence is SEQ ID NO: 289. In certain embodiments, the targeting sequence is SEQ ID NO: 290. In certain embodiments, the targeting sequence is SEQ ID NO: 291. In certain embodiments, the targeting sequence is SEQ ID NO: 292. In certain embodiments, the targeting sequence is SEQ ID NO: 293. In certain embodiments, the targeting sequence is SEQ ID NO: 294. In certain embodiments, the targeting sequence is SEQ ID NO: 295. In certain embodiments, the targeting sequence is SEQ ID NO: 296. In certain embodiments, the targeting sequence is SEQ ID NO: 297.
- the targeting sequence is SEQ ID NO: 298. In certain embodiments, the targeting sequence is SEQ ID NO: 299. In certain embodiments, the targeting sequence is SEQ ID NO: 300. In certain embodiments, the targeting sequence is SEQ ID NO: 301. In certain embodiments, the targeting sequence is SEQ ID NO: 302. In certain embodiments, the targeting sequence is SEQ ID NO: 303. In certain embodiments, the targeting sequence is SEQ ID NO: 304. In certain embodiments, the targeting sequence is SEQ ID NO: 305. In certain embodiments, the targeting sequence is SEQ ID NO: 306. In certain embodiments, the targeting sequence is SEQ ID NO: 307. In certain embodiments, the targeting sequence is SEQ ID NO: 308.
- the targeting sequence is SEQ ID NO: 309. In certain embodiments, the targeting sequence is SEQ ID NO: 310. In certain embodiments, the targeting sequence is SEQ ID NO: 311. In certain embodiments, the targeting sequence is SEQ ID NO: 312. In certain embodiments, the targeting sequence is SEQ ID NO: 313. In certain embodiments, the targeting sequence is SEQ ID NO: 314. In certain embodiments, the targeting sequence is SEQ ID NO: 315. In certain embodiments, the targeting sequence is SEQ ID NO: 316. In certain embodiments, the targeting sequence is SEQ ID NO: 317. In certain embodiments, the targeting sequence is SEQ ID NO: 318. In certain embodiments, the targeting sequence is SEQ ID NO: 319.
- the target region is within the exon internal region of exon 2 (SEQ ID NO: 2), the intron 1/exon 2 junction, the exon 2/intron 2 junction, or the intron internal region of intron 2 (SEQ ID NO: 11) of the human DNM2 gene pre-mRNA.
- the target region of the human DNM2 gene pre-mRNA is within the 21 st nucleotide to the 47 th nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2).
- the target region is a region within SEQ ID NO: 13.
- the target region is a region within DNM2 H2A(+21+47).
- the target region of the human DNM2 gene pre-mRNA is within the 23 rd nucleotide to the 51 st nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 14. In certain embodiments, the target region is a region within DNM2 H2A(+23+51). [0419] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 46 th nucleotide to the 72 nd nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2).
- the target region is a region within SEQ ID NO: 15. In certain embodiments, the target region is a region within DNM2 H2A(+46+72). [0420] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 13 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 24 th nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 16. In certain embodiments, the target region is a region within DNM2 H2D(+13-24).
- the target region of the human DNM2 gene pre-mRNA is within the 13 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 14 th nucleotide of intron 2 measured from the 3′ end of exon 2.
- the target region is a region within SEQ ID NO: 17.
- the target region is a region within DNM2 H2D(+13-14).
- the target region of the human DNM2 gene pre-mRNA is within the 10 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 19 th nucleotide of intron 2 measured from the 3′ end of exon 2.
- the target region is a region within SEQ ID NO: 18.
- the target region is a region within DNM2 H2D(+10-19).
- the target region of the human DNM2 gene pre-mRNA is within the 9 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 18 th nucleotide of intron 2 measured from the 3′ end of exon 2.
- the target region is a region within SEQ ID NO: 19.
- the target region is a region within DNM2 H2D(+9-18).
- the target region of the human DNM2 gene pre-mRNA is within the 6 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23 rd nucleotide of intron 2 measured from the 3′ end of exon 2.
- the target region is a region within SEQ ID NO: 20.
- the target region is a region within DNM2 H2D(+6-23).
- the target region of the human DNM2 gene pre-mRNA is within the 4 th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23 rd nucleotide of intron 2 measured from the 3′ end of exon 2.
- the target region is a region within SEQ ID NO: 21.
- the target region is a region within DNM2 H2D(+4-23).
- the target region of the human DNM2 gene pre-mRNA is within the 1 st nucleotide to the 30 th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 22. In certain embodiments, the target region is a region within DNM2 H2D(-1-30).
- the target region of the human DNM2 gene pre-mRNA is within the 1 st nucleotide to the 29 th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 23. In certain embodiments, the target region is a region within DNM2 H2D(-1-29).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12- 13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2 H2A (-10+15),
- the target region of the human DNM2 gene pre-mRNA is DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9- 16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23) , DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6- 30).
- the target region is DNM2 H2A(+21+45) or DNM2 H2A(+23+47). In certain embodiments, the target region is DNM2 H2A(+23+47) or DNM2 H2A(+27+51). In certain embodiments, the target region is DNM2 H2A(+46+70) or DNM2 H2A(+48+72).
- the target region is DNM2 H2D(+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), or DNM2 H2D(+1-24).
- the target region is DNM2 H2D (+13-12), DNM2 H2D(+12-13), or DNM2 H2D(+11-14). [0432] In certain embodiments, the target region is DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), or DNM2 H2D(+6-19). In certain embodiments, the target region is DNM2 H2D(+9-16), DNM2 H2D(+8-17), or DNM2 H2D(+7-18). In certain embodiments, the target region is DNM2 H2D(+8-17).
- the target region is DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23). In certain embodiments, the target region is DNM2 H2D(+4-21), DNM2 H2D(+3-22), or DNM2 H2D(+2-23). In certain embodiments, the target region is DNM2 H2D(+3-22).
- the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30). In certain embodiments, the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), or DNM2 H2D(-5-29). [0435] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(-7+17). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H2A(+3+27). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+15+37). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+37+61). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2D(+20-5).
- the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGGCCGC); SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC); SEQ ID NO: 77 (TTTACCTGTTT
- the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTC
- the targeting sequence is selected from: SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); and SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC). [0439] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); and SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA). [0440] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); and SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAA).
- the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTC
- the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); and SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG).
- the targeting sequence is selected from: SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); and SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT).
- the targeting sequence is selected from: SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); and SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT). [0445] In certain embodiments, the targeting sequence is SEQ ID NO: 83.
- the targeting sequence is selected from: SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); and SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT).
- the targeting sequence is selected from: SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); and SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT). [0448] In certain embodiments, the targeting sequence is SEQ ID NO: 88.
- the targeting sequence is selected from: SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT); and SEQ ID NO: 96 (CCTGCTGAACCTCAGGCCGCCCCAT).
- the targeting sequence is selected from: SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); and SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT).
- the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGAGGCCGC); and SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC).
- the target region is within the exon internal region of exon 9 (SEQ ID NO: 6), the intron 8/exon 9 junction, or the exon 9/intron 9 junction of the human DNM2 gene pre-mRNA.
- the target region of the human DNM2 gene pre-mRNA is within the 36 th nucleotide to the 61 st nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6).
- the target region is a region within SEQ ID NO: 24.
- the target region is a region within DNM2 H9A(+36+61).
- the target region of the human DNM2 gene pre-mRNA is within the 42 nd nucleotide to the 68 th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6). In certain embodiments, the target region is a region within SEQ ID NO: 25. In certain embodiments, the target region is a region within DNM2 H9A(+42+70). [0455] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 43 rd nucleotide to the 68 th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6).
- the target region is a region within SEQ ID NO: 26. In certain embodiments, the target region is a region within DNM2 H9A(+42+68). [0456] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 24 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15 th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 27. In certain embodiments, the target region is a region within DNM2 H9D(+24-15).
- the target region of the human DNM2 gene pre-mRNA is within the 24 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 5 th nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 28.
- the target region is a region within DNM2 H9D(+24-5).
- the target region of the human DNM2 gene pre-mRNA is within the 19 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 10 th nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 29.
- the target region is a region within DNM2 H9D(+19-10).
- the target region of the human DNM2 gene pre-mRNA is within the 14 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15 th nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 30.
- the target region is a region within DNM2 H9D(+14-15).
- the target region of the human DNM2 gene pre-mRNA is within the 9 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22 nd nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 31.
- the target region is a region within DNM2 H9D(+9-22).
- the target region of the human DNM2 gene pre-mRNA is within the 9 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 20 th nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 32.
- the target region is a region within DNM2 H9D(+9-20).
- the target region of the human DNM2 gene pre-mRNA is within the 9 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 18 th nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 33.
- the target region is a region within DNM2 H9D(+9-18).
- the target region of the human DNM2 gene pre-mRNA is within the 8 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 21 st nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 34.
- the target region is a region within DNM2 H9D(+8-21).
- the target region of the human DNM2 gene pre-mRNA is within the 6 th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22 nd nucleotide of intron 9 measured from the 3′ end of exon 9.
- the target region is a region within SEQ ID NO: 35.
- the target region is a region within DNM2 H9D(+6-22).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13), DNM2 H9A(-5+20), DNM2 H9A(+1+25),DNM2 H9A(+6+30), DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(N-N-N-(+
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+43+67), DNM2 H9A(+42+66), DNM2 H9A(+44+68), DNM2 H9D(+24-1), DNM2 H9D(+23- 2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+36+60) or DNM2 H9A(+37+61).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), or DNM2 H9A(+48+67).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), or DNM2 H9A(+44+68).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+67).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11- 14), or DNM2 H9D(+10-15).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), or DNM2 H9D(+20-5). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), or DNM2 H9D(+15-10).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6- 17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6- 17), DNM2 H9D(+5-17), or DNM2 H9D(+5-20).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), or DNM2 H9D(+7-18).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), or DNM2 H9D(+4-21).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-5+20).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H9A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+6+30). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+25+49).
- the targeting sequence is selected from: SEQ ID NO: 158 (CGTCAAACTCCATCTGAGAAGAGAC); SEQ ID NO: 159 (TCCTTCTCGTCAAACTCCATCTGAG); SEQ ID NO: 160 (GTAAGTCCTTCTCGTCAAACTCCAT); SEQ ID NO: 161 (CCGTCGTAAGTCCTTCTCGTCAAAC); SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG); SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT); SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (C
- the targeting sequence is selected from: SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG); SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT); SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 181 (TGCCTGACT
- the targeting sequence is selected from: SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); and SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT).
- the targeting sequence is selected from: SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG);
- the targeting sequence is SEQ ID NO: 172. [0483] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT); SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG); SEQ ID NO: 189
- the targeting sequence is selected from: SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); and SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT).
- the targeting sequence is selected from: SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); and SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG).
- the targeting sequence is selected from: SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); and SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT).
- the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACT
- the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); and SEQ ID NO: 205 (ACTCTCCTCGTGGA
- the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); and SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC).
- the targeting sequence is selected from: SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); and SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA).
- the targeting sequence is SEQ ID NO: 199.
- the targeting sequence is selected from: SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 208 (CTCCTCGTGGAACTTGCCTG); and SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG).
- the targeting sequence is selected from: SEQ ID NO: 158 (CGTCAAACTCCATCTGAGAAGAGAC); SEQ ID NO: 159 (TCCTTCTCGTCAAACTCCATCTGAG); SEQ ID NO: 160 (GTAAGTCCTTCTCGTCAAACTCCAT); SEQ ID NO: 161 (CCGTCGTAAGTCCTTCTCGTCAAAC); SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); and SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG).
- the target region is within the exon internal region of exon 12 (SEQ ID NO: 7), the intron 11/exon 12 junction, or the exon 12/intron 12 junction of the human DNM2 gene pre-mRNA.
- the target region of the human DNM2 gene pre-mRNA is within the 4 th nucleotide to the 31 st nucleotide measured of exon 12 from the 5′ end of exon 12 (SEQ ID NO: 7).
- the target region is a region within SEQ ID NO: 36.
- the target region is a region within DNM2 H12A(+4+31).
- the target region of the human DNM2 gene pre-mRNA is within the 10 th nucleotide to the 44 th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 37. In certain embodiments, the target region is a region within DNM2 H12A(+10+44). [0497] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 10 th nucleotide to the 38 th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
- the target region is a region within SEQ ID NO: 38. In certain embodiments, the target region is a region within DNM2 H12A(+10+38). [0498] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 16 th nucleotide to the 44 th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 39. In certain embodiments, the target region is a region within DNM2 H12A(+16+44).
- the target region of the human DNM2 gene pre-mRNA is within the 14 th nucleotide to the 42 nd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 40. In certain embodiments, the target region is a region within DNM2 H12A(+14+42). [0500] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 25 th nucleotide to the 54 th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
- the target region is a region within SEQ ID NO: 41. In certain embodiments, the target region is a region within DNM2 H12A(+25+54). [0501] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 25 th nucleotide to the 53 rd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 42. In certain embodiments, the target region is a region within DNM2 H12A(+25+53).
- the target region of the human DNM2 gene pre-mRNA is within the 26 th nucleotide to the 54 th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 43. In certain embodiments, the target region is a region within DNM2 H12A(+26+54). [0503] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 36 th nucleotide to the 61 st nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
- the target region is a region within SEQ ID NO: 44. In certain embodiments, the target region is a region within DNM2 H12A(+36+61). [0504] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 22nd nucleotide of exon 12 measured from the 3′ end of exon 12 (SEQ ID NO: 7) to the 5th nucleotide of intron 12 measured from the 3′ end of exon 12. In certain embodiments, the target region is a region within SEQ ID NO: 45. In certain embodiments, the target region is a region within DNM2 H12D(+22-5).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-13+12), DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12
- the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+49), DNM2 H12A(+29+53), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H
- the target region is DNM2 H12A(+4+28) or DNM2 H12A(+7+31). [0508] In certain embodiments, the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44).
- the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), or DNM2 H12A(+14+38).
- the target region is DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), or DNM2 H12A(+20+44).
- the target region is DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), or DNM2 H12A(+18+42).
- the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54).
- the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), or DNM2 H12A(+29+53).
- the target region is DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54).
- the target region is DNM2 H12A(+25+49).
- the target region is DNM2 H12A(+36+60) or DNM2 H12A(+37+61).
- the target region is DNM2 H12D(+22-3) or DNM2 H12D(+20-5).
- the target region is DNM2 H12A(-13+12). In certain embodiments, the target region is DNM2 H12A(-3+22). In certain embodiments, the target region is DNM2 H12A(+43+67). In certain embodiments, the target region is DNM2 H12D(+15-10).
- the targeting sequence is selected from: SEQ ID NO: 210 (CAGCAGAAGAATCTGGBGGAGAGCG); SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG); SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 210 (CAG
- the targeting sequence is selected from: SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG); SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ
- the targeting sequence is selected from: SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); and SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG).
- the targeting sequence is selected from: SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222
- the targeting sequence is selected from: SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); and SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA).
- the targeting sequence is selected from: SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); and SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA).
- the targeting sequence is selected from: SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); and SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG).
- the targeting sequence is selected from: SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAGG);
- the targeting sequence is selected from: SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); and SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG).
- the targeting sequence is selected from: SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); and SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG).
- the targeting sequence is SEQ ID NO: 230
- the targeting sequence is selected from: SEQ ID NO: 238 (GATGAAGTCCTCATGGTTCGTGTTG); and SEQ ID NO: 239 (CGATGAAGTCCTCATGGTTCGTGTT).
- the targeting sequence is selected from: SEQ ID NO: 241 (TACTTGGCAAACCCGATGAAGTCCT); and SEQ ID NO: 242 (CCTACTTGGCAAACCCGATGAAGTC).
- the targeting sequence is selected from: SEQ ID NO: 210 (CAGCAGAAGAATCTGGBGGAGAGCG); SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 240 (CAAACCCGATGAAGTCCTCATGGTT); and SEQ ID NO: 243 (AAGTACCTACTTGGCAAACCCGATG).
- the target region is within the exon internal region of exon 13 (SEQ ID NO: 8), the intron 12/exon 13 junction, the exon 13/intron 13 junction, or the intron internal region of intron 12 (SEQ ID NO: 12).
- the target region of the human DNM2 gene pre-mRNA is within the 27 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12 th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 46. In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within DNM2 H13A(-27+12).
- the target region of the human DNM2 gene pre-mRNA is within the 27 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 2 nd nucleotide of exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 47.
- the target region is a region within DNM2 H13A(-27+2).
- the target region of the human DNM2 gene pre-mRNA is within the 22 nd nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 7 th nucleotide of exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 48.
- the target region is a region within DNM2 H13A(-22+7).
- the target region of the human DNM2 gene pre-mRNA is within the 17 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12 th nucleotide of exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 49.
- the target region is a region within DNM2 H13A(-17+12).
- the target region of the human DNM2 gene pre-mRNA is within the 11 th nucleotide of intron 12 (SEQ ID NO: 12) measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24 th nucleotide of exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 50.
- the target region is a region within DNM2 H13A(-11+24).
- the target region of the human DNM2 gene pre-mRNA is within the 11 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 18 th nucleotide of exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 51.
- the target region is a region within DNM2 H13A(-11+18).
- the target region of the human DNM2 gene pre-mRNA is within the 5 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24 th nucleotide exon 13 measured from the 5′ end of exon 13.
- the target region is a region within SEQ ID NO: 52.
- the target region is a region within DNM2 H13A(-5+24).
- the target region of the human DNM2 gene pre-mRNA is within the 8 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 21 st nucleotide exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 53. In certain embodiments, the target region is a region within DNM2 H13A(-8+21). [0539] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8).
- the target region is a region within SEQ ID NO: 54. In certain embodiments, the target region is a region within DNM2 H13A(+1+30). [0540] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 28th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 55. In certain embodiments, the target region is a region within DNM2 H13A(+1+28).
- the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 56. In certain embodiments, the target region is a region within DNM2 H13A(+6+30). [0542] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 3rd nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8).
- the target region is a region within SEQ ID NO: 57. In certain embodiments, the target region is a region within DNM2 H13A(+3+30). [0543] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15),
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), or DNM2 H13A(- 23+2).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), or DNM2 H13A(- 18+7).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(- 7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(-1+24).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), or DNM2 H13A(-7+18).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(- 1+24).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), or DNM2 H13A(- 4+21).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), or DNM2 H13A(+5+28).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), or DNM2 H13A(+6+30).
- the target region of the human DNM2 gene pre-mRNA DNM2 H13A(+4+28).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+16+40). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H13D(+22-3). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13D(+7-18).
- the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 254 (GCTGG
- the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 254 (GCTGG
- the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 254 (GCTGG
- the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); and SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG).
- the targeting sequence is selected from: SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); and SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA).
- the targeting sequence is selected from: SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); and SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA).
- the targeting sequence is selected from: SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); and SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC).
- the targeting sequence is selected from: SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); and SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC).
- the targeting sequence is selected from: SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); and SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC).
- the targeting sequence is selected from: SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); and SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA).
- the targeting sequence is selected from: SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (CAGCTGCGTGCTCC
- the targeting sequence is selected from: SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); and SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG).
- the targeting sequence is selected from: SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 282 (GTTCAGCTGCGTGCTCCTCTGC); SEQ ID NO: 283 (GTTCAGCTGCGTGCTCCTCTG); and SEQ ID NO: 284 (GTTCAGCTGCGTGCTCCTCT).
- the targeting sequence is selected from: SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); and SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT).
- the targeting sequence is selected from: SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 285 (GGCTCTCTTCTTGTTCAGCTGCGTG); SEQ ID NO: 286 (TACCTGATTGGBGATGGCTCTCTTC); and SEQ ID NO: 287 (TGCAGAGGBGTGTGCTACCTGATTG).
- the target region is within the exon internal region of exon 6 (SEQ ID NO: 3), the intron 5/exon 6 junction, the exon 6/intron 6 junction, or the intron internal region of intron 6 (SEQ ID NO: 11).
- the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 37th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 58. In certain embodiments, the target region is a region within DNM2 H6A(+10+37). [0575] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 32nd nucleotide to the 59th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3).
- the target region is a region within SEQ ID NO: 59. In certain embodiments, the target region is a region within DNM2 H6A(+32+59). [0576] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 60th nucleotide to the 87th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 60. In certain embodiments, the target region is a region within DNM2 H6A(+60+87).
- the target region of the human DNM2 gene pre-mRNA is within the 130th nucleotide to the 156th nucleotide measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 61. In certain embodiments, the target region is a region within DNM2 H6A(+130+156). [0578] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 17th nucleotide of exon 6 measured from the 3′ end of exon 6 (SEQ ID NO: 3) to the 10th nucleotide of intron 6 measured from the 3′ end of exon 6.
- the target region is a region within SEQ ID NO: 62. In certain embodiments, the target region is a region within DNM2 H6D(+17-10). [0579] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5), DNM2 H6A(-6+19), DNM2 H6A(-2+23), DNM2 H6A(+3+26), DNM2 H6A(+10+3), DNM2 H6A(+13+37), DNM2 H6A(+23+47), DNM2 H6A(+32+56), DNM2 H6A(+35+59), DNM2 H6A(+44+68), DNM2 H6A(+54+78), DNM2 H6A(+60+84), DNM2 H6A(+63+87), DNM2 H6A(+88+112), DNM2 H6A(+96+119), DNM2 H6A(+105
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+10+3) or DNM2 H6A(+13+37).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+32+56) or DNM2 H6A(+35+59).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+60+84) or DNM2 H6A(+63+87).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+130+154) or DNM2 H6A(+132+156).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+15-10) or DNM2 H6D (+17-8).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-6+19).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-2+23).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+3+26). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+23+47). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+44+68). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+54+78). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+88+112).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H6A(+96+119). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+105+129). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+121+145). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+6-19).
- the targeting sequence is selected from: SEQ ID NO: 97 (GTAGCCTGTGGGGAGAGGCAGTCAG); SEQ ID NO: 98 (TTCACCACGCCAATGTAGCCTGTGG); SEQ ID NO: 99 (GCGGTTCACCACGCCAATGTAGCCT); SEQ ID NO: 100 (GCTGCGGTTCACCACGCCAATGTA); SEQ ID NO: 101 (TCCTTCTGGCTGCGGTTCACCACGC); SEQ ID NO: 102 (ATATCCTTCTGGCTGCGGTTCACCA); SEQ ID NO: 103 (CTTGCCCTCAATATCCTTCTGGCTG); SEQ ID NO: 104 (GATGTCCTTCTTGCCCTCAATATCC); SEQ ID NO: 105 (ACGGATGTCCTTCTTGCCCTCAATA); SEQ ID NO: 106 (CAGTGCTGCACGGATGTCCTTCTTG); SEQ ID NO: 107 (TCTCAGCTGCCAGTGCTGCACGGAT); SEQ ID NO: 107 (TCTCAGCTG
- the target region is within the exon internal region of exon 7 (SEQ ID NO: 4), the intron 6/exon 7 junction, or the exon 7/intron 7 junction.
- the target region of the human DNM2 gene pre-mRNA is within the 72nd nucleotide to the 98th nucleotide of exon 7 measured from the 5′ end of exon 7 (SEQ ID NO: 4).
- the target region is a region within SEQ ID NO: 63.
- the target region is a region within DNM2 H7A(+72+98).
- the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 7 measured from the 3′ end of exon 7 (SEQ ID NO: 4) to the 16th nucleotide of intron 7 measured from the 3′ end of exon 7.
- the target region is a region within SEQ ID NO: 64.
- the target region is a region within DNM2 H7D(+9-16).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+1+25), DNM2 H7A(+6+30), DNM2 H7A(+11+35), DNM2 H7A(+22+46), DNM2 H7A(+29+53), DNM2 H7A(+41+65), DNM2 H7A(+51+75), DNM2 H7A(+57+81), DNM2 H7A(+62+86), DNM2 H7A(+72+96), DNM2 H7A(+74+98), DNM2 H7A(+81+105), DNM2 H7A(+91+115), or DNM2 H7A(+110+134).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+72+96) or DNM2 H7A(+74+98).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+8-16) or DNM2 H7D(+9-16).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-15+10). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-5+20).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H7A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+6+30). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+11+35). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+22+46). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+29+53). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+41+65).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+51+75). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+57+81). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+62+86). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H7A(+81+105). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+91+115).
- the target region is within the exon internal region of exon 8 (SEQ ID NO: 5), the intron 7/exon 8 junction, or the exon 8/intron 8 junction.
- the target region of the human DNM2 gene pre-mRNA is within the 7th nucleotide of intron 7 measured from the 5′ end of exon 8 (SEQ ID NO: 5) to the 19th nucleotide of exon 8 measured from the 5′ end of exon 8.
- the target region is a region within SEQ ID NO: 65.
- the target region is a region within DNM2 H8A(-7+19).
- the target region of the human DNM2 gene pre-mRNA is within the 18th nucleotide to the 45th nucleotide of exon 8 measured from the 5′ end of exon 8 (SEQ ID NO: 5). In certain embodiments, the target region is a region within SEQ ID NO: 66. In certain embodiments, the target region is a region within DNM2 H8A(+18+45).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-20+5), DNM2 H8A(-14+11), DNM2 H8A(-7+18), DNM2 H8A(-6+19), DNM2 H8A(-2+23), DNM2 H8A(+5+29), DNM2 H8A(+18+42), DNM2 H8A(+21+45), DNM2 H8A(+27+51), DNM2 H8A(+36+60), DNM2 H8A(+46+70), DNM2 H8A(+51+75), DNM2 H8A(+56+80), DNM2 H8A(+65+88), DNM2 H8A(+76+100), DNM2 H8A(+83+107), DNM2 H8A(+96+119), DNM2 H8A(+105+129), DNM2 H8D(+18-7), or DNM2 H8D(+
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-7+18) or DNM2 H8A(-6+19). [0599] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+18+42) or DNM2 H8A(+21+45). [0600] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-20+5). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-14+11). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H8A(-2+23).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+5+29). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+27+51). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+36+60). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+46+70). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+51+75). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+56+80).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+65+88). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+76+100). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H8A(+83+107). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+96+119). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+105+129). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+18-7).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+10-15).
- the targeting sequence is selected from: SEQ ID NO: 138 (CCATCCTATGAGGAAAAGGAAAAAG); SEQ ID NO: 139 (GCTGGACCATCCTATGAGGAAAAGG); SEQ ID NO: 140 (CCAAACTGCTGGACCATCCTATGAG); SEQ ID NO: 141 (CCCAAACTGCTGGACCATCCTATGA); SEQ ID NO: 142 (CCACCCCAAACTGCTGGACCATCCT); SEQ ID NO: 143 (CAAAATCCACCCCAAACTGCTGGAC); SEQ ID NO: 144 (TCGATCCTCTTCTCAAAATCCACCC); SEQ ID NO: 145 (CCCTCGATCCTCTTCTCAAAATCCA); SEQ ID NO: 146 (CCTGAGCCCTCGATCCTCTTCTCAA); SEQ ID NO: 147 (ACCTGATCTCCTG
- the target region is within the exon internal region of exon 16 (SEQ ID NO: 9), the intron 15/exon 16 junction, or the exon 16/intron 16 junction.
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3), DNM2 H16A(-13+12), DNM2 H16A(-6+19), DNM2 H16A(+6+30), DNM2 H16A(+15+39), DNM2 H16A(+26+50), DNM2 H16A(+36+60), DNM2 H16A(+45+69), DNM2 H16A(+55+79), DNM2 H16A(+66+90), DNM2 H16A(+75+99), DNM2 H16A(+82+106), DNM2 H16D(+19-6), or DNM2 H16D(+11-14).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-13+12).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H16A(-6+19).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+6+30).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+15+39).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+26+50). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+36+60). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+45+69). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+55+79). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+66+90). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+75+99).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+82+106). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16D(+19-6). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16D(+11-14).
- the targeting sequence is selected from: SEQ ID NO: 288 (CTCCTGAGGGTGGAGGGAAAAGAG); SEQ ID NO: 289 (CTTCTCTTTCTCCTGAGGGTGGAGG); SEQ ID NO: 290 (TGTACTTCTTCTCTTTCTCCTGAGG); SEQ ID NO: 291 (CAGAGGCAGCATGTACTTCTTCTCT); SEQ ID NO: 292 (GAGGTTGTCCAGAGGCAGCATGTAC); SEQ ID NO: 293 (TCACGGATCTTGAGGTTGTCCAGAG); SEQ ID NO: 294 (CTTCTCCACATCACGGATCTTGAGG); SEQ ID NO: 295 (CATGAAGCCCTTCTCCACATCACGG); SEQ ID NO: 296 (GCTTGTTGGACATGAAGCCCTTCTC); SEQ ID NO: 297 (GGCGAAGACGTGCTTGTTGGACATG); SEQ ID NO: 298 (GTTGAAGATGGCGAAGACGTGCTTG); SEQ ID NO: 299 (GGCGAAGA
- the target region is within the exon internal region of exon 19 (SEQ ID NO: 9), the intron 18/exon 19 junction, or the exon 18/intron 18 junction.
- the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14), DNM2 H19A(-4+21), DNM2 H19A(+1+25), DNM2 H19A(+7+31), DNM2 H19A(+16+40), DNM2 H19A(+26+50), DNM2 H19A(+37+61), DNM2 H19A(+47+71), DNM2 H19A(+56+80), DNM2 H19A(+66+90), DNM2 H19A(+76+100), DNM2 H19A(+86+110), DNM2 H19A(+96+120), DNM2 H19A(+106+130), DNM2 H19A(+117+141), DNM
- the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-4+21). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+7+31). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+16+40).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+26+50). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+37+61). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+47+71). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+56+80). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+66+90). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+76+100).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+86+110). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+96+120). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+106+130). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+117+141). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+127+151).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+137+161). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+147+171). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+157+181). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+166+189). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+191+215).
- the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+200+224). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+23-2). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+17-8). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+10-15).
- the targeting sequence is selected from: SEQ ID NO: 302 (ATGAAGGCCTTCGTCTGCGGGCAGG); SEQ ID NO: 303 (GTGGTGGATGAAGGCCTTCGTCTGC); SEQ ID NO: 304 (GCTCGTGGTGGATGAAGGCCTTCGT); SEQ ID NO: 305 (CCAGCAGCTCGTGGTGGATGAAGGC); SEQ ID NO: 306 (ATAGGTAGGCCAGCAGCTCGTGGTG); SEQ ID NO: 307 (GCCGAGGAGTATAGGTAGGCCAGCA); SEQ ID NO: 308 (TGCTCTGGTCTGCCGAGGAGTATAG); SEQ ID NO: 309 (TCCATGAGGCTGCTCTGGTCTGCCG); SEQ ID NO: 310 (GCCGACTCCTCCATGAGGCTGCTCT); SEQ ID NO: 311 (TGCCTGGTCAGCCGACTCCTCCATG); SEQ ID NO: 312 (CCCGCCGCTGTGCCTGGTCAGCCGA); SEQ ID NO:
- the targeting sequence is at least 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99 percent complementary to the target region.
- the targeting sequence is at least 84%, at least 88%, or at least 92% complementary to the target region.
- the targeting sequence is at least 90% complementary to the target region.
- the targeting sequence is at least 95% complementary to the target region.
- the targeting sequence is 100% complementary to the target region.
- the antisense oligomer of the present disclosure can have a modified chemical backbone as described herein or known in the art.
- the antisense oligomer may be a peptide nucleic acid (PNA), a locked nucleic acid, a phosphorodiamidate morpholino oligomer, a 2′-OMe phosphorothioate oligomer, or a combination thereof.
- the antisense oligomer is a phosphorodiamidate morpholino oligomer.
- the antisense oligomer is a peptide nucleic acid.
- the antisense oligomer is a locked nucleic acid.
- the antisense oligomer is a 2′-OMe phosphorothioate oligomer. In still another embodiment, the antisense oligomer is a combination of any of the above oligonucleotides having a modified chemical backbone. [0612] In some embodiments, the antisense oligomer is conjugated to, e.g., covalently attached to, a delivery moiety such as, for example, a cell-penetrating peptide, an antibody, a fragment of an antibody, or an antigen fragment of an antibody. In an embodiment, the antisense oligomer is conjugated to a cell-penetrating peptide.
- the antisense oligomer is conjugated to a cell-penetrating peptide, wherein the cell-penetrating peptide is selected from rTAT, Tat, R 9 F 2 , R 5 F 2 R 4 , R 4 , R 5 , R 6 , R 7 , R 8 , R 9 , (RXR) 4 , (RXR) 5 , (RXRRBR) 2 , (RAR) 4 F 2 , (RGR) 4 F 2 , and RBRBYLIQFRBRRBR, wherein A represents alanine, B represents beta-alanine, F represents phenylalanine, G represents glycine, I represents isoleucine, L represents leucine, Q represents glutamine, R represents arginine, X represent 6-aminohexanoic acid, and Y represents tyrosine.
- the cell-penetrating peptide is selected from rTAT, Tat, R 9 F 2 , R 5 F 2
- t is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28.
- each R 2 taken together forms a targeting sequence of 13-30 bases.
- each R 2 taken together forms a targeting sequence of 11-28 bases.
- each R 2 taken together forms a targeting sequence of 18-27 bases.
- each R 2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases.
- the pharmaceutically acceptable salt is an HCl salt.
- E′ is selected from H, -C1-6-alkyl, -C(O)C1-6-alkyl, benzoyl, stearoyl, trityl, monomethoxytrityl, dimethoxytrityl, trimethoxytrityl, and .
- E′ is selected from -H, -C(O)CH3, benzoyl, stearoyl, trityl, 4- methoxytrityl, and .
- A′ is selected from: .
- A′ is selected from: ; and E′ is .
- A′ is [0620]
- E′ is selected from H, -C(O)CH 3 , trityl, 4-methoxytrityl, benzoyl, and stearoyl.
- each R 1 is -N(CH3)2.
- L is glycine, proline, or b-alanine.
- L is glycine.
- L is proline.
- L is b-alanine.
- the pharmaceutically acceptable salt is an HCl salt.
- each R 2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases.
- the pharmaceutically acceptable salt is an HCl salt.
- an antisense oligomer which is an oligomer of Formula (II): or a pharmaceutically acceptable salt thereof, wherein t, G, and R 2 are as described herein.
- t is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28.
- each R 2 taken together forms a targeting sequence of 13-30 bases.
- each R 2 taken together forms a targeting sequence of 11-28 bases.
- each R 2 taken together forms a targeting sequence of 18-27 bases.
- each R 2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases.
- the pharmaceutically acceptable salt is an HCl salt.
- an antisense oligomer which is an oligomer of Formula (III): (III), or a pharmaceutically acceptable salt thereof, wherein n is 11-28 and R 2 is as described herein.
- n is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28.
- each R 2 taken together forms a targeting sequence of 13- 30 bases.
- each R 2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt. [0627] In some embodiments, provided herein is an antisense oligomer, which is an oligomer of Formula (IV): (IV), wherein n is 11-28 and R 2 is as described herein. [0628] In certain embodiments, n is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28.
- each R 2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R 2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. [0629] In certain embodiments, each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 83. In certain embodiments, each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 88. In certain embodiments, each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 172.
- R 2 taken together, forms a targeting sequence having SEQ ID NO: 199. In certain embodiments, each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 230. In certain embodiments, each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 278. [0630] In certain embodiments, the antisense oligomer is selected from:
- the antisense oligomer has the structure: .
- the antisense oligomer has the structure:
- the antisense oligomer has the structure:
- the antisense oligomer has the structure:
- the antisense oligomer has the structure:
- the antisense oligomer has the structure: . IV. Target Sequences and Target Regions [0637]
- the oligomer can be 100% complementary to the nucleic acid target sequence (excluding at least one abasic subunit), or it may include mismatches, e.g., to accommodate variants, as long as a heteroduplex formed between the oligomer and nucleic acid target sequence is sufficiently stable to withstand the action of cellular nucleases and other modes of degradation which may occur in vivo. Mismatches, if present, are less destabilizing toward the end regions of the hybrid duplex than in the middle.
- the number of mismatches allowed will depend on the length of the oligomer, the percentage of G: C base pairs in the duplex, and the position of the mismatch(es) in the duplex, according to well-understood principles of duplex stability.
- an antisense oligomer is not necessarily 100% complementary to the nucleic acid target sequence, it is effective to stably and specifically bind to the target sequence, such that a biological activity of the nucleic acid target, e.g., expression of the encoded protein(s), is modulated.
- the stability of the duplex formed between an oligomer and the target sequence is a function of the binding T m and the susceptibility of the duplex to cellular enzymatic cleavage.
- each antisense oligomer has a binding T m , with respect to a complementary-sequence RNA, of greater than body temperature or in other embodiments greater than 50 °C. In other embodiments, T m s are in the range of 60-80 °C or greater.
- the T m of an oligomer compound, with respect to a complementary-based RNA hybrid can be increased by increasing the ratio of C: G paired bases in the duplex, and/or by increasing the length (in base pairs) of the heteroduplex.
- compounds show high T m (50 °C or greater) at a length of 20 bases or less.
- longer oligomers for example longer than 20 bases, may have certain advantages.
- the targeting sequence bases may be normal DNA bases or analogues thereof, e.g., uracil and inosine that are capable of Watson-Crick base pairing to target-sequence RNA bases.
- An antisense oligomer can be designed to block or inhibit or modulate translation of mRNA or to inhibit or modulate pre-mRNA splice processing, or induce degradation of targeted mRNAs, and may be said to be “directed to” or “targeted against” a target sequence with which it hybridizes.
- the target sequence includes a region including a 3′ or 5′ splice site of a pre-processed mRNA, a branch point, or other sequence involved in the regulation of splicing.
- the target sequence may be within an exon or within an intron or spanning an intron/exon junction.
- An antisense oligomer having a sufficient sequence complementarity to a target RNA sequence to modulate splicing of the target RNA means that the antisense agent has a sequence sufficient to trigger the masking of a binding site for a native protein that would otherwise modulate splicing and/or alters the three-dimensional structure of the targeted RNA.
- an oligomer reagent having a sufficient sequence complementary to a target RNA sequence to modulate splicing of the target RNA means that the oligomer reagent has a sequence sufficient to trigger the masking of a binding site for a native protein that would otherwise modulate splicing and/or alters the three-dimensional structure of the targeted RNA.
- the antisense oligomer has sufficient length and complementarity to target region within a pre-mRNA of the human dynamin 2 (DNM2) gene (SEQ ID NO: 1).
- the human DNM2 gene (SEQ ID NO:1) and the exon 2 (SEQ ID NO: 2), exon 6 (SEQ ID NO: 3), exon 7 (SEQ ID NO: 4), exon 8 (SEQ ID NO: 5), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), exon 13 (SEQ ID NO: 8), exon 16 (SEQ ID NO: 9), and exon 19 (SEQ ID NO: 10) sequences for human DNM2 gene pre-mRNA are shown in Table 2. Table 2. Target Regions for DNM2-targeted oligomers.
- oligomers as long as 40 bases may be suitable, where at least a minimum number of bases, e.g., 10-12 bases, are complementary to the target sequence.
- facilitated or active uptake in cells is optimized at oligomer lengths of less than about 30 bases.
- an optimum balance of binding stability and uptake generally occurs at lengths of 18-25 bases.
- antisense oligomers e.g., PMOs, PMO-X, PNAs, LNAs, 2′-OMe
- PMOs, PMO-X, PNAs, LNAs, 2′-OMe e.g., PMOs, PMO-X, PNAs, LNAs, 2′-OMe
- PNAs e.g., PNAs, PNAs, LNAs, 2′-OMe
- 2′-OMe antisense oligomers that consist of about 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, or 40 contiguous or non-contiguous bases that are complementary to the desired target sequences.
- antisense oligomers may be 100% complementary to the target sequence (excluding at least one abasic nucleotide), or may include mismatches, e.g., to accommodate variants, as long as a heteroduplex formed between the oligomer and target sequence is sufficiently stable to withstand the action of cellular nucleases and other modes of degradation which may occur in vivo.
- certain oligomers may have substantial complementarity, meaning, about or at least about 70% sequence complementarity, e.g., 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% sequence complementarity, between the oligomer (excluding at least one abasic nucleotide) and the target sequence. Oligomer backbones that are less susceptible to cleavage by nucleases are discussed herein.
- Mismatches are typically less destabilizing toward the end regions of the hybrid duplex than in the middle.
- the number of mismatches allowed will depend on the length of the oligomer, the percentage of G: C base pairs in the duplex, and the position of the mismatch(es) in the duplex, according to well-understood principles of duplex stability.
- an antisense oligomer is not necessarily 100% complementary to the target sequence, it is effective to stably and specifically bind to the target sequence, such that splicing of the target pre-RNA is modulated.
- the stability of the duplex formed between an oligomer and a target sequence is a function of the binding Tm and the susceptibility of the duplex to cellular enzymatic cleavage.
- the Tm of an oligomer with respect to complementary-sequence RNA may be measured by conventional methods, such as those described by Hames et al., Nucleic Acid Hybridization, IRL Press, 1985, pp.107-108 or as described in Miyada C. G. and Wallace R. B., 1987, Oligomer Hybridization Techniques, Methods Enzymol. Vol.154 pp.94-107.
- antisense oligomers may have a binding Tm, with respect to a complementary- sequence RNA, of greater than body temperature and greater than about 45 °C or 50 °C. T m s in the range 60-80 °C or greater are also included.
- Tm of an oligomer, with respect to a complementary-based RNA hybrid can be increased by increasing the ratio of C: G paired bases in the duplex, and/or by increasing the length (in base pairs) of the heteroduplex.
- antisense targeting sequences are designed to hybridize to a region of one or more of the target sequences listed in Table 2. Selected antisense targeting sequences can be made shorter, e.g., about 12 bases, or longer, e.g., about 40 bases, and include a small number of mismatches, as long as the sequence is sufficiently complementary to effect splice modulation upon hybridization to the target sequence, and optionally forms with the RNA a heteroduplex having a Tm of 45 °C or greater. V.
- the antisense oligomer described herein is conjugated to, e.g., covalently attached to, a delivery moiety. Any delivery moiety that increases cellular uptake of the antisense oligomer may be suitable for use herein.
- the antisense oligomer may be covalently attached to delivery moiety such as a cell-penetrating peptide, an antibody, a fragment of an antibody, or an antigen fragment of an antibody.
- the antisense oligomer is conjugated to an antibody or antibody fragment.
- Antibodies and antibody fragments that affect cellular uptake are known in the art, and any such antibody or antibody fragment may be covalently linked to the oligomer of the present disclosure.
- the antisense oligomer is conjugated to an antibody is an anti-TfRl antibody.
- anti-TfRI antibodies means of conjugation to an antisense oligomer, complexes comprising antisense oligomers covalently linked to an anti-transferrin receptor 1 (TfRl) antibody, and formulations thereof for use in the instant disclosure can be found in WO/2023/283623 and WO/2023/283624, both of which are incorporated by reference in their entirety.
- the antisense oligomer is an antibody-peptide-oligomer conjugate, for example, of formula A-(Xi-B-X2-D)n or A-(Xi-D-X2-B)n, wherein A is an antibody or antigen binding fragment thereof; B is a polynucleotide; D is an endosomolytic peptide or a membrane penetrating peptide; Xi is a bond or a first non-polymeric linker; X2 is an optional bond or an optional second linker; and n is an integer > 1, as described in WO/2022/212886, which is incorporated by reference in its entirety.
- the antisense oligomer is conjugated to a cell-penetrating peptide.
- the antisense oligomer of the present disclosure may be covalently linked to a cell- penetrating peptide as described herein or as known in the art.
- arginine-rich cell-penetrating peptides CPP
- arginine-rich peptides CPP
- exemplary arginine-rich peptides are provided in Table 3.
- Table 3 Arginine-Rich Peptide Transporters (J).
- an antisense oligomer disclosed herein is a conjugate of D-Pep 1.9b wherein L is isoglutamine (IsoGln). Examples of such conjugates can be found in WO/2022/171972 which is incorporated by reference in its entirety.
- exemplary cell-penetrating peptides within the scope of substituent J are provided in Table 4.
- the point of connection to substituent “L” is as shown in Table 4.
- Table 4 Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) (RFF)3; CP0407 RFFRFFRFF-L RTR RTRTRFLRRT-L RFFR RFFRFFRFFR-L KTR KTRTKFLKKT-L KFF KFFKFFKFF-L KFFK KFFKFFKFFK-L (RFF)2 RFFRFF-L (RFF)2R RFFRFFR-L R ⁇ R ⁇ R ⁇ R-L (R ⁇ R)4; P007 R ⁇ RR ⁇ RR ⁇ RR ⁇ R-L RBRBYLIQFRBRRBR-L Tat47_58 YGRKKRRQRRR-L Tat48_58 GRKKRRQRRR-L Tat49_58 RKKRRQRRR-L Penetratin RQI
- Linker “L” may additionally be an Isoglutamine amino acid residue (IsoGln) linking the carrier peptide to the oligomer by an amide bond between the carboxyl of IsoGln and the oligomer and an amide bond between the amino of IsoGln and the carboxy terminus of the peptide.
- IsoGln is an Isoglutamine amino acid residue linking the carrier peptide to the oligomer by an amide bond between the carboxyl of IsoGln and the oligomer and an amide bond between the amino of IsoGln and the carboxy terminus of the peptide. Examples of use of IsoGln as a linker can be found in, for example, WO/2022/171972, which is incorporated by reference in its entirety.
- compositions [0655] The present disclosure also provides for the formulation and delivery of the disclosed antisense oligomers. Accordingly, an aspect of the present disclosure is a pharmaceutical composition comprising antisense oligomers as disclosed herein and a pharmaceutically acceptable carrier. [0656] Effective delivery of the antisense oligomers to the target nucleic acid is an important aspect of treatment. Routes of antisense oligomer delivery include, but are not limited to, various systemic routes, including oral and parenteral routes, e.g., intravenous, subcutaneous, intraperitoneal, and intramuscular, as well as inhalation, transdermal, and topical delivery. The appropriate route may be determined by one of skill in the art, as appropriate to the condition of the subject under treatment.
- an appropriate route for delivery of an antisense oligomer in the treatment of a viral infection of the skin is topical delivery, while the delivery of an antisense oligomer for the treatment of a viral respiratory infection can be intravenous or by inhalation.
- the antisense oligomer may also be delivered directly to any particular site of viral infection.
- the antisense oligomer can be administered in any convenient vehicle which is physiologically and/or pharmaceutically acceptable.
- Such a composition can include any of a variety of standard pharmaceutically acceptable carriers employed by those of ordinary skill in the art.
- Examples include, but are not limited to, saline, phosphate-buffered saline (PBS), water (e.g., sterile water for injection), aqueous ethanol, emulsions such as oil/water emulsions or triglyceride emulsions, tablets, and capsules.
- PBS phosphate-buffered saline
- water e.g., sterile water for injection
- aqueous ethanol emulsions such as oil/water emulsions or triglyceride emulsions
- tablets e.g., triglyceride emulsions
- suitable physiologically acceptable carrier will vary dependent upon the chosen mode of administration.
- the compounds disclosed herein e.g., an antisense oligomer
- the compounds disclosed herein may be used in the form of acid or base addition salts.
- Acid addition salts of the free amino compounds may be prepared by methods well known in the art and may be formed from organic and inorganic acids.
- Suitable organic acids include maleic, fumaric, benzoic, ascorbic, succinic, methanesulfonic, acetic, trifluoroacetic, oxalic, propionic, tartaric, salicylic, citric, gluconic, lactic, mandelic, cinnamic, aspartic, stearic, palmitic, glycolic, glutamic, and benzenesulfonic acids.
- Suitable inorganic acids include hydrochloric, hydrobromic, sulfuric, phosphoric, and nitric acids.
- Base addition salts included those salts that form with the carboxylate anion and include salts formed with organic and inorganic cations such as those chosen from the alkali and alkaline earth metals (for example, lithium, sodium, potassium, magnesium, barium and calcium), as well as the ammonium ion and substituted derivatives thereof (for example, dibenzylammonium, benzylammonium, 2-hydroxyethylammonium, and the like).
- the term “pharmaceutically acceptable salt” of Formulae (I), (IA), (II), and (III) is intended to encompass any and all acceptable salt forms.
- prodrugs are also included within the context of this disclosure.
- Prodrugs are any covalently bonded carriers that release a compound of structure (I) in vivo when such a prodrug is administered to a patient.
- Prodrugs are generally prepared by modifying functional groups in a way such that the modification is cleaved, either by routine manipulation or in vivo, yielding the parent compound.
- Prodrugs include, for example, compounds of this disclosure wherein hydroxy, amine, or sulfhydryl groups are bonded to any group that, when administered to a patient, cleaves to form the hydroxy, amine, or sulfhydryl groups.
- representative examples of prodrugs include (but are not limited to) acetate, formate, and benzoate derivatives of alcohol and amine functional groups of the compounds of structure (I).
- the morpholino subunits may be prepared from the corresponding ribonucleoside (1) as shown.
- the morpholino subunit (2) may be optionally protected by reaction with a suitable protecting group precursor, for example, trityl chloride.
- the 3′ protecting group is generally removed during solid-state oligomer synthesis as described in more detail below.
- the base pairing moiety may be suitably protected for sold phase oligomer synthesis.
- Suitable protecting groups include benzoyl for adenine and cytosine, phenylacetyl for guanine, and pivaloyloxymethyl for hypoxanthine (I).
- the pivaloyloxymethyl group can be introduced onto the N1 position of the hypoxanthine heterocyclic base. Although an unprotected hypoxanthine subunit may be employed, yields in activation reactions are far superior when the base is protected.
- Other suitable protecting groups include those disclosed in U.S. Patent No. 8,076,476, which is hereby incorporated by reference in its entirety. [0663] Reaction of 3 with the activated phosphorous compound 4 results in morpholino subunits having the desired linkage moiety 5.
- the protecting group e.g., trityl
- the free amine is reacted with an activated phosphorous moiety of a second compound of structure 5. This sequence is repeated until the desired length of oligo is obtained.
- the protecting group in the terminal 5′ end may either be removed or left on if a 5′-modification is desired.
- the preparation of modified morpholino subunits and morpholino oligomers are described in more detail in the Examples.
- the morpholino oligomers containing any number of modified linkages may be prepared using methods described herein, methods known in the art and/or described by reference herein.
- PMO with a 3′ trityl modification are synthesized essentially as described in PCT publication number WO 2009/064471 with the exception that the detritylation step is omitted.
- Methods of Use Provided herein is a method of reducing the expression of a pre-mRNA of the human DNM2 gene and/or protein using the antisense oligomers of the present disclosure for therapeutic purposes (e.g., treating subjects with a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein, including but not limited to a centronuclear myopathy).
- DNM2 dynamin 2
- the method comprises administering to a patient in need thereof a therapeutically effective amount of an antisense oligomer disclosed herein or a pharmaceutical composition thereof.
- the disease is a centronuclear myopathy.
- the antisense oligomer comprising a nucleotide sequence of sufficient length and complementarity to specifically hybridize to a region within the pre-mRNA of the human DNM2 gene, wherein binding of the antisense oligomer to the region induces exon skipping in the human DNM2 gene pre-mRNA, thereby triggering nonsense-mediated decay of DNM2 in a cell and/or tissue of the subject.
- Exemplary antisense targeting sequences are shown in Tables 5 and 6 herein.
- the disease or disorder is caused by at least one mutation in a gene selected from DNM2 gene, MTM1 gene, BIN1 gene, RYR1 gene, TTN gene, SPEG gene, CACNA1S gene, MTMR14 gene, ZAK gene, and CCDC78 gene.
- the disease or disorder is autosomal dominant, autosomal recessive, or X-linked.
- the disease or disorder is a genetic muscle disorder.
- the disease or disorder is a neuromuscular disorder.
- the disease or disorder is a centronuclear myopathy (CNM).
- the disease or disorder is myotubular myopathy (XLMTM or XLCNM), autosomal dominant congenital myopathy (ADCNM), autosomal recessive centronuclear myopathy (ARCNM), or Charcot-Marie-Tooth disease (CMT).
- CMT is DNM2-implicated CMT.
- the Charcot-Marie-Tooth disease (CMT) is Charcot-Marie-Tooth disease, dominant intermediate B (CMTDIB or DI-CMTB).
- the Charcot- Marie-Tooth disease is Charcot-Marie-Tooth disease, axonal type 2M (CMT2M).
- the disease or disorder is a hereditary spastic paraplegia (HSP).
- HSP hereditary spastic paraplegia
- the disease or disorder is prostate cancer or pancreatic cancer.
- CCM centronuclear myopathy
- the antisense oligomer compound comprises a non- natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of the human dynamin 2 (DNM2) gene (SEQ ID NO: 1).
- the target region is an intron/exon junction, exon/intron junction, an exon internal region, or intron internal region of the human DNM2 gene pre-mRNA.
- the antisense oligomer compound comprises a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of the human dynamin 2 (DNM2) gene (SEQ ID NO: 1).
- the target region is within an exon internal region of the human DNM2 gene pre- mRNA selected from exon 2 (SEQ ID NO: 2), exon 6 (SEQ ID NO: 3), exon 7 (SEQ ID NO: 4), exon 8 (SEQ ID NO: 5), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), exon 13 (SEQ ID NO: 8), exon 16 (SEQ ID NO: 9), and exon 19 (SEQ ID NO: 10).
- exon 2 SEQ ID NO: 2
- exon 6 SEQ ID NO: 3
- exon 7 SEQ ID NO: 4
- exon 8 SEQ ID NO: 5
- exon 9 SEQ ID NO: 6
- exon 12 SEQ ID NO: 7
- exon 13 SEQ ID NO: 8
- exon 16 SEQ ID NO: 9
- exon 19 SEQ ID NO: 10
- the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), and exon 13 (SEQ ID NO: 8).
- the target region is within an intron-exon junction or an exon- intron junction region of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 5/exon 6 junction, exon 6/intron 6 junction, intron 6/exon 7 junction, exon 7/intron 7 junction, intron 7/exon 8 junction, exon 8/intron 8 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, exon 13/intron 13 junction, intron 15/exon 16 junction, exon 16/intron 16 junction, intron 18/exon 19 junction, and exon 19/intron 19 junction [0678] In certain embodiments, the target region is the target region is within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon
- the target region is the target region is within an intron internal region of the human DNM2 gene pre-mRNA selected from intron 2 (SEQ ID NO: 11) and intron 12 (SEQ ID NO: 12).
- the antisense oligomer compound comprises a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of the human dynamin 2 (DNM2) gene (SEQ ID NO: 1).
- the target region is an intron/exon junction or an exon internal region of the human DNM2 gene pre-mRNA, and wherein the antisense oligomer is an antisense oligomer of Formula (I), Formula (IA), Formula (II), or Formula (III).
- a subject has aberrant expression and/or aggregation of DNM2 protein.
- DNM2 expression is reduced by about or at least about 5%, 6%, 7%, 8%, 9%, 10%, 11%, 12%, 13%, 14%, 15%, 16%, 17%, 18%, 19%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100% relative to a control, for example, a control cell/subject, a control composition without the antisense oligomer, the absence of treatment, and/or an earlier time-point. In some instances, DNM2 expression is reduced by about 5% to about 100%, relative to a control.
- DNM2 expression is reduced by about 10% to about 95%, relative to a control. In some instances, DNM2 expression is reduced by about 20% to about 95%, relative to a control. In some instances, DNM2 expression is reduced by about 30% to about 95%, relative to a control. In some instances, DNM2 expression is reduced by about 30% to about 90%, relative to a control. In some instances, DNM2 expression is reduced by about 40% to about 90%, relative to a control. In some instances, DNM2 expression is reduced by about 50% to about 90%, relative to a control. In some instances, DNM2 expression is reduced by about 45% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 50% to about 85%, relative to a control.
- DNM2 expression is reduced by about 55% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 60% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 65% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 50% to about 80%, relative to a control. In some instances, DNM2 expression is reduced by about 60% to about 80%, relative to a control. In some instances, DNM2 expression is reduced by about 50% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 50% to about 70%, relative to a control.
- DNM2 expression is reduced by about 60% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 65% to about 75%, relative to a control. [0682] In some instances, DNM2 expression is reduced by about 30% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 30% to about 80%, relative to a control. In some instances, DNM2 expression is reduced by about 85% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 40% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 40% to about 80%, relative to a control.
- DNM2 expression is reduced by about 40% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 20% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 20% to about 80%, relative to a control. In some instances, DNM2 expression is reduced by about 20% to about 85%, relative to a control. [0683] Also included are methods of reducing one or more symptoms of a centronuclear myopathy in a subject in need thereof. Examples include symptoms such as elevated blood creatinine levels, interstitial fibrosis, tubular atrophy, gout, hyperuricemia, and end stage kidney disease (ESKD).
- EKD end stage kidney disease
- the antisense oligomers of the disclosure can be administered to subjects to treat (prophylactically or therapeutically) centronuclear myopathy.
- pharmacogenomics i.e., the study of the relationship between an individual’s genotype and that individual’s response to a foreign compound or drug
- Differences in the metabolism of therapeutics can lead to severe toxicity or therapeutic failure by altering the relation between dose and blood concentration of the pharmacologically active drug.
- a physician or clinician may consider applying knowledge obtained in relevant pharmacogenomics studies in determining whether to administer a therapeutic agent as well as tailoring the dosage and/or therapeutic regimen of treatment with a therapeutic agent.
- Routes of antisense oligomer delivery include, but are not limited to, various systemic routes, including oral and parenteral routes, e.g., intravenous, subcutaneous, intraperitoneal, and intramuscular, as well as inhalation, transdermal, and topical delivery.
- the appropriate route may be determined by one of skill in the art, as appropriate to the condition of the subject under treatment.
- Vascular or extravascular circulation, the blood or lymph system, and cerebrospinal fluid are some non-limiting sites where the RNA may be introduced.
- Direct CNS delivery may be employed, for instance, intracerebral ventricular or intrathecal administration may be used as routes of administration.
- the antisense oligomer(s) are administered to the subject by intramuscular injection (IM), i.e., they are administered or delivered intramuscularly.
- IM intramuscular injection
- intramuscular injection sites include the deltoid muscle of the arm, the vastus lateralis muscle of the leg, and the ventrogluteal muscles of the hips, and dorsogluteal muscles of the buttocks.
- a PMO, PMO-X, or PPMO is administered by IM.
- the antisense oligomers of the disclosure can be delivered by transdermal methods (e.g., via incorporation of the antisense oligomers into, e.g., emulsions, with such antisense oligomers optionally packaged into liposomes).
- transdermal and emulsion/liposome-mediated methods of delivery are described for delivery of antisense oligomers in the art, e.g., in U.S. Pat. No.6,965,025, the contents of which are incorporated in their entirety by reference herein.
- the antisense oligomers described herein may also be delivered via an implantable device.
- Antisense oligomers can be introduced into cells using art-recognized techniques (e.g., transfection, electroporation, fusion, liposomes, colloidal polymeric particles, and viral and non- viral vectors as well as other means known in the art). The method of delivery selected will depend at least on the oligomer chemistry, the cells to be treated and the location of the cells and will be apparent to the skilled artisan.
- antisense oligomers may be delivered using, e.g., methods involving liposome-mediated uptake, exosome-mediated uptake, lipid conjugates, polylysine-mediated uptake, nanoparticle-mediated uptake, and receptor-mediated endocytosis, as well as additional non-endocytic modes of delivery, such as microinjection, permeabilization (e.g., streptolysin-O permeabilization, anionic peptide permeabilization), electroporation, and various non-invasive non-endocytic methods of delivery that are known in the art (refer to Dokka and Rojanasakul, Advanced Drug Delivery Reviews 44, 35-49, incorporated by reference in its entirety).
- the antisense oligomers may be administered in any convenient vehicle or carrier which is physiologically and/or pharmaceutically acceptable.
- a composition may include any of a variety of standard pharmaceutically acceptable carriers employed by those of ordinary skill in the art. Examples include, but are not limited to, saline, phosphate-buffered saline (PBS), water, aqueous ethanol, emulsions, such as oil/water emulsions or triglyceride emulsions, tablets, and capsules.
- PBS phosphate-buffered saline
- emulsions such as oil/water emulsions or triglyceride emulsions, tablets, and capsules.
- the choice of a suitable physiologically acceptable carrier will vary dependent upon the chosen mode of administration.
- “Pharmaceutically acceptable carrier” is intended to include any and all solvents, dispersion media, coatings, antibacterial and antifungal agents, isotonic and absorption delaying agents, and the like, compatible with pharmaceutical administration.
- the use of such media and agents for pharmaceutically active substances is well known in the art. Except insofar as any conventional media or agent is incompatible with the active compound, use thereof in the compositions is contemplated. Supplementary active compounds can also be incorporated into the compositions.
- the compounds (e.g., antisense oligomers) of the present disclosure may generally be utilized as the free acid or free base. Alternatively, the compounds of this disclosure may be used in the form of acid or base addition salts.
- Acid addition salts of the free amino compounds of the present disclosure may be prepared by methods well known in the art and may be formed from organic and inorganic acids.
- Suitable organic acids include maleic, fumaric, benzoic, ascorbic, succinic, methanesulfonic, acetic, trifluoroacetic, oxalic, propionic, tartaric, salicylic, citric, gluconic, lactic, mandelic, cinnamic, aspartic, stearic, palmitic, glycolic, glutamic, and benzenesulfonic acids.
- Suitable inorganic acids include hydrochloric, hydrobromic, sulfuric, phosphoric, and nitric acids.
- Base addition salts included those salts that form with the carboxylate anion and include salts formed with organic and inorganic cations such as those chosen from the alkali and alkaline earth metals (for example, lithium, sodium, potassium, magnesium, barium and calcium), as well as the ammonium ion and substituted derivatives thereof (for example, dibenzylammonium, benzylammonium, 2-hydroxyethylammonium, and the like).
- the term “pharmaceutically acceptable salt” is intended to encompass any and all acceptable salt forms.
- prodrugs are also included within the context of this disclosure. Prodrugs are any covalently bonded carriers that release a compound in vivo when such prodrug is administered to a patient.
- esters may be employed, such as methyl esters, ethyl esters, and the like.
- liposomes may be employed to facilitate uptake of the antisense oligomer into cells (see, e.g., Williams, S.A., Leukemia 10(12):1980-1989, 1996; Lappalainen et al., Antiviral Res.23:119, 1994; Uhlmann et al., antisense oligomers: a new therapeutic principle, Chemical Reviews, Volume 90, No.4, 25 pages 544-584, 1990; Gregoriadis, G., Chapter 14, Liposomes, Drug Carriers in Biology and Medicine, pp.287-341, Academic Press, 1979).
- Hydrogels may also be used as vehicles for antisense oligomer administration, for example, as described in WO 93/01286.
- the oligomers may be administered in microspheres or microparticles. (See, e.g., Wu, G.Y. and Wu, C.H., J. Biol. Chem.262:4429- 4432, 301987).
- the use of gas-filled microbubbles complexed with the antisense oligomers can enhance delivery to target tissues, as described in US Patent No.6,245,747.
- Sustained-release compositions may also be used. These may include semipermeable polymeric matrices in the form of shaped articles such as films or microcapsules.
- the antisense oligomer is administered to a mammalian subject, e.g., human or domestic animal, exhibiting the symptoms of a kidney disease, in a suitable pharmaceutical carrier.
- the subject is a human subject, e.g., a patient diagnosed as having a centronuclear myopathy.
- the antisense oligomer is contained in a pharmaceutically acceptable carrier and is delivered orally.
- the oligomer is contained in a pharmaceutically acceptable carrier and is delivered intravenously (i.v.).
- the antisense compound is administered in an amount and manner effective to result in a peak blood concentration of at least 200-400 nM antisense oligomer.
- one or more doses of antisense oligomer are administered, generally at regular intervals, for a period of about one to two weeks.
- Doses for oral administration are from about 1- 1000 mg oligomer per 70 kg. In some cases, doses of greater than 1000 mg oligomer/patient may be necessary. For i.v. administration, doses are from about 0.5 mg to 1000 mg oligomer per 70 kg.
- the antisense oligomer may be administered at regular intervals for a short time period, e.g., daily for two weeks or less.
- the oligomer is administered intermittently over a longer period of time. Administration may be followed by, or concurrent with, administration of an antibiotic or other therapeutic treatment.
- the treatment regimen may be adjusted (dose, frequency, route, etc.) as indicated, based on the results of immunoassays, other biochemical tests, and physiological examination of the subject under treatment.
- the method is an in vitro method. In certain other embodiments, the method is an in vivo method.
- the host cell is a mammalian cell. In certain embodiments, the host cell is a non-human primate cell. In certain embodiments, the host cell is a human cell. [0701] In certain embodiments, the host cell is a naturally occurring cell.
- the host cell is an engineered cell.
- the antisense oligomer is administered to a mammalian subject, e.g., a human or a laboratory or domestic animal, in a suitable pharmaceutical carrier.
- the antisense oligomer is administered to a mammalian subject, e.g., a human or laboratory or domestic animal, together with an additional agent.
- the antisense oligomer and the additional agent can be administered simultaneously or sequentially, via the same or different routes and/or sites of administration.
- the antisense oligomer and the additional agent can be co-formulated and administered together.
- the antisense oligomer is administered in an amount and manner effective to result in a peak blood concentration of at least 200 nM antisense oligomer. In one embodiment, the antisense oligomer is administered in an amount and manner effective to result in a peak plasma concentration of at least 200 nM antisense oligomer. In one embodiment, the antisense oligomer is administered in an amount and manner effective to result in a peak serum concentration of at least 200 nM antisense oligomer.
- the antisense oligomer is administered in an amount and manner effective to result in a peak blood concentration of at least 400 nM antisense oligomer. In one embodiment, the antisense oligomer is administered in an amount and manner effective to result in a peak plasma concentration of at least 400 nM antisense oligomer. In one embodiment, the antisense oligomer is administered in an amount and manner effective to result in a peak serum concentration of at least 400 nM antisense oligomer. [0710] Typically, one or more doses of the antisense oligomer are administered, generally at regular intervals, for a period of about one to two weeks.
- Doses for oral administration are from about 0.01-15 mg antisense oligomer per kg body weight. In some cases, doses of greater than 15 mg antisense oligomer /kg may be necessary. For i.v. administration, doses are from about 0.005 mg to 15 mg antisense oligomer per kg body weight.
- the antisense oligomer may be administered at regular intervals for a short time period, e.g., daily for two weeks or less. However, in some cases, the antisense oligomer is administered intermittently over a longer period of time. Administration may be followed by or accompanied by administration of an antibiotic or other therapeutic treatment.
- the treatment regimen may be adjusted (dose, frequency, route, etc.) as indicated, based on the results of immunoassays, other biochemical tests, and physiological examination of the subject under treatment.
- An effective in vivo treatment regimen using the antisense oligomer may vary according to the duration, dose, frequency, and route of administration, as well as the condition of the subject under treatment (i.e., prophylactic administration versus administration in response to localized or systemic infection). Accordingly, such in vivo therapy will often require monitoring by tests under treatment, and corresponding adjustments in the dose or treatment regimen, to achieve an optimal therapeutic outcome.
- the antisense oligomer is actively taken up by mammalian cells.
- the antisense oligomer can be conjugated to a transport moiety (e.g., transport peptide) as described herein to facilitate such uptake.
- a transport moiety e.g., transport peptide
- An antisense oligomer comprising a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of human dynamin 2 (DNM2) gene represented by SEQ ID NO: 1, wherein the target region is within an exon internal region, an intron internal region, an intron-exon junction, or an exon-intron junction of the human DNM2 gene pre-mRNA.
- DNM2 human dynamin 2
- exon 2 SEQ ID NO: 2
- exon 6 SEQ ID NO: 3
- exon 7 SEQ ID NO: 4
- exon 8 SEQ ID NO: 5
- exon 9 SEQ ID NO: 6
- exon 12 SEQ ID NO: 7
- exon 13 SEQ ID NO: 8
- exon 16 SEQ ID NO: 9
- exon 19 SEQ ID NO: 10
- exon 2 SEQ ID NO: 2
- exon 9 SEQ ID NO: 6
- exon 12 SEQ ID NO: 7
- exon 13 SEQ ID NO: 8
- Embodiment 5 The antisense oligomer of embodiment 1 or 4, wherein the target region is within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, and exon 13/intron 13 junction.
- Embodiment 6 Embodiment 6.
- Embodiment 8 The antisense oligomer of embodiment 7, wherein the targeting sequence is selected from SEQ ID NOs: 67-326.
- Embodiment 9. The antisense oligomer of any one of embodiments 1-6, wherein the target region is within the exon internal region of exon 2 (SEQ ID NO: 2), the intron 1/exon 2 junction, the exon 2/intron 2 junction, or the intron internal region of intron 2 (SEQ ID NO: 11) of the human DNM2 gene pre-mRNA.
- Embodiment 10 Embodiment 10.
- Embodiment 11 The antisense oligomer of embodiment 10, wherein the target region is a region within SEQ ID NO: 13.
- Embodiment 13 The antisense oligomer of embodiment 12, wherein the target region is a region within SEQ ID NO: 14.
- Embodiment 15 The antisense oligomer of embodiment 14, wherein the target region is a region within SEQ ID NO: 15.
- Embodiment 17 The antisense oligomer of embodiment 16, wherein the target region is a region within SEQ ID NO: 16.
- Embodiment 19 The antisense oligomer of embodiment 18, wherein the target region is a region within SEQ ID NO: 17.
- Embodiment 20 Embodiment 20.
- Embodiment 21 The antisense oligomer of embodiment 20, wherein the target region is a region within SEQ ID NO: 18.
- Embodiment 22 Embodiment 22.
- Embodiment 23 The antisense oligomer of embodiment 22, wherein the target region is a region within SEQ ID NO: 19.
- Embodiment 24 Embodiment 24.
- Embodiment 25 The antisense oligomer of embodiment 24, wherein the target region is a region within SEQ ID NO: 20.
- Embodiment 26 Embodiment 26.
- Embodiment 27 The antisense oligomer of embodiment 26, wherein the target region is a region within SEQ ID NO: 21.
- Embodiment 28 Embodiment 28.
- Embodiment 29. The antisense oligomer of embodiment 28, wherein the target region is a region within SEQ ID NO: 22.
- Embodiment 30 Embodiment 30.
- Embodiment 31 The antisense oligomer of embodiment 30, wherein the target region is a region within SEQ ID NO: 23.
- Embodiment 32 Embodiment 32.
- Embodiment 33 The antisense oligomer of embodiment 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2- 23) , DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30).
- Embodiment 34 The antisense oligomer of any one of embodiments 1-3, 7, 9-11, or 32, wherein the target region is DNM2 H2A(+21+45) or DNM2 H2A(+23+47).
- Embodiment 35 The antisense oligomer of any one of embodiments 1-3, 7, 9, 12-13, or 32, wherein the target region is DNM2 H2A(+23+47) or DNM2 H2A(+27+51).
- Embodiment 36 Embodiment 36.
- Embodiment 37 The antisense oligomer of any one of embodiments 1-3, 7, 9, 14-15, or 32, wherein the target region is DNM2 H2A(+46+70) or DNM2 H2A(+48+72). [0751] Embodiment 37.
- Embodiment 38 Embodiment 38.
- Embodiment 39 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-21, 32, or 37, wherein the target region is DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), or DNM2 H2D(+6-19).
- Embodiment 40 Embodiment 40.
- Embodiment 41 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-23, 32, 37, or 39, wherein the target region is DNM2 H2D(+9-16), DNM2 H2D(+8-17), or DNM2 H2D(+7-18).
- Embodiment 41 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-23, 32, 35, 37, or 39-40, wherein the target region is DNM2 H2D(+8-17).
- Embodiment 42 Embodiment 42.
- Embodiment 43 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42, wherein the target region is DNM2 H2D(+4-21), DNM2 H2D(+3-22), or DNM2 H2D(+2-23).
- Embodiment 44 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42, wherein the target region is DNM2 H2D(+4-21), DNM2 H2D(+3-22), or DNM2 H2D(+2-23).
- Embodiment 45 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42-43, wherein the target region is DNM2 H2D(+3-22).
- Embodiment 45 The antisense oligomer of any one of embodiments 1, 6-7, 9, 28-29, or 32, wherein the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30).
- Embodiment 46 Embodiment 46.
- Embodiment 47 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15).
- Embodiment 48 Embodiment 48.
- Embodiment 49 The antisense oligomer of any one of embodiments 1-3, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+3+27).
- Embodiment 50 The antisense oligomer of any one of embodiments 1-3, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+15+37).
- Embodiment 51 Embodiment 51.
- Embodiment 52 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2D(+20-5).
- Embodiment 53 The antisense oligomer of any one of embodiments 1-32, wherein the targeting sequence is selected from SEQ ID NOs: 67-96.
- Embodiment 54 The antisense oligomer of any one of embodiments 1-32, wherein the targeting sequence is selected from SEQ ID NOs: 67-96.
- Embodiment 55 The antisense oligomer of any one of embodiments 1-3, 7, 9-11, 32, or 34, wherein the targeting sequence is selected from SEQ ID NOs: 71-72.
- Embodiment 56 The antisense oligomer of any one of embodiments 1-3, 7, 9, 12-13, 32, or 35, wherein the targeting sequence is selected from SEQ ID NOs: 72-73.
- Embodiment 57 The antisense oligomer of any one of embodiments 1, 7, or 32-33, wherein the targeting sequence is selected from SEQ ID NOs: 78-96.
- Embodiment 55 The antisense oligomer of any one of embodiments 1-3, 7, 9-11, 32, or 34, wherein the targeting sequence is selected from SEQ ID NOs: 71-72.
- Embodiment 56 The antisense oligomer of any one of embodiments 1-3, 7, 9, 12-13, 32, or 35, wherein the targeting sequence is selected from SEQ ID NOs: 72-73.
- Embodiment 60 The antisense oligomer of any one of embodiments 1-3, 7, 9, 14-15, 32, or 36, wherein the targeting sequence is selected from SEQ ID NOs: 75-76.
- Embodiment 58 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 32, or 37, wherein the targeting sequence is selected from SEQ ID NOs: 78-90.
- Embodiment 59 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-19, 32, or 37-38, wherein the targeting sequence is selected from SEQ ID NOs: 78-80.
- Embodiment 60 Embodiment 60.
- Embodiment 61 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-21, 32, 37, or 39, wherein the targeting sequence is selected from SEQ ID NOs: 81-85.
- Embodiment 61 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-23, 32, 37, or 39-40, wherein the targeting sequence is selected from SEQ ID NOs: 82-84.
- Embodiment 62 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-23, 32, 35, 37, or 39-41, wherein the targeting sequence is SEQ ID NO: 83.
- Embodiment 63 Embodiment 63.
- Embodiment 64 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42, wherein the targeting sequence is selected from SEQ ID NOs: 87-89.
- Embodiment 65 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42-44, wherein the targeting sequence is SEQ ID NO: 88.
- Embodiment 66 Embodiment 66.
- Embodiment 67 The antisense oligomer of any one of embodiments 1, 6-7, 9, 28-29, 32, or 45, wherein the targeting sequence is selected from SEQ ID NOs: 91-96.
- Embodiment 67 The antisense oligomer of any one of embodiments 1, 6-7, 9, 28-32, or 45-46, wherein the targeting sequence is selected from SEQ ID NOs: 91-95.
- Embodiment 68 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 32, or 47, wherein the targeting sequence is SEQ ID NO: 67.
- Embodiment 69 Embodiment 69.
- Embodiment 70 The antisense oligomer of any one of embodiments 1, 4-5, or 7, 9, 32, or 48, wherein the targeting sequence is SEQ ID NO: 68.
- Embodiment 70 The antisense oligomer of any one of embodiments 1-3, 7, 9, 32, or 49, wherein the targeting sequence is SEQ ID NO: 69.
- Embodiment 71 The antisense oligomer of any one of embodiments 1-3, 7, 9, 32, or 50, wherein the targeting sequence is SEQ ID NO: 70.
- Embodiment 72 The antisense oligomer of any one of embodiments 1-3, 7, 9, 32, or 51, wherein the targeting sequence is SEQ ID NO: 74.
- Embodiment 73 The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 32, or 52, wherein the targeting sequence is SEQ ID NO: 77.
- Embodiment 74 The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 9 (SEQ ID NO: 6), the intron 8/exon 9 junction, or the exon 9/intron 9 junction of the human DNM2 gene pre-mRNA.
- Embodiment 75 Embodiment 75.
- Embodiment 76. The antisense oligomer of embodiment 75, wherein the target region is a region within SEQ ID NO: 24.
- Embodiment 77 Embodiment 77.
- Embodiment 78. The antisense oligomer of embodiment 77, wherein the target region is a region within SEQ ID NO: 25.
- Embodiment 80 The antisense oligomer of embodiment 79, wherein the target region is a region within SEQ ID NO: 26.
- Embodiment 81 Embodiment 81.
- Embodiment 82 The antisense oligomer of embodiment 81, wherein the target region is a region within SEQ ID NO: 27.
- Embodiment 83 Embodiment 83.
- Embodiment 84 The antisense oligomer of embodiment 83, wherein the target region is a region within SEQ ID NO: 28.
- Embodiment 85 Embodiment 85.
- Embodiment 86. The antisense oligomer of embodiment 85, wherein the target region is a region within SEQ ID NO: 29.
- Embodiment 87 Embodiment 87.
- Embodiment 88. The antisense oligomer of embodiment 87, wherein the target region is a region within SEQ ID NO: 30.
- Embodiment 89 Embodiment 89.
- Embodiment 90 The antisense oligomer of embodiment 89, wherein the target region is a region within SEQ ID NO: 31.
- Embodiment 91 Embodiment 91.
- Embodiment 92 The antisense oligomer of embodiment 91, wherein the target region is a region within SEQ ID NO: 32.
- Embodiment 93 Embodiment 93.
- Embodiment 94 The antisense oligomer of embodiment 93, wherein the target region is a region within SEQ ID NO: 33.
- Embodiment 95 Embodiment 95.
- Embodiment 96 The antisense oligomer of embodiment 95, wherein the target region is a region within SEQ ID NO: 34.
- Embodiment 97 Embodiment 97.
- Embodiment 98. The antisense oligomer of embodiment 97, wherein the target region is a region within SEQ ID NO: 35. [0813] Embodiment 99.
- Embodiment 100 The antisense oligomer of embodiment 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+43+67), DNM2 H9A(+42+66), DNM2 H9A(+44+68), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM
- Embodiment 101 The antisense oligomer of any one of embodiments 1-5, 7, 74-76, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+36+60) or DNM2 H9A(+37+61).
- Embodiment 102 Embodiment 102.
- Embodiment 103 The antisense oligomer of any one of embodiments 1-3, 7, 74, 77-80, 99, or 102, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), or DNM2 H9A(+44+68).
- Embodiment 104 Embodiment 104.
- Embodiment 105 The antisense oligomer of any one of embodiments 1-3, 7, 74, 77-80, 99, or 102-103, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+67).
- Embodiment 106 The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 84, 99, or 105, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), or DNM2 H9D(+20-5).
- Embodiment 107 Embodiment 107.
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), or DNM2 H9D(+15- 10).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), or DNM2 H9D(+10-15).
- Embodiment 110 The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 92, 99, or 109, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6- 17), DNM2 H9D(+5-17), or DNM2 H9D(+5-20).
- Embodiment 111 The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 94, 99, or 109-110, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), or DNM2 H9D(+7-18).
- Embodiment 113 Embodiment 113.
- Embodiment 114 The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 96, 99, or 109-112, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17).
- Embodiment 115 Embodiment 115.
- Embodiment 116 The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13).
- Embodiment 116 The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-5+20).
- Embodiment 117 The antisense oligomer of any one of embodiments 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+1+25).
- Embodiment 118 Embodiment 118.
- Embodiment 119 The antisense oligomer of any one of embodiments 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+6+30).
- Embodiment 119 The antisense oligomer of any one of embodiments 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40).
- Embodiment 120 The antisense oligomer of any one of embodiments 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+25+49).
- Embodiment 121 Embodiment 121.
- Embodiment 122 The antisense oligomer of any one of embodiments 1, 7, or 99-100, wherein the targeting sequence is selected from SEQ ID NOs: 162-166, 172, 174, 179-194, 199- 200, 205, 207, and 209.
- Embodiment 123 The antisense oligomer of any one of embodiments 1-5, 7, 74-76, 99, or 101, wherein the targeting sequence is selected from SEQ ID NOs: 164-165.
- Embodiment 124 Embodiment 124.
- Embodiment 125 The antisense oligomer of any one of embodiments 1-3, 7, 74, 77-80, 99, or 102-103, wherein the targeting sequence is selected from SEQ ID NOs: 167-174.
- Embodiment 126 The antisense oligomer of any one of embodiments 1-3, 7, 74, 77-80, 99, or 102-104, wherein the targeting sequence is SEQ ID NO: 172.
- Embodiment 127 Embodiment 127.
- Embodiment 129 The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 82, 85-86, 99, or 107, wherein the targeting sequence is selected from SEQ ID NOs: 184-188.
- Embodiment 130 The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 82, 87-88, 99, or 108, wherein the targeting sequence is selected from SEQ ID NOs: 189-193.
- Embodiment 131 The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 90, 99, or 109, wherein the targeting sequence is selected from SEQ ID NOs: 194-209.
- Embodiment 132 Embodiment 132.
- Embodiment 135. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 96, 99, or 109-113, wherein the targeting sequence is SEQ ID NO: 199.
- Embodiment 136. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 90, 97-99, 109, or 114, wherein the targeting sequence is selected from SEQ ID NOs: 202-209.
- Embodiment 137. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 99, or 115, wherein the targeting sequence is SEQ ID NO: 158.
- Embodiment 140 The antisense oligomer of any one of embodiments 1-3, 7, 74, 99, or 118, wherein the targeting sequence is SEQ ID NO: 161.
- Embodiment 141 The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 99, or 116, wherein the targeting sequence is SEQ ID NO: 159.
- Embodiment 139 The antisense oligomer of any one of embodiments 1-3, 7, 74, 99, or 117, wherein the targeting sequence is SEQ ID NO: 160.
- Embodiment 140 The antisense oligomer of any one of embodiments 1-3, 7, 74, 99, or 118, wherein the targeting sequence is SEQ ID NO: 161.
- Embodiment 141 Embodiment 141.
- Embodiment 142 The antisense oligomer of any one of embodiments 1-3, 7, 74, 99, or 119, wherein the targeting sequence is SEQ ID NO: 162.
- Embodiment 142 The antisense oligomer of any one of embodiments 1-3, 7, 74, 99, or 120, wherein the targeting sequence is SEQ ID NO: 163.
- Embodiment 143 The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 12 (SEQ ID NO: 7), the intron 11/exon 12 junction, or the exon 12/intron 12 junction of the human DNM2 gene pre-mRNA.
- Embodiment 144 Embodiment 144.
- Embodiment 145 The antisense oligomer of embodiment 144, wherein the target region is a region within SEQ ID NO: 36.
- Embodiment 146 Embodiment 146.
- Embodiment 147. The antisense oligomer of embodiment 146, wherein the target region is a region within SEQ ID NO: 37.
- Embodiment 148 Embodiment 148.
- Embodiment 149 The antisense oligomer of embodiment 148, wherein the target region is a region within SEQ ID NO: 38.
- Embodiment 150 Embodiment 150.
- Embodiment 151 The antisense oligomer of embodiment 150, wherein the target region is a region within SEQ ID NO: 39.
- Embodiment 152 Embodiment 152.
- Embodiment 153 The antisense oligomer of embodiment 152, wherein the target region is a region within SEQ ID NO: 40.
- Embodiment 154 Embodiment 154.
- Embodiment 155 The antisense oligomer of embodiment 154, wherein the target region is a region within SEQ ID NO: 41.
- Embodiment 156 Embodiment 156.
- Embodiment 157 The antisense oligomer of embodiment 156, wherein the target region is a region within SEQ ID NO: 42.
- Embodiment 158 Embodiment 158.
- Embodiment 159 The antisense oligomer of embodiment 158, wherein the target region is a region within SEQ ID NO: 43.
- Embodiment 160 Embodiment 160.
- Embodiment 161 The antisense oligomer of embodiment 160, wherein the target region is a region within SEQ ID NO: 44.
- Embodiment 163 The antisense oligomer of embodiment 162, wherein the target region is a region within SEQ ID NO: 45.
- Embodiment 164 Embodiment 164.
- Embodiment 165 The antisense oligomer of any one of embodiment 164, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+49), DNM2 H12A(+29+53), DNM2 H12A(+30+54), DNM2 H12A(+
- Embodiment 166 The antisense oligomer of any one of embodiments 1-3, 7, 143-145, or 164, wherein the target region is DNM2 H12A(+4+28) or DNM2 H12A(+7+31).
- Embodiment 167 Embodiment 167.
- Embodiment 168 Embodiment 168.
- Embodiment 169 Embodiment 169.
- Embodiment 170 Embodiment 170.
- Embodiment 171 is DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), or DNM2 H12A(+18+42).
- Embodiment 172 The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 157, 164, or 171, wherein the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), or DNM2 H12A(+29+53).
- Embodiment 173 The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 155, 158-159, 164, or 171, wherein the target region is DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54).
- Embodiment 174 Embodiment 174.
- Embodiment 175. The antisense oligomer of any one of embodiments 1-3, 7, 143, 160- 161, or 164, wherein the target region is DNM2 H12A(+36+60) or DNM2 H12A(+37+61).
- Embodiment 177 The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, or 162-164, wherein the target region is DNM2 H12D(+22-3) or DNM2 H12D(+20-5).
- Embodiment 177 The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, or 164, wherein the target region is DNM2 H12A(-13+12).
- Embodiment 178 The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, or 164, wherein the target region is DNM2 H12A(-3+22).
- Embodiment 179 Embodiment 179.
- Embodiment 180 The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, or 164, wherein the target region is DNM2 H12D(+15-10).
- Embodiment 181. The antisense oligomer of any one of embodiments 1-5, 7, or 143-164, wherein the targeting sequence is selected from SEQ ID NOs: 210-243.
- Embodiment 183. The antisense oligomer of any one of embodiments 1-3, 7, 143-145, 164, or 166, wherein the targeting sequence is selected from SEQ ID NOs: 212-213.
- Embodiment 187 The antisense oligomer of any one of embodiments 1-3, 7, 143, 146- 153, 164, 167-170, wherein the targeting sequence is selected from SEQ ID NOs: 218-222.
- Embodiment 188 The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 155, 164, or 171, wherein the targeting sequence is selected from SEQ ID NOs: 225-237.
- Embodiment 189 The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 157, 164, or 171-172, wherein the targeting sequence is selected from SEQ ID NOs: 225-235.
- Embodiment 190 Embodiment 190.
- Embodiment 194 The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, 164, or 177, wherein the targeting sequence is SEQ ID NO: 210.
- Embodiment 195 The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, 164, or 178, wherein the targeting sequence is SEQ ID NO: 211.
- Embodiment 196 Embodiment 196.
- Embodiment 197 The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, 164, or 180, wherein the targeting sequence is SEQ ID NO: 243.
- Embodiment 198 The antisense oligomer of any one of embodiments 1-6, wherein the target region is within the exon internal region of exon 13 (SEQ ID NO: 8), the intron 12/exon 13 junction, the exon 13/intron 13 junction, or the intron internal region of intron 12 (SEQ ID NO: 12).
- Embodiment 199 The antisense oligomer of any one of embodiments 1, 4-6, or 198, wherein the target region of the human DNM2 gene pre-mRNA is within the 27 th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12 th nucleotide of exon 13 measured from the 5′ end of exon 13.
- Embodiment 200 The antisense oligomer of embodiment 199, wherein the target region is a region within SEQ ID NO: 46.
- Embodiment 201 Embodiment 201.
- Embodiment 202 The antisense oligomer of embodiment 201, wherein the target region is a region within SEQ ID NO: 47.
- Embodiment 203 Embodiment 203.
- Embodiment 204 The antisense oligomer of embodiment 203, wherein the target region is a region within SEQ ID NO: 48.
- Embodiment 205 is a region within SEQ ID NO: 48.
- Embodiment 206 The antisense oligomer of embodiment 205, wherein the target region is a region within SEQ ID NO: 49.
- Embodiment 207 Embodiment 207.
- SEQ ID NO: 12 11 th nucleotide of intron 12
- SEQ ID NO: 8 the antisense oligomer of embodiment 207, wherein the target region is a region within SEQ ID NO: 50.
- Embodiment 210 The antisense oligomer of embodiment 209, wherein the target region is a region within SEQ ID NO: 51.
- Embodiment 211 Embodiment 211.
- Embodiment 212 The antisense oligomer of embodiment 211, wherein the target region is a region within SEQ ID NO: 52.
- Embodiment 214 The antisense oligomer of embodiment 213, wherein the target region is a region within SEQ ID NO: 53.
- Embodiment 216 The antisense oligomer of embodiment 215, wherein the target region is a region within SEQ ID NO: 54.
- Embodiment 217 Embodiment 217.
- Embodiment 218. The antisense oligomer of embodiment 217, wherein the target region is a region within SEQ ID NO: 55.
- Embodiment 220 The antisense oligomer of embodiment 219, wherein the target region is a region within SEQ ID NO: 56.
- Embodiment 224 The antisense oligomer of embodiment 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(- 25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(- 11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A
- Embodiment 225 The antisense oligomer of any one of embodiments 1, 4-7, 198-200, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(- 18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2
- Embodiment 226 The antisense oligomer of any one of embodiments 1, 4-7, 198-202, 223, or 225, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27- 3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), or DNM2 H13A(-23+2).
- Embodiment 227 Embodiment 227.
- Embodiment 228 Embodiment 228.
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12).
- Embodiment 230 Embodiment 230.
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), or DNM2 H13A(-7+18).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(-1+24).
- the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), or DNM2 H13A(-4+21).
- Embodiment 23 The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 218, 223, or 233, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), or DNM2 H13A(+5+28).
- Embodiment 235 Embodiment 235.
- Embodiment 236 Embodiment 236.
- Embodiment 237 Embodiment 237.
- Embodiment 240 The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13D(+7-18).
- Embodiment 241 The antisense oligomer of any one of embodiments 1-7, 198-221, or 223, wherein the targeting sequence is selected from SEQ ID NOs: 244-287.
- Embodiment 242. The antisense oligomer of any one of embodiments 1, 7, or 223-224, wherein the targeting sequence is selected from SEQ ID NOs: 244-278.
- Embodiment 243 Embodiment 243.
- Embodiment 244. The antisense oligomer of any one of embodiments 1, 4-7, 198-202, 223, or 225-226, wherein the targeting sequence is selected from SEQ ID NOs: 244-248.
- Embodiment 245. The antisense oligomer of any one of embodiments 1, 4-7, 198-200, 203-204, 223, 225, or 227, wherein the targeting sequence is selected from SEQ ID NOs: 249- 253.
- Embodiment 247 The antisense oligomer of any one of embodiments 1, 4-7, 198, 207- 208, 223, or 229, wherein the targeting sequence is selected from SEQ ID NOs: 259-269.
- Embodiment 248 The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 207-210, 223, or 229-230, wherein the targeting sequence is selected from SEQ ID NOs: 259- 263.
- Embodiment 250 The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 207-214, 223, or 229-232, wherein the targeting sequence is selected from SEQ ID NOs: 262- 265.
- Embodiment 253 The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 216, 219-220, 223, 233, or 235, wherein the targeting sequence is selected from SEQ ID NOs: 280-284.
- Embodiment 254 The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 223, or 228-236, wherein the targeting sequence is selected from SEQ ID NOs: 272-281.
- Embodiment 255 The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 223, or 228-237, wherein the targeting sequence is SEQ ID NO: 278.
- Embodiment 256 The antisense oligomer of any one of embodiments 1-3, 7, 198, 223, or 238, wherein the targeting sequence is SEQ ID NO: 285.
- Embodiment 257 The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 223, or 239, wherein the targeting sequence is SEQ ID NO: 286.
- Embodiment 258 The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 223, or 239, wherein the targeting sequence is SEQ ID NO: 286.
- Embodiment 261. The antisense oligomer of embodiment 260, wherein the target region is a region within SEQ ID NO: 58.
- Embodiment 263 The antisense oligomer of embodiment 262, wherein the target region is a region within SEQ ID NO: 59.
- Embodiment 264 Embodiment 264.
- Embodiment 265. The antisense oligomer of embodiment 264, wherein the target region is a region within SEQ ID NO: 60.
- Embodiment 266 Embodiment 266.
- Embodiment 267 The antisense oligomer of embodiment 266, wherein the target region is a region within SEQ ID NO: 61.
- Embodiment 268 Embodiment 268.
- Embodiment 269. The antisense oligomer of embodiment 268, wherein the target region is a region within SEQ ID NO: 62.
- Embodiment 270 Embodiment 270.
- Embodiment 27 The antisense oligomer of any one of embodiments 1-3, 7, 260-261, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+10+3) or DNM2 H6A(+13+37).
- Embodiment 272. The antisense oligomer of any one of embodiments 1-3, 7, 259, 262- 263, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+32+56) or DNM2 H6A(+35+59).
- Embodiment 274. The antisense oligomer of any one of embodiments 1-3, 7, 259, 266- 267, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+130+154) or DNM2 H6A(+132+156).
- Embodiment 275 Embodiment 275.
- Embodiment 276 The antisense oligomer of any one of embodiments 1, 4-5, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+15-10) or DNM2 H6D (+17-8).
- Embodiment 276 The antisense oligomer of any one of embodiments 1, 4-5, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5).
- Embodiment 277 Embodiment 277.
- Embodiment 278. The antisense oligomer of any one of embodiments 1, 4-5, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-2+23).
- Embodiment 280 The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+23+47).
- Embodiment 281. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+44+68).
- Embodiment 283. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+88+112).
- Embodiment 284. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+96+119).
- Embodiment 286 The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+105+129).
- Embodiment 286 The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+121+145).
- Embodiment 287 The antisense oligomer of any one of embodiments 1, 4-5, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+6-19).
- Embodiment 288 The antisense oligomer of any one of embodiments 259-287, wherein the targeting sequence is selected from SEQ ID NOs: 97-118.
- Embodiment 289. The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 7 (SEQ ID NO: 4), the intron 6/exon 7 junction, or the exon 7/intron 7 junction.
- Embodiment 290 Embodiment 290.
- Embodiment 291. The antisense oligomer of embodiment 290, wherein the target region is a region within SEQ ID NO: 63.
- Embodiment 293 The antisense oligomer of embodiment 292, wherein the target region is a region within SEQ ID NO: 64.
- Embodiment 294 Embodiment 294.
- Embodiment 295. The antisense oligomer of any one of embodiments 1-3, 7, 289-291, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+72+96) or DNM2 H7A(+74+98).
- Embodiment 296. The antisense oligomer of any one of embodiments 1, 4-5, 7, 289, or 292-294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+8-16) or DNM2 H7D(+9-16).
- Embodiment 298. The antisense oligomer of any one of embodiments 1, 4-5, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-5+20).
- Embodiment 301 The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+6+30).
- Embodiment 301 The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+11+35).
- Embodiment 302. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+22+46).
- Embodiment 303 Embodiment 303.
- Embodiment 304 The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+29+53).
- Embodiment 304 The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+41+65).
- Embodiment 305 The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+51+75).
- Embodiment 307. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+62+86).
- Embodiment 308 The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+81+105).
- Embodiment 310 The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+91+115).
- Embodiment 310 The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+110+134).
- Embodiment 311 The antisense oligomer of any one of embodiments 1, 4-5, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+21-4).
- Embodiment 313 The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 8 (SEQ ID NO: 5), the intron 7/exon 8 junction, or the exon 8/intron 8 junction.
- Embodiment 315 The antisense oligomer of embodiment 314, wherein the target region is a region within SEQ ID NO: 65.
- Embodiment 316 Embodiment 316.
- Embodiment 317 The antisense oligomer of embodiment 316, wherein the target region is a region within SEQ ID NO: 66.
- Embodiment 318 Embodiment 318.
- Embodiment 322. The antisense oligomer of any one of embodiments 1, 4-5, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-14+11).
- Embodiment 324 Embodiment 324.
- Embodiment 325 The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+5+29).
- Embodiment 326 The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+36+60).
- Embodiment 327 Embodiment 327.
- Embodiment 331 The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+65+88).
- Embodiment 332 The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+83+107).
- Embodiment 333 Embodiment 333.
- Embodiment 334 The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+96+119).
- Embodiment 335 The antisense oligomer of any one of embodiments 1, 4-5, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+18-7).
- Embodiment 336 The antisense oligomer of any one of embodiments 1, 4-5, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+18-7).
- Embodiment 337 The antisense oligomer of any one of embodiments 313-336, wherein the targeting sequence is selected from SEQ ID NOs: 138-157.
- Embodiment 338 The antisense oligomer of any one of embodiments 1-5 wherein the target region is within the exon internal region of exon 16 (SEQ ID NO: 9), the intron 15/exon 16 junction, or the exon 16/intron 16 junction.
- Embodiment 339 Embodiment 339.
- Embodiment 340 The antisense oligomer of any one of embodiments 1, 4-5, 7, or 338- 339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3).
- Embodiment 341. The antisense oligomer of any one of embodiments 1, 4-5, 7, or 338- 339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-13+12).
- Embodiment 343 The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+6+30).
- Embodiment 344 The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+15+39).
- Embodiment 345 Embodiment 345.
- Embodiment 346 The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+26+50).
- Embodiment 346 The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+36+60).
- Embodiment 347 The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+45+69).
- Embodiment 348 Embodiment 348.
- Embodiment 349 The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+55+79).
- Embodiment 349 The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+66+90).
- Embodiment 350 The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+75+99).
- Embodiment 352 The antisense oligomer of any one of embodiments 1, 4-5, 7, or 338- 339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16D(+19-6).
- Embodiment 353. The antisense oligomer of any one of embodiments 1, 4-5, 7, or 338- 339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16D(+11-14).
- Embodiment 355. The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 19 (SEQ ID NO: 9), the intron 18/exon 19 junction, or the exon 18/intron 18 junction.
- Embodiment 356 Embodiment 356.
- Embodiment 357 The antisense oligomer of any one of embodiments 1, 4-5, or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14).
- Embodiment 358 The antisense oligomer of any one of embodiments 1, 4-5, or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-4+21).
- Embodiment 360 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+7+31).
- Embodiment 361. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+16+40).
- Embodiment 363 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+37+61).
- Embodiment 364. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+47+71).
- Embodiment 365 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+56+80).
- Embodiment 366 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+66+90).
- Embodiment 367 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+76+100).
- Embodiment 368 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+86+110).
- Embodiment 372 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+127+151).
- Embodiment 373 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+137+161).
- Embodiment 374 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+147+171).
- Embodiment 375 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+157+181).
- Embodiment 376 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+166+189).
- Embodiment 377 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+191+215).
- Embodiment 378 The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+200+224).
- Embodiment 379 The antisense oligomer of any one of embodiments 1, 4-5, or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+23-2).
- Embodiment 380 The antisense oligomer of any one of embodiments 1, 4-5, or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+17-8).
- Embodiment 38 The antisense oligomer of any one of embodiments 1, 4-5, or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+10-15).
- Embodiment 382. The antisense oligomer of any one of embodiments 355-381, wherein the targeting sequence is selected from SEQ ID NOs: 302-326.
- Embodiment 385 Embodiment 385.
- Embodiment 386. The antisense oligomer of any one of embodiments 1-385, wherein the antisense oligomer is covalently linked to a cell-penetrating peptide.
- Embodiment 387 Embodiment 387.
- the antisense oligomer of embodiment 386 wherein the antisense oligomer is covalently linked to the cell-penetrating peptide by a linker selected from a direct bond, a glycine amino acid, a proline amino acid, glutamic acid amino acid, or an isoglutamine amino acid.
- a linker selected from a direct bond, a glycine amino acid, a proline amino acid, glutamic acid amino acid, or an isoglutamine amino acid.
- Embodiment 389 The antisense oligomer of any one of embodiments 1-382 having a structure of Formula (I): or a pharmaceutically acceptable salt thereof, wherein: R 5 is -C(O)(O-alkyl)x-OH, wherein x is 3-10 and each alkyl group is, independently at each occurrence, C 2-6 -alkyl, or R 5 is selected from H, -C(O)C 1-6 -alkyl, trityl, monomethoxytrityl, -(C 1-6 -alkyl)-R 6 , -(C 1-6 - heteroalkyl)-R 6 , -C 6-10 -aryl-R 6 , 5-10 membered heteroaryl-R 6 , -C(O)O-(C 1-6 -alkyl)-R 6 , - C(O)O-aryl-R 6 , -C(O)O-(5-10 membered heteroaryl
- Embodiment 390 The antisense oligomer of or antisense oligomer embodiment 389, wherein E′ is selected from -H, -C 1-6 -alkyl, -C(O)C 1-6 -alkyl, benzoyl, stearoyl, trityl, monomethoxytrityl, dimethoxytrityl, trimethoxytrityl, and .
- Embodiment 391. The antisense oligomer of embodiment 389 or 390, wherein E′ is selected from -H, -C(O)CH3, benzoyl, stearoyl, trityl, 4-methoxytrityl, and .
- Embodiment 392 The antisense oligomer of any one of embodiments 389-391, wherein A′ is selected from: .
- Embodiment 393. The antisense oligomer of any one of embodiments 389-392, wherein at least one of the following is true: [1107] Embodiment 394.
- Embodiment 395 Embodiment 395.
- Embodiment 394 The antisense oligomer of any one of embodiments 389-394, wherein A′ is: E′ is selected from H, -C(O)CH3, trityl, 4-methoxytrityl, benzoyl, and stearoyl.
- Embodiment 396 The antisense oligomer of any one of embodiments 389-395, wherein each R 1 is -N(CH 3 ) 2 .
- Embodiment 397 The antisense oligomer of any one of embodiments 389-396, wherein L is glycine, proline, or ⁇ -alanine.
- Embodiment 398 Embodiment 398.
- Embodiment 399 The antisense oligomer of any one of embodiments 389-397, wherein L is proline.
- Embodiment 400 The antisense oligomer of any one of embodiments 389-397, wherein L is ⁇ -alanine.
- Embodiment 401 Embodiment 401.
- Embodiment 403. The antisense oligomer of any one of embodiments 389-402, wherein G is -H or -C(O)CH 3 .
- Embodiment 404. The antisense oligomer of any one of embodiments 389-403, wherein G is -H.
- Embodiment 405. The antisense oligomer of any one of embodiments 389-403, wherein G is -C(O)CH 3 .
- Embodiment 408 The antisense oligomer of any one of embodiments 389-384, 396-397, 400-403, or 405-407, wherein the antisense oligomer is of Formula (III):
- Embodiment 409 The antisense oligomer of any one of embodiments 389-384, 396-397, 400-403, or 405-408, or a pharmaceutically acceptable salt thereof, wherein the antisense oligomer is of Formula (IV): (IV), wherein n is 11-28.
- Embodiment 410 The antisense oligomer of embodiment 408 or 409, or a pharmaceutically acceptable salt thereof, wherein each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 83.
- Embodiment 412 The antisense oligomer of embodiment 408 or 409, or a pharmaceutically acceptable salt thereof, wherein each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 172.
- Embodiment 413 The antisense oligomer of embodiment 408 or 409, or a pharmaceutically acceptable salt thereof, wherein each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 199. [1127] Embodiment 414.
- Embodiment 415 The antisense oligomer of embodiment 408 or 409, or a pharmaceutically acceptable salt thereof, wherein each R 2 , taken together, forms a targeting sequence having SEQ ID NO: 278.
- Embodiment 416 An antisense oligomer selected from: ,
- Embodiment 417 The antisense oligomer of embodiment 416 that is:
- Embodiment 418 The antisense oligomer of embodiment 416 that is:
- Embodiment 419 The antisense oligomer of embodiment 416 that is:
- Embodiment 420 The antisense oligomer of embodiment 416 that is:
- Embodiment 421 The antisense oligomer of embodiment 416 that is:
- Embodiment 422. The antisense oligomer of embodiment 416 that is:
- Embodiment 423 A pharmaceutical composition, comprising the antisense oligomer, or a pharmaceutically acceptable salt thereof, of any one of embodiments 1-422 and a pharmaceutically acceptable carrier.
- Embodiment 424 A method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein, comprising administering to a subject in need thereof a therapeutically effective amount of the antisense oligomer, or a pharmaceutically acceptable salt thereof, of any one of embodiments 1-422 or the pharmaceutical composition of embodiment 423.
- Embodiment 425 The method of embodiment 424, wherein the disease or disorder is a genetic neuromuscular disorder.
- Embodiment 426 The method of embodiment 424 or 425, wherein the disease or disorder is caused by at least one mutation in a gene selected from DNM2 gene, MTM1 gene, BIN1 gene, RYR1 gene, TTN gene, SPEG gene, CACNA1S gene, MTMR14 gene, ZAK gene, and CCDC78 gene.
- Embodiment 427 The method of any one of embodiments 424-426, wherein the disease or disorder is autosomal dominant, autosomal recessive, or X-linked.
- Embodiment 428 The method of any one of embodiments 424-427, wherein the disease or disorder is a centronuclear myopathy (CNM).
- CCM centronuclear myopathy
- Embodiment 429 The method of any one of embodiments 424-428, wherein the disease or disorder is myotubular myopathy (XLMTM or XLCNM), autosomal dominant congenital myopathy (ADCNM), autosomal recessive centronuclear myopathy (ARCNM), or Charcot-Marie- Tooth disease.
- XLMTM or XLCNM myotubular myopathy
- ADCNM autosomal dominant congenital myopathy
- ARCNM autosomal recessive centronuclear myopathy
- Charcot-Marie- Tooth disease Charcot-Marie- Tooth disease.
- antisense oligomers will induce exon skipping to create a premature stop codon (PTC) and thereby trigger nonsense-mediated decay to degrade DNM2 mRNA in a diseased cell. Reduction of DNM2 expression to normal or near- normal levels would then reduce disease-causing aggregation or increased DNM2 activity in the cell, thereby providing a clinical benefit to patients.
- Exemplary oligomers comprising a targeting sequence as set forth in Tables 5 and 6 were prepared as PMOs. As described below, these antisense oligomers were introduced into Rhabdomyosarcoma (RD) cells and analyzed for DNM2 expression. The knockdown data is shown in Tables 5 and 6.
- the PMOs used in this example are of Formula (V) below having the sequences indicated in Tables 5 and 6.
- Example 2 Macrowalk Screen [1146] Rhabdomyosarcoma (RD) cells were seeded at 15,000-20,000 cells per well in 96-well plates, in DMEM (Gibco, 10569010) with 10% FBS (Cytiva, SH30071.03) and treated a day later with a combination of 1 ⁇ M Endo-Porter (Gene tools) and 20 ⁇ M of each PMO diluted in DMEM + 10% FBS. Cells were harvested after 48 hours of treatment and RNA extracted (Quick RNA 96 kit, Zymo Research) (PBS- Vehicle/no treatment control, Negative control PMO 1, positive control PPMO 1).
- RT-PCR was performed using MaximaTM H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTimeTM Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to manufacturer’s instructions.
- PrimeTime qPCR Probe Assays are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems. The results of the macrowalk screen are shown in Table 5 and FIGs.1A-1E and FIGs.2A-2F.
- the PMOs used in this example are of Formula (V) above having the sequences indicated in Table 5.
- RT-PCR conditions were as follows: 1.Incubate at 25° C for 10 minutes. 2.Incubate at 50 °C for 15 minutes. 3.Incubate at 85 °C for 5 minutes. qPCR conditions were as follows: Table 5. Macrowalk screen- mRNA reduction of human DNM2 by PMOs in Rhabdomyosarcoma (RD) cells.
- RD Rhabdomyosarcoma
- Example 3 Microwalk Screen [1147] Rhabdomyosarcoma (RD) cells were prepared and treated with PMOs according to the protocol described in Example 2. The results of the microwalk screen are shown in Table 6 and FIGs.3A-3J. The PMOs used in this example are of Formula (V) above having the sequences indicated in Table 6. Table 6. Microwalk lead optimization screen – mRNA reduction of human DNM2 by PMOs in rhabdomyosarcoma (RD) cells. “B” represents an abasic subunit or guanine.
- Example 4 Dose response of PMOs in rhabdomyosarcoma (RD) cells.
- RD rhabdomyosarcoma
- RNA extracted Quick RNA 96 kit, Zymo Research
- PBS- Vehicle/no treatment control Negative control PMO 1, positive control PPMO 1
- RT-PCR was performed using MaximaTM H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTimeTM Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to the manufacturer’s instructions.
- PrimeTimeTM qPCR Probe Assays (IDT) are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems. The results of the PMO dose response are shown in FIGs.4A-4G.
- the PMOs used in this example are of Formula (V) above having the sequences disclosed in Table 7. Table 7.
- RT-PCR conditions were as follows: 1.Incubate at 25° C for 10 minutes. 2.Incubate at 50 °C for 15 minutes. 3.Incubate at 85 °C for 5 minutes. qPCR conditions were as follows: Example 5 – Exon-skipping induced by PPMOs targeting exon 2 and exon 9 in rhabdomyosarcoma (RD) cells.
- RD rhabdomyosarcoma
- Rhabdomyosarcoma (RD) cells were seeded at 15,000-20,000 cells per well in 96-well plates, in DMEM (Gibco, 10569010) with 10% FBS (Cytiva, SH30071.03) and treated a day later with a combination of 1 ⁇ M Endo-Porter (Gene tools) and 20 ⁇ M of each PPMO diluted in DMEM + 10% FBS. Cells were harvested after 48 hours of treatment and RNA extracted (Quick RNA 96 kit, Zymo Research).
- RT-PCR was performed using MaximaTM H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by a PCR utilizing PlatinumTM II Hot- Start PCR Master Mix (2X) (Fisher Scientific 14000013) for 36 cycles. Samples were run on a FlashGel system (Lonza, 57067) utilizing a 2.2% DNA cassette (57031). The results of PPMO induced exon-skipping are shown in FIGs.5A-5B.
- the PPMOs used in this example are of Formula (IV) above having the sequences indicated in Table 8. Table 8.
- RT-PCR conditions were as follows: 1.Incubate at 25° C for 10 minutes. 2.Incubate at 50 °C for 15 minutes.
- cell lysates were prepared in M-PER buffer (ThermoFisher, 78501) supplemented with HaltTM Protease and Phosphatase Inhibitor Cocktail, EDTA-free (ThermoFischer, 78447). Protein concentration was measured using PierceTM Rapid Gold BCA Protein Assay Kit (ThermoFisher, A53225). Cell lysates were prepared using the sample preparation kit (Proteinsimple) for an automated capillary Western blot system, JESS system (Proteinsimple), and utilizing the protein normalization module.
- M-PER buffer ThermoFisher, 78501
- HaltTM Protease and Phosphatase Inhibitor Cocktail EDTA-free
- Protein concentration was measured using PierceTM Rapid Gold BCA Protein Assay Kit (ThermoFisher, A53225).
- Cell lysates were prepared using the sample preparation kit (Proteinsimple) for an automated capillary Western blot system, J
- FIGs.6A-6B The results of protein reduction by PPMO 1 (i.e., a PPMO of Formula (IV)) are shown in FIGs.6A-6B.
- AB1190C16PV human immortalized WT myoblasts cells derived from paravertebral muscle of 16-year-old male see Mamchaoui, K., et al., Immortalized pathological human myoblasts: towards a universal tool for the study of neuromuscular disorders.
- RT-PCR was performed using MaximaTM H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTimeTM Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to the manufacturer’s instructions.
- PrimeTimeTM qPCR Probe Assays are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems. The results of AB1190C16PV myoblast immortalized cell line treated with PPMO 1, PPMO 3, PPMO 4, PPMO 5, PPMO 6, PPMO 7, and PPMO 8 (at various concentrations) are shown in Table 9 and FIG 7.
- PPMOs used in this example are of Formula (IV) having the sequences indicated in Table 9.
- RT-PCR conditions were as follows: 1.Incubate at 25° C for 10 minutes. 2.Incubate at 50 °C for 15 minutes. 3.Incubate at 85 °C for 5 minutes.
- qPCR conditions were as follows: Table 9. mRNA reduction of human DNM2 by PPMOs in differentiated myoblast immortalized cell line.
- NTC non targeting control.
- the NTC is a PMO that targets exon 2, but does not induce skipping .
- AB1190C16PV human immortalized WT myoblasts cells (derived from paravertebral muscle of 16-year-old male) were seeded at 10,000 cells per well in 96-well collagen Type I coated plates, in PromoCell skeletal muscle growth media (C-23060). Cells were allowed to proliferate for two days after which media was replaced to differentiation media (C-23061) (method 2).
- RNA extracted Quality of DNA 96 kit, Zymo Research
- RT-PCR was performed using MaximaTM H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTimeTM Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to manufacturer’s instructions.
- IDTT PrimeTime qPCR Probe Assays
- RT-PCR conditions were as follows: 1.Incubate at 25° C for 10 minutes. 2.Incubate at 50 °C for 15 minutes. 3.Incubate at 85 °C for 5 minutes.
- qPCR conditions were as follows: Table 10. mRNA reduction of human DNM2 by PPMOs in differentiated myoblast immortalized cell line. NTC – non targeting control. The NTC is a PMO that targets exon 2, but does not induce skipping. Example 9 – mRNA reduction in human DNM2 following PPMO treatment in differentiated healthy human skeletal muscle derived iPSCs.
- iPSC reprogramming (iXCell) [1153] Human fibroblast cells were reprogrammed using integration-free episomal vectors carrying Oct-4, Sox2, Lin28, L-Myc, Klf4, mp53DD, and EBNA from the Ei5TM Episomal iPSC Reprogramming kit (Life Technology, Cat.no. A15960). Primary fibroblast cells were cultured at a density of 105 cells, electrotransfected (Neon, Themofisher), and seeded on plates coated with Matrigel (Corning, Cat.no.354230).
- iPSCs were cultured on a mouse feeder layer until day 15 that was replaced with mTesR medium (StemCell Technologies, Cat.no.100-0276) until iPSC colonies appeared. The colonies were isolated, and their pluripotency was validated by immunostaining with markers Oct3/4, NANOG, TRA-1-60. iPSCs were confirmed to retain normal karyotype and alkaline phosphatase activity. The iPSCs were maintained in the same mTesR medium on Matrigel-coated dishes and passaged using ReLeSR (StemCell Technologies, Cat.no.05873).
- Myogenic progenitor generation (iXCell) [1154] Myogenic progenitors were differentiated from hiPSCs according to the protocols described in Chal, J., et al., Differentiation of pluripotent stem cells to muscle fiber to model Duchenne muscular dystrophy. Nat. Biotech.2015, 33, 962–969; and Chal, J., et al. Generation of human muscle fibers and satellite-like cells from human pluripotent stem cells in vitro. Nat. Protoc.2016, 11, 1833–1850. Briefly, myogenic progenitors were generated through a multi- step small molecule differentiation protocol, and later subcultured in hSKM Expansion Medium and Differentiation Medium (iXCells Biotechnologies, San Diego, US).
- iPSC Human normal induced pluripotent stem cells
- iXCell 40HU- 176
- MD-0102A iPSC-derived Myoblast Expansion Medium
- MD-0102B iPSC-derived Myoblast Differentiation Medium
- RT-PCR and qPCR conditions were as described in Example 8.
- the results of healthy human iPSC derived skeletal muscle treated with PPMO 1, PPMO 3, PPMO 4, PPMO 5, PPMO 6, PPMO 7, and PPMO 8 (at various concentrations) are shown in Table 11 and FIG 9.
- the PPMOs used in this example are of Formula (IV) above having the sequences indicated in Table 11.
- Table 11 mRNA reduction of human DNM2 by PPMOs in differentiated healthy human skeletal muscle derived iPSCs.
- NTC non targeting control.
- the NTC is a PMO that targets exon 2, but does not induce skipping.
- Example 10 – mRNA reduction of human DNM2 following PPMO treatment in MTM1 patient-derived iPSC myotubes, GM06813 C1.
- iPSC reprogramming iXCell
- Human fibroblast cells were reprogrammed using integration-free episomal vectors carrying Oct-4, Sox2, Lin28, L-Myc, Klf4, mp53DD, and EBNA from the Ei5TM Episomal iPSC Reprogramming kit (Life Technology, Cat.no. A15960).
- fibroblast cells were cultured at a density of 105 cells, electrotransfected (Neon, Themofisher), and seeded on plates coated with Matrigel (Corning, Cat.no.354230). Cells were cultured on a mouse feeder layer until day 15 that was replaced with mTesR medium (StemCell Technologies, Cat.no.100-0276) until iPSC colonies appeared. The colonies were isolated, and their pluripotency was validated by immunostaining with markers Oct3/4, NANOG, TRA-1-60. iPSCs were confirmed to retain normal karyotype and alkaline phosphatase activity.
- the iPSCs were maintained in the same mTesR medium on Matrigel-coated dishes and passaged using ReLeSR (StemCell Technologies, Cat.no.05873).
- Myogenic progenitor generation (iXCell) [1157] Myogenic progenitors were differentiated from hiPSCs according to the protocol described in Example 9. Myogenic lineage was confirmed by immunofluorescence of the myoblast markers Desmin and MyoD and expression. Terminal differentiation of the myoblasts was performed over 3-6 days of culturing and confirmed by expression of the myogenic markers MHC and MyoG measured by immunofluorescence.
- fibroblast lines Two patient untransformed fibroblast lines were purchased from the Coriell Institute, GM06813 and GM24509A, and reprogrammed at iXCell into iPSC derived myoblasts as detailed above. Two clones of each line were validated as detailed above. Cells from clone 1 of GM06813 (GM06813 C1) and clone 4 of GM24509A (GM24509A C4) were seeded at 20,000 cells per well in 96-well collagen Type I coated plates, in iPSC-derived Myoblast Expansion Medium (MD-0102A). Cells expanded in expansion media for one day before media was replaced for iPSC-derived Myoblast Differentiation Medium (MD-0102B).
- MD-0102A iPSC-derived Myoblast Expansion Medium
- RNA extraction Quantick RNA 96 kit, Zymo Research
- RT-PCR was performed using MaximaTM H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTimeTM Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to manufacturer’s instructions.
- PrimeTime qPCR Probe Assays are detailed in the table below.
- NTC non targeting control.
- the NTC is a PMO that targets exon 2, but does not induce skipping.
- the assay conditions for this experiment were as described in Example 10.
- the results of healthy human iPSC derived skeletal muscle treated with PPMO2, PPMO 3, PPMO 4, PPMO 5, PPMO 6, PPMO 7, and PPMO 8 (at various concentrations) are shown in Table 13 and FIG 11.
- the PPMOs used in this example are of Formula (IV) having the sequences indicated in Table 13. Table 13.
- NTC non targeting control.
- the NTC is a PMO that targets exon 2, but does not induce skipping.
- Example 12 Dose response with select PPMO compounds in GM06813 C1 myotubes.
- iPSC reprogramming (iXCell) [1160] Human fibroblast cells were reprogrammed using integration-free episomal vectors carrying Oct-4, Sox2, Lin28, L-Myc, Klf4, mp53DD, and EBNA from the Ei5TM Episomal iPSC Reprogramming kit (Life Technology, Cat.no. A15960). Primary fibroblast cells were cultured at a density of 105 cells, electrotransfected (Neon, Themofisher), and seeded on plates coated with Matrigel (Corning, Cat.no.354230).
- iPSCs were cultured on a mouse feeder layer until day 15 that was replaced with mTesR medium (StemCell Technologies, Cat.no.100-0276) until iPSC colonies appeared. The colonies were isolated, and their pluripotency was validated by immunostaining with markers Oct3/4, NANOG, TRA-1-60. iPSCs were confirmed to retain normal karyotype and alkaline phosphatase activity. The iPSCs were maintained in the same mTesR medium on Matrigel-coated dishes and passaged using ReLeSR (StemCell Technologies, Cat.no.05873).
- Myogenic progenitor generation (iXCell) [1161] Myogenic progenitors were differentiated from hiPSCs according to the protocol described in Example 9. Myogenic lineage was confirmed by immunofluorescence of the myoblast markers Desmin and MyoD and expression. Terminal differentiation of the myoblasts was performed over 3-6 days of culturing and confirmed by expression of the myogenic markers MHC and MyoG measured by immunofluorescence. The results of the dose responses on protein reduction by PPMO 2, PPMO 3, PPMO 4, PPMO 5, PPMO 6, PPMO 7, and PPMO 8 in normal myotubes differentiated for three days are shown in FIGs.12A-12G.
- the PPMOs used in this example are of Formula (IV) having the sequences indicated in Table 14.
- NTC is a non- targeting control.
- the NTC is a PMO that targets exon 2, but does not induce skipping (is not active).
- MD- 0102A iPSC-derived Myoblast Expansion Medium
- MD-0102B iPSC-derived Myoblast Differentiation Medium
- RNA extraction Quick RNA 96 kit, Zymo Research
- RT-PCR was performed using MaximaTM H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTimeTM Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to the manufacturer’s instructions.
- PrimeTime qPCR Probe Assays are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems.
- Example 13 Exon-skipping of human DNM2 by PPMOs in healthy myotubes.
- Healthy human myoblast cells were allowed to differentiate in differentiation media (5% HS DMEM-low glucose) for three days followed by treatment for three days with PPMOs diluted in Opti-MEM to the indicated doses (10, 20, 30 or 15 ⁇ M), free uptake (untreated control, NTC control (Gene Tools), negative- no template).
- the NTC is a 25mer PMO that is not complimentary to any portion of DMN2.
- RT-PCR was performed on 50 ng of the RNA template using Superscript III One-Step RT-PCR System with Platinum® Taq DNA Polymerase (ThermoFisher Scientific, Australia). Cycling conditions are listed in Table below. They include 55 °C for 30 minutes, 94 °C for 2 minutes followed by various cycles of 94 °C for 30 seconds, 60 °C for 30 seconds and 68 °C for 1 minute. [1165] Densitometry was performed and the exon skipping was calculated as a ratio of skipped transcriptions to total transcripts.
- the PPMOs used in this example are of Formula (VI) below having the sequences indicated in Table 15, where R5G is a cell penetrating peptide having the sequence Arg–Arg–Arg–Arg–Arg–Gly–, where z is 5 in Formula (VI); and R7G is a cell penetrating peptide having the sequence Arg–Arg–Arg–Arg–Arg–Arg–Gly–, where z is 7 in Formula (VI).
- Table 15 Summary of exon-skipping of human DNM2 by PPMOs in healthy myotubes.
- R7G is a peptide Arg–Arg–Arg–Arg–Arg–Arg–Arg–Gly–.
- Example 14 Protein reduction of human DNM2 by PPMOs in healthy myotubes.
- Normal human myoblast cells were allowed to differentiate in differentiation media (5% HS DMEM-low glucose) for three days followed by treatment for three days with PPMOs diluted in Opti-MEM at 15 ⁇ M, free uptake (untreated control, NTC control (Gene Tools), negative- no template).
- the NTC is a 25mer PMO that is not complimentary to any portion of DMN2.
- DNM2 was detected with polyclonal anti-DNM2 (Abcam) applied at a dilution of 1:2000 for 24 hours at 4 °C.
- ⁇ tubulin was detected with polyclonal anti ⁇ Tubulin (Thermo Fisher) at a dilution of 1:2500 for 24 hours at 4 °C.
- HRP-labelled anti-rabbit secondary antibody was applied respectively for 1 hour at room temperature. The blots were detected using Immobilon western chemiluminescent HRP substrate (Merck Millipore) and images were captured using a Fusion FX gel documentation system (Vilber Lourmat) with FusionCapt Advance software. Image J software (NIH) was used for densitometric analysis.
- the PPMOs used in this example are of Formula (VI) below having the sequences indicated in Table 16, where R5G is a cell penetrating peptide having the sequence Arg–Arg–Arg–Arg–Arg–Gly–, where z is 5 in Formula (VI); and R7G is a cell penetrating peptide having the sequence Arg–Arg–Arg–Arg–Arg–Arg–Gly–, where z is 7 in Formula (VI). Table 16. Protein reduction of human DNM2 by PPMOs in healthy myotubes. R7G is a peptide Arg–Arg–Arg–Arg–Arg–Arg–Gly–. Table 17. DNM2 Target Regions and Targeting Sequences.
- B represents an abasic subunit or guanine.
Landscapes
- Life Sciences & Earth Sciences (AREA)
- Genetics & Genomics (AREA)
- Health & Medical Sciences (AREA)
- Engineering & Computer Science (AREA)
- Biomedical Technology (AREA)
- General Engineering & Computer Science (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Organic Chemistry (AREA)
- Chemical & Material Sciences (AREA)
- Biotechnology (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Molecular Biology (AREA)
- Microbiology (AREA)
- Plant Pathology (AREA)
- Virology (AREA)
- Biophysics (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Physics & Mathematics (AREA)
- Pharmaceuticals Containing Other Organic And Inorganic Compounds (AREA)
Abstract
Provided herein are oligonucleotides/oligomers, peptide-oligonucleotide-conjugates, and targeting sequences complementary to a target region within a pre-mRNA of the human dynamin 2 (DNM2) gene. Also provided herein are methods of treating a disease or disorder associated with aberrant expression of DNM2 in a subject in need thereof, comprising administering to the subject the antisense oligomer, or a pharmaceutically acceptable salt thereof, or pharmaceutical composition comprising the antisense oligomer, or a pharmaceutically acceptable salt thereof.
Description
ANTISENSE OLIGOMERS FOR TREATMENT OF CENTRONUCLEAR MYOPATHIES RELATED APPLICATIONS [0001] This application claims priority to U.S. Provisional Application No.: 63/591,174 filed on October 18, 2023, the contents of which are incorporated in their entirety. SEQUENCE LISTING [0002] This application contains a Sequence Listing in computer readable format. The Sequence Listing is provided as a file entitled 4140_107PC01_Sequencelisting_ST2, created October 18, 2024 which is 360,337 bytes in size. The information in the computer readable format of the Sequence Listing is incorporated herein by reference in its entirety. BACKGROUND [0003] Dynamin 2 is a ubiquitously expressed protein that belongs to the superfamily of large GTPases. Dynamin 2 (DNM2) acts as a mechanochemical scaffolding molecule that deforms biological membranes leading to the release of nascent vesicles. At the plasma membrane DNM2 is involved in clathrin-dependent and clathrin-independent endocytosis. This protein is also involved in the formation of vesicles from endosomes and trans-Golgi network. Several studies have highlighted the role of Dynamin 2 as regulator of actin and microtubule cytoskeletons. [0004] Several dominant genetic diseases are caused by heterozygous mutations of the DNM2 gene coding for the dynamin 2 protein. Autosomal dominant centronuclear myopathy (AD-CNM) results from mutations in the DNM2 gene. AD-CNM is a rare congenital myopathy characterized by the high incidence of centrally placed nuclei in muscle fibers in absence of regenerative process. The AD-CNM is associated with a wide clinical spectrum from severe-neonatal to mild- adult forms. In general, motor milestones are delayed, and diffuse skeletal muscle weakness mainly involves facial and limb muscles. Muscle weakness is slowly progressive, but loss of independent ambulation may occur during the fifth decade. In the severe and early-onset CNM, pediatric patients usually have generalized weakness, hypotonia, moderate degree of facial weakness with open mouth, ptosis and ophthalmoplegia. No curative treatment is available for AD-CNM. [0005] Mutations in the DNM2 gene are also involved in rare cases of Charcot-Marie-Tooth disease and Hereditary Spastic Paraplegia. Charcot-Marie-Tooth disease (CMT) is a hereditary motor and sensory neuropathy characterized by progressive loss of muscle tissue and touch sensation across various parts of the body. Hereditary spastic paraplegia (HSP) is an inherited
disease characterized by lower extremity spasticity and weakness occurring in variable proportion. [0006] About thirty heterozygous mutations in the DNM2 gene were identified as involved in AD- CNM, CMT, or HSP diseases, for which there is no available curative treatment. [0007] Overexpression of DNM2 in absence of mutation is also involved in some pathophysiological mechanisms of other diseases such X-linked myotubular myopathy or cancers, for example prostate cancer and pancreatic cancer. SUMMARY [0008] The present disclosure relates to antisense oligomers and related compositions and methods for the treatment of a disease or disorder associated with aberrant DNM2 expression including, but not limited to, a genetic muscle disease or neuromuscular disease. In certain embodiments, the disease or disorder associated with aberrant DNM2 expression is a centronuclear myopathy (CNM). [0009] In an aspect provided herein, is an antisense oligomer comprising a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of human dynamin 2 (DNM2) gene represented by SEQ ID NO: 1, wherein the target region is within an exon internal region, an intron internal region, an intron-exon junction, or an exon-intron junction of the human DNM2 gene pre-mRNA. [0010] In certain embodiments, the target region is within the exon internal region of exon 2 (SEQ ID NO: 2), the intron 1/exon 2 junction, the exon 2/intron 2 junction, or the intron internal region of intron 2 (SEQ ID NO: 11) of the human DNM2 gene pre-mRNA. [0011] In certain embodiments, the target region is within the exon internal region of exon 9 (SEQ ID NO: 6), the intron 8/exon 9 junction, or the exon 9/intron 9 junction of the human DNM2 gene pre-mRNA. [0012] In certain embodiments, the target region is within the exon internal region of exon 12 (SEQ ID NO: 7), the intron 11/exon 12 junction, or the exon 12/intron 12 junction of the human DNM2 gene pre-mRNA. [0013] In certain embodiments, the target region is within the exon internal region of exon 13 (SEQ ID NO: 8), the intron 12/exon 13 junction, the exon 13/intron 13 junction, or the intron internal region of intron 12 (SEQ ID NO: 12). [0014] In certain embodiments, the target region is within the exon internal region of exon 6 (SEQ ID NO: 3), the intron 5/exon 6 junction, the exon 6/intron 6 junction, or the intron internal region of intron 6 (SEQ ID NO: 11). [0015] In certain embodiments, the target region is within the exon internal region of exon 7 (SEQ ID NO: 4), the intron 6/exon 7 junction, or the exon 7/intron 7 junction.
[0016] In certain embodiments, the target region is within the exon internal region of exon 8 (SEQ ID NO: 5), the intron 7/exon 8 junction, or the exon 8/intron 8 junction. [0017] In certain embodiments, the target region is within the exon internal region of exon 16 (SEQ ID NO: 9), the intron 15/exon 16 junction, or the exon 16/intron 16 junction. [0018] In certain embodiments, the target region is within the exon internal region of exon 19 (SEQ ID NO: 9), the intron 18/exon 19 junction, or the exon 18/intron 18 junction. [0019] In an aspect provided herein, is an antisense oligomer, or a pharmaceutically acceptable salt thereof, comprising: a non-natural chemical backbone selected from a peptide nucleic acid, a locked nucleic acid, a phosphorodiamidate morpholino oligomer, a 2′-OMe phosphorothioate oligomer, or a combination thereof; and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of human dynamin 2 (DNM2) gene represented by SEQ ID NO: 1, wherein the target region is within an exon internal region, an intron internal region, an intron-exon junction, or an exon-intron junction of the human DNM2 gene pre-mRNA. [0020] In another aspect provided herein, is an antisense oligomer, or a pharmaceutically acceptable salt thereof, having a structure of Formula (I):
(I), wherein: A′ is selected from -OH,
wherein: R5 is -C(O)(O-alkyl)x-OH, wherein x is 3-10 and each alkyl group is, independently at each occurrence, C2-6-alkyl, or R5 is selected from H, -C(O)C1-6-alkyl, trityl, monomethoxytrityl, -(C1-6-alkyl)-R6, -(C1-6- heteroalkyl)-R6, -C6-10-aryl-R6, 5-10 membered heteroaryl-R6, -C(O)O-(C1-6-alkyl)-R6, - C(O)O-aryl-R6, -C(O)O-(5-10 membered heteroaryl)-R6, and
; R6 is selected from -OH, -SH, and -NH2, or R6 is O, S, or NH, each of which is covalently linked to a solid support; R9 is C1-6 alkyl; each R1 is independently selected from -OH and -N(R3)(R4), wherein each R3 and R4 is, independently at each occurrence, -H or -C1-6-alkyl; each R2 is independently selected from a naturally or non-naturally occurring nucleobase, which, when taken together, forms a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of human dynamin 2 (DNM2) gene represented by SEQ ID NO: 1, wherein the target region is within an exon internal region, an intron internal region, an intron-exon junction, or an exon-intron junction of the human DNM2 gene pre-mRNA; t is 11-28; E′ is selected from -H, -C1-6-alkyl, -C(O)C1-6-alkyl, benzoyl, stearoyl, trityl, monomethoxytrityl, dimethoxytrityl, trimethoxytrityl,
; wherein: Q is -C(O)(CH2)6C(O)- or -C(O)(CH2)2S2(CH2)2C(O)-; R7 is -(CH2)2OC(O)N(R8)2, wherein R8 is -(CH2)6NHC(=NH)NH2; L is a linking amino acid, wherein L is covalently-linked by an amide bond to the C-terminus of J; J is a cell-penetrating peptide; and G is selected from -H, -C(O)C1-6-alkyl, benzoyl, and stearoyl, wherein G is covalently-linked to J. [0021] In an aspect provided herein, is an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formula (IA):
(IA),
wherein A′, R2, and t are as described herein. In some embodiments, A′ is a moiety selected from:
[0022] In another aspect provided herein, is antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formula (II):
(II), wherein G, R2, and t are as described herein.
[0023] In an aspect provided herein, is an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formula (III):
wherein R2 is as described herein, and n is 11-28. [0024] In an aspect provided herein, is antisense oligomer of Formula (IV):
(IV), wherein R2 and n are as described herein. [0025] In an aspect provided herein, is a pharmaceutical composition comprising: an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formulae (I), (IA), (II), (III), (IV) and/or (V); and a pharmaceutically acceptable carrier. [0026] In an aspect provided herein, is a method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein, comprising administering to a subject in need thereof a therapeutically effective amount of an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formula (I):
(I), wherein A′, R1, R2, t, E′, and G are as described herein. [0027] In another aspect provided herein, is a method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein, comprising administering to a subject in need thereof a therapeutically effective amount of an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formula (IA):
(IA), wherein A′, R2, and t are as described herein. [0028] In an aspect provided herein, is a method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein, comprising administering to a subject in need thereof a therapeutically effective amount of an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formula (II):
wherein G, R2, and t are as described herein. [0029] In an aspect provided herein, is a method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein, comprising administering to a subject in need thereof a therapeutically effective amount of an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formula (III):
wherein R2 is as described herein, and n is 11-28. [0030] In another aspect provided herein, is a method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein, comprising administering to a subject in need thereof a therapeutically effective amount of an antisense oligomer, or a pharmaceutically acceptable salt thereof, of In an aspect provided herein, is antisense oligomer of Formula (IV):
(IV), wherein R2 and n are as described herein. [0031] In certain embodiments, the disease or disorder is a genetic muscle disorder. In certain embodiments, the disease or disorder is a neuromuscular disorder. In certain embodiments, the disease or disorder is a centronuclear myopathy (CNM). In certain embodiments, the disease or disorder is myotubular myopathy (XLMTM or XLCNM), autosomal dominant congenital myopathy (ADCNM), autosomal recessive centronuclear myopathy (ARCNM), or Charcot-Marie- Tooth disease (CMT). In certain embodiments, the CMT is DNM2-implicated CMT. [0032] In certain embodiments, the Charcot-Marie-Tooth disease (CMT) is Charcot-Marie-Tooth disease, dominant intermediate B (CMTDIB or DI-CMTB). In certain embodiments, the Charcot-
Marie-Tooth disease is Charcot-Marie-Tooth disease, axonal type 2M (CMT2M). In certain embodiments, the disease or disorder is a hereditary spastic paraplegia (HSP). [0033] The antisense oligomers disclosed herein are useful in methods of inducing exon skipping to create premature stop codon in a human DNM2 pre-mRNA, thereby triggering nonsense-mediated decay and reducing DNM2-aggregation in diseased cells. Accordingly, the antisense oligomers described herein are useful for the treatment for various diseases in a subject in need thereof, including, but not limited to, diseases such as a centronuclear myopathy. BRIEF DESCRIPTION OF THE FIGURES [0034] FIGs.1A-1E show normalized DNM2 transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exons 6, 8 and 16.1A-1B: RT-qPCR after treatment with indicated PMO compounds for exon 8.1C: RT-qPCR after treatment with indicated PMO compounds for exon 16.1D-1E: RT-qPCR after treatment with indicated PMO compounds for exon 6. [0035] FIGs.2A-2F show normalized DNM2 transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exons 2, 7, 9, 12, 13, and 19.2A: RT-qPCR after treatment with indicated PMO compounds for exon 2.2B: RT-qPCR after treatment with indicated PMO compounds for exon 9.2C: RT-qPCR after treatment with indicated PMO compounds for exon 12.2D: RT-qPCR after treatment with indicated PMO compounds for exon 13.2E: RT-qPCR after treatment with indicated PMO compounds for exon 19.2F: RT-qPCR after treatment with indicated PMO compounds for exon 7. [0036] FIGs.3A-3J show normalized DNM2 transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exons, 2, 9, 12, and 13.3A-3B: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 2.3C-3E: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 9.3F-3G: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 12.3H-3J: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 13. [0037] FIGs.4A-4G show the dose response of PMOs targeted against exons 2, 9, 12, and 13 as measured by RT-qPCR.4A-4B and G: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 2.4C-4D: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 9.4E: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 12.4F: transcript levels as measured by RT-qPCR after treatment with indicated PMO compounds for exon 13.
[0038] FIGs.5A-5B show DNM2 exon skipping confirmation after treatment with select compounds targeting exon 2 and exon 9 as measured by RT-PCR.5A: photographic depiction of DNA agarose gel electrophoresis of DNM2 exon skipping confirmation after treatment with select compounds targeting exon 2.5B: photographic depiction of DNA agarose gel electrophoresis of DNM2 exon skipping confirmation after treatment with select compounds targeting exon 9. [0039] FIGs.6A-6B show treatment with select exon 2 targeting compound results in DNM2 protein reduction.6A: photographic depiction of capillary electrophoresis of DNM2 protein reduction in RD Cells after treatment with PPMOs.6B: is a graphic representation of exon 2 targeting compound results in DNM2 protein reduction. [0040] FIG.7 shows normalized DNM2 transcript levels as measured by RT-qPCR after PPMO treatment in AB1190C16PV myoblast immortalized cell line, differentiation method 1. [0041] FIG.8 shows normalized DNM2 transcript levels as measured by RT-qPCR after PPMO treatment in AB1190C16PV myoblast immortalized cell line, differentiation method 2. [0042] FIG.9 shows normalized DNM2 transcript levels as measured by RT-qPCR after PPMO treatment in healthy human iPSC derived skeletal muscle. [0043] FIG.10 shows normalized DNM2 transcript levels as measured by RT-qPCR after PPMO treatment in MTM1 patient derived iPSC myotubes, GM06813 C1. [0044] FIG.11 shows normalized DNM2 transcript levels as measured by RT-qPCR after PPMO treatment in MTM1 patient derived iPSC myotubes, GM24509 C4. [0045] FIGs.12A-12G show dose response of normalized DNM2 transcript levels as measured by RT-qPCR with select compounds in GM06813 C1 myotubes.12A: dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 2 in GM06813 C1 myotubes.12B: dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 3 in GM06813 C1 myotubes.12C: dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 4 in GM06813 C1 myotubes.12D: dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 5 in GM06813 C1 myotubes.12E: dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 6 in GM06813 C1 myotubes.12F: dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 7 in GM06813 C1 myotubes.12G: dose response of normalized DNM2 transcript levels as measured by RT-qPCR with PPMO 8 in GM06813 C1 myotubes. [0046] FIGs.13A-13D show DNM2 exon skipping confirmation in myotubes after treatment with indicated PPMO compounds as measured by RT-PCR.13A: DNM2 exon skipping confirmation in myotubes after treatment with PPMO 10 as measured by RT-PCR.13B: DNM2 exon skipping
confirmation in myotubes after treatment with PPMOs 14, 22, and 28 as measured by RT-PCR. 13C: DNM2 exon skipping confirmation in myotubes after treatment with PPMO 12 as measured by RT-PCR.13D: DNM2 exon skipping confirmation in myotubes after treatment with PPMOs 27 and 28 as measured by RT-PCR. [0047] FIGs.14A-14D show DNM2 exon skipping confirmation in myotubes after treatment with indicated PPMO compounds as measured by RT-PCR.14A: DNM2 exon skipping confirmation in myotubes after treatment with PPMO 29 as measured by RT-PCR.14B: DNM2 exon skipping confirmation in myotubes after treatment with PPMOs 30, 22, and 28 as measured by RT-PCR. 14C: DNM2 exon skipping confirmation in myotubes after treatment with PPMOs 1 and 29 as measured by RT-PCR.14D: DNM2 exon skipping confirmation in myotubes after treatment with PPMO 35 as measured by RT-PCR. [0048] FIGs.15A-15B show DNM2 protein reduction in myotubes after treatment with indicated PPMO compounds.15A: photographic depiction of a Western blot of DNM2 protein reduction in myotubes after treatment with PPMOs 10, 27, and 28.15B: a graphic representation of DNM2 protein reduction in myotubes after treatment with PPMOs 10, 27, and 28. [0049] FIGs.16A-16B show DNM2 protein reduction in myotubes after treatment with indicated PPMO compounds.16A: photographic depiction of a Western blot of DNM2 protein reduction in myotubes after treatment with PPMOs 1, 29, and 35.16B: a graphic representation of DNM2 protein reduction in myotubes after treatment with PPMOs 1, 29, and 35. DETAILED DESCRIPTION [0050] Provided herein is an antisense oligomer or pharmaceutically acceptable salt thereof, comprising a modified chemical backbone and a targeting sequence of 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of the human DNM2 gene (SEQ ID NO: 1). The target region may be an intron/exon junction or an exon internal region of the human DNM2 gene pre-mRNA. Such antisense oligomers are useful for the treatment of various diseases in a subject in need thereof, including, but not limited to, centronuclear myopathies (CNM). [0051] Certain embodiments relate to methods of inducing exon skipping to create a premature stop codon on a human DNM2 pre-mRNA in a cell, comprising contacting the cell with an antisense oligomer of sufficient length and complementarity to specifically hybridize to a region within the human DNM2 gene, such that the expression of DNM2 is reduced. In some embodiments, the cell is in a subject, and the method comprises administering the antisense oligomer to the subject. In some embodiments, the cell is a diseased cell. [0052] In some embodiments, the antisense oligomer further comprises a cell-penetrating peptide, an antibody, an antibody fragment, or an antigen fragment of an antibody. In particular,
the antisense oligomer comprises a cell penetrating peptide, wherein the cell-penetrating peptide is any of the peptides provided herein or known in the art. [0053] Also provided herein are pharmaceutical compositions and methods for treating diseases or disorders, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein. I. Definitions [0054] Unless otherwise indicated, the following terms used herein have the following meanings: [0055] The term “about” will be understood by persons of ordinary skill in the art and will vary to some extent on the context in which it is used. As used herein when referring to a measurable value such as an amount, a temporal duration, and the like, the term “about” is meant to encompass variations of ±10%, including ±5%, ±1%, and ±0.1% from the specified value, as such variations are appropriate to perform the disclosed methods. [0056] The term “alkyl” refers to saturated, straight- or branched-chain hydrocarbon moieties containing, in certain embodiments, between one and six, or one and eight carbon atoms, respectively. Examples of C1-6-alkyl moieties include, but are not limited to, methyl, ethyl, propyl, isopropyl, n-butyl, tert-butyl, neopentyl, n-hexyl moieties; and examples of C1-8-alkyl moieties include, but are not limited to, methyl, ethyl, propyl, isopropyl, n-butyl, tert-butyl, neopentyl, n- hexyl, heptyl, and octyl moieties. [0057] The number of carbon atoms in an alkyl substituent can be indicated by the prefix “Cx-y,” or “Cx-Cy” where x is the minimum and y is the maximum number of carbon atoms in the substituent. Likewise, a Cx chain means an alkyl chain containing x carbon atoms. [0058] The term “heteroalkyl” by itself or in combination with another term means, unless otherwise stated, a stable straight or branched chain alkyl group consisting of the stated number of carbon atoms and up to five heteroatoms selected from O, N, S, and P, wherein the nitrogen, sulfur, and phosphorus atoms may be optionally oxidized, and the nitrogen heteroatom may be optionally quaternized. The heteroatom(s) may be placed at any position of the heteroalkyl group, including between the rest of the heteroalkyl group and the fragment to which it is attached, as well as attached to the most distal carbon atom in the heteroalkyl group. Examples include: -O-CH2-CH2-, -O-CH2-CH2-CH3, -CH2-CH2-CH2-OH, -(O-CH2-CH2)3-OH-, -CH2-CH2-NH- CH3, -CH2-S-CH2-CH3, and -CH2-CH2-S(=O)-CH3. Up to three heteroatoms may be consecutive, such as, for example, -CH2-NH-OCH3, or -CH2-CH2-S-S-CH3. [0059] The term “aryl,” employed alone or in combination with other terms, means, unless otherwise stated, a carbocyclic aromatic system containing one or more rings (typically one, two, or three rings), wherein such rings may be attached together in a pendent manner, such as a
biphenyl, or may be fused, such as naphthalene. Examples of aryl groups include phenyl, anthracyl, and naphthyl. In various embodiments, examples of an aryl group may include phenyl (i.e., C6-aryl) and biphenyl (i.e., C12-aryl). In some embodiments, aryl groups have from six to sixteen carbon atoms (i.e., C6-16-aryl). In some embodiments, aryl groups have from six to twelve carbon atoms (i.e., C6-12-aryl). In some embodiments, aryl groups have six carbon atoms (i.e., C6-aryl). [0060] As used herein, the term “heteroaryl” or “heteroaromatic” refers to a heterocycle having aromatic character. Heteroaryl substituents may be described by the number of carbon atoms, e.g., C1-9-heteroaryl indicates the number of carbon atoms contained in the heteroaryl group without including the number of heteroatoms. For example, a C1-9-heteroaryl will include an additional one to four heteroatoms. A polycyclic heteroaryl may include one or more rings that are partially saturated. Non-limiting examples of heteroaryls include pyridyl, pyrazinyl, pyrimidinyl (including, e.g., 2- and 4-pyrimidinyl), pyridazinyl, thienyl, furyl, pyrrolyl (including, e.g., 2-pyrrolyl), imidazolyl, thiazolyl, oxazolyl, pyrazolyl (including, e.g., 3- and 5-pyrazolyl), isothiazolyl, 1,2,3-triazolyl, 1,2,4-triazolyl, 1,3,4-triazolyl, tetrazolyl, 1,2,3-thiadiazolyl, 1,2,3-oxadiazolyl, 1,3,4-thiadiazolyl and 1,3,4-oxadiazolyl. [0061] Non-limiting examples of polycyclic heterocycles and heteroaryls include indolyl (including, e.g., 3-, 4-, 5-, 6- and 7-indolyl), indolinyl, quinolyl, tetrahydroquinolyl, isoquinolyl (including, e.g., 1- and 5-isoquinolyl), 1,2,3,4-tetrahydroisoquinolyl, cinnolinyl, quinoxalinyl (including, e.g., 2- and 5-quinoxalinyl), quinazolinyl, phthalazinyl, 1,8-naphthyridinyl, 1,4-benzodioxanyl, coumarin, dihydrocoumarin, 1,5-naphthyridinyl, benzofuryl (including, e.g., 3-, 4-, 5-, 6- and 7-benzofuryl), 2,3-dihydrobenzofuryl, 1,2-benzisoxazolyl, benzothienyl (including, e.g., 3-, 4-, 5-, 6-, and 7-benzothienyl), benzoxazolyl, benzothiazolyl (including, e.g., 2-benzothiazolyl and 5-benzothiazolyl), purinyl, benzimidazolyl (including, e.g., 2-benzimidazolyl), benzotriazolyl, thioxanthinyl, carbazolyl, carbolinyl, acridinyl, pyrrolizidinyl, and quinolizidinyl. [0062] The term “protecting group” or “chemical protecting group” refers to chemical moieties that block some or all reactive moieties of a compound and prevent such moieties from participating in chemical reactions until the protective group is removed, for example, those moieties listed and described in T.W. Greene, P.G.M. Wuts, Protective Groups in Organic Synthesis, 3rd ed. John Wiley & Sons (1999). It may be advantageous, where different protecting groups are employed, that each (different) protective group be removable by a different means. Protective groups that are cleaved under totally disparate reaction conditions allow differential removal of such protecting groups. For example, protective groups can be removed by acid, base, and hydrogenolysis. Groups such as trityl, monomethoxytrityl,
dimethoxytrityl, acetal and tert-butyldimethylsilyl are acid labile and may be used to protect carboxy and hydroxy reactive moieties in the presence of amino groups protected with Cbz groups, which are removable by hydrogenolysis, and Fmoc groups, which are base labile. Carboxylic acid moieties may be blocked with base labile groups such as, without limitation, methyl, or ethyl, and hydroxy reactive moieties may be blocked with base labile groups such as acetyl in the presence of amines blocked with acid labile groups such as tert-butyl carbamate or with carbamates that are both acid and base stable but hydrolytically removable. [0063] Carboxylic acid and hydroxyl reactive moieties may also be blocked with hydrolytically removable protective groups such as the benzyl group, while amine groups may be blocked with base labile groups such as Fmoc. A particularly useful amine protecting group for the synthesis of compound of Formula (I) is trifluoroacetamide. Carboxylic acid reactive moieties may be blocked with oxidatively-removable protective groups such as 2,4-dimethoxybenzyl, while coexisting amino groups may be blocked with fluoride labile silyl carbamates. [0064] Allyl blocking groups are useful in the presence of acid- and base-protecting groups since the former are stable and can be subsequently removed by metal or pi-acid catalysts. For example, an allyl-blocked carboxylic acid can be deprotected with a palladium(0)-catalyzed reaction in the presence of acid labile t-butyl carbamate or base-labile acetate amine protecting groups. Yet another form of protecting group is a resin to which a compound or intermediate may be attached. Provided that the residue is attached to the resin, that functional group is blocked and cannot react. Once released from the resin, the functional group is available to react. [0065] The term “nucleobase,” “base pairing moiety,” “nucleobase-pairing moiety,” or “base” refers to the heterocyclic ring portion of a nucleoside, nucleotide, and/or morpholino subunit. Nucleobases may be naturally occurring (e.g., uracil, thymine, adenine, cytosine, and guanine), or may be modified or analogs of these naturally occurring nucleobases, e.g., one or more nitrogen atoms of the nucleobase may be independently at each occurrence replaced by carbon. Exemplary analogs include hypoxanthine (the base component of the nucleoside inosine); 2, 6-diaminopurine; 5-methyl cytosine; C5-propynyl-modified pyrimidines; 10-(9- (aminoethoxy)phenoxazinyl) (G-clamp) and the like. [0066] Further examples of base pairing moieties include, but are not limited to, uracil, thymine, adenine, cytosine, guanine and hypoxanthine having their respective amino groups protected by acyl protecting groups, 2-fluorouracil, 2-fluorocytosine, 5-bromouracil, 5-iodouracil, 2,6- diaminopurine, azacytosine, pyrimidine analogs such as pseudoisocytosine and pseudouracil and other modified nucleobases such as 8-substituted purines, xanthine, or hypoxanthine (the latter two being the natural degradation products). The modified nucleobases disclosed in Chiu
and Rana (2003) RNA 9:1034-1048, Limbach et al. (1994) Nucleic Acids Res.22:2183-2196 and Revankar and Rao, Comprehensive Natural Products Chemistry, vol.7, 313, are also contemplated, the contents of which are incorporated herein by reference. [0067] Further examples of base pairing moieties include, but are not limited to, expanded-size nucleobases in which one or more benzene rings have been added. Nucleic base replacements described in the Glen Research catalog (www.glenresearch.com); Krueger AT et al. (2007) Acc. Chem. Res.40:141-150; Kool ET (2002) Acc. Chem. Res.35:936-943; Benner SA et al. (2005) Nat. Rev. Genet.6:553-543; Romesberg FE et al. (2003) Curr. Opin. Chem. Biol.7:723-733; Hirao, I (2006) Curr. Opin. Chem. Biol.10:622-627, the contents of which are incorporated herein by reference, are contemplated as useful for the synthesis of the oligomers described herein. Examples of expanded-size nucleobases are shown below:
[0068] As used herein, the terms “-G-R6” and “-G-R6-Ac” and “R6G” are used interchangeably and refer to a peptide moiety conjugated to an antisense oligomer of the disclosure. In various embodiments, “G” represents a glycine residue conjugated to “R6” by an amide bond, and each “R” represents an arginine residue conjugated together by amide bonds such that “R6” means six (6) arginine residues conjugated together by amide bonds. The arginine residues can have any stereo configuration, for example, the arginine residues can be L-arginine residues, D-
arginine residues, or a mixture of D- and L-arginine residues. In certain embodiments, “-G-R6” or “-G-R6-Ac” is conjugated to the morpholine ring nitrogen of the 3′ most morpholino subunit of a PMO antisense oligomer of the disclosure. In some embodiments, “-G-R6” or “-G-R6-Ac” is conjugated to the 3′ end of an antisense oligomer of the disclosure and is of the following formula:
. [0069] The terms “oligonucleotide” or “oligomer” refer to a compound comprising a plurality of linked nucleosides, nucleotides, or a combination of both nucleosides and nucleotides. In specific embodiments provided herein, an oligonucleotide is a morpholino oligonucleotide. [0070] The term “nucleoside” means a compound comprising a nucleobase and a sugar moiety. The nucleobase and sugar moiety are each, independently, unmodified or modified. “Modified nucleoside” means a nucleoside comprising a modified nucleobase and/or a modified sugar moiety. Modified nucleosides include abasic nucleosides, which lack a nucleobase. In certain embodiments, the antisense oligonucleotides comprising an abasic subunit have the partial structure:
[0071] In certain embodiments, modified oligonucleotides comprise one or more nucleosides comprising a modified nucleobase. In certain embodiments, modified oligonucleotides comprise one or more nucleosides that do not comprise a nucleobase, referred to as an abasic nucleoside. With reference to an oligonucleotide, “subunit” refers to a nucleotide, nucleoside, nucleobase, or sugar, or a modified nucleotide, nucleoside, nucleobase, or sugar as provided herein. [0072] As used herein, the terms “antisense oligomer” or “antisense compound” are used interchangeably and refer to a sequence of subunits, each having a base carried on a backbone subunit composed of ribose or other pentose sugar or morpholino group, and where the backbone groups are linked by intersubunit linkages that allow the bases in the compound to hybridize to a target sequence in a nucleic acid (typically an RNA) by Watson-Crick base pairing, to form a nucleic acid:oligomer heteroduplex within the target sequence. The oligomer may have exact sequence complementarity to the target sequence or nearly exact complementarity. Such antisense oligomers are designed to block or inhibit translation of the mRNA containing the target sequence and may be said to be “directed to” a sequence with which it hybridizes. [0073] Also contemplated herein as types of “antisense oligomers” or “antisense compounds” are phosphorothioate-modified oligomers, peptide nucleic acids (PNAs), locked nucleic acids (LNAs), 2′-fluoro-modified oligomers, 2′-O,4′-C-ethylene-bridged nucleic acids (ENAs), tricyclo- DNAs, tricylo-DNA phosphorothioate-modified oligomers, 2′-O-[2-(N-methylcarbamoyl) ethyl] modified oligomers, 2′-O-methyl phosphorothioate modified oligomers, 2′-O-methoxyethyl (2′-O- MOE) modified oligomers, and 2′-O-Methyl oligonucleotides, or combinations thereof, as well as other antisense agents known in the art.
[0074] An antisense oligomer “specifically hybridizes” to a target polynucleotide if the oligomer hybridizes to the target under physiological conditions, with a Tm greater than 37 °C, greater than 45 °C, at least 50 °C, and typically 60 °C-80 °C or higher. The “Tm” of an oligomer is the temperature at which 50% hybridizes to a complementary polynucleotide. Tm is determined under standard conditions in physiological saline, as described, for example, in Miyada et al. (1987) Methods Enzymol.154:94-107. Such hybridization may occur with “near” or “substantial” complementarity of the antisense oligomer to the target sequence, as well as with exact complementarity. [0075] The terms “complementary” and “complementarity” refer to oligonucleotides (i.e., a sequence of nucleotides) related by base-pairing rules. For example, the sequence “T-G-A (5′- 3′)” is complementary to the sequence “T-C-A (5′-3′).” Complementarity may be “partial,” in which only some of the nucleic acids’ bases are matched according to base pairing rules, or there may be “complete,” “total,” or “perfect” (100%) complementarity between the nucleic acids. The degree of complementarity between nucleic acid strands has significant effects on the efficiency and strength of hybridization between nucleic acid strands. In some embodiments, oligonucleotides can include one or more mismatches (e.g., 6, 5, 4, 3, 2, or 1 mismatches) with respect to the target RNA. Such hybridization may occur with “near” or “substantial” complementarity of the antisense oligomer to the target sequence, as well as with exact complementarity. In some embodiments, an oligomer may hybridize to a target sequence with about 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 99% or 100% complementarity. Variations at any location within the oligomer are included. In certain embodiments, variations in sequence near the termini of an oligomer, if present, are typically within about 6, 5, 4, 3, 2, or 1 nucleotides of the 5′-terminus, 3′-terminus, or both termini. [0076] The terms “TEG,” “EG3,” or “triethylene glycol tail” refer to triethylene glycol moieties conjugated to the oligomer, e.g., at its 3′- or 5′-end. For example, in some embodiments, “TEG” includes, for example, wherein A′ of the conjugate of Formula (I) is of the formula:
. [0077] Naturally occurring nucleotide bases include adenine, guanine, cytosine, thymine, and uracil, which have the symbols A, G, C, T, and U, respectively. Nucleotide bases can also encompass analogs of naturally occurring nucleotide bases. Base pairing typically occurs between purine A and pyrimidine T or U, and between purine G and pyrimidine C.
[0078] Oligonucleotides may also include nucleobase modifications or substitutions. Oligonucleotides containing a modified or substituted base include oligonucleotides in which one or more purine or pyrimidine bases most found in nucleic acids are replaced with less common or non-natural bases. In some embodiments, the nucleobase is covalently linked at the N9 atom of the purine base, or at the N1 atom of the pyrimidine base, to the morpholine ring of a nucleotide or nucleoside. [0079] Purine bases comprise a pyrimidine ring fused to an imidazole ring, as described by the general formula:
. [0080] Adenine and guanine are the two purine nucleobases most found in nucleic acids. These may be substituted with other naturally occurring purines, including, but not limited to, N6- methyladenine, N2-methylguanine, hypoxanthine, and 7-methylguanine. [0081] Pyrimidine bases comprise a six-membered pyrimidine ring as described by the general formula:
. [0082] Cytosine, uracil, and thymine are the pyrimidine bases most found in nucleic acids. These may be substituted with other naturally occurring pyrimidines, including, but not limited to, 5-methylcytosine, 5-hydroxymethylcytosine, pseudouracil, and 4-thiouracil. In one embodiment, the oligonucleotides described herein contain thymine bases in place of uracil. [0083] Other modified or substituted bases include, but are not limited to, 2,6-diaminopurine, orotic acid, agmatidine, lysidine, 2-thiopyrimidine (e.g.2-thiouracil, 2-thiothymine), G-clamp and its derivatives, 5-substituted pyrimidine (e.g.5-halouracil, 5-propynyluracil, 5-propynylcytosine, 5-aminomethyluracil, 5-hydroxymethyluracil, 5-aminomethylcytosine, 5-hydroxymethylcytosine, Super T), 7-deazaguanine, 7-deazaadenine, 7-aza-2,6-diaminopurine, 8-aza-7-deazaguanine, 8-aza-7-deazaadenine, 8-aza-7-deaza-2,6-diaminopurine, Super G, Super A, and N4- ethylcytosine, or derivatives thereof; N2-cyclopentylguanine (cPent-G), N2-cyclopentyl-2- aminopurine (cPent-AP), and N2-propyl-2-aminopurine (Pr-AP), pseudouracil or derivatives thereof; and degenerate or universal bases, like 2,6-difluorotoluene or absent bases like abasic sites (e.g.1-deoxyribose, 1,2-dideoxyribose, 1-deoxy-2-O-methylribose; or pyrrolidine derivatives in which the ring oxygen has been replaced with nitrogen (azaribose)). Pseudouracil
is a naturally occurring isomerized version of uracil, with a C-glycoside rather than the regular N- glycoside as in uridine. [0084] In some embodiments, modified or substituted nucleobases are useful for facilitating the purification of antisense oligonucleotides. For example, in certain embodiments, antisense oligonucleotides may contain three or more (e.g., 3, 4, 5, 6, or more) consecutive guanine bases. In certain antisense oligonucleotides, a string of three or more consecutive guanine bases can result in aggregation of the oligonucleotides, complicating purification. In such antisense oligonucleotides, one or more of the consecutive guanines can be substituted with hypoxanthine. The substitution of hypoxanthine for one or more guanines in a string of three or more consecutive guanine bases can reduce aggregation of the antisense oligonucleotide, thereby facilitating purification. [0085] The oligonucleotides provided herein are synthesized and do not include antisense compositions of biological origin. The molecules of the disclosure may also be mixed, encapsulated, conjugated, or otherwise associated with other molecules, molecule structures, or mixtures of compounds, as for example, liposomes, receptor-targeted molecules, oral, rectal, topical, or other formulations, for assisting in uptake, distribution, or absorption, or a combination thereof. [0086] As used herein, a “nucleic acid analog” refers to a non-naturally occurring nucleic acid molecule. A nucleic acid is a polymer of nucleotide subunits linked together into a linear structure. Each nucleotide consists of a nitrogen-containing aromatic base attached to a pentose (five-carbon) sugar, which is in turn attached to a phosphate group. Successive phosphate groups are linked together through phosphodiester bonds to form the polymer. The two common forms of naturally occurring nucleic acids are deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). One end of the chain carries a free phosphate group attached to the 5′-carbon atom of a sugar moiety; this is called the 5′ end of the molecule. The other end has a free hydroxyl (-OH) group at the 3′-carbon of a sugar moiety and is called the 3′ end of the molecule. A nucleic acid analog can include one or more non-naturally occurring nucleobases, sugars, and/or internucleotide linkages, for example, a phosphorodiamidate morpholino oligomer (PMO). As disclosed herein, in certain embodiments, a “nucleic acid analog” is a PMO, and in certain embodiments, a “nucleic acid analog” is a positively charged cationic PMO. [0087] A “morpholino oligomer” or “PMO” refers to a polymeric molecule having a backbone that supports bases capable of hydrogen bonding to typical polynucleotides, wherein the polymer lacks a pentose sugar backbone moiety, and more specifically a ribose backbone linked by phosphodiester bonds which is typical of nucleotides and nucleosides, but instead contains a ring nitrogen with coupling through the ring nitrogen. An exemplary “morpholino” oligomer comprises
morpholino subunit structures linked together by phosphoramidate or phosphorodiamidate linkages, joining the morpholino nitrogen of one subunit to the 5′ exocyclic carbon of an adjacent subunit, each subunit comprising a purine or pyrimidine base-pairing moiety effective to bind, by base-specific hydrogen bonding, to a base in a polynucleotide. Morpholino oligomers (including antisense oligomers) are detailed, for example, in U.S. Pat. Nos.5,034,506; 5,142,047; 5,166,315; 5,185,444; 5,217,866; 5,506,337; 5,521,063; 5,698,685; 8,076,476; and 8,299,206; and PCT publication number WO 2009/064471, all of which are incorporated herein by reference in their entirety. [0088] A preferred morpholino oligomer is a phosphorodiamidate-linked morpholino oligomer, referred to herein as a PMO. Such oligomers are composed of morpholino subunit structures such as those shown below:
where X is NH2, NHR, or NR2 (where R is lower alkyl, (e.g., methyl)), Y1 is O, and Z is O, and Pi and Pj are purine or pyrimidine base-pairing moieties effective to bind, by base-specific hydrogen bonding, to a base in a polynucleotide. Also preferred are structures having an alternate phosphorodiamidate linkage, where X is lower alkoxy, such as methoxy or ethoxy, Y1 is NH or NR, where R is lower alkyl, and Z is O. [0089] Representative PMOs include PMOs wherein the intersubunit linkages are linkage (A1). See Table 1.
Table 1. Representative Intersubunit Linkages
[0090] A “phosphoramidate” group comprises phosphorus having three attached oxygen atoms and one attached nitrogen atom, while a “phosphorodiamidate” group comprises phosphorus having two attached oxygen atoms and two attached nitrogen atoms. A representative phosphorodiamidate example is below:
where each Pi is independently selected from H, a nucleobase, and a nucleobase functionalized with a chemical protecting-group, wherein the nucleobase independently at each occurrence comprises a pyridine, a pyrimidine, a triazinane, purine, or a deaza-purine; and n is an integer of 6-38 (i.e., n is 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, or 38). The ring nitrogen of the subunit at the 3′ terminus of a PMO may be capped with a capping group such as acetyl or may be uncapped with a free hydrogen. [0091] In the uncharged or the modified intersubunit linkages of the antisense oligomers described herein, one nitrogen is always pendant to the backbone chain. The second nitrogen, in a phosphorodiamidate linkage, is typically the ring nitrogen in a morpholino ring structure. [0092] PMOs are water-soluble, uncharged, or substantially uncharged antisense molecules that inhibit gene expression by preventing binding or progression of splicing or translational machinery components. PMOs have also been shown to inhibit or block viral replication (Stein, Skilling et al.2001; McCaffrey, Meuse et al.2003), and are highly resistant to enzymatic digestion (Hudziak, Barofsky et al.1996). PMOs have demonstrated high antisense specificity and efficacy in vitro in cell-free and cell culture models (Stein, Foster et al.1997; Summerton and Weller 1997), and in vivo in zebrafish, frog, and sea urchin embryos (Heasman, Kofron et al.2000; Nasevicius and Ekker 2000), as well as in adult animal models, such as rats, mice, rabbits, dogs, and pigs (see e.g. Arora and Iversen 2000; Qin, Taylor et al.2000; Iversen 2001;
Kipshidze, Keane et al.2001; Devi 2002; Devi, Oldenkamp et al.2002; Kipshidze, Kim et al. 2002; Ricker, Mata et al.2002). [0093] Antisense PMO oligomers have been shown to be taken up into cells and to be more consistently effective in vivo, with fewer nonspecific effects, than other widely used antisense oligonucleotides (see e.g., P. Iversen, “Phosphoramidite Morpholino Oligomers,” in Antisense Drug Technology, S.T. Crooke, ed., Marcel Dekker, Inc., New York, 2001). Conjugation of PMOs to arginine-rich peptides has been shown to increase their cellular uptake (see e.g., U.S. Patent No.7,468,418, incorporated herein by reference in its entirety). [0094] “Charged,” “uncharged,” “cationic,” and “anionic” as used herein refer to the predominant state of a chemical moiety at near-neutral pH, e.g., about 6 to 8. For example, the term may refer to the predominant state of the chemical moiety at physiological pH, that is, at about 7.4. [0095] A “cationic PMO” or “PMO+” refers to a phosphorodiamidate morpholino oligomer comprising any number of (1-piperazino)phosphinylideneoxy, (1-(4-(^-guanidino-alkanoyl))- piperazino)phosphinylideneoxy linkages (A2 and A3; see Table 1) that have been described previously (see e.g., PCT publication WO 2008/036127 which is incorporated herein by reference in its entirety). [0096] The “backbone” of an antisense oligomer (e.g., an uncharged oligonucleotide analog) refers to the structure supporting the base-pairing moieties, e.g., for a morpholino oligomer, as described herein, the “backbone” includes morpholino ring structures connected by intersubunit linkages (e.g., phosphorus-containing linkages). A “substantially uncharged backbone” refers to the backbone of an oligonucleotide analogue wherein less than 50% of the intersubunit linkages are charged at near-neutral pH. For example, a substantially uncharged backbone may comprise less than 50%, less than 40%, less than 30%, less than 20%, less than 10%, less than 5% or even 0% intersubunit linkages which are charged at near neutral pH. In some embodiments, the substantially uncharged backbone comprises at most one charged (at physiological pH) intersubunit linkage for every four uncharged (at physiological pH) linkages, at most one for every eight or at most one for every sixteen uncharged linkages. In some embodiments, the nucleic acid analogs described herein are fully uncharged. [0097] In certain embodiments, the compounds described herein comprise or consist of an oligonucleotide comprising a targeting base sequence that is complementary to a target nucleic acid. In certain embodiments, the target nucleic acid is an endogenous RNA molecule. In certain embodiments, the target nucleic acid encodes a protein. In certain embodiments, the target nucleic acid is non-coding. In certain such embodiments, the target nucleic acid is selected from an mRNA and a pre-mRNA, including intronic, exonic and untranslated regions. In certain embodiments, the target RNA is an mRNA. In certain embodiments, the target nucleic acid is a
pre-mRNA. In certain such embodiments, the target region is entirely within an exon. In certain such embodiments, the target region is entirely within an intron. In certain embodiments, the target region spans an intron/exon junction. In certain embodiments, the target region is at least 50% within an intron. [0098] The term “targeting base sequence” or “targeting sequence” is the (nucleobase) sequence in the antisense oligomer that is complementary or substantially complementary to a target nucleic acid. The entire sequence, or only a portion, of the oligomer compound may be complementary to the target sequence. [0099] The “target region” refers to a sequence in a nucleic acid that is targeted by, and is therefore complementary, substantially complementary, or partially complementary, to the targeting sequence of the antisense oligomer. The entire sequence, or only a portion, of the target region may be complementary to the targeting sequence of the oligomer compound described herein. In certain embodiments, a plurality of antisense compounds is directed to a single target region. In certain embodiments, a nomenclature system according to gene, species, exon number, and annealing coordinates is used herein to identify or indicate a target region or annealing coordinates of a targeted nucleic acid (e.g., a nucleic acid transcript) as follows: G SpZA/D (±X±Y), where G is gene name; Sp is species (e.g., “H” for human, “M” for mouse); Z is exon number; A/D is selected from acceptor (A) and donor (D); The symbol “–“ denotes an intronic position; The symbol “+” denotes an exonic position; X is the position of the first 5′ base relative to A or D site; and Y is the position of the first 3′ base relative to A or D site. [0100] The nomenclature begins with the name of the transcript (e.g., dynamin 2; DNM2), then the species of the target mRNA (e.g., H: human or M: mouse), followed by the target exon number of the specified transcript and specification of an acceptor (A) or donor (D) site. The annealing coordinates are shown in brackets within the pre-mRNA transcript. The intronic bases are designated with a negative prefix (-) and the exonic position with a positive (+) symbol. The annealing coordinates are the positions of bases relative to the bases relative to the acceptor or donor sites of the reference transcript as denoted by National Center for Biotechnology Information and Ensembl genome browser 96. See, for example, international application
WO2006/000057; Aung-Htut, et al. (2019) Int. J. Mol. Sci.20:5030; and Mann, et al. (2002) J Gene Med.4:644. [0101] As an example, entirely exonic annealing coordinates represented by H5A(+76+100) indicates the site within the 76th and 100th nucleotides as measured from the 5′ end of an exon 5 (e.g., exon 5 of human DNM2 gene pre-mRNA) (i.e., the region within the 76th and 100th nucleotides as measured from the start of exon 5 at the 5′ end). The closest splice site is the acceptor; therefore, these coordinates are preceded with “A.” [0102] As another example, DNM2 H2D (+13-12) indicates the last 13 exonic bases and the first 12 intronic bases of the target exon donor splice site (i.e., the target region within the 13th nucleotide of exon 2 measured from the 3′ end of exon 2 to the 12th nucleotide of intron 2 measured from the 3′ end of exon 2 (e.g., human exon 2 of the human DNM2 gene pre-mRNA). The closest splice site is the donor; therefore, these coordinates are preceded with “D.” [0103] As another example, DNM2 H13A(-8+17) indicates the last 8 bases of the intron preceding the target exon 2 and the first 17 bases of the target exon 2 (i.e., the target region within the 8th nucleotide of intron 12 measured from the 5′ end of exon 13 to the 17th nucleotide of exon 13 measured from the 5′ end of exon 13). The closest splice site is the acceptor; therefore, these coordinates are preceded with “A.” [0104] The term “peptide” refers to a compound comprising a plurality of linked amino acids. In certain embodiments, the peptides provided herein are cell-penetrating peptides (CPP). [0105] As used herein, a “cell-penetrating peptide” (CPP) or “carrier peptide” is a relatively short peptide capable of promoting the uptake of PMOs by cells, thereby delivering the PMOs to the interior (cytoplasm) of the cells. The CPP or carrier peptide typically is about 12 to about 40 amino acids long. The length of the carrier peptide is not particularly limited and varies in different embodiments. In some embodiments, the carrier peptide comprises from 4 to 40 amino acid subunits. In other embodiments, the carrier peptide comprises from 6 to 30, from 6 to 20, from 8 to 25, or from 10 to 20 amino acid subunits. In various embodiments, the CPPs disclosed herein comprise an arginine-rich peptide as described further below. [0106] As used herein, a “peptide-conjugated phosphorodiamidate-linked morpholino oligomer” or “PPMO” refers to a PMO covalently linked to a peptide, such as a cell-penetrating peptide (CPP) or carrier peptide. The cell-penetrating peptide promotes the uptake of the PMO by cells, thereby delivering the PMO to the interior (cytoplasm) of the cells. Depending on its amino acid sequence, a CPP can be generally effective, or it can be specifically or selectively effective for PMO delivery to a particular type or particular types of cells. PMOs and CPPs are typically linked at their ends, e.g., the C-terminal end of the CPP can be linked to the 5′ end of the PMO, or the 3′ end of the PMO can be linked to the N-terminal end of the CPP. PPMOs can include
uncharged PMOs, charged (e.g., cationic) PMOs, and mixtures thereof. In an embodiment, the linking moiety of the conjugates described herein may be cleaved to release a PPMO. [0107] The carrier peptide may be linked to the nucleic acid analog either directly or via an optional linker, e.g., one or more additional naturally occurring amino acids, e.g., cysteine (C), glycine (G), or proline (P), or additional amino acid analogs, e.g., 6-aminohexanoic acid (X), beta-alanine (B), or XB. Other linking moieties known in the art may also be employed. [0108] An “amino acid subunit” is generally an ^-amino acid residue (–CO-CHR-NH-); but may also be a ^- or other amino acid residue (e.g., -CO-CH2CHR-NH-), where R is an amino acid side chain. [0109] The term “naturally occurring amino acid” refers to an amino acid present in proteins found in nature; examples include Alanine (A), Cysteine (C), Aspartic acid (D), Glutamic acid (E), Phenyalanine (F), Glycine (G), Histidine (H), Isoleucine (I), Lysine (K), Leucine (L). Methionine (M), Asparagine (N), Proline (P), Glutamine (Q), Arginine (R), Serine (S), Threonine (T), Valine (V), Tryptophan (W), and Tyrosine (Y). The term “non-natural amino acids” refers to those amino acids not present in proteins found in nature; examples include beta-alanine (β-Ala) and 6-aminohexanoic acid (Ahx). [0110] An agent is “actively taken up by mammalian cells” when the agent can enter the cell by a mechanism other than passive diffusion across the cell membrane. The agent may be transported, for example, by “active transport,” referring to transport of agents across a mammalian cell membrane by, e.g., an ATP-dependent transport mechanism, or by “facilitated transport,” referring to transport of antisense agents across the cell membrane by a transport mechanism that requires binding of the agent to a transport protein, which then facilitates passage of the bound agent across the membrane. [0111] As used herein, an “effective amount” refers to any amount of a substance that is sufficient to achieve a desired biological result. A “therapeutically effective amount” refers to any amount of a substance that is sufficient to achieve a desired therapeutic result. [0112] As used herein, a “subject” is a mammal, which can include a mouse, rat, hamster, guinea pig, rabbit, goat, sheep, cat, dog, pig, cow, horse, monkey, non-human primate, or human. In certain embodiments, a subject is a human. [0113] “Treatment” of an individual (e.g., a mammal, such as a human) or a cell is any type of intervention used to alter the natural course of the individual or cell. Treatment includes, but is not limited to, administration of a pharmaceutical composition, and may be performed either prophylactically or subsequent to the initiation of a pathologic event or contact with an etiologic agent.
II. Peptide-oligonucleotides [0114] Provided herein is an antisense oligomer comprising a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of human dynamin 2 (DNM2) gene represented by SEQ ID NO: 1, wherein the target region is within an exon internal region, an intron internal region, an intron-exon junction, or an exon-intron junction of the human DNM2 gene pre-mRNA. [0115] In certain embodiments, the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 6 (SEQ ID NO: 3), exon 7 (SEQ ID NO: 4), exon 8 (SEQ ID NO: 5), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), exon 13 (SEQ ID NO: 8), exon 16 (SEQ ID NO: 9), and exon 19 (SEQ ID NO: 10). [0116] In certain embodiments, the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), and exon 13 (SEQ ID NO: 8). In certain embodiments, the target region within an exon internal region of the human DNM2 gene pre-mRNA is exon 2 (SEQ ID NO: 2). In certain embodiments, the target region within an exon internal region of the human DNM2 gene pre-mRNA is exon 9 (SEQ ID NO: 6). In certain embodiments, the target region within an exon internal region of the human DNM2 gene pre-mRNA is exon 12 (SEQ ID NO: 7). In certain embodiments, the target region within an exon internal region of the human DNM2 gene pre- mRNA and exon 13 (SEQ ID NO: 8). [0117] In certain embodiments, the target region is within an intron-exon junction or an exon- intron junction region of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 5/exon 6 junction, exon 6/intron 6 junction, intron 6/exon 7 junction, exon 7/intron 7 junction, intron 7/exon 8 junction, exon 8/intron 8 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, exon 13/intron 13 junction, intron 15/exon 16 junction, exon 16/intron 16 junction, intron 18/exon 19 junction, and exon 19/intron 19 junction. [0118] In certain embodiments, the target region is within an intron-exon junction or an exon- intron junction of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, and exon 13/intron 13 junction. [0119] In certain embodiments, the target region within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA is intron 1/exon 2 junction. In certain embodiments, the target region within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA is exon 2/intron 2 junction. In certain embodiments, the target region within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-
mRNA is intron 8/exon 9 junction. In certain embodiments, the target region within an intron- exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA is exon 9/intron 9 junction. In certain embodiments, the target region within an intron-exon junction or an exon- intron junction of the human DNM2 gene pre-mRNA is intron 11/exon 12 junction. In certain embodiments, the target region within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA is exon 12/intron 12 junction, intron 12/exon 13 junction. In certain embodiments, the target region within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA is exon 13/intron 13 junction. [0120] In certain embodiments, the target region is within an intron internal region of the human DNM2 gene pre-mRNA selected from intron 2 (SEQ ID NO: 11) and intron 12 (SEQ ID NO: 12). In certain embodiments, the target region within an intron internal region of the human DNM2 gene pre-mRNA is intron 2 (SEQ ID NO: 11). In certain embodiments, the target region within an intron internal region of the human DNM2 gene pre-mRNA is intron 12 (SEQ ID NO: 12). [0121] In certain embodiments, the antisense oligomer is 13-30 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-30, 14-30, 15-30, 16-30, 17-30, 18-30, 19-30, 20-30, 21-30, 22-30, 23-30, 24-30, 25-30, 26-30, 27-30, 28-30, or 29-30 subunits in length. [0122] In certain embodiments, the antisense oligomer is 13-29 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-29, 14-29, 15-29, 16-29, 17-29, 18-29, 19-29, 20-29, 21-29, 22-29, 23-29, 24-29, 25-29, 26-29, 27-29, or 28-29 subunits in length. [0123] In certain embodiments, the antisense oligomer is 13-28 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-28, 14-28, 15-28, 16-28, 17-28, 18-28, 19-28, 20-28, 21-28, 22-28, 23-28, 24-28, 25-28, 26-28, or 27-28 subunits in length. [0124] In certain embodiments, the antisense oligomer is 13-27 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-27, 14-27, 15-27, 16-27, 17-27, 18-27, 19-27, 20-27, 21-27, 22-27, 23-27, 24-27, 25-27, or 26-27 subunits in length. [0125] In certain embodiments, the antisense oligomer is 13-26 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-26, 14-26, 15-26, 16-26, 17-26, 18-26, 19-26, 20-26, 21-26, 22-26, 23-26, 24-26, or 25-26 subunits in length. [0126] In certain embodiments, the antisense oligomer is 13-25 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-25, 14-25, 15-25, 16-25, 17-25, 18-25, 19-25, 20-25, 21-25, 22-25, 23-25, or 24-25 subunits in length. [0127] In certain embodiments, the antisense oligomer is 13-24 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-24, 14-24, 15-24, 16-24, 17-24, 18-24, 19-24, 20-24, 21-24, 22-24, or 23-24 subunits in length.
[0128] In certain embodiments, the antisense oligomer is 13-23 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-23, 14-23, 15-23, 16-23, 17-23, 18-23, 19-23, 20-23, 21-23, or 22-23 subunits in length. [0129] In certain embodiments, the antisense oligomer is 13-22 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-22, 14-22, 15-22, 16-22, 17-22, 18-22, 19-22, 20-22, or 21-22 subunits in length. [0130] In certain embodiments, the antisense oligomer is 13-21 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-21, 14-21, 15-21, 16-21, 17-21, 18-21, 19-21, or 20-21 subunits in length. [0131] In certain embodiments, the antisense oligomer is 13-20 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-20, 14-20, 15-20, 16-20, 17-20, 18-20, or 19- 20 subunits in length. [0132] In certain embodiments, the antisense oligomer is 13-19 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-19, 14-19, 15-19, 16-19, 17-19, or 18-19 subunits in length. [0133] In certain embodiments, the antisense oligomer is 13-18 subunits in length. In certain embodiments, the antisense oligonucleotide is 13-18, 14-18, 15-18, 16-18, or 17-18 subunits in length. [0134] In certain embodiments, the antisense oligonucleotide is 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 subunits in length. In certain embodiments, the antisense oligonucleotide is 13 subunits in length. In certain embodiments, the antisense oligonucleotide is 14 subunits in length. In certain embodiments, the antisense oligonucleotide is 15 subunits in length. In certain embodiments, the antisense oligonucleotide is 16 subunits in length. In certain embodiments, the antisense oligonucleotide is 17 subunits in length. In certain embodiments, the antisense oligonucleotide is 18 subunits in length. In certain embodiments, the antisense oligonucleotide is 19 subunits in length. In certain embodiments, the antisense oligonucleotide is 20 subunits in length. In certain embodiments, the antisense oligonucleotide is 21 subunits in length. In certain embodiments, the antisense oligonucleotide is 22 subunits in length. In certain embodiments, the antisense oligonucleotide is 23 subunits in length. In certain embodiments, the antisense oligonucleotide is 24 subunits in length. In certain embodiments, the antisense oligonucleotide is 25 subunits in length. In certain embodiments, the antisense oligonucleotide is 26 subunits in length. In certain embodiments, the antisense oligonucleotide is 27 subunits in length. In certain embodiments, the antisense oligonucleotide is 28 subunits in length. In certain embodiments, the antisense oligonucleotide is 29 subunits in length. In certain embodiments, the antisense oligonucleotide is 30 subunits in length.
[0135] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12- 13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3- 27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), DNM2 H2D(-6-30), DNM2 H9A(-12+13), DNM2 H9A(-5+20), DNM2 H9A(+1+25),DNM2 H9A(+6+30), DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), DNM2 H9A(+48+67), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11- 14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4- 17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), DNM2 H9D(+3-22), DNM2 H12A(-13+12), DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), DNM2 H12D(+15-10), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2
H12A(+37+61), DNM2 H12A(+43+67), DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(- 25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(- 11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), DNM2 H13A(+9+28), DNM2 H13A(+16+40), DNM2 H13D(+22-3), or DNM2 H13D(+7-18). [0136] In certain embodiments, the target region is DNM2 H2A (-10+15). In certain embodiments, the target region is DNM2 H2A(-7+17). In certain embodiments, the target region is DNM2 H2A(+3+27). In certain embodiments, the target region is DNM2 H2A(+15+37). In certain embodiments, the target region is DNM2 H2A(+21+45). In certain embodiments, the target region is DNM2 H2A(+23+47). In certain embodiments, the target region is DNM2 H2A(+27+51). In certain embodiments, the target region is DNM2 H2A(+37+61). In certain embodiments, the target region is DNM2 H2A(+46+70). In certain embodiments, the target region is DNM2 H2A (+48+72). In certain embodiments, the target region is DNM2 H2D(+20-5). In certain embodiments, the target region is DNM2 H2D (+13-12). In certain embodiments, the target region is DNM2 H2D(+12-13). In certain embodiments, the target region is DNM2 H2D(+11-14). In certain embodiments, the target region is DNM2 H2D(+10-15). In certain embodiments, the target region is DNM2 H2D(+9-16). In certain embodiments, the target region is DNM2 H2D(+8-17). In certain embodiments, the target region is DNM2 H2D(+7-18). In certain embodiments, the target region is DNM2 H2D(+6-19). In certain embodiments, the target region is DNM2 H2D(+5-20). In certain embodiments, the target region is DNM2 H2D(+4-21). In certain embodiments, the target region is DNM2 H2D(+3-22). In certain embodiments, the target region is DNM2 H2D(+2-23) . In certain embodiments, the target region is DNM2 H2D(+1-24). In certain embodiments, the target region is DNM2 H2D(-1-25). In certain embodiments, the target region is DNM2 H2D(-2-26). In certain embodiments, the target region is DNM2 H2D(-3-27). In certain embodiments, the target region is DNM2 H2D(-4-28). In certain embodiments, the target region is DNM2 H2D(-5-29). In certain embodiments, the target region is DNM2 H2D(-6-30). In certain embodiments, the target region is DNM2 H6A(-20+5). In certain embodiments, the target region is DNM2 H6A(-6+19). In certain embodiments, the target region is DNM2 H6A(-2+23). In certain embodiments, the target region is DNM2 H6A(+3+26). In certain embodiments, the
target region is DNM2 H6A(+10+3). In certain embodiments, the target region is DNM2 H6A(+13+37). In certain embodiments, the target region is DNM2 H6A(+23+47). In certain embodiments, the target region is DNM2 H6A(+32+56). In certain embodiments, the target region is DNM2 H6A(+35+59). In certain embodiments, the target region is DNM2 H6A(+44+68). In certain embodiments, the target region is DNM2 H6A(+54+78). In certain embodiments, the target region is DNM2 H6A(+60+84). In certain embodiments, the target region is DNM2 H6A(+63+87). In certain embodiments, the target region is DNM2 H6A(+88+112). In certain embodiments, the target region is DNM2 H6A(+96+119). In certain embodiments, the target region is DNM2 H6A(+105+129). In certain embodiments, the target region is DNM2 H6A(+121+145). In certain embodiments, the target region is DNM2 H6A(+130+154). In certain embodiments, the target region is DNM2 H6A(+132+156). In certain embodiments, the target region is DNM2 H6D (+6-19). In certain embodiments, the target region is DNM2 H6D (+15-10). In certain embodiments, the target region is DNM2 H6D (+17-8). In certain embodiments, the target region is DNM2 H7A(-15+10). In certain embodiments, the target region is DNM2 H7A(- 5+20). In certain embodiments, the target region is DNM2 H7A(+1+25). In certain embodiments, the target region is DNM2 H7A(+6+30). In certain embodiments, the target region is. In certain embodiments, the target region is DNM2 H7A(+22+46). In certain embodiments, the target region is DNM2 H7A(+29+53). In certain embodiments, the target region is DNM2 H7A(+41+65). In certain embodiments, the target region is DNM2 H7A(+51+75). In certain embodiments, the target region is DNM2 H7A(+57+81). In certain embodiments, the target region is DNM2 H7A(+62+86). In certain embodiments, the target region is DNM2 H7A(+72+96). In certain embodiments, the target region is DNM2 H7A(+74+98). In certain embodiments, the target region is DNM2 H7A(+81+105). In certain embodiments, the target region is DNM2 H7A(+91+115). In certain embodiments, the target region is DNM2 H7A(+110+134). In certain embodiments, the target region is DNM2 H7D(+8-16). In certain embodiments, the target region is DNM2 H7D(+9-16). In certain embodiments, the target region is DNM2 H7D(+21-4). In certain embodiments, the target region is DNM2 H8A(-20+5). In certain embodiments, the target region is DNM2 H8A(-14+11). In certain embodiments, the target region is DNM2 H8A(-7+18). In certain embodiments, the target region is DNM2 H8A(-6+19). In certain embodiments, the target region is DNM2 H8A(-2+23). In certain embodiments, the target region is DNM2 H8A(+5+29). In certain embodiments, the target region is DNM2 H8A(+18+42). In certain embodiments, the target region is DNM2 H8A(+21+45). In certain embodiments, the target region is DNM2 H8A(+27+51). In certain embodiments, the target region is DNM2 H8A(+36+60). In certain embodiments, the target region is DNM2 H8A(+46+70). In certain embodiments, the target region is DNM2 H8A(+51+75). In certain embodiments, the target region is DNM2 H8A(+56+80).
In certain embodiments, the target region is DNM2 H8A(+65+88). In certain embodiments, the target region is DNM2 H8A(+76+100). In certain embodiments, the target region is DNM2 H8A(+83+107). In certain embodiments, the target region is DNM2 H8A(+96+119). In certain embodiments, the target region is DNM2 H8A(+105+129). In certain embodiments, the target region is DNM2 H8D(+18-7). In certain embodiments, the target region is DNM2 H8D(+10-15). In certain embodiments, the target region is DNM2 H9A(-12+13). In certain embodiments, the target region is DNM2 H9A(-5+20). In certain embodiments, the target region is DNM2 H9A(+1+25). In certain embodiments, the target region is DNM2 H9A(+6+30). In certain embodiments, the target region is DNM2 H9A(+16+40). In certain embodiments, the target region is DNM2 H9A(+25+49). In certain embodiments, the target region is DNM2 H9A(+36+60). In certain embodiments, the target region is DNM2 H9A(+37+61). In certain embodiments, the target region is DNM2 H9A(+42+66). In certain embodiments, the target region is DNM2 H9A(+43+62). In certain embodiments, the target region is DNM2 H9A(+43+63). In certain embodiments, the target region is DNM2 H9A(+43+64). In certain embodiments, the target region is DNM2 H9A(+43+65). In certain embodiments, the target region is DNM2 H9A(+43+66). In certain embodiments, the target region is DNM2 H9A(+43+67). In certain embodiments, the target region is DNM2 H9A(+44+67). In certain embodiments, the target region is DNM2 H9A(+44+68). In certain embodiments, the target region is DNM2 H9A(+45+67). In certain embodiments, the target region is DNM2 H9A (+46+67). In certain embodiments, the target region is DNM2 H9A(+47+67). In certain embodiments, the target region is DNM2 H9A(+48+67). In certain embodiments, the target region is DNM2 H9D(+24-1). In certain embodiments, the target region is DNM2 H9D(+23-2). In certain embodiments, the target region is DNM2 H9D(+22-3). In certain embodiments, the target region is DNM2 H9D(+21-4). In certain embodiments, the target region is DNM2 H9D(+20-5). In certain embodiments, the target region is DNM2 H9D(+19-6). In certain embodiments, the target region is DNM2 H9D (+18-7). In certain embodiments, the target region is DNM2 H9D(+17-8). In certain embodiments, the target region is DNM2 H9D(+16-9). In certain embodiments, the target region is DNM2 H9D(+15-10). In certain embodiments, the target region is DNM2 H9D(+14-11). In certain embodiments, the target region is DNM2 H9D(+13-12). In certain embodiments, the target region is DNM2 H9D(+12-13). In certain embodiments, the target region is DNM2 H9D(+11-14). In certain embodiments, the target region is DNM2 H9D(+10-15). In certain embodiments, the target region is DNM2 H9D(+9-16). In certain embodiments, the target region is DNM2 H9D(+8-12). In certain embodiments, the target region is DNM2 H9D(+8-13). In certain embodiments, the target region is DNM2 H9D(+8-14). In certain embodiments, the target region is DNM2 H9D(+8-15). In certain embodiments, the target region is DNM2 H9D(+8-17). In certain embodiments, the target
region is DNM2 H9D(+7-18). In certain embodiments, the target region is DNM2 H9D(+7-17). In certain embodiments, the target region is DNM2 H9D(+6-19). In certain embodiments, the target region is DNM2 H9D(+6-17). In certain embodiments, the target region is DNM2 H9D(+5-17). In certain embodiments, the target region is DNM2 H9D(+5-20). In certain embodiments, the target region is DNM2 H9D(+4-17). In certain embodiments, the target region is DNM2 H9D(+4-21). In certain embodiments, the target region is DNM2 H9D(+3-17). In certain embodiments, the target region is DNM2 H9D(+3-22). In certain embodiments, the target region is DNM2 H12A(-13+12). In certain embodiments, the target region is DNM2 H12A(-3+22). In certain embodiments, the target region is DNM2 H12A(+4+28). In certain embodiments, the target region is DNM2 H12A(+7+31). In certain embodiments, the target region is DNM2 H12A(+10+34). In certain embodiments, the target region is DNM2 H12A(+11+35). In certain embodiments, the target region is DNM2 H12A(+12+36). In certain embodiments, the target region is DNM2 H12A(+13+37). In certain embodiments, the target region is DNM2 H12A(+14+38). In certain embodiments, the target region is DNM2 H12A(+15+39). In certain embodiments, the target region is DNM2 H12A(+16+40). In certain embodiments, the target region is DNM2 H12A(+17+41). In certain embodiments, the target region is DNM2 H12A(+18+42). In certain embodiments, the target region is DNM2 H12A(+19+43). In certain embodiments, the target region is DNM2 H12A(+20+44). In certain embodiments, the target region is DNM2 H12A(+25+44). In certain embodiments, the target region is DNM2 H12A(+25+45). In certain embodiments, the target region is DNM2 H12A(+25+46). In certain embodiments, the target region is DNM2 H12A(+25+47). In certain embodiments, the target region is DNM2 H12A(+25+48). In certain embodiments, the target region is DNM2 H12A(+25+49). In certain embodiments, the target region is DNM2 H12A(+26+49). In certain embodiments, the target region is DNM2 H12A(+27+49). In certain embodiments, the target region is DNM2 H12A(+28+49). In certain embodiments, the target region is DNM2 H12A(+29+49). In certain embodiments, the target region is DNM2 H12A(+29+53). In certain embodiments, the target region is DNM2 H12A(+30+49). In certain embodiments, the target region is DNM2 H12A(+30+54). In certain embodiments, the target region is DNM2 H12A(+36+60). In certain embodiments, the target region is DNM2 H12A(+37+61). In certain embodiments, the target region is DNM2 H12A(+43+67). In certain embodiments, the target region is DNM2 H12D(+22- 3). In certain embodiments, the target region is DNM2 H12D(+20-5). In certain embodiments, the target region is DNM2 H12D(+15-10). In certain embodiments, the target region is DNM2 H13A(-27-3). In certain embodiments, the target region is DNM2 H13A(-26-2). In certain embodiments, the target region is DNM2 H13A(-25-1). In certain embodiments, the target region is DNM2 H13A(-24+1). In certain embodiments, the target region is DNM2 H13A(-23+2). In
certain embodiments, the target region is DNM2 H13A(-22+3). In certain embodiments, the target region is DNM2 H13A(-21+4). In certain embodiments, the target region is DNM2 H13A(- 20+5). In certain embodiments, the target region is DNM2 H13A(-19+6). In certain embodiments, the target region is DNM2 H13A(-18+7). In certain embodiments, the target region is DNM2 H13A(-17+8). In certain embodiments, the target region is DNM2 H13A(-16+9). In certain embodiments, the target region is DNM2 H13A(-15+10). In certain embodiments, the target region is DNM2 H13A(-14+11). In certain embodiments, the target region is DNM2 H13A(- 13+12). In certain embodiments, the target region is DNM2 H13A(-11+14). In certain embodiments, the target region is DNM2 H13A(-10+15). In certain embodiments, the target region is DNM2 H13A(-9+16). In certain embodiments, the target region is DNM2 H13A(-8+17). In certain embodiments, the target region is DNM2 H13A(-7+18). In certain embodiments, the target region is DNM2 H13A(-6+19). In certain embodiments, the target region is DNM2 H13A(- 5+20). In certain embodiments, the target region is DNM2 H13A(-4+21). In certain embodiments, the target region is DNM2 H13A(-3+22). In certain embodiments, the target region is DNM2 H13A(-2+23). In certain embodiments, the target region is DNM2 H13A(-1+24). In certain embodiments, the target region is DNM2 H13A(+1+25). In certain embodiments, the target region is DNM2 H13A(+2+25). In certain embodiments, the target region is DNM2 H13A(+3+27). In certain embodiments, the target region is DNM2 H13A(+4+23). In certain embodiments, the target region is DNM2 H13A(+4+24). In certain embodiments, the target region is DNM2 H13A(+4+25). In certain embodiments, the target region is DNM2 H13A(+4+26). In certain embodiments, the target region is DNM2 H13A(+4+27). In certain embodiments, the target region is DNM2 H13A(+4+28). In certain embodiments, the target region is DNM2 H13A(+5+28). In certain embodiments, the target region is DNM2 H13A(+6+28). In certain embodiments, the target region is DNM2 H13A(+6+30). In certain embodiments, the target region is DNM2 H13A(+7+28). In certain embodiments, the target region is DNM2 H13A(+8+28). In certain embodiments, the target region is DNM2 H13A(+9+28). In certain embodiments, the target region is DNM2 H13A(+16+40). In certain embodiments, the target region is DNM2 H13D(+22-3). In certain embodiments, the target region is DNM2 H13D(+7-18). In certain embodiments, the target region is DNM2 H16A(-21+3). In certain embodiments, the target region is DNM2 H16A(-13+12). In certain embodiments, the target region is DNM2 H16A(- 6+19). In certain embodiments, the target region is DNM2 H16A(+6+30). In certain embodiments, the target region is DNM2 H16A(+15+39). In certain embodiments, the target region is DNM2 H16A(+26+50). In certain embodiments, the target region is DNM2 H16A(+36+60). In certain embodiments, the target region is DNM2 H16A(+45+69). In certain embodiments, the target region is DNM2 H16A(+55+79). In certain embodiments, the target
region is DNM2 H16A(+66+90). In certain embodiments, the target region is DNM2 H16A(+75+99). In certain embodiments, the target region is DNM2 H16A(+82+106). In certain embodiments, the target region is DNM2 H16D(+19-6). In certain embodiments, the target region is DNM2 H16D(+11-14). In certain embodiments, the target region is DNM2 H19A(- 11+14). In certain embodiments, the target region is DNM2 H19A(-4+21). In certain embodiments, the target region is DNM2 H19A(+1+25). In certain embodiments, the target region is DNM2 H19A(+7+31). In certain embodiments, the target region is DNM2 H19A(+16+40). In certain embodiments, the target region is DNM2 H19A(+26+50). In certain embodiments, the target region is DNM2 H19A(+37+61). In certain embodiments, the target region is DNM2 H19A(+47+71). In certain embodiments, the target region is DNM2 H19A(+56+80). In certain embodiments, the target region is DNM2 H19A(+66+90). In certain embodiments, the target region is DNM2 H19A(+76+100). In certain embodiments, the target region is DNM2 H19A(+86+110). In certain embodiments, the target region is DNM2 H19A(+96+120). In certain embodiments, the target region is DNM2 H19A(+106+130). In certain embodiments, the target region is DNM2 H19A(+117+141). In certain embodiments, the target region is DNM2 H19A(+127+151). In certain embodiments, the target region is DNM2 H19A(+137+161). In certain embodiments, the target region is DNM2 H19A(+147+171). In certain embodiments, the target region is DNM2 H19A(+157+181). In certain embodiments, the target region is DNM2 H19A(+166+189). In certain embodiments, the target region is DNM2 H19A(+191+215). In certain embodiments, the target region is DNM2 H19A(+200+224). In certain embodiments, the target region is DNM2 H19D(+23-2). In certain embodiments, the target region is DNM2 H19D(+17-8). In certain embodiments, the target region is DNM2 H19D(+10-15). [0137] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGAGGCCGC); SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAA); SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC);
SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT); SEQ ID NO: 90 (GAACCTCAGGCCGCCCCATTTTACC); SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT); SEQ ID NO: 96 (CCTGCTGAACCTCAGGCCGCCCCAT); SEQ ID NO: 97 (GTAGCCTGTGGGGAGAGGCAGTCAG); SEQ ID NO: 98 (TTCACCACGCCAATGTAGCCTGTGG); SEQ ID NO: 99 (GCGGTTCACCACGCCAATGTAGCCT); SEQ ID NO: 100 (GCTGCGGTTCACCACGCCAATGTA); SEQ ID NO: 101 (TCCTTCTGGCTGCGGTTCACCACGC); SEQ ID NO: 102 (ATATCCTTCTGGCTGCGGTTCACCA); SEQ ID NO: 103 (CTTGCCCTCAATATCCTTCTGGCTG); SEQ ID NO: 104 (GATGTCCTTCTTGCCCTCAATATCC); SEQ ID NO: 105 (ACGGATGTCCTTCTTGCCCTCAATA); SEQ ID NO: 106 (CAGTGCTGCACGGATGTCCTTCTTG); SEQ ID NO: 107 (TCTCAGCTGCCAGTGCTGCACGGAT); SEQ ID NO: 108 (ACTTCCTCTCAGCTGCCAGTGCTGC); SEQ ID NO: 109 (AGAACTTCCTCTCAGCTGCCAGTGC); SEQ ID NO: 110 (TGCCGGTAGGCCGGGTGGGAGAGGA); SEQ ID NO: 111 (GGCCATGTGCCGGTAGGCCGGGTG); SEQ ID NO: 112 (CCATGCGGTCGGCCATGTGCCGGTA);
SEQ ID NO: 113 (TGCAGATGTGGCGTGCCCATGCGGT); SEQ ID NO: 114 (AGCGTCTTCTGCAGATGTGGCGTGC); SEQ ID NO: 115 (TCAGCGTCTTCTGCAGATGTGGCGT); SEQ ID NO: 116 (GTGCAAACCCTTGCAGTACCTGATT); SEQ ID NO: 117 (CTTGCAGTACCTGATTCAGCGTCTT); SEQ ID NO: 118 (TGCAGTACCTGATTCAGCGTCTTCT); SEQ ID NO: 119 (TGGTCAGTTGCTGCAGGAGAGAGGT); SEQ ID NO: 120 (CGGATGTGGTTGGTCAGTTGCTGCA); SEQ ID NO: 121 (ACTCCCGGATGTGGTTGGTCAGTTG); SEQ ID NO: 122 (CAGCGACTCCCGGATGTGGTTGGTC); SEQ ID NO: 123 (GCCGGCAGCGACTCCCGGATGTGGT); SEQ ID NO: 124 (TGCTACGTAGGGCCGGCAGCGACTC); SEQ ID NO: 125 (TGTAGTTTGCTACGTAGGGCCGGCA); SEQ ID NO: 126 (AGCAGCTGGCTCTGTAGTTTGCTAC); SEQ ID NO: 127 (CTCCAGGGACAGCAGCTGGCTCTGT); SEQ ID NO: 128 (CTCCTTCTCCAGGGACAGCAGCTGG); SEQ ID NO: 129 (TCCACCTCCTTCTCCAGGGACAGCA); SEQ ID NO: 130 (CTTGTACTCCTCCACCTCCTTCTCC); SEQ ID NO: 131 (TTCTTGTACTCCTCCACCTCCTTCT); SEQ ID NO: 132 (CCGAAAGTTCTTGTACTCCTCCACC); SEQ ID NO: 133 (GGTCGTCGGGCCGAAAGTTCTTGTA); SEQ ID NO: 134 (GCTTTGGTTTTGCGGGTGGGGTCGT); SEQ ID NO: 135 (GTGCCGBGGTACATACTGCAGCAG); SEQ ID NO: 136 (GTGCCGGGGTACATACTGCAGCAGG); SEQ ID NO: 137 (ATACTGCAGCAGGGCTTTGGTTTTG); SEQ ID NO: 138 (CCATCCTATGAGGAAAAGGAAAAAG); SEQ ID NO: 139 (GCTGGACCATCCTATGAGGAAAAGG); SEQ ID NO: 140 (CCAAACTGCTGGACCATCCTATGAG); SEQ ID NO: 141 (CCCAAACTGCTGGACCATCCTATGA); SEQ ID NO: 142 (CCACCCCAAACTGCTGGACCATCCT); SEQ ID NO: 143 (CAAAATCCACCCCAAACTGCTGGAC); SEQ ID NO: 144 (TCGATCCTCTTCTCAAAATCCACCC); SEQ ID NO: 145 (CCCTCGATCCTCTTCTCAAAATCCA); SEQ ID NO: 146 (CCTGAGCCCTCGATCCTCTTCTCAA); SEQ ID NO: 147 (ACCTGATCTCCTGAGCCCTCGATCC);
SEQ ID NO: 148 (CAGAGTGTCCACCTGATCTCCTGAG); SEQ ID NO: 149 (AGCTCCAGAGTGTCCACCTGATCTC); SEQ ID NO: 150 (CGGAGAGCTCCAGAGTGTCCACCTG); SEQ ID NO: 151 (GGCGCCCCCGGAGAGCTCCAGAGT); SEQ ID NO: 152 (GCGATTGATTCGGGCGCCCCCGGAG); SEQ ID NO: 153 (GGAAGATGCGATTGATTCGGGCGCC); SEQ ID NO: 154 (GGAACCGCTCGTGGAAGATGCGAT); SEQ ID NO: 155 (AGCTCAAATGGGAACCGCTCGTGGA); SEQ ID NO: 156 (GCACTACCTTCACCAGCTCAAATGG); SEQ ID NO: 157 (CCCGGGGGGCACTACCTTCACCAGC); SEQ ID NO: 158 (CGTCAAACTCCATCTGAGAAGAGAC); SEQ ID NO: 159 (TCCTTCTCGTCAAACTCCATCTGAG); SEQ ID NO: 160 (GTAAGTCCTTCTCGTCAAACTCCAT); SEQ ID NO: 161 (CCGTCGTAAGTCCTTCTCGTCAAAC); SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG); SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT); SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 175 (TGACTCCATGGATGTTCTTAATG); SEQ ID NO: 176 (TGACTCCATGGATGTTCTTAAT); SEQ ID NO: 177 (TGACTCCATGGATGTTCTTAA); SEQ ID NO: 178 (TGACTCCATGGATGTTCTTA); SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA);
SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT); SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG); SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT); SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 208 (CTCCTCGTGGAACTTGCCTG); SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG); SEQ ID NO: 210 (CAGCAGAAGAATCTGGBGGAGAGCG); SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG); SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT);
SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA); SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG); SEQ ID NO: 238 (GATGAAGTCCTCATGGTTCGTGTTG); SEQ ID NO: 239 (CGATGAAGTCCTCATGGTTCGTGTT); SEQ ID NO: 240 (CAAACCCGATGAAGTCCTCATGGTT); SEQ ID NO: 241 (TACTTGGCAAACCCGATGAAGTCCT); SEQ ID NO: 242 (CCTACTTGGCAAACCCGATGAAGTC); SEQ ID NO: 243 (AAGTACCTACTTGGCAAACCCGATG); SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC);
SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA); SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC); SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 282 (GTTCAGCTGCGTGCTCCTCTGC); SEQ ID NO: 283 (GTTCAGCTGCGTGCTCCTCTG); SEQ ID NO: 284 (GTTCAGCTGCGTGCTCCTCT); SEQ ID NO: 285 (GGCTCTCTTCTTGTTCAGCTGCGTG); SEQ ID NO: 286 (TACCTGATTGGBGATGGCTCTCTTC); SEQ ID NO: 287 (TGCAGAGGBGTGTGCTACCTGATTG);
SEQ ID NO: 288 (CTCCTGAGGGTGGAGGGAAAAGAG); SEQ ID NO: 289 (CTTCTCTTTCTCCTGAGGGTGGAGG); SEQ ID NO: 290 (TGTACTTCTTCTCTTTCTCCTGAGG); SEQ ID NO: 291 (CAGAGGCAGCATGTACTTCTTCTCT); SEQ ID NO: 292 (GAGGTTGTCCAGAGGCAGCATGTAC); SEQ ID NO: 293 (TCACGGATCTTGAGGTTGTCCAGAG); SEQ ID NO: 294 (CTTCTCCACATCACGGATCTTGAGG); SEQ ID NO: 295 (CATGAAGCCCTTCTCCACATCACGG); SEQ ID NO: 296 (GCTTGTTGGACATGAAGCCCTTCTC); SEQ ID NO: 297 (GGCGAAGACGTGCTTGTTGGACATG); SEQ ID NO: 298 (GTTGAAGATGGCGAAGACGTGCTTG); SEQ ID NO: 299 (GCTCCGTGTTGAAGATGGCGAAGAC); SEQ ID NO: 300 (CCTCACCTCTGCTCCGTGTTGAAGA); SEQ ID NO: 301 (CCTGGGCCCCTCACCTCTGCTCCGT); SEQ ID NO: 302 (ATGAAGGCCTTCGTCTGCGGGCAGG); SEQ ID NO: 303 (GTGGTGGATGAAGGCCTTCGTCTGC); SEQ ID NO: 304 (GCTCGTGGTGGATGAAGGCCTTCGT); SEQ ID NO: 305 (CCAGCAGCTCGTGGTGGATGAAGGC); SEQ ID NO: 306 (ATAGGTAGGCCAGCAGCTCGTGGTG); SEQ ID NO: 307 (GCCGAGGAGTATAGGTAGGCCAGCA); SEQ ID NO: 308 (TGCTCTGGTCTGCCGAGGAGTATAG); SEQ ID NO: 309 (TCCATGAGGCTGCTCTGGTCTGCCG); SEQ ID NO: 310 (GCCGACTCCTCCATGAGGCTGCTCT); SEQ ID NO: 311 (TGCCTGGTCAGCCGACTCCTCCATG); SEQ ID NO: 312 (CCCGCCGCTGTGCCTGGTCAGCCGA); SEQ ID NO: 313 (AGCATGTCGTCCCGCCGCTGTGCCT); SEQ ID NO: 314 (GTACATGCGCAGCATGTCGTCCCGC); SEQ ID NO: 315 (TGAGGGCATGGTACATGCGCAGCAT); SEQ ID NO: 316 (GAGCGCCTCCTTGAGGGCATGGTAC); SEQ ID NO: 317 (CGATGATGTTGAGCGCCTCCTTGAG); SEQ ID NO: 318 (CTGATGTCACCGATGATGTTGAGCG); SEQ ID NO: 319 (AGTGCTGGTGCTGATGTCACCGATG); SEQ ID NO: 320 (GCGTGGACACAGTGCTGGTGCTGAT); SEQ ID NO: 321 (GGGTACAGGCGTGGACACAGTGCT); SEQ ID NO: 322 (TGGAGCCAGGTGTCATCGACAGGCG);
SEQ ID NO: 323 (CTGGCGCTCTGGAGCCAGGTGTCAT); SEQ ID NO: 324 (ACCTGTGGCTGCTGGCGCTCTGGAG); SEQ ID NO: 325 (TTCCGGACCTGTGGCTGCTGGCGCT); and SEQ ID NO: 326 (ACCAGGCTTCCGGACCTGTGGCTGC), wherein “B” represents an abasic subunit or guanine. In certain embodiments, B is an abasic subunit. [0138] In certain embodiments, the targeting sequence is SEQ ID NO: 67. In certain embodiments, the targeting sequence is SEQ ID NO: 68. In certain embodiments, the targeting sequence is SEQ ID NO: 69. In certain embodiments, the targeting sequence is SEQ ID NO: 70. [0139] In certain embodiments, the targeting sequence is SEQ ID NO: 71. In certain embodiments, the targeting sequence is SEQ ID NO: 72. In certain embodiments, the targeting sequence is SEQ ID NO: 73. In certain embodiments, the targeting sequence is SEQ ID NO: 74. In certain embodiments, the targeting sequence is SEQ ID NO: 75. In certain embodiments, the targeting sequence is SEQ ID NO: 76. In certain embodiments, the targeting sequence is SEQ ID NO: 77. In certain embodiments, the targeting sequence is SEQ ID NO: 78. In certain embodiments, the targeting sequence is SEQ ID NO: 79. In certain embodiments, the targeting sequence is SEQ ID NO: 80. In certain embodiments, the targeting sequence is SEQ ID NO: 81. In certain embodiments, the targeting sequence is SEQ ID NO: 82. In certain embodiments, the targeting sequence is SEQ ID NO: 83. In certain embodiments, the targeting sequence is SEQ ID NO: 84. In certain embodiments, the targeting sequence is SEQ ID NO: 85. In certain embodiments, the targeting sequence is SEQ ID NO: 86. In certain embodiments, the targeting sequence is SEQ ID NO: 87. In certain embodiments, the targeting sequence is SEQ ID NO: 88. In certain embodiments, the targeting sequence is SEQ ID NO: 89. In certain embodiments, the targeting sequence is SEQ ID NO: 90. In certain embodiments, the targeting sequence is SEQ ID NO: 91. In certain embodiments, the targeting sequence is SEQ ID NO: 92. In certain embodiments, the targeting sequence is SEQ ID NO: 93. In certain embodiments, the targeting sequence is SEQ ID NO: 94. In certain embodiments, the targeting sequence is SEQ ID NO: 95. In certain embodiments, the targeting sequence is SEQ ID NO: 96. In certain embodiments, the targeting sequence is SEQ ID NO: 97. In certain embodiments, the targeting sequence is SEQ ID NO: 98. In certain embodiments, the targeting sequence is SEQ ID NO: 99. In certain embodiments, the targeting sequence is SEQ ID NO: 100. In certain embodiments, the targeting sequence is SEQ ID NO: 101. In certain embodiments, the targeting sequence is SEQ ID NO: 102. In certain embodiments, the targeting sequence is SEQ ID NO: 103. In certain embodiments, the targeting sequence is SEQ ID NO: 104. In certain embodiments, the targeting sequence is SEQ ID NO: 105. In certain embodiments, the targeting sequence is SEQ ID NO:
106. In certain embodiments, the targeting sequence is SEQ ID NO: 107. In certain embodiments, the targeting sequence is SEQ ID NO: 108. In certain embodiments, the targeting sequence is SEQ ID NO: 109. In certain embodiments, the targeting sequence is SEQ ID NO: 110. In certain embodiments, the targeting sequence is SEQ ID NO: 111. In certain embodiments, the targeting sequence is SEQ ID NO: 112. In certain embodiments, the targeting sequence is SEQ ID NO: 113. In certain embodiments, the targeting sequence is SEQ ID NO: 114. In certain embodiments, the targeting sequence is SEQ ID NO: 115. In certain embodiments, the targeting sequence is SEQ ID NO: 116. In certain embodiments, the targeting sequence is SEQ ID NO: 117. In certain embodiments, the targeting sequence is SEQ ID NO: 118. In certain embodiments, the targeting sequence is SEQ ID NO: 119. In certain embodiments, the targeting sequence is SEQ ID NO: 120. In certain embodiments, the targeting sequence is SEQ ID NO: 121. In certain embodiments, the targeting sequence is SEQ ID NO: 122. In certain embodiments, the targeting sequence is SEQ ID NO: 123. In certain embodiments, the targeting sequence is SEQ ID NO: 124. In certain embodiments, the targeting sequence is SEQ ID NO: 125. In certain embodiments, the targeting sequence is SEQ ID NO: 126. In certain embodiments, the targeting sequence is SEQ ID NO: 127. In certain embodiments, the targeting sequence is SEQ ID NO: 128. In certain embodiments, the targeting sequence is SEQ ID NO: 129. In certain embodiments, the targeting sequence is SEQ ID NO: 130. In certain embodiments, the targeting sequence is SEQ ID NO: 131. In certain embodiments, the targeting sequence is SEQ ID NO: 132. In certain embodiments, the targeting sequence is SEQ ID NO: 133. In certain embodiments, the targeting sequence is SEQ ID NO: 134. In certain embodiments, the targeting sequence is SEQ ID NO: 135. In certain embodiments, the targeting sequence is SEQ ID NO: 136. In certain embodiments, the targeting sequence is SEQ ID NO: 137. In certain embodiments, the targeting sequence is SEQ ID NO: 138. In certain embodiments, the targeting sequence is SEQ ID NO: 139. In certain embodiments, the targeting sequence is SEQ ID NO: 140. In certain embodiments, the targeting sequence is SEQ ID NO: 141. In certain embodiments, the targeting sequence is SEQ ID NO: 142. In certain embodiments, the targeting sequence is SEQ ID NO: 143. In certain embodiments, the targeting sequence is SEQ ID NO: 144. In certain embodiments, the targeting sequence is SEQ ID NO: 145. In certain embodiments, the targeting sequence is SEQ ID NO: 146. In certain embodiments, the targeting sequence is SEQ ID NO: 147. In certain embodiments, the targeting sequence is SEQ ID NO: 148. In certain embodiments, the targeting sequence is SEQ ID NO: 149. In certain embodiments, the targeting sequence is SEQ ID NO: 150. In certain embodiments, the targeting sequence is SEQ ID NO: 151. In certain embodiments, the targeting sequence is SEQ ID NO: 152. In certain embodiments, the targeting
sequence is SEQ ID NO: 153. In certain embodiments, the targeting sequence is SEQ ID NO: 154. In certain embodiments, the targeting sequence is SEQ ID NO: 155. In certain embodiments, the targeting sequence is SEQ ID NO: 156. In certain embodiments, the targeting sequence is SEQ ID NO: 157. In certain embodiments, the targeting sequence is SEQ ID NO: 158. In certain embodiments, the targeting sequence is SEQ ID NO: 159. In certain embodiments, the targeting sequence is SEQ ID NO: 160. In certain embodiments, the targeting sequence is SEQ ID NO: 161. In certain embodiments, the targeting sequence is SEQ ID NO: 162. In certain embodiments, the targeting sequence is SEQ ID NO: 163. In certain embodiments, the targeting sequence is SEQ ID NO: 164. In certain embodiments, the targeting sequence is SEQ ID NO: 165. In certain embodiments, the targeting sequence is SEQ ID NO: 166. In certain embodiments, the targeting sequence is SEQ ID NO: 167. In certain embodiments, the targeting sequence is SEQ ID NO: 168. In certain embodiments, the targeting sequence is SEQ ID NO: 169. In certain embodiments, the targeting sequence is SEQ ID NO: 170. In certain embodiments, the targeting sequence is SEQ ID NO: 171. In certain embodiments, the targeting sequence is SEQ ID NO: 172. In certain embodiments, the targeting sequence is SEQ ID NO: 173. In certain embodiments, the targeting sequence is SEQ ID NO: 174. In certain embodiments, the targeting sequence is SEQ ID NO: 175. In certain embodiments, the targeting sequence is SEQ ID NO: 176. In certain embodiments, the targeting sequence is SEQ ID NO: 177. In certain embodiments, the targeting sequence is SEQ ID NO: 178. In certain embodiments, the targeting sequence is SEQ ID NO: 179. In certain embodiments, the targeting sequence is SEQ ID NO: 180. In certain embodiments, the targeting sequence is SEQ ID NO: 181. In certain embodiments, the targeting sequence is SEQ ID NO: 182. In certain embodiments, the targeting sequence is SEQ ID NO: 183. In certain embodiments, the targeting sequence is SEQ ID NO: 184. In certain embodiments, the targeting sequence is SEQ ID NO: 185. In certain embodiments, the targeting sequence is SEQ ID NO: 186. In certain embodiments, the targeting sequence is SEQ ID NO: 187. In certain embodiments, the targeting sequence is SEQ ID NO: 188. In certain embodiments, the targeting sequence is SEQ ID NO: 189. In certain embodiments, the targeting sequence is SEQ ID NO: 190. In certain embodiments, the targeting sequence is SEQ ID NO: 191. In certain embodiments, the targeting sequence is SEQ ID NO: 192. In certain embodiments, the targeting sequence is SEQ ID NO: 193. In certain embodiments, the targeting sequence is SEQ ID NO: 194. In certain embodiments, the targeting sequence is SEQ ID NO: 195. In certain embodiments, the targeting sequence is SEQ ID NO: 196. In certain embodiments, the targeting sequence is SEQ ID NO: 197. In certain embodiments, the targeting sequence is SEQ ID NO: 198. In certain embodiments, the targeting sequence is SEQ ID NO: 199. In certain
embodiments, the targeting sequence is SEQ ID NO: 200. In certain embodiments, the targeting sequence is SEQ ID NO: 201. In certain embodiments, the targeting sequence is SEQ ID NO: 202. In certain embodiments, the targeting sequence is SEQ ID NO: 203. In certain embodiments, the targeting sequence is SEQ ID NO: 204. In certain embodiments, the targeting sequence is SEQ ID NO: 205. In certain embodiments, the targeting sequence is SEQ ID NO: 206. In certain embodiments, the targeting sequence is SEQ ID NO: 207. In certain embodiments, the targeting sequence is SEQ ID NO: 208. In certain embodiments, the targeting sequence is SEQ ID NO: 209. In certain embodiments, the targeting sequence is SEQ ID NO: 210. In certain embodiments, the targeting sequence is SEQ ID NO: 211. In certain embodiments, the targeting sequence is SEQ ID NO: 212. In certain embodiments, the targeting sequence is SEQ ID NO: 213. In certain embodiments, the targeting sequence is SEQ ID NO: 214. In certain embodiments, the targeting sequence is SEQ ID NO: 215. In certain embodiments, the targeting sequence is SEQ ID NO: 216. In certain embodiments, the targeting sequence is SEQ ID NO: 217. In certain embodiments, the targeting sequence is SEQ ID NO: 218. In certain embodiments, the targeting sequence is SEQ ID NO: 219. In certain embodiments, the targeting sequence is SEQ ID NO: 220. In certain embodiments, the targeting sequence is SEQ ID NO: 221. In certain embodiments, the targeting sequence is SEQ ID NO: 222. In certain embodiments, the targeting sequence is SEQ ID NO: 223. In certain embodiments, the targeting sequence is SEQ ID NO: 224. In certain embodiments, the targeting sequence is SEQ ID NO: 225. In certain embodiments, the targeting sequence is SEQ ID NO: 226. In certain embodiments, the targeting sequence is SEQ ID NO: 227. In certain embodiments, the targeting sequence is SEQ ID NO: 228. In certain embodiments, the targeting sequence is SEQ ID NO: 229. In certain embodiments, the targeting sequence is SEQ ID NO: 230. In certain embodiments, the targeting sequence is SEQ ID NO: 231. In certain embodiments, the targeting sequence is SEQ ID NO: 232. In certain embodiments, the targeting sequence is SEQ ID NO: 233. In certain embodiments, the targeting sequence is SEQ ID NO: 234. In certain embodiments, the targeting sequence is SEQ ID NO: 235. In certain embodiments, the targeting sequence is SEQ ID NO: 236. In certain embodiments, the targeting sequence is SEQ ID NO: 237. In certain embodiments, the targeting sequence is SEQ ID NO: 238. In certain embodiments, the targeting sequence is SEQ ID NO: 239. In certain embodiments, the targeting sequence is SEQ ID NO: 240. In certain embodiments, the targeting sequence is SEQ ID NO: 241. In certain embodiments, the targeting sequence is SEQ ID NO: 242. In certain embodiments, the targeting sequence is SEQ ID NO: 243. In certain embodiments, the targeting sequence is SEQ ID NO: 244. In certain embodiments, the targeting sequence is SEQ ID NO: 245. In certain embodiments, the targeting sequence is SEQ ID NO:
246. In certain embodiments, the targeting sequence is SEQ ID NO: 247. In certain embodiments, the targeting sequence is SEQ ID NO: 248. In certain embodiments, the targeting sequence is SEQ ID NO: 249. In certain embodiments, the targeting sequence is SEQ ID NO: 250. In certain embodiments, the targeting sequence is SEQ ID NO: 251. In certain embodiments, the targeting sequence is SEQ ID NO: 252. In certain embodiments, the targeting sequence is SEQ ID NO: 253. In certain embodiments, the targeting sequence is SEQ ID NO: 254. In certain embodiments, the targeting sequence is SEQ ID NO: 255. In certain embodiments, the targeting sequence is SEQ ID NO: 256. In certain embodiments, the targeting sequence is SEQ ID NO: 257. In certain embodiments, the targeting sequence is SEQ ID NO: 258. In certain embodiments, the targeting sequence is SEQ ID NO: 259. In certain embodiments, the targeting sequence is SEQ ID NO: 260. In certain embodiments, the targeting sequence is SEQ ID NO: 261. In certain embodiments, the targeting sequence is SEQ ID NO: 262. In certain embodiments, the targeting sequence is SEQ ID NO: 263. In certain embodiments, the targeting sequence is SEQ ID NO: 264. In certain embodiments, the targeting sequence is SEQ ID NO: 265. In certain embodiments, the targeting sequence is SEQ ID NO: 266. In certain embodiments, the targeting sequence is SEQ ID NO: 267. In certain embodiments, the targeting sequence is SEQ ID NO: 268. In certain embodiments, the targeting sequence is SEQ ID NO: 269. In certain embodiments, the targeting sequence is SEQ ID NO: 270. In certain embodiments, the targeting sequence is SEQ ID NO: 271. In certain embodiments, the targeting sequence is SEQ ID NO: 272. In certain embodiments, the targeting sequence is SEQ ID NO: 273. In certain embodiments, the targeting sequence is SEQ ID NO: 274. In certain embodiments, the targeting sequence is SEQ ID NO: 275. In certain embodiments, the targeting sequence is SEQ ID NO: 276. In certain embodiments, the targeting sequence is SEQ ID NO: 277. In certain embodiments, the targeting sequence is SEQ ID NO: 278. In certain embodiments, the targeting sequence is SEQ ID NO: 279. In certain embodiments, the targeting sequence is SEQ ID NO: 280. In certain embodiments, the targeting sequence is SEQ ID NO: 281. In certain embodiments, the targeting sequence is SEQ ID NO: 282. In certain embodiments, the targeting sequence is SEQ ID NO: 283. In certain embodiments, the targeting sequence is SEQ ID NO: 284. In certain embodiments, the targeting sequence is SEQ ID NO: 285. In certain embodiments, the targeting sequence is SEQ ID NO: 286. In certain embodiments, the targeting sequence is SEQ ID NO: 287. In certain embodiments, the targeting sequence is SEQ ID NO: 288. In certain embodiments, the targeting sequence is SEQ ID NO: 289. In certain embodiments, the targeting sequence is SEQ ID NO: 290. In certain embodiments, the targeting sequence is SEQ ID NO: 291. In certain embodiments, the targeting sequence is SEQ ID NO: 292. In certain embodiments, the targeting
sequence is SEQ ID NO: 293. In certain embodiments, the targeting sequence is SEQ ID NO: 294. In certain embodiments, the targeting sequence is SEQ ID NO: 295. In certain embodiments, the targeting sequence is SEQ ID NO: 296. In certain embodiments, the targeting sequence is SEQ ID NO: 297. In certain embodiments, the targeting sequence is SEQ ID NO: 298. In certain embodiments, the targeting sequence is SEQ ID NO: 299. In certain embodiments, the targeting sequence is SEQ ID NO: 300. In certain embodiments, the targeting sequence is SEQ ID NO: 301. In certain embodiments, the targeting sequence is SEQ ID NO: 302. In certain embodiments, the targeting sequence is SEQ ID NO: 303. In certain embodiments, the targeting sequence is SEQ ID NO: 304. In certain embodiments, the targeting sequence is SEQ ID NO: 305. In certain embodiments, the targeting sequence is SEQ ID NO: 306. In certain embodiments, the targeting sequence is SEQ ID NO: 307. In certain embodiments, the targeting sequence is SEQ ID NO: 308. In certain embodiments, the targeting sequence is SEQ ID NO: 309. In certain embodiments, the targeting sequence is SEQ ID NO: 310. In certain embodiments, the targeting sequence is SEQ ID NO: 311. In certain embodiments, the targeting sequence is SEQ ID NO: 312. In certain embodiments, the targeting sequence is SEQ ID NO: 313. In certain embodiments, the targeting sequence is SEQ ID NO: 314. In certain embodiments, the targeting sequence is SEQ ID NO: 315. In certain embodiments, the targeting sequence is SEQ ID NO: 316. In certain embodiments, the targeting sequence is SEQ ID NO: 317. In certain embodiments, the targeting sequence is SEQ ID NO: 318. In certain embodiments, the targeting sequence is SEQ ID NO: 319. In certain embodiments, the targeting sequence is SEQ ID NO: 320. In certain embodiments, the targeting sequence is SEQ ID NO: 321. In certain embodiments, the targeting sequence is SEQ ID NO: 322. In certain embodiments, the targeting sequence is SEQ ID NO: 323. In certain embodiments, the targeting sequence is SEQ ID NO: 324. In certain embodiments, the targeting sequence is SEQ ID NO: 325. In certain embodiments, the targeting sequence is SEQ ID NO: 326. [0140] In certain embodiments, the target region is within the exon internal region of exon 2 (SEQ ID NO: 2), the intron 1/exon 2 junction, the exon 2/intron 2 junction, or the intron internal region of intron 2 (SEQ ID NO: 11) of the human DNM2 gene pre-mRNA. In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 21st nucleotide to the 47th nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 13. In certain embodiments, the target region is a region within DNM2 H2A(+21+47). [0141] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 23rd nucleotide to the 51st nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID
NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 14. In certain embodiments, the target region is a region within DNM2 H2A(+23+51). [0142] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 46th nucleotide to the 72nd nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 15. In certain embodiments, the target region is a region within DNM2 H2A(+46+72). [0143] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 13th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 24th nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 16. In certain embodiments, the target region is a region within DNM2 H2D(+13-24). [0144] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 13th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 14th nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 17. In certain embodiments, the target region is a region within DNM2 H2D(+13-14). [0145] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 19th nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 18. In certain embodiments, the target region is a region within DNM2 H2D(+10-19). [0146] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 18th nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 19. In certain embodiments, the target region is a region within DNM2 H2D(+9-18). [0147] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23rd nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 20. In certain embodiments, the target region is a region within DNM2 H2D(+6-23). [0148] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 4th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23rd nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target
region is a region within SEQ ID NO: 21. In certain embodiments, the target region is a region within DNM2 H2D(+4-23). [0149] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 30th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 22. In certain embodiments, the target region is a region within DNM2 H2D(-1-30). [0150] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 29th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 23. In certain embodiments, the target region is a region within DNM2 H2D(-1-29). [0151] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12- 13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3- 27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30). [0152] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9- 16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23) , DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6- 30). [0153] In certain embodiments, the target region is DNM2 H2A(+21+45) or DNM2 H2A(+23+47). In certain embodiments, the target region is DNM2 H2A(+23+47) or DNM2 H2A(+27+51). In certain embodiments, the target region is DNM2 H2A(+46+70) or DNM2 H2A(+48+72). [0154] In certain embodiments, the target region is DNM2 H2D(+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), or DNM2 H2D(+1-24). In certain embodiments, the target region is DNM2 H2D (+13-12), DNM2 H2D(+12-13), or DNM2 H2D(+11-14). [0155] In certain embodiments, the target region is DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), or DNM2 H2D(+6-19). In certain embodiments, the
target region is DNM2 H2D(+9-16), DNM2 H2D(+8-17), or DNM2 H2D(+7-18). In certain embodiments, the target region is DNM2 H2D(+8-17). [0156] In certain embodiments, the target region is DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23). In certain embodiments, the target region is DNM2 H2D(+4-21), DNM2 H2D(+3-22), or DNM2 H2D(+2-23). In certain embodiments, the target region is DNM2 H2D(+3-22). [0157] In certain embodiments, the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30). In certain embodiments, the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), or DNM2 H2D(-5-29). [0158] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(-7+17). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H2A(+3+27). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+15+37). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+37+61). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2D(+20-5). [0159] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGAGGCCGC); SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAA); SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC); SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT);
SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT); SEQ ID NO: 90 (GAACCTCAGGCCGCCCCATTTTACC); SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT); and SEQ ID NO: 96 (CCTGCTGAACCTCAGGCCGCCCCAT), wherein “B” represents an abasic subunit or guanine. In certain embodiments, B is an abasic subunit. [0160] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT); SEQ ID NO: 90 (GAACCTCAGGCCGCCCCATTTTACC); SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT); and SEQ ID NO: 96 (CCTGCTGAACCTCAGGCCGCCCCAT).
[0161] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); and SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC). [0162] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); and SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA). [0163] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); and SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAA). [0164] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT); and SEQ ID NO: 90 (GAACCTCAGGCCGCCCCATTTTACC). [0165] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); and SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG). [0166] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); and SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT). [0167] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG);
SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); and SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT). [0168] In certain embodiments, the targeting sequence is SEQ ID NO: 83. [0169] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); and SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT). [0170] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); and SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT). [0171] In certain embodiments, the targeting sequence is SEQ ID NO: 88. [0172] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT); and SEQ ID NO: 96 (CCTGCTGAACCTCAGGCCGCCCCAT). [0173] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); and SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT). [0174] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGAGGCCGC); and SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC), wherein “B” represents an abasic subunit or guanine. In certain embodiments, B is an abasic subunit.
[0175] In certain embodiments, the target region is within the exon internal region of exon 9 (SEQ ID NO: 6), the intron 8/exon 9 junction, or the exon 9/intron 9 junction of the human DNM2 gene pre-mRNA. [0176] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 36th nucleotide to the 61st nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6). In certain embodiments, the target region is a region within SEQ ID NO: 24. In certain embodiments, the target region is a region within DNM2 H9A(+36+61). [0177] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 42nd nucleotide to the 68th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6). In certain embodiments, the target region is a region within SEQ ID NO: 25. In certain embodiments, the target region is a region within DNM2 H9A(+42+70). [0178] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 43rd nucleotide to the 68th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6). In certain embodiments, the target region is a region within SEQ ID NO: 26. In certain embodiments, the target region is a region within DNM2 H9A(+42+68). [0179] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 24th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 27. In certain embodiments, the target region is a region within DNM2 H9D(+24-15). [0180] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 24th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 5th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 28. In certain embodiments, the target region is a region within DNM2 H9D(+24-5). [0181] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 19th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 10th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 29. In certain embodiments, the target region is a region within DNM2 H9D(+19-10). [0182] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 14th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 30. In certain embodiments, the target region is a region within DNM2 H9D(+14-15).
[0183] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22nd nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 31. In certain embodiments, the target region is a region within DNM2 H9D(+9-22). [0184] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 20th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 32. In certain embodiments, the target region is a region within DNM2 H9D(+9-20). [0185] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 18th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 33. In certain embodiments, the target region is a region within DNM2 H9D(+9-18). [0186] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 8th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 21st nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 34. In certain embodiments, the target region is a region within DNM2 H9D(+8-21). [0187] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22nd nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 35. In certain embodiments, the target region is a region within DNM2 H9D(+6-22). [0188] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13), DNM2 H9A(-5+20), DNM2 H9A(+1+25),DNM2 H9A(+6+30), DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), DNM2 H9A(+48+67), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11- 14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2
H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4- 17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22). [0189] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+43+67), DNM2 H9A(+42+66), DNM2 H9A(+44+68), DNM2 H9D(+24-1), DNM2 H9D(+23- 2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+6-19), DNM2 H9D(+5-20), DNM2 H9D(+4-21), or DNM2 H9D(+3-22). [0190] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+36+60) or DNM2 H9A(+37+61). [0191] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), or DNM2 H9A(+48+67). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), or DNM2 H9A(+44+68). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+67). [0192] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11- 14), or DNM2 H9D(+10-15). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), or DNM2 H9D(+20-5). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), or DNM2 H9D(+15-10). [0193] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), or DNM2 H9D(+10-15). [0194] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15),
DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6- 17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22). [0195] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6- 17), DNM2 H9D(+5-17), or DNM2 H9D(+5-20). [0196] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), or DNM2 H9D(+7-18). [0197] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), or DNM2 H9D(+4-21). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17). [0198] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22). [0199] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-5+20). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H9A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+6+30). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+25+49). [0200] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 158 (CGTCAAACTCCATCTGAGAAGAGAC); SEQ ID NO: 159 (TCCTTCTCGTCAAACTCCATCTGAG); SEQ ID NO: 160 (GTAAGTCCTTCTCGTCAAACTCCAT); SEQ ID NO: 161 (CCGTCGTAAGTCCTTCTCGTCAAAC); SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG); SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT); SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC);
SEQ ID NO: 168 (TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 175 (TGACTCCATGGATGTTCTTAATG); SEQ ID NO: 176 (TGACTCCATGGATGTTCTTAAT); SEQ ID NO: 177 (TGACTCCATGGATGTTCTTAA); SEQ ID NO: 178 (TGACTCCATGGATGTTCTTA); SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT); SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG); SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT); SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT);
SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 208 (CTCCTCGTGGAACTTGCCTG); and SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG). [0201] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG); SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT); SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT); SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG); SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 (TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT); SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); and
SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG). [0202] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); and SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT). [0203] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 175 (TGACTCCATGGATGTTCTTAATG); SEQ ID NO: 176 (TGACTCCATGGATGTTCTTAAT); SEQ ID NO: 177 (TGACTCCATGGATGTTCTTAA); and SEQ ID NO: 178 (TGACTCCATGGATGTTCTTA). [0204] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); and SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG). [0205] In certain embodiments, the targeting sequence is SEQ ID NO: 172. [0206] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT); SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT);
SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG); SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); and SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT). [0207] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); and SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT). [0208] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); and SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG). [0209] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 (TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); and SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT). [0210] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC);
SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 208 (CTCCTCGTGGAACTTGCCTG); and SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG). [0211] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); and SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC). [0212] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); and SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC). [0213] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT);
SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); and SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA). [0214] In certain embodiments, the targeting sequence is SEQ ID NO: 199. [0215] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 208 (CTCCTCGTGGAACTTGCCTG); and SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG). [0216] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 158 (CGTCAAACTCCATCTGAGAAGAGAC); SEQ ID NO: 159 (TCCTTCTCGTCAAACTCCATCTGAG); SEQ ID NO: 160 (GTAAGTCCTTCTCGTCAAACTCCAT); SEQ ID NO: 161 (CCGTCGTAAGTCCTTCTCGTCAAAC); SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); and SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG). [0217] In certain embodiments, the target region is within the exon internal region of exon 12 (SEQ ID NO: 7), the intron 11/exon 12 junction, or the exon 12/intron 12 junction of the human DNM2 gene pre-mRNA. [0218] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 4th nucleotide to the 31st nucleotide measured of exon 12 from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 36. In certain embodiments, the target region is a region within DNM2 H12A(+4+31). [0219] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 44th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 37. In certain embodiments, the target region is a region within DNM2 H12A(+10+44). [0220] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 38th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ
ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 38. In certain embodiments, the target region is a region within DNM2 H12A(+10+38). [0221] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 16th nucleotide to the 44th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 39. In certain embodiments, the target region is a region within DNM2 H12A(+16+44). [0222] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 14th nucleotide to the 42nd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 40. In certain embodiments, the target region is a region within DNM2 H12A(+14+42). [0223] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 25th nucleotide to the 54th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 41. In certain embodiments, the target region is a region within DNM2 H12A(+25+54). [0224] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 25th nucleotide to the 53rd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 42. In certain embodiments, the target region is a region within DNM2 H12A(+25+53). [0225] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 26th nucleotide to the 54th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 43. In certain embodiments, the target region is a region within DNM2 H12A(+26+54). [0226] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 36th nucleotide to the 61st nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 44. In certain embodiments, the target region is a region within DNM2 H12A(+36+61). [0227] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 22nd nucleotide of exon 12 measured from the 3′ end of exon 12 (SEQ ID NO: 7) to the 5th nucleotide of intron 12 measured from the 3′ end of exon 12. In certain embodiments, the target region is a region within SEQ ID NO: 45. In certain embodiments, the target region is a region within DNM2 H12D(+22-5). [0228] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-13+12), DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2
H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), or DNM2 H12D(+15-10). [0229] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+49), DNM2 H12A(+29+53), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), or DNM2 H12D(+15-10). [0230] In certain embodiments, the target region is DNM2 H12A(+4+28) or DNM2 H12A(+7+31). [0231] In certain embodiments, the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44). [0232] In certain embodiments, the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), or DNM2 H12A(+14+38). [0233] In certain embodiments, the target region is DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), or DNM2 H12A(+20+44). [0234] In certain embodiments, the target region is DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), or DNM2 H12A(+18+42). [0235] In certain embodiments, the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54). [0236] In certain embodiments, the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), or DNM2 H12A(+29+53). [0237] In certain embodiments, the target region is DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2
H12A(+30+49), or DNM2 H12A(+30+54). In certain embodiments, the target region is DNM2 H12A(+25+49). In certain embodiments, the target region is DNM2 H12A(+36+60) or DNM2 H12A(+37+61). [0238] In certain embodiments, the target region is DNM2 H12D(+22-3) or DNM2 H12D(+20-5). In certain embodiments, the target region is DNM2 H12A(-13+12). In certain embodiments, the target region is DNM2 H12A(-3+22). In certain embodiments, the target region is DNM2 H12A(+43+67). In certain embodiments, the target region is DNM2 H12D(+15-10). [0239] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 210 (CAGCAGAAGAATCTGGBGGAGAGCG); SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG); SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA); SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG);
SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG); SEQ ID NO: 238 (GATGAAGTCCTCATGGTTCGTGTTG); SEQ ID NO: 239 (CGATGAAGTCCTCATGGTTCGTGTT); SEQ ID NO: 240 (CAAACCCGATGAAGTCCTCATGGTT); SEQ ID NO: 241 (TACTTGGCAAACCCGATGAAGTCCT); SEQ ID NO: 242 (CCTACTTGGCAAACCCGATGAAGTC); and SEQ ID NO: 243 (AAGTACCTACTTGGCAAACCCGATG). [0240] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG); SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG); SEQ ID NO: 238 (GATGAAGTCCTCATGGTTCGTGTTG); SEQ ID NO: 240 (CAAACCCGATGAAGTCCTCATGGTT); SEQ ID NO: 241 (TACTTGGCAAACCCGATGAAGTCCT); SEQ ID NO: 242 (CCTACTTGGCAAACCCGATGAAGTC); and SEQ ID NO: 243 (AAGTACCTACTTGGCAAACCCGATG). [0241] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); and SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG).
[0242] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); and SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA). [0243] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); and SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA). [0244] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); and SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA). [0245] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); and SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG). [0246] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG);
SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); and SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG). [0247] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); and SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG). [0248] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); and SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG). [0249] In certain embodiments, the targeting sequence is SEQ ID NO: 230 [0250] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 238 (GATGAAGTCCTCATGGTTCGTGTTG); and SEQ ID NO: 239 (CGATGAAGTCCTCATGGTTCGTGTT). [0251] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 241 (TACTTGGCAAACCCGATGAAGTCCT); and
SEQ ID NO: 242 (CCTACTTGGCAAACCCGATGAAGTC). [0252] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 210 (CAGCAGAAGAATCTGGBGGAGAGCG); SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 240 (CAAACCCGATGAAGTCCTCATGGTT); and SEQ ID NO: 243 (AAGTACCTACTTGGCAAACCCGATG). [0253] In certain embodiments, the target region is within the exon internal region of exon 13 (SEQ ID NO: 8), the intron 12/exon 13 junction, the exon 13/intron 13 junction, or the intron internal region of intron 12 (SEQ ID NO: 12). [0254] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 27th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 46. In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within DNM2 H13A(-27+12). [0255] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 27th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 2nd nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 47. In certain embodiments, the target region is a region within DNM2 H13A(-27+2). [0256] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 22nd nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 7th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 48. In certain embodiments, the target region is a region within DNM2 H13A(-22+7). [0257] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 17th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 49. In certain embodiments, the target region is a region within DNM2 H13A(-17+12). [0258] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 11th nucleotide of intron 12 (SEQ ID NO: 12) measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 50. In certain embodiments, the target region is a region within DNM2 H13A(-11+24).
[0259] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 11th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 18th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 51. In certain embodiments, the target region is a region within DNM2 H13A(-11+18). [0260] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 5th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24th nucleotide exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 52. In certain embodiments, the target region is a region within DNM2 H13A(-5+24). [0261] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 8th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 21st nucleotide exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 53. In certain embodiments, the target region is a region within DNM2 H13A(-8+21). [0262] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 54. In certain embodiments, the target region is a region within DNM2 H13A(+1+30). [0263] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 28th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 55. In certain embodiments, the target region is a region within DNM2 H13A(+1+28). [0264] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 56. In certain embodiments, the target region is a region within DNM2 H13A(+6+30). [0265] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 3rd nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 57. In certain embodiments, the target region is a region within DNM2 H13A(+3+30). [0266] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2
H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(- 5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), DNM2 H13A(+9+28), DNM2 H13A(+16+40), DNM2 H13D(+22-3), or DNM2 H13D(+7-18). [0267] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(- 5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), or DNM2 H13A(+4+28). [0268] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12). [0269] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), or DNM2 H13A(- 23+2). [0270] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), or DNM2 H13A(- 18+7). [0271] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12). [0272] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(- 7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(-1+24).
[0273] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), or DNM2 H13A(-7+18). [0274] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(- 1+24). [0275] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), or DNM2 H13A(- 4+21). [0276] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28). [0277] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), or DNM2 H13A(+5+28). [0278] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28). [0279] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), or DNM2 H13A(+6+30). [0280] In certain embodiments, the target region of the human DNM2 gene pre-mRNA DNM2 H13A(+4+28). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+16+40). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H13D(+22-3). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13D(+7-18). [0281] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT);
SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA); SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC); SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 282 (GTTCAGCTGCGTGCTCCTCTGC);
SEQ ID NO: 283 (GTTCAGCTGCGTGCTCCTCTG); SEQ ID NO: 284 (GTTCAGCTGCGTGCTCCTCT); SEQ ID NO: 285 (GGCTCTCTTCTTGTTCAGCTGCGTG); SEQ ID NO: 286 (TACCTGATTGGBGATGGCTCTCTTC); and SEQ ID NO: 287 (TGCAGAGGBGTGTGCTACCTGATTG), wherein “B” represents an abasic subunit or guanine. In certain embodiments, B is an abasic subunit. [0282] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA); SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC); SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC);
SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); and SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG). [0283] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); and SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA). [0284] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); and SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG). [0285] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); and SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA).
[0286] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); and SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA). [0287] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); and SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC). [0288] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); and SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC). [0289] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); and SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC). [0290] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); and SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA).
[0291] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 282 (GTTCAGCTGCGTGCTCCTCTGC); SEQ ID NO: 283 (GTTCAGCTGCGTGCTCCTCTG); and SEQ ID NO: 284 (GTTCAGCTGCGTGCTCCTCT). [0292] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); and SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG). [0293] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 282 (GTTCAGCTGCGTGCTCCTCTGC); SEQ ID NO: 283 (GTTCAGCTGCGTGCTCCTCTG); and SEQ ID NO: 284 (GTTCAGCTGCGTGCTCCTCT). [0294] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG);
SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); and SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT). [0295] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 285 (GGCTCTCTTCTTGTTCAGCTGCGTG); SEQ ID NO: 286 (TACCTGATTGGBGATGGCTCTCTTC); and SEQ ID NO: 287 (TGCAGAGGBGTGTGCTACCTGATTG). [0296] In certain embodiments, the target region is within the exon internal region of exon 6 (SEQ ID NO: 3), the intron 5/exon 6 junction, the exon 6/intron 6 junction, or the intron internal region of intron 6 (SEQ ID NO: 11). [0297] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 37th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 58. In certain embodiments, the target region is a region within DNM2 H6A(+10+37). [0298] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 32nd nucleotide to the 59th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 59. In certain embodiments, the target region is a region within DNM2 H6A(+32+59). [0299] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 60th nucleotide to the 87th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 60. In certain embodiments, the target region is a region within DNM2 H6A(+60+87). [0300] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 130th nucleotide to the 156th nucleotide measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 61. In certain embodiments, the target region is a region within DNM2 H6A(+130+156). [0301] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 17th nucleotide of exon 6 measured from the 3′ end of exon 6 (SEQ ID NO: 3) to the 10th
nucleotide of intron 6 measured from the 3′ end of exon 6. In certain embodiments, the target region is a region within SEQ ID NO: 62. In certain embodiments, the target region is a region within DNM2 H6D(+17-10). [0302] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5), DNM2 H6A(-6+19), DNM2 H6A(-2+23), DNM2 H6A(+3+26), DNM2 H6A(+10+3), DNM2 H6A(+13+37), DNM2 H6A(+23+47), DNM2 H6A(+32+56), DNM2 H6A(+35+59), DNM2 H6A(+44+68), DNM2 H6A(+54+78), DNM2 H6A(+60+84), DNM2 H6A(+63+87), DNM2 H6A(+88+112), DNM2 H6A(+96+119), DNM2 H6A(+105+129), DNM2 H6A(+121+145), DNM2 H6A(+130+154), DNM2 H6A(+132+156), DNM2 H6D (+6-19), DNM2 H6D (+15-10), or DNM2 H6D (+17-8). [0303] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+10+3) or DNM2 H6A(+13+37). [0304] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+32+56) or DNM2 H6A(+35+59). [0305] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+60+84) or DNM2 H6A(+63+87). [0306] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+130+154) or DNM2 H6A(+132+156). [0307] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+15-10) or DNM2 H6D (+17-8). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-6+19). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-2+23). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+3+26). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+23+47). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+44+68). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+54+78). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+88+112). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H6A(+96+119). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+105+129). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+121+145). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+6-19). [0308] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 97 (GTAGCCTGTGGGGAGAGGCAGTCAG);
SEQ ID NO: 98 (TTCACCACGCCAATGTAGCCTGTGG); SEQ ID NO: 99 (GCGGTTCACCACGCCAATGTAGCCT); SEQ ID NO: 100 (GCTGCGGTTCACCACGCCAATGTA); SEQ ID NO: 101 (TCCTTCTGGCTGCGGTTCACCACGC); SEQ ID NO: 102 (ATATCCTTCTGGCTGCGGTTCACCA); SEQ ID NO: 103 (CTTGCCCTCAATATCCTTCTGGCTG); SEQ ID NO: 104 (GATGTCCTTCTTGCCCTCAATATCC); SEQ ID NO: 105 (ACGGATGTCCTTCTTGCCCTCAATA); SEQ ID NO: 106 (CAGTGCTGCACGGATGTCCTTCTTG); SEQ ID NO: 107 (TCTCAGCTGCCAGTGCTGCACGGAT); SEQ ID NO: 108 (ACTTCCTCTCAGCTGCCAGTGCTGC); SEQ ID NO: 109 (AGAACTTCCTCTCAGCTGCCAGTGC); SEQ ID NO: 110 (TGCCGGTAGGCCGGGTGGGAGAGGA); SEQ ID NO: 111 (GGCCATGTGCCGGTAGGCCGGGTG); SEQ ID NO: 112 (CCATGCGGTCGGCCATGTGCCGGTA); SEQ ID NO: 113 (TGCAGATGTGGCGTGCCCATGCGGT); SEQ ID NO: 114 (AGCGTCTTCTGCAGATGTGGCGTGC); SEQ ID NO: 115 (TCAGCGTCTTCTGCAGATGTGGCGT); SEQ ID NO: 116 (GTGCAAACCCTTGCAGTACCTGATT); SEQ ID NO: 117 (CTTGCAGTACCTGATTCAGCGTCTT); and SEQ ID NO: 118 (TGCAGTACCTGATTCAGCGTCTTCT). [0309] In certain embodiments, the target region is within the exon internal region of exon 7 (SEQ ID NO: 4), the intron 6/exon 7 junction, or the exon 7/intron 7 junction. [0310] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 72nd nucleotide to the 98th nucleotide of exon 7 measured from the 5′ end of exon 7 (SEQ ID NO: 4). In certain embodiments, the target region is a region within SEQ ID NO: 63. In certain embodiments, the target region is a region within DNM2 H7A(+72+98). [0311] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 7 measured from the 3′ end of exon 7 (SEQ ID NO: 4) to the 16th nucleotide of intron 7 measured from the 3′ end of exon 7. In certain embodiments, the target region is a region within SEQ ID NO: 64. In certain embodiments, the target region is a region within DNM2 H7D(+9-16). [0312] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+1+25), DNM2 H7A(+6+30), DNM2 H7A(+11+35), DNM2 H7A(+22+46), DNM2 H7A(+29+53), DNM2 H7A(+41+65), DNM2 H7A(+51+75), DNM2 H7A(+57+81), DNM2
H7A(+62+86), DNM2 H7A(+72+96), DNM2 H7A(+74+98), DNM2 H7A(+81+105), DNM2 H7A(+91+115), or DNM2 H7A(+110+134). [0313] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+72+96) or DNM2 H7A(+74+98). [0314] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+8-16) or DNM2 H7D(+9-16). [0315] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-15+10). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-5+20). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H7A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+6+30). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+11+35). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+22+46). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+29+53). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+41+65). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+51+75). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+57+81). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+62+86). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H7A(+81+105). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+91+115). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+110+134). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+21-4). [0316] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 119 (TGGTCAGTTGCTGCAGGAGAGAGGT); SEQ ID NO: 120 (CGGATGTGGTTGGTCAGTTGCTGCA); SEQ ID NO: 121 (ACTCCCGGATGTGGTTGGTCAGTTG); SEQ ID NO: 122 (CAGCGACTCCCGGATGTGGTTGGTC); SEQ ID NO: 123 (GCCGGCAGCGACTCCCGGATGTGGT); SEQ ID NO: 124 (TGCTACGTAGGGCCGGCAGCGACTC); SEQ ID NO: 125 (TGTAGTTTGCTACGTAGGGCCGGCA); SEQ ID NO: 126 (AGCAGCTGGCTCTGTAGTTTGCTAC); SEQ ID NO: 127 (CTCCAGGGACAGCAGCTGGCTCTGT); SEQ ID NO: 128 (CTCCTTCTCCAGGGACAGCAGCTGG); SEQ ID NO: 129 (TCCACCTCCTTCTCCAGGGACAGCA);
SEQ ID NO: 130 (CTTGTACTCCTCCACCTCCTTCTCC); SEQ ID NO: 131 (TTCTTGTACTCCTCCACCTCCTTCT); SEQ ID NO: 132 (CCGAAAGTTCTTGTACTCCTCCACC); SEQ ID NO: 133 (GGTCGTCGGGCCGAAAGTTCTTGTA); SEQ ID NO: 134 (GCTTTGGTTTTGCGGGTGGGGTCGT); SEQ ID NO: 135 (GTGCCGBGGTACATACTGCAGCAG); SEQ ID NO: 136 (GTGCCGGGGTACATACTGCAGCAGG); and SEQ ID NO: 137 (ATACTGCAGCAGGGCTTTGGTTTTG), wherein “B” represents an abasic subunit or guanine. In certain embodiments, B is an abasic subunit. [0317] In certain embodiments, the target region is within the exon internal region of exon 8 (SEQ ID NO: 5), the intron 7/exon 8 junction, or the exon 8/intron 8 junction. [0318] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 7th nucleotide of intron 7 measured from the 5′ end of exon 8 (SEQ ID NO: 5) to the 19th nucleotide of exon 8 measured from the 5′ end of exon 8. In certain embodiments, the target region is a region within SEQ ID NO: 65. In certain embodiments, the target region is a region within DNM2 H8A(-7+19). [0319] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 18th nucleotide to the 45th nucleotide of exon 8 measured from the 5′ end of exon 8 (SEQ ID NO: 5). In certain embodiments, the target region is a region within SEQ ID NO: 66. In certain embodiments, the target region is a region within DNM2 H8A(+18+45). [0320] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-20+5), DNM2 H8A(-14+11), DNM2 H8A(-7+18), DNM2 H8A(-6+19), DNM2 H8A(-2+23), DNM2 H8A(+5+29), DNM2 H8A(+18+42), DNM2 H8A(+21+45), DNM2 H8A(+27+51), DNM2 H8A(+36+60), DNM2 H8A(+46+70), DNM2 H8A(+51+75), DNM2 H8A(+56+80), DNM2 H8A(+65+88), DNM2 H8A(+76+100), DNM2 H8A(+83+107), DNM2 H8A(+96+119), DNM2 H8A(+105+129), DNM2 H8D(+18-7), or DNM2 H8D(+10-15). [0321] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-7+18) or DNM2 H8A(-6+19). [0322] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+18+42) or DNM2 H8A(+21+45). [0323] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-20+5). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-14+11). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H8A(-2+23). In certain embodiments, the target region of the human DNM2
gene pre-mRNA is DNM2 H8A(+5+29). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+27+51). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+36+60). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+46+70). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+51+75). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+56+80). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+65+88). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+76+100). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H8A(+83+107). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+96+119). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+105+129). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+18-7). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+10-15). [0324] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 138 (CCATCCTATGAGGAAAAGGAAAAAG); SEQ ID NO: 139 (GCTGGACCATCCTATGAGGAAAAGG); SEQ ID NO: 140 (CCAAACTGCTGGACCATCCTATGAG); SEQ ID NO: 141 (CCCAAACTGCTGGACCATCCTATGA); SEQ ID NO: 142 (CCACCCCAAACTGCTGGACCATCCT); SEQ ID NO: 143 (CAAAATCCACCCCAAACTGCTGGAC); SEQ ID NO: 144 (TCGATCCTCTTCTCAAAATCCACCC); SEQ ID NO: 145 (CCCTCGATCCTCTTCTCAAAATCCA); SEQ ID NO: 146 (CCTGAGCCCTCGATCCTCTTCTCAA); SEQ ID NO: 147 (ACCTGATCTCCTGAGCCCTCGATCC); SEQ ID NO: 148 (CAGAGTGTCCACCTGATCTCCTGAG); SEQ ID NO: 149 (AGCTCCAGAGTGTCCACCTGATCTC); SEQ ID NO: 150 (CGGAGAGCTCCAGAGTGTCCACCTG); SEQ ID NO: 151 (GGCGCCCCCGGAGAGCTCCAGAGT); SEQ ID NO: 152 (GCGATTGATTCGGGCGCCCCCGGAG); SEQ ID NO: 153 (GGAAGATGCGATTGATTCGGGCGCC); SEQ ID NO: 154 (GGAACCGCTCGTGGAAGATGCGAT); SEQ ID NO: 155 (AGCTCAAATGGGAACCGCTCGTGGA); SEQ ID NO: 156 (GCACTACCTTCACCAGCTCAAATGG); and SEQ ID NO: 157 (CCCGGGGGGCACTACCTTCACCAGC).
[0325] In certain embodiments, the target region is within the exon internal region of exon 16 (SEQ ID NO: 9), the intron 15/exon 16 junction, or the exon 16/intron 16 junction. [0326] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3), DNM2 H16A(-13+12), DNM2 H16A(-6+19), DNM2 H16A(+6+30), DNM2 H16A(+15+39), DNM2 H16A(+26+50), DNM2 H16A(+36+60), DNM2 H16A(+45+69), DNM2 H16A(+55+79), DNM2 H16A(+66+90), DNM2 H16A(+75+99), DNM2 H16A(+82+106), DNM2 H16D(+19-6), or DNM2 H16D(+11-14). DNM2 H16A(-21+3). [0327] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-13+12). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H16A(-6+19). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+6+30). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+15+39). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+26+50). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+36+60). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+45+69). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+55+79). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+66+90). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+75+99). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+82+106). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16D(+19-6). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16D(+11-14). [0328] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 288 (CTCCTGAGGGTGGAGGGAAAAGAG); SEQ ID NO: 289 (CTTCTCTTTCTCCTGAGGGTGGAGG); SEQ ID NO: 290 (TGTACTTCTTCTCTTTCTCCTGAGG); SEQ ID NO: 291 (CAGAGGCAGCATGTACTTCTTCTCT); SEQ ID NO: 292 (GAGGTTGTCCAGAGGCAGCATGTAC); SEQ ID NO: 293 (TCACGGATCTTGAGGTTGTCCAGAG); SEQ ID NO: 294 (CTTCTCCACATCACGGATCTTGAGG); SEQ ID NO: 295 (CATGAAGCCCTTCTCCACATCACGG); SEQ ID NO: 296 (GCTTGTTGGACATGAAGCCCTTCTC); SEQ ID NO: 297 (GGCGAAGACGTGCTTGTTGGACATG); SEQ ID NO: 298 (GTTGAAGATGGCGAAGACGTGCTTG);
SEQ ID NO: 299 (GCTCCGTGTTGAAGATGGCGAAGAC); SEQ ID NO: 300 (CCTCACCTCTGCTCCGTGTTGAAGA); and SEQ ID NO: 301 (CCTGGGCCCCTCACCTCTGCTCCGT). [0329] In certain embodiments, the target region is within the exon internal region of exon 19 (SEQ ID NO: 9), the intron 18/exon 19 junction, or the exon 18/intron 18 junction. [0330] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14), DNM2 H19A(-4+21), DNM2 H19A(+1+25), DNM2 H19A(+7+31), DNM2 H19A(+16+40), DNM2 H19A(+26+50), DNM2 H19A(+37+61), DNM2 H19A(+47+71), DNM2 H19A(+56+80), DNM2 H19A(+66+90), DNM2 H19A(+76+100), DNM2 H19A(+86+110), DNM2 H19A(+96+120), DNM2 H19A(+106+130), DNM2 H19A(+117+141), DNM2 H19A(+127+151), DNM2 H19A(+137+161), DNM2 H19A(+147+171), DNM2 H19A(+157+181), DNM2 H19A(+166+189), DNM2 H19A(+191+215), DNM2 H19A(+200+224), DNM2 H19D(+23-2), DNM2 H19D(+17-8), or DNM2 H19D(+10-15). [0331] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-4+21). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+7+31). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+16+40). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+26+50). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+37+61). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+47+71). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+56+80). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+66+90). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+76+100). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+86+110). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+96+120). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+106+130). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+117+141). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+127+151). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+137+161). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+147+171). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+157+181). In certain embodiments, the
target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+166+189). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+191+215). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+200+224). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+23-2). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+17-8). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+10-15). [0332] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 302 (ATGAAGGCCTTCGTCTGCGGGCAGG); SEQ ID NO: 303 (GTGGTGGATGAAGGCCTTCGTCTGC); SEQ ID NO: 304 (GCTCGTGGTGGATGAAGGCCTTCGT); SEQ ID NO: 305 (CCAGCAGCTCGTGGTGGATGAAGGC); SEQ ID NO: 306 (ATAGGTAGGCCAGCAGCTCGTGGTG); SEQ ID NO: 307 (GCCGAGGAGTATAGGTAGGCCAGCA); SEQ ID NO: 308 (TGCTCTGGTCTGCCGAGGAGTATAG); SEQ ID NO: 309 (TCCATGAGGCTGCTCTGGTCTGCCG); SEQ ID NO: 310 (GCCGACTCCTCCATGAGGCTGCTCT); SEQ ID NO: 311 (TGCCTGGTCAGCCGACTCCTCCATG); SEQ ID NO: 312 (CCCGCCGCTGTGCCTGGTCAGCCGA); SEQ ID NO: 313 (AGCATGTCGTCCCGCCGCTGTGCCT); SEQ ID NO: 314 (GTACATGCGCAGCATGTCGTCCCGC); SEQ ID NO: 315 (TGAGGGCATGGTACATGCGCAGCAT); SEQ ID NO: 316 (GAGCGCCTCCTTGAGGGCATGGTAC); SEQ ID NO: 317 (CGATGATGTTGAGCGCCTCCTTGAG); SEQ ID NO: 318 (CTGATGTCACCGATGATGTTGAGCG); SEQ ID NO: 319 (AGTGCTGGTGCTGATGTCACCGATG); SEQ ID NO: 320 (GCGTGGACACAGTGCTGGTGCTGAT); SEQ ID NO: 321 (GGGTACAGGCGTGGACACAGTGCT); SEQ ID NO: 322 (TGGAGCCAGGTGTCATCGACAGGCG); SEQ ID NO: 323 (CTGGCGCTCTGGAGCCAGGTGTCAT); SEQ ID NO: 324 (ACCTGTGGCTGCTGGCGCTCTGGAG); SEQ ID NO: 325 (TTCCGGACCTGTGGCTGCTGGCGCT); and SEQ ID NO: 326 (ACCAGGCTTCCGGACCTGTGGCTGC). [0333] In certain embodiments, each R2, taken together, forms a targeting sequence having SEQ ID NO: 83. In certain embodiments, each R2, taken together, forms a targeting sequence
having SEQ ID NO: 88. In certain embodiments, each R2, taken together, forms a targeting sequence having SEQ ID NO: 172. In certain embodiments, R2, taken together, forms a targeting sequence having SEQ ID NO: 199. In certain embodiments, each R2, taken together, forms a targeting sequence having SEQ ID NO: 230. In certain embodiments, each R2, taken together, forms a targeting sequence having SEQ ID NO: 278. [0334] In one embodiment, the targeting sequence is at least 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99 percent complementary to the target region. In another embodiment, the targeting sequence is at least 84%, at least 88%, or at least 92% complementary to the target region. In still another embodiment, the targeting sequence is at least 90% complementary to the target region. In yet another embodiment, the targeting sequence is at least 95% complementary to the target region. In still another embodiment, the targeting sequence is 100% complementary to the target region. [0335] The antisense oligomer of the present disclosure can have a modified chemical backbone as described herein or known in the art. For example, the antisense oligomer may be a peptide nucleic acid (PNA), a locked nucleic acid, a phosphorodiamidate morpholino oligomer, a 2′-OMe phosphorothioate oligomer, or a combination thereof. In an embodiment, the antisense oligomer is a phosphorodiamidate morpholino oligomer. In another embodiment, the antisense oligomer is a peptide nucleic acid. In yet another embodiment, the antisense oligomer is a locked nucleic acid. In still another embodiment, the antisense oligomer is a 2′-OMe phosphorothioate oligomer. In still another embodiment, the antisense oligomer is a combination of any of the above oligonucleotides having a modified chemical backbone. [0336] In some embodiments, the antisense oligomer is conjugated to, e.g., covalently attached to, a delivery moiety such as, for example, a cell-penetrating peptide, an antibody, a fragment of an antibody, or an antigen fragment of an antibody. In an embodiment, the antisense oligomer is conjugated to a cell-penetrating peptide. In another embodiment, the antisense oligomer is conjugated to a cell-penetrating peptide, wherein the cell-penetrating peptide is selected from rTAT, Tat, R9F2, R5F2R4, R4, R5, R6, R7, R8, R9, (RXR)4, (RXR)5, (RXRRBR)2, (RAR)4F2, (RGR)4F2, and RBRBYLIQFRBRRBR, wherein A represents alanine, B represents beta-alanine, F represents phenylalanine, G represents glycine, I represents isoleucine, L represents leucine, Q represents glutamine, R represents arginine, X represent 6-aminohexanoic acid, and Y represents tyrosine. In yet another embodiment, the antisense oligomer is conjugated to an antibody, an antibody fragment, or an antigen fragment of an antibody. [0337] The antisense oligomer of the present disclosure may be attached to the delivery moiety via a linker. In an embodiment, the linker is a direct bond from the antisense oligomer to the
delivery moiety. In another embodiment, the linker is an amino acid such as, for example, a glycine amino acid, a proline amino acid, or a glutamic amino acid. In a particular embodiment, the linker is a glycine amino acid or a proline amino acid. [0338] Provided herein is an antisense oligomer, or a pharmaceutically acceptable salt thereof, having a structure of Formula (I):
(I), wherein A′, R1, R2, t, E′, and G are as described herein. [0259] In certain embodiments, t is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28. In some embodiments, each R2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt. [0260] In an embodiment, E′ is selected from H, -C1-6-alkyl, -C(O)C1-6-alkyl, benzoyl, stearoyl, trityl, monomethoxytrityl, dimethoxytrityl, trimethoxytrityl, and
.
[0339] In a further embodiment, E′ is selected from -H, -C(O)CH3, benzoyl, stearoyl, trityl, 4- methoxytrityl, and
. [0341] In an embodiment, at least one of the following is true:
[0342] In an embodiment, A′ is selected from:
and E′ is
. [0343] In an embodiment, A′ is
[0344] E′ is selected from H, -C(O)CH3, trityl, 4-methoxytrityl, benzoyl, and stearoyl. In an embodiment, each R1 is -N(CH3)2. In an embodiment, L is glycine, proline, or b-alanine. In an embodiment, L is glycine. In an embodiment, L is proline. In an embodiment, L is b-alanine. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt. [0345] In some embodiments, provided herein is an antisense oligomer, which is an oligomer of Formula (IA):
or a pharmaceutically acceptable salt thereof, wherein: A′ is a moiety selected from:
are as described herein. [0346] In certain embodiments, t is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28. In some embodiments, each R2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt.
[0347] In some embodiments, provided herein is an antisense oligomer, which is an oligomer of Formula (II):
or a pharmaceutically acceptable salt thereof, wherein t, G, and R2 are as described herein. [0348] In certain embodiments, t is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28. In some embodiments, each R2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt.
[0349] In some embodiments, provided herein is an antisense oligomer, which is an oligomer of Formula (III):
or a pharmaceutically acceptable salt thereof, wherein n is 11-28 and R2 is as described herein. [0350] In certain embodiments, n is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28. In some embodiments, each R2 taken together forms a targeting sequence of 13- 30 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt. [0351] In some embodiments, provided herein is an antisense oligomer, which is an oligomer of Formula (IV):
(IV), wherein n is 11-28 and R2 is as described herein.
[0352] In certain embodiments, n is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28. In some embodiments, each R2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. [0353] In an embodiment, J is selected from rTAT, TAT, R9F2, R5F2R4, R4, R5, R6, R7, R8, R9, (RXR)4, (RXR)5, (RXRRBR)2, (RAR)4F2, (RGR)4F2, and RBRBYLIQFRBRRBR, wherein A represents alanine, B represents βbeta-alanine, F represents phenylalanine, G represents glycine, R represents arginine, and X represent 6-aminohexanoic acid. [0354] In an embodiment, G is selected from H, C(O)CH3, benzoyl, and stearoyl. In an embodiment, G is H or -C(O)CH3. In an embodiment, G is H. In an embodiment, G is -C(O)CH3. [0355] In certain embodiments, the antisense oligomer is selected from:
,
[0356] In certain embodiments, the antisense oligomer has the structure:
. [0357] In certain embodiments, the antisense oligomer has the structure:
[0361] In certain embodiments, the antisense oligomer has the structure:
. [0362] Provided herein is a pharmaceutical composition, comprising: an antisense oligomer, or a pharmaceutically acceptable salt thereof, of Formulae (I), (IA), (II), (III), (IV) and/or (V); and at least one pharmaceutically acceptable carrier. [0363] Provided herein is a method of treating a disease in a subject in need thereof, the method comprising administering a therapeutically effective amount of an antisense oligomer, including an oligomer of Formulae (I), (IA), (II), (III), (IV), and/or (V), or a pharmaceutical composition disclosed herein to the subject. [0364] In an embodiment, the disease is associated with aberrant expression of dynamin 2 (DNM2) protein. In certain embodiments, the disease is a centronuclear myopathy (CNM). In one embodiment, the subject is a human.
III. Oligomer Chemistry Features [0365] Also provided herein are antisense oligomers useful for targeting the human DNM2 gene pre-mRNA, wherein the antisense oligomer is a modified antisense oligomer. Examples of modified antisense oligomers include, without limitation, morpholino oligomers, phosphorothioate modified oligomers, 2′-O-methyl modified oligomers, peptide nucleic acid (PNA), locked nucleic acid (LNA), phosphorothioate oligomers, 2′-O-MOE modified oligomers, 2′-fluoro-modified oligomer, 2′-O,4′-C-ethylene-bridged nucleic acids (ENAs), tricyclo-DNAs, tricyclo-DNA phosphorothioate subunits, 2′-O-[2-(N- methylcarbamoyl)ethyl] modified oligomers, including combinations of any of the foregoing. Phosphorothioate and 2′-O-Me-modified chemistries can be combined to generate a 2′-O-Me-phosphorothioate backbone. See, e.g., PCT Publication Nos. WO/2013/112053 and WO/2009/008725, which are hereby incorporated by reference in their entireties. [0366] In some embodiments, the nucleobases of the modified antisense oligomer are linked to morpholino ring structures, wherein the morpholino ring structures are joined by phosphoros- containing intersubunit linkages joining a morpholino nitrogen of one ring structure to a 5′ exocyclic carbon of an adjacent ring structure. [0367] In some embodiments, the nucleobases of the antisense oligomer are linked to a peptide nucleic acid (PNA), wherein the phosphate-sugar polynucleotide backbone is replaced by a flexible pseudo-peptide polymer to which the nucleobases are linked. In some aspects, at least one of the nucleobases of the antisense oligomer is linked to a locked nucleic acid (LNA), wherein the locked nucleic acid structure is a nucleotide analog that is chemically modified where the ribose moiety has an extra bridge connecting the 2′ oxygen and the 4′ carbon. [0368] In some embodiments, at least one of the nucleobases of the antisense oligomer is linked to a bridged nucleic acid (BNA), wherein the sugar conformation is restricted or locked by the introduction of an additional bridged structure to the furanose skeleton. In some aspects, at least one of the nucleobases of the antisense oligomer is linked to a 2′-O,4′-C-ethylene- bridged nucleic acid (ENA). [0369] In some embodiments, the modified antisense oligomer may contain unlocked nucleic acid (UNA) subunits. UNAs and UNA oligomers are an analogue of RNA in which the C2′-C3′ bond of the subunit has been cleaved. [0370] In some embodiments, the modified antisense oligomer contains one or more phosphorothioates (or S-oligos), in which one of the nonbridging oxygens is replaced by sulfur. In some aspects, the modified antisense oligomer contains one or more 2′ O-Methyl, 2′ O-MOE, MCE, and 2′-F in which the 2′-OH of the ribose is substituted with a methyl, methoxy ethyl, 2-(N- methylcarbamoyl)ethyl, or fluoro group, respectively.
[0371] In some embodiments, the modified antisense oligomer is a tricyclo-DNA (tc-DNA) which is a constrained DNA analog in which each nucleotide is modified by the introduction of a cyclopropane ring to restrict conformational flexibility of the backbone and to optimize the backbone geometry of the torsion angle g. [0372] In some embodiments, at least one of the nucleobases of the antisense oligomer is linked to a bridged nucleic acid (BNA), wherein the sugar conformation is restricted or locked by the introduction of an additional bridged structure to the furanose skeleton. In some aspects, at least one of the nucleobases of the antisense oligomer is linked to a 2′-O,4′-C-ethylene- bridged nucleic acid (ENA). In such aspects, each nucleobase which is linked to a BNA or ENA comprises a 5-methyl group. Exemplary embodiments of oligomer chemistries of the disclosure are further described below. 1. Peptide Nucleic Acids (PNAs) [0373] Peptide nucleic acids (PNAs) are analogs of DNA in which the backbone is structurally homomorphous with a deoxyribose backbone, consisting of N-(2-aminoethyl) glycine units to which pyrimidine or purine bases are attached. PNAs containing natural pyrimidine and purine bases hybridize to complementary oligomers obeying Watson-Crick base-pairing rules, and mimic DNA in terms of base-pair recognition. The backbone of PNAs is formed by peptide bonds rather than phosphodiester bonds, making them well-suited for antisense applications (see structure below). The backbone is uncharged, resulting in PNA/DNA or PNA/RNA duplexes that exhibit greater than normal thermal stability. PNAs are not recognized by nucleases or proteases. A non-limiting example of a PNA is depicted below.
[0374] Despite a radical structural change to the natural structure, PNAs are capable of sequence-specific binding in a helix form to DNA or RNA. Characteristics of PNAs include a high binding affinity to complementary DNA or RNA, a destabilizing effect caused by single-base mismatch, resistance to nucleases and proteases, hybridization with DNA or RNA independent
of salt concentration, and triplex formation with homopurine DNA. PANAGENE™ has developed its proprietary Bts PNA monomers (Bts; benzothiazole-2-sulfonyl group) and proprietary oligomerization process. The PNA oligomerization using Bts PNA monomers is composed of repetitive cycles of deprotection, coupling, and capping. PNAs can be produced synthetically using any technique known in the art. See, e.g., U.S. Pat. Nos.: 6,969,766; 7,211,668; 7,022,851; 7,125,994; 7,145,006; and 7,179,896. See also U.S. Pat. Nos.: 5,539,082; 5,714,331; and 5,719,262 for the preparation of PNAs. Further teaching of PNA compounds can be found in Nielsen et al., Science, 254:1497-1500, 1991. Each of the foregoing is incorporated by reference in its entirety. 2. Locked Nucleic Acids (LNAs) [0375] Antisense oligomers may also contain "locked nucleic acid" subunits (LNAs). "LNAs" are a member of a class of modifications called bridged nucleic acid (BNA). BNA is characterized by a covalent linkage that locks the conformation of the ribose ring in a C30-endo (northern) sugar pucker. For LNA, the bridge is composed of a methylene between the 2′-O and the 4′-C positions. LNA enhances backbone preorganization and base stacking to increase hybridization and thermal stability. [0376] The structures of LNAs can be found, for example, in Wengel, et al., Chemical Communications (1998) 455; Koshkin et al., Tetrahedron (1998) 54:3607; Jesper Wengel, Accounts of Chem. Research (1999) 32:301; Obika, et al., Tetrahedron Letters (1997) 38:8735; Obika, et al., Tetrahedron Letters (1998) 39:5401; and Obika, et al., Bioorganic Medicinal Chemistry (2008) 16:9230, which are hereby incorporated by reference in their entirety. A non- limiting example of an LNA is depicted below.
[0377] Antisense oligomers of the disclosure may incorporate one or more LNAs; in some cases, the antisense oligomers may be entirely composed of LNAs. Methods for the synthesis of
individual LNA nucleoside subunits and their incorporation into oligomers are described, for example, in U.S. Pat.: Nos.7,572,582; 7,569,575; 7,084,125; 7,060,809; 7,053,207; 7,034,133; 6,794,499; and 6,670,461; each of which is incorporated by reference in its entirety. Typical intersubunit linkers include phosphodiester and phosphorothioate moieties; alternatively, non- phosphorous-containing linkers may be employed. Further embodiments include an LNA containing antisense oligomer where each LNA subunit is separated by a DNA subunit. Certain antisense oligomers are composed of alternating LNA and DNA subunits where the intersubunit linker is phosphorothioate. 3. Ethylene-Bridged Nucleic Acids (ENAs) [0378] 2′-O,4′C-ethylene-bridged nucleic acids (ENAs) are another member of the class of BNAs. A non-limiting example is depicted below.
[0379] ENA oligomers and their preparation are described in Obika et al., Tetrahedron Lett (1997) 38 (50): 8735, which is hereby incorporated by reference in its entirety. Antisense oligomers of the disclosure may incorporate one or more ENA subunits. 4. Unlocked nucleic acid (UNA) [0380] Antisense oligomers may also contain unlocked nucleic acid (UNA) subunits. UNAs and UNA oligomers are an analog of RNA in which the C2′-C3′ bond of the subunit has been cleaved. Whereas LNA is conformationally restricted (relative to DNA and RNA), UNA is very flexible. UNAs are disclosed, for example, in WO 2016/070166. A non-limiting example of an UNA is depicted below.
[0381] Typical intersubunit linkers include phosphodiester and phosphorothioate moieties; alternatively, non-phosphorous containing linkers may be employed. 5. Phosphorothioates [0382] "Phosphorothioates" (or S-oligos) are a variant of normal DNA in which one of the nonbridging oxygens is replaced by sulfur. A non-limiting example of a phosphorothioate is depicted below.
[0383] The sulfurization of the internucleotide bond reduces the action of endo-and exonucleases including 5′ to 3′ and 3′ to 5′ DNA POL 1 exonuclease, nucleases S1 and P1, RNases, serum nucleases, and snake venom phosphodiesterase. Phosphorothioates are made by two principal routes: by the action of a solution of elemental sulfur in carbon disulfide on a hydrogen phosphonate, or by the method of sulfurizing phosphite triesters with either tetraethylthiuram disulfide (TETD) or 3H-1, 2-benzodithiol-3-one 1, 1-dioxide (BDTD) (see, e.g., Iyer et al., J. Org. Chem.55, 4693-4699, 1990, which is hereby incorporated by reference in its entirety). The latter methods avoid the problem of elemental sulfur's insolubility in most organic solvents and the toxicity of carbon disulfide. The TETD and BDTD methods also yield higher purity phosphorothioates. 6. Tricyclo-DNAs and Tricyclo-Phosphorothioate Subunits [0384] Tricyclo-DNAs (tc-DNA) are a class of constrained DNA analogs in which each nucleotide is modified by the introduction of a cyclopropane ring to restrict conformational flexibility of the backbone and to optimize the backbone geometry of the torsion angle γ. Homobasic adenine- and thymine-containing tc-DNAs form extraordinarily stable A-T base pairs with complementary RNAs. Tricyclo-DNAs and their synthesis are described in International Patent Application Publication No. WO 2010/115993, which is hereby incorporated by reference in its entirety. Antisense oligomers of the disclosure may incorporate one or more tricycle-DNA
subunits; in some cases, the antisense oligomers may be entirely composed of tricycle-DNA subunits. [0385] Tricyclo-phosphorothioate subunits are tricyclo-DNA subunits with phosphorothioate intersubunit linkages. Tricyclo-phosphorothioate subunits and their synthesis are described in International Patent Application Publication No. WO 2013/053928, which is hereby incorporated by reference in its entirety. Antisense oligomers of the disclosure may incorporate one or more tricycle-DNA subunits; in some cases, the antisense oligomers may be entirely composed of tricycle-DNA subunits. A non-limiting example of a tricycle-DNA/tricycle- phosphorothioate subunit is depicted below.
7. 2′-O-Methyl, 2′-O-MOE, and 2′-F Oligomers [0386] "2′-O-Me oligomer" molecules carry a methyl group at the 2′-OH residue of the ribose molecule.2′-O-Me-RNAs show the same (or similar) behavior as DNA but are protected against nuclease degradation.2′-O-Me-RNAs can also be combined with phosphorothioate oligomers (PTOs) for further stabilization.2′O-Me oligomers (phosphodiester or phosphorothioate) can be synthesized according to routine techniques in the art (see, e.g., Yoo et al., Nucleic Acids Res. 32:2008-16, 2004, which is hereby incorporated by reference in its entirety). A non-limiting example of a 2′-O-Me oligomer is depicted below.
2′-O-Me [0387] 2′-O-Methoxyethyl Oligomers (2′-O-MOE) carry a methoxyethyl group at the 2′-OH residue of the ribose molecule and are discussed in Martin et al., Helv. Chim. Acta, 78, 486-504,
1995, which is hereby incorporated by reference in its entirety. A non-limiting example of a 2′-O- MOE subunit is depicted below.
[0388] 2′-Fluoro (2′-F) oligomers have a fluoro radical in at the 2′ position in place of the 2′-OH. A non-limiting example of a 2′-F oligomer is depicted below.
[0389] 2′-fluoro oligomers are further described in WO 2004/043977, which is hereby incorporated by reference in its entirety. [0390] 2′-O-Methyl, 2′-O-MOE, and 2′-F oligomers may also comprise one or more phosphorothioate (PS) linkages as depicted below.
2′-O-Methyl PS 2′-O-MOE PS 2′-F PS [0391] Additionally, 2′-O-Methyl, 2′-O-MOE, and 2′-F oligomers may comprise PS intersubunit linkages throughout the oligomer, for example, as in the 2′-O-methyl PS oligomer drisapersen depicted below.
[0392] Alternatively, 2′-O-Methyl, 2′-O-MOE, and/or 2′-F oligomers may comprise PS linkages at the ends of the oligomer, as depicted below:
wherein: R is CH2CH2OCH3 (methoxyethyl or MOE); and X, Y, and Z denote the number of nucleotides contained within each of the designated 5′-wing, central gap, and 3′-wing regions, respectively. [0393] Antisense oligomers of the disclosure may incorporate one or more 2′-O-methyl, 2′-O- MOE, and 2′-F subunits and may utilize any of the intersubunit linkages described here. In some instances, an antisense oligomer of the disclosure may be composed of entirely 2′-O-methyl, 2′- O-MOE, or 2′-F subunits. One embodiment of the antisense oligomers of the disclosure is composed entirely of 2′-O-methyl subunits. 8. 2′-O-[2-(N-methylcarbamoyl)ethyl] Oligomers (MCEs) [0394] MCEs are another example of 2′-O modified ribonucleosides useful in the antisense oligomers of the disclosure. Here, the 2′-OH is derivatized to a 2-(N-methylcarbamoyl)ethyl moiety to increase nuclease resistance. A non-limiting example of an MCE oligomer is depicted below.
[0395] MCEs and their synthesis are described in Yamada et al., J. Org. Chem. (2011) 76(9):3042-53, which is hereby incorporated by reference in its entirety. Antisense oligomers of the disclosure may incorporate one or more MCE subunits. 9. Stereo-Specific Oligomers [0396] Stereo-specific oligomers are those in which the stereo chemistry of each phosphorous- containing linkage is fixed by the method of synthesis such that a substantially stereo-pure oligomer is produced. A non-limiting example of a stereo-specific oligomer is depicted below.
[0397] In the above example, each phosphorous of the oligomer has the same stereo configuration. Additional examples include the oligomers described herein. For example, LNAs, ENAs, Tricyclo-DNAs, MCEs, 2′-O-Methyl, 2′-O-MOE, 2′-F, and morpholino-based oligomers can be prepared with stereo-specific phosphorous-containing internucleoside linkages such as, for example, phosphorothioate, phosphodiester, phosphoramidate, phosphorodiamidate, or
other phosphorous-containing internucleoside linkages. Stereo specific oligomers, methods of preparation, chiral controlled synthesis, chiral design, and chiral auxiliaries for use in the preparation of such oligomers are detailed, for example, in WO2017192664, WO2017192679, WO2017062862, WO2017015575, WO2017015555, WO2015107425, WO2015108048, WO2015108046, WO2015108047, WO2012039448, WO2010064146, WO2011034072, WO2014010250, WO2014012081, WO20130127858, and WO2011005761, each of which is hereby incorporated by reference in its entirety. [0398] Stereo-specific oligomers can have phosphorous-containing internucleoside linkages in an RP or SP configuration. Chiral phosphorous-containing linkages in which the stereo configuration of the linkages is controlled is referred to as "stereopure," while chiral phosphorous-containing linkages in which the stereo configuration of the linkages is uncontrolled is referred to as "stereorandom." In certain embodiments, the oligomers of the disclosure comprise a plurality of stereopure and stereorandom linkages, such that the resulting oligomer has stereopure subunits at pre-specified positions of the oligomer. An example of the location of the stereopure subunits is provided in international patent application publication number WO 2017/062862 A2 in Figures 7A and 7B. In an embodiment, all the chiral phosphorous-containing linkages in an oligomer are stereorandom. In an embodiment, all the chiral phosphorous-containing linkages in an oligomer are stereopure. [0399] In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), all n of the chiral phosphorous-containing linkages in the oligomer are stereorandom. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), all n of the chiral phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 10% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 20% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous- containing linkages (where n is an integer of 1 or greater), at least 30% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 40% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 50% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an
oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 60% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 70% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 80% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), at least 90% (to the nearest integer) of the n phosphorous-containing linkages in the oligomer are stereopure. [0400] In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 2 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 3 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 4 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 5 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 6 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 7 contiguous stereopure phosphorous- containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 8 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 9 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 10 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or
RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 11 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 12 contiguous stereopure phosphorous- containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 13 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 14 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 15 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 16 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 17 contiguous stereopure phosphorous- containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 18 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 19 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 20 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP). [0401] In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 2 contiguous stereopure phosphorous-containing linkages of the same stereo orientation (i.e., either SP or RP) and at least 2 contiguous stereopure phosphorous-containing linkages of the other stereo orientation. For example, the oligomer can contain at least 2 contiguous stereopure phosphorous-containing
linkages of the SP orientation and at least 2 contiguous stereopure phosphorous-containing linkages of the RP orientation. [0402] In an embodiment of an oligomer with n chiral phosphorous-containing linkages (where n is an integer of 1 or greater), the oligomer contains at least 2 contiguous stereopure phosphorous-containing linkages of the same stereo orientation in an alternating pattern. For example, the oligomer can contain the following in order: 2 or more RP, 2 or more SP, and 2 or more RP, etc. 10. Morpholino Oligomers [0403] Exemplary embodiments of the disclosure relate to phosphorodiamidate morpholino oligomers of the following general structure:
and as described in Figure 2 of Summerton, J., et al., Antisense & Nucleic Acid Drug Development, 7: 187-195 (1997). Morpholinos as described herein are intended to cover all stereoisomers and tautomers of the foregoing general structure. The synthesis, structures, and binding characteristics of morpholino oligomers are detailed in U.S. Patent Nos.: 5,698,685; 5,217,866; 5,142,047; 5,034,506; 5,166,315; 5,521,063; 5,506,337; 8,076,476; and 8,299,206, all of which are incorporated herein by reference. [0404] In certain embodiments, a morpholino is conjugated at the 5′ or 3′ end of the oligomer with a "tail" moiety to increase its stability and/or solubility. Exemplary tails include: [0405] In various aspects, the disclosure provides antisense oligomers according to Formulae (I), (IA), (II), (III), (IV), and/or (V), and pharmaceutically acceptable salts thereof. [0406] In an embodiment, the antisense oligomers disclosed herein, including antisense oligomers of Formulae (I), (IA), (II), (III), (IV), and/or (V), comprise a targeting sequence complementary to a target region within a pre-mRNA of the human DNM2 gene represented by SEQ ID NO: 1. [0407] In some embodiments of the antisense oligomers disclosed herein, including antisense oligomers of Formula (I), Formula (IA), Formula (II), Formula (III), Formula (IV), and/or Formula
(V), the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 6 (SEQ ID NO: 3), exon 7 (SEQ ID NO: 4), exon 8 (SEQ ID NO: 5), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), exon 13 (SEQ ID NO: 8), exon 16 (SEQ ID NO: 9), and exon 19 (SEQ ID NO: 10). [0408] In some embodiments of the antisense oligomers disclosed herein, including antisense oligomers of Formula (I), Formula (IA), Formula (II), Formula (III), and Formula (IV), the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), and exon 13 (SEQ ID NO: 8). [0409] In some embodiments of the antisense oligomers disclosed herein, including antisense oligomers of Formula (I), Formula (IA), Formula (II), Formula (III), Formula (IV), and/or Formula (V), the target region is within an intron-exon junction or an exon-intron junction region of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 5/exon 6 junction, exon 6/intron 6 junction, intron 6/exon 7 junction, exon 7/intron 7 junction, intron 7/exon 8 junction, exon 8/intron 8 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, exon 13/intron 13 junction, intron 15/exon 16 junction, exon 16/intron 16 junction, intron 18/exon 19 junction, and exon 19/intron 19 junction. [0410] In some embodiments of the antisense oligomers disclosed herein, including antisense oligomers of Formula (I), Formula (IA), Formula (II), Formula (III), Formula (IV), and/or Formula (V), the target region is within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, and exon 13/intron 13 junction. [0411] In some embodiments of the antisense oligomers disclosed herein, including antisense oligomers of Formula (I), Formula (IA), Formula (II), Formula (III), Formula (IV), and/or Formula (V), the target region is within an intron internal region of the human DNM2 gene pre-mRNA selected from intron 2 (SEQ ID NO: 11) and intron 12 (SEQ ID NO: 12). [0412] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12- 13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-
27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), DNM2 H2D(-6-30), DNM2 H9A(-12+13), DNM2 H9A(-5+20), DNM2 H9A(+1+25),DNM2 H9A(+6+30), DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), DNM2 H9A(+48+67), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11- 14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4- 17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), DNM2 H9D(+3-22), DNM2 H12A(-13+12), DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), DNM2 H12D(+15-10), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(- 25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(- 11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2
H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), DNM2 H13A(+9+28), DNM2 H13A(+16+40), DNM2 H13D(+22-3), or DNM2 H13D(+7-18). [0413] In certain embodiments, the target region is DNM2 H2A (-10+15). In certain embodiments, the target region is DNM2 H2A(-7+17). In certain embodiments, the target region is DNM2 H2A(+3+27). In certain embodiments, the target region is DNM2 H2A(+15+37). In certain embodiments, the target region is DNM2 H2A(+21+45). In certain embodiments, the target region is DNM2 H2A(+23+47). In certain embodiments, the target region is DNM2 H2A(+27+51). In certain embodiments, the target region is DNM2 H2A(+37+61). In certain embodiments, the target region is DNM2 H2A(+46+70). In certain embodiments, the target region is DNM2 H2A (+48+72). In certain embodiments, the target region is DNM2 H2D(+20-5). In certain embodiments, the target region is DNM2 H2D (+13-12). In certain embodiments, the target region is DNM2 H2D(+12-13). In certain embodiments, the target region is DNM2 H2D(+11-14). In certain embodiments, the target region is DNM2 H2D(+10-15). In certain embodiments, the target region is DNM2 H2D(+9-16). In certain embodiments, the target region is DNM2 H2D(+8-17). In certain embodiments, the target region is DNM2 H2D(+7-18). In certain embodiments, the target region is DNM2 H2D(+6-19). In certain embodiments, the target region is DNM2 H2D(+5-20). In certain embodiments, the target region is DNM2 H2D(+4-21). In certain embodiments, the target region is DNM2 H2D(+3-22). In certain embodiments, the target region is DNM2 H2D(+2-23) . In certain embodiments, the target region is DNM2 H2D(+1-24). In certain embodiments, the target region is DNM2 H2D(-1-25). In certain embodiments, the target region is DNM2 H2D(-2-26). In certain embodiments, the target region is DNM2 H2D(-3-27). In certain embodiments, the target region is DNM2 H2D(-4-28). In certain embodiments, the target region is DNM2 H2D(-5-29). In certain embodiments, the target region is DNM2 H2D(-6-30). In certain embodiments, the target region is DNM2 H6A(-20+5). In certain embodiments, the target region is DNM2 H6A(-6+19). In certain embodiments, the target region is DNM2 H6A(-2+23). In certain embodiments, the target region is DNM2 H6A(+3+26). In certain embodiments, the target region is DNM2 H6A(+10+3). In certain embodiments, the target region is DNM2 H6A(+13+37). In certain embodiments, the target region is DNM2 H6A(+23+47). In certain embodiments, the target region is DNM2 H6A(+32+56). In certain embodiments, the target region is DNM2 H6A(+35+59). In certain embodiments, the target region is DNM2 H6A(+44+68). In certain embodiments, the target region is DNM2 H6A(+54+78). In certain embodiments, the target region is DNM2 H6A(+60+84). In certain embodiments, the target region is DNM2 H6A(+63+87). In certain embodiments, the target region is DNM2 H6A(+88+112). In certain
embodiments, the target region is DNM2 H6A(+96+119). In certain embodiments, the target region is DNM2 H6A(+105+129). In certain embodiments, the target region is DNM2 H6A(+121+145). In certain embodiments, the target region is DNM2 H6A(+130+154). In certain embodiments, the target region is DNM2 H6A(+132+156). In certain embodiments, the target region is DNM2 H6D (+6-19). In certain embodiments, the target region is DNM2 H6D (+15-10). In certain embodiments, the target region is DNM2 H6D (+17-8). In certain embodiments, the target region is DNM2 H7A(-15+10). In certain embodiments, the target region is DNM2 H7A(- 5+20). In certain embodiments, the target region is DNM2 H7A(+1+25). In certain embodiments, the target region is DNM2 H7A(+6+30). In certain embodiments, the target region is. In certain embodiments, the target region is DNM2 H7A(+22+46). In certain embodiments, the target region is DNM2 H7A(+29+53). In certain embodiments, the target region is DNM2 H7A(+41+65). In certain embodiments, the target region is DNM2 H7A(+51+75). In certain embodiments, the target region is DNM2 H7A(+57+81). In certain embodiments, the target region is DNM2 H7A(+62+86). In certain embodiments, the target region is DNM2 H7A(+72+96). In certain embodiments, the target region is DNM2 H7A(+74+98). In certain embodiments, the target region is DNM2 H7A(+81+105). In certain embodiments, the target region is DNM2 H7A(+91+115). In certain embodiments, the target region is DNM2 H7A(+110+134). In certain embodiments, the target region is DNM2 H7D(+8-16). In certain embodiments, the target region is DNM2 H7D(+9-16). In certain embodiments, the target region is DNM2 H7D(+21-4). In certain embodiments, the target region is DNM2 H8A(-20+5). In certain embodiments, the target region is DNM2 H8A(-14+11). In certain embodiments, the target region is DNM2 H8A(-7+18). In certain embodiments, the target region is DNM2 H8A(-6+19). In certain embodiments, the target region is DNM2 H8A(-2+23). In certain embodiments, the target region is DNM2 H8A(+5+29). In certain embodiments, the target region is DNM2 H8A(+18+42). In certain embodiments, the target region is DNM2 H8A(+21+45). In certain embodiments, the target region is DNM2 H8A(+27+51). In certain embodiments, the target region is DNM2 H8A(+36+60). In certain embodiments, the target region is DNM2 H8A(+46+70). In certain embodiments, the target region is DNM2 H8A(+51+75). In certain embodiments, the target region is DNM2 H8A(+56+80). In certain embodiments, the target region is DNM2 H8A(+65+88). In certain embodiments, the target region is DNM2 H8A(+76+100). In certain embodiments, the target region is DNM2 H8A(+83+107). In certain embodiments, the target region is DNM2 H8A(+96+119). In certain embodiments, the target region is DNM2 H8A(+105+129). In certain embodiments, the target region is DNM2 H8D(+18-7). In certain embodiments, the target region is DNM2 H8D(+10-15). In certain embodiments, the target region is DNM2 H9A(-12+13). In certain embodiments, the target region is DNM2 H9A(-5+20). In certain embodiments, the target region is DNM2
H9A(+1+25). In certain embodiments, the target region is DNM2 H9A(+6+30). In certain embodiments, the target region is DNM2 H9A(+16+40). In certain embodiments, the target region is DNM2 H9A(+25+49). In certain embodiments, the target region is DNM2 H9A(+36+60). In certain embodiments, the target region is DNM2 H9A(+37+61). In certain embodiments, the target region is DNM2 H9A(+42+66). In certain embodiments, the target region is DNM2 H9A(+43+62). In certain embodiments, the target region is DNM2 H9A(+43+63). In certain embodiments, the target region is DNM2 H9A(+43+64). In certain embodiments, the target region is DNM2 H9A(+43+65). In certain embodiments, the target region is DNM2 H9A(+43+66). In certain embodiments, the target region is DNM2 H9A(+43+67). In certain embodiments, the target region is DNM2 H9A(+44+67). In certain embodiments, the target region is DNM2 H9A(+44+68). In certain embodiments, the target region is DNM2 H9A(+45+67). In certain embodiments, the target region is DNM2 H9A (+46+67). In certain embodiments, the target region is DNM2 H9A(+47+67). In certain embodiments, the target region is DNM2 H9A(+48+67). In certain embodiments, the target region is DNM2 H9D(+24-1). In certain embodiments, the target region is DNM2 H9D(+23-2). In certain embodiments, the target region is DNM2 H9D(+22-3). In certain embodiments, the target region is DNM2 H9D(+21-4). In certain embodiments, the target region is DNM2 H9D(+20-5). In certain embodiments, the target region is DNM2 H9D(+19-6). In certain embodiments, the target region is DNM2 H9D (+18-7). In certain embodiments, the target region is DNM2 H9D(+17-8). In certain embodiments, the target region is DNM2 H9D(+16-9). In certain embodiments, the target region is DNM2 H9D(+15-10). In certain embodiments, the target region is DNM2 H9D(+14-11). In certain embodiments, the target region is DNM2 H9D(+13-12). In certain embodiments, the target region is DNM2 H9D(+12-13). In certain embodiments, the target region is DNM2 H9D(+11-14). In certain embodiments, the target region is DNM2 H9D(+10-15). In certain embodiments, the target region is DNM2 H9D(+9-16). In certain embodiments, the target region is DNM2 H9D(+8-12). In certain embodiments, the target region is DNM2 H9D(+8-13). In certain embodiments, the target region is DNM2 H9D(+8-14). In certain embodiments, the target region is DNM2 H9D(+8-15). In certain embodiments, the target region is DNM2 H9D(+8-17). In certain embodiments, the target region is DNM2 H9D(+7-18). In certain embodiments, the target region is DNM2 H9D(+7-17). In certain embodiments, the target region is DNM2 H9D(+6-19). In certain embodiments, the target region is DNM2 H9D(+6-17). In certain embodiments, the target region is DNM2 H9D(+5-17). In certain embodiments, the target region is DNM2 H9D(+5-20). In certain embodiments, the target region is DNM2 H9D(+4-17). In certain embodiments, the target region is DNM2 H9D(+4-21). In certain embodiments, the target region is DNM2 H9D(+3-17). In certain embodiments, the target region is DNM2 H9D(+3-22). In certain embodiments, the target region is DNM2 H12A(-13+12).
In certain embodiments, the target region is DNM2 H12A(-3+22). In certain embodiments, the target region is DNM2 H12A(+4+28). In certain embodiments, the target region is DNM2 H12A(+7+31). In certain embodiments, the target region is DNM2 H12A(+10+34). In certain embodiments, the target region is DNM2 H12A(+11+35). In certain embodiments, the target region is DNM2 H12A(+12+36). In certain embodiments, the target region is DNM2 H12A(+13+37). In certain embodiments, the target region is DNM2 H12A(+14+38). In certain embodiments, the target region is DNM2 H12A(+15+39). In certain embodiments, the target region is DNM2 H12A(+16+40). In certain embodiments, the target region is DNM2 H12A(+17+41). In certain embodiments, the target region is DNM2 H12A(+18+42). In certain embodiments, the target region is DNM2 H12A(+19+43). In certain embodiments, the target region is DNM2 H12A(+20+44). In certain embodiments, the target region is DNM2 H12A(+25+44). In certain embodiments, the target region is DNM2 H12A(+25+45). In certain embodiments, the target region is DNM2 H12A(+25+46). In certain embodiments, the target region is DNM2 H12A(+25+47). In certain embodiments, the target region is DNM2 H12A(+25+48). In certain embodiments, the target region is DNM2 H12A(+25+49). In certain embodiments, the target region is DNM2 H12A(+26+49). In certain embodiments, the target region is DNM2 H12A(+27+49). In certain embodiments, the target region is DNM2 H12A(+28+49). In certain embodiments, the target region is DNM2 H12A(+29+49). In certain embodiments, the target region is DNM2 H12A(+29+53). In certain embodiments, the target region is DNM2 H12A(+30+49). In certain embodiments, the target region is DNM2 H12A(+30+54). In certain embodiments, the target region is DNM2 H12A(+36+60). In certain embodiments, the target region is DNM2 H12A(+37+61). In certain embodiments, the target region is DNM2 H12A(+43+67). In certain embodiments, the target region is DNM2 H12D(+22- 3). In certain embodiments, the target region is DNM2 H12D(+20-5). In certain embodiments, the target region is DNM2 H12D(+15-10). In certain embodiments, the target region is DNM2 H13A(-27-3). In certain embodiments, the target region is DNM2 H13A(-26-2). In certain embodiments, the target region is DNM2 H13A(-25-1). In certain embodiments, the target region is DNM2 H13A(-24+1). In certain embodiments, the target region is DNM2 H13A(-23+2). In certain embodiments, the target region is DNM2 H13A(-22+3). In certain embodiments, the target region is DNM2 H13A(-21+4). In certain embodiments, the target region is DNM2 H13A(- 20+5). In certain embodiments, the target region is DNM2 H13A(-19+6). In certain embodiments, the target region is DNM2 H13A(-18+7). In certain embodiments, the target region is DNM2 H13A(-17+8). In certain embodiments, the target region is DNM2 H13A(-16+9). In certain embodiments, the target region is DNM2 H13A(-15+10). In certain embodiments, the target region is DNM2 H13A(-14+11). In certain embodiments, the target region is DNM2 H13A(-
13+12). In certain embodiments, the target region is DNM2 H13A(-11+14). In certain embodiments, the target region is DNM2 H13A(-10+15). In certain embodiments, the target region is DNM2 H13A(-9+16). In certain embodiments, the target region is DNM2 H13A(-8+17). In certain embodiments, the target region is DNM2 H13A(-7+18). In certain embodiments, the target region is DNM2 H13A(-6+19). In certain embodiments, the target region is DNM2 H13A(- 5+20). In certain embodiments, the target region is DNM2 H13A(-4+21). In certain embodiments, the target region is DNM2 H13A(-3+22). In certain embodiments, the target region is DNM2 H13A(-2+23). In certain embodiments, the target region is DNM2 H13A(-1+24). In certain embodiments, the target region is DNM2 H13A(+1+25). In certain embodiments, the target region is DNM2 H13A(+2+25). In certain embodiments, the target region is DNM2 H13A(+3+27). In certain embodiments, the target region is DNM2 H13A(+4+23). In certain embodiments, the target region is DNM2 H13A(+4+24). In certain embodiments, the target region is DNM2 H13A(+4+25). In certain embodiments, the target region is DNM2 H13A(+4+26). In certain embodiments, the target region is DNM2 H13A(+4+27). In certain embodiments, the target region is DNM2 H13A(+4+28). In certain embodiments, the target region is DNM2 H13A(+5+28). In certain embodiments, the target region is DNM2 H13A(+6+28). In certain embodiments, the target region is DNM2 H13A(+6+30). In certain embodiments, the target region is DNM2 H13A(+7+28). In certain embodiments, the target region is DNM2 H13A(+8+28). In certain embodiments, the target region is DNM2 H13A(+9+28). In certain embodiments, the target region is DNM2 H13A(+16+40). In certain embodiments, the target region is DNM2 H13D(+22-3). In certain embodiments, the target region is DNM2 H13D(+7-18). In certain embodiments, the target region is DNM2 H16A(-21+3). In certain embodiments, the target region is DNM2 H16A(-13+12). In certain embodiments, the target region is DNM2 H16A(- 6+19). In certain embodiments, the target region is DNM2 H16A(+6+30). In certain embodiments, the target region is DNM2 H16A(+15+39). In certain embodiments, the target region is DNM2 H16A(+26+50). In certain embodiments, the target region is DNM2 H16A(+36+60). In certain embodiments, the target region is DNM2 H16A(+45+69). In certain embodiments, the target region is DNM2 H16A(+55+79). In certain embodiments, the target region is DNM2 H16A(+66+90). In certain embodiments, the target region is DNM2 H16A(+75+99). In certain embodiments, the target region is DNM2 H16A(+82+106). In certain embodiments, the target region is DNM2 H16D(+19-6). In certain embodiments, the target region is DNM2 H16D(+11-14). In certain embodiments, the target region is DNM2 H19A(- 11+14). In certain embodiments, the target region is DNM2 H19A(-4+21). In certain embodiments, the target region is DNM2 H19A(+1+25). In certain embodiments, the target region is DNM2 H19A(+7+31). In certain embodiments, the target region is DNM2
H19A(+16+40). In certain embodiments, the target region is DNM2 H19A(+26+50). In certain embodiments, the target region is DNM2 H19A(+37+61). In certain embodiments, the target region is DNM2 H19A(+47+71). In certain embodiments, the target region is DNM2 H19A(+56+80). In certain embodiments, the target region is DNM2 H19A(+66+90). In certain embodiments, the target region is DNM2 H19A(+76+100). In certain embodiments, the target region is DNM2 H19A(+86+110). In certain embodiments, the target region is DNM2 H19A(+96+120). In certain embodiments, the target region is DNM2 H19A(+106+130). In certain embodiments, the target region is DNM2 H19A(+117+141). In certain embodiments, the target region is DNM2 H19A(+127+151). In certain embodiments, the target region is DNM2 H19A(+137+161). In certain embodiments, the target region is DNM2 H19A(+147+171). In certain embodiments, the target region is DNM2 H19A(+157+181). In certain embodiments, the target region is DNM2 H19A(+166+189). In certain embodiments, the target region is DNM2 H19A(+191+215). In certain embodiments, the target region is DNM2 H19A(+200+224). In certain embodiments, the target region is DNM2 H19D(+23-2). In certain embodiments, the target region is DNM2 H19D(+17-8). In certain embodiments, the target region is DNM2 H19D(+10-15). [0414] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGAGGCCGC); SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAA); SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC); SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT);
SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT); SEQ ID NO: 90 (GAACCTCAGGCCGCCCCATTTTACC); SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT); SEQ ID NO: 96 (CCTGCTGAACCTCAGGCCGCCCCAT); SEQ ID NO: 97 (GTAGCCTGTGGGGAGAGGCAGTCAG); SEQ ID NO: 98 (TTCACCACGCCAATGTAGCCTGTGG); SEQ ID NO: 99 (GCGGTTCACCACGCCAATGTAGCCT); SEQ ID NO: 100 (GCTGCGGTTCACCACGCCAATGTA); SEQ ID NO: 101 (TCCTTCTGGCTGCGGTTCACCACGC); SEQ ID NO: 102 (ATATCCTTCTGGCTGCGGTTCACCA); SEQ ID NO: 103 (CTTGCCCTCAATATCCTTCTGGCTG); SEQ ID NO: 104 (GATGTCCTTCTTGCCCTCAATATCC); SEQ ID NO: 105 (ACGGATGTCCTTCTTGCCCTCAATA); SEQ ID NO: 106 (CAGTGCTGCACGGATGTCCTTCTTG); SEQ ID NO: 107 (TCTCAGCTGCCAGTGCTGCACGGAT); SEQ ID NO: 108 (ACTTCCTCTCAGCTGCCAGTGCTGC); SEQ ID NO: 109 (AGAACTTCCTCTCAGCTGCCAGTGC); SEQ ID NO: 110 (TGCCGGTAGGCCGGGTGGGAGAGGA); SEQ ID NO: 111 (GGCCATGTGCCGGTAGGCCGGGTG); SEQ ID NO: 112 (CCATGCGGTCGGCCATGTGCCGGTA); SEQ ID NO: 113 (TGCAGATGTGGCGTGCCCATGCGGT); SEQ ID NO: 114 (AGCGTCTTCTGCAGATGTGGCGTGC); SEQ ID NO: 115 (TCAGCGTCTTCTGCAGATGTGGCGT); SEQ ID NO: 116 (GTGCAAACCCTTGCAGTACCTGATT); SEQ ID NO: 117 (CTTGCAGTACCTGATTCAGCGTCTT); SEQ ID NO: 118 (TGCAGTACCTGATTCAGCGTCTTCT); SEQ ID NO: 119 (TGGTCAGTTGCTGCAGGAGAGAGGT);
SEQ ID NO: 120 (CGGATGTGGTTGGTCAGTTGCTGCA); SEQ ID NO: 121 (ACTCCCGGATGTGGTTGGTCAGTTG); SEQ ID NO: 122 (CAGCGACTCCCGGATGTGGTTGGTC); SEQ ID NO: 123 (GCCGGCAGCGACTCCCGGATGTGGT); SEQ ID NO: 124 (TGCTACGTAGGGCCGGCAGCGACTC); SEQ ID NO: 125 (TGTAGTTTGCTACGTAGGGCCGGCA); SEQ ID NO: 126 (AGCAGCTGGCTCTGTAGTTTGCTAC); SEQ ID NO: 127 (CTCCAGGGACAGCAGCTGGCTCTGT); SEQ ID NO: 128 (CTCCTTCTCCAGGGACAGCAGCTGG); SEQ ID NO: 129 (TCCACCTCCTTCTCCAGGGACAGCA); SEQ ID NO: 130 (CTTGTACTCCTCCACCTCCTTCTCC); SEQ ID NO: 131 (TTCTTGTACTCCTCCACCTCCTTCT); SEQ ID NO: 132 (CCGAAAGTTCTTGTACTCCTCCACC); SEQ ID NO: 133 (GGTCGTCGGGCCGAAAGTTCTTGTA); SEQ ID NO: 134 (GCTTTGGTTTTGCGGGTGGGGTCGT); SEQ ID NO: 135 (GTGCCGBGGTACATACTGCAGCAG); SEQ ID NO: 136 (GTGCCGGGGTACATACTGCAGCAGG); SEQ ID NO: 137 (ATACTGCAGCAGGGCTTTGGTTTTG); SEQ ID NO: 138 (CCATCCTATGAGGAAAAGGAAAAAG); SEQ ID NO: 139 (GCTGGACCATCCTATGAGGAAAAGG); SEQ ID NO: 140 (CCAAACTGCTGGACCATCCTATGAG); SEQ ID NO: 141 (CCCAAACTGCTGGACCATCCTATGA); SEQ ID NO: 142 (CCACCCCAAACTGCTGGACCATCCT); SEQ ID NO: 143 (CAAAATCCACCCCAAACTGCTGGAC); SEQ ID NO: 144 (TCGATCCTCTTCTCAAAATCCACCC); SEQ ID NO: 145 (CCCTCGATCCTCTTCTCAAAATCCA); SEQ ID NO: 146 (CCTGAGCCCTCGATCCTCTTCTCAA); SEQ ID NO: 147 (ACCTGATCTCCTGAGCCCTCGATCC); SEQ ID NO: 148 (CAGAGTGTCCACCTGATCTCCTGAG); SEQ ID NO: 149 (AGCTCCAGAGTGTCCACCTGATCTC); SEQ ID NO: 150 (CGGAGAGCTCCAGAGTGTCCACCTG); SEQ ID NO: 151 (GGCGCCCCCGGAGAGCTCCAGAGT); SEQ ID NO: 152 (GCGATTGATTCGGGCGCCCCCGGAG); SEQ ID NO: 153 (GGAAGATGCGATTGATTCGGGCGCC); SEQ ID NO: 154 (GGAACCGCTCGTGGAAGATGCGAT);
SEQ ID NO: 155 (AGCTCAAATGGGAACCGCTCGTGGA); SEQ ID NO: 156 (GCACTACCTTCACCAGCTCAAATGG); SEQ ID NO: 157 (CCCGGGGGGCACTACCTTCACCAGC); SEQ ID NO: 158 (CGTCAAACTCCATCTGAGAAGAGAC); SEQ ID NO: 159 (TCCTTCTCGTCAAACTCCATCTGAG); SEQ ID NO: 160 (GTAAGTCCTTCTCGTCAAACTCCAT); SEQ ID NO: 161 (CCGTCGTAAGTCCTTCTCGTCAAAC); SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG); SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT); SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 175 (TGACTCCATGGATGTTCTTAATG); SEQ ID NO: 176 (TGACTCCATGGATGTTCTTAAT); SEQ ID NO: 177 (TGACTCCATGGATGTTCTTAA); SEQ ID NO: 178 (TGACTCCATGGATGTTCTTA); SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT); SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG); SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT);
SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT); SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 208 (CTCCTCGTGGAACTTGCCTG); SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG); SEQ ID NO: 210 (CAGCAGAAGAATCTGGBGGAGAGCG); SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG); SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA);
SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG); SEQ ID NO: 238 (GATGAAGTCCTCATGGTTCGTGTTG); SEQ ID NO: 239 (CGATGAAGTCCTCATGGTTCGTGTT); SEQ ID NO: 240 (CAAACCCGATGAAGTCCTCATGGTT); SEQ ID NO: 241 (TACTTGGCAAACCCGATGAAGTCCT); SEQ ID NO: 242 (CCTACTTGGCAAACCCGATGAAGTC); SEQ ID NO: 243 (AAGTACCTACTTGGCAAACCCGATG); SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA); SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA);
SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC); SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 282 (GTTCAGCTGCGTGCTCCTCTGC); SEQ ID NO: 283 (GTTCAGCTGCGTGCTCCTCTG); SEQ ID NO: 284 (GTTCAGCTGCGTGCTCCTCT); SEQ ID NO: 285 (GGCTCTCTTCTTGTTCAGCTGCGTG); SEQ ID NO: 286 (TACCTGATTGGBGATGGCTCTCTTC); SEQ ID NO: 287 (TGCAGAGGBGTGTGCTACCTGATTG); SEQ ID NO: 288 (CTCCTGAGGGTGGAGGGAAAAGAG); SEQ ID NO: 289 (CTTCTCTTTCTCCTGAGGGTGGAGG); SEQ ID NO: 290 (TGTACTTCTTCTCTTTCTCCTGAGG); SEQ ID NO: 291 (CAGAGGCAGCATGTACTTCTTCTCT); SEQ ID NO: 292 (GAGGTTGTCCAGAGGCAGCATGTAC); SEQ ID NO: 293 (TCACGGATCTTGAGGTTGTCCAGAG); SEQ ID NO: 294 (CTTCTCCACATCACGGATCTTGAGG);
SEQ ID NO: 295 (CATGAAGCCCTTCTCCACATCACGG); SEQ ID NO: 296 (GCTTGTTGGACATGAAGCCCTTCTC); SEQ ID NO: 297 (GGCGAAGACGTGCTTGTTGGACATG); SEQ ID NO: 298 (GTTGAAGATGGCGAAGACGTGCTTG); SEQ ID NO: 299 (GCTCCGTGTTGAAGATGGCGAAGAC); SEQ ID NO: 300 (CCTCACCTCTGCTCCGTGTTGAAGA); SEQ ID NO: 301 (CCTGGGCCCCTCACCTCTGCTCCGT); SEQ ID NO: 302 (ATGAAGGCCTTCGTCTGCGGGCAGG); SEQ ID NO: 303 (GTGGTGGATGAAGGCCTTCGTCTGC); SEQ ID NO: 304 (GCTCGTGGTGGATGAAGGCCTTCGT); SEQ ID NO: 305 (CCAGCAGCTCGTGGTGGATGAAGGC); SEQ ID NO: 306 (ATAGGTAGGCCAGCAGCTCGTGGTG); SEQ ID NO: 307 (GCCGAGGAGTATAGGTAGGCCAGCA); SEQ ID NO: 308 (TGCTCTGGTCTGCCGAGGAGTATAG); SEQ ID NO: 309 (TCCATGAGGCTGCTCTGGTCTGCCG); SEQ ID NO: 310 (GCCGACTCCTCCATGAGGCTGCTCT); SEQ ID NO: 311 (TGCCTGGTCAGCCGACTCCTCCATG); SEQ ID NO: 312 (CCCGCCGCTGTGCCTGGTCAGCCGA); SEQ ID NO: 313 (AGCATGTCGTCCCGCCGCTGTGCCT); SEQ ID NO: 314 (GTACATGCGCAGCATGTCGTCCCGC); SEQ ID NO: 315 (TGAGGGCATGGTACATGCGCAGCAT); SEQ ID NO: 316 (GAGCGCCTCCTTGAGGGCATGGTAC); SEQ ID NO: 317 (CGATGATGTTGAGCGCCTCCTTGAG); SEQ ID NO: 318 (CTGATGTCACCGATGATGTTGAGCG); SEQ ID NO: 319 (AGTGCTGGTGCTGATGTCACCGATG); SEQ ID NO: 320 (GCGTGGACACAGTGCTGGTGCTGAT); SEQ ID NO: 321 (GGGTACAGGCGTGGACACAGTGCT); SEQ ID NO: 322 (TGGAGCCAGGTGTCATCGACAGGCG); SEQ ID NO: 323 (CTGGCGCTCTGGAGCCAGGTGTCAT); SEQ ID NO: 324 (ACCTGTGGCTGCTGGCGCTCTGGAG); SEQ ID NO: 325 (TTCCGGACCTGTGGCTGCTGGCGCT); and SEQ ID NO: 326 (ACCAGGCTTCCGGACCTGTGGCTGC). [0415] In certain embodiments, the targeting sequence is SEQ ID NO: 67. In certain embodiments, the targeting sequence is SEQ ID NO: 68. In certain embodiments, the targeting sequence is SEQ ID NO: 69. In certain embodiments, the targeting sequence is SEQ ID NO: 70.
[0416] In certain embodiments, the targeting sequence is SEQ ID NO: 71. In certain embodiments, the targeting sequence is SEQ ID NO: 72. In certain embodiments, the targeting sequence is SEQ ID NO: 73. In certain embodiments, the targeting sequence is SEQ ID NO: 74. In certain embodiments, the targeting sequence is SEQ ID NO: 75. In certain embodiments, the targeting sequence is SEQ ID NO: 76. In certain embodiments, the targeting sequence is SEQ ID NO: 77. In certain embodiments, the targeting sequence is SEQ ID NO: 78. In certain embodiments, the targeting sequence is SEQ ID NO: 79. In certain embodiments, the targeting sequence is SEQ ID NO: 80. In certain embodiments, the targeting sequence is SEQ ID NO: 81. In certain embodiments, the targeting sequence is SEQ ID NO: 82. In certain embodiments, the targeting sequence is SEQ ID NO: 83. In certain embodiments, the targeting sequence is SEQ ID NO: 84. In certain embodiments, the targeting sequence is SEQ ID NO: 85. In certain embodiments, the targeting sequence is SEQ ID NO: 86. In certain embodiments, the targeting sequence is SEQ ID NO: 87. In certain embodiments, the targeting sequence is SEQ ID NO: 88. In certain embodiments, the targeting sequence is SEQ ID NO: 89. In certain embodiments, the targeting sequence is SEQ ID NO: 90. In certain embodiments, the targeting sequence is SEQ ID NO: 91. In certain embodiments, the targeting sequence is SEQ ID NO: 92. In certain embodiments, the targeting sequence is SEQ ID NO: 93. In certain embodiments, the targeting sequence is SEQ ID NO: 94. In certain embodiments, the targeting sequence is SEQ ID NO: 95. In certain embodiments, the targeting sequence is SEQ ID NO: 96. In certain embodiments, the targeting sequence is SEQ ID NO: 97. In certain embodiments, the targeting sequence is SEQ ID NO: 98. In certain embodiments, the targeting sequence is SEQ ID NO: 99. In certain embodiments, the targeting sequence is SEQ ID NO: 100. In certain embodiments, the targeting sequence is SEQ ID NO: 101. In certain embodiments, the targeting sequence is SEQ ID NO: 102. In certain embodiments, the targeting sequence is SEQ ID NO: 103. In certain embodiments, the targeting sequence is SEQ ID NO: 104. In certain embodiments, the targeting sequence is SEQ ID NO: 105. In certain embodiments, the targeting sequence is SEQ ID NO: 106. In certain embodiments, the targeting sequence is SEQ ID NO: 107. In certain embodiments, the targeting sequence is SEQ ID NO: 108. In certain embodiments, the targeting sequence is SEQ ID NO: 109. In certain embodiments, the targeting sequence is SEQ ID NO: 110. In certain embodiments, the targeting sequence is SEQ ID NO: 111. In certain embodiments, the targeting sequence is SEQ ID NO: 112. In certain embodiments, the targeting sequence is SEQ ID NO: 113. In certain embodiments, the targeting sequence is SEQ ID NO: 114. In certain embodiments, the targeting sequence is SEQ ID NO: 115. In certain embodiments, the targeting sequence is SEQ ID NO: 116. In certain embodiments, the targeting sequence is SEQ ID NO: 117. In certain embodiments, the targeting sequence is SEQ ID NO:
118. In certain embodiments, the targeting sequence is SEQ ID NO: 119. In certain embodiments, the targeting sequence is SEQ ID NO: 120. In certain embodiments, the targeting sequence is SEQ ID NO: 121. In certain embodiments, the targeting sequence is SEQ ID NO: 122. In certain embodiments, the targeting sequence is SEQ ID NO: 123. In certain embodiments, the targeting sequence is SEQ ID NO: 124. In certain embodiments, the targeting sequence is SEQ ID NO: 125. In certain embodiments, the targeting sequence is SEQ ID NO: 126. In certain embodiments, the targeting sequence is SEQ ID NO: 127. In certain embodiments, the targeting sequence is SEQ ID NO: 128. In certain embodiments, the targeting sequence is SEQ ID NO: 129. In certain embodiments, the targeting sequence is SEQ ID NO: 130. In certain embodiments, the targeting sequence is SEQ ID NO: 131. In certain embodiments, the targeting sequence is SEQ ID NO: 132. In certain embodiments, the targeting sequence is SEQ ID NO: 133. In certain embodiments, the targeting sequence is SEQ ID NO: 134. In certain embodiments, the targeting sequence is SEQ ID NO: 135. In certain embodiments, the targeting sequence is SEQ ID NO: 136. In certain embodiments, the targeting sequence is SEQ ID NO: 137. In certain embodiments, the targeting sequence is SEQ ID NO: 138. In certain embodiments, the targeting sequence is SEQ ID NO: 139. In certain embodiments, the targeting sequence is SEQ ID NO: 140. In certain embodiments, the targeting sequence is SEQ ID NO: 141. In certain embodiments, the targeting sequence is SEQ ID NO: 142. In certain embodiments, the targeting sequence is SEQ ID NO: 143. In certain embodiments, the targeting sequence is SEQ ID NO: 144. In certain embodiments, the targeting sequence is SEQ ID NO: 145. In certain embodiments, the targeting sequence is SEQ ID NO: 146. In certain embodiments, the targeting sequence is SEQ ID NO: 147. In certain embodiments, the targeting sequence is SEQ ID NO: 148. In certain embodiments, the targeting sequence is SEQ ID NO: 149. In certain embodiments, the targeting sequence is SEQ ID NO: 150. In certain embodiments, the targeting sequence is SEQ ID NO: 151. In certain embodiments, the targeting sequence is SEQ ID NO: 152. In certain embodiments, the targeting sequence is SEQ ID NO: 153. In certain embodiments, the targeting sequence is SEQ ID NO: 154. In certain embodiments, the targeting sequence is SEQ ID NO: 155. In certain embodiments, the targeting sequence is SEQ ID NO: 156. In certain embodiments, the targeting sequence is SEQ ID NO: 157. In certain embodiments, the targeting sequence is SEQ ID NO: 158. In certain embodiments, the targeting sequence is SEQ ID NO: 159. In certain embodiments, the targeting sequence is SEQ ID NO: 160. In certain embodiments, the targeting sequence is SEQ ID NO: 161. In certain embodiments, the targeting sequence is SEQ ID NO: 162. In certain embodiments, the targeting sequence is SEQ ID NO: 163. In certain embodiments, the targeting sequence is SEQ ID NO: 164. In certain embodiments, the targeting
sequence is SEQ ID NO: 165. In certain embodiments, the targeting sequence is SEQ ID NO: 166. In certain embodiments, the targeting sequence is SEQ ID NO: 167. In certain embodiments, the targeting sequence is SEQ ID NO: 168. In certain embodiments, the targeting sequence is SEQ ID NO: 169. In certain embodiments, the targeting sequence is SEQ ID NO: 170. In certain embodiments, the targeting sequence is SEQ ID NO: 171. In certain embodiments, the targeting sequence is SEQ ID NO: 172. In certain embodiments, the targeting sequence is SEQ ID NO: 173. In certain embodiments, the targeting sequence is SEQ ID NO: 174. In certain embodiments, the targeting sequence is SEQ ID NO: 175. In certain embodiments, the targeting sequence is SEQ ID NO: 176. In certain embodiments, the targeting sequence is SEQ ID NO: 177. In certain embodiments, the targeting sequence is SEQ ID NO: 178. In certain embodiments, the targeting sequence is SEQ ID NO: 179. In certain embodiments, the targeting sequence is SEQ ID NO: 180. In certain embodiments, the targeting sequence is SEQ ID NO: 181. In certain embodiments, the targeting sequence is SEQ ID NO: 182. In certain embodiments, the targeting sequence is SEQ ID NO: 183. In certain embodiments, the targeting sequence is SEQ ID NO: 184. In certain embodiments, the targeting sequence is SEQ ID NO: 185. In certain embodiments, the targeting sequence is SEQ ID NO: 186. In certain embodiments, the targeting sequence is SEQ ID NO: 187. In certain embodiments, the targeting sequence is SEQ ID NO: 188. In certain embodiments, the targeting sequence is SEQ ID NO: 189. In certain embodiments, the targeting sequence is SEQ ID NO: 190. In certain embodiments, the targeting sequence is SEQ ID NO: 191. In certain embodiments, the targeting sequence is SEQ ID NO: 192. In certain embodiments, the targeting sequence is SEQ ID NO: 193. In certain embodiments, the targeting sequence is SEQ ID NO: 194. In certain embodiments, the targeting sequence is SEQ ID NO: 195. In certain embodiments, the targeting sequence is SEQ ID NO: 196. In certain embodiments, the targeting sequence is SEQ ID NO: 197. In certain embodiments, the targeting sequence is SEQ ID NO: 198. In certain embodiments, the targeting sequence is SEQ ID NO: 199. In certain embodiments, the targeting sequence is SEQ ID NO: 200. In certain embodiments, the targeting sequence is SEQ ID NO: 201. In certain embodiments, the targeting sequence is SEQ ID NO: 202. In certain embodiments, the targeting sequence is SEQ ID NO: 203. In certain embodiments, the targeting sequence is SEQ ID NO: 204. In certain embodiments, the targeting sequence is SEQ ID NO: 205. In certain embodiments, the targeting sequence is SEQ ID NO: 206. In certain embodiments, the targeting sequence is SEQ ID NO: 207. In certain embodiments, the targeting sequence is SEQ ID NO: 208. In certain embodiments, the targeting sequence is SEQ ID NO: 209. In certain embodiments, the targeting sequence is SEQ ID NO: 210. In certain embodiments, the targeting sequence is SEQ ID NO: 211. In certain
embodiments, the targeting sequence is SEQ ID NO: 212. In certain embodiments, the targeting sequence is SEQ ID NO: 213. In certain embodiments, the targeting sequence is SEQ ID NO: 214. In certain embodiments, the targeting sequence is SEQ ID NO: 215. In certain embodiments, the targeting sequence is SEQ ID NO: 216. In certain embodiments, the targeting sequence is SEQ ID NO: 217. In certain embodiments, the targeting sequence is SEQ ID NO: 218. In certain embodiments, the targeting sequence is SEQ ID NO: 219. In certain embodiments, the targeting sequence is SEQ ID NO: 220. In certain embodiments, the targeting sequence is SEQ ID NO: 221. In certain embodiments, the targeting sequence is SEQ ID NO: 222. In certain embodiments, the targeting sequence is SEQ ID NO: 223. In certain embodiments, the targeting sequence is SEQ ID NO: 224. In certain embodiments, the targeting sequence is SEQ ID NO: 225. In certain embodiments, the targeting sequence is SEQ ID NO: 226. In certain embodiments, the targeting sequence is SEQ ID NO: 227. In certain embodiments, the targeting sequence is SEQ ID NO: 228. In certain embodiments, the targeting sequence is SEQ ID NO: 229. In certain embodiments, the targeting sequence is SEQ ID NO: 230. In certain embodiments, the targeting sequence is SEQ ID NO: 231. In certain embodiments, the targeting sequence is SEQ ID NO: 232. In certain embodiments, the targeting sequence is SEQ ID NO: 233. In certain embodiments, the targeting sequence is SEQ ID NO: 234. In certain embodiments, the targeting sequence is SEQ ID NO: 235. In certain embodiments, the targeting sequence is SEQ ID NO: 236. In certain embodiments, the targeting sequence is SEQ ID NO: 237. In certain embodiments, the targeting sequence is SEQ ID NO: 238. In certain embodiments, the targeting sequence is SEQ ID NO: 239. In certain embodiments, the targeting sequence is SEQ ID NO: 240. In certain embodiments, the targeting sequence is SEQ ID NO: 241. In certain embodiments, the targeting sequence is SEQ ID NO: 242. In certain embodiments, the targeting sequence is SEQ ID NO: 243. In certain embodiments, the targeting sequence is SEQ ID NO: 244. In certain embodiments, the targeting sequence is SEQ ID NO: 245. In certain embodiments, the targeting sequence is SEQ ID NO: 246. In certain embodiments, the targeting sequence is SEQ ID NO: 247. In certain embodiments, the targeting sequence is SEQ ID NO: 248. In certain embodiments, the targeting sequence is SEQ ID NO: 249. In certain embodiments, the targeting sequence is SEQ ID NO: 250. In certain embodiments, the targeting sequence is SEQ ID NO: 251. In certain embodiments, the targeting sequence is SEQ ID NO: 252. In certain embodiments, the targeting sequence is SEQ ID NO: 253. In certain embodiments, the targeting sequence is SEQ ID NO: 254. In certain embodiments, the targeting sequence is SEQ ID NO: 255. In certain embodiments, the targeting sequence is SEQ ID NO: 256. In certain embodiments, the targeting sequence is SEQ ID NO: 257. In certain embodiments, the targeting sequence is SEQ ID NO:
258. In certain embodiments, the targeting sequence is SEQ ID NO: 259. In certain embodiments, the targeting sequence is SEQ ID NO: 260. In certain embodiments, the targeting sequence is SEQ ID NO: 261. In certain embodiments, the targeting sequence is SEQ ID NO: 262. In certain embodiments, the targeting sequence is SEQ ID NO: 263. In certain embodiments, the targeting sequence is SEQ ID NO: 264. In certain embodiments, the targeting sequence is SEQ ID NO: 265. In certain embodiments, the targeting sequence is SEQ ID NO: 266. In certain embodiments, the targeting sequence is SEQ ID NO: 267. In certain embodiments, the targeting sequence is SEQ ID NO: 268. In certain embodiments, the targeting sequence is SEQ ID NO: 269. In certain embodiments, the targeting sequence is SEQ ID NO: 270. In certain embodiments, the targeting sequence is SEQ ID NO: 271. In certain embodiments, the targeting sequence is SEQ ID NO: 272. In certain embodiments, the targeting sequence is SEQ ID NO: 273. In certain embodiments, the targeting sequence is SEQ ID NO: 274. In certain embodiments, the targeting sequence is SEQ ID NO: 275. In certain embodiments, the targeting sequence is SEQ ID NO: 276. In certain embodiments, the targeting sequence is SEQ ID NO: 277. In certain embodiments, the targeting sequence is SEQ ID NO: 278. In certain embodiments, the targeting sequence is SEQ ID NO: 279. In certain embodiments, the targeting sequence is SEQ ID NO: 280. In certain embodiments, the targeting sequence is SEQ ID NO: 281. In certain embodiments, the targeting sequence is SEQ ID NO: 282. In certain embodiments, the targeting sequence is SEQ ID NO: 283. In certain embodiments, the targeting sequence is SEQ ID NO: 284. In certain embodiments, the targeting sequence is SEQ ID NO: 285. In certain embodiments, the targeting sequence is SEQ ID NO: 286. In certain embodiments, the targeting sequence is SEQ ID NO: 287. In certain embodiments, the targeting sequence is SEQ ID NO: 288. In certain embodiments, the targeting sequence is SEQ ID NO: 289. In certain embodiments, the targeting sequence is SEQ ID NO: 290. In certain embodiments, the targeting sequence is SEQ ID NO: 291. In certain embodiments, the targeting sequence is SEQ ID NO: 292. In certain embodiments, the targeting sequence is SEQ ID NO: 293. In certain embodiments, the targeting sequence is SEQ ID NO: 294. In certain embodiments, the targeting sequence is SEQ ID NO: 295. In certain embodiments, the targeting sequence is SEQ ID NO: 296. In certain embodiments, the targeting sequence is SEQ ID NO: 297. In certain embodiments, the targeting sequence is SEQ ID NO: 298. In certain embodiments, the targeting sequence is SEQ ID NO: 299. In certain embodiments, the targeting sequence is SEQ ID NO: 300. In certain embodiments, the targeting sequence is SEQ ID NO: 301. In certain embodiments, the targeting sequence is SEQ ID NO: 302. In certain embodiments, the targeting sequence is SEQ ID NO: 303. In certain embodiments, the targeting sequence is SEQ ID NO: 304. In certain embodiments, the targeting
sequence is SEQ ID NO: 305. In certain embodiments, the targeting sequence is SEQ ID NO: 306. In certain embodiments, the targeting sequence is SEQ ID NO: 307. In certain embodiments, the targeting sequence is SEQ ID NO: 308. In certain embodiments, the targeting sequence is SEQ ID NO: 309. In certain embodiments, the targeting sequence is SEQ ID NO: 310. In certain embodiments, the targeting sequence is SEQ ID NO: 311. In certain embodiments, the targeting sequence is SEQ ID NO: 312. In certain embodiments, the targeting sequence is SEQ ID NO: 313. In certain embodiments, the targeting sequence is SEQ ID NO: 314. In certain embodiments, the targeting sequence is SEQ ID NO: 315. In certain embodiments, the targeting sequence is SEQ ID NO: 316. In certain embodiments, the targeting sequence is SEQ ID NO: 317. In certain embodiments, the targeting sequence is SEQ ID NO: 318. In certain embodiments, the targeting sequence is SEQ ID NO: 319. In certain embodiments, the targeting sequence is SEQ ID NO: 320. In certain embodiments, the targeting sequence is SEQ ID NO: 321. In certain embodiments, the targeting sequence is SEQ ID NO: 322. In certain embodiments, the targeting sequence is SEQ ID NO: 323. In certain embodiments, the targeting sequence is SEQ ID NO: 324. In certain embodiments, the targeting sequence is SEQ ID NO: 325. In certain embodiments, the targeting sequence is SEQ ID NO: 326. [0417] In certain embodiments, the target region is within the exon internal region of exon 2 (SEQ ID NO: 2), the intron 1/exon 2 junction, the exon 2/intron 2 junction, or the intron internal region of intron 2 (SEQ ID NO: 11) of the human DNM2 gene pre-mRNA. In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 21st nucleotide to the 47th nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 13. In certain embodiments, the target region is a region within DNM2 H2A(+21+47). [0418] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 23rd nucleotide to the 51st nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 14. In certain embodiments, the target region is a region within DNM2 H2A(+23+51). [0419] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 46th nucleotide to the 72nd nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 15. In certain embodiments, the target region is a region within DNM2 H2A(+46+72). [0420] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 13th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 24th nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target
region is a region within SEQ ID NO: 16. In certain embodiments, the target region is a region within DNM2 H2D(+13-24). [0421] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 13th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 14th nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 17. In certain embodiments, the target region is a region within DNM2 H2D(+13-14). [0422] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 19th nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 18. In certain embodiments, the target region is a region within DNM2 H2D(+10-19). [0423] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 18th nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 19. In certain embodiments, the target region is a region within DNM2 H2D(+9-18). [0424] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23rd nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 20. In certain embodiments, the target region is a region within DNM2 H2D(+6-23). [0425] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 4th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23rd nucleotide of intron 2 measured from the 3′ end of exon 2. In certain embodiments, the target region is a region within SEQ ID NO: 21. In certain embodiments, the target region is a region within DNM2 H2D(+4-23). [0426] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 30th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 22. In certain embodiments, the target region is a region within DNM2 H2D(-1-30). [0427] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 29th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2). In certain embodiments, the target region is a region within SEQ ID NO: 23. In certain embodiments, the target region is a region within DNM2 H2D(-1-29).
[0428] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12- 13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3- 27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30). [0429] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9- 16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23) , DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6- 30). [0430] In certain embodiments, the target region is DNM2 H2A(+21+45) or DNM2 H2A(+23+47). In certain embodiments, the target region is DNM2 H2A(+23+47) or DNM2 H2A(+27+51). In certain embodiments, the target region is DNM2 H2A(+46+70) or DNM2 H2A(+48+72). [0431] In certain embodiments, the target region is DNM2 H2D(+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), or DNM2 H2D(+1-24). In certain embodiments, the target region is DNM2 H2D (+13-12), DNM2 H2D(+12-13), or DNM2 H2D(+11-14). [0432] In certain embodiments, the target region is DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), or DNM2 H2D(+6-19). In certain embodiments, the target region is DNM2 H2D(+9-16), DNM2 H2D(+8-17), or DNM2 H2D(+7-18). In certain embodiments, the target region is DNM2 H2D(+8-17). [0433] In certain embodiments, the target region is DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23). In certain embodiments, the target region is DNM2 H2D(+4-21), DNM2 H2D(+3-22), or DNM2 H2D(+2-23). In certain embodiments, the target region is DNM2 H2D(+3-22). [0434] In certain embodiments, the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30). In certain embodiments, the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), or DNM2 H2D(-5-29).
[0435] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(-7+17). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H2A(+3+27). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+15+37). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+37+61). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H2D(+20-5). [0436] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGAGGCCGC); SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAA); SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC); SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT); SEQ ID NO: 90 (GAACCTCAGGCCGCCCCATTTTACC); SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT);
SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT); and SEQ ID NO: 96 (CCTGCTGAACCTCAGGCCGCCCCAT), wherein B is an abasic subunit or guanine. [0437] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT); SEQ ID NO: 90 (GAACCTCAGGCCGCCCCATTTTACC); SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT); and SEQ ID NO: 96 (CCTGCTGAACCTCAGGCCGCCCCAT). [0438] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 71 (AGAGGCCGCCGGGTGACGATTCCTG); and SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC). [0439] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 72 (TGAGAGGCCGCCGGGTGACGATTCC); and SEQ ID NO: 73 (AGAATGAGAGGCCGCCGGGTGACGA). [0440] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 75 (TTTTGAGAAGATGAGCTGCAGAATG); and SEQ ID NO: 76 (GTTTTTGAGAAGATGAGCTGCAGAA). [0441] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA);
SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG); SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT); and SEQ ID NO: 90 (GAACCTCAGGCCGCCCCATTTTACC). [0442] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 78 (GCCCCATTTTACCTGTTTTTGAGAA); SEQ ID NO: 79 (CGCCCCATTTTACCTGTTTTTGAGA); and SEQ ID NO: 80 (CCGCCCCATTTTACCTGTTTTTGAG). [0443] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 81 (GCCGCCCCATTTTACCTGTTTTTGA); SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT); and SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT). [0444] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 82 (GGCCGCCCCATTTTACCTGTTTTTG); SEQ ID NO: 83 (AGGCCGCCCCATTTTACCTGTTTTT); and SEQ ID NO: 84 (CAGGCCGCCCCATTTTACCTGTTTT). [0445] In certain embodiments, the targeting sequence is SEQ ID NO: 83. [0446] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 85 (TCAGGCCGCCCCATTTTACCTGTTT); SEQ ID NO: 86 (CTCAGGCCGCCCCATTTTACCTGTT); SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT); SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); and SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT). [0447] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 87 (CCTCAGGCCGCCCCATTTTACCTGT);
SEQ ID NO: 88 (ACCTCAGGCCGCCCCATTTTACCTG); and SEQ ID NO: 89 (AACCTCAGGCCGCCCCATTTTACCT). [0448] In certain embodiments, the targeting sequence is SEQ ID NO: 88. [0449] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT); and SEQ ID NO: 96 (CCTGCTGAACCTCAGGCCGCCCCAT). [0450] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 91 (TGAACCTCAGGCCGCCCCATTTTAC); SEQ ID NO: 92 (CTGAACCTCAGGCCGCCCCATTTTA); SEQ ID NO: 93 (GCTGAACCTCAGGCCGCCCCATTTT); SEQ ID NO: 94 (TGCTGAACCTCAGGCCGCCCCATTT); and SEQ ID NO: 95 (CTGCTGAACCTCAGGCCGCCCCATT). [0451] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 67 (CGGGGAAGGAAGTCCCTGTGAGGGG); SEQ ID NO: 68 (CGCGGBGAAGGAAGTCCCTGTGAG); SEQ ID NO: 69 (ATTCCTGAACCGCGBGGAAGGAAGT); SEQ ID NO: 70 (CCGGGTGACGATTCCTGAACCGC); SEQ ID NO: 74 (GATGAGCTGCAGAATGAGAGGCCGC); and SEQ ID NO: 77 (TTTACCTGTTTTTGAGAAGATGAGC). [0452] In certain embodiments, the target region is within the exon internal region of exon 9 (SEQ ID NO: 6), the intron 8/exon 9 junction, or the exon 9/intron 9 junction of the human DNM2 gene pre-mRNA. [0453] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 36th nucleotide to the 61st nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6). In certain embodiments, the target region is a region within SEQ ID NO: 24. In certain embodiments, the target region is a region within DNM2 H9A(+36+61). [0454] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 42nd nucleotide to the 68th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6). In certain embodiments, the target region is a region within SEQ ID NO: 25. In certain embodiments, the target region is a region within DNM2 H9A(+42+70).
[0455] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 43rd nucleotide to the 68th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6). In certain embodiments, the target region is a region within SEQ ID NO: 26. In certain embodiments, the target region is a region within DNM2 H9A(+42+68). [0456] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 24th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 27. In certain embodiments, the target region is a region within DNM2 H9D(+24-15). [0457] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 24th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 5th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 28. In certain embodiments, the target region is a region within DNM2 H9D(+24-5). [0458] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 19th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 10th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 29. In certain embodiments, the target region is a region within DNM2 H9D(+19-10). [0459] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 14th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 30. In certain embodiments, the target region is a region within DNM2 H9D(+14-15). [0460] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22nd nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 31. In certain embodiments, the target region is a region within DNM2 H9D(+9-22). [0461] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 20th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 32. In certain embodiments, the target region is a region within DNM2 H9D(+9-20).
[0462] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 18th nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 33. In certain embodiments, the target region is a region within DNM2 H9D(+9-18). [0463] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 8th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 21st nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 34. In certain embodiments, the target region is a region within DNM2 H9D(+8-21). [0464] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22nd nucleotide of intron 9 measured from the 3′ end of exon 9. In certain embodiments, the target region is a region within SEQ ID NO: 35. In certain embodiments, the target region is a region within DNM2 H9D(+6-22). [0465] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13), DNM2 H9A(-5+20), DNM2 H9A(+1+25),DNM2 H9A(+6+30), DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), DNM2 H9A(+48+67), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11- 14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4- 17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22). [0466] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+43+67), DNM2 H9A(+42+66), DNM2 H9A(+44+68), DNM2 H9D(+24-1), DNM2 H9D(+23- 2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), DNM2 H9D(+10-15), DNM2
H9D(+9-16), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+6-19), DNM2 H9D(+5-20), DNM2 H9D(+4-21), or DNM2 H9D(+3-22). [0467] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+36+60) or DNM2 H9A(+37+61). [0468] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), or DNM2 H9A(+48+67). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), or DNM2 H9A(+44+68). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+67). [0469] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11- 14), or DNM2 H9D(+10-15). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), or DNM2 H9D(+20-5). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), or DNM2 H9D(+15-10). [0470] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), or DNM2 H9D(+10-15). [0471] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6- 17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22). [0472] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6- 17), DNM2 H9D(+5-17), or DNM2 H9D(+5-20).
[0473] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), or DNM2 H9D(+7-18). [0474] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), or DNM2 H9D(+4-21). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17). [0475] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22). [0476] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-5+20). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H9A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+6+30). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+25+49). [0477] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 158 (CGTCAAACTCCATCTGAGAAGAGAC); SEQ ID NO: 159 (TCCTTCTCGTCAAACTCCATCTGAG); SEQ ID NO: 160 (GTAAGTCCTTCTCGTCAAACTCCAT); SEQ ID NO: 161 (CCGTCGTAAGTCCTTCTCGTCAAAC); SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG); SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT); SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG);
SEQ ID NO: 175 (TGACTCCATGGATGTTCTTAATG); SEQ ID NO: 176 (TGACTCCATGGATGTTCTTAAT); SEQ ID NO: 177 (TGACTCCATGGATGTTCTTAA); SEQ ID NO: 178 (TGACTCCATGGATGTTCTTA); SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT); SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG); SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT); SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 208 (CTCCTCGTGGAACTTGCCTG); and SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG).
[0478] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG); SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT); SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT); SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG); SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT); SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); and SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG). [0479] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 164 (ATGGATGTTCTTAATGGCATAGCTG); and SEQ ID NO: 165 (CATGGATGTTCTTAATGGCATAGCT). [0480] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 166 (GACTCCATGGATGTTCTTAATGGCA); SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC);
SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG); SEQ ID NO: 175 (TGACTCCATGGATGTTCTTAATG); SEQ ID NO: 176 (TGACTCCATGGATGTTCTTAAT); SEQ ID NO: 177 (TGACTCCATGGATGTTCTTAA); and SEQ ID NO: 178 (TGACTCCATGGATGTTCTTA). [0481] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 167 (CCATGGATGTTCTTAATGGC); SEQ ID NO: 168 TCCATGGATGTTCTTAATGGC); SEQ ID NO: 169 (CTCCATGGATGTTCTTAATGGC); SEQ ID NO: 170 (ACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 171 (GACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 172 (TGACTCCATGGATGTTCTTAATGGC); SEQ ID NO: 173 (TGACTCCATGGATGTTCTTAATGG); and SEQ ID NO: 174 (CTGACTCCATGGATGTTCTTAATGG). [0482] In certain embodiments, the targeting sequence is SEQ ID NO: 172. [0483] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT); SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG); SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 TCGTGGAACTTGCCTGACTCCATGG);
SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); and SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT). [0484] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 179 (CCTGACTCCATGGATGTTCTTAATG); SEQ ID NO: 180 (GCCTGACTCCATGGATGTTCTTAAT); SEQ ID NO: 181 (TGCCTGACTCCATGGATGTTCTTAA); SEQ ID NO: 182 (TTGCCTGACTCCATGGATGTTCTTA); and SEQ ID NO: 183 (CTTGCCTGACTCCATGGATGTTCTT). [0485] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 184 (ACTTGCCTGACTCCATGGATGTTCT); SEQ ID NO: 185 (AACTTGCCTGACTCCATGGATGTTC); SEQ ID NO: 186 (GAACTTGCCTGACTCCATGGATGTT); SEQ ID NO: 187 (GGAACTTGCCTGACTCCATGGATGT); and SEQ ID NO: 188 TGGAACTTGCCTGACTCCATGGATG). [0486] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 189 (GTGGAACTTGCCTGACTCCATGGAT); SEQ ID NO: 190 (CGTGGAACTTGCCTGACTCCATGGA); SEQ ID NO: 191 TCGTGGAACTTGCCTGACTCCATGG); SEQ ID NO: 192 (CTCGTGGAACTTGCCTGACTCCATG); and SEQ ID NO: 193 (CCTCGTGGAACTTGCCTGACTCCAT). [0487] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA);
SEQ ID NO: 208 (CTCCTCGTGGAACTTGCCTG); and SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG). [0488] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); and SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC). [0489] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 194 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 195 (TCCTCGTGGAACTTGCCTGACTCCA); SEQ ID NO: 196 (TCGTGGAACTTGCCTGACTCC); SEQ ID NO: 197 (CTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 198 (CCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); and SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC). [0490] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 199 (CTCCTCGTGGAACTTGCCTGACTCC); SEQ ID NO: 200 (TCTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 201 (CTCCTCGTGGAACTTGCCTGACTC); SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); and SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA). [0491] In certain embodiments, the targeting sequence is SEQ ID NO: 199. [0492] In certain embodiments, the targeting sequence is selected from:
SEQ ID NO: 202 (CTCTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 203 (CTCCTCGTGGAACTTGCCTGACT); SEQ ID NO: 204 (CTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 205 (ACTCTCCTCGTGGAACTTGCCTGAC); SEQ ID NO: 206 (CTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 207 (GACTCTCCTCGTGGAACTTGCCTGA); SEQ ID NO: 208 (CTCCTCGTGGAACTTGCCTG); and SEQ ID NO: 209 (TGACTCTCCTCGTGGAACTTGCCTG). [0493] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 158 (CGTCAAACTCCATCTGAGAAGAGAC); SEQ ID NO: 159 (TCCTTCTCGTCAAACTCCATCTGAG); SEQ ID NO: 160 (GTAAGTCCTTCTCGTCAAACTCCAT); SEQ ID NO: 161 (CCGTCGTAAGTCCTTCTCGTCAAAC); SEQ ID NO: 162 (AGCTGATCTCCCGTCGTAAGTCCTT); and SEQ ID NO: 163 (TAATGGCATAGCTGATCTCCCGTCG). [0494] In certain embodiments, the target region is within the exon internal region of exon 12 (SEQ ID NO: 7), the intron 11/exon 12 junction, or the exon 12/intron 12 junction of the human DNM2 gene pre-mRNA. [0495] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 4th nucleotide to the 31st nucleotide measured of exon 12 from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 36. In certain embodiments, the target region is a region within DNM2 H12A(+4+31). [0496] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 44th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 37. In certain embodiments, the target region is a region within DNM2 H12A(+10+44). [0497] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 38th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 38. In certain embodiments, the target region is a region within DNM2 H12A(+10+38). [0498] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 16th nucleotide to the 44th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 39. In certain embodiments, the target region is a region within DNM2 H12A(+16+44).
[0499] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 14th nucleotide to the 42nd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 40. In certain embodiments, the target region is a region within DNM2 H12A(+14+42). [0500] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 25th nucleotide to the 54th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 41. In certain embodiments, the target region is a region within DNM2 H12A(+25+54). [0501] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 25th nucleotide to the 53rd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 42. In certain embodiments, the target region is a region within DNM2 H12A(+25+53). [0502] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 26th nucleotide to the 54th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 43. In certain embodiments, the target region is a region within DNM2 H12A(+26+54). [0503] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 36th nucleotide to the 61st nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). In certain embodiments, the target region is a region within SEQ ID NO: 44. In certain embodiments, the target region is a region within DNM2 H12A(+36+61). [0504] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 22nd nucleotide of exon 12 measured from the 3′ end of exon 12 (SEQ ID NO: 7) to the 5th nucleotide of intron 12 measured from the 3′ end of exon 12. In certain embodiments, the target region is a region within SEQ ID NO: 45. In certain embodiments, the target region is a region within DNM2 H12D(+22-5). [0505] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-13+12), DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), or DNM2 H12D(+15-10).
[0506] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+49), DNM2 H12A(+29+53), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), or DNM2 H12D(+15-10). [0507] In certain embodiments, the target region is DNM2 H12A(+4+28) or DNM2 H12A(+7+31). [0508] In certain embodiments, the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44). [0509] In certain embodiments, the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), or DNM2 H12A(+14+38). [0510] In certain embodiments, the target region is DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), or DNM2 H12A(+20+44). [0511] In certain embodiments, the target region is DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), or DNM2 H12A(+18+42). [0512] In certain embodiments, the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54). [0513] In certain embodiments, the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), or DNM2 H12A(+29+53). [0514] In certain embodiments, the target region is DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54). In certain embodiments, the target region is DNM2 H12A(+25+49). In certain embodiments, the target region is DNM2 H12A(+36+60) or DNM2 H12A(+37+61). [0515] In certain embodiments, the target region is DNM2 H12D(+22-3) or DNM2 H12D(+20-5). In certain embodiments, the target region is DNM2 H12A(-13+12). In certain embodiments, the
target region is DNM2 H12A(-3+22). In certain embodiments, the target region is DNM2 H12A(+43+67). In certain embodiments, the target region is DNM2 H12D(+15-10). [0516] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 210 (CAGCAGAAGAATCTGGBGGAGAGCG); SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG); SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA); SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG); SEQ ID NO: 238 (GATGAAGTCCTCATGGTTCGTGTTG); SEQ ID NO: 239 (CGATGAAGTCCTCATGGTTCGTGTT); SEQ ID NO: 240 (CAAACCCGATGAAGTCCTCATGGTT); SEQ ID NO: 241 (TACTTGGCAAACCCGATGAAGTCCT);
SEQ ID NO: 242 (CCTACTTGGCAAACCCGATGAAGTC); and SEQ ID NO: 243 (AAGTACCTACTTGGCAAACCCGATG). [0517] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG); SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG); SEQ ID NO: 238 (GATGAAGTCCTCATGGTTCGTGTTG); SEQ ID NO: 240 (CAAACCCGATGAAGTCCTCATGGTT); SEQ ID NO: 241 (TACTTGGCAAACCCGATGAAGTCCT); SEQ ID NO: 242 (CCTACTTGGCAAACCCGATGAAGTC); and SEQ ID NO: 243 (AAGTACCTACTTGGCAAACCCGATG). [0518] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 212 (ACTGCTCAATGTCGATCAGCAGAAG); and SEQ ID NO: 213 (AGGACTGCTCAATGTCGATCAGCAG). [0519] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA);
SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); and SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA). [0520] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 214 (TGTAGGACTGCTCAATGTCGATCAG); SEQ ID NO: 215 (ATGTAGGACTGCTCAATGTCGATCA); SEQ ID NO: 216 (GATGTAGGACTGCTCAATGTCGATC); SEQ ID NO: 217 (TGATGTAGGACTGCTCAATGTCGAT); and SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA). [0521] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG); SEQ ID NO: 223 (TCGTGTTGATGTAGGACTGCTCAAT); and SEQ ID NO: 224 (TTCGTGTTGATGTAGGACTGCTCAA). [0522] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 218 (TTGATGTAGGACTGCTCAATGTCGA); SEQ ID NO: 219 (GTTGATGTAGGACTGCTCAATGTCG); SEQ ID NO: 220 (TGTTGATGTAGGACTGCTCAATGTC); SEQ ID NO: 221 (GTGTTGATGTAGGACTGCTCAATGT); and SEQ ID NO: 222 (CGTGTTGATGTAGGACTGCTCAATG). [0523] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG);
SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); and SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG). [0524] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 225 (TTCGTGTTGATGTAGGACTG); SEQ ID NO: 226 (GTTCGTGTTGATGTAGGACTG); SEQ ID NO: 227 (GGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 228 (TGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 229 (ATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 230 (CATGGTTCGTGTTGATGTAGGACTG); SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); and SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG). [0525] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 231 (CATGGTTCGTGTTGATGTAGGACT); SEQ ID NO: 232 (CATGGTTCGTGTTGATGTAGGAC); SEQ ID NO: 233 (CATGGTTCGTGTTGATGTAGGA); SEQ ID NO: 234 (CATGGTTCGTGTTGATGTAGG); SEQ ID NO: 235 (TCCTCATGGTTCGTGTTGATGTAGG); SEQ ID NO: 236 (CATGGTTCGTGTTGATGTAG); and SEQ ID NO: 237 (GTCCTCATGGTTCGTGTTGATGTAG). [0526] In certain embodiments, the targeting sequence is SEQ ID NO: 230 [0527] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 238 (GATGAAGTCCTCATGGTTCGTGTTG); and SEQ ID NO: 239 (CGATGAAGTCCTCATGGTTCGTGTT). [0528] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 241 (TACTTGGCAAACCCGATGAAGTCCT); and SEQ ID NO: 242 (CCTACTTGGCAAACCCGATGAAGTC). [0529] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 210 (CAGCAGAAGAATCTGGBGGAGAGCG); SEQ ID NO: 211 (CAATGTCGATCAGCAGAAGAATCTG); SEQ ID NO: 240 (CAAACCCGATGAAGTCCTCATGGTT); and SEQ ID NO: 243 (AAGTACCTACTTGGCAAACCCGATG).
[0530] In certain embodiments, the target region is within the exon internal region of exon 13 (SEQ ID NO: 8), the intron 12/exon 13 junction, the exon 13/intron 13 junction, or the intron internal region of intron 12 (SEQ ID NO: 12). [0531] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 27th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 46. In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within DNM2 H13A(-27+12). [0532] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 27th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 2nd nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 47. In certain embodiments, the target region is a region within DNM2 H13A(-27+2). [0533] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 22nd nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 7th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 48. In certain embodiments, the target region is a region within DNM2 H13A(-22+7). [0534] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 17th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 49. In certain embodiments, the target region is a region within DNM2 H13A(-17+12). [0535] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 11th nucleotide of intron 12 (SEQ ID NO: 12) measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 50. In certain embodiments, the target region is a region within DNM2 H13A(-11+24). [0536] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 11th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 18th nucleotide of exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 51. In certain embodiments, the target region is a region within DNM2 H13A(-11+18). [0537] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 5th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24th
nucleotide exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 52. In certain embodiments, the target region is a region within DNM2 H13A(-5+24). [0538] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 8th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 21st nucleotide exon 13 measured from the 5′ end of exon 13. In certain embodiments, the target region is a region within SEQ ID NO: 53. In certain embodiments, the target region is a region within DNM2 H13A(-8+21). [0539] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 54. In certain embodiments, the target region is a region within DNM2 H13A(+1+30). [0540] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 28th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 55. In certain embodiments, the target region is a region within DNM2 H13A(+1+28). [0541] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 56. In certain embodiments, the target region is a region within DNM2 H13A(+6+30). [0542] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 3rd nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). In certain embodiments, the target region is a region within SEQ ID NO: 57. In certain embodiments, the target region is a region within DNM2 H13A(+3+30). [0543] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(- 5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2
H13A(+7+28), DNM2 H13A(+8+28), DNM2 H13A(+9+28), DNM2 H13A(+16+40), DNM2 H13D(+22-3), or DNM2 H13D(+7-18). [0544] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(- 5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), or DNM2 H13A(+4+28). [0545] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12). [0546] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), or DNM2 H13A(- 23+2). [0547] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), or DNM2 H13A(- 18+7). [0548] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12). [0549] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(- 7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(-1+24). [0550] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), or DNM2 H13A(-7+18). [0551] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(- 1+24).
[0552] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), or DNM2 H13A(- 4+21). [0553] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28). [0554] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), or DNM2 H13A(+5+28). [0555] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28). [0556] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), or DNM2 H13A(+6+30). [0557] In certain embodiments, the target region of the human DNM2 gene pre-mRNA DNM2 H13A(+4+28). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+16+40). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H13D(+22-3). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H13D(+7-18). [0558] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA);
SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA); SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC); SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 282 (GTTCAGCTGCGTGCTCCTCTGC); SEQ ID NO: 283 (GTTCAGCTGCGTGCTCCTCTG); SEQ ID NO: 284 (GTTCAGCTGCGTGCTCCTCT); SEQ ID NO: 285 (GGCTCTCTTCTTGTTCAGCTGCGTG); SEQ ID NO: 286 (TACCTGATTGGBGATGGCTCTCTTC); and SEQ ID NO: 287 (TGCAGAGGBGTGTGCTACCTGATTG). [0559] In certain embodiments, the targeting sequence is selected from:
SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA); SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC); SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); and SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG).
[0560] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG); SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA); SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); and SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA). [0561] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 244 (GAAACCAAAGAACAGACAGGGTTCA); SEQ ID NO: 245 (TGAAACCAAAGAACAGACAGGGTTC); SEQ ID NO: 246 (CTGAAACCAAAGAACAGACAGGGTT); SEQ ID NO: 247 (ACTGAAACCAAAGAACAGACAGGGT); and SEQ ID NO: 248 (CACTGAAACCAAAGAACAGACAGGG). [0562] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 249 (GCACTGAAACCAAAGAACAGACAGG); SEQ ID NO: 250 (GGCACTGAAACCAAAGAACAGACAG); SEQ ID NO: 251 (GGGCACTGAAACCAAAGAACAGACA); SEQ ID NO: 252 (TGGGCACTGAAACCAAAGAACAGAC); and SEQ ID NO: 253 (CTGGGCACTGAAACCAAAGAACAGA). [0563] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 254 (GCTGGGCACTGAAACCAAAGAACAG); SEQ ID NO: 255 (TGCTGGGCACTGAAACCAAAGAACA); SEQ ID NO: 256 (CTGCTGGGCACTGAAACCAAAGAAC); SEQ ID NO: 257 (TCTGCTGGGCACTGAAACCAAAGAA); and SEQ ID NO: 258 (CTCTGCTGGGCACTGAAACCAAAGA). [0564] In certain embodiments, the targeting sequence is selected from:
SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); and SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC). [0565] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 259 (TCCTCTGCTGGGCACTGAAACCAAA); SEQ ID NO: 260 (CTCCTCTGCTGGGCACTGAAACCAA); SEQ ID NO: 261 (GCTCCTCTGCTGGGCACTGAAACCA); SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); and SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC). [0566] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA); SEQ ID NO: 266 (TGCGTGCTCCTCTGCTGGGCACTGA); SEQ ID NO: 267 (CTGCGTGCTCCTCTGCTGGGCACTG); SEQ ID NO: 268 (GCTGCGTGCTCCTCTGCTGGGCACT); and SEQ ID NO: 269 (AGCTGCGTGCTCCTCTGCTGGGCAC). [0567] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 262 (TGCTCCTCTGCTGGGCACTGAAACC); SEQ ID NO: 263 (GTGCTCCTCTGCTGGGCACTGAAAC); SEQ ID NO: 264 (CGTGCTCCTCTGCTGGGCACTGAAA); and SEQ ID NO: 265 (GCGTGCTCCTCTGCTGGGCACTGAA). [0568] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG);
SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG); SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 282 (GTTCAGCTGCGTGCTCCTCTGC); SEQ ID NO: 283 (GTTCAGCTGCGTGCTCCTCTG); and SEQ ID NO: 284 (GTTCAGCTGCGTGCTCCTCT). [0569] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 270 (CAGCTGCGTGCTCCTCTGCTGGGCA); SEQ ID NO: 271 (TCAGCTGCGTGCTCCTCTGCTGGGC); SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); and SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG). [0570] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT); SEQ ID NO: 282 (GTTCAGCTGCGTGCTCCTCTGC); SEQ ID NO: 283 (GTTCAGCTGCGTGCTCCTCTG); and SEQ ID NO: 284 (GTTCAGCTGCGTGCTCCTCT). [0571] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 272 (TTCAGCTGCGTGCTCCTCTGCTGGG); SEQ ID NO: 273 (GCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 274 (AGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 275 (CAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 276 (TCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 277 (TTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 279 (GTTCAGCTGCGTGCTCCTCTGCTG);
SEQ ID NO: 280 (GTTCAGCTGCGTGCTCCTCTGCT); and SEQ ID NO: 281 (TTGTTCAGCTGCGTGCTCCTCTGCT). [0572] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 278 (GTTCAGCTGCGTGCTCCTCTGCTGG); SEQ ID NO: 285 (GGCTCTCTTCTTGTTCAGCTGCGTG); SEQ ID NO: 286 (TACCTGATTGGBGATGGCTCTCTTC); and SEQ ID NO: 287 (TGCAGAGGBGTGTGCTACCTGATTG). [0573] In certain embodiments, the target region is within the exon internal region of exon 6 (SEQ ID NO: 3), the intron 5/exon 6 junction, the exon 6/intron 6 junction, or the intron internal region of intron 6 (SEQ ID NO: 11). [0574] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 37th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 58. In certain embodiments, the target region is a region within DNM2 H6A(+10+37). [0575] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 32nd nucleotide to the 59th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 59. In certain embodiments, the target region is a region within DNM2 H6A(+32+59). [0576] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 60th nucleotide to the 87th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 60. In certain embodiments, the target region is a region within DNM2 H6A(+60+87). [0577] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 130th nucleotide to the 156th nucleotide measured from the 5′ end of exon 6 (SEQ ID NO: 3). In certain embodiments, the target region is a region within SEQ ID NO: 61. In certain embodiments, the target region is a region within DNM2 H6A(+130+156). [0578] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 17th nucleotide of exon 6 measured from the 3′ end of exon 6 (SEQ ID NO: 3) to the 10th nucleotide of intron 6 measured from the 3′ end of exon 6. In certain embodiments, the target region is a region within SEQ ID NO: 62. In certain embodiments, the target region is a region within DNM2 H6D(+17-10). [0579] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5), DNM2 H6A(-6+19), DNM2 H6A(-2+23), DNM2 H6A(+3+26), DNM2 H6A(+10+3), DNM2 H6A(+13+37), DNM2 H6A(+23+47), DNM2 H6A(+32+56), DNM2 H6A(+35+59), DNM2 H6A(+44+68), DNM2 H6A(+54+78), DNM2 H6A(+60+84), DNM2 H6A(+63+87), DNM2
H6A(+88+112), DNM2 H6A(+96+119), DNM2 H6A(+105+129), DNM2 H6A(+121+145), DNM2 H6A(+130+154), DNM2 H6A(+132+156), DNM2 H6D (+6-19), DNM2 H6D (+15-10), or DNM2 H6D (+17-8). [0580] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+10+3) or DNM2 H6A(+13+37). [0581] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+32+56) or DNM2 H6A(+35+59). [0582] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+60+84) or DNM2 H6A(+63+87). [0583] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+130+154) or DNM2 H6A(+132+156). [0584] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+15-10) or DNM2 H6D (+17-8). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-6+19). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-2+23). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+3+26). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+23+47). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+44+68). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+54+78). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+88+112). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H6A(+96+119). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+105+129). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+121+145). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+6-19). [0585] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 97 (GTAGCCTGTGGGGAGAGGCAGTCAG); SEQ ID NO: 98 (TTCACCACGCCAATGTAGCCTGTGG); SEQ ID NO: 99 (GCGGTTCACCACGCCAATGTAGCCT); SEQ ID NO: 100 (GCTGCGGTTCACCACGCCAATGTA); SEQ ID NO: 101 (TCCTTCTGGCTGCGGTTCACCACGC); SEQ ID NO: 102 (ATATCCTTCTGGCTGCGGTTCACCA); SEQ ID NO: 103 (CTTGCCCTCAATATCCTTCTGGCTG); SEQ ID NO: 104 (GATGTCCTTCTTGCCCTCAATATCC);
SEQ ID NO: 105 (ACGGATGTCCTTCTTGCCCTCAATA); SEQ ID NO: 106 (CAGTGCTGCACGGATGTCCTTCTTG); SEQ ID NO: 107 (TCTCAGCTGCCAGTGCTGCACGGAT); SEQ ID NO: 108 (ACTTCCTCTCAGCTGCCAGTGCTGC); SEQ ID NO: 109 (AGAACTTCCTCTCAGCTGCCAGTGC); SEQ ID NO: 110 (TGCCGGTAGGCCGGGTGGGAGAGGA); SEQ ID NO: 111 (GGCCATGTGCCGGTAGGCCGGGTG); SEQ ID NO: 112 (CCATGCGGTCGGCCATGTGCCGGTA); SEQ ID NO: 113 (TGCAGATGTGGCGTGCCCATGCGGT); SEQ ID NO: 114 (AGCGTCTTCTGCAGATGTGGCGTGC); SEQ ID NO: 115 (TCAGCGTCTTCTGCAGATGTGGCGT); SEQ ID NO: 116 (GTGCAAACCCTTGCAGTACCTGATT); SEQ ID NO: 117 (CTTGCAGTACCTGATTCAGCGTCTT); and SEQ ID NO: 118 (TGCAGTACCTGATTCAGCGTCTTCT). [0586] In certain embodiments, the target region is within the exon internal region of exon 7 (SEQ ID NO: 4), the intron 6/exon 7 junction, or the exon 7/intron 7 junction. [0587] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 72nd nucleotide to the 98th nucleotide of exon 7 measured from the 5′ end of exon 7 (SEQ ID NO: 4). In certain embodiments, the target region is a region within SEQ ID NO: 63. In certain embodiments, the target region is a region within DNM2 H7A(+72+98). [0588] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 7 measured from the 3′ end of exon 7 (SEQ ID NO: 4) to the 16th nucleotide of intron 7 measured from the 3′ end of exon 7. In certain embodiments, the target region is a region within SEQ ID NO: 64. In certain embodiments, the target region is a region within DNM2 H7D(+9-16). [0589] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+1+25), DNM2 H7A(+6+30), DNM2 H7A(+11+35), DNM2 H7A(+22+46), DNM2 H7A(+29+53), DNM2 H7A(+41+65), DNM2 H7A(+51+75), DNM2 H7A(+57+81), DNM2 H7A(+62+86), DNM2 H7A(+72+96), DNM2 H7A(+74+98), DNM2 H7A(+81+105), DNM2 H7A(+91+115), or DNM2 H7A(+110+134). [0590] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+72+96) or DNM2 H7A(+74+98). [0591] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+8-16) or DNM2 H7D(+9-16).
[0592] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-15+10). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-5+20). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H7A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+6+30). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+11+35). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+22+46). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+29+53). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+41+65). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+51+75). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+57+81). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+62+86). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H7A(+81+105). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+91+115). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+110+134). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+21-4). [0593] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 119 (TGGTCAGTTGCTGCAGGAGAGAGGT); SEQ ID NO: 120 (CGGATGTGGTTGGTCAGTTGCTGCA); SEQ ID NO: 121 (ACTCCCGGATGTGGTTGGTCAGTTG); SEQ ID NO: 122 (CAGCGACTCCCGGATGTGGTTGGTC); SEQ ID NO: 123 (GCCGGCAGCGACTCCCGGATGTGGT); SEQ ID NO: 124 (TGCTACGTAGGGCCGGCAGCGACTC); SEQ ID NO: 125 (TGTAGTTTGCTACGTAGGGCCGGCA); SEQ ID NO: 126 (AGCAGCTGGCTCTGTAGTTTGCTAC); SEQ ID NO: 127 (CTCCAGGGACAGCAGCTGGCTCTGT); SEQ ID NO: 128 (CTCCTTCTCCAGGGACAGCAGCTGG); SEQ ID NO: 129 (TCCACCTCCTTCTCCAGGGACAGCA); SEQ ID NO: 130 (CTTGTACTCCTCCACCTCCTTCTCC); SEQ ID NO: 131 (TTCTTGTACTCCTCCACCTCCTTCT); SEQ ID NO: 132 (CCGAAAGTTCTTGTACTCCTCCACC); SEQ ID NO: 133 (GGTCGTCGGGCCGAAAGTTCTTGTA); SEQ ID NO: 134 (GCTTTGGTTTTGCGGGTGGGGTCGT); SEQ ID NO: 135 (GTGCCGBGGTACATACTGCAGCAG);
SEQ ID NO: 136 (GTGCCGGGGTACATACTGCAGCAGG); and SEQ ID NO: 137 (ATACTGCAGCAGGGCTTTGGTTTTG). [0594] In certain embodiments, the target region is within the exon internal region of exon 8 (SEQ ID NO: 5), the intron 7/exon 8 junction, or the exon 8/intron 8 junction. [0595] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 7th nucleotide of intron 7 measured from the 5′ end of exon 8 (SEQ ID NO: 5) to the 19th nucleotide of exon 8 measured from the 5′ end of exon 8. In certain embodiments, the target region is a region within SEQ ID NO: 65. In certain embodiments, the target region is a region within DNM2 H8A(-7+19). [0596] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is within the 18th nucleotide to the 45th nucleotide of exon 8 measured from the 5′ end of exon 8 (SEQ ID NO: 5). In certain embodiments, the target region is a region within SEQ ID NO: 66. In certain embodiments, the target region is a region within DNM2 H8A(+18+45). [0597] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-20+5), DNM2 H8A(-14+11), DNM2 H8A(-7+18), DNM2 H8A(-6+19), DNM2 H8A(-2+23), DNM2 H8A(+5+29), DNM2 H8A(+18+42), DNM2 H8A(+21+45), DNM2 H8A(+27+51), DNM2 H8A(+36+60), DNM2 H8A(+46+70), DNM2 H8A(+51+75), DNM2 H8A(+56+80), DNM2 H8A(+65+88), DNM2 H8A(+76+100), DNM2 H8A(+83+107), DNM2 H8A(+96+119), DNM2 H8A(+105+129), DNM2 H8D(+18-7), or DNM2 H8D(+10-15). [0598] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-7+18) or DNM2 H8A(-6+19). [0599] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+18+42) or DNM2 H8A(+21+45). [0600] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-20+5). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-14+11). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H8A(-2+23). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+5+29). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+27+51). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+36+60). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+46+70). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+51+75). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+56+80). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+65+88). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is
DNM2 H8A(+76+100). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H8A(+83+107). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+96+119). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+105+129). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+18-7). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+10-15). [0601] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 138 (CCATCCTATGAGGAAAAGGAAAAAG); SEQ ID NO: 139 (GCTGGACCATCCTATGAGGAAAAGG); SEQ ID NO: 140 (CCAAACTGCTGGACCATCCTATGAG); SEQ ID NO: 141 (CCCAAACTGCTGGACCATCCTATGA); SEQ ID NO: 142 (CCACCCCAAACTGCTGGACCATCCT); SEQ ID NO: 143 (CAAAATCCACCCCAAACTGCTGGAC); SEQ ID NO: 144 (TCGATCCTCTTCTCAAAATCCACCC); SEQ ID NO: 145 (CCCTCGATCCTCTTCTCAAAATCCA); SEQ ID NO: 146 (CCTGAGCCCTCGATCCTCTTCTCAA); SEQ ID NO: 147 (ACCTGATCTCCTGAGCCCTCGATCC); SEQ ID NO: 148 (CAGAGTGTCCACCTGATCTCCTGAG); SEQ ID NO: 149 (AGCTCCAGAGTGTCCACCTGATCTC); SEQ ID NO: 150 (CGGAGAGCTCCAGAGTGTCCACCTG); SEQ ID NO: 151 (GGCGCCCCCGGAGAGCTCCAGAGT); SEQ ID NO: 152 (GCGATTGATTCGGGCGCCCCCGGAG); SEQ ID NO: 153 (GGAAGATGCGATTGATTCGGGCGCC); SEQ ID NO: 154 (GGAACCGCTCGTGGAAGATGCGAT); SEQ ID NO: 155 (AGCTCAAATGGGAACCGCTCGTGGA); SEQ ID NO: 156 (GCACTACCTTCACCAGCTCAAATGG); and SEQ ID NO: 157 (CCCGGGGGGCACTACCTTCACCAGC). [0602] In certain embodiments, the target region is within the exon internal region of exon 16 (SEQ ID NO: 9), the intron 15/exon 16 junction, or the exon 16/intron 16 junction. [0603] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3), DNM2 H16A(-13+12), DNM2 H16A(-6+19), DNM2 H16A(+6+30), DNM2 H16A(+15+39), DNM2 H16A(+26+50), DNM2 H16A(+36+60), DNM2 H16A(+45+69), DNM2 H16A(+55+79), DNM2 H16A(+66+90), DNM2 H16A(+75+99), DNM2 H16A(+82+106), DNM2 H16D(+19-6), or DNM2 H16D(+11-14). DNM2 H16A(-21+3).
[0604] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-13+12). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H16A(-6+19). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+6+30). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+15+39). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+26+50). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+36+60). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+45+69). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+55+79). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+66+90). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+75+99). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+82+106). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16D(+19-6). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H16D(+11-14). [0605] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 288 (CTCCTGAGGGTGGAGGGAAAAGAG); SEQ ID NO: 289 (CTTCTCTTTCTCCTGAGGGTGGAGG); SEQ ID NO: 290 (TGTACTTCTTCTCTTTCTCCTGAGG); SEQ ID NO: 291 (CAGAGGCAGCATGTACTTCTTCTCT); SEQ ID NO: 292 (GAGGTTGTCCAGAGGCAGCATGTAC); SEQ ID NO: 293 (TCACGGATCTTGAGGTTGTCCAGAG); SEQ ID NO: 294 (CTTCTCCACATCACGGATCTTGAGG); SEQ ID NO: 295 (CATGAAGCCCTTCTCCACATCACGG); SEQ ID NO: 296 (GCTTGTTGGACATGAAGCCCTTCTC); SEQ ID NO: 297 (GGCGAAGACGTGCTTGTTGGACATG); SEQ ID NO: 298 (GTTGAAGATGGCGAAGACGTGCTTG); SEQ ID NO: 299 (GCTCCGTGTTGAAGATGGCGAAGAC); SEQ ID NO: 300 (CCTCACCTCTGCTCCGTGTTGAAGA); and SEQ ID NO: 301 (CCTGGGCCCCTCACCTCTGCTCCGT). [0606] In certain embodiments, the target region is within the exon internal region of exon 19 (SEQ ID NO: 9), the intron 18/exon 19 junction, or the exon 18/intron 18 junction. [0607] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14), DNM2 H19A(-4+21), DNM2 H19A(+1+25), DNM2 H19A(+7+31), DNM2
H19A(+16+40), DNM2 H19A(+26+50), DNM2 H19A(+37+61), DNM2 H19A(+47+71), DNM2 H19A(+56+80), DNM2 H19A(+66+90), DNM2 H19A(+76+100), DNM2 H19A(+86+110), DNM2 H19A(+96+120), DNM2 H19A(+106+130), DNM2 H19A(+117+141), DNM2 H19A(+127+151), DNM2 H19A(+137+161), DNM2 H19A(+147+171), DNM2 H19A(+157+181), DNM2 H19A(+166+189), DNM2 H19A(+191+215), DNM2 H19A(+200+224), DNM2 H19D(+23-2), DNM2 H19D(+17-8), or DNM2 H19D(+10-15). [0608] In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-4+21). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+1+25). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+7+31). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+16+40). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+26+50). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+37+61). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+47+71). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+56+80). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+66+90). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+76+100). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+86+110). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+96+120). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+106+130). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+117+141). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+127+151). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+137+161). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+147+171). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+157+181). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+166+189). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+191+215). In certain embodiments, the target region of the human DNM2 gene pre- mRNA is DNM2 H19A(+200+224). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+23-2). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+17-8). In certain embodiments, the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+10-15).
[0609] In certain embodiments, the targeting sequence is selected from: SEQ ID NO: 302 (ATGAAGGCCTTCGTCTGCGGGCAGG); SEQ ID NO: 303 (GTGGTGGATGAAGGCCTTCGTCTGC); SEQ ID NO: 304 (GCTCGTGGTGGATGAAGGCCTTCGT); SEQ ID NO: 305 (CCAGCAGCTCGTGGTGGATGAAGGC); SEQ ID NO: 306 (ATAGGTAGGCCAGCAGCTCGTGGTG); SEQ ID NO: 307 (GCCGAGGAGTATAGGTAGGCCAGCA); SEQ ID NO: 308 (TGCTCTGGTCTGCCGAGGAGTATAG); SEQ ID NO: 309 (TCCATGAGGCTGCTCTGGTCTGCCG); SEQ ID NO: 310 (GCCGACTCCTCCATGAGGCTGCTCT); SEQ ID NO: 311 (TGCCTGGTCAGCCGACTCCTCCATG); SEQ ID NO: 312 (CCCGCCGCTGTGCCTGGTCAGCCGA); SEQ ID NO: 313 (AGCATGTCGTCCCGCCGCTGTGCCT); SEQ ID NO: 314 (GTACATGCGCAGCATGTCGTCCCGC); SEQ ID NO: 315 (TGAGGGCATGGTACATGCGCAGCAT); SEQ ID NO: 316 (GAGCGCCTCCTTGAGGGCATGGTAC); SEQ ID NO: 317 (CGATGATGTTGAGCGCCTCCTTGAG); SEQ ID NO: 318 (CTGATGTCACCGATGATGTTGAGCG); SEQ ID NO: 319 (AGTGCTGGTGCTGATGTCACCGATG); SEQ ID NO: 320 (GCGTGGACACAGTGCTGGTGCTGAT); SEQ ID NO: 321 (GGGTACAGGCGTGGACACAGTGCT); SEQ ID NO: 322 (TGGAGCCAGGTGTCATCGACAGGCG); SEQ ID NO: 323 (CTGGCGCTCTGGAGCCAGGTGTCAT); SEQ ID NO: 324 (ACCTGTGGCTGCTGGCGCTCTGGAG); SEQ ID NO: 325 (TTCCGGACCTGTGGCTGCTGGCGCT); and SEQ ID NO: 326 (ACCAGGCTTCCGGACCTGTGGCTGC). [0610] In one embodiment, the targeting sequence is at least 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99 percent complementary to the target region. In another embodiment, the targeting sequence is at least 84%, at least 88%, or at least 92% complementary to the target region. In still another embodiment, the targeting sequence is at least 90% complementary to the target region. In yet another embodiment, the targeting sequence is at least 95% complementary to the target region. In still another embodiment, the targeting sequence is 100% complementary to the target region.
[0611] The antisense oligomer of the present disclosure can have a modified chemical backbone as described herein or known in the art. For example, the antisense oligomer may be a peptide nucleic acid (PNA), a locked nucleic acid, a phosphorodiamidate morpholino oligomer, a 2′-OMe phosphorothioate oligomer, or a combination thereof. In an embodiment, the antisense oligomer is a phosphorodiamidate morpholino oligomer. In another embodiment, the antisense oligomer is a peptide nucleic acid. In yet another embodiment, the antisense oligomer is a locked nucleic acid. In still another embodiment, the antisense oligomer is a 2′-OMe phosphorothioate oligomer. In still another embodiment, the antisense oligomer is a combination of any of the above oligonucleotides having a modified chemical backbone. [0612] In some embodiments, the antisense oligomer is conjugated to, e.g., covalently attached to, a delivery moiety such as, for example, a cell-penetrating peptide, an antibody, a fragment of an antibody, or an antigen fragment of an antibody. In an embodiment, the antisense oligomer is conjugated to a cell-penetrating peptide. In another embodiment, the antisense oligomer is conjugated to a cell-penetrating peptide, wherein the cell-penetrating peptide is selected from rTAT, Tat, R9F2, R5F2R4, R4, R5, R6, R7, R8, R9, (RXR)4, (RXR)5, (RXRRBR)2, (RAR)4F2, (RGR)4F2, and RBRBYLIQFRBRRBR, wherein A represents alanine, B represents beta-alanine, F represents phenylalanine, G represents glycine, I represents isoleucine, L represents leucine, Q represents glutamine, R represents arginine, X represent 6-aminohexanoic acid, and Y represents tyrosine. In yet another embodiment, the antisense oligomer is conjugated to an antibody, an antibody fragment, or an antigen fragment of an antibody. [0613] The antisense oligomer of the present disclosure may be attached to the delivery moiety via a linker. In an embodiment, the linker is a direct bond from the antisense oligomer to the delivery moiety. In another embodiment, the linker is an amino acid such as, for example, a glycine amino acid, a proline amino acid, or a glutamic amino acid. In a particular embodiment, the linker is a glycine amino acid or a proline amino acid. [0614] Provided herein is an antisense oligomer, or a pharmaceutically acceptable salt thereof, having a structure of Formula (I):
wherein A′, R1, R2, t, E′, and G are as described herein. [0261] In certain embodiments, t is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28. In some embodiments, each R2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt. [0262] In an embodiment, E′ is selected from H, -C1-6-alkyl, -C(O)C1-6-alkyl, benzoyl, stearoyl, trityl, monomethoxytrityl, dimethoxytrityl, trimethoxytrityl, and
. [0615] In a further embodiment, E′ is selected from -H, -C(O)CH3, benzoyl, stearoyl, trityl, 4- methoxytrityl, and
. [0616] In an embodiment, A′ is selected from:
. [0617] In an embodiment, at least one of the following is true:
[0618] In an embodiment, A′ is selected from:
; and E′ is
. [0619] In an embodiment, A′ is
[0620] E′ is selected from H, -C(O)CH3, trityl, 4-methoxytrityl, benzoyl, and stearoyl. In an embodiment, each R1 is -N(CH3)2. In an embodiment, L is glycine, proline, or b-alanine. In an embodiment, L is glycine. In an embodiment, L is proline. In an embodiment, L is b-alanine. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt. [0621] In some embodiments, provided herein is an antisense oligomer, which is an oligomer of Formula (IA):
or a pharmaceutically acceptable salt thereof, wherein: A′ is a moiety selected from:
are as described herein. [0622] In certain embodiments, t is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28. In some embodiments, each R2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt.
[0623] In some embodiments, provided herein is an antisense oligomer, which is an oligomer of Formula (II):
or a pharmaceutically acceptable salt thereof, wherein t, G, and R2 are as described herein. [0624] In certain embodiments, t is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28. In some embodiments, each R2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt.
[0625] In some embodiments, provided herein is an antisense oligomer, which is an oligomer of Formula (III):
(III), or a pharmaceutically acceptable salt thereof, wherein n is 11-28 and R2 is as described herein. [0626] In certain embodiments, n is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28. In some embodiments, each R2 taken together forms a targeting sequence of 13- 30 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. In certain embodiments, the pharmaceutically acceptable salt is an HCl salt. [0627] In some embodiments, provided herein is an antisense oligomer, which is an oligomer of Formula (IV):
(IV), wherein n is 11-28 and R2 is as described herein.
[0628] In certain embodiments, n is 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, or 28. In some embodiments, each R2 taken together forms a targeting sequence of 13-30 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11-28 bases. In some embodiments, each R2 taken together forms a targeting sequence of 18-27 bases. In some embodiments, each R2 taken together forms a targeting sequence of 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30 bases. [0629] In certain embodiments, each R2, taken together, forms a targeting sequence having SEQ ID NO: 83. In certain embodiments, each R2, taken together, forms a targeting sequence having SEQ ID NO: 88. In certain embodiments, each R2, taken together, forms a targeting sequence having SEQ ID NO: 172. In certain embodiments, R2, taken together, forms a targeting sequence having SEQ ID NO: 199. In certain embodiments, each R2, taken together, forms a targeting sequence having SEQ ID NO: 230. In certain embodiments, each R2, taken together, forms a targeting sequence having SEQ ID NO: 278. [0630] In certain embodiments, the antisense oligomer is selected from:
[0631] In certain embodiments, the antisense oligomer has the structure:
. [0632] In certain embodiments, the antisense oligomer has the structure:
[0636] In certain embodiments, the antisense oligomer has the structure:
. IV. Target Sequences and Target Regions [0637] In some embodiments for antisense applications, the oligomer can be 100% complementary to the nucleic acid target sequence (excluding at least one abasic subunit), or it may include mismatches, e.g., to accommodate variants, as long as a heteroduplex formed between the oligomer and nucleic acid target sequence is sufficiently stable to withstand the action of cellular nucleases and other modes of degradation which may occur in vivo. Mismatches, if present, are less destabilizing toward the end regions of the hybrid duplex than in the middle. The number of mismatches allowed will depend on the length of the oligomer, the percentage of G: C base pairs in the duplex, and the position of the mismatch(es) in the duplex, according to well-understood principles of duplex stability. Although such an antisense oligomer is not necessarily 100% complementary to the nucleic acid target sequence, it is effective to
stably and specifically bind to the target sequence, such that a biological activity of the nucleic acid target, e.g., expression of the encoded protein(s), is modulated. [0638] The stability of the duplex formed between an oligomer and the target sequence is a function of the binding Tm and the susceptibility of the duplex to cellular enzymatic cleavage. The Tm of an antisense compound with respect to complementary-sequence RNA may be measured by conventional methods, such as those described by Hames et al., Nucleic Acid Hybridization, IRL Press, 1985, pp.107-108 or as described in Miyada CG. and Wallace RB (1987) Oligonucleotide hybridization techniques, Methods Enzymol. Vol.154 pp.94-107. [0639] In some embodiments, each antisense oligomer has a binding Tm, with respect to a complementary-sequence RNA, of greater than body temperature or in other embodiments greater than 50 °C. In other embodiments, Tms are in the range of 60-80 °C or greater. According to well-known principles, the Tm of an oligomer compound, with respect to a complementary-based RNA hybrid, can be increased by increasing the ratio of C: G paired bases in the duplex, and/or by increasing the length (in base pairs) of the heteroduplex. At the same time, for purposes of optimizing cellular uptake, it may be advantageous to limit the size of the oligomer. In certain embodiments, compounds show high Tm (50 °C or greater) at a length of 20 bases or less. For some applications, longer oligomers, for example longer than 20 bases, may have certain advantages. [0640] The targeting sequence bases may be normal DNA bases or analogues thereof, e.g., uracil and inosine that are capable of Watson-Crick base pairing to target-sequence RNA bases. [0641] An antisense oligomer can be designed to block or inhibit or modulate translation of mRNA or to inhibit or modulate pre-mRNA splice processing, or induce degradation of targeted mRNAs, and may be said to be “directed to” or “targeted against” a target sequence with which it hybridizes. In certain embodiments, the target sequence includes a region including a 3′ or 5′ splice site of a pre-processed mRNA, a branch point, or other sequence involved in the regulation of splicing. The target sequence may be within an exon or within an intron or spanning an intron/exon junction. [0642] An antisense oligomer having a sufficient sequence complementarity to a target RNA sequence to modulate splicing of the target RNA means that the antisense agent has a sequence sufficient to trigger the masking of a binding site for a native protein that would otherwise modulate splicing and/or alters the three-dimensional structure of the targeted RNA. Likewise, an oligomer reagent having a sufficient sequence complementary to a target RNA sequence to modulate splicing of the target RNA means that the oligomer reagent has a sequence sufficient to trigger the masking of a binding site for a native protein that would otherwise modulate splicing and/or alters the three-dimensional structure of the targeted RNA.
[0643] In certain embodiments, the antisense oligomer has sufficient length and complementarity to target region within a pre-mRNA of the human dynamin 2 (DNM2) gene (SEQ ID NO: 1). The human DNM2 gene (SEQ ID NO:1) and the exon 2 (SEQ ID NO: 2), exon 6 (SEQ ID NO: 3), exon 7 (SEQ ID NO: 4), exon 8 (SEQ ID NO: 5), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), exon 13 (SEQ ID NO: 8), exon 16 (SEQ ID NO: 9), and exon 19 (SEQ ID NO: 10) sequences for human DNM2 gene pre-mRNA are shown in Table 2. Table 2. Target Regions for DNM2-targeted oligomers.
[0644] In certain embodiments, the degree of complementarity between the target sequence and antisense targeting sequence is sufficient to form a stable duplex. The region of complementarity of the antisense oligomers, excluding the abasic units, with the target RNA sequence may be as short as 8-11 bases, but can be 12-15 bases or more, e.g., 10-40 bases, 12-30 bases, 12-25 bases, 15-25 bases, 12-20 bases, or 15-20 bases, including all integers in between these ranges. An antisense oligomer of about 14-15 bases is generally long enough to
have a unique complementary sequence. In certain embodiments, a minimum length of complementary bases may be required to achieve the requisite binding Tm, as discussed herein. [0645] In certain embodiments, oligomers as long as 40 bases may be suitable, where at least a minimum number of bases, e.g., 10-12 bases, are complementary to the target sequence. In some embodiments, facilitated or active uptake in cells is optimized at oligomer lengths of less than about 30 bases. For PMO oligomers, described further herein, an optimum balance of binding stability and uptake generally occurs at lengths of 18-25 bases. Included in the disclosure are antisense oligomers (e.g., PMOs, PMO-X, PNAs, LNAs, 2′-OMe) that consist of about 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, or 40 bases, in which at least about 6, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, or 40 contiguous or non-contiguous bases that are complementary to the desired target sequences. [0646] In certain embodiments, antisense oligomers may be 100% complementary to the target sequence (excluding at least one abasic nucleotide), or may include mismatches, e.g., to accommodate variants, as long as a heteroduplex formed between the oligomer and target sequence is sufficiently stable to withstand the action of cellular nucleases and other modes of degradation which may occur in vivo. Hence, certain oligomers may have substantial complementarity, meaning, about or at least about 70% sequence complementarity, e.g., 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% sequence complementarity, between the oligomer (excluding at least one abasic nucleotide) and the target sequence. Oligomer backbones that are less susceptible to cleavage by nucleases are discussed herein. Mismatches, if present, are typically less destabilizing toward the end regions of the hybrid duplex than in the middle. The number of mismatches allowed will depend on the length of the oligomer, the percentage of G: C base pairs in the duplex, and the position of the mismatch(es) in the duplex, according to well-understood principles of duplex stability. Although such an antisense oligomer is not necessarily 100% complementary to the target sequence, it is effective to stably and specifically bind to the target sequence, such that splicing of the target pre-RNA is modulated. The stability of the duplex formed between an oligomer and a target sequence is a function of the binding Tm and the susceptibility of the duplex to cellular enzymatic cleavage. The Tm of an oligomer with respect to complementary-sequence RNA may be measured by conventional methods, such as those described by Hames et al., Nucleic Acid Hybridization, IRL Press, 1985, pp.107-108 or as described in Miyada C. G. and Wallace R. B., 1987, Oligomer Hybridization Techniques, Methods Enzymol. Vol.154 pp.94-107. In certain
embodiments, antisense oligomers may have a binding Tm, with respect to a complementary- sequence RNA, of greater than body temperature and greater than about 45 °C or 50 °C. Tms in the range 60-80 °C or greater are also included. According to well-known principles, the Tm of an oligomer, with respect to a complementary-based RNA hybrid, can be increased by increasing the ratio of C: G paired bases in the duplex, and/or by increasing the length (in base pairs) of the heteroduplex. At the same time, for purposes of optimizing cellular uptake, it may be advantageous to limit the size of the oligomer. In certain embodiments, the compounds show high Tm (45-50 °C or greater) at a length of 25 bases or less. [0647] In certain embodiments, antisense targeting sequences are designed to hybridize to a region of one or more of the target sequences listed in Table 2. Selected antisense targeting sequences can be made shorter, e.g., about 12 bases, or longer, e.g., about 40 bases, and include a small number of mismatches, as long as the sequence is sufficiently complementary to effect splice modulation upon hybridization to the target sequence, and optionally forms with the RNA a heteroduplex having a Tm of 45 °C or greater. V. Cell-Penetrating Peptides (CPPs) [0648] In an aspect, the antisense oligomer described herein is conjugated to, e.g., covalently attached to, a delivery moiety. Any delivery moiety that increases cellular uptake of the antisense oligomer may be suitable for use herein. For example, the antisense oligomer may be covalently attached to delivery moiety such as a cell-penetrating peptide, an antibody, a fragment of an antibody, or an antigen fragment of an antibody. In an embodiment, the antisense oligomer is conjugated to an antibody or antibody fragment. Antibodies and antibody fragments that affect cellular uptake are known in the art, and any such antibody or antibody fragment may be covalently linked to the oligomer of the present disclosure. In some embodiments, the antisense oligomer is conjugated to an antibody is an anti-TfRl antibody. Examples of such anti-TfRI antibodies, means of conjugation to an antisense oligomer, complexes comprising antisense oligomers covalently linked to an anti-transferrin receptor 1 (TfRl) antibody, and formulations thereof for use in the instant disclosure can be found in WO/2023/283623 and WO/2023/283624, both of which are incorporated by reference in their entirety. [0649] In some embodiments, the antisense oligomer is an antibody-peptide-oligomer conjugate, for example, of formula A-(Xi-B-X2-D)n or A-(Xi-D-X2-B)n, wherein A is an antibody or antigen binding fragment thereof; B is a polynucleotide; D is an endosomolytic peptide or a membrane penetrating peptide; Xi is a bond or a first non-polymeric linker; X2 is an optional bond or an optional second linker; and n is an integer > 1, as described in WO/2022/212886, which is incorporated by reference in its entirety.
[0650] In another embodiment, the antisense oligomer is conjugated to a cell-penetrating peptide. The antisense oligomer of the present disclosure may be covalently linked to a cell- penetrating peptide as described herein or as known in the art. [0651] In particular, arginine-rich cell-penetrating peptides (CPP) discussed herein, e.g., within the scope of substituent J, can be effective in enhancing penetration of antisense oligomers into a cell and to cause exon skipping in different muscle groups in animal models. [0652] Exemplary arginine-rich peptides are provided in Table 3. Table 3: Arginine-Rich Peptide Transporters (J).
aSequences assigned to SEQ ID NOs do not include the linkage portion “-L”. In the amino acid sequence, A represents alanine, B represents β-alanine, F represents phenylalanine, G represents glycine, L represents leucine, R represents arginine, and X represent 6- aminohexanoic acid. In some embodiments, an antisense oligomer disclosed herein is a conjugate of D-Pep 1.9b wherein L is isoglutamine (IsoGln). Examples of such conjugates can be found in WO/2022/171972 which is incorporated by reference in its entirety. [0653] In another aspect, exemplary cell-penetrating peptides within the scope of substituent J are provided in Table 4. The point of connection to substituent “L” is as shown in Table 4.
Table 4 Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) (RFF)3; CP0407 RFFRFFRFF-L RTR RTRTRFLRRT-L RFFR RFFRFFRFFR-L KTR KTRTKFLKKT-L KFF KFFKFFKFF-L KFFK KFFKFFKFFK-L (RFF)2 RFFRFF-L (RFF)2R RFFRFFR-L Rα RααRααR-L (RαR)4; P007 RαRRαRRαRRαR-L RBRBYLIQFRBRRBR-L Tat47_58 YGRKKRRQRRR-L Tat48_58 GRKKRRQRRR-L Tat49_58 RKKRRQRRR-L Penetratin RQIKIWFQNRRMKWKKGG-L Transportan GWTLNSAGYLLGKINLKALAALAKKIL-L 2αHph-1 YARVRRRGPRGYARVRRRGPRR-L Hph-1 YARVRRRGPRR-L Sim-2 AK AARQAAR-L HSVl VP22 DAATATRGRSAASRPTERPRAPARSASRPRRPVE- L Pep-1 KETWWETWWTEWSQPKKKRKV-L Pep-2 KETWFETWFTEWSQPKKKRKV-L ANTP RQIKIWFQNRRMKWKK-L R6Pen RRRRRR-RQIKIWFQNRRMKWKKGG-L rTat RRRQRRKKRC-L pTat CYGRKKRRQRRR-L R9F2 RRRRRRRRRFFC-L R9CF2 RRRRRRRRRCFF-L R8CF2R RRRRRRRRCFFR-L R6CF2R3 RRRRRRCFFRRR-L R5FCFR4 RRRRRFCFRRRR-L R5F2R4 RRRRRFFRRRR-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) R4CF2R5 RRRRCFFRRRRR-L R2CF2R7 RRCFFRRRRRRR-L CF2R9 CFFRRRRRRRRR-L CR9F2 CRRRRRRRRRFF-L F2R9 FFRRRRRRRRR-L R5F2CF2R4 RRRRRFFCFFRRRR-L R9I2 RRRRRRRRRII-L R8F3 RRRRRRRRFFF-L R9F4 RRRRRRRRRFFFF-L RsF2 RRRRRRRRFF-L R6F2 RRRRRRFF-L R5F2 RRRRRFF-L (RRα)3RR RRαRRαRRαRR-L (RαR)4 RαRRαRRαRRαR-L (αRR)4 αRRαRRαRRαRR-L (Rα)5RR RαRαRαRαRαR-L (RαR)3 RαRRαRRαR-L (RαR)2R RαRRαRR-L (RαR)2 RαRRαR-L (RKα)3RK RKαRKαRKαRK-L (RHα)3RH RHαRHαRHαRH-L R8CF2R RRRRRRRRCFFR-L (RRα)3RR RRαRRαRRαRR-L (RαR)4; P007 RαRRαRRαRRαR-L (αRR)4 αRRαRRαRRαRR-L (Rα)5R RαRαRαRαRαR-L (Rα)7R RαRαRαRαRαRαR-L (RαR)5 RαRRαRRαRRαRRαR-L (RαRRβR)2; B RαRRβRRαRRβR-L (RαR)3RβR RαRRαRRαRRβR-L (Rβ)5RαRβR RβRβRβRβRβRαRβR-L RβRβRβRαRβRβRβR RβRβRβRαRβRβRβR-L α(Rβ)3Rα(Rβ)3R-α αRβRβRβRαRβRβRβR-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) (RβRα)4 RβRαRβRαRβRαRβR-L (Rβ)4(Rα)3R RβRβRβRβRαRαRαR-L Rα(Rβ)2Rα(Rβ)3R RαRβRβRαRβRβRβR-L (Rβ)7R RβRβRβRβRβRβRβR-L R4 tg-RRRR-L R5 tg-RRRRR-L R6 tg-RRRRRR-L R7 tg-RRRRRRR-L R8 tg-RRRRRRRR-L R5GR4 tg-RRRRRGRRRR-L R5F2R4 tg-RRRRRFFRRRR-L Tat tg-RKKRRQRRR-L rTat tg-RRRQRRKKR-L RαRRαR-L RβRRβR-L RαRRβR-L RβRRαR-L RαRYbRαR-L RβRYbRβR-L RαRYbRβR-L RβRYbRαR-L RαRILFQYRαR-L RβRILFQYRβR-L RαRILFQYRβR-L RβRILFQYRαR-L RαRRαRRαR-L RβRRβRRβR-L RαRRβRRαR-L RαRRβRRβR-L RαRRαRRβR-L RβRRαRRβR-L RβRRαRRαR-L RβRRβRRαR-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) RαRYbRαRRαR-L RαRRαRYbRαR-L RαRILFQYRαRRαR-L RαRRαRILFQYRαR-L RαRYbRαRYbRαR-L RαRILFQYRαRILFQYRαR-L RαRILFQYRαRYbRαR-L RαRYbRαRILFQYRαR-L RβRYbRβRRβR-L RβRRβRYbRβR-L RβRILFQYRβRRβR-L RβRRβRILFQYRβR-L RβRYRβRYbRβR-L RβRILFQYRβRILFQYRβR-L RβRYbRβRILFQYRβR-L RβRILFQYRβRYbRβR-L RαRYbRβRRαR-L RαRRβRYbRαR-L RαRILFQYRβRRαR-L RαRRβRILFQYRαR-L RαRYbRβRYbRαR-L RαRILFQYRβRILFQYRαR-L RαRYbRβRILFQYRαR-L RαRILFQYRβRYbRαR-L RαRYbRβRRβR-L RαRRβRYbRβR-L RαRILFQYRβRRβR-L RαRRβRILFQYRβR-L RαRYbRβRYbRβR-L RαRILFQYRβRILFQYRβR-L RαRYbRβRILFQYRβR-L RαRILFQYRβRYbRβR-L RαRYbRαRRβR-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) RαRRαRYbRβR-L RαRILFQYRαRRβR-L RαRRαRILFQYRβR-L RαRYbRαRYbRβR-L RαRILFQYRαRILFQYRβR-L RαRYbRαRILFQYRβR-L RαRILFQYRαRYbRβR-L RβRYbRαRRβR-L RβRRαRYbRβR-L RβRILFQYRαRRβR-L RβRRαRILFQYRβR-L RβRYbRαRYbRβR-L RβRILFQYRαRILFQYRβR-L RβRYbRαRILFQYRβR-L RβRILFQYRαRYbRβR-L RβRYbRαRRαR-L RβRRαRYbRαR-L RβRILFQYRαRRαR-L RβRRαRILFQYRαR-L RβRYbRαRYbRαR-L RβRILFQYRαRILFQYRαR-L RβRYbRαRILFQYRαR-L RβRILFQYRαRYbRαR-L RβRYbRβRRαR-L RβRRβRYbRαR-L RβRILFQYRβRRαR-L RβRRβRILFQYRαR-L RβRYbRβRYbRαR-L RβRILFQYRβRILFQYRαR-L RβRYbRβRILFQYRαR-L RβRILFQYRβRYbRαR-L RαRRαRRαRRαR-L RαRRβRRαRILFQYRαRβRαR-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) RαRRβRRαRRβR-L YGRKKRRQRRRP-L RαRRαRRαRRαRαβASSLNIAαC-L RαRRβRRαRILFQYRαRβRαRβASSLNIAαC-L RαRRβRRαRASSLNIARαRβRαRβC-L RαRRβRRαRRβRαβASSLNIA-L THRPPMWSPVWP-L HRPPMWSPVWP-L THRPPMWSPV-L THRPPMWSP-L THRPPMWSPVFP-L THRPPMWSPVYP-L THRPPMWSPAWP-L THRPPMWSPLWP-L THRPPMWSPIWP-L THRPPMWTPVVWP-L THRPPMFSPVWP-L THRPPMWS-L HRPPMWSPVW-L THRPPMYSPVWP-L THRPPnleWSPVWP-L (nle = norleucine) THKPPMWSPVWP-L SHRPPMWSPVWP-L STFTHPR-L YDIDNRR-L AYKPVGR-L HAIYPRH-L HTPNSTH-L ASSPVHR-L SSLPLRK-L KKRS-L KRSK-L KKRSK-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) KSRK-L SRKR-L RKRK-L KSRKR-L QHPPWRV-L THPPTTH-L YKHTPTT-L QGMHRGT-L SRKRK-L KSRKRK-L PKKKRKV-L GKKRSKV-L KSRKRKL-L HSPSKIP-L HMATFHY-L AQPNKFK-L NLTRLHT-L KKKR-L KKRK-L KKKRK-L RRRRRRQIKIWFQNRRMKWKKGGC-L RRRRRRRQIKIWFQNRRMKWKKGGC-L RQIKIWFQNRRMKWKKGGC-L RRRRRRRQIKIWFQNRRMKWKKC-L RαRRαRRαRRQIKIWFQNRRMKWKKGGC-L RRRRRRRQIKILFQNRαRαRαRαC-L RαRRαRRαRRαRC-L RαRRαRRαRRαRαC-L RαRRαRRαRIKILFQNRRMKWKKGGC-L RαRRαRRαRIKILFQNRRMKWKKC-L RαRRαRRαRIKILFQNRMKWKKC-L RαRRαRRαRIKILFQNαRMKWKKC-L RαRRαRRαRIKILFQNHRMKWKKC-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) RαRRαRRαRIKILFQNαRMKWKKC-L RαRRαRRαRIKILFQNαRMKWKKC-L RαRRαRRαRIKILFQNαRMKWKAC-L RαRRαRRαRIKILFQNαRMKWHKAC-L RαRRαRRαRIKILFQNαRMKWHRC-L RαRαRαRαRIKILFQNRRMKWKKC-L RARARARARIKILFQNRRMKWKKC-L RαRRαRRαRIαILFQNαRMKWHKAC-L RαRRαRRαRIHILFQNαRMKWHKAC-L RαRRαRRαRIRILFQNαRMKWHKAC-L RαRRαRRαRIαILFQYαRMKWHKAC-L RαRRαRRαRLYSPLSFQαRMKWHKAC-L RαRRαRRαRISILFQYαRMKWHKAC-L RαRRαRRαRILFQYαRMKWHKAC-L RαRRαRIαILFQYαRMKWHKAC-L RαRRARRαRIHILFQYαRMKWHKAC-L RARRαRRARIHILFQYαRMKWHKAC-L RαRRαRRαRIHILFQYαRMKWHKAC-L RαRRαRRαRIαILFQNαRMKWHKAC-L RαRRαRRαRIHILFQNαRMKWHKAC-L RαRRαRRαRIKILFQNRRMKWHK-L RαRRαRRαRIKILFQNαRMKWHK-L RαRRαRRαRIαILFQNRRMKWHK-L RαRRαRRαRIαILFQNαRMKWHK-L RαRRαRRαRIHILFQNRRMKWHK-L RαRRαRRαRIHILFQNαRMKWHK-L RαRRαRRαRIRILFQNRRMKWHK-L RαRRαRRαRIRILFQNαRMKWHK-L RαRRαRRαRIILFQNRRMKWHK-L RαRRαRRαRIILFQNαRMKWHK-L RαRRαRRαRKILFQNRRMKWHK-L RαRRαRRαRKILFQNαRMKWHK-L RαRRαRRαRαILFQNRRMKWHK-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) RαRRαRRαRαILFQNαRMKWHK-L RαRRαRRαRHILFQNRRMKWHK-L RαRRαRRαRHILFQNαRMKWHK-L RαRRαRRαRRILFQNRRMKWHK-L RαRRαRRαRRILFQNαRMKWHK-L RαRRαRRαRILFQNRRMKWHK-L RαRRαRRαRILFQNαRMKWHK-L RαRRαRRαRIKILFQYRRMKWHK-L RαRRαRRαRIKILFQYαRMKWHK-L RαRRαRRαRIαILFQYRRMKWHK-L RαRRαRRαRIαILFQYαRMKWHK-L RαRRαRRαRIHILFQYRRMKWHK-L RαRRαRRαRIHILFQYαRMKWHK-L RαRRαRRαRIRILFQYRRMKWHK-L RαRRαRRαRIRILFQYαRMKWHK-L RαRRαRRαRIILFQYRRMKWHK-L RαRRαRRαRIILFQYαRMKWHK-L RαRRαRRαRKILFQYRRMKWHK-L RαRRαRRαRKILFQYαRMKWHK-L RαRRαRRαRαILFQYRRMKWHK-L RαRRαRRαRαILFQYαRMKWHK-L RαRRαRRαRHILFQYRRMKWHK-L RαRRαRRαRHILFQYαRMKWHK-L RαRRαRRαRRILFQYRRMKWHK-L RαRRαRRαRRILFQYαRMKWHK-L RαRRαRRαRILFQYRRMKWHK-L RαRRαRRαRILFQYαRMKWHK-L RαRRαRRαR-L RαRRαRRαRRαR-L RARRAR-L RARRARRAR-L RARRARRARRAR-L RαRRαRI-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) RαRRARRαR-L RARRαRRAR-L RRRRR-L RRRRRR-L RRRRRRR-L RαRRαRRαRRαRC-L RαRRαRRαRRαRαC-L RαRRαRRαRIKILFQNRRMKWKKGGC-L RαRRαRRαRIKILFQNRRMKWKKC-L RαRRαRRαRIKILFQNRMKWKKC-L RαRRαRRαRIKILFQNαRMKWKKC-L RαRRαRRαRIKILFQNHRMKWKKC-L RαRRαRRαRIKILFQNαRMKWKKC-L RαRRαRRαRIKILFQNαRMKWKKC-L RαRRαRRαRIKILFQNαRMKWKAC-L RαRRαRRαRIKILFQNαRMKWHKAC-L RαRRαRRαRIKILFQNαRMKWHRC-L RαRαRαRαRIKILFQNRRMKWKKC-L RARARARARIKILFQNRRMKWKKC-L RαRRαRRαRIαILFQNαRMKWHKAC-L RαRRαRRαRIHILFQNαRMKWHKAC-L RαRRαRRαRIRILFQNαRMKWHKAC-L RαRRαRRαRIαILFQYαRMKWHKAC-L RαRRαRRαRLYSPLSFQαRMKWHKAC-L RRMKWHK-L αRMKWHK-L ααααααααααααααILFQααRMKWHK-L ααααααααααααααILFQααRMKWHK-L RRRRRRRQIKILFQNPKKKRKVGGC-L HHFFRRRRRRRRRFFC-L HHHHHHRRRRRRRRRFFC-L HHHHHHFFRRRRRRRRRFFC-L HHHHHααRRRRRRRRRFFC-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) HHHHHHααFFRRRRRRRRRFFC-L HHHαRRRRRRRRRFFαHHHC-L αRMKWHK-L αRWKWHK-L RαRARαR-L RαRαRαR-L RARαRAR-L RαRAR-L ααααααααααααααILFQααHMKWHK-L ααααααααααααααILFQααRWKWHK-L ααααααααααααααILFQααHWKWHK-L ααααααααααααααILFQαRαRARαR-L ααααααααααααααILFQαRαRαRαR-L ααααααααααααααILFQαRαRRαR-L ααααααααααααααILFQαRARαRAR-L ααααααααααααααILFQαRαRARαR-L ααααααααααααααILFQαRαRAR-L ααααααααααααααILIQααRMKWHK-L ααααααααααααααILIQααHMKWHK-L ααααααααααααααILIQααRWKWHK-L ααααααααααααααILIQααHWKWHK-L ααααααααααααααILIQαRαRARαR-L ααααααααααααααILIQαRαRαRαR-L ααααααααααααααILIQαRαRRαR-L ααααααααααααααILIQαRARαRAR-L ααααααααααααααILIQαRαRARαR-L ααααααααααααααILIQαRαRAR-L ααααααααααααααILFQααHMKWHK-L ααααααααααααααILFQααRWKWHK-L ααααααααααααααILFQααHWKWHK-L ααααααααααααααILFQαRαRARαR-L ααααααααααααααILFQαRαRαRαR-L ααααααααααααααILFQαRαRRαR-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) ααααααααααααααILFQαRARαRAR-L ααααααααααααααILFQαRαRARαR-L ααααααααααααααILFQαRαRAR-L ααααααααααααααILIQααRMKWHK-L ααααααααααααααILIQααHMKWHK-L ααααααααααααααILIQααRWKWHK-L ααααααααααααααILIQααHWKWHK-L ααααααααααααααILIQαRαRARαR-L ααααααααααααααILIQαRαRαRαR-L ααααααααααααααILIQαRαRRαR-L ααααααααααααααILIQαRARαRAR-L ααααααααααααααILIQαRαRARαR-L ααααααααααααααILIQαRαRAR-L RαRRARRαRRARαA-L RαRRARRαRILFQYαHMKWHKAC-L RαRRARRαRILFQYαRMKWHKAC-L RαRRARRαRILFQYαRWKWHKAC-L RαRRαRRαRRαRC-L RαRRαRRαRIαILFQNαRMKWHKAC-L RαRRαRRαRIHILFQNαRMKWHKAC-L RαRRαRRαRIαILFQYαRMKWHKAC-L RαRRαRRαRLYSPLSFQαRMKWHKAC-L RαRRαRRαRILFQYαRMKWHKAC-L RαRRαRIαILFQYαRMKWHKAC-L RARRαRRARILFQYαRMKWHKAC-L RαRRARRαRILFQYαRMKWHKAC-L RARRαRRARILFQYαRMKWHKAC-L RαRRARRαRILFQYαRMKWHKAC-L RαRRARRαRILFQYαHMKWHKAC-L RαRRARRαRILFQYαRMKWHKAC-L RαRRARRαRILFQYαRWKWHKAC-L RαRRARRαRILFQYαHWKWHKAC-L RαRRARRαRILFQYRαRARαRAC-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) RαRRARRαRILFQYRαRαRαRAC-L RαRRARRαRILIQYαRMKWHKAC-L RαRRαRILFQYRαRRαRC-L RαRRARRαRILFQYRαRARαRAC-L RαRRARRαRILFQYRαRαRαRAC-L RαRRARRαRILIQYαRMKWHKAC-L RαRRαRILFQYRαRRαRCYS-L RARRαRRARILFQYRARαRARAC-L RARRαRRARILFQYRαRARαRAC-L RARRαRRARILFQYRαRRαRAC-L RARRαRRARILFQYRαRARαAC-L RαRRARRαRILFQYRαRRαRAC-L RαRRARRαRILFQYRαRARαAC-L RαRRARRαRIHILFQNαRMKWHKAC-L RαRRARRαRRARαAC-L RαRRARRαRILFQYαHMKWHK-L RαRRARRαRILFQYαRMKWHK-L RαRRARRαRILFQYαRWKWHK-L RαRRARRαRILFQYαRMKWHK-L RαRRARRαRILFQYRαRARαR-L RαRRARRαRILFQYRαRαRαR-L RαRRARRαRILFQYRαRRαR-L RαRRARRαRILFQYRARαRAR-L RαRRARRαRILFQYRαRAR-L RαRRARRαRILIQYαHMKWHK-L RαRRARRαRILIQYαRMKWHK-L RαRRARRαRILIQYαRWKWHK-L RαRRARRαRILIQYαRMKWHK-L RαRRARRαRILIQYRαRARαR-L RαRRARRαRILIQYRαRαRαR-L RαRRARRαRILIQYRαRRαR-L RαRRARRαRILIQYRARαRAR-L RαRRARRαRILIQYRαRAR-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) RARRαRRARILFQYαHMKWHK-L RARRαRRARILFQYαRMKWHK-L RARRαRRARILFQYαRWKWHK-L RARRαRRARILFQYαRMKWHK-L RARRαRRARILFQYRαRARαR-L RARRαRRARILFQYRαRαRαR-L RARRαRRARILFQYRαRRαR-L RARRαRRARILFQYRARαRAR-L RARRαRRARILFQYRαRAR-L RARRαRRARILIQYαHMKWHK-L RARRαRRARILIQYαRMKWHK-L RARRαRRARILIQYαRWKWHK-L RARRαRRARILIQYαRMKWHK-L RARRαRRARILIQYRαRARαR-L RARRαRRARILIQYRαRαRαR-L RARRαRRARILIQYRαRRαR-L RARRαRRARILIQYRARαRAR-L RARRαRRARILIQYRαRAR-L RαRRαRILFQYαHMKWHK-L RαRRαRILFQYαRMKWHK-L RαRRαRILFQYαRWKWHK-L RαRRαRILFQYαRMKWHK-L RαRRαRILFQYRαRARαR-L RαRRαRILFQYRαRαRαR-L RαRRαRILFQYRαRRαR-L RαRRαRILFQYRARαRAR-L RαRRαRILFQYRαRAR-L RαRRαRILIQYαHMKWHK-L RαRRαRILIQYαRMKWHK-L RαRRαRILIQYαRWKWHK-L RαRRαRILIQYαRMKWHK-L RαRRαRILIQYRαRARαR-L RαRRαRILIQYRαRαRαR-L
Exemplary Carrier Peptide Sequences Name Sequence (Amino to Carboxy Terminus) RαRRαRILIQYRαRRαR-L RαRRαRILIQYRARαRAR-L RαRRαRILIQYRαRAR-L PRPαααααααααααPRG-L RRRRRRRR-L RRMKWKK-L PKKKRKV-L CKDEPQRRSARLSAKPAPPKPEPKPKKAPAKK-L RKKRRQRRR-L RKKRRQRR-L RKKRRQR-L KKRRQRRR-L KKRRQRRR-L AKKRRQRRR-L RAKRRQRRR-L RKARRQRRR-L RKKARQRRR-L [0654] L = as recited in Formula (I) and as described throughout the specification; β = 3-alanine; α = 6-aminohexanoic acid; tg = unmodified amino terminus, or the amino-terminal capped with an acetyl, benzoyl or stearoyl group (i.e., an acetyl amide, benzoyl amide or stearoyl amide) and Y is NH-(CHR)-C(O) - wherein n is 2 to 7 and each R is independently, at each occurrence, hydrogen or methyl. For simplicity, not all sequences are noted with a terminal td group; however, each of the above sequences may comprise an unmodified amino terminus or an amino terminus capped with an acetyl, benzoyl, or stearoyl group. Linker “L” may additionally be an Isoglutamine amino acid residue (IsoGln) linking the carrier peptide to the oligomer by an amide bond between the carboxyl of IsoGln and the oligomer and an amide bond between the amino of IsoGln and the carboxy terminus of the peptide. Examples of use of IsoGln as a linker can be found in, for example, WO/2022/171972, which is incorporated by reference in its entirety. VI. Pharmaceutical Compositions [0655] The present disclosure also provides for the formulation and delivery of the disclosed antisense oligomers. Accordingly, an aspect of the present disclosure is a pharmaceutical
composition comprising antisense oligomers as disclosed herein and a pharmaceutically acceptable carrier. [0656] Effective delivery of the antisense oligomers to the target nucleic acid is an important aspect of treatment. Routes of antisense oligomer delivery include, but are not limited to, various systemic routes, including oral and parenteral routes, e.g., intravenous, subcutaneous, intraperitoneal, and intramuscular, as well as inhalation, transdermal, and topical delivery. The appropriate route may be determined by one of skill in the art, as appropriate to the condition of the subject under treatment. For example, an appropriate route for delivery of an antisense oligomer in the treatment of a viral infection of the skin is topical delivery, while the delivery of an antisense oligomer for the treatment of a viral respiratory infection can be intravenous or by inhalation. The antisense oligomer may also be delivered directly to any particular site of viral infection. [0657] The antisense oligomer can be administered in any convenient vehicle which is physiologically and/or pharmaceutically acceptable. Such a composition can include any of a variety of standard pharmaceutically acceptable carriers employed by those of ordinary skill in the art. Examples include, but are not limited to, saline, phosphate-buffered saline (PBS), water (e.g., sterile water for injection), aqueous ethanol, emulsions such as oil/water emulsions or triglyceride emulsions, tablets, and capsules. The choice of a suitable physiologically acceptable carrier will vary dependent upon the chosen mode of administration. [0658] The compounds disclosed herein (e.g., an antisense oligomer) can generally be utilized as the free acid or free base. Alternatively, the compounds disclosed herein may be used in the form of acid or base addition salts. Acid addition salts of the free amino compounds may be prepared by methods well known in the art and may be formed from organic and inorganic acids. Suitable organic acids include maleic, fumaric, benzoic, ascorbic, succinic, methanesulfonic, acetic, trifluoroacetic, oxalic, propionic, tartaric, salicylic, citric, gluconic, lactic, mandelic, cinnamic, aspartic, stearic, palmitic, glycolic, glutamic, and benzenesulfonic acids. Suitable inorganic acids include hydrochloric, hydrobromic, sulfuric, phosphoric, and nitric acids. Base addition salts included those salts that form with the carboxylate anion and include salts formed with organic and inorganic cations such as those chosen from the alkali and alkaline earth metals (for example, lithium, sodium, potassium, magnesium, barium and calcium), as well as the ammonium ion and substituted derivatives thereof (for example, dibenzylammonium, benzylammonium, 2-hydroxyethylammonium, and the like). Thus, the term “pharmaceutically acceptable salt” of Formulae (I), (IA), (II), and (III) is intended to encompass any and all acceptable salt forms.
[0659] In addition, prodrugs are also included within the context of this disclosure. Prodrugs are any covalently bonded carriers that release a compound of structure (I) in vivo when such a prodrug is administered to a patient. Prodrugs are generally prepared by modifying functional groups in a way such that the modification is cleaved, either by routine manipulation or in vivo, yielding the parent compound. Prodrugs include, for example, compounds of this disclosure wherein hydroxy, amine, or sulfhydryl groups are bonded to any group that, when administered to a patient, cleaves to form the hydroxy, amine, or sulfhydryl groups. Thus, representative examples of prodrugs include (but are not limited to) acetate, formate, and benzoate derivatives of alcohol and amine functional groups of the compounds of structure (I). Further, in the case of a carboxylic acid (-COOH), esters may be employed, such as methyl esters, ethyl esters, and the like. VII. Synthesis Details [0660] Preparation of Oligomers with Basic Nitrogen Internucleoside Linkers [0661] Morpholino subunits, the modified intersubunit linkages, and oligomers comprising the same can be prepared as described, for example, in U.S. Patent Nos.5,185,444, and 7,943,762, which are incorporated by reference in their entireties. The morpholino subunits can be prepared according to the following general Reaction Scheme 1. Reaction Scheme 1. Preparation of Morpholino Subunit
[0662] Referring to Reaction Scheme 1, wherein B′ represents a base-pairing moiety and PG represents a protecting group, the morpholino subunits may be prepared from the corresponding ribonucleoside (1) as shown. The morpholino subunit (2) may be optionally protected by reaction with a suitable protecting group precursor, for example, trityl chloride. The 3′ protecting group is generally removed during solid-state oligomer synthesis as described in more detail below. The base pairing moiety may be suitably protected for sold phase oligomer synthesis. Suitable
protecting groups include benzoyl for adenine and cytosine, phenylacetyl for guanine, and pivaloyloxymethyl for hypoxanthine (I). The pivaloyloxymethyl group can be introduced onto the N1 position of the hypoxanthine heterocyclic base. Although an unprotected hypoxanthine subunit may be employed, yields in activation reactions are far superior when the base is protected. Other suitable protecting groups include those disclosed in U.S. Patent No. 8,076,476, which is hereby incorporated by reference in its entirety. [0663] Reaction of 3 with the activated phosphorous compound 4 results in morpholino subunits having the desired linkage moiety 5. Compounds of structure 4 can be prepared using any number of methods known to those of skill in the art. For example, such compounds may be prepared by reaction of the corresponding amine and phosphorous oxychloride. In this regard, the amine starting material can be prepared using any method known in the art, for example those methods described in the Examples and in U.S. Patent No.7,943,762. [0664] Compounds of structure 5 can be used in solid-phase automated oligomer synthesis for the preparation of oligomers comprising the intersubunit linkages. Such methods are well known in the art. Briefly, a compound of structure 5 may be modified at the 5′ end to contain a linker to a solid support. For example, compound 5 may be linked to a solid support by a linker. Once supported, the protecting group (e.g., trityl) is removed and the free amine is reacted with an activated phosphorous moiety of a second compound of structure 5. This sequence is repeated until the desired length of oligo is obtained. The protecting group in the terminal 5′ end may either be removed or left on if a 5′-modification is desired. [0665] The preparation of modified morpholino subunits and morpholino oligomers are described in more detail in the Examples. The morpholino oligomers containing any number of modified linkages may be prepared using methods described herein, methods known in the art and/or described by reference herein. Also described in the examples are global modifications of morpholino oligomers prepared as previously described (see e.g., PCT publication WO 2008/036127). [0666] Synthesis of PMO, PMO+, PPMO, and PMO-X containing further linkage modifications as described herein was done using methods known in the art and described in pending U.S. Patent Nos.8,299,206 and 8,076,476 and PCT publication numbers WO 2009/064471, WO 2011/150408 and WO 2012/150960, which are hereby incorporated by reference in their entirety. [0667] PMO with a 3′ trityl modification are synthesized essentially as described in PCT publication number WO 2009/064471 with the exception that the detritylation step is omitted. VIII. Methods of Use [0668] Provided herein is a method of reducing the expression of a pre-mRNA of the human DNM2 gene and/or protein using the antisense oligomers of the present disclosure for
therapeutic purposes (e.g., treating subjects with a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein, including but not limited to a centronuclear myopathy). The method comprises administering to a patient in need thereof a therapeutically effective amount of an antisense oligomer disclosed herein or a pharmaceutical composition thereof. In an embodiment, the disease is a centronuclear myopathy. In some embodiments, the antisense oligomer comprising a nucleotide sequence of sufficient length and complementarity to specifically hybridize to a region within the pre-mRNA of the human DNM2 gene, wherein binding of the antisense oligomer to the region induces exon skipping in the human DNM2 gene pre-mRNA, thereby triggering nonsense-mediated decay of DNM2 in a cell and/or tissue of the subject. Exemplary antisense targeting sequences are shown in Tables 5 and 6 herein. In certain embodiments, the disease or disorder is caused by at least one mutation in a gene selected from DNM2 gene, MTM1 gene, BIN1 gene, RYR1 gene, TTN gene, SPEG gene, CACNA1S gene, MTMR14 gene, ZAK gene, and CCDC78 gene. In certain embodiments, the disease or disorder is autosomal dominant, autosomal recessive, or X-linked. [0669] In certain embodiments, the disease or disorder is a genetic muscle disorder. In certain embodiments, the disease or disorder is a neuromuscular disorder. In certain embodiments, the disease or disorder is a centronuclear myopathy (CNM). [0670] In certain embodiments, the disease or disorder is myotubular myopathy (XLMTM or XLCNM), autosomal dominant congenital myopathy (ADCNM), autosomal recessive centronuclear myopathy (ARCNM), or Charcot-Marie-Tooth disease (CMT). In certain embodiments, the CMT is DNM2-implicated CMT. [0671] In certain embodiments, the Charcot-Marie-Tooth disease (CMT) is Charcot-Marie-Tooth disease, dominant intermediate B (CMTDIB or DI-CMTB). In certain embodiments, the Charcot- Marie-Tooth disease is Charcot-Marie-Tooth disease, axonal type 2M (CMT2M). [0672] In certain embodiments, the disease or disorder is a hereditary spastic paraplegia (HSP). [0673] In certain embodiments, the disease or disorder is prostate cancer or pancreatic cancer. [0674] Also included are antisense oligomers for use in the preparation of a medicament for the treatment of a centronuclear myopathy (CNM), comprising a nucleotide sequence of sufficient length and complementarity to specifically hybridize to a target region within the pre-mRNA of the human DNM2 gene, wherein binding of the antisense oligomer to the target region decreases the expression of DNM2. [0675] In some embodiments of the method of treating a centronuclear myopathy (CNM) or the medicament for the treatment of CNM, the antisense oligomer compound comprises a non- natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of the human dynamin 2 (DNM2) gene
(SEQ ID NO: 1). In certain embodiments, the target region is an intron/exon junction, exon/intron junction, an exon internal region, or intron internal region of the human DNM2 gene pre-mRNA. [0676] In some embodiments of the method of treating CNM or the medicament for the treatment of CNM, the antisense oligomer compound comprises a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of the human dynamin 2 (DNM2) gene (SEQ ID NO: 1). In certain embodiments, the target region is within an exon internal region of the human DNM2 gene pre- mRNA selected from exon 2 (SEQ ID NO: 2), exon 6 (SEQ ID NO: 3), exon 7 (SEQ ID NO: 4), exon 8 (SEQ ID NO: 5), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), exon 13 (SEQ ID NO: 8), exon 16 (SEQ ID NO: 9), and exon 19 (SEQ ID NO: 10). In certain embodiments, the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), and exon 13 (SEQ ID NO: 8). [0677] In certain embodiments, the target region is within an intron-exon junction or an exon- intron junction region of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 5/exon 6 junction, exon 6/intron 6 junction, intron 6/exon 7 junction, exon 7/intron 7 junction, intron 7/exon 8 junction, exon 8/intron 8 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, exon 13/intron 13 junction, intron 15/exon 16 junction, exon 16/intron 16 junction, intron 18/exon 19 junction, and exon 19/intron 19 junction [0678] In certain embodiments, the target region is the target region is within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, and exon 13/intron 13 junction. [0679] In certain embodiments, the target region is the target region is within an intron internal region of the human DNM2 gene pre-mRNA selected from intron 2 (SEQ ID NO: 11) and intron 12 (SEQ ID NO: 12). [0680] In some embodiments of the method of treating CNM or the medicament for the treatment of CNM, the antisense oligomer compound comprises a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of the human dynamin 2 (DNM2) gene (SEQ ID NO: 1). In certain embodiments, the target region is an intron/exon junction or an exon internal region of the human DNM2 gene pre-mRNA, and wherein the antisense oligomer is an antisense oligomer of
Formula (I), Formula (IA), Formula (II), or Formula (III). In certain embodiments, a subject has aberrant expression and/or aggregation of DNM2 protein. [0681] Certain embodiments relate to methods of reducing DNM2 expression in a cell, tissue, and/or subject, as described herein. In some instances, DNM2 expression is reduced by about or at least about 5%, 6%, 7%, 8%, 9%, 10%, 11%, 12%, 13%, 14%, 15%, 16%, 17%, 18%, 19%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100% relative to a control, for example, a control cell/subject, a control composition without the antisense oligomer, the absence of treatment, and/or an earlier time-point. In some instances, DNM2 expression is reduced by about 5% to about 100%, relative to a control. In some instances, DNM2 expression is reduced by about 10% to about 95%, relative to a control. In some instances, DNM2 expression is reduced by about 20% to about 95%, relative to a control. In some instances, DNM2 expression is reduced by about 30% to about 95%, relative to a control. In some instances, DNM2 expression is reduced by about 30% to about 90%, relative to a control. In some instances, DNM2 expression is reduced by about 40% to about 90%, relative to a control. In some instances, DNM2 expression is reduced by about 50% to about 90%, relative to a control. In some instances, DNM2 expression is reduced by about 45% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 50% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 55% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 60% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 65% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 50% to about 80%, relative to a control. In some instances, DNM2 expression is reduced by about 60% to about 80%, relative to a control. In some instances, DNM2 expression is reduced by about 50% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 50% to about 70%, relative to a control. In some instances, DNM2 expression is reduced by about 60% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 65% to about 75%, relative to a control. [0682] In some instances, DNM2 expression is reduced by about 30% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 30% to about 80%, relative to a control. In some instances, DNM2 expression is reduced by about 85% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 40% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by about 40% to about 80%, relative to a control. In some instances, DNM2 expression is reduced by about 40% to about 85%, relative to a control. In some instances, DNM2 expression is reduced by about 20% to about 75%, relative to a control. In some instances, DNM2 expression is reduced by
about 20% to about 80%, relative to a control. In some instances, DNM2 expression is reduced by about 20% to about 85%, relative to a control. [0683] Also included are methods of reducing one or more symptoms of a centronuclear myopathy in a subject in need thereof. Examples include symptoms such as elevated blood creatinine levels, interstitial fibrosis, tubular atrophy, gout, hyperuricemia, and end stage kidney disease (ESKD). [0684] The antisense oligomers of the disclosure can be administered to subjects to treat (prophylactically or therapeutically) centronuclear myopathy. In conjunction with such treatment, pharmacogenomics (i.e., the study of the relationship between an individual’s genotype and that individual’s response to a foreign compound or drug) may be considered. Differences in the metabolism of therapeutics can lead to severe toxicity or therapeutic failure by altering the relation between dose and blood concentration of the pharmacologically active drug. [0685] Thus, a physician or clinician may consider applying knowledge obtained in relevant pharmacogenomics studies in determining whether to administer a therapeutic agent as well as tailoring the dosage and/or therapeutic regimen of treatment with a therapeutic agent. [0686] Effective delivery of the antisense oligomer to the target nucleic acid is one aspect of treatment. Routes of antisense oligomer delivery include, but are not limited to, various systemic routes, including oral and parenteral routes, e.g., intravenous, subcutaneous, intraperitoneal, and intramuscular, as well as inhalation, transdermal, and topical delivery. The appropriate route may be determined by one of skill in the art, as appropriate to the condition of the subject under treatment. Vascular or extravascular circulation, the blood or lymph system, and cerebrospinal fluid are some non-limiting sites where the RNA may be introduced. Direct CNS delivery may be employed, for instance, intracerebral ventricular or intrathecal administration may be used as routes of administration. [0687] In certain embodiments, the antisense oligomer(s) are administered to the subject by intramuscular injection (IM), i.e., they are administered or delivered intramuscularly. Non-limiting examples of intramuscular injection sites include the deltoid muscle of the arm, the vastus lateralis muscle of the leg, and the ventrogluteal muscles of the hips, and dorsogluteal muscles of the buttocks. In specific embodiments, a PMO, PMO-X, or PPMO is administered by IM. [0688] In certain embodiments, the antisense oligomers of the disclosure can be delivered by transdermal methods (e.g., via incorporation of the antisense oligomers into, e.g., emulsions, with such antisense oligomers optionally packaged into liposomes). Such transdermal and emulsion/liposome-mediated methods of delivery are described for delivery of antisense oligomers in the art, e.g., in U.S. Pat. No.6,965,025, the contents of which are incorporated in their entirety by reference herein.
[0689] The antisense oligomers described herein may also be delivered via an implantable device. Design of such a device is an art-recognized process, with, e.g., synthetic implant design described in, e.g., U.S. Pat. No.6,969,400, the contents of which are incorporated in their entirety by reference herein. [0690] Antisense oligomers can be introduced into cells using art-recognized techniques (e.g., transfection, electroporation, fusion, liposomes, colloidal polymeric particles, and viral and non- viral vectors as well as other means known in the art). The method of delivery selected will depend at least on the oligomer chemistry, the cells to be treated and the location of the cells and will be apparent to the skilled artisan. For instance, localization can be achieved by liposomes with specific markers on the surface to direct the liposome, direct injection into tissue containing target cells, specific receptor-mediated uptake, or the like. [0691] As known in the art, antisense oligomers may be delivered using, e.g., methods involving liposome-mediated uptake, exosome-mediated uptake, lipid conjugates, polylysine-mediated uptake, nanoparticle-mediated uptake, and receptor-mediated endocytosis, as well as additional non-endocytic modes of delivery, such as microinjection, permeabilization (e.g., streptolysin-O permeabilization, anionic peptide permeabilization), electroporation, and various non-invasive non-endocytic methods of delivery that are known in the art (refer to Dokka and Rojanasakul, Advanced Drug Delivery Reviews 44, 35-49, incorporated by reference in its entirety). [0692] The antisense oligomers may be administered in any convenient vehicle or carrier which is physiologically and/or pharmaceutically acceptable. Such a composition may include any of a variety of standard pharmaceutically acceptable carriers employed by those of ordinary skill in the art. Examples include, but are not limited to, saline, phosphate-buffered saline (PBS), water, aqueous ethanol, emulsions, such as oil/water emulsions or triglyceride emulsions, tablets, and capsules. The choice of a suitable physiologically acceptable carrier will vary dependent upon the chosen mode of administration. “Pharmaceutically acceptable carrier” is intended to include any and all solvents, dispersion media, coatings, antibacterial and antifungal agents, isotonic and absorption delaying agents, and the like, compatible with pharmaceutical administration. The use of such media and agents for pharmaceutically active substances is well known in the art. Except insofar as any conventional media or agent is incompatible with the active compound, use thereof in the compositions is contemplated. Supplementary active compounds can also be incorporated into the compositions. [0693] The compounds (e.g., antisense oligomers) of the present disclosure may generally be utilized as the free acid or free base. Alternatively, the compounds of this disclosure may be used in the form of acid or base addition salts. Acid addition salts of the free amino compounds of the present disclosure may be prepared by methods well known in the art and may be formed
from organic and inorganic acids. Suitable organic acids include maleic, fumaric, benzoic, ascorbic, succinic, methanesulfonic, acetic, trifluoroacetic, oxalic, propionic, tartaric, salicylic, citric, gluconic, lactic, mandelic, cinnamic, aspartic, stearic, palmitic, glycolic, glutamic, and benzenesulfonic acids. [0694] Suitable inorganic acids include hydrochloric, hydrobromic, sulfuric, phosphoric, and nitric acids. Base addition salts included those salts that form with the carboxylate anion and include salts formed with organic and inorganic cations such as those chosen from the alkali and alkaline earth metals (for example, lithium, sodium, potassium, magnesium, barium and calcium), as well as the ammonium ion and substituted derivatives thereof (for example, dibenzylammonium, benzylammonium, 2-hydroxyethylammonium, and the like). Thus, the term “pharmaceutically acceptable salt” is intended to encompass any and all acceptable salt forms. [0695] In addition, prodrugs are also included within the context of this disclosure. Prodrugs are any covalently bonded carriers that release a compound in vivo when such prodrug is administered to a patient. Prodrugs are generally prepared by modifying functional groups in a way such that the modification is cleaved, either by routine manipulation or in vivo, yielding the parent compound. Prodrugs include, for example, compounds of this disclosure wherein hydroxy, amine, or sulfhydryl groups are bonded to any group that, when administered to a patient, cleaves to form the hydroxy, amine, or sulfhydryl groups. Thus, representative examples of prodrugs include (but are not limited to) acetate, formate, and benzoate derivatives of alcohol and amine functional groups of the antisense oligomers of the disclosure. Further, in the case of a carboxylic acid (-COOH), esters may be employed, such as methyl esters, ethyl esters, and the like. [0696] In some instances, liposomes may be employed to facilitate uptake of the antisense oligomer into cells (see, e.g., Williams, S.A., Leukemia 10(12):1980-1989, 1996; Lappalainen et al., Antiviral Res.23:119, 1994; Uhlmann et al., antisense oligomers: a new therapeutic principle, Chemical Reviews, Volume 90, No.4, 25 pages 544-584, 1990; Gregoriadis, G., Chapter 14, Liposomes, Drug Carriers in Biology and Medicine, pp.287-341, Academic Press, 1979). Hydrogels may also be used as vehicles for antisense oligomer administration, for example, as described in WO 93/01286. Alternatively, the oligomers may be administered in microspheres or microparticles. (See, e.g., Wu, G.Y. and Wu, C.H., J. Biol. Chem.262:4429- 4432, 301987). Alternatively, the use of gas-filled microbubbles complexed with the antisense oligomers can enhance delivery to target tissues, as described in US Patent No.6,245,747. Sustained-release compositions may also be used. These may include semipermeable polymeric matrices in the form of shaped articles such as films or microcapsules.
[0697] In one embodiment, the antisense oligomer is administered to a mammalian subject, e.g., human or domestic animal, exhibiting the symptoms of a kidney disease, in a suitable pharmaceutical carrier. In one aspect of the method, the subject is a human subject, e.g., a patient diagnosed as having a centronuclear myopathy. In one embodiment, the antisense oligomer is contained in a pharmaceutically acceptable carrier and is delivered orally. In another embodiment, the oligomer is contained in a pharmaceutically acceptable carrier and is delivered intravenously (i.v.). [0698] In one embodiment, the antisense compound is administered in an amount and manner effective to result in a peak blood concentration of at least 200-400 nM antisense oligomer. Typically, one or more doses of antisense oligomer are administered, generally at regular intervals, for a period of about one to two weeks. Doses for oral administration are from about 1- 1000 mg oligomer per 70 kg. In some cases, doses of greater than 1000 mg oligomer/patient may be necessary. For i.v. administration, doses are from about 0.5 mg to 1000 mg oligomer per 70 kg. The antisense oligomer may be administered at regular intervals for a short time period, e.g., daily for two weeks or less. However, in some cases the oligomer is administered intermittently over a longer period of time. Administration may be followed by, or concurrent with, administration of an antibiotic or other therapeutic treatment. The treatment regimen may be adjusted (dose, frequency, route, etc.) as indicated, based on the results of immunoassays, other biochemical tests, and physiological examination of the subject under treatment. [0699] In certain embodiments, the method is an in vitro method. In certain other embodiments, the method is an in vivo method. [0700] In certain embodiments, the host cell is a mammalian cell. In certain embodiments, the host cell is a non-human primate cell. In certain embodiments, the host cell is a human cell. [0701] In certain embodiments, the host cell is a naturally occurring cell. In certain other embodiments, the host cell is an engineered cell. [0702] In certain embodiments, the antisense oligomer is administered to a mammalian subject, e.g., a human or a laboratory or domestic animal, in a suitable pharmaceutical carrier. [0703] In certain embodiments, the antisense oligomer is administered to a mammalian subject, e.g., a human or laboratory or domestic animal, together with an additional agent. The antisense oligomer and the additional agent can be administered simultaneously or sequentially, via the same or different routes and/or sites of administration. In certain embodiments, the antisense oligomer and the additional agent can be co-formulated and administered together. In certain embodiments, the antisense oligomer and the additional agent can be provided together in a kit. [0704] In one embodiment, the antisense oligomer, contained in a pharmaceutically acceptable carrier, is delivered orally.
[0705] In one embodiment, the antisense oligomer, contained in a pharmaceutically acceptable carrier, is delivered intravenously (i.v.). [0706] Additional routes of administration, e.g., subcutaneous, intraperitoneal, and pulmonary, are also contemplated herein. [0707] In another application of the method, the subject is a livestock animal, e.g., a pig, cow, or goat, etc., and the treatment is either prophylactic or therapeutic. Also contemplated is, in a method of feeding livestock with a food substance, an improvement in which the food substance is supplemented with an effective amount of an antisense oligomer composition as described above. [0708] In an embodiment, the antisense oligomer is administered in an amount and manner effective to result in a peak blood concentration of at least 200 nM antisense oligomer. In one embodiment, the antisense oligomer is administered in an amount and manner effective to result in a peak plasma concentration of at least 200 nM antisense oligomer. In one embodiment, the antisense oligomer is administered in an amount and manner effective to result in a peak serum concentration of at least 200 nM antisense oligomer. [0709] In an embodiment, the antisense oligomer is administered in an amount and manner effective to result in a peak blood concentration of at least 400 nM antisense oligomer. In one embodiment, the antisense oligomer is administered in an amount and manner effective to result in a peak plasma concentration of at least 400 nM antisense oligomer. In one embodiment, the antisense oligomer is administered in an amount and manner effective to result in a peak serum concentration of at least 400 nM antisense oligomer. [0710] Typically, one or more doses of the antisense oligomer are administered, generally at regular intervals, for a period of about one to two weeks. Doses for oral administration are from about 0.01-15 mg antisense oligomer per kg body weight. In some cases, doses of greater than 15 mg antisense oligomer /kg may be necessary. For i.v. administration, doses are from about 0.005 mg to 15 mg antisense oligomer per kg body weight. The antisense oligomer may be administered at regular intervals for a short time period, e.g., daily for two weeks or less. However, in some cases, the antisense oligomer is administered intermittently over a longer period of time. Administration may be followed by or accompanied by administration of an antibiotic or other therapeutic treatment. The treatment regimen may be adjusted (dose, frequency, route, etc.) as indicated, based on the results of immunoassays, other biochemical tests, and physiological examination of the subject under treatment. [0711] An effective in vivo treatment regimen using the antisense oligomer may vary according to the duration, dose, frequency, and route of administration, as well as the condition of the subject under treatment (i.e., prophylactic administration versus administration in response to
localized or systemic infection). Accordingly, such in vivo therapy will often require monitoring by tests under treatment, and corresponding adjustments in the dose or treatment regimen, to achieve an optimal therapeutic outcome. [0712] In some embodiments, the antisense oligomer is actively taken up by mammalian cells. In further embodiments, the antisense oligomer can be conjugated to a transport moiety (e.g., transport peptide) as described herein to facilitate such uptake. INCORPORATION BY REFERENCE [0713] All publications, patents, and patent applications mentioned in this specification are herein incorporated by reference to the same extent as if each individual publication, patent, or patent application was specifically and individually indicated to be incorporated by reference. VIII. Additional Embodiments [0714] Additional embodiments include embodiments 1 to 429 following. [0715] Embodiment 1. An antisense oligomer comprising a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre-mRNA of human dynamin 2 (DNM2) gene represented by SEQ ID NO: 1, wherein the target region is within an exon internal region, an intron internal region, an intron-exon junction, or an exon-intron junction of the human DNM2 gene pre-mRNA. [0716] Embodiment 2. The antisense oligomer of embodiment 1, wherein the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 6 (SEQ ID NO: 3), exon 7 (SEQ ID NO: 4), exon 8 (SEQ ID NO: 5), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), exon 13 (SEQ ID NO: 8), exon 16 (SEQ ID NO: 9), and exon 19 (SEQ ID NO: 10). [0717] Embodiment 3. The antisense oligomer of embodiment 1 or 2, wherein the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), and exon 13 (SEQ ID NO: 8). [0718] Embodiment 4. The antisense oligomer of embodiment 1, wherein the target region is within an intron-exon junction or an exon-intron junction region of the human DNM2 gene pre- mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 5/exon 6 junction, exon 6/intron 6 junction, intron 6/exon 7 junction, exon 7/intron 7 junction, intron 7/exon 8 junction, exon 8/intron 8 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, exon 13/intron 13 junction, intron 15/exon 16 junction, exon 16/intron 16 junction, intron 18/exon 19 junction, and exon 19/intron 19 junction. [0719] Embodiment 5. The antisense oligomer of embodiment 1 or 4, wherein the target region is within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA
selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, and exon 13/intron 13 junction. [0720] Embodiment 6. The antisense oligomer of embodiment 1, wherein the target region is within an intron internal region of the human DNM2 gene pre-mRNA selected from intron 2 (SEQ ID NO: 11) and intron 12 (SEQ ID NO: 12). [0721] Embodiment 7. The antisense oligomer of any one of embodiments 1-6, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10- 15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5- 29), DNM2 H2D(-6-30), DNM2 H9A(-12+13), DNM2 H9A(-5+20), DNM2 H9A(+1+25),DNM2 H9A(+6+30), DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), DNM2 H9A(+48+67), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22- 3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13- 12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), DNM2 H9D(+3-22), DNM2 H12A(-13+12), DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), DNM2 H12D(+15-10), DNM2 H12A(+4+28), DNM2
H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(- 23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(- 5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), DNM2 H13A(+9+28), DNM2 H13A(+16+40), DNM2 H13D(+22-3), or DNM2 H13D(+7-18). [0722] Embodiment 8. The antisense oligomer of embodiment 7, wherein the targeting sequence is selected from SEQ ID NOs: 67-326. [0723] Embodiment 9. The antisense oligomer of any one of embodiments 1-6, wherein the target region is within the exon internal region of exon 2 (SEQ ID NO: 2), the intron 1/exon 2 junction, the exon 2/intron 2 junction, or the intron internal region of intron 2 (SEQ ID NO: 11) of the human DNM2 gene pre-mRNA. [0724] Embodiment 10. The antisense oligomer of any one of embodiments 1-3 or 9, wherein the target region of the human DNM2 gene pre-mRNA is within the 21st nucleotide to the 47th nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2). [0725] Embodiment 11. The antisense oligomer of embodiment 10, wherein the target region is a region within SEQ ID NO: 13. [0726] Embodiment 12. The antisense oligomer of any one of embodiments 1-3 or 9, wherein the target region of the human DNM2 gene pre-mRNA is within the 23rd nucleotide to the 51st nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2). [0727] Embodiment 13. The antisense oligomer of embodiment 12, wherein the target region is a region within SEQ ID NO: 14.
[0728] Embodiment 14. The antisense oligomer of any one of embodiments 1-3 or 9, wherein the target region of the human DNM2 gene pre-mRNA is within the 46th nucleotide to the 72nd nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2). [0729] Embodiment 15. The antisense oligomer of embodiment 14, wherein the target region is a region within SEQ ID NO: 15. [0730] Embodiment 16. The antisense oligomer of any one of embodiments 1, 4-5, or 9, wherein the target region of the human DNM2 gene pre-mRNA is within the 13th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 24th nucleotide of intron 2 measured from the 3′ end of exon 2. [0731] Embodiment 17. The antisense oligomer of embodiment 16, wherein the target region is a region within SEQ ID NO: 16. [0732] Embodiment 18. The antisense oligomer of any one of embodiments 1, 4-5, 9, or 16-17, wherein the target region of the human DNM2 gene pre-mRNA is within the 13th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 14th nucleotide of intron 2 measured from the 3′ end of exon 2. [0733] Embodiment 19. The antisense oligomer of embodiment 18, wherein the target region is a region within SEQ ID NO: 17. [0734] Embodiment 20. The antisense oligomer of any one of embodiments 1, 4-5, 9, or 16-17, wherein the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 19th nucleotide of intron 2 measured from the 3′ end of exon 2. [0735] Embodiment 21. The antisense oligomer of embodiment 20, wherein the target region is a region within SEQ ID NO: 18. [0736] Embodiment 22. The antisense oligomer of any one of embodiments 1, 4-5, 9, 16-17, or 20-21, wherein the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 18th nucleotide of intron 2 measured from the 3′ end of exon 2. [0737] Embodiment 23. The antisense oligomer of embodiment 22, wherein the target region is a region within SEQ ID NO: 19. [0738] Embodiment 24. The antisense oligomer of any one of embodiments 1, 4-5, 9, or 16-17, wherein the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23rd nucleotide of intron 2 measured from the 3′ end of exon 2. [0739] Embodiment 25. The antisense oligomer of embodiment 24, wherein the target region is a region within SEQ ID NO: 20.
[0740] Embodiment 26. The antisense oligomer of any one of embodiments 1, 4-5, 9, 16-17, or 24-25, wherein the target region of the human DNM2 gene pre-mRNA is within the 4th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23rd nucleotide of intron 2 measured from the 3′ end of exon 2. [0741] Embodiment 27. The antisense oligomer of embodiment 26, wherein the target region is a region within SEQ ID NO: 21. [0742] Embodiment 28. The antisense oligomer of any one of embodiments 1, 6, or 9, wherein the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 30th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2). [0743] Embodiment 29. The antisense oligomer of embodiment 28, wherein the target region is a region within SEQ ID NO: 22. [0744] Embodiment 30. The antisense oligomer of any one of embodiments 1, 6, 9, or 28-29, wherein the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 29th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2). [0745] Embodiment 31. The antisense oligomer of embodiment 30, wherein the target region is a region within SEQ ID NO: 23. [0746] Embodiment 32. The antisense oligomer of any one of embodiments 1-31, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10- 15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5- 29), or DNM2 H2D(-6-30). [0747] Embodiment 33. The antisense oligomer of embodiment 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2- 23) , DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30). [0748] Embodiment 34. The antisense oligomer of any one of embodiments 1-3, 7, 9-11, or 32, wherein the target region is DNM2 H2A(+21+45) or DNM2 H2A(+23+47). [0749] Embodiment 35. The antisense oligomer of any one of embodiments 1-3, 7, 9, 12-13, or 32, wherein the target region is DNM2 H2A(+23+47) or DNM2 H2A(+27+51).
[0750] Embodiment 36. The antisense oligomer of any one of embodiments 1-3, 7, 9, 14-15, or 32, wherein the target region is DNM2 H2A(+46+70) or DNM2 H2A(+48+72). [0751] Embodiment 37. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, or 32, wherein the target region is DNM2 H2D(+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11- 14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), or DNM2 H2D(+1-24). [0752] Embodiment 38. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-19, 32, or 37, wherein the target region is DNM2 H2D (+13-12), DNM2 H2D(+12-13), or DNM2 H2D(+11-14). [0753] Embodiment 39. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-21, 32, or 37, wherein the target region is DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), or DNM2 H2D(+6-19). [0754] Embodiment 40. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-23, 32, 37, or 39, wherein the target region is DNM2 H2D(+9-16), DNM2 H2D(+8-17), or DNM2 H2D(+7-18). [0755] Embodiment 41. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-23, 32, 35, 37, or 39-40, wherein the target region is DNM2 H2D(+8-17). [0756] Embodiment 42. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-25, 32, or 37, wherein the target region is DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23). [0757] Embodiment 43. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42, wherein the target region is DNM2 H2D(+4-21), DNM2 H2D(+3-22), or DNM2 H2D(+2-23). [0758] Embodiment 44. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42-43, wherein the target region is DNM2 H2D(+3-22). [0759] Embodiment 45. The antisense oligomer of any one of embodiments 1, 6-7, 9, 28-29, or 32, wherein the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30). [0760] Embodiment 46. The antisense oligomer of any one of embodiments 1, 6-7, 9, 28-32, or 45, wherein the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), or DNM2 H2D(-5-29). [0761] Embodiment 47. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15).
[0762] Embodiment 48. The antisense oligomer of any one of embodiments 1, 4-5, or 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(-7+17). [0763] Embodiment 49. The antisense oligomer of any one of embodiments 1-3, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+3+27). [0764] Embodiment 50. The antisense oligomer of any one of embodiments 1-3, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+15+37). [0765] Embodiment 51. The antisense oligomer of any one of embodiments 1-3, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+37+61). [0766] Embodiment 52. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2D(+20-5). [0767] Embodiment 53. The antisense oligomer of any one of embodiments 1-32, wherein the targeting sequence is selected from SEQ ID NOs: 67-96. [0768] Embodiment 54. The antisense oligomer of any one of embodiments 1, 7, or 32-33, wherein the targeting sequence is selected from SEQ ID NOs: 78-96. [0769] Embodiment 55. The antisense oligomer of any one of embodiments 1-3, 7, 9-11, 32, or 34, wherein the targeting sequence is selected from SEQ ID NOs: 71-72. [0770] Embodiment 56. The antisense oligomer of any one of embodiments 1-3, 7, 9, 12-13, 32, or 35, wherein the targeting sequence is selected from SEQ ID NOs: 72-73. [0771] Embodiment 57. The antisense oligomer of any one of embodiments 1-3, 7, 9, 14-15, 32, or 36, wherein the targeting sequence is selected from SEQ ID NOs: 75-76. [0772] Embodiment 58. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 32, or 37, wherein the targeting sequence is selected from SEQ ID NOs: 78-90. [0773] Embodiment 59. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-19, 32, or 37-38, wherein the targeting sequence is selected from SEQ ID NOs: 78-80. [0774] Embodiment 60. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-21, 32, 37, or 39, wherein the targeting sequence is selected from SEQ ID NOs: 81-85. [0775] Embodiment 61. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-23, 32, 37, or 39-40, wherein the targeting sequence is selected from SEQ ID NOs: 82-84. [0776] Embodiment 62. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 20-23, 32, 35, 37, or 39-41, wherein the targeting sequence is SEQ ID NO: 83. [0777] Embodiment 63. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-25, 32, 37, or 42, wherein the targeting sequence is selected from SEQ ID NOs: 85-89. [0778] Embodiment 64. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42-43, wherein the targeting sequence is selected from SEQ ID NOs: 87-89.
[0779] Embodiment 65. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42-44, wherein the targeting sequence is SEQ ID NO: 88. [0780] Embodiment 66. The antisense oligomer of any one of embodiments 1, 6-7, 9, 28-29, 32, or 45, wherein the targeting sequence is selected from SEQ ID NOs: 91-96. [0781] Embodiment 67. The antisense oligomer of any one of embodiments 1, 6-7, 9, 28-32, or 45-46, wherein the targeting sequence is selected from SEQ ID NOs: 91-95. [0782] Embodiment 68. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 32, or 47, wherein the targeting sequence is SEQ ID NO: 67. [0783] Embodiment 69. The antisense oligomer of any one of embodiments 1, 4-5, or 7, 9, 32, or 48, wherein the targeting sequence is SEQ ID NO: 68. [0784] Embodiment 70. The antisense oligomer of any one of embodiments 1-3, 7, 9, 32, or 49, wherein the targeting sequence is SEQ ID NO: 69. [0785] Embodiment 71. The antisense oligomer of any one of embodiments 1-3, 7, 9, 32, or 50, wherein the targeting sequence is SEQ ID NO: 70. [0786] Embodiment 72. The antisense oligomer of any one of embodiments 1-3, 7, 9, 32, or 51, wherein the targeting sequence is SEQ ID NO: 74. [0787] Embodiment 73. The antisense oligomer of any one of embodiments 1, 4-5, 7, 9, 32, or 52, wherein the targeting sequence is SEQ ID NO: 77. [0788] Embodiment 74. The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 9 (SEQ ID NO: 6), the intron 8/exon 9 junction, or the exon 9/intron 9 junction of the human DNM2 gene pre-mRNA. [0789] Embodiment 75. The antisense oligomer of any one of embodiments 1-3 or 74, wherein the target region of the human DNM2 gene pre-mRNA is within the 36th nucleotide to the 61st nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6). [0790] Embodiment 76. The antisense oligomer of embodiment 75, wherein the target region is a region within SEQ ID NO: 24. [0791] Embodiment 77. The antisense oligomer of any one of embodiments 1-3 or 74, wherein the target region of the human DNM2 gene pre-mRNA is within the 42nd nucleotide to the 68th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6). [0792] Embodiment 78. The antisense oligomer of embodiment 77, wherein the target region is a region within SEQ ID NO: 25. [0793] Embodiment 79. The antisense oligomer of any one of embodiments 1-3, 74, or 77-78, wherein the target region of the human DNM2 gene pre-mRNA is within the 43rd nucleotide to the 68th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6).
[0794] Embodiment 80. The antisense oligomer of embodiment 79, wherein the target region is a region within SEQ ID NO: 26. [0795] Embodiment 81. The antisense oligomer of any one of embodiments 1, 4-5, or 74, wherein the target region of the human DNM2 gene pre-mRNA is within the 24th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15th nucleotide of intron 9 measured from the 3′ end of exon 9. [0796] Embodiment 82. The antisense oligomer of embodiment 81, wherein the target region is a region within SEQ ID NO: 27. [0797] Embodiment 83. The antisense oligomer of any one of embodiments 1, 4-5, 74, or 81-82, wherein the target region of the human DNM2 gene pre-mRNA is within the 24th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 5th nucleotide of intron 9 measured from the 3′ end of exon 9. [0798] Embodiment 84. The antisense oligomer of embodiment 83, wherein the target region is a region within SEQ ID NO: 28. [0799] Embodiment 85. The antisense oligomer of any one of embodiments 1, 4-5, 74, or 81-82, wherein the target region of the human DNM2 gene pre-mRNA is within the 19th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 10th nucleotide of intron 9 measured from the 3′ end of exon 9. [0800] Embodiment 86. The antisense oligomer of embodiment 85, wherein the target region is a region within SEQ ID NO: 29. [0801] Embodiment 87. The antisense oligomer of any one of embodiments 1, 4-5, 74, or 81-82, wherein the target region of the human DNM2 gene pre-mRNA is within the 14th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15th nucleotide of intron 9 measured from the 3′ end of exon 9. [0802] Embodiment 88. The antisense oligomer of embodiment 87, wherein the target region is a region within SEQ ID NO: 30. [0803] Embodiment 89. The antisense oligomer of any one of embodiments 1, 4-5, or 74, wherein the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22nd nucleotide of intron 9 measured from the 3′ end of exon 9. [0804] Embodiment 90. The antisense oligomer of embodiment 89, wherein the target region is a region within SEQ ID NO: 31. [0805] Embodiment 91. The antisense oligomer of any one of embodiments 1, 4-5, 74, or 89-90, wherein the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of
exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 20th nucleotide of intron 9 measured from the 3′ end of exon 9. [0806] Embodiment 92. The antisense oligomer of embodiment 91, wherein the target region is a region within SEQ ID NO: 32. [0807] Embodiment 93. The antisense oligomer of any one of embodiments 1, 4-5, 74, or 89-92, wherein the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 18th nucleotide of intron 9 measured from the 3′ end of exon 9. [0808] Embodiment 94. The antisense oligomer of embodiment 93, wherein the target region is a region within SEQ ID NO: 33. [0809] Embodiment 95. The antisense oligomer of any one of embodiments 1, 4-5, 74, or 89-90, wherein the target region of the human DNM2 gene pre-mRNA is within the 8th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 21st nucleotide of intron 9 measured from the 3′ end of exon 9. [0810] Embodiment 96. The antisense oligomer of embodiment 95, wherein the target region is a region within SEQ ID NO: 34. [0811] Embodiment 97. The antisense oligomer of any one of embodiments 1, 4-5, 74, or 89-90, wherein the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22nd nucleotide of intron 9 measured from the 3′ end of exon 9. [0812] Embodiment 98. The antisense oligomer of embodiment 97, wherein the target region is a region within SEQ ID NO: 35. [0813] Embodiment 99. The antisense oligomer of any one of embodiments 1-5, 7, or 74-98, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13), DNM2 H9A(-5+20), DNM2 H9A(+1+25),DNM2 H9A(+6+30), DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), DNM2 H9A(+48+67), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11- 14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17),
DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4- 17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22). [0814] Embodiment 100. The antisense oligomer of embodiment 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+43+67), DNM2 H9A(+42+66), DNM2 H9A(+44+68), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12- 13), DNM2 H9D(+11-14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+6-19), DNM2 H9D(+5-20), DNM2 H9D(+4-21), or DNM2 H9D(+3-22). [0815] Embodiment 101. The antisense oligomer of any one of embodiments 1-5, 7, 74-76, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+36+60) or DNM2 H9A(+37+61). [0816] Embodiment 102. The antisense oligomer of any one of embodiments 1-3, 7, 74, 77-78, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), or DNM2 H9A(+48+67). [0817] Embodiment 103. The antisense oligomer of any one of embodiments 1-3, 7, 74, 77-80, 99, or 102, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), or DNM2 H9A(+44+68). [0818] Embodiment 104. The antisense oligomer of any one of embodiments 1-3, 7, 74, 77-80, 99, or 102-103, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+67). [0819] Embodiment 105. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 82, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), or DNM2 H9D(+10-15). [0820] Embodiment 106. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 84, 99, or 105, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), or DNM2 H9D(+20-5).
[0821] Embodiment 107. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 82, 85-86, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), or DNM2 H9D(+15- 10). [0822] Embodiment 108. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 82, 87-88, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), or DNM2 H9D(+10-15). [0823] Embodiment 109. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 90, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8- 17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22). [0824] Embodiment 110. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 92, 99, or 109, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6- 17), DNM2 H9D(+5-17), or DNM2 H9D(+5-20). [0825] Embodiment 111. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 94, 99, or 109-110, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), or DNM2 H9D(+7-18). [0826] Embodiment 112. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 90, 95-96, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), or DNM2 H9D(+4-21). [0827] Embodiment 113. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 96, 99, or 109-112, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17). [0828] Embodiment 114. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 90, 97-99, or 109, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22).
[0829] Embodiment 115. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13). [0830] Embodiment 116. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-5+20). [0831] Embodiment 117. The antisense oligomer of any one of embodiments 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+1+25). [0832] Embodiment 118. The antisense oligomer of any one of embodiments 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+6+30). [0833] Embodiment 119. The antisense oligomer of any one of embodiments 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40). [0834] Embodiment 120. The antisense oligomer of any one of embodiments 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+25+49). [0835] Embodiment 121. The antisense oligomer of any one of embodiments 1-5, 7, or 74-99, wherein the targeting sequence is selected from SEQ ID NOs: 158-209. [0836] Embodiment 122. The antisense oligomer of any one of embodiments 1, 7, or 99-100, wherein the targeting sequence is selected from SEQ ID NOs: 162-166, 172, 174, 179-194, 199- 200, 205, 207, and 209. [0837] Embodiment 123. The antisense oligomer of any one of embodiments 1-5, 7, 74-76, 99, or 101, wherein the targeting sequence is selected from SEQ ID NOs: 164-165. [0838] Embodiment 124. The antisense oligomer of any one of embodiments 1-3, 7, 74, 77-78, 99, or 102, wherein the targeting sequence is selected from SEQ ID NOs: 166-178. [0839] Embodiment 125. The antisense oligomer of any one of embodiments 1-3, 7, 74, 77-80, 99, or 102-103, wherein the targeting sequence is selected from SEQ ID NOs: 167-174. [0840] Embodiment 126. The antisense oligomer of any one of embodiments 1-3, 7, 74, 77-80, 99, or 102-104, wherein the targeting sequence is SEQ ID NO: 172. [0841] Embodiment 127. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 82, 99, or 105, wherein the targeting sequence is selected from SEQ ID NOs: 179-193. [0842] Embodiment 128. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 84, 99, or 105-106, wherein the targeting sequence is selected from SEQ ID NOs: 179-183. [0843] Embodiment 129. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 82, 85-86, 99, or 107, wherein the targeting sequence is selected from SEQ ID NOs: 184-188. [0844] Embodiment 130. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 81- 82, 87-88, 99, or 108, wherein the targeting sequence is selected from SEQ ID NOs: 189-193. [0845] Embodiment 131. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 90, 99, or 109, wherein the targeting sequence is selected from SEQ ID NOs: 194-209.
[0846] Embodiment 132. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 92, 99, or 109-110, wherein the targeting sequence is selected from SEQ ID NOs: 194-205. [0847] Embodiment 133. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 94, 99, or 109-111, wherein the targeting sequence is selected from SEQ ID NOs: 194-200. [0848] Embodiment 134. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 90, 95-96, 99, or 112, wherein the targeting sequence is selected from SEQ ID NOs: 199-207. [0849] Embodiment 135. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 96, 99, or 109-113, wherein the targeting sequence is SEQ ID NO: 199. [0850] Embodiment 136. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 89- 90, 97-99, 109, or 114, wherein the targeting sequence is selected from SEQ ID NOs: 202-209. [0851] Embodiment 137. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 99, or 115, wherein the targeting sequence is SEQ ID NO: 158. [0852] Embodiment 138. The antisense oligomer of any one of embodiments 1, 4-5, 7, 74, 99, or 116, wherein the targeting sequence is SEQ ID NO: 159. [0853] Embodiment 139. The antisense oligomer of any one of embodiments 1-3, 7, 74, 99, or 117, wherein the targeting sequence is SEQ ID NO: 160. [0854] Embodiment 140. The antisense oligomer of any one of embodiments 1-3, 7, 74, 99, or 118, wherein the targeting sequence is SEQ ID NO: 161. [0855] Embodiment 141. The antisense oligomer of any one of embodiments 1-3, 7, 74, 99, or 119, wherein the targeting sequence is SEQ ID NO: 162. [0856] Embodiment 142. The antisense oligomer of any one of embodiments 1-3, 7, 74, 99, or 120, wherein the targeting sequence is SEQ ID NO: 163. [0857] Embodiment 143. The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 12 (SEQ ID NO: 7), the intron 11/exon 12 junction, or the exon 12/intron 12 junction of the human DNM2 gene pre-mRNA. [0858] Embodiment 144. The antisense oligomer of any one of embodiments 1-3 or 143, wherein the target region of the human DNM2 gene pre-mRNA is within the 4th nucleotide to the 31st nucleotide measured of exon 12 from the 5′ end of exon 12 (SEQ ID NO: 7). [0859] Embodiment 145. The antisense oligomer of embodiment 144, wherein the target region is a region within SEQ ID NO: 36. [0860] Embodiment 146. The antisense oligomer of any one of embodiments 1-3 or 143, wherein the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 44th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). [0861] Embodiment 147. The antisense oligomer of embodiment 146, wherein the target region is a region within SEQ ID NO: 37.
[0862] Embodiment 148. The antisense oligomer of any one of embodiments 1-3, 143, or 146- 147, wherein the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 38th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). [0863] Embodiment 149. The antisense oligomer of embodiment 148, wherein the target region is a region within SEQ ID NO: 38. [0864] Embodiment 150. The antisense oligomer of any one of embodiments 1-3, 143, or 146- 147, wherein the target region of the human DNM2 gene pre-mRNA is within the 16th nucleotide to the 44th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). [0865] Embodiment 151. The antisense oligomer of embodiment 150, wherein the target region is a region within SEQ ID NO: 39. [0866] Embodiment 152. The antisense oligomer of any one of embodiments 1-3, 143, or 146- 151, wherein the target region of the human DNM2 gene pre-mRNA is within the 14th nucleotide to the 42nd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). [0867] Embodiment 153. The antisense oligomer of embodiment 152, wherein the target region is a region within SEQ ID NO: 40. [0868] Embodiment 154. The antisense oligomer of any one of embodiments 1-3 or 143, wherein the target region of the human DNM2 gene pre-mRNA is within the 25th nucleotide to the 54th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). [0869] Embodiment 155. The antisense oligomer of embodiment 154, wherein the target region is a region within SEQ ID NO: 41. [0870] Embodiment 156. The antisense oligomer of any one of embodiments 1-3, 143, or 154- 155, wherein the target region of the human DNM2 gene pre-mRNA is within the 25th nucleotide to the 53rd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). [0871] Embodiment 157. The antisense oligomer of embodiment 156, wherein the target region is a region within SEQ ID NO: 42. [0872] Embodiment 158. The antisense oligomer of any one of embodiments 1-3, 143, or 154- 155, wherein the target region of the human DNM2 gene pre-mRNA is within the 26th nucleotide to the 54th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). [0873] Embodiment 159. The antisense oligomer of embodiment 158, wherein the target region is a region within SEQ ID NO: 43. [0874] Embodiment 160. The antisense oligomer of any one of embodiments 1-3 or 143, wherein the target region of the human DNM2 gene pre-mRNA is within the 36th nucleotide to the 61st nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7). [0875] Embodiment 161. The antisense oligomer of embodiment 160, wherein the target region is a region within SEQ ID NO: 44.
[0876] Embodiment 162. The antisense oligomer of any one of embodiments 1, 4-5, or 143, wherein the target region of the human DNM2 gene pre-mRNA is within the 22nd nucleotide of exon 12 measured from the 3′ end of exon 12 (SEQ ID NO: 7) to the 5th nucleotide of intron 12 measured from the 3′ end of exon 12. [0877] Embodiment 163. The antisense oligomer of embodiment 162, wherein the target region is a region within SEQ ID NO: 45. [0878] Embodiment 164. The antisense oligomer of any one of embodiments 1-5, 7, or 143-163, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-13+12), DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), or DNM2 H12D(+15-10). [0879] Embodiment 165. The antisense oligomer of any one of embodiment 164, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+49), DNM2 H12A(+29+53), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), or DNM2 H12D(+15-10). [0880] Embodiment 166. The antisense oligomer of any one of embodiments 1-3, 7, 143-145, or 164, wherein the target region is DNM2 H12A(+4+28) or DNM2 H12A(+7+31). [0881] Embodiment 167. The antisense oligomer of any one of embodiments 1-3, 7, 143, 146- 147, or 164, wherein the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44). [0882] Embodiment 168. The antisense oligomer of any one of embodiments 1-3, 7, 143, 146- 149, 164, or 167, wherein the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), or DNM2 H12A(+14+38).
[0883] Embodiment 169. The antisense oligomer of any one of embodiments 1-3, 7, 143, 146- 147, 150-151, 164, or 167, wherein the target region is DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), or DNM2 H12A(+20+44). [0884] Embodiment 170. The antisense oligomer of any one of embodiments 1-3, 7, 143, 146- 153, 164, or 167-169, wherein the target region is DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), or DNM2 H12A(+18+42). [0885] Embodiment 171. The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 155, or 164, wherein the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54). [0886] Embodiment 172. The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 157, 164, or 171, wherein the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), or DNM2 H12A(+29+53). [0887] Embodiment 173. The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 155, 158-159, 164, or 171, wherein the target region is DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54). [0888] Embodiment 174. The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 155, 158-159, 164, 171, or 173, wherein the target region is DNM2 H12A(+25+49). [0889] Embodiment 175. The antisense oligomer of any one of embodiments 1-3, 7, 143, 160- 161, or 164, wherein the target region is DNM2 H12A(+36+60) or DNM2 H12A(+37+61). [0890] Embodiment 176. The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, or 162-164, wherein the target region is DNM2 H12D(+22-3) or DNM2 H12D(+20-5). [0891] Embodiment 177. The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, or 164, wherein the target region is DNM2 H12A(-13+12). [0892] Embodiment 178. The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, or 164, wherein the target region is DNM2 H12A(-3+22). [0893] Embodiment 179. The antisense oligomer of any one of embodiments 1-3, 7, 143, or 164, wherein the target region is DNM2 H12A(+43+67). [0894] Embodiment 180. The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, or 164, wherein the target region is DNM2 H12D(+15-10).
[0895] Embodiment 181. The antisense oligomer of any one of embodiments 1-5, 7, or 143-164, wherein the targeting sequence is selected from SEQ ID NOs: 210-243. [0896] Embodiment 182. The antisense oligomer of any one of embodiments 1, 7, or 164-165, wherein the targeting sequence is selected from SEQ ID NOs: 211-224, 230, 235, 237-238, and 240-243. [0897] Embodiment 183. The antisense oligomer of any one of embodiments 1-3, 7, 143-145, 164, or 166, wherein the targeting sequence is selected from SEQ ID NOs: 212-213. [0898] Embodiment 184. The antisense oligomer of any one of embodiments 1-3, 7, 143, 146- 147, 164, or 167, wherein the targeting sequence is selected from SEQ ID NOs: 214-224. Embodiment 185. The antisense oligomer of any one of embodiments 1-3, 7, 143, 146-149, 164, or 167-168, wherein the targeting sequence is selected from SEQ ID NOs: 214-218. [0899] Embodiment 186. The antisense oligomer of any one of embodiments 1-3, 7, 143, 146- 147, 150-151, 164, or 169, wherein the targeting sequence is selected from SEQ ID NOs: 220- 224. [0900] Embodiment 187. The antisense oligomer of any one of embodiments 1-3, 7, 143, 146- 153, 164, 167-170, wherein the targeting sequence is selected from SEQ ID NOs: 218-222. [0901] Embodiment 188. The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 155, 164, or 171, wherein the targeting sequence is selected from SEQ ID NOs: 225-237. [0902] Embodiment 189. The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 157, 164, or 171-172, wherein the targeting sequence is selected from SEQ ID NOs: 225-235. [0903] Embodiment 190. The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 155, 158-159, 164, or 173, wherein the targeting sequence is selected from SEQ ID NOs: 231- 237 [0904] Embodiment 191. The antisense oligomer of any one of embodiments 1-3, 7, 143, 154- 155, 158-159, 164, or 174, wherein the targeting sequence is SEQ ID NO: 230 [0905] Embodiment 192. The antisense oligomer of any one of embodiments 1-3, 7, 143, 160- 161, 164, or 175, wherein the targeting sequence is selected from SEQ ID NOs: 238-239. [0906] Embodiment 193. The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, 162-164, or 176, wherein the targeting sequence is selected from SEQ ID NOs: 241-242. [0907] Embodiment 194. The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, 164, or 177, wherein the targeting sequence is SEQ ID NO: 210. [0908] Embodiment 195. The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, 164, or 178, wherein the targeting sequence is SEQ ID NO: 211. [0909] Embodiment 196. The antisense oligomer of any one of embodiments 1-3, 7, 143, 164, or 179, wherein the targeting sequence is SEQ ID NO: 240.
[0910] Embodiment 197. The antisense oligomer of any one of embodiments 1, 4-5, 7, 143, 164, or 180, wherein the targeting sequence is SEQ ID NO: 243. [0911] Embodiment 198. The antisense oligomer of any one of embodiments 1-6, wherein the target region is within the exon internal region of exon 13 (SEQ ID NO: 8), the intron 12/exon 13 junction, the exon 13/intron 13 junction, or the intron internal region of intron 12 (SEQ ID NO: 12). [0912] Embodiment 199. The antisense oligomer of any one of embodiments 1, 4-6, or 198, wherein the target region of the human DNM2 gene pre-mRNA is within the 27th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12th nucleotide of exon 13 measured from the 5′ end of exon 13. [0913] Embodiment 200. The antisense oligomer of embodiment 199, wherein the target region is a region within SEQ ID NO: 46. [0914] Embodiment 201. The antisense oligomer of any one of embodiments 1, 4-6, or 198-200, wherein the target region of the human DNM2 gene pre-mRNA is within the 27th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 2nd nucleotide of exon 13 measured from the 5′ end of exon 13. [0915] Embodiment 202. The antisense oligomer of embodiment 201, wherein the target region is a region within SEQ ID NO: 47. [0916] Embodiment 203. The antisense oligomer of any one of embodiments 1, 4-6, or 198-200, wherein the target region of the human DNM2 gene pre-mRNA is within the 22nd nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 7th nucleotide of exon 13 measured from the 5′ end of exon 13. [0917] Embodiment 204. The antisense oligomer of embodiment 203, wherein the target region is a region within SEQ ID NO: 48. [0918] Embodiment 205. The antisense oligomer of any one of embodiments 1, 4-6, or 198-200, wherein the target region of the human DNM2 gene pre-mRNA is within the 17th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12th nucleotide of exon 13 measured from the 5′ end of exon 13. [0919] Embodiment 206. The antisense oligomer of embodiment 205, wherein the target region is a region within SEQ ID NO: 49. [0920] Embodiment 207. The antisense oligomer of any one of embodiments 1, 4-5, or 198, wherein the target region of the human DNM2 gene pre-mRNA is within the 11th nucleotide of intron 12 (SEQ ID NO: 12) measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24th nucleotide of exon 13 measured from the 5′ end of exon 13.
[0921] Embodiment 208. The antisense oligomer of embodiment 207, wherein the target region is a region within SEQ ID NO: 50. [0922] Embodiment 209. The antisense oligomer of any one of embodiments 1, 4-5, 198, or 207-208, wherein the target region of the human DNM2 gene pre-mRNA is within the 11th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 18th nucleotide of exon 13 measured from the 5′ end of exon 13. [0923] Embodiment 210. The antisense oligomer of embodiment 209, wherein the target region is a region within SEQ ID NO: 51. [0924] Embodiment 211. The antisense oligomer of any one of embodiments 1, 4-5, 198, or 207-208, wherein the target region of the human DNM2 gene pre-mRNA is within the 5th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24th nucleotide exon 13 measured from the 5′ end of exon 13. [0925] Embodiment 212. The antisense oligomer of embodiment 211, wherein the target region is a region within SEQ ID NO: 52. [0926] Embodiment 213. The antisense oligomer of any one of embodiments 1, 4-5, 191, or 207-212, wherein the target region of the human DNM2 gene pre-mRNA is within the 8th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 21st nucleotide exon 13 measured from the 5′ end of exon 13. [0927] Embodiment 214. The antisense oligomer of embodiment 213, wherein the target region is a region within SEQ ID NO: 53. [0928] Embodiment 215. The antisense oligomer of any one of embodiments 1-3 or 198, wherein the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). [0929] Embodiment 216. The antisense oligomer of embodiment 215, wherein the target region is a region within SEQ ID NO: 54. [0930] Embodiment 217. The antisense oligomer of any one of embodiments 1-3, 198, or 215- 216, wherein the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 28th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). [0931] Embodiment 218. The antisense oligomer of embodiment 217, wherein the target region is a region within SEQ ID NO: 55. [0932] Embodiment 219. The antisense oligomer of any one of embodiments 1-3, 198, or 215- 216, wherein the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). [0933] Embodiment 220. The antisense oligomer of embodiment 219, wherein the target region is a region within SEQ ID NO: 56.
[0934] Embodiment 221. The antisense oligomer of any one of embodiments 1-3, 198, or 215- 220, wherein the target region of the human DNM2 gene pre-mRNA is within the 3rd nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8). [0935] Embodiment 222. The antisense oligomer of embodiment 221, wherein the target region is a region within SEQ ID NO: 57. [0936] Embodiment 223. The antisense oligomer of any one of embodiments 1-7 or 198-221, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(- 22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(- 8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), DNM2 H13A(+9+28), DNM2 H13A(+16+40), DNM2 H13D(+22-3), or DNM2 H13D(+7-18). [0937] Embodiment 224. The antisense oligomer of embodiment 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(- 25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(- 11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), or DNM2 H13A(+4+28). [0938] Embodiment 225. The antisense oligomer of any one of embodiments 1, 4-7, 198-200, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(- 18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12).
[0939] Embodiment 226. The antisense oligomer of any one of embodiments 1, 4-7, 198-202, 223, or 225, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27- 3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), or DNM2 H13A(-23+2). [0940] Embodiment 227. The antisense oligomer of any one of embodiments 1, 4-7, 198-200, 203-204, 223, or 225, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), or DNM2 H13A(- 18+7). [0941] Embodiment 228. The antisense oligomer of any one of embodiments 1, 4-7, 198-200, 205-206, 223, or 225, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12). [0942] Embodiment 229. The antisense oligomer of any one of embodiments 1, 4-7, 198, 207- 208, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(- 11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(-1+24). [0943] Embodiment 230. The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 207-210, 223, or 229, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), or DNM2 H13A(-7+18). [0944] Embodiment 231. The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 207-208, 211-212, 223, or 229, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(-1+24). [0945] Embodiment 232. The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 207-214, 223, or 229-231, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), or DNM2 H13A(-4+21). [0946] Embodiment 233. The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 216, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28).
[0947] Embodiment 234. The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 218, 223, or 233, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), or DNM2 H13A(+5+28). [0948] Embodiment 235. The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 216, 219-220, 223, or 233, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28). [0949] Embodiment 236. The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 223, or 228-235, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), or DNM2 H13A(+6+30). [0950] Embodiment 237. The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 223, or 228-236, wherein the target region of the human DNM2 gene pre-mRNA DNM2 H13A(+4+28). [0951] Embodiment 238. The antisense oligomer of any one of embodiments 1-3, 7, 198, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+16+40). [0952] Embodiment 239. The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13D(+22-3). [0953] Embodiment 240. The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13D(+7-18). [0954] Embodiment 241. The antisense oligomer of any one of embodiments 1-7, 198-221, or 223, wherein the targeting sequence is selected from SEQ ID NOs: 244-287. [0955] Embodiment 242. The antisense oligomer of any one of embodiments 1, 7, or 223-224, wherein the targeting sequence is selected from SEQ ID NOs: 244-278. [0956] Embodiment 243. The antisense oligomer of any one of embodiments 1, 4-7, 198-200, 223, or 225, wherein the targeting sequence is selected from SEQ ID NOs: 244-258. [0957] Embodiment 244. The antisense oligomer of any one of embodiments 1, 4-7, 198-202, 223, or 225-226, wherein the targeting sequence is selected from SEQ ID NOs: 244-248. [0958] Embodiment 245. The antisense oligomer of any one of embodiments 1, 4-7, 198-200, 203-204, 223, 225, or 227, wherein the targeting sequence is selected from SEQ ID NOs: 249- 253.
[0959] Embodiment 246. The antisense oligomer of any one of embodiments 1, 4-7, 198-200, 205-206, 223, 225, or 228, wherein the targeting sequence is selected from SEQ ID NOs: 254- 258 [0960] Embodiment 247. The antisense oligomer of any one of embodiments 1, 4-7, 198, 207- 208, 223, or 229, wherein the targeting sequence is selected from SEQ ID NOs: 259-269. [0961] Embodiment 248. The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 207-210, 223, or 229-230, wherein the targeting sequence is selected from SEQ ID NOs: 259- 263. [0962] Embodiment 249. The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 207-208, 211-212, or 231, wherein the targeting sequence is selected from SEQ ID NOs: 265- 269. [0963] Embodiment 250. The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 207-214, 223, or 229-232, wherein the targeting sequence is selected from SEQ ID NOs: 262- 265. [0964] Embodiment 251. The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 216, 223, or 233, wherein the targeting sequence is selected from SEQ ID NOs: 270-284. [0965] Embodiment 252. The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 218, 223, or 233-234, wherein the targeting sequence is selected from SEQ ID NOs: 270-279. [0966] Embodiment 253. The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 216, 219-220, 223, 233, or 235, wherein the targeting sequence is selected from SEQ ID NOs: 280-284. [0967] Embodiment 254. The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 223, or 228-236, wherein the targeting sequence is selected from SEQ ID NOs: 272-281. [0968] Embodiment 255. The antisense oligomer of any one of embodiments 1-3, 7, 198, 215- 223, or 228-237, wherein the targeting sequence is SEQ ID NO: 278. [0969] Embodiment 256. The antisense oligomer of any one of embodiments 1-3, 7, 198, 223, or 238, wherein the targeting sequence is SEQ ID NO: 285. [0970] Embodiment 257. The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 223, or 239, wherein the targeting sequence is SEQ ID NO: 286. [0971] Embodiment 258. The antisense oligomer of any one of embodiments 1, 4-5, 7, 198, 223, or 240, wherein the targeting sequence is SEQ ID NO: 287. [0972] Embodiment 259. The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 6 (SEQ ID NO: 3), the intron 5/exon 6 junction, the exon 6/intron 6 junction, or the intron internal region of intron 6 (SEQ ID NO: 11).
[0973] Embodiment 260. The antisense oligomer of any one of embodiments 1-3 or 259, wherein the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 37th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). [0974] Embodiment 261. The antisense oligomer of embodiment 260, wherein the target region is a region within SEQ ID NO: 58. [0975] Embodiment 262. The antisense oligomer of any one of embodiments 1-3 or 259, wherein the target region of the human DNM2 gene pre-mRNA is within the 32nd nucleotide to the 59th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). [0976] Embodiment 263. The antisense oligomer of embodiment 262, wherein the target region is a region within SEQ ID NO: 59. [0977] Embodiment 264. The antisense oligomer of any one of embodiments 1-3 or 259, wherein the target region of the human DNM2 gene pre-mRNA is within the 60th nucleotide to the 87th nucleotide of exon 6 measured from the 5′ end of exon 6 (SEQ ID NO: 3). [0978] Embodiment 265. The antisense oligomer of embodiment 264, wherein the target region is a region within SEQ ID NO: 60. [0979] Embodiment 266. The antisense oligomer of any one of embodiments 1-3 or 259, wherein the target region of the human DNM2 gene pre-mRNA is within the 130th nucleotide to the 156th nucleotide measured from the 5′ end of exon 6 (SEQ ID NO: 3). [0980] Embodiment 267. The antisense oligomer of embodiment 266, wherein the target region is a region within SEQ ID NO: 61. [0981] Embodiment 268. The antisense oligomer of any one of embodiments 1, 4-5, or 259, wherein the target region of the human DNM2 gene pre-mRNA is within the 17th nucleotide of exon 6 measured from the 3′ end of exon 6 (SEQ ID NO: 3) to the 10th nucleotide of intron 6 measured from the 3′ end of exon 6. [0982] Embodiment 269. The antisense oligomer of embodiment 268, wherein the target region is a region within SEQ ID NO: 62. [0983] Embodiment 270. The antisense oligomer of any one of embodiments 1-5, 7, or 259, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5), DNM2 H6A(-6+19), DNM2 H6A(-2+23), DNM2 H6A(+3+26), DNM2 H6A(+10+3), DNM2 H6A(+13+37), DNM2 H6A(+23+47), DNM2 H6A(+32+56), DNM2 H6A(+35+59), DNM2 H6A(+44+68), DNM2 H6A(+54+78), DNM2 H6A(+60+84), DNM2 H6A(+63+87), DNM2 H6A(+88+112), DNM2 H6A(+96+119), DNM2 H6A(+105+129), DNM2 H6A(+121+145), DNM2 H6A(+130+154), DNM2 H6A(+132+156), DNM2 H6D (+6-19), DNM2 H6D (+15-10), or DNM2 H6D (+17-8).
[0984] Embodiment 271. The antisense oligomer of any one of embodiments 1-3, 7, 260-261, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+10+3) or DNM2 H6A(+13+37). [0985] Embodiment 272. The antisense oligomer of any one of embodiments 1-3, 7, 259, 262- 263, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+32+56) or DNM2 H6A(+35+59). [0986] Embodiment 273. The antisense oligomer of any one of embodiments 1-3, 7, 259, 264- 265, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+60+84) or DNM2 H6A(+63+87). [0987] Embodiment 274. The antisense oligomer of any one of embodiments 1-3, 7, 259, 266- 267, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+130+154) or DNM2 H6A(+132+156). [0988] Embodiment 275. The antisense oligomer of any one of embodiments 1, 4-5, 7, 259, or 268-270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+15-10) or DNM2 H6D (+17-8). [0989] Embodiment 276.The antisense oligomer of any one of embodiments 1, 4-5, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-20+5). [0990] Embodiment 277. The antisense oligomer of any one of embodiments 1, 4-5, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-6+19). [0991] Embodiment 278. The antisense oligomer of any one of embodiments 1, 4-5, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(-2+23). [0992] Embodiment 279. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+3+26). [0993] Embodiment 280. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+23+47). [0994] Embodiment 281. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+44+68). [0995] Embodiment 282. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+54+78). [0996] Embodiment 283. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+88+112). [0997] Embodiment 284. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+96+119). [0998] Embodiment 285. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+105+129).
[0999] Embodiment 286. The antisense oligomer of any one of embodiments 1-3, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6A(+121+145). [1000] Embodiment 287. The antisense oligomer of any one of embodiments 1, 4-5, 7, 259, or 270, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H6D (+6-19). [1001] Embodiment 288.The antisense oligomer of any one of embodiments 259-287, wherein the targeting sequence is selected from SEQ ID NOs: 97-118. [1002] Embodiment 289. The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 7 (SEQ ID NO: 4), the intron 6/exon 7 junction, or the exon 7/intron 7 junction. [1003] Embodiment 290. The antisense oligomer of any one of embodiments 1-3 or 289, wherein the target region of the human DNM2 gene pre-mRNA is within the 72nd nucleotide to the 98th nucleotide of exon 7 measured from the 5′ end of exon 7 (SEQ ID NO: 4). [1004] Embodiment 291. The antisense oligomer of embodiment 290, wherein the target region is a region within SEQ ID NO: 63. [1005] Embodiment 292. The antisense oligomer of any one of embodiments 1, 4-5, or 289, wherein the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 7 measured from the 3′ end of exon 7 (SEQ ID NO: 4) to the 16th nucleotide of intron 7 measured from the 3′ end of exon 7. [1006] Embodiment 293. The antisense oligomer of embodiment 292, wherein the target region is a region within SEQ ID NO: 64. [1007] Embodiment 294. The antisense oligomer of any one of embodiments 1-5, 7, or 289-293, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+1+25), DNM2 H7A(+6+30), DNM2 H7A(+11+35), DNM2 H7A(+22+46), DNM2 H7A(+29+53), DNM2 H7A(+41+65), DNM2 H7A(+51+75), DNM2 H7A(+57+81), DNM2 H7A(+62+86), DNM2 H7A(+72+96), DNM2 H7A(+74+98), DNM2 H7A(+81+105), DNM2 H7A(+91+115), or DNM2 H7A(+110+134). [1008] Embodiment 295. The antisense oligomer of any one of embodiments 1-3, 7, 289-291, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+72+96) or DNM2 H7A(+74+98). [1009] Embodiment 296. The antisense oligomer of any one of embodiments 1, 4-5, 7, 289, or 292-294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+8-16) or DNM2 H7D(+9-16). [1010] Embodiment 297. The antisense oligomer of any one of embodiments 1, 4-5, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-15+10).
[1011] Embodiment 298. The antisense oligomer of any one of embodiments 1, 4-5, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(-5+20). [1012] Embodiment 299. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+1+25). [1013] Embodiment 300. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+6+30). [1014] Embodiment 301. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+11+35). [1015] Embodiment 302. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+22+46). [1016] Embodiment 303. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+29+53). [1017] Embodiment 304. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+41+65). [1018] Embodiment 305. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+51+75). [1019] Embodiment 306. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+57+81). [1020] Embodiment 307. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+62+86). [1021] Embodiment 308. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+81+105). [1022] Embodiment 309. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+91+115). [1023] Embodiment 310. The antisense oligomer of any one of embodiments 1-3, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7A(+110+134). [1024] Embodiment 311. The antisense oligomer of any one of embodiments 1, 4-5, 7, 289, or 294, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H7D(+21-4). [1025] Embodiment 312. The antisense oligomer of any one of embodiments 289-311, wherein the targeting sequence is selected from SEQ ID NOs: 119-137. [1026] Embodiment 313. The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 8 (SEQ ID NO: 5), the intron 7/exon 8 junction, or the exon 8/intron 8 junction. [1027] Embodiment 314. The antisense oligomer of any one of embodiments 1, 4-5, or 313, wherein the target region of the human DNM2 gene pre-mRNA is within the 7th nucleotide of
intron 7 measured from the 5′ end of exon 8 (SEQ ID NO: 5) to the 19th nucleotide of exon 8 measured from the 5′ end of exon 8. [1028] Embodiment 315. The antisense oligomer of embodiment 314, wherein the target region is a region within SEQ ID NO: 65. [1029] Embodiment 316. The antisense oligomer of any one of embodiments 1-3 or 313, wherein the target region of the human DNM2 gene pre-mRNA is within the 18th nucleotide to the 45th nucleotide of exon 8 measured from the 5′ end of exon 8 (SEQ ID NO: 5). [1030] Embodiment 317. The antisense oligomer of embodiment 316, wherein the target region is a region within SEQ ID NO: 66. [1031] Embodiment 318. The antisense oligomer of any one of embodiments 1-5, 7, or 313-317, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-20+5), DNM2 H8A(-14+11), DNM2 H8A(-7+18), DNM2 H8A(-6+19), DNM2 H8A(-2+23), DNM2 H8A(+5+29), DNM2 H8A(+18+42), DNM2 H8A(+21+45), DNM2 H8A(+27+51), DNM2 H8A(+36+60), DNM2 H8A(+46+70), DNM2 H8A(+51+75), DNM2 H8A(+56+80), DNM2 H8A(+65+88), DNM2 H8A(+76+100), DNM2 H8A(+83+107), DNM2 H8A(+96+119), DNM2 H8A(+105+129), DNM2 H8D(+18-7), or DNM2 H8D(+10-15). [1032] Embodiment 319. The antisense oligomer of any one of embodiments 1, 4-5, 7, 313-315, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-7+18) or DNM2 H8A(-6+19). [1033] Embodiment 320. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 316-317, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+18+42) or DNM2 H8A(+21+45). [1034] Embodiment 321. The antisense oligomer of any one of embodiments 1, 4-5, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-20+5). [1035] Embodiment 322. The antisense oligomer of any one of embodiments 1, 4-5, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-14+11). [1036] Embodiment 323. The antisense oligomer of any one of embodiments 1, 4-5, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(-2+23). [1037] Embodiment 324. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+5+29). [1038] Embodiment 325. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+27+51). [1039] Embodiment 326. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+36+60).
[1040] Embodiment 327. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+46+70). [1041] Embodiment 328. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+51+75). [1042] Embodiment 329. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+56+80). [1043] Embodiment 330. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+65+88). [1044] Embodiment 331. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+76+100). [1045] Embodiment 332. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+83+107). [1046] Embodiment 333. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+96+119). [1047] Embodiment 334. The antisense oligomer of any one of embodiments 1-3, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8A(+105+129). [1048] Embodiment 335. The antisense oligomer of any one of embodiments 1, 4-5, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+18-7). [1049] Embodiment 336. The antisense oligomer of any one of embodiments 1, 4-5, 7, 313, or 318, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H8D(+10-15). [1050] Embodiment 337. The antisense oligomer of any one of embodiments 313-336, wherein the targeting sequence is selected from SEQ ID NOs: 138-157. [1051] Embodiment 338. The antisense oligomer of any one of embodiments 1-5 wherein the target region is within the exon internal region of exon 16 (SEQ ID NO: 9), the intron 15/exon 16 junction, or the exon 16/intron 16 junction. [1052] Embodiment 339. The antisense oligomer of any one of embodiments 1-5, 7, or 338, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3), DNM2 H16A(-13+12), DNM2 H16A(-6+19), DNM2 H16A(+6+30), DNM2 H16A(+15+39), DNM2 H16A(+26+50), DNM2 H16A(+36+60), DNM2 H16A(+45+69), DNM2 H16A(+55+79), DNM2 H16A(+66+90), DNM2 H16A(+75+99), DNM2 H16A(+82+106), DNM2 H16D(+19-6), or DNM2 H16D(+11-14). DNM2 H16A(-21+3). [1053] Embodiment 340. The antisense oligomer of any one of embodiments 1, 4-5, 7, or 338- 339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-21+3). [1054] Embodiment 341. The antisense oligomer of any one of embodiments 1, 4-5, 7, or 338- 339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-13+12).
[1055] Embodiment 342. The antisense oligomer of any one of embodiments 1, 4-5, 7, or 338- 339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(-6+19). [1056] Embodiment 343. The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+6+30). [1057] Embodiment 344. The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+15+39). [1058] Embodiment 345. The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+26+50). [1059] Embodiment 346. The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+36+60). [1060] Embodiment 347. The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+45+69). [1061] Embodiment 348. The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+55+79). [1062] Embodiment 349. The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+66+90). [1063] Embodiment 350. The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+75+99). [1064] Embodiment 351. The antisense oligomer of any one of embodiments 1-3, 7, or 338-339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16A(+82+106). [1065] Embodiment 352. The antisense oligomer of any one of embodiments 1, 4-5, 7, or 338- 339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16D(+19-6). [1066] Embodiment 353. The antisense oligomer of any one of embodiments 1, 4-5, 7, or 338- 339, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H16D(+11-14). [1067] Embodiment 354. The antisense oligomer of any one of embodiments 338-353, wherein the targeting sequence is selected from SEQ ID NOs: 288-301. [1068] Embodiment 355. The antisense oligomer of any one of embodiments 1-5, wherein the target region is within the exon internal region of exon 19 (SEQ ID NO: 9), the intron 18/exon 19 junction, or the exon 18/intron 18 junction. [1069] Embodiment 356. The antisense oligomer of any one of embodiments 1-5, 7, or 355, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14), DNM2 H19A(-4+21), DNM2 H19A(+1+25), DNM2 H19A(+7+31), DNM2 H19A(+16+40), DNM2 H19A(+26+50), DNM2 H19A(+37+61), DNM2 H19A(+47+71), DNM2 H19A(+56+80), DNM2 H19A(+66+90), DNM2 H19A(+76+100), DNM2 H19A(+86+110), DNM2 H19A(+96+120), DNM2 H19A(+106+130), DNM2 H19A(+117+141), DNM2 H19A(+127+151), DNM2 H19A(+137+161),
DNM2 H19A(+147+171), DNM2 H19A(+157+181), DNM2 H19A(+166+189), DNM2 H19A(+191+215), DNM2 H19A(+200+224), DNM2 H19D(+23-2), DNM2 H19D(+17-8), or DNM2 H19D(+10-15). [1070] Embodiment 357. The antisense oligomer of any one of embodiments 1, 4-5, or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-11+14). [1071] Embodiment 358. The antisense oligomer of any one of embodiments 1, 4-5, or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(-4+21). [1072] Embodiment 359. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+1+25). [1073] Embodiment 360. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+7+31). [1074] Embodiment 361. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+16+40). [1075] Embodiment 362. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+26+50). [1076] Embodiment 363. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+37+61). [1077] Embodiment 364. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+47+71). [1078] Embodiment 365. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+56+80). [1079] Embodiment 366. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+66+90). [1080] Embodiment 367. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+76+100). [1081] Embodiment 368. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+86+110). [1082] Embodiment 369. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+96+120). [1083] Embodiment 370. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+106+130). [1084] Embodiment 371. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+117+141). [1085] Embodiment 372. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+127+151).
[1086] Embodiment 373. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+137+161). [1087] Embodiment 374. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+147+171). [1088] Embodiment 375. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+157+181). [1089] Embodiment 376. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+166+189). [1090] Embodiment 377. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+191+215). [1091] Embodiment 378. The antisense oligomer of any one of embodiments 1-3 or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19A(+200+224). [1092] Embodiment 379. The antisense oligomer of any one of embodiments 1, 4-5, or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+23-2). [1093] Embodiment 380. The antisense oligomer of any one of embodiments 1, 4-5, or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+17-8). [1094] Embodiment 381. The antisense oligomer of any one of embodiments 1, 4-5, or 355-356, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H19D(+10-15). [1095] Embodiment 382. The antisense oligomer of any one of embodiments 355-381, wherein the targeting sequence is selected from SEQ ID NOs: 302-326. [1096] Embodiment 383. The antisense oligomer of any one of embodiments 1-382, wherein the antisense oligomer is selected from a peptide nucleic acid, a locked nucleic acid, a phosphorodiamidate morpholino oligomer, a 2′-OMe phosphorothioate oligomer, or a combination thereof. [1097] Embodiment 384. The antisense oligomer of any one of embodiments 1-383, wherein the antisense oligomer is a phosphorodiamidate morpholino oligomer. [1098] Embodiment 385. The antisense oligomer of any one of embodiments 1-384, wherein the antisense oligomer is covalently linked to a delivery moiety selected from a cell-penetrating peptide, an antibody, a fragment of an antibody, and an antigen binding agent. [1099] Embodiment 386. The antisense oligomer of any one of embodiments 1-385, wherein the antisense oligomer is covalently linked to a cell-penetrating peptide. [1100] Embodiment 387. The antisense oligomer of embodiment 386, wherein the antisense oligomer is covalently linked to the cell-penetrating peptide by a linker selected from a direct bond, a glycine amino acid, a proline amino acid, glutamic acid amino acid, or an isoglutamine amino acid.
[1101] Embodiment 388. The antisense oligomer of embodiment 386 or 387, wherein the cell- penetrating peptide is selected from rTAT, Tat, R9F2, R5F2R4, R4, R5, R6, R7, R8, R9, (RXR)4, (RXR)5, (RXRRBR)2, (RAR)4F2, (RGR)4F2, and RBRBYLIQFRBRRBR, wherein A represents alanine, B represents β-alanine, F represents phenylalanine, G represents glycine, I represents isoleucine, L represents leucine, Q represents glutamine, R represents arginine, X represents 6- aminohexanoic acid, and Y represents tyrosine. [1102] Embodiment 389. The antisense oligomer of any one of embodiments 1-382 having a structure of Formula (I):
or a pharmaceutically acceptable salt thereof, wherein:
R5 is -C(O)(O-alkyl)x-OH, wherein x is 3-10 and each alkyl group is, independently at each occurrence, C2-6-alkyl, or R5 is selected from H, -C(O)C1-6-alkyl, trityl, monomethoxytrityl, -(C1-6-alkyl)-R6, -(C1-6- heteroalkyl)-R6, -C6-10-aryl-R6, 5-10 membered heteroaryl-R6, -C(O)O-(C1-6-alkyl)-R6, - C(O)O-aryl-R6, -C(O)O-(5-10 membered heteroaryl)-R6, and
; R6 is selected from -OH, -SH, and -NH2, or R6 is O, S, or NH, each of which is covalently linked to a solid support; R9 is C1-6 alkyl; each R1 is independently selected from -OH and -N(R3)(R4), wherein each R3 and R4 is, independently at each occurrence, -H or -C1-6-alkyl; each R2 is independently selected from a naturally or non-naturally occurring nucleobase, which, when taken together, forms the targeting sequence; t is 11-28; E′ is selected from -H, -C1-6-alkyl, -C(O)C1-6-alkyl, benzoyl, stearoyl, trityl, monomethoxytrityl, dimethoxytrityl, trimethoxytrityl,
wherein Q is -C(O)(CH2)6C(O)- or -C(O)(CH2)2S2(CH2)2C(O)-; R7 is -(CH2)2OC(O)N(R8)2, wherein R8 is -(CH2)6NHC(=NH)NH2; L is a linking amino acid, wherein L is covalently-linked by an amide bond to the C-terminus of J; J is a cell-penetrating peptide; and G is selected from -H, -C(O)C1-6-alkyl, benzoyl, and stearoyl, wherein G is covalently-linked to J. [1103] Embodiment 390. The antisense oligomer of or antisense oligomer embodiment 389, wherein E′ is selected from -H, -C1-6-alkyl, -C(O)C1-6-alkyl, benzoyl, stearoyl, trityl, monomethoxytrityl, dimethoxytrityl, trimethoxytrityl, and
. [1104] Embodiment 391. The antisense oligomer of embodiment 389 or 390, wherein E′ is selected from -H, -C(O)CH3, benzoyl, stearoyl, trityl, 4-methoxytrityl, and
. [1105] Embodiment 392. The antisense oligomer of any one of embodiments 389-391, wherein A′ is selected from:
. [1106] Embodiment 393. The antisense oligomer of any one of embodiments 389-392, wherein at least one of the following is true:
[1107] Embodiment 394. The antisense oligomer of any one of embodiments 389-393, wherein A′ is selected from:
; and E′ is
. [1108] Embodiment 395. The antisense oligomer of any one of embodiments 389-394, wherein A′ is:
E′ is selected from H, -C(O)CH3, trityl, 4-methoxytrityl, benzoyl, and stearoyl. [1109] Embodiment 396. The antisense oligomer of any one of embodiments 389-395, wherein each R1 is -N(CH3)2. [1110] Embodiment 397. The antisense oligomer of any one of embodiments 389-396, wherein L is glycine, proline, or β-alanine. [1111] Embodiment 398. The antisense oligomer of any one of embodiments 389-397, wherein L is glycine. [1112] Embodiment 399. The antisense oligomer of any one of embodiments 389-397, wherein L is proline. [1113] Embodiment 400. The antisense oligomer of any one of embodiments 389-397, wherein L is β-alanine. [1114] Embodiment 401. The antisense oligomer of any one of embodiments 389-400, wherein J is selected from rTAT, Tat, R9F2, R5F2R4, R4, R5, R6, R7, R8, R9, (RXR)4, (RXR)5, (RXRRBR)2, (RAR)4F2, and (RGR)4F2, wherein A represents alanine, B represents β-alanine, F represents
phenylalanine, G represents glycine, R represents arginine, and X represent 6-aminohexanoic acid. [1115] Embodiment 402. The antisense oligomer of any one of embodiments 389-401, wherein G is selected from -H, -C(O)CH3, benzoyl, and stearoyl. [1116] Embodiment 403. The antisense oligomer of any one of embodiments 389-402, wherein G is -H or -C(O)CH3. [1117] Embodiment 404. The antisense oligomer of any one of embodiments 389-403, wherein G is -H. [1118] Embodiment 405. The antisense oligomer of any one of embodiments 389-403, wherein G is -C(O)CH3. [1119] Embodiment 406. The antisense oligomer of any one of embodiments 389-397, 400-403, or 405, wherein the antisense oligomer is of Formula (IA): ,
or a pharmaceutically acceptable salt thereof, wherein: A′ is a moiety selected from:
[1120] Embodiment 407. The antisense oligomer of any one of embodiments 389-394 or 396- 406, wherein the antisense oligomer is of Formula (II):
(II), or a pharmaceutically acceptable salt thereof. [1121] Embodiment 408. The antisense oligomer of any one of embodiments 389-384, 396-397, 400-403, or 405-407, wherein the antisense oligomer is of Formula (III):
(III), or a pharmaceutically acceptable salt thereof, wherein n is 11-28. [1122] Embodiment 409. The antisense oligomer of any one of embodiments 389-384, 396-397, 400-403, or 405-408, or a pharmaceutically acceptable salt thereof, wherein the antisense oligomer is of Formula (IV):
(IV), wherein n is 11-28. [1123] Embodiment 410. The antisense oligomer of embodiment 408 or 409, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 83. [1124] Embodiment 411. The antisense oligomer of embodiment 408 or 409, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 88. [1125] Embodiment 412. The antisense oligomer of embodiment 408 or 409, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 172.
[1126] Embodiment 413. The antisense oligomer of embodiment 408 or 409, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 199. [1127] Embodiment 414. The antisense oligomer of embodiment 408 or 409, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 230. [1128] Embodiment 415. The antisense oligomer of embodiment 408 or 409, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 278. [1129] Embodiment 416. An antisense oligomer selected from:
,
. [1136] Embodiment 423. A pharmaceutical composition, comprising the antisense oligomer, or a pharmaceutically acceptable salt thereof, of any one of embodiments 1-422 and a pharmaceutically acceptable carrier. [1137] Embodiment 424. A method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein, comprising administering to a subject in need thereof a therapeutically effective amount of the antisense oligomer, or a pharmaceutically acceptable salt thereof, of any one of embodiments 1-422 or the pharmaceutical composition of embodiment 423. [1138] Embodiment 425. The method of embodiment 424, wherein the disease or disorder is a genetic neuromuscular disorder. [1139] Embodiment 426. The method of embodiment 424 or 425, wherein the disease or disorder is caused by at least one mutation in a gene selected from DNM2 gene, MTM1 gene,
BIN1 gene, RYR1 gene, TTN gene, SPEG gene, CACNA1S gene, MTMR14 gene, ZAK gene, and CCDC78 gene. [1140] Embodiment 427. The method of any one of embodiments 424-426, wherein the disease or disorder is autosomal dominant, autosomal recessive, or X-linked. [1141] Embodiment 428. The method of any one of embodiments 424-427, wherein the disease or disorder is a centronuclear myopathy (CNM). [1142] Embodiment 429. The method of any one of embodiments 424-428, wherein the disease or disorder is myotubular myopathy (XLMTM or XLCNM), autosomal dominant congenital myopathy (ADCNM), autosomal recessive centronuclear myopathy (ARCNM), or Charcot-Marie- Tooth disease. EXAMPLES [1143] Examples have been set forth below for the purpose of illustration and to describe certain specific embodiments of the disclosure. However, the scope of the claims is not to be in any way limited by the examples set forth herein. Various changes and modifications to the disclosed embodiments will be apparent to those skilled in the art and such changes and modifications including, without limitation, those relating to the chemical structures, substituents, derivatives, formulations, or methods of the disclosure may be made without departing from the spirit of the disclosure and the scope of the appended claims. Definitions of the variables in the structures in the schemes herein are commensurate with those of corresponding positions in the formulae presented herein. Example 1 – Design of Antisense Targeting Sequences [1144] Antisense oligomer targeting sequences were designed for therapeutic applications related to the pre-mRNA of the human DNM2 gene. Here, antisense oligomers will induce exon skipping to create a premature stop codon (PTC) and thereby trigger nonsense-mediated decay to degrade DNM2 mRNA in a diseased cell. Reduction of DNM2 expression to normal or near- normal levels would then reduce disease-causing aggregation or increased DNM2 activity in the cell, thereby providing a clinical benefit to patients. [1145] Exemplary oligomers comprising a targeting sequence as set forth in Tables 5 and 6 were prepared as PMOs. As described below, these antisense oligomers were introduced into Rhabdomyosarcoma (RD) cells and analyzed for DNM2 expression. The knockdown data is shown in Tables 5 and 6. The PMOs used in this example are of Formula (V) below having the sequences indicated in Tables 5 and 6.
Example 2 – Macrowalk Screen [1146] Rhabdomyosarcoma (RD) cells were seeded at 15,000-20,000 cells per well in 96-well plates, in DMEM (Gibco, 10569010) with 10% FBS (Cytiva, SH30071.03) and treated a day later with a combination of 1 µM Endo-Porter (Gene tools) and 20 µM of each PMO diluted in DMEM + 10% FBS. Cells were harvested after 48 hours of treatment and RNA extracted (Quick RNA 96 kit, Zymo Research) (PBS- Vehicle/no treatment control, Negative control PMO 1, positive control PPMO 1). RT-PCR was performed using Maxima™ H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTime™ Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to manufacturer’s instructions. PrimeTime qPCR Probe Assays (IDT) are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems. The results of the macrowalk screen are shown in Table 5 and FIGs.1A-1E and FIGs.2A-2F. The PMOs used in this example are of Formula (V) above having the sequences indicated in Table 5.
RT-PCR conditions were as follows:
1.Incubate at 25° C for 10 minutes. 2.Incubate at 50 °C for 15 minutes. 3.Incubate at 85 °C for 5 minutes. qPCR conditions were as follows:
Table 5. Macrowalk screen- mRNA reduction of human DNM2 by PMOs in Rhabdomyosarcoma (RD) cells.
ND – mRNA reduction (knock down) not detected. “B” represents an abasic subunit or guanine. Example 3 – Microwalk Screen [1147] Rhabdomyosarcoma (RD) cells were prepared and treated with PMOs according to the protocol described in Example 2. The results of the microwalk screen are shown in Table 6 and FIGs.3A-3J. The PMOs used in this example are of Formula (V) above having the sequences indicated in Table 6.
Table 6. Microwalk lead optimization screen – mRNA reduction of human DNM2 by PMOs in rhabdomyosarcoma (RD) cells.
“B” represents an abasic subunit or guanine. Example 4 – Dose response of PMOs in rhabdomyosarcoma (RD) cells. [1148] Rhabdomyosarcoma (RD) cells were seeded at 15,000-20,000 cells per well in 96-well plates, in DMEM (Gibco, 10569010) with 10% FBS (Cytiva, SH30071.03) and treated a day later with a combination of 1uM Endo-Porter (Gene tools) and the indicated concentrations (1, 2.5, 5, 10, 20, 30, 40, 50 µM) of each PMO diluted in DMEM + 10% FBS. Cells were harvested after 48 hours of treatment and RNA extracted (Quick RNA 96 kit, Zymo Research) (PBS- Vehicle/no treatment control, Negative control PMO 1, positive control PPMO 1). RT-PCR was performed using Maxima™ H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTime™ Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to the manufacturer’s instructions. PrimeTime™ qPCR Probe Assays (IDT) are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems. The results of the PMO dose response are shown in FIGs.4A-4G. The PMOs used in this example are of Formula (V) above having the sequences disclosed in Table 7. Table 7.
RT-PCR conditions were as follows:
1.Incubate at 25° C for 10 minutes. 2.Incubate at 50 °C for 15 minutes. 3.Incubate at 85 °C for 5 minutes. qPCR conditions were as follows:
Example 5 – Exon-skipping induced by PPMOs targeting exon 2 and exon 9 in rhabdomyosarcoma (RD) cells. [1149] Rhabdomyosarcoma (RD) cells were seeded at 15,000-20,000 cells per well in 96-well plates, in DMEM (Gibco, 10569010) with 10% FBS (Cytiva, SH30071.03) and treated a day later with a combination of 1 µM Endo-Porter (Gene tools) and 20 µM of each PPMO diluted in DMEM + 10% FBS. Cells were harvested after 48 hours of treatment and RNA extracted (Quick RNA 96 kit, Zymo Research). RT-PCR was performed using Maxima™ H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by a PCR utilizing Platinum™ II Hot- Start PCR Master Mix (2X) (Fisher Scientific 14000013) for 36 cycles. Samples were run on a FlashGel system (Lonza, 57067) utilizing a 2.2% DNA cassette (57031). The results of PPMO induced exon-skipping are shown in FIGs.5A-5B. The PPMOs used in this example are of Formula (IV) above having the sequences indicated in Table 8. Table 8.
RT-PCR conditions were as follows:
1.Incubate at 25° C for 10 minutes. 2.Incubate at 50 °C for 15 minutes.
3.Incubate at 85 °C for 5 minutes. PCR conditions were as follows:
Example 6 – DNM2 protein reduction in rhabdomyosarcoma (RD) cells induced by a PPMO targeting exon 2. [1150] Rhabdomyosarcoma (RD) cells were seeded at 90,000 cells per well in 24-well plates, in DMEM (Gibco, 10569010) with 10% FBS (Cytiva, SH30071.03) and treated a day later with a combination of 1uM Endo-Porter (Gene tools) and 20uM of the indicated PPMO/PMO compound diluted in DMEM + 10% FBS (PBS- Vehicle/no treatment control, Negative control PMO 1,3 wells per compound). After 48 hours of treatment, cell lysates were prepared in M-PER buffer (ThermoFisher, 78501) supplemented with Halt™ Protease and Phosphatase Inhibitor Cocktail, EDTA-free (ThermoFischer, 78447). Protein concentration was measured using Pierce™ Rapid Gold BCA Protein Assay Kit (ThermoFisher, A53225). Cell lysates were prepared using the sample preparation kit (Proteinsimple) for an automated capillary Western blot system, JESS system (Proteinsimple), and utilizing the protein normalization module. Cell lysates were diluted to the same protein concentrations using the 0.1X sample buffer and mixed with 5X fluorescence master mix followed by denaturation according to the manufacturer’s instructions. JESS was run using a 1:200 diluted anti-DNM2 primary antibody (Sigma HPA054246). Antibody specificity was confirmed separately by DNM2 targeting siRNA treatment. Signal intensity (peak area) of the protein was normalized to the peak area of the total protein included in the capillary well using the protein normalization kit and analysis on
Compass Software. Quantitative analysis of DNM2 protein band was performed using the Compass Software (ProteinSimple), presented as the mean +/- SD. The results of protein reduction by PPMO 1 (i.e., a PPMO of Formula (IV)) are shown in FIGs.6A-6B. Example 7 – mRNA reduction in human DNM2 following PPMO treatment in AB1190C16PV myoblast immortalized cell line, differentiation method 1. [1151] AB1190C16PV human immortalized WT myoblasts cells derived from paravertebral muscle of 16-year-old male (see Mamchaoui, K., et al., Immortalized pathological human myoblasts: towards a universal tool for the study of neuromuscular disorders. Skeletal muscle 1, 34 (2011)) were seeded at 10,000 cells per well in 96-well collagen Type I coated plates, in PromoCell™ skeletal muscle growth media (C-23060). Cells were allowed to differentiate in growth media for six days (method 1) followed by treatment with compounds at two doses, 20 µM and 60 µM, free uptake. Differentiation was monitored by Desmin and MHC immunostaining. Cells were harvested after 48 hours of treatment and RNA extracted (Quick RNA 96 kit, Zymo Research) (PBS- Vehicle/no treatment control, negative control PPMO 9, positive control PPMO 1). RT-PCR was performed using Maxima™ H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTime™ Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to the manufacturer’s instructions. PrimeTime™ qPCR Probe Assays (IDT) are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems. The results of AB1190C16PV myoblast immortalized cell line treated with PPMO 1, PPMO 3, PPMO 4, PPMO 5, PPMO 6, PPMO 7, and PPMO 8 (at various concentrations) are shown in Table 9 and FIG 7. The PPMOs used in this example are of Formula (IV) having the sequences indicated in Table 9.
RT-PCR conditions were as follows:
1.Incubate at 25° C for 10 minutes. 2.Incubate at 50 °C for 15 minutes. 3.Incubate at 85 °C for 5 minutes. qPCR conditions were as follows:
Table 9. mRNA reduction of human DNM2 by PPMOs in differentiated myoblast immortalized cell line.
ND – mRNA reduction (knock down) not detected. NTC – non targeting control. The NTC is a PMO that targets exon 2, but does not induce skipping . Example 8 – mRNA reduction in human DNM2 following PPMO treatment in AB1190C16PV myoblast immortalized cell line, differentiation method 2. [1152] AB1190C16PV human immortalized WT myoblasts cells (derived from paravertebral muscle of 16-year-old male) were seeded at 10,000 cells per well in 96-well collagen Type I coated plates, in PromoCell skeletal muscle growth media (C-23060). Cells were allowed to proliferate for two days after which media was replaced to differentiation media (C-23061) (method 2). After four days of differentiation, cells were treated with compounds at two doses, 20 µM and 60 µM, free uptake. Differentiation was monitored by Desmin and MHC immunostaining. Cells were harvested after 48 hours of treatment and RNA extracted (Quick RNA 96 kit, Zymo Research) (PBS- Vehicle/no treatment control, negative control PPMO 9, positive control PPMO 12). RT-PCR was performed using Maxima™ H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTime™ Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to manufacturer’s instructions. PrimeTime qPCR Probe Assays (IDT) are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems. The results of AB1190C16PV myoblast immortalized cell line treated with PPMO 1, PPMO 3, PPMO 4, PPMO 5, PPMO 6, PPMO 7, and PPMO 8 (at various concentrations) are shown in Table 10 and FIG 8. The PPMOs used in this example are of Formula (IV) having the sequences indicated in Table 10.
RT-PCR conditions were as follows:
1.Incubate at 25° C for 10 minutes. 2.Incubate at 50 °C for 15 minutes. 3.Incubate at 85 °C for 5 minutes. qPCR conditions were as follows:
Table 10. mRNA reduction of human DNM2 by PPMOs in differentiated myoblast immortalized cell line.
NTC – non targeting control. The NTC is a PMO that targets exon 2, but does not induce skipping. Example 9 – mRNA reduction in human DNM2 following PPMO treatment in differentiated healthy human skeletal muscle derived iPSCs. iPSC reprogramming (iXCell) [1153] Human fibroblast cells were reprogrammed using integration-free episomal vectors carrying Oct-4, Sox2, Lin28, L-Myc, Klf4, mp53DD, and EBNA from the Ei5TM Episomal iPSC Reprogramming kit (Life Technology, Cat.no. A15960). Primary fibroblast cells were cultured at a density of 105 cells, electrotransfected (Neon, Themofisher), and seeded on plates coated with Matrigel (Corning, Cat.no.354230). Cells were cultured on a mouse feeder layer until day 15 that was replaced with mTesR medium (StemCell Technologies, Cat.no.100-0276) until iPSC colonies appeared. The colonies were isolated, and their pluripotency was validated by immunostaining with markers Oct3/4, NANOG, TRA-1-60. iPSCs were confirmed to retain normal karyotype and alkaline phosphatase activity. The iPSCs were maintained in the same
mTesR medium on Matrigel-coated dishes and passaged using ReLeSR (StemCell Technologies, Cat.no.05873). Myogenic progenitor generation (iXCell) [1154] Myogenic progenitors were differentiated from hiPSCs according to the protocols described in Chal, J., et al., Differentiation of pluripotent stem cells to muscle fiber to model Duchenne muscular dystrophy. Nat. Biotech.2015, 33, 962–969; and Chal, J., et al. Generation of human muscle fibers and satellite-like cells from human pluripotent stem cells in vitro. Nat. Protoc.2016, 11, 1833–1850. Briefly, myogenic progenitors were generated through a multi- step small molecule differentiation protocol, and later subcultured in hSKM Expansion Medium and Differentiation Medium (iXCells Biotechnologies, San Diego, US). [1155] Human normal induced pluripotent stem cells (iPSC) derived myoblasts (iXCell, 40HU- 176) were seeded at 20,000 cells per well in 96-well collagen Type I coated plates, in iPSC- derived Myoblast Expansion Medium (MD-0102A). Cells expanded in expansion media for one day before media was replaced for iPSC-derived Myoblast Differentiation Medium (MD-0102B). Differentiation was monitored by Desmin and MHC immunostaining. After 3 days of differentiation, cells were treated with compounds at two doses, 20 µM and 60 µM, free uptake, in differentiation media and harvested after 48 hours of treatment, followed by RNA extraction (Quick RNA 96 kit, Zymo Research) (PBS- no treatment control, negative control PPMO 9, positive control PPMO 12). RT-PCR was performed using Maxima™ H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTime™ Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to the manufacturer’s instructions. PrimeTime qPCR Probe Assays (IDT) are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems. RT-PCR and qPCR conditions were as described in Example 8. The results of healthy human iPSC derived skeletal muscle treated with PPMO 1, PPMO 3, PPMO 4, PPMO 5, PPMO 6, PPMO 7, and PPMO 8 (at various concentrations) are shown in Table 11 and FIG 9. The PPMOs used in this example are of Formula (IV) above having the sequences indicated in Table 11. Table 11. mRNA reduction of human DNM2 by PPMOs in differentiated healthy human skeletal muscle derived iPSCs.
ND – mRNA reduction (knock down) not detected. NTC – non targeting control. The NTC is a PMO that targets exon 2, but does not induce skipping. Example 10 – mRNA reduction of human DNM2 following PPMO treatment in MTM1 patient-derived iPSC myotubes, GM06813 C1. iPSC reprogramming (iXCell) [1156] Human fibroblast cells were reprogrammed using integration-free episomal vectors carrying Oct-4, Sox2, Lin28, L-Myc, Klf4, mp53DD, and EBNA from the Ei5TM Episomal iPSC Reprogramming kit (Life Technology, Cat.no. A15960). Primary fibroblast cells were cultured at a density of 105 cells, electrotransfected (Neon, Themofisher), and seeded on plates coated with Matrigel (Corning, Cat.no.354230). Cells were cultured on a mouse feeder layer until day 15 that was replaced with mTesR medium (StemCell Technologies, Cat.no.100-0276) until iPSC colonies appeared. The colonies were isolated, and their pluripotency was validated by immunostaining with markers Oct3/4, NANOG, TRA-1-60. iPSCs were confirmed to retain normal karyotype and alkaline phosphatase activity. The iPSCs were maintained in the same mTesR medium on Matrigel-coated dishes and passaged using ReLeSR (StemCell Technologies, Cat.no.05873). Myogenic progenitor generation (iXCell) [1157] Myogenic progenitors were differentiated from hiPSCs according to the protocol described in Example 9. Myogenic lineage was confirmed by immunofluorescence of the myoblast markers Desmin and MyoD and expression. Terminal differentiation of the myoblasts
was performed over 3-6 days of culturing and confirmed by expression of the myogenic markers MHC and MyoG measured by immunofluorescence. [1158] Two patient untransformed fibroblast lines were purchased from the Coriell Institute, GM06813 and GM24509A, and reprogrammed at iXCell into iPSC derived myoblasts as detailed above. Two clones of each line were validated as detailed above. Cells from clone 1 of GM06813 (GM06813 C1) and clone 4 of GM24509A (GM24509A C4) were seeded at 20,000 cells per well in 96-well collagen Type I coated plates, in iPSC-derived Myoblast Expansion Medium (MD-0102A). Cells expanded in expansion media for one day before media was replaced for iPSC-derived Myoblast Differentiation Medium (MD-0102B). Differentiation was monitored by Desmin and MHC immunostaining. After 3 days of differentiation, cells were treated with compounds at two doses, 20 µM and 60 µM, free uptake, in differentiation media and harvested after 48 hours of treatment, followed by RNA extraction (Quick RNA 96 kit, Zymo Research) (PBS- no treatment control, Negative PPMO 9). RT-PCR was performed using Maxima™ H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTime™ Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to manufacturer’s instructions. PrimeTime qPCR Probe Assays (IDT) are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems. RT-PCR and qPCR conditions were as in Example 8. The results of healthy human iPSC derived skeletal muscle treated with PPMO 2, PPMO 3, PPMO 4, PPMO 5, PPMO 6, PPMO 7, and PPMO 8 (at various concentrations) are shown in Table 12 and FIG 10. The PPMOs used in this example are of Formula (IV) having the sequences indicated in Table 12. Table 12. mRNA reduction of human DNM2 by PPMOs in MTM1 patient-derived iPSC myotubes, GM06813 C1.
ND – mRNA reduction (knock down) not detected. NTC – non targeting control. The NTC is a PMO that targets exon 2, but does not induce skipping. Example 11 – PPMO treatment in MTM1 patient-derived iPSC myotubes, GM24509 C4. [1159] The assay conditions for this experiment were as described in Example 10. The results of healthy human iPSC derived skeletal muscle treated with PPMO2, PPMO 3, PPMO 4, PPMO 5, PPMO 6, PPMO 7, and PPMO 8 (at various concentrations) are shown in Table 13 and FIG 11. The PPMOs used in this example are of Formula (IV) having the sequences indicated in Table 13. Table 13. mRNA reduction of human DNM2 by PPMOs in MTM1 patient-derived iPSC myotubes, GM06813 C1.
ND – mRNA reduction (knock down) not detected. NTC – non targeting control. The NTC is a PMO that targets exon 2, but does not induce skipping.
Example 12 – Dose response with select PPMO compounds in GM06813 C1 myotubes. iPSC reprogramming (iXCell) [1160] Human fibroblast cells were reprogrammed using integration-free episomal vectors carrying Oct-4, Sox2, Lin28, L-Myc, Klf4, mp53DD, and EBNA from the Ei5TM Episomal iPSC Reprogramming kit (Life Technology, Cat.no. A15960). Primary fibroblast cells were cultured at a density of 105 cells, electrotransfected (Neon, Themofisher), and seeded on plates coated with Matrigel (Corning, Cat.no.354230). Cells were cultured on a mouse feeder layer until day 15 that was replaced with mTesR medium (StemCell Technologies, Cat.no.100-0276) until iPSC colonies appeared. The colonies were isolated, and their pluripotency was validated by immunostaining with markers Oct3/4, NANOG, TRA-1-60. iPSCs were confirmed to retain normal karyotype and alkaline phosphatase activity. The iPSCs were maintained in the same mTesR medium on Matrigel-coated dishes and passaged using ReLeSR (StemCell Technologies, Cat.no.05873). Myogenic progenitor generation (iXCell) [1161] Myogenic progenitors were differentiated from hiPSCs according to the protocol described in Example 9. Myogenic lineage was confirmed by immunofluorescence of the myoblast markers Desmin and MyoD and expression. Terminal differentiation of the myoblasts was performed over 3-6 days of culturing and confirmed by expression of the myogenic markers MHC and MyoG measured by immunofluorescence. The results of the dose responses on protein reduction by PPMO 2, PPMO 3, PPMO 4, PPMO 5, PPMO 6, PPMO 7, and PPMO 8 in normal myotubes differentiated for three days are shown in FIGs.12A-12G. The PPMOs used in this example are of Formula (IV) having the sequences indicated in Table 14. NTC is a non- targeting control. The NTC is a PMO that targets exon 2, but does not induce skipping (is not active). [1162] Cells from clone 1 of GM06813 (GM06813 C1) were seeded at 20,000 cells per well in 96-well collagen Type I coated plates, in iPSC-derived Myoblast Expansion Medium (MD- 0102A). Cells expanded in expansion media for one day before media was replaced for iPSC- derived Myoblast Differentiation Medium (MD-0102B). Differentiation was monitored by Desmin and MHC immunostaining. After 3 days of differentiation, cells were treated with compounds at 8 doses as indicated (1, 2.5, 5, 10, 20, 30, 40, 60 µM), free uptake, in differentiation media and harvested after 48 hours of treatment, followed by RNA extraction (Quick RNA 96 kit, Zymo Research) (PBS- no treatment control, Negative control PPMO 9). RT-PCR was performed using Maxima™ H Minus cDNA Synthesis Master Mix (Thermo Scientific, M1662) followed by qPCR using PrimeTime™ Gene Expression Master Mix (IDT, 1055772). Fast cycling conditions were applied according to the manufacturer’s instructions. PrimeTime qPCR Probe Assays (IDT)
are detailed in the table below. Analysis was performed with the Design and Analysis Software Version 2.6 for QuantStudio 6/7 Pro systems. RT-PCR and qPCR conditions were as described in Example 8. Table 14. PPMO compounds used in dose experiments.
Example 13 – Exon-skipping of human DNM2 by PPMOs in healthy myotubes. [1163] Healthy human myoblast cells were allowed to differentiate in differentiation media (5% HS DMEM-low glucose) for three days followed by treatment for three days with PPMOs diluted in Opti-MEM to the indicated doses (10, 20, 30 or 15 µM), free uptake (untreated control, NTC control (Gene Tools), negative- no template). The NTC is a 25mer PMO that is not complimentary to any portion of DMN2. [1164] RT-PCR was performed on 50 ng of the RNA template using Superscript III One-Step RT-PCR System with Platinum® Taq DNA Polymerase (ThermoFisher Scientific, Australia). Cycling conditions are listed in Table below. They include 55 °C for 30 minutes, 94 °C for 2 minutes followed by various cycles of 94 °C for 30 seconds, 60 °C for 30 seconds and 68 °C for 1 minute. [1165] Densitometry was performed and the exon skipping was calculated as a ratio of skipped transcriptions to total transcripts. “–“ indicates <20% exon skipping; “+” indicates between 20- 40% exon skipping; “++” indicates between 40-80% exon skipping; and “+++” indicates >80% exon skipping. The results for exon-skipping of human DNM2 by PPMOs in normal myotubes are shown in Table 15 and FIGs.13A-13D and 14A-14D. The PPMOs used in this example are
of Formula (VI) below having the sequences indicated in Table 15, where R5G is a cell penetrating peptide having the sequence Arg–Arg–Arg–Arg–Arg–Gly–, where z is 5 in Formula (VI); and R7G is a cell penetrating peptide having the sequence Arg–Arg–Arg–Arg–Arg–Arg– Arg–Gly–, where z is 7 in Formula (VI).
Table 15. Summary of exon-skipping of human DNM2 by PPMOs in healthy myotubes.
R7G is a peptide Arg–Arg–Arg–Arg–Arg–Arg–Arg–Gly–. [1166] In certain embodiments, the PPMOs described herein are of Formula (VI):
(VI), or a pharmaceutically acceptable salt thereof, wherein t, n, and R2 are as described herein. Example 14 – Protein reduction of human DNM2 by PPMOs in healthy myotubes. [1167] Normal human myoblast cells were allowed to differentiate in differentiation media (5% HS DMEM-low glucose) for three days followed by treatment for three days with PPMOs diluted in Opti-MEM at 15 µM, free uptake (untreated control, NTC control (Gene Tools), negative- no template). The NTC is a 25mer PMO that is not complimentary to any portion of DMN2. [1168] Approximately 25 µg total protein was used for each sample and transferred onto PDVF membrane using iBlot™ 2 Gel Transfer Device (Thermo Fisher). DNM2 was detected with polyclonal anti-DNM2 (Abcam) applied at a dilution of 1:2000 for 24 hours at 4 °C. β tubulin was detected with polyclonal anti β Tubulin (Thermo Fisher) at a dilution of 1:2500 for 24 hours at 4 °C. HRP-labelled anti-rabbit secondary antibody was applied respectively for 1 hour at room temperature. The blots were detected using Immobilon western chemiluminescent HRP
substrate (Merck Millipore) and images were captured using a Fusion FX gel documentation system (Vilber Lourmat) with FusionCapt Advance software. Image J software (NIH) was used for densitometric analysis. [1169] Densitometry was performed and relative protein quantities were calculated by normalizing to β tubulin. The relative change from the NTC samples is used to show the protein reduction of human DNM2. “–“ indicates no protein reduction; “+” indicates between <20% protein reduction; “++” indicates between 20-50% protein reduction; and “+++” indicates >50% protein reduction. The results for protein reduction of human DNM2 by PPMOs in normal myotubes. The results for protein reduction of human DNM2 by PPMOs in normal myotubes are shown in Table 16 and FIGs.15A-15B and 16A-16B. The PPMOs used in this example are of Formula (VI) below having the sequences indicated in Table 16, where R5G is a cell penetrating peptide having the sequence Arg–Arg–Arg–Arg–Arg–Gly–, where z is 5 in Formula (VI); and R7G is a cell penetrating peptide having the sequence Arg–Arg–Arg–Arg–Arg–Arg–Arg–Gly–, where z is 7 in Formula (VI). Table 16. Protein reduction of human DNM2 by PPMOs in healthy myotubes.
R7G is a peptide Arg–Arg–Arg–Arg–Arg–Arg–Arg–Gly–. Table 17. DNM2 Target Regions and Targeting Sequences.
Claims
CLAIMS WHAT IS CLAIMED IS: 1. An antisense oligomer comprising a non-natural chemical backbone and a targeting sequence 13 to 30 bases in length that is complementary to a target region within a pre- mRNA of human dynamin 2 (DNM2) gene represented by SEQ ID NO: 1, wherein the target region is within an exon internal region, an intron internal region, an intron-exon junction, or an exon-intron junction of the human DNM2 gene pre-mRNA.
2. The antisense oligomer of claim 1, wherein the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 6 (SEQ ID NO: 3), exon 7 (SEQ ID NO: 4), exon 8 (SEQ ID NO: 5), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), exon 13 (SEQ ID NO: 8), exon 16 (SEQ ID NO: 9), and exon 19 (SEQ ID NO: 10).
3. The antisense oligomer of claim 1 or 2, wherein the target region is within an exon internal region of the human DNM2 gene pre-mRNA selected from exon 2 (SEQ ID NO: 2), exon 9 (SEQ ID NO: 6), exon 12 (SEQ ID NO: 7), and exon 13 (SEQ ID NO: 8).
4. The antisense oligomer of claim 1, wherein the target region is within an intron-exon junction or an exon-intron junction region of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 5/exon 6 junction, exon 6/intron 6 junction, intron 6/exon 7 junction, exon 7/intron 7 junction, intron 7/exon 8 junction, exon 8/intron 8 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, exon 13/intron 13 junction, intron 15/exon 16 junction, exon 16/intron 16 junction, intron 18/exon 19 junction, and exon 19/intron 19 junction.
5. The antisense oligomer of claim 1 or 4, wherein the target region is within an intron-exon junction or an exon-intron junction of the human DNM2 gene pre-mRNA selected from intron 1/exon 2 junction, exon 2/intron 2 junction, intron 8/exon 9 junction, exon 9/intron 9 junction, intron 11/exon 12 junction, exon 12/intron 12 junction, intron 12/exon 13 junction, and exon 13/intron 13 junction.
6. The antisense oligomer of claim 1, wherein the target region is within an intron internal region of the human DNM2 gene pre-mRNA selected from intron 2 (SEQ ID NO: 11) and intron 12 (SEQ ID NO: 12).
7. The antisense oligomer of any one of claims 1-6, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5),
DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4- 28), DNM2 H2D(-5-29), DNM2 H2D(-6-30), DNM2 H9A(-12+13), DNM2 H9A(-5+20), DNM2 H9A(+1+25),DNM2 H9A(+6+30), DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), DNM2 H9A(+48+67), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), DNM2 H9D(+3-22), DNM2 H12A(-13+12), DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), DNM2 H12D(+15-10), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2
H12A(+43+67), DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), DNM2 H13A(+9+28), DNM2 H13A(+16+40), DNM2 H13D(+22-3), or DNM2 H13D(+7-18).
8. The antisense oligomer of claim 7, wherein the targeting sequence is selected from SEQ ID NOs: 67-326.
9. The antisense oligomer of any one of claims 1-6, wherein the target region is within the exon internal region of exon 2 (SEQ ID NO: 2), the intron 1/exon 2 junction, the exon 2/intron 2 junction, or the intron internal region of intron 2 (SEQ ID NO: 11) of the human DNM2 gene pre-mRNA.
10. The antisense oligomer of any one of claims 1-3 or 9, wherein the target region of the human DNM2 gene pre-mRNA is within the 21st nucleotide to the 47th nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2).
11. The antisense oligomer of claim 10, wherein the target region is a region within SEQ ID NO: 13.
12. The antisense oligomer of any one of claims 1-3 or 9, wherein the target region of the human DNM2 gene pre-mRNA is within the 23rd nucleotide to the 51st nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2).
13. The antisense oligomer of claim 12, wherein the target region is a region within SEQ ID NO: 14.
14. The antisense oligomer of any one of claims 1-3 or 9, wherein the target region of the human DNM2 gene pre-mRNA is within the 46th nucleotide to the 72nd nucleotide of exon 2 measured from the 5′ end of exon 2 (SEQ ID NO: 2).
15. The antisense oligomer of claim 14, wherein the target region is a region within SEQ ID NO: 15.
17. The antisense oligomer of claim 16, wherein the target region is a region within SEQ ID NO: 16.
18. The antisense oligomer of any one of claims 1, 4-5, 9, or 16-17, wherein the target region of the human DNM2 gene pre-mRNA is within the 13th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 14th nucleotide of intron 2 measured from the 3′ end of exon 2.
19. The antisense oligomer of claim 18, wherein the target region is a region within SEQ ID NO: 17.
20. The antisense oligomer of any one of claims 1, 4-5, 9, or 16-17, wherein the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 19th nucleotide of intron 2 measured from the 3′ end of exon 2.
21. The antisense oligomer of claim 20, wherein the target region is a region within SEQ ID NO: 18.
22. The antisense oligomer of any one of claims 1, 4-5, 9, 16-17, or 20-21, wherein the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 18th nucleotide of intron 2 measured from the 3′ end of exon 2.
23. The antisense oligomer of claim 22, wherein the target region is a region within SEQ ID NO: 19.
24. The antisense oligomer of any one of claims 1, 4-5, 9, or 16-17, wherein the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23rd nucleotide of intron 2 measured from the 3′ end of exon 2.
25. The antisense oligomer of claim 24, wherein the target region is a region within SEQ ID NO: 20.
26. The antisense oligomer of any one of claims 1, 4-5, 9, 16-17, or 24-25, wherein the target region of the human DNM2 gene pre-mRNA is within the 4th nucleotide of exon 2 measured from the 3′ end of exon 2 (SEQ ID NO: 2) to the 23rd nucleotide of intron 2 measured from the 3′ end of exon 2.
27. The antisense oligomer of claim 26, wherein the target region is a region within SEQ ID NO: 21.
28. The antisense oligomer of any one of claims 1, 6, or 9, wherein the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 30th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2).
29. The antisense oligomer of claim 28, wherein the target region is a region within SEQ ID NO: 22.
30. The antisense oligomer of any one of claims 1, 6, 9, or 28-29, wherein the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 29th nucleotide of intron 2 (SEQ ID NO: 11) measured from the 3′ end of exon 2 (SEQ ID NO: 2).
31. The antisense oligomer of claim 30, wherein the target region is a region within SEQ ID NO: 23.
32. The antisense oligomer of any one of claims 1-31, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15), DNM2 H2A(-7+17), DNM2 H2A(+3+27), DNM2 H2A(+15+37), DNM2 H2A(+21+45), DNM2 H2A(+23+47), DNM2 H2A(+27+51), DNM2 H2A(+37+61), DNM2 H2A(+46+70), DNM2 H2A (+48+72), DNM2 H2D(+20-5), DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3- 27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30).
33. The antisense oligomer of claim 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2D (+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23) , DNM2 H2D(+1-24), DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3- 27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30).
34. The antisense oligomer of any one of claims 1-3, 7, 9-11, or 32, wherein the target region is DNM2 H2A(+21+45) or DNM2 H2A(+23+47).
35. The antisense oligomer of any one of claims 1-3, 7, 9, 12-13, or 32, wherein the target region is DNM2 H2A(+23+47) or DNM2 H2A(+27+51).
36. The antisense oligomer of any one of claims 1-3, 7, 9, 14-15, or 32, wherein the target region is DNM2 H2A(+46+70) or DNM2 H2A(+48+72).
37. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, or 32, wherein the target region is DNM2 H2D(+13-12), DNM2 H2D(+12-13), DNM2 H2D(+11-14), DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), DNM2
H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23), or DNM2 H2D(+1-24).
38. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-19, 32, or 37, wherein the target region is DNM2 H2D (+13-12), DNM2 H2D(+12-13), or DNM2 H2D(+11-14).
39. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 20-21, 32, or 37, wherein the target region is DNM2 H2D(+10-15), DNM2 H2D(+9-16), DNM2 H2D(+8-17), DNM2 H2D(+7-18), or DNM2 H2D(+6-19).
40. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 20-23, 32, 37, or 39, wherein the target region is DNM2 H2D(+9-16), DNM2 H2D(+8-17), or DNM2 H2D(+7- 18).
41. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 20-23, 32, 35, 37, or 39- 40, wherein the target region is DNM2 H2D(+8-17).
42. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 24-25, 32, or 37, wherein the target region is DNM2 H2D(+6-19), DNM2 H2D(+5-20), DNM2 H2D(+4-21), DNM2 H2D(+3-22), DNM2 H2D(+2-23).
43. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42, wherein the target region is DNM2 H2D(+4-21), DNM2 H2D(+3-22), or DNM2 H2D(+2- 23).
44. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42-43, wherein the target region is DNM2 H2D(+3-22).
45. The antisense oligomer of any one of claims 1, 6-7, 9, 28-29, or 32, wherein the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), DNM2 H2D(-5-29), or DNM2 H2D(-6-30).
46. The antisense oligomer of any one of claims 1, 6-7, 9, 28-32, or 45, wherein the target region is DNM2 H2D(-1-25), DNM2 H2D(-2-26), DNM2 H2D(-3-27), DNM2 H2D(-4-28), or DNM2 H2D(-5-29).
47. The antisense oligomer of any one of claims 1, 4-5, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A (-10+15).
48. The antisense oligomer of any one of claims 1, 4-5, or 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(-7+17).
49. The antisense oligomer of any one of claims 1-3, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+3+27).
50. The antisense oligomer of any one of claims 1-3, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+15+37).
51. The antisense oligomer of any one of claims 1-3, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2A(+37+61).
52. The antisense oligomer of any one of claims 1, 4-5, 7, 9, or 32, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H2D(+20-5).
53. The antisense oligomer of any one of claims 1-32, wherein the targeting sequence is selected from SEQ ID NOs: 67-96.
54. The antisense oligomer of any one of claims 1, 7, or 32-33, wherein the targeting sequence is selected from SEQ ID NOs: 78-96.
55. The antisense oligomer of any one of claims 1-3, 7, 9-11, 32, or 34, wherein the targeting sequence is selected from SEQ ID NOs: 71-72.
56. The antisense oligomer of any one of claims 1-3, 7, 9, 12-13, 32, or 35, wherein the targeting sequence is selected from SEQ ID NOs: 72-73.
57. The antisense oligomer of any one of claims 1-3, 7, 9, 14-15, 32, or 36, wherein the targeting sequence is selected from SEQ ID NOs: 75-76.
58. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 32, or 37, wherein the targeting sequence is selected from SEQ ID NOs: 78-90.
59. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-19, 32, or 37-38, wherein the targeting sequence is selected from SEQ ID NOs: 78-80.
60. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 20-21, 32, 37, or 39, wherein the targeting sequence is selected from SEQ ID NOs: 81-85.
61. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 20-23, 32, 37, or 39-40, wherein the targeting sequence is selected from SEQ ID NOs: 82-84.
62. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 20-23, 32, 35, 37, or 39- 41, wherein the targeting sequence is SEQ ID NO: 83.
63. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 24-25, 32, 37, or 42, wherein the targeting sequence is selected from SEQ ID NOs: 85-89.
64. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42-43, wherein the targeting sequence is selected from SEQ ID NOs: 87-89.
65. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 16-17, 24-27, 32, 37, or 42-44, wherein the targeting sequence is SEQ ID NO: 88.
66. The antisense oligomer of any one of claims 1, 6-7, 9, 28-29, 32, or 45, wherein the targeting sequence is selected from SEQ ID NOs: 91-96.
67. The antisense oligomer of any one of claims 1, 6-7, 9, 28-32, or 45-46, wherein the targeting sequence is selected from SEQ ID NOs: 91-95.
68. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 32, or 47, wherein the targeting sequence is SEQ ID NO: 67.
69. The antisense oligomer of any one of claims 1, 4-5, or 7, 9, 32, or 48, wherein the targeting sequence is SEQ ID NO: 68.
70. The antisense oligomer of any one of claims 1-3, 7, 9, 32, or 49, wherein the targeting sequence is SEQ ID NO: 69.
71. The antisense oligomer of any one of claims 1-3, 7, 9, 32, or 50, wherein the targeting sequence is SEQ ID NO: 70.
72. The antisense oligomer of any one of claims 1-3, 7, 9, 32, or 51, wherein the targeting sequence is SEQ ID NO: 74.
73. The antisense oligomer of any one of claims 1, 4-5, 7, 9, 32, or 52, wherein the targeting sequence is SEQ ID NO: 77.
74. The antisense oligomer of any one of claims 1-5, wherein the target region is within the exon internal region of exon 9 (SEQ ID NO: 6), the intron 8/exon 9 junction, or the exon 9/intron 9 junction of the human DNM2 gene pre-mRNA.
75. The antisense oligomer of any one of claims 1-3 or 74, wherein the target region of the human DNM2 gene pre-mRNA is within the 36th nucleotide to the 61st nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6).
76. The antisense oligomer of claim 75, wherein the target region is a region within SEQ ID NO: 24.
77. The antisense oligomer of any one of claims 1-3 or 74, wherein the target region of the human DNM2 gene pre-mRNA is within the 42nd nucleotide to the 68th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6).
78. The antisense oligomer of claim 77, wherein the target region is a region within SEQ ID NO: 25.
79. The antisense oligomer of any one of claims 1-3, 74, or 77-78, wherein the target region of the human DNM2 gene pre-mRNA is within the 43rd nucleotide to the 68th nucleotide of exon 9 measured from the 5′ end of exon 9 (SEQ ID NO: 6).
80. The antisense oligomer of claim 79, wherein the target region is a region within SEQ ID NO: 26.
81. The antisense oligomer of any one of claims 1, 4-5, or 74, wherein the target region of the human DNM2 gene pre-mRNA is within the 24th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15th nucleotide of intron 9 measured from the 3′ end of exon 9.
82. The antisense oligomer of claim 81, wherein the target region is a region within SEQ ID NO: 27.
83. The antisense oligomer of any one of claims 1, 4-5, 74, or 81-82, wherein the target region of the human DNM2 gene pre-mRNA is within the 24th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 5th nucleotide of intron 9 measured from the 3′ end of exon 9.
84. The antisense oligomer of claim 83, wherein the target region is a region within SEQ ID NO: 28.
85. The antisense oligomer of any one of claims 1, 4-5, 74, or 81-82, wherein the target region of the human DNM2 gene pre-mRNA is within the 19th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 10th nucleotide of intron 9 measured from the 3′ end of exon 9.
86. The antisense oligomer of claim 85, wherein the target region is a region within SEQ ID NO: 29.
87. The antisense oligomer of any one of claims 1, 4-5, 74, or 81-82, wherein the target region of the human DNM2 gene pre-mRNA is within the 14th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 15th nucleotide of intron 9 measured from the 3′ end of exon 9.
88. The antisense oligomer of claim 87, wherein the target region is a region within SEQ ID NO: 30.
89. The antisense oligomer of any one of claims 1, 4-5, or 74, wherein the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22nd nucleotide of intron 9 measured from the 3′ end of exon 9.
90. The antisense oligomer of claim 89, wherein the target region is a region within SEQ ID NO: 31.
91. The antisense oligomer of any one of claims 1, 4-5, 74, or 89-90, wherein the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 20th nucleotide of intron 9 measured from the 3′ end of exon 9.
92. The antisense oligomer of claim 91, wherein the target region is a region within SEQ ID NO: 32.
93. The antisense oligomer of any one of claims 1, 4-5, 74, or 89-92, wherein the target region of the human DNM2 gene pre-mRNA is within the 9th nucleotide of exon 9
measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 18th nucleotide of intron 9 measured from the 3′ end of exon 9.
94. The antisense oligomer of claim 93, wherein the target region is a region within SEQ ID NO: 33.
95. The antisense oligomer of any one of claims 1, 4-5, 74, or 89-90, wherein the target region of the human DNM2 gene pre-mRNA is within the 8th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 21st nucleotide of intron 9 measured from the 3′ end of exon 9.
96. The antisense oligomer of claim 95, wherein the target region is a region within SEQ ID NO: 34.
97. The antisense oligomer of any one of claims 1, 4-5, 74, or 89-90, wherein the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide of exon 9 measured from the 3′ end of exon 9 (SEQ ID NO: 6) to the 22nd nucleotide of intron 9 measured from the 3′ end of exon 9.
98. The antisense oligomer of claim 97, wherein the target region is a region within SEQ ID NO: 35.
99. The antisense oligomer of any one of claims 1-5, 7, or 74-98, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13), DNM2 H9A(-5+20), DNM2 H9A(+1+25),DNM2 H9A(+6+30), DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), DNM2 H9A(+48+67), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22).
100. The antisense oligomer of claim 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40), DNM2 H9A(+25+49), DNM2 H9A(+36+60), DNM2 H9A(+37+61), DNM2 H9A(+43+67), DNM2 H9A(+42+66), DNM2 H9A(+44+68), DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2
H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), DNM2 H9D(+10-15), DNM2 H9D(+9-16), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+6-19), DNM2 H9D(+5-20), DNM2 H9D(+4-21), or DNM2 H9D(+3-22).
101. The antisense oligomer of any one of claims 1-5, 7, 74-76, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+36+60) or DNM2 H9A(+37+61).
102. The antisense oligomer of any one of claims 1-3, 7, 74, 77-78, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+42+66), DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), DNM2 H9A(+44+68), DNM2 H9A(+45+67), DNM2 H9A (+46+67), DNM2 H9A(+47+67), or DNM2 H9A(+48+67).
103. The antisense oligomer of any one of claims 1-3, 7, 74, 77-80, 99, or 102, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+62), DNM2 H9A(+43+63), DNM2 H9A(+43+64), DNM2 H9A(+43+65), DNM2 H9A(+43+66), DNM2 H9A(+43+67), DNM2 H9A(+44+67), or DNM2 H9A(+44+68).
104. The antisense oligomer of any one of claims 1-3, 7, 74, 77-80, 99, or 102-103, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+43+67).
105. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 81-82, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), DNM2 H9D(+20-5), DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), DNM2 H9D(+15-10), DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), or DNM2 H9D(+10-15).
106. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 81-84, 99, or 105, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+24-1), DNM2 H9D(+23-2), DNM2 H9D(+22-3), DNM2 H9D(+21-4), or DNM2 H9D(+20-5).
107. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 81-82, 85-86, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+19-6), DNM2 H9D (+18-7), DNM2 H9D(+17-8), DNM2 H9D(+16-9), or DNM2 H9D(+15-10).
108. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 81-82, 87-88, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+14-11), DNM2 H9D(+13-12), DNM2 H9D(+12-13), DNM2 H9D(+11-14), or DNM2 H9D(+10-15).
109. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-90, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22).
110. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-92, 99, or 109, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), or DNM2 H9D(+5-20).
111. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-94, 99, or 109-110, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+9-16), DNM2 H9D(+8-12), DNM2 H9D(+8-13), DNM2 H9D(+8-14), DNM2 H9D(+8-15), DNM2 H9D(+8-17), or DNM2 H9D(+7-18).
112. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-90, 95-96, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17), DNM2 H9D(+7-18), DNM2 H9D(+7-17), DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), or DNM2 H9D(+4-21).
113. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-96, 99, or 109-112, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+8-17).
114. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-90, 97-99, or 109, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9D(+6-19), DNM2 H9D(+6-17), DNM2 H9D(+5-17), DNM2 H9D(+5-20), DNM2 H9D(+4-17), DNM2 H9D(+4-21), DNM2 H9D(+3-17), or DNM2 H9D(+3-22).
115. The antisense oligomer of any one of claims 1, 4-5, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-12+13).
116. The antisense oligomer of any one of claims 1, 4-5, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(-5+20).
117. The antisense oligomer of any one of claims 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+1+25).
118. The antisense oligomer of any one of claims 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+6+30).
119. The antisense oligomer of any one of claims 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+16+40).
120. The antisense oligomer of any one of claims 1-3, 7, 74, or 99, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H9A(+25+49).
121. The antisense oligomer of any one of claims 1-5, 7, or 74-99, wherein the targeting sequence is selected from SEQ ID NOs: 158-209.
122. The antisense oligomer of any one of claims 1, 7, or 99-100, wherein the targeting sequence is selected from SEQ ID NOs: 162-166, 172, 174, 179-194, 199-200, 205, 207, and 209.
123. The antisense oligomer of any one of claims 1-5, 7, 74-76, 99, or 101, wherein the targeting sequence is selected from SEQ ID NOs: 164-165.
124. The antisense oligomer of any one of claims 1-3, 7, 74, 77-78, 99, or 102, wherein the targeting sequence is selected from SEQ ID NOs: 166-178.
125. The antisense oligomer of any one of claims 1-3, 7, 74, 77-80, 99, or 102-103, wherein the targeting sequence is selected from SEQ ID NOs: 167-174.
126. The antisense oligomer of any one of claims 1-3, 7, 74, 77-80, 99, or 102-104, wherein the targeting sequence is SEQ ID NO: 172.
127. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 81-82, 99, or 105, wherein the targeting sequence is selected from SEQ ID NOs: 179-193.
128. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 81-84, 99, or 105-106, wherein the targeting sequence is selected from SEQ ID NOs: 179-183.
129. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 81-82, 85-86, 99, or 107, wherein the targeting sequence is selected from SEQ ID NOs: 184-188.
130. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 81-82, 87-88, 99, or 108, wherein the targeting sequence is selected from SEQ ID NOs: 189-193.
131. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-90, 99, or 109, wherein the targeting sequence is selected from SEQ ID NOs: 194-209.
132. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-92, 99, or 109-110, wherein the targeting sequence is selected from SEQ ID NOs: 194-205.
133. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-94, 99, or 109-111, wherein the targeting sequence is selected from SEQ ID NOs: 194-200.
134. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-90, 95-96, 99, or 112, wherein the targeting sequence is selected from SEQ ID NOs: 199-207.
135. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-96, 99, or 109-113, wherein the targeting sequence is SEQ ID NO: 199.
136. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 89-90, 97-99, 109, or 114, wherein the targeting sequence is selected from SEQ ID NOs: 202-209.
137. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 99, or 115, wherein the targeting sequence is SEQ ID NO: 158.
138. The antisense oligomer of any one of claims 1, 4-5, 7, 74, 99, or 116, wherein the targeting sequence is SEQ ID NO: 159.
139. The antisense oligomer of any one of claims 1-3, 7, 74, 99, or 117, wherein the targeting sequence is SEQ ID NO: 160.
140. The antisense oligomer of any one of claims 1-3, 7, 74, 99, or 118, wherein the targeting sequence is SEQ ID NO: 161.
141. The antisense oligomer of any one of claims 1-3, 7, 74, 99, or 119, wherein the targeting sequence is SEQ ID NO: 162.
142. The antisense oligomer of any one of claims 1-3, 7, 74, 99, or 120, wherein the targeting sequence is SEQ ID NO: 163.
143. The antisense oligomer of any one of claims 1-5, wherein the target region is within the exon internal region of exon 12 (SEQ ID NO: 7), the intron 11/exon 12 junction, or the exon 12/intron 12 junction of the human DNM2 gene pre-mRNA.
144. The antisense oligomer of any one of claims 1-3 or 143, wherein the target region of the human DNM2 gene pre-mRNA is within the 4th nucleotide to the 31st nucleotide measured of exon 12 from the 5′ end of exon 12 (SEQ ID NO: 7).
145. The antisense oligomer of claim 144, wherein the target region is a region within SEQ ID NO: 36.
146. The antisense oligomer of any one of claims 1-3 or 143, wherein the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 44th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
147. The antisense oligomer of claim 146, wherein the target region is a region within SEQ ID NO: 37.
148. The antisense oligomer of any one of claims 1-3, 143, or 146-147, wherein the target region of the human DNM2 gene pre-mRNA is within the 10th nucleotide to the 38th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
149. The antisense oligomer of claim 148, wherein the target region is a region within SEQ ID NO: 38.
150. The antisense oligomer of any one of claims 1-3, 143, or 146-147, wherein the target region of the human DNM2 gene pre-mRNA is within the 16th nucleotide to the 44th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
151. The antisense oligomer of claim 150, wherein the target region is a region within SEQ ID NO: 39.
152. The antisense oligomer of any one of claims 1-3, 143, or 146-151, wherein the target region of the human DNM2 gene pre-mRNA is within the 14th nucleotide to the 42nd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
153. The antisense oligomer of claim 152, wherein the target region is a region within SEQ ID NO: 40.
154. The antisense oligomer of any one of claims 1-3 or 143, wherein the target region of the human DNM2 gene pre-mRNA is within the 25th nucleotide to the 54th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
155. The antisense oligomer of claim 154, wherein the target region is a region within SEQ ID NO: 41.
156. The antisense oligomer of any one of claims 1-3, 143, or 154-155, wherein the target region of the human DNM2 gene pre-mRNA is within the 25th nucleotide to the 53rd nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
157. The antisense oligomer of claim 156, wherein the target region is a region within SEQ ID NO: 42.
158. The antisense oligomer of any one of claims 1-3, 143, or 154-155, wherein the target region of the human DNM2 gene pre-mRNA is within the 26th nucleotide to the 54th nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
159. The antisense oligomer of claim 158, wherein the target region is a region within SEQ ID NO: 43.
160. The antisense oligomer of any one of claims 1-3 or 143, wherein the target region of the human DNM2 gene pre-mRNA is within the 36th nucleotide to the 61st nucleotide of exon 12 measured from the 5′ end of exon 12 (SEQ ID NO: 7).
161. The antisense oligomer of claim 160, wherein the target region is a region within SEQ ID NO: 44.
162. The antisense oligomer of any one of claims 1, 4-5, or 143, wherein the target region of the human DNM2 gene pre-mRNA is within the 22nd nucleotide of exon 12 measured from the 3′ end of exon 12 (SEQ ID NO: 7) to the 5th nucleotide of intron 12 measured from the 3′ end of exon 12.
163. The antisense oligomer of claim 162, wherein the target region is a region within SEQ ID NO: 45.
164. The antisense oligomer of any one of claims 1-5, 7, or 143-163, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-13+12), DNM2 H12A(- 3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38),
DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), or DNM2 H12D(+15- 10).
165. The antisense oligomer of any one of claim 164, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H12A(-3+22), DNM2 H12A(+4+28), DNM2 H12A(+7+31), DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44), DNM2 H12A(+25+49), DNM2 H12A(+29+53), DNM2 H12A(+30+54), DNM2 H12A(+36+60), DNM2 H12A(+37+61), DNM2 H12A(+43+67), DNM2 H12D(+22-3), DNM2 H12D(+20-5), or DNM2 H12D(+15-10).
166. The antisense oligomer of any one of claims 1-3, 7, 143-145, or 164, wherein the target region is DNM2 H12A(+4+28) or DNM2 H12A(+7+31).
167. The antisense oligomer of any one of claims 1-3, 7, 143, 146-147, or 164, wherein the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), DNM2 H12A(+20+44).
168. The antisense oligomer of any one of claims 1-3, 7, 143, 146-149, 164, or 167, wherein the target region is DNM2 H12A(+10+34), DNM2 H12A(+11+35), DNM2 H12A(+12+36), DNM2 H12A(+13+37), or DNM2 H12A(+14+38).
169. The antisense oligomer of any one of claims 1-3, 7, 143, 146-147, 150-151, 164, or 167, wherein the target region is DNM2 H12A(+16+40), DNM2 H12A(+17+41), DNM2 H12A(+18+42), DNM2 H12A(+19+43), or DNM2 H12A(+20+44).
170. The antisense oligomer of any one of claims 1-3, 7, 143, 146-153, 164, or 167-169, wherein the target region is DNM2 H12A(+14+38), DNM2 H12A(+15+39), DNM2 H12A(+16+40), DNM2 H12A(+17+41), or DNM2 H12A(+18+42).
171. The antisense oligomer of any one of claims 1-3, 7, 143, 154-155, or 164, wherein the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2
H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54).
172. The antisense oligomer of any one of claims 1-3, 7, 143, 154-157, 164, or 171, wherein the target region is DNM2 H12A(+25+44), DNM2 H12A(+25+45), DNM2 H12A(+25+46), DNM2 H12A(+25+47), DNM2 H12A(+25+48), DNM2 H12A(+25+49), DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), or DNM2 H12A(+29+53).
173. The antisense oligomer of any one of claims 1-3, 7, 143, 154-155, 158-159, 164, or 171, wherein the target region is DNM2 H12A(+26+49), DNM2 H12A(+27+49), DNM2 H12A(+28+49), DNM2 H12A(+29+49), DNM2 H12A(+29+53), DNM2 H12A(+30+49), or DNM2 H12A(+30+54).
174. The antisense oligomer of any one of claims 1-3, 7, 143, 154-155, 158-159, 164, 171, or 173, wherein the target region is DNM2 H12A(+25+49).
175. The antisense oligomer of any one of claims 1-3, 7, 143, 160-161, or 164, wherein the target region is DNM2 H12A(+36+60) or DNM2 H12A(+37+61).
176. The antisense oligomer of any one of claims 1, 4-5, 7, 143, or 162-164, wherein the target region is DNM2 H12D(+22-3) or DNM2 H12D(+20-5).
177. The antisense oligomer of any one of claims 1, 4-5, 7, 143, or 164, wherein the target region is DNM2 H12A(-13+12).
178. The antisense oligomer of any one of claims 1, 4-5, 7, 143, or 164, wherein the target region is DNM2 H12A(-3+22).
179. The antisense oligomer of any one of claims 1-3, 7, 143, or 164, wherein the target region is DNM2 H12A(+43+67).
180. The antisense oligomer of any one of claims 1, 4-5, 7, 143, or 164, wherein the target region is DNM2 H12D(+15-10).
181. The antisense oligomer of any one of claims 1-5, 7, or 143-164, wherein the targeting sequence is selected from SEQ ID NOs: 210-243.
182. The antisense oligomer of any one of claims 1, 7, or 164-165, wherein the targeting sequence is selected from SEQ ID NOs: 211-224, 230, 235, 237-238, and 240-243.
183. The antisense oligomer of any one of claims 1-3, 7, 143-145, 164, or 166, wherein the targeting sequence is selected from SEQ ID NOs: 212-213.
184. The antisense oligomer of any one of claims 1-3, 7, 143, 146-147, 164, or 167, wherein the targeting sequence is selected from SEQ ID NOs: 214-224.
185. The antisense oligomer of any one of claims 1-3, 7, 143, 146-149, 164, or 167-168, wherein the targeting sequence is selected from SEQ ID NOs: 214-218.
186. The antisense oligomer of any one of claims 1-3, 7, 143, 146-147, 150-151, 164, or 169, wherein the targeting sequence is selected from SEQ ID NOs: 220-224.
187. The antisense oligomer of any one of claims 1-3, 7, 143, 146-153, 164, 167-170, wherein the targeting sequence is selected from SEQ ID NOs: 218-222.
188. The antisense oligomer of any one of claims 1-3, 7, 143, 154-155, 164, or 171, wherein the targeting sequence is selected from SEQ ID NOs: 225-237.
189. The antisense oligomer of any one of claims 1-3, 7, 143, 154-157, 164, or 171-172, wherein the targeting sequence is selected from SEQ ID NOs: 225-235.
190. The antisense oligomer of any one of claims 1-3, 7, 143, 154-155, 158-159, 164, or 173, wherein the targeting sequence is selected from SEQ ID NOs: 231-237.
191. The antisense oligomer of any one of claims 1-3, 7, 143, 154-155, 158-159, 164, or 174, wherein the targeting sequence is SEQ ID NO: 230.
192. The antisense oligomer of any one of claims 1-3, 7, 143, 160-161, 164, or 175, wherein the targeting sequence is selected from SEQ ID NOs: 238-239.
193. The antisense oligomer of any one of claims 1, 4-5, 7, 143, 162-164, or 176, wherein the targeting sequence is selected from SEQ ID NOs: 241-242.
194. The antisense oligomer of any one of claims 1, 4-5, 7, 143, 164, or 177, wherein the targeting sequence is SEQ ID NO: 210.
195. The antisense oligomer of any one of claims 1, 4-5, 7, 143, 164, or 178, wherein the targeting sequence is SEQ ID NO: 211.
196. The antisense oligomer of any one of claims 1-3, 7, 143, 164, or 179, wherein the targeting sequence is SEQ ID NO: 240.
197. The antisense oligomer of any one of claims 1, 4-5, 7, 143, 164, or 180, wherein the targeting sequence is SEQ ID NO: 243.
198. The antisense oligomer of any one of claims 1-6, wherein the target region is within the exon internal region of exon 13 (SEQ ID NO: 8), the intron 12/exon 13 junction, the exon 13/intron 13 junction, or the intron internal region of intron 12 (SEQ ID NO: 12).
199. The antisense oligomer of any one of claims 1, 4-6, or 198, wherein the target region of the human DNM2 gene pre-mRNA is within the 27th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12th nucleotide of exon 13 measured from the 5′ end of exon 13.
200. The antisense oligomer of claim 199, wherein the target region is a region within SEQ ID NO: 46.
201. The antisense oligomer of any one of claims 1, 4-6, or 198-200, wherein the target region of the human DNM2 gene pre-mRNA is within the 27th nucleotide of intron 12
measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 2nd nucleotide of exon 13 measured from the 5′ end of exon 13.
202. The antisense oligomer of claim 201, wherein the target region is a region within SEQ ID NO: 47.
203. The antisense oligomer of any one of claims 1, 4-6, or 198-200, wherein the target region of the human DNM2 gene pre-mRNA is within the 22nd nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 7th nucleotide of exon 13 measured from the 5′ end of exon 13.
204. The antisense oligomer of claim 203, wherein the target region is a region within SEQ ID NO: 48.
205. The antisense oligomer of any one of claims 1, 4-6, or 198-200, wherein the target region of the human DNM2 gene pre-mRNA is within the 17th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 12th nucleotide of exon 13 measured from the 5′ end of exon 13.
206. The antisense oligomer of claim 205, wherein the target region is a region within SEQ ID NO: 49.
207. The antisense oligomer of any one of claims 1, 4-5, or 198, wherein the target region of the human DNM2 gene pre-mRNA is within the 11th nucleotide of intron 12 (SEQ ID NO: 12) measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24th nucleotide of exon 13 measured from the 5′ end of exon 13.
208. The antisense oligomer of claim 207, wherein the target region is a region within SEQ ID NO: 50.
209. The antisense oligomer of any one of claims 1, 4-5, 198, or 207-208, wherein the target region of the human DNM2 gene pre-mRNA is within the 11th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 18th nucleotide of exon 13 measured from the 5′ end of exon 13.
210. The antisense oligomer of claim 209, wherein the target region is a region within SEQ ID NO: 51.
211. The antisense oligomer of any one of claims 1, 4-5, 198, or 207-208, wherein the target region of the human DNM2 gene pre-mRNA is within the 5th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 24th nucleotide exon 13 measured from the 5′ end of exon 13.
212. The antisense oligomer of claim 211, wherein the target region is a region within SEQ ID NO: 52.
213. The antisense oligomer of any one of claims 1, 4-5, 191, or 207-212, wherein the target region of the human DNM2 gene pre-mRNA is within the 8th nucleotide of intron 12 measured from the 5′ end of exon 13 (SEQ ID NO: 8) to the 21st nucleotide exon 13 measured from the 5′ end of exon 13.
214. The antisense oligomer of claim 213, wherein the target region is a region within SEQ ID NO: 53.
215. The antisense oligomer of any one of claims 1-3 or 198, wherein the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8).
216. The antisense oligomer of claim 215, wherein the target region is a region within SEQ ID NO: 54.
217. The antisense oligomer of any one of claims 1-3, 198, or 215-216, wherein the target region of the human DNM2 gene pre-mRNA is within the 1st nucleotide to the 28th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8).
218. The antisense oligomer of claim 217, wherein the target region is a region within SEQ ID NO: 55.
219. The antisense oligomer of any one of claims 1-3, 198, or 215-216, wherein the target region of the human DNM2 gene pre-mRNA is within the 6th nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8).
220. The antisense oligomer of claim 219, wherein the target region is a region within SEQ ID NO: 56.
221. The antisense oligomer of any one of claims 1-3, 198, or 215-220, wherein the target region of the human DNM2 gene pre-mRNA is within the 3rd nucleotide to the 30th nucleotide of exon 13 measured from the 5′ end of exon 13 (SEQ ID NO: 8).
222. The antisense oligomer of claim 221, wherein the target region is a region within SEQ ID NO: 57.
223. The antisense oligomer of any one of claims 1-7 or 198-221, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2
H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), DNM2 H13A(+9+28), DNM2 H13A(+16+40), DNM2 H13D(+22-3), or DNM2 H13D(+7-18).
224. The antisense oligomer of claim 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), DNM2 H13A(-13+12), DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), DNM2 H13A(-1+24), DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), or DNM2 H13A(+4+28).
225. The antisense oligomer of any one of claims 1, 4-7, 198-200, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(-26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), DNM2 H13A(-23+2), DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), DNM2 H13A(-18+7), DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(-13+12).
226. The antisense oligomer of any one of claims 1, 4-7, 198-202, 223, or 225, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-27-3), DNM2 H13A(- 26-2), DNM2 H13A(-25-1), DNM2 H13A(-24+1), or DNM2 H13A(-23+2).
227. The antisense oligomer of any one of claims 1, 4-7, 198-200, 203-204, 223, or 225, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-22+3), DNM2 H13A(-21+4), DNM2 H13A(-20+5), DNM2 H13A(-19+6), or DNM2 H13A(-18+7).
228. The antisense oligomer of any one of claims 1, 4-7, 198-200, 205-206, 223, or 225, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-17+8), DNM2 H13A(-16+9), DNM2 H13A(-15+10), DNM2 H13A(-14+11), or DNM2 H13A(- 13+12).
229. The antisense oligomer of any one of claims 1, 4-7, 198, 207-208, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(-1+24).
230. The antisense oligomer of any one of claims 1, 4-5, 7, 198, 207-210, 223, or 229, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-11+14), DNM2 H13A(-10+15), DNM2 H13A(-9+16), DNM2 H13A(-8+17), or DNM2 H13A(-7+18).
231. The antisense oligomer of any one of claims 1, 4-5, 7, 198, 207-208, 211-212, 223, or 229, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(- 5+20), DNM2 H13A(-4+21), DNM2 H13A(-3+22), DNM2 H13A(-2+23), or DNM2 H13A(- 1+24).
232. The antisense oligomer of any one of claims 1, 4-5, 7, 198, 207-214, 223, or 229-231, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(-8+17), DNM2 H13A(-7+18), DNM2 H13A(-6+19), DNM2 H13A(-5+20), or DNM2 H13A(-4+21).
233. The antisense oligomer of any one of claims 1-3, 7, 198, 215-216, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28).
234. The antisense oligomer of any one of claims 1-3, 7, 198, 215-218, 223, or 233, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+1+25), DNM2 H13A(+2+25), DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), or DNM2 H13A(+5+28).
235. The antisense oligomer of any one of claims 1-3, 7, 198, 215-216, 219-220, 223, or 233, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+6+28), DNM2 H13A(+6+30), DNM2 H13A(+7+28), DNM2 H13A(+8+28), or DNM2 H13A(+9+28).
236. The antisense oligomer of any one of claims 1-3, 7, 198, 215-223, or 228-235, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+3+27), DNM2 H13A(+4+23), DNM2 H13A(+4+24), DNM2 H13A(+4+25), DNM2 H13A(+4+26), DNM2 H13A(+4+27), DNM2 H13A(+4+28), DNM2 H13A(+5+28), DNM2 H13A(+6+28), or DNM2 H13A(+6+30).
237. The antisense oligomer of any one of claims 1-3, 7, 198, 215-223, or 228-236, wherein the target region of the human DNM2 gene pre-mRNA DNM2 H13A(+4+28).
238. The antisense oligomer of any one of claims 1-3, 7, 198, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13A(+16+40).
239. The antisense oligomer of any one of claims 1, 4-5, 7, 198, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13D(+22-3).
240. The antisense oligomer of any one of claims 1, 4-5, 7, 198, or 223, wherein the target region of the human DNM2 gene pre-mRNA is DNM2 H13D(+7-18).
241. The antisense oligomer of any one of claims 1-7, 198-221, or 223, wherein the targeting sequence is selected from SEQ ID NOs: 244-287.
242. The antisense oligomer of any one of claims 1, 7, or 223-224, wherein the targeting sequence is selected from SEQ ID NOs: 244-278.
243. The antisense oligomer of any one of claims 1, 4-7, 198-200, 223, or 225, wherein the targeting sequence is selected from SEQ ID NOs: 244-258.
244. The antisense oligomer of any one of claims 1, 4-7, 198-202, 223, or 225-226, wherein the targeting sequence is selected from SEQ ID NOs: 244-248.
245. The antisense oligomer of any one of claims 1, 4-7, 198-200, 203-204, 223, 225, or 227, wherein the targeting sequence is selected from SEQ ID NOs: 249-253.
246. The antisense oligomer of any one of claims 1, 4-7, 198-200, 205-206, 223, 225, or 228, wherein the targeting sequence is selected from SEQ ID NOs: 254-258.
247. The antisense oligomer of any one of claims 1, 4-7, 198, 207-208, 223, or 229, wherein the targeting sequence is selected from SEQ ID NOs: 259-269.
248. The antisense oligomer of any one of claims 1, 4-5, 7, 198, 207-210, 223, or 229-230, wherein the targeting sequence is selected from SEQ ID NOs: 259-263.
249. The antisense oligomer of any one of claims 1, 4-5, 7, 198, 207-208, 211-212, or 231, wherein the targeting sequence is selected from SEQ ID NOs: 265-269.
250. The antisense oligomer of any one of claims 1, 4-5, 7, 198, 207-214, 223, or 229-232, wherein the targeting sequence is selected from SEQ ID NOs: 262-265.
251. The antisense oligomer of any one of claims 1-3, 7, 198, 215-216, 223, or 233, wherein the targeting sequence is selected from SEQ ID NOs: 270-284.
252. The antisense oligomer of any one of claims 1-3, 7, 198, 215-218, 223, or 233-234, wherein the targeting sequence is selected from SEQ ID NOs: 270-279.
253. The antisense oligomer of any one of claims 1-3, 7, 198, 215-216, 219-220, 223, 233, or 235, wherein the targeting sequence is selected from SEQ ID NOs: 280-284.
254. The antisense oligomer of any one of claims 1-3, 7, 198, 215-223, or 228-236, wherein the targeting sequence is selected from SEQ ID NOs: 272-281.
255. The antisense oligomer of any one of claims 1-3, 7, 198, 215-223, or 228-237, wherein the targeting sequence is SEQ ID NO: 278.
256. The antisense oligomer of any one of claims 1-3, 7, 198, 223, or 238, wherein the targeting sequence is SEQ ID NO: 285.
257. The antisense oligomer of any one of claims 1, 4-5, 7, 198, 223, or 239, wherein the targeting sequence is SEQ ID NO: 286.
258. The antisense oligomer of any one of claims 1, 4-5, 7, 198, 223, or 240, wherein the targeting sequence is SEQ ID NO: 287.
259. The antisense oligomer of any one of claims 1-258, wherein the antisense oligomer is selected from a peptide nucleic acid, a locked nucleic acid, a phosphorodiamidate morpholino oligomer, a 2′-OMe phosphorothioate oligomer, or a combination thereof.
260. The antisense oligomer of any one of claims 1-259, wherein the antisense oligomer is a phosphorodiamidate morpholino oligomer.
261. The antisense oligomer of any one of claims 1-260, wherein the antisense oligomer is covalently linked to a delivery moiety selected from a cell-penetrating peptide, an antibody, a fragment of an antibody, and an antigen binding agent.
262. The antisense oligomer of any one of claims 1-261, wherein the antisense oligomer is covalently linked to a cell-penetrating peptide.
263. The antisense oligomer of claim 262, wherein the antisense oligomer is covalently linked to the cell-penetrating peptide by a linker selected from a direct bond, a glycine amino acid, a proline amino acid, glutamic acid amino acid, or an isoglutamine amino acid.
264. The antisense oligomer of claim 262 or 263, wherein the cell-penetrating peptide is selected from rTAT, Tat, R9F2, R5F2R4, R4, R5, R6, R7, R8, R9, (RXR)4, (RXR)5, (RXRRBR)2, (RAR)4F2, (RGR)4F2, and RBRBYLIQFRBRRBR, wherein A represents alanine, B represents β-alanine, F represents phenylalanine, G represents glycine, I represents isoleucine, L represents leucine, Q represents glutamine, R represents arginine, X represent 6-aminohexanoic acid, and Y represents tyrosine.
265. The antisense oligomer, or a pharmaceutically acceptable salt thereof, of any one of claims 1-258 having a structure of Formula (I):
(I), wherein: A′ is selected from -OH,
R5 is -C(O)(O-alkyl)x-OH, wherein x is 3-10 and each alkyl group is, independently at each occurrence, C2-6-alkyl, or R5 is selected from H, -C(O)C1-6-alkyl, trityl, monomethoxytrityl, -(C1-6-alkyl)-R6, -(C1-6- heteroalkyl)-R6, -C6-10-aryl-R6, 5-10 membered heteroaryl-R6, -C(O)O-(C1-6-alkyl)-R6, - C(O)O-aryl-R6, -C(O)O-(5-10 membered heteroaryl)-R6, and
;
R6 is selected from -OH, -SH, and -NH2, or R6 is O, S, or NH, each of which is covalently linked to a solid support; R9 is C1-6 alkyl; each R1 is independently selected from -OH and -N(R3)(R4), wherein each R3 and R4 is, independently at each occurrence, -H or -C1-6-alkyl; each R2 is independently selected from a naturally or non-naturally occurring nucleobase, which, when taken together, forms the targeting sequence; t is 11-28; E′ is selected from -H, -C1-6-alkyl, -C(O)C1-6-alkyl, benzoyl, stearoyl, trityl, monomethoxytrityl, dimethoxytrityl, trimethoxytrityl,
; Q is -C(O)(CH2)6C(O)- or -C(O)(CH2)2S2(CH2)2C(O)-; R7 is -(CH2)2OC(O)N(R8)2, wherein R8 is -(CH2)6NHC(=NH)NH2; L is a linking amino acid, wherein L is covalently-linked by an amide bond to the C-terminus of J; J is a cell-penetrating peptide; and G is selected from -H, -C(O)C1-6-alkyl, benzoyl, and stearoyl, wherein G is covalently-linked to J.
267. The antisense oligomer of claim 265 or 266, wherein E′ is selected from -H, -C(O)CH3, benzoyl, stearoyl, trityl, 4-methoxytrityl, and
272. The antisense oligomer of any one of claims 265-271, wherein each R1 is -N(CH3)2.
273. The antisense oligomer of any one of claims 265-272, wherein L is glycine, proline, or β- alanine.
274. The antisense oligomer of any one of claims 265-273, wherein L is glycine.
275. The antisense oligomer of any one of claims 265-273, wherein L is proline.
276. The antisense oligomer of any one of claims 265-273, wherein L is β-alanine.
277. The antisense oligomer of any one of claims 265-276, wherein J is selected from rTAT, Tat, R9F2, R5F2R4, R4, R5, R6, R7, R8, R9, (RXR)4, (RXR)5, (RXRRBR)2, (RAR)4F2, and (RGR)4F2, wherein A represents alanine, B represents β-alanine, F represents phenylalanine, G represents glycine, R represents arginine, and X represents 6- aminohexanoic acid.
278. The antisense oligomer of any one of claims 265-277, wherein G is selected from -H, - C(O)CH3, benzoyl, and stearoyl.
279. The antisense oligomer of any one of claims 265-278, wherein G is -H or -C(O)CH3.
280. The antisense oligomer of any one of claims 265-278, wherein G is -H.
281. The antisense oligomer of any one of claims 265-278, wherein G is -C(O)CH3.
282. The antisense oligomer of any one of claims 265-273, 276-279, or 281, wherein the antisense oligomer is of Formula (IA):
283. The antisense oligomer of any one of claims 265-270 or 272-282, wherein the antisense oligomer is of Formula (II):
286. The antisense oligomer of claim 284 or 285, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 83.
287. The antisense oligomer of claim 284 or 285, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 88.
288. The antisense oligomer of claim 284 or 285, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 172.
289. The antisense oligomer of claim 284 or 285, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 199.
290. The antisense oligomer of claim 284 or 285, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 230.
291. The antisense oligomer of claim 284 or 285, or a pharmaceutically acceptable salt thereof, wherein each R2, taken together, forms a targeting sequence having SEQ ID NO: 278.
299. A pharmaceutical composition comprising the antisense oligomer of any one of claims 1- 298 and a pharmaceutically acceptable carrier.
300. A method of treating a disease or disorder, or sequelae thereof, associated with aberrant expression of dynamin 2 (DNM2) protein, comprising administering to a subject in need thereof a therapeutically effective amount of the oligomer of any one of claims 1-298 or the pharmaceutical composition of claim 299.
301. The method of claim 300, wherein the disease or disorder is a genetic neuromuscular disorder.
302. The method of claim 300 or 301, wherein the disease or disorder is caused by at least one mutation in a gene selected from DNM2 gene, MTM1 gene, BIN1 gene, RYR1 gene, TTN gene, SPEG gene, CACNA1S gene, MTMR14 gene, ZAK gene, and CCDC78 gene.
303. The method of any one of claims 300-302, wherein the disease or disorder is autosomal dominant, autosomal recessive, or X-linked.
304. The method of any one of claims 300-303, wherein the disease or disorder is a centronuclear myopathy (CNM).
305. The method of any one of claims 300-304, wherein the disease or disorder is myotubular myopathy (XLMTM or XLCNM), autosomal dominant congenital myopathy (ADCNM), autosomal recessive centronuclear myopathy (ARCNM), or Charcot-Marie-Tooth disease.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363591174P | 2023-10-18 | 2023-10-18 | |
| US63/591,174 | 2023-10-18 |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| WO2025085810A2 true WO2025085810A2 (en) | 2025-04-24 |
| WO2025085810A3 WO2025085810A3 (en) | 2025-05-30 |
Family
ID=93455982
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/052063 Pending WO2025085810A2 (en) | 2023-10-18 | 2024-10-18 | Antisense oligomers for treatment of centronuclear myopathies |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025085810A2 (en) |
Citations (56)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5034506A (en) | 1985-03-15 | 1991-07-23 | Anti-Gene Development Group | Uncharged morpholino-based polymers having achiral intersubunit linkages |
| US5142047A (en) | 1985-03-15 | 1992-08-25 | Anti-Gene Development Group | Uncharged polynucleotide-binding polymers |
| US5166315A (en) | 1989-12-20 | 1992-11-24 | Anti-Gene Development Group | Sequence-specific binding polymers for duplex nucleic acids |
| WO1993001286A2 (en) | 1991-06-28 | 1993-01-21 | Massachusetts Institute Of Technology | Localized oligonucleotide therapy |
| US5185444A (en) | 1985-03-15 | 1993-02-09 | Anti-Gene Deveopment Group | Uncharged morpolino-based polymers having phosphorous containing chiral intersubunit linkages |
| US5217866A (en) | 1985-03-15 | 1993-06-08 | Anti-Gene Development Group | Polynucleotide assay reagent and method |
| US5506337A (en) | 1985-03-15 | 1996-04-09 | Antivirals Inc. | Morpholino-subunit combinatorial library and method |
| US5521063A (en) | 1985-03-15 | 1996-05-28 | Antivirals Inc. | Polynucleotide reagent containing chiral subunits and methods of use |
| US5539082A (en) | 1993-04-26 | 1996-07-23 | Nielsen; Peter E. | Peptide nucleic acids |
| US5714331A (en) | 1991-05-24 | 1998-02-03 | Buchardt, Deceased; Ole | Peptide nucleic acids having enhanced binding affinity, sequence specificity and solubility |
| US5719262A (en) | 1993-11-22 | 1998-02-17 | Buchardt, Deceased; Ole | Peptide nucleic acids having amino acid side chains |
| US6245747B1 (en) | 1996-03-12 | 2001-06-12 | The Board Of Regents Of The University Of Nebraska | Targeted site specific antisense oligodeoxynucleotide delivery method |
| US6670461B1 (en) | 1997-09-12 | 2003-12-30 | Exiqon A/S | Oligonucleotide analogues |
| WO2004043977A2 (en) | 2002-11-05 | 2004-05-27 | Isis Pharmaceuticals, Inc. | 2’-fluoro substituted oligomeric compounds and compositions for use in gene modulations |
| US6965025B2 (en) | 2001-12-10 | 2005-11-15 | Isis Pharmaceuticals, Inc. | Antisense modulation of connective tissue growth factor expression |
| US6969400B2 (en) | 1995-12-18 | 2005-11-29 | Cohesion Technologies, Inc. | Synthetic implant with nonimmunogenicity coating |
| US6969766B2 (en) | 2002-04-26 | 2005-11-29 | Panagene, Inc. | PNA monomer and precursor |
| WO2006000057A1 (en) | 2004-06-28 | 2006-01-05 | SMITHKLINE BEECHAM CORPORATION, doing business as GLAXOSMITHKLINE | Antisense oligonucleotides for inducing exon skipping and methods of use thereof |
| US7022851B2 (en) | 2002-01-24 | 2006-04-04 | Panagene, Inc. | PNA monomer and precursor |
| US7053207B2 (en) | 1999-05-04 | 2006-05-30 | Exiqon A/S | L-ribo-LNA analogues |
| US7060809B2 (en) | 2001-09-04 | 2006-06-13 | Exiqon A/S | LNA compositions and uses thereof |
| US7084125B2 (en) | 1999-03-18 | 2006-08-01 | Exiqon A/S | Xylo-LNA analogues |
| US7211668B2 (en) | 2003-07-28 | 2007-05-01 | Panagene, Inc. | PNA monomer and precursor |
| WO2008036127A2 (en) | 2006-05-10 | 2008-03-27 | Avi Biopharma, Inc. | Oligonucleotide analogs having cationic intersubunit linkages |
| US7468418B2 (en) | 2003-04-29 | 2008-12-23 | Avi Biopharma., Inc. | Compositions for enhancing transport of molecules into cells |
| WO2009008725A2 (en) | 2007-07-12 | 2009-01-15 | Prosensa Technologies B.V. | Molecules for targeting compounds to various selected organs, tissues or tumor cells |
| WO2009064471A1 (en) | 2007-11-15 | 2009-05-22 | Avi Biopharma, Inc. | Method of synthesis of morpholino oligomers |
| US7569575B2 (en) | 2002-05-08 | 2009-08-04 | Santaris Pharma A/S | Synthesis of locked nucleic acid derivatives |
| US7572582B2 (en) | 1997-09-12 | 2009-08-11 | Exiqon A/S | Oligonucleotide analogues |
| WO2010064146A2 (en) | 2008-12-02 | 2010-06-10 | Chiralgen, Ltd. | Method for the synthesis of phosphorus atom modified nucleic acids |
| WO2010115993A1 (en) | 2009-04-10 | 2010-10-14 | Association Institut De Myologie | Tricyclo-dna antisense oligonucleotides, compositions, and methods for the treatment of disease |
| WO2011005761A1 (en) | 2009-07-06 | 2011-01-13 | Ontorii, Inc | Novel nucleic acid prodrugs and methods use thereof |
| WO2011034072A1 (en) | 2009-09-16 | 2011-03-24 | 株式会社キラルジェン | Novel protecting group for synthesizing rna and derivative thereof |
| WO2011150408A2 (en) | 2010-05-28 | 2011-12-01 | Avi Biopharma, Inc. | Oligonucleotide analogues having modified intersubunit linkages and/or terminal groups |
| US8076476B2 (en) | 2007-11-15 | 2011-12-13 | Avi Biopharma, Inc. | Synthesis of morpholino oligomers using doubly protected guanine morpholino subunits |
| WO2012039448A1 (en) | 2010-09-24 | 2012-03-29 | 株式会社キラルジェン | Asymmetric auxiliary group |
| US8299206B2 (en) | 2007-11-15 | 2012-10-30 | Avi Biopharma, Inc. | Method of synthesis of morpholino oligomers |
| WO2012150960A1 (en) | 2011-05-05 | 2012-11-08 | Avi Biopharma, Inc. | Peptide oligonucleotide conjugates |
| WO2013053928A1 (en) | 2011-10-13 | 2013-04-18 | Association Institut De Myologie | Tricyclo-phosphorothioate dna |
| WO2013112053A1 (en) | 2012-01-27 | 2013-08-01 | Prosensa Technologies B.V. | Rna modulating oligonucleotides with improved characteristics for the treatment of duchenne and becker muscular dystrophy |
| WO2013127858A1 (en) | 2012-02-29 | 2013-09-06 | Roesler Klaus-Dieter | Method and device for editing forms using a data processing unit |
| WO2014010250A1 (en) | 2012-07-13 | 2014-01-16 | Chiralgen, Ltd. | Asymmetric auxiliary group |
| WO2014012081A2 (en) | 2012-07-13 | 2014-01-16 | Ontorii, Inc. | Chiral control |
| WO2015108047A1 (en) | 2014-01-15 | 2015-07-23 | 株式会社新日本科学 | Chiral nucleic acid adjuvant having immunity induction activity, and immunity induction activator |
| WO2015108046A1 (en) | 2014-01-15 | 2015-07-23 | 株式会社新日本科学 | Chiral nucleic acid adjuvant having anti-allergic activity, and anti-allergic agent |
| WO2015108048A1 (en) | 2014-01-15 | 2015-07-23 | 株式会社新日本科学 | Chiral nucleic acid adjuvant having antitumor effect and antitumor agent |
| WO2015107425A2 (en) | 2014-01-16 | 2015-07-23 | Wave Life Sciences Pte. Ltd. | Chiral design |
| WO2016070166A2 (en) | 2014-11-02 | 2016-05-06 | Arcturus Therapeutics, Inc. | Messenger una molecules and uses thereof |
| WO2017015555A1 (en) | 2015-07-22 | 2017-01-26 | Wave Life Sciences Ltd. | Oligonucleotide compositions and methods thereof |
| WO2017062862A2 (en) | 2015-10-09 | 2017-04-13 | Wave Life Sciences Ltd. | Oligonucleotide compositions and methods thereof |
| WO2017192679A1 (en) | 2016-05-04 | 2017-11-09 | Wave Life Sciences Ltd. | Methods and compositions of biologically active agents |
| WO2017192664A1 (en) | 2016-05-04 | 2017-11-09 | Wave Life Sciences Ltd. | Oligonucleotide compositions and methods thereof |
| WO2022171972A1 (en) | 2021-02-12 | 2022-08-18 | Oxford University Innovation Limited | Cell-penetrating peptide conjugates and methods of their use |
| WO2022212886A1 (en) | 2021-04-02 | 2022-10-06 | Avidity Biosciences, Inc. | Antibody-oligonucleotide conjugate and antibody-peptide-oligonucleotide conjugate compositions and methods of inducing exon skipping |
| WO2023283623A1 (en) | 2021-07-09 | 2023-01-12 | Dyne Therapeutics, Inc. | Muscle targeting complexes and formulations for treating dystrophinopathies |
| WO2023283624A2 (en) | 2021-07-09 | 2023-01-12 | Dyne Therapeutics, Inc. | Muscle targeting complexes and uses thereof for treating dystrophinopathies |
Family Cites Families (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| EP3085785A1 (en) * | 2015-04-22 | 2016-10-26 | Université de Strasbourg | Dynamin 2 inhibitor for the treatment of duchenne's muscular dystrophy |
| WO2018115477A1 (en) * | 2016-12-23 | 2018-06-28 | Universite De Strasbourg | Dynamin 2 inhibitor for the treatment of myotonic dystrophy |
| WO2018189208A1 (en) * | 2017-04-10 | 2018-10-18 | Genethon | Antisense targeting dynamin 2 and use for the treatment of centronuclear myopathies and neuropathies |
| WO2019140452A1 (en) * | 2018-01-15 | 2019-07-18 | Ionis Pharmaceuticals, Inc. | Modulators of dnm2 expression |
| CN111020710A (en) * | 2018-10-10 | 2020-04-17 | 珠海铂华生物工程有限公司 | ctDNA high-throughput detection of hematopoietic and lymphoid tissue tumors |
| US20230226212A1 (en) * | 2020-01-10 | 2023-07-20 | Dyne Therapeutics, Inc. | Muscle-targeting complexes and uses thereof |
| EP4215614A1 (en) * | 2022-01-24 | 2023-07-26 | Dynacure | Combination therapy for dystrophin-related diseases |
-
2024
- 2024-10-18 WO PCT/US2024/052063 patent/WO2025085810A2/en active Pending
Patent Citations (64)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5142047A (en) | 1985-03-15 | 1992-08-25 | Anti-Gene Development Group | Uncharged polynucleotide-binding polymers |
| US5185444A (en) | 1985-03-15 | 1993-02-09 | Anti-Gene Deveopment Group | Uncharged morpolino-based polymers having phosphorous containing chiral intersubunit linkages |
| US5217866A (en) | 1985-03-15 | 1993-06-08 | Anti-Gene Development Group | Polynucleotide assay reagent and method |
| US5506337A (en) | 1985-03-15 | 1996-04-09 | Antivirals Inc. | Morpholino-subunit combinatorial library and method |
| US5521063A (en) | 1985-03-15 | 1996-05-28 | Antivirals Inc. | Polynucleotide reagent containing chiral subunits and methods of use |
| US5034506A (en) | 1985-03-15 | 1991-07-23 | Anti-Gene Development Group | Uncharged morpholino-based polymers having achiral intersubunit linkages |
| US5698685A (en) | 1985-03-15 | 1997-12-16 | Antivirals Inc. | Morpholino-subunit combinatorial library and method |
| US5166315A (en) | 1989-12-20 | 1992-11-24 | Anti-Gene Development Group | Sequence-specific binding polymers for duplex nucleic acids |
| US5714331A (en) | 1991-05-24 | 1998-02-03 | Buchardt, Deceased; Ole | Peptide nucleic acids having enhanced binding affinity, sequence specificity and solubility |
| WO1993001286A2 (en) | 1991-06-28 | 1993-01-21 | Massachusetts Institute Of Technology | Localized oligonucleotide therapy |
| US5539082A (en) | 1993-04-26 | 1996-07-23 | Nielsen; Peter E. | Peptide nucleic acids |
| US5719262A (en) | 1993-11-22 | 1998-02-17 | Buchardt, Deceased; Ole | Peptide nucleic acids having amino acid side chains |
| US6969400B2 (en) | 1995-12-18 | 2005-11-29 | Cohesion Technologies, Inc. | Synthetic implant with nonimmunogenicity coating |
| US6245747B1 (en) | 1996-03-12 | 2001-06-12 | The Board Of Regents Of The University Of Nebraska | Targeted site specific antisense oligodeoxynucleotide delivery method |
| US6670461B1 (en) | 1997-09-12 | 2003-12-30 | Exiqon A/S | Oligonucleotide analogues |
| US7034133B2 (en) | 1997-09-12 | 2006-04-25 | Exiqon A/S | Oligonucleotide analogues |
| US6794499B2 (en) | 1997-09-12 | 2004-09-21 | Exiqon A/S | Oligonucleotide analogues |
| US7572582B2 (en) | 1997-09-12 | 2009-08-11 | Exiqon A/S | Oligonucleotide analogues |
| US7084125B2 (en) | 1999-03-18 | 2006-08-01 | Exiqon A/S | Xylo-LNA analogues |
| US7053207B2 (en) | 1999-05-04 | 2006-05-30 | Exiqon A/S | L-ribo-LNA analogues |
| US7060809B2 (en) | 2001-09-04 | 2006-06-13 | Exiqon A/S | LNA compositions and uses thereof |
| US6965025B2 (en) | 2001-12-10 | 2005-11-15 | Isis Pharmaceuticals, Inc. | Antisense modulation of connective tissue growth factor expression |
| US7022851B2 (en) | 2002-01-24 | 2006-04-04 | Panagene, Inc. | PNA monomer and precursor |
| US7125994B2 (en) | 2002-04-26 | 2006-10-24 | Panagene, Inc. | PNA monomer and precursor |
| US6969766B2 (en) | 2002-04-26 | 2005-11-29 | Panagene, Inc. | PNA monomer and precursor |
| US7145006B2 (en) | 2002-04-26 | 2006-12-05 | Panagene, Inc. | PNA monomer and precursor |
| US7179896B2 (en) | 2002-04-26 | 2007-02-20 | Panagene, Inc. | Method of making PNA oligomers |
| US7569575B2 (en) | 2002-05-08 | 2009-08-04 | Santaris Pharma A/S | Synthesis of locked nucleic acid derivatives |
| WO2004043977A2 (en) | 2002-11-05 | 2004-05-27 | Isis Pharmaceuticals, Inc. | 2’-fluoro substituted oligomeric compounds and compositions for use in gene modulations |
| US7468418B2 (en) | 2003-04-29 | 2008-12-23 | Avi Biopharma., Inc. | Compositions for enhancing transport of molecules into cells |
| US7211668B2 (en) | 2003-07-28 | 2007-05-01 | Panagene, Inc. | PNA monomer and precursor |
| WO2006000057A1 (en) | 2004-06-28 | 2006-01-05 | SMITHKLINE BEECHAM CORPORATION, doing business as GLAXOSMITHKLINE | Antisense oligonucleotides for inducing exon skipping and methods of use thereof |
| WO2008036127A2 (en) | 2006-05-10 | 2008-03-27 | Avi Biopharma, Inc. | Oligonucleotide analogs having cationic intersubunit linkages |
| US7943762B2 (en) | 2006-05-10 | 2011-05-17 | Avi Biopharma, Inc. | Oligonucleotide analogs having cationic intersubunit linkages |
| WO2009008725A2 (en) | 2007-07-12 | 2009-01-15 | Prosensa Technologies B.V. | Molecules for targeting compounds to various selected organs, tissues or tumor cells |
| US8076476B2 (en) | 2007-11-15 | 2011-12-13 | Avi Biopharma, Inc. | Synthesis of morpholino oligomers using doubly protected guanine morpholino subunits |
| WO2009064471A1 (en) | 2007-11-15 | 2009-05-22 | Avi Biopharma, Inc. | Method of synthesis of morpholino oligomers |
| US8299206B2 (en) | 2007-11-15 | 2012-10-30 | Avi Biopharma, Inc. | Method of synthesis of morpholino oligomers |
| WO2010064146A2 (en) | 2008-12-02 | 2010-06-10 | Chiralgen, Ltd. | Method for the synthesis of phosphorus atom modified nucleic acids |
| WO2010115993A1 (en) | 2009-04-10 | 2010-10-14 | Association Institut De Myologie | Tricyclo-dna antisense oligonucleotides, compositions, and methods for the treatment of disease |
| WO2011005761A1 (en) | 2009-07-06 | 2011-01-13 | Ontorii, Inc | Novel nucleic acid prodrugs and methods use thereof |
| WO2011034072A1 (en) | 2009-09-16 | 2011-03-24 | 株式会社キラルジェン | Novel protecting group for synthesizing rna and derivative thereof |
| WO2011150408A2 (en) | 2010-05-28 | 2011-12-01 | Avi Biopharma, Inc. | Oligonucleotide analogues having modified intersubunit linkages and/or terminal groups |
| WO2012039448A1 (en) | 2010-09-24 | 2012-03-29 | 株式会社キラルジェン | Asymmetric auxiliary group |
| WO2012150960A1 (en) | 2011-05-05 | 2012-11-08 | Avi Biopharma, Inc. | Peptide oligonucleotide conjugates |
| WO2013053928A1 (en) | 2011-10-13 | 2013-04-18 | Association Institut De Myologie | Tricyclo-phosphorothioate dna |
| WO2013112053A1 (en) | 2012-01-27 | 2013-08-01 | Prosensa Technologies B.V. | Rna modulating oligonucleotides with improved characteristics for the treatment of duchenne and becker muscular dystrophy |
| WO2013127858A1 (en) | 2012-02-29 | 2013-09-06 | Roesler Klaus-Dieter | Method and device for editing forms using a data processing unit |
| WO2014012081A2 (en) | 2012-07-13 | 2014-01-16 | Ontorii, Inc. | Chiral control |
| WO2014010250A1 (en) | 2012-07-13 | 2014-01-16 | Chiralgen, Ltd. | Asymmetric auxiliary group |
| WO2015108047A1 (en) | 2014-01-15 | 2015-07-23 | 株式会社新日本科学 | Chiral nucleic acid adjuvant having immunity induction activity, and immunity induction activator |
| WO2015108046A1 (en) | 2014-01-15 | 2015-07-23 | 株式会社新日本科学 | Chiral nucleic acid adjuvant having anti-allergic activity, and anti-allergic agent |
| WO2015108048A1 (en) | 2014-01-15 | 2015-07-23 | 株式会社新日本科学 | Chiral nucleic acid adjuvant having antitumor effect and antitumor agent |
| WO2015107425A2 (en) | 2014-01-16 | 2015-07-23 | Wave Life Sciences Pte. Ltd. | Chiral design |
| WO2016070166A2 (en) | 2014-11-02 | 2016-05-06 | Arcturus Therapeutics, Inc. | Messenger una molecules and uses thereof |
| WO2017015555A1 (en) | 2015-07-22 | 2017-01-26 | Wave Life Sciences Ltd. | Oligonucleotide compositions and methods thereof |
| WO2017015575A1 (en) | 2015-07-22 | 2017-01-26 | Wave Life Sciences Ltd. | Oligonucleotide compositions and methods thereof |
| WO2017062862A2 (en) | 2015-10-09 | 2017-04-13 | Wave Life Sciences Ltd. | Oligonucleotide compositions and methods thereof |
| WO2017192679A1 (en) | 2016-05-04 | 2017-11-09 | Wave Life Sciences Ltd. | Methods and compositions of biologically active agents |
| WO2017192664A1 (en) | 2016-05-04 | 2017-11-09 | Wave Life Sciences Ltd. | Oligonucleotide compositions and methods thereof |
| WO2022171972A1 (en) | 2021-02-12 | 2022-08-18 | Oxford University Innovation Limited | Cell-penetrating peptide conjugates and methods of their use |
| WO2022212886A1 (en) | 2021-04-02 | 2022-10-06 | Avidity Biosciences, Inc. | Antibody-oligonucleotide conjugate and antibody-peptide-oligonucleotide conjugate compositions and methods of inducing exon skipping |
| WO2023283623A1 (en) | 2021-07-09 | 2023-01-12 | Dyne Therapeutics, Inc. | Muscle targeting complexes and formulations for treating dystrophinopathies |
| WO2023283624A2 (en) | 2021-07-09 | 2023-01-12 | Dyne Therapeutics, Inc. | Muscle targeting complexes and uses thereof for treating dystrophinopathies |
Non-Patent Citations (28)
| Title |
|---|
| AUNG-HTUT ET AL., INT. J. MOL. SCI., vol. 20, 2019, pages 5030 |
| BENNER SA ET AL., NAT. REV. GENET., vol. 6, 2005, pages 553 - 543 |
| DOKKAROJANASAKUL, ADVANCED DRUG DELIVERY REVIEWS, vol. 44, pages 35 - 49 |
| GREGORIADIS, G.: "Drug Carriers in Biology and Medicine", 1979, ACADEMIC PRESS, article "Liposomes", pages: 287 - 341 |
| HAMES ET AL.: "Nucleic Acid Hybridization", 1985, IRL PRESS, pages: 107 - 108 |
| HIRAO, I, CURR. OPIN. CHEM. BIOL., vol. 10, 2006, pages 622 - 627 |
| IYER ET AL., J. ORG. CHEM., vol. 55, 1990, pages 4693 - 4699 |
| KOOL ET, ACC. CHEM. RES., vol. 35, 2002, pages 936 - 943 |
| KRUEGER AT ET AL., ACC. CHEM. RES., vol. 40, 2007, pages 141 - 150 |
| LAPPALAINEN ET AL., ANTIVIRAL RES, vol. 23, 1994, pages 119 |
| LIMBACH ET AL., NUCLEIC ACIDS RES, vol. 22, 1994, pages 2183 - 2196 |
| MANN ET AL., J GENE MED, vol. 4, 2002, pages 644 |
| MARTIN ET AL., HELV. CHIM. ACTA, vol. 78, 1995, pages 486 - 504 |
| MIYADA C. G.WALLACE R. B.: "Oligomer Hybridization Techniques", METHODS ENZYMOL, vol. 154, 1987, pages 94 - 107 |
| MIYADA CG.WALLACE RB: "Oligonucleotide hybridization techniques", METHODS ENZYMOL., vol. 154, 1987, pages 94 - 107, XP009096793, DOI: 10.1016/0076-6879(87)54072-1 |
| NIELSEN ET AL., SCIENCE, vol. 254, 1991, pages 1497 - 1500 |
| OBIKA ET AL., TETRAHEDRON LETT, vol. 38, no. 50, 1997, pages 8735 |
| P. IVERSEN: "Antisense Drug Technology", 2001, MARCEL DEKKER, INC., article "Phosphoramidite Morpholino Oligomers" |
| RANA, RNA, vol. 9, 2003, pages 1034 - 1048 |
| REVANKARRAO, COMPREHENSIVE NATURAL PRODUCTS CHEMISTRY, vol. 7, pages 313 |
| ROMESBERG FE ET AL., CURR. OPIN. CHEM. BIOL., vol. 7, 2003, pages 723 - 733 |
| SUMMERTON, J. ET AL., ANTISENSE & NUCLEIC ACID DRUG DEVELOPMENT, vol. 7, 1997, pages 187 - 195 |
| T.W. GREENEP.G.M. WUTS: "Protective Groups in Organic Synthesis", 1999, JOHN WILEY & SONS |
| UHLMANN ET AL.: "antisense oligomers: a new therapeutic principle", CHEMICAL REVIEWS, vol. 90, no. 4, 25, 1990, pages 544 - 584 |
| WILLIAMS, S.A., LEUKEMIA, vol. 10, no. 12, 1996, pages 1980 - 1989 |
| WU, G.Y.WU, C.H., J. BIOL. CHEM., vol. 262, 1987, pages 4429 - 4432 |
| YAMADA ET AL., J. ORG. CHEM., vol. 76, no. 9, 2011, pages 3042 - 53 |
| YOO ET AL., NUCLEIC ACIDS RES., vol. 32, 2004, pages 2008 - 16 |
Also Published As
| Publication number | Publication date |
|---|---|
| WO2025085810A3 (en) | 2025-05-30 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| AU2022206704B2 (en) | Nanoparticle formulations for delivery of nucleic acid complexes | |
| JP6671449B2 (en) | Antisense-induced exon 2 inclusion in acid α-glucosidase | |
| US20230407301A1 (en) | Antisense oligomer compounds | |
| JP2016528897A (en) | Compositions and methods for modulating RNA | |
| WO2024064237A2 (en) | Dmd antisense oligonucleotide-mediated exon skipping efficiency | |
| US20240415857A1 (en) | Antisense-induced exon exclusion in type vii collagen | |
| JP2021048877A (en) | Modified antisense oligomers for exon inclusion in spinal muscular atrophy | |
| WO2025085810A2 (en) | Antisense oligomers for treatment of centronuclear myopathies | |
| JP2024539223A (en) | Morpholino oligomers for the treatment of peripheral myelin protein 22-associated diseases | |
| US20240425864A1 (en) | Antisense oligonucleotides having one or more abasic units | |
| US20250066776A1 (en) | Antisense oligomers for treatment of chronic kidney disease | |
| CN118019847A (en) | Antisense oligonucleotides having one or more abasic units | |
| JP2025539260A (en) | Thiomorpholino antisense oligonucleotides for treating PTP1B-related disorders |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24804631 Country of ref document: EP Kind code of ref document: A2 |