WO2024102434A1 - Rna compositions comprising lipid nanoparticles or lipid reconstructed natural messenger packs - Google Patents
Rna compositions comprising lipid nanoparticles or lipid reconstructed natural messenger packs Download PDFInfo
- Publication number
- WO2024102434A1 WO2024102434A1 PCT/US2023/037077 US2023037077W WO2024102434A1 WO 2024102434 A1 WO2024102434 A1 WO 2024102434A1 US 2023037077 W US2023037077 W US 2023037077W WO 2024102434 A1 WO2024102434 A1 WO 2024102434A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- independently
- branched
- lipid
- alkyl
- unbranched
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/87—Introduction of foreign genetic material using processes not otherwise provided for, e.g. co-transformation
- C12N15/88—Introduction of foreign genetic material using processes not otherwise provided for, e.g. co-transformation using microencapsulation, e.g. using amphiphile liposome vesicle
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
- C12N9/16—Hydrolases (3) acting on ester bonds (3.1)
- C12N9/22—Ribonucleases [RNase]; Deoxyribonucleases [DNase]
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K9/00—Medicinal preparations characterised by special physical form
- A61K9/48—Preparations in capsules, e.g. of gelatin, of chocolate
- A61K9/50—Microcapsules having a gas, liquid or semi-solid filling; Solid microparticles or pellets surrounded by a distinct coating layer, e.g. coated microspheres, coated drug crystals
- A61K9/51—Nanocapsules; Nanoparticles
- A61K9/5107—Excipients; Inactive ingredients
- A61K9/5123—Organic compounds, e.g. fats, sugars
Definitions
- RNA polynucleotides are typically based on whole microorganisms, protein antigens, peptides, polysaccharides or deoxyribonucleic acid (DNA) vaccines and their combinations.
- DNA deoxyribonucleic acid
- RNA composition for gene editing comprising one or more polynucleotides encoding one or more components of a gene editing system or one or more gene editing systems.
- the one or more polynucleotides are formulated within a complex lipid particle (CLP), such as a lipid reconstructed natural messenger packs (LNMPs) comprising natural lipids and an ionizable lipid.
- CLP complex lipid particle
- LNMPs lipid reconstructed natural messenger packs
- the ionizable lipid has two or more of the characteristics listed below:
- each of the lipid tails is at least 6 carbon atoms in length
- RNA composition in another aspect, comprises reconstituting a film comprising purified NMP lipids in the presence of an ionizable lipid to produce a lipid reconstructed natural messenger packs (LNMP) comprising the ionizable lipid described herein.
- LNMP lipid reconstructed natural messenger packs
- the method further comprises loading into the LNMPs with one or more polynucleotides encoding one or more components of a gene editing system or one or more gene editing systems.
- RNA composition for gene editing comprising: one or more polynucleotides encoding one or more components of a gene editing system or one or more gene editing systems, formulated within a plurality of lipid nanoparticles (LNPs) comprising synthetic structural lipids and an ionizable lipid.
- LNPs lipid nanoparticles
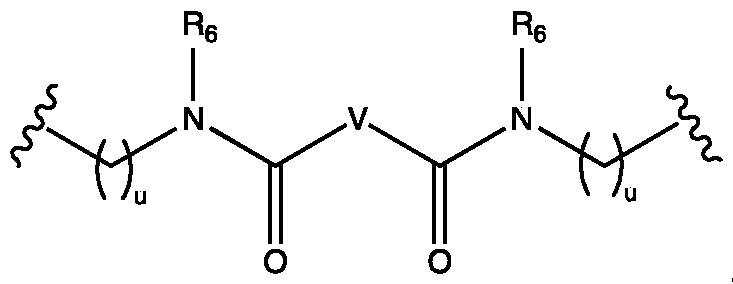
- the ionizable lipid may be selected from one the following groups of compounds: i) a compound of formula pharmaceutically acceptable salt thereof, or a stereoisomer of any of the foregoing, wherein: each A is independently C1-C16 branched or unbranched alkyl or C1-C16 branched or unbranched alkenyl, optionally substituted with heteroatom or substituted with OH, SH, or halogen; each B is independently C1-C16 branched or unbranched alkyl or C1-C16 branched or unbranched alkenyl, optionally substituted with heteroatom or substituted with OH, SH, or halogen; each X is independently a biodegradable moiety; and
- each R 6 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, or cycloalkyl
- each R 7 and each R 8 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, halogen, OH, SH, NR10R11, wherein each R10 and Rn is independently H, C1-C3 alkyl, or R10 and Rn are taken together to form a heterocyclic ring; or R 7 and R 8 are taken together to form a ring; each s is independently 1 , 2, 3, 4, or 5; each u is independently 1 , 2, 3, 4, or 5; t is 1 , 2, 3, 4 or 5; each Z is independently absent, O, S, or NR12, wherein R12 is H, C1-C7 branched or unbranched alkyl, or C2-
- Q is O, S, or NR13, wherein each R13 is H, C1-C5 alkyl;
- Y is alkyl, hydroxy, hydroxyalkyl or
- A is absent, -O-, -N R 7 )-, -O-alkylene-, -alkylene-O-, -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, -N(R 7 )C(O)N(R 7 )-, -S-, -S-S-, or a bivalent heterocycle; each of X and Z is independently absent, -O-, -CO-, -N(R 7 )-, -O-alkylene-, -alkylene-O-, -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, or -S-; each R 7 is independently H, alkyl, alkenyl, cycloalkyl, hydroxy, hydroxyalkyl, or amino
- W is hydroxyl, substituted or unsubstituted hydroxyalkyl, substituted or unsubstituted amino, substituted or unsubstituted aminocarbonyl, or substituted or unsubstituted heterocylyl or heteroaryl; and iii) a compound of formula pharmaceutically acceptable salts thereof, and stereoisomers of any of the foregoing, wherein:
- R20 and R30 are each independently H, C1-C5 branched or unbranched alkyl, or C2-C5 branched or unbranched alkenyl, or R20 and R30 together with the adjacent N atom form a 3 to 7 membered cyclic ring, optionally substituted with R a ;
- R a is H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, halogen, OH, or SH; each Ri and each R 2 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, OH, halogen, SH, or NR10R11, or
- R1 and R 2 are taken together to form a cyclic ring; each R10 and Rn is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, or R10 and Rn are taken together to form a heterocyclic ring; n is 0, 1 , 2, 3 or 4;
- Y is O or S
- E is each independently -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, -C(O-RI 3 )-O-, -C(O)O(CH 2 )r, -C(O)N(R 7 )(CH 2 )r, -S-S-, or -C(O-Ri 3 )-O-(CH 2 )r, wherein each R 7 is independently H, alkyl, alkenyl, cycloalkyl, hydroxyalkyl, or aminoalkyl;
- R13 is branched or unbranched C3-C10 alkyl; r is 1 , 2, 3, 4, or 5;
- R a is each independently C1-C5 alkyl, C2-C5 alkenyl, or C2-C5 alkynyl; u1 and u2 are each independently 0, 1 , 2, 3, 4, 5, 6, or 7;
- R* is each independently H, C1-C16 branched or unbranched alkyl or C1-C16 branched or unbranched alkenyl, optionally interrupted with heteroatom or substituted with OH, SH, or halogen, or cycloalkyl or substituted cycloalkyl; represents the bond connecting the tail group to the head group; and wherein the lipid has a pKa from about 4 to about 8.
- the ionizable lipid is a compound of group i), represented by a formula pharmaceutically acceptable salts thereof, and stereoisomers of any of the foregoing, wherein: each Ri and each R 2 is independently H, C1-C3 branched or unbranched alkyl, OH, halogen, SH, or NR10R11 , or each R1 and each R 2 are independently taken together with the carbon atom(s) to which they are attached to form a cyclic ring; each R10 and Rn is independently H, C1-C3 branched or unbranched alkyl, or R10 and Rn are taken together to form a heterocyclic ring; each R 3 and each R 4 is independently H, C2-C14 branched or unbranched alkyl (e.g., C3-C10 branched or unbranched alkyl), or C3-C10 branched or unbranched alkenyl, provided that at least one of R 3 and R 4 is not
- V is a branched or unbrachned C2-
- the ionizable lipid is a compound of group i), represented by a formula pharmaceutically acceptable salts thereof, and stereoisomers of any of the foregoing, wherein: each R1 and each R 2 is independently H, C1-C3 branched or unbranched alkyl, OH, halogen, SH, or NR10R11 , or each R1 and each R 2 are independently taken together with the carbon atom(s) to which they are attached to form a cyclic ring; each R10 and Rn is independently H, C1-C3 branched or unbranched alkyl, or R10 and Rn are taken together to form a heterocyclic ring; each R 3 and each R 4 is independently H, C2-C14 branched or unbranched alkyl (e.g., C3-C10 branched or unbranched alkyl), or C3-C10 branched or unbranched alkenyl, provided that at least one of R 3 and R 4 is
- T is -NHC(O)O-, -OC(O)NH-, or a divalent heterocyclic optionally substituted with one or more -(CH2) V OH, -(CH2) V SH, -(CH2) v -halogen groups
- each R 7 and each R 8 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, halogen, OH, SH, (CH2) v Ri7, or NR10R11, wherein R17 is OH, SH, or N(CH 3 ) 2 ;
- each v is independently 0, 1 , 2, 3, 4, or 5; and each m is independently 1 , 2, 3, 4, 5, 6, 7, 8, 9 or 10.
- T is a divalent piperazine or a divalent dioxopiperazine.
- X is -OCO-, -COO-, -NHCO-, or -CONH-.
- the ionizable lipid is a compound of group ii), represented by one of the following formulas: each 11 is independently an integer from 4 to 8, m2 and I2 are each independently an integer from 0 to 3,
- Rso and R90 are each independently unsubstituted Cs-Cs alkyl or alkenyl; or Rso is H or unsubstituted C1-C4 alkyl or alkenyl, and R90 is unsubstituted C5-C11 alkyl or alkenyl; and
- R110 and R120 are each independently unsubstituted Cs-Cs alkyl or alkenyl; or Rno is H or unsubstituted C1-C4 alkyl or alkenyl, and R120 is unsubstituted C5-C11 alkyl or alkenyl.
- M is -OC(O)- or -C(0)0-;
- the ionizable lipid is a compound of group iii), wherein R1 and R 2 are each H, or each R1 is H, and one of the R 2 variables is OH; and X is -OC(O)- or -C(0)0-.
- the ionizable lipid is a compound of group iii), represented by formula III), wherein R 20 and R30 are each independently H or C1-C3 branched or unbranched alkyl; or R 20 and R30 together with the adjacent N atom form a 3 to 7 membered cyclic ring, optionally substituted with R a ; R a is H or OH; Z is absent, S, O, or NH; and n is 0, 1 , or 2.
- the ionizable lipid is a compound of group iii), represented by formula V),
- the ionizable lipid is a compound of group iv), wherein the lipid comprises at least one head group and at least one tail group, wherein: the tail group has a structure of formula (Tl) or formula Tl’ the head group has a structure of one of the following formulas: wherein:
- R20 and R30 are each independently H, C1-C5 branched or unbranched alkyl, or C2-C5 branched or unbranched alkenyl, optionally interrupted with one or more heteroatoms or substituted with OH, SH, halogen, or cycloalkyl groups; or
- R20 and R30 together with the adjacent N atom, form a 3 to 7 membered heterocylic or heteroaromatic ring containing one or more heteroatoms, optionally substituted with one or more OH, SH, halogen, alkyl, or cycloalkyl groups; each of Ri and R2 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, OH, halogen, SH, or NR10R11 ; or Ri and R2 together form a cyclic ring; each of Rw and Rn is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl; or Rw and Rn together form a heterocyclic ring; n is 0, 1 , 2, 3 or 4; and
- Z is absent, O, S, or NR12, wherein R12 is H or C1-C7 branched or unbranched alkyl; provided that when Z is not absent, the adjacent Ri and R2 cannot be OH, wherein:
- Ri is H, C1-C3 alkyl, OH, halogen, SH, or NR10R11;
- R2 is OH, halogen, SH, or NR10R11; or Ri and R2 can be taken together to form a cyclic ring;
- R10 and R11 are each independently H or C1-C3 alkyl; or Rw and Rn can be taken together to form a heterocyclic ring;
- R20 and R30 are each independently H, C1-C5 branched or unbranched alkyl, C2-C5 branched or unbranched alkenyl; or R20 and R30 can be taken together to form a cyclic ring; and each of v and y is independently 1 , 2, 3, or 4; wherein W is
- each R20 is independently H, or C1-C3 branched or unbranched alkyl;
- Ru is a heterocyclic, NR10R11, C(O)NRwRn, NRwC(0)NRioRn, or NRioC(S)NRioRii, wherein each Rw and Rn is independently H, C1-C3 alkyl, C3-C7 cycloalkyl, C3-C7 cycloalkenyl, optionally substituted with one or more NH and/or oxo groups, or Rw and Rn are taken together to form a heterocyclic ring;
- Q is O, S, CH2, or NR13, wherein each R13 is H, C1-C5 alkyl;
- V is branched or unbrachned C2-C10 alkylene, C2-C10 alkenylene, C2-C10 alkynylene, or C2-C10 heteroalkylene, optionally substituted with one or more OH, SH, and/or halogen groups; and divalent heterocyclic; and oiety;
- Y is alkyl, hydroxy, hydroxyalkyl
- A is absent, -O-, -N(R 7 )-, -O-alkylene-, -alkylene-O-, -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, -N(R 7 )C(O)N(R 7 )-, -S-, or -S-S-; each of X and Z is independently absent, -O-, -C(O)-, -N(R 7 )-, -O-alkylene-, -alkylene-O-, -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, or -S-; each R 7 is independently H, alkyl, alkenyl, cycloalkyl, hydroxy, alkoxy, hydroxyalkyl, al
- W is hydroxyl, substituted or unsubstituted hydroxyalkyl, substituted or unsubstituted amino, substituted or unsubstituted aminocarbonyl, or substituted or unsubstituted heterocyclyl or heteroaryl; and wherein the lipid has a pKa from about 4 to about 8.
- the ionizable lipid is a compound of group iv), and wherein at least one tail group of the lipid has one of the following formulas:
- R 7 is each independently H or methyl
- R b is in each occasion independently H or C1-C4 alkyl; and u3 and u4 are each independently 0, 1 , 2, 3, 4, 5, 6, or 7; and the head group has a structure of one of the following formulas:
- At least one tail group has the structure of formula (Til), (Till), (TIV), (TV), (TH’), and/or (Till’), wherein u1 is 3-5, u2 is 0-3, u3 and u4 are each independently 1-7, and R a is each independently methyl.
- the tail group has the structure of formula (Til) or formula (Till), wherein each R a is methyl; u1 is 3-5, u2 is 0-3; and u3 and u4 are each independently 1-4.
- the head group has the structure of one of the following formulas rein each R20 and R30 are independently C1-C3 alkyl. , wherein: each RB, R7, and Rs are independently H or methyl; and each of u and t is independently 1 , 2, or 3; or
- R14 is a nitrogen-containing 5- or 6- membered heterocyclic, NR10R11, C(O)NRwRii, NRwC(0)NRioRii, or NRioC(S)NRioRn, wherein each R10 and Rn is independently H or C1-C3 alkyl; and each of u and v is independently 1 , 2, or 3; or .
- each RB is independently H or methyl; each u is independently 1 , 2, or 3; and V is C2-C6 alkylene or C2-C6 alkenylene; or , wherein: each Re is independently H or methyl; each R7 is independently H; each Rs is methyl; each u is independently 1 , 2, or 3; and
- V is C2-C6 alkylene or C2-C6 alkenylene; or , wherein: each u is independently 1 , 2, or 3; and
- T is a divalent nitrogen-containing 5- or 6- membered heterocyclic; or , wherein: each u is independently 1 , 2, or 3;
- Q is O; each Z is independently NR12; and
- R12 is H or C1-C3 alkyl
- W is hydroxyl, substituted or unsubstituted hydroxyalkyl, one of the following moieties:
- each Q is independently absent, -O-, -C(O)-, -C(S)-, -C(O)O-, -(CH2) q C(R 7 )2-, -C(O)N(R 7 )-, -C(S)N(R 7 )-, or -N R 7 );
- R 6 is independently H, alkyl, hydroxyl, hydroxyalkyl, alkoxy, -O-alkylene-O-alkyl, -O-alkylene-N(R 7 )2, amino, alkylamino, aminoalkyl, thiol, thiolalkyl, or N + (R 7 )3-alkylene-Q-; each R 8 is independently H, alkyl, hydroxyalkyl, amino, aminoalkyl, alkylamino, thiol, thiolalkyl, heterocyclyl, heteroaryl; or two R 8 together with the nitrogen atom form a ring, optionally substituted with one or more alkyl, hydroxy, hydroxyalkyl, alkoxy, alkylaminoalkyl, alkylamino, or aminoalkyl; q is 0, 1 , 2, 3, 4, or 5; and p is 0, 1 , 2, 3, 4, or 5. [0018] In some embodiments, the ionizable lipid is a
- the ionizable lipid is Lipid No. 2252, 2272, 2320, 2439, 2356, 2243, 2431 , 2455, 2454, 2424, 2425, 2433, 2275, 2220, 2335, or 2282.
- more than one ionizable lipid can be used for the ionizable lipid component: one or more of the ionizable lipids from the compounds of formulas in groups i)-iv) can be used alone or in combination with a different ionizable lipid from the compounds of formulas in groups i)-iv).
- the CLP is a LNMP
- the CLP formulation encapsulating one or more polynucleotides is a LNMP.
- the polynucleotide is encapsulated by the lipid reconstructed natural messenger packs (LNMPs). In some embodiments, the polynucleotide is encapsulated by the lipid reconstructed plant messenger packs (LPMPs). In some embodiments, the polynucleotide is embedded on the surface of the LNMPs. In some embodiments, the polynucleotide is conjugated to the surface of the LNMPs.
- LNMPs lipid reconstructed natural messenger packs
- LPMPs lipid reconstructed plant messenger packs
- the LNMP is produced by a method comprising lipid extrusion. In some embodiments, the LNMP is produced by a method comprising processing a solution comprising a lipid extract of the NMPs in a microfluidics device comprising an aqueous phase, thereby producing the LNMPs. In some embodiments, the aqueous phase comprises the polynucleotides.
- the natural lipids of the LPMPs are extracted from a plant source, such as lemon or algae.
- the ionizable lipids from the compounds of formulas in groups i)-iv) can be used in combination with one or more other ionizable lipids.
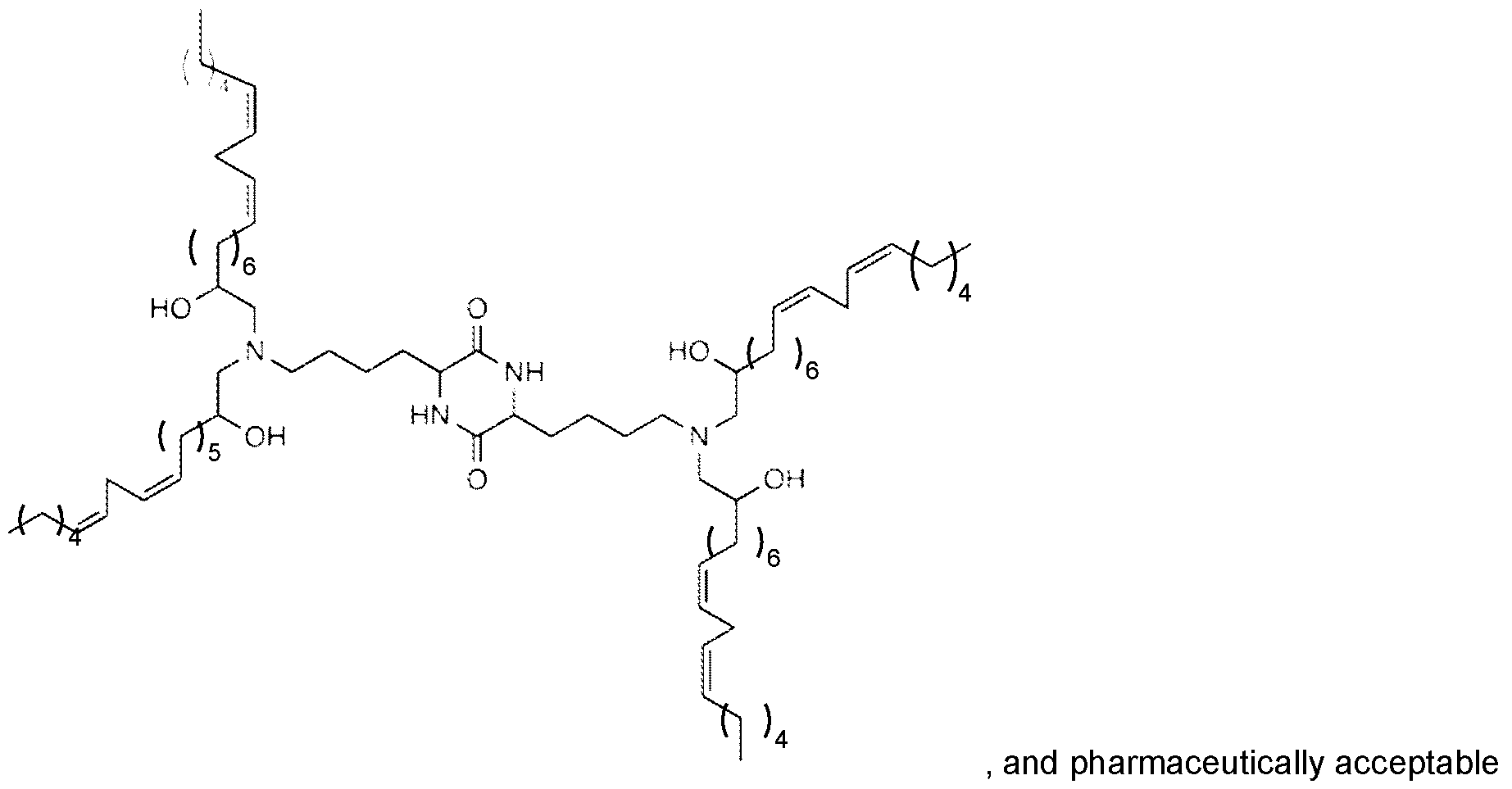
- the ionizable lipid of the LNMPs is selected from the group consisting of 1 ,1 ’-((2-(4-(2-((2-(bis(2-hydroxydodecyl)amino)ethyl) (2-hydroxydodecyl)amino)ethyl)piperazin-1- yl)ethyl)azanediyl)bis(dodecan-2-ol) (C12-200), MD1 (CKK-E12), OF2, EPC, ZA3-Ep10, TT3, LP01 , 5A2-SC8, Lipid 5, SM-102 (Lipid H), and ALC-315.
- the ionizable lipid is C12- 200.
- the ionizable lipid is wherein R is C8-C14 alkyl group.
- the reconstitution is performed in the presence of a sterol, thereby producing a LNMP that comprises natural lipids, an ionizable lipid, and a sterol.
- the reconstitution is performed in the presence of a PEGylated lipid (or a PEG-lipid conjugate), thereby producing a LNMP that comprises natural lipids, a ionizable lipid, and a PEG-lipid conjugate.
- the LNMPs further comprise a sterol and a polyethylene glycol (PEG)- lipid conjugate.
- the LNPs further comprise a sterol and a polyethylene glycol (PEG)-lipid conjugate.
- the sterol is cholesterol or sitosterol.
- PEG lipid conjugate comprises a PEG-2k. In some embodiments, the PEG-lipid conjugate is C14-PEG2k, C18-PEG2k, or DMPE-PEG2k. In some embodiments, the PEG- lipid conjugate is PEG-DMG or PEG-PE. In some embodiments, the PEG lipid conjugate is PEG2000-DMG or PEG2000-PE.
- the amount of the PEG lipid conjugate is about 1 .5-2.5 mol%.
- the amount of the ionizable lipid is about 30-50 mol%, about 30-40 mol%, or about 45-55%. In one embodiment, the amount of the ionizable lipid is about 50 or 35 mol%.
- the LNMP comprises: about 20 mol% to about 50 mol% of the ionizable lipid, about 20 mol% to about 60 mol% of the natural lipids, about 7 mol% to about 50 mol% of the sterol, and about 0.5 mol% to about 3 mol% of the polyethylene glycol (PEG)-lipid conjugate.
- PEG polyethylene glycol
- the LNMPs comprise the ionizable lipid znatural lipids:sterol:PEG-lipid at a molar ratio of about 35:50:12.5:2.5. In one embodiment, the LNMPs comprise the ionizable lipid :natural lipids:sterol:PEG-lipid at a molar ratio of about 35:20:42.5:2.5. In one embodiment, the LNMPs comprise the ionizable lipid :natural lipids:sterol: PEG-lipid at a molar ratio of about 35:16:46.5:2.5.
- the LNMPs comprise the ionizable lipid :natural lipids:sterol:PEG-lipid at a molar ratio of about 50:10:38.5:1.5. In one embodiment, the LNMPs comprise the ionizable lipid :natural lipids:sterol: PEG-lipid at a molar ratio of about 50:20:28.5:1.5.
- the LNMPs comprise: natural lipids extracted from lemon or algae, the ionizable lipid (e.g., from Table I, Table II, Table III, or Table IV), cholesterol, and DMPE-PEG2k.
- the LNMPs comprise: natural lipids extracted from lemon, the ionizable lipid from Table I, Table II, Table III, or Table IV, cholesterol, and
- the LNMPs may comprise the ionizable lipid:lemon lipids:cholesterol: DMPE- PEG2k at a molar ratio of about 35:50:12.5:2.5, about 35:20:42.5:2.5, about 35:16:46.5:2.5, about 50:10:38.5:1 .5, or about 50:20:28.5:1 .5.
- the LNMPs comprise: natural lipids extracted from algae, the ionizable lipid (e.g., from Table I, Table II, Table III, or Table IV), cholesterol, and DMPE-PEG2k.
- the LNMPs may comprise the ionizable lipid:algae lipids:cholesterol: DMPE- PEG2k at a molar ratio of about 35:20:42.5:2.5, about 35:50:12.5:2.5, about 35:16:46.5:2.5, about 50:10:38.5:1 .5, or about 50:20:28.5:1 .5.
- the LNP comprises: about 20 mol% to about 50 mol% of the ionizable lipid, about 5 mol% to about 40 mol% of the synthetic structural lipids, about 20 mol% to about 50 mol% of the sterol, and about 0.5 mol% to about 3 mol% of the polyethylene glycol (PEG)-lipid conjugate.
- PEG polyethylene glycol
- the LNP comprises ionizable lipid: synthetic structural lipid:sterol:PEG- lipid at a molar ratio of about 35:50:12.5:2.5, about 35:20:42.5:2.5, about 35:16:46.5:2.5, about 50:10:38.5:1 .5, or about 50:20:28.5:1 .5.
- the LNPs comprise: synthetic structural lipids, an ionizable lipid (e.g., from Table I, Table II, Table III, or Table IV), cholesterol, and a PEG-lipid.
- the (e.g. LNPs) comprise: synthetic structural lipids, an ionizable lipid (e.g., 2252, 2272, 2320, 2439, 2356, 2243, 2431 , 2455, 2454, 2424, 2425, 2433, 2275, 2220, 2335, or 2282), cholesterol, and
- the LNMP is a lipophilic moiety selected from the group consisting of a lipoplex, a liposome, a lipid nanoparticle, a polymer-based carrier, an exosome, a lamellar body, a micelle, and an emulsion.
- the LNMP is a liposome selected from the group consisting of a cationic liposome, a nanoliposome, a proteoliposome, a unilamellar liposome, a multilamellar liposome, a ceramide-containing nanoliposome, and a multivesicular liposome.
- the LNMP is a lipid nanoparticle.
- the LNMP has a size of less than about 200 nm. In one embodiment, the LNMP has a size of less than about 150 nm. In one embodiment, the LNMP has a size of less than about 100 nm. In one embodiment, the LNMP has a size of about 55 nm to about 80 nm. In one embodiment, the LNMP has a size of about 80 nm to about 100 nm.
- the LNMP or LNP has an N:P ratio of at least 3, for instance, an N:P ratio of 3 to 100, 3 to 50, 3 to 30, 3 to 20, 3 to 15, 3 to 12, 6 to 30, 6 to 20, 6 to 15, or 6 to 12.
- the N/P ratio is 6 ⁇ 1 .
- the N/P ratio is 3 ⁇ 1 .
- the N/P ratio is 15 ⁇ 1 .
- the polypeptide is erythropoietin or Epogen, or a fragment or subunit thereof.
- one or more polynucleotides encode one or more components of a gene editing system. In some embodiments, one or more polynucleotides encoded one or more gene editing systems. In some embodiments, the gene editing system comprises an RNA-guided DNA- binding agent.
- the polynucleotide may be a mRNA, an siRNA or siRNA precursor, a microRNA (miRNA) or miRNA precursor, a plasmid, a Dicer substrate small interfering RNA (dsiRNA), a short hairpin RNA (shRNA), an asymmetric interfering RNA (aiRNA), a peptide nucleic acid (PNA), a morpholino, a locked nucleic acid (LNA), a piwi-interacting RNA (piRNA), a ribozyme, a deoxyribozyme (DNAzyme), an aptamer, a circular RNA (circRNA), a guide RNA (gRNA), a singleguide RNA (sgRNA), or a DNA molecule encoding any of these RNAs.
- dsiRNA Dicer substrate small interfering RNA
- shRNA short hairpin RNA
- aiRNA asymmetric interfering RNA
- PNA peptide nucleic
- the one or more polynucleotides comprise an mRNA or modified mRNA.
- the mRNA is derived from (a) a DNA molecule, or (b) an RNA molecule.
- T is optionally substituted with U.
- the mRNA is derived from a DNA molecule.
- the DNA molecule can further comprise a promoter.
- the promoter is a T7 promoter, a T3 promoter, or an SP6 promoter.
- the promoter is located at the 5’ UTR.
- the mRNA is derived from an RNA molecule.
- the RNA molecule may be a self-replicating RNA molecule.
- the mRNA is an RNA molecule.
- the RNA molecule may further comprise a 5’ cap.
- the 5’ cap can have a Cap 1 structure, a Cap 1 (m6A) structure, a Cap 2 structure, a Cap 3 structure, a Cap 0 structure, or any combination thereof.
- the mRNA comprises a 5' untranslated region (UTR) and/or a 3' UTR.
- UTR 5' untranslated region
- the mRNA comprises a 5' UTR.
- the 5' UTR may comprise a Kozak sequence.
- the mRNA comprises a 3' UTR.
- the 3’ UTR comprises one or more sequences derived from an amino-terminal enhancer of split (AES).
- the 3’ UTR comprises a sequence derived from mitochondrially encoded 12S rRNA (mtRNRI).
- the mRNA comprises a poly(A) sequence.
- the poly(A) sequence is a 110-nucleotide sequence consisting of a sequence of 30 adenosine residues, a 10-nucleotide linker sequence, and a sequence of 70 adenosine residues.
- the RNA-guided DNA-binding agent of the gene editing system is a Cas nuclease mRNA.
- the Cas nuclease mRNA is a Class II Cas nuclease mRNA.
- the Class II Cas nuclease is a Cas9 nuclease mRNA.
- the one or more polynucleotides comprise a gRNA or modified gRNA.
- the gRNA is a dual-guide RNA (dgRNA) or an sgRNA.
- the gRNA is a modified gRNA comprising a modification selected from the group consisting of 2'-O- methyl (2'-O-Me) modified nucleotide, a phosphorothioate (PS) bond between nucleotides, and a 2'- fiuoro (2'-F) modified nucleotide.
- the gRNA is a modified gRNA comprising a modification at one or more of the first five nucleotides at the 5' end or the 3' end. In some embodiments, the gRNA is a modified gRNA comprising PS bonds between the first four nucleotides or the last four nucleotides. In some embodiments, the modified gRNA further comprises 2'-O-Me modified nucleotides at the first three nucleotides at the 5' end or the 3' end.
- the one or more polynucleotides comprise a gRNA and a Class II Cas nuclease mRNA.
- the gRNA and Class 2 Cas nuclease mRNA are present in a ratio ranging from about 10:1 to about 1 :10 by weight.
- the gRNA and Class 2 Cas nuclease mRNA are present in a ratio ranging from about 5:1 to about 1 :5 by weight.
- the gRNA and Class 2 Cas nuclease mRNA are present in a ratio ranging from about 2:1 to about 1 :2 by weight.
- the gRNA and Class 2 Cas nuclease mRNA are present in a ratio of about 2:1 by weight or about 1 :1 by weight.
- the one or more polynucleotides comprise an RNA comprising an open reading frame encoding an RNA-guided DNA-binding agent, wherein the open reading frame has a uridine content ranging from its minimum uridine content to 150% of the minimum uridine content.
- the one or more polynucleotides comprise an mRNA comprising an open reading frame encoding an RNA-guided DNA-binding agent, wherein the open reading frame has a uridine dinucleotide content ranging from its minimum uridine dinucleotide content to 150% of the minimum uridine dinucleotide content.
- the RNA composition further comprises at least one template nucleic acid.
- the RNA composition has a total lipid: polynucleotide weight ratio of about 50:1 to about 10:1. In one embodiment, the RNA composition has a total lipid:polynucleotide weight ratio of about 44: 1 to about 24: 1 . In one embodiment, the RNA composition has a total lipid : polynucleotide weight ratio of about 40: 1 to about 28: 1 . In one embodiment, the RNA composition has a total lipid :polynucleotide weight ratio of about 38: 1 to about 30:1 . In one embodiment, the RNA composition has a total lipid :polynucleotide weight ratio of about 37: 1 to about 33:1.
- the RNA composition e.g., the aqueous phase, further comprises a HEPES or TRIS buffer.
- the HEPES or TRIS buffer may have a pH of about 7.0 to about 8.5.
- the HEPES or TRIS buffer can be at a concentration of about 7 mg/mL to about 15 mg/mL.
- the aqueous phase may further comprise about 2.0 mg/mL to about 4.0 mg/mL of NaCI.
- the RNA composition e.g., the aqueous phase comprises water, PBS, or a citrate buffer.
- the aqueous phase comprises a citrate buffer having a pH of about 3.2.
- the aqueous phase and the lipid solution are mixed at a 3:1 volumetric ratio.
- the RNA composition further comprises one or more cryoprotectants.
- the one or more cryoprotectants may be sucrose, glycerol, or a combination thereof.
- the RNA composition comprises a combination of sucrose at a concentration of about 70 mg/mL to about 110 mg/mL and glycerol at a concentration of about 50 mg/mL to about 70 mg/mL.
- the RNA composition is a lyophilized composition.
- the lyophilized RNA composition may comprise one or more lyoprotectants.
- the lyophilized RNA composition may comprise a poloxamer, potassium sorbate, sucrose, or any combination thereof.
- the lyophilized RNA composition comprises a poloxamer, e.g., poloxamer 188.
- the RNA composition is a lyophilized composition.
- the lyophilized RNA composition comprises about 0.01 to about 1 .0 % w/w of the polynucleotides.
- the lyophilized RNA composition comprises about 1.0 to about 5.0 % w/w lipids.
- the lyophilized RNA composition comprises about 0.5 to about 2.5 % w/w of TRIS buffer.
- the lyophilized RNA composition comprises about 0.75 to about 2.75 % w/w of NaCI.
- the lyophilized RNA composition comprises about 85 to about 95 % w/w of a sugar, e.g., sucrose.
- the lyophilized RNA composition comprises about 0.01 to about 1 .0 % w/w of a poloxamer, e.g., poloxamer 188. In one embodiment, the lyophilized RNA composition comprises about 1 .0 to about 5.0 % w/w of potassium sorbate.
- RNA composition comprising: one or more polynucleotides encoding one or more components of a gene editing system or one or more gene editing systems, formulated within
- LNPs lipid nanoparticles
- lipid reconstructed natural messenger packs comprising natural lipids and an ionizable lipid, wherein the ionizable lipid has two or more of the characteristics listed below:
- each of the lipid tails is at least 6 carbon atoms in length;
- a method of gene editing in a cell or a subject comprising contacting the cell with or administering to the subject: one or more polynucleotides encoding one or more components of a gene editing system or one or more gene editing systems, formulated within
- LNPs lipid nanoparticles
- lipid reconstructed natural messenger packs comprising natural lipids and an ionizable lipid, wherein the ionizable lipid has two or more of the characteristics listed below:
- each of the lipid tails is at least 6 carbon atoms in length;
- the RNA composition is administered at least one time.
- the RNA composition is administered at least twice, at least three times, at least four times, at least five times, at least six times, at least seven times, at least eight times, at least nine times, at least ten times, at least fifteen times, at least twenty times, or more. In some embodiments, the RNA composition is administered 2-8 times. In some embodiments, the delivery of the gene editing system, or the result of gene editing improves upon multiple administrations.
- the RNA composition is administered to the subject once, twice, three times, four times, five times, six times, seven times, eight times, nine times, ten times or more. In some embodiments, the RNA composition is administered to the subject eight times. In some embodiments, the RNA composition is administered to the subject five times a week apart. In some embodiments, the RNA composition is administered to the subject twice, three times, four times, five times, six times, seven times, eight times, ten times or more, wherein the RNA composition is administered to the subject one week apart, two weeks apart, three weeks apart, four weeks apart, five weeks apart, or more.
- the RNA composition further comprises at least one template nucleic acid.
- RNA compositions are administered into the subject or contacted with the cell: a first RNA composition comprising a mRNA and the second RNA composition comprising a guide RNA nucleic acid.
- first and second RNA compositions are administered simultaneously.
- the first and second RNA compositions are administered sequentially.
- a single RNA composition is contacted with the cell or administered into the subject, wherein the single RNA composition comprises an mRNA and a guide RNA nucleic acid.
- the RNA composition may be administered by oral, intravenous, intradermal, intramuscular, intranasal, intraocular, or rectal, and/or subcutaneous administration.
- the RNA composition is administered by oral, intravenous, intramuscular, and/or subcutaneous administration.
- the RNA composition is administered at a dosage level sufficient to deliver about 0.01 mg/kg to about 4 mg/kg of the mRNA to the subject. In some embodiments, the RNA composition is administered at a dosage level sufficient to deliver 0.125 mg/kg, 0.3 mg/kg, 0.5 mg/kg, 1 mg/kg, 1 .5 mg/kg, or 3 mg/kg of the mRNA to the subject.
- the method further comprises administering an additional therapeutic agent to the subject.
- the additional therapeutic agent is administered prior to, concurrent with, or after the administration of the RNA composition.
- the RNA composition may be formulated with b) CLPs such as LNMPs, as described herein.
- the ionizable lipid of the LNMPs is selected from the group consisting of 1 ,1 ’-((2-(4-(2-((2-(bis(2-hydroxydodecyl)amino)ethyl) (2-hydroxydodecyl)amino)ethyl)piperazin-1- yl)ethyl)azanediyl)bis(dodecan-2-ol) (C12-200), MD1 (CKK-E12), OF2, EPC, ZA3-Ep10, TT3, LP01 , 5A2-SC8, Lipid 5, SM-102 (Lipid H), and ALC-315.
- the ionizable lipid is C12- 200.
- the ionizable lipid is wherein R is C8-C14 alkyl group.
- the RNA composition may be formulated with a) LNPs, as described herein.
- the ionizable lipid in CLP such as LNMP
- LNP may be selected from one of the following groups of compounds: i) a compound of formula pharmaceutically acceptable salt thereof, or a stereoisomer of any of the foregoing, wherein: each A is independently Ci-C branched or unbranched alkyl or C1-C16 branched or unbranched alkenyl, optionally substituted with heteroatom or substituted with OH, SH, or halogen; each B is independently Ci-C branched or unbranched alkyl or Ci-C branched or unbranched alkenyl, optionally substituted with heteroatom or substituted with OH, SH, or halogen
- each R 6 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, or cycloalkyl
- each R 7 and each R 8 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, halogen, OH, SH, NR10R11, wherein each R10 and Rn is independently H, C1-C3 alkyl, or R10 and Rn are taken together to form a heterocyclic ring
- each s is independently 1 , 2, 3, 4, or 5
- each u is independently 1 , 2, 3, 4, or 5
- t is 1 , 2, 3, 4 or 5
- each Z is independently absent, O, S, or NR12, wherein R12 is H, C1-C7 branched or unbranched alkyl, or C2-C7 branched or unbranched alkenyl; and
- Q is O, S, or NR13, wherein each R13 is H, C1-C5 alkyl; ii) a compound of formula pharmaceutically acceptable salt thereof, or a stereoisomer of any of the foregoing, wherein: cyclic or heterocyclic moiety;
- Y is alkyl, hydroxy, hydroxyalkyl or
- A is absent, -O-, -N(R 7 )-, -O-alkylene-, -alkylene-O-, -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, -N(R 7 )C(O)N(R 7 )-, -S-, -S-S-, or a bivalent heterocycle; each of X and Z is independently absent, -O-, -CO-, -N(R 7 )-, -O-alkylene-;
- W is hydroxyl, substituted or unsubstituted hydroxyalkyl, substituted or unsubstituted amino, substituted or unsubstituted aminocarbonyl, or substituted or unsubstituted heterocylyl or heteroaryl; and iii) a compound of formula (V), pharmaceutically acceptable salts thereof, and stereoisomers of any of the foregoing, wherein:
- R20 and R30 are each independently H, C1-C5 branched or unbranched alkyl, or C2-C5 branched or unbranched alkenyl, or R20 and R30 together with the adjacent N atom form a 3 to 7 membered cyclic ring, optionally substituted with R a ;
- R a is H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, halogen, OH, or SH; each Ri and each R 2 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, OH, halogen, SH, or NR10R11, or
- Ri and R 2 are taken together to form a cyclic ring; each R10 and Rn is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, or R10 and Rn are taken together to form a heterocyclic ring; n is 0, 1 , 2, 3 or 4;
- Y is O or S
- E is each independently -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, -C(O-RI 3 )-O-, -C(O)O(CH 2 )r, -C(O)N(R 7 ) (CH 2 )r, -S-S-, or -C(O-Ri 3 )-O-(CH 2 )r, wherein each R 7 is independently H, alkyl, alkenyl, cycloalkyl, hydroxyalkyl, or aminoalkyl;
- RI 3 is branched or unbranched C 3 -Cw alkyl; r is 1 , 2, 3, 4, or 5;
- R a is each independently C1-C5 alkyl, C 2 -Cs alkenyl, or C 2 -Cs alkynyl; u1 and u2 are each independently 0, 1 , 2, 3, 4, 5, 6, or 7;
- R* is each independently H, C1-C16 branched or unbranched alkyl or C1-C16 branched or unbranched alkenyl, optionally interrupted with heteroatom or substituted with OH, SH, or halogen, or cycloalkyl or substituted cycloalkyl; represents the bond connecting the tail group to the head group; and wherein the lipid has a pKa from about 4 to about 8.
- the ionizable lipid is a compound in Table I, Table II, Table III, or Table IV. In some embodiments, the ionizable lipid is 2272, 2320, 2439, 2356, 2243, 2431 , 2455, 2454, 2424, 2433, 2425, 2275, 2220, or 2335.
- RNA composition comprising: one or more polynucleotides (e.g., those encoding one or more components of a gene editing system or one or more gene editing systems), formulated within
- LNPs lipid nanoparticles
- lipid reconstructed natural messenger packs comprising natural lipids and an ionizable lipid, wherein the ionizable lipid has two or more of the characteristics listed below:
- each of the lipid tails is at least 6 carbon atoms in length;
- the RNA composition is administered at least twice, at least three times, at least four times, at least five times, at least six times, at least seven times, at least eight times, at least nine times, at least ten times, at least fifteen times, at least twenty times, or more. In some embodiments, the RNA composition is administered 2-8 times. In some embodiments, the delivery of the RNA composition or the result of dosing improves upon multiple administrations. [0089] In some embodiments, the RNA composition is administered to the subject once, twice, three times, four times, five times, six times, seven times, eight times, nine times, ten times or more. In some embodiments, the RNA composition is administered to the subject eight times.
- the RNA composition is administered to the subject five times a week apart. In some embodiments, the RNA composition is administered to the subject twice, three times, four times, five times, six times, seven times, eight times, ten times or more, wherein the RNA composition is administered to the subject one week apart, two weeks apart, three weeks apart, four weeks apart, five weeks apart, or more.
- the term “effective amount,” “effective concentration,” or “concentration effective to” refers to an amount of a LNMP, or nucleic acid composition, sufficient to effect the recited result or to reach a target level (e.g., a predetermined or threshold level) in or on a target organism.
- the term “therapeutic agent” refers to an agent that can act on an animal, e.g., a mammal (e.g., a human), an animal pathogen, or a pathogen vector, such as an antifungal agent, an antibacterial agent, a virucidal agent, an anti-viral agent, an insecticidal agent, a nematicidal agent, an antiparasitic agent, or an insect repellent.
- nucleic acid and “polynucleotide” are interchangeable and refer to RNA or DNA that is linear or branched, single or double stranded, or a hybrid thereof, regardless of length (e.g., at least 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 30, 40, 50, 100, 150, 200, 250, 500, 1000, or more nucleic acids).
- the term also encompasses RNA/DNA hybrids.
- Nucleotides are typically linked in a nucleic acid by phosphodiester bonds, although the term “nucleic acid” also encompasses nucleic acid analogs having other types of linkages or backbones (e.g., phosphoramide, phosphorothioate, phosphorodithioate, O-methylphosphoroamidate, morpholino, locked nucleic acid (LNA), glycerol nucleic acid (GNA), threose nucleic acid (TNA), and peptide nucleic acid (PNA) linkages or backbones, among others).
- the nucleic acids may be single-stranded, double-stranded, or contain portions of both single-stranded and double-stranded sequence.
- a nucleic acid can contain any combination of deoxyribonucleotides and ribonucleotides, as well as any combination of bases, including, for example, adenine, thymine, cytosine, guanine, uracil, and modified or non-canonical bases (including, e.g., hypoxanthine, xanthine, 7-methylguanine, 5,6-dihydrouracil, 5-methylcytosine, and 5 hydroxymethylcytosine).
- bases including, for example, adenine, thymine, cytosine, guanine, uracil, and modified or non-canonical bases (including, e.g., hypoxanthine, xanthine, 7-methylguanine, 5,6-dihydrouracil, 5-methylcytosine, and 5 hydroxymethylcytosine).
- peptide encompasses any chain of naturally or non-naturally occurring amino acids (either D- or L-amino acids), regardless of length (e.g., at least 2, 3, 4, 5, 6, 7, 10, 12, 14, 16, 18, 20, 25, 30, 40, 50, 100, 150, 200, 250, 300, 350, 400, 450, 500, 550, 600, 650, 700, 750, 800, 850, 900, 950, 1000, or more than 1000 amino acids), the presence or absence of post-translational modifications (e.g., glycosylation or phosphorylation), or the presence of, e.g., one or more non-amino acyl groups (for example, sugar, lipid, etc.) covalently linked to the peptide, and includes, for example, natural proteins, synthetic, or recombinant polypeptides and peptides, hybrid molecules, peptoids, or peptidomimetics.
- amino acids either D- or L-amino acids
- length e.g., at least 2, 3,
- the polypeptide may be, e.g., at least 0.1 , at least 1 , at least 5, at least 10, at least 15, at least 20, at least 30, at least 40, at least 50, or more than 50 kD in size.
- the polypeptide may be a full-length protein.
- the polypeptide may comprise one or more domains of a protein.
- animal refers to humans and non-human animals (including for example, dogs, cats, horses, rabbits, zoo animals, cows, pigs, sheep, chickens, and non-human primates).
- pathogen refers to an organism, such as a microorganism or an invertebrate, which causes disease or disease symptoms in an animal by, e.g., (i) directly infecting the animal, (ii) producing agents that causes disease or disease symptoms in an animal (e.g., bacteria that produce pathogenic toxins and the like), and/or (iii) by eliciting an immune (e.g., inflammatory response) in animals (e.g., biting insects, e.g., bedbugs).
- an immune e.g., inflammatory response
- pathogens include, but are not limited to, bacteria, protozoa, parasites, fungi, nematodes, insects, viroids and viruses, or any combination thereof, wherein each pathogen is capable, either by itself or in concert with another pathogen, of eliciting disease or symptoms in humans.
- heterologous refers to an agent (e.g., a polypeptide) that is either (1) exogenous to the plant (e.g., originating from a source that is not the plant or plant part from which the PMP is produced) (e.g., an agent which is added to the PMP using loading approaches described herein) or (2) endogenous to the plant cell or tissue from which the PMP is produced, but present in the PMP (e.g., added to the PMP using loading approaches described herein, genetic engineering, as well as in vitro or in vivo approaches) at a concentration that is higher than that found in nature (e.g., higher than a concentration found in a naturally-occurring plant extracellular vesicle).
- agent e.g., a polypeptide
- percent identity between two sequences is determined by the BLAST 2.0 algorithm, which is described in Altschul et al., (1990) J. Mol. Biol. 215:403-410. Software for performing BLAST analyses is publicly available through the National Center for Biotechnology Information.
- modified NMPs or “modified LNMPs” refers to a composition including a plurality of NMPs or LNMPs that include one or more heterologous agents (e.g., one or more exogenous lipids, such as a ionizable lipids, e.g., a NMP or LNMP comprising an ionizable lipid and a sterol and/or a PEGylated lipid) capable of increasing cell uptake (e.g., animal cell uptake, plant cell uptake, bacterial cell uptake, or fungal cell uptake) of the NMP or LNMP, or a portion or component thereof, relative to an unmodified NMP or LNMP; capable of enabling or increasing delivery of a heterologous functional agent (e.g., an agricultural or therapeutic agent) by the NMP or LNMP to a cell, and/or capable of enabling or increasing loading (e.g., loading efficiency or loading capacity)
- heterologous agents e.g., one
- the NMPs or LNMPs may be modified in vitro or in vivo.
- the term “unmodified NMPs” or “unmodified LNMPs” refers to a composition including a plurality of NMPs or LNMPs that lack a heterologous cell uptake agent capable of increasing cell uptake (e.g., animal cell uptake, plant cell uptake, bacterial cell uptake, or fungal cell uptake) of the NMP.
- modified PMPs or “modified LPMPs” refers to a composition including a plurality of PMPs or LPMPs that include one or more heterologous agents (e.g., one or more exogenous lipids, such as a ionizable lipids, e.g., a PMP or LPMP comprising an ionizable lipid and a sterol and/or a PEGylated lipid) capable of increasing cell uptake (e.g., animal cell uptake, plant cell uptake, bacterial cell uptake, or fungal cell uptake) of the PMP or LPMP, or a portion or component thereof, relative to an unmodified PMP or LPMP; capable of enabling or increasing delivery of a heterologous functional agent (e.g., an agricultural or therapeutic agent) by the PMP or LPMP to a cell, and/or capable of enabling or increasing loading (e.g., loading efficiency or loading capacity)
- heterologous agents e.g., one
- unmodified PMPs or “unmodified LPMPs” refers to a composition including a plurality of PMPs or LPMPs that lack a heterologous cell uptake agent capable of increasing cell uptake (e.g., animal cell uptake, plant cell uptake, bacterial cell uptake, or fungal cell uptake) of the PMP.
- a heterologous cell uptake agent capable of increasing cell uptake (e.g., animal cell uptake, plant cell uptake, bacterial cell uptake, or fungal cell uptake) of the PMP.
- the term “cell uptake” refers to uptake of a NMP or LNMP or a portion or component thereof (e.g., a polynucleotide carried by the NMP or LNMP) by a cell, such as an animal cell, a plant cell, bacterial cell, or fungal cell.
- uptake can involve transfer of the NMP (e.g., LNMP) or a portion of component thereof from the extracellular environment into or across the cell membrane, the cell wall, the extracellular matrix, or into the intracellular environment of the cell).
- NMPs e.g., LNMPs
- Cell uptake includes aspects in which the entire NMP (e.g., LNMP) is taken up by a cell, e.g., taken up by endocytosis.
- one or more polynucleotides are exposed to the cytoplasm of the target cell following endocytosis and endosomal escape.
- a modified LNMP e.g., a LNMP comprising an ionizable lipid, e.g., a LNMP comprising an ionizable lipid and a sterol and/or a PEGylated lipid
- Cell uptake also includes aspects in which the NMP (e.g., LNMP) fuses with the membrane of the target cell.
- the NMP e.g., LNMP
- one or more polynucleotides are exposed to the cytoplasm of the target cell following membrane fusion.
- a LNMPs has an increased rate of fusion with the membrane of the target cell (e.g., is more fusogenic) relative to an unmodified LNMP.
- cell-penetrating agent refers to agents that alter properties (e.g., permeability) of the cell wall, extracellular matrix, or cell membrane of a cell (e.g., an animal cell, a plant cell, a bacterial cell, or a fungal cell) in a manner that promotes increased cell uptake relative to a cell that has not been contacted with the agent.
- a cell e.g., an animal cell, a plant cell, a bacterial cell, or a fungal cell
- plant refers to whole plants, plant organs, plant tissues, seeds, plant cells, seeds, and progeny of the same.
- Plant cells include, without limitation, cells from seeds, suspension cultures, embryos, meristematic regions, callus tissue, leaves, roots, shoots, gametophytes, sporophytes, pollen, and microspores.
- Plant parts include differentiated and undifferentiated tissues including, but not limited to the following: roots, stems, shoots, leaves, pollen, seeds, fruit, harvested produce, tumor tissue, and various forms of cells and culture (e.g., single cells, protoplasts, embryos, and callus tissue).
- the plant tissue may be in a plant or in a plant organ, tissue, or cell culture.
- a plant may be genetically engineered to produce a heterologous protein or RNA.
- Bacteria refers to whole bacteria or parts of bacteria. Further divisions of bacteria can be classified as coccals, bacillus, spirillum, or vibrio, and varying phylums include but are not limited to Proteobacteria, Firmicutes, Bacteroids, sphingobacteria, Flavobacteria, Fusobacteria, Spirochaetes, Chlorobia, Cyanobacteria, Thermomicrobia, Xenobacteria, or Aquificae.
- Example of specific bacteria species include Staphylococcus aureus, Escherichia coli, Streptococcus pneumoniae, and Pseudomonas aeruginosa.
- Parts of bacteria include cellular components such as peptidoglycan, outer membranes, inner membranes, cell walls, RNA polymerase, metabolic products, polypeptides, proteins.
- a bacteria may be genetically engineered to produce a heterologous protein or RNA, or may be genetically engineered to not produce an endogenous protein or RNA.
- Arthropod refers to any animal within the phylum Arthropoda, or any animal section, part, organ, tissue, egg, cell, or progeny of the same.
- Example animals include insects, spiders, and crustaceans.
- Arthropod cells include, without limitation, cells from eggs, suspension cultures, embryos, tissue, organs, exoskeletion, segments, and appendages.
- Arthropod parts include body segments, appendages, exoskeleton, eggs, organs, embryos, and various forms of cells and culture.
- Arthropod tissue may be in an arthropod or in an organ, tissue, or cell culture.
- An arthropod may be genetically engineered to produce a heterologous protein or RNA.
- An arthropod may be genetically engineered to not produce an endogenous protein or RNA.
- Fungi refers to whole fungi, fungi organs, fungi tissue, spores, fungi cells, and progeny of the same.
- Example fungi include yeasts, mushrooms, molds, and mildews.
- Fungi cells include without limitation cells from spores, suspension cultures, mycelium, hyphae, thallus, cell walls, tissue, gametophytes, sporophytes, and organs.
- Fungal tissue may be in a fungus or in an organ, tissue, or cell culture.
- a fungus may be genetically engineered to produce a heterologous protein or RNA.
- a fungus may be genetically engineered to not produce an endogenous protein or RNA.
- the term “Archaea” refers to whole archaea or parts of archaea.
- Example archaea include euryarchaeota, crenarchaeota, and koraarchaeota.
- Parts of archaea include cellular components such as RNA polymerases, glycerol-ether lipids, membranes, cell walls, polypeptides, proteins, and metabolic products.
- An archaea may be genetically engineered to produce a heterologous protein or RNA, or may be genetically engineered to not produce an endogenous protein or RNA.
- the term “extracellular vesicle” or “EV” refers to an enclosed lipid-bilayer structure naturally occurring in an organism or cell.
- the EV includes one or more EV markers.
- the term “EV marker” refers to a component that is naturally associated with a specific organism, such as a protein, a nucleic acid, a small molecule, a lipid, or a combination thereof. In some instances, the EV marker is an identifying marker of an EV but is not a pesticidal agent.
- the EV marker is an identifying marker of an EV and also a pesticidal agent (e.g., either associated with or encapsulated by the plurality of NMPs, or not directly associated with or encapsulated by the plurality of NMPs).
- a pesticidal agent e.g., either associated with or encapsulated by the plurality of NMPs, or not directly associated with or encapsulated by the plurality of NMPs.
- the term “natural messenger pack” or “NMP” refers to a lipid structure (e.g., a lipid bilayer, unilamellar, multilamellar structure; e.g., a vesicular lipid structure), that is about 5-2000 nm (e.g., at least 5-1000 nm, at least 5-500 nm, at least 400-500 nm, at least 25-250 nm, at least 50- 150 nm, or at least 70-120 nm) in diameter that is derived from (e.g., enriched, isolated or purified from) a natural source or segment, portion, or extract thereof, including lipid or non-lipid components (e.g., peptides, nucleic acids, or small molecules) associated therewith and that has been enriched, isolated or purified from a natural source, such as a Plant, Arthropod, Fungi, Archaea, or Bacteria ; an Arthropod, Plant, F
- NMPs are also able to be isolated from other natural sources, such as algae or animal organs.
- NMPs may be highly purified preparations of naturally occurring EVs.
- at least 1% of contaminants or undesired components from the source are removed (e.g., at least 2%, 5%, 10%, 15%, 20%, 25%, 30%, 40%, 45%, 50%, 55%, 60%, 70%, 80%, 90%, 95%, 96%, 98%, 99%, or 100%) of one or more contaminants or undesired components from the source, e.g., cell wall components; membrane components; chitin; organelles (e.g., mitochondria; nucleolus; golgi complex; ribosomes, endoplasmic reticulum and nuclei); chromatin; or molecular aggregates (e.g., protein aggregates, protein-nucleic acid aggregates, lipoprotein aggregates, or lipido-proteic structures).
- organelles e.g., mitochondria; nucleolus
- a NMP is at least 30% pure (e.g., at least 40% pure, at least 50% pure, at least 60% pure, at least 70% pure, at least 80% pure, at least 90% pure, at least 99% pure, or 100% pure) relative to the one or more contaminants or undesired components from the source as measured by weight (w/w), spectral imaging (% transmittance), or conductivity (S/m).
- the term “plant extracellular vesicle”, “plant EV”, or “EV” refers to an enclosed lipid-bilayer structure naturally occurring in a plant.
- the plant EV includes one or more plant EV markers.
- plant EV marker refers to a component that is naturally associated with a plant, such as a plant protein, a plant nucleic acid, a plant small molecule, a plant lipid, or a combination thereof, including but not limited to any of the plant EV markers listed in the Appendix.
- the plant EV marker is an identifying marker of a plant EV but is not a pesticidal agent.
- the plant EV marker is an identifying marker of a plant EV and also a pesticidal agent (e.g., either associated with or encapsulated by the plurality of PMPs or LPMPs, or not directly associated with or encapsulated by the plurality of PMPs or LPMPs).
- a pesticidal agent e.g., either associated with or encapsulated by the plurality of PMPs or LPMPs, or not directly associated with or encapsulated by the plurality of PMPs or LPMPs.
- the term “plant messenger pack” or “PMP” refers to a lipid structure (e.g., a lipid bilayer, unilamellar, multilamellar structure; e.g., a vesicular lipid structure), that is about 5-2000 nm (e.g., at least 5-1000 nm, at least 5-500 nm, at least 400-500 nm, at least 25-250 nm, at least 50- 150 nm, or at least 70-120 nm) in diameter that is derived from (e.g., enriched, isolated or purified from) a plant source or segment, portion, or extract thereof, including lipid or non-lipid components (e.g., peptides, nucleic acids, or small molecules) associated therewith and that has been enriched, isolated or purified from a plant, a plant part, or a plant cell, the enrichment or isolation removing one or more contaminants or undesired components from the source plant.
- lipid structure e.g
- PMPs may be highly purified preparations of naturally occurring EVs.
- at least 1% of contaminants or undesired components from the source plant are removed (e.g., at least 2%, 5%, 10%, 15%, 20%, 25%, 30%, 40%, 45%, 50%, 55%, 60%, 70%, 80%, 90%, 95%, 96%, 98%, 99%, or 100%) of one or more contaminants or undesired components from the source plant, e.g., plant cell wall components; pectin; plant organelles (e.g., mitochondria; plastids such as chloroplasts, leucoplasts or amyloplasts; and nuclei); plant chromatin (e.g., a plant chromosome); or plant molecular aggregates (e.g., protein aggregates, protein-nucleic acid aggregates, lipoprotein aggregates, or lipido-proteic structures).
- a PMP is at least 30% pure (e.g., at least 40% pure, at least 50% pure, at least 60% pure, at least 70% pure, at least 80% pure, at least 90% pure, at least 99% pure, or 100% pure) relative to the one or more contaminants or undesired components from the source plant as measured by weight (w/w), spectral imaging (% transmittance), or conductivity (S/m).
- a lipid reconstructed NMP is used herein.
- a lipid reconstructed PMP LPMP
- the terms “lipid reconstructed NMP” and “LNMP” refer to a NMP that has been derived from a lipid structure (e.g., a lipid bilayer, unilamellar, multilamellar structure; e.g., a vesicular lipid structure) derived from (e.g., enriched, isolated or purified from) an Arthropod, Plant, Fungi, Archaea, or Bacteria source, wherein the lipid structure is disrupted (e.g., disrupted by lipid extraction) and reassembled or reconstituted in a liquid phase (e.g., a liquid phase containing a cargo) using standard methods, e.g., reconstituted by a method comprising lipid film hydration, multifluidics, and/or solvent injection, to produce the L
- a lipid structure e.g.,
- the method may, if desired, further comprise sonication, freeze/thaw treatment, and/or lipid extrusion, e.g., to reduce the size of the reconstituted NMPs.
- LNMPs may be produced using a microfluidic device (such as a NanoAssemblr® IGNITETM microfluidic instrument (Precision NanoSystems)).
- a lipid reconstructed PMP is used herein as a subcategory of lipid reconstructed NMPs.
- the terms “lipid reconstructed PMP” and “LPMP” refer to a PMP that has been derived from a lipid structure (e.g., a lipid bilayer, unilamellar, multilamellar structure; e.g., a vesicular lipid structure) derived from (e.g., enriched, isolated or purified from) a plant source, wherein the lipid structure is disrupted (e.g., disrupted by lipid extraction) and reassembled or reconstituted in a liquid phase (e.g., a liquid phase containing a cargo) using standard methods, e.g., reconstituted by a method comprising lipid film hydration and/or solvent injection, to produce the LPMP, as is described herein.
- a lipid structure e.g., a lipid bilayer, unilamellar, multi
- the method may, if desired, further comprise sonication, freeze/thaw treatment, and/or lipid extrusion, e.g., to reduce the size of the reconstituted LPMPs.
- LPMPs may be produced using a microfluidic device (such as a NanoAssemblr® IGNITETM microfluidic instrument (Precision NanoSystems)).
- the term “pure” refers to a PMP preparation in which at least a portion (e.g., at least 20%, 25%, 30%, 40%, 45%, 50%, 55%, 60%, 70%, 80%, 90%, 95%, 96%, 98%, 99%, or 100%) of plant cell wall components, plant organelles (e.g., mitochondria, chloroplasts, and nuclei), or plant molecule aggregates (protein aggregates, protein-nucleic acid aggregates, lipoprotein aggregates, or lipido-proteic structures) have been removed relative to the initial sample isolated from a plant, or part thereof.
- plant organelles e.g., mitochondria, chloroplasts, and nuclei
- plant molecule aggregates protein aggregates, protein-nucleic acid aggregates, lipoprotein aggregates, or lipido-proteic structures
- complex lipid particle refers to a lipid particle that has a complexity characterized by comprising a wide variety of lipids, including structural lipids extracted from one or more natural sources (such as plants or bacteria), and optionally at least one exogenous ionizable lipid.
- the complex lipid particle may comprise between 10% w/w and 99% w/w structural lipids derived from a lipid structure from one or more natural sources, e.g., it may contain at least 10% w/w, at least 20% w/w, at least 30% w/w, at least 40% w/w, at least 50% w/w, at least 60% w/w, at least 70% w/w, at least 80% w/w, at least 90% w/w, at least 95% w/w, or about 99% w/w lipids derived from a lipid structure from one or more natural sources.
- a complex lipid particle incorporating natural lipid extracts may also be referred to as a natural messenger pack (NMP).
- a complex lipid particle incorporating plant lipid extracts may also be referred to as a plant messenger pack (PMP).
- PMP plant messenger pack
- a complex lipid particle incorporating natural lipid extracts and at least one exogenous ionizable lipid may also be referred to as a lipid reconstructed natural messenger pack (LNMP).
- LNMP lipid reconstructed natural messenger pack
- LPMP lipid reconstructed plant messenger pack
- the complex lipid particle may contain 3-1000 lipids extracted from one or more natural (e.g., plant, bacteria) sources.
- the complex lipid particle may contain natural (e.g., plant, bacteria) lipids from at least 1 , at least 2, at least 3, at least 4, at least 5, at least 6, at least 7, at least 8, at least 9, or at least 10 different classes or sub-classes of lipids from the natural (e.g., plant, bacteria) source.
- the complex lipid particle may comprise all or a fraction of the lipid species present in the lipid structure from the natural (e.g., plant, bacteria) source, e.g., it may contain at least 10%, at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, or virtually 100% of the lipid species present in the lipid structure from the natural source.
- the complex lipid particle may comprise all or a fraction of the lipid species present in the lipid structure from a particular natural source.
- it may contain at least 10%, at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, or virtually 100% of the lipid species present in the lipid structure from a plant source or from a bacteria source.
- the complex lipid particle may comprise reduced or minimized protein matter endogenous to the one or more natural (i.e. plant, bacteria) sources, e.g., it may contain 0% w/w, less than 1 % w/w, less than 5% w/w, less than 10% w/w, less than 15% w/w, less than 20% w/w, less than 30% w/w, less than 40% w/w, or less than 50% w/w of the protein matter endogenous to the one or more natural (e.g., plant, bacteria) sources.
- the lipid bilayer of the complex lipid particle does not contain proteins.
- the complex lipid particle may also include synthetic structural lipids such as neutral lipids as the structural lipid component.
- the structural lipid component of the complex lipid particle may comprise between 10% w/w and 99% w/w structural lipids derived from a synthetic lipid structure (as opposed to the lipids extracted from a natural source), e.g., it may contain at least 10% w/w, at least 20% w/w, at least 30% w/w, at least 40% w/w, at least 50% w/w, at least 60% w/w, at least 70% w/w, at least 80% w/w, at least 90% w/w, at least 95% w/w, or about 99% w/w lipids derived from a synthetic lipid structure.
- the complex lipid particle may further comprise at least two exogenous lipids.
- the complex lipid particle may include at least 1% w/w, at least 2% w/w, at least 5% w/w, at least 10% w/w, at least 15% w/w, at least 20% w/w, at least 25% w/w, at least 30% w/w, at least 40% w/w, at least 50% w/w, at least 60% w/w, at least 70% w/w, at least 80% w/w, or about 90% w/w exogenous lipids.
- exogenous lipids include sterols and PEG-lipid conjugate.
- the complex lipid particle may be used to encapsulate one or more exogenous nucleic acids or polynucleotides encoding one or more peptides, polypeptides, or proteins, to enable delivery of the exogenous nucleic acids or polynucleotides to a target cell or tissue.
- exogenous lipid refers to a lipid that is exogenous to the natural source (e.g., plant, bacteria), i.e., a lipid originates from a source that is not the natural source from which the lipids are extracted (e.g., a lipid that is added to the complex lipid particle formulation using method described herein).
- the term “exogenous lipid” does not exclude a natural-derived lipid (such as a plant-derived sterol).
- an exogenous lipid can be a natural-derived lipid (such as a plant-derived sterol that is exogenous to the plant source from which the lipids are extracted, e.g., an exogenous lipid can be a plant derived sterol that is added to the complex lipid particle formulation).
- an exogenous lipid can be a natural-derived lipid that is exogenous to the particular natural source from which the lipids are extracted (e.g., a bacteria-derived lipid that is exogenous to the plant source from which the lipids are extracted, or vice versa).
- An exogenous lipid may be a cell-penetrating agent, may be capable of increasing delivery of one or more polynucleotides by the complex lipid formulation to a cell, and/or may be capable of increasing loading (e.g., loading efficiency or loading capacity) of a polynucleotide.
- the exogenous lipid may be a stabilizing lipid.
- the exogenous lipid may be a structural lipid (e.g., a synthetic structural lipid).
- Exemplary exogenous lipids include ionizable lipids, synthetic structural lipids, sterols, and PEGylated lipids.
- cationic lipid refers to an amphiphilic molecule (e.g., a lipid or a lipidoid) that is positively charged, containing a cationic group (e.g., a cationic head group).
- a cationic group e.g., a cationic head group
- ionizable lipid refers to an amphiphilic molecule (e.g., a lipid or a lipidoid, e.g., a synthetic lipid or lipidoid) containing a group (e.g., a head group) that can be ionized, e.g., dissociated to produce one or more electrically charged species, under a given condition (e.g., pH). It has been surprisingly found that ionizable lipids comprising alkyl chains with multiple sites of unsaturation, e.g., at least two or three sites of unsaturation, are particularly useful for forming lipid particles with increased membrane fluidity.
- ionizable lipids and related analogs suitable for use herein, have been described in U.S. Patent Publication Nos. 20060083780 and 20060240554; U.S. Pat. Nos. 5,208,036; 5,264,618; 5,279,833; 5,283,185; 5,753,613; and 5,785,992; and PCT Publication No. WO 96/10390, the disclosures of which are herein incorporated by reference in their entirety for all purposes.
- ionizable lipids are ionizable such that they can dissociate to exist in a positively charged form depending on pH.
- the ionization of an ionizable lipid affects the surface charge of a lipid nanoparticle comprising the ionizable lipid under different pH conditions.
- the surface charge of the lipid nanoparticle in turn can influence its plasma protein absorption, blood clearance, and tissue distribution (Semple, S.C., et al., Adv. Drug Deliv Rev 32:3-17 (1998)) as well as its ability to form endosomolytic non-bilayer structures (Hafez, I.M., et al., Gene Ther 8: 1188-1196 (2001)) that can influence the intracellular delivery of nucleic acids.
- ionizable lipids are those that are generally neutral, e.g., at physiological pH (e.g., pH about 7), but can carry net charge(s) at an acidic pH or basic pH. In one embodiment, ionizable lipids are those that are generally neutral at pH about 7, but can carry net charge(s) at an acidic pH. In one embodiment, ionizable lipids are those that are generally neutral at pH about 7, but can carry net charge(s) at a basic pH. In some embodiments, ionizable lipids do not include those cationic lipids or anionic lipids that generally carry net charge(s) at physiological pH (e.g., pH about 7).
- lipidoid refers to a molecule having one or more characteristics of a lipid.
- stable LNMP formulation or “stable CLP formulation” refers to a CLP formulation or a LNMP composition that over a period of time (e.g., at least 24 hours, at least 48 hours, at least 1 week, at least 2 weeks, at least 3 weeks, at least 4 weeks, at least 30 days, at least 60 days, or at least 90 days) retains at least 5% (e.g., at least 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100%) of the initial number of CLPs or LNMPs (e.g., CLPs or LNMPs per mL of solution) relative to the number of CLPs or LNMPs in the CLP formulation or LNMP formulation (e.g., at the time of production or formulation) optionally at a defined temperature range (e.g., a temperature of at
- the expression refers to a CLP formulation or LNMP composition that over a period of time (e.g., at least 24 hours, at least 48 hours, at least 1 week, at least 2 weeks, at least 3 weeks, at least 4 weeks, at least 30 days, at least 60 days, or at least 90 days) retains at least 5% (e.g., at least 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, or 100%) of its activity relative to the initial activity of the CLP formulation or LNMP formulation (e.g., at the time of production or formulation) optionally at a defined temperature range (e.g., a temperature of at least 24°C (e.g., at least 24°C, 25°C, 26°C, 27°C, 28°C, 29°C, or 30°C), at least 20°C (e.g., at least 20°C, 21
- the expression refers to a CLP formulation or a LNMP formulation that over a period of time (e.g., at least 24 hours, at least 48 hours, at least 1 week, at least 2 weeks, at least 3 weeks, at least 4 weeks, at least 30 days, at least 60 days, or at least 90 days) retains their particle size, i.e., the particle size does not increase, or has an increase of no more than 5% (e.g., no more than 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 100%, 150%, 2-fold, 2.5-fold, or 3-fold) relative to the initial size of the CLPs or LNMPs (e.g., at the time of production or formulation) optionally at a defined temperature range (e.g., a temperature of at least 24°C (e.g., at least 24°C, 25°C, 26°C
- the stable CLP or LNMP formulation continues to encapsulate or remains associated with an exogenous peptide, polypeptide, or protein with which the CLP or LNMP formulation has been loaded, e.g., continues to encapsulate or remains associated with an exogenous peptide, polypeptide, or protein for at least 24 hours, at least 48 hours, at least 1 week, at least 2 weeks, at least 3 weeks, at least 4 weeks, at least 30 days, at least 60 days, at least 90 days, or 90 or more days.
- treatment refers to administering a pharmaceutical composition to an animal for prophylactic and/or therapeutic purposes.
- gene editing system or “genome editing system” refers to any genome editing technology including the CRISPR-Cas nuclease system, transcription activator-like effector nucleases (TALENS), zinc-finger nucleases (ZFNs), or other known means of inducing gene editing.
- CRISPR-Cas nuclease system transcription activator-like effector nucleases (TALENS), zinc-finger nucleases (ZFNs), or other known means of inducing gene editing.
- TALENS transcription activator-like effector nucleases
- ZFNs zinc-finger nucleases
- RNA-guided DNA binding agent means a polypeptide or complex of polypeptides having RNA and DNA binding activity, or a DNA-binding subunit of such a complex, wherein the DNA binding activity is sequence-specific and depends on the sequence of the RNA.
- RNA-guided DNA binding agents include Cas cleavases/nickases and inactivated forms thereof ("dCas DNA binding agents").
- Cas nuclease as used herein, encompasses Cas cleavases, Cas nickases, and dCas DNA binding agents.
- Cas cleavases/nickases and dCas DNA binding agents include a Csm or Cmr complex of a type III CRISPR system, the Cas 10, Csm1 , or Cmr2 subunit thereof, a Cascade complex of a type I CRISPR system, the Cas3 subunit thereof, and Class II Cas nucleases.
- a "Class II Cas nuclease” is a single-chain polypeptide with RNA-guided DNA binding activity.
- Class II Cas nucleases include Class II Cas cleavases/nickases (e.g., H840A, D 10A, or N863A variants), which further have RNA-guided DNA cleavase or nickase activity, and Class II dCas DNA binding agents, in which cleavase/nickase activity is inactivated.
- Class II Cas cleavases/nickases e.g., H840A, D 10A, or N863A variants
- Class II dCas DNA binding agents in which cleavase/nickase activity is inactivated.
- Class II Cas nucleases include, for example, Cas9, Cpf1 , C2c1 , C2c2, C2e3, HF Cas9 (e.g., N497A, R661A, Q695A, Q926A variants), HypaCas9 (e.g., N692A, M694A, Q695A, H69SA variants), eSPCas9(1.0) (e.g, K810A, K1003A, R1060A variants), and eSPCas9(1.1) (e.g., K848A, K1003A, R1060A variants) proteins and modifications thereof.
- Cas9, Cpf1 , C2c1 , C2c2, C2e3, HF Cas9 e.g., N497A, R661A, Q695A, Q926A variants
- HypaCas9 e.g., N692A, M
- Figure 1 B shows the amount of gene editing (%) of the target gene (TTR) in the bone marrow, spleen, and the targeted organ, the liver, with an intravenous dose of either an LPMP delivering CRISPR/Cas gene editing components (3 mg/kg recLemon LPMP; 1 :1 Cas9 mRNA:sgRNA mTTR_G211) or an equivalent volume of PBS.
- N 5/group.
- RNA compositions e.g., mRNA/gRNA encoding gene editing systems
- the RNA compositions may be dosed repeatedly (e.g., 5 times, 6 times, 7 times, 8 times).
- RNA compositions include one or more polynucleotides (e.g., RNA such as messenger RNA (mRNA or gRNA)) encoding one or more components of a gene editing system, or one or more gene editing systems, formulated within a lipid nanoparticle (LNP) or a complex lipid particle (CLP).
- LNP lipid nanoparticle
- CLP complex lipid particle
- the CLP is a lipid reconstructed natural messenger packs (LNMPs) comprising lipids extracted from one or more natural sources (i.e., natural lipids) and an ionizable lipid.
- NMPs are lipid assemblies produced wholly or in part from natural extracellular vesicles (EVs), or segments, portions, or extracts thereof.
- PMPs are lipid assemblies produced wholly or in part from plant extracellular vesicles (EVs), or segments, portions, or extracts thereof.
- LNMPs are NMPs derived from a lipid structure wherein the lipid structure is disrupted and reassembled or reconstituted in a liquid phase.
- LPMPs are PMPs derived from a lipid structure wherein the lipid structure is disrupted and reassembled or reconstituted in a liquid phase.
- this disclosure also includes a method of repeatable dosing to a subject in need thereof, the method comprising administering to the subject the RNA composition comprising one or more polynucleotides, formulated within an LNP or LNMP, as described herein.
- the disclosure also includes a method for making an RNA composition, comprising reconstituting a film comprising purified NMP lipids in the presence of an ionizable lipid to produce a LNMP comprising the ionizable lipid, and loading into the LNMPs with one or more polynucleotides encoding one or more components of a gene editing system or one or more gene editing systems.
- Complex lipid particles (CLPs) described herein comprise a wide variety of lipids, including structural lipids extracted from one or more natural sources (such as plants or bacteria).
- a complex lipid particle is a natural messenger pack (NMP) incorporating natural lipid extracts.
- a complex lipid particle is a lipid reconstructed natural messenger pack (LNMP) incorporating natural lipid extracts and at least one exogenous ionizable lipid.
- the complex lipid particles may also comprise at least exogenous ionizable lipid.
- the ionizable lipid has two or more of the characteristics listed below:
- each of the lipid tails is at least 6 carbon atoms in length;
- the complex lipid particle may comprise between 10% w/w and 99% w/w structural lipids derived from a lipid structure from one or more natural sources, e.g., it may contain at least 10% w/w, at least 20% w/w, at least 30% w/w, at least 40% w/w, at least 50% w/w, at least 60% w/w, at least 70% w/w, at least 80% w/w, at least 90% w/w, at least 95% w/w, or about 99% w/w lipids derived from a lipid structure from one or more natural sources.
- the complex lipid particle comprises about 10-95% w/w of the natural (e.g., plant, bacteria) lipids.
- the complex lipid particle comprises about 25-95% w/w, about 30-95% w/w, about 35-95% w/w, about 40-95% w/w, about 45-95% w/w, about 50-95% w/w, about 55-95% w/w, about 60-95% w/w, about 65-95% w/w, about 70-95% w/w, about 75-95% w/w, about 80-95% w/w, or about 85-95% w/w of the natural (e.g., plant, bacteria) lipids based on the amounts of total lipids in the complex lipid formulation.
- the complex lipid particle may contain 3-1000 lipids extracted from one or more natural (e.g., plant, bacteria) sources.
- the natural source is a plant, plant extract, or fragment or part of a plant.
- the natural source is a bacteria, bacteria fragment or part of a bacteria.
- the natural source is lemon.
- the natural source is soy.
- the natural source is E. coli.
- the complex lipid particle contains at least 10 natural lipids belonging to one or more of the classes selected from the group consisting of fatty acyls (FA), fatty acyl conjugates, phospholipids, glycerolipids, glycolipids, glycerophospholipids, sphingolipids, waxes, and sterol.
- FA fatty acyls
- fatty acyl conjugates phospholipids, glycerolipids, glycolipids, glycerophospholipids, sphingolipids, waxes, and sterol.
- the complex lipid particle contains at least 15, at least 20, at least 25, at least 30, at least 35, at least 40, at least 45, at least 50, at least 60, at least 70, at least 80, at least 90, at least 100, at least 150, at least 200, at least 250, at least 300, at least 400, at least 500, at least 600, at least 700, or at least 800 natural lipids belonging to one or more of the classes selected from the group consisting of fatty acyls (FA), fatty acyl conjugates, phospholipids, glycerolipids, glycolipids, glycerophospholipids, sphingolipids, waxes, and sterol.
- the complex lipid particle contains lipids from at least two or at least three of these different classes.
- the complex lipid particle contains at least 10 natural lipids belonging to one or more of the classes selected from the group consisting of glycerolipid, sphingolipid, and sterol.
- the complex lipid particle contains at least 15, at least 20, at least 25, at least 30, at least 35, at least 40, at least 45, at least 50, at least 60, at least 70, at least 80, at least 90, at least 100, at least 150, at least 200, at least 250, at least 300, at least 400, at least 500, at least 600, at least 700, or at least 800 natural lipids belonging to one or more of the classes selected from the group consisting of glycerolipid, sphingolipid, and sterol.
- the complex lipid particle contains lipids from at least two or at least three of these different classes.
- the complex lipid particle may contain one or more glycerolipids (GL) or glycerophospholipids (GP), which may also include glycolipids.
- GL glycerolipids
- GP glycerophospholipids
- the complex lipid particle may contain one or more glycerolipids selected from the group consisting of phospholipids (PL), galactolipids, triacylglycerols (TG), and sulfolipids (SL).
- the CLPs contains one or more glycerophospholipids (GP) selected from the group consisting of phosphatidylcholines (PC), phosphatidylethanolamines (PE), phosphatidylserines (PS), and phosphatidylinositols (PI).
- GP glycerophospholipids
- the complex lipid particle contains one or more sphingolipids (SP) selected from the group consisting of sulfolipids (SL), glycosyl inositol phosphoryl ceramides (GIPC), glucosylceramides (GCer), ceramides (Cer), and free long-chain bases (LCB).
- SP sphingolipids
- the complex lipid particle contains one or more phytosterols selected from the group consisting of campesterol, stigmasterol, p-sitosterol, A5-avenasterol, brassicasterol, avenasterol, 4-desmethyl sterol, 4a-monomethyl sterol, A5-sterol, A7-sterol, a-spinasterol, A5,A7-sterol, phytostanol, and sitosterol.
- phytosterols selected from the group consisting of campesterol, stigmasterol, p-sitosterol, A5-avenasterol, brassicasterol, avenasterol, 4-desmethyl sterol, 4a-monomethyl sterol, A5-sterol, A7-sterol, a-spinasterol, A5,A7-sterol, phytostanol, and sitosterol.
- the CLP may contain one or more natural lipids belonging to one or more classes or sub- classes selected from the group consisting of fatty acids, fatty esters, fatty aldehydes, fatty amides, acyclic oxylipins, cyclic oxylipins, glycerolipids, monoradylglycerols, diradylglycerols, triradylglycerols, estolides, glycosylmonoacylglycerols, sulfoquinovosylmonoacylglycerols, monogalactosylmonoacylglycerol, digalactosylmonoacylglycerol, sulfoquinovosyldiacylglycerols, monogalactosyldiacylglycerol, digalactosyldiacylglycerol, glycosyldiacylglycerols, glycerop
- the complex lipid particle may contain one or more natural lipids belonging to one or more of the classes or sub-classes selected from the group consisting of the classes or sub-classes listed above.
- the complex lipid particle contains at least 15, at least 20, at least 25, at least 30, at least 35, at least 40, at least 45, at least 50, at least 60, at least 70, at least 80, at least 90, at least 100, at least 150, at least 200, at least 250, at least 300, at least 400, at least 500, at least 600, at least 700, or at least 800 natural lipids belonging to one or more of the classes or sub-classes selected from the group consisting of the classes or sub-classes listed above.
- the CLP contains one or more natural lipids belonging to one or more of the sub-classes selected from the group consisting of acyl diacylglyceryl glucuronides, acylhexosylceramides, acylsterylglycosides, bile acids, acyl carnitines, cholesteryl esters, ceramides, cardiolipins, coenzyme Qs, diacylglycerols, digalactosyldiacylglycerols, diacylglyceryl glucuronides, dilysocardiolipins, fatty acids, fatty acid esters of hydroxyl fatty acids, hemibismonoacylglycerophosphates, hexosylceramides, lysophosphatidic acids, lysophophatidylcholines, lysophosphatidylethanolamines, N-acyl-lysophosphatidy
- the complex lipid particle contains at least 10 natural (e.g., plant, bacteria) lipids belonging to one or more of the subclasses selected from the group consisting of the sub-classes listed above.
- the complex lipid particle contains at least 15, at least 20, at least 25, at least 30, at least 35, at least 40, at least 45, at least 50, at least 60, at least 70, at least 80, at least 90, at least 100, at least 150, at least 200, at least 250, at least 300, at least 400, at least 500, at least 600, at least 700, or at least 800 natural lipids belonging to one or more of the sub-classes selected from the group consisting of the subclasses listed above.
- the complex lipid particle may contain 10 or more natural lipids belonging to one or more of the sub-classes selected from the group consisting of acylsterylglycosides, ceramides, digalactosyldiacylglycerols, diacylglyceryl glucuronides, hemibismonoacylglycerophosphates, hexosylceramides, lysophophatidylcholines, lysophosphatidylethanolamines, monogalactosyldiacylglycerols, phosphatidylcholines, phosphatidylethanolamines, phosphatidylethanols, phosphatidylglycerols, phosphatidylinositols, sulfoquinovosyl diacylglycerosl, and sterols.
- the complex lipid particle contains at least 15, at least 20, at least 25, at least 30, at least 35, at least 40, at least 45, at least 50, at least 60, at least 70, at least 80, at least 90, at least 100, at least 150, at least 200, at least 250, at least 300, at least 400, at least 400, at least 500, at least 600, at least 700, or at least 800 natural lipids belonging to one or more of the subclasses selected from the group consisting of the sub-classes listed above.
- the complex lipid particle may contain natural lipids from at least 5, at least 6, at least 7, at least 8, at least 9, or at least 10 different classes or sub-classes of lipids from the natural sources.
- the complex lipid particle contains natural lipids from at least 5, at least 6, at least 7, at least 8, at least 9, or at least 10 different classes or sub-classes of lipids from a single natural source (e.g., from only the plant source, or from only the bacteria source).
- the CLP may contain natural lipids from only one class or only one sub-class of lipids from the natural sources.
- the identity (and class and subclass) and the amounts of the lipids extracted from the natural source(s) can be analyzed by lipidomics analysis by solubilizing the lipid extracts or complex lipid particles in compatible solvents and analyzing by a mass spectrometry (e.g., MS/MS).
- lipidomics analysis by solubilizing the lipid extracts or complex lipid particles in compatible solvents and analyzing by a mass spectrometry (e.g., MS/MS).
- Other known methods such as charged aerosol detection (CAD) (e.g., HPLC-CAD, normal-phase high- performance liquid chromatography (NP-HPLC-CAD), or reversed-phase high-performance liquid chromatography (RP-HPLC-CAD)), may also be used.
- CAD charged aerosol detection
- HPLC-CAD normal-phase high- performance liquid chromatography
- RP-HPLC-CAD reversed-phase high-performance liquid chromatography
- the complex lipid particle may comprise all or a fraction of the lipid species present in the lipid structure from the particular natural source(s), e.g., it may contain at least 10%, at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, or virtually 100% of the lipid species present in the lipid structure from the particular natural source(s).
- the complex lipid particle may comprise reduced or minimized protein matter endogenous to the one or more natural sources.
- the complex lipid particle may contain less than 50% w/w, less than 45% w/w, less than 40% w/w, less than 35% w/w, less than 30% w/w, less than 25% w/w, less than 20% w/w, less than 15% w/w, less than 10% w/w, less than 9% w/w, less than 8% w/w, less than 7% w/w, less than 6% w/w, less than 5% w/w, less than 4% w/w, less than 3% w/w, less than 2% w/w, less than 1 % w/w, less than 0.5% w/w, less than 0.1 % w/w, or essentially free of protein matter endogenous to the one or more natural sources.
- the lipid bilayer of the complex lipid particle does not contain proteins.
- protein concentration is divided by the concentration of the natural lipid extract and then multiplied by 100.
- %w/w is calculated as the percent of the mass of total protein endogenous to the one or more natural sources based on the mass of the total lipid extract.
- the complex lipid particle may comprise reduced or minimized residual dsDNA matter endogenous to the one or more natural sources.
- the complex lipid particle may contain less than 15% w/w, less than 10% w/w, less than 5% w/w, less than 1 % w/w, less than 0.5% w/w, less than 0.1 % w/w, less than 0.05% w/w, less than 0.01 % w/w, less than 0.005% w/w, less than 0.001 % w/w, or essentially free of residual dsDNA matter endogenous to the one or more natural sources.
- the lipid bilayer of the complex lipid particle does not contain residual dsDNA.
- total adjusted dsDNA concentration is divided by the concentration of the natural lipid extract and then multiplied by 100.
- %w/w is calculated as the percent of the mass of total residual dsDNA endogenous to the one or more natural sources based on the mass of the total lipid extract.
- the complex lipid particle further incorporates a synthetic structural lipid such as a neutral lipid.
- the structural lipid component of the complex lipid particle may comprise between 10% w/w and 99% w/w structural lipids derived from a synthetic lipid structure, e.g., it may contain at least 10% w/w, at least 20% w/w, at least 30% w/w, at least 40% w/w, at least 50% w/w, at least 60% w/w, at least 70% w/w, at least 80% w/w, at least 90% w/w, at least 95% w/w, or about 99% w/w lipids derived from a synthetic lipid structure.
- the complex lipid particle may further comprise at least two other exogenous lipids.
- the complex lipid particle may include at least 1 % w/w, at least 2% w/w, at least 5% w/w, at least 10% w/w, at least 15% w/w, at least 20% w/w, at least 25% w/w, at least 30% w/w, at least 40% w/w, at least 50% w/w, at least 60% w/w, at least 70% w/w, at least 80% w/w, at least 90% w/w, or about 95% w/w exogenous lipids.
- exogenous lipids include ionizable lipids, synthetic structural lipids, sterols, and PEG-lipid conjugate.
- the complex lipid particle may further comprise at least two exogenous lipids.
- the complex lipid particle contains an ionizable lipid, a sterol, and PEG-lipid conjugate. Additional exogenous lipids suitable for being included in the complex lipid particle are described herein below.
- the CLPs contain natural lipids comprising fatty acid-derived tails, said fatty acid-derived tails of the natural lipids being: about 5 to 20% of fatty acid 16:0 about 0 to 10% of fatty acid 18:1 (C9) about 0 to 10% of fatty acid 18:1 (C7) about 5 to 30% of fatty acid 18:2 about 2 to 20% of fatty acid 18:3.
- the CLPs contain phosphatidylcholine (PC) lipids comprising fatty acid-derived tails, said fatty acid-derived tails of the PC lipids being: about 10 to 20% of fatty acid 16:0 about 2 to 5% of fatty acid 18:0 about 7 to 15% of fatty acid 18:1 about 50 to 75% of fatty acid 18:2 about 2 to 10% of fatty acid 18:3.
- PC phosphatidylcholine
- the CLPs contain phosphatidylethanolamines (PE) lipids comprising fatty acid-derived tails, said fatty acid-derived tails of the PE lipids being: about 0.25 to 5% of fatty acid 14:0 about 25 to 45% of fatty acid 16:0 about 5 to 15% of fatty acid 16:1 about 10 to 25% of fatty acid 17:0 about 25 to 45% of fatty acid 18:1 about 2 to 7% of fatty acid 19:0.
- PE phosphatidylethanolamines
- the CLPs contain natural lipids belonging to the sub-classes of phosphatidylethanolamines, phosphatidylglycerol, and cardiolipin, and comprising: about 50 to 75 wt/wt% of phosphatidylethanolamines (PE) about 15 to 30 wt/wt% of phosphatidylglycerol (PG) about 5 to 15 wt/wt% of cardiolipin (CL).
- PE phosphatidylethanolamines
- PG phosphatidylglycerol
- CL cardiolipin
- the CLPs contain natural lipids comprising : about 10 to 50 wt/wt% of phosphatidylcholines (PC) about 5 to 50 wt/wt% of phosphatidylethanolamines (PE) about 0 to 15 wt/wt% of triacylglycerol (TG) about 5 to 35 wt/wt% of hexosylceramides (HexCer) about 0 to 5 wt/wt% of phosphatidylglycerol (PG) about 0 to 7 wt/wt% of phosphatidylserines (PS) about 0 to 10 wt/wt% of phosphatidylinositols (PI) about 0 to 5 wt/wt% of cardiolipin (CL).
- PC phosphatidylcholines
- PE phosphatidylethanolamines
- TG triacylglycerol
- PG hexosyl
- the complex lipid particle contains less than 12% w/w of chloroplast endogenous to the one or more natural sources. In some embodiments, the complex lipid particle contains less than 20% w/w, less than 15% w/w, less than 10% w/w, less than 5% w/w, less than 1% w/w, less than 0.5% w/w, or less than 0.1% w/w of chloroplast endogenous to the one or more natural sources.
- the complex lipid particle contains less than 5% w/w of exogenous antioxidant.
- the CLPs contain natural lipids comprising about 0 to 20 wt/wt% of cardiolipin (CL).
- a plurality of NMPs in a modified NMP formulation may be loaded with the exogenous peptide, polypeptide, or protein such that at least 5%, at least 10%, at least 15%, at least 25%, at least 50%, at least 75%, at least 90%, or at least 95% of NMPs in the plurality of NMPs encapsulate the exogenous peptide, polypeptide, or protein.
- the NMP is derived from an arthropod, fungi, archaea, or bacteria.
- NMPs can include Arthropod, Plant, Fungi, Archaea, or Bacteria EVs, or segments, portions, or extracts, thereof, in which the EVs are about 5-2000 nm in diameter.
- the NMP can include an EV, or segment, portion, or extract thereof, that has a mean diameter of about 5-50 nm, about 50-100 nm, about 100-150 nm, about 150-200 nm, about 200-250 nm, about 250-300 nm, about 300-350 nm, about 350-400 nm, about 400-450 nm, about 450-500 nm, about 500-550 nm, about 550-600 nm, about 600-650 nm, about 650-700 nm, about 700-750 nm, about 750-800 nm, about 800-850 nm, about 850-900 nm, about 900-950 nm, about 950-1 OOOnm, about 1000-12
- the NMP includes a Arthropod, Plant, Fungi, Archaea, or Bacteria EV, or segment, portion, or extract thereof, that has a mean diameter of about 5-1400 nm, 5-950 nm, about 5-900 nm, about 5-850 nm, about 5- 800 nm, about 5-750 nm, about 5-700 nm, about 5-650 nm, about 5-600 nm, about 5-550 nm, about 5-500 nm, about 5-450 nm, about 5-400 nm, about 5-350 nm, about 5-300 nm, about 5-250 nm, about 5-200 nm, about 5-150 nm, about 5-100 nm, about 5-50 nm, or about 5-25 nm.
- the Arthropod, Plant, Fungi, Archaea, or Bacteria EV, or segment, portion, or extract thereof has a mean diameter of about 50-200 nm. In certain instances, the EV, or segment, portion, or extract thereof, has a mean diameter of about 50-300 nm. In certain instances, the EV, or segment, portion, or extract thereof, has a mean diameter of about 200-500 nm. In certain instances, the EV, or segment, portion, or extract thereof, has a mean diameter of about 30-150 nm.
- the NMP may include a Arthropod, Plant, Fungi, Archaea, or Bacteria EV, or segment, portion, or extract thereof, that has a mean diameter of at least 5 nm, at least 50 nm, at least 100 nm, at least 150 nm, at least 200 nm, at least 250 nm, at least 300 nm, at least 350 nm, at least 400 nm, at least 450 nm, at least 500 nm, at least 550 nm, at least 600 nm, at least 650 nm, at least 700 nm, at least 750 nm, at least 800 nm, at least 850 nm, at least 900 nm, at least 950 nm, at least 1000 nm, or at least 1300.
- the NMP includes a Arthropod, Fungi, Archaea, or Bacteria EV, or segment, portion, or extract thereof, that has a mean diameter less than 1400 nm, less than 1000 nm, less than 950 nm, less than 900 nm, less than 850 nm, less than 800 nm, less than 750 nm, less than 700 nm, less than 650 nm, less than 600 nm, less than 550 nm, less than 500 nm, less than 450 nm, less than 400 nm, less than 350 nm, less than 300 nm, less than 250 nm, less than 200 nm, less than 150 nm, less than 100 nm, or less than 50 nm.
- a variety of methods e.g., a dynamic light scattering method
- a variety of methods can be used to measure the particle diameter of the EVs, or segment, portion, or extract thereof.
- the NMP may include an Arthropod, Plant, Fungi, Archaea, or Bacteria EV, or segment, portion, or extract thereof, that has a mean surface area of 77 nm 2 to 3.2 x10 6 nm 2 (e.g., 77-100 nm 2 , 100-1000 nm 2 , 1000-1x10 4 nm 2 , 1x10 4 - 1x10 5 nm 2 , 1x10 5 -1x10 6 nm 2 , or 1x10 6 - 3.2x10 6 nm 2 ).
- 77-100 nm 2 100-1000 nm 2 , 1000-1x10 4 nm 2 , 1x10 4 - 1x10 5 nm 2 , 1x10 5 -1x10 6 nm 2 , or 1x10 6 - 3.2x10 6 nm 2 ).
- the NMP may include a Arthropod, Fungi, Archaea, or Bacteria EV, or segment, portion, or extract thereof, that has a mean volume of 65 nm 3 to 5.3x10 8 nm 3 (e.g., 65- 100 nm 3 , 100-1000 nm 3 , 1000-1x10 4 nm 3 , 1x10 4 - 1x10 5 nm 3 , 1x10 5 -1x10 e nm 3 , 1x10 e -1x10 7 nm 3 , 1x10 7 -1x10 8 nm 3 , 1x10 8 -5.3x10 8 nm 3 ).
- the NMP may include a Arthropod, Plant, Fungi, Archaea, or Bacteria EV, or segment, portion, or extract thereof, that has a mean surface area of at least 77 nm 2 , (e.g., at least 77 nm 2 , at least 100 nm 2 , at least 1000 nm 2 , at least 1x10 4 nm 2 , at least 1x10 5 nm 2 , at least 1x10 6 nm 2 , or at least 2x10 6 nm 2 ).
- the NMP may include a Arthropod, Fungi, Archaea, or Bacteria EV, or segment, portion, or extract thereof, that has a mean volume of at least 65 nm 3 (e.g., at least 65 nm 3 , at least 100 nm 3 , at least 1000 nm 3 , at least 1x10 4 nm 3 , at least 1x10 5 nm 3 , at least 1x10 6 nm 3 , at least 1x10 7 nm 3 , at least 1x10 8 nm 3 , at least 2x10 8 nm 3 , at least 3x10 8 nm 3 , at least 4x10 8 nm 3 , or at least 5x10 8 nm 3 .
- at least 65 nm 3 e.g., at least 65 nm 3 , at least 100 nm 3 , at least 1000 nm 3 , at least 1x10 4 nm 3 , at least 1x10 5 n
- the NMP can have the same size as the Arthropod, Plant, Fungi, Archaea, or Bacteria EV or segment, extract, or portion thereof.
- the NMP may have a different size than the initial EV from which the NMP is produced.
- the NMP may have a diameter of about 5-2000 nm in diameter.
- the NMP can have a mean diameter of about 5-50 nm, about 50-100 nm, about 100-150 nm, about 150-200 nm, about 200-250 nm, about 250-300 nm, about 300-350 nm, about 350-400 nm, about 400-450 nm, about 450-500 nm, about 500-550 nm, about 550-600 nm, about 600-650 nm, about 650-700 nm, about 700-750 nm, about 750-800 nm, about 800-850 nm, about 850-900 nm, about 900-950 nm, about 950-1 OOOnm, about 1000-1200 nm, about 1200-1400 nm, about 1400-1600 nm, about 1600 - 1800 nm, or about 1800 - 2000 nm.
- the NMP may have a mean diameter of at least 5 nm, at least 50 nm, at least 100 nm, at least 150 nm, at least 200 nm, at least 250 nm, at least 300 nm, at least 350 nm, at least 400 nm, at least 450 nm, at least 500 nm, at least 550 nm, at least 600 nm, at least 650 nm, at least 700 nm, at least 750 nm, at least 800 nm, at least 850 nm, at least 900 nm, at least 950 nm, at least 1000 nm, at least 1200 nm, at least 1400 nm, at least 1600 nm, at least 1800 nm, or about 2000 nm.
- a variety of methods can be used to measure the particle diameter of the NMPs.
- the size of the NMP is determined following loading of heterologous functional agents, or following other modifications to the NMPs.
- the NMP may have a mean surface area of 77 nm 2 to 1 .3 x10 7 nm 2 (e.g., 77-100 nm 2 , 100-1000 nm 2 , 1000-1x10 4 nm 2 , 1x10 4 - 1x10 5 nm 2 , 1x10 5 -1x10 e nm 2 , or 1x10 e -1 .3x10 7 nm 2 ).
- the NMP may have a mean volume of 65 nm 3 to 4.2 x10 9 nm 3 (e.g., 65-100 nm 3 , 100-1000 nm 3 , 1000-1x10 4 nm 3 , 1x10 4 - 1x10 5 nm 3 , 1x10 5 -1x10 6 nm 3 , 1x10 6 -1x10 7 nm 3 , 1x10 7 - 1x10 8 nm 3 , 1x10 8 -1x10 9 nm 3 , or 1x10 9 - 4.2 x10 9 nm 3 ).
- 65-100 nm 3 100-1000 nm 3 , 1000-1x10 4 nm 3 , 1x10 4 - 1x10 5 nm 3 , 1x10 5 -1x10 6 nm 3 , 1x10 6 -1x10 7 nm 3 , 1x10 7 - 1x10 8 nm 3 , 1x10 8 -1
- the NMP has a mean surface area of at least 77 nm 2 , (e.g., at least 77 nm 2 , at least 100 nm 2 , at least 1000 nm 2 , at least 1x10 4 nm 2 , at least 1x10 5 nm 2 , at least 1x10 6 nm 2 , or at least 1x10 7 nm 2 ).
- the NMP has a mean volume of at least 65 nm 3 (e.g., at least 65 nm 3 , at least 100 nm 3 , at least 1000 nm 3 , at least 1x10 4 nm 3 , at least 1x10 5 nm 3 , at least 1x10 6 nm 3 , at least 1x10 7 nm 3 , at least 1x10 8 nm 3 , at least 1x10 9 nm 3 , at least 2x10 9 nm 3 , at least 3x10 9 nm 3 , or at least 4x10 9 nm 3 ).
- at least 65 nm 3 e.g., at least 65 nm 3 , at least 100 nm 3 , at least 1000 nm 3 , at least 1x10 4 nm 3 , at least 1x10 5 nm 3 , at least 1x10 6 nm 3 , at least 1x10 7 nm 3 , at least 1x10 8
- the NMP may include an intact Arthropod, Plant, Fungi, Archaea, or Bacteria EV.
- the NMP may include a non-plant natural source such as algae or animal-derived organs EV, or segment, portion, or extract thereof.
- the NMP may include a segment, portion, or extract of the full surface area of the vesicle (e.g., a segment, portion, or extract including less than 100% (e.g., less than 90%, less than 80%, less than 70%, less than 60%, less than 50%, less than 40%, less than 30%, less than 20%, less than 10%, less than 10%, less than 5%, or less than 1 %) of the full surface area of the vesicle) of a EV.
- the segment, portion, or extract may be any shape, such as a circumferential segment, spherical segment (e.g., hemisphere), curvilinear segment, linear segment, or flat segment.
- the spherical segment may represent one that arises from the splitting of a spherical vesicle along a pair of parallel lines, or one that arises from the splitting of a spherical vesicle along a pair of non-parallel lines.
- the plurality of NMPs can include a plurality of intact EVs, a plurality of EV segments, portions, or extracts, or a mixture of intact and segments of EVs.
- the ratio of intact to segmented EVs will depend on the particular isolation method used.
- grinding or blending a Arthropod, Fungi, Plant, Archaea, or Bacteria, or part thereof, may produce NMPs that contain a higher percentage of EV segments, portions, or extracts than a non-destructive extraction method, such as vacuum-infiltration.
- the NMP includes a segment, portion, or extract of a Arthropod, Fungi, Archaea, or Bacteria EV
- the EV segment, portion, or extract may have a mean surface area less than that of an intact vesicle, e.g., a mean surface area less than 77 nm 2 , 100 nm 2 , 1000 nm 2 , 1x10 4 nm 2 , 1x10 5 nm 2 , 1x10 6 nm 2 , or 3.2x10 6 nm 2 ).
- the EV segment, portion, or extract has a surface area of less than 70 nm 2 , 60 nm 2 , 50 nm 2 , 40 nm 2 , 30 nm 2 , 20 nm 2 , or 10 nm 2 ).
- the NMP may include a Arthropod, Fungi, Archaea, or Bacteria EV, or segment, portion, or extract thereof, that has a mean volume less than that of an intact vesicle, e.g., a mean volume of less than 65 nm 3 , 100 nm 3 , 1000 nm 3 , 1x10 4 nm 3 , 1x10 5 nm 3 , 1x10 6 nm 3 , 1x10 7 nm 3 , 1x10 8 nm 3 , or 5.3x10 8 nm 3 ).
- the NMP includes an extract of a Arthropod, Plant, Fungi, Archaea, or Bacteria EV
- the NMP may include at least 1 %, 2%, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, 99%, or more than 99% of lipids extracted (e.g., with chloroform or with ethanol) from a Arthropod, Fungi, Archaea, or Bacteria EV.
- the NMPs in the plurality may include Arthropod, Plant, Fungi, Archaea, or Bacteria EV segments and/or EV-extracted lipids or a mixture thereof.
- a PMP is a lipid (e.g., lipid bilayer, unilamellar, or multilamellar structure) structure that includes a plant EV, or segment, portion, or extract (e.g., lipid extract) thereof.
- Plant EVs refer to an enclosed lipid-bilayer structure that naturally occurs in a plant and that is about 5-2000 nm in diameter. Plant EVs can originate from a variety of plant biogenesis pathways. In nature, plant EVs can be found in the intracellular and extracellular compartments of plants, such as the plant apoplast, the compartment located outside the plasma membrane and formed by a continuum of cell walls and the extracellular space.
- PMPs can be enriched plant EVs found in cell culture media upon secretion from plant cells. Plant EVs can be separated from plants, thereby providing PMPs, by a variety of methods further described herein. Further, the PMPs can optionally include a therapeutic agent, which can be introduced in vivo or in vitro.
- PMPs can include plant EVs, or segments, portions, or extracts, thereof.
- PMPs can also include exogenous lipids (e.g., sterols (e.g., cholesterol or sitosterol), ionizable lipids, and/or PEGylated lipids) in addition to lipids derived from plant EVs.
- exogenous lipids e.g., sterols (e.g., cholesterol or sitosterol), ionizable lipids, and/or PEGylated lipids
- the plant EVs are about 5-1000 nm in diameter.
- the PMP can include a plant EV, or segment, portion, or extract thereof, that has a mean diameter of about 5-50 nm, about 50-100 nm, about 100-150 nm, about 150-200 nm, about 200-250 nm, about 250-300 nm, about 300-350 nm, about 350-400 nm, about 400-450 nm, about 450-500 nm, about 500-550 nm, about 550-600 nm, about 600-650 nm, about 650-700 nm, about 700-750 nm, about 750-800 nm, about 800-850 nm, about 850-900 nm, about 900-950 nm, about 950-1 OOOnm, about 1000-1250nm, about 1250-1500nm, about 1500-
- the PMP includes a plant EV, or segment, portion, or extract thereof, that has a mean diameter of about 5-950 nm, about 5-900 nm, about 5-850 nm, about 5-800 nm, about 5-750 nm, about 5-700 nm, about 5-650 nm, about 5-600 nm, about 5-550 nm, about 5-500 nm, about 5-450 nm, about 5-400 nm, about 5-350 nm, about 5-300 nm, about 5-250 nm, about 5-200 nm, about 5-150 nm, about 5-100 nm, about 5-50 nm, or about 5-25 nm.
- the plant EV, or segment, portion, or extract thereof has a mean diameter of about 50-200 nm. In certain instances, the plant EV, or segment, portion, or extract thereof, has a mean diameter of about 50-300 nm. In certain instances, the plant EV, or segment, portion, or extract thereof, has a mean diameter of about 200-500 nm. In certain instances, the plant EV, or segment, portion, or extract thereof, has a mean diameter of about 30-150 nm.
- the PMP may include a plant EV, or segment, portion, or extract thereof, that has a mean diameter of at least 5 nm, at least 50 nm, at least 100 nm, at least 150 nm, at least
- the PMP includes a plant EV, or segment, portion, or extract thereof, that has a mean diameter less than 1000 nm, less than 950 nm, less than 900 nm, less than 850 nm, less than 800 nm, less than 750 nm, less than 700 nm, less than 650 nm, less than 600 nm, less than 550 nm, less than 500 nm, less than 450 nm, less than 400 nm, less than 350 nm, less than 300 nm, less than 250 nm, less than 200 nm, less than 150 nm, less than 100 nm, or less than 50 nm.
- a variety of methods e.g., a dynamic light scattering method
- a variety of methods can be used to measure the particle diameter of the plant EV, or segment, portion,
- the PMP may include a plant EV, or segment, portion, or extract thereof, that has a mean surface area of 77 nm 2 to 3.2 x10 6 nm 2 (e.g., 77-100 nm 2 , 100-1000 nm 2 , 1000-1x10 4 nm 2 , 1x10 4 - 1x10 5 nm 2 , 1x10 5 -1x10 6 nm 2 , or 1x10 6 -3.2x10 6 nm 2 ).
- the PMP may include a plant EV, or segment, portion, or extract thereof, that has a mean volume of 65 nm 3 to 5.3x10 8 nm 3 (e.g., 65-100 nm 3 , 100-1000 nm 3 , 1000-1x10 4 nm 3 , 1x10 4 - 1x10 5 nm 3 , 1x10 5 -1x10 6 nm 3 , 1x10 6 -1x10 7 nm 3 , 1x10 7 -1x10 8 nm 3 , 1x10 8 -5.3x10 8 nm 3 ).
- 65-100 nm 3 100-1000 nm 3 , 1000-1x10 4 nm 3 , 1x10 4 - 1x10 5 nm 3 , 1x10 5 -1x10 6 nm 3 , 1x10 6 -1x10 7 nm 3 , 1x10 7 -1x10 8 nm 3 , 1x10 8 -5.3x10 8
- the PMP may include a plant EV, or segment, portion, or extract thereof, that has a mean surface area of at least 77 nm 2 , (e.g., at least 77 nm 2 , at least 100 nm 2 , at least 1000 nm 2 , at least 1x10 4 nm 2 , at least 1x10 5 nm 2 , at least 1x10 6 nm 2 , or at least 2x10 6 nm 2 ).
- the PMP may include a plant EV, or segment, portion, or extract thereof, that has a mean volume of at least 65 nm 3 (e.g., at least 65 nm 3 , at least 100 nm 3 , at least 1000 nm 3 , at least 1x10 4 nm 3 , at least 1x10 5 nm 3 , at least 1x10 6 nm 3 , at least 1x10 7 nm 3 , at least 1x10 8 nm 3 , at least 2x10 8 nm 3 , at least 3x10 8 nm 3 , at least 4x10 8 nm 3 , or at least 5x10 8 nm 3 .
- at least 65 nm 3 e.g., at least 65 nm 3 , at least 100 nm 3 , at least 1000 nm 3 , at least 1x10 4 nm 3 , at least 1x10 5 nm 3 , at least 1x10 6 nm 3 ,
- the PMP can have the same size as the plant EV or segment, extract, or portion thereof.
- the PMP may have a different size than the initial plant EV from which the PMP is produced.
- the PMP may have a diameter of about 5-2000 nm in diameter.
- the PMP can have a mean diameter of about 5-50 nm, about 50-100 nm, about 100-150 nm, about 150-200 nm, about 200-250 nm, about 250-300 nm, about 300-350 nm, about 350-400 nm, about 400-450 nm, about 450-500 nm, about 500-550 nm, about 550-600 nm, about 600-650 nm, about 650-700 nm, about 700-750 nm, about 750-800 nm, about 800-850 nm, about 850-900 nm, about 900-950 nm, about 950-1 OOOnm, about 1000-1200 nm, about 1200-1400 nm, about 1400-1600 nm, about 1600 - 1800 nm, or about 1800 - 2000 nm.
- the PMP may have a mean diameter of at least 5 nm, at least 50 nm, at least 100 nm, at least 150 nm, at least 200 nm, at least 250 nm, at least 300 nm, at least 350 nm, at least 400 nm, at least 450 nm, at least 500 nm, at least 550 nm, at least 600 nm, at least 650 nm, at least 700 nm, at least 750 nm, at least 800 nm, at least 850 nm, at least 900 nm, at least 950 nm, at least 1000 nm, at least 1200 nm, at least 1400 nm, at least 1600 nm, at least 1800 nm, or about 2000 nm.
- a variety of methods can be used to measure the particle diameter of the PMPs.
- the size of the PMP is determined following loading of a therapeutic agent, or following other modifications to the PMPs.
- the PMP may have a mean surface area of 77 nm 2 to 1 .3 x10 7 nm 2 (e.g., 77-100 nm 2 , 100-1000 nm 2 , 1000-1x10 4 nm 2 , 1x10 4 - 1x10 5 nm 2 , 1x10 5 -1x10 6 nm 2 , or 1x10 6 -1 .3x10 7 nm 2 ).
- the PMP may have a mean volume of 65 nm 3 to 4.2 x10 9 nm 3 (e.g., 65-100 nm 3 , 100-1000 nm 3 , 1000-1x10 4 nm 3 , 1x10 4 - 1x10 5 nm 3 , 1x10 5 -1x10 e nm 3 , 1x10 e -1x10 7 nm 3 , 1x10 7 - 1x10 8 nm 3 , 1x10 8 -1x10 9 nm 3 , or 1x10 9 - 4.2 x10 9 nm 3 ).
- 65-100 nm 3 100-1000 nm 3 , 1000-1x10 4 nm 3 , 1x10 4 - 1x10 5 nm 3 , 1x10 5 -1x10 e nm 3 , 1x10 e -1x10 7 nm 3 , 1x10 7 - 1x10 8 nm 3 , 1x
- the PMP has a mean surface area of at least 77 nm 2 , (e.g., at least 77 nm 2 , at least 100 nm 2 , at least 1000 nm 2 , at least 1x10 4 nm 2 , at least 1x10 5 nm 2 , at least 1x10 6 nm 2 , or at least 1x10 7 nm 2 ).
- the PMP has a mean volume of at least 65 nm 3 (e.g., at least 65 nm 3 , at least 100 nm 3 , at least 1000 nm 3 , at least 1x10 4 nm 3 , at least 1x10 5 nm 3 , at least 1x10 6 nm 3 , at least 1x10 7 nm 3 , at least 1x10 8 nm 3 , at least 1x10 9 nm 3 , at least 2x10 9 nm 3 , at least 3x10 9 nm 3 , or at least 4x10 9 nm 3 ).
- at least 65 nm 3 e.g., at least 65 nm 3 , at least 100 nm 3 , at least 1000 nm 3 , at least 1x10 4 nm 3 , at least 1x10 5 nm 3 , at least 1x10 6 nm 3 , at least 1x10 7 nm 3 , at least 1x10 8
- the PMP may include an intact plant EV.
- the PMP may include a segment, portion, or extract of the full surface area of the vesicle (e.g., a segment, portion, or extract including less than 100% (e.g., less than 90%, less than 80%, less than 70%, less than 60%, less than 50%, less than 40%, less than 30%, less than 20%, less than 10%, less than 10%, less than 5%, or less than 1 %) of the full surface area of the vesicle) of a plant EV.
- the segment, portion, or extract may be any shape, such as a circumferential segment, spherical segment (e.g., hemisphere), curvilinear segment, linear segment, or flat segment.
- the spherical segment may represent one that arises from the splitting of a spherical vesicle along a pair of parallel lines, or one that arises from the splitting of a spherical vesicle along a pair of non-parallel lines.
- the plurality of PMPs can include a plurality of intact plant EVs, a plurality of plant EV segments, portions, or extracts, or a mixture of intact and segments of plant EVs.
- a ratio of intact to segmented plant EVs will depend on the particular isolation method used. For example, grinding or blending a plant, or part thereof, may produce PMPs that contain a higher percentage of plant EV segments, portions, or extracts than a non-destructive extraction method, such as vacuum-infiltration.
- the PMP includes a segment, portion, or extract of a plant EV
- the EV segment, portion, or extract may have a mean surface area less than that of an intact vesicle, (e.g., a mean surface area less than 77 nm 2 , 100 nm 2 , 1000 nm 2 , 1x10 4 nm 2 , 1x10 5 nm 2 , 1x10 6 nm 2 , or 3.2x10 6 nm 2 ).
- the EV segment, portion, or extract has a surface area of less than 70 nm 2 , 60 nm 2 , 50 nm 2 , 40 nm 2 , 30 nm 2 , 20 nm 2 , or 10 nm 2 .
- the PMP may include a plant EV, or segment, portion, or extract thereof, that has a mean volume less than that of an intact vesicle, (e.g., a mean volume of less than 65 nm 3 , 100 nm 3 , 1000 nm 3 , 1x10 4 nm 3 , 1x10 5 nm 3 , 1x10 6 nm 3 , 1x10 7 nm 3 , 1x10 8 nm 3 , or 5.3x10 8 nm 3 ).
- a mean volume less than that of an intact vesicle e.g., a mean volume of less than 65 nm 3 , 100 nm 3 , 1000 nm 3 , 1x10 4 nm 3 , 1x10 5 nm 3 , 1x10 6 nm 3 , 1x10 7 nm 3 , 1x10 8 nm 3 , or 5.3x10 8 nm 3 ).
- the PMP includes an extract of a plant EV
- the PMP may include at least 1%, 2%, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, 99%, or more than 99%, of lipids extracted (e.g., with chloroform or ethanol) from a plant EV.
- the PMPs in the plurality may include plant EV segments and/or plant EV-extracted lipids or a mixture thereof.
- NMPs may be produced from Arthropod, Fungi, Archaea, or Bacteria EVs, or a segment, portion or extract (e.g., lipid extract) thereof, that occur naturally in Arthropod, Fungi, Archaea, or Bacteria, or parts thereof, including tissues or cells.
- the NMP may include a non-plant natural source such as algae or animal-derived organs EV, or segment, portion, or extract thereof.
- An exemplary method for producing NMPs includes (a) providing an initial sample from a source, or a part thereof, wherein the source or part thereof comprises EVs; and (b) isolating a crude NMP fraction from the initial sample, wherein the crude NMP fraction has a decreased level of at least one contaminant or undesired component from the source or part thereof relative to the level in the initial sample.
- the method can further include an additional step (c) comprising purifying the crude NMP fraction, thereby producing a plurality of pure NMPs, wherein the plurality of pure NMPs have a decreased level of at least one contaminant or undesired component from the Arthropod, Fungi, Archaea, or Bacteria or part thereof relative to the level in the crude EV fraction.
- an additional step (c) comprising purifying the crude NMP fraction, thereby producing a plurality of pure NMPs, wherein the plurality of pure NMPs have a decreased level of at least one contaminant or undesired component from the Arthropod, Fungi, Archaea, or Bacteria or part thereof relative to the level in the crude EV fraction.
- Each production step is discussed in further detail, below. Exemplary methods regarding the isolation and purification of NMPs is found, for example, in Rutter and Innes, Plant Physiol. 173(1): 728-741 , 2017; Rutter et al, Bio. Proto
- a plurality of NMPs may be isolated from a Arthropod, Fungi, Archaea, or Bacteria by a process which includes the steps of: (a) providing an initial sample from a source, or a part thereof, wherein the source or part thereof comprises EVs; (b) isolating a crude NMP fraction from the initial sample, wherein the crude NMP fraction has a decreased level of at least one contaminant or undesired component from the Arthropod, Fungi, Archaea, or Bacteria or part thereof relative to the level in the initial sample (e.g., a level that is decreased by at least 1%, 2%, 5%, 10%, 15%, 20%, 25%, 30%, 40%, 45%, 50%, 55%, 60%, 70%, 80%, 90%, 95%, 96%, 98%, 99%, or 100%); and (c) purifying the crude NMP fraction, thereby producing a plurality of pure NMPs, wherein the plurality of pure NMPs have
- the NMPs provided herein can include a Arthropod, Fungi, Archaea, or Bacteria EV, or segment, portion, or extract thereof, isolated from a variety of sources.
- NMPs may be isolated from any genera of Arthropod, Fungi, Archaea, or Bacteria , including but not limited to crabs, crawfish, shrimp, spiders, scorpions, crickets, grasshoppers, beetles, millipedes, ticks, mites, centipedes, ants, wasps, dragonflies, flies, gnats, other insects and crustaceans, yeast, mushrooms, puffballs, stinkhorns, boletes, smuts, bunts, bracket fungi, jelly fungi, toadstools, molds, rusts, earth stars, chanterelles, ergot, pyroIobus, picrophilus, methanogens, crenarchaeota, nanoarchaeota,
- NMPs may be produced from a whole Arthropod, Fungi, Archaea, or Bacteria (e.g., a whole insect, spider, crustacean, fungi, or single-cell of archaea or bacteria) or alternatively from one or more source parts (e.g., segments, organs, eggs, spores, mycelium, tissue, membrane or cell wall).
- a whole Arthropod, Fungi, Archaea, or Bacteria e.g., a whole insect, spider, crustacean, fungi, or single-cell of archaea or bacteria
- source parts e.g., segments, organs, eggs, spores, mycelium, tissue, membrane or cell wall.
- NMPs can be produced from organs/structures/tissues/cell cultures (e.g., body segments, appendages, organs, eggs, exoskeleton, embryos, spores, mycelium, hyphae, thallus, suspension cultures, cell walls, inner or outer membranes, gametophytes, sporophytes, polymerases, glycerol-ether lipids, metabolic products, flagella, pili, ribosomes or organelles) or progeny of same.
- the source may be at any stage of development.
- the NMP is produced from an insect or fungi, (e.g. cricket, yeast, or mushroom).
- the NMP is produced from a bacteria or archaea (e.g. E. coli). In some embodiments, the NMP is produced from an algae (e.g. kelp or chlorella). In some embodiments, the NMP is produced from an animal organ (e.g brain or blood).
- a bacteria or archaea e.g. E. coli
- the NMP is produced from an algae (e.g. kelp or chlorella).
- the NMP is produced from an animal organ (e.g brain or blood).
- NMPs can be produced from a Arthropod, Fungi, Archaea, or Bacteria, or part thereof, by a variety of methods. Any method that allows release of the EV-containing fraction of a source, or an otherwise extracellular fraction that contains NMPs comprising secreted EVs (e.g., cell culture media) is suitable in the present methods. EVs can be separated from the source or source part by either destructive (e.g., grinding or blending) or non-destructive (washing or vacuum infiltration) methods.
- the Arthropod, Fungi, Archaea, or Bacteria, or part thereof can be vacuum-infiltrated, ground, blended, or a combination thereof to isolate EVs from the source or source part, thereby producing NMPs.
- the isolating step may involve (b) isolating a crude NMP fraction from the initial sample (e.g., a Arthropod, Fungi, Archaea, or Bacteria or part, or a sample derived from a Arthropod, Fungi, Archaea, or Bacteria or part), wherein the crude NMP fraction has a decreased level of at least one contaminant or undesired component from the source or part thereof relative to the level in the initial sample;, wherein the isolating step involves vacuum infiltrating the Arthropod, Fungi, Archaea, or Bacteria (e.g., with a vesicle isolation buffer) to release and collect the desired fraction.
- the isolating step involves vacuum infiltra
- the NMPs can be separated or collected into a crude NMP fraction.
- the separating step may involve separating the plurality of NMPs into a crude NMP fraction using centrifugation (e.g., differential centrifugation or ultracentrifugation) and/or filtration to separate the NMP-containing fraction from large contaminants, including tissue debris, cells, or cell organelles.
- centrifugation e.g., differential centrifugation or ultracentrifugation
- filtration e.g., differential centrifugation or ultracentrifugation
- the crude NMP fraction will have a decreased number of large contaminants, including, for example, tissue debris, cells, or cell organelles (e.g., nuclei, mitochondria, etc), as compared to the initial sample from the source or source part.
- the crude NMP fraction can be further purified by additional purification methods to produce a plurality of pure NMPs.
- the crude NMP fraction can be separated from other source components by ultracentrifugation, e.g., using a density gradient (iodixanol or sucrose), sizeexclusion, and/or use of other approaches to remove aggregated components (e.g., precipitation or size-exclusion chromatography).
- the resulting pure NMPs may have a decreased level of contaminants or undesired components from the source (e.g., one or more non-NMP components, such as protein aggregates, nucleic acid aggregates, protein-nucleic acid aggregates, free lipoproteins, lipido-proteic structures), nuclei, cell wall components, cell organelles, or a combination thereof) relative to one or more fractions generated during the earlier separation steps, or relative to a pre-established threshold level, e.g., a commercial release specification.
- a pre-established threshold level e.g., a commercial release specification.
- the pure NMPs may have a decreased level (e.g., by about 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or more than 100%; or by about 2x fold, 4x fold, 5x fold, 10x fold, 20x fold, 25x fold, 50x fold, 75x fold, 100x fold, or more than WOx fold) of source organelles or cell wall components relative to the level in the initial sample.
- a decreased level e.g., by about 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or more than 100%; or by about 2x fold, 4x fold, 5x fold, 10x fold, 20x fold, 25x fold, 50x fold, 75x fold, 100x fold, or more than WOx fold
- the pure NMPs are substantially free (e.g., have undetectable levels) of one or more non-NMP components, such as protein aggregates, nucleic acid aggregates, protein-nucleic acid aggregates, free lipoproteins, lipido-proteic structures), nuclei, cell wall components, cell organelles, or a combination thereof.
- non-NMP components such as protein aggregates, nucleic acid aggregates, protein-nucleic acid aggregates, free lipoproteins, lipido-proteic structures
- nuclei cell wall components, cell organelles, or a combination thereof.
- the NMPs may be at a concentration of, e.g., 1x10 9 , 5x10 9 , 1x10 10 , 5x10 10 , 5x10 10 , 1x10 11 , 2x10 11 , 3x10 11 , 4x10 11 , 5x10 11 , 6x10 11 , 7x10 11 , 8x10 11 , 9x10 11 , 1x10 12 , 2x10 12 , 3x10 12 , 4x10 12 , 5x10 12 , 6x10 12 , 7x10 12 , 8x10 12 , 9x10 12 , 1x10 13 , or more than 1x10 13 NMPs/mL.
- protein aggregates may be removed from isolated NMPs.
- the isolated NMP solution can be taken through a range of pHs (e.g., as measured using a pH probe) to precipitate out protein aggregates in solution.
- the pH can be adjusted to, e.g., pH 3, pH 5, pH 7, pH 9, or pH 11 with the addition of, e.g., sodium hydroxide or hydrochloric acid.
- the isolated NMP solution can be flocculated using the addition of charged polymers, such as Polymin-P or Praestol 2640. Briefly, Polymin-P or Praestol 2640 is added to the solution and mixed with an impeller.
- the solution can then be filtered to remove particulates.
- aggregates can be solubilized by increasing salt concentration.
- NaCI can be added to the isolated NMP solution until it is at, e.g., 1 mol/L.
- the solution can then be filtered to isolate the NMPs.
- aggregates are solubilized by increasing the temperature.
- the isolated NMPs can be heated under mixing until the solution has reached a uniform temperature of, e.g., 50°C for 5 minutes. The NMP mixture can then be filtered to isolate the NMPs.
- soluble contaminants from NMP solutions can be separated by size-exclusion chromatography column according to standard procedures, where NMPs elute in the first fractions, whereas proteins and ribonucleoproteins and some lipoproteins are eluted later.
- the efficiency of protein aggregate removal can be determined by measuring and comparing the protein concentration before and after removal of protein aggregates via BCA/Bradford protein quantification.
- protein aggregates are removed before the exogenous peptide, polypeptide, or protein is encapsulated by the NMP.
- protein aggregates are removed after the exogenous peptide, polypeptide, or protein is encapsulated by the NMP.
- the preparation of NMPs from natural sources is through an ethanol extraction method.
- a 3:2 ethyl acetate:ethanol solvent aids in extraction.
- the preparation of NMPs from natural sources is through a modified Matyash extraction method.
- a 1 :2 MeOH:MTBE solvent aids in extraction.
- NMPs may be characterized by a variety of analysis methods to estimate NMP yield, NMP concentration, NMP purity, NMP composition, or NMP sizes.
- NMPs can be evaluated by a number of methods known in the art that enable visualization, quantitation, or qualitative characterization (e.g., identification of the composition) of the NMPs, such as microscopy (e.g., transmission electron microscopy), dynamic light scattering, nanoparticle tracking, spectroscopy (e.g., Fourier transform infrared analysis), or mass spectrometry (protein and lipid analysis).
- the NMPs can additionally be labelled or stained.
- the NMPs can be stained with 3,3’-dihexyloxacarbocyanine iodide (DIOCe), a fluorescent lipophilic dye, PKH67 (Sigma Aldrich); Alexa Fluor® 488 (Thermo Fisher Scientific), or DyLightTM 800 (Thermo Fisher).
- DIOCe 3,3’-dihexyloxacarbocyanine iodide
- PKH67 Sigma Aldrich
- Alexa Fluor® 488 Thermo Fisher Scientific
- DyLightTM 800 Thermo Fisher
- the NMPs can optionally be prepared such that the NMPs are at an increased concentration (e.g., by about 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or more than 100%; or by about 2x fold, 4x fold, 5x fold, 10x fold, 20x fold, 25x fold, 50x fold, 75x fold, 100x fold, or more than 100x fold) relative to the EV level in a control or initial sample.
- concentration e.g., by about 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or more than 100%; or by about 2x fold, 4x fold, 5x fold, 10x fold, 20x fold, 25x fold, 50x fold, 75x fold, 100x fold, or more than 100x fold
- the isolated NMPs may make up about 0.1% to about 100% of the NMP composition, such as any one of about 0.01% to about 100%, about 1% to about 99.9%, about 0.1% to about 10%, about 1% to about 25%, about 10% to about 50%, about 50% to about 99%.
- the composition described herein includes at least any of 0.1%, 0.5%, 1 %, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, or more NMPs, e.g., as measured by wt/vol, percent NMP protein composition, and/or percent lipid composition (e.g., by measuring fluorescently labelled lipids)).
- the concentrated agents are used as commercial products, e.g., the final user may use diluted agents, which have a substantially lower concentration of active ingredient.
- the composition described herein is formulated as a NMP concentrate formulation, e.g., an ultra-low- volume concentrate formulation.
- the NMPs in the composition are at a concentration effective to increase the fitness of an organism, e.g., a plant, an animal, an insect, a bacterium, or a fungus.
- the NMPs in the composition are at a concentration effective to decrease the fitness of an organism, e.g., a plant, an animal, an insect, a bacterium, or a fungus.
- NMPs can be produced from a variety of Arthropod, Fungi, Archaea, or Bacteria, or one or more parts thereof (e.g., segments, organs, eggs, spores, mycelium, tissue, membrane or cell wall).
- NMPs can be produced from organs/structures/tissues/cell cultures (e.g., body segments, appendages, organs, eggs, exoskeleton, embryos, spores, mycelium, hyphae, thallus, suspension cultures, cell walls, inner or outer membranes, gametophytes, sporophytes, polymerases, glycerol-ether lipids, metabolic products, flagella, pili, ribosomes or organelles) or progeny of same.
- the source may be at any stage of development.
- the NMP is produced from an insect or fungi, (e.g. cricket, yeast, or mushroom).
- the NMP is produced from a bacteria or archaea (e.g. E. coli). In some embodiments, the NMP is produced from an algae (e.g. kelp or chlorella). In some embodiments, the NMP is produced from an animal organ (e.g brain or blood).
- a bacteria or archaea e.g. E. coli
- the NMP is produced from an algae (e.g. kelp or chlorella).
- the NMP is produced from an animal organ (e.g brain or blood).
- NMPs can be produced and purified by a variety of methods, for example, by using a density gradient (iodixanol or sucrose) in conjunction with ultracentrifugation and/or methods to remove aggregated contaminants, e.g., precipitation or size-exclusion chromatography.
- a density gradient iodixanol or sucrose
- ultracentrifugation and/or methods to remove aggregated contaminants e.g., precipitation or size-exclusion chromatography.
- the NMPs of the present compositions and methods can be isolated from a Arthropod, Fungi, Archaea, or Bacteria, or part thereof, and used without further modification to the NMP. In other instances, the NMP can be modified prior to use, as outlined further herein.
- PMPs may be produced from plant EVs, or a segment, portion or extract (e.g., lipid extract) thereof, that occur naturally in plants, or parts thereof, including plant tissues or plant cells.
- An exemplary method for producing PMPs includes (a) providing an initial sample from a plant, or a part thereof, wherein the plant or part thereof comprises EVs; and (b) isolating a crude PMP fraction from the initial sample, wherein the crude PMP fraction has a decreased level of at least one contaminant or undesired component from the plant or part thereof relative to the level in the initial sample.
- the method can further include an additional step (c) comprising purifying the crude PMP fraction, thereby producing a plurality of pure PMPs, wherein the plurality of pure PMPs have a decreased level of at least one contaminant or undesired component from the plant or part thereof relative to the level in the crude EV fraction.
- an additional step (c) comprising purifying the crude PMP fraction, thereby producing a plurality of pure PMPs, wherein the plurality of pure PMPs have a decreased level of at least one contaminant or undesired component from the plant or part thereof relative to the level in the crude EV fraction.
- a plurality of PMPs may be isolated from a plant by a process which includes the steps of: (a) providing an initial sample from a plant, or a part thereof, wherein the plant or part thereof comprises EVs; (b) isolating a crude PMP fraction from the initial sample, wherein the crude PMP fraction has a decreased level of at least one contaminant or undesired component from the plant or part thereof relative to the level in the initial sample (e.g., a level that is decreased by at least 1%, 2%, 5%, 10%, 15%, 20%, 25%, 30%, 40%, 45%, 50%, 55%, 60%, 70%, 80%, 90%, 95%, 96%, 98%, 99%, or 100%); and (c) purifying the crude PMP fraction, thereby producing a plurality of pure PMPs, wherein the plurality of pure PMPs have a decreased level of at least one contaminant or undesired component from the plant or part thereof relative to the level in the crude sample
- the PMPs can include a plant EV, or segment, portion, or extract thereof, produced from a variety of plants.
- PMPs may be produced from any genera of plants (vascular or nonvascular), including but not limited to angiosperms (monocotyledonous and dicotyledonous plants), gymnosperms, ferns, selaginellas, horsetails, psilophytes, lycophytes, algae (e.g., unicellular or multicellular, e.g., archaeplastida), or bryophytes.
- PMPs can be produced using a vascular plant, for example monocotyledons or dicotyledons or gymnosperms.
- PMPs can be produced using alfalfa, apple, Arabidopsis, banana, barley, a Brassica species (e.g., Arabidopsis thaliana or Brassica napus), canola, castor bean, chicory, chrysanthemum, clover, cocoa, coffee, cotton, cottonseed, corn, crambe, cranberry, cucumber, dendrobium, dioscorea, eucalyptus, fescue, flax, gladiolus, liliacea, linseed, millet, muskmelon, mustard, oat, oil palm, oilseed rape, papaya, peanut, pineapple, ornamental plants, Phaseolus, potato, rapeseed, rice, rye, ryegrass, safflower, sesame, sorghum, soybean, sugarbeet, sugarcane, sunflower, strawberry, tobacco, tomato, turfgrass, wheat or vegetable crops such as lettuce, celery, broccoli,
- PMPs may be produced using a whole plant (e.g., a whole rosettes or seedlings) or alternatively from one or more plant parts (e.g., leaf, seed, root, fruit, vegetable, pollen, phloem sap, or xylem sap).
- a whole plant e.g., a whole rosettes or seedlings
- plant parts e.g., leaf, seed, root, fruit, vegetable, pollen, phloem sap, or xylem sap.
- PMPs can be produced using shoot vegetative organs/structures (e.g., leaves, stems, or tubers), roots, flowers and floral organs/structures (e.g., pollen, bracts, sepals, petals, stamens, carpels, anthers, or ovules), seed (including embryo, endosperm, or seed coat), fruit (the mature ovary), sap (e.g., phloem or xylem sap), plant tissue (e.g., vascular tissue, ground tissue, tumor tissue, or the like), and cells (e.g., single cells, protoplasts, embryos, callus tissue, guard cells, egg cells, or the like), or progeny of same.
- shoot vegetative organs/structures e.g., leaves, stems, or tubers
- roots e.g., flowers and floral organs/structures (e.g., pollen, bracts, sepals, petals, stamens, carpels, anthers
- the isolation step may involve (a) providing a plant, or a part thereof.
- the plant part is an Arabidopsis leaf.
- the plant may be at any stage of development.
- the PMPs can be produced using seedlings, e.g., 1 week, 2 week, 3 week, 4 week, 5 week, 6 week, 7 week, or 8 week old seedlings (e.g., Arabidopsis seedlings).
- PMPs can include PMPs produced using roots (e.g., ginger roots), fruit juice (e.g., grapefruit juice), vegetables (e.g., broccoli), pollen (e.g., olive pollen), phloem sap (e.g., Arabidopsis phloem sap), or xylem sap (e.g., tomato plant xylem sap).
- roots e.g., ginger roots
- fruit juice e.g., grapefruit juice
- vegetables e.g., broccoli
- pollen e.g., olive pollen
- phloem sap e.g., Arabidopsis phloem sap
- xylem sap e.g., tomato plant xylem sap
- the PMPs are produced from algae or lemon.
- PMPs can be produced using a plant, or part thereof, by a variety of methods. Any method that allows release of the EV-containing apoplastic fraction of a plant, or an otherwise extracellular fraction that contains PMPs comprising secreted EVs (e.g., cell culture media) is suitable in the present methods.
- EVs can be separated from the plant or plant part by either destructive (e.g., grinding or blending of a plant, or any plant part) or non-destructive (washing or vacuum infiltration of a plant or any plant part) methods. For instance, the plant, or part thereof, can be vacuum-infiltrated, ground, blended, or a combination thereof to isolate EVs from the plant or plant part, thereby producing PMPs.
- the isolating step may involve vacuum infiltrating the plant (e.g., with a vesicle isolation buffer) to release and collect the apoplastic fraction.
- the isolating step may involve grinding or blending the plant to release the EVs, thereby producing PMPs.
- the PMPs can be separated or collected into a crude PMP fraction (e.g., an apoplastic fraction).
- the separating step may involve separating the plurality of PMPs into a crude PMP fraction using centrifugation (e.g., differential centrifugation or ultracentrifugation) and/or filtration to separate the plant PMP-containing fraction from large contaminants, including plant tissue debris or plant cells.
- centrifugation e.g., differential centrifugation or ultracentrifugation
- the crude PMP fraction will have a decreased number of large contaminants, including plant tissue debris or plant cells, as compared to the initial sample from the plant or plant part.
- the crude PMP fraction may additionally comprise a decreased level of plant cell organelles (e.g., nuclei, mitochondria or chloroplasts), as compared to the initial sample from the plant or plant part.
- the isolating step may involve separating the plurality of PMPs into a crude PMP fraction using centrifugation (e.g., differential centrifugation or ultracentrifugation) and/or filtration to separate the PMP-containing fraction from plant cells or cellular debris.
- centrifugation e.g., differential centrifugation or ultracentrifugation
- the crude PMP fraction will have a decreased number of plant cells or cellular debris, as compared to the initial sample from the source plant or plant part.
- the crude PMP fraction can be further purified by additional purification methods to produce a plurality of pure PMPs.
- the crude PMP fraction can be separated from other plant components by ultracentrifugation, e.g., using a density gradient (iodixanol or sucrose) and/or use of other approaches to remove aggregated components (e.g., precipitation or size-exclusion chromatography).
- the resulting pure PMPs may have a decreased level of contaminants or other undesired components from the source plant (e.g., one or more non-PMP components, such as protein aggregates, nucleic acid aggregates, protein-nucleic acid aggregates, free lipoproteins, lipido- proteic structures), nuclei, cell wall components, cell organelles, or a combination thereof) relative to one or more fractions generated during the earlier separation steps, or relative to a pre-established threshold level, e.g., a commercial release specification.
- one or more non-PMP components such as protein aggregates, nucleic acid aggregates, protein-nucleic acid aggregates, free lipoproteins, lipido- proteic structures
- nuclei cell wall components
- cell organelles e.g., cell organelles, or a combination thereof
- the pure PMPs may have a decreased level (e.g., by about 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or more than 100%; or by about 2x fold, 4x fold, 5x fold, 10x fold, 20x fold, 25x fold, 50x fold, 75x fold, 10Ox fold, or more than 10Ox fold) of plant organelles or cell wall components relative to the level in the initial sample.
- a decreased level e.g., by about 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or more than 100%; or by about 2x fold, 4x fold, 5x fold, 10x fold, 20x fold, 25x fold, 50x fold, 75x fold, 10Ox fold, or more than 10Ox fold
- the pure PMPs are substantially free (e.g., have undetectable levels) of one or more non-PMP components, such as protein aggregates, nucleic acid aggregates, protein-nucleic acid aggregates, free lipoproteins, lipido-proteic structures), nuclei, cell wall components, cell organelles, or a combination thereof.
- non-PMP components such as protein aggregates, nucleic acid aggregates, protein-nucleic acid aggregates, free lipoproteins, lipido-proteic structures
- nuclei cell wall components, cell organelles, or a combination thereof.
- the PMPs may be at a concentration of, e.g., 1x10 9 , 5x10 9 , 1x10 10 , 5x10 10 , 5x10 10 , 1x10 11 , 2x10 11 , 3x10 11 , 4x10 11 , 5x10 11 , 6x10 11 , 7x10 11 , 8x10 11 , 9x10 11 , 1x10 12 , 2x10 12 , 3x10 12 , 4x10 12 , 5x10 12 , 6x10 12 , 7x10 12 , 8x10 12 , 9x10 12 , 1x10 13 , or more than 1x10 13 PMPs/mL.
- protein aggregates may be removed from PMPs.
- the PMPs can be taken through a range of pHs (e.g., as measured using a pH probe) to precipitate out protein aggregates in solution.
- the pH can be adjusted to, e.g., pH 3, pH 5, pH 7, pH 9, or pH 1 1 with the addition of, e.g., sodium hydroxide or hydrochloric acid.
- the solution Once the solution is at the specified pH, it can be filtered to remove particulates.
- the PMPs can be flocculated using the addition of charged polymers, such as Polymin-P or Praestol 2640. Briefly, Polymin-P or Praestol 2640 is added to the solution and mixed with an impeller.
- the solution can then be filtered to remove particulates.
- aggregates can be solubilized by increasing salt concentration. For example, NaCI can be added to the PMPs until it is at, e.g., 1 mol/L. The solution can then be filtered to isolate the PMPs.
- aggregates are solubilized by increasing the temperature. For example, the PMPs can be heated under mixing until the solution has reached a uniform temperature of, e.g., 50°C for 5 minutes. The PMP mixture can then be filtered to isolate the PMPs.
- soluble contaminants from PMP solutions can be separated by size-exclusion chromatography column according to standard procedures, where PMPs elute in the first fractions, whereas proteins and ribonucleoproteins and some lipoproteins are eluted later.
- the efficiency of protein aggregate removal can be determined by measuring and comparing the protein concentration before and after removal of protein aggregates via BCA/Bradford protein quantification.
- PMPs may be characterized by a variety of analysis methods to estimate PMP yield, PMP concentration, PMP purity, PMP composition, or PMP sizes.
- PMPs can be evaluated by a number of methods known in the art that enable visualization, quantitation, or qualitative characterization (e.g., identification of the composition) of the PMPs, such as microscopy (e.g., transmission electron microscopy), dynamic light scattering, nanoparticle tracking, spectroscopy (e.g., Fourier transform infrared analysis), or mass spectrometry (protein and lipid analysis).
- the PMPs can additionally be labelled or stained.
- the PMPs can be stained with 3,3’-dihexyloxacarbocyanine iodide (DIOCe), a fluorescent lipophilic dye, PKH67 (Sigma Aldrich); Alexa Fluor® 488 (Thermo Fisher Scientific), or DyLightTM 800 (Thermo Fisher).
- DIOCe 3,3’-dihexyloxacarbocyanine iodide
- PKH67 Sigma Aldrich
- Alexa Fluor® 488 Thermo Fisher Scientific
- DyLightTM 800 Thermo Fisher
- the PMPs can optionally be prepared such that the PMPs are at an increased concentration (e.g., by about 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or more than 100%; or by about 2x fold, 4x fold, 5x fold, 10x fold, 20x fold, 25x fold, 50x fold, 75x fold, 100x fold, or more than 100x fold) relative to the EV level in a control or initial sample.
- concentration e.g., by about 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or more than 100%; or by about 2x fold, 4x fold, 5x fold, 10x fold, 20x fold, 25x fold, 50x fold, 75x fold, 100x fold, or more than 100x fold
- the PMPs may make up about 0.1% to about 100% of the PMP composition, such as any one of about 0.01% to about 100%, about 1% to about 99.9%, about 0.1% to about 10%, about 1% to about 25%, about 10% to about 50%, about 50% to about 99%, or about 75% to about 100%.
- the composition described herein includes at least any of 0.1%, 0.5%, 1%, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, or more PMPs, e.g., as measured by wt/vol, percent PMP protein composition, and/or percent lipid composition (e.g., by measuring fluorescently labelled lipids).
- the concentrated agents are used as commercial products, e.g., the final user may use diluted agents, which have a substantially lower concentration of active ingredient.
- the composition described herein is formulated as an agricultural concentrate formulation, e.g., an ultra-low-volume concentrate formulation.
- LNMP refers to a NMP that has been derived from a lipid structure (e.g., a lipid bilayer, unilamellar, multilamellar structure; e.g., a vesicular lipid structure) derived from (e.g., enriched, isolated or purified from) a natural source, wherein the lipid structure is disrupted (e.g., disrupted by lipid extraction) and reassembled or reconstituted in a liquid phase (e.g., a liquid phase containing a cargo) using standard methods, e.g., reconstituted by a method comprising lipid film hydration and/or solvent injection, to produce the LNMP, as is described herein.
- a lipid structure e.g., a lipid bilayer, unilamellar, multilamellar structure; e.g., a vesicular lipid structure
- a liquid phase e.g., a liquid phase containing a
- the method may, if desired, further comprise sonication, freeze/thaw treatment, and/or lipid extrusion, e.g., to reduce the size of the reconstituted LNMPs.
- LNMPs may be produced using a microfluidic device (such as a NanoAssemblr® IGNITETM microfluidic instrument (Precision NanoSystems)).
- the LNMPs are produced by a process which comprises the steps of (a) providing a plurality of purified NMPs (e.g., NMPs purified as described in Section IA herein); (b) processing the plurality of NMPs to produce a lipid film; (c) reconstituting the lipid film in an organic solvent or solvent combination, thereby producing a lipid solution; and (d) processing the lipid solution of step (c) in a microfluidics device comprising an aqueous phase, thereby producing the LNMPs.
- a plurality of purified NMPs e.g., NMPs purified as described in Section IA herein
- processing the plurality of NMPs to produce a lipid film includes extracting lipids from the plurality of NMPs, e.g., extracting lipids using the Bligh-Dyer method (Bligh and Dyer, J Biolchem Physiol, 37: 911-917, 1959).
- the extracted lipids may be provided as a stock solution, e.g., a solution in chloroform:methanol.
- Producing the lipid film may comprise, e.g., evaporation of the solvent with a stream of inert gas (e.g., nitrogen).
- LPMP refers to a PMP that has been derived from a lipid structure (e.g., a lipid bilayer, unilamellar, multilamellar structure; e.g., a vesicular lipid structure) derived from (e.g., enriched, isolated or purified from) a plant source, wherein the lipid structure is disrupted (e.g., disrupted by lipid extraction) and reassembled or reconstituted in a liquid phase (e.g., a liquid phase containing a cargo) using standard methods, e.g., reconstituted by a method comprising lipid film hydration and/or solvent injection, to produce the LPMP, as is described herein.
- a lipid structure e.g., a lipid bilayer, unilamellar, multilamellar structure; e.g., a vesicular lipid structure
- a liquid phase e.g., a liquid phase containing a
- the method may, if desired, further comprise sonication, freeze/thaw treatment, and/or lipid extrusion, e.g., to reduce the size of the reconstituted LPMPs.
- LPMPs may be produced using a microfluidic device (such as a NanoAssemblr® IGNITETM microfluidic instrument (Precision NanoSystems)).
- the LPMPs are produced by a process which comprises the steps of (a) providing a plurality of purified PMPs (e.g., PMPs purified as described in Section IA herein); (b) processing the plurality of PMPs to produce a lipid film; (c) reconstituting the lipid film in an organic solvent or solvent combination, thereby producing a lipid solution; and (d) processing the lipid solution of step (c) in a microfluidics device comprising an aqueous phase, thereby producing the LPMPs.
- a plurality of purified PMPs e.g., PMPs purified as described in Section IA herein
- processing the plurality of PMPs to produce a lipid film
- (c) reconstituting the lipid film in an organic solvent or solvent combination thereby producing a lipid solution
- processing the lipid solution of step (c) in a microfluidics device comprising an aqueous phase, thereby producing the LPMPs
- processing the plurality of PMPs to produce a lipid film includes extracting lipids from the plurality of PMPs, e.g., extracting lipids using the Bligh-Dyer method (Bligh and Dyer, J Biolchem Physiol, 37: 911-917, 1959).
- the extracted lipids may be provided as a stock solution, e.g., a solution in chloroform:methanol.
- Producing the lipid film may comprise, e.g., evaporation of the solvent with a stream of inert gas (e.g., nitrogen).
- a LNMP may comprise between 10% and 100% lipids derived from the lipid structure from the natural source (e.g., lemon or algae), e.g., may contain at least 10%, at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, or 100% lipids derived from the lipid structure from the natural source.
- the natural source e.g., lemon or algae
- a LNMP may comprise all or a fraction of the lipid species present in the lipid structure from the source (e.g., lemon or algae), e.g., it may contain at least 10%, at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, or 100% of the lipid species present in the lipid structure from the source.
- the source e.g., lemon or algae
- a LNMP may comprise none, a fraction, or all of the protein species present in the lipid structure from the source (e.g., lemon or algae), e.g., may contain 0%, less than 1%, less than 5%, less than 10%, less than 15%, less than 20%, less than 30%, less than 40%, less than 50%, less than 60%, less than 70%, less than 80%, less than 90%, less than 100%, or 100% of the protein species present in the lipid structure from the natural source (e.g., lemon or algae).
- the lipid bilayer of the LNMP does not contain proteins.
- the lipid structure of the LNMP contains a reduced amount of proteins relative to the lipid structure from the natural source.
- the natural lipids of the LNMPs are extracted from a plant source, such as lemon or algae.
- the natural lipids are replaced with synthetic structural lipids to form a LNP formulation.
- the LNMPs may be modified to contain a heterologous agent (e.g., a cell-penetrating agent) that is capable of increasing cell uptake (e.g., animal cell uptake (e.g., mammalian cell uptake, e.g., human cell uptake), plant cell uptake, bacterial cell uptake, or fungal cell uptake) relative to an unmodified LNMP.
- a heterologous agent e.g., a cell-penetrating agent
- cell uptake e.g., animal cell uptake (e.g., mammalian cell uptake, e.g., human cell uptake), plant cell uptake, bacterial cell uptake, or fungal cell uptake
- the modified LNMPs may include (e.g., be loaded with, e.g., encapsulate or be conjugated to) or be formulated with (e.g., be suspended or resuspended in a solution comprising) a plant cell-penetrating agent, such as an ionizable lipid.
- a plant cell-penetrating agent such as an ionizable lipid.
- Each of the modified LNMPs may comprise at least 1%, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or more than 90% ionizable lipid.
- LNMPs may include one or more exogenous lipids, e.g., lipids that are exogenous to the plant (e.g., originating from a source that is not the plant or plant part from which the LNMP is produced).
- the lipid composition of the LNMP may include 0%, less than 1%, or at least 1 %, 2%, 5%, 10%, 15%, 20%, 25%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, or more than 95% exogenous lipid.
- the exogenous lipid e.g., ionizable lipid
- the exogenous lipid is added to amount to 25% or 40% (w/w) of total lipids in the preparation.
- the exogenous lipid is added to the preparation prior to step (b), e.g., mixed with extracted NMP lipids prior to step (b).
- Exemplary exogenous lipids include ionizable lipids.
- the ionizable lipids in the LNMP compositions herein include one or more from the compounds of groups i)-iv) as described herein.
- Exogenous lipids may also include cationic lipids.
- the exogenous lipid may also include an ionizable lipid or cationic lipid chosen from 1 ,1 ‘-((2-(4-(2-((2-(bis(2-hydroxydodecyl)amino)ethyl) (2- hydroxydodecyl)amino)ethyl)piperazin-1-yl)ethyl)azanediyl)bis(dodecan-2-ol) (C12-200), DLin-MC3- DMA (MC3), dioleoyl-3-trimethylammonium propane (DODAP), DC-cholesterol, DOTAP, Ethyl PC, GL67, DLin-KC2-DMA (KC2), MD1 (CKK-E12), OF2, EPC, ZA3-Ep10, TT3, LP01 , 5A2-SC8, Lipid 5 (Moderna), a cationic sulfonamide amino lipid, an amphi
- the exogenous lipid may also include an ionizable lipid or cationic lipid chosen from C12-200, MC3, DODAP, DC-cholesterol, DOTAP, Ethyl PC, GL67, KC2, MD1 , OF2, EPC, ZA3-Ep10, TT3, LP01 , 5A2-SC8, Lipid 5 (Moderna), a cationic sulfonamide amino lipid, and an amphiphilic zwitterionic amino lipid or a combination thereof.
- the ionizable lipid is chosen from C12-200, MC3, DODAP, and DC-cholesterol or combinations thereof.
- the ionizable lipid is an ionizable lipid. In some embodiments, the ionizable lipid is 1 ,1 ‘-((2- (4-(2-((2-(bis(2-hydroxydodecyl)amino)ethyl) (2-hydroxydodecyl)amino)ethyl)piperazin-1- yl)ethyl)azanediyl)bis(dodecan-2-ol) (C12-200) or (6Z,9Z,28Z,31Z)-Heptatriaconta-6,9,28,31-tetraen- 19-yl 4-(dimethylamino)butanoate, DLin-MC3-DMA (MC3). In some instances, the exogenous lipid is a cationic lipid. In some embodiments, the cationic lipid is DC-cholesterol or dioleoyl-3- trimethylammonium propane (DOTAP).
- DOTAP di
- the LNMPs comprise at least 1%, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or more than 90% ionizable lipid.
- the LNMPs comprise a molar ratio of least 0.1%, 1%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75% 80%, 85%, 90%, or more than 90% ionizable lipid, e.g., 1 %-10%, 10%-20%, 20%-30%, 30%-40%, 40%-50%, 50%-60%, 60%-70%, 70%-80%, or 80%-90% ionizable lipid, e.g., about 30%-75% ionizable lipid (e.g., about 30%-75% ionizable lipid).
- the LNMP comprises 25% ionizable lipid.
- the LNMP comprises a molar ratio of 35% ionizable lipid. In some embodiments, the LNMP comprises a molar ratio of 50% ionizable lipid. In some embodiments, the LNMP comprises 40% MC3. In some embodiments, the LNMP comprises a molar ratio of 50% ionizable lipid. In some embodiments, the LNMP comprises 20% or 40% DC-cholesterol. In some embodiments, the LNMP comprises 25% or 40% DOTAP.
- the agent may increase uptake of the LNMP as a whole or may increase uptake of a portion or component of the LNMP (e.g., the mRNA therapeutic) carried by the LNMP.
- the degree to which cell uptake is increased may vary depending on the plant or plant part to which the composition described herein is delivered, the LNMP formulation, and other modifications made to the LNMP,
- the modified LNMPs may have an increased cell uptake (e.g., animal cell uptake, plant cell uptake, bacterial cell uptake, or fungal cell uptake) of at least 1%, 2%, 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% relative to an unmodified LNMP.
- the increased cell uptake is an increased cell uptake of at least 2x-fold, 4x-fold, 5x-fold, 10x-fold, 100x- fold, or 1000x-fold relative to an unmodified LNMP.
- a LNMP that has been modified with a ionizable lipid more efficiently encapsulates a negatively charged a polynucleotide than a LNMP that has not been modified with an ionizable lipid.
- a LNMP that has been modified with an ionizable lipid has altered biodistribution relative to a LNMP that has not been modified with an ionizable lipid.
- a LNMP that has been modified with an ionizable lipid has altered (e.g., increased) fusion with an endosomal membrane of a target cell relative to a LNMP that has not been modified with an ionizable lipid.
- the ionizable lipid has at least one (e.g., one, two, three, four or all five) of the characteristics listed below: (i) at least 2 ionizable amines (e.g., at least 2, at least 3, at least 4, at least 5, at least 6, or more than 6 ionizable amines, e.g., 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 , 12, or more than 12 ionizable amines);
- each of the lipid tails is independently at least 6 carbon atoms in length (e.g., at least 6, at least 7, at least 8, at least 9, at least 10, at least 11 , at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18, or more than 18 carbon atoms in length, e.g., 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24, 25, or more than 25 carbon atoms in length);
- an acid dissociation constant of from about 4.5 to about 7.5 (e.g., a pKa of about 4.5, 4.6, 4.7, 4.8, 4.9, 5.0, 5.1 , 5.2, 5.3, 5.4, 5.5, 5.6, 5.7, 5.8, 5.9, 6.0, 6.1 , 6.2, 6.3, 6.4, 6.5, 6.6, 6.7, 6.8, 6.9, 7.0, 7.1 , 7.2, 7.3, 7.4, or 7.5 (e.g., a pKa of from about 6.5 and about 7.5 (e.g., a pKa of about 6.5, 6.6, 6.7, 6.8, 6.9, 7.0, 7.1 , 7.2, 7.3, 7.4, or 7.5));
- the ionizable lipid is an ionizable amine and a heteroorganic group.
- the heteroorganic group is hydroxyl.
- the heteroorganic group comprises a hydrogen bond donor.
- the heteroorganic group comprises a hydrogen bond acceptor.
- the heteroorganic group is -OH, -SH, -(CO)H, - CO2H, -NH2, -CONH2, optionally substituted C1-C6 alkoxy, or fluorine.
- the ionizable lipid is an ionizable amine and a heteroorganic group separated by a chain of at least two atoms
- the ionizable lipid in the LNMP compositions included one of the compounds from group i) to group iv) as discussed below.
- the ionizable lipid is represented by the following formula I: a pharmaceutically acceptable salt thereof, or a stereoisomer of any of the foregoing, wherein each A is independently C1-C16 branched or unbranched alkyl or C1-C16 branched or unbranched alkenyl, optionally substituted with heteroatom or substituted with OH, SH, or halogen; each B is independently C1-C16 branched or unbranched alkyl or C1-C16 branched or unbranched alkenyl, optionally substituted with heteroatom or substituted with OH, SH, or halogen; each X is independently a biodegradable moiety; and
- each R 6 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, or cycloalkyl
- each R 7 and each R 8 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, halogen, OH, SH, NR10R11, wherein each R10 and Rn is independently H, C1-C3 alkyl, or R10 and Rn are taken together to form a heterocyclic ring
- each s is independently 1 , 2, 3, 4, or 5
- each u is independently 1 , 2, 3, 4, or 5
- t is 1 , 2, 3, 4 or 5
- each Z is independently absent, O, S, or NR12, wherein R12 is H, C1-C7 branched or unbranched alkyl, or C2-C7 branched or unbranched alkenyl, provided that
- Q is O, S, or NR13, wherein each R13 is H, C1-C5 alkyl.
- B is C3-C20 alkyl.
- W in formula (I) may alternatively wherein:
- R17 is OH, SH, or N(CH 3 ) 2 ; and each u is independently 1 , 2, 3, 4, or 5.
- W in formula (I) may alternatively be , wherein:
- V is C2-C10 alkenylene, C2-C10 alkynylene, or C2-Cw heteroalkylene; each Re is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, or cycloalkyl; and each u is independently 1 , 2, 3, 4, or 5.
- W in formula (I) may alternatively wherein:
- Ri 4 is a heterocyclic; each v is independently 0, 1 , 2, 3, 4, or 5; and each u is independently 1 , 2, 3, 4, or 5.
- W in formula (I) may alternatively be 0 0 wherein:
- Z is O, S, -C((CH 2 )VN(RI 5 )2)-, or N(RI 5 ), wherein R15 is H, C1-C4 branched or unbranched alkyl, and v is 0, 1 , 2, 3, 4, or 5; each R10 is independently H, or C1-C3 alkyl; and each u is independently 0, 1 , 2, 3, 4, or 5.
- W in formula (I) may alternatively be Q , wherein: each Y is a divalent heterocyclic;
- Q is O, S, or NH; and each u is independently 1 , 2, 3, 4, or 5.
- W in formula (I) may alternatively be
- Ri 4 is a heterocyclic, NR10R11 , C(O)NRi 0 Rn, or C(S)NRi 0 Rn, wherein each R10 and Rn is independently H, C1-C3 alkyl, C3-C7 cycloalkyl, C3-C7 cycloalkenyl, optionally substituted with one or more NH and/or oxo groups, or R10 and Rn are taken together to form a heterocyclic ring;
- W in formula (I) may alternatively be , wherein:
- T is -NHC(O)O-, -OC(O)NH-, or a divalent heterocyclic optionally substituted with one or more -(CH2) V OH, -(CH2) V SH, and/or -(CH2) v -halogen groups; each R 7 and each R 8 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, halogen, OH, SH, (CH2) v Ri7, or NR10R11, wherein each R10 and Rn is independently H, C1-C3 alkyl, or R10 and Rn are taken together to form a heterocyclic ring;
- R17 is OH, SH, or N(CH 3 ) 2 ; each v is independently 0, 1 , 2, 3, 4, or 5; and each u is independently 1 , 2, 3, 4, or 5.
- W in formula (I) may alternatively be , wherein:
- T is -NHC(O)O-, -OC(O)NH-, or a divalent heterocyclic; and each u is independently 1 , 2, 3, 4, or 5.
- the adjacent R1 and R 2 when Z is not absent, cannot be OH, NR10R11, or SH.
- the heterocyclic is a piperazine, piperazine dione, piperazine-2,5- dione, piperidine, pyrrolidine, piperidinol, dioxopiperazine, bis-piperazine, aromatic or heteroaromatic.
- the ionizable lipid is represented by formula (IX): pharmaceutically acceptable salts thereof, and stereoisomers of any of the foregoing, wherein each R1 and each R 2 is independently H, C1-C3 branched or unbranched alkyl, OH, halogen, SH, or NR10R11 , or each R1 and each R 2 are independently taken together with the carbon atom(s) to which they are attached to form a cyclic ring; each R10 and Rn is independently H, C1-C3 branched or unbranched alkyl, or R10 and Rn are taken together to form a heterocyclic ring; each R 3 and each R 4 is independently H, C2-C14 branched or unbranched alkyl (e.g., C3-C10 branched or unbranched alkyl), or C3-C10 branched or unbranched alkenyl, provided that at least one of R 3 and R 4 is not H; each X
- V is a branched or unbrachned C2-C3 alkylene. In some embodiments, V is a C2-C3 alkylene substituted with OH. In some embodiments, V is a branched or unbrachned C2-C3 alkenylene. In some embodiments, each R 6 is independently H or methyl.
- the ionizable lipid is represented by one of the following formulas for the variables are the same as those in formula (X).
- the disclosure relates to ionizable lipids of Formula (XI): and stereoisomers of any of the foregoing, wherein each Ri and each R 2 is independently H, C1-C3 branched or unbranched alkyl, OH, halogen, SH, or NR10R11 , or each R1 and each R 2 are independently taken together with the carbon atom(s) to which they are attached to form a cyclic ring; each R10 and Rn is independently H, C1-C3 branched or unbranched alkyl, or R10 and Rn are taken together to form a heterocyclic ring; each R 3 and each R 4 is independently H, C2-C14 branched or unbranched alkyl (e.g., C3-C10 branched or unbranched alkyl), or C3-C10 branched or unbranched alkenyl, provided that at least one of R 3 and R 4 is not H; each X is independently a biofluorate, a biol
- T is -NHC(O)O-, -OC(O)NH-, or a divalent heterocyclic optionally substituted with one or more -(CH2) V OH, -(CH2) V SH, -(CH2) v -halogen groups
- each R 7 and each R 8 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, halogen, OH, SH, (CH2) v Ri7, or NR10R11, wherein R17 is OH, SH, or N(CH 3 ) 2 ; each v is independently 0, 1 , 2, 3, 4, or 5; and each m is independently 1 , 2, 3, 4, 5, 6, 7, 8, 9 or 10.
- T is a divalent heterocyclic (e.g., a divalent piperazine, or a divalent dioxopiperazine) optionally substituted with -(CH2) V OH, wherein v is independently 0, 1 , or 2.
- a divalent heterocyclic e.g., a divalent piperazine, or a divalent dioxopiperazine
- v is independently 0, 1 , or 2.
- X is -OC(O)-, -C(O)O-, -SS-, -N(R 18 )C(O)-, -C(O)N(R 18 )-, -C(O-RI 3 )-O-, -C(O)O(CH 2 ) a -, -OC(O)(CH 2 ) a -, -C(O)N(R 18 )(CH 2 ) a -, -N(R 18 )C(O)(CH2) a -, -C(O-Ri 3 )-O-(CH2) a -, wherein each R 18 is independently H, alkyl, alkenyl, cycloalkyl, hydroxyalkyl, or aminoalkyl, each RI 3 is independently C3-C10 alkyl, and each a is independently 0-16. In one embodiment, each X is independently -OC(O)-, -C(O)O-, -
- ionizable lipid of formula (I), in the Ionizable lipid compounds group i) may be found in PCT Application No. PCT/US22/50725, filed on November 22, 2022, the content of which is incorporated herein by reference in its entirety.
- all the ionizable lipids of formulas (l)-(XII) of PCT Application No. PCT/US22/50725 are suitable for use as the ionizable lipids in this disclosure, and are incorporated herein by reference in its entirety.
- the ionizable lipid is represented by the following formula II: a pharmaceutically acceptable salt thereof, or a stereoisomer of any of the foregoing, wherein: cyclic or heterocyclic moiety; Y is alkyl, hydroxy, hydroxyalkyl or ;
- A is absent, -O-, -N(R 7 )-, -O-alkylene-, -alkylene-O-, -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, -N(R 7 )C(O)N(R 7 )-, -S-, -S-S-, or a bivalent heterocycle; each of X and Z is independently absent, -O-, -CO-, -N(R 7 )-, -O-alkylene-; -alkylene-O-, -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, or -S-; each R 7 is independently H, alkyl, alkenyl, cycloalkyl, hydroxy, hydroxyalkyl, or
- W is hydroxyl, substituted or unsubstituted hydroxyalkyl, substituted or unsubstituted amino, substituted or unsubstituted aminocarbonyl, or substituted or unsubstituted heterocylyl or heteroaryl.
- Y is hydroxyl or _ b ? A ⁇ w f Cycle )
- the ionizable lipid is represented by formula: All the variables in this formula have been defined and exemplified as those described in the above embodiments.
- the ionizable lipid is represented by formula: wherein: each ml is independently an integer from 3 to 6, each 11 is independently an integer from 4 to 8, m2 and I2 are each independently an integer from 0 to 3,
- Rso and R90 are each independently unsubstituted Cs-Cs alkyl; or Rso is H or unsubstituted Ci- 04 alkyl, and R90 is unsubstituted C5-C11 alkyl; and
- Rno and R120 are each independently unsubstituted Cs-Cs alkyl; or Rno is H or unsubstituted C1-C4 alkyl, and R120 is unsubstituted C5-C11 alkyl. All the other variables in these formulas have been defined and exemplified as those described in the above embodiments. In some embodiments, in these formulas, Rso is H or unsubstituted C1-C2 alkyl, and R90 is unsubstituted Ce-C alkyl; and Rno and R120 are each independently unsubstituted Cs-Cs alkyl. In some embodiments, Rso, R90, Rno, and R120 are each independently unsubstituted Cs-Cs alkyl.
- A is absent, -O-, -N(R 7 )-, N(R 7 )C(O)-,
- R 6 is independently H, alkyl, hydroxyl, hydroxyalkyl, amino, aminoalkyl, thiol, thiolalkyl, or N + (R 7 )s-alkylene-Q-; and R 7 is H or C1-C3 alkyl.
- t1 is 0, 1 , 2, 3 or 4; and t is 0, 1 , or 2.
- W is hydroxyl, hydroxyalkyl, or one of the following:
- each Q is independently absent, -O-, -C(O)-, -C(S)-, -C(O)O-, -C(R 7 )2-, -C(O)N(R 7 )-, -C(S)N(R 7 )-, or -N(R 7 )-;
- each R 6 is independently H, alkyl, hydroxyl, hydroxyalkyl, alkoxy, amino, aminoalkyl, alkylamino, thiol, thiolalkyl, or N + (R 7 )3-alkylene-Q-;
- each R 8 is independently H, alkyl, hydroxyalkyl, amino, aminoalkyl, thiol, or thiolalkyl, or two R 8 together with the nitrogen atom may form a ring;
- R70 is H; and each of Rso and R90 is independently H or C1-C12 branched or unbranched alkyl;
- R100 is H; and each of Rno and R120 is independently H or C1-C12 branched or unbranched alkyl, provided that at least one of Rso and R90 is not H, and at least one of Rno and R120 is not H;
- I is from 3 to 7; and m is from 1 to 5.
- ionizable lipid of formula (II), in the Ionizable lipid compounds group ii) may be found in PCT Application No. PCT/US23/16300, filed on March 24, 2023, the content of which is incorporated herein by reference in its entirety.
- all the ionizable lipids of formulas (I), (IA-1), (IA-2), (IIA)-(IIC), (IIA-1), (IIIA)-(IIIIE), (IIIC-1), (IVA-1)-(IVA-3), (IVC-1)-(IVC-2), (VA-1)-(VA-9), (VC-1)-(VC-6) of PCT Application No. PCT/US23/16300 are suitable for use as the ionizable lipids in this disclosure, and are incorporated herein by reference in its entirety.
- Table II Exemplary ionizable lipid compounds.
- the ionizable lipid is represented by formula pharmaceutically acceptable salts thereof, and stereoisomers of any of the foregoing, wherein:
- R20 and R30 are each independently H, C1-C5 branched or unbranched alkyl, or C2-C5 branched or unbranched alkenyl, or
- R20 and R30 together with the adjacent N atom form a 3 to 7 membered cyclic ring, optionally substituted with R a ;
- R a is H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, halogen, OH, or SH; each R1 and each R 2 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, OH, halogen, SH, or NR10R11, or
- R1 and R 2 are taken together to form a cyclic ring; each R10 and Rn is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, or
- R10 and Rn are taken together to form a heterocyclic ring; n is 0, 1 , 2, 3 or 4;
- Y is O or S;
- Z is absent, O, S, or N(RI 2 )(RI 2 ), wherein each RI 2 is independently H, C1-C7 branched or unbranched alkyl, or C2-C7 branched or unbranched alkenyl, provided that when Z is not absent, the adjacent R1 and R 2 cannot be OH, NR10R11, or SH;
- each A is each independently C1-C16 branched or unbranched alkyl, or C2-C16 branched or unbranched alkenyl, optionally interrupted with one or more heteroatoms or optionally substituted with OH, SH, or halogen;
- each B is each independently C1-C16 branched or unbranched alkyl, or C2-Ci6 branched or unbranched alkenyl, optionally interrupted with one or more heteroatoms or optionally substituted with OH, SH, or halogen;
- each X is independently a biodegradable moiety.
- R 20 and R30 are each independently H or C1-C3 branched or unbranched alkyl. In some embodiments, R 20 and R30 together with the adjacent N atom form a 3 to 7 membered cyclic ring, optionally substituted with R a . In some embodiments, R a is H, C1-C3 branched or unbranched alkyl or OH. In one embodiment, R a is H or OH.
- Z is absent, S, O, or NH.
- n is 0, 1 , or 2.
- the ionizable lipid is represented by formula (V): (V), pharmaceutically acceptable salts thereof, and stereoisomers of any of the foregoing, wherein
- R1 is H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, OH, halogen, SH, or NR10R11, and
- R 2 is H, OH, halogen, SH, or NR10R11, or
- R1 and R 2 are taken together to form a cyclic ring
- R10 and Rn are each independently H or C1-C3 alkyl, or R10 and Rn are taken together to form a heterocyclic ring;
- Q is OH or -(OCH 2 CH2)uNR 2 oR 3 o
- R 20 and R 3 o are each independently H, C1-C5 branched or unbranched alkyl, or C2-C5 branched or unbranched alkenyl, or
- R 20 and R30 together with the adjacent N atom form a 3 to 7 membered cyclic ring optionally substituted with R a ;
- the disclosure relates to ionizable lipids of one of the following formulas: , wherein: u is 0, 1 , 2, 3, 4, 5, 6, 7, or 8; v is 0, 1 , 2, 3, or 4; and y is 0, 1 , 2, 3, or 4.
- Other variables are defined as in formulas III) and V) above.
- X is -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, -C(O-RI 3 )-O-, -C(O)O(CH 2 ) S -, -OC(O)(CH 2 ) S -, -C(O)N(R 7 )(CH 2 ) S -, -N(R 7 )C(O)(CH 2 ) S -, -C(O-RI 3 )-O-(CH 2 ) S -, wherein each R 7 is independently H, alkyl, alkenyl, cycloalkyl, hydroxyalkyl, or aminoalkyl, each R13 is independently C 3 -Cw alkyl, and each s is independently 0-16.
- X is -OC(O)-, -C(O)O-, -C(O)O(CH 2 ) S -, or -OC(O)(CH 2 ) S -.
- s is 0, 1 , 2, 3, 4, 5, 6, 7, 8, 9, or 10.
- ionizable lipid of formula (III) or (V), in the Ionizable lipid compounds group iii) may be found in PCT Application No. PCT/US22/50111 , filed on November 16, 2022, the content of which is incorporated herein by reference in its entirety.
- PCT/US22/50111 are suitable for use as the ionizable lipids in this disclosure, and are incorporated herein by reference in its entirety.
- Table III Exemplary ionizable lipid compounds.
- the ionizable lipid is a lipid comprising at least one head group and at least one tail group of formula (Tl) or (TH): pharmaceutically acceptable salt thereof, or a stereoisomer of any of the foregoing, wherein:
- E is each independently a biodegradable group
- R a is each independently C1-C5 alkyl, C2-C5 alkenyl, or C2-C5 alkynyl; u1 and u2 are each independently 0, 1 , 2, 3, 4, 5, 6, or 7;
- R* is each independently H, C1-C16 branched or unbranched alkyl or C1-C16 branched or unbranched alkenyl, optionally interrupted with heteroatom or substituted with OH, SH, or halogen, or cycloalkyl or substituted cycloalkyl; represents the bond connecting the tail group to the head group; and wherein the lipid has a pKa from about 4 to about 8.
- E is each independently -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, -C(O-RI 3 )-O-, -C(O)O(CH 2 )r, -C(O)N(R 7 ) (CH 2 )r, -S-S-, or -C(O-Ri 3 )-O-(CH 2 )r, wherein each R 7 is independently H, alkyl, alkenyl, cycloalkyl, hydroxyalkyl, or aminoalkyl; R13 is branched or unbranched C3-C10 alkyl; and r is 1 , 2, 3, 4, or 5.
- E is each independently -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, or -C(0)N(R 7 )-, wherein R 7 is independently H, alkyl, alkenyl, cycloalkyl, hydroxyalkyl, or aminoalkyl.
- the lipid comprises at least one head group and at least one tail group of formula (TH): wherein u3 and u4 are each independently 0, 1 , 2, 3, or 4.
- TH tail group of formula
- the lipid comprises at least one head group and at least one tail group of formula (Till):
- the lipid comprises at least one head group and at least one tail group
- the lipid comprises at least one head group and at least one tail group independently H or methyl; and R b is in each occasion independently H or C1-C4 alkyl.
- the definitions of other variables in (TV) are the same as those defined above in (Tl).
- the lipid comprises at least one head group and at least one tail group is in each occasion independently H or C1-C4 alkyl.
- the definitions of other variables in (TH’) are the same as those defined above in (Tl’).
- the lipid comprises at least one head group and at least one tail group each independently H or methyl; and R b is in each occasion independently H or C1-C4 alkyl.
- R b is in each occasion independently H or C1-C4 alkyl.
- the lipid comprises at least one tail group of the formulas (TH), (Till), (TIV), (TV), (TH’), and (Till’), wherein
- R 7 is each independently H or methyl
- R b is in each occasion independently H or C1-C4 alkyl; u3 and u4 are each independently 0, 1 , 2, 3, 4, 5, 6, or 7; and wherein the lipid has a pKa from about 4 to about 8.
- the lipid comprises tw, three, four, 0 or more tail groups that have a formula of (T), (Tl), (Til), (Till), (TIV), (TV), (TH’), and/or (Till’), and each tail group may be the same or different.
- R a is each independently C1-C5 branched or unbranched alkyl, C2-C5 branched or unbranched alkenyl, or C2-C5 branched or unbranched alkynyl. In some embodiments, R a is each independently C1-C3 branched or unbranched alkyl. In one embodiment, each R a is methyl.
- u1 is 3, 4, or 5.
- u2 is 0, 1 , 2, or 3.
- u3 and u4 are each independently 1-7, for instance, u3 and u4 are each independently 1 , 2, 3, or 4.
- the lipid comprises at least one tail of formula (Till), wherein each R a is methyl; R b is in each occasion independently H, ethyl, or butyl; u1 is 3-5, u2 is 0-3, and u3 is 1-7 (e.g., 1-4).
- the lipid comprises at least one two tails of formula (Till), wherein the two tails of formula (Till) are the same or different.
- the lipid comprises at least three tails of formula (Till), wherein each tail may be the same or different.
- the lipid has four tails of formula (Till), wherein each tail may be the same or different.
- each R a is methyl, and u1 is 3, u2 is 2, and u3 is 4.
- the lipid comprises at least one tail of formula (Til), wherein each R a is methyl, u1 is 3-5, u2 is 0-3, u3 is 1-4, and u4 is 1-4.
- the lipid has at least two tails of formula (Til), wherein the two tails of formula (Til) are the same.
- the lipid has at least two tails of formula (Til), wherein the two tails of formula (Til) are or different.
- the lipid comprises at least three tails of formula (Til), wherein each tail may be the same or different.
- the lipid has four tails of formula (Til), wherein each tail may be the same or different.
- the lipid comprises at least one tail of formula (TIV), wherein each R a is methyl, u1 is 3-5, u2 is 0-3, u3 is 1-4, and u4 is 1-4.
- the lipid comprises at least two tails of formula (TIV), wherein each tail may be the same or different.
- the lipid comprises at least three tails of formula (TIV), wherein each tail may be the same or different.
- the lipid comprises at least four tails of formula (TIV), wherein each tail may be the same or different.
- the lipid comprises at least two tails of formula (TV), wherein each tail may be the same or different. In some embodiments, the lipid comprises at least three tails of formula (TV), wherein each tail may be the same or different. In some embodiments, the lipid comprises at least four tails of formula (TV), wherein each tail may be the same or different.
- the lipid has at least two tails of formula (TH’), wherein each tail may be the same or different. In some embodiments, the lipid has at least three tails of formula (TH’), wherein each tail may be the same or different. In some embodiments, the lipid has at least four tails of formula (TH’), wherein each tail may be the same or different.
- the lipid has at least two tails of formula (Till’), wherein each tail may be the same or different. In some embodiments, the lipid has at least three tails of formula (Till’), wherein each tail may be the same or different. In some embodiments, the lipid has at least four tails of formula (Till’), wherein each tail may be the same or different.
- the lipid has at least one tail of formula (Til) and/or at least one tail of formula (Till); the lipid further comprises at least one tail that does not have a formula (T), (Tl), (Til), (Till), (TIV), (TV), (TH’) , and/or (Till’). That is to say, the lipid further comprises at least one tail that does not contain a gem-di functional groups bonded to the same carbon next to E (e.g., -C(O)O-).
- the lipid further comprises at least one tail that does not have a formula (T), (Tl), (TH), (Till), (TIV), (TV), (Tl ), (TIT), and/or (Till’). That is to say, the lipid further comprises at least one tail that does not contain a gem-di functional groups bonded to the same carbon next to E.
- the lipid further comprises at least one tail that does not have a formula (T), (Tl), (TH), (Till), (TIV), (TV), (Tl ), (TIT), and/or (Till’). That is to say, the lipid further comprises at least one tail that does not contain a gem-di functional groups bonded to the same carbon next to E.
- the lipid further comprises at least one tail of formula (TNG-I): wherein
- E is each independently a biodegradable group as described herein (e.g., -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -S-S-, or -C(O)N(R 7 )-); u1 and u2 are each independently 0, 1 , 2, 3, 4, 5, 6, or 7; and
- R 7 is independently H, alkyl, alkenyl, cycloalkyl, hydroxyalkyl, or aminoalkyl.
- TNG-I the at least one tail of formula (TNG-I) can be represented by
- R b is in each occasion independently H or C1-C4 alkyl.
- the lipid further comprises at least two tails that do not have a formula (T), (Tl), (TH), (Till), (TIV), (TV), (Tl ), (TH’), and/or (Till’).
- the lipid comprises two tail groups of formula (TNG-II) or (TNG-III), and wherein each tail group may be the same or different,
- the lipid further comprises at least three tails that do not have a formula (T), (Tl), (TH), (Till), (TIV), (TV), (Tl ), (TH’), and/or (Till’).
- the lipid comprises three tail groups of formula (TNG-II) or (TNG-III), and wherein each tail group may be the same or different,
- the head group of the lipid has a structure of formula (HA-I): wherein:
- R20 and R30 are each independently H, C1-C5 branched or unbranched alkyl, or C2-C5 branched or unbranched alkenyl, optionally interrupted with one or more heteroatoms or substituted with OH, SH, halogen, or cycloalkyl groups; or
- R20 and R30 together with the adjacent N atom, form a 3 to 7 membered heterocylic or heteroaromatic ring containing one or more heteroatoms, optionally substituted with one or more OH, SH, halogen, alkyl, or cycloalkyl groups; each of R1 and R2 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, OH, halogen, SH, or NR10R11 ; or R1 and R2 together form a cyclic ring; each of Rw and Rn is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl; or Rw and Rn together form a heterocyclic ring; n is 0, 1 , 2, 3 or 4;
- Z is absent, O, S, or NR12, wherein R12 is H or C1-C7 branched or unbranched alkyl; provided that when Z is not absent, the adjacent R1 and R2 cannot be OH, NR10R11, SH; and represents the bond connecting the head group to the tail group.
- R20 and R30 together with the adjacent N atom form a 3 to 7 membered heterocylic or heteroaromatic ring containing one or more heteroatoms, optionally substituted with one or more OH, SH, halogen, alkyl, or cycloalkyl groups.
- the head group of the ionizable lipid has a structure of formula (HA-I): wherein: each of R1 and R2 is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl, OH, halogen, SH, or NR10R11 ; or R1 and R2 are taken together to form a cyclic ring; each of Rw and Rn is independently H, C1-C3 branched or unbranched alkyl, C2-C3 branched or unbranched alkenyl; or Rw and Rn are taken together to form a heterocyclic ring; m is 1 , 2, 3, 4, 5, 6, 7 or 8; n is 0, 1 , 2, 3 or 4;
- Z is absent, O, S, or NR12, wherein R12 is H or C1-C7 branched or unbranched alkyl; provided that when Z is not absent, the adjacent R1 and R2 cannot be OH, NR10R11, SH; and represents the bond connecting the head group to the tail group.
- the head group of the ionizable lipid has a structure of formula (HA- wherein Z is absent, O, S, or NR12; and R12 is H or C1-C7 branched or unbranched alkyl.
- Z is absent, O, S, or NR12; and R12 is H or C1-C7 branched or unbranched alkyl.
- R12 is H or C1-C7 branched or unbranched alkyl.
- the head group has a structure of: , wherein:
- Rc is H or alkyl, optionally substituted with OH; and ml is 1 , 2, or 3.
- the head group of the ionizable lipid has a structure of formula (HA-V): wherein:
- Ri is H, C1-C3 alkyl, OH, halogen, SH, or NR10R11;
- R2 is OH, halogen, SH, or NR10R11; or R1 and R2 can be taken together to form a cyclic ring;
- R10 and R11 are each independently H or Ci-Cs alkyl; or Rio and Rn can be taken together to form a heterocyclic ring;
- R20 and R30 are each independently H, C1-C5 branched or unbranched alkyl, C2-C5 branched or unbranched alkenyl; or R20 and R30 can be taken together to form a cyclic ring; and each of v and y is independently 1 , 2, 3, or 4.
- the head group of the ionizable lipid has a structure of formula (HA-
- each R20 and R30 are independently C1-C3 alkyl. In one embodiment, each R20 and R30 are independently methyl.
- the head group of the ionizable lipid has a structure of formula (HB-I): wherein
- each R20 is independently H, or C1-C3 branched or unbranched alkyl;
- Ru is a heterocyclic, NR10R11, C(O)NRwRn, NRwC(O)NRwRn , or NRwC(S)NRwRn, wherein each Rw and Rn is independently H, C1-C3 alkyl, C3-C7 cycloalkyl, C3-C7 cycloalkenyl, optionally substituted with one or more NH and/or oxo groups, or Rw and Rn are taken together to form a heterocyclic ring;
- Q is O, S, CH2, or NR13, wherein each R13 is H, C1-C5 alkyl;
- V is branched or unbrachned C2-C10 alkylene, C2-C10 alkenylene, C2-Cw alkynylene, or C2-Cw heteroalkylene, optionally substituted with one or more OH, SH, and/or halogen groups; and T is -NHC(O)O-, -OC(O)NH-, or a divalent heterocyclic.
- each Re, R7, and Rs are independently H or methyl; and each of u and t is independently 1 , 2, or 3. [0317] In some embodiments, in formula wherein:
- Ru is a nitrogen-containing 5- or 6- membered heterocyclic, NRwRn, C(O)NRwRn, NRwC(0)NRioRii , or NRioC(S)NRioRn, wherein each R and Ru is independently H or C1-C3 alkyl; and each of u and v is independently 1 , 2, or 3.
- W is , wherein: each Rs is independently H or methyl; each u is independently 1 , 2, or 3; and
- V is C2-C6 alkylene or C2-C6 alkenylene.
- each Re is independently H or methyl
- each Ry is independently H
- each Rs is methyl
- each u is independently 1 , 2, or 3;
- V is C2-C6 alkylene or C2-C6 alkenylene.
- T is a divalent nitrogen-containing 5- or 6- membered heterocyclic.
- each u is independently 1 , 2, or 3;
- Q is O; each Z is independently NR12; and
- R12 is H or C1-C3 alkyl.
- the head group has the structure of: independently 1 or 2.
- the head group of the ionizable lipid has a structure of formula (HC-I):
- A is absent, -O-, -N R 7 )-, -O-alkylene-, -alkylene-O-, -OC(O)-, -C(O)O-, -N(R 7 )C(O)-, -C(O)N(R 7 )-, -N(R 7 )C(O)N(R 7 )-, -S-, or -S-S-; each of X and Z is independently absent, -O-, -C(O)-, -N(R 7 )-, -O-alkylene-, -alkylene-O-,
- each R 7 is independently H, alkyl, alkenyl, cycloalkyl, hydroxy, alkoxy, hydroxyalkyl, alkylamino, alkylaminoalkyl, or aminoalkyl; t is 0, 1 , 2, or 3; t1 is an integer from 0 to 10; and
- W is hydroxyl, substituted or unsubstituted hydroxyalkyl, substituted or unsubstituted amino, substituted or unsubstituted aminocarbonyl, or substituted or unsubstituted heterocyclyl or heteroaryl.
- the head group has a structure of formula
- Z is -O-, -C
- the head group has a structure of formula wherein t1 is 0, 1 , 2, or 3.
- W is hydroxyl, substituted or unsubstituted hydroxyalkyl, or one of the following moieties:
- each Q is independently absent, -O-, -C(O)-, -C(S)-, -C(O)O-, -(CH2) q C(R 7 )2-, -C(O)N(R 7 )-, -C(S)N(R 7 )-, or -N R 7 );
- R 6 is independently H, alkyl, hydroxyl, hydroxyalkyl, alkoxy, -O-alkylene-O-alkyl, -O-alkylene-N(R 7 )2, amino, alkylamino, aminoalkyl, thiol, thiolalkyl, or N + (R 7 )3-alkylene-Q-; each R 8 is independently H, alkyl, hydroxyalkyl, amino, aminoalkyl, alkylamino, thiol, or thiolalkyl, heterocyclyl, heteroaryl, or two R 8 together with the nitrogen atom may form a ring, optionally substituted with one or more alkyl, hydroxy, hydroxyalkyl, alkoxy, alkylaminoalkyl, alkylamino, aminoalkyl; q is 0, 1 , 2, 3, 4, or 5; and p is 0, 1 , 2, 3, 4, or 5.
- W is one of the following:
- Ri is each independently H, C1-C3 alkyl, OH, halogen, SH, or NR10R11; R1 and R2 can be taken together to form a cyclic ring; R and Rn are each independently H, C1-C3 alkyl, and Rio and R11 can be taken together to form a heterocyclic ring;
- R2 is each independently H, C1-C3 alkyl, OH, halogen, SH, or NR10R11; R1 and R2 can be taken together to form a cyclic ring; Rw and Rn are each independently H, C1-C3 alkyl, and Rw and R11 can be taken together to form a heterocyclic ring; m is 1 , 2, 3, 4, 5, 6, 7 or 8; n is 0, 1 , 2, 3 or 4; r is each independently 0, 1 , 2, 3, 4, 5, 6, 7 or 8;
- R3 is each independently H, or C3-C10 alkyl
- R4 is each independently H, or C3-C10 alkyl; provided that at least one of R3 and R4 is not H; Z is absent, O, S, or NRI 2 ; wherein RI 2 is C1-C7 alkyl;
- X’ is a biodegradable moiety.
- X is N-(2-aminoethyl)-2-aminoethyl-N-(2-aminoethyl)-2-aminoethyl-N-(2-aminoethyl)-2-aminoethyl-N-(2-aminoethyl)-2-aminoethyl-N-(2-aminoethyl)-2-aminoethyl)-2-aminoethyl-N-(2-aminoethyl)-2-aminoethyl-N-(2-aminoethyl)-2-aminoethyl-N-(2-aminoethyl)-2-aminoethyl-N-(2-aminoethyl)-2-aminoethyl-N-(2-aminoethyl)-2-aminoethyl-N-(2-aminoethyl)-2-aminoethyl
- Z is S. In some embodiments, Z is O. In some embodiments, Z is NH.
- r is 3. In some embodiments, r is 4.
- More embodiments of the above ionizable lipids comprising at least one head group (e.g., head group of formulas (HA-I) to (HA-VII), (HB-I), and (HC-I) to (HC-IIIE)), and at least one tail group of formula (Tl) or (T1 ’) (e.g., tail group of formula (Til), (Till), TIV, TV, TH’, or Till’), in the Ionizable lipid compounds group iv), may be found in PCT Application No. PCT/US23/31669, filed on August 31 , 2023, the content of which is incorporated herein by reference in their entirety.
- a lipid membrane of the LNMPs comprises at least 35% of the lipid compound from group i), e.g., at least 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75% 80%, 85%, 90%, or more than 90% of the lipid compound from group i), e.g., 35%-40%, 40%-50%, 50%- 60%, 60%-70%, 70%-80%, or 80%-90% of the lipid compound from group i).
- a lipid membrane of the LNMPs comprises at least 35% of the lipid compound from group ii), e.g., at least 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75% 80%, 85%, 90%, or more than 90% of the lipid compound from group ii), e.g., 35%-40%, 40%-50%, 50%- 60%, 60%-70%, 70%-80%, or 80%-90% of the lipid compound from group ii).
- a lipid membrane of the LNMPs comprises at least 35% of the lipid compound from group iii), e.g., at least 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75% 80%, 85%, 90%, or more than 90% of the lipid compound from group iii), e.g., 35%-40%, 40%-50%, 50%- 60%, 60%-70%, 70%-80%, or 80%-90% of the lipid compound from group iii).
- a lipid membrane of the LNMPs comprises at least 35% of the lipid compound from group iii), e.g., at least 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75% 80%, 85%, 90%, or more than 90% of the lipid compound from group iii), e.g., 35%-40%, 40%-50%, 50%- 60%, 60%-70%, 70%-80%, or 80%-90% of the lipid compound from group iv).
- the LNMPs comprise at least 1 %, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or more than 90% ionizable lipid.
- the LNMPs comprise a molar ratio of at least 0.1 %, 1 %, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75% 80%, 85%, 90%, or more than 90% ionizable lipid, e.g., 1 %-10%, 10%-20%, 20%-30%, 30%-40%, 40%-50%, 50%-60%, 60%-70%, 70%-80%, or 80%-90% ionizable lipid, e.g., about 25%-75% ionizable lipid (e.g., about 25%-75% ionizable lipid).
- more than one ionizable lipid can be used for the ionizable lipid component: one or more of the ionizable lipids from the compounds of formulas in groups i)-iv) can be used alone or in combination with a different ionizable lipid from the compounds of formulas in groups i)-iv).
- the ionizable lipid is not selected from 1 ‘-((2-(4-(2-((2-(bis(2- hydroxydodecyl)amino)ethyl) (2-hydroxydodecyl)amino)ethyl)piperazin-1- yl)ethyl)azanediyl)bis(dodecan-2-ol) (C12-200), MD1 (CKK-E12), OF2, EPC, ZA3-Ep10, TT3, LP01 , 5A2-SC8, Lipid 5 (Moderna), and 98N12-5.
- the additional ionizable lipid is selected from the group consisting of 1 ,1 ’-((2-(4-(2-((2-(bis(2-hydroxydodecyl)amino)ethyl) (2-hydroxydodecyl)amino)ethyl)piperazin-1- yl)ethyl)azanediyl)bis(dodecan-2-ol) (C12-200), MD1 (CKK-E12), OF2, EPC, ZA3-Ep10, TT3, LP01 , 5A2-SC8, Lipid 5, SM-102 (Lipid H), and ALC-315.
- the additional ionizable lipid is represented by the following formula III: (lll), wherein R is Cs-C alkyl group.
- the ionizable lipid described herein may include an amine core described herein substituted with one or more (e.g., 1 , 2, 3, 4, 5, or 6) lipid tails.
- the ionizable lipid described herein include at least 3 lipid tails.
- a lipid tail may be a Cs-C hydrocarbon (e.g., Ce-Cis alkyl or CB-CIS alkanoyl).
- An amine core may be substituted with one or more lipid tails at a nitrogen atom (e.g., one hydrogen atom attached to the nitrogen atom may be replaced with a lipid tail).
- the amine core has a structure of:
- the amine core has a structure of:
- the amine core has a structure of:
- the amine core has a structure of:
- the amine core has a structure of:
- the amine core has a structure of:
- the amine core has a structure of:
- the amine core has a structure of:
- RNA compositions and methods for making and using thereof include the lipids as described in International Patent Publication WO 2016/118725, which is incorporated herein by reference in its entirety.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- RNA compositions and methods for making and using thereof include the lipids as described in International Patent Publication WO 2016/118724, which is incorporated herein by reference in its entirety.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- Suitable lipids for use in the RNA composition and methods for making and using thereof include a lipid having the formula of 14,25-ditridecyl 15,18,21 ,24-tetraaza-octatriacontane, and pharmaceutically acceptable salts thereof.
- Suitable lipids for use in the RNA composition and methods for making and using thereof include the lipids as described in International Patent Publications WO 2013/063468 and WO 2016/205691 , each of which is incorporated herein by reference in its entirety.
- the RNA composition and methods for making and using thereof include a lipid of the following formula: r pharmaceutically acceptable salts thereof, wherein each instance of R L is independently optionally substituted C6-C40 alkenyl.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of:
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of:
- RNA composition and methods for making and using thereof include the lipids as described in International Patent Publication WO 2015/184256, which is incorporated herein by reference in its entirety.
- the RNA composition and methods for making and using thereof include a lipid of the following formula: pharmaceutically acceptable salt thereof, wherein each X independently is O or S; each Y independently is O or S; each m independently is 0 to 20; each n independently is 1 to 6; each RA is independently hydrogen, optionally substituted C1-50 alkyl, optionally substituted C2-50 alkenyl, optionally substituted C2-50 alkynyl, optionally substituted C3-10 carbocyclyl, optionally substituted 3-14 membered heterocyclyl, optionally substituted C6-14 aryl, optionally substituted 5-14 membered heteroaryl or halogen; and each RB is independently hydrogen, optionally substituted C1-50 alkyl, optionally substituted C2-50 alkenyl, optionally
- the RNA composition and methods for making and using thereof include a lipid, “Target 23”, having a compound structure of: , (Target 23) and pharmaceutically acceptable salts thereof.
- RNA composition and methods for making and using thereof include the lipids as described in International Patent Publication WO 2016/004202, which is incorporated herein by reference in its entirety.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salt thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salt thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: salt thereof.
- Other suitable lipids for use in the RNA composition and methods for making and using thereof include lipids as described in United States Provisional Patent Application Serial Number 62/758,179, which is incorporated herein by reference in its entirety.
- the RNA composition and methods for making and using thereof include a lipid of the following formula: , or a pharmaceutically acceptable salt thereof, wherein each R 1 and R 2 is independently H or C1-C6 aliphatic; each m is independently an integer having a value of 1 to 4; each A is independently a covalent bond or arylene; each L 1 is independently an ester, thioester, disulfide, or anhydride group; each L 2 is independently C2-C10 aliphatic; each X 1 is independently H or OH; and each R 3 is independently C6-C20 aliphatic.
- each R 1 and R 2 is independently H or C1-C6 aliphatic
- each m is independently an integer having a value of 1 to 4
- each A is independently a covalent bond or arylene
- each L 1 is independently an ester, thioester, disulfide, or anhydride group
- each L 2 is independently C2-C10 aliphatic
- each X 1 is independently
- the RNA composition and methods for making and using thereof include a lipid of the following formula: (Compound 1), or a pharmaceutically acceptable salt thereof.
- the RNA composition and methods for making and using thereof include a lipid off the ffollowing fformula: pharmaceutically acceptable salt thereof.
- the RNA composition and methods for making and using thereof include a lipid of the following formula: pharmaceutically acceptable salt thereof.
- RNA compositions and methods for making and using thereof include the lipids as described in J. McClellan, M. C. King, Cell 2010, 141 , 210-217 and in Whitehead et al., Nature Communications (2014) 5:4277, which is incorporated herein by reference in its entirety.
- the lipids of the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- RNA compositions and methods for making and using thereof include the lipids as described in International Patent Publication WO 2015/199952, which is incorporated herein by reference in its entirety.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure:
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: and pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: , and pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- RNA compositions and methods for making and using thereof include the lipids as described in International Patent Publication WO 2017/004143, which is incorporated herein by reference in its entirety.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: thereof.
- the RNA composition and methods for making and using thereof salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: thereof.
- the RNA composition and methods for making and using thereof acceptable salts thereof are provided.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: o. .0.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: thereof.
- the RNA composition and methods for making and using thereof salts thereof.
- RNA compositions and methods for making and using thereof include the lipids as described in International Patent Publication WO 2017/075531 , which is incorporated herein by reference in its entirety.
- G 1 and G 2 are each independently unsubstituted Ci- 012 alkylene or Ci-Ci2 alkenylene;
- G 3 is C1-C24 alkylene, C1-C24 alkenylene, C3-C8 cycloalkylene, C3- Cs cycloalkenylene;
- R a is H or C1-C12 alkyl;
- R 1 and R 2 are each independently C6-C24 alkyl or Ce- C 2 4 alkenyl;
- R 4 is C1-C12 alkyl;
- R 5 is H or
- Ci-C 6 alkyl and x is 0, 1 or 2.
- RNA composition and methods for making and using thereof include the lipids as described in International Patent Publication WO 2017/117528, which is incorporated herein by reference in its entirety.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof acceptable salts thereof are provided.
- the RNA composition and methods for making and using thereof include a lipid having the compound structure: pharmaceutically acceptable salts thereof.
- RNA compositions and methods for making and using thereof include the lipids as described in International Patent Publication WO 2017/049245, which is incorporated herein by reference in its entirety.
- the lipids of the RNA composition and methods for making and using thereof include a compound of one of the following formulas: pharmaceutically acceptable salts thereof.
- R4 is independently selected from -(CH2)nQ and -(CH2)nCHQR;
- Q is selected from the group consisting of -OR, -OH, -O(CH2)nN(R)2, -OC(O)R, -CX3, -CN, -N(R)C(O)R, -N(H)C(O)R, -N(R)S(O) 2 R, -N(H)S(O) 2 R, -N(R)C(O)N(R) 2 , -N(H)C(O)N(R) 2 , -N(H)C(O)N(R) 2 , -N(H)C(O)N(H)(R), -N(R)C(S)N(R)2, -N(H)C(S)N(R) 2I -N(H)C(S)N(H)(R), and a heterocycle; and n is 1 , 2, or 3.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- RNA composition and methods for making and using thereof include the lipids as described in International Patent Publication WO 2017/173054 and WO 2015/095340, each of which is incorporated herein by reference in its entirety.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: pharmaceutically acceptable salts thereof.
- the RNA composition and methods for making and using thereof include a lipid having a compound structure of: described in, may be formulated as described in, or may comprise or be comprised by a composition as described in WO2016118724, WO2016118725, WO2016187531 , WO2017176974, WO2018078053, WO2019027999, WO2019036030, WO2019089828, WO2019099501 , W02020072605, W02020081938, W02020118041 , W02020146805, or W02020219876, each of which is incorporated by reference herein in its entirety.
- the ionizable lipids disclosed herein may be used to form the LNMP composition together with one or more natural lipids disclosed herein.
- the LNMP composition is formulated to further comprise one or more therapeutic agents.
- the LNMP composition is a lipid nanoparticle that encapsulates or is associated with the one or more therapeutic agents.
- the therapeutic agents are one or more polynucleotides encoding one or more components of a gene editing system or one or more gene editing systems.
- the RNA composition disclosed herein has an N/P ratio of at least 3, for instance, an N/P ratio of 3 to 100, 3 to 50, 3 to 30, 3 to 20, 3 to 15, 3 to 12, about 3 to about 10, , 6 to 30, 6 to 20, 6 to 15, or 6 to 12.
- the N/P ratio may be about 6 ⁇ 1 , or the N/P ratio may be about 6 ⁇ 0.5.
- the N/P ratio is about 6. .
- the N/P ratio is about 3 (e.g., 3 ⁇ 1 or 3 ⁇ 0.5).
- the RNA composition has an N/P ratio of about 12 to about 17, for example the N/P ratio is about 15 ⁇ 1 , or the N/P ratio is about 15 ⁇ 0.5. In some embodiments, the N/P ratio is about 15. In some embodiments, the N/P ratio is about 12 (e.g., 12 ⁇ 1 or 12 ⁇ 0.5).
- the disclosure relates to a composition
- a composition comprising (i) one or more compounds chosen from the ionizable lipids of Formula (l)-(lll), pharmaceutically acceptable salts thereof, and stereoisomers of any of the foregoing and (ii) a lipid component.
- the composition comprises 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, or 95% of the one or more compounds.
- the disclosure relates to a composition
- a composition comprising (i) one or more lipid nanoparticles and (ii) one or more lipid components.
- one or more lipid components comprise one or more helper lipids and one or more PEG lipids.
- the lipid component(s) comprise(s) one or more helper lipids, one or more PEG lipids, and one or more neutral lipids.
- Non-limiting examples of neutral lipids include phospholipids such as lecithin, phosphatidylethanolamine, lysolecithin, lysophosphatidylethanolamine, phosphatidylserine, phosphatidylinositol, sphingomyelin, egg sphingomyelin (ESM), cephalin, cardiolipin, phosphatidic acid, cerebrosides, dicetylphosphate, distearoylphosphatidylcholine (DSPC), dioleoylphosphatidylcholine (DOPC), dipalmitoylphosphatidylcholine (DPPC), dioleoylphosphatidylglycerol (DOPG), dipalmitoylphosphatidylglycerol (DPPG), dioleoylphosphatidylethanolamine (DOPE), palmitoyloleoyl-phosphatidylcholine (POPC), palmitoyl
- acyl groups in these lipids may be acyl groups derived from fatty acids having C10-C24 carbon chains, e.g. , lauroyl, myristoyl, palmitoyl, stearoyl, or oleoyl.
- the RNA composition comprises a phytosterol or a combination of a phytosterol and cholesterol.
- the phytosterol is selected from the group consisting of b-sitosterol, stigmasterol, b-sitostanol, campesterol, brassicasterol, and combinations thereof.
- the phytosterol is selected from the group consisting of b-sitosterol, b- sitostanol, campesterol, brassicasterol, Compound S-140, Compound S-151 , Compound S-156, Compound S-157, Compound S-159, Compound S-160, Compound S-164, Compound S-165, Compound S-170, Compound S-173, Compound S-175 and combinations thereof.
- the phytosterol is selected from the group consisting of Compound S-140, Compound S-151 , Compound S-156, Compound S-157, Compound S-159, Compound S-160, Compound S-164, Compound S-165, Compound S-170, Compound S-173, Compound S-175, and combinations thereof.
- the phytosterol is a combination of Compound S-141 , Compound S-140, Compound S-143 and Compound S-148.
- the phytosterol comprises a sitosterol or a salt or an ester thereof.
- the phytosterol comprises a stigmasterol or a salt or an ester thereof.
- the exogenous lipid may be a cell-penetrating agent, may be capable of increasing delivery of a polypeptide by the LNMP to a cell, and/or may be capable of increasing loading (e.g., loading efficiency or loading capacity) of a polypeptide.
- Further exemplary exogenous lipids include sterols and PEGylated lipids.
- the LNMPs can be modified with other components (e.g., lipids, e.g., sterols, e.g., cholesterol; or small molecules) to further alter the functional and structural characteristics of the LNMP.
- the LNMPs can be further modified with stabilizing molecules that increase the stability of the LNMPs (e.g., for at least one day at room temperature, and/or stable for at least one week at 4°C).
- the LNMP is modified with a sterol, e.g., sitosterol, sitostanol, B- sitosterol, 7a-hydroxycholesterol, pregnenolone, cholesterol (e.g., ovine cholesterol or cholesterol isolated from plants), stigmasterol, campesterol, fucosterol, or an analog (e.g., a glycoside, ester, or peptide) of any sterol.
- a sterol e.g., sitosterol, sitostanol, B- sitosterol, 7a-hydroxycholesterol, pregnenolone, cholesterol (e.g., ovine cholesterol or cholesterol isolated from plants), stigmasterol, campesterol, fucosterol, or an analog (e.g., a glycoside, ester, or peptide) of any sterol.
- the exogenous sterol is added to the preparation prior to step (b), e.g., mixed with extracted NMP lipids prior to step (b).
- the exogenous sterol may be added to amount to, e.g., 1%, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or more than 90% (w/w) of total lipids and sterols in the preparation.
- the sterol is cholesterol or sitosterol.
- the LNMPs comprise a molar ratio of least 1%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, or more than 60% sterol (e.g., cholesterol or sitosterol), e.g., 1 %-10%, 10%-20%, 20%-30%, 30%-40%, 40%-50%, or 50%-60% sterol.
- the LNMP comprises a molar ratio of about 35%-50% sterol (e.g., cholesterol or sitosterol), e.g., about 36%, 38.5%, 42.5%, or 46.5% sterol.
- the LNMP comprises a molar ratio of about 20%-40% sterol.
- a LNMP that has been modified with a sterol has altered stability (e.g., increased stability) relative to a LNMP that has not been modified with a sterol.
- a LNMP that has been modified with a sterol has a greater rate of fusion with a membrane of a target cell relative to a LNMP that has not been modified with a sterol.
- the LNMPs comprise an exogenous lipid and an exogenous sterol.
- the LNMP is modified with a PEGylated lipid.
- Polyethylene glycol (PEG) length can vary from 1 kDa to 10kDa; in some aspects, PEG having a length of 2kDa is used.
- the PEGylated lipid is C14-PEG2k, C18-PEG2k, or DMPE-PEG2k.
- the LNMPs comprise a molar ratio of at least 0.1%, 0.2%, 0.3%, 0.4%, 0.5%, 0.6%, 0.7%, 0.8%, 0.9%, 1%, 1.1%, 1.2%, 1.3%, 1.4%, 1.5%, 1.6%, 1.7%, 1.8%, 1.9%, 2%, 2.1%, 2.2%, 2.3%, 2.4%, 2.5%, 2.6%, 2.7%, 2.8%, 2.9%, 3%, 3.5%, 4%, 4.5%, 5%, 10%, 20%, 30%, 40%, 50%, or more than 50% PEGylated lipid (e.g., C14-PEG2k, C18-PEG2k, or DMPE-PEG2k), e.g., 0.1%-0.5%, 0.5%- 1%, 1%-1 .5%, 1.5%-2.5%, 2.5%-3.5%, 3.5%-5%, 5%-10%, 10%-20%, 20%-30%, 30%-40%, or 30%- 50% PEGylated lipid.
- PEGylated lipid e.g
- the LNMP comprises a molar ratio of about about 0.1 %- 10% PEGylated lipid (e.g., C14-PEG2k, C18-PEG2k, or DMPE-PEG2k), e.g., about 1%-3% PEGylated lipid, e.g., about 1.5% or about 2.5% PEGylated lipid.
- a LNMP that has been modified with a PEGylated lipid has altered stability (e.g., increased stability) relative to a LNMP that has not been modified with a PEGylated lipid.
- a LNMP that has been modified with a PEGylated lipid has altered particle size relative to a LNMP that has not been modified with a PEGylated lipid.
- a LNMP that has been modified with a PEGylated lipid is less likely to be phagocytosed than a LNMP that has not been modified with a PEGylated lipid.
- the addition of PEGylated lipids can also affect stability in Gl tract and enhance particle migration through mucus.
- PEG may be used as a method to attach targeting moieties.
- the LNMPs are modified with an ionizable lipid and one or both of a sterol (e.g., cholesterol or sitosterol) and a PEGylated lipid (e.g., C14-PEG2k, C18-PEG2k, or DMPE- PEG2k).
- a sterol e.g., cholesterol or sitosterol
- a PEGylated lipid e.g., C14-PEG2k, C18-PEG2k, or DMPE- PEG2k.
- the LNMPs comprise a molar ratio of about 5%-50% natural lipids (e.g., about 10%-20% natural lipids, e.g., about 10%, 12.5%, 16%, or 20% natural lipids); about 30%- 75% ionizable lipids (e.g., about 35% or about 50% ionizable lipids); about 35%-50% sterol (e.g., about 36%, 38.5%, 42.5%, or 46.5% sterol); and about 0.1 %-10% PEGylated lipid (e.g., about 1 %-3% PEGylated lipid, e.g., about 1.5% or about 2.5% PEGylated lipid).
- natural lipids e.g., about 10%-20% natural lipids, e.g., about 10%, 12.5%, 16%, or 20% natural lipids
- about 30%- 75% ionizable lipids e.g., about 35% or about 50% i
- the modified LNMPs comprise a molar ratio of about 5%-60% natural lipids (e.g., about 10%-20%, 20%-30%, 30%-40%, 40%-50%, or 50%-60% natural lipids, e.g., about 10%, 12.5%, 16%, 20%, 30%, 40%, 50%, or 60% natural lipids); about 25%-75% ionizable lipids (e.g., about 35% or about 50% ionizable lipids); about 10%-50% sterol (e.g., about 10%, 12.5%, 14%, 16%, 18%, 20%, 36%, 38.5%, 42.5%, or 46.5% sterol); and about 0.1 %-10% PEGylated lipid (e.g., about 0.5%-5% PEGylated lipid, e.g., about 1 %-3% PEGylated lipid, or about 1 .5% or about 2.5% PEGylated lipid).
- 5%-60% natural lipids e.g.
- the ionizable lipids, natural lipids, sterol, and PEGylated lipid comprise about 25%-75%, about 20%-60%, about 10%-45%, and about 0.5%-5%, respectively, of the lipids in the modified NMP.
- the ionizable lipids, natural lipids, sterol, and PEGylated lipid comprise about 30%-75%, about 20%-50%, about 10%-45%, and about 1 %-5%, respectively, of the lipids in the modified NMP.
- the ionizable lipids, natural lipids, sterol, and PEGylated lipid comprise about 35%-75%, about 20%-50%, about 10%-45%, and about 1 %-5%, respectively, of the lipids in the modified NMP.
- the ionizable lipids, natural lipids, sterol, and PEGylated lipid are formulated at a molar ratio of about 35:50:12.5:2.5.
- the ionizable lipids, natural lipids, sterol, and PEGylated lipid are formulated at a molar ratio of about 35:50:1 1 .5:3.5.
- the ionizable lipids, natural lipids, sterol, and PEGylated lipid are formulated at a molar ratio of about 35:20:42.5:2.5.
- a LNMP that has been modified with a cationic lipid and a sterol and/or a PEGylated lipid more efficiently encapsulates a negatively charged cargo (e.g., a nucleic acid) than a LNMP that has not been modified with a cationic lipid and a sterol and/or a PEGylated lipid.
- a negatively charged cargo e.g., a nucleic acid
- the modified LNMP may have an encapsulation efficiency for the cargo (e.g., nucleic acid, e.g., RNA or DNA) that is at least 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 95%, 98%, 99%, or more than 99%, e.g., may have an encapsulation efficiency of 5%-30%, 30%-50%, 50%-70%, 70%-80%, 80%-90%, 90%-95%, or 95%-100%.
- the cargo e.g., nucleic acid, e.g., RNA or DNA
- Cell uptake of the modified LNMPs can be measured by a variety of methods known in the art.
- the LNMPs, or a component thereof can be labelled with a marker (e.g., a fluorescent marker) that can be detected in isolated cells to confirm uptake.
- a marker e.g., a fluorescent marker
- a LNMP formulation provided herein comprises two or more different modified LNMPs, e.g., comprises modified LNMPs derived from different unmodified LNMPs (e.g., unmodified LNMPs from two or more different natural sources) and/or comprises modified LNMPs comprising different species and/or different ratios of ionizable lipids, sterols, and/or PEGylated lipids.
- the organic solvent in which the lipid film is dissolved is dimethylformamide:methanol (DMF:MeOH).
- the organic solvent or solvent combination may be, e.g., acetonitrile, acetone, ethanol, methanol, dimethylformamide, tetrahydrofuran, 1- buthanol, dimethyl sulfoxide, acetonitrile:ethanol, acetonitrile:methanol, acetone:methanol, methyl tert-butyl etherpropanol, tetrahydrofuran:methanol, dimethyl sulfoxide:methanol, or dimethylformamide:methanol.
- the aqueous phase may be any suitable solution, e.g., a citrate buffer (e.g., a citrate buffer having a pH of about 3.2), water, or phosphate-buffered saline (PBS).
- the aqueous phase may further comprise a nucleic acid (e.g., an siRNA or siRNA precursor (e.g., dsRNA), miRNA or miRNA precursor, mRNA, or plasmid (pDNA)) or a small molecule.
- a nucleic acid e.g., an siRNA or siRNA precursor (e.g., dsRNA), miRNA or miRNA precursor, mRNA, or plasmid (pDNA)
- the lipid solution and the aqueous phase may be mixed in the microfluidics device at any suitable ratio.
- aqueous phase and the lipid solution are mixed at a 3:1 volumetric ratio.
- LNMPs may optionally include additional agents, e.g., cell-penetrating agents, therapeutic agents, polynucleotides, polypeptides, or small molecules.
- the LNMPs can carry or associate with additional agents in a variety of ways to enable delivery of the agent to a target plant, e.g., by encapsulating the agent, incorporation of the agent in the lipid bilayer structure, or association of the agent (e.g., by conjugation) with the surface of the lipid bilayer structure.
- Nucleic acid molecules can be incorporated into the LNMPs either in vivo (e.g., in planta) or in vitro (e.g., in tissue culture, in cell culture, or synthetically incorporated).
- the LNMPs comprising an ionizable lipid, and optionally a cationic lipid may have, e.g., a zeta potential of greater than -30 mV when in the absence of cargo, greater than -20 mV, greater than -5mV, greater than 0 mV, or about 30 mvwhen in the absence of cargo.
- the LNMP has a negative zeta potential, e.g., a zeta potential of less than 0 mV, less than -10 mV, less than -20 mV, less than -30 mV, less than -40 mV, or less than -50 mV when in the absence of cargo.
- the LNMP has a positive zeta potential, e.g., a zeta potential of greater than 0 mV, greater than 10 mV, greater than 20 mV, greater than 30 mV, greater than 40 mV, or greater than 50 mV when in the absence of cargo.
- the LNMP has a zeta potential of about 0.
- the zeta potential of the LNMP may be measured using any method known in the art. Zeta potentials are generally measured indirectly, e.g., calculated using theoretical models from the data obtained using methods and techniques known in the art, e.g., electrophoretic mobility or dynamic electrophoretic mobility. Electrophoretic mobility is typically measured using microelectrophoresis, electrophoretic light scattering, or tunable resistive pulse sensing. Electrophoretic light scattering is based on dynamic light scattering. Typically, zeta potentials are accessible from dynamic light scattering (DLS) measurements, also known as photon correlation spectroscopy or quasi-elastic light scattering.
- DLS dynamic light scattering
- the LPMPs in the RNA composition and methods of making and using thereof may have a range of markers that identify the LPMPs as being produced using a plant EV, and/or including a segment, portion, or extract thereof.
- plant EV-marker refers to a component that is naturally associated with a plant and incorporated into or onto the plant EV in planta, such as a plant protein, a plant nucleic acid, a plant small molecule, a plant lipid, or a combination thereof. Examples of plant EV-markers can be found, for example, in Rutter and Innes, Plant Physiol. 173(1): 728-741 , 2017; Raimondo et al., Oncotarget.
- the bacterial components in the bacteria-derived lipid composition and methods of making and using thereof may have a range of markers that identify the bacterial component as being produced.
- the term “bacterial EV-marker” refers to a component that is naturally associated with a bacterium and incorporated into or onto the bacterial EV, such as a bacterial protein, a bacterial nucleic acid, a bacterial small molecule, a bacterial lipid, or a combination thereof.
- the NMPs may have a range of markers that identify the NMP as being produced from a specific source EV, and/or including a segment, portion, or extract thereof.
- EV-marker refers to a component that is naturally associated with a specific source and incorporated into or onto the EV in vivo, such as a protein, a nucleic acid, a small molecule, a lipid, or a combination thereof.
- source EV-markers include but are not limited to peptidoglycan, lipopolysaccharide, ester-linked lipids, ether-linked lipids, circular DNA, chitin, beta-glucan, pekilo, mycoprotein, cerato-platanins, exotoxins, diacylglycerol, triglycerides, phosphatidylcholine, phosphatidylinositol, ornithine lipids, glycolipids, sphingolipids, hopanoids, or ergosterol.
- the source EV marker can include a lipid.
- lipid markers that may be found in the NMP include lipid A, lipopolysaccharide, ergosterol, ornithine lipids (OLs), sulfolipids, diacylglyceryl- N,N,N-trimethylhomoserine (DGTS), glycolipids (GLs), diacylglycerol (DAG), hopanoids (HOPs), glucosyleramide, sterylglycosides, ether-linked lipids, or a combination thereof.
- lipid A lipopolysaccharide, ergosterol, ornithine lipids (OLs), sulfolipids, diacylglyceryl- N,N,N-trimethylhomoserine (DGTS), glycolipids (GLs), diacylglycerol (DAG), hopanoids (HOPs), glucosyleramide, sterylglycosides, ether-linked
- EV markers may include lipids that accumulate in sources in response to abiotic or biotic stressors.
- the source EV marker may include a protein.
- the protein EV marker may be an antimicrobial or antiviral protein naturally produced by the source, including proteins that are secreted in response to abiotic or biotic stressors.
- Some examples of protein EV markers include but are not limited to cecropins, moricins, defensins, proline- and glycine-rich peptides, fungal immunomodulatory proteins, flagellin, encapsulin, streptavidin, internalin, pilin, halocin, or archaeocins.
- the EV marker can include a protein involved in lipid metabolism,.
- the protein EV marker is a cellular trafficking protein in the source.
- the protein marker may lack a signal peptide that is typically associated with secreted proteins.
- Unconventional secretory proteins seem to share several common features like (i) lack of a leader sequence, (ii) absence of PTMs specific for ER or Golgi apparatus, and/or (iii) secretion not affected by brefeldin A which blocks the classical ER/Golgi- dependent secretion pathway.
- One skilled in the art can use a variety of tools freely accessible to the public to evaluate a protein for a signal sequence, or lack thereof.
- the protein may have an amino acid sequence having at least 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 98%, 99%, or 100% sequence identity to a known EV marker.
- the EV marker includes a nucleic acid encoded in the source, e.g., a Arthropod, Plant, Fungi, Archaea, or Bacteria RNA, DNA, or PNA.
- the NMP may include dsRNA, mRNA, a viral RNA, a microRNA (miRNA), or a small interfering RNA (siRNA) encoded by the source.
- the nucleic acid may be one that is associated with a protein that facilitates the long-distance transport of RNA.
- the nucleic acid EV marker may be one involved in host-induced gene silencing (HIGS), which is the process by which a source silences foreign transcripts of pathogens.
- the nucleic acid may be a microRNA.
- the nucleic acid may have a nucleotide sequence having at least 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 98%, 99%, or 100% sequence identity to a known EV marker.
- the EV marker includes a compound produced by the source.
- the compound may a component of the cell wall (e.g. lipopolysaccharide).
- the compound may be a defense compound produced in response to abiotic or biotic stressors, such as pathogens or extreme environmental stress.
- the NMP may also be identified as being produced from a source EV based on the lack of certain markers (e.g., lipids, polypeptides, or polynucleotides) that are not typically produced by these sources, but are generally associated with other organisms (e.g., markers of animal EVs or plant EVs).
- markers e.g., lipids, polypeptides, or polynucleotides
- the NMP lacks lipids typically found in animal EVs or plant EVs.
- EV markers can be identified using any approaches known in the art that enable identification of small molecules (e.g., mass spectroscopy, mass spectrometry), lipds (e.g., mass spectroscopy, mass spectrometry), proteins (e.g., mass spectroscopy, immunoblotting), or nucleic acids (e.g., PCR analysis).
- a NMP composition described herein includes a detectable amount, e.g., a pre-determined threshold amount, of an EV marker described herein.
- Agents e.g., nucleic acids
- the LNMPs are modified to include a RNA agent (e.g., gene editing system; a nucleic acid molecule like mRNA or gRNA) to form the RNA composition.
- a RNA agent e.g., gene editing system; a nucleic acid molecule like mRNA or gRNA
- the LNMPs can carry or associate with such agents by a variety of means to enable delivery of the agent to a target organism (e.g., a target animal), e.g., by encapsulating the agent, incorporation of the component in the lipid bilayer structure, or association of the component (e.g., by conjugation) with the surface of the lipid bilayer structure of the LNMP.
- the agent is included in the LNMP formulation, as described herein.
- the agent can be incorporated or loaded into or onto the LNMPs by any methods known in the art that allow association, directly or indirectly, between the LNMPs and agent.
- the agents can be incorporated into the LNMPs by an in vivo method (e.g., in planta, e.g., through production of LNMPs from a transgenic plant or source that comprises the agent), or in vitro (e.g., in tissue culture, or in cell culture), or both in vivo and in vitro methods.
- the LNMPs are loaded in vitro.
- the substance may be loaded onto or into (e.g., may be encapsulated by) the LNMPs using, but not limited to, physical, chemical, and/or biological methods (e.g., in tissue culture or in cell culture).
- the agent may be introduced into LNMPs by one or more of electroporation, sonication, passive diffusion, stirring, lipid extraction, or extrusion.
- the agent is incorporated into the LNMP using a microfluidic device, e.g., using a method in which natural lipids are provided in an organic phase, the heterologous functional agent is provided in an aqueous phase, and the organic and aqueous phases are combined in the microfluidics device to produce a LNMP comprising the heterologous functional agent.
- Loaded LNMPs can be assessed to confirm the presence or level of the loaded agent using a variety of methods, such as HPLC (e.g., to assess small molecules), immunoblotting (e.g., to assess proteins); and/or quantitative PCR (e.g., to assess nucleotides).
- HPLC e.g., to assess small molecules
- immunoblotting e.g., to assess proteins
- quantitative PCR e.g., to assess nucleotides
- the agent can be conjugated to the LNMP, in which the agent is connected or joined, indirectly or directly, to the LNMP.
- one or more agents can be chemically linked to a LNMP, such that the one or more agents are joined (e.g., by covalent or ionic bonds) directly to the lipid bilayer of the LNMP.
- the conjugation of various agents to the LNMPs can be achieved by first mixing the one or more agents with an appropriate crosslinking agent (e.g., N-ethylcarbo- diimide (“EDC”), which is generally utilized as a carboxyl activating agent for amide bonding with primary amines and also reacts with phosphate groups) in a suitable solvent.
- EDC N-ethylcarbo- diimide
- the cross-linking agent/ agent mixture can then be combined with the LNMPs and, after another period of incubation, subjected to a sucrose gradient (e.g., and 8, 30, 45, and 60% sucrose gradient) to separate the free agent and free LNMPs from the agent conjugated to the LNMPs.
- a sucrose gradient e.g., and 8, 30, 45, and 60% sucrose gradient
- the LNMPs conjugated to the agent are then seen as a band in the sucrose gradient, such that the conjugated LNMPs can then be collected, washed, and dissolved in a suitable solution for use as described herein.
- the LNMPs are stably associated with the agent prior to and following delivery of the LNMP, e.g., to a plant or animal.
- the LNMPs are associated with the agent such that the agent becomes dissociated from the LNMPs following delivery of the LNMP, e.g., to a plant or animal.
- the LNMPs can be loaded or the LNMP can be formulated with various concentrations of the agent, depending on the particular agent or use.
- the LNMPs are loaded or the LNMP is formulated such that the LNMP formulation disclosed herein includes about 0.001 , 0.01 , 0.1 , 1.0, 2, 3, 4, 5, 6, 7, 8, 9, 10, 15, 20, 30, 40, 50, 60, 70, 80, 90, or 95 (or any range between about 0.001 and 95) or more wt% of an agent.
- the LNMPs are loaded or the LNMP is formulated such that the LNMP formulation includes about 95, 90, 80, 70, 60, 50, 40, 30, 20, 15, 10, 9, 8, 7, 6, 5, 4, 3, 2, 1 .0, 0.1 , 0.01 , 0.001 (or any range between about 95 and 0.001) or less wt% of an agent.
- the LNMP formulation can include about 0.001 to about 0.01 wt%, about 0.01 to about 0.1 wt%, about 0.1 to about 1 wt%, about 1 to about 5 wt%, or about 5 to about 10 wt%, about 10 to about 20 wt% of the agent.
- the LNMP can be loaded or the LNMP is formulated with about 1 , 5, 10, 50, 100, 200, or 500, 1 ,000, 2,000 (or any range between about 1 and 2,000) or more pg/ml of an agent.
- a LNMP of the invention can be loaded or a LNMP can be formulated with about 2,000, 1 ,000, 500, 200, 100, 50, 10, 5, 1 (or any range between about 2,000 and 1) or less pg/ml of an agent.
- the LNMPs are loaded or the LNMP is formulated such that the LNMP formulation disclosed herein includes at least 0.001 wt%, at least 0.01 wt%, at least 0.1 wt%, at least 1 .0 wt%, at least 2 wt%, at least 3 wt%, at least 4 wt%, at least 5 wt%, at least 6 wt%, at least 7 wt%, at least 8 wt%, at least 9 wt%, at least 10 wt%, at least 15 wt%, at least 20 wt%, at least 30 wt%, at least 40 wt%, at least 50 wt%, at least 60 wt%, at least 70 wt%, at least 80 wt%, at least 90 wt%, or at least 95 wt% of an agent.
- the LNMP can be loaded or the LNMP can be formulated with at least 1 pg/ml, at least 5 pg/ml, at least 10 pg/ml, at least 50 pg/ml, at least 100 pg/ml, at least 200 pg/ml, at least 500 pg/ml, at least 1 ,000 pg/ml, at least 2,000 pg/ml of an agent.
- the LNMP is formulated with the agent by suspending the LNMPs in a solution comprising or consisting of the agent, e.g., suspending or resuspending the LNMPs by vigorous mixing.
- the agent e.g., cell-penetrating agent, e.g., nucleic acids, enzyme, detergent, ionic, fluorous, or zwitterionic liquid, or ionizable lipid may comprise, e.g., less than 1% or at least 1%, 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% of the solution.
- the modified LNMPs are formulated into pharmaceutical compositions (i.e., a RNA composition), e.g., for administration to an animal (e.g., a human).
- the pharmaceutical composition may be administered to an animal (e.g., human) with a pharmaceutically acceptable diluent, carrier, and/or excipient.
- the pharmaceutical composition of the methods described herein will be formulated into suitable pharmaceutical compositions to permit facile delivery.
- the single dose may be in a unit dose form as needed.
- the LNMP I RNA composition may be formulated for e.g., oral administration, intravenous administration (e.g., injection or infusion), intramuscular, or subcutaneous administration to an animal.
- intravenous administration e.g., injection or infusion
- intramuscular e.g., intramuscular
- subcutaneous administration e.g., intramuscular, or subcutaneous administration to an animal.
- various effective pharmaceutical carriers are known in the art (See, e.g., Remington: The Science and Practice of Pharmacy, 22 nd ed., (2012) and ASHP Handbook on Injectable Drugs, 18 th ed., (2014)).
- Suitable pharmaceutically acceptable carriers and excipients are nontoxic to recipients at the dosages and concentrations employed.
- Acceptable carriers and excipients may include buffers such as phosphate, citrate, HEPES, and TAE, antioxidants such as ascorbic acid and methionine, preservatives such as hexamethonium chloride, octadecyldimethylbenzyl ammonium chloride, resorcinol, and benzalkonium chloride, proteins such as human serum albumin, gelatin, dextran, and immunoglobulins, hydrophilic polymers such as polyvinylpyrrolidone, amino acids such as glycine, glutamine, histidine, and lysine, and carbohydrates such as glucose, mannose, sucrose, and sorbitol.
- buffers such as phosphate, citrate, HEPES, and TAE
- antioxidants such as ascorbic acid and methionine
- preservatives such as hexamethon
- the LNMP I RNA composition may be formulated according to conventional pharmaceutical practice.
- concentration of the compound in the formulation will vary depending upon a number of factors, including the dosage of the active agent (e.g., LNMPs and nucleic acids) to be administered, and the route of administration.
- the active agent e.g., LNMPs and nucleic acids
- the LNMP I RNA composition can be prepared in the form of an oral formulation.
- Formulations for oral use can include tablets, caplets, capsules, syrups, or oral liquid dosage forms containing the active ingredient(s) in a mixture with non-toxic pharmaceutically acceptable excipients.
- excipients may be, for example, inert diluents or fillers (e.g., sucrose, sorbitol, sugar, mannitol, microcrystalline cellulose, starches including potato starch, calcium carbonate, sodium chloride, lactose, calcium phosphate, calcium sulfate, or sodium phosphate); granulating and disintegrating agents (e.g., cellulose derivatives including microcrystalline cellulose, starches including potato starch, croscarmellose sodium, alginates, or alginic acid); binding agents (e.g., sucrose, glucose, sorbitol, acacia, alginic acid, sodium alginate, gelatin, starch, pregelatinized starch, microcrystalline cellulose, magnesium aluminum silicate, carboxymethylcellulose sodium, methylcellulose, hydroxypropyl methylcellulose, ethylcellulose, polyvinylpyrrolidone, or polyethylene glycol); and lubricating agents, glidants, and antiad
- compositions for oral use may also be provided in unit dosage form as chewable tablets, non-chewable tablets, caplets, capsules (e.g., as hard gelatin capsules wherein the active ingredient is mixed with an inert solid diluent, or as soft gelatin capsules wherein the active ingredient is mixed with water or an oil medium).
- the compositions disclosed herein may also further include an immediate-release, extended release or delayed-release formulation.
- the LNMP I RNA compositions may be formulated in the form of liquid solutions or suspensions and administered by a parenteral route (e.g., subcutaneous, intravenous, or intramuscular).
- the pharmaceutical composition can be formulated for injection or infusion.
- Pharmaceutical compositions for parenteral administration can be formulated using a sterile solution or any pharmaceutically acceptable liquid as a vehicle.
- Pharmaceutically acceptable vehicles include, but are not limited to, sterile water, physiological saline, or cell culture media (e.g., Dulbecco’s Modified Eagle Medium (DMEM), a-Modified Eagles Medium (a-MEM), and F-12 medium).
- DMEM Modified Eagle Medium
- a-MEM a-Modified Eagles Medium
- F-12 medium F-12 medium
- the LNMP I RNA composition includes one or more nucleic acid molecules, e.g., polynucleotides, which encode one or more wild type or engineered proteins, peptides, or polypeptides.
- exemplary polynucleotides e.g., polynucleotide constructs, include gene editing system - encoding RNA polynucleotides, e.g., mRNAs and gRNAs.
- polypeptides that can be used herein can include an enzyme (e.g., a metabolic recombinase, a helicase, an integrase, a RNAse, a DNAse, or an ubiquitination protein), a poreforming protein, a signaling ligand, a cell penetrating peptide, a transcription factor, a receptor, an antibody, a nanobody, a gene editing protein (e.g., CRISPR-Cas system, TALEN, or zinc finger), riboprotein, a protein aptamer, or a chaperone.
- an enzyme e.g., a metabolic recombinase, a helicase, an integrase, a RNAse, a DNAse, or an ubiquitination protein
- a poreforming protein e.g., a signaling ligand, a cell penetrating peptide, a transcription factor, a receptor,
- Polypeptides included herein may include naturally occurring polypeptides or recombinantly produced variants.
- the polypeptide may be a functional fragments or variants thereof (e.g., an enzymatically active fragment or variant thereof).
- the polypeptide may be a functionally active variant of any of the polypeptides described herein with at least 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81 %, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity, e.g., over a specified region or over the entire sequence, to a sequence of a polypeptide described herein or a naturally occurring polypeptide.
- the polypeptide may have at least 50% (e.g., at least 50%, 60%, 70%, 80%, 90%, 95%, 97%, 99%, or greater) identity to a protein of interest.
- the LNMP I RNA composition may include any number or type (e.g., classes) of polypeptides, such as at least about any one of 1 polypeptide, 2, 3, 4, 5, 10, 15, 20, or more polypeptides.
- a suitable concentration of each polypeptide in the LNMP I RNA composition depends on factors such as efficacy, stability of the polypeptide, number of distinct polypeptides in the formulation, and methods of application of the formulation.
- each polypeptide in a liquid formulation is from about 0.1 ng/mL to about 100 mg/mL.
- each polypeptide in a solid formulation is from about 0.1 ng/g to about 100 mg/g.
- the LNMP I RNA composition include a heterologous nucleic acid encoding a polypeptide.
- Nucleic acids encoding a polypeptide may have a length from about 10 to about 50,000 nucleotides (nts), about 25 to about 100 nts, about 50 to about 150 nts, about 100 to about 200 nts, about 150 to about 250 nts, about 200 to about 300 nts, about 250 to about 350 nts, about 300 to about 500 nts, about 10 to about 1000 nts, about 50 to about 1000 nts, about 100 to about 1000 nts, about 1000 to about 2000 nts, about 2000 to about 3000 nts, about 3000 to about 4000 nts, about 4000 to about 5000 nts, about 5000 to about 6000 nts, about 6000 to about 7000 nts, about 7000 to about 8000 nts, about 8000 to about
- the LNMP I RNA composition may also include active variants of a nucleic acid sequence of interest.
- the variant of the nucleic acids has at least 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity, e.g., over a specified region or overthe entire sequence, to a sequence of a nucleic acid of interest.
- the invention includes an active polypeptide encoded by a nucleic acid variant as described herein.
- the active polypeptide encoded by the nucleic acid variant has at least 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91 %, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% identity, e.g., over a specified region or over the entire amino acid sequence, to a sequence of a polypeptide of interest or the naturally derived polypeptide sequence.
- Certain methods for expressing a nucleic acid encoding a protein may involve expression in cells, including insect, yeast, plant, bacteria, or other cells under the control of appropriate promoters.
- Expression vectors may include nontranscribed elements, such as an origin of replication, a suitable promoter and enhancer, and other 5’ or 3’ flanking nontranscribed sequences, and 5’ or 3’ nontranslated sequences such as necessary ribosome binding sites, a polyadenylation site, splice donor and acceptor sites, and termination sequences.
- DNA sequences derived from the SV40 viral genome for example, SV40 origin, early promoter, enhancer, splice, and polyadenylation sites may be used to provide the other genetic elements required for expression of a heterologous DNA sequence.
- Appropriate cloning and expression vectors for use with bacterial, fungal, yeast, and mammalian cellular hosts are described in Green et al., Molecular Cloning: A Laboratory Manual, Fourth Edition, Cold Spring Harbor Laboratory Press, 2012.
- a nucleic acid sequence coding for a desired gene can be obtained using recombinant methods known in the art, such as, for example by screening libraries from cells expressing the gene, by deriving the gene from a vector known to include the same, or by isolating directly from cells and tissues containing the same, using standard techniques.
- a gene of interest can be produced synthetically, rather than cloned.
- Expression of natural or synthetic nucleic acids is typically achieved by operably linking a nucleic acid encoding the gene of interest to a promoter, and incorporating the construct into an expression vector.
- Expression vectors can be suitable for replication and expression in bacteria.
- Expression vectors can also be suitable for replication and integration in eukaryotes.
- Typical cloning vectors contain transcription and translation terminators, initiation sequences, and promoters useful for expression of the desired nucleic acid sequence.
- Additional promoter elements e.g., enhancers, regulate the frequency of transcriptional initiation.
- these are located in the region 30-110 basepairs (bp) upstream of the start site, although a number of promoters have recently been shown to contain functional elements downstream of the start site as well.
- the spacing between promoter elements frequently is flexible, so that promoter function is preserved when elements are inverted or moved relative to one another.
- tk thymidine kinase
- the spacing between promoter elements can be increased to 50 bp apart before activity begins to decline.
- individual elements can function either cooperatively or independently to activate transcription.
- a suitable promoter is the immediate early cytomegalovirus (CMV) promoter sequence. This promoter sequence is a strong constitutive promoter sequence capable of driving high levels of expression of any polynucleotide sequence operatively linked thereto.
- CMV immediate early cytomegalovirus
- EF-1a Elongation Growth Factor-1a
- constitutive promoter sequences may also be used, including, but not limited to the simian virus 40 (SV40) early promoter, mouse mammary tumor virus (MMTV), human immunodeficiency virus (HIV) long terminal repeat (LTR) promoter, MoMuLV promoter, an avian leukemia virus promoter, an Epstein-Barr virus immediate early promoter, a Rous sarcoma virus promoter, as well as human gene promoters such as, but not limited to, the actin promoter, the myosin promoter, the hemoglobin promoter, and the creatine kinase promoter.
- SV40 simian virus 40
- MMTV mouse mammary tumor virus
- HSV human immunodeficiency virus
- LTR long terminal repeat
- MoMuLV promoter MoMuLV promoter
- an avian leukemia virus promoter an Epstein-Barr virus immediate early promoter
- Rous sarcoma virus promoter as well as human gene promoters such as
- the promoter may be an inducible promoter.
- an inducible promoter provides a molecular switch capable of turning on expression of the polynucleotide sequence to which it is operatively linked when such expression is desired, or turning off the expression when expression is not desired.
- inducible promoters include, but are not limited to a metallothionine promoter, a glucocorticoid promoter, a progesterone promoter, and a tetracycline promoter.
- the expression vector to be introduced can also contain either a selectable marker gene or a reporter gene or both to facilitate identification and selection of expressing cells from the population of cells sought to be transfected or infected through viral vectors.
- the selectable marker may be carried on a separate piece of DNA and used in a co-transfection procedure. Both selectable markers and reporter genes may be flanked with appropriate regulatory sequences to enable expression in the host cells.
- Useful selectable markers include, for example, antibioticresistance genes, such as neo and the like.
- Reporter genes may be used for identifying potentially transformed cells and for evaluating the functionality of regulatory sequences.
- a reporter gene is a gene that is not present in or expressed by the recipient source and that encodes a polypeptide whose expression is manifested by some easily detectable property, e.g., enzymatic activity. Expression of the reporter gene is assayed at a suitable time after the DNA has been introduced into the recipient cells.
- Suitable reporter genes may include genes encoding luciferase, beta-galactosidase, chloramphenicol acetyl transferase, secreted alkaline phosphatase, or the green fluorescent protein gene (e.g., Ui-Tei et al., FEBS Letters 479:79-82, 2000). Suitable expression systems are well known and may be prepared using known techniques or obtained commercially. In general, the construct with the minimal 5’ flanking region showing the highest level of expression of reporter gene is identified as the promoter. Such promoter regions may be linked to a reporter gene and used to evaluate agents for the ability to modulate promoter-driven transcription.
- an organism may be genetically modified to alter expression of one or more proteins. Expression of the one or more proteins may be modified for a specific time, e.g., development or differentiation state of the organism. In one instance, provided is a composition to alter expression of one or more proteins, e.g., proteins that affect activity, structure, or function. Expression of the one or more proteins may be restricted to a specific location(s) or widespread throughout the organism.
- mRNA messenger RNA
- the LNMP / RNA composition may include a mRNA molecule, e.g., a mRNA molecule encoding a polypeptide.
- the mRNA molecule can be synthetic and modified (e.g., chemically).
- the mRNA molecule can be chemically synthesized or transcribed in vitro.
- the mRNA molecule can be disposed on a plasmid, e.g., a viral vector, bacterial vector, or eukaryotic expression vector.
- the mRNA molecule can be delivered to cells by transfection, electroporation, or transduction (e.g., adenoviral or lentiviral transduction).
- the modified RNA agent of interest described herein has modified nucleosides or nucleotides. Such modifications are known and are described, e.g., in WO 2012/019168. Additional modifications are described, e.g., in WO 2015/038892; WO 2015/038892; WO 2015/089511 ; WO 2015/196130; WO 2015/196118 and WO 2015/196128 A2, which are herein incorporated by reference in their entirety.
- the modified RNA encoding a polypeptide of interest has one or more terminal modification, e.g., a 5’ cap structure and/or a poly-A tail (e.g., of between 100-200 nucleotides in length).
- the 5’ cap structure may be selected from the group consisting of CapO, Capl, ARCA, inosine, Nl-methyl-guanosine, 2’fluoro- guanosine, 7-deaza-guanosine, 8-oxo-guanosine, 2- amino-guanosine, LNA-guanosine, and 2-azido- guanosine.
- the modified RNAs also contain a 5‘ UTR including at least one Kozak sequence, and a 3‘ UTR.
- modifications are known and are described, e.g., in WO 2012/135805 and WO 2013/052523, which are incorporated herein by reference in their entirety. Additional terminal modifications are described, e.g., in WO 2014/164253 and WO 2016/011306, WO 2012/045075, and WO 2014/093924, which are incorporated herein by reference in their entirety.
- Chimeric enzymes for synthesizing capped RNA molecules e.g., modified mRNA
- modified mRNA which may include at least one chemical modification are described in WO 2014/028429, which is incorporated herein by reference in its entirety.
- a modified mRNA may be cyclized, or concatemerized, to generate a translation competent molecule to assist interactions between poly-A binding proteins and 5 ‘-end binding proteins.
- the mechanism of cyclization or concatemerization may occur through at least 3 different routes: 1) chemical, 2) enzymatic, and 3) ribozyme catalyzed.
- the newly formed 5’- /3’- linkage may be intramolecular or intermolecular.
- modifications are described, e.g., in WO 2013/151736.
- modified RNAs are made using only in vitro transcription (IVT) enzymatic synthesis.
- IVT in vitro transcription
- Methods of making IVT polynucleotides are known in the art and are described in WO 2013/151666, WO 2013/151668, WO 2013/151663, WO 2013/151669, WO 2013/151670, WO 2013/151664, WO 2013/151665, WO 2013/151671 , WO 2013/151672, WO 2013/151667 and WO 2013/151736, which are incorporated herein by reference in their entirety.
- Methods of purification include purifying an RNA transcript including a polyA tail by contacting the sample with a surface linked to a plurality of thymidines or derivatives thereof and/or a plurality of uracils or derivatives thereof (polyT/U) under conditions such that the RNA transcript binds to the surface and eluting the purified RNA transcript from the surface (WO 2014/152031); using ion (e.g., anion) exchange chromatography that allows for separation of longer RNAs up to 10,000 nucleotides in length via a scalable method (WO 2014/144767); and subjecting a modified mRNA sample to DNAse treatment (WO 2014/152030).
- ion e.g., anion
- Formulations of modified RNAs are known and are described, e.g., in WO 2013/090648.
- the formulation may be, but is not limited to, nanoparticles, poly(lactic-co-glycolic acid)(PLGA) microspheres, lipidoids, lipoplex, liposome, polymers, carbohydrates (including simple sugars), cationic lipids, fibrin gel, fibrin hydrogel, fibrin glue, fibrin sealant, fibrinogen, thrombin, rapidly eliminated lipid nanoparticles (reLNPs) and combinations thereof.
- RNAs encoding polypeptides in the fields of human disease, antibodies, viruses, and a variety of in vivo settings are known and are disclosed in for example, Table 6 of International Publication Nos. WO 2013/151666, WO 2013/151668, WO 2013/151663, WO 2013/151669, WO 2013/151670, WO 2013/151664, WO 2013/151665, WO 2013/151736; Tables 6 and 7 International Publication No. WO 2013/151672; Tables 6, 178 and 179 of International Publication No. WO 2013/151671 ; Tables 6, 185 and 186 of International Publication No WO 2013/151667; which are incorporated herein by reference in their entirety. Any of the foregoing may be synthesized as an IVT polynucleotide, chimeric polynucleotide or a circular polynucleotide, and each may include one or more modified nucleotides or terminal modifications.
- the LNMP I RNA composition includes an inhibitory RNA molecule, e.g., that acts via the RNA interference (RNAi) pathway.
- the inhibitory RNA molecule decreases the level of gene expression in a plant and/or decreases the level of a protein in the plant.
- the inhibitory RNA molecule inhibits expression of a plant gene.
- an inhibitory RNA molecule may include a short interfering RNA or its precursor, short hairpin RNA, and/or a microRNA or its precursor that targets a gene in the plant. Certain RNA molecules can inhibit gene expression through the biological process of RNA interference (RNAi).
- RNAi RNA interference
- RNAi molecules include RNA or RNA-like structures typically containing 15-50 base pairs (such as about 18-25 base pairs) and having a nucleobase sequence identical (or complementary) or nearly identical (or substantially complementary) to a coding sequence in an expressed target gene within the cell.
- RNAi molecules include, but are not limited to short interfering RNAs (siRNAs), double-strand RNAs (dsRNA), short hairpin RNAs (shRNA), meroduplexes, dicer substrates, and multivalent RNA interference (U.S. Pat. Nos. 8,084,599 8,349,809, 8,513,207 and 9,200,276, which are incorporated herein by reference in their entirety).
- the inhibitory RNA molecule can be chemically synthesized or transcribed in vitro.
- inhibitory RNA molecules include those described in details in International Patent Application Publication No. WO 2021/041301 , which is incorporated herein by reference in its entirety.
- the LNP I RNA compositions or LNMP I RNA compositions may include a component of a gene editing system.
- the agent may introduce an alteration (e.g., insertion, deletion (e.g., knockout), translocation, inversion, single point mutation, or other mutation) in a gene in the target cell.
- exemplary gene editing systems include the zinc finger nucleases (ZFNs), Transcription Activator-Like Effector-based Nucleases (TALEN), and the clustered regulatory interspaced short palindromic repeat (CRISPR) system. ZFNs, TALENs, and CRISPR-based methods are described, e.g., in Gaj et al., Trends Biotechnol. 31 (7):397-405, 2013.
- an endonuclease is directed to a target nucleotide sequence (e.g., a site in the genome that is to be sequence-edited) by sequence-specific, non-coding guide RNAs (eg sgRNA) that target single- or double-stranded DNA sequences.
- sequence-specific, non-coding guide RNAs eg sgRNA
- l-lll Three classes (l-lll) of CRISPR systems are known, and class II CRISPR systems use a single Cas endonuclease (rather than multiple Cas proteins).
- One class II CRISPR system includes a type II Cas endonuclease such as Cas9, a CRISPR RNA (crRNA), and a trans-activating crRNA (tracrRNA).
- the crRNA contains a guide RNA, i.e., typically about 20-nucleotide RNA sequence that corresponds to a target DNA sequence.
- the crRNA also contains a region that binds to the tracrRNA to form a partially double-stranded structure which is cleaved by RNase III, resulting in a crRNA/tracrRNA hybrid.
- the RNAs serve as guides to direct Cas proteins to silence specific DNA/RNA sequences, depending on the spacer sequence.
- the target DNA sequence must generally be adjacent to a protospacer adjacent motif (PAM) that is specific for a given Cas endonuclease; however, PAM sequences appear throughout a given genome.
- PAM protospacer adjacent motif
- CRISPR endonucleases identified from various prokaryotic species have unique PAM sequence requirements; examples of PAM sequences include 5’-NGG (Streptococcus pyogenes), 5’- NNAGAA (Streptococcus thermophilus CRISPR1), 5’-NGGNG (Streptococcus thermophilus CRISPR3), and 5’-NNNGATT (Neisseria meningiditis).
- 5’-NGG Streptococcus pyogenes
- 5’- NNAGAA Streptococcus thermophilus CRISPR1
- 5’-NGGNG Streptococcus thermophilus CRISPR3
- 5’-NNNGATT Neisseria meningiditis
- Some endonucleases are associated with G-rich PAM sites, e.g., 5’-NGG, and perform blunt-end cleaving of the target DNA at a location 3 nucleotides upstream from (5’ from) the PAM site.
- Another class II CRISPR system includes the type V endonuclease Cpfl (e.g. AsCpfl (from Acidaminococcus sp.) and LbCpfl (from Lachnospiraceae sp.)), which is smaller than Cas9.
- Cpfl -associated CRISPR arrays are processed into mature crRNAs without the requirement of a tracrRNA.
- Cpfl endonucleases are associated with T-rich PAM sites, e.g., 5’-TTN, but can also recognize a 5’-CTA PAM motif.
- Cpfl cleaves the target DNA by introducing an offset or staggered double-strand break with a 4- or 5-nucleotide 5’ overhang, for example, cleaving a target DNA with a 5-nucleotide offset or staggered cut located 18 nucleotides downstream from (3’ from) from the PAM site on the coding strand and 23 nucleotides downstream from the PAM site on the complimentary strand; the 5-nucleotide overhang that results from such offset cleavage allows more precise genome editing by DNA insertion by homologous recombination than by insertion at blunt-end cleaved DNA.
- CRISPR arrays can be designed to contain one or multiple guide RNA sequences corresponding to a desired target DNA sequence; see, for example, Cong et al., Science 339:819-823, 2013; Ran et al., Nature Protocols 8:2281-2308, 2013. At least about 16 or 17 nucleotides of gRNA sequence are required by Cas9 for DNA cleavage to occur; for Cpfl at least about 16 nucleotides of gRNA sequence is needed to achieve detectable DNA cleavage.
- guide RNA sequences are generally designed to have a length of between 17-24 nucleotides (e.g., 19, 20, or 21 nucleotides) and complementarity to the targeted gene or nucleic acid sequence.
- Custom gRNA generators and algorithms are available commercially for use in the design of effective guide RNAs.
- Gene editing has also been achieved using a chimeric single guide RNA (sgRNA), an engineered (synthetic) single RNA molecule that mimics a naturally occurring crRNA-tracrRNA complex and contains both a tracrRNA (for binding the nuclease) and at least one crRNA (to guide the nuclease to the sequence targeted for editing).
- sgRNA chimeric single guide RNA
- Chemically modified sgRNAs have also been demonstrated to be effective in genome editing; see, for example, Hendel et al., Nature Biotechnol. 985-991 , 2015.
- dCas9 can further be fused with an effector to repress (CRISPRi) or activate (CRISPRa) expression of a target gene.
- Cas9 can be fused to a transcriptional repressor (e.g., a KRAB domain) or a transcriptional activator (e.g., a dCas9-VP64 fusion).
- a catalytically inactive Cas9 (dCas9) fused to Fokl nuclease (dCas9-Fokl) can be used to generate DSBs at target sequences homologous to two gRNAs. See, e.g., the numerous CRISPR/Cas9 plasmids disclosed in and publicly available from the Addgene repository (Addgene, 75 Sidney St., Suite 550A, Cambridge, MA 02139; addgene.org/crispr/).
- a double nickase Cas9 that introduces two separate double-strand breaks, each directed by a separate guide RNA, is described as achieving more accurate genome editing by Ran et al., Cell 154:1380-1389, 2013.
- CRISPR technology for editing the genes of eukaryotes is disclosed in US Patent Application Publications US 2016/0138008 A1 and US 2015/0344912 A1 , and in US Patents 8,697,359, 8,771 ,945, 8,945,839, 8,999,641 , 8,993,233, 8,895,308, 8,865,406, 8,889,418, 8,871 ,445, 8,889,356, 8,932,814, 8,795,965, and 8,906,616.
- Cpfl endonuclease and corresponding guide RNAs and PAM sites are disclosed in US Patent Application Publication 2016/0208243 A1 .
- the desired genome modification involves homologous recombination, wherein one or more double-stranded DNA breaks in the target nucleotide sequence is generated by the RNA-guided nuclease and guide RNA(s), followed by repair of the break(s) using a homologous recombination mechanism (homology-directed repair).
- a donor template that encodes the desired nucleotide sequence to be inserted or knocked-in at the double-stranded break is provided to the cell or subject; suitable templates include single-stranded DNA templates and double-stranded DNA templates.
- a donor template encoding a nucleotide change over a region of less than about 50 nucleotides is provided in the form of single-stranded DNA; larger donor templates (e.g., more than 100 nucleotides) are often provided as double-stranded DNA plasmids.
- the donor template is provided to the cell or subject in a quantity that is sufficient to achieve the desired homology-directed repair but that does not persist in the cell or subject after a given period of time (e.g., after one or more cell division cycles).
- a donor template has a core nucleotide sequence that differs from the target nucleotide sequence (e.g., a homologous endogenous genomic region) by at least 1 , at least 5, at least 10, at least 20, at least 30, at least 40, at least 50, or more nucleotides.
- This core sequence is flanked by homology arms or regions of high sequence identity with the targeted nucleotide sequence; in some instances, the regions of high identity include at least 10, at least 50, at least 100, at least 150, at least 200, at least 300, at least 400, at least 500, at least 600, at least 750, or at least 1000 nucleotides on each side of the core sequence.
- the core sequence is flanked by homology arms including at least 10, at least 20, at least 30, at least 40, at least 50, at least 60, at least 70, at least 80, or at least 100 nucleotides on each side of the core sequence.
- the core sequence is flanked by homology arms including at least 500, at least 600, at least 700, at least 800, at least 900, or at least 1000 nucleotides on each side of the core sequence.
- two separate doublestrand breaks are introduced into the cell or subject’s target nucleotide sequence with a double nickase Cas9 (see Ran et al., Cell 154:1380-1389, 2013), followed by delivery of the donor template.
- the RNA composition includes a gRNA and a targeted nuclease, e.g., a Cas9, e.g., a wild type Cas9, a nickase Cas9 (e.g., Cas9 D10A), a dead Cas9 (dCas9), eSpCas9, Cpfl , C2C1 , or C2C3, or a nucleic acid encoding such a nuclease.
- a Cas9 e.g., a wild type Cas9, a nickase Cas9 (e.g., Cas9 D10A), a dead Cas9 (dCas9), eSpCas9, Cpfl , C2C1 , or C2C3, or a nucleic acid encoding such a nuclease.
- a Cas9 e.g., a wild type Cas9, a nickase Ca
- nuclease and gRNA(s) are determined by whether the targeted mutation is a deletion, substitution, or addition of nucleotides, e.g., a deletion, substitution, or addition of nucleotides to a targeted sequence.
- Fusions of a catalytically inactive endonuclease e.g., a dead Cas9 (dCas9, e.g., D10A; H840A) tethered with all or a portion of (e.g., biologically active portion of) an (one or more) effector domain create chimeric proteins that can be linked to the polypeptide to guide the RNA composition to specific DNA sites by one or more RNA sequences (sgRNA) to modulate activity and/or expression of one or more target nucleic acids sequences.
- dCas9 dead Cas9
- H840A dead Cas9
- sgRNA RNA sequences
- the agent includes a guide RNA (gRNA) for use in a CRISPR system for gene editing.
- the agent includes a zinc finger nuclease (ZFN), or a mRNA encoding a ZFN, that targets (e.g., cleaves) a nucleic acid sequence (e.g., DNA sequence) of a gene.
- the agent includes a TALEN, or an mRNA encoding a TALEN, that targets (e.g., cleaves) a nucleic acid sequence (e.g., DNA sequence) in a gene in the target organism.
- the gRNA can be used in a CRISPR system to engineer an alteration in a gene in the target organism.
- the ZFN and/or TALEN can be used to engineer an alteration in a gene.
- Exemplary alterations include insertions, deletions (e.g., knockouts), translocations, inversions, single point mutations, or other mutations.
- the alteration can be introduced in the gene in a cell, e.g., in vitro, ex vivo, or in vivo. In some examples, the alteration increases the level and/or activity of a gene in the target organism.
- the alteration decreases the level and/or activity of (e.g., knocks down or knocks out) a gene in the target organism.
- the alteration corrects a defect (e.g., a mutation causing a defect), in a gene in the target organism.
- the CRISPR system is used to edit (e.g., to add or delete a base pair) a target gene in the target organism.
- the CRISPR system is used to introduce a premature stop codon, e.g., thereby decreasing the expression of a target gene.
- the CRISPR system is used to turn off a target gene in a reversible manner, e.g., similarly to RNA interference.
- the CRISPR system is used to direct Cas to a promoter of a gene, thereby blocking an RNA polymerase sterically.
- a CRISPR system can be generated to edit a gene in the target organism, using technology described in, e.g., U.S. Publication No. 20140068797, Cong, Science 339: 819-823, 2013; Tsai, Nature Biotechnol. 32:6569-576, 2014; U.S. Patent Nos.: 8,871 ,445; 8,865,406; 8,795,965; 8,771 ,945; and 8,697,359.
- the CRISPR interference (CRISPRi) technique can be used for transcriptional repression of specific genes in the target organism.
- an engineered Cas9 protein e.g., nuclease-null dCas9, or dCas9 fusion protein, e.g., dCas9-KRAB ordCas9-SID4X fusion
- sgRNA sequence specific guide RNA
- the Cas9-gRNA complex can block RNA polymerase, thereby interfering with transcription elongation.
- the complex can also block transcription initiation by interfering with transcription factor binding.
- CRISPRi method is specific with minimal off-target effects and is multiplexable, e.g., can simultaneously repress more than one gene (e.g., using multiple gRNAs). Also, the CRISPRi method permits reversible gene repression.
- CRISPR-mediated gene activation can be used for transcriptional activation of a gene in the target organism.
- dCas9 fusion proteins recruit transcriptional activators.
- dCas9 can be fused to polypeptides (e.g., activation domains) such as VP64 or the p65 activation domain (p65D) and used with sgRNA (e.g., a single sgRNA or multiple sgRNAs), to activate a gene or genes in the target organism.
- activation domains e.g., VP64 or the p65 activation domain (p65D)
- sgRNA e.g., a single sgRNA or multiple sgRNAs
- Multiple activators can be recruited by using multiple sgRNAs - this can increase activation efficiency.
- a variety of activation domains and single or multiple activation domains can be used.
- sgRNAs can also be engineered to recruit activators.
- RNA aptamers can be incorporated into a sgRNA to recruit proteins (e.g., activation domains) such as VP64.
- proteins e.g., activation domains
- the synergistic activation mediator (SAM) system can be used for transcriptional activation.
- SAM synergistic activation mediator
- MS2 aptamers are added to the sgRNA.
- MS2 recruits the MS2 coat protein (MCP) fused to p65AD and heat shock factor 1 (HSF1).
- MCP MS2 coat protein
- HSF1 heat shock factor 1
- RNA-guided DNA-binding agent is a Class 2 Cas nuclease.
- the RNA-guided DNA-binding agent has cleavase activity, which can also be referred to as double-strand endonuclease activity.
- the RNA-guided DNA- binding agent comprises a Cas nuclease, such as a Class 2 Cas nuclease (which may be, e.g., a Cas nuclease of Type II V, or VI).
- Class 2 Cas nucleases include, for example, Cas9, Cpf1 , C2c1 , C2c2, and C2c3 proteins and modifications thereof. Examples of Cas9 nucleases include those of the type II CRISPR systems of S. pyogenes, S.
- the Cas nuclease may be from a Type-IIA, Type-IIB, or Type-IIC system.
- Non-limiting exemplary species that the Cas nuclease can be derived from include Streptococcus pyogenes, Streptococcus thermophilus, Streptococcus sp., Staphylococcus aureus, Listeria innocua, Lactobacillus gasseri, Francisella novicida, Wolinella succinogenes, Sutterella wadsworthensis, Gammaproteobacterium, Neisseria meningitidis.
- Campylobacter jejuni Pasteurella multocida, Fibrobacter succinogene, Rhodospirillum rubrum, Nocardiopsis rougei, Streptomyces pristinaespiralis, Streptomyces viridochromogenes, Streptomyces viridochromogenes, Streptosporangium roseum, Streptosporangium roseum, Alicyclobacillus acidocaldarius.
- Bacillus pseudomycoides Bacillus selenitireducens, Exiguobacterium sibiricum, Lactobacillus delbrueckii, Lactobacillus salivarius, Lactobacillus buchneri, Treponema denticola, Microscilla marina, Burkholderiales bacterium, Polaromonas naphthalenivorans, Polaromonas sp., Crocosphaera watsonii, Cyanothece sp., Microcystis aeruginosa, Synechococcus sp., Acetohalobium arabaticum, Ammonifex degensii, Caldicommeosiruptor becscii, Candidatus Desulforudis, Clostridium botulinum, Clostridium difficile, Finegoldia magna, Natranaerobius thermophilus, Pelotomaculum thermopropionicum, Ac idithio bacillus caldus,
- Streptococcus pasteurianus Neisseria cinerea, Campylobacter lari, Parvibaculum lavamentivorans, Corynebacterium diphtheria, Acidaminococcus sp., Lachnospiraceae bacterium ND2006, and Acaryochloris marina.
- the Cas nuclease is the Cas9 nuclease from Streptococcus pyogenes. In some embodiments, the Cas nuclease is the Cas9 nuclease from Streptococcus thermophilus. In some embodiments, the Cas nuclease is the Cas9 nuclease from Neisseria meningitidis. In some embodiments, the Cas nuclease is the Cas9 nuclease is from Staphylococcus aureus. In some embodiments, the Cas nuclease is the Cpf1 nuclease from Francisella novicida.
- the Cas nuclease is the Cpf1 nuclease from Acidaminococcus sp. In some embodiments, the Cas nuclease is the Cpf1 nuclease from Lachnospiraceae bacterium ND2006.
- the Cas nuclease is the Cpfl nuclease from Francisella tularensis, Lachnospiraceae bacterium, Butyrivibrio proteoclasticus, Peregrinibacteria bacterium, Parcubacferia bacterium, Smithella, Acidaminococcus, Candidatus Methanoplasma termitum, Eubacterium eligens, Moraxella bovoculi, Leptospira inadai, Porphyromonas crevioricanis, Prevotella disiens, or Porphyromonas macacae.
- the Cas nuclease is a Cpf1 nuclease from an Acidaminococcus or Lachnospiraceae.
- the Cas9 nuclease comprises more than one RuvC domain and/or more than one HNH domain.
- the Cas9 nuclease is a wild type Cas9.
- the Cas9 is capable of inducing a double strand break in target DNA.
- the Cas nuclease may cleave dsDNA, it may cleave one strand of dsDNA, or it may not have DNA cleavase or nickase activity.
- chimeric Cas nucleases are used, where one domain or region of the protein is replaced by a portion of a different protein.
- a Cas nuclease domain may be replaced with a domain from a different nuclease such as Fok1.
- a Cas nuclease may be a modified nuclease.
- the Cas nuclease may be from a Type-1 CRISPR/Cas system.
- the Cas nuclease may be a component of the Cascade complex of a Type-I CRISPR/Cas system.
- the Cas nuclease may have an RNA cleavage activity.
- the RNA-guided DNA-binding agent has single-strand nickase activity, i.e., can cut one DNA strand to produce a single-strand break, also known as a "nick.”
- the RNA-guided DNA-bmding agent comprises a Cas nickase.
- a nickase is an enzyme that creates a stick in dsDNA, i.e., cuts one strand but not the other of the DNA double helix.
- a Cas nickase is a version of a Cas nuclease (e.g., a Cas nuclease discussed above) in which an endonucleolytic active site is inactivated, e.g., by one or more alterations (e.g., point mutations) in a catalytic domain. See, e.g., U.S. Pat. No. 8,889,356 for discussion of Cas nickases and exemplary catalytic domain alterations.
- a Cas nickase such as a Cas9 nickase has an inactivated RuvC or HNH domain.
- the RNA-guided DNA- binding agent lacks cleavase and nickase activity.
- the RNA-guided DNA- binding agent comprises a dCas DNA-binding polypeptide.
- a dCas polypeptide has DNA-binding activity while essentially lacking catalytic (cleavase/nickase) activity.
- the dCas polypeptide is a dCas9 polypeptide.
- the RNA-guided DNA-binding agent lacking cleavase and nickase activity or the dCas DNA-binding polypeptide is a version of a Cas nuclease (e.g., a Cas nuclease discussed above) in which its endonucleolytic active sites are inactivated, e.g., by one or more alterations (e.g., point mutations) in its catalytic domains. See, e.g., U.S. 2014/0186958 A1 ; U.S. 2015/0166980 A1 .
- alterations e.g., point mutations
- the RNA-guided DNA-binding agent comprises one or more heterologous functional domains (e.g., is or comprises a fusion polypeptide).
- the heterologous functional domain may be an effector domain.
- the effector domain may modify or affect the target sequence,
- the effector domain may be chosen from a nucleic acid binding domain, a nuclease domain (e.g., a non- Cas nuclease domain), an epigenetic modification domain, a transcriptional activation domain, or a transcriptional repressor domain.
- the heterologous functional domain is a nuclease, such as a Fokl nuclease. See, e.g., U.S. Pat. No. 9,023,649.
- the heterologous functional domain is a transcriptional activator or repressor.
- a transcriptional activator or repressor See, e.g., Qi et al., "Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression," Cell 152:1173-83 (2013); Perez-Pinera et al., "RNA-guided gene activation by CRISPR-Cas9-based transcription factors," Nat. Methods 10:973-6 (2013); Mali et al., “CASO transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering," Nat. Biotechnol.
- the RNA-guided DNA-binding agent essentially becomes a transcription factor that can be directed to bind a desired target sequence using a guide RNA.
- the DNA modification domain is a methylation domain, such as a demethylation or methyltransferase domain.
- the effector domain is a DNA modification domain, such as a base-editing domain.
- the DNA modification domain is a nucleic acid editing domain that introduces a specific modification into the DNA, such as a deaminase domain. See, e.g., WO 2015/089406; U.S.
- the nuclease may comprise at least one domain that interacts with a guide RNA ("gRNA"). Additionally, the nuclease may be directed to a target sequence by a gRNA. In Class 2 Cas nuclease systems, the gRNA interacts with the nuclease as well as the target sequence, such that it directs binding to the target sequence. In some embodiments, the gRNA provides the specificity for the targeted cleavage, and the nuclease may be universal and paired with different gRNAs to cleave different target sequences. In some embodiments of the present disclosure, the cargo for the LNMP formulation includes at least one gRNA.
- the gRNA may guide the Cas nuclease or Class 2 Cas nuclease to a target sequence on a target nucleic acid molecule.
- a gRNA binds with and provides specificity of cleavage by a Class 2 Cas nuclease.
- the gRNA and the Cas nuclease may form a ribonucleoprotein (RNP), e.g., a CRISPR/Cas complex such as a CRISPR/Cas9 complex which may be delivered by the LNMP composition.
- the CRISPR/Cas complex may be a Type- II CRISPR/Cas9 complex.
- the CRISPR/Cas complex may be a Type-V CRISPR/Cas complex, such as a Cpf1/guide RNA complex.
- Cas nucleases and cognate gRNAs may be paired.
- the gRNA scaffold structures that pair with each Class 2 Cas nuclease vary with the specific CRISPR/Cas system.
- RNA Ribonucleic acid
- gRNA gRNA
- tracrRNA RNA trRNA
- the crRNA and trRNA may be associated as a single RNA molecule (single guide RNA, sgRNA) or in two separate RNA molecules (dual guide RNA, dgRNA).
- sgRNA single guide RNA
- dgRNA dual guide RNA
- trRNA gRNA
- the trRNA may be a naturally-occurring sequence, or a CrRNA sequence with modifications or variations compared to naturally-occurring sequences.
- a "guide sequence” refers to a sequence within a guide RNA that is complementary to a target sequence and functions to direct a guide RNA to a target sequence for binding or modification (e.g., cleavage) by an RNA-guided DNA binding agent.
- a “guide sequence” may also be referred to as a "targeting sequence,” or a "spacer sequence.”
- a guide sequence can be 20 base pairs in length, e.g., in the case of Streptococcus pyogenes (i.e., Spy Cas9) and related Cas9 homologs/orthologs.
- shorter or longer sequences can also be used as guides, e.g., 15-, 16-, 17-, 18-, 19-, 21-, 22-, 23-, 24-, or 25- nucleotides in length.
- the target sequence is in a gene or on a chromosome, for example, and is complementary to the guide sequence.
- the degree of complementarity or identity between a guide sequence and its corresponding target sequence may be about 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 100%.
- the guide sequence and the target region may be 100% complementary or identical. In other embodiments, the guide sequence and the target region may contain at least one mismatch.
- the guide sequence and the target sequence may contain 1 , 2, 3, or 4 mismatches, where the total length of the target sequence is at least 17, 18, 19, 20 or more base pairs.
- the guide sequence and the target region may contain 1-4 mismatches where the guide sequence comprises at least 17, 18, 19, 20 or more nucleotides.
- the guide sequence and the target region may contain 1 , 2, 3, or 4 mismatches where the guide sequence comprises 20 nucleotides.
- Target sequences for Cas proteins include both the positive and negative strands of genomic DNA (i.e., the sequence given and the sequence's reverse compliment), as a nucleic acid substrate for a Cas protein is a double stranded nucleic acid.
- a guide sequence may direct a guide RNA to bind to the reverse complement of a target sequence.
- the guide sequence is identical to certain nucleotides of the target sequence (e.g., the target sequence not including the PAM) except for the substitution of U for T in the guide sequence.
- the length of the targeting sequence may depend on the CRISPR/Cas system and components used. For example, different Class 2 Cas nucleases from different bacterial species have varying optimal targeting sequence lengths.
- the targeting sequence may comprise 5, 6, 7, 8, 9, 10, 11 , 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 ,22, 23, 24, 25, 26, 27, 28, 29, 30, 35, 40, 45, 50, or more than 50 nucleotides in length.
- the targeting sequence length is 0, 1 , 2, 3, 4, or 5 nucleotides longer or shorter than the guide sequence of a naturally-occurring CRISPR/Cas system.
- the Cas nuclease and gRNA scaffold will be derived from the same CRISPR/Cas system.
- the targeting sequence may comprise or consist of 18-24 nucleotides.
- the targeting sequence may comprise or consist of 19-21 nucleotides.
- the targeting sequence may comprise or consist of 20 nucleotides.
- the sgRNA is a "Cas9 sgRNA" capable of mediating RNA-guided DNA cleavage by a Cas9 protein.
- the gRNA comprises a crRNA and tracr RNA sufficient for forming an active complex with a Cas9 protein and mediating RNA-guided DNA cleavage.
- the gRNA comprises a crRNA sufficient for forming an active complex with a Cpf1 protein and mediating RNA-guided DNA cleavage. See Zetsche 2015. Certain embodiments of the invention also provide nucleic acids, e.g., expression cassettes, encoding the gRNA described herein.
- guide RNA nucleic acid is used herein to refer to a guide UNA (e.g. an sgRNA or a dgRNA) and a guide RNA expression cassette, which is a nucleic acid that encodes one or more guide RNAs.
- a guide UNA e.g. an sgRNA or a dgRNA
- a guide RNA expression cassette which is a nucleic acid that encodes one or more guide RNAs.
- the nucleic acid may comprise a nucleotide sequence encoding a crRNA.
- the nucleotide sequence encoding the crRNA comprises a targeting sequence flanked by all or a portion of a repeat sequence from a naturally-occurring CRISPR/Cas system.
- the nucleic acid may comprise a nucleotide sequence encoding a tracr RNA.
- the crRNA and the tracr RNA may be encoded by two separate nucleic acids. In other embodiments, the crRNA and the tracr RNA may be encoded by a single nucleic acid.
- the crRNA and the tracr RNA may be encoded by opposite strands of a single nucleic acid. In other embodiments, the crRNA and the tracr RNA may be encoded by the same strand of a single nucleic acid.
- the gRNA nucleic acid encodes an sgRNA. In some embodiments, the gRNA nucleic acid encodes a Cas9 nuclease sgRNA. In come embodiments, the gRNA nucleic acid encodes a Cpf1 nuclease sgRNA.
- the nucleotide sequence encoding the guide RNA may be operabiy linked to at least one transcriptional or regulatory control sequence, such as a promoter, a 3' UTR, or a 5' UTR.
- the promoter may be a tRNA promoter, e.g., tRNA L v s3 , or a tRNA chimera. See Mefferd et al., RNA. 201521 :1683-9; Scherer et al., Nucleic Acids Res. 2007 35: 2620-2628.
- the promoter may be recognized by RNA polymerase III (Pol III).
- Non-limiting examples of Pol III promoters also include U6 and H1 promoters.
- the nucleotide sequence encoding the guide RNA may be operably linked to a mouse or human U6 promoter.
- the gRNA nucleic acid is a modified nucleic acid.
- the gRNA nucleic acid includes a modified nucleoside or nucleotide.
- the gRNA nucleic acid includes a 5' end modification, for example a modified nucleoside or nucleotide io stabilize and prevent integration of the nucleic acid.
- the gRNA nucleic acid comprises a double-stranded DNA having a 5' end modification on each strand.
- the gRNA nucleic acid includes an inverted dideoxy-T or an inverted abasic nucleoside or nucleotide as the 5' end modification.
- gRNA nucleic acid includes a label such as biotin, desthiobioten-TEG, digoxigenin, and fluorescent markers, including, for example, FAM, ROX, TAMRA, and AlexaFluor.
- more than one gRNA nucleic acid, such as a gRNA can be used with a CRISPR/Cas nuclease system.
- Each gRNA nucleic acid may contain a different targeting sequence, such that the CRISPR/Cas system cleaves more than one target sequence.
- one or more gRNAs may have the same or differing properties such as activity or stability within a CRISPR/Cas complex. Where more than one gRNA is used, each gRNA can be encoded on the same or on different gRNA nucleic acid. The promoters used to drive expression of the more than one gRNA may be the same or different.
- the RNA composition or formulation disclosed herein comprises an mRNA comprising an open reading frame (ORF) encoding an RNA-guided DNA binding agent, such as a Cas nuclease, or Class 2 Cas nuclease as described herein.
- an mRNA comprising an ORF encoding an RNA-guided DNA binding agent, such as a Cas nuclease or Class 2 Cas nuclease is provided, used, or administered.
- the ORF encoding an RNA- guided DNA binding agent is a "modified RNA-guided DNA binding agent ORF" or simply a "modified ORF,” which is used as shorthand to indicate that the ORF is modified in one or more of the following ways: (1) the modified ORF has a uridine content ranging from its minimum uridine content to 150% of the minimum uridine content; (2) the modified ORF has a uridine dinucleotide content ranging from its minimum uridine dinucleotide content to 150% of the minimum uridine dinucleotide content; (3) the modified ORF consists of a set of codons of which at least 75% of the codons are minimal uridine codon(s) for a given amino acid, e.g.
- the modified ORF comprises at least one modified uridine.
- the modified ORF is modified in at least two, three, or four of the foregoing ways.
- the modified ORF comprises at least one modified uridine and is modified in at least one, two, three, or all of (1)-(3) above.
- Modified uridine is used herein to refer to a nucleoside other than thymidine with the same hydrogen bond acceptors as uridine and one or more structural differences from uridine.
- a modified uridine is a substituted uridine, i.e., a uridine in which one or more non-proton substituents (e.g., alkoxy, such as methoxy) takes the place of a proton.
- a modified uridine is pseudouridine.
- a modified uridine is a substituted pseudouridine, i.e., a pseudouridine in which one or more non-proton substituents (e.g., alkyl, such as methyl) takes the place of a proton.
- a modified uridine is any of a substituted uridine, pseudouridine, or a substituted pseudouridine.
- Uridine position refers to a position in a polynucleotide occupied by a uridine or a modified uridine.
- a polynucleotide in which "100% of the uridine positions are modified uridines" contains a modified uridine at every position that would be a uridine in a conventional RNA (where all bases are standard A, U, C, or G bases) of the same sequence.
- a U in a polynucleotide sequence of a sequence listing in, or accompanying, this disclosure can be a uridine or a modified uridine.
- a cap can be included co-transcriptionally.
- ARCA anti-reverse cap analog; Thermo Fisher Scientific Cat. No. AM8045
- ARCA is a cap analog comprising a 7- methylguanine 3'- methoxy-5'-triphosphate linked to the 5' position of a guanine ribonucleotide which can be incorporated in vitro into a transcript at initiation.
- ARCA results in a CapO cap in which the 2' position of the first cap-proximal nucleotide is hydroxy).
- RNA 7: 1486-1495 See, e.g., Stepinski et al., (2001) "Synthesis and properties of mRNAs containing the novel 'anti- reverse' cap analogs 7-methyl(3'-O-methyl)GpppG and 7-methyl(3'deoxy)GpppG," RNA 7: 1486-1495.
- CleanCapTM AG (m7G(5')ppp(5')(2'OMeA)pG; TriLink Biotechnologies Cat. No. N-7113) or CleanCapTM GG (m7G(5')ppp(5')(2'OMeG)pG; TriLink Biotechnologies Cat. No. N-7133) can be used to provide a Cap1 structure co-transcriptionally.
- 3'-O-methylated versions of CleanCapTM AG and CleanCapTM GG are also available from TriLink Biotechnologies as Cat. Nos. N-7413 and N-7433, respectively.
- the LNMPs described herein may include gene editing composition sequences as described in WO2019067992, WO2019090173, WO2019090175, WO2022159741 , WO2022150608, WO2016201155, WO2022086846, WO2015153789, WO2017165826, WO2016154596, WO2018209158, WG2021050512, WO2021113494, WO2022229851 , WO2020264254, each of which is incorporated by reference herein in its entirety.
- the LNP I RNA composition or LNMP I RNA composition comprises one or more polynucleotides (e.g., mRNA, gRNA) encoding one or more gene editing systems for multiple purposes (e.g. therapeutic).
- the one or more polynucleotides e.g., mRNA, gRNA
- identity refers to a relationship between the sequences of two or more polypeptides (e.g. antigens, anti-cancer proteins or signaling proteins) or polynucleotides (nucleic acids), as determined by comparing the sequences. Identity also refers to the degree of sequence relatedness between or among sequences as determined by the number of matches between strings of two or more amino acid residues or nucleic acid residues. Identity measures the percent of identical matches between the smaller of two or more sequences with gap alignments (if any) addressed by a particular mathematical model or computer program (e.g., “algorithms”). Identity of related antigens, proteins, or nucleic acids can be readily calculated by known methods.
- Percent (%) identity as it applies to polypeptide or polynucleotide sequences is defined as the percentage of residues (amino acid residues or nucleic acid residues) in the candidate amino acid or nucleic acid sequence that are identical with the residues in the amino acid sequence or nucleic acid sequence of a second sequence after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent identity. Methods and computer programs for the alignment are well known in the art. It is understood that identity depends on a calculation of percent identity but may differ in value due to gaps and penalties introduced in the calculation.
- variants of a particular polynucleotide or polypeptide have at least 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% but less than 100% sequence identity to that particular reference polynucleotide or polypeptide as determined by sequence alignment programs and parameters described herein and known to those skilled in the art.
- Such tools for alignment include those of the BLAST suite (Stephen F.
- sequence tags or amino acids such as one or more lysines
- Sequence tags can be used for peptide detection, purification, or localization.
- Lysines can be used to increase peptide solubility or to allow for biotinylation.
- amino acid residues located at the carboxy and amino terminal regions of the amino acid sequence of a peptide or protein may optionally be deleted providing for truncated sequences.
- Certain amino acids e.g., C-terminal or N-terminal residues
- sequences for (or encoding) signal sequences, termination sequences, transmembrane domains, linkers, multimerization domains (such as, e.g., foldon regions) and the like may be substituted with alternative sequences that achieve the same or a similar function.
- cavities in the core of proteins can be filled to improve stability, e.g., by introducing larger amino acids.
- buried hydrogen bond networks may be replaced with hydrophobic resides to improve stability.
- glycosylation sites may be removed and replaced with appropriate residues.
- sequences are readily identifiable to one of skill in the art. It should also be understood that some of the sequences provided herein contain sequence tags or terminal peptide sequences (e.g., at the N-terminal or C-terminal ends) that may be deleted, for example, prior to use in the preparation of an RNA (e.g., mRNA) vaccine.
- RNA e.g., mRNA
- the polynucleotide is an mRNA.
- Messenger RNA is any RNA that encodes a (at least one) protein (a naturally- occurring, non-naturally-occurring, or modified polymer of amino acids) and can be translated to produce the encoded protein in vitro, in vivo, in situ, or ex vivo.
- mRNA messenger RNA
- nucleic acid sequences set forth in the instant application may recite “T”s in a representative DNA sequence but where the sequence represents RNA (e.g., mRNA), the “T”s would be substituted for “U”s.
- any of the DNAs disclosed and identified by a particular sequence identification number herein also disclose the corresponding RNA (e.g., mRNA) sequence complementary to the DNA, where each “T” of the DNA sequence is substituted with “U.”
- the polynucleotide e.g., mRNA
- the polynucleotide e.g., mRNA
- An open reading frame (ORF) is a continuous stretch of DNA or RNA beginning with a start codon (e.g., methionine (ATG or AUG)) and ending with a stop codon (e.g., TAA, TAG or TGA, or UAA, UAG or UGA).
- An ORF typically encodes a protein.
- the sequences may further comprise additional elements, e.g., 5' and 3' UTRs.
- the RNA (e.g., mRNA) further comprises a 5' UTR, 3' UTR, a poly(A) tail and/or a 5' cap analog.
- the mRNA comprises a 5' untranslated region (UTR) and/or a 3' UTR.
- the mRNA is derived from (a) a DNA molecule; or (b) an RNA molecule. In the mRNA, T is optionally substituted with U.
- the mRNA is derived from a DNA molecule.
- the DNA molecule can further comprise a promoter.
- the promoter is a T7 promoter, a T3 promoter, or an SP6 promoter.
- the promoter is located at the 5’ UTR.
- the mRNA is an RNA molecule.
- the RNA molecule may be a selfreplicating RNA molecule.
- the mRNA is an RNA molecule.
- the RNA molecule may further comprise a 5’ cap.
- the 5’ cap can have a Cap 1 structure, a Cap 1 (m6A) structure, a Cap 2 structure, a Cap 3 structure, a Cap 0 structure, or any combination thereof.
- the polynucleotide is an mRNA which encodes an IL-2 molecule.
- the IL-2 molecule comprises a naturally occurring IL-2 molecule, a fragment of a naturally occurring IL-2 molecule, or a variant thereof.
- the IL-2 molecule comprises a variant of a naturally occurring IL-2 molecule (e.g., an IL-2 variant, e.g., as described herein), or a fragment thereof.
- the polynucleotide is an mRNA which encodes an IL-2 molecule comprising an amino acid sequence having at least 85%, 90%, 95%, 96%, 97%, 98%, 99% or 100% identity to the amino acid sequence of an IL-2 molecule provided in any one of Tables l-lll.
- the mRNA comprises a 5' untranslated region (UTR) and/or a 3' UTR.
- the mRNA comprises a 5' UTR.
- the 5' UTR may comprise a Kozak sequence.
- the mRNA comprises a 3' UTR.
- the 3’ UTR comprises one or more sequences derived from an amino-terminal enhancer of split (AES).
- the 3’ UTR comprises a sequence derived from mitochondrially encoded 12S rRNA (mtRNRI).
- the mRNA comprises a poly(A) sequence.
- the poly(A) sequence is a 110-nucleotide sequence consisting of a sequence of 30 adenosine residues, a 10-nucleotide linker sequence, and a sequence of 70 adenosine residues.
- Naturally occurring eukaryotic mRNA molecules can contain stabilizing elements, including, but not limited to untranslated regions (UTR) at their 5 '-end (5' UTR) and/or at their 3 '-end (3' UTR), in addition to other structural features, such as a 5 '-cap structure or a 3'-poly(A) tail.
- UTR untranslated regions
- Both the 5' UTR and the 3' UTR are typically transcribed from the genomic DNA and are elements of the premature mRNA. Characteristic structural features of mature mRNA, such as the 5 '-cap and the 3'-poly(A) tail are usually added to the transcribed (premature) mRNA during mRNA processing.
- the polynucleotide has an open reading frame encoding at least one gene editing system having at least one modification, at least one 5' terminal cap, and is formulated within a lipid nanoparticle.
- 5 '-capping of polynucleotides may be completed concomitantly during the in vitro-transcription reaction using the following chemical RNA cap analogs to generate the 5'- guanosine cap structure according to manufacturer protocols: 3'-O-Me-m7G(5')ppp(5') G [the ARCA cap];G(5')ppp(5')A; G(5')ppp(5')G; m7G(5')ppp(5')A; m7G(5')ppp(5')G (New England BioLabs, Ipswich, MA).
- 5'- capping of modified RNA may be completed post-transcriptionally using a Vaccinia Vims Capping Enzyme to generate the “Cap 0” structure: m7G(5')ppp(5')G (New England BioLabs, Ipswich, MA).
- Cap 1 structure may be generated using both Vaccinia Vims Capping Enzyme and a 2'- 0 methyl-transferase to generate m7G(5')ppp(5')G-2 '-O-methyl.
- Cap 2 structure may be generated from the Cap 1 stmcture followed by the 2'-0-methylation of the 5'- antepenultimate nucleotide using a 2'-O methyl-transferase.
- Cap 3 structure may be generated from the Cap 2 stmcture followed by the 2'-0-methylation of the 5'-preantepenultimate nucleotide using a 2'-O methyl-transferase.
- Enzymes may be derived from a recombinant source.
- the 3 '-poly(A) tail is typically a stretch of adenine nucleotides added to the 3 '-end of the transcribed mRNA. It can, in some instances, comprise up to about 400 adenine nucleotides. In some embodiments, the length of the 3'-poly(A) tail may be an essential element with respect to the stability of the individual mRNA.
- the polynucleotide includes a stabilizing element. Stabilizing elements may include for instance a histone stem-loop. A stem-loop binding protein (SLBP), a 32 kDa protein has been identified.
- SLBP stem-loop binding protein
- SLBP RNA binding domain of SLBP is conserved through metazoa and protozoa; its binding to the histone stem-loop depends on the structure of the loop.
- the minimum binding site includes at least three nucleotides 5' and two nucleotides 3' relative to the stem- loop.
- the polynucleotide (e.g., mRNA) includes a coding region, at least one histone stem-loop, and optionally, a poly(A) sequence or polyadenylation signal.
- the poly(A) sequence or polyadenylation signal generally should enhance the expression level of the encoded protein.
- the encoded protein in some embodiments, is not a histone protein, a reporter protein (e.g. Luciferase, GFP, EGFP, b-Galactosidase, EGFP), or a marker or selection protein (e.g. alpha-Globin, Galactokinase and Xanthine:guanine phosphoribosyl transferase (GPT)).
- a reporter protein e.g. Luciferase, GFP, EGFP, b-Galactosidase, EGFP
- a marker or selection protein e.g. alpha-Globin, Galactokina
- the polynucleotide (e.g., mRNA) includes the combination of a poly(A) sequence or polyadenylation signal and at least one histone stem-loop, even though both represent alternative mechanisms in nature, acts synergistically to increase the protein expression beyond the level observed with either of the individual elements.
- the synergistic effect of the combination of poly(A) and at least one histone stem-loop does not depend on the order of the elements or the length of the poly(A) sequence.
- an RNA (e.g., mRNA) does not include a histone downstream element (HDE).
- Histone downstream element includes a purine-rich polynucleotide stretch of approximately 15 to 20 nucleotides 3' of naturally occurring stem-loops, representing the binding site for the U7 snRNA, which is involved in processing of histone pre-mRNA into mature histone mRNA.
- the nucleic acid does not include an intron.
- the polynucleotide may or may not contain an enhancer and/or promoter sequence, which may be modified or unmodified or which may be activated or inactivated.
- the histone stem-loop is generally derived from histone genes and includes an intramolecular base pairing of two neighbored partially or entirely reverse complementary sequences separated by a spacer, consisting of a short sequence, which forms the loop of the structure.
- the unpaired loop region is typically unable to base pair with either of the stem loop elements. It occurs more often in RNA, as is a key component of many RNA secondary structures but may be present in single- stranded DNA as well.
- the Stability of the stem-loop structure generally depends on the length, number of mismatches or bulges, and base composition of the paired region.
- wobble base pairing non-Watson-Crick base pairing
- the at least one histone stem-loop sequence comprises a length of 15 to 45 nucleotides.
- the polynucleotide e.g., mRNA
- the polynucleotide has one or more AU-rich sequences removed. These sequences, sometimes referred to as AURES are destabilizing sequences found in the 3'UTR.
- the AURES may be removed from the RNA vaccines. Alternatively, the AURES may remain in the RNA vaccine.
- the polynucleotide e.g., mRNA
- the polynucleotide has an ORF that encodes a signal peptide fused to the antigen.
- Signal peptides comprising the N- terminal 15-60 amino acids of proteins, are typically needed for the translocation across the membrane on the secretory pathway and, thus, universally control the entry of most proteins both in eukaryotes and in prokaryotes to the secretory pathway.
- the signal peptide of a nascent precursor protein pre-protein
- ER endoplasmic reticulum
- ER processing produces mature proteins, wherein the signal peptide is cleaved from precursor proteins, typically by an ER-resident signal peptidase of the host cell, or they remain uncleaved and function as a membrane anchor.
- a signal peptide may also facilitate the targeting of the protein to the cell membrane.
- a signal peptide may have a length of 15-60 amino acids.
- a signal peptide may have a length of 15, 16, 17, 18, 19, 20, 21 , 22, 23, 24, 25, 26, 27, 28, 29, 30, 31 , 32, 33, 34, 35, 36, 37, 38, 39, 40, 41 , 42, 43, 44, 45, 46, 47, 48, 49, 50, 51 , 52, 53, 54, 55, 56, 57, 58, 59, or 60 amino acids.
- a signal peptide has a length of 20-60, 25-60, 30-60, 35- 60, 40-60, 45- 60, 50-60, 55-60, 15-55, 20-55, 25-55, 30-55, 35- 55, 40-55, 45-55, 50-55, 15-50, 20-50, 25-50, 30-50, 35-50, 40-50, 45-50, 15-45, 20-45, 25-45, 30-45, 35-45, 40-45, 15-40, 20- 40, 25-40, 30-40, 35-40, 15-35, 20-35, 25-35, 30-35, 15-30, 20-30, 25-30, 15-25, 20-25, or 15-20 amino acids.
- signal peptides from heterologous genes are known in the art and can be tested for desired properties and then incorporated into a nucleic acid of the disclosure.
- the signal peptide may comprise those described in WO 2021/154763, which is incorporated by reference in its entirety.
- the polynucleotide encodes an antigenic fusion protein.
- the encoded antigen or antigens may include two or more proteins (e.g., protein and/or protein fragment) joined together.
- the protein to which a protein antigen is fused does not promote a strong immune response to itself, but rather to the coronavirus antigen.
- Antigenic fusion proteins retain the functional property from each original protein.
- the polynucleotide in some embodiments, encodes fusion proteins that comprise coronavirus antigens linked to scaffold moieties.
- such scaffold moieties impart desired properties to an antigen encoded by a nucleic acid of the disclosure.
- scaffold proteins may improve the immunogenicity of an antigen, e.g., by altering the structure of the antigen, altering the uptake and processing of the antigen, and/or causing the antigen to bind to a binding partner.
- the scaffold moiety is protein that can self-assemble into protein nanoparticles that are highly symmetric, stable, and structurally organized, with diameters of 10- 150 nm, a highly suitable size range for optimal interactions with various cells of the immune system.
- bacterial protein platforms may be used. Non-limiting examples of these self-assembling proteins include ferritin, lumazine and encapsulin.
- Ferritin is a protein whose main function is intracellular iron storage. Ferritin is made of 24 subunits, each composed of a four-alpha-helix bundle, that self-assemble in a quaternary structure with octahedral symmetry (Cho K.J. et al. J Mol Biol. 2009; 390:83-98). Several high- resolution structures of ferritin have been determined, confirming that Helicobacter pylori ferritin is made of 24 identical protomers, whereas in animals, there are ferritin light and heavy chains that can assemble alone or combine with different ratios into particles of 24 subunits (Granier T. et al. J Biol Inorg Chem.
- Ferritin self-assembles into nanoparticles with robust thermal and chemical stability.
- the ferritin nanoparticle is well suited to carry and expose antigens.
- Fumazine synthase is also well suited as a nanoparticle platform for antigen display.
- FS which is responsible for the penultimate catalytic step in the biosynthesis of riboflavin, is an enzyme present in a broad variety of organisms, including archaea, bacteria, fungi, plants, and eubacteria (Weber S.E. Flavins and Flavoproteins. Methods and Protocols, Series: Methods in Molecular Biology. 2014).
- the FS monomer is 150 amino acids long, and consists of beta- sheets along with tandem alpha-helices flanking its sides.
- Encapsulin a novel protein cage nanoparticle isolated from thermophile Thermotoga maritima, may also be used as a platform to present antigens on the surface of self-assembling nanoparticles.
- the polynucleotide encodes a coronavirus antigen (e.g., SARS-CoV-2 S protein) fused to a foldon domain.
- the foldon domain may be, for example, obtained from bacteriophage T4 fibritin (see, e.g., Tao Y, et al. Structure. 1997 Jun 15; 5(6):789-98).
- the polynucleotide encodes more than one polypeptide, referred to herein as fusion proteins.
- the mRNA further encodes a linker located between at least one or each domain of the fusion protein.
- the linker can be, for example, a cleavable linker or protease-sensitive linker.
- the linker is selected from the group consisting of F2A linker, P2A linker, T2A linker, E2A linker, and combinations thereof. This family of self-cleaving peptide linkers, referred to as 2 A peptides, has been described in the art (see for example, Kim, J.H.
- the linker is an F2A linker.
- the fusion protein contains three domains with intervening linkers, having the structure: domain-linker-domain-linker-domain.
- Cleavable linkers known in the art may be used in connection with the disclosure.
- Exemplary such linkers include F2A linkers, T2A linkers, P2A linkers, E2A linkers (See, e.g., WO2017/127750, which is incorporated herein by reference in its entirety).
- linkers include F2A linkers, T2A linkers, P2A linkers, E2A linkers (See, e.g., WO2017/127750, which is incorporated herein by reference in its entirety).
- linkers may be suitable for use in the constructs of the disclosure (e.g., encoded by the nucleic acids of the disclosure).
- polycistronic constructs mRNA encoding more than one antigen/polypeptide separately within the same molecule may be suitable for use as provided herein.
- an ORF encoding one or more polypeptides of the disclosure is codon optimized. Codon optimization methods are known in the art. For example, an ORF of any one or more of the sequences provided herein may be codon optimized.
- Codon optimization may be used to match codon frequencies in target and host organisms to ensure proper folding; bias GC content to increase mRNA stability or reduce secondary structures; minimize tandem repeat codons or base runs that may impair gene construction or expression; customize transcriptional and translational control regions; insert or remove protein trafficking sequences; remove/add post translation modification sites in encoded protein (e.g., glycosylation sites); add, remove or shuffle protein domains; insert or delete restriction sites; modify ribosome binding sites and mRNA degradation sites; adjust translational rates to allow the various domains of the protein to fold properly; or reduce or eliminate problem secondary structures within the polynucleotide.
- encoded protein e.g., glycosylation sites
- add, remove or shuffle protein domains add or delete restriction sites
- modify ribosome binding sites and mRNA degradation sites adjust translational rates to allow the various domains of the protein to fold properly; or reduce or eliminate problem secondary structures within the polynucleotide.
- Codon optimization tools, algorithms and services are known in the art - non-limiting examples include services from GeneArt (Life Technologies), DNA2.0 (Menlo Park CA) and/or proprietary methods.
- the open reading frame (ORF) sequence is optimized using optimization algorithms.
- a codon optimized sequence shares less than 95% sequence identity to a naturally-occurring or wild-type sequence ORF (e.g., a naturally-occurring or wild- type mRNA sequence encoding a coronavirus antigen). In some embodiments, a codon optimized sequence shares less than 90% sequence identity to a naturally occurring or wild-type sequence (e.g., a naturally occurring or wild-type mRNA sequence encoding a coronavirus antigen). In some embodiments, a codon optimized sequence shares less than 85% sequence identity to a naturally occurring or wild-type sequence (e.g., a naturally occurring or wild-type mRNA sequence encoding a coronavirus antigen).
- a codon optimized sequence shares less than 80% sequence identity to a naturally occurring or wild-type sequence (e.g., a naturally occurring or wildtype mRNA sequence encoding a coronavirus antigen). In some embodiments, a codon optimized sequence shares less than 75% sequence identity to a naturally occurring or wild-type sequence (e.g., a naturally occurring or wild-type mRNA sequence encoding a coronavirus antigen).
- a codon optimized sequence shares between 65% and 85% (e.g., between about 67% and about 85% or between about 67% and about 80%) sequence identity to a naturally occurring or wild-type sequence (e.g., a naturally occurring or wild-type mRNA sequence encoding a coronavirus antigen). In some embodiments, a codon optimized sequence shares between 65% and 75% or about 80% sequence identity to a naturally occurring or wild- type sequence (e.g., a naturally occurring or wild-type mRNA sequence encoding a coronavirus antigen).
- a codon-optimized sequence encodes an antigen that is as immunogenic as, or more immunogenic than (e.g., at least 10%, at least 20%, at least 30%, at least 40%, at least 50%, at least 100%, or at least 200% more), a coronavirus antigen encoded by a non- codon-optimized sequence.
- the modified mRNAs When transfected into mammalian host cells, the modified mRNAs have a stability of between 12-18 hours, or greater than 18 hours, e.g., 24, 36, 48, 60, 72, or greater than 72 hours and are capable of being expressed by the mammalian host cells.
- a codon optimized RNA may be one in which the levels of G/C are enhanced.
- the G/C-content of nucleic acid molecules may influence the stability of the RNA.
- RNA having an increased amount of guanine (G) and/or cytosine (C) residues may be functionally more stable than RNA containing a large amount of adenine (A) and thymine (T) or uracil (U) nucleotides.
- WO02/098443 discloses a pharmaceutical composition containing an mRNA stabilized by sequence modifications in the translated region. Due to the degeneracy of the genetic code, the modifications work by substituting existing codons for those that promote greater RNA stability without changing the resulting amino acid. The approach is limited to coding regions of the RNA.
- the polynucleotide e.g., mRNA
- the polynucleotide is not chemically modified and comprises the standard ribonucleotides consisting of adenosine, guanosine, cytosine and uridine.
- nucleotides and nucleosides of the polynucleotide comprise standard nucleoside residues such as those present in transcribed RNA (e.g. A, G, C, or U).
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Genetics & Genomics (AREA)
- Chemical & Material Sciences (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Wood Science & Technology (AREA)
- Zoology (AREA)
- Biomedical Technology (AREA)
- Biotechnology (AREA)
- General Engineering & Computer Science (AREA)
- Molecular Biology (AREA)
- Microbiology (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- Medicinal Chemistry (AREA)
- Physics & Mathematics (AREA)
- Biophysics (AREA)
- Plant Pathology (AREA)
- Pharmaceuticals Containing Other Organic And Inorganic Compounds (AREA)
- Medicinal Preparation (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
Abstract
Description
Claims
Priority Applications (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202380084391.7A CN120359023A (en) | 2022-11-10 | 2023-11-09 | RNA compositions comprising lipid nanoparticles or lipid reconstituted natural messenger packages |
| EP23821780.6A EP4615424A1 (en) | 2022-11-10 | 2023-11-09 | Rna compositions comprising lipid nanoparticles or lipid reconstructed natural messenger packs |
| JP2025526705A JP2025537750A (en) | 2022-11-10 | 2023-11-09 | RNA compositions containing lipid nanoparticles or lipid-reconstituted natural messenger packs |
Applications Claiming Priority (6)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202263424208P | 2022-11-10 | 2022-11-10 | |
| US63/424,208 | 2022-11-10 | ||
| US202363447494P | 2023-02-22 | 2023-02-22 | |
| US63/447,494 | 2023-02-22 | ||
| US202363527780P | 2023-07-19 | 2023-07-19 | |
| US63/527,780 | 2023-07-19 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024102434A1 true WO2024102434A1 (en) | 2024-05-16 |
Family
ID=89168273
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2023/037077 Ceased WO2024102434A1 (en) | 2022-11-10 | 2023-11-09 | Rna compositions comprising lipid nanoparticles or lipid reconstructed natural messenger packs |
Country Status (4)
| Country | Link |
|---|---|
| EP (1) | EP4615424A1 (en) |
| JP (1) | JP2025537750A (en) |
| CN (1) | CN120359023A (en) |
| WO (1) | WO2024102434A1 (en) |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2025042786A1 (en) | 2023-08-18 | 2025-02-27 | Flagship Pioneering Innovations Vi, Llc | Compositions comprising circular polyribonucleotides and uses thereof |
| WO2025106915A1 (en) | 2023-11-17 | 2025-05-22 | Sail Biomedicines, Inc. | Circular polyribonucleotides encoding glucagon-like peptide 1 (glp-1) and uses thereof |
| WO2025106930A1 (en) | 2023-11-17 | 2025-05-22 | Sail Biomedicines, Inc. | Circular polyribonucleotides encoding glucagon-like peptide 2 (glp-2) and uses thereof |
| WO2025179198A1 (en) | 2024-02-23 | 2025-08-28 | Sail Biomedicines, Inc. | Circular polyribonucleotides and unmodified linear rnas with reduced immunogenicity |
Citations (101)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5208036A (en) | 1985-01-07 | 1993-05-04 | Syntex (U.S.A.) Inc. | N-(ω, (ω-1)-dialkyloxy)- and N-(ω, (ω-1)-dialkenyloxy)-alk-1-yl-N,N,N-tetrasubstituted ammonium lipids and uses therefor |
| US5264618A (en) | 1990-04-19 | 1993-11-23 | Vical, Inc. | Cationic lipids for intracellular delivery of biologically active molecules |
| US5279833A (en) | 1990-04-04 | 1994-01-18 | Yale University | Liposomal transfection of nucleic acids into animal cells |
| US5283185A (en) | 1991-08-28 | 1994-02-01 | University Of Tennessee Research Corporation | Method for delivering nucleic acids into cells |
| WO1996010390A1 (en) | 1994-09-30 | 1996-04-11 | Inex Pharmaceuticals Corp. | Novel compositions for the introduction of polyanionic materials into cells |
| US5785992A (en) | 1994-09-30 | 1998-07-28 | Inex Pharmaceuticals Corp. | Compositions for the introduction of polyanionic materials into cells |
| WO2002098443A2 (en) | 2001-06-05 | 2002-12-12 | Curevac Gmbh | Stabilised mrna with an increased g/c content and optimised codon for use in gene therapy |
| US20060083780A1 (en) | 2004-06-07 | 2006-04-20 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods of use |
| US20060240554A1 (en) | 2005-02-14 | 2006-10-26 | Sirna Therapeutics, Inc. | Lipid nanoparticle based compositions and methods for the delivery of biologically active molecules |
| US20090226470A1 (en) | 2007-12-11 | 2009-09-10 | Mauro Vincent P | Compositions and methods related to mRNA translational enhancer elements |
| US20100129877A1 (en) | 2005-09-28 | 2010-05-27 | Ugur Sahin | Modification of RNA, Producing an Increased Transcript Stability and Translation Efficiency |
| US20100293625A1 (en) | 2007-09-26 | 2010-11-18 | Interexon Corporation | Synthetic 5'UTRs, Expression Vectors, and Methods for Increasing Transgene Expression |
| US8084599B2 (en) | 2004-03-15 | 2011-12-27 | City Of Hope | Methods and compositions for the specific inhibition of gene expression by double-stranded RNA |
| WO2012019168A2 (en) | 2010-08-06 | 2012-02-09 | Moderna Therapeutics, Inc. | Engineered nucleic acids and methods of use thereof |
| WO2012045075A1 (en) | 2010-10-01 | 2012-04-05 | Jason Schrum | Modified nucleosides, nucleotides, and nucleic acids, and uses thereof |
| US8278063B2 (en) | 2007-06-29 | 2012-10-02 | Commonwealth Scientific And Industrial Research Organisation | Methods for degrading toxic compounds |
| WO2012135805A2 (en) | 2011-03-31 | 2012-10-04 | modeRNA Therapeutics | Delivery and formulation of engineered nucleic acids |
| US8349809B2 (en) | 2008-12-18 | 2013-01-08 | Dicerna Pharmaceuticals, Inc. | Single stranded extended dicer substrate agents and methods for the specific inhibition of gene expression |
| WO2013052523A1 (en) | 2011-10-03 | 2013-04-11 | modeRNA Therapeutics | Modified nucleosides, nucleotides, and nucleic acids, and uses thereof |
| WO2013063468A1 (en) | 2011-10-27 | 2013-05-02 | Massachusetts Institute Of Technology | Amino acid derivates functionalized on the n- terminal capable of forming drug incapsulating microspheres |
| WO2013090648A1 (en) | 2011-12-16 | 2013-06-20 | modeRNA Therapeutics | Modified nucleoside, nucleotide, and nucleic acid compositions |
| US8513207B2 (en) | 2008-12-18 | 2013-08-20 | Dicerna Pharmaceuticals, Inc. | Extended dicer substrate agents and methods for the specific inhibition of gene expression |
| WO2013151670A2 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of nuclear proteins |
| WO2013151666A2 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of biologics and proteins associated with human disease |
| WO2013185069A1 (en) | 2012-06-08 | 2013-12-12 | Shire Human Genetic Therapies, Inc. | Pulmonary delivery of mrna to non-lung target cells |
| WO2014028429A2 (en) | 2012-08-14 | 2014-02-20 | Moderna Therapeutics, Inc. | Enzymes and polymerases for the synthesis of rna |
| US20140068797A1 (en) | 2012-05-25 | 2014-03-06 | University Of Vienna | Methods and compositions for rna-directed target dna modification and for rna-directed modulation of transcription |
| US8697359B1 (en) | 2012-12-12 | 2014-04-15 | The Broad Institute, Inc. | CRISPR-Cas systems and methods for altering expression of gene products |
| WO2014093924A1 (en) | 2012-12-13 | 2014-06-19 | Moderna Therapeutics, Inc. | Modified nucleic acid molecules and uses thereof |
| US20140186958A1 (en) | 2012-12-12 | 2014-07-03 | Feng Zhang | Engineering and optimization of systems, methods and compositions for sequence manipulation with functional domains |
| US20140206753A1 (en) | 2011-06-08 | 2014-07-24 | Shire Human Genetic Therapies, Inc. | Lipid nanoparticle compositions and methods for mrna delivery |
| US8795965B2 (en) | 2012-12-12 | 2014-08-05 | The Broad Institute, Inc. | CRISPR-Cas component systems, methods and compositions for sequence manipulation |
| WO2014144767A1 (en) | 2013-03-15 | 2014-09-18 | Moderna Therapeutics, Inc. | Ion exchange purification of mrna |
| WO2014144196A1 (en) | 2013-03-15 | 2014-09-18 | Shire Human Genetic Therapies, Inc. | Synergistic enhancement of the delivery of nucleic acids via blended formulations |
| WO2014152027A1 (en) | 2013-03-15 | 2014-09-25 | Moderna Therapeutics, Inc. | Manufacturing methods for production of rna transcripts |
| WO2014152030A1 (en) | 2013-03-15 | 2014-09-25 | Moderna Therapeutics, Inc. | Removal of dna fragments in mrna production process |
| WO2014152031A1 (en) | 2013-03-15 | 2014-09-25 | Moderna Therapeutics, Inc. | Ribonucleic acid purification |
| WO2014164253A1 (en) | 2013-03-09 | 2014-10-09 | Moderna Therapeutics, Inc. | Heterologous untranslated regions for mrna |
| US8865406B2 (en) | 2012-12-12 | 2014-10-21 | The Broad Institute Inc. | Engineering and optimization of improved systems, methods and enzyme compositions for sequence manipulation |
| US8889356B2 (en) | 2012-12-12 | 2014-11-18 | The Broad Institute Inc. | CRISPR-Cas nickase systems, methods and compositions for sequence manipulation in eukaryotes |
| US8906616B2 (en) | 2012-12-12 | 2014-12-09 | The Broad Institute Inc. | Engineering of systems, methods and optimized guide compositions for sequence manipulation |
| WO2015024667A1 (en) | 2013-08-21 | 2015-02-26 | Curevac Gmbh | Method for increasing expression of rna-encoded proteins |
| WO2015038892A1 (en) | 2013-09-13 | 2015-03-19 | Moderna Therapeutics, Inc. | Polynucleotide compositions containing amino acids |
| US9012219B2 (en) | 2005-08-23 | 2015-04-21 | The Trustees Of The University Of Pennsylvania | RNA preparations comprising purified modified RNA for reprogramming cells |
| US9023649B2 (en) | 2012-12-17 | 2015-05-05 | President And Fellows Of Harvard College | RNA-guided human genome engineering |
| WO2015062738A1 (en) | 2013-11-01 | 2015-05-07 | Curevac Gmbh | Modified rna with decreased immunostimulatory properties |
| WO2015089511A2 (en) | 2013-12-13 | 2015-06-18 | Moderna Therapeutics, Inc. | Modified nucleic acid molecules and uses thereof |
| US20150166980A1 (en) | 2013-12-12 | 2015-06-18 | President And Fellows Of Harvard College | Fusions of cas9 domains and nucleic acid-editing domains |
| WO2015095340A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2015101415A1 (en) | 2013-12-30 | 2015-07-09 | Curevac Gmbh | Artificial nucleic acid molecules |
| WO2015101414A2 (en) | 2013-12-30 | 2015-07-09 | Curevac Gmbh | Artificial nucleic acid molecules |
| WO2015153789A1 (en) | 2014-04-01 | 2015-10-08 | Editas Medicine, Inc. | Crispr/cas-related methods and compositions for treating herpes simplex virus type 1 (hsv-1) |
| US9200276B2 (en) | 2009-06-01 | 2015-12-01 | Halo-Bio Rnai Therapeutics, Inc. | Polynucleotides for multivalent RNA interference, compositions and methods of use thereof |
| US20150344912A1 (en) | 2012-10-23 | 2015-12-03 | Toolgen Incorporated | Composition for cleaving a target dna comprising a guide rna specific for the target dna and cas protein-encoding nucleic acid or cas protein, and use thereof |
| WO2015184256A2 (en) | 2014-05-30 | 2015-12-03 | Shire Human Genetic Therapies, Inc. | Biodegradable lipids for delivery of nucleic acids |
| WO2015196130A2 (en) | 2014-06-19 | 2015-12-23 | Moderna Therapeutics, Inc. | Alternative nucleic acid molecules and uses thereof |
| WO2015196118A1 (en) | 2014-06-19 | 2015-12-23 | Moderna Therapeutics, Inc. | Alternative nucleic acid molecules and uses thereof |
| WO2015196128A2 (en) | 2014-06-19 | 2015-12-23 | Moderna Therapeutics, Inc. | Alternative nucleic acid molecules and uses thereof |
| WO2015199952A1 (en) | 2014-06-25 | 2015-12-30 | Acuitas Therapeutics Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2016004202A1 (en) | 2014-07-02 | 2016-01-07 | Massachusetts Institute Of Technology | Polyamine-fatty acid derived lipidoids and uses thereof |
| WO2016011306A2 (en) | 2014-07-17 | 2016-01-21 | Moderna Therapeutics, Inc. | Terminal modifications of polynucleotides |
| US20160208243A1 (en) | 2015-06-18 | 2016-07-21 | The Broad Institute, Inc. | Novel crispr enzymes and systems |
| WO2016118725A1 (en) | 2015-01-23 | 2016-07-28 | Moderna Therapeutics, Inc. | Lipid nanoparticle compositions |
| WO2016118724A1 (en) | 2015-01-21 | 2016-07-28 | Moderna Therapeutics, Inc. | Lipid nanoparticle compositions |
| WO2016154596A1 (en) | 2015-03-25 | 2016-09-29 | Editas Medicine, Inc. | Crispr/cas-related methods, compositions and components |
| WO2016187531A1 (en) | 2015-05-21 | 2016-11-24 | Ohio State Innovation Foundation | Benzene-1,3,5-tricarboxamide derivatives and uses thereof |
| WO2016201155A1 (en) | 2015-06-10 | 2016-12-15 | Caribou Biosciences, Inc. | Crispr-cas compositions and methods |
| WO2016205691A1 (en) | 2015-06-19 | 2016-12-22 | Massachusetts Institute Of Technology | Alkenyl substituted 2,5-piperazinediones and their use in compositions for delivering an agent to a subject or cell |
| WO2017004143A1 (en) | 2015-06-29 | 2017-01-05 | Acuitas Therapeutics Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017049245A2 (en) | 2015-09-17 | 2017-03-23 | Modernatx, Inc. | Compounds and compositions for intracellular delivery of therapeutic agents |
| WO2017075531A1 (en) | 2015-10-28 | 2017-05-04 | Acuitas Therapeutics, Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017117528A1 (en) | 2015-12-30 | 2017-07-06 | Acuitas Therapeutics, Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017127750A1 (en) | 2016-01-22 | 2017-07-27 | Modernatx, Inc. | Messenger ribonucleic acids for the production of intracellular binding polypeptides and methods of use thereof |
| WO2017165826A1 (en) | 2016-03-25 | 2017-09-28 | Editas Medicine, Inc. | Genome editing systems comprising repair-modulating enzyme molecules and methods of their use |
| WO2017173054A1 (en) | 2016-03-30 | 2017-10-05 | Intellia Therapeutics, Inc. | Lipid nanoparticle formulations for crispr/cas components |
| WO2017176974A1 (en) | 2016-04-06 | 2017-10-12 | Ohio State Innovation Foundation | Biodegradable amino-ester nanomaterials for nucleic acid delivery |
| WO2018053209A1 (en) | 2016-09-14 | 2018-03-22 | Modernatx, Inc. | High purity rna compositions and methods for preparation thereof |
| WO2018078053A1 (en) | 2016-10-26 | 2018-05-03 | Curevac Ag | Lipid nanoparticle mrna vaccines |
| WO2018209158A2 (en) | 2017-05-10 | 2018-11-15 | Editas Medicine, Inc. | Crispr/rna-guided nuclease systems and methods |
| WO2019027999A1 (en) | 2017-07-31 | 2019-02-07 | Ohio State Innovation Foundation | Biomimetic nanomaterials and uses thereof |
| WO2019036030A1 (en) | 2017-08-17 | 2019-02-21 | Acuitas Therapeutics, Inc. | Lipids for use in lipid nanoparticle formulations |
| WO2019036682A1 (en) | 2017-08-18 | 2019-02-21 | Modernatx, Inc. | Rna polymerase variants |
| WO2019067992A1 (en) | 2017-09-29 | 2019-04-04 | Intellia Therapeutics, Inc. | Formulations |
| WO2019089828A1 (en) | 2017-10-31 | 2019-05-09 | Acuitas Therapeutics, Inc. | Lamellar lipid nanoparticles |
| WO2019090175A1 (en) | 2017-11-02 | 2019-05-09 | Arbor Biotechnologies, Inc. | Novel crispr-associated transposon systems and components |
| WO2019099501A1 (en) | 2017-11-14 | 2019-05-23 | Ohio State Innovation Foundation | Benzene-1,3,5-tricarboxamide derived ester lipids and uses thereof |
| WO2020072605A1 (en) | 2018-10-02 | 2020-04-09 | Intellia Therapeutics, Inc. | Ionizable amine lipids |
| WO2020081938A1 (en) | 2018-10-18 | 2020-04-23 | Acuitas Therapeutics, Inc. | Lipids for lipid nanoparticle delivery of active agents |
| WO2020118041A1 (en) | 2018-12-05 | 2020-06-11 | Intellia Therapeutics, Inc. | Modified amine lipids |
| WO2020146805A1 (en) | 2019-01-11 | 2020-07-16 | Acuitas Therapeutics, Inc. | Lipids for lipid nanoparticle delivery of active agents |
| WO2020219876A1 (en) | 2019-04-25 | 2020-10-29 | Intellia Therapeutics, Inc. | Ionizable amine lipids and lipid nanoparticles |
| WO2020264254A1 (en) | 2019-06-28 | 2020-12-30 | Crispr Therapeutics Ag | Materials and methods for controlling gene editing |
| WO2021041301A1 (en) | 2019-08-24 | 2021-03-04 | Flagship Pioneering Innovations Vi, Llc | Modification of plant messenger packs with charged lipids |
| WO2021050512A1 (en) | 2019-09-09 | 2021-03-18 | Beam Therapeutics Inc. | Novel crispr enzymes, methods, systems and uses thereof |
| WO2021113494A1 (en) | 2019-12-03 | 2021-06-10 | Beam Therapeutics Inc. | Synthetic guide rna, compositions, methods, and uses thereof |
| WO2021154763A1 (en) | 2020-01-28 | 2021-08-05 | Modernatx, Inc. | Coronavirus rna vaccines |
| WO2022086846A2 (en) | 2020-10-19 | 2022-04-28 | Caribou Biosciences, Inc. | Dna-containing polynucleotides and guides for crispr type v systems, and methods of making and using the same |
| WO2022150608A1 (en) | 2021-01-08 | 2022-07-14 | Arbor Biotechnologies, Inc. | Compositions comprising a variant crispr nuclease polypeptide and uses thereof |
| WO2022159741A1 (en) | 2021-01-22 | 2022-07-28 | Arbor Biotechnologies, Inc. | Compositions comprising a nuclease and uses thereof |
| WO2022229851A1 (en) | 2021-04-26 | 2022-11-03 | Crispr Therapeutics Ag | Compositions and methods for using slucas9 scaffold sequences |
| WO2023122080A1 (en) * | 2021-12-20 | 2023-06-29 | Senda Biosciences, Inc. | Compositions comprising mrna and lipid reconstructed plant messenger packs |
-
2023
- 2023-11-09 EP EP23821780.6A patent/EP4615424A1/en active Pending
- 2023-11-09 WO PCT/US2023/037077 patent/WO2024102434A1/en not_active Ceased
- 2023-11-09 JP JP2025526705A patent/JP2025537750A/en active Pending
- 2023-11-09 CN CN202380084391.7A patent/CN120359023A/en active Pending
Patent Citations (122)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5208036A (en) | 1985-01-07 | 1993-05-04 | Syntex (U.S.A.) Inc. | N-(ω, (ω-1)-dialkyloxy)- and N-(ω, (ω-1)-dialkenyloxy)-alk-1-yl-N,N,N-tetrasubstituted ammonium lipids and uses therefor |
| US5279833A (en) | 1990-04-04 | 1994-01-18 | Yale University | Liposomal transfection of nucleic acids into animal cells |
| US5264618A (en) | 1990-04-19 | 1993-11-23 | Vical, Inc. | Cationic lipids for intracellular delivery of biologically active molecules |
| US5283185A (en) | 1991-08-28 | 1994-02-01 | University Of Tennessee Research Corporation | Method for delivering nucleic acids into cells |
| US5785992A (en) | 1994-09-30 | 1998-07-28 | Inex Pharmaceuticals Corp. | Compositions for the introduction of polyanionic materials into cells |
| US5753613A (en) | 1994-09-30 | 1998-05-19 | Inex Pharmaceuticals Corporation | Compositions for the introduction of polyanionic materials into cells |
| WO1996010390A1 (en) | 1994-09-30 | 1996-04-11 | Inex Pharmaceuticals Corp. | Novel compositions for the introduction of polyanionic materials into cells |
| WO2002098443A2 (en) | 2001-06-05 | 2002-12-12 | Curevac Gmbh | Stabilised mrna with an increased g/c content and optimised codon for use in gene therapy |
| US8084599B2 (en) | 2004-03-15 | 2011-12-27 | City Of Hope | Methods and compositions for the specific inhibition of gene expression by double-stranded RNA |
| US20060083780A1 (en) | 2004-06-07 | 2006-04-20 | Protiva Biotherapeutics, Inc. | Cationic lipids and methods of use |
| US20060240554A1 (en) | 2005-02-14 | 2006-10-26 | Sirna Therapeutics, Inc. | Lipid nanoparticle based compositions and methods for the delivery of biologically active molecules |
| US9012219B2 (en) | 2005-08-23 | 2015-04-21 | The Trustees Of The University Of Pennsylvania | RNA preparations comprising purified modified RNA for reprogramming cells |
| US20100129877A1 (en) | 2005-09-28 | 2010-05-27 | Ugur Sahin | Modification of RNA, Producing an Increased Transcript Stability and Translation Efficiency |
| US8278063B2 (en) | 2007-06-29 | 2012-10-02 | Commonwealth Scientific And Industrial Research Organisation | Methods for degrading toxic compounds |
| US20100293625A1 (en) | 2007-09-26 | 2010-11-18 | Interexon Corporation | Synthetic 5'UTRs, Expression Vectors, and Methods for Increasing Transgene Expression |
| US20090226470A1 (en) | 2007-12-11 | 2009-09-10 | Mauro Vincent P | Compositions and methods related to mRNA translational enhancer elements |
| US8349809B2 (en) | 2008-12-18 | 2013-01-08 | Dicerna Pharmaceuticals, Inc. | Single stranded extended dicer substrate agents and methods for the specific inhibition of gene expression |
| US8513207B2 (en) | 2008-12-18 | 2013-08-20 | Dicerna Pharmaceuticals, Inc. | Extended dicer substrate agents and methods for the specific inhibition of gene expression |
| US9200276B2 (en) | 2009-06-01 | 2015-12-01 | Halo-Bio Rnai Therapeutics, Inc. | Polynucleotides for multivalent RNA interference, compositions and methods of use thereof |
| WO2012019168A2 (en) | 2010-08-06 | 2012-02-09 | Moderna Therapeutics, Inc. | Engineered nucleic acids and methods of use thereof |
| WO2012045075A1 (en) | 2010-10-01 | 2012-04-05 | Jason Schrum | Modified nucleosides, nucleotides, and nucleic acids, and uses thereof |
| WO2012135805A2 (en) | 2011-03-31 | 2012-10-04 | modeRNA Therapeutics | Delivery and formulation of engineered nucleic acids |
| US20140206753A1 (en) | 2011-06-08 | 2014-07-24 | Shire Human Genetic Therapies, Inc. | Lipid nanoparticle compositions and methods for mrna delivery |
| WO2013052523A1 (en) | 2011-10-03 | 2013-04-11 | modeRNA Therapeutics | Modified nucleosides, nucleotides, and nucleic acids, and uses thereof |
| WO2013063468A1 (en) | 2011-10-27 | 2013-05-02 | Massachusetts Institute Of Technology | Amino acid derivates functionalized on the n- terminal capable of forming drug incapsulating microspheres |
| WO2013090648A1 (en) | 2011-12-16 | 2013-06-20 | modeRNA Therapeutics | Modified nucleoside, nucleotide, and nucleic acid compositions |
| WO2013151736A2 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | In vivo production of proteins |
| WO2013151667A1 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides |
| WO2013151669A1 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of cytoplasmic and cytoskeletal proteins |
| WO2013151666A2 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of biologics and proteins associated with human disease |
| WO2013151671A1 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of cosmetic proteins and peptides |
| WO2013151668A2 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of secreted proteins |
| WO2013151664A1 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of proteins |
| WO2013151663A1 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of membrane proteins |
| WO2013151670A2 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of nuclear proteins |
| WO2013151665A2 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of proteins associated with human disease |
| WO2013151672A2 (en) | 2012-04-02 | 2013-10-10 | modeRNA Therapeutics | Modified polynucleotides for the production of oncology-related proteins and peptides |
| US20140068797A1 (en) | 2012-05-25 | 2014-03-06 | University Of Vienna | Methods and compositions for rna-directed target dna modification and for rna-directed modulation of transcription |
| US20160138008A1 (en) | 2012-05-25 | 2016-05-19 | The Regents Of The University Of California | Methods and compositions for rna-directed target dna modification and for rna-directed modulation of transcription |
| WO2013185069A1 (en) | 2012-06-08 | 2013-12-12 | Shire Human Genetic Therapies, Inc. | Pulmonary delivery of mrna to non-lung target cells |
| WO2014028429A2 (en) | 2012-08-14 | 2014-02-20 | Moderna Therapeutics, Inc. | Enzymes and polymerases for the synthesis of rna |
| US20150344912A1 (en) | 2012-10-23 | 2015-12-03 | Toolgen Incorporated | Composition for cleaving a target dna comprising a guide rna specific for the target dna and cas protein-encoding nucleic acid or cas protein, and use thereof |
| US8993233B2 (en) | 2012-12-12 | 2015-03-31 | The Broad Institute Inc. | Engineering and optimization of systems, methods and compositions for sequence manipulation with functional domains |
| US8771945B1 (en) | 2012-12-12 | 2014-07-08 | The Broad Institute, Inc. | CRISPR-Cas systems and methods for altering expression of gene products |
| US8697359B1 (en) | 2012-12-12 | 2014-04-15 | The Broad Institute, Inc. | CRISPR-Cas systems and methods for altering expression of gene products |
| US20140186958A1 (en) | 2012-12-12 | 2014-07-03 | Feng Zhang | Engineering and optimization of systems, methods and compositions for sequence manipulation with functional domains |
| US8999641B2 (en) | 2012-12-12 | 2015-04-07 | The Broad Institute Inc. | Engineering and optimization of systems, methods and compositions for sequence manipulation with functional domains |
| US8795965B2 (en) | 2012-12-12 | 2014-08-05 | The Broad Institute, Inc. | CRISPR-Cas component systems, methods and compositions for sequence manipulation |
| US8945839B2 (en) | 2012-12-12 | 2015-02-03 | The Broad Institute Inc. | CRISPR-Cas systems and methods for altering expression of gene products |
| US8865406B2 (en) | 2012-12-12 | 2014-10-21 | The Broad Institute Inc. | Engineering and optimization of improved systems, methods and enzyme compositions for sequence manipulation |
| US8871445B2 (en) | 2012-12-12 | 2014-10-28 | The Broad Institute Inc. | CRISPR-Cas component systems, methods and compositions for sequence manipulation |
| US8889418B2 (en) | 2012-12-12 | 2014-11-18 | The Broad Institute Inc. | Engineering and optimization of improved systems, methods and enzyme compositions for sequence manipulation |
| US8889356B2 (en) | 2012-12-12 | 2014-11-18 | The Broad Institute Inc. | CRISPR-Cas nickase systems, methods and compositions for sequence manipulation in eukaryotes |
| US8895308B1 (en) | 2012-12-12 | 2014-11-25 | The Broad Institute Inc. | Engineering and optimization of improved systems, methods and enzyme compositions for sequence manipulation |
| US8906616B2 (en) | 2012-12-12 | 2014-12-09 | The Broad Institute Inc. | Engineering of systems, methods and optimized guide compositions for sequence manipulation |
| US8932814B2 (en) | 2012-12-12 | 2015-01-13 | The Broad Institute Inc. | CRISPR-Cas nickase systems, methods and compositions for sequence manipulation in eukaryotes |
| WO2014093924A1 (en) | 2012-12-13 | 2014-06-19 | Moderna Therapeutics, Inc. | Modified nucleic acid molecules and uses thereof |
| US9023649B2 (en) | 2012-12-17 | 2015-05-05 | President And Fellows Of Harvard College | RNA-guided human genome engineering |
| WO2014164253A1 (en) | 2013-03-09 | 2014-10-09 | Moderna Therapeutics, Inc. | Heterologous untranslated regions for mrna |
| WO2014152031A1 (en) | 2013-03-15 | 2014-09-25 | Moderna Therapeutics, Inc. | Ribonucleic acid purification |
| WO2014152030A1 (en) | 2013-03-15 | 2014-09-25 | Moderna Therapeutics, Inc. | Removal of dna fragments in mrna production process |
| WO2014144767A1 (en) | 2013-03-15 | 2014-09-18 | Moderna Therapeutics, Inc. | Ion exchange purification of mrna |
| WO2014152027A1 (en) | 2013-03-15 | 2014-09-25 | Moderna Therapeutics, Inc. | Manufacturing methods for production of rna transcripts |
| WO2014144196A1 (en) | 2013-03-15 | 2014-09-18 | Shire Human Genetic Therapies, Inc. | Synergistic enhancement of the delivery of nucleic acids via blended formulations |
| WO2015024667A1 (en) | 2013-08-21 | 2015-02-26 | Curevac Gmbh | Method for increasing expression of rna-encoded proteins |
| WO2015038892A1 (en) | 2013-09-13 | 2015-03-19 | Moderna Therapeutics, Inc. | Polynucleotide compositions containing amino acids |
| WO2015062738A1 (en) | 2013-11-01 | 2015-05-07 | Curevac Gmbh | Modified rna with decreased immunostimulatory properties |
| US20150166980A1 (en) | 2013-12-12 | 2015-06-18 | President And Fellows Of Harvard College | Fusions of cas9 domains and nucleic acid-editing domains |
| WO2015089406A1 (en) | 2013-12-12 | 2015-06-18 | President And Fellows Of Harvard College | Cas variants for gene editing |
| WO2015089511A2 (en) | 2013-12-13 | 2015-06-18 | Moderna Therapeutics, Inc. | Modified nucleic acid molecules and uses thereof |
| WO2015095340A1 (en) | 2013-12-19 | 2015-06-25 | Novartis Ag | Lipids and lipid compositions for the delivery of active agents |
| WO2015101415A1 (en) | 2013-12-30 | 2015-07-09 | Curevac Gmbh | Artificial nucleic acid molecules |
| WO2015101414A2 (en) | 2013-12-30 | 2015-07-09 | Curevac Gmbh | Artificial nucleic acid molecules |
| WO2015153789A1 (en) | 2014-04-01 | 2015-10-08 | Editas Medicine, Inc. | Crispr/cas-related methods and compositions for treating herpes simplex virus type 1 (hsv-1) |
| WO2015184256A2 (en) | 2014-05-30 | 2015-12-03 | Shire Human Genetic Therapies, Inc. | Biodegradable lipids for delivery of nucleic acids |
| WO2015196130A2 (en) | 2014-06-19 | 2015-12-23 | Moderna Therapeutics, Inc. | Alternative nucleic acid molecules and uses thereof |
| WO2015196118A1 (en) | 2014-06-19 | 2015-12-23 | Moderna Therapeutics, Inc. | Alternative nucleic acid molecules and uses thereof |
| WO2015196128A2 (en) | 2014-06-19 | 2015-12-23 | Moderna Therapeutics, Inc. | Alternative nucleic acid molecules and uses thereof |
| WO2015199952A1 (en) | 2014-06-25 | 2015-12-30 | Acuitas Therapeutics Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2016004202A1 (en) | 2014-07-02 | 2016-01-07 | Massachusetts Institute Of Technology | Polyamine-fatty acid derived lipidoids and uses thereof |
| WO2016011306A2 (en) | 2014-07-17 | 2016-01-21 | Moderna Therapeutics, Inc. | Terminal modifications of polynucleotides |
| WO2016118724A1 (en) | 2015-01-21 | 2016-07-28 | Moderna Therapeutics, Inc. | Lipid nanoparticle compositions |
| WO2016118725A1 (en) | 2015-01-23 | 2016-07-28 | Moderna Therapeutics, Inc. | Lipid nanoparticle compositions |
| WO2016154596A1 (en) | 2015-03-25 | 2016-09-29 | Editas Medicine, Inc. | Crispr/cas-related methods, compositions and components |
| WO2016187531A1 (en) | 2015-05-21 | 2016-11-24 | Ohio State Innovation Foundation | Benzene-1,3,5-tricarboxamide derivatives and uses thereof |
| WO2016201155A1 (en) | 2015-06-10 | 2016-12-15 | Caribou Biosciences, Inc. | Crispr-cas compositions and methods |
| US20160208243A1 (en) | 2015-06-18 | 2016-07-21 | The Broad Institute, Inc. | Novel crispr enzymes and systems |
| WO2016205691A1 (en) | 2015-06-19 | 2016-12-22 | Massachusetts Institute Of Technology | Alkenyl substituted 2,5-piperazinediones and their use in compositions for delivering an agent to a subject or cell |
| WO2017004143A1 (en) | 2015-06-29 | 2017-01-05 | Acuitas Therapeutics Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017049245A2 (en) | 2015-09-17 | 2017-03-23 | Modernatx, Inc. | Compounds and compositions for intracellular delivery of therapeutic agents |
| WO2017075531A1 (en) | 2015-10-28 | 2017-05-04 | Acuitas Therapeutics, Inc. | Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017117528A1 (en) | 2015-12-30 | 2017-07-06 | Acuitas Therapeutics, Inc. | Lipids and lipid nanoparticle formulations for delivery of nucleic acids |
| WO2017127750A1 (en) | 2016-01-22 | 2017-07-27 | Modernatx, Inc. | Messenger ribonucleic acids for the production of intracellular binding polypeptides and methods of use thereof |
| WO2017165826A1 (en) | 2016-03-25 | 2017-09-28 | Editas Medicine, Inc. | Genome editing systems comprising repair-modulating enzyme molecules and methods of their use |
| WO2017173054A1 (en) | 2016-03-30 | 2017-10-05 | Intellia Therapeutics, Inc. | Lipid nanoparticle formulations for crispr/cas components |
| WO2017176974A1 (en) | 2016-04-06 | 2017-10-12 | Ohio State Innovation Foundation | Biodegradable amino-ester nanomaterials for nucleic acid delivery |
| WO2018053209A1 (en) | 2016-09-14 | 2018-03-22 | Modernatx, Inc. | High purity rna compositions and methods for preparation thereof |
| WO2018078053A1 (en) | 2016-10-26 | 2018-05-03 | Curevac Ag | Lipid nanoparticle mrna vaccines |
| WO2018209158A2 (en) | 2017-05-10 | 2018-11-15 | Editas Medicine, Inc. | Crispr/rna-guided nuclease systems and methods |
| WO2019027999A1 (en) | 2017-07-31 | 2019-02-07 | Ohio State Innovation Foundation | Biomimetic nanomaterials and uses thereof |
| WO2019036030A1 (en) | 2017-08-17 | 2019-02-21 | Acuitas Therapeutics, Inc. | Lipids for use in lipid nanoparticle formulations |
| WO2019036682A1 (en) | 2017-08-18 | 2019-02-21 | Modernatx, Inc. | Rna polymerase variants |
| WO2019067992A1 (en) | 2017-09-29 | 2019-04-04 | Intellia Therapeutics, Inc. | Formulations |
| WO2019089828A1 (en) | 2017-10-31 | 2019-05-09 | Acuitas Therapeutics, Inc. | Lamellar lipid nanoparticles |
| WO2019090175A1 (en) | 2017-11-02 | 2019-05-09 | Arbor Biotechnologies, Inc. | Novel crispr-associated transposon systems and components |
| WO2019090173A1 (en) | 2017-11-02 | 2019-05-09 | Arbor Biotechnologies, Inc. | Novel crispr-associated transposon systems and components |
| WO2019099501A1 (en) | 2017-11-14 | 2019-05-23 | Ohio State Innovation Foundation | Benzene-1,3,5-tricarboxamide derived ester lipids and uses thereof |
| WO2020072605A1 (en) | 2018-10-02 | 2020-04-09 | Intellia Therapeutics, Inc. | Ionizable amine lipids |
| WO2020081938A1 (en) | 2018-10-18 | 2020-04-23 | Acuitas Therapeutics, Inc. | Lipids for lipid nanoparticle delivery of active agents |
| WO2020118041A1 (en) | 2018-12-05 | 2020-06-11 | Intellia Therapeutics, Inc. | Modified amine lipids |
| WO2020146805A1 (en) | 2019-01-11 | 2020-07-16 | Acuitas Therapeutics, Inc. | Lipids for lipid nanoparticle delivery of active agents |
| WO2020219876A1 (en) | 2019-04-25 | 2020-10-29 | Intellia Therapeutics, Inc. | Ionizable amine lipids and lipid nanoparticles |
| WO2020264254A1 (en) | 2019-06-28 | 2020-12-30 | Crispr Therapeutics Ag | Materials and methods for controlling gene editing |
| WO2021041301A1 (en) | 2019-08-24 | 2021-03-04 | Flagship Pioneering Innovations Vi, Llc | Modification of plant messenger packs with charged lipids |
| WO2021050512A1 (en) | 2019-09-09 | 2021-03-18 | Beam Therapeutics Inc. | Novel crispr enzymes, methods, systems and uses thereof |
| WO2021113494A1 (en) | 2019-12-03 | 2021-06-10 | Beam Therapeutics Inc. | Synthetic guide rna, compositions, methods, and uses thereof |
| WO2021154763A1 (en) | 2020-01-28 | 2021-08-05 | Modernatx, Inc. | Coronavirus rna vaccines |
| WO2022086846A2 (en) | 2020-10-19 | 2022-04-28 | Caribou Biosciences, Inc. | Dna-containing polynucleotides and guides for crispr type v systems, and methods of making and using the same |
| WO2022150608A1 (en) | 2021-01-08 | 2022-07-14 | Arbor Biotechnologies, Inc. | Compositions comprising a variant crispr nuclease polypeptide and uses thereof |
| WO2022159741A1 (en) | 2021-01-22 | 2022-07-28 | Arbor Biotechnologies, Inc. | Compositions comprising a nuclease and uses thereof |
| WO2022229851A1 (en) | 2021-04-26 | 2022-11-03 | Crispr Therapeutics Ag | Compositions and methods for using slucas9 scaffold sequences |
| WO2023122080A1 (en) * | 2021-12-20 | 2023-06-29 | Senda Biosciences, Inc. | Compositions comprising mrna and lipid reconstructed plant messenger packs |
Non-Patent Citations (46)
| Title |
|---|
| ALTSCHUL ET AL., J. MOL. BIOL., vol. 215, 1990, pages 403 - 410 |
| BLIGHDYER, J BIOLCHEM PHYSIOL, vol. 37, 1959, pages 911 - 917 |
| CHO K.J. ET AL., J MOL BIOL., vol. 390, 2009, pages 83 - 98 |
| CONG ET AL., SCIENCE, vol. 339, 2013, pages 819 - 823 |
| DOMINGUEZ ET AL., NAT. REV. MOL. CELL BIOL., vol. 17, 2016, pages 5 - 15 |
| FAWSON D.M. ET AL., NATURE, vol. 349, 1991, pages 541 - 544 |
| GAJ ET AL., TRENDS BIOTECHNOL., vol. 31, no. 7, 2013, pages 397 - 405 |
| GILBERT ET AL.: "CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes", CELL, vol. 154, 2013, pages 442 - 51, XP055115843, DOI: 10.1016/j.cell.2013.06.044 |
| GRANIER T. ET AL., J BIOL INORG CHEM., vol. 8, 2003, pages 105 - 111 |
| GREEN ET AL.: "Remington: The Science and Practice of Pharmacy", 2012, COLD SPRING HARBOR LABORATORY PRESS |
| HAFEZ, I.M. ET AL., GENE THER, vol. 8, 2001, pages 1188 - 1196 |
| HAN XUEXIANG ET AL: "An ionizable lipid toolbox for RNA delivery", NATURE COMMUNICATIONS, vol. 12, no. 1, 1 December 2021 (2021-12-01), XP055938542, Retrieved from the Internet <URL:https://www.nature.com/articles/s41467-021-27493-0.pdf> DOI: 10.1038/s41467-021-27493-0 * |
| HENDEL ET AL., NATURE BIOTECHNOL., 2015, pages 985 - 991 |
| HORVATH ET AL., SCIENCE, vol. 327, 2010, pages 167 - 170 |
| HOU XUCHENG ET AL: "Lipid nanoparticles for mRNA delivery", NATURE REVIEWS MATERIALS, 10 August 2021 (2021-08-10), XP055861763, ISSN: 2058-8437, DOI: 10.1038/s41578-021-00358-0 * |
| J. MCCLELLANM. C. KING, CELL, vol. 141, 2010, pages 210 - 217 |
| JONATHAN D. FINN ET AL: "A Single Administration of CRISPR/Cas9 Lipid Nanoparticles Achieves Robust and Persistent In Vivo Genome Editing", CELL REPORTS, vol. 22, no. 9, 1 February 2018 (2018-02-01), US, pages 2227 - 2235, XP055527484, ISSN: 2211-1247, DOI: 10.1016/j.celrep.2018.02.014 * |
| JU ET AL., MOL. THERAPY., vol. 21, no. 7, 2013, pages 1345 - 1357 |
| KIM, J.H. ET AL., PLOS ONE, vol. 6, 2011, pages el8556 |
| MAKAROVA ET AL., BIOLOGY DIRECT, vol. 1, 2006, pages 7 |
| MALI ET AL.: "CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering", NAT., vol. 31, 2013, pages 833 - 8, XP055693153, DOI: 10.1038/nbt.2675 |
| MU ET AL., MOL. NUTR. FOOD RES., vol. 58, 2014, pages 1561 - 1573 |
| NEEDLEMAN, S.B.WUNSCH, C.D.: "A general method applicable to the search for similarities in the amino acid sequences of two proteins", J. MOL. BIOL., vol. 48, 1970, pages 443 - 453 |
| PEREZ-PINERA ET AL.: "RNA-guided gene activation by CRISPR-Cas9-based transcription factors", NAT. METHODS, vol. 10, 2013, pages 973 - 6, XP055181249, DOI: 10.1038/nmeth.2600 |
| RAHMANPOUR R. ET AL., FEBS J., vol. 280, 2013, pages 2097 - 2104 |
| RAIMONDO ET AL., ONCOTARGET, vol. 6, no. 23, 2015, pages 19514 |
| RAN ET AL., NATURE PROTOCOLS, vol. 8, 2013, pages 2281 - 2308 |
| RAN ET AL.: "Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression", CELL, vol. 152, 2013, pages 1173 - 1389 |
| REGENTE ET AL., FEBS LETTERS, vol. 583, 2009, pages 3363 - 3366 |
| REGENTE ET AL., J OF EXP. BIOL., vol. 68, no. 20, 2017, pages 5485 - 5496 |
| RUTTER ET AL., BIO. PROTOC., vol. 7, no. 17, 2017, pages e2533 |
| RUTTERINNES, PLANT PHYSIOL., vol. 173, no. 1, 2017, pages 728 - 741 |
| SCHERER ET AL., NUCLEIC ACIDS RES., vol. 35, 2007, pages 2620 - 2628 |
| SEMPLE, S.C. ET AL., ADV. DRUG DELIV REV, vol. 32, 1998, pages 3 - 17 |
| SMITH, T.F.WATERMAN, M.S.: "identification of common molecular subsequences", J. MOL. BIOL., vol. 147, 1981, pages 195 - 197, XP024015032, DOI: 10.1016/0022-2836(81)90087-5 |
| STEPHEN F. ALTSCHUL ET AL.: "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", NUCLEIC ACIDS RES., vol. 25, no. N-7433, 1997, pages 3389 - 3402, XP002905950, DOI: 10.1093/nar/25.17.3389 |
| STEPINSKI ET AL.: "Synthesis and properties of mRNAs containing the novel 'anti- reverse' cap analogs 7-methyl(3'-O-methyl)GpppG and 7-methyl(3'deoxy)GpppG", RNA, vol. 201521, 2001, pages 1486 - 1495 |
| SUTTER M. ET AL., NAT STRUCT MOL BIOL., vol. 15, 2008, pages 939 - 947 |
| TAO Y ET AL., STRUCTURE., vol. 5, no. 6, 15 June 1997 (1997-06-15), pages 789 - 98 |
| TSAI, NATURE BIOTECHNOL., vol. 32, 2014, pages 6569 - 576 |
| UI-TEI ET AL., FEBS LETTERS, vol. 479, 2000, pages 79 - 82 |
| WANG ET AL., MOLECULAR THERAPY., vol. 22, no. 3, 2014, pages 522 - 534 |
| WHITEHEAD ET AL., NATURE COMMUNICATIONS, vol. 5, 2014, pages 4277 |
| YAN JINGYUE ET AL: "Harnessing lipid nanoparticles for efficient CRISPR delivery", BIOMATERIALS SCIENCE, vol. 9, no. 18, 14 September 2021 (2021-09-14), GB, pages 6001 - 6011, XP093041709, ISSN: 2047-4830, Retrieved from the Internet <URL:https://pubs.rsc.org/en/content/articlepdf/2021/bm/d1bm00537e> DOI: 10.1039/D1BM00537E * |
| ZETSCHE ET AL., CELL, vol. 163, 2015, pages 759 - 771 |
| ZHANG X. ET AL., J MOL BIOL., vol. 362, 2006, pages 753 - 770 |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2025042786A1 (en) | 2023-08-18 | 2025-02-27 | Flagship Pioneering Innovations Vi, Llc | Compositions comprising circular polyribonucleotides and uses thereof |
| WO2025106915A1 (en) | 2023-11-17 | 2025-05-22 | Sail Biomedicines, Inc. | Circular polyribonucleotides encoding glucagon-like peptide 1 (glp-1) and uses thereof |
| WO2025106930A1 (en) | 2023-11-17 | 2025-05-22 | Sail Biomedicines, Inc. | Circular polyribonucleotides encoding glucagon-like peptide 2 (glp-2) and uses thereof |
| WO2025179198A1 (en) | 2024-02-23 | 2025-08-28 | Sail Biomedicines, Inc. | Circular polyribonucleotides and unmodified linear rnas with reduced immunogenicity |
Also Published As
| Publication number | Publication date |
|---|---|
| JP2025537750A (en) | 2025-11-20 |
| CN120359023A (en) | 2025-07-22 |
| EP4615424A1 (en) | 2025-09-17 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP7581405B2 (en) | Lipid Nanoparticle Formulations for CRISPR/CAS Components | |
| WO2024102434A1 (en) | Rna compositions comprising lipid nanoparticles or lipid reconstructed natural messenger packs | |
| JP7759311B2 (en) | Recombinase compositions and methods of use | |
| US20250367322A1 (en) | Nucleobase editing system and method of using same for modifying nucleic acid sequences | |
| US20250057980A1 (en) | Co-Delivery of a Gene Editor Construct and a Donor Template | |
| US20250295755A1 (en) | Mrna vaccine composition | |
| AU2023311964A1 (en) | Gene editing components, systems, and methods of use | |
| CA3221566A1 (en) | Integrase compositions and methods | |
| AU2024272060A1 (en) | Gene editing systems and compositions for treatment of hemoglobinopathies and methods of using the same | |
| WO2023225670A2 (en) | Ex vivo programmable gene insertion | |
| WO2023215831A1 (en) | Guide rna compositions for programmable gene insertion | |
| WO2024220712A2 (en) | Vaccine compositions | |
| WO2024234006A1 (en) | Systems, compositions, and methods for targeting liver sinusodial endothelial cells (lsecs) | |
| WO2024138194A1 (en) | Platforms, compositions, and methods for in vivo programmable gene insertion | |
| US20240084274A1 (en) | Gene editing components, systems, and methods of use | |
| WO2025081042A1 (en) | Nickase-retron template-based precision editing system and methods of use | |
| WO2025050069A1 (en) | Programmable gene insertion using engineered integration enzymes | |
| WO2025224182A2 (en) | Single construct platform for simultaneous delivery of gene editing machinery and nucleic acid cargo | |
| HK40005508A (en) | Lipid nanoparticle formulations for crispr/cas components |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 23821780 Country of ref document: EP Kind code of ref document: A1 |
|
| ENP | Entry into the national phase |
Ref document number: 2025526705 Country of ref document: JP Kind code of ref document: A |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2025526705 Country of ref document: JP |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 202380084391.7 Country of ref document: CN |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2023821780 Country of ref document: EP |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2023821780 Country of ref document: EP Effective date: 20250610 |
|
| WWP | Wipo information: published in national office |
Ref document number: 202380084391.7 Country of ref document: CN |
|
| WWP | Wipo information: published in national office |
Ref document number: 2023821780 Country of ref document: EP |