WO2025122927A1 - Antibodies directed to claudin 18.2, including bispecific formats thereof - Google Patents
Antibodies directed to claudin 18.2, including bispecific formats thereof Download PDFInfo
- Publication number
- WO2025122927A1 WO2025122927A1 PCT/US2024/058961 US2024058961W WO2025122927A1 WO 2025122927 A1 WO2025122927 A1 WO 2025122927A1 US 2024058961 W US2024058961 W US 2024058961W WO 2025122927 A1 WO2025122927 A1 WO 2025122927A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- heavy chain
- seq
- variant
- sequence
- cdr2
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/2803—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily
- C07K16/2809—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily against the T-cell receptor (TcR)-CD3 complex
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/31—Immunoglobulins specific features characterized by aspects of specificity or valency multispecific
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/60—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments
- C07K2317/62—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments comprising only variable region components
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/60—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments
- C07K2317/62—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments comprising only variable region components
- C07K2317/622—Single chain antibody (scFv)
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/60—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments
- C07K2317/64—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments comprising a combination of variable region and constant region components
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/75—Agonist effect on antigen
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/92—Affinity (KD), association rate (Ka), dissociation rate (Kd) or EC50 value
Definitions
- compositions that bind Claudin 18.2 and related methods are directed to compositions that bind Claudin 18.2 and related methods.
- CLDN claudin
- the present disclosure relates to a composition that specifically binds to Claudin 18.2 and CD3.
- the present disclosure describes the isolation and characterization of antibodies, antibody fragments, and antibody variants specific for Claudin 18.2 and CD3.
- the antibody is a tandem single-chain variable fragment (scFv) specific for Claudin 18.2 and CD3 (FIG. 1A).
- the antibody is a hetero VH IgG (FIG. 1B) specific for Claudin 18.2 and CD3.
- the antibody is in the format of an scFv-Fab IgG antibody, which is illustrated as, for example, in FIG. 1C and is specific for Claudin 18.2 and CD3.
- the antibody in the format of an IgG-scFv, which is illustrated as, for example, in FIG. 1D and is specific for Claudin 18.2 and CD3.
- the antibody is in an IgG- (scFv) 2 format , which is illustrated, for example, in FIG. 1E and is specific for Claudin 18.2 and CD3.
- the antibody is in an IgG-(scFv) format, which is, for example, illustrated in FIG. 1F and is specific for Claudin 18.2 and CD3.
- the antibody specific for Claudin 18.2 and CD3 binds Claudin 18.2 and CD3 contemporaneously.
- the bispecific antibody will bind to a tumor cell expressing Claudin 18.2 and an immune cell, such as a cytotoxic T cell, expressing CD3.
- an immune cell such as a cytotoxic T cell, expressing CD3.
- compositions comprising an antibody in an scFv-Fab IgG antibody that binds to Claudin 18.2 and CD3.
- the antibody comprises a first heavy chain, a second heavy chain, and a light chain, wherein
- the first heavy chain is selected from:
- CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, or
- QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4767), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, or (iii)
- the second heavy chain is selected from:
- KGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPP VLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4168), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and
- DKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3983), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAK1RPY1FK1AGQYDY (SEQ ID NO: 18) or a variant thereof, and
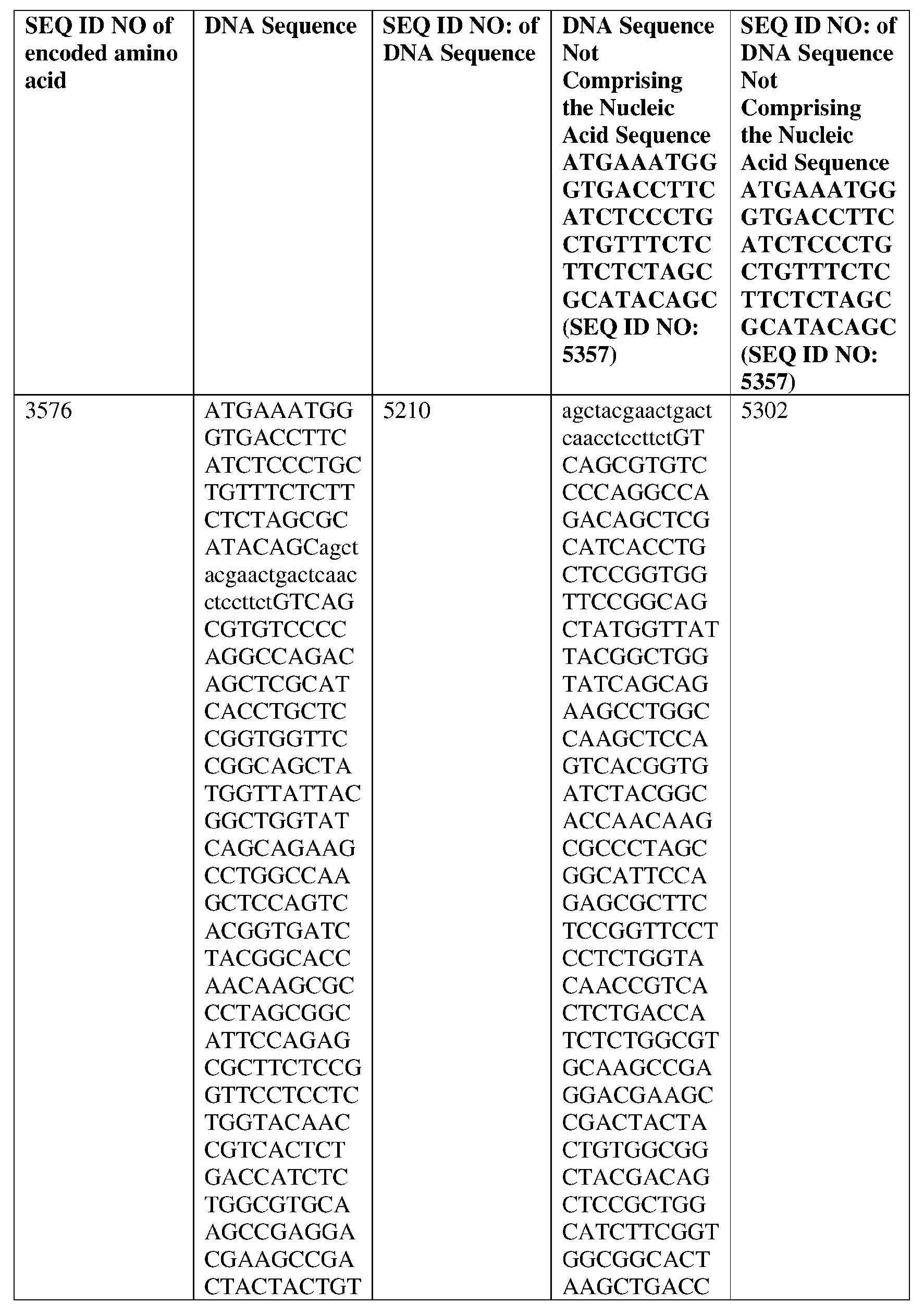
- S S (SEQ ID NO: 3576), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or (ii)
- an antibody is provided in an IgG-scFv antibody format that binds to Claudin 18.2 and CD3, wherein the antibody comprises: (a) a first heavy chain and a second heavy chain, wherein
- the first heavy chain comprises:
- QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and (ii) the second heavy chain comprises:
- K SEQ ID NO: 4778
- amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
- the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof
- the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof
- the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof
- the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof
- the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof
- the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof
- LHNHYTQKSLSLSPGK (SEQ ID NO: 4803), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
- CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF
- PGK (SEQ ID NO: 4806); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
- CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSN
- the first heavy chain comprises:
- LHNHYTQKSLSLSPGK (SEQ ID NO: 4804), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
- a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences
- the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof
- the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof
- the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof
- a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences
- the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof
- the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof
- the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and/or a
- SPGK (SEQ ID NO: 4807); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
- heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences
- the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof
- the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof
- the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof
- a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences
- the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof
- the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof
- the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, and/or heavy chain
- the first heavy chain comprises:
- the second heavy chain is selected from:
- the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof
- the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof
- the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof
- the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof
- the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof
- the antibody or composition comprising the same is an antibody an IgG-(scFv) 2 antibody format that binds to Claudin 18.2 and CD3, wherein the antibody comprises: (a) a heavy chain selected from:
- the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), (it) MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC
- GGGSGGGGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAPKRWIY DTSKLASGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTFGQGTKLQITR (SEQ ID NO: 4770), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR
- the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof
- the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof
- the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof
- the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof
- the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof
- the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof
- the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof
- the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof
- VSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWV QQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYS NRWVFGGGTKLTVL (SEQ ID NO: 4772), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region
- R (SEQ ID NO: 4773), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO
- FTISRDNAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 4774), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO
- GGTKLTVL (SEQ ID NO: 4775), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO
- amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or
- a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, optionally wherein the first or second heavy chain comprises a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N- or C -terminus; and/or optionally wherein the composition comprises a linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof.
- the disclosure comprises a linker
- the first heavy chain comprises:
- the second heavy chain is selected from:
- VSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWV QQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYS NRWVFGGGTKLTVL (SEQ ID NO: 5202), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR
- the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
- the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof
- the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof
- the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof
- the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof
- the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof
- the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof
- the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof
- the first heavy chain comprises:
- QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4767), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
- the heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences
- the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof
- the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof
- the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof
- the second heavy chain is selected from:
- the first heavy chain comprises:
- the second heavy chain is selected from:
- GGTKLTVL (SEQ ID NO: 5204), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
- the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof
- the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof
- the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof
- the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof
- the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof
- the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof
- the Fc can be from IgG.
- the IgG is human IgG.
- the human IgG is selected from IgGl, IgG2, IgG3, and IgG4.
- the antibody is formed through a knob-in-hole interaction in the Fc region.
- the human IgG Fc comprises one or mutations to promote knob-in-hole interaction.
- the mutations are selected from (i) T366Y or T366W, and (ii) Y407T, Y407A, or Y407V. In certain embodiments of any of the above aspects, the mutations are selected from:
- the human IgG Fc comprises one or mutations to reduce or eliminate the effector function of the Fc domains.
- the mutations are L234A and L235A (LALA) substitutions in human IgGl.
- the human IgG Fc comprises one or mutations to stabilize a hinge region in the Fc domain.
- the mutation is S228P.
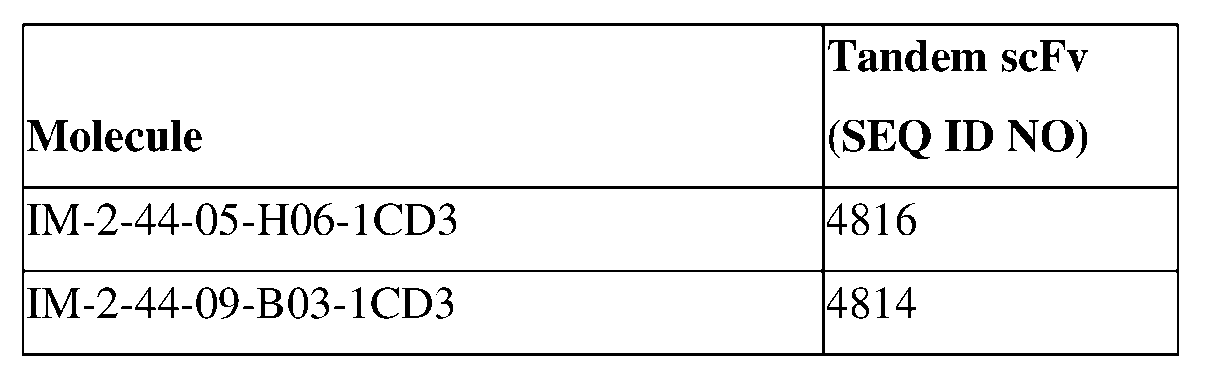
- composition comprising a tandem singlechain variable fragment (scFv) specific for Claudin 18.2 and CD3 comprising one or more of: (a) an amino acid sequence of:
- WVFGGGTKLTVL (SEQ ID: 4816), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or a variant thereof, the CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or a variant thereof, and the CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ
- CVLWYSNRWVFGGGTKLTVL (SEQ ID NO. 4814), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or a variant thereof, the CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or a variant thereof, and the CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the CDR3 sequence is CAKDASNGYCW
- the composition of any of the foregoing binds Claudin 18.2 and CD3 contemporaneously.
- the composition binds Claudin 18.2 with an affinity of less than 10 nM and with at least 100 fold greater affinity than Claudin 9, Claudin 3, and/or Claudin 4.
- the first heavy chain, the second heavy chain, the light chain, and/or the single-chain variable fragment does not comprise amino acids 1-18 of its respective SEQ ID NO.
- the first heavy chain, the second heavy chain, the light chain, and/or the single-chain variable fragment does not comprise the amino acid sequence MKWVTFISLLFLFSSAYS (SEQ ID NO: 5356).
- the leader sequence such as the amino acid sequence of SEQ ID NO: 5356, is removed during the expression of the antibody.
- the leader sequence of SEQ ID NO: 5356 can also be replaced with a different leader sequence to facilitate expression and/or secretion of the antibody.
- the antibody in a composition such as a pharmaceutical composition, does not comprise a leader sequence, including, but not limited to the amino acid sequence of SEQ ID NO: 5356.
- the disclosure relates to a pharmaceutical composition
- a pharmaceutical composition comprising an isolated antibody disclosed herein, or a nucleic acid molecule encoding the same.
- the pharmaceutical composition is an injectable pharmaceutical composition.
- the pharmaceutical composition is sterile.
- the pharmaceutical composition is pyrogen free.
- the disclosure relates to a nucleic acid molecule encoding an antibody or an amino acid sequence as disclosed herein.
- the disclosure relates to a vector comprising a nucleic acid molecule disclosed herein.
- the disclosure relates to a cell comprising a nucleic acid molecule or vector as disclosure herein.
- the disclosure relates to a method for modulating and/or targeting Claudin 18.2 and CD3 in a biological cell, comprising contacting the cell with a composition or pharmaceutical composition as disclosed herein.
- the disclosure relates to a method for modulating Claudin 18.2 activity in a biological cell comprising contacting a cell expressing Claudin 18.2 with a composition or pharmaceutical composition as disclosed herein.
- the disclosure relates to a method for inhibiting the function of Claudin 18.2 in a biological cell comprising contacting a cell expressing Claudin 18.2 with a composition or pharmaceutical composition as disclosed herein.
- the disclosure relates to a method for treating or preventing cancer comprising administering an effective amount of the composition or pharmaceutical composition of any one of claims 1 to 52 as disclosed herein to a subject in need thereof.
- the disclosure relates to a use of the composition or pharmaceutical composition as disclosed herein for the preparation of a medicament for the treatment of prevention of cancer.
- the cancer is selected from one or more of basal cell carcinoma, biliary tract cancer; bladder cancer; bone cancer; brain and central nervous system cancer; breast cancer; cancer of the peritoneum; cervical cancer; choriocarcinoma; colon and rectum cancer; connective tissue cancer; cancer of the digestive system; endometrial cancer; esophageal cancer; eye cancer; cancer of the head and neck; gastric cancer (including gastrointestinal cancer); glioblastoma; glioma; hepatic carcinoma; hepatoma; intra-epithelial neoplasm; kidney or renal cancer; larynx cancer; leukemia; liver cancer; lung cancer (e.g., small-cell lung cancer, non-small cell lung cancer, adenocarcinoma of the lung, and squamous carcinoma of the lung); melanoma; myeloma; neuroblastoma; oral cavity cancer (lip, tongue, mouth, and pharynx); ova
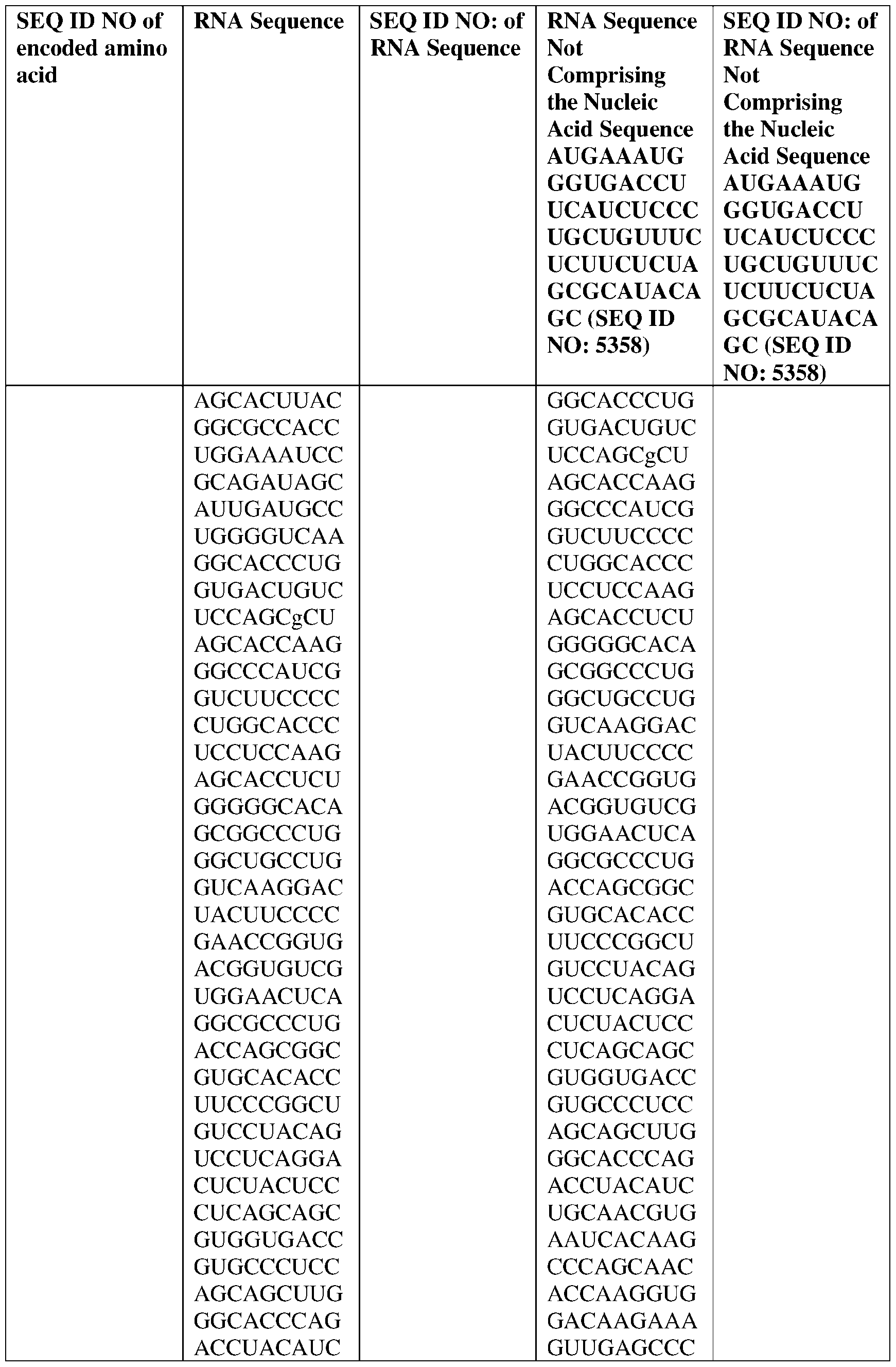
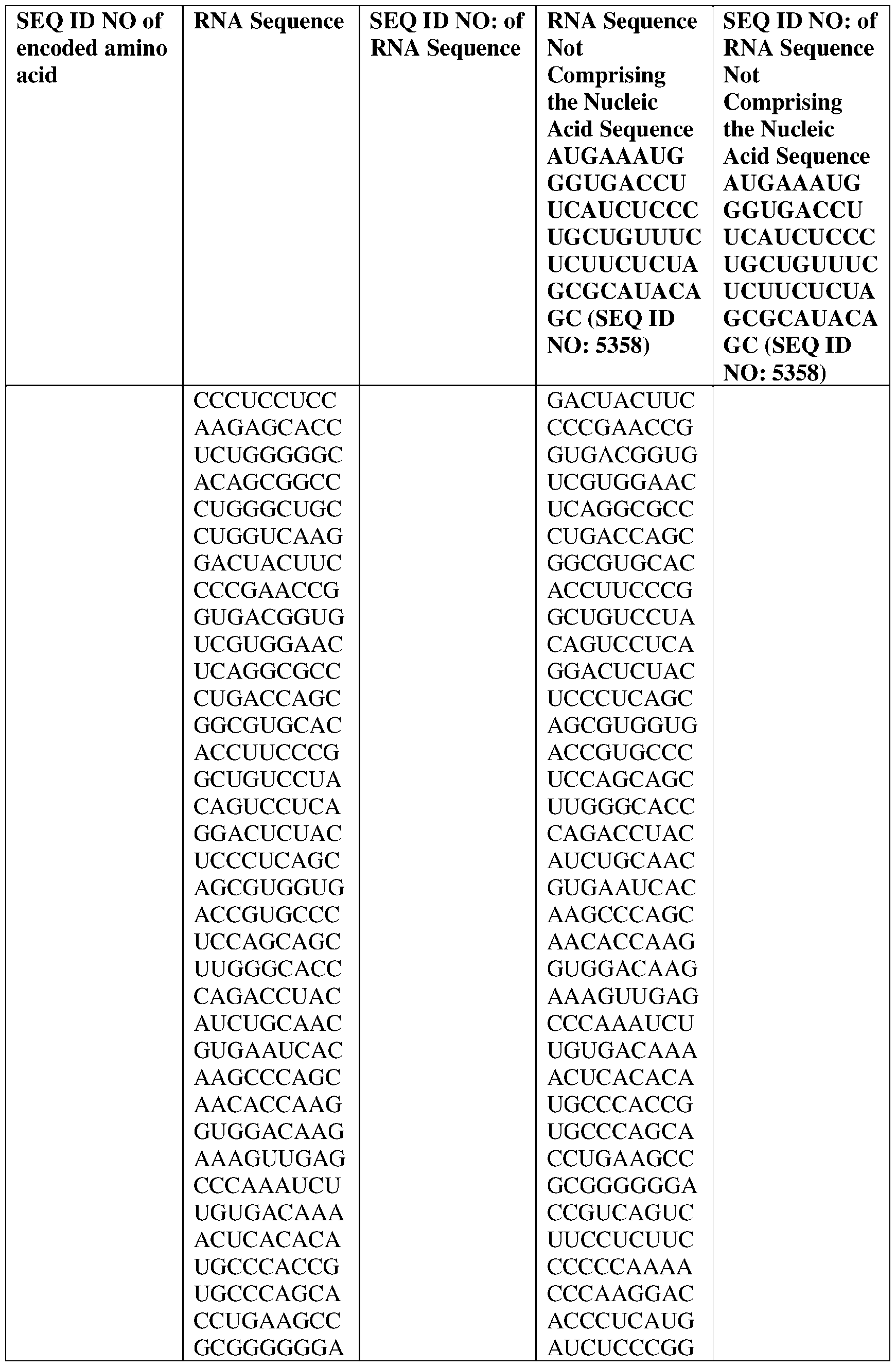
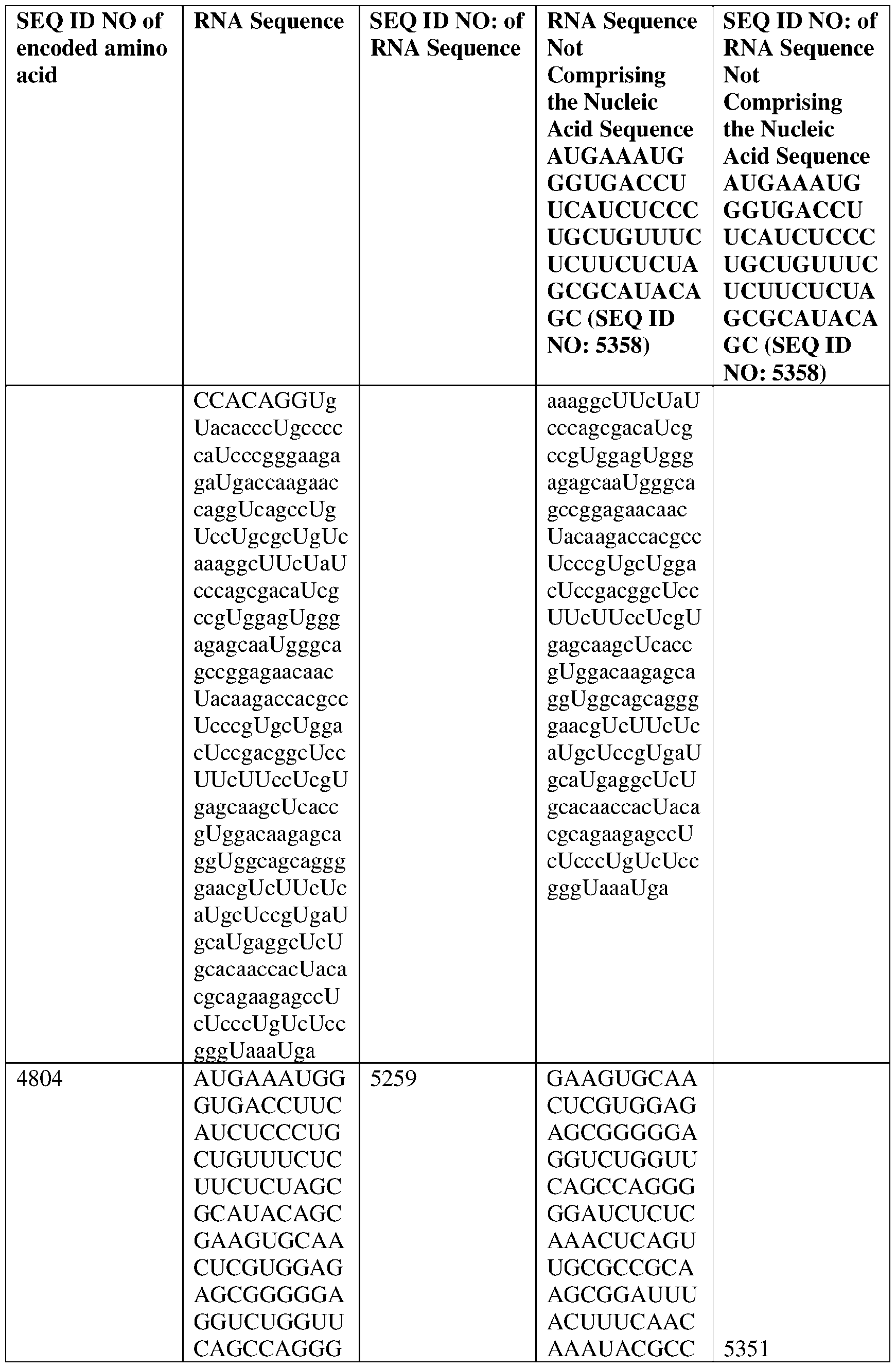
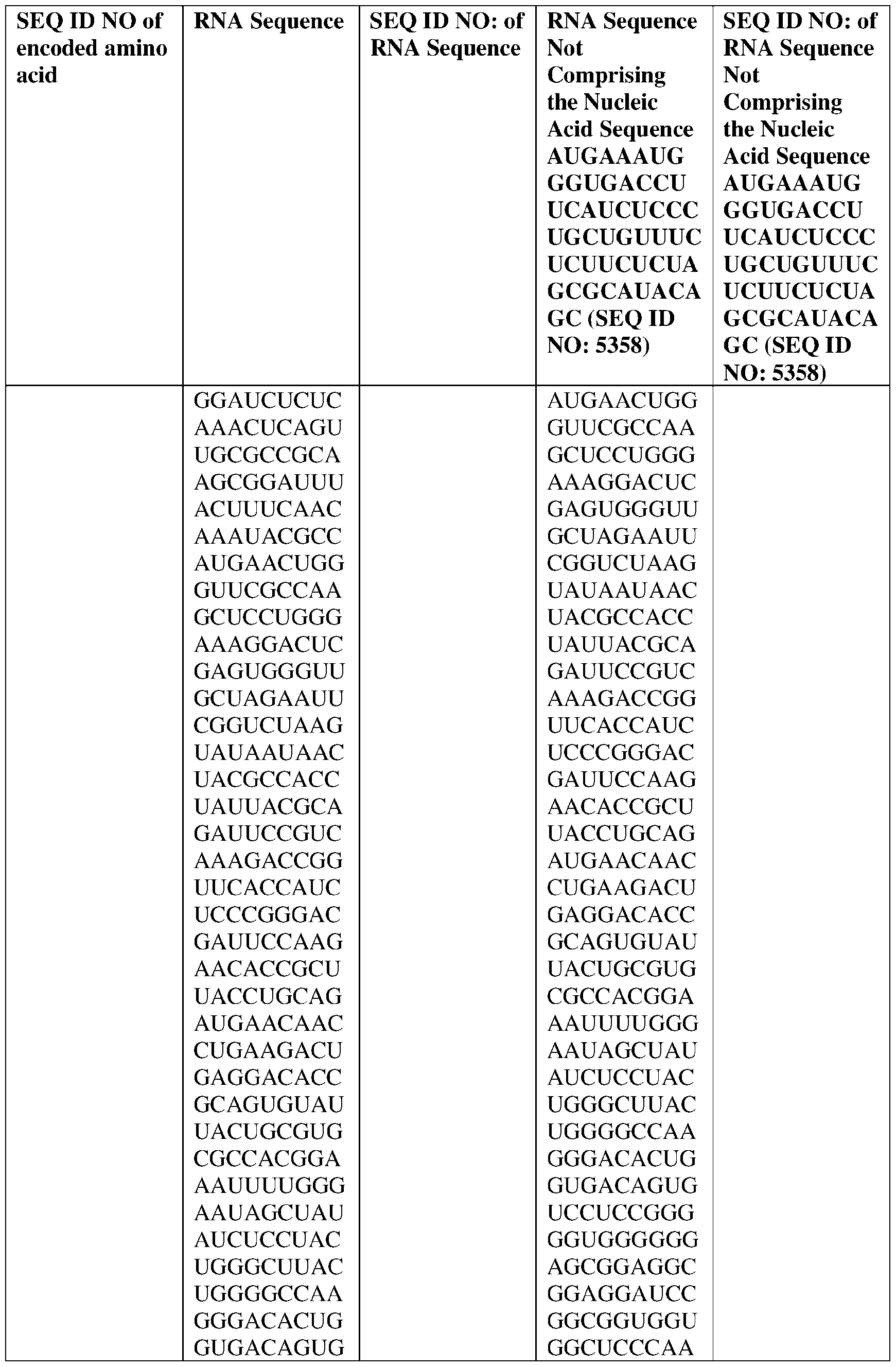
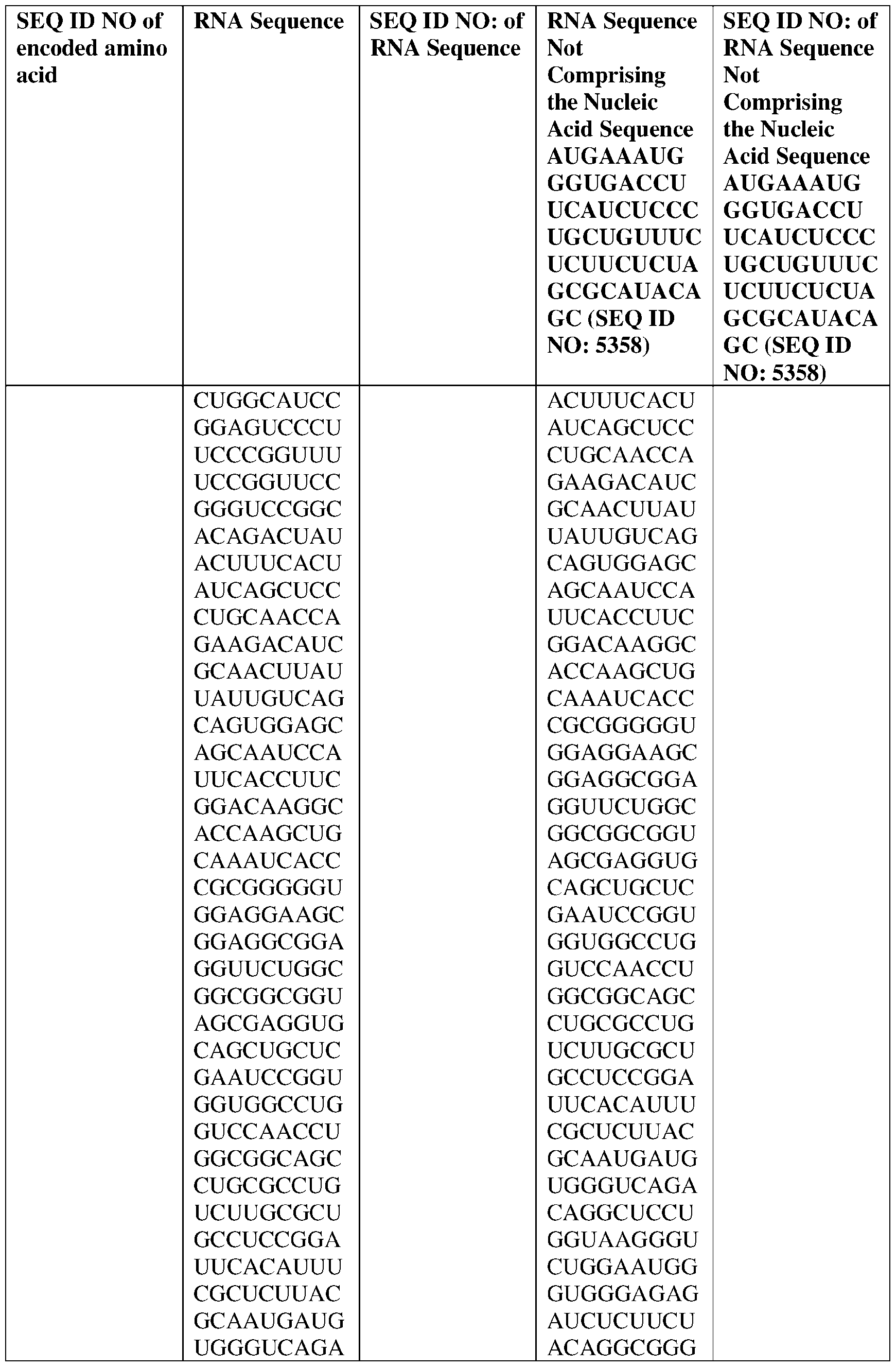
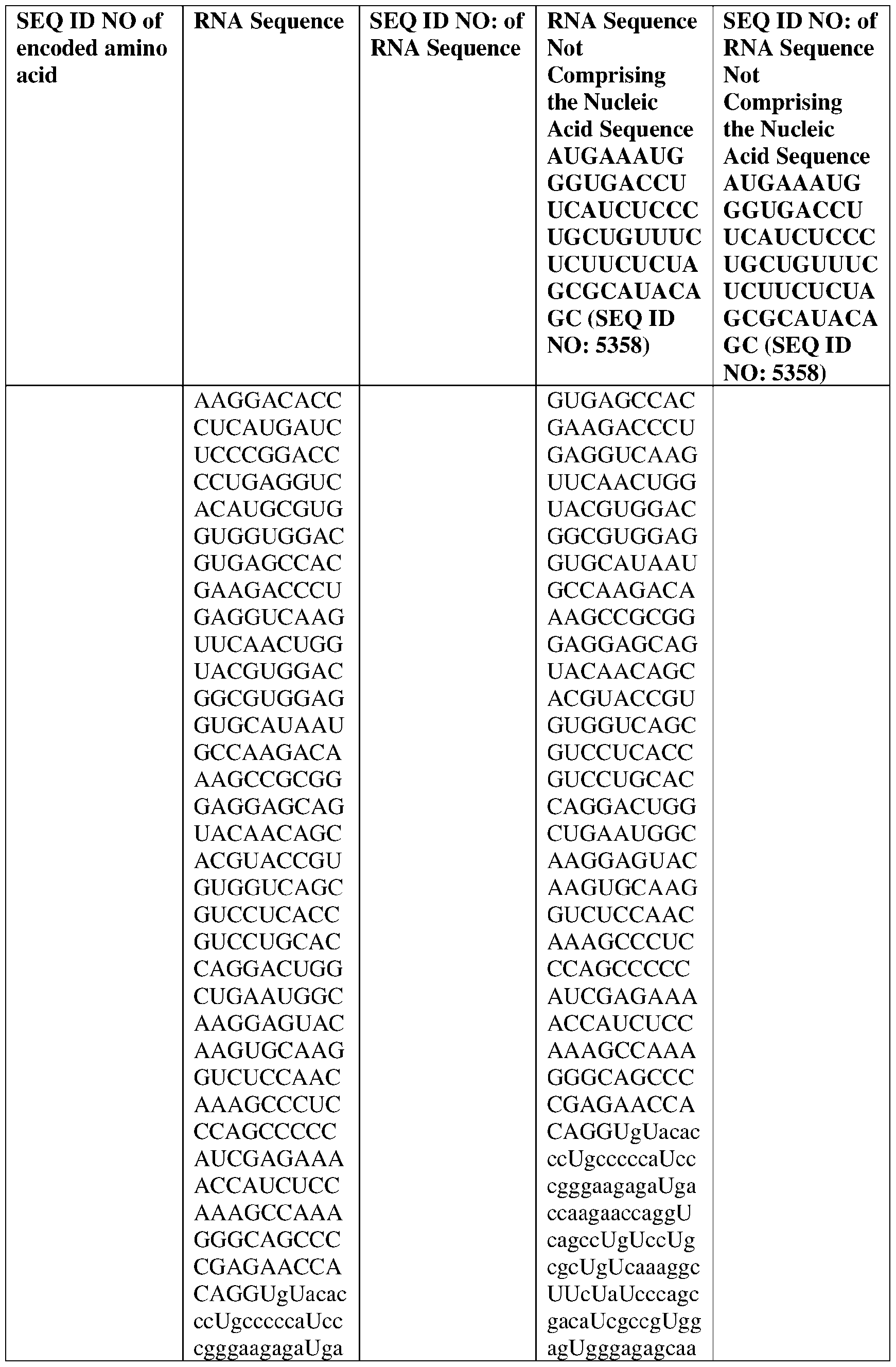
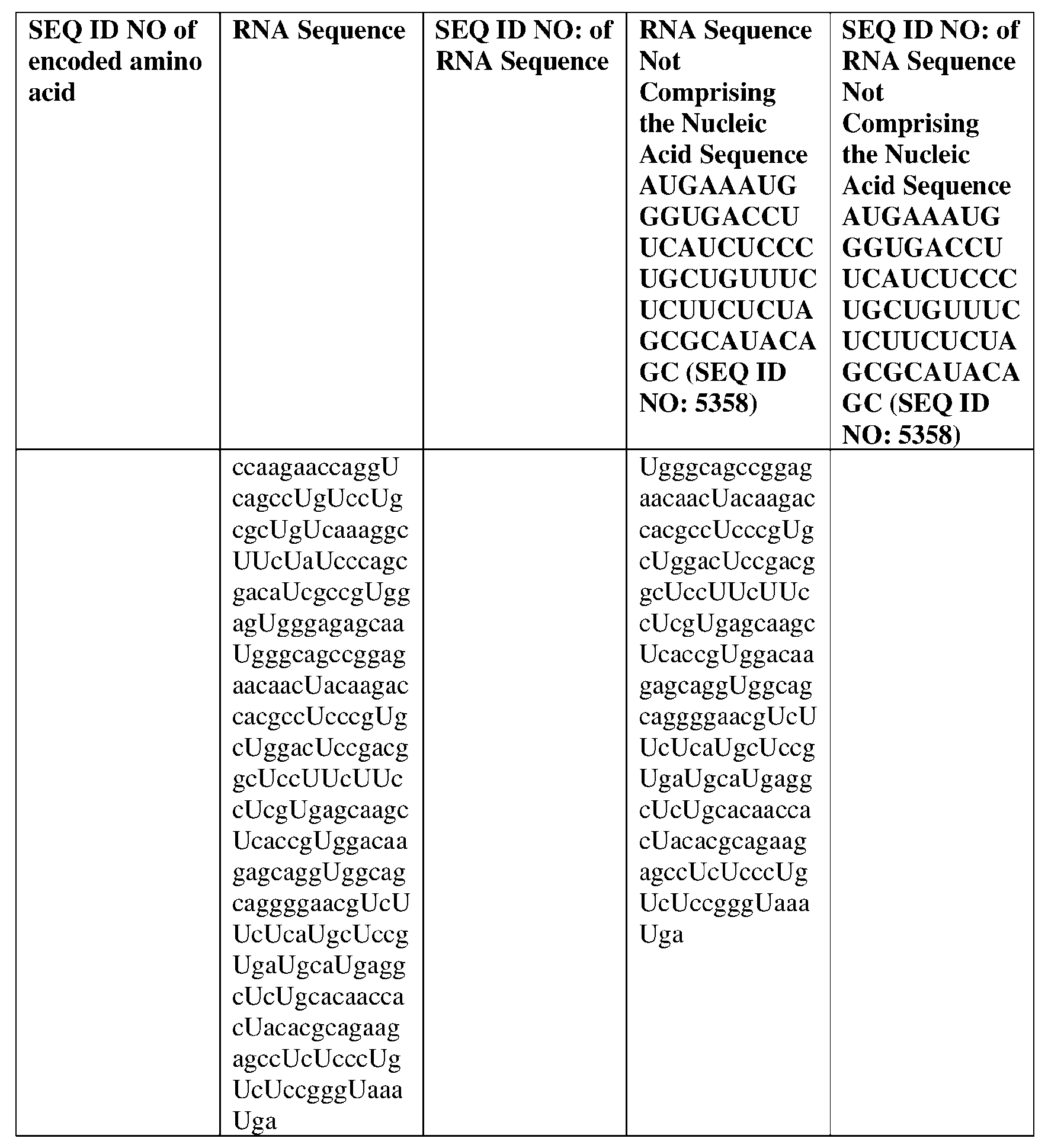
- the disclosure relates to a method of treating or preventing cancer comprising administering an effective amount of one or more nucleic acid molecules (e.g., RNA) described herein to a subject in need thereof.
- nucleic acid molecules e.g., RNA
- the cancer is selected from one or more of basal cell carcinoma, biliary tract cancer; bladder cancer; bone cancer; brain and central nervous system cancer; breast cancer; cancer of the peritoneum; cervical cancer; choriocarcinoma; colon and rectum cancer; connective tissue cancer; cancer of the digestive system; endometrial cancer; esophageal cancer; eye cancer; cancer of the head and neck; gastric cancer (including gastrointestinal cancer); glioblastoma; glioma; hepatic carcinoma; hepatoma; intra-epithelial neoplasm; kidney or renal cancer; larynx cancer; leukemia; liver cancer; lung cancer (e.g., small-cell lung cancer, non-small cell lung cancer, adenocarcinoma of the lung, and squamous carcinoma of the lung); melanoma; myeloma; neuroblastoma; oral cavity cancer (lip, tongue, mouth, and pharynx); ova
- the nucleic acid molecule comprises a sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity to any one of SEQ ID NOs: 5329-5355.
- FIGs. 1A-1F show images of illustrative bispecific formats, including a tandem scFv (FIG. 1A), a Hetero VH IgG (FIG. 1B), an scFv-Fab IgG (FIG. 1C), IgG-scFv (FIG. 1D), a IgG-(scFv) 2 (FIG. 1E), and IgG-(scFv) (FIG. 1F).
- FIG. 1A shows images of illustrative bispecific formats, including a tandem scFv (FIG. 1A), a Hetero VH IgG (FIG. 1B), an scFv-Fab IgG (FIG. 1C), IgG-scFv (FIG. 1D), a IgG-(scFv) 2 (FIG. 1E), and IgG-(scFv) (FIG. 1F).
- FIG. 2 is graph of the IM-16-44-09-B03-1CD3 antibody (Format 16 (scFv-Fab IgG)) binding to HEK 293F cells expressing human P2X3, CD3, or Claudin (CLDN) 18.2.
- FIG. 3 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 - or CLDN 18.2-expressing cells (HEK293F) in the presence of IMC-16-C2 (control) or IM-16-44-09-B03-1CD3 antibodies (Format 16 (scFv-Fab IgG)). Percent (%) specific cytotoxicity was assessed 48 hours (48hr) later.
- IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-16-44-09-B03-1CD3 on CLDN 18.1 (diamond), and IM-16-44-09-B03-1CD3 on CLDN 18.2 (triangle).
- FIGs. 4A, 4B, 4C, 4D, and 4E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IMC-16-C2 or IM-16-44-09-B03-1CD3 antibodies.
- FIG. 4A illustrates the results for IL-2 production
- FIG. 4B illustrates the results for IL-6 production
- FIG. 4C illustrates the results for IL- 10 production
- FIG. 4D illustrates the results for IFN- ⁇ (INFgamma) production
- FIG. 4E illustrates the results for TNF- ⁇ (TNFalpha) production.
- IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-16-44-09-B03-1CD3 on CLDN 18.1 (diamond), and IM-16-44-09-B03-1CD3 on CLDN 18.2 (triangle).
- FIGs. 5A and 5B show size exclusion chromatography (SEC) and differential scanning fluorimetry (DSF) experimental results for the IM-16-44-09-B03-1CD3, IM-16-44- 09-B03-2CD3, and IM-16-44-09-B03-3CD3 antibodies.
- FIG. 6 is graph of the IM-21-44-01 -E10-3CD3 antibody (Format 21 (IgG-(scFv) 2 )) binding to HEK 293F cells expressing human P2X3, CD3, or CLDN 18.2.
- FIG. 7 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IMC-16-C2 (control) or IM-21 -44-01-E10-3CD3 (Format 21 (IgG- (scFv) 2 )). Percent (%) specific cytotoxicity was assessed 48 hours (48hr) later.
- IMC- 16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-21-44-01 -E10- 3CD3 on CLDN 18.1 (diamond), and IM-21-44-01-E10-3CD3 on CLDN 18.2 (triangle).
- FIGs. 8A, 8B, 8C, 8D, and 8E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IMC-16-C2 or IM-21-44-01-E10-3CD3.
- FIG. 8A illustrates the results for IL-2 production
- FIG. 8B illustrates the results for IL-6 production
- FIG. 8C illustrates the results for IL- 10 production
- FIG. 8D illustrates the results for IFN- ⁇ production
- FIG. 8E illustrates the results for TNF- ⁇ production.
- IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-21-44-01-E10-3CD3 on CLDN 18.1 (diamond), and IM-21-44-01-E10-3CD3 on CLDN 18.2 (triangle).
- FIGs. 9 A and 9B show size exclusion chromatography (SEC) and differential scanning fluorimetry (DSF) experimental results for IM-21 -44-01 -E10-1CD3 and IM-21-44- 01 -E10-3CD3 antibodies.
- FIG. 10 is graph of the IM-21 -44-09-B03-3CD3 antibody (Format 21 (IgG-(scFv) 2 )) binding to HEK 293F cells expressing human P2X3, CD3, or CLDN 18.2.
- FIG. 11 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IMC-16-C2 (control) or IM-21 -44-09-B03-3CD3 (Format 21 (IgG- (scFv) 2 )). Percent (%) specific cytotoxicity was assessed 48 hours (48hr) later.
- FIGs. 12A, 12B, 12C, 12D, and 12E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IMC-16-C2 or IM-21-44-09-B03-3CD3.
- FIG. 12A illustrates the results for IL-2 production
- FIG. 12B illustrates the results for IL-6 production
- FIG. 12C illustrates the results for IL- 10 production
- FIG. 12D illustrates the results for IFN- ⁇ production
- FIG. 12E illustrates the results for TNF- a production.
- IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-21 -44-09-B03-3CD3 on CLDN 18.1 (diamond), and IM-21-44- 09-B03-3CD3 on CLDN 18.2 (triangle).
- FIGs. 13A and 13B show size exclusion chromatography (SEC) and differential scanning fluorimetry (DSF) experimental results for IM-21-44-09-B03-1CD3 and IM-21-44- 09-B03-3CD3 antibodies.
- FIG. 14 is graph of the IM-2-44-05-H06-1CD3 antibody (Format 2 (tandem scFv)) binding to HEK 293F cells expressing human P2X3, CD3, or CLDN 18.2.
- FIG. 15 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1- or CLDN 18.2-expressing HEK 293F cells in the presence of IMC-16-C2 (control) or IM-2-44-05-H06-1CD3 (Format 2 (tandem scFv)). Percent (%) specific cytotoxicity was assessed 48 hours (48hr) later.
- IMC-16-C2 on CLDN 18.1 circles
- IMC-16-C2 on CLDN 18.2 squares
- IM-2-44-05-H06-1CD3 on CLDN 18.1 (diamond)
- IM-2-44-05-H06-1CD3 on CLDN 18.2 (triangle).
- FIGs. 16A, 16B, 16C, 16D, and 16E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IMC-16-C2 or IM-2-44-05-H06-1CD3.
- FIG. 16A illustrates the results for IL-2 production
- FIG. 16B illustrates the results for IL-6 production
- FIG. 16C illustrates the results for IL- 10 production
- FIG. 16D illustrates the results for IFN- ⁇ production
- FIG. 16E illustrates the results for TNF- ⁇ production.
- IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-2-44-05-H06-1CD3 on CLDN 18.1 (diamond), and IM-2-44- 05-H06-1CD3 on CLDN 18.2 (triangle).
- FIG. 17 is graph of the IM-2-44-09-B03-1CD3 antibody (Format 2 (tandem scFv)) binding to HEK 293F cells expressing human P2X3, CD3, or CLDN 18.2.
- FIG. 18 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IMC-16-C2 (control) or IM-2-44-09-B03-1CD3 (Format 2 (tandem scFv)). Percent (%) specific cytotoxicity was assessed 48 hours (48hr) later.
- FIGs. 19A, 19B, 19C, 19D, and 19E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IMC-16-C2 or IM-2-44-09-B03-1CD3.
- FIG. 19A illustrates the results for IL-2 production
- FIG. 19B illustrates the results for IL-6 production
- FIG. 19C illustrates the results for IL- 10 production
- FIG. 19D illustrates the results for IFN- ⁇ production
- FIG. 19E illustrates the results for TNF- ⁇ production.
- IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-2-44-09-B03-1CD3 on CLDN 18.1 (diamond), and IM-2-44- 09-B03-1CD3 on CLDN 18.2 (triangle).
- FIG. 20 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IMC-16-C2 (control) antibodies.
- FIG. 21 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-42-44-09-B03-1CD3 (Format 42 EgG-(scFv)) antibodies.
- FIGs. 22A, 22B, 22C, 22D, and 22E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 09-B03-1CD3 antibodies.
- FIG. 22A illustrates the results for IL-2 production
- FIG. 22B illustrates the results for IL-6 production
- FIG. 22C illustrates the results for IL- 10 production
- FIG. 22D illustrates the results for IFN- ⁇ production
- FIG. 22E illustrates the results for TNF- ⁇ production.
- FIGs. 23A, 23B, 23C, 23D, and 23E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 09-B03-1CD3 antibodies.
- FIG. 23A illustrates the results for IL-2 production
- FIG. 23B illustrates the results for IL-6 production
- FIG. 23C illustrates the results for IL- 10 production
- FIG. 23D illustrates the results for IFN- ⁇ production
- FIG. 23E illustrates the results for TNF- ⁇ production.
- FIG. 24 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-42-44-09-B03-3CD3 antibodies (Format 42 IgG-(scFv)).
- FIGs. 25A, 25B, 25C, 25D, and 25E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 09-B03-1CD3.
- FIG. 25A illustrates the results for IL-2 production
- FIG. 25B illustrates the results for IL-6 production
- FIG. 25C illustrates the results for IL- 10 production
- FIG. 25D illustrates the results for IFN- ⁇ production
- FIG. 25E illustrates the results for TNF- ⁇ production.
- FIGs. 26A26B, 26C, 26D, and 26E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 09-B03-3CD3.
- FIG. 26A illustrates the results for IL-2 production
- FIG. 26B illustrates the results for IL-6 production
- FIG. 26C illustrates the results for IL- 10 production
- FIG. 26D illustrates the results for IFN- ⁇ production
- FIG. 26E illustrates the results for TNF- ⁇ production.
- FIG. 27 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-42-44-01-E10-1CD3 antibodies (Format 42 IgG-(scFv)).
- FIGs. 28A, 28B, 28C, 28D, and 28E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 01 -E10-1 CD3.
- FIG. 28A illustrates the results for IL-2 production
- FIG. 28B illustrates the results for IL-6 production
- FIG. 28C illustrates the results for IL- 10 production
- FIG. 28D illustrates the results for IFN- ⁇ production
- FIG. 28E illustrates the results for TNF- ⁇ production.
- FIGs. 29A, 29B, 29C, 29D, and 29E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 01 -E10-1 CD3.
- FIG. 29A illustrates the results for IL-2 production
- FIG. 29B illustrates the results for IL-6 production
- FIG. 29C illustrates the results for IL- 10 production
- FIG. 29D illustrates the results for IFN- ⁇ production
- FIG. 29E illustrates the results for TNF- ⁇ production.
- FIG. 30 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM -42-44-01 -E10-3CD3 antibodies (Format 42 IgG-(scFv)).
- FIGs. 31A, 31B, 31C, 31D, and 31E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 01-E10-3CD3.
- FIG. 31A illustrates the results for IL-2 production
- FIG. 31B illustrates the results for IL-6 production
- FIG. 31C illustrates the results for IL- 10 production
- FIG. 31D illustrates the results for IFN- ⁇ production
- FIG. 31E illustrates the results for TNF- ⁇ production.
- IM-42-44-01 -E10-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
- FIGs. 32A, 32B, 32C, 32D, and 32E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293 F cells with IM-42-44- 01 -E10-3CD3.
- FIG. 32A illustrates the results for IL-2 production
- FIG. 32B illustrates the results for IL-6 production
- FIG. 32C illustrates the results for IL- 10 production
- FIG. 32D illustrates the results for IFN- ⁇ production
- FIG. 32E illustrates the results for TNF- ⁇ production.
- FIG. 33 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-42-44-05-H06-1CD3 antibodies (Format 42 IgG-(scFv)).
- FIGs. 34A, 34B, 34C, 34D, and 34E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 05-H06-1CD3.
- FIG. 34A illustrates the results for IL-2 production
- FIG. 34B illustrates the results for IL-6 production
- FIG. 34C illustrates the results for IL- 10 production
- FIG. 34D illustrates the results for IFN- ⁇ production
- FIG. 34E illustrates the results for TNF- ⁇ production.
- FIGs. 35A, 35B, 35C, 35D, and 35E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 05 -H06-1CD3.
- FIG. 35 A illustrates the results for IL-2 production
- FIG. 35B illustrates the results for IL-6 production
- FIG. 35C illustrates the results for IL- 10 production
- FIG. 35D illustrates the results for IFN- ⁇ production
- FIG. 35E illustrates the results for TNF- ⁇ production.
- FIG. 36 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-42-44-05-H06-3CD3 antibodies (Format 42 IgG-(scFv)).
- FIGs. 37A, 37B, 37C, 37D, and 37E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 05-H06-3CD3.
- FIG. 37 A illustrates the results for IL-2 production
- FIG. 37B illustrates the results for IL-6 production
- FIG. 37C illustrates the results for IL- 10 production
- FIG. 37D illustrates the results for IFN- ⁇ production
- FIG. 37E illustrates the results for TNF- ⁇ production.
- IM-42-44-05-H06-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
- FIGs. 38A, 38B, 38C, 38D, and 38E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293 F cells with IM-42-44- 05-H06-3CD3.
- FIG. 38A illustrates the results for IL-2 production
- FIG. 38B illustrates the results for IL-6 production
- FIG. 38C illustrates the results for IL- 10 production
- FIG. 38D illustrates the results for IFN- ⁇ production
- FIG. 38E illustrates the results for TNF- ⁇ production.
- IM-42-44-05-H06-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
- FIG. 39 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM -21 -44-09-B03-3CD3 antibodies (Format 21 (IgG-(scFv) 2 )).
- FIGs. 40A, 40B, 40C, 40D, and 40E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-21 -44- 09-B03-3CD3 antibodies.
- FIG. 40A illustrates the results for IL-2 production
- FIG. 40B illustrates the results for IL-6 production
- FIG. 40C illustrates the results for IL- 10 production
- FIG. 40D illustrates the results for IFN- ⁇ production
- FIG. 40E illustrates the results for TNF- ⁇ production.
- FIGs. 41A, 41B, 41C, 41D, and 41E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-21 -44- 09-B03-3CD3.
- FIG. 41A illustrates the results for IL-2 production
- FIG. 41B illustrates the results for IL-6 production
- FIG. 41C illustrates the results for IL- 10 production
- FIG. 4 ID illustrates the results for IFN- ⁇ production
- FIG. 41E illustrates the results for TNF- ⁇ production.
- IM-21 -44-09-B03-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
- FIG. 42 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-21-44-01-E10-3CD3 antibodies (Format 21 (IgG-(scFv) 2 )).
- FIGs. 43A, 43B, 43C, 43D, and 43E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-21-44- 01 -E10-3CD3 antibodies.
- FIG. 43 A illustrates the results for IL-2 production
- FIG. 43B illustrates the results for IL-6 production
- FIG. 43C illustrates the results for IL- 10 production
- FIG. 43D illustrates the results for IFN- ⁇ production
- FIG. 43E illustrates the results for TNF- ⁇ production.
- FIGs. 44A, 44B, 44C, 44D, and 44E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293 F cells with IM-21-44- 01 -E10-3CD3.
- FIG. 44A illustrates the results for IL-2 production
- FIG. 44B illustrates the results for IL-6 production
- FIG. 44C illustrates the results for IL- 10 production
- FIG. 44D illustrates the results for IFN- ⁇ production
- FIG. 44E illustrates the results for TNF- ⁇ production.
- FIGs. 46A, 46B, 46C, 46D, and 46E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-2-44- 05-H06-1CD3 antibodies.
- FIG. 46A illustrates the results for IL-2 production
- FIG. 46B illustrates the results for IL-6 production
- FIG. 46C illustrates the results for IL- 10 production
- FIG. 46D illustrates the results for IFN- ⁇ production
- FIG. 46E illustrates the results for TNF- ⁇ production.
- FIGs. 47A, 47B, 47C, 47D, and 47E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-2-44- 05 -H06-1CD3.
- FIG. 47 A illustrates the results for IL-2 production
- FIG. 47B illustrates the results for IL-6 production
- FIG. 47C illustrates the results for IL- 10 production
- FIG. 47D illustrates the results for IFN- ⁇ production
- FIG. 47E illustrates the results for TNF-a production.
- FIG. 48 is a set of graphs of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of mRNA encoding IM-2-44-09B03-1CD3 or IM-16-44-09B03- 1CD3 antibodies.
- the present embodiments are based, in part, on the surprising discovery of antibodies specific for Claudin 18.2 and CD3.
- the present disclosure describes, in part, the isolation and characterization of antibodies, antibody fragments, and antibody variants specific for Claudin 18.2 and CD3.
- the antibody is a tandem single-chain variable fragment (scFv) specific for Claudin 18.2 and CD3, which is for example, illustrated in FIG. 1A.
- the antibody is a hetero VH IgG, such as illustrated in FIG. 1B and specific for Claudin 18.2 and CD3.
- the antibody is an scFv-Fab IgG antibody format, which, for example, is illustrated in FIG.
- the antibody is an IgG-scFv format, which, for example, is illustrated in FIG. 1D and specifically binds to Claudin 18.2 and CD3.
- the antibody is an IgG-(scFv) 2 antibody format, which, for example, is illustrated in FIG. 1E and specifically binds to Claudin 18.2 and CD3.
- the antibody is a IgG-(scFv) format, which, for example, is illustrated in FIG. 1F and specifically binds to Claudin 18.2 and CD3.
- the antibody specific for Claudin 18.2 and CD3 binds Claudin 18.2 and CD3 contemporaneously.
- the bispecific antibody will bind to a tumor cell expressing Claudin 18.2 and an immune cell, such as a cytotoxic T cell, expressing CD3. By bringing the cells together, the T cell is activated and targets the tumor cell expressing Claudin 18.2 for destruction.
- the antibody specific for Claudin 18.2 and CD3 is a tandem scFv.
- An exemplary tandem scFv is shown schematically in FIG. 1A.
- a tandem scFv antibody specific for Claudin 18.2 and CD3 can be selected from an amino acid sequence of SEQ ID NO: 4816 and SEQ ID NO: 4814, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the antibody specific for Claudin 18.2 and CD3 is a hetero VH IgG antibody comprising two heavy chains and two light chains.
- the light chains optionally can comprise the same amino acid sequence.
- An exemplary hetero VH IgG is shown schematically in FIG. 1B.
- a hetero VH IgG antibody specific for Claudin 18.2 and CD3 can comprise a first heavy chain having an amino acid sequence of SEQ ID NO: 3869, SEQ ID NO: 4767, or SEQ ID NO: 4768 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a second heavy chain having an amino acid sequence comprising the heavy chain CDRs of any anti-CD3 scFv provided herein.
- the light chain comprises an amino acid sequence of SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the antibody specific for Claudin 18.2 and CD3 is an scFv-Fab IgG antibody comprising a first heavy chain, a second heavy chain, and a light chain.
- An exemplary scFv-Fab IgG antibody is shown schematically in FIG. 1C.
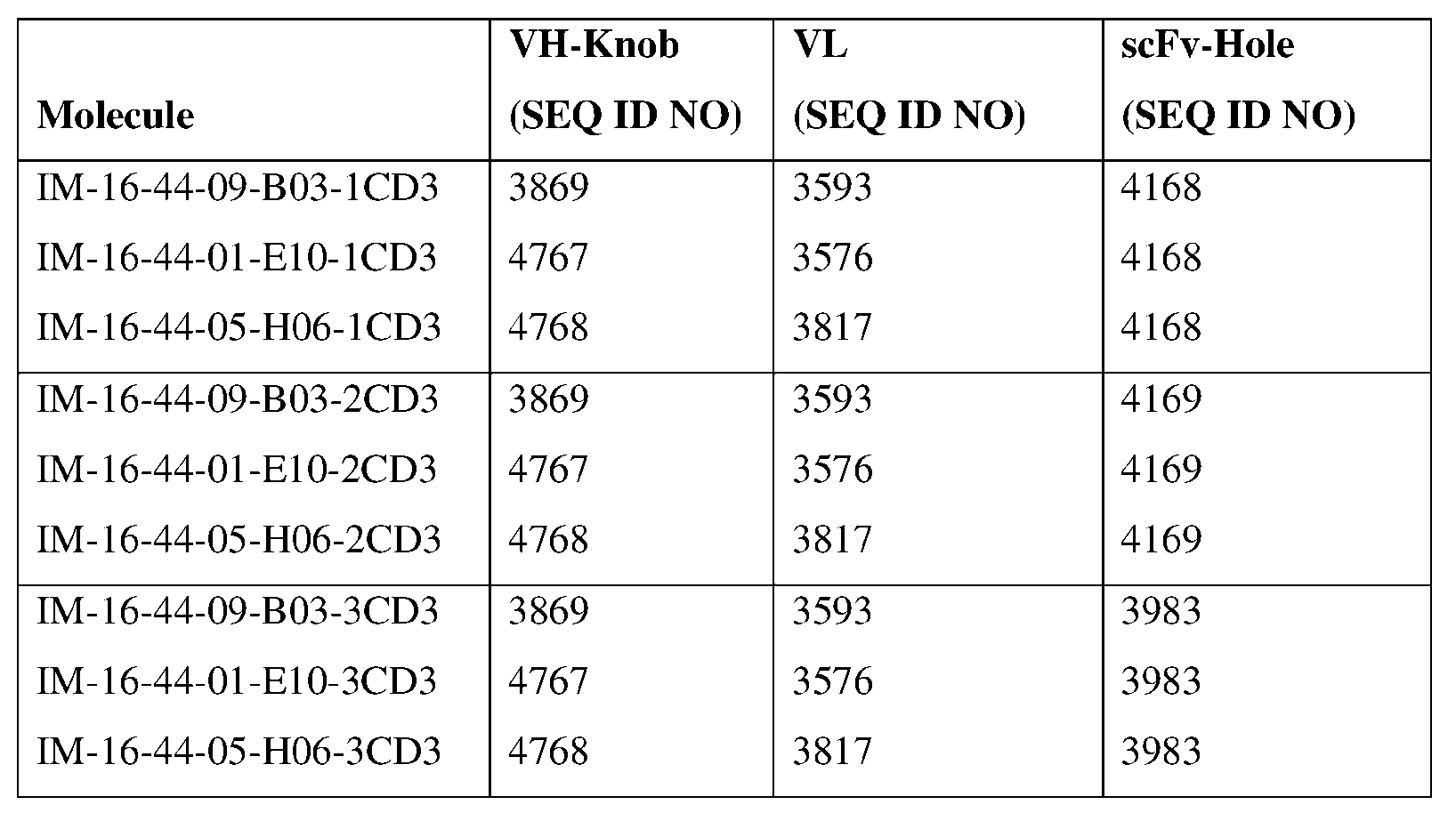
- An antibody specific for Claudin 18.2 and CD3 can comprise a first heavy chain having an amino acid sequence of SEQ ID NO: 3869, SEQ ID NO: 4767, or SEQ ID NO: 4768, a second heavy chain having an amino acid sequence of SEQ ID NO: 4168, SEQ ID NO: 4169, or SEQ ID NO: 3983, and a light chain having an amino acid sequence of SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817.
- the first heavy chain comprises an amino acid sequence of SEQ ID NO: 3869, SEQ ID NO: 4767, or SEQ ID NO: 4768, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the second heavy comprises an amino acid sequence of SEQ ID NO: 4168, SEQ ID NO: 4169, or SEQ ID NO: 3983, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the light chain comprises an amino acid sequence of SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the antibody specific for Claudin 18.2 and CD3 is a IgG-scFv antibody comprising two heavy chains (a first and second heavy chain) and two light chains.
- the two heavy chains comprising amino acid sequences that are different from each other, and the two light chains comprise amino acid sequences that are identical.
- An exemplary IgG-scFv is shown schematically in FIG. 1D.
- the first heavy chain has an amino acid sequence of SEQ ID NO: 3869, SEQ ID NO: 4767, or SEQ ID NO: 4768 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the second heavy chain has an amino acid sequence of SEQ ID NO: 4778, SEQ ID NO: 4779, SEQ ID NO: 4780, SEQ ID NO: 4803, SEQ ID NO: 4804, SEQ ID NO: 4805, SEQ ID NO: 4806, SEQ ID NO: 4807, SEQ ID NO: 4808 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 3869 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a second heavy chain selected from SEQ ID NO: 4778, SEQ ID NO: 4803, and SEQ ID NO: 4806 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto
- the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 4767 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a second heavy chain selected from SEQ ID NO: 4779, SEQ ID NO: 4804, and SEQ ID NO: 4807 or an amino acid sequence having at least about 90%, about 93%,
- the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 4768 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a second heavy chain selected from SEQ ID NO: 4780, SEQ ID NO: 4805, and SEQ ID NO: 4808 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the two light chains can be selected from SEQ ID NO: 3593, SEQ ID NO: 3576, and SEQ ID NO: 3817 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the two light chains can comprise the same amino acid sequence.
- the antibody specific for Claudin 18.2 and CD3 is an IgG- (SCFV) 2 antibody comprising two heavy chains and two light chains, wherein the heavy chains are identical to each other and the light chains are identical to each other.
- IgG- (SCFV) 2 antibody is shown schematically in FIG. 1E.
- the antibody comprises a heavy chain selected comprising an amino acid sequence of SEQ ID NO: 4769, SEQ ID NO: 4770, SEQ ID NO: 4771, SEQ ID NO: 4772, SEQ ID NO: 4773, SEQ ID NO: 4774, SEQ ID NO: 4775, SEQ ID NO: 4776, or SEQ ID NO: 4777, and a light chain selected from an amino acid sequence of SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817.
- the heavy chain comprises an amino acid sequence of SEQ ID NO: 4769, SEQ ID NO: 4770, SEQ ID NO: 4771, SEQ ID NO: 4772, SEQ ID NO: 4773, SEQ ID NO: 4774, SEQ ID NO: 4775, SEQ ID NO: 4776, or SEQ ID NO: 4777,, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the light chain comprises an amino acid sequence of SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the antibody specific for Claudin 18.2 and CD3 is a IgG- (scFv) antibody comprising two heavy chains (a first and second heavy chain) and two light chains.
- the two heavy chains comprising amino acid sequences that are different from each other, and the two light chains comprise amino acid sequences that are identical.
- An exemplary IgG-scFv is shown schematically in FIG. 1E.
- the first heavy chain has an amino acid sequence of SEQ ID NO: 3869, SEQ ID NO: 4767, or SEQ ID NO: 4768, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
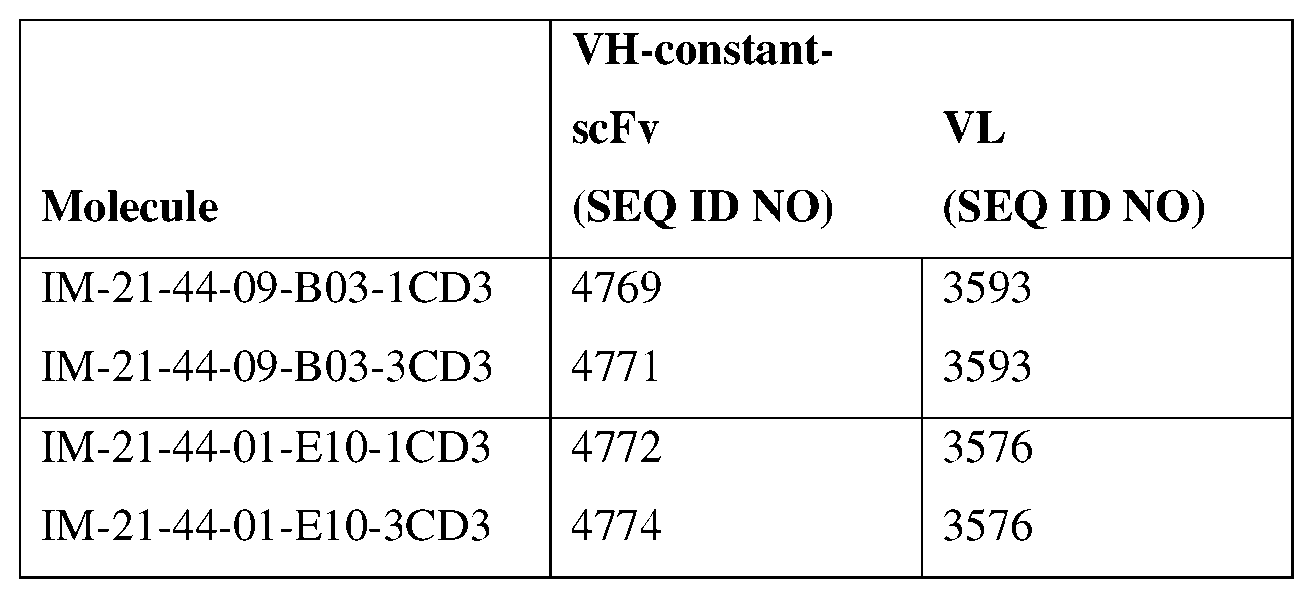
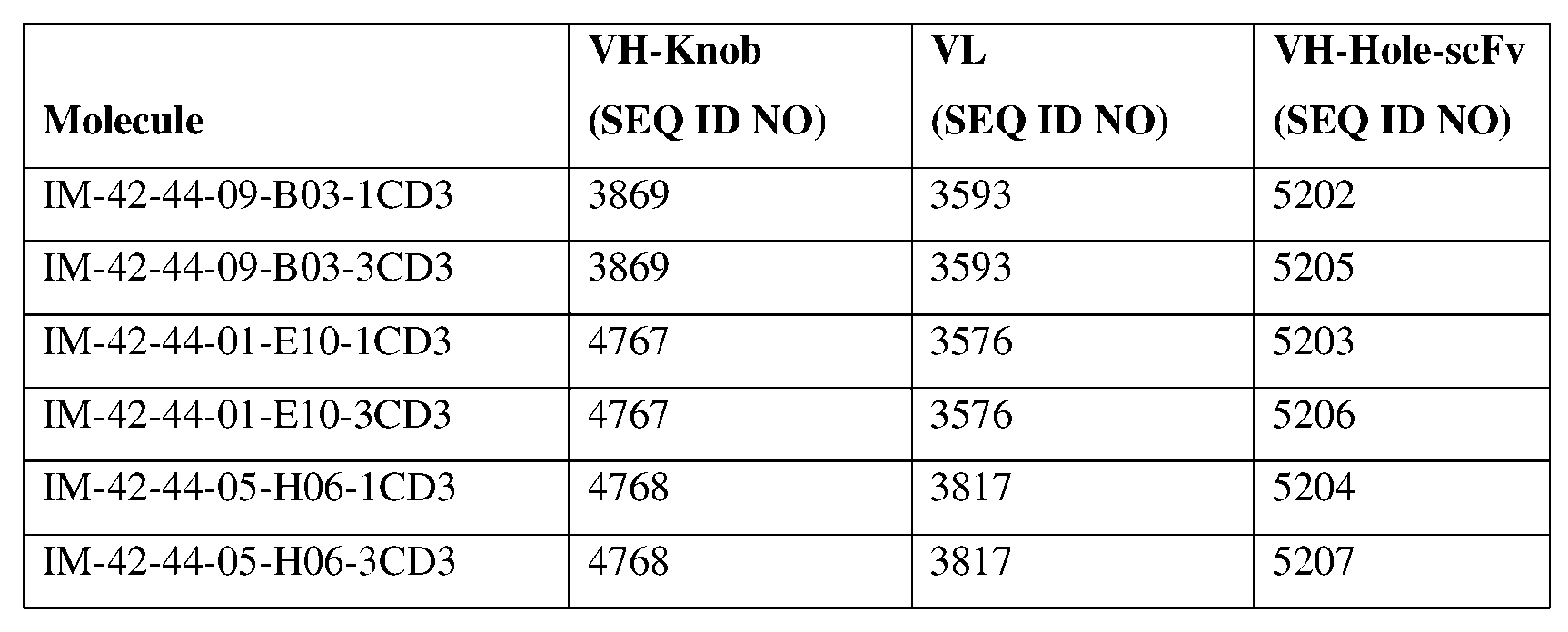
- the second heavy chain has an amino acid sequence of SEQ ID NO: 5202, SEQ ID NO: 5205, SEQ ID NO: 5203, SEQ ID NO: 5206, SEQ ID NO: 5204, or SEQ ID NO: 5207, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the two light chains each have an amino acid sequence comprising SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the two light chains can comprise the same amino acid sequence.
- the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 3869 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, a second heavy chain having an amino acid sequence of SEQ ID NO: 5202 or 5205 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a light chain (e.g., a first and second light chain) having an amino acid sequence of SEQ ID NO: 3593 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- a light chain e.g., a first and second light chain
- the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 4767 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, a second heavy chain having an amino acid sequence of SEQ ID NO: 5203 or SEQ ID NO: 5206 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a light chain (e.g., a first and second light chain) having an amino acid sequence of SEQ ID NO: 3576 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- a light chain e.g., a first and second light chain
- the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 4768 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a second heavy chain having an amino acid sequence of SEQ ID NO: 5204 or 5207 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a light chain having an amino acid sequence of SEQ ID NO: 3817 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
- the antibody e.g., a tandem scFv antibody, a hetero VH IgG antibody, an scFv-Fab IgG antibody, an IgG-scFv antibody, IgG-(scFv)? antibody, and a IgG- (scFv) antibody
- the antibody may comprise an amino acid sequence having one or more amino acid mutations (e.g., substitutions or deletions) relative to any of the sequences disclosed herein.
- the one or more amino acid mutations may be independently selected from substitutions, insertions, deletions, and truncations.
- the antibody e.g., a tandem scFv antibody, a hetero VH IgG antibody, an scFv-Fab IgG antibody, an IgG-scFv antibody, IgG-(scFv) 2 antibody, and a IgG- (scFv) antibody
- the antibody comprises a sequence that has about 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 or more amino acid mutations with respect to any one of the amino acid sequences disclosed herein.
- the antibody or antibody format (e.g. , a tandem scFv antibody, a hetero VH IgG antibody, an scFv-Fab IgG antibody, an IgG-scFv antibody, IgG- (SCFV) 2 antibody, and a IgG-(scFv) antibody), or fragment thereof, or variant thereof, may comprise an amino acid sequence having al least about 50%, about 60%, about 70%, about 80%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99% sequence homology to the amino acid sequences disclosed herein.
- the present invention provides for variants or fragments comprising any of the sequences described herein, for instance, a sequence having at least about 60%, or at least about 61%, or at least about 62%, or at least about 63%, or at least about 64%, or at least about 65%, or at least about 66%, or at least about 67%, or at least about 68%, or at least about 69%, or at least about 70%, or at least about 71%, or at least about 72%, or at least about 73%, or at least about 74%, or at least about 75%, or at least about 76%, or at least about 77%, or at least about 78%, or at least about 79%, or at least about 80%, or at least about 81%, or at least about 82%, or at least about 83%, or at least about 84%, or at least about 85%, or at least about 86%, or at least about 87%, or at least about 88%, or at least about 89%, or at least about 90%, or at least about 9
- variants are those that have conservative amino acid substitutions made at one or more predicted non-essential amino acid residues.
- a “conservative amino acid substitution” is one in which the amino acid residue is replaced with an amino acid residue having a similar side chain.
- the amino acid mutations are amino acid substitutions, and may include conservative and/or non-conservative substitutions.
- “Conservative substitutions” may be made, for instance, on the basis of similarity in polarity, charge, size, solubility, hydrophobicity, hydrophilicity, and/or the amphipathic nature of the amino acid residues involved.
- the 20 naturally occurring amino acids can be grouped into the following six standard amino acid groups: (1) hydrophobic: Met, Ala, Vai, Leu, Ile; (2) neutral hydrophilic: Cys, Ser, Thr; Asn, Gin; (3) acidic: Asp, Glu; (4) basic: His, Lys, Arg; (5) residues that influence chain orientation: Gly, Pro; and (6) aromatic: Trp, Tyr, Phe.
- “conservative substitutions” are defined as exchanges of an amino acid by another amino acid listed within the same group of the six standard amino acid groups shown above. For example, the exchange of Asp by Glu retains one negative charge in the so modified polypeptide.
- glycine and proline may be substituted for one another based on their ability to disrupt ⁇ -helices.
- non-conservative substitutions are defined as exchanges of an amino acid by another amino acid listed in a different group of the six standard amino acid groups (1) to (6) shown above.
- antibody refers to a broad sense and includes immunoglobulin or antibody molecules including polyclonal antibodies, monoclonal antibodies including murine, human, humanized and chimeric monoclonal antibodies and antibody fragments, such as ScFv (PLOS Biology
- antibodies are proteins or polypeptides that exhibit binding specificity to a specific antigen.
- Intact antibodies are heterotetrameric glycoproteins, composed of two identical light chains and two identical heavy chains. Typically, each light chain is linked to a heavy chain by one covalent disulfide bond, while the number of disulfide linkages varies between the heavy chains of different immunoglobulin isotypes. Each heavy and light chain also has regularly spaced intrachain disulfide bridges. Each heavy chain has at one end a variable domain (VH) followed by a number of constant domains.
- VH variable domain
- Each light chain has a variable domain at one end (VL) and a constant domain at its other end; the constant domain of the light chain is aligned with the first constant domain of the heavy chain and the light chain variable domain is aligned with the variable domain of the heavy chain.
- Antibody light chains of any vertebrate species can be assigned to one of two clearly distinct types, namely kappa and lambda, based on the amino acid sequences of their constant domains.
- Immunoglobulins can be assigned to five major classes, namely IgA, IgD, IgE, IgG and IgM, depending on the heavy chain constant domain amino acid sequence.
- IgA and IgG are further sub-classified as the isotypes IgAl, IgA2, IgGl, IgG2, IgG3 and IgG4.
- antibody fragment refers to an intact antibody, generally the antigen binding or variable region of the intact antibody.
- antibody fragments include Fab, Fab', F(ab')2 and Fv fragments, diabodies, single chain antibody molecules and multispecific antibodies formed from at least two intact antibodies or fragments thereof.
- antigen refers to any molecule that has the ability to generate antibodies either directly or indirectly.
- telomere binding refers to antibody binding to a predetermined antigen (e.g., Claudin 18.2) or epitope present on the antigen.
- the antibody binds with a dissociation constant (KD) of about 10 -10 M or less, of about 10 -9 M or less, of about 10 -8 M or less, of about 10 -7 M or less, of about 10 -6 M or less, of about 10 -5 M or less, and binds to the predetermined antigen with a KD that is at least two-fold less than its KD for binding to a non-specific antigen (e.g., BSA, casein, or another non-specific polypeptide) other than the predetermined antigen.
- KD dissociation constant
- an antibody recognizing Claudin 18.2 and “an antibody specific for Claudin 18.2” are used interchangeably herein with the term “an antibody which binds immunospecifically to Claudin 18.2.” Reference in the present disclosure may be made to Claudin 18.2. In some embodiments, the antibody is specific for Claudin 18.2 and does not specifically bind to Claudin 3, Claudin 4, and/or Claudin 9.
- CDRs are referred to as the complementarity determining region amino acid sequences of an antibody which are the hypervariable regions of immunoglobulin heavy and light chains. See, e.g., Kabat et al., Sequences of Proteins of Immunological Interest, 4th ed., U.S. Department of Health and Human Services, National Institutes of Health (1987). There are three heavy chain and three light chain CDRs or CDR regions in the variable portion of an immunoglobulin. Thus, “CDRs” as used herein refers to all three heavy chain CDRs, or all three light chain CDRs or both all heavy and all light chain CDRs, if appropriate.
- Each variable region comprises three hypervariable regions also known as complementarity determining regions (CDRs) flanked by four relatively conserved framework regions (FRs).
- CDR1, CDR2, and CDR3 contribute to the antibody binding specificity, as the CDRs provide the majority of contact residues for the binding of the antibody to the antigen or epitope.
- CDRs of interest can be derived from donor antibody variable heavy and light chain sequences, and include analogs of the naturally occurring CDRs, which analogs also share or retain the same antigen binding specificity and/or neutralizing ability as the donor antibody from which they were derived.
- the antibody is a chimeric antibody.
- the antibody is a humanized antibody.
- CDRs are based on sequence variability (Wu and Kabat, J. Exp. Med. 132:211-250, 1970). There are six CDRs— three in the variable heavy chain, or VH, and are typically designated H-CDR1, H-CDR2, and H-CDR3, and three CDRs in the variable light chain, or VL, and are typically designated L-CDR1 , L-CDR2, and L-CDR3 (Kabat et al. , Sequences of Proteins of Immunological Interest, Sth Ed. Public Health Service, National Institutes of Health, Bethesda, Md., 1991).
- HVR human variable region
- HV human variable domain
- Chothia and Lesk Chothia and Lesk, Mol. Biol. 196:901-917, 1987.
- an “isolated antibody,” as used herein, refers to an antibody that is substantially free of other antibodies having different antigenic specificities (e.g., an isolated antibody that specifically binds Claudin 18.2 is substantially free of antibodies that specifically bind antigens other than Claudin 18.2). Moreover, an isolated antibody may be substantially free of other cellular material and/or chemicals. An isolated antibody can also be sterile or pyrogen free or formulated as injectable pharmaceutical as described herein.
- the source for the DNA encoding a non-human antibody include cell lines which produce antibody, such as hybrid cell lines commonly known as hybridomas.
- the antibody or antibody fragment or variant disclosed herein comprises a CD3 engager.
- the CD3 engager can be selected from muOKT3, huOKT3, huSP34, huUCHTl and a CD3 nanobody (VHH).
- the antibody or antibody fragment or variant is an antibody provided in TABLES 1-5, or an antibody fragment or variant thereof having at least about 60%, or at least about 61%, or at least about 62%, or at least about 63%, or at least about 64%, or at least about 65%, or at least about 66%, or at least about 67%, or at least about 68%, or at least about 69%, or at least about 70%, or at least about 71%, or at least about 72%, or at least about 73%, or at least about 74%, or at least about 75%, or at least about 76%, or at least about 77%, or at least about 78%, or at least about 79%, or at least about
- the leader sequence such as the amino acid sequence of SEQ ID NO: 5356
- the leader sequence of SEQ ID NO: 5356 can also be replaced with a different leader sequence to facilitate expression and/or secretion of the antibody.
- the antibody in a composition such as a pharmaceutical composition, does not comprise a leader sequence, including, but not limited to the amino acid sequence of SEQ ID NO: 5356.
- the antibody or antibody fragment or variant is an antibody provided in TABLES 1-5, wherein the first heavy chain, the second heavy chain, the light chain, and/or the single-chain variable fragment does not comprise the signal sequence of amino acids 1-18 relative to the respective SEQ ID NO (e.g., in some embodiments, a fragment of the antibody or variant comprises a sequence provided in TABLE A).
- the antibody or antibody fragment or variant is an antibody provided in TABLES 1-5, wherein the first heavy chain, the second heavy chain, the light chain, and/or the single-chain variable fragment does not comprise the amino acid sequence MKWVTFISLLFLFSSAYS (SEQ ID NO: 5356) (e.g., in some embodiments, a fragment of the antibody or variant comprises a sequence provided in TABLE A).
- the disclosure relates to a composition
- a composition comprising an scFv-Fab IgG antibody specific for Claudin 18.2 and CD3 comprising a first heavy chain, a second heavy chain, and a light chain, wherein:
- the first heavy chain is selected from:
- QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
- CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, or (it)
- QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4767), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, or (iii)
- VLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4168), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof
- KLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4169), o an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain
- GGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTEC S (SEQ ID NO: 3576), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSA
- the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof
- the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof
- the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or
- AGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or wherein: any of the above sequences in subsections (a) optionally can have
- the first heavy chain comprises:
- CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and
- the second heavy chain is selected from:
- the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof
- the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof
- the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof
- the heavy chain CDR1 sequence GFTFSSYGMGWV SEQ ID NO: 1 or a variant thereof
- the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof
- the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
- PIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPE NNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS PGK (SEQ ID NO: 4806); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR
- the first heavy chain comprises:
- CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof
- the second heavy chain is selected from:
- LHNHYTQKSLSLSPGK (SEQ ID NO: 4804), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNS Y1SYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP
- SPGK (SEQ ID NO: 4807); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof,
- the first heavy chain comprises:
- the second heavy chain is selected from:
- RWVFGGGTKLTVL (SEQ ID NO: 4769), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
- the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof
- the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof
- the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof
- the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof
- the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof
- the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant
- the heavy chain CDR1 sequence is GFTFSS Y GMGWV (SEQ ID NO: 1) or a variant thereof
- the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof
- the heavy chain CDR3 sequence is
- CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, (iv)
- the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof
- the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof
- the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof
- the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof
- the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or
- GGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLA SGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTFGQGTKLQITR (SEQ ID NO: 4776), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and C
- SCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAK NTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 4777), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein
- the first heavy chain comprises:
- QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
- CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and
- the second heavy chain is selected from:
- NRWVFGGGTKLTVL (SEQ ID NO: 5202), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (S
- the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof
- the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof
- the heavy chain CDR3 sequence is
- CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, or
- the first heavy chain comprises:
- FTISRDNAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 5206) or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO
- the first heavy chain comprises:
- the second heavy chain is selected from:
- PRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFG GGTKLTVL (SEQ ID NO: 5204), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO:
- TKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKA GVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3593), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG(SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ
- CVLWYSNRWVFGGGTKLTVL (SEQ ID NO. 4814), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or a variant thereof, the CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or a variant thereof, and the CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or a variant thereof, and/or a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the CDR3 sequence is CAKDASNG
- the antibody or antibody fragment or variant disclosed herein comprise a linker having one or more glycines and serines replaced with a functionally equivalent variation thereof.
- the linker is identified in the underlined text above.
- the linker is selected from the amino acid sequence of GGGGSGGGGSGGGGS (SEQ ID NO: 50), GGGGS (SEQ ID NO: 51), GGSGGSGGSGGSGGVD (SEQ ID NO: 52), and GKPGSGKPGSGKPGS (SEQ ID NO: 53).
- the linker may be derived from naturally-occurring multidomain proteins or are empirical linkers as described, for example, in Chichili et al., (2013), Protein Sci. 22(2): 153- 167, Chen et al., (2013), Adv Drug Deliv Rev. 65(10): 1357-1369, the entire contents of which are hereby incorporated by reference.
- the linker may be designed using linker designing databases and computer programs such as those described in Chen et al., (2013), Adv Drug Deliv Rev. 65(10):1357-1369 and Crasto et. al., (2000), Protein Eng. 13(5):309-312, the entire contents of which are hereby incorporated by reference.
- the linker is as provided in Moore et al., (2019), Methods 154:38-50, the entire contents of which are hereby incorporated by reference.
- the linker is a polypeptide. In some embodiments, the linker is less than about 100 amino acids long. For example, the linker may be less than about 100, about 95, about 90, about 85, about 80, about 75, about 70, about 65, about 60, about 55, about 50, about 45, about 40, about 35, about 30, about 25, about 20, about 19, about 18, about 17, about 16, about 15, about 14, about 13, about 12, about 11, about 10, about 9, about 8, about 7, about 6, about 5, about 4, about 3, or about 2 amino acids long. In some embodiments, the linker is flexible. In another embodiment, the linker is rigid. In various embodiments, the linker is substantially comprised of glycine and serine residues (e.g. about 30%, or about 40%, or about 50%, or about 60%, or about 70%, or about 80%, or about 90%, or about 95%, or about 97% glycines and serines).
- glycine and serine residues e.g. about 30%, or about 40%, or about 50%, or about 60%, or about 70%
- the linker may be functional.
- the linker may function to improve the folding and/or stability, improve the expression, improve the pharmacokinetics, and/or improve the bioactivity of the present compositions.
- the linker may function to target the compositions to a particular cell type or location.
- constant domain refers to the Fc domain.
- the constant domains exemplified above are optional embodiments and other constant domains can be substituted for the constant domains described herein.
- the Fc is from IgG.
- the IgG is human IgG.
- the human IgG is selected from IgGl, IgG2, IgG3, and IgG4.
- the antibody is formed through a knob-in-hole interaction in the Fc region.
- the human IgG Fc comprises one or mutations to promote knob-in-hole interaction.
- the mutations are selected from (i) T366Y or T366W, and (ii) Y407T, Y407A, or Y407V.
- the mutations are:
- the human IgG Fc comprises one or mutations to reduce or eliminate the effector function of the Fc domains.
- the mutations are L234A and L235A (LALA) substitutions in human IgGl.
- the human IgG Fc comprises one or mutations to stabilize a hinge region in the Fc domain.
- the mutation is S228P.
- the composition binds Claudin 18.2 and CD3 contemporaneously. [00122] In some embodiments, the composition binds Claudin 18.2 with an affinity of less than 10 nM and with at least 100 fold greater affinity than Claudin 9, Claudin 3, and/or Claudin 4.
- the composition induces cellular cytotoxicity.
- the composition induces T cell cytotoxicity.
- the composition induces T cell dependent cytotoxicity.
- the composition increases the expression and/or the release of one or more cytokines. In any of the embodiments disclosed herein, the composition increases the expression and/or the release of one or more cytokines selected from IL-2, IL-6, IL-10, IFN- ⁇ , and TNF- ⁇ .
- a pharmaceutical composition comprising an isolated antibody of any one of the preceding embodiments, or a nucleic acid molecule encoding the same.
- the composition is an injectable pharmaceutical composition.
- the composition is sterile.
- the composition is pyrogen free.
- compositions that composition as described herein, in combination with a pharmaceutically acceptable carrier.
- a “pharmaceutically acceptable carrier” is a pharmaceutically acceptable solvent, suspending agent, stabilizing agent, or any other pharmacologically inert vehicle for delivering one or more therapeutic compounds to a subject (e.g., a mammal, such as a human, non-human primate, dog, cat, sheep, pig, horse, cow, mouse, rat, or rabbit), which is nontoxic to the cell or subject being exposed thereto at the dosages and concentrations employed.
- Pharmaceutically acceptable carriers can be liquid or solid, and can be selected with the planned manner of administration in mind so as to provide for the desired bulk, consistency, and other pertinent transport and chemical properties, when combined with one or more of therapeutic compounds and any other components of a given pharmaceutical composition.
- Typical pharmaceutically acceptable carriers that do not deleteriously react with amino acids include, by way of example and not limitation: water, saline solution, binding agents (e.g., polyvinylpyrrolidone or hydroxypropyl methylcellulose), fillers (e.g., lactose and other sugars, gelatin, or calcium sulfate), lubricants (e.g., starch, polyethylene glycol, or sodium acetate), disintegrates (e.g., starch or sodium starch glycolate), and wetting agents (e.g., sodium lauryl sulfate).
- binding agents e.g., polyvinylpyrrolidone or hydroxypropyl methylcellulose
- fillers e.g., lacto
- Pharmaceutically acceptable carriers also include aqueous pH buffered solutions or liposomes (small vesicles composed of various types of lipids, phospholipids and/or surfactants which are useful for delivery of a drug to a mammal).
- Further examples of pharmaceutically acceptable carriers include buffers such as phosphate, citrate, and other organic acids, antioxidants such as ascorbic acid, low molecular weight (less than about 10 residues) polypeptides, proteins such as serum albumin, gelatin, or immunoglobulins, hydrophilic polymers such as polyvinylpyrrolidone, amino acids such as glycine, glutamine, asparagine, arginine or lysine, monosaccharides, disaccharides, and other carbohydrates including glucose, mannose or dextrins, chelating agents such as EDTA, sugar alcohols such as mannitol or sorbitol, saltforming counterions such as sodium, and/or nonionic surfactants such as TWEENTM, poly
- compositions can be formulated by mixing one or more active agents with one or more physiologically acceptable carriers, diluents, and/or adjuvants, and optionally other agents that are usually incorporated into formulations to provide improved transfer, delivery, tolerance, and the like.
- a pharmaceutical composition can be formulated, e.g., in lyophilized formulations, aqueous solutions, dispersions, or solid preparations, such as tablets, dragees or capsules.
- a multitude of appropriate formulations can be found in the formulary known to all pharmaceutical chemists: Remington’s Pharmaceutical Sciences (18th ed, Mack Publishing Company, Easton, PA (1990)), particularly Chapter 87 by Block, Lawrence, therein.
- formulations include, for example, powders, pastes, ointments, jellies, waxes, oils, lipids, lipid (cationic or anionic) containing vesicles (such as LIPOFECTINTM), DNA conjugates, anhydrous absorption pastes, oil-in-water and water-in- oil emulsions, emulsions carbowax (polyethylene glycols of various molecular weights), semi-solid gels, and semi-solid mixtures containing carbowax. Any of the foregoing mixtures may be appropriate in treatments and therapies as described herein, provided that the active agent in the formulation is not inactivated by the formulation and the formulation is physiologically compatible and tolerable with the route of administration.
- compositions include, without limitation, solutions, emulsions, aqueous suspensions, and liposome-containing formulations. These compositions can be generated from a variety of components that include, for example, preformed liquids, selfemulsifying solids and self-emulsifying semisolids.
- Emulsions are often biphasic systems comprising of two immiscible liquid phases intimately mixed and dispersed with each other; in general, emulsions are either of the water-in-oil (w/o) or oil-in-water (o/w) variety.
- Emulsion formulations have been widely used for oral delivery of therapeutics due to their ease of formulation and efficacy of solubilization, absorption, and bioavailability.
- compositions and formulations can contain sterile aqueous solutions, which also can contain buffers, diluents and other suitable additives (e.g., penetration enhancers, carrier compounds and other pharmaceutically acceptable carriers). Compositions additionally can contain other adjunct components conventionally found in pharmaceutical compositions. Thus, the compositions also can include compatible, pharmaceutically active materials such as, for example, antipruritics, astringents, local anesthetics or anti-inflammatory agents, or additional materials useful in physically formulating various dosage forms of the compositions provided herein, such as dyes, flavoring agents, preservatives, antioxidants, opacifiers, thickening agents and stabilizers.
- suitable additives e.g., penetration enhancers, carrier compounds and other pharmaceutically acceptable carriers.
- Compositions additionally can contain other adjunct components conventionally found in pharmaceutical compositions.
- the compositions also can include compatible, pharmaceutically active materials such as, for example, antipruritics, astringents, local anesthetics or anti-inflammatory agents,
- composition can be mixed with auxiliary agents, e.g., lubricants, preservatives, stabilizers, wetting agents, emulsifiers, salts for influencing osmotic pressure, buffers, colorings, flavorings, and aromatic substances.
- auxiliary agents e.g., lubricants, preservatives, stabilizers, wetting agents, emulsifiers, salts for influencing osmotic pressure, buffers, colorings, flavorings, and aromatic substances.
- compositions provided herein When added, however, such materials should not unduly interfere with the biological activities of the polypeptide components within the compositions provided herein.
- the formulations can be sterilized if desired.
- a composition containing a composition as provided herein can be in the form of a solution or powder with or without a diluent to make an injectable suspension.
- the composition may contain additional ingredients including, without limitation, pharmaceutically acceptable vehicles, such as saline, water, lactic acid, mannitol, or combinations thereof, for example.
- Administration can be, for example, parenteral (e.g., by subcutaneous, intrathecal, intraventricular, intramuscular, or intraperitoneal injection, or by intravenous drip). Administration can be rapid (e.g., by injection) or can occur over a period of time (e.g., by slow infusion or administration of slow release formulations). In some embodiments, administration can be topical (e.g., transdermal, sublingual, ophthalmic, or intranasal), pulmonary (e.g. , by inhalation or insufflation of powders or aerosols), or oral.
- parenteral e.g., by subcutaneous, intrathecal, intraventricular, intramuscular, or intraperitoneal injection, or by intravenous drip.
- Administration can be rapid (e.g., by injection) or can occur over a period of time (e.g., by slow infusion or administration of slow release formulations).
- administration can be topical (e.g., transdermal, sublingual, ophthal
- a composition containing a composition as described herein can be administered prior to, after, or in lieu of surgical resection of a tumor.
- the antibodies, compositions, or pharmaceutical compositions provided for herein may be administered at any appropriate interval to achieve the desired effect in a subject.
- the antibodies, compositions, or pharmaceutical compositions are administered daily, every other day, weekly, biweekly, once every three weeks, or monthly (i.e., once every four weeks).
- a method as provided for herein comprises administering an antibody, composition, or pharmaceutical composition to a cell or to a subject in need thereof at an interval of daily, every other day, weekly, biweekly, once every three weeks, or monthly (i.e., once every four weeks).
- nucleic acid molecule encoding an antibody or an amino acid sequence of any of the preceding embodiments.
- a cell comprising the nucleic acid molecule of any of the preceding embodiments, or the vector of any of the preceding embodiments.
- a method for modulating and/or targeting Claudin 18.2 and CD3 in a biological cell comprising contacting the cell with a composition of any of the preceding embodiments.
- a method for modulating Claudin 18.2 activity in a biological cell comprising contacting a cell expressing Claudin 18.2 with a composition of any of the preceding embodiments.
- a method for inhibiting the function of Claudin 18.2 in a biological cell comprising contacting a cell expressing Claudin 18.2 with a composition of any of the preceding embodiments.
- a method for treating or preventing cancer comprising administering an effective amount of the composition of any of the preceding embodiments to a subject in need thereof.
- composition of any of the preceding embodiments for the preparation of a medicament for the treatment of prevention of cancer.
- the cancer is selected from one or more of basal cell carcinoma, biliary tract cancer; bladder cancer; bone cancer; brain and central nervous system cancer; breast cancer; cancer of the peritoneum; cervical cancer; choriocarcinoma; colon and rectum cancer; connective tissue cancer; cancer of the digestive system; endometrial cancer; esophageal cancer; eye cancer; cancer of the head and neck; gastric cancer (including gastrointestinal cancer); glioblastoma; glioma; hepatic carcinoma; hepatoma; intra-epithelial neoplasm; kidney or renal cancer; larynx cancer; leukemia; liver cancer; lung cancer (e.g., small-cell lung cancer, non-small cell lung cancer, adenocarcinoma of the lung, and squamous carcinoma of the lung); melanoma; myeloma; neuroblastom
- the prevention of an onset, the presence, and/or the evaluation of the progression of a cancer in a subject can be assessed according to the Tumor/Nodes/Metastases (TNM) system of classification (International Union against Cancer, 6th edition, 2002), or the Whitmore- Jewett staging system (American Urological Association).
- TMM Tumor/Nodes/Metastases
- Whitmore- Jewett staging system American Urological Association
- cancers are staged using a combination of physical examination, blood tests, and medical imaging. If tumor tissue is obtained via biopsy or surgery, examination of the tissue under a microscope can also provide pathologic staging. In some embodiments, the stage or grade of a cancer assists a practitioner in determining the prognosis for the cancer and in selecting the appropriate modulating therapy.
- the prevention of an onset, or progression, of cancer is assessed using the overall stage grouping as a non-limiting example: Stage I cancers are localized to one part of the body, typically in a small area; Stage II cancers are locally advanced and have grown into nearby tissues or lymph nodes, as are Stage III cancers. Whether a cancer is designated as Stage II or Stage III can depend on the specific type of cancer. The specific criteria for Stages II and III can differ according to diagnosis. Stage IV cancers have often metastasized or spread to other organs or throughout the body.
- the onset or progression of cancer can be assessed using conventional methods available to one of skill in the art, such as a physical exam, blood tests, and imaging scans (e.g., X-rays, MRI, CT scans, ultrasound etc.).
- administering refers to a treatment/therapy from which a subject receives a beneficial effect, such as the reduction, decrease, attenuation, diminishment, stabilization, remission, suppression, inhibition or arrest of the development or progression of cancer, or a symptom thereof.
- the treatment/therapy that a subject receives, or the prevention in the onset of cancer results in at least one or more of the following effects: (1) the reduction or amelioration of the severity of cancer and/or a genetic disease or disorder, and/or a symptom associated therewith; (2) the reduction in the duration of a symptom associated with cancer and/or a genetic disease or disorder; (3) the prevention in the recurrence of a symptom associated with cancer and/or a genetic disease or disorder; (4) the regression of cancer and/or a genetic disease or disorder, and/or a symptom associated therewith; (5) the reduction in hospitalization of a subject; (6) the reduction in hospitalization length; (7) the increase in the survival of a subject; (8) the inhibition of the progression of cancer and/or a genetic disease or disorder and/or a symptom associated therewith; (9) the enhancement or improvement the therapeutic effect of another therapy; (10) a reduction or elimination in the cancer cell population, and/or a cell population associated with a genetic disease or disorder
- the treatment/therapy that a subject receives does not cure cancer, but prevents the progression or worsening of the disease. In certain embodiments, the treatment/therapy that a subject receives does not prevent the onset/development of cancer, but may prevent the onset of cancer symptoms.
- the subject that is treated does not develop cytokine release syndrome (CRS), or does not develop significant CRS -associated clinical symptoms or toxicity.
- CRS cytokine release syndrome
- symptoms include, but are not limited to fever, chills, fatigue, weakness, loss of appetite, nausea, vomiting, diarrhea, headache, joint or muscle aches, skin rash, low blood pressure, increased heart rate, irregular heartbeat, tachycardia, decreased heart function, swelling, buildup of fluids (edema), confusion, dizziness, seizures, hallucinations, decreased coordination, problems talking or swallowing, shaking, problems controlling movements, cough, shortness of breath, tachypnoea, decreased lung function, reduced oxygen levels, decreased kidney or liver function, increased cytokine levels in the blood, change in electrolytes, change in blood clotting.
- IL-6 interleukin-6
- IL-10 interleukin- 10
- IFN interferon
- MCP-1 monocyte chemoattractant protein 1
- GM-CSF monocyte chemoattractant protein 1
- TNF tumor necrosis factor
- IL-1 IL-2
- IL-2-receptor- ⁇ IL-8
- “preventing” an onset or progression of cancer in a subject in need thereof is inhibiting or blocking the cancer or disorder.
- the methods disclosed herein prevent, or inhibit, the cancer or disorder at any amount or level.
- the methods disclosed herein prevent or inhibit the cancer or genetic disease or disorder by at least or about a 10% inhibition (e.g., at least or about a 20% inhibition, at least or about a 30% inhibition, at least or about a 40% inhibition, at least or about a 50% inhibition, at least or about a 60% inhibition, at least or about a 70% inhibition, at least or about a 80% inhibition, at least or about a 90% inhibition, at least or about a 95% inhibition, at least or about a 98% inhibition, or at least or about a 100% inhibition).
- a 10% inhibition e.g., at least or about a 20% inhibition, at least or about a 30% inhibition, at least or about a 40% inhibition, at least or about a 50% inhibition, at least or about a 60% inhibition, at least or about a 70% inhibition, at least or about
- disclosed herein is an isolated antibody comprising one or more of the sequences disclosed herein.
- nucleic acid sequences encoding an antibody described herein or components thereof are nucleic acid sequences encoding an antibody described herein or components thereof.
- the nucleic acid encoding the antibody described herein or components thereof is a DNA, for example a linear DNA, a plasmid DNA, or a minicircle DNA.
- the nucleic acid encoding the antibody described herein or components thereof is an RNA, for example a mRNA.
- the nucleic acid encoding the antibody described herein or components thereof is delivered by a nucleic acid-based vector.
- the nucleic acid-based vector is a plasmid (e.g., circular DNA molecules that can autonomously replicate inside a cell), cosmid (e.g., pWE or sCos vectors), artificial chromosome, human artificial chromosome (HAC), yeast artificial chromosomes (YAC), bacterial artificial chromosome (BAC), Pl-derived artificial chromosomes (PAC), phagemid, phage derivative, bacmid, or virus.
- cosmid e.g., pWE or sCos vectors
- HAC human artificial chromosome
- YAC yeast artificial chromosomes
- BAC bacterial artificial chromosome
- PAC Pl-derived artificial chromosomes
- the nucleic acid-based vector is selected from the list consisting of: pSF-CMV-NEO-NH2-PPT-3XFLAG, pSF-CMV-NEO-COOH-3XFLAG, pSF-CMV-PURO-NH2-GST-TEV, pSF-OXB20-COOH-TEV-FLAG(R)-6His, pCEP4 pDEST27, pSF-CMV-Ub-KrYFP, pSF-CMV-FMDV-daGFP, pEFla-mCherry-Nl vector, pEFla-tdTomato vector, pSF-CMV-FMDV-Hygro, pSF-CMV-PGK-Puro, pMCP-tag(m), pSF-CMV-PURO-NH2-CMYC, pSF-OXB20-BetaGal,pSF-OXB20-Fluc, pSF-OXB20,
- the nucleic acid-based vector comprises a promoter.
- the promoter is selected from the group consisting of a mini promoter, an inducible promoter, a constitutive promoter, and derivatives thereof.
- the promoter is selected from the group consisting of CMV, CBA, EFla, CAG, PGK, TRE, U6, UAS, T7, Sp6, lac, araBad, trp, Ptac, p5, pl9, p40, Synapsin, CaMKII, GRK1, and derivatives thereof.
- the promoter is a U6 promoter.
- the promoter is a CAG promoter.
- the nucleic acid-based vector is a virus.
- the virus is an alphavirus, a parvovirus, an adenovirus, an AAV, a baculovirus, a Dengue virus, a lentivirus, a herpesvirus, a poxvirus, an anellovirus, a bocavirus, a vaccinia virus, or a retrovirus.
- the virus is an alphavirus.
- the virus is a parvovirus.
- the virus is an adenovirus.
- the virus is an AAV.
- the virus is a baculovirus.
- the virus is a Dengue virus. In some embodiments, the virus is a lentivirus. In some embodiments, the virus is a herpesvirus. In some embodiments, the virus is a poxvirus. In some embodiments, the virus is an anellovirus. In some embodiments, the virus is a bocavirus. In some embodiments, the virus is a vaccinia virus. In some embodiments, the virus is or a retrovirus.
- the AAV is AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV12, AAV13, AAV14, AAV15, AAV16, AAV-rh8, AAV-rhlO, AAV-rh20, AAV-rh39, AAV-rh74, AAV-rhM4-l, AAV-hu37, AAV-
- the herpesvirus is HSV type 1, HSV-2, VZV, EBV, CMV, HHV-6, HHV-7, or HHV-8.
- the nucleic acid encoding the antibody described herein or components thereof is delivered by a non-nucleic acid-based delivery system (e.g., a non- viral delivery system).
- a non-viral delivery system is a liposome.
- the nucleic acid is associated with a lipid.
- the nucleic acid associated with a lipid in some embodiments, is encapsulated in the aqueous interior of a liposome, interspersed within the lipid bilayer of a liposome, attached to a liposome via a linking molecule that is associated with both the liposome and the nucleic acid, entrapped in a liposome, complexed with a liposome, dispersed in a solution containing a lipid, mixed with a lipid, combined with a lipid, contained as a suspension in a lipid, contained or complexed with a micelle, or otherwise associated with a lipid.
- the nucleic acid is comprised in a lipid nanoparticle (LNP).
- the antibody described herein or components thereof is introduced into the cell in any suitable way, either stably or transiently.
- the antibody described herein or components thereof is transfected into the cell.
- the cell is transduced or transfected with a nucleic acid construct that encodes the antibody described herein or components thereof.
- a cell is transduced (e.g., with a virus encoding the antibody described herein or components thereof), or transfected (e.g., with a plasmid encoding the antibody described herein or components thereof) with a nucleic acid that encodes the antibody described herein or components thereof, or the translated the antibody described herein or components thereof.
- the transduction is a stable or transient transduction.
- a plasmid expressing the antibody described herein or components thereof is introduced into cells through electroporation, transient (e.g., lipofection) and stable genome integration (e.g., piggybac) and viral transduction (for example lentivirus or AAV) or other methods known to those of skill in the art.
- the gene editing system is introduced into the cell as one or more polypeptides.
- delivery is achieved through the use of RNP complexes. Delivery methods to cells for polypeptides and/or RNPs are known in the art, for example by electroporation or by cell squeezing.
- Exemplary methods of delivery of nucleic acids include lipofection, nucleofection, electroporation, stable genome integration (e.g., piggybac), microinjection, biolistics, virosomes, liposomes, immunoliposomes, polycation or lipid nucleic acid conjugates, naked DNA, artificial virions, and agent-enhanced uptake of DNA.
- lipofection is described in e.g., U.S. Pat. Nos.
- lipofection reagents are sold commercially (e.g., TransfectamTM, LipofectinTM and SF Cell Line 4D-Nucleofector X KitTM (Lonza)).
- Cationic and neutral lipids that are suitable for efficient receptor-recognition lipofection of polynucleotides include those of WO 91/17424 and WO 91/16024.
- the delivery is to cells (e.g., in vitro or ex vivo administration) or target tissues (e.g., in vivo administration).
- the nucleic acid is comprised in a liposome or a nanoparticle that specifically targets a host cell.
- a nucleic acid molecule encoding an antibody or a component thereof described herein includes one or more of the DNA sequences provided in TABLE B.
- the nucleic acid molecule comprises a sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity to any one of SEQ ID NOs: 5210-5236.
- a nucleic acid molecule encoding an antibody or a component thereof described herein includes one or more of the RNA sequences provided in TABLE C.
- the nucleic acid molecule comprises a sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity to any one of SEQ ID NOs: 5237-5263.
- TABLE B DNA Sequences Encoding Antibody or a Component Thereof Described
- lipid nanoparticles comprising an antibody or components thereof of the disclosure for delivery of the antibody into a cell.
- the lipid nanoparticle comprises the antibody or a nucleic acid encoding the antibody or components thereof.
- the lipid nanoparticle is tethered to the antibody or a nucleic acid encoding the antibody or components thereof.
- Lipid nanoparticles as described herein can be 4-component lipid nanoparticles. Such nanoparticles can be configured for delivery of RNA or other nucleic acids (e.g., synthetic RNA, mRNA, or in vitro-synthesized mRNA) and can be generally formulated as described in WO2012135805A2.
- Such nanoparticles can generally comprise: (a) a cationic lipid, (b) a neutral lipid (e.g., DSPC or DOPE), (c) a sterol (e.g., cholesterol or a cholesterol analog), or (d) a PEG-modified lipid (e.g., PEG-DMG).
- a cationic lipid e.g., DSPC or DOPE
- a neutral lipid e.g., DSPC or DOPE
- a sterol e.g., cholesterol or a cholesterol analog
- PEG-DMG PEG-modified lipid
- Cationic lipid formulations can include particles comprising either 3 or 4 or more components in addition to polynucleotide, primary construct, or RNA (e.g., mRNA).
- RNA e.g., mRNA
- formulations with certain cationic lipids include, but are not limited to, 98N12-5 and may contain 42% lipidoid, 48% cholesterol and 10% PEG (C14 or greater alkyl chain length).
- formulations with certain lipidoids include, but are not limited to, C12-200 and may contain 50% cationic lipid, 10% disteroylphosphatidyl choline, 38.5% cholesterol, and 1.5% PEG-DMG.
- the cationic lipid nanoparticle comprises a cationic lipid, a PEG-modified lipid, a sterol, and a non-cationic lipid.
- the cationic lipid nanoparticle has a molar ratio of about 20-60% cationic lipid: about 5-25% non-cationic lipid: about 25-55% sterol; and about 0.5-15% PEG-modified lipid.
- the cationic lipid nanoparticle comprises a molar ratio of about 50% cationic lipid, about 1.5% PEG-modified lipid, about 38.5% cholesterol, and about 10% non-cationic lipid.
- the cationic lipid nanoparticle comprises a molar ratio of about 55% cationic lipid, about 2.5% PEG-modified lipid, about 32.5% cholesterol, and about 10% noncationic lipid.
- the cationic lipid is an ionizable cationic lipid
- the noncationic lipid is a neutral lipid
- the sterol is a cholesterol.
- the cationic lipid nanoparticle has a molar ratio of 50:38.5: 10: 1 .5 of cationic lipid: cholesterol: PEG2000-DMG:DSPC or DMG:DOPE.
- lipid nanoparticles as described herein can comprise cholesterol, 1,2-dioleoyl-sn-glycero-3 -phosphoethanolamine (DOPE), 1 , 1 ‘-((2-(4-(2-((2-(bis(2-hydroxydodecyl)amino)ethyl)(2- hydroxydodecyl)amino)ethyl)piperazin- 1 -yl)ethyl)azanediyl)bis(dodecan-2-ol) (C 12-200), and DMG-PEG-2000 at molar ratios of 47.5:16:35:1.5.
- DOPE 1,2-dioleoyl-sn-glycero-3 -phosphoethanolamine
- the lipid nanoparticle comprises 1,1'-[[2-[4-[2-[[2-[bis(2- hydroxydodecyl)amino]ethyl](2-hydroxydodecyl)amino]ethyl]-l- piperazinyl]ethyl]imino]bis-2-dodecanol (C12-200), a sterol (e.g., cholesterol, campesterol, or ⁇ -sitosterol), 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE), and 1,2- dimyristoyl-sn-glycero-3-phosphoethanolamine-N-[methoxy(polyethylene glycol)-2000] (ammonium salt) (C14:0 PEG2000 PE).
- a sterol e.g., cholesterol, campesterol, or ⁇ -sitosterol
- DOPE 1,2-dioleoyl-sn-glycero-3-
- the lipid nanoparticle comprises 1,1'-[[2-[4-[2-[[2-[bis(2-hydroxydodecyl)amino]ethyl](2- hydroxydodecyl)amino]ethyl]-l-piperazinyl]ethyl]imino]bis-2-dodecanol (C12-200), sterol (e.g., cholesterol, campesterol, or ⁇ -sitosterol), 1,2-dioleoyl-sn-glycero-3- phosphoethanolamine (DOPE), 1 ,2-dimyristoyl-sn-glycero-3-phosphoethanolamine-N- [methoxy(polyethylene glycol)-2000] (ammonium salt) (C14:0 PEG2000 PE), and 1 ,2- distearoyl-sn-glycero-3-phosphoethanolamine-N-[tris-GalNAc-Y-aminobutyric acid- (
- the lipid nanoparticle comprises (9Z,12Z)-3-((4,4- Bis(octyloxy)butanoyl)oxy)-2-((((3-(diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9, 12-dienoate (LP01), a sterol (e.g., cholesterol, campesterol, or ⁇ -sitosterol), 1,2- dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE), 1 ,2-dioctadecanoyl-sn-glycero-3- phosphocholine (DS PC), and 1,2-dimyristoyl-rac-glycero-3-methoxypolyethylene glycol- 2000 (DMG-PEG 2000).
- a sterol e.g., cholesterol, campesterol, or ⁇ -sitosterol
- DOPE 1,2- dioleoyl-sn-glycero
- the lipid nanoparticle comprises (9Z,12Z)-3- ((4,4-Bis(octyloxy)butanoyl)oxy)-2-((((3- (diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9, 12-dienoate (LP01), a sterol (e.g., cholesterol, campesterol, or ⁇ -sitosterol), 1,2-dioleoyl-sn-glycero-3- phosphoethanolamine (DOPE), 1,2-dioctadecanoyl-sn-glycero-3 -phosphocholine (DSPC), 1,2-dimyristoyl-rac-glycero-3-methoxypolyethylene glycol-2000 (DMG-PEG 2000), and 1,2- distearoyl-sn-glycero-3-phosphoethanolamine-N-[tris-GalNAc-Y-aminobutyric acid- (L)),
- the lipid nanoparticle comprises (9Z,12Z)-3-((4,4-Bis(octyloxy)butanoyl)oxy)-2-((((3- (diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9, 12-dienoate (LP01 ) , cholesterol, 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE), 1,2-dioctadecanoyl-sn- glycero-3-phosphocholine (DSPC), and 1,2-dimyristoyl-rac-glycero-3-methoxypolyethylene glycol-2000 (DMG-PEG 2000).
- DOPE 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine
- DSPC 1,2-dioctadecanoyl-sn- glycero-3-phosphocholine
- DMG-PEG 2000 1,2-dim
- the lipid nanoparticle comprises (9Z, 12Z)-3-((4,4-Bis(octyloxy)butanoyl)oxy)-2-((((3- (diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9, 12-dienoate (LP01 ) , cholesterol, 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE), 1,2-dioctadecanoyl-sn- glycero-3-phosphocholine (DSPC), 1 ,2-dimyristoyl-rac-glycero-3-methoxypolyethylene glycol-2000 (DMG-PEG 2000), and 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N- [tris-GalNAc-Y-aminobutyric acid-(polyethylene glycol)-2000.
- DOPE 1,2-dioleoyl-s
- the lipid nanoparticles comprises (9Z,12Z)-3-((4,4- Bis(octyloxy)butanoyl)oxy)-2-((((3-(diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9,12-dienoate (LP-01), cholesterol, 1,2-dioctadecanoyl-sn-glycero-3- phosphocholine (DSPC) and 1 ,2-dimyristoyl-rac-glycero-3 -methoxypolyethylene glycol- 2000 (DMG-PEG 2000).
- the lipid nanoparticles comprises one of the phospholipids selected from 1,2-dioleoyl-sn-glycero-3 -phosphoethanolamine (DOPE), 1,2-dilinolenoyl-sn- glycero-3-phosphoethanolamine (18:3 PE), 1,2-didocosahexaenoyl-sn-glycero-3- phosphoethanolamine (22:6 PE), and 1,2-dioleoyl-sn-glycero-3 -phosphocholine (DOPC).
- DOPE 1,2-dioleoyl-sn-glycero-3 -phosphoethanolamine
- DOPC 1,2-dilinolenoyl-sn- glycero-3-phosphoethanolamine
- the lipid nanoparticle comprises of different lipids comprising (9Z,12Z)-3-((4,4-Bis(octyloxy)butanoyl)oxy)-2-((((3- (diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9, 12-dienoate (LP-01 ), cholesterol, 1,2-dioctadecanoyl-sn-glycero-3-phosphocholine (DSPC), 1 ,2-dimyristoyl-rac- glycero-3-methoxypolyethylene glycol-2000 (DMG-PEG 2000), 1 ,2-dioleoyl-sn-glycero-3- phosphoethanolamine (DOPE) or 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC), and 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-
- the word “include,” and its variants, is intended to be non-limiting, such that recitation of items in a list is not to the exclusion of other like items that may also be useful in the materials, compositions, devices, and methods of this technology.
- the terms “can” and “may” and their variants are intended to be non-limiting, such that recitation that an embodiment can or may comprise certain elements or features does not exclude other embodiments of the present technology that do not contain those elements or features.
- Example 1 Overview and Development of Antibodies Specific for Claudin 18.2 and CD3
- the experiments in this example demonstrated certain features of antibodies specific for Claudin 18.2 and CD3.
- antibodies specific for Claudin 18.2 and CD3 were engineered and characterized.
- This example provided tandem scFv antibodies, scFv-Fab IgG antibodies, IgG-(scFv) 2 antibodies, and IgG-(scFv) antibodies specific for Claudin 18.2 and CD3 (FIG. 1).
- Claudin 18.2 and CD3 To develop bispecific antibodies to Claudin 18.2 and CD3, a panel of constructs were generated using the following antibody formats: tandem scFv, scFv-Fab IgG, IgG-scFv, IgG-(scFv) 2 , and IgG-(scFv).
- Potential Claudin 18.2-specific arms included those from 4409- B03, 4401 -E10, and 4405-H06 antibodies, and specific CD3 arms included those from hsOKT3 and pasotuxizumab antibodies, as well as a nanobody.
- Antibodies tested are provided in TABLEs 6-9, as follows:
- FIG. 2 shows that IM-16-44-09-B03-1CD3 bound to both human Claudin 18.2
- CDN 18.2 (“CLDN 18.2”) and human CD3.
- Flow cytometry experiments were performed to test the ability of the IM-16-44-09-B03-1CD3 antibody to bind HEK293F cells expressing human CLDN 18.2, human CD3, or human P2X3.
- HEK293F cells were transfected by lipofectamine with target and control plasmids. At 18-24 hours post-transfection, cells were stained with bispecific and control MAbs followed by APC -labelled goat anti-human antibody.
- IM-16-44-09-B03-1CD3 selectively bound to both human CLDN 18.2 and human CD3, and not P2X3.
- FIG. 3 shows cellular toxicity data from T-cell dependent cellular cytotoxicity experiments.
- Human T cells 25,000 cells/well
- CLDN 18.1 human Claudin 18.1
- CLDN 18.2-expressing HEK293F cells (2500 cells/well) also expressing Firefly Luciferase (ffLuciferase) in the presence of IMC-16-C2 (control antibody) or IM- 16- 44-09-B03-1CD3 antibodies.
- IMC-16-C2 control antibody
- IM- 16- 44-09-B03-1CD3 antibodies expressing Firefly Luciferase activity
- FIG. 3 killing was only potently induced against CLDN 18.2- expressing gastric cancer cells by the IM-16-44-09-B03-1CD3 antibody. No killing activity was observed using the control antibody or against cells
- FIGs. 4A, 4B, 4C, 4D, and 4E show cytokines produced by human peripheral blood mononuclear cells (PBMC) cultured with IMC-16-C2 or IM-16-44-09-B03-1CD3 antibodies and human CLDN 18.1 - or 18.2-expressing cells (HEK293F).
- FIG. 4A shows the levels of IL-2
- FIG. 4B shows the levels of IL-6
- FIG. 4C shows the levels of IL- 10
- FIG. 4D shows the levels of IFN- ⁇ (gamma)
- 4E shows the levels of TNF- ⁇ (alpha) produced by human PBMCs in the presence of human CLDN 18.1- or 18.2-expressing cells and IMC-16- C2 or IM-16-44-09-B03-1CD3 antibodies.
- FIGs. 5A and 5B show size exclusion chromatography (SEC) and differential scanning fhiorimetry (DSF) experimental results for the IM-16-44-09-B03-1CD3, IM-16-44- 09-B03-2CD3, and IM-16-44-09-B03-3CD3 antibodies.
- SEC was performed using an AKTA FPLC with a Superdex 200 increase 3.2/300 SEC column. Samples (0.1 ml) were run in PBS at a concentration of 0.5 to 1 mg/ml.
- DSF was performed by mixing 20 ⁇ L antibody (0.1 mg/ml) with 10 ⁇ L of Sypro Orange (BioRad).
- FIG. 6 shows that 1M-21-44-01-E10-3CD3 bound to both human CLDN 18.2 and human CD3.
- Flow cytometry experiments were performed to test the ability of the IM-21-44- 01-E10-3CD3 antibody to bind HEK293F cells expressing human CLDN 18.2, human CD3, or human P2X3.
- HEK293F cells were transfected by lipofectamine with target and control plasmids. At 18-24 hours post-transfection, cells were stained with bispecific and control MAbs followed by APC-labelled goat anti-human antibody.
- FIG. 7 shows cellular toxicity data from T-cell dependent cellular cytotoxicity experiments.
- Human T cells 25,000 cells/well
- human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IMC-16-C2 (control antibody) or IM-21 -44-01-E10-3CD3 antibodies.
- IMC-16-C2 control antibody
- IM-21 -44-01-E10-3CD3 antibodies IM-21 -44-01-E10-3CD3 antibodies
- luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
- killing was only potently induced against CLDN 18.2-expressing cells by the IM-21-44-01 -E10-3CD3 antibody. No killing activity was observed using the control antibody or against cells expressing CLDN 18.1.
- FIGs. 8A, 8B, 8C, 8D, and 8E show cytokines produced by human PBMC cultured with IMC-16-C2 or IM-16-44-09-B03-1CD3 antibodies and human CLDN 18.1- or 18.2- expressing cells.
- FIG. 8A shows the levels of IL-2
- FIG. 8B shows the levels of IL-6
- FIG. 8A shows the levels of IL-6
- FIG. 8C shows the levels of IL-10
- FIG. 8D shows the levels of IFN- ⁇ (gamma)
- FIG. 8E shows the levels of TNF- ⁇ (alpha) produced by human PBMCs in the presence of human CLDN 18.1- or 18.2-expressing cells and the IMC-16-C2 or IM-21-44-01-E10-3CD3 antibodies.
- FIGs. 9 A and 9B show SEC and DSF experimental results for IM-21-44-01 -E10- 1CD3, IM-21 -44-01-E10-2CD3, and IM-21-44-01-E10-3CD3 antibodies.
- SEC was performed using an AKTA FPLC with a Superdex 200 increase 3.2/300 SEC column.
- FIG. 10 shows that IM-21-44-09-B03-3CD3 bound to both human CLDN 18.2 and human CD3. Flow cytometry experiments were performed to test the ability of the IM-21-44- 01-E10-3CD3 antibody to bind HEK-293F cells expressing human CLDN 18.2, human CD3, or human P2X3.
- HEK-293F cells were transfected by lipofectamine with target and control plasmids. At 18-24 hours post transfection, cells were stained with bispecific and control MAbs followed by APC-labelled goat anti-human antibody. Fluorescence was detected using an Intellicyt high-throughput flow cytometer (iQue; Intellicyt, Albuquerque, NM). As shown in FIG. 10, IM-21-44-09-B03-3CD3 selectively bound to both human CLDN 18.2 and human CD3, and not P2X3.
- FIG. 11 shows cellular toxicity data from T-cell dependent cellular cytotoxicity experiments.
- Human T cells 25,000 cells/well
- human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IMC-16-C2 (control antibody) or IM-21-44-09-B03-3CD3 antibodies.
- IMC-16-C2 control antibody
- IM-21-44-09-B03-3CD3 antibodies IM-21-44-09-B03-3CD3 antibodies
- luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
- killing was only potently induced against CLDN 18.2-expressing cells by the IM-21-44-09-B03-3CD3 antibody. No killing activity was observed using the control antibody or against cells expressing CLDN 18.1.
- FIGs. 12A, 12B, 12C, 12D, and 12E show cytokines produced by human PBMC cultured with the IMC-16-C2 or IM-21-44-09-B03-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells.
- FIG. 12A shows the levels of IL-2
- FIG. 12B shows the levels of IL-6
- FIG. 12C shows the levels of IL-10
- FIG. 12D shows the levels of IFN- ⁇ (gamma)
- FIG. 12E shows the levels of TNF- ⁇ (alpha) produced by human PBMCs in the presence of human CLDN 18.1- or 18.2-expressing cells and IMC-16-C2 or IM-21-44-09-B03-3CD3 antibodies.
- FIGs. 13A and 13B show SEC and DSF experimental results for IM-21 -44-09-B03- 1CD3, IM-21-44-09-B03-2CD3, and IM-21 -44-09-B03-3CD3 antibodies.
- SEC was performed using an AKTA FPLC with a Superdex 200 increase 3.2/300 SEC column.
- FIG. 14 shows that IM-2-44-05-H06-1CD3 bound to both human CLDN 18.2 and human CD3.
- Flow cytometry experiments were performed to test the ability of the IM-2-44- 05-H06-1CD3 antibody to bind HEK293F cells expressing human CLDN 18.2, human CD3, or human P2X3.
- HEK293F cells were transfected by lipofectamine with target and control plasmids. At 18-24 hours post transfection, cells were stained with bispecific and control MAbs followed by APC-labelled goat anti-human antibody. Fluorescence was detected using an Intellicyt high-throughput flow cytometer (iQue; Intellicyt, Albuquerque, NM). As shown in FIG.
- FIG. 15 shows cellular toxicity data from T-cell dependent cellular cytotoxicity experiments.
- Human T cells 25,000 cells/well
- human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IMC-16-C2 (control antibody) or IM-2-44-05-H06-1CD3 antibodies.
- IMC-16-C2 control antibody
- IM-2-44-05-H06-1CD3 antibodies After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
- killing was only potently induced against CLDN 18.2-expressing cells by the IM-2-44-05 -H06-1CD3 antibody. No killing activity was observed using the control antibody or against cells expressing CLDN 18.1.
- FIGs. 16A, 16B, 16C, 16D, and 16E show cytokines produced by human PBMC cultured with the IMC-16-C2 or IM-2-44-05 -H06-1CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells.
- FIG. 16A shows the levels of IL-2
- FIG. 16B shows the levels of IL-6
- FIG. 16C shows the levels of IL- 10
- FIG. 16D shows the levels of IFN- ⁇ (gamma)
- FIG. 16E shows the levels of TNF- ⁇ (alpha) produced by human PBMCs in the presence of human CLDN 18.1- or 18.2-expressing cells and IMC-16-C2 or IM-2-44-05-H06-1CD3 antibodies.
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Immunology (AREA)
- Organic Chemistry (AREA)
- Medicinal Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- General Health & Medical Sciences (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Genetics & Genomics (AREA)
- Molecular Biology (AREA)
- Biophysics (AREA)
- Biochemistry (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Pharmacology & Pharmacy (AREA)
- Animal Behavior & Ethology (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
- Peptides Or Proteins (AREA)
Abstract
The disclosure is directed to an antibody specific for Claudin 18.2 and CD3. The present disclosure describes the isolation and characterization of antibodies, antibody fragments, and antibody variants specific for Claudin 18.2 and CD3.
Description
ANTIBODIES DIRECTED TO CLAUDIN 18.2, INCLUDING BISPECIFIC FORMATS THEREOF
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] The invention claims the benefit of and priority to U.S. Provisional Patent Application No. 63/606,967, filed December 6, 2023, the disclosure of which is hereby incorporated by reference in its entirety for all purposes.
FIELD OF THE DISCLOSURE
[0002] The disclosure is directed to compositions that bind Claudin 18.2 and related methods.
BACKGROUND
[0003] Cell adhesion proteins are critical for maintaining tissue integrity, as well as regulating diverse cellular events in a wide variety of physiological and pathological processes. Among cell adhesion proteins, some members of the claudin (CLDN) family are often aberrantly expressed in various cancers. Clinical application of CLDN therapeutics has been difficult because of lack of antibody specificity for particular CLDN proteins and widespread expression of closely related CLDN family members in normal cells. Thus, there remains a significant need for improved compositions and methods that can modulate the activity of CLDN family members to treat various cancers and diseases.
SUMMARY
[0004] Accordingly, in various aspects, the present disclosure relates to a composition that specifically binds to Claudin 18.2 and CD3. The present disclosure describes the isolation and characterization of antibodies, antibody fragments, and antibody variants specific for Claudin 18.2 and CD3. In some embodiments, the antibody is a tandem single-chain variable fragment (scFv) specific for Claudin 18.2 and CD3 (FIG. 1A). In some embodiments, the antibody is a hetero VH IgG (FIG. 1B) specific for Claudin 18.2 and CD3. In some embodiments, the antibody is in the format of an scFv-Fab IgG antibody, which is illustrated as, for example, in FIG. 1C and is specific for Claudin 18.2 and CD3. In some embodiments, the antibody in the format of an IgG-scFv, which is illustrated as, for example, in FIG. 1D and is specific for Claudin 18.2 and CD3. In some embodiments, the antibody is in an IgG- (scFv)2 format , which is illustrated, for example, in FIG. 1E and is specific for Claudin 18.2 and CD3. In some embodiments, the antibody is in an IgG-(scFv) format, which is, for example, illustrated in FIG. 1F and is specific for Claudin 18.2 and CD3. In some embodiments, the antibody specific for Claudin 18.2 and CD3 binds Claudin 18.2 and CD3 contemporaneously. Without being bound to any particular theory, the bispecific antibody
will bind to a tumor cell expressing Claudin 18.2 and an immune cell, such as a cytotoxic T cell, expressing CD3. By bringing the cells together, the T cell is activated and targets the tumor cell expressing Claudin 18.2 for destruction.
[0005] In one aspect, compositions are provided comprising an antibody in an scFv-Fab IgG antibody that binds to Claudin 18.2 and CD3. In some embodiments, the antibody comprises a first heavy chain, a second heavy chain, and a light chain, wherein
(a) the first heavy chain is selected from:
(i)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ
APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC
AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ VSLWCLVKGFYPSD1AVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, or
(it)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ
VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ
QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4767), or
an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, or (iii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR
QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLWCLVKG FYPSD1AVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4768), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(b) the second heavy chain is selected from:
(i)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAMNWVR
QAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQMNNLKTEDT
AVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGGSGGGGSQTVVTQEP
SLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFS
GSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLASPKSSDKTHTC
PPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVE
VHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKA
KGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPP
VLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4168), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), or
(it)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFTRYTMHWVR QAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKSTAFLQMDSLRPEDTAVYY CARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGGGGSDIQMTQSPSSLSASVG DRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTI SSLQPEDIATYYCQQWSSNPFTFGQGTKLQITRASPKSSDKTHTCPPCPAPEAAGGPS VFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQ YNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTL PPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVS KLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4169), o an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant
thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, or
(iii)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSMGWFR
QAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTSLKPEDSATY
YCAAKIRPYIFKIAGQYDYWGQGTQVTVSSASPKSSDKTHTCPPCPAPEAAGGPSVFL
FPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNS
TYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSR
EEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTV
DKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3983), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAK1RPY1FK1AGQYDY (SEQ ID NO: 18) or a variant thereof, and
(b) the light chain is selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWYQQKP
GQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIF
GGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP
VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTEC
S (SEQ ID NO: 3576), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQKPGQ APVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIFGGG TKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKA GVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3593), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQQKPGQ APVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGGYDSSAGIFGG GTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVK AGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, optionally wherein the first or second heavy chain comprises a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N- or C -terminus; and/or optionally wherein the composition comprises a linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof.
[0006] In some embodiments, an antibody is provided in an IgG-scFv antibody format that binds to Claudin 18.2 and CD3, wherein the antibody comprises:
(a) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ
APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC
AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS
GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL
GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL
TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ
VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ
QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and (ii) the second heavy chain comprises:
(A)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSMGWFR QAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTSLKPEDSATY
YCAAKIRPYIFKIAGQYDYWGQGTQVTVSSGGGGSGGGGSGGGGSEVQLLESGGGL
VQPGGSLRLSCAASGFTFSSYGMGWVRQAPGKGLEWVAAITRDSSTNYGAAVKGR
ATISRDNSKNTLYLQMNSLRAEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDA
WGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGAL
TSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCD
KTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY
VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIE
KTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENN
YKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPG
K (SEQ ID NO: 4778), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
98%, or about 99% identity thereto, or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof,
(B)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAMNWVR
QAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQMNNLKTEDT
AVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGGSGGGGSQTVVTQEP
SLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFS
GSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLGGGGSGGGGSG
GGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQAPGKGLEWVAAIT
RDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKDASNGYCWYT
GRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDY
FPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPS
NTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVV
DVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKE
YKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPS
DIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEA
LHNHYTQKSLSLSPGK (SEQ ID NO: 4803), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a
variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
(C)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFTRYTMHWVR
QAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKSTAFLQMDSLRPEDTAVYY
CARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGGGGSDIQMTQSPSSLSASVG
DRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTI
SSLQPEDIATYYCQQWSSNPFTFGQGTKLQITRGGGGSGGGGSGGGGSEVQLLESGG
GLVQPGGSLRLSCAASGFTFSSYGMGWVRQAPGKGLEWVAAITRDSSTNYGAAVK
GRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHI
DAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSG
ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKS
CDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFN
WYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPA
PIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPE
NNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS
PGK (SEQ ID NO: 4806); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11)
or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, or
(b) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV LTVLHQDWLNGKEYKCKVSNKALPAPIEKT1SKAKGQPREPQVYTLPPSREEMTKNQ VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4767), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and (ii) the second heavy chain comprises:
(A)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSMGWFR QAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTSLKPEDSATY YCAAKIRPYIFKIAGQYDYWGQGTQVTVSSGGGGSGGGGSGGGGSEVQLLESGGGL VQPGGSLRLSCAASGFTFSSYGMSWVRQAPGKGLEWVAGIGSSGSDTWYADSVKG RFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHHID AWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGA LTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSC
DKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNW
YVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPI EKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPEN NYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK (SEQ ID NO: 4779), or an amino acid sequence having al least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof,
(B)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAMNWVR
QAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQMNNLKTEDT
AVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGGSGGGGSQTVVTQEP
SLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFS
GSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLGGGGSGGGGSG
GGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQAPGKGLEWVAGIG
SSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKDATNGYCWY
TGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKD
YFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKP
SNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVV
DVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKE
YKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPS
DIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEA
LHNHYTQKSLSLSPGK (SEQ ID NO: 4804), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
98%, or about 99% identity thereto, or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and (C)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFTRYTMHWVR QAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKSTAFLQMDSLRPEDTAVYY CARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGGGGSDIQMTQSPSSLSASVG DRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTI SSLQPEDIATYYCQQWSSNPFTFGQGTKLQITRGGGGSGGGGSGGGGSEVQLLESGG GLVQPGGSLRLSCAASGFTFSSYGMSWVRQAPGKGLEWVAGIGSSGSDTWYADSV
KGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHH
IDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNS
GALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEP
KSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKF
NWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP
APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQP
ENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL
SPGK (SEQ ID NO: 4807); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a
variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, and/or heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, or (c) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTF1SLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLWCLVKG FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4768), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSMGWFR
QAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTSLKPEDSATY
YCAAKIRPYIFKIAGQYDYWGQGTQVTVSSGGGGSGGGGSGGGGSEVQLLESGGGL
VQPGGSLRLSCAASGFTFRSYAMMWVRQAPGKGLEWVGEISSTGGSTYYGSAVKG
RATISRDNSKNTLYLQMNSLRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVT
VSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPA
VLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCP
APEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN
AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQ
PREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLD
SDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID
NO: 4780), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof,
(B)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAMNWVR
QAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQMNNLKTEDT
AVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGGSGGGGSQTVVTQEP
SLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFS
GSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLGGGGSGGGGSG
GGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVRQAPGKGLEWVGEIS
STGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKSTYGATWKS
ADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVS
WNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKK
VEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPE
VKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNK ALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESN GQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKS LSLSPGK (SEQ ID NO: 4805), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(C)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFTRYTMHWVR
QAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKSTAFLQMDSLRPEDTAVYY
CARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGGGGSDIQMTQSPSSLSASVG
DRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTI
SSLQPEDIATYYCQQWSSNPFTFGQGTKLQITRGGGGSGGGGSGGGGSEVQLLESGG
GLVQPGGSLRLSCAASGFTFRSYAMMWVRQAPGKGLEWVGEISSTGGSTYYGSAV
KGRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTL
VTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTF
PAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPP
CPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVH
NAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKG
QPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVL
DSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4808); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(b) a light chain selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWYQQKP GQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIF GGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTEC S (SEQ ID NO: 3576), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof,
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQKPGQ APVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIFGGG TKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKA GVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3593), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or (iii) MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQQKPGQ APVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGGYDSSAGIFGG GTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVK AGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, optionally wherein the first or second heavy chain comprises a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N- or C -terminus; and/or optionally wherein the composition comprises a linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof.
[0007] In some embodiments, the antibody or composition comprising the same, is an antibody an IgG-(scFv)2 antibody format that binds to Claudin 18.2 and CD3, wherein the antibody comprises:
(a) a heavy chain selected from:
(i)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ
APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC
AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS
GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL
GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL
TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQV
SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQG
NVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLVQ PGGSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKD RFTISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTV SSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQ QKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSN RWVFGGGTKLTVL (SEQ ID NO: 4769), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO:
5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), (it) MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ
APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC
AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS
GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL
GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL
TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQV
SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQG
NVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSQVQLVQSGGGVVQ
PGRSLRLSCKASGYTFTRYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFT
ISTDKSKSTAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGGSG
GGGSGGGGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAPKRWIY DTSKLASGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTFGQGTKLQITR (SEQ ID NO: 4770), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof,
(iii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ
APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC
AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS
GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL
GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL
TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQV
SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQG
NVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPVQ
AGGSLRLSCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRF
TTSRDNAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS
(SEQ ID NO: 4771), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, (iv)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPffiKTISKAKGQPREPQVYTLPPSRDELTKNQ
VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQ
GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLV
QPGGSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVK
DRFTISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVT
VSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWV
QQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYS NRWVFGGGTKLTVL (SEQ ID NO: 4772), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), (v) MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQ
VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQ
GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSQVQLVQSGGGVV
QPGRSLRLSCKASGYTFTRYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRF
TISTDKSKSTAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGGS
GGGGSGGGGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAPKRWI
YDTSKLASGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTFGQGTKLQrr
R (SEQ ID NO: 4773), or
an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof.
(vi)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQ GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPV
QAGGSLRLSCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGR
FTISRDNAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 4774), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a
variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof,
(vii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR
QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL
NGKEYKCKVSNKALPAPIEKT1SKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKG
FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV
MHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLVQPGGSLKL
SCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRD
DSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGG
SGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQA
PRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFG
GGTKLTVL (SEQ ID NO: 4775), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO:
5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), (viii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR
QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKG FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSQVQLVQSGGGVVQPGRSLRL SCKASGYTFTRYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSK STAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGG GGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLA SGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTFGQGTKLQITR (SEQ ID NO: 4776), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, and
(ix.)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR
QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKG FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPVQAGGSLRL SCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFT1SRDNAK NTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 4777), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and
(b) a light chain selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWYQQKP
GQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIF GGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTEC S (SEQ ID NO: 3576), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof,
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQKPGQ APVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIFGGG TKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCL1SDFYPGAVTVAWKADSSPVKA GVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS
(SEQ ID NO: 3593), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID
NO: 33) or a variant thereof, and
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQQKPGQ
APVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGGYDSSAGIFGG
GTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVK
AGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS
(SEQ ID NO: 3817),
an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or
[0008] a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, optionally wherein the first or second heavy chain comprises a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N- or C -terminus; and/or optionally wherein the composition comprises a linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof. In another aspect, the disclosure relates to a composition comprising an IgG-(scFv) antibody specific for Claudin 18.2 and CD3 comprising:
(a) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTF1SLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ
APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC
AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS
GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL
GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL
TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ
VSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQ
GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLV
QPGGSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVK
DRFTISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVT
VSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWV QQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYS NRWVFGGGTKLTVL (SEQ ID NO: 5202), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO:
5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
(B)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ
APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC
AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS
GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL
GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL
TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ
VSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQ
GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPV
QAGGSLRLSCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGR
FTISRDNAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS
(SEQ ID NO: 5205), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, or
(b) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ
VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ
QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4767), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
98%, or about 99% identity thereto, or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and (ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ VSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQ GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLV QPGGSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVK DRFTISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVT VSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWV QQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYS NRWVFGGGTKLTVL (SEQ ID NO: 5203), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
(B)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ VSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQ GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPV QAGGSLRLSCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGR FTISRDNAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 5206) or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, or
(c) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLWCLVKG FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4768), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, or
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL
NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKG
FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSV
MHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLVQPGGSLKL
SCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRD
DSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGG
SGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQA
PRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFG
GGTKLTVL (SEQ ID NO: 5204), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a
variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof,
(B)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR
QAPGKGLEWVGE1SSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKG FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPVQAGGSLRL SCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAK NTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 5207), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and
(b) a light chain selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWYQQKP GQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIF GGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTEC S (SEQ ID NO: 3576), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof,
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQKPGQ APVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIFGGG TKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKA GVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3593), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, and
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQQKPGQ APVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGGYDSSAGIFGG GTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVK AGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, optionally wherein the first or second heavy chain comprises a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N- or C -terminus; and/or optionally wherein the composition comprises a linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof
[0009] In certain embodiments of any of the above aspects, the Fc can be from IgG. In certain embodiments the IgG is human IgG. In certain embodiments, the human IgG is selected from IgGl, IgG2, IgG3, and IgG4.
[0010] In certain embodiments of any of the above aspects, the antibody is formed through a knob-in-hole interaction in the Fc region. In certain embodiments, the human IgG Fc comprises one or mutations to promote knob-in-hole interaction. In certain embodiments of any of the above aspects, the mutations are selected from (i) T366Y or T366W, and (ii) Y407T, Y407A, or Y407V. In certain embodiments of any of the above aspects, the mutations are selected from:
(a) T366Y and Y407T or T366Y/F405A and T394W/Y407T in human IgGl, and
(b) T366W/D399C and T366S/L368A/K392C/Y407V;
T366W/K392C and T366S/L368A/D399C/Y407V;
S354C/T366W and Y349C/T366S/L368A/Y407V;
Y349C/T366W and S354C/T366S/L368A/Y407V;
E356C/T366W and Y349C/T366S/L368A/Y407V;
Y349C/T366W and E356C/T366S/L368A/Y407V;
E357C/T366W and Y349C/T366S/L368A/Y407V; or
Y349C/T366W and E357C/T366S/L368A/Y407V, in human IgGl.
[0011] In certain embodiments of any of the above aspects, the human IgG Fc comprises one or mutations to reduce or eliminate the effector function of the Fc domains. In certain embodiments the mutations are L234A and L235A (LALA) substitutions in human IgGl.
[0012] In certain embodiments the human IgG Fc comprises one or mutations to stabilize a hinge region in the Fc domain. In certain embodiments, the mutation is S228P.
[0013] In another aspect, the disclosure relates to a composition comprising a tandem singlechain variable fragment (scFv) specific for Claudin 18.2 and CD3 comprising one or more of: (a) an amino acid sequence of:
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQQKPGQ APVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGGYDSSAGIFGG GTKLTVLGGGGSGGGGSGGGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAM MWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAE DTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSGGGGSEVQLVESGGGLVQPG GSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVAR1RSKYNNYATYYADSVKDRF nSRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVSS GGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQK PGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSNR
WVFGGGTKLTVL (SEQ ID: 4816), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or a variant thereof, the CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or a variant thereof, and the CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or
a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, or
(b) an amino acid sequence of:
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQKPGQ APVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIFGGG TKLTVLGGGGSGGGGSGGGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMG WVRQAPGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDT AVYYCAKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSGGGGSEVQLVES GGGLVQPGGSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATY YADSVKDRFTISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWG QGTLVTVSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGN YPNWVQQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYY
CVLWYSNRWVFGGGTKLTVL (SEQ ID NO. 4814), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or a variant thereof, the CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or a variant thereof, and the CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), wherein:
any of the above sequences in subsections (a) optionally can have a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N or C-terminus and any of the above sequences optionally have the linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof. [0014] In certain embodiments, the composition of any of the foregoing binds Claudin 18.2 and CD3 contemporaneously. In certain embodiments, the composition binds Claudin 18.2 with an affinity of less than 10 nM and with at least 100 fold greater affinity than Claudin 9, Claudin 3, and/or Claudin 4.
[0015] In certain embodiments, the first heavy chain, the second heavy chain, the light chain, and/or the single-chain variable fragment does not comprise amino acids 1-18 of its respective SEQ ID NO. In certain embodiments, the first heavy chain, the second heavy chain, the light chain, and/or the single-chain variable fragment does not comprise the amino acid sequence MKWVTFISLLFLFSSAYS (SEQ ID NO: 5356). In certain embodiments, the leader sequence, such as the amino acid sequence of SEQ ID NO: 5356, is removed during the expression of the antibody. The leader sequence of SEQ ID NO: 5356 can also be replaced with a different leader sequence to facilitate expression and/or secretion of the antibody. As provided for herein, in some embodiments, the antibody in a composition, such as a pharmaceutical composition, does not comprise a leader sequence, including, but not limited to the amino acid sequence of SEQ ID NO: 5356.
[0016] In another aspect, the disclosure relates to a pharmaceutical composition comprising an isolated antibody disclosed herein, or a nucleic acid molecule encoding the same. In certain embodiments, the pharmaceutical composition is an injectable pharmaceutical composition. In certain embodiments, the pharmaceutical composition is sterile. In certain embodiments, the pharmaceutical composition is pyrogen free.
[0017] In another aspect, the disclosure relates to a nucleic acid molecule encoding an antibody or an amino acid sequence as disclosed herein.
[0018] In another aspect, the disclosure relates to a vector comprising a nucleic acid molecule disclosed herein.
[0019] In another aspect, the disclosure relates to a cell comprising a nucleic acid molecule or vector as disclosure herein.
[0020] In another aspect, the disclosure relates to a method for modulating and/or targeting Claudin 18.2 and CD3 in a biological cell, comprising contacting the cell with a composition or pharmaceutical composition as disclosed herein.
[0021] In another aspect, the disclosure relates to a method for modulating Claudin 18.2 activity in a biological cell comprising contacting a cell expressing Claudin 18.2 with a composition or pharmaceutical composition as disclosed herein.
[0022] In another aspect, the disclosure relates to a method for inhibiting the function of Claudin 18.2 in a biological cell comprising contacting a cell expressing Claudin 18.2 with a composition or pharmaceutical composition as disclosed herein.
[0023] In another aspect, the disclosure relates to a method for treating or preventing cancer comprising administering an effective amount of the composition or pharmaceutical composition of any one of claims 1 to 52 as disclosed herein to a subject in need thereof. [0024] In another aspect, the disclosure relates to a use of the composition or pharmaceutical composition as disclosed herein for the preparation of a medicament for the treatment of prevention of cancer.
[0025] In certain embodiments, the cancer is selected from one or more of basal cell carcinoma, biliary tract cancer; bladder cancer; bone cancer; brain and central nervous system cancer; breast cancer; cancer of the peritoneum; cervical cancer; choriocarcinoma; colon and rectum cancer; connective tissue cancer; cancer of the digestive system; endometrial cancer; esophageal cancer; eye cancer; cancer of the head and neck; gastric cancer (including gastrointestinal cancer); glioblastoma; glioma; hepatic carcinoma; hepatoma; intra-epithelial neoplasm; kidney or renal cancer; larynx cancer; leukemia; liver cancer; lung cancer (e.g., small-cell lung cancer, non-small cell lung cancer, adenocarcinoma of the lung, and squamous carcinoma of the lung); melanoma; myeloma; neuroblastoma; oral cavity cancer (lip, tongue, mouth, and pharynx); ovarian cancer; pancreatic cancer; prostate cancer; retinoblastoma; rhabdomyosarcoma; malignant rhabdoid tumor; rectal cancer; cancer of the respiratory system; salivary gland carcinoma; sarcoma; skin cancer; squamous cell cancer; stomach cancer; testicular cancer; thyroid cancer; uterine or endometrial cancer; cancer of the urinary system; vulvar cancer; lymphoma including Hodgkin’s and non-Hodgkin's lymphoma, as well as B-cell lymphoma (including low grade/follicular non-Hodgkin's lymphoma (NHL); small lymphocytic (SL) NHL; intermediate grade/follicular NHL; intermediate grade diffuse NHL; high grade immunoblastic NHL; high grade lymphoblastic NHL; high grade small non-cleaved cell NHL; bulky disease NHL; mantle cell lymphoma; AIDS-related lymphoma; and Waldenstrom’s Macroglobulinemia; chronic lymphocytic leukemia (CLL); acute lymphoblastic leukemia (ALL); Hairy cell leukemia; chronic myeloblastic leukemia; as well as other carcinomas and sarcomas; and post-transplant
lymphoproliferative disorder (PTLD), as well as abnormal vascular proliferation associated with phakomatoses; edema (e.g. that associated with brain tumors); and Meigs’ syndrome. [0026] In another aspect, the disclosure relates to an isolated antibody comprising one or more of the sequences disclosed herein.
[0027] In another aspect, the disclosure relates to a method of treating or preventing cancer comprising administering an effective amount of one or more nucleic acid molecules (e.g., RNA) described herein to a subject in need thereof.
[0028] In certain embodiments, the cancer is selected from one or more of basal cell carcinoma, biliary tract cancer; bladder cancer; bone cancer; brain and central nervous system cancer; breast cancer; cancer of the peritoneum; cervical cancer; choriocarcinoma; colon and rectum cancer; connective tissue cancer; cancer of the digestive system; endometrial cancer; esophageal cancer; eye cancer; cancer of the head and neck; gastric cancer (including gastrointestinal cancer); glioblastoma; glioma; hepatic carcinoma; hepatoma; intra-epithelial neoplasm; kidney or renal cancer; larynx cancer; leukemia; liver cancer; lung cancer (e.g., small-cell lung cancer, non-small cell lung cancer, adenocarcinoma of the lung, and squamous carcinoma of the lung); melanoma; myeloma; neuroblastoma; oral cavity cancer (lip, tongue, mouth, and pharynx); ovarian cancer; pancreatic cancer; prostate cancer; retinoblastoma; rhabdomyosarcoma; malignant rhabdoid tumor; rectal cancer; cancer of the respiratory system; salivary gland carcinoma; sarcoma; skin cancer; squamous cell cancer; stomach cancer; testicular cancer; thyroid cancer; uterine or endometrial cancer; cancer of the urinary system; vulvar cancer; lymphoma including Hodgkin’s and non-Hodgkin’s lymphoma, as well as B-cell lymphoma (including low grade/follicular non-Hodgkin’s lymphoma (NHL); small lymphocytic (SL) NHL; intermediate grade/follicular NHL; intermediate grade diffuse NHL; high grade immunoblastic NHL; high grade lymphoblastic NHL; high grade small non-cleaved cell NHL; bulky disease NHL; mantle cell lymphoma; AIDS-related lymphoma; and Waldenstrom’s Macroglobulinemia; chronic lymphocytic leukemia (CLL); acute lymphoblastic leukemia (ALL); Hairy cell leukemia; chronic myeloblastic leukemia; as well as other carcinomas and sarcomas; and post-transplant lymphoproliferative disorder (PTLD), as well as abnormal vascular proliferation associated with phakomatoses; edema (e.g. that associated with brain tumors); and Meigs’ syndrome. [0029] In certain embodiments, the nucleic acid molecule comprises a sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity to any one of SEQ ID NOs: 5329-5355.
BRIEF DESCRIPTION OF THE DRAWINGS
[0030] FIGs. 1A-1F show images of illustrative bispecific formats, including a tandem scFv (FIG. 1A), a Hetero VH IgG (FIG. 1B), an scFv-Fab IgG (FIG. 1C), IgG-scFv (FIG. 1D), a IgG-(scFv)2 (FIG. 1E), and IgG-(scFv) (FIG. 1F).
[0031] FIG. 2 is graph of the IM-16-44-09-B03-1CD3 antibody (Format 16 (scFv-Fab IgG)) binding to HEK 293F cells expressing human P2X3, CD3, or Claudin (CLDN) 18.2.
[0032] FIG. 3 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 - or CLDN 18.2-expressing cells (HEK293F) in the presence of IMC-16-C2 (control) or IM-16-44-09-B03-1CD3 antibodies (Format 16 (scFv-Fab IgG)). Percent (%) specific cytotoxicity was assessed 48 hours (48hr) later. Legend: IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-16-44-09-B03-1CD3 on CLDN 18.1 (diamond), and IM-16-44-09-B03-1CD3 on CLDN 18.2 (triangle).
[0033] FIGs. 4A, 4B, 4C, 4D, and 4E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IMC-16-C2 or IM-16-44-09-B03-1CD3 antibodies. FIG. 4A illustrates the results for IL-2 production, FIG. 4B illustrates the results for IL-6 production, FIG. 4C illustrates the results for IL- 10 production, FIG. 4D illustrates the results for IFN-γ (INFgamma) production, and FIG. 4E illustrates the results for TNF-α (TNFalpha) production. Legend: IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-16-44-09-B03-1CD3 on CLDN 18.1 (diamond), and IM-16-44-09-B03-1CD3 on CLDN 18.2 (triangle).
[0034] FIGs. 5A and 5B show size exclusion chromatography (SEC) and differential scanning fluorimetry (DSF) experimental results for the IM-16-44-09-B03-1CD3, IM-16-44- 09-B03-2CD3, and IM-16-44-09-B03-3CD3 antibodies.
[0035] FIG. 6 is graph of the IM-21-44-01 -E10-3CD3 antibody (Format 21 (IgG-(scFv)2)) binding to HEK 293F cells expressing human P2X3, CD3, or CLDN 18.2.
[0036] FIG. 7 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IMC-16-C2 (control) or IM-21 -44-01-E10-3CD3 (Format 21 (IgG- (scFv)2)). Percent (%) specific cytotoxicity was assessed 48 hours (48hr) later. Legend: IMC- 16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-21-44-01 -E10- 3CD3 on CLDN 18.1 (diamond), and IM-21-44-01-E10-3CD3 on CLDN 18.2 (triangle).
[0037] FIGs. 8A, 8B, 8C, 8D, and 8E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IMC-16-C2
or IM-21-44-01-E10-3CD3. FIG. 8A illustrates the results for IL-2 production, FIG. 8B illustrates the results for IL-6 production, FIG. 8C illustrates the results for IL- 10 production, FIG. 8D illustrates the results for IFN-γ production, and FIG. 8E illustrates the results for TNF-α production. Legend: IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-21-44-01-E10-3CD3 on CLDN 18.1 (diamond), and IM-21-44-01-E10-3CD3 on CLDN 18.2 (triangle).
[0038] FIGs. 9 A and 9B show size exclusion chromatography (SEC) and differential scanning fluorimetry (DSF) experimental results for IM-21 -44-01 -E10-1CD3 and IM-21-44- 01 -E10-3CD3 antibodies.
[0039] FIG. 10 is graph of the IM-21 -44-09-B03-3CD3 antibody (Format 21 (IgG-(scFv)2)) binding to HEK 293F cells expressing human P2X3, CD3, or CLDN 18.2.
[0040] FIG. 11 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IMC-16-C2 (control) or IM-21 -44-09-B03-3CD3 (Format 21 (IgG- (scFv)2)). Percent (%) specific cytotoxicity was assessed 48 hours (48hr) later.
[0041] FIGs. 12A, 12B, 12C, 12D, and 12E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IMC-16-C2 or IM-21-44-09-B03-3CD3. FIG. 12A illustrates the results for IL-2 production, FIG. 12B illustrates the results for IL-6 production, FIG. 12C illustrates the results for IL- 10 production, FIG. 12D illustrates the results for IFN-γ production, and FIG. 12E illustrates the results for TNF- a production. Legend: IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-21 -44-09-B03-3CD3 on CLDN 18.1 (diamond), and IM-21-44- 09-B03-3CD3 on CLDN 18.2 (triangle).
[0042] FIGs. 13A and 13B show size exclusion chromatography (SEC) and differential scanning fluorimetry (DSF) experimental results for IM-21-44-09-B03-1CD3 and IM-21-44- 09-B03-3CD3 antibodies.
[0043] FIG. 14 is graph of the IM-2-44-05-H06-1CD3 antibody (Format 2 (tandem scFv)) binding to HEK 293F cells expressing human P2X3, CD3, or CLDN 18.2.
[0044] FIG. 15 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1- or CLDN 18.2-expressing HEK 293F cells in the presence of IMC-16-C2 (control) or IM-2-44-05-H06-1CD3 (Format 2 (tandem scFv)). Percent (%) specific cytotoxicity was assessed 48 hours (48hr) later. Legend: IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-2-44-05-H06-1CD3 on CLDN 18.1 (diamond), and IM-2-44-05-H06-1CD3 on CLDN 18.2 (triangle).
[0045] FIGs. 16A, 16B, 16C, 16D, and 16E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IMC-16-C2 or IM-2-44-05-H06-1CD3. FIG. 16A illustrates the results for IL-2 production, FIG. 16B illustrates the results for IL-6 production, FIG. 16C illustrates the results for IL- 10 production, FIG. 16D illustrates the results for IFN-γ production, and FIG. 16E illustrates the results for TNF-α production. Legend: IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-2-44-05-H06-1CD3 on CLDN 18.1 (diamond), and IM-2-44- 05-H06-1CD3 on CLDN 18.2 (triangle).
[0046] FIG. 17 is graph of the IM-2-44-09-B03-1CD3 antibody (Format 2 (tandem scFv)) binding to HEK 293F cells expressing human P2X3, CD3, or CLDN 18.2.
[0047] FIG. 18 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IMC-16-C2 (control) or IM-2-44-09-B03-1CD3 (Format 2 (tandem scFv)). Percent (%) specific cytotoxicity was assessed 48 hours (48hr) later. legend: IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-2-44-09-B03-1CD3 on CLDN 18.1 (diamond), and 1M-2-44-09-B03-1CD3 on CLDN 18.2 (triangle).
[0048] FIGs. 19A, 19B, 19C, 19D, and 19E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IMC-16-C2 or IM-2-44-09-B03-1CD3. FIG. 19A illustrates the results for IL-2 production, FIG. 19B illustrates the results for IL-6 production, FIG. 19C illustrates the results for IL- 10 production, FIG. 19D illustrates the results for IFN-γ production, and FIG. 19E illustrates the results for TNF-α production. Legend: IMC-16-C2 on CLDN 18.1 (circles), IMC-16-C2 on CLDN 18.2 (squares), IM-2-44-09-B03-1CD3 on CLDN 18.1 (diamond), and IM-2-44- 09-B03-1CD3 on CLDN 18.2 (triangle).
[0049] FIG. 20 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IMC-16-C2 (control) antibodies.
[0050] FIG. 21 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-42-44-09-B03-1CD3 (Format 42 EgG-(scFv)) antibodies. legend: IM- 42-44-09-B03-1CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0051] FIGs. 22A, 22B, 22C, 22D, and 22E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 09-B03-1CD3 antibodies. FIG. 22A illustrates the results for IL-2 production, FIG. 22B
illustrates the results for IL-6 production, FIG. 22C illustrates the results for IL- 10 production, FIG. 22D illustrates the results for IFN-γ production, and FIG. 22E illustrates the results for TNF-α production. Legend: IM-42-44-09-B03-1CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0052] FIGs. 23A, 23B, 23C, 23D, and 23E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 09-B03-1CD3 antibodies. FIG. 23A illustrates the results for IL-2 production, FIG. 23B illustrates the results for IL-6 production, FIG. 23C illustrates the results for IL- 10 production, FIG. 23D illustrates the results for IFN-γ production, and FIG. 23E illustrates the results for TNF-α production.
[0053] FIG. 24 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-42-44-09-B03-3CD3 antibodies (Format 42 IgG-(scFv)).
[0054] FIGs. 25A, 25B, 25C, 25D, and 25E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 09-B03-1CD3. FIG. 25A illustrates the results for IL-2 production, FIG. 25B illustrates the results for IL-6 production, FIG. 25C illustrates the results for IL- 10 production, FIG. 25D illustrates the results for IFN-γ production, and FIG. 25E illustrates the results for TNF-α production. legend: IM-42-44-09-B03-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0055] FIGs. 26A26B, 26C, 26D, and 26E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 09-B03-3CD3. FIG. 26A illustrates the results for IL-2 production, FIG. 26B illustrates the results for IL-6 production, FIG. 26C illustrates the results for IL- 10 production, FIG. 26D illustrates the results for IFN-γ production, and FIG. 26E illustrates the results for TNF-α production.
[0056] FIG. 27 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-42-44-01-E10-1CD3 antibodies (Format 42 IgG-(scFv)).
[0057] FIGs. 28A, 28B, 28C, 28D, and 28E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 01 -E10-1 CD3. FIG. 28A illustrates the results for IL-2 production, FIG. 28B illustrates the results for IL-6 production, FIG. 28C illustrates the results for IL- 10 production, FIG. 28D illustrates the results for IFN-γ production, and FIG. 28E illustrates the results for TNF-α
production. Legend: IM-42-44-01-E10-1CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0058] FIGs. 29A, 29B, 29C, 29D, and 29E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 01 -E10-1 CD3. FIG. 29A illustrates the results for IL-2 production, FIG. 29B illustrates the results for IL-6 production, FIG. 29C illustrates the results for IL- 10 production, FIG. 29D illustrates the results for IFN-γ production, and FIG. 29E illustrates the results for TNF-α production. Legend: IM-42-44-01 -E10-1CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0059] FIG. 30 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM -42-44-01 -E10-3CD3 antibodies (Format 42 IgG-(scFv)).
[0060] FIGs. 31A, 31B, 31C, 31D, and 31E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 01-E10-3CD3. FIG. 31A illustrates the results for IL-2 production, FIG. 31B illustrates the results for IL-6 production, FIG. 31C illustrates the results for IL- 10 production, FIG. 31D illustrates the results for IFN-γ production, and FIG. 31E illustrates the results for TNF-α production. Legend: IM-42-44-01 -E10-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0061] FIGs. 32A, 32B, 32C, 32D, and 32E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293 F cells with IM-42-44- 01 -E10-3CD3. FIG. 32A illustrates the results for IL-2 production, FIG. 32B illustrates the results for IL-6 production, FIG. 32C illustrates the results for IL- 10 production, FIG. 32D illustrates the results for IFN-γ production, and FIG. 32E illustrates the results for TNF-α production. Legend: IM-42-44-01-E10-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0062] FIG. 33 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-42-44-05-H06-1CD3 antibodies (Format 42 IgG-(scFv)).
[0063] FIGs. 34A, 34B, 34C, 34D, and 34E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 05-H06-1CD3. FIG. 34A illustrates the results for IL-2 production, FIG. 34B illustrates the results for IL-6 production, FIG. 34C illustrates the results for IL- 10 production, FIG. 34D illustrates the results for IFN-γ production, and FIG. 34E illustrates the results for TNF-α
production. Legend: IM-42-44-05 -H06-1CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0064] FIGs. 35A, 35B, 35C, 35D, and 35E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 05 -H06-1CD3. FIG. 35 A illustrates the results for IL-2 production, FIG. 35B illustrates the results for IL-6 production, FIG. 35C illustrates the results for IL- 10 production, FIG. 35D illustrates the results for IFN-γ production, and FIG. 35E illustrates the results for TNF-α production. Legend: IM-42-44-05-H06-1CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0065] FIG. 36 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-42-44-05-H06-3CD3 antibodies (Format 42 IgG-(scFv)).
[0066] FIGs. 37A, 37B, 37C, 37D, and 37E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-42-44- 05-H06-3CD3. FIG. 37 A illustrates the results for IL-2 production, FIG. 37B illustrates the results for IL-6 production, FIG. 37C illustrates the results for IL- 10 production, FIG. 37D illustrates the results for IFN-γ production, and FIG. 37E illustrates the results for TNF-α production. Legend: IM-42-44-05-H06-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0067] FIGs. 38A, 38B, 38C, 38D, and 38E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293 F cells with IM-42-44- 05-H06-3CD3. FIG. 38A illustrates the results for IL-2 production, FIG. 38B illustrates the results for IL-6 production, FIG. 38C illustrates the results for IL- 10 production, FIG. 38D illustrates the results for IFN-γ production, and FIG. 38E illustrates the results for TNF-α production. Legend: IM-42-44-05-H06-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0068] FIG. 39 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM -21 -44-09-B03-3CD3 antibodies (Format 21 (IgG-(scFv)2)).
[0069] FIGs. 40A, 40B, 40C, 40D, and 40E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-21 -44- 09-B03-3CD3 antibodies. FIG. 40A illustrates the results for IL-2 production, FIG. 40B illustrates the results for IL-6 production, FIG. 40C illustrates the results for IL- 10 production, FIG. 40D illustrates the results for IFN-γ production, and FIG. 40E illustrates
the results for TNF-α production. Legend: IM-21-44-09-B03-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0070] FIGs. 41A, 41B, 41C, 41D, and 41E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-21 -44- 09-B03-3CD3. FIG. 41A illustrates the results for IL-2 production, FIG. 41B illustrates the results for IL-6 production, FIG. 41C illustrates the results for IL- 10 production, FIG. 4 ID illustrates the results for IFN-γ production, and FIG. 41E illustrates the results for TNF-α production. Legend: IM-21 -44-09-B03-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0071] FIG. 42 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-21-44-01-E10-3CD3 antibodies (Format 21 (IgG-(scFv)2)).
[0072] FIGs. 43A, 43B, 43C, 43D, and 43E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-21-44- 01 -E10-3CD3 antibodies. FIG. 43 A illustrates the results for IL-2 production, FIG. 43B illustrates the results for IL-6 production, FIG. 43C illustrates the results for IL- 10 production, FIG. 43D illustrates the results for IFN-γ production, and FIG. 43E illustrates the results for TNF-α production. Legend: IM-21-44-01-E10-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0073] FIGs. 44A, 44B, 44C, 44D, and 44E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293 F cells with IM-21-44- 01 -E10-3CD3. FIG. 44A illustrates the results for IL-2 production, FIG. 44B illustrates the results for IL-6 production, FIG. 44C illustrates the results for IL- 10 production, FIG. 44D illustrates the results for IFN-γ production, and FIG. 44E illustrates the results for TNF-α production. Legend: IM-21-44-01-E10-3CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0074] FIG. 45 is a graph of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of IM-2-44-05-H06-1CD3 antibodies (Format 2 (tandem scFv)).
[0075] FIGs. 46A, 46B, 46C, 46D, and 46E show cytokines produced 24hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-2-44- 05-H06-1CD3 antibodies. FIG. 46A illustrates the results for IL-2 production, FIG. 46B illustrates the results for IL-6 production, FIG. 46C illustrates the results for IL- 10 production, FIG. 46D illustrates the results for IFN-γ production, and FIG. 46E illustrates
the results for TNF-α production. Legend: IM-2-44-05-H06-1CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0076] FIGs. 47A, 47B, 47C, 47D, and 47E show cytokines produced 48hr after co-culturing human PBMCs and human CLDN 18.1- or 18.2-expressing HEK 293F cells with IM-2-44- 05 -H06-1CD3. FIG. 47 A illustrates the results for IL-2 production, FIG. 47B illustrates the results for IL-6 production, FIG. 47C illustrates the results for IL- 10 production, FIG. 47D illustrates the results for IFN-γ production, and FIG. 47E illustrates the results for TNF-a production. Legend: IM-2-44-05-H06-1CD3 on CLDN 18.1 (circles) and CLDN 18.2 (triangles).
[0077] FIG. 48 is a set of graphs of T-cell dependent cellular cytotoxicity experiments where human T-cells were co-cultured with human CLDN 18.1 or CLDN 18.2 expressing HEK 293F cells in the presence of mRNA encoding IM-2-44-09B03-1CD3 or IM-16-44-09B03- 1CD3 antibodies.
DETAILED DESCRIPTION
Antibodies Specific for Claudin 18.2 and CD3
[0078] The present embodiments are based, in part, on the surprising discovery of antibodies specific for Claudin 18.2 and CD3. The present disclosure describes, in part, the isolation and characterization of antibodies, antibody fragments, and antibody variants specific for Claudin 18.2 and CD3. In some embodiments, the antibody is a tandem single-chain variable fragment (scFv) specific for Claudin 18.2 and CD3, which is for example, illustrated in FIG. 1A. In some embodiments, the antibody is a hetero VH IgG, such as illustrated in FIG. 1B and specific for Claudin 18.2 and CD3. In some embodiments, the antibody is an scFv-Fab IgG antibody format, which, for example, is illustrated in FIG. 1C and specifically binds to Claudin 18.2 and CD3. In some embodiments, the antibody is an IgG-scFv format, which, for example, is illustrated in FIG. 1D and specifically binds to Claudin 18.2 and CD3. In some embodiments, the antibody is an IgG-(scFv)2 antibody format, which, for example, is illustrated in FIG. 1E and specifically binds to Claudin 18.2 and CD3. In some embodiments, the antibody is a IgG-(scFv) format, which, for example, is illustrated in FIG. 1F and specifically binds to Claudin 18.2 and CD3. In some embodiments, the antibody specific for Claudin 18.2 and CD3 binds Claudin 18.2 and CD3 contemporaneously. Without being bound to any particular theory, the bispecific antibody will bind to a tumor cell expressing Claudin 18.2 and an immune cell, such as a cytotoxic T cell, expressing CD3. By bringing the cells together, the T cell is activated and targets the tumor cell expressing Claudin 18.2 for destruction.
[0079] For example, in some embodiments, the antibody specific for Claudin 18.2 and CD3 is a tandem scFv. An exemplary tandem scFv is shown schematically in FIG. 1A. A tandem scFv antibody specific for Claudin 18.2 and CD3 can be selected from an amino acid sequence of SEQ ID NO: 4816 and SEQ ID NO: 4814, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
[0080] In some embodiments, the antibody specific for Claudin 18.2 and CD3 is a hetero VH IgG antibody comprising two heavy chains and two light chains. The light chains optionally can comprise the same amino acid sequence. An exemplary hetero VH IgG is shown schematically in FIG. 1B. A hetero VH IgG antibody specific for Claudin 18.2 and CD3 can comprise a first heavy chain having an amino acid sequence of SEQ ID NO: 3869, SEQ ID NO: 4767, or SEQ ID NO: 4768 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a second heavy chain having an amino acid sequence comprising the heavy chain CDRs of any anti-CD3 scFv provided herein. In some embodiments, the light chain comprises an amino acid sequence of SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
[0081] In some embodiments, the antibody specific for Claudin 18.2 and CD3 is an scFv-Fab IgG antibody comprising a first heavy chain, a second heavy chain, and a light chain. An exemplary scFv-Fab IgG antibody is shown schematically in FIG. 1C. An antibody specific for Claudin 18.2 and CD3 can comprise a first heavy chain having an amino acid sequence of SEQ ID NO: 3869, SEQ ID NO: 4767, or SEQ ID NO: 4768, a second heavy chain having an amino acid sequence of SEQ ID NO: 4168, SEQ ID NO: 4169, or SEQ ID NO: 3983, and a light chain having an amino acid sequence of SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817. In some embodiments, the first heavy chain comprises an amino acid sequence of SEQ ID NO: 3869, SEQ ID NO: 4767, or SEQ ID NO: 4768, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. In some embodiments, the second heavy comprises an amino acid sequence of SEQ ID NO: 4168, SEQ ID NO: 4169, or SEQ ID NO: 3983, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. In some embodiments, the light chain comprises an amino acid sequence of SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817, or an amino acid sequence
having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
[0082] In some embodiments, the antibody specific for Claudin 18.2 and CD3 is a IgG-scFv antibody comprising two heavy chains (a first and second heavy chain) and two light chains. In some embodiments, the two heavy chains comprising amino acid sequences that are different from each other, and the two light chains comprise amino acid sequences that are identical. An exemplary IgG-scFv is shown schematically in FIG. 1D. In some embodiments, the first heavy chain has an amino acid sequence of SEQ ID NO: 3869, SEQ ID NO: 4767, or SEQ ID NO: 4768 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. In some embodiments, the second heavy chain has an amino acid sequence of SEQ ID NO: 4778, SEQ ID NO: 4779, SEQ ID NO: 4780, SEQ ID NO: 4803, SEQ ID NO: 4804, SEQ ID NO: 4805, SEQ ID NO: 4806, SEQ ID NO: 4807, SEQ ID NO: 4808 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. In certain embodiments, the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 3869 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a second heavy chain selected from SEQ ID NO: 4778, SEQ ID NO: 4803, and SEQ ID NO: 4806 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto In certain embodiments, the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 4767 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a second heavy chain selected from SEQ ID NO: 4779, SEQ ID NO: 4804, and SEQ ID NO: 4807 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. In certain embodiments, the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 4768 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a second heavy chain selected from SEQ ID NO: 4780, SEQ ID NO: 4805, and SEQ ID NO: 4808 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. The two light chains can be selected from SEQ ID NO: 3593, SEQ ID NO: 3576, and SEQ ID NO: 3817 or an amino acid sequence having at least about 90%,
about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. The two light chains can comprise the same amino acid sequence.
[0083] In some embodiments, the antibody specific for Claudin 18.2 and CD3 is an IgG- (SCFV)2 antibody comprising two heavy chains and two light chains, wherein the heavy chains are identical to each other and the light chains are identical to each other. An exemplary IgG- (SCFV)2 antibody is shown schematically in FIG. 1E. In some embodiments, the antibody comprises a heavy chain selected comprising an amino acid sequence of SEQ ID NO: 4769, SEQ ID NO: 4770, SEQ ID NO: 4771, SEQ ID NO: 4772, SEQ ID NO: 4773, SEQ ID NO: 4774, SEQ ID NO: 4775, SEQ ID NO: 4776, or SEQ ID NO: 4777, and a light chain selected from an amino acid sequence of SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817. In some embodiments, the heavy chain comprises an amino acid sequence of SEQ ID NO: 4769, SEQ ID NO: 4770, SEQ ID NO: 4771, SEQ ID NO: 4772, SEQ ID NO: 4773, SEQ ID NO: 4774, SEQ ID NO: 4775, SEQ ID NO: 4776, or SEQ ID NO: 4777,, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. In some embodiments, the light chain comprises an amino acid sequence of SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
[0084] In some embodiments, the antibody specific for Claudin 18.2 and CD3 is a IgG- (scFv) antibody comprising two heavy chains (a first and second heavy chain) and two light chains. In some embodiments, the two heavy chains comprising amino acid sequences that are different from each other, and the two light chains comprise amino acid sequences that are identical. An exemplary IgG-scFv is shown schematically in FIG. 1E. In some embodiments, the first heavy chain has an amino acid sequence of SEQ ID NO: 3869, SEQ ID NO: 4767, or SEQ ID NO: 4768, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. In some embodiments, the second heavy chain has an amino acid sequence of SEQ ID NO: 5202, SEQ ID NO: 5205, SEQ ID NO: 5203, SEQ ID NO: 5206, SEQ ID NO: 5204, or SEQ ID NO: 5207, or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. In some embodiments, the two light chains each have an amino acid sequence comprising SEQ ID NO: 3593, SEQ ID NO: 3576, or SEQ ID NO: 3817 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. The two light chains can comprise the same amino acid sequence. In certain embodiments, the antibody specific for
Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 3869 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, a second heavy chain having an amino acid sequence of SEQ ID NO: 5202 or 5205 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a light chain (e.g., a first and second light chain) having an amino acid sequence of SEQ ID NO: 3593 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. In certain embodiments, the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 4767 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, a second heavy chain having an amino acid sequence of SEQ ID NO: 5203 or SEQ ID NO: 5206 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a light chain (e.g., a first and second light chain) having an amino acid sequence of SEQ ID NO: 3576 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto. In certain embodiments, the antibody specific for Claudin 18.2 and CD3 comprises a first heavy chain having an amino acid sequence of SEQ ID NO: 4768 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a second heavy chain having an amino acid sequence of SEQ ID NO: 5204 or 5207 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, and a light chain having an amino acid sequence of SEQ ID NO: 3817 or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto.
[0085] In various embodiments, the antibody (e.g., a tandem scFv antibody, a hetero VH IgG antibody, an scFv-Fab IgG antibody, an IgG-scFv antibody, IgG-(scFv)? antibody, and a IgG- (scFv) antibody), or fragment thereof, or variant thereof, may comprise an amino acid sequence having one or more amino acid mutations (e.g., substitutions or deletions) relative to any of the sequences disclosed herein. In some embodiments, the one or more amino acid mutations may be independently selected from substitutions, insertions, deletions, and truncations. In embodiments, the antibody (e.g., a tandem scFv antibody, a hetero VH IgG antibody, an scFv-Fab IgG antibody, an IgG-scFv antibody, IgG-(scFv)2 antibody, and a IgG- (scFv) antibody), or fragment thereof, or variant thereof, comprises a sequence that has about
1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 or more amino acid mutations with respect to any one of the amino acid sequences disclosed herein.
[0086] In various embodiments, the antibody or antibody format (e.g. , a tandem scFv antibody, a hetero VH IgG antibody, an scFv-Fab IgG antibody, an IgG-scFv antibody, IgG- (SCFV)2 antibody, and a IgG-(scFv) antibody), or fragment thereof, or variant thereof, may comprise an amino acid sequence having al least about 50%, about 60%, about 70%, about 80%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99% sequence homology to the amino acid sequences disclosed herein.
[0087] In various embodiments, the present invention provides for variants or fragments comprising any of the sequences described herein, for instance, a sequence having at least about 60%, or at least about 61%, or at least about 62%, or at least about 63%, or at least about 64%, or at least about 65%, or at least about 66%, or at least about 67%, or at least about 68%, or at least about 69%, or at least about 70%, or at least about 71%, or at least about 72%, or at least about 73%, or at least about 74%, or at least about 75%, or at least about 76%, or at least about 77%, or at least about 78%, or at least about 79%, or at least about 80%, or at least about 81%, or at least about 82%, or at least about 83%, or at least about 84%, or at least about 85%, or at least about 86%, or at least about 87%, or at least about 88%, or at least about 89%, or at least about 90%, or at least about 91%, or at least about 92%, or at least about 93%, or at least about 94%, or at least about 95%, or at least about 96%, or at least about 97%, or at least about 98%, or at least about 99%) sequence identity with any of the sequences disclosed herein.
[0088] In some embodiments, variants are those that have conservative amino acid substitutions made at one or more predicted non-essential amino acid residues. For example, a “conservative amino acid substitution” is one in which the amino acid residue is replaced with an amino acid residue having a similar side chain.
[0089] In embodiments, the amino acid mutations are amino acid substitutions, and may include conservative and/or non-conservative substitutions.
[0090] “Conservative substitutions” may be made, for instance, on the basis of similarity in polarity, charge, size, solubility, hydrophobicity, hydrophilicity, and/or the amphipathic nature of the amino acid residues involved. The 20 naturally occurring amino acids can be grouped into the following six standard amino acid groups: (1) hydrophobic: Met, Ala, Vai, Leu, Ile; (2) neutral hydrophilic: Cys, Ser, Thr; Asn, Gin; (3) acidic: Asp, Glu; (4) basic: His, Lys, Arg; (5) residues that influence chain orientation: Gly, Pro; and (6) aromatic: Trp, Tyr, Phe.
[0091] As used herein, “conservative substitutions” are defined as exchanges of an amino acid by another amino acid listed within the same group of the six standard amino acid groups shown above. For example, the exchange of Asp by Glu retains one negative charge in the so modified polypeptide. In addition, glycine and proline may be substituted for one another based on their ability to disrupt α-helices.
[0092] As used herein, “non-conservative substitutions” are defined as exchanges of an amino acid by another amino acid listed in a different group of the six standard amino acid groups (1) to (6) shown above.
[0093] As disclosed herein, the term “antibody” refers to a broad sense and includes immunoglobulin or antibody molecules including polyclonal antibodies, monoclonal antibodies including murine, human, humanized and chimeric monoclonal antibodies and antibody fragments, such as ScFv (PLOS Biology | D01:10.1371/joumal.pbio.l002344 January 6, 2016, which is hereby incorporated by reference in its entirety).
[0094] In general, antibodies are proteins or polypeptides that exhibit binding specificity to a specific antigen. Intact antibodies are heterotetrameric glycoproteins, composed of two identical light chains and two identical heavy chains. Typically, each light chain is linked to a heavy chain by one covalent disulfide bond, while the number of disulfide linkages varies between the heavy chains of different immunoglobulin isotypes. Each heavy and light chain also has regularly spaced intrachain disulfide bridges. Each heavy chain has at one end a variable domain (VH) followed by a number of constant domains. Each light chain has a variable domain at one end (VL) and a constant domain at its other end; the constant domain of the light chain is aligned with the first constant domain of the heavy chain and the light chain variable domain is aligned with the variable domain of the heavy chain. Antibody light chains of any vertebrate species can be assigned to one of two clearly distinct types, namely kappa and lambda, based on the amino acid sequences of their constant domains.
Immunoglobulins can be assigned to five major classes, namely IgA, IgD, IgE, IgG and IgM, depending on the heavy chain constant domain amino acid sequence. IgA and IgG are further sub-classified as the isotypes IgAl, IgA2, IgGl, IgG2, IgG3 and IgG4.
[0095] As disclosed herein, the term “antibody fragment” refers to an intact antibody, generally the antigen binding or variable region of the intact antibody. Examples of antibody fragments include Fab, Fab', F(ab')2 and Fv fragments, diabodies, single chain antibody molecules and multispecific antibodies formed from at least two intact antibodies or fragments thereof.
[0096] As disclosed herein, the term “antigen” refers to any molecule that has the ability to generate antibodies either directly or indirectly.
[0097] As disclosed herein, the term “specific binding” or “immunospecific binding” or “binds immunospecifically” refers to antibody binding to a predetermined antigen (e.g., Claudin 18.2) or epitope present on the antigen. In some embodiments, the antibody binds with a dissociation constant (KD) of about 10-10 M or less, of about 10-9 M or less, of about 10-8 M or less, of about 10-7 M or less, of about 10-6 M or less, of about 10-5 M or less, and binds to the predetermined antigen with a KD that is at least two-fold less than its KD for binding to a non-specific antigen (e.g., BSA, casein, or another non-specific polypeptide) other than the predetermined antigen. The phrases “an antibody recognizing Claudin 18.2” and “an antibody specific for Claudin 18.2” are used interchangeably herein with the term “an antibody which binds immunospecifically to Claudin 18.2.” Reference in the present disclosure may be made to Claudin 18.2. In some embodiments, the antibody is specific for Claudin 18.2 and does not specifically bind to Claudin 3, Claudin 4, and/or Claudin 9.
[0098] “CDRs” are referred to as the complementarity determining region amino acid sequences of an antibody which are the hypervariable regions of immunoglobulin heavy and light chains. See, e.g., Kabat et al., Sequences of Proteins of Immunological Interest, 4th ed., U.S. Department of Health and Human Services, National Institutes of Health (1987). There are three heavy chain and three light chain CDRs or CDR regions in the variable portion of an immunoglobulin. Thus, “CDRs” as used herein refers to all three heavy chain CDRs, or all three light chain CDRs or both all heavy and all light chain CDRs, if appropriate.
[0099] Each variable region comprises three hypervariable regions also known as complementarity determining regions (CDRs) flanked by four relatively conserved framework regions (FRs). The three CDRs, referred to as CDR1, CDR2, and CDR3, contribute to the antibody binding specificity, as the CDRs provide the majority of contact residues for the binding of the antibody to the antigen or epitope. CDRs of interest can be derived from donor antibody variable heavy and light chain sequences, and include analogs of the naturally occurring CDRs, which analogs also share or retain the same antigen binding specificity and/or neutralizing ability as the donor antibody from which they were derived. In some embodiments, the antibody is a chimeric antibody. In some embodiments, the antibody is a humanized antibody.
[00100] CDRs are based on sequence variability (Wu and Kabat, J. Exp. Med. 132:211-250, 1970). There are six CDRs— three in the variable heavy chain, or VH, and are typically designated H-CDR1, H-CDR2, and H-CDR3, and three CDRs in the variable light chain, or
VL, and are typically designated L-CDR1 , L-CDR2, and L-CDR3 (Kabat et al. , Sequences of Proteins of Immunological Interest, Sth Ed. Public Health Service, National Institutes of Health, Bethesda, Md., 1991). “Hypervariable region”, “HVR”, or “HV” refer to the regions of an antibody variable domain which are variable in structure as defined by Chothia and Lesk (Chothia and Lesk, Mol. Biol. 196:901-917, 1987). There are six HVRs, three in VH (Hl, H2, H3) and three in VL (LI, L2, L3). Chothia and Lesk refer to structurally conserved HVs as “canonical structures.” Another method of describing the regions that form the antigen-binding site has been proposed by Lefranc (Lefranc et al., Developmental & Comparative Immunology 27:55-77, 2003) based on the comparison of V domains from immunoglobulins and T-cell receptors (Lefranc et al., Developmental & Comparative Immunology 27:55-77, 2003). The antigen-binding site can also be delineated based on “Specificity Determining Residue Usage (SDRU)", according to Almagro (Almagro, Mol. Recognit. 17:132-43, 2004), where SDRU refers to amino acid residues of an immunoglobulin that are directly involved in antigen contact.
[00101] An “isolated antibody,” as used herein, refers to an antibody that is substantially free of other antibodies having different antigenic specificities (e.g., an isolated antibody that specifically binds Claudin 18.2 is substantially free of antibodies that specifically bind antigens other than Claudin 18.2). Moreover, an isolated antibody may be substantially free of other cellular material and/or chemicals. An isolated antibody can also be sterile or pyrogen free or formulated as injectable pharmaceutical as described herein.
[00102] In some embodiments, the source for the DNA encoding a non-human antibody include cell lines which produce antibody, such as hybrid cell lines commonly known as hybridomas.
[00103] In some embodiments, the antibody or antibody fragment or variant disclosed herein (e.g., a tandem scFv antibody, a hetero VH IgG antibody, an scFv-Fab IgG antibody, an IgG- scFv antibody, IgG-(scFv)2 antibody, and a IgG-(scFv) antibody) comprises a CD3 engager. In some embodiments, the CD3 engager can be selected from muOKT3, huOKT3, huSP34, huUCHTl and a CD3 nanobody (VHH).
[00104] In some embodiments, the antibody or antibody fragment or variant is an antibody provided in TABLES 1-5, or an antibody fragment or variant thereof having at least about 60%, or at least about 61%, or at least about 62%, or at least about 63%, or at least about 64%, or at least about 65%, or at least about 66%, or at least about 67%, or at least about 68%, or at least about 69%, or at least about 70%, or at least about 71%, or at least about 72%, or at least about 73%, or at least about 74%, or at least about 75%, or at least about
76%, or at least about 77%, or at least about 78%, or at least about 79%, or at least about
80%, or at least about 81%, or at least about 82%, or at least about 83%, or at least about
84%, or at least about 85%, or at least about 86%, or at least about 87%, or at least about
88%, or at least about 89%, or at least about 90%, or at least about 91%, or at least about
92%, or at least about 93%, or at least about 94%, or at least about 95%, or at least about
96%, or al least about 97%, or al least about 98%, or al least about 99%) sequence identity with an antibody disclosed therein.
[00105] In certain embodiments, the leader sequence, such as the amino acid sequence of SEQ ID NO: 5356, is removed during the expression of the antibody. The leader sequence of SEQ ID NO: 5356 can also be replaced with a different leader sequence to facilitate expression and/or secretion of the antibody. As provided for herein, in some embodiments, the antibody in a composition, such as a pharmaceutical composition, does not comprise a leader sequence, including, but not limited to the amino acid sequence of SEQ ID NO: 5356. [00106] In some embodiments, the antibody or antibody fragment or variant is an antibody provided in TABLES 1-5, wherein the first heavy chain, the second heavy chain, the light chain, and/or the single-chain variable fragment does not comprise the signal sequence of amino acids 1-18 relative to the respective SEQ ID NO (e.g., in some embodiments, a fragment of the antibody or variant comprises a sequence provided in TABLE A). In some embodiments, the antibody or antibody fragment or variant is an antibody provided in TABLES 1-5, wherein the first heavy chain, the second heavy chain, the light chain, and/or the single-chain variable fragment does not comprise the amino acid sequence MKWVTFISLLFLFSSAYS (SEQ ID NO: 5356) (e.g., in some embodiments, a fragment of the antibody or variant comprises a sequence provided in TABLE A).
TABLE A: Antibody Components Described in TABLES 1-5 Not Comprising the Amino Acid Sequence MKWVTFISLLFLFSSAYS (SEQ ID NO: 5356)
[00107] In one aspect, the disclosure relates to a composition comprising an scFv-Fab IgG antibody specific for Claudin 18.2 and CD3 comprising a first heavy chain, a second heavy chain, and a light chain, wherein:
(a) the first heavy chain is selected from:
(i)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ
VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ
QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, or
(it)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ
VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ
QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4767), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, or (iii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL
NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLWCLVKG FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4768), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(b) the second heavy chain is selected from:
(i)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAMNWVR
QAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQMNNLKTEDT
AVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGGSGGGGSQTVVTQEP
SLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFS
GSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLASPKSSDKTHTC
PPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVE
VHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKA
KGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPP
VLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4168), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), or
(ii)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFTRYTMHWVR
QAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKSTAFLQMDSLRPEDTAVYY
CARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGGGGSDIQMTQSPSSLSASVG
DRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTI
SSLQPEDIATYYCQQWSSNPFTFGQGTKLQITRASPKSSDKTHTCPPCPAPEAAGGPS
VFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQ
YNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTL
PPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVS
KLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4169), o
an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, or
(iii)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSMGWFR QAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTSLKPEDSATY YCAAK1RPY1FK1AGQYDYWGQGTQVTVSSASPKSSDKTHTCPPCPAPEAAGGPSVFL FPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNS TYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSR EEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTV DKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3983), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and
(c) the light chain selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWYQQKP
GQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIF
GGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP
VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTEC S (SEQ ID NO: 3576), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID
NO: 33) or a variant thereof, or
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQKPGQ APVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIFGGG TKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKA GVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS
(SEQ ID NO: 3593), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQQKPGQ
APVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGGYDSSAGIFGG
GTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVK
AGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or
a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or wherein: any of the above sequences in subsections (a) optionally can have a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N or C-terminus and any of the above sequences optionally have the linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof. [00108] In another aspect, the disclosure relates to a composition comprising an IgG-scFv antibody specific for Claudin 18.2 and CD3 comprising:
(a) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTF1SLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSMGWFR
QAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTSLKPEDSATY
YCAAKIRPYIFKIAGQYDYWGQGTQVTVSSGGGGSGGGGSGGGGSEVQLLESGGGL
VQPGGSLRLSCAASGFTFSSYGMGWVRQAPGKGLEWVAAITRDSSTNYGAAVKGR
ATISRDNSKNTLYLQMNSLRAEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDA
WGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGAL TSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCD KTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY
VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIE
KTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENN
YKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPG
K (SEQ ID NO: 4778), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof,
(B)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAMNWVR
QAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQMNNLKTEDT
AVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGGSGGGGSQTVVTQEP
SLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFS
GSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLGGGGSGGGGSG
GGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQAPGKGLEWVAAIT
RDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKDASNGYCWYT
GRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDY
FPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPS
NTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVV
DVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKE YKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPS DIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEA LHNHYTQKSLSLSPGK (SEQ ID NO: 4803), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO:
5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
(C)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFTRYTMHWVR
QAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKSTAFLQMDSLRPEDTAVYY
CARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGGGGSDIQMTQSPSSLSASVG
DRVTJTCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTI
SSLQPEDIATYYCQQWSSNPFTFGQGTKLQITRGGGGSGGGGSGGGGSEVQLLESGG
GLVQPGGSLRLSCAASGFTFSSYGMGWVRQAPGKGLEWVAAITRDSSTNYGAAVK
GRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHI
DAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSG
ALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKS
CDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFN
WYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPA
PIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPE
NNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS PGK (SEQ ID NO: 4806); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, or
(b) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4767), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a
variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSMGWFR
QAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTSLKPEDSATY
YCAAKIRPYIFKIAGQYDYWGQGTQVTVSSGGGGSGGGGSGGGGSEVQLLESGGGL
VQPGGSLRLSCAASGFTFSSYGMSWVRQAPGKGLEWVAGIGSSGSDTWYADSVKG
RFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHHID
AWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGA
LTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSC
DKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNW
YVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPI EKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPEN NYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSP GK (SEQ ID NO: 4779), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, (B)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAMNWVR QAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQMNNLKTEDT AVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGGSGGGGSQTVVTQEP SLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFS
GSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLGGGGSGGGGSG
GGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQAPGKGLEWVAGIG
SSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKDATNGYCWY
TGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKD
YFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKP
SNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVV
DVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKE
YKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPS
DIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEA
LHNHYTQKSLSLSPGK (SEQ ID NO: 4804), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNS Y1SYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and (C)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFTRYTMHWVR QAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKSTAFLQMDSLRPEDTAVYY CARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGGGGSDIQMTQSPSSLSASVG DRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTI SSLQPEDIATYYCQQWSSNPFTFGQGTKLQITRGGGGSGGGGSGGGGSEVQLLESGG GLVQPGGSLRLSCAASGFTFSSYGMSWVRQAPGKGLEWVAGIGSSGSDTWYADSV
KGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHH
IDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNS
GALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEP
KSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKF
NWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP
APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQP
ENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSL
SPGK (SEQ ID NO: 4807); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, and/or heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, or (c) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL
NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLWCLVKG
FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4768), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSMGWFR QAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTSLKPEDSATY YCAAKIRPYIFKIAGQYDYWGQGTQVTVSSGGGGSGGGGSGGGGSEVQLLESGGGL VQPGGSLRLSCAASGFTFRSYAMMWVRQAPGKGLEWVGEISSTGGSTYYGSAVKG RAT1SRDNSKNTLYLQMNSLRAEDTAVYYCAKSTYGATWKSADS1DAWGQGTLVT VSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPA
VLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCP
APEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN
AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQ PREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLD SDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4780), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23)
or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA
(SEQ ID NO: 24) or a variant thereof,
(B)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAMNWVR
QAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQMNNLKTEDT
AVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGGSGGGGSQTVVTQEP
SLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFS GSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLGGGGSGGGGSG
GGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVRQAPGKGLEWVGEIS
STGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKSTYGATWKS
ADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVS
WNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKK
VEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPE
VKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNK ALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESN GQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKS LSLSPGK (SEQ ID NO: 4805), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(C)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFTRYTMHWVR
QAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKSTAFLQMDSLRPEDTAVYY
CARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGGGGSDIQMTQSPSSLSASVG
DRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTI
SSLQPEDIATYYCQQWSSNPFTFGQGTKLQITRGGGGSGGGGSGGGGSEVQLLESGG
GLVQPGGSLRLSCAASGFTFRSYAMMWVRQAPGKGLEWVGEISSTGGSTYYGSAV
KGRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTL
VTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTF
PAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPP
CPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVH
NAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKG QPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVL DSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4808); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(b) a light chain selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWYQQKP GQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIF GGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTEC S (SEQ ID NO: 3576), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof,
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTAR1TCSGGYNGHYGWYQQKPGQ APVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIFGGG TKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKA GVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3593), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQQKPGQ APVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGGYDSSAGIFGG GTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVK AGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), or
an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, wherein: any of the above sequences in subsections (a) optionally can have a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N or C-terminus and any of the above sequences optionally have the linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof. [00109] In another aspect, the disclosure relates to a composition comprising an IgG-(scFv)2 antibody specific for Claudin 18.2 and CD3 comprising:
(a) a heavy chain selected from:
(i)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL
TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQV
SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQG
NVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLVQ
PGGSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKD
RFTISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTV
SSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQ
QKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSN
RWVFGGGTKLTVL (SEQ ID NO: 4769), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about
98%, or about 99% identity thereto, or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), (ii) MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQV SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQG NVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSQVQLVQSGGGVVQ PGRSLRLSCKASGYTFTRYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFT ISTDKSKSTAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGGSG GGGSGGGGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAPKRWIY DTSKLASGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTFGQGTKLQITR (SEQ ID NO: 4770), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant
thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof,
(iii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ
APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC
AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS
GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL
GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL
TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQV
SLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQG
NVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPVQ
AGGSLRLSCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRF
TISRDNAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS
(SEQ ID NO: 4771), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSS Y GMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a
variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, (iv)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPffiKTISKAKGQPREPQVYTLPPSRDELTKNQ VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQ GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLV QPGGSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVK DRFTISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVT VSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWV QQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYS NRWVFGGGTKLTVL (SEQ ID NO: 4772), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9),
(v)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQ
VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQ
GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSQVQLVQSGGGVV
QPGRSLRLSCKASGYTFTRYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRF TISTDKSKSTAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGGS GGGGSGGGGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAPKRWI YDTSKLASGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTFGQGTKLQIT R (SEQ ID NO: 4773), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof.
(vi)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQ
VSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQ
GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPV
QAGGSLRLSCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGR
FTISRDNAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS
(SEQ ID NO: 4774), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, (vii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR
QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL
NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKG
FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV
MHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLVQPGGSLKL
SCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRD
DSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGG SGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQA PRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFG GGTKLTVL (SEQ ID NO: 4775), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), (viii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR
QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL
NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKG
FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV
MHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSQVQLVQSGGGVVQPGRSLRL
SCKASGYTFTRYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSK
STAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGG
GGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLA
SGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTFGQGTKLQITR (SEQ ID NO: 4776), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, and
(ix.)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR
QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKG FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPVQAGGSLRL
SCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAK NTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 4777), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and
(b) a light chain selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWYQQKP GQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIF GGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCL1SDFYPGAVTVAWKADSSP VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTEC
S (SEQ ID NO: 3576), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof,
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQKPGQ
APVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIFGGG TKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKA GVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3593), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or
a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, and
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQQKPGQ APVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGGYDSSAGIFGG GTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVK AGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or wherein: any of the above sequences in subsections (a) optionally can have a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N or C-terminus and any of the above sequences optionally have the linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof. [00110] In another aspect, the disclosure relates to a composition comprising an IgG-(scFv) antibody specific for Claudin 18.2 and CD3 comprising:
(a) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL
GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ
QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ APGKGLEWVAAITRDSSTNYGAAVKGRAT1SRDNSKNTLYLQMNSLRAEDTAVYYC AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ VSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQ
GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLV
QPGGSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVK
DRFTISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVT VSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWV QQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYS
NRWVFGGGTKLTVL (SEQ ID NO: 5202), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant
thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
(B)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQ
APGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYC
AKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS
GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSL
GTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVL
TVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ
VSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQ
GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPV
QAGGSLRLSCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGR
FTISRDNAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS
(SEQ ID NO: 5205), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a
variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, or
(b) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ VSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4767), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and (ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ
VSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQ
GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLV
QPGGSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVK
DRFTISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVT
VSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWV QQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYS NRWVFGGGTKLTVL (SEQ ID NO: 5203), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
(B)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQ
APGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYY
CAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSS
LGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKD
TLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSV
LTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ
VSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQ
GNVFSCSVMHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPV
QAGGSLRLSCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGR
FTISRDNAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 5206) or
an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, or
(c) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR QAPGKGLEWVGE1SSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLWCLVKG FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4768), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, or
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL
NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKG
FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSV
MHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLVQPGGSLKL
SCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRD
DSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGG
SGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQA
PRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFG GGTKLTVL (SEQ ID NO: 5204), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof,
(B)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVR
QAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVY
YCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVT
CVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKG FYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSV MHEALHNHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPVQAGGSLRL SCAASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAK NTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 5207), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and
(b) a light chain selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWYQQKP GQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIF GGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSP VKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTEC S (SEQ ID NO: 3576), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the
light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof,
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQKPGQ
APVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIFGGG
TKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVKA GVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3593), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG(SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, and
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQQKPGQ APVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGGYDSSAGIFGG GTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKADSSPVK AGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQVTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, wherein: any of the above sequences in subsections (a) optionally can have a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N or C-terminus and
any of the above sequences optionally have the linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof. [00111] In another aspect, the disclosure relates to a composition comprising a tandem single-chain variable fragment (scFv) specific for Claudin 18.2 and CD3 comprising one or more of:
(a) an amino acid sequence of:
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQQKPGQ APVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGGYDSSAGIFGG GTKLTVLGGGGSGGGGSGGGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAM MWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAE DTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSGGGGSEVQLVESGGGLVQPG GSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRF TISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVSS GGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQK PGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSNR WVFGGGTKLTVL (SEQ ID: 4816), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or a variant thereof, the CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or a variant thereof, and the CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, or
(b) an amino acid sequence of:
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQKPGQ APVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGYDSSAGIFGGG TKLTVLGGGGSGGGGSGGGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMG WVRQAPGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDT AVYYCAKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSGGGGSEVQLVES GGGLVQPGGSLKLSCAASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATY YADSVKDRFTISRDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWG QGTLVTVSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGN YPNWVQQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYY
CVLWYSNRWVFGGGTKLTVL (SEQ ID NO. 4814), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or a variant thereof, the CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or a variant thereof, and the CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or a variant thereof, and/or a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), wherein: any of the above sequences in subsections (a) optionally can have a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N or C-terminus and any of the above sequences optionally have the linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof.
[00112] In some embodiments, the antibody or antibody fragment or variant disclosed herein (e.g., a tandem scFv antibody, a hetero VH IgG antibody, an scFv-Fab IgG antibody, an IgG- scFv antibody, IgG-(scFv)z antibody, and a IgG-(scFv) antibody) comprise a linker having one or more glycines and serines replaced with a functionally equivalent variation thereof. In some embodiments, the linker is identified in the underlined text above. For example, in some embodiments, the linker is selected from the amino acid sequence of GGGGSGGGGSGGGGS (SEQ ID NO: 50), GGGGS (SEQ ID NO: 51), GGSGGSGGSGGSGGVD (SEQ ID NO: 52), and GKPGSGKPGSGKPGSGKPGS (SEQ ID NO: 53).
[00113] In some embodiments, the linker may be derived from naturally-occurring multidomain proteins or are empirical linkers as described, for example, in Chichili et al., (2013), Protein Sci. 22(2): 153- 167, Chen et al., (2013), Adv Drug Deliv Rev. 65(10): 1357-1369, the entire contents of which are hereby incorporated by reference. In some embodiments, the linker may be designed using linker designing databases and computer programs such as those described in Chen et al., (2013), Adv Drug Deliv Rev. 65(10):1357-1369 and Crasto et. al., (2000), Protein Eng. 13(5):309-312, the entire contents of which are hereby incorporated by reference. In some embodiments, the linker is as provided in Moore et al., (2019), Methods 154:38-50, the entire contents of which are hereby incorporated by reference.
[00114] In some embodiments, the linker is a polypeptide. In some embodiments, the linker is less than about 100 amino acids long. For example, the linker may be less than about 100, about 95, about 90, about 85, about 80, about 75, about 70, about 65, about 60, about 55, about 50, about 45, about 40, about 35, about 30, about 25, about 20, about 19, about 18, about 17, about 16, about 15, about 14, about 13, about 12, about 11, about 10, about 9, about 8, about 7, about 6, about 5, about 4, about 3, or about 2 amino acids long. In some embodiments, the linker is flexible. In another embodiment, the linker is rigid. In various embodiments, the linker is substantially comprised of glycine and serine residues (e.g. about 30%, or about 40%, or about 50%, or about 60%, or about 70%, or about 80%, or about 90%, or about 95%, or about 97% glycines and serines).
[00115] Additional illustrative linkers include, but are not limited to, linkers having the sequence LE, GGGGS (SEQ ID NO: 51), (GGGGS)n(n=1-4) (SEQ ID NO: 5264),(Gly)8 (SEQ ID NO: 5265), (Gly)6(SEQ ID NO: 5266), (EAAAK)n (n=1-3) (SEQ ID NO: 5267), A(EAAAK)nA (n = 2-5) (SEQ ID NO: 5268), AEAAAKEAAAKA (SEQ ID NO: 5269), A(EAAAK)4ALEA(EAAAK)4A (SEQ ID NO: 5270), PAPAP (SEQ ID NO: 5271), KESGSVSSEQLAQFRSLD (SEQ ID NO: 5272), EGKSSGSGSESKST (SEQ ID NO:
5273), GSAGSAAGSGEF (SEQ ID NO: 5274), and (XP)n, with X designating any amino acid, e.g., Ala, Lys, or Glu.
[00116] In various embodiments, the linker may be functional. For example, without limitation, the linker may function to improve the folding and/or stability, improve the expression, improve the pharmacokinetics, and/or improve the bioactivity of the present compositions. In another example, the linker may function to target the compositions to a particular cell type or location.
[00117] As used herein, the term constant domain refers to the Fc domain. The constant domains exemplified above are optional embodiments and other constant domains can be substituted for the constant domains described herein.
[00118] In some embodiments, the Fc is from IgG. In some embodiments, the IgG is human IgG. In some embodiments, the human IgG is selected from IgGl, IgG2, IgG3, and IgG4. [00119] In some embodiments, the antibody is formed through a knob-in-hole interaction in the Fc region. In some embodiments, the human IgG Fc comprises one or mutations to promote knob-in-hole interaction. In some embodiments, the mutations are selected from (i) T366Y or T366W, and (ii) Y407T, Y407A, or Y407V. In some embodiments, the mutations are:
(a) T366Y and Y407T or T366Y/F405A and T394W/Y407T in human IgGl or
(b) T366W/D399C and T366S/L368A/K392C/Y407V;
T366W/K392C and T366S/L368A/D399C/Y407V;
S354C/T366W and Y349C/T366S/L368A/Y407V;
Y349C/T366W and S354C/T366S/L368A/Y407V;
E356C/T366W and Y349C/T366S/L368A/Y407V;
Y349C/T366W and E356C/T366S/L368A/Y407V;
E357C/T366W and Y349C/T366S/L368A/Y407V; or Y349C/T366W and E357C/T366S/L368A/Y407V, in human IgGl.
[00120] In some embodiments, the human IgG Fc comprises one or mutations to reduce or eliminate the effector function of the Fc domains. In some embodiments, the mutations are L234A and L235A (LALA) substitutions in human IgGl. In some embodiments, the human IgG Fc comprises one or mutations to stabilize a hinge region in the Fc domain. In some embodiments, the mutation is S228P.
[00121] In some embodiments, the composition binds Claudin 18.2 and CD3 contemporaneously.
[00122] In some embodiments, the composition binds Claudin 18.2 with an affinity of less than 10 nM and with at least 100 fold greater affinity than Claudin 9, Claudin 3, and/or Claudin 4.
[00123] In any of the embodiments disclosed herein, the composition induces cellular cytotoxicity.
[00124] In any of the embodiments disclosed herein, the composition induces T cell cytotoxicity.
[00125] In any of the embodiments disclosed herein, the composition induces T cell dependent cytotoxicity.
[00126] In any of the embodiments disclosed herein, the composition increases the expression and/or the release of one or more cytokines. In any of the embodiments disclosed herein, the composition increases the expression and/or the release of one or more cytokines selected from IL-2, IL-6, IL-10, IFN-γ, and TNF-α.
[00127] In some embodiments, disclosed herein is a pharmaceutical composition comprising an isolated antibody of any one of the preceding embodiments, or a nucleic acid molecule encoding the same. In some embodiments, the composition is an injectable pharmaceutical composition. In some embodiments, the composition is sterile. In some embodiments, the composition is pyrogen free.
[00128] In addition, this document also provides pharmaceutical compositions that composition as described herein, in combination with a pharmaceutically acceptable carrier. A “pharmaceutically acceptable carrier” (also referred to as an “excipient” or a “carrier”) is a pharmaceutically acceptable solvent, suspending agent, stabilizing agent, or any other pharmacologically inert vehicle for delivering one or more therapeutic compounds to a subject (e.g., a mammal, such as a human, non-human primate, dog, cat, sheep, pig, horse, cow, mouse, rat, or rabbit), which is nontoxic to the cell or subject being exposed thereto at the dosages and concentrations employed. Pharmaceutically acceptable carriers can be liquid or solid, and can be selected with the planned manner of administration in mind so as to provide for the desired bulk, consistency, and other pertinent transport and chemical properties, when combined with one or more of therapeutic compounds and any other components of a given pharmaceutical composition. Typical pharmaceutically acceptable carriers that do not deleteriously react with amino acids include, by way of example and not limitation: water, saline solution, binding agents (e.g., polyvinylpyrrolidone or hydroxypropyl methylcellulose), fillers (e.g., lactose and other sugars, gelatin, or calcium sulfate), lubricants (e.g., starch, polyethylene glycol, or sodium acetate), disintegrates (e.g., starch or sodium
starch glycolate), and wetting agents (e.g., sodium lauryl sulfate). Pharmaceutically acceptable carriers also include aqueous pH buffered solutions or liposomes (small vesicles composed of various types of lipids, phospholipids and/or surfactants which are useful for delivery of a drug to a mammal). Further examples of pharmaceutically acceptable carriers include buffers such as phosphate, citrate, and other organic acids, antioxidants such as ascorbic acid, low molecular weight (less than about 10 residues) polypeptides, proteins such as serum albumin, gelatin, or immunoglobulins, hydrophilic polymers such as polyvinylpyrrolidone, amino acids such as glycine, glutamine, asparagine, arginine or lysine, monosaccharides, disaccharides, and other carbohydrates including glucose, mannose or dextrins, chelating agents such as EDTA, sugar alcohols such as mannitol or sorbitol, saltforming counterions such as sodium, and/or nonionic surfactants such as TWEEN™, polyethylene glycol (PEG), and PLURONICS™.
[00129] Pharmaceutical compositions can be formulated by mixing one or more active agents with one or more physiologically acceptable carriers, diluents, and/or adjuvants, and optionally other agents that are usually incorporated into formulations to provide improved transfer, delivery, tolerance, and the like. A pharmaceutical composition can be formulated, e.g., in lyophilized formulations, aqueous solutions, dispersions, or solid preparations, such as tablets, dragees or capsules. A multitude of appropriate formulations can be found in the formulary known to all pharmaceutical chemists: Remington’s Pharmaceutical Sciences (18th ed, Mack Publishing Company, Easton, PA (1990)), particularly Chapter 87 by Block, Lawrence, therein. These formulations include, for example, powders, pastes, ointments, jellies, waxes, oils, lipids, lipid (cationic or anionic) containing vesicles (such as LIPOFECTIN™), DNA conjugates, anhydrous absorption pastes, oil-in-water and water-in- oil emulsions, emulsions carbowax (polyethylene glycols of various molecular weights), semi-solid gels, and semi-solid mixtures containing carbowax. Any of the foregoing mixtures may be appropriate in treatments and therapies as described herein, provided that the active agent in the formulation is not inactivated by the formulation and the formulation is physiologically compatible and tolerable with the route of administration. See, also, Baldrick, Regul Toxicol Pharmacol 32:210-218, 2000; Wang, IntJ Pharm 203:1-60, 2000; Charman J Pharm Sci 89:967-978, 2000; and Powell et al. PDA J Pharm Sci Technol 52:238-311, 1998), and the citations therein for additional information related to formulations, excipients and carriers well known to pharmaceutical chemists.
[00130] Pharmaceutical compositions include, without limitation, solutions, emulsions, aqueous suspensions, and liposome-containing formulations. These compositions can be
generated from a variety of components that include, for example, preformed liquids, selfemulsifying solids and self-emulsifying semisolids. Emulsions are often biphasic systems comprising of two immiscible liquid phases intimately mixed and dispersed with each other; in general, emulsions are either of the water-in-oil (w/o) or oil-in-water (o/w) variety. Emulsion formulations have been widely used for oral delivery of therapeutics due to their ease of formulation and efficacy of solubilization, absorption, and bioavailability.
[00131] Compositions and formulations can contain sterile aqueous solutions, which also can contain buffers, diluents and other suitable additives (e.g., penetration enhancers, carrier compounds and other pharmaceutically acceptable carriers). Compositions additionally can contain other adjunct components conventionally found in pharmaceutical compositions. Thus, the compositions also can include compatible, pharmaceutically active materials such as, for example, antipruritics, astringents, local anesthetics or anti-inflammatory agents, or additional materials useful in physically formulating various dosage forms of the compositions provided herein, such as dyes, flavoring agents, preservatives, antioxidants, opacifiers, thickening agents and stabilizers. Furthermore, the composition can be mixed with auxiliary agents, e.g., lubricants, preservatives, stabilizers, wetting agents, emulsifiers, salts for influencing osmotic pressure, buffers, colorings, flavorings, and aromatic substances.
When added, however, such materials should not unduly interfere with the biological activities of the polypeptide components within the compositions provided herein. The formulations can be sterilized if desired.
[00132] In some embodiments, a composition containing a composition as provided herein can be in the form of a solution or powder with or without a diluent to make an injectable suspension. The composition may contain additional ingredients including, without limitation, pharmaceutically acceptable vehicles, such as saline, water, lactic acid, mannitol, or combinations thereof, for example.
[00133] Any appropriate method can be used to administer a composition as described herein to a mammal. Administration can be, for example, parenteral (e.g., by subcutaneous, intrathecal, intraventricular, intramuscular, or intraperitoneal injection, or by intravenous drip). Administration can be rapid (e.g., by injection) or can occur over a period of time (e.g., by slow infusion or administration of slow release formulations). In some embodiments, administration can be topical (e.g., transdermal, sublingual, ophthalmic, or intranasal), pulmonary (e.g. , by inhalation or insufflation of powders or aerosols), or oral. In addition, a composition containing a composition as described herein can be administered prior to, after, or in lieu of surgical resection of a tumor.
[00134] The antibodies, compositions, or pharmaceutical compositions provided for herein may be administered at any appropriate interval to achieve the desired effect in a subject. In some embodiments, the antibodies, compositions, or pharmaceutical compositions, are administered daily, every other day, weekly, biweekly, once every three weeks, or monthly (i.e., once every four weeks). In some embodiments, a method as provided for herein comprises administering an antibody, composition, or pharmaceutical composition to a cell or to a subject in need thereof at an interval of daily, every other day, weekly, biweekly, once every three weeks, or monthly (i.e., once every four weeks).
[00135] In some embodiments, disclosed herein is a nucleic acid molecule encoding an antibody or an amino acid sequence of any of the preceding embodiments.
[00136] In some embodiments, disclosed herein is a vector comprising the nucleic acid molecule of any of the preceding embodiments.
[00137] In some embodiments, disclosed herein is a cell comprising the nucleic acid molecule of any of the preceding embodiments, or the vector of any of the preceding embodiments.
[00138] In some embodiments, disclosed herein is a method for modulating and/or targeting Claudin 18.2 and CD3 in a biological cell, comprising contacting the cell with a composition of any of the preceding embodiments.
[00139] In some embodiments, disclosed herein is a method for modulating Claudin 18.2 activity in a biological cell comprising contacting a cell expressing Claudin 18.2 with a composition of any of the preceding embodiments.
[00140] In some embodiments, disclosed herein is a method for inhibiting the function of Claudin 18.2 in a biological cell comprising contacting a cell expressing Claudin 18.2 with a composition of any of the preceding embodiments.
[00141] In some embodiments, disclosed herein is a method for treating or preventing cancer comprising administering an effective amount of the composition of any of the preceding embodiments to a subject in need thereof.
[00142] In some embodiments, disclosed herein is a use of the composition of any of the preceding embodiments for the preparation of a medicament for the treatment of prevention of cancer.
[00143] In some embodiments, disclosed herein is a method or use of any one of the preceding embodiments, wherein the cancer is selected from one or more of basal cell carcinoma, biliary tract cancer; bladder cancer; bone cancer; brain and central nervous system cancer; breast cancer; cancer of the peritoneum; cervical cancer; choriocarcinoma; colon and
rectum cancer; connective tissue cancer; cancer of the digestive system; endometrial cancer; esophageal cancer; eye cancer; cancer of the head and neck; gastric cancer (including gastrointestinal cancer); glioblastoma; glioma; hepatic carcinoma; hepatoma; intra-epithelial neoplasm; kidney or renal cancer; larynx cancer; leukemia; liver cancer; lung cancer (e.g., small-cell lung cancer, non-small cell lung cancer, adenocarcinoma of the lung, and squamous carcinoma of the lung); melanoma; myeloma; neuroblastoma; oral cavity cancer (lip, tongue, mouth, and pharynx); ovarian cancer; pancreatic cancer; prostate cancer; retinoblastoma; rhabdomyosarcoma; malignant rhabdoid tumor; rectal cancer; cancer of the respiratory system; salivary gland carcinoma; sarcoma; skin cancer; squamous cell cancer; stomach cancer; testicular cancer; thyroid cancer; uterine or endometrial cancer; cancer of the urinary system; vulvar cancer; lymphoma including Hodgkin's and non-Hodgkin's lymphoma, as well as B-cell lymphoma (including low grade/follicular non-Hodgkin's lymphoma (NHL); small lymphocytic (SL) NHL; intermediate grade/follicular NHL; intermediate grade diffuse NHL; high grade immunoblastic NHL; high grade lymphoblastic NHL; high grade small non-cleaved cell NHL; bulky disease NHL; mantle cell lymphoma; AIDS-related lymphoma; and Waldenstrom's Macroglobulinemia; chronic lymphocytic leukemia (CLL); acute lymphoblastic leukemia (ALL); Hairy cell leukemia; chronic myeloblastic leukemia; as well as other carcinomas and sarcomas; and post-transplant lymphoproliferative disorder (PTLD), as well as abnormal vascular proliferation associated with phakomatoses; edema (e.g. that associated with brain tumors); and Meigs’ syndrome. [00144] As a non-limiting example, the prevention of an onset, the presence, and/or the evaluation of the progression of a cancer in a subject can be assessed according to the Tumor/Nodes/Metastases (TNM) system of classification (International Union Against Cancer, 6th edition, 2002), or the Whitmore- Jewett staging system (American Urological Association). Typically, cancers are staged using a combination of physical examination, blood tests, and medical imaging. If tumor tissue is obtained via biopsy or surgery, examination of the tissue under a microscope can also provide pathologic staging. In some embodiments, the stage or grade of a cancer assists a practitioner in determining the prognosis for the cancer and in selecting the appropriate modulating therapy.
[00145] In some embodiments, the prevention of an onset, or progression, of cancer is assessed using the overall stage grouping as a non-limiting example: Stage I cancers are localized to one part of the body, typically in a small area; Stage II cancers are locally advanced and have grown into nearby tissues or lymph nodes, as are Stage III cancers. Whether a cancer is designated as Stage II or Stage III can depend on the specific type of
cancer. The specific criteria for Stages II and III can differ according to diagnosis. Stage IV cancers have often metastasized or spread to other organs or throughout the body. The onset or progression of cancer can be assessed using conventional methods available to one of skill in the art, such as a physical exam, blood tests, and imaging scans (e.g., X-rays, MRI, CT scans, ultrasound etc.).
[00146] As disclosed herein, administering, or administering a treatment/therapy, refers to a treatment/therapy from which a subject receives a beneficial effect, such as the reduction, decrease, attenuation, diminishment, stabilization, remission, suppression, inhibition or arrest of the development or progression of cancer, or a symptom thereof.
[00147] In some embodiments, the treatment/therapy that a subject receives, or the prevention in the onset of cancer results in at least one or more of the following effects: (1) the reduction or amelioration of the severity of cancer and/or a genetic disease or disorder, and/or a symptom associated therewith; (2) the reduction in the duration of a symptom associated with cancer and/or a genetic disease or disorder; (3) the prevention in the recurrence of a symptom associated with cancer and/or a genetic disease or disorder; (4) the regression of cancer and/or a genetic disease or disorder, and/or a symptom associated therewith; (5) the reduction in hospitalization of a subject; (6) the reduction in hospitalization length; (7) the increase in the survival of a subject; (8) the inhibition of the progression of cancer and/or a genetic disease or disorder and/or a symptom associated therewith; (9) the enhancement or improvement the therapeutic effect of another therapy; (10) a reduction or elimination in the cancer cell population, and/or a cell population associated with a genetic disease or disorder; (11) a reduction in the growth of a tumor or neoplasm; (12) a decrease in tumor size; (13) a reduction in the formation of a tumor; (14) eradication, removal, or control of primary, regional and/or metastatic cancer; (15) a decrease in the number or size of metastases; (16) a reduction in mortality; (17) an increase in cancer-free survival rate of a subject; (18) an increase in relapse-free survival; (19) an increase in the number of subjects in remission; (20) a decrease in hospitalization rate; (21) the size of the tumor is maintained and does not increase in size or increases the size of the tumor by less 5% or 10% after administration of a therapy as measured by conventional methods available to one of skill in the art, e.g., X-rays, MRI, CAT scan, ultrasound etc.; (22) the prevention of the development or onset of cancer and/or a genetic disease or disorder, and/or a symptom associated therewith; (23) an increase in the length of remission for a subject; (24) the reduction in the number of symptoms associated with cancer and/or a genetic disease or disorder; (25) an increase in symptom-free survival of a cancer subject and/or a subject associated with a
genetic disease or disorder; and/or (26) limitation of or reduction in metastasis. In some embodiments, the treatment/therapy that a subject receives does not cure cancer, but prevents the progression or worsening of the disease. In certain embodiments, the treatment/therapy that a subject receives does not prevent the onset/development of cancer, but may prevent the onset of cancer symptoms.
[00148] In some embodiments, the subject that is treated does not develop cytokine release syndrome (CRS), or does not develop significant CRS -associated clinical symptoms or toxicity. These symptoms include, but are not limited to fever, chills, fatigue, weakness, loss of appetite, nausea, vomiting, diarrhea, headache, joint or muscle aches, skin rash, low blood pressure, increased heart rate, irregular heartbeat, tachycardia, decreased heart function, swelling, buildup of fluids (edema), confusion, dizziness, seizures, hallucinations, decreased coordination, problems talking or swallowing, shaking, problems controlling movements, cough, shortness of breath, tachypnoea, decreased lung function, reduced oxygen levels, decreased kidney or liver function, increased cytokine levels in the blood, change in electrolytes, change in blood clotting. In some embodiments, there is no increase or no significant increase in the blood levels of interleukin-6 (IL-6), interleukin- 10 (IL-10), interferon (IFN)-γ, monocyte chemoattractant protein 1 (MCP-1) granulocyte-macrophage colony-stimulating factor (GM-CSF), tumor necrosis factor (TNF), IL-1, IL-2, IL-2-receptor- α, or IL-8 after administration of an antibody as provided for herein.
[00149] In some embodiments, “preventing” an onset or progression of cancer in a subject in need thereof, is inhibiting or blocking the cancer or disorder. In some embodiments, the methods disclosed herein prevent, or inhibit, the cancer or disorder at any amount or level. In some embodiments, the methods disclosed herein prevent or inhibit the cancer or genetic disease or disorder by at least or about a 10% inhibition (e.g., at least or about a 20% inhibition, at least or about a 30% inhibition, at least or about a 40% inhibition, at least or about a 50% inhibition, at least or about a 60% inhibition, at least or about a 70% inhibition, at least or about a 80% inhibition, at least or about a 90% inhibition, at least or about a 95% inhibition, at least or about a 98% inhibition, or at least or about a 100% inhibition).
[00150] In some embodiments, disclosed herein is an isolated antibody comprising one or more of the sequences disclosed herein.
Delivery and. Vectors
[00151] Disclosed herein, in some embodiments, are nucleic acid sequences encoding an antibody described herein or components thereof.
[00152] In some embodiments, the nucleic acid encoding the antibody described herein or components thereof is a DNA, for example a linear DNA, a plasmid DNA, or a minicircle DNA. In some embodiments, the nucleic acid encoding the antibody described herein or components thereof is an RNA, for example a mRNA.
[00153] In some embodiments, the nucleic acid encoding the antibody described herein or components thereof is delivered by a nucleic acid-based vector. In some embodiments, the nucleic acid-based vector is a plasmid (e.g., circular DNA molecules that can autonomously replicate inside a cell), cosmid (e.g., pWE or sCos vectors), artificial chromosome, human artificial chromosome (HAC), yeast artificial chromosomes (YAC), bacterial artificial chromosome (BAC), Pl-derived artificial chromosomes (PAC), phagemid, phage derivative, bacmid, or virus. In some embodiments, the nucleic acid-based vector is selected from the list consisting of: pSF-CMV-NEO-NH2-PPT-3XFLAG, pSF-CMV-NEO-COOH-3XFLAG, pSF-CMV-PURO-NH2-GST-TEV, pSF-OXB20-COOH-TEV-FLAG(R)-6His, pCEP4 pDEST27, pSF-CMV-Ub-KrYFP, pSF-CMV-FMDV-daGFP, pEFla-mCherry-Nl vector, pEFla-tdTomato vector, pSF-CMV-FMDV-Hygro, pSF-CMV-PGK-Puro, pMCP-tag(m), pSF-CMV-PURO-NH2-CMYC, pSF-OXB20-BetaGal,pSF-OXB20-Fluc, pSF-OXB20, pSF- Tac, pRI 101-AN DNA, pCambia2301, pTYB21, pKLAC2, pAc5.1/V5-His A, and pDEST8. [00154] In some embodiments, the nucleic acid-based vector comprises a promoter. In some embodiments, the promoter is selected from the group consisting of a mini promoter, an inducible promoter, a constitutive promoter, and derivatives thereof. In some embodiments, the promoter is selected from the group consisting of CMV, CBA, EFla, CAG, PGK, TRE, U6, UAS, T7, Sp6, lac, araBad, trp, Ptac, p5, pl9, p40, Synapsin, CaMKII, GRK1, and derivatives thereof. In some embodiments the promoter is a U6 promoter. In some embodiments, the promoter is a CAG promoter.
[00155] In some embodiments, the nucleic acid-based vector is a virus. In some embodiments, the virus is an alphavirus, a parvovirus, an adenovirus, an AAV, a baculovirus, a Dengue virus, a lentivirus, a herpesvirus, a poxvirus, an anellovirus, a bocavirus, a vaccinia virus, or a retrovirus. In some embodiments, the virus is an alphavirus. In some embodiments, the virus is a parvovirus. In some embodiments, the virus is an adenovirus. In some embodiments, the virus is an AAV. In some embodiments, the virus is a baculovirus. In some embodiments, the virus is a Dengue virus. In some embodiments, the virus is a lentivirus. In some embodiments, the virus is a herpesvirus. In some embodiments, the virus is a poxvirus. In some embodiments, the virus is an anellovirus. In some embodiments, the virus is a
bocavirus. In some embodiments, the virus is a vaccinia virus. In some embodiments, the virus is or a retrovirus.
[00156] In some embodiments, the AAV is AAV1, AAV2, AAV3, AAV4, AAV5, AAV6, AAV7, AAV8, AAV9, AAV10, AAV11, AAV12, AAV13, AAV14, AAV15, AAV16, AAV-rh8, AAV-rhlO, AAV-rh20, AAV-rh39, AAV-rh74, AAV-rhM4-l, AAV-hu37, AAV-
Anc80, AAV-Anc80L65, AAV-7m8, AAV-PHP-B, AAV-PHP-EB, AAV-2.5, AAV-21YF, AAV-3B, AAV-LK03, AAV-HSC1, AAV-HSC2, AAV-HSC3, AAV-HSC4, AAV-HSC5, AAV-HSC6, AAV-HSC7, AAV-HSC8, AAV-HSC9, AAV-HSC10, AAV-HSC11 , AAV-
HSC12, AAV-HSC13, AAV-HSC14, AAV-HSC15, AAV-TT, AAV-DJ/8, AAV-Myo, AAV-NP40, AAV-NP59, AAV-NP22, AAV-NP66, AAV-HSC16, or a derivative thereof. In some embodiments, the herpesvirus is HSV type 1, HSV-2, VZV, EBV, CMV, HHV-6, HHV-7, or HHV-8.
[00157] In some embodiments, the nucleic acid encoding the antibody described herein or components thereof is delivered by a non-nucleic acid-based delivery system (e.g., a non- viral delivery system). In some embodiments, the non-viral delivery system is a liposome. In some embodiments, the nucleic acid is associated with a lipid. The nucleic acid associated with a lipid, in some embodiments, is encapsulated in the aqueous interior of a liposome, interspersed within the lipid bilayer of a liposome, attached to a liposome via a linking molecule that is associated with both the liposome and the nucleic acid, entrapped in a liposome, complexed with a liposome, dispersed in a solution containing a lipid, mixed with a lipid, combined with a lipid, contained as a suspension in a lipid, contained or complexed with a micelle, or otherwise associated with a lipid. In some embodiments, the nucleic acid is comprised in a lipid nanoparticle (LNP).
[00158] In some embodiments, the antibody described herein or components thereof is introduced into the cell in any suitable way, either stably or transiently. In some embodiments, the antibody described herein or components thereof is transfected into the cell. In some embodiments, the cell is transduced or transfected with a nucleic acid construct that encodes the antibody described herein or components thereof. For example, a cell is transduced (e.g., with a virus encoding the antibody described herein or components thereof), or transfected (e.g., with a plasmid encoding the antibody described herein or components thereof) with a nucleic acid that encodes the antibody described herein or components thereof, or the translated the antibody described herein or components thereof. In some embodiments, the transduction is a stable or transient transduction. In some embodiments, a plasmid expressing the antibody described herein or components thereof is introduced into
cells through electroporation, transient (e.g., lipofection) and stable genome integration (e.g., piggybac) and viral transduction (for example lentivirus or AAV) or other methods known to those of skill in the art. In some embodiments, the gene editing system is introduced into the cell as one or more polypeptides. In some embodiments, delivery is achieved through the use of RNP complexes. Delivery methods to cells for polypeptides and/or RNPs are known in the art, for example by electroporation or by cell squeezing.
[00159] Exemplary methods of delivery of nucleic acids include lipofection, nucleofection, electroporation, stable genome integration (e.g., piggybac), microinjection, biolistics, virosomes, liposomes, immunoliposomes, polycation or lipid nucleic acid conjugates, naked DNA, artificial virions, and agent-enhanced uptake of DNA. Lipofection is described in e.g., U.S. Pat. Nos. 5,049,386; 4,946,787; and 4,897,355) and lipofection reagents are sold commercially (e.g., Transfectam™, Lipofectin™ and SF Cell Line 4D-Nucleofector X Kit™ (Lonza)). Cationic and neutral lipids that are suitable for efficient receptor-recognition lipofection of polynucleotides include those of WO 91/17424 and WO 91/16024. In some embodiments, the delivery is to cells (e.g., in vitro or ex vivo administration) or target tissues (e.g., in vivo administration). In some embodiments, the nucleic acid is comprised in a liposome or a nanoparticle that specifically targets a host cell.
[00160] Additional methods for the delivery of nucleic acids to cells are known to those skilled in the art. See, for example, US 2003/0087817.
[00161] In some embodiments, a nucleic acid molecule encoding an antibody or a component thereof described herein includes one or more of the DNA sequences provided in TABLE B. In some embodiments, the nucleic acid molecule comprises a sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity to any one of SEQ ID NOs: 5210-5236.
[00162] In some embodiments, a nucleic acid molecule encoding an antibody or a component thereof described herein includes one or more of the RNA sequences provided in TABLE C. In some embodiments, the nucleic acid molecule comprises a sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity to any one of SEQ ID NOs: 5237-5263.
TABLE B: DNA Sequences Encoding Antibody or a Component Thereof Described
TABLE C: RNA Sequences Encoding Antibody or a Component Thereof Described
Herein
Lipid nanoparticles
[00163] Disclosed herein, in certain embodiments, are lipid nanoparticles comprising an antibody or components thereof of the disclosure for delivery of the antibody into a cell.
[00164] In some embodiments, the lipid nanoparticle comprises the antibody or a nucleic acid encoding the antibody or components thereof.
[00165] In some embodiments, the lipid nanoparticle is tethered to the antibody or a nucleic acid encoding the antibody or components thereof.
[00166] Lipid nanoparticles as described herein can be 4-component lipid nanoparticles. Such nanoparticles can be configured for delivery of RNA or other nucleic acids (e.g., synthetic RNA, mRNA, or in vitro-synthesized mRNA) and can be generally formulated as described in WO2012135805A2. Such nanoparticles can generally comprise: (a) a cationic lipid, (b) a neutral lipid (e.g., DSPC or DOPE), (c) a sterol (e.g., cholesterol or a cholesterol analog), or (d) a PEG-modified lipid (e.g., PEG-DMG).
[00167] The cationic lipid referred to herein as “C 12-200” is disclosed by Love et al., Proc Natl Acad Sci USA. 2010 107:1864-1869 and Liu and Huang, Molecular Therapy. 2010 669- 670. Cationic lipid formulations can include particles comprising either 3 or 4 or more components in addition to polynucleotide, primary construct, or RNA (e.g., mRNA). As an example, formulations with certain cationic lipids, include, but are not limited to, 98N12-5 and may contain 42% lipidoid, 48% cholesterol and 10% PEG (C14 or greater alkyl chain length). As another example, formulations with certain lipidoids include, but are not limited to, C12-200 and may contain 50% cationic lipid, 10% disteroylphosphatidyl choline, 38.5% cholesterol, and 1.5% PEG-DMG.
[00168] In some embodiments, the cationic lipid nanoparticle comprises a cationic lipid, a PEG-modified lipid, a sterol, and a non-cationic lipid. In some embodiments, the cationic lipid nanoparticle has a molar ratio of about 20-60% cationic lipid: about 5-25% non-cationic lipid: about 25-55% sterol; and about 0.5-15% PEG-modified lipid. In some embodiments, the cationic lipid nanoparticle comprises a molar ratio of about 50% cationic lipid, about 1.5% PEG-modified lipid, about 38.5% cholesterol, and about 10% non-cationic lipid. In some embodiments, the cationic lipid nanoparticle comprises a molar ratio of about 55% cationic lipid, about 2.5% PEG-modified lipid, about 32.5% cholesterol, and about 10% noncationic lipid. In some embodiments, the cationic lipid is an ionizable cationic lipid, the noncationic lipid is a neutral lipid, and the sterol is a cholesterol. In some embodiments, the cationic lipid nanoparticle has a molar ratio of 50:38.5: 10: 1 .5 of cationic lipid: cholesterol: PEG2000-DMG:DSPC or DMG:DOPE. In some embodiments, lipid nanoparticles as described herein can comprise cholesterol, 1,2-dioleoyl-sn-glycero-3 -phosphoethanolamine (DOPE), 1 , 1 ‘-((2-(4-(2-((2-(bis(2-hydroxydodecyl)amino)ethyl)(2- hydroxydodecyl)amino)ethyl)piperazin- 1 -yl)ethyl)azanediyl)bis(dodecan-2-ol) (C 12-200), and DMG-PEG-2000 at molar ratios of 47.5:16:35:1.5.
[00169] In some embodiments, the lipid nanoparticle comprises 1,1'-[[2-[4-[2-[[2-[bis(2- hydroxydodecyl)amino]ethyl](2-hydroxydodecyl)amino]ethyl]-l- piperazinyl]ethyl]imino]bis-2-dodecanol (C12-200), a sterol (e.g., cholesterol, campesterol,
or β-sitosterol), 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE), and 1,2- dimyristoyl-sn-glycero-3-phosphoethanolamine-N-[methoxy(polyethylene glycol)-2000] (ammonium salt) (C14:0 PEG2000 PE). In some embodiments, the lipid nanoparticle comprises 1,1'-[[2-[4-[2-[[2-[bis(2-hydroxydodecyl)amino]ethyl](2- hydroxydodecyl)amino]ethyl]-l-piperazinyl]ethyl]imino]bis-2-dodecanol (C12-200), sterol (e.g., cholesterol, campesterol, or β-sitosterol), 1,2-dioleoyl-sn-glycero-3- phosphoethanolamine (DOPE), 1 ,2-dimyristoyl-sn-glycero-3-phosphoethanolamine-N- [methoxy(polyethylene glycol)-2000] (ammonium salt) (C14:0 PEG2000 PE), and 1 ,2- distearoyl-sn-glycero-3-phosphoethanolamine-N-[tris-GalNAc-Y-aminobutyric acid- (polyethylene glycol)-2000.
[00170] In some embodiments, the lipid nanoparticle comprises (9Z,12Z)-3-((4,4- Bis(octyloxy)butanoyl)oxy)-2-((((3-(diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9, 12-dienoate (LP01), a sterol (e.g., cholesterol, campesterol, or β-sitosterol), 1,2- dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE), 1 ,2-dioctadecanoyl-sn-glycero-3- phosphocholine (DS PC), and 1,2-dimyristoyl-rac-glycero-3-methoxypolyethylene glycol- 2000 (DMG-PEG 2000). In some embodiments, the lipid nanoparticle comprises (9Z,12Z)-3- ((4,4-Bis(octyloxy)butanoyl)oxy)-2-((((3- (diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9, 12-dienoate (LP01), a sterol (e.g., cholesterol, campesterol, or β-sitosterol), 1,2-dioleoyl-sn-glycero-3- phosphoethanolamine (DOPE), 1,2-dioctadecanoyl-sn-glycero-3 -phosphocholine (DSPC), 1,2-dimyristoyl-rac-glycero-3-methoxypolyethylene glycol-2000 (DMG-PEG 2000), and 1,2- distearoyl-sn-glycero-3-phosphoethanolamine-N-[tris-GalNAc-Y-aminobutyric acid- (polyethylene glycol)-2000. In some embodiments, the lipid nanoparticle comprises (9Z,12Z)-3-((4,4-Bis(octyloxy)butanoyl)oxy)-2-((((3- (diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9, 12-dienoate (LP01 ) , cholesterol, 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE), 1,2-dioctadecanoyl-sn- glycero-3-phosphocholine (DSPC), and 1,2-dimyristoyl-rac-glycero-3-methoxypolyethylene glycol-2000 (DMG-PEG 2000). In some embodiments, the lipid nanoparticle comprises (9Z, 12Z)-3-((4,4-Bis(octyloxy)butanoyl)oxy)-2-((((3- (diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9, 12-dienoate (LP01 ) , cholesterol, 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (DOPE), 1,2-dioctadecanoyl-sn- glycero-3-phosphocholine (DSPC), 1 ,2-dimyristoyl-rac-glycero-3-methoxypolyethylene glycol-2000 (DMG-PEG 2000), and 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N- [tris-GalNAc-Y-aminobutyric acid-(polyethylene glycol)-2000.
[00171] In some embodiments, the lipid nanoparticles comprises (9Z,12Z)-3-((4,4- Bis(octyloxy)butanoyl)oxy)-2-((((3-(diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9,12-dienoate (LP-01), cholesterol, 1,2-dioctadecanoyl-sn-glycero-3- phosphocholine (DSPC) and 1 ,2-dimyristoyl-rac-glycero-3 -methoxypolyethylene glycol- 2000 (DMG-PEG 2000).
[00172] In some embodiments, the lipid nanoparticles comprises one of the phospholipids selected from 1,2-dioleoyl-sn-glycero-3 -phosphoethanolamine (DOPE), 1,2-dilinolenoyl-sn- glycero-3-phosphoethanolamine (18:3 PE), 1,2-didocosahexaenoyl-sn-glycero-3- phosphoethanolamine (22:6 PE), and 1,2-dioleoyl-sn-glycero-3 -phosphocholine (DOPC). [00173] In some embodiments, the lipid nanoparticle comprises of different lipids comprising (9Z,12Z)-3-((4,4-Bis(octyloxy)butanoyl)oxy)-2-((((3- (diethylamino)propoxy)carbonyl)oxy)methyl)propyl octadeca-9, 12-dienoate (LP-01 ), cholesterol, 1,2-dioctadecanoyl-sn-glycero-3-phosphocholine (DSPC), 1 ,2-dimyristoyl-rac- glycero-3-methoxypolyethylene glycol-2000 (DMG-PEG 2000), 1 ,2-dioleoyl-sn-glycero-3- phosphoethanolamine (DOPE) or 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC), and 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-|tris-GalNAc-Y-aminobutyric acid- (polyethylene glycol)-2000, Tris-β-GalNAc-GABA-PEG2000-DSPE (GalNAc).
[00174] As used herein, the word “include,” and its variants, is intended to be non-limiting, such that recitation of items in a list is not to the exclusion of other like items that may also be useful in the materials, compositions, devices, and methods of this technology. Similarly, the terms “can” and “may” and their variants are intended to be non-limiting, such that recitation that an embodiment can or may comprise certain elements or features does not exclude other embodiments of the present technology that do not contain those elements or features. Although the open-ended term “comprising,” as a synonym of terms such as including, containing, or having, is used herein to describe and claim the disclosure, the present technology, or embodiments thereof, may alternatively be described using more limiting terms such as “consisting of” or “consisting essentially of” the recited ingredients. [00175] Unless defined otherwise, all technical and scientific terms herein have the same meaning as commonly understood by one of ordinary skill in the art to which this disclosure belongs. Although any methods and materials, similar or equivalent to those described herein, can be used in the practice or testing of the present disclosure, the preferred methods and materials are described herein. All publications, patents, and patent publications cited are incorporated by reference herein in their entirety for all purposes.
[00176] This disclosure is further illustrated by the following non-limiting examples.
EXAMPLES
[00177] The subject matter is now described with reference to the following examples. These examples are provided for the purpose of illustration only and the claims should in no way be construed as being limited to these examples, but rather should be construed to encompass any and all variations which become evident as a result of the teaching provided herein. Those of skill in the art will readily recognize a variety of non-crilical parameters that could be changed or modified to yield essentially similar results.
Example 1: Overview and Development of Antibodies Specific for Claudin 18.2 and CD3 [00178] The experiments in this example demonstrated certain features of antibodies specific for Claudin 18.2 and CD3. In this example, antibodies specific for Claudin 18.2 and CD3 were engineered and characterized. This example provided tandem scFv antibodies, scFv-Fab IgG antibodies, IgG-(scFv)2 antibodies, and IgG-(scFv) antibodies specific for Claudin 18.2 and CD3 (FIG. 1).
[00179] To develop bispecific antibodies to Claudin 18.2 and CD3, a panel of constructs were generated using the following antibody formats: tandem scFv, scFv-Fab IgG, IgG-scFv, IgG-(scFv)2, and IgG-(scFv). Potential Claudin 18.2-specific arms included those from 4409- B03, 4401 -E10, and 4405-H06 antibodies, and specific CD3 arms included those from hsOKT3 and pasotuxizumab antibodies, as well as a nanobody. Antibodies tested are provided in TABLEs 6-9, as follows:
Example 2: Functional and Developability Characterization of IM-16-44-09-B03-1CD3
[00180] FIG. 2 shows that IM-16-44-09-B03-1CD3 bound to both human Claudin 18.2
(“CLDN 18.2”) and human CD3. Flow cytometry experiments were performed to test the ability of the IM-16-44-09-B03-1CD3 antibody to bind HEK293F cells expressing human CLDN 18.2, human CD3, or human P2X3. HEK293F cells were transfected by lipofectamine with target and control plasmids. At 18-24 hours post-transfection, cells were stained with bispecific and control MAbs followed by APC -labelled goat anti-human antibody.
Fluorescence was detected using an Intellicyt high-throughput flow cytometer (iQue; Intellicyt, Albuquerque, NM). As shown in FIG. 2, IM-16-44-09-B03-1CD3 selectively bound to both human CLDN 18.2 and human CD3, and not P2X3.
[00181] FIG. 3 shows cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) were co-cultured with human Claudin 18.1 (“CLDN 18.1”) or CLDN 18.2-expressing HEK293F cells (2500 cells/well) also expressing Firefly Luciferase (ffLuciferase) in the presence of IMC-16-C2 (control antibody) or IM- 16-
44-09-B03-1CD3 antibodies. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated. As shown in FIG. 3, killing was only potently induced against CLDN 18.2- expressing gastric cancer cells by the IM-16-44-09-B03-1CD3 antibody. No killing activity was observed using the control antibody or against cells expressing CLDN 18.1.
[00182] FIGs. 4A, 4B, 4C, 4D, and 4E show cytokines produced by human peripheral blood mononuclear cells (PBMC) cultured with IMC-16-C2 or IM-16-44-09-B03-1CD3 antibodies and human CLDN 18.1 - or 18.2-expressing cells (HEK293F). FIG. 4A shows the levels of IL-2, FIG. 4B shows the levels of IL-6, FIG. 4C shows the levels of IL- 10, FIG. 4D shows the levels of IFN-γ (gamma), and FIG. 4E shows the levels of TNF-α (alpha) produced by human PBMCs in the presence of human CLDN 18.1- or 18.2-expressing cells and IMC-16- C2 or IM-16-44-09-B03-1CD3 antibodies. These experiments showed that IL-2, IL-6, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMC by the IM-16-44-09-B03-1CD3 antibody in the presence of CLDN 18.2-expressing cells.
[00183] FIGs. 5A and 5B show size exclusion chromatography (SEC) and differential scanning fhiorimetry (DSF) experimental results for the IM-16-44-09-B03-1CD3, IM-16-44- 09-B03-2CD3, and IM-16-44-09-B03-3CD3 antibodies. SEC was performed using an AKTA FPLC with a Superdex 200 increase 3.2/300 SEC column. Samples (0.1 ml) were run in PBS at a concentration of 0.5 to 1 mg/ml. DSF was performed by mixing 20 μL antibody (0.1 mg/ml) with 10 μL of Sypro Orange (BioRad). Fluorescence was monitored as a function of temperature (45 °C to 95 °C) using a real-time PCR machine. These experiments were utilized for developability assessment (including, without limitation, large-scale yield). TABLE 10 details developability parameters for each of the aforementioned antibodies.
TABLE 10: Developability parameters based on SEC and DSF experiments with IM- 16-44- 09-B03-1CD3, IM-16-44-09-B03-2CD3, and IM-16-44-09-B03-3CD3 antibodies
Example 3: Functional and Developability Characterization of I M-21-44-01 -E10-3 CD3 [00184] FIG. 6 shows that 1M-21-44-01-E10-3CD3 bound to both human CLDN 18.2 and human CD3. Flow cytometry experiments were performed to test the ability of the IM-21-44- 01-E10-3CD3 antibody to bind HEK293F cells expressing human CLDN 18.2, human CD3, or human P2X3. HEK293F cells were transfected by lipofectamine with target and control plasmids. At 18-24 hours post-transfection, cells were stained with bispecific and control MAbs followed by APC-labelled goat anti-human antibody. Fluorescence was detected using an Intellicyt high-throughput flow cytometer (iQue; Intellicyt, Albuquerque, NM). As shown in FIG. 6, IM-21 -44-01-E10-3CD3 selectively bound to both human CLDN 18.2 and human CD3, and not P2X3.
[00185] FIG. 7 shows cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) were co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IMC-16-C2 (control antibody) or IM-21 -44-01-E10-3CD3 antibodies. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated. As shown in FIG. 7, killing was only potently induced against CLDN 18.2-expressing cells by the IM-21-44-01 -E10-3CD3 antibody. No killing activity was observed using the control antibody or against cells expressing CLDN 18.1.
[00186] FIGs. 8A, 8B, 8C, 8D, and 8E show cytokines produced by human PBMC cultured with IMC-16-C2 or IM-16-44-09-B03-1CD3 antibodies and human CLDN 18.1- or 18.2- expressing cells. FIG. 8A shows the levels of IL-2, FIG. 8B shows the levels of IL-6, FIG.
8C shows the levels of IL-10, FIG. 8D shows the levels of IFN-γ (gamma), and FIG. 8E shows the levels of TNF-α (alpha) produced by human PBMCs in the presence of human
CLDN 18.1- or 18.2-expressing cells and the IMC-16-C2 or IM-21-44-01-E10-3CD3 antibodies. These experiments showed that IL-2, IL-6, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMC by the IM-21-44-01-E10-3CD3 18.2 antibody in the presence of CLDN 18.2-expressing cells.
[00187] FIGs. 9 A and 9B show SEC and DSF experimental results for IM-21-44-01 -E10- 1CD3, IM-21 -44-01-E10-2CD3, and IM-21-44-01-E10-3CD3 antibodies. SEC was performed using an AKTA FPLC with a Superdex 200 increase 3.2/300 SEC column.
Samples (0.1 ml) were run in PBS at a concentration of 0.5 to 1 mg/ml. DSF was performed by mixing 20 μL antibody (0.1 mg/ml) with 10 μL of Sypro Orange (BioRad). Fluorescence was monitored as a function of temperature (45 °C to 95 °C) using a real-time PCR machine. These experiments were utilized for developability assessment (including, without limitation, large-scale yield). TABLE 11 details developability parameters for each of the aforementioned antibodies.
TABLE 11: Developability parameters based on SEC and DSF experiments with IM-21 -44- 01-E10-1CD3, IM-21 -44-01-E10-2CD3, and IM-21 -44-01-E10-3CD3 antibodies
Example 4: Functional and. Developability Characterization of IM-21-44-09-B03-3CD3 [00188] FIG. 10 shows that IM-21-44-09-B03-3CD3 bound to both human CLDN 18.2 and human CD3. Flow cytometry experiments were performed to test the ability of the IM-21-44- 01-E10-3CD3 antibody to bind HEK-293F cells expressing human CLDN 18.2, human CD3, or human P2X3. HEK-293F cells were transfected by lipofectamine with target and control plasmids. At 18-24 hours post transfection, cells were stained with bispecific and control MAbs followed by APC-labelled goat anti-human antibody. Fluorescence was detected using an Intellicyt high-throughput flow cytometer (iQue; Intellicyt, Albuquerque, NM). As shown in FIG. 10, IM-21-44-09-B03-3CD3 selectively bound to both human CLDN 18.2 and human CD3, and not P2X3.
[00189] FIG. 11 shows cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) were co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IMC-16-C2 (control antibody) or IM-21-44-09-B03-3CD3 antibodies. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated. As shown in FIG. 11, killing was only potently induced against CLDN 18.2-expressing cells by the IM-21-44-09-B03-3CD3 antibody. No killing activity was observed using the control antibody or against cells expressing CLDN 18.1.
[00190] FIGs. 12A, 12B, 12C, 12D, and 12E show cytokines produced by human PBMC cultured with the IMC-16-C2 or IM-21-44-09-B03-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 12A shows the levels of IL-2, FIG. 12B shows the levels of IL-6, FIG. 12C shows the levels of IL-10, FIG. 12D shows the levels of IFN-γ (gamma), and FIG. 12E shows the levels of TNF-α (alpha) produced by human PBMCs in the presence of human CLDN 18.1- or 18.2-expressing cells and IMC-16-C2 or IM-21-44-09-B03-3CD3 antibodies. These experiments showed that IL-2, IL-6, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMC by the IM-21-44-09-B03-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
[00191] FIGs. 13A and 13B show SEC and DSF experimental results for IM-21 -44-09-B03- 1CD3, IM-21-44-09-B03-2CD3, and IM-21 -44-09-B03-3CD3 antibodies. SEC was performed using an AKTA FPLC with a Superdex 200 increase 3.2/300 SEC column.
Samples (0.1 ml) were run in PBS at a concentration of 0.5 to 1 mg/ml. DSF was performed by mixing 20 μl antibody (0.1 mg/ml) with 10 μl of Sypro Orange (BioRad). Fluorescence was monitored as a function of temperature (45 °C to 95 °C) using a real-time PCR machine.
These experiments were utilized for developability assessment (including, without limitation, large-scale yield). TABLE 12 details developability parameters for each of the aforementioned antibodies.
TABLE 12: Developability parameters based on SEC and DSF experiments with IM-21 -44- 09-B03-1CD3, IM-21 -44-09-B03-2CD3, and IM-21 -44-09-B03-3CD3 antibodies
Example 5: Functional Characterization of IM-2-44-05-H06-1CD3
[00192] FIG. 14 shows that IM-2-44-05-H06-1CD3 bound to both human CLDN 18.2 and human CD3. Flow cytometry experiments were performed to test the ability of the IM-2-44- 05-H06-1CD3 antibody to bind HEK293F cells expressing human CLDN 18.2, human CD3, or human P2X3. HEK293F cells were transfected by lipofectamine with target and control plasmids. At 18-24 hours post transfection, cells were stained with bispecific and control MAbs followed by APC-labelled goat anti-human antibody. Fluorescence was detected using an Intellicyt high-throughput flow cytometer (iQue; Intellicyt, Albuquerque, NM). As shown in FIG. 14, IM-2-44-05-H06-1CD3 selectively bound to both human CLDN 18.2 and human CD3, and not P2X3.
[00193] FIG. 15 shows cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) were co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IMC-16-C2 (control antibody) or IM-2-44-05-H06-1CD3 antibodies. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated. As shown in FIG. 15, killing was only potently induced against CLDN 18.2-expressing cells by the IM-2-44-05 -H06-1CD3 antibody. No killing activity was observed using the control antibody or against cells expressing CLDN 18.1.
[00194] FIGs. 16A, 16B, 16C, 16D, and 16E show cytokines produced by human PBMC cultured with the IMC-16-C2 or IM-2-44-05 -H06-1CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 16A shows the levels of IL-2, FIG. 16B shows the levels of IL-6, FIG. 16C shows the levels of IL- 10, FIG. 16D shows the levels of IFN-γ (gamma), and FIG. 16E shows the levels of TNF-α (alpha) produced by human PBMCs in the presence of human CLDN 18.1- or 18.2-expressing cells and IMC-16-C2 or IM-2-44-05-H06-1CD3 antibodies. These experiments showed, inter alia, that IL-2, IL-6, IL-10, IFN-γ, and TNF-α were selectively induced in human PBMC by the IM-2-44-05 -H06-1CD3 in the presence of human CLDN 18.2-expressing cells.
Example 6: Functional Characterization of IM-2-44-09-B03-1CD3
[00195] FIG. 17 shows that IM-2-44-09-B03-1CD3 bound to both human CLDN 18.2 and human CD3. Flow cytometry experiments were performed to test the ability of the IM-2-44- 09-B03-1CD3 antibody to bind HEK-293F cells expressing human CLDN 18.2, human CD3, or human P2X3. HEK293F cells were transfected by lipofectamine with target and control plasmids. At 18-24 hours post transfection, cells were stained with bispecific and control MAbs followed by APC-labelled goat anti-human antibody. Fluorescence was detected using an Intellicyt high-throughput flow cytometer (iQue; Intellicyt, Albuquerque, NM).-As shown in FIG. 17, IM-2-44-09-B03-1CD3 selectively bound to both human CLDN 18.2 and human CD3, and not P2X3.
[00196] FIG. 18 shows cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) were co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IMC-16-C2 (control antibody) or IM-2-44-09-B03-1CD3 antibodies. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit
(Promega) and percent cell killing was calculated. As shown in FIG. 18, killing was only potently induced against CLDN 18.2 expressing cells by the IM-2-44-09-B03-1CD3 antibody. No killing activity was observed using the control antibody or against cells expressing CLDN 18.2.
[00197] FIGs. 19A, 19B, 19C, 19D, and 19E show cytokines produced by human PBMC cultured with the IMC-16-C2 or IM-2-44-09-B03-1CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 19A shows the levels of IL-2, FIG. 19B shows the levels of IL-6, FIG. 19C shows the levels of IL-10, FIG. 19D shows the levels of IFN-γ (gamma), and FIG. 19E shows the levels of TNF-α (alpha) produced by human PBMCs in the presence of human CLDN 18.1- or 18.2-expressing cells and IMC-16-C2 or IM-2-44-09-B03-1CD3 antibodies. These experiments showed that IL-2 , IL-6, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMC by the IM-2-44-09-B03-1CD3 antibody in the presence of human CLDN 18.1 or 18.2-expressing cells .
Example 7: Cytotoxicity Characterization ofIMC-16-C2 (Control Antibody)
[00198] FIG. 20 is a graph of cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. FIG. 20 shows cytotoxicity data of human T cells (25,000 cells/well) co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IMC-16-C2 control antibodies. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated. These experiments showed that killing was not induced against human CLDN 18.1- or 18.2-expressing cells by the IMC-16-C2 control antibody.
Example 8: Functional Characterization of IM-42-44-09-B03-1CD3
[00199] FIG. 21 is a graph of cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) co-cultured with human CLDN
18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well)in the presence of IM-42-44-09-B03-1CD3 antibodies induced killing against human CLDN 18.2-expressing cells but not human CLDN 18.1 -expressing cells. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
[00200] FIGs. 22A, 22B, 22C, 22D, and 22E show cytokines produced by human PBMC cultured with IM-42-44-09-B03-1CD3 antibodies and human CLDN 18.1- or 18.2-expressing
cells. FIG. 22A shows the levels of IL-2, FIG. 22B shows the levels of IL-6, FIG. 22C shows the levels of IL-10, FIG. 22D shows the levels of IFN-γ (gamma), and FIG. 22E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-42-44-09-B03-1CD3 antibodies at the 24 hour (24hr) timepoint. These experiments showed that TNF-α was selectively induced in human PBMCs by the IM-42-44-09-B03-1CD3 antibody in the presence of human CLDN 18.2- expressing cells.
[00201] FIGs. 23A, 23B, 23C, 23D, and 23E shows cytokines produced by human PBMC cultured with IM-42-44-09-B03-1CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 23A shows the levels of IL-2, FIG. 23B shows the levels of IL-6, FIG. 23C shows the levels of IL-10, FIG. 23D shows the levels of IFN-γ (gamma), and FIG. 23E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-42-44-09-B03-1CD3 antibodies at the 48 hour (48hr) timepoint. These experiments showed that IFN-γ was selectively induced in human PBMCs by the IM-42-44-09-B03-1CD3 antibody in the presence of human CLDN 18.2- expressing cells.
Example 9: Functional Characterization of IM-42-44-09-B03-3CD3
[00202] FIG. 24 is a graph of cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence IM-42-44-09-B03-3CD3 antibodies induced killing against human CLDN 18.2 expressing cells but not human CLDN 18.1 expressing cells. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
[00203] FIGs. 25A, 25B, 25C, 25D, and 25E shows cytokines produced by human PBMC cultured with IM-42-44-09-B03-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 25A shows the levels of IL-2, FIG. 25B shows the levels of IL-6, FIG. 25C shows the levels of IL-10, FIG. 25D shows the levels of IFN-γ (gamma), and FIG. 25E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-42-44-09-B03-3CD3 antibodies at the 24 hour (24hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs by the IM-42-44-09-B03-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
[00204] FIGs. 26A, 26B, 26C, 26D, and 26E shows cytokines produced by human PBMC cultured with IM-42-44-09-B03-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 26A shows the levels of IL-2, FIG. 26B shows the levels of IL-6, FIG. 26C shows the levels of IL-10, FIG. 26D shows the levels of IFN-γ (gamma), and FIG. 26E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-42-44-09-B03-3CD3 antibodies al the 48 hour (48hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs by the IM-42-44-09-B03-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
Example 10: Functional Characterization of IM-42-44-01-E10-1CD3
[00205] FIG. 27 is a graph of cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IM-42-44-01-E10-1CD3 induced killing against CLDN 18.2-expressing cells but not human CLDN 18.1 -expressing cells. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
[00206] FIGs. 28A, 28B, 28C, 28D, and 28E shows cytokines produced by human PBMC cultured with IM-42-44-01-E10-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 28A shows the levels of IL-2, FIG. 28B shows the levels of IL-6, FIG. 28C shows the levels of IL-10, FIG. 28D shows the levels of IFN-γ (gamma), and FIG. 28E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM -42-44-01 -E10-3CD3 antibodies at the 24 hour (24hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs by the IM-42-44-01-E10-1CD3 antibody in the presence of human CLDN 18.2-expressing cells.
[00207] FIGs. 29A, 29B, 29C, 29D, and 29E shows cytokines produced by human PBMC cultured with IM-42-44-01-E10-1CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 29A shows the levels of IL-2, FIG. 29B shows the levels of IL-6, FIG. 29C shows the levels of IL-10, FIG. 29D shows the levels of IFN-γ (gamma), and FIG. 29E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-42-44-01-E10-1CD3 antibodies at the 48 hour (48hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were
selectively induced in human PBMCs by the IM-42-44-01-E10-1CD3 antibody in the presence of human CLDN 18.2-expressing cells.
Example 11: Functional Characterization of IM-42-44-01-E10-3CD3
[00208] FIG. 30 is a graph of cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IM-42-44-01 -E10-3CD3 antibodies induced killing against human CLDN 18.2-expressing cells but not human CLDN 18.1 -expressing cells. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
[00209] FIGs. 31A, 31B, 31C, 31D, and 31E shows cytokines produced by human PBMC cultured with IM-42-44-01-E10-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 31A shows the levels of IL-2, FIG. 31B shows the levels of IL-6, FIG. 31C shows the levels of IL-10, FIG. 31D shows the levels of IFN-γ (gamma), and FIG. 31E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM -42-44-01 -E10-3CD3 antibodies at the 24 hour (24hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs by the IM-42-44-01 -E10-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
[00210] FIGs. 32A, 32B, 32C, 32D, and 32E shows cytokines produced by human PBMC cultured with IM-42-44-01-E10-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 32A shows the levels of IL-2, FIG. 32B shows the levels of IL-6, FIG. 32C shows the levels of IL-10, FIG. 32D shows the levels of IFN-γ (gamma), and FIG. 32E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM -42-44-01 -E10-3CD3 antibodies at the 48 hour (48hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs by the IM-42-44-01 -E10-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
Example 12: Functional Characterization of IM-42-44-05-H06-1CD3
[00211] FIG. 33 is a graph of cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence
of IM-42-44-05-H06-1CD3 antibodies induced killing against human CLDN 18.2-expressing cells but not human CLDN 18.1 -expressing cells. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
[00212] FIGs. 34A, 34B, 34C, 34D, and 34E shows cytokines produced by human PBMC cultured with IM-42-44-05-H06-1CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 34A shows the levels of IL-2, FIG. 34B shows the levels of IL-6, FIG. 34C shows the levels of IL-10, FIG. 34D shows the levels of IFN-γ (gamma), and FIG. 34E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-42-44-05-H06-1CD3 antibodies at the 24 hour (24hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs by IM-42-44-05 -H06-1CD3 antibody in the presence of human CLDN 18.2-expressing cells.
[00213] FIGs. 35A, 35B, 35C, 35D, and 35E shows cytokines produced by human PBMC cultured with IM-42-44-05-H06-1CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 35A shows the levels of IL-2, FIG. 35B shows the levels of IL-6, FIG. 35C shows the levels of IL-10, FIG. 35D shows the levels of IFN-γ (gamma), and FIG. 35E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-42-44-05-H06-1CD3 antibodies at the 48 hour (48hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs by the IM-42-44-05 -H06-1CD3 antibody in the presence of human CLDN 18.2-expressing cells.
Example 13: Functional Characterization of IM-42-44-05-H06-3CD3
[00214] FIG. 36 is a graph of cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence of IM-42-44-05-H06-3CD3 antibodies induced killing against human CLDN 18.2-expressing cells but not human CLDN 18.1 -expressing cells. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
[00215] FIGs. 37A, 37B, 37C, 37D, and 37E show cytokines produced by human PBMC cultured with IM-42-44-05-H06-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 37A shows the levels of IL-2, FIG. 37B shows the levels of IL-6, FIG. 37C
shows the levels of IL-10, FIG. 37D shows the levels of IFN-γ (gamma), and FIG. 37E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-42-44-05-H06-3CD3 antibodies at the 24 hour (24hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs by IM-42-44-05-H06-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
[00216] FIGs. 38A, 38B, 38C, 38D, and 38E shows cytokines produced by human PBMC cultured with IM-42-44-05-H06-3CD3 antibodies and human CLDN 18.1 - or 18.2-expressing cells. FIG. 38A shows the levels of IL-2, FIG. 38B shows the levels of IL-6, FIG. 38C shows the levels of IL-10, FIG. 38D shows the levels of IFN-γ (gamma), and FIG. 38E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-42-44-05-H06-3CD3 antibodies at the 48 hour (48hr) timepoint. These experiments showed that IL-2, IL- 10, and IFN-γ were selectively induced in human PBMCs by the IM-42-44-05-H06-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
Example 14: Functional Characterization of IM-21-44-09-B03-3CD3
[00217] FIG. 39 is a graph of cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence IM-21 -44-09-B03-3CD3 antibodies induced killing against CLDN 18.2-expressing cells but not human CLDN 18.1 -expressing cells. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
[00218] FIGs. 40A, 40B, 40C, 40D, and 40E shows cytokines produced by human PBMC cultured with IM-21-44-09-B03-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 40A shows the levels of IL-2, FIG. 40B shows the levels of IL-6, FIG. 40C shows the levels of IL-10, FIG. 40D shows the levels of IFN-γ (gamma), and FIG. 40E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-21-44-09-B03-3CD3 antibodies at the 24 hour (24hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs IM-21 -44-09-B03-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
[00219] FIGs. 41A, 41B, 41C, 41D, and 41E shows cytokines produced by human PBMC cultured with IM-21-44-09-B03-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 41A shows the levels of IL-2, FIG. 41B shows the levels of IL-6, FIG. 41C shows the levels of IL-10, FIG. 41D shows the levels of IFN-γ (gamma), and FIG. 41E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-21-44-09-B03-3CD3 antibodies al the 48 hour (48hr) timepoint. These experiments showed that IL-2, IL-6, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs by IM-21 -44-09-B03-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
Example 15: Functional Characterization of IM-21-44-01-E10-3CD3
[00220] FIG. 42 is a graph of cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence IM-21-44-01-E10-3CD3 antibodies induced killing against human CLDN 18.2-expressing cells but not human CLDN 18.1 -expressing cells. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
[00221] FIGs. 43A, 43B, 43C, 43D, and 43E shows cytokines produced by human PBMC cultured with IM-21-44-01-E10-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 43A shows the levels of IL-2, FIG. 43B shows the levels of IL-6, FIG. 43C shows the levels of IL-10, FIG. 43D shows the levels of IFN-γ (gamma), and FIG. 43E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-21-44-01-E10-3CD3 antibodies at the 24 hour (24hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs IM-21-44-01-E10-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
[00222] FIGs. 44A, 44B, 44C, 44D, and 44E shows cytokines produced by human PBMC cultured with IM-21-44-01-E10-3CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 44A shows the levels of IL-2, FIG. 44B shows the levels of IL-6, FIG. 44C shows the levels of IL-10, FIG. 44D shows the levels of IFN-γ (gamma), and FIG. 44E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-21-44-01-E10-3CD3 antibodies at the 48 hour (48hr) timepoint. These experiments showed that IL-2, IL-6, IL- 10, IFN-γ, and TNF-α were
selectively induced in human PBMCs by IM-21-44-01-E10-3CD3 antibody in the presence of human CLDN 18.2-expressing cells.
Example 16: Functional Characterization of IM -2-44-05- H06-1 CD3
[00223] FIG. 45 is a graph of cellular toxicity data from T-cell dependent cellular cytotoxicity experiments. Human T cells (25,000 cells/well) co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells (2500 cells/well) in the presence IM-2-44-05-H06-1CD3 antibodies induced killing against human CLDN 18.2-expressing cells but not human CLDN 18.1 -expressing cells. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
[00224] FIGs. 46A, 46B, 46C, 46D, and 46E shows cytokines produced by human PBMC cultured with IM-2-44-05-H06-1CD3 antibodies and human CLDN 18.1 or 18.2 expressing cells. FIG. 46A shows the levels of IL-2, FIG. 46B shows the levels of IL-6, FIG. 46C shows the levels of IL-10, FIG. 46D shows the levels of IFN-γ (gamma), and FIG. 46E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-2-44-05-H06-1CD3 antibodies at the 24 hour (24hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs IM-2-44-01-H06-1CD3 antibody in the presence of human CLDN 18.2-expressing cells.
[00225] FIGs. 47A, 47B, 47C, 47D, and 47E shows cytokines produced by human PBMC cultured with IM-2-44-05-H06-1CD3 antibodies and human CLDN 18.1- or 18.2-expressing cells. FIG. 47A shows the levels of IL-2, FIG. 47B shows the levels of IL-6, FIG. 47C shows the levels of IL-10, FIG. 47D shows the levels of IFN-γ (gamma), and FIG. 47E shows the levels of TNF-α (alpha) produced by human PBMC in the presence of human CLDN 18.1- or 18.2-expressing cells and IM-2-44-05-H06-1CD3 antibodies at the 48 hour (48hr) timepoint. These experiments showed that IL-2, IL- 10, IFN-γ, and TNF-α were selectively induced in human PBMCs by IM-2-44-05-H06-1CD3 antibody in the presence of human CLDN 18.2-expressing cells.
Example 17: mRNA Production of Antibodies Specific for Claudin 18.2 in T-cell
Dependent Cellular Cytotoxicity Assay
Materials and Methods
[00226] HEK293T cells were plated at 1,250 cells (control Cells: CLDN18.1 -positive HEK293T w/ ffLuc, P29; target Cells: CLDN18.2-positive HEK293T w/ ffLuc, P34) per well in a 384 well plate. The next day, the cells were transfected with mRNA encoding a respective antibody described in TABLE 13 starting at 0.05 μg and titrated by 1/4 A-H in two columns for each mRNA product. Columns that be administered antibody protein were treated with the same titration of lipofectamine without mRNA. The same day as transfection, antibody protein controls were added to the plate along with the T-cells (Lot 08152022) in a T-cell Dependent Cellular Cytotoxicity (TDCC) assay, as described throughout Examples 2- 16. Briefly, in the TDCC assay, control and target cells were harvested, and seeded at 1,250 cells/well. On the next day, mRNA in lipofectamine or just lipofectamine was added to cells. A column for each mRNA construct was left without T-cells to determine if mRNA transfection caused cell death. In wells that were not administered mRNA, protein controls were added. Next, T-cells were thawed and 25,000 were seeded. At 24 hours, 5 μL/well was collected for cytokine analysis. At 48 hours, 25 μL/well was collected for cytokine analysis. The Bright-Glo Substrate was added, and luminescence was read to measure killing. For data analysis, data was processed for control correction and organization using R and all curve fitting and graphing was done in GraphPad Prism 9. Non-linear Regression (curve fit) - log(agonist) vs. response (three parameters) for fitting were used. Testing conditions are listed in TABLE 14 and the results of the TDCC assay are provided in FIG. 48 and TABLE 15.
[00227] FIG. 48 is a graph of cellular toxicity data from TDCC experiments. Human T cells co-cultured with human CLDN 18.1 or CLDN 18.2-expressing ffLuciferase HEK293F cells in the presence of mRNA encoding IM-2-44-09B03-1CD3 or IM-16-44-09B03-1CD3 antibodies induced killing against human CLDN 18.2-expressing cells but not human CLDN 18.1-expressing cells. After 48 hour incubation at 37 °C, luciferase activity was detected using a Bright-Glo luciferase assay kit (Promega) and percent cell killing was calculated.
[00228] The data show that both mRNA molecules were able to kill their target cell lines at a comparable rate with the IM-16-44-09B03-1CD3 being slightly more efficient than IM-2-44- 09B03-1CD3. This result was unexpected, as the IM-16-44-09B03-1CD3 protein reference is considerably better at killing, however the finding could be due to expression since the IM-2- 44-09B03-1CD3 is just one plasmid so more mRNA is being transfected. Meanwhile, reference and control killing was observed, as expected.
[00229] All of the features disclosed herein may be combined in any combination. Each feature disclosed in this specification may be replaced by an alternative feature serving the same, equivalent, or similar purpose. Thus, unless expressly stated otherwise, each feature disclosed is only an example of a generic series of equivalent or similar features.
[00230] From the above description, one skilled in the art can easily ascertain the essential characteristics of the present disclosure, and without departing from the spirit and scope thereof, can make various changes and modifications of the disclosure to adapt it to various usages and conditions. Thus, other embodiments are also within the claims.
Claims
1. A composition comprising an scFv-Fab IgG antibody specific for Claudin 18.2 and CD3 comprising: a first heavy chain, a second heavy chain, and a light chain, wherein:
(a) the first heavy chain is selected from:
(i)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSS
YGMGWVRQAPGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNT
LYLQMNSLRAEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDAW
GQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVT
VSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVN
HKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTL
MISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYN STYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR EPQVYTLPPSREEMTKNQVSLWCLVKGFYPSDIAVEWESNGQPENNY KTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQ KSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, or (ii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSS
YGMSWVRQAPGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKN
TLYLQMNSLRAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHHIDA
WGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPV
TVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTYICNV
NHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQY
NSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQP
REPQVYTLPPSREEMTKNQVSLWCLVKGFYPSDIAVEWESNGQPENN
YKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYT QKSLSLSPGK (SEQ ID NO: 4767), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof;, or
(iii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRS
YAMMWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKN
TLYLQMNSLRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVS
SASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALT
SGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQTY1CNVNHKPSNTKVD
KKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTC
VVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLT VLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSR EEMTKNQVSLWCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDG SFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4768), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(b) the second heavy chain is selected from:
(i)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFN
KYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDD
SKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLV
TVSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAV
TSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALT
LSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLASPKSSDKTHTCPP
CPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFN
WYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKC
KVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAV
KGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRW
QQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4168), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSY1SYWAY (SEQ ID NO: 6) or a variant thereof and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), or
(ii)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFT
RYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKS
TAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGG
SGGGGSGGGGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQ
TPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTISSLQPEDIATYY
CQQWSSNPFTFGQGTKLQITRASPKSSDKTHTCPPCPAPEAAGGPSVFL
FPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKT
KPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTIS
KAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESN
GQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEA
LHNHYTQKSLSLSPGK (SEQ ID NO: 4169),
an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, or (iii)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYR
GYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAK
NTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVS
SASPKSSDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCV
VVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTV LHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSRE EMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGS FFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 3983), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and
(b) the light chain is selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWY
QQKPGQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCG GYDSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPG AVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSC QVTHEGSTVEKTVAPTECS (SEQ ID NO: 3576) or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof,
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQ KPGQAPVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGY DSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAV TVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQV THEGSTVEKTVAPTECS (SEQ ID NO: 3593) or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQ QKPGQAPVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGG YDSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGA VTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQ VTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or
a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof; optionally wherein the first or second heavy chain comprises a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N- or C -terminus; and/or optionally wherein the composition comprises a linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof.
2. The composition of claim 1, wherein the Fc is from IgG.
3. The composition of claim 2, wherein the IgG is human IgG.
4. The composition of claim 3, wherein the human IgG is selected from IgGl, IgG2,
IgG3, and IgG4.
5. The composition of any one of claims 2 to 4, wherein the antibody is formed through a knob-in-hole interaction in the Fc region.
6. The composition of any one of claims 3 to 5, wherein the human IgG Fc comprises one or mutations to promote knob-in-hole interactions.
7. The composition of claim 6, wherein the mutations are selected from (i) T366Y or T366W, and (ii) Y407T, Y407A, or Y407V.
8. The composition of claim 6, wherein the mutations are selected from:
(a) T366Y and Y407T; or T366Y/F405A and T394W/Y407T in human
IgGl; or
(b) T366W/D399C and T366S/L368A/K392C/Y407V;
T366W/K392C and T366S/L368A/D399C/Y407V;
S354C/T366W and Y349C/T366S/L368A/Y407V;
Y349C/T366W and S354C/T366S/L368A/Y407V;
E356C/T366W and Y349C/T366S/L368A/Y407V;
Y349C/T366W and E356C/T366S/L368A/Y407V;
E357C/T366W and Y349C/T366S/L368A/Y407V; or
Y349C/T366W and E357C/T366S/L368A/Y407V in human IgGl.
9. The composition of claim 3, wherein the human IgG Fc comprises one or mutations to reduce or eliminate the effector function of the Fc domains.
10. The composition of claim 9, wherein the mutations are L234A and L235A (LALA) substitutions in human IgGl.
11. The composition of claim 3, wherein the human IgG Fc comprises one or mutations to stabilize a hinge region in the Fc domain.
12. The composition of claim 11, wherein the mutation is S228P.
13. A composition comprising an IgG-scFv antibody specific for Claudin 18.2 and CD3 comprising:
(a) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMG
WVRQAPGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLR
AEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTK
GPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL
QSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCP
PCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLWCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALH NHYTQKSLSLSPGK (SEQ ID NO: 3869), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and
(ii) the second heavy chain comprises:
(A)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSM
GWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTS
LKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSSGGGGSGGGGSGG
GGSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQAPGKGLEWV
AAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKDA
SNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS
GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTV
PSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVF
LFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPRE
EQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPRE
PQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPV
LDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4778), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, or
(B)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFN
KYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDD
SKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLV
TVSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAV
TSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALT
LSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLGGGGSGGGGSGGG
GSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMGWVRQAPGKGL
EWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAV
YYCAKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGP
SVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFP
AVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKS
CDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSH
EDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWL NGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQ VSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKL TVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4803), or an amino acid sequence having al least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, or
(C)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFT
RYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKS
TAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGG
SGGGGSGGGGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQ
TPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTISSLQPEDIATYY
CQQWSSNPFTFGQGTKLQITRGGGGSGGGGSGGGGSEVQLLESGGGL
VQPGGSLRLSCAASGFTFSSYGMGWVRQAPGKGLEWVAAITRDSSTN
YGAAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKDASNGYC
WYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSG
GTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSV
VTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPE
AAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVD
GVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNK ALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPS DIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNV FSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4806); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof; and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, or
(b) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMS WVRQAPGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSL RAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSAST KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAV
LQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTC
PPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLWCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALH NHYTQKSLSLSPGK (SEQ ID NO: 4767), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSM
GWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTS
LKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSSGGGGSGGGGSGG
GGSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQAPGKGLEWV
AGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKD
ATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKST
SGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVT
VPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSV
FLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPR
EEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR
EPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPP VLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4779), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain
CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof,
(B)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFN
KYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDD
SKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLV
TVSSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAV
TSGNYPNWVQQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALT
LSGVQPEDEAEYYCVLWYSNRWVFGGGTKLTVLGGGGSGGGGSGGG
GSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMSWVRQAPGKGL
EWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTA
VYYCAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKG
PSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTF
PAVLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPK
SCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVS
HEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQD
WLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTK
NQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLVS
KLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4804), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and
(C)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFT
RYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKS
TAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGG
SGGGGSGGGGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQ
TPGKAPKRWIYDTSKLASGVPSRFSGSGSGTDYTFTISSLQPEDIATYY
CQQWSSNPFTFGQGTKLQITRGGGGSGGGGSGGGGSEVQLLESGGGL
VQPGGSLRLSCAASGFTFSSYGMSWVRQAPGKGLEWVAGIGSSGSDT
WYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKDATNGY
CWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTS
GGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSS
VVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAP
EAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVD
GVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNK ALPAPEEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPS DIAVEWESNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNV FSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4807); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain
CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, and/or heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, or
(c) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAM MWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNS LRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAP SSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLS SVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAG GPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAK
TKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK GQPREPQVYTLPPSREEMTKNQVSLWCLVKGFYPSDIAVEWESNGQPENNYK TTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS PGK (SEQ ID NO: 4768), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYSM
GWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQMTS
LKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSSGGGGSGGGGSGG
GGSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWVRQAPGKGLEW
VGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAEDTAVYYCAKS
TYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKSTSGGTAALGC
LVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVVTVPSSSLGTQT
YICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDT
LMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKPREEQYNSTYR
VVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPP SREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFF LVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4780), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof,
(B)
MKWVTFISLLFLFSSAYSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAM
NWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQ
MNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGG
SGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAP
RGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRW
VFGGGTKLTVLGGGGSGGGGSGGGGSEVQLLESGGGLVQPGGSLRLSCAAS
GFTFRSYAMMWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSK
NTLYLQMNSLRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSAST
KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAV
LQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTC
PPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALH NHYTQKSLSLSPGK (SEQ ID NO: 4805), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(C)
MKWVTFISLLFLFSSAYSQVQLVQSGGGVVQPGRSLRLSCKASGYTFTRYTM HWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKSTAFLQMDSL RPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGGGGSDI QMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSKLA
SGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTFGQGTKLQITRG
GGGSGGGGSGGGGSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAMMWV
RQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNSLRAE
DTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAPSSKS
TSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLSSVV
TVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPS
VFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKTKP
REEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQP REPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYKTTP PVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLSPGK (SEQ ID NO: 4808); or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(b) a light chain selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWY
QQKPGQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCG
GYDSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPG AVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSC QVTHEGSTVEKTVAPTECS (SEQ ID NO: 3576), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof,
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQ KPGQAPVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGY DSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAV TVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQV THEGSTVEKTVAPTECS (SEQ ID NO: 3593), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQ QKPGQAPVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGG YDSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGA VTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQ VTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or
a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof; optionally wherein the first or second heavy chain comprises a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N- or C -terminus; and/or optionally wherein the composition comprises a linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof.
14. The composition of claim 13, wherein the Fc is from IgG.
15. The composition of claim 14, wherein the IgG is human IgG.
16. The composition of claim 15, wherein the human IgG is selected from IgGl, IgG2,
IgG3, and IgG4.
17. The composition of any one of claims 13 to 16, wherein the antibody is formed through a knob-in-hole interaction in the Fc region.
18. The composition of any one of claims 15-17, wherein the human IgG Fc comprises one or mutations to promote knob-in-hole interaction.
19. The composition of claim 18, wherein the mutations are selected from:
(i) T366Y or T366W; and
(ii) Y407T, Y407A, or Y407V.
20. The composition of claim 18, wherein the mutations are selected from:
(a) T366Y and Y407T; or T366Y/F405A and T394W/Y407T in human
IgGl; or
(b) T366W/D399C and T366S/L368A/K392C/Y407V;
T366W/K392C and T366S/L368A/D399C/Y407V;
S354C/T366W and Y349C/T366S/L368A/Y407V;
Y349C/T366W and S354C/T366S/L368A/Y407V;
E356C/T366W and Y349C/T366S/L368A/Y407V;
Y349C/T366W and E356C/T366S/L368A/Y407V;
E357C/T366W and Y349C/T366S/L368A/Y407V; or
Y349C/T366W and E357C/T366S/L368A/Y407V, in human IgGl.
21. The composition of claim 15, wherein the human IgG Fc comprises one or mutations to reduce or eliminate the effector function of the Fc domains.
22. The composition of claim 21, wherein the mutations are L234A and L235A (LALA) substitutions in human IgGl .
23. The composition of claim 15, wherein the human IgG Fc comprises one or mutations to stabilize a hinge region in the Fc domain.
24. The composition of claim 23, wherein the mutation is S228P.
25. A composition comprising an IgG-(scFv)2 antibody specific for Claudin 18.2 and
CD3 comprising:
(a) a heavy chain selected from:
(i)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMG
WVRQAPGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLR
AEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTK
GPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL
QSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCP
PCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP
APIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES
NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHN
HYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLVQPGGSLKLSCA
ASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISR
DDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVS
SGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPN
WVQQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEY YCVLWYSNRWVFGGGTKLTVL (SEQ ID NO: 4769), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), (ii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMG WVRQAPGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLR AEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTK GPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL
QSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCP
PCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP
APIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES
NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHN
HYTQKSLSLSPGKGGGGSGGGGTGGGGSQVQLVQSGGGVVQPGRSLRLSCK
ASGYTFTRYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKS
KSTAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGGSG
GGGSGGGGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAP KRWIYDTSKLASGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTF GQGTKLQITR (SEQ ID NO: 4770), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, (iii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMG WVRQAPGKGLEWVAAITRDSSTNYGAAVKGRAT1SRDNSKNTLYLQMNSLR AEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTK GPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL
QSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCP
PCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHN HYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPVQAGGSLRLSCA ASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDN AKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS
(SEQ ID NO: 4771), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is
CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, (iv)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMS
WVRQAPGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSL
RAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSAST
KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAV
LQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTC
PPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP
APIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES
NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHN
HYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLVQPGGSLKLSCA
ASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISR
DDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVS
SGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPN
WVQQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEY
YCVLWYSNRWVFGGGTKLTVL (SEQ ID NO: 4772), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is
VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9),
(v)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMS
WVRQAPGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSL
RAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSAST
KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAV
LQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTC
PPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP
APIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES
NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHN
HYTQKSLSLSPGKGGGGSGGGGTGGGGSQVQLVQSGGGVVQPGRSLRLSCK
ASGYTFTRYTMHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKS
KSTAFLQMDSLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGGSG
GGGSGGGGSDIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAP
KRWIYDTSKLASGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTF
GQGTKLQITR (SEQ ID NO: 4773), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID
NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, (vi)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMS
WVRQAPGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSL
RAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSAST
KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAV
LQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTC
PPCPAPEAAGGPSVFLFPPKPKDTLM1SRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWES NGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHN HYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPVQAGGSLRLSCA ASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDN AKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 4774), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is
VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, (vii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAM MWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNS LRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAP
SSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLS SVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAG GPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAK
TKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK
GQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYK TTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS PGKGGGGSGGGGTGGGGSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYA MNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYL QMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGG GSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQ APRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSN RWVFGGGTKLTVL (SEQ ID NO: 4775), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ
ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9), (viii)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAM MWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNS LRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAP SSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLS
SVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAG
GPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAK
TKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK GQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYK TTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS PGKGGGGSGGGGTGGGGSQVQLVQSGGGVVQPGRSLRLSCKASGYTFTRYT MHWVRQAPGKGLEWIGYINPSRGYTNYNQKVKDRFTISTDKSKSTAFLQMD SLRPEDTAVYYCARYYDDHYCLDYWGQGTTLTVSSGGGGSGGGGSGGGGS DIQMTQSPSSLSASVGDRVTITCSASSSVSYMNWYQQTPGKAPKRWIYDTSK LASGVPSRFSGSGSGTDYTFTISSLQPEDIATYYCQQWSSNPFTFGQGTKLQIT R (SEQ ID NO: 4776), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GYTFTRYTMHWV (SEQ ID NO: 10) or a variant thereof, the heavy chain CDR2 sequence is IGYINPSRGYTNYNQ (SEQ ID NO: 11) or a variant thereof, and the heavy chain CDR3 sequence is CARYYDDHYCLDY (SEQ ID NO: 12) or a variant thereof, and/or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CSASSSVSYMN (SEQ ID NO: 13) or a variant thereof, the heavy chain CDR2 sequence is IYDTSKLASGVP (SEQ ID NO: 14) or a variant thereof, and the heavy chain CDR3 sequence is CQQWSSNPFTF (SEQ ID NO: 15) or a variant thereof, and (ix.)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAM MWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNS
LRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAP
SSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLS
SVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAG
GPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAK
TKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK GQPREPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYK TTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS PGKGGGGSGGGGTGGGGSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYS MGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQM TSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 4777), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and
(b) a light chain selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWY QQKPGQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCG GYDSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPG AVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSC QVTHEGSTVEKTVAPTECS (SEQ ID NO: 3576), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1 , CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof,
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQ KPGQAPVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGY DSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAV TVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQV THEGSTVEKTVAPTECS (SEQ ID NO: 3593), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, and
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQ
QKPGQAPVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGG
YDSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGA
VTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQ VTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1 , CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof; optionally wherein the first or second heavy chain comprises a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N- or C -terminus; and/or optionally wherein the composition comprises a linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof.
26. The composition of claim 25, wherein the Fc is from IgG.
27. The composition of claim 26, wherein the IgG is human IgG.
28. The composition of claim 27, wherein the human IgG is selected from IgGl, IgG2,
IgG3, and IgG4.
29. The composition of claim 26, wherein the human IgG Fc comprises one or mutations to reduce or eliminate the effector function of the Fc domains.
30. The composition of claim 29, wherein the mutations are L234A and L235A (LALA) substitutions in human IgGl.
31. The composition of claim 27, wherein the human IgG Fc comprises one or mutations to stabilize a hinge region in the Fc domain.
32. The composition of claim 31, wherein the mutation is S228P.
33. A composition comprising an IgG-(scFv) antibody specific for Claudin 18.2 and CD3 comprising:
(a) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMG
WVRQAPGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLR
AEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTK
GPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL
QSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCP
PCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLWCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALH NHYTQKSLSLSPGK (SEQ ID NO: 3869), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMG
WVRQAPGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLR
AEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTK
GPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL
QSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCP
PCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP
APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWE
SNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALH
NHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLVQPGGSLKLSC
AASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTIS
RDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTV
SSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYP
NWVQQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAE
YYCVLWYSNRWVFGGGTKLTVL (SEQ ID NO: 5202), an amino acid sequence having at least about 90%, about 93%, about 95%, about
97%, about 98%, or about 99% identity thereto, or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
(B)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMG WVRQAPGKGLEWVAAITRDSSTNYGAAVKGRATISRDNSKNTLYLQMNSLR AEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSASTK GPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVL
QSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCP
PCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALH
NHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPVQAGGSLRLSC AASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRD NAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS
(SEQ ID NO: 5205), or an amino acid sequence having at least about 90%, about 93%, about 95%, about
97%, about 98%, or about 99% identity thereto, or
a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence GFIFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the heavy chain CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the heavy chain CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, or
(b) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMS
WVRQAPGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSL
RAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSAST
KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAV
LQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTC
PPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLWCLVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALH NHYTQKSLSLSPGK (SEQ ID NO: 4767), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMS
WVRQAPGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSL
RAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSAST
KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAV
LQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTC
PPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP
APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWE
SNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALH
NHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGLVQPGGSLKLSC AASGFTFNKYAMNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTIS RDDSKNTAYLQMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTV SSGGGGSGGGGSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYP NWVQQKPGQAPRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAE YYCVLWYSNRWVFGGGTKLTVL (SEQ ID NO: 5203), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP
(SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
(B)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFSSYGMS
WVRQAPGKGLEWVAGIGSSGSDTWYADSVKGRFTISRDNSKNTLYLQMNSL RAEDTAVYYCAKDATNGYCWYTGRGSYSCGAHHIDAWGQGTLVTVSSAST
KGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAV
LQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTC
PPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYV
DGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALP
APIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWE SNGQPENNYKTTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALH NHYTQKSLSLSPGKGGGGSGGGGTGGGGSEVQLVESGGGPVQAGGSLRLSC AASGRTYRGYSMGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRD NAKNTMYLQMTSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 5206) or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1 , CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFSSYGMSWV (SEQ ID NO: 19) or a variant thereof, the heavy chain CDR2 sequence is VAGIGSSGSDTWYA (SEQ ID NO: 20) or a variant thereof, and the heavy chain CDR3 sequence is CAKDATNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 21) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, or
(c) a first heavy chain and a second heavy chain, wherein
(i) the first heavy chain comprises:
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAM
MWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNS
LRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAP
SSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLS
SVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAG
GPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAK
TKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK
GQPREPQVYTLPPSREEMTKNQVSLWCLVKGFYPSDIAVEWESNGQPENNYK
TTPPVLDSDGSFFLYSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS
PGK (SEQ ID NO: 4768), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and
(ii) the second heavy chain is selected from:
(A)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAM
MWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNS
LRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAP
SSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLS
SVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAG
GPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAK
TKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK
GQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYK
TTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS
PGKGGGGSGGGGTGGGGSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYA
MNWVRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYL
QMNNLKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGG
GSGGGGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQ
APRGLIGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSN
RWVFGGGTKLTVL (SEQ ID NO: 5204), or
an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the heavy chain CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the heavy chain CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the heavy chain CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the heavy chain CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, and
(B)
MKWVTFISLLFLFSSAYSEVQLLESGGGLVQPGGSLRLSCAASGFTFRSYAM
MWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATISRDNSKNTLYLQMNS
LRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVTVSSASTKGPSVFPLAP
SSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSGVHTFPAVLQSSGLYSLS
SVVTVPSSSLGTQTYICNVNHKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAG
GPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAK
TKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAK
GQPREPQVYTLPPSREEMTKNQVSLSCAVKGFYPSDIAVEWESNGQPENNYK
TTPPVLDSDGSFFLVSKLTVDKSRWQQGNVFSCSVMHEALHNHYTQKSLSLS PGKGGGGSGGGGTGGGGSEVQLVESGGGPVQAGGSLRLSCAASGRTYRGYS MGWFRQAPGKEREFVAAIVWSGGNTYYEDSVKGRFTISRDNAKNTMYLQM TSLKPEDSATYYCAAKIRPYIFKIAGQYDYWGQGTQVTVSS (SEQ ID NO: 5207), or
an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the heavy chain CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the heavy chain CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a heavy chain variable region comprising heavy chain CDR1, CDR2, and CDR3 sequences, wherein the heavy chain CDR1 sequence is GRTYRGYSMGW (SEQ ID NO: 16) or a variant thereof, the heavy chain CDR2 sequence is VAAIVWSGGNTYYE (SEQ ID NO: 17) or a variant thereof, and the heavy chain CDR3 sequence is CAAKIRPYIFKIAGQYDY (SEQ ID NO: 18) or a variant thereof, and
(b) a light chain selected from:
(i)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGSGSYGYYGWY QQKPGQAPVTVIYGTNKRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCG GYDSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPG AVTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSC QVTHEGSTVEKTVAPTECS (SEQ ID NO: 3576), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGSGSYGYYG (SEQ ID NO: 25) or SGGSGSYGYYG (SEQ ID NO: 30) or a variant thereof, the light chain CDR2 sequence is IYGTNKRP (SEQ ID NO: 26) or GTNKRPS (SEQ ID NO: 32) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof,
(ii)
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQ KPGQAPVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGY DSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAV
TVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQV THEGSTVEKTVAPTECS (SEQ ID NO: 3593), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1 , CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof, or
(iii)
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQ QKPGQAPVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGG YDSSAGIFGGGTKLTVLGQPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGA VTVAWKADSSPVKAGVETTTPSKQSNNKYAASSYLSLTPEQWKSHRSYSCQ VTHEGSTVEKTVAPTECS (SEQ ID NO: 3817), an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a light chain variable region comprising light chain CDR1, CDR2, and CDR3 sequences, wherein the light chain CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or SGGYNGHYG (SEQ ID NO: 28) or a variant thereof, the light chain CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or SNNQRPS (SEQ ID NO: 31) or a variant thereof, and the light chain CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or GGYDSSAGI (SEQ ID NO: 33) or a variant thereof; optionally wherein the first or second heavy chain comprises a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N- or C -terminus; and/or optionally wherein the composition comprises a linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof.
34. The composition of claim 33, wherein the Fc is from IgG.
35. The composition of claim 34, wherein the IgG is human IgG.
36. The composition of claim 35, wherein the human IgG is selected from IgGl, IgG2,
IgG3, and IgG4.
37. The composition of any one of claims 33 to 36, wherein the antibody is formed through a knob-in-hole interaction in the Fc region.
38. The composition of claim 34 or claim 36, wherein the human IgG Fc comprises one or mutations to promote knob-in-hole interaction.
39. The composition of claim 38, wherein the mutations are selected from (i) T366Y or T366W, and (ii) Y407T, Y407A, or Y407V.
40. The composition of claim 38, wherein the mutations are selected from:
(a) T366Y and Y407T or T366Y/F405A and T394W/Y407T in human
IgGl, and
(b) T366W/D399C and T366S/L368A/K392C/Y407V;
T366W/K392C and T366S/L368A/D399C/Y407V;
S354C/T366W and Y349C/T366S/L368A/Y407V;
Y349C/T366W and S354C/T366S/L368A/Y407V;
E356C/T366W and Y349C/T366S/L368A/Y407V;
Y349C/T366W and E356C/T366S/L368A/Y407V;
E357C/T366W and Y349C/T366S/L368A/Y407V; or
Y349C/T366W and E357C/T366S/L368A/Y407V, in human IgGl.
41. The composition of claim 35, wherein the human IgG Fc comprises one or mutations to reduce or eliminate the effector function of the Fc domains.
42. The composition of claim 41, wherein the mutations are L234A and L235A (LALA) substitutions in human IgGl.
43. The composition of claim 35, wherein the human IgG Fc comprises one or mutations to stabilize a hinge region in the Fc domain.
44. The composition of claim 43, wherein the mutation is S228P.
45. A composition comprising a tandem single-chain variable fragment (scFv) specific for
Claudin 18.2 and CD3 comprising one or more of:
(a) an amino acid sequence of:
MKWVTFISLLFLFSSAYSSYVLTQPPSVSVAPGKTARITCSGGYNGHYGWYQ
QKPGQAPVLVIYSNNQRPSGIPERFSGSNSGNTATLTISRVEAGDEADYYCGG
YDSSAGIFGGGTKLTVLGGGGSGGGGSGGGGSEVQLLESGGGLVQPGGSLRL
SCAASGFTFRSYAMMWVRQAPGKGLEWVGEISSTGGSTYYGSAVKGRATIS
RDNSKNTLYLQMNSLRAEDTAVYYCAKSTYGATWKSADSIDAWGQGTLVT
VSSGGGGSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAMNWVRQAPGK
GLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQMNNLKTEDTA VYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGGSGGGGSQTVV TQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAPRGLIGGTKFL APGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFGGGTKLT VL (SEQ ID: 4816), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or a variant thereof, the CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or a variant thereof, and the CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFRSYAMMWV (SEQ ID NO: 22) or a variant thereof, the CDR2 sequence is VGEISSTGGSTYYG (SEQ ID NO: 23) or a variant thereof, and the CDR3 sequence is CAKSTYGATWKSADSIDA (SEQ ID NO: 24) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9) or a variant thereof, or (b) an amino acid sequence of:
MKWVTFISLLFLFSSAYSSYELTQPPSVSVSPGQTARITCSGGYNGHYGWYQQ KPGQAPVTVIYSNNQRPSGIPERFSGSSSGTTVTLTISGVQAEDEADYYCGGY DSSAGIFGGGTKLTVLGGGGSGGGGSGGGGSEVQLLESGGGLVQPGGSLRLS CAASGFTFSSYGMGWVRQAPGKGLEWVAAITRDSSTNYGAAVKGRATISRD NSKNTLYLQMNSLRAEDTAVYYCAKDASNGYCWYTGRGSYSCGAHHIDAW GQGTLVTVSSGGGGSEVQLVESGGGLVQPGGSLKLSCAASGFTFNKYAMNW VRQAPGKGLEWVARIRSKYNNYATYYADSVKDRFTISRDDSKNTAYLQMNN LKTEDTAVYYCVRHGNFGNSYISYWAYWGQGTLVTVSSGGGGSGGGGSGG
GGSQTVVTQEPSLTVSPGGTVTLTCGSSTGAVTSGNYPNWVQQKPGQAPRGL IGGTKFLAPGTPARFSGSLLGGKAALTLSGVQPEDEAEYYCVLWYSNRWVFG GGTKLTVL (SEQ ID NO. 4814), or an amino acid sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity thereto, or a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is CSGGYNGHYG (SEQ ID NO: 34) or a variant thereof, the CDR2 sequence is IYSNNQRP (SEQ ID NO: 35) or a variant thereof, and the CDR3 sequence is CGGYDSSAGIF (SEQ ID NO: 27) or a variant thereof, and/or a variable region comprising CDR1 , CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFSSYGMGWV (SEQ ID NO: 1) or a variant thereof, the CDR2 sequence is VAAITRDSSTNYG (SEQ ID NO: 2) or a variant thereof, and the CDR3 sequence is CAKDASNGYCWYTGRGSYSCGAHHIDA (SEQ ID NO: 3) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is GFTFNKYAMNWV (SEQ ID NO: 4) or a variant thereof, the CDR2 sequence is VARIRSKYNNYATYYA (SEQ ID NO: 5) or a variant thereof, and the CDR3 sequence is CVRHGNFGNSYISYWAY (SEQ ID NO: 6) or a variant thereof, and/or a variable region comprising CDR1, CDR2, and CDR3 sequences, wherein the CDR1 sequence is CGSSTGAVTSGNYP (SEQ ID NO: 7) or a variant thereof, the CDR2 sequence is IGGTKFLAP (SEQ ID NO: 8) or a variant thereof, and the CDR3 sequence is CVLWYSNRWVF (SEQ ID NO: 9); optionally wherein the first or second heavy chain comprises a His6 tag (e.g., HHHHHH (SEQ ID NO: 46)) added to the N- or C -terminus; and/or optionally wherein the composition comprises a linker comprising one or more glycines and serines replaced with another peptide linker or functionally equivalent variation thereof.
46. The composition of any one of claims 1 to 45, wherein the composition binds Claudin 18.2 and CD3 contemporaneously.
47. The composition of any one of claims 1 to 46, wherein the composition binds Claudin
18.2 with an affinity of less than 10 nM and with at least 100-fold greater affinity than Claudin 9, Claudin 3, and/or Claudin 4.
48. The composition of any one of claims 1 to 45, wherein the first heavy chain, the second heavy chain, the light chain, and/or the single-chain variable fragment does not comprise: (a) amino acids 1-18 relative to the respective SEQ ID NO; and (b) the amino acid sequence MKWVTFISLLFLFSSAYS (SEQ ID NO: 5356).
49. A pharmaceutical composition comprising the antibody of any one of claims 1 to 48 or a nucleic acid molecule encoding the same.
50. The pharmaceutical composition of claim 49, wherein the composition is an injectable pharmaceutical composition.
51. The pharmaceutical composition of claim 49 or 50, wherein the pharmaceutical composition is sterile.
52. The pharmaceutical composition of any one of claims 49 to 51, wherein the pharmaceutical composition is pyrogen free.
53. A nucleic acid molecule encoding an antibody or an amino acid sequence of any of the preceding claims.
54. A vector comprising the nucleic acid molecule of claim 53.
55. A cell comprising the nucleic acid molecule of claim 53 or the vector of claim 54.
56. A method for modulating and/or targeting Claudin 18.2 and CD3 in a biological cell comprising contacting the cell with the composition or pharmaceutical composition of any one of claims 1 to 52.
57. A method for modulating Claudin 18.2 activity in a biological cell comprising contacting a cell expressing Claudin 18.2 with the composition or pharmaceutical composition of any one of claims 1 to 52.
58. A method for inhibiting the function of Claudin 18.2 in a biological cell comprising contacting a cell expressing Claudin 18.2 with the composition or pharmaceutical composition of any one of claims 1 to 52.
59. A method for treating or preventing cancer comprising administering an effective amount of the composition or pharmaceutical composition of any one of claims 1 to 52 to a subject in need thereof.
60. A use of the composition or pharmaceutical composition of any one of claims 1 to 52 for the preparation of a medicament for the treatment of prevention of cancer.
61. The method or use of claim 60, wherein the cancer is selected from one or more of basal cell carcinoma, biliary tract cancer; bladder cancer; bone cancer; brain and central nervous system cancer; breast cancer; cancer of the peritoneum; cervical cancer; choriocarcinoma; colon and rectum cancer; connective tissue cancer; cancer of the digestive
system; endometrial cancer; esophageal cancer; eye cancer; cancer of the head and neck; gastric cancer (including gastrointestinal cancer); glioblastoma; glioma; hepatic carcinoma; hepatoma; intra-epithelial neoplasm; kidney or renal cancer; larynx cancer; leukemia; liver cancer; lung cancer (e.g., small-cell lung cancer, non-small cell lung cancer, adenocarcinoma of the lung, and squamous carcinoma of the lung); melanoma; myeloma; neuroblastoma; oral cavity cancer (lip, tongue, mouth, and pharynx); ovarian cancer; pancreatic cancer; prostate cancer; retinoblastoma; rhabdomyosarcoma; malignant rhabdoid tumor; rectal cancer; cancer of the respiratory system; salivary gland carcinoma; sarcoma; skin cancer; squamous cell cancer; stomach cancer; testicular cancer; thyroid cancer; uterine or endometrial cancer; cancer of the urinary system; vulvar cancer; lymphoma including Hodgkin's and nonHodgkin's lymphoma, as well as B-cell lymphoma (including low grade/follicular nonHodgkin's lymphoma (NHL); small lymphocytic (SL) NHL; intermediate grade/follicular NHL; intermediate grade diffuse NHL; high grade immunoblastic NHL; high grade lymphoblastic NHL; high grade small non-cleaved cell NHL; bulky disease NHL; mantle cell lymphoma; AIDS-related lymphoma; and Waldenstrom's Macroglobulinemia; chronic lymphocytic leukemia (CLL); acute lymphoblastic leukemia (ALL); Hairy cell leukemia; chronic myeloblastic leukemia; as well as other carcinomas and sarcomas; and posttransplant lymphoproliferative disorder (PTLD), as well as abnormal vascular proliferation associated with phakomatoses; edema (e.g. that associated with brain tumors); and Meigs’ syndrome.
62. An isolated antibody comprising one or more of the sequences disclosed herein.
63. A method of treating or preventing cancer comprising administering an effective amount of one or more of the nucleic acid molecules of claim 53 to a subject in need thereof. 64. The method of claim 63, wherein the cancer is selected from one or more of basal cell carcinoma, biliary tract cancer; bladder cancer; bone cancer; brain and central nervous system cancer; breast cancer; cancer of the peritoneum; cervical cancer; choriocarcinoma; colon and rectum cancer; connective tissue cancer; cancer of the digestive system; endometrial cancer; esophageal cancer; eye cancer; cancer of the head and neck; gastric cancer (including gastrointestinal cancer); glioblastoma; glioma; hepatic carcinoma; hepatoma; intra-epithelial neoplasm; kidney or renal cancer; larynx cancer; leukemia; liver cancer; lung cancer (e.g., small-cell lung cancer, non-small cell lung cancer, adenocarcinoma of the lung, and squamous carcinoma of the lung); melanoma; myeloma; neuroblastoma; oral cavity cancer (lip, tongue, mouth, and pharynx); ovarian cancer; pancreatic cancer; prostate cancer; retinoblastoma; rhabdomyosarcoma; malignant rhabdoid tumor; rectal cancer; cancer of the
respiratory system; salivary gland carcinoma; sarcoma; skin cancer; squamous cell cancer; stomach cancer; testicular cancer; thyroid cancer; uterine or endometrial cancer; cancer of the urinary system; vulvar cancer; lymphoma including Hodgkin’s and non-Hodgkin’s lymphoma, as well as B-cell lymphoma (including low grade/follicular non-Hodgkin’s lymphoma (NHL); small lymphocytic (SL) NHL; intermediate grade/follicular NHL; intermediate grade diffuse NHL; high grade immunoblastic NHL; high grade lymphoblastic NHL; high grade small non-cleaved cell NHL; bulky disease NHL; mantle cell lymphoma; AIDS-related lymphoma; and Waldenstrom's Macroglobulinemia; chronic lymphocytic leukemia (CLL); acute lymphoblastic leukemia (ALL); Hairy cell leukemia; chronic myeloblastic leukemia; as well as other carcinomas and sarcomas; and post-transplant lymphoproliferative disorder (PTLD), as well as abnormal vascular proliferation associated with phakomatoses; edema (e.g. that associated with brain tumors); and Meigs’ syndrome. 65. The method of claim 63 or 64, wherein the nucleic acid molecule comprises a sequence having at least about 90%, about 93%, about 95%, about 97%, about 98%, or about 99% identity to any one of SEQ ID NOs: 5329-5355.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363606967P | 2023-12-06 | 2023-12-06 | |
| US63/606,967 | 2023-12-06 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025122927A1 true WO2025122927A1 (en) | 2025-06-12 |
Family
ID=95980476
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/058961 Pending WO2025122927A1 (en) | 2023-12-06 | 2024-12-06 | Antibodies directed to claudin 18.2, including bispecific formats thereof |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025122927A1 (en) |
Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20180044434A1 (en) * | 2015-03-06 | 2018-02-15 | Public University Corporation Yokohama City University | Novel anti-pad4 antibody |
| US20180142021A1 (en) * | 2015-04-17 | 2018-05-24 | Amgen Research (Munich) Gmbh | Bispecific antibody constructs for cdh3 and cd3 |
| US20190144565A1 (en) * | 2016-06-10 | 2019-05-16 | Ucb Biopharma Sprl | ANTI-IgE ANTIBODIES |
| US20220348659A1 (en) * | 2021-03-24 | 2022-11-03 | Twist Bioscience Corporation | Variant nucleic acid libraries for cd3 |
| WO2022266219A1 (en) * | 2021-06-15 | 2022-12-22 | Xencor, Inc. | Heterodimeric antibodies that bind claudin18.2 and cd3 |
-
2024
- 2024-12-06 WO PCT/US2024/058961 patent/WO2025122927A1/en active Pending
Patent Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20180044434A1 (en) * | 2015-03-06 | 2018-02-15 | Public University Corporation Yokohama City University | Novel anti-pad4 antibody |
| US20180142021A1 (en) * | 2015-04-17 | 2018-05-24 | Amgen Research (Munich) Gmbh | Bispecific antibody constructs for cdh3 and cd3 |
| US20190144565A1 (en) * | 2016-06-10 | 2019-05-16 | Ucb Biopharma Sprl | ANTI-IgE ANTIBODIES |
| US20220348659A1 (en) * | 2021-03-24 | 2022-11-03 | Twist Bioscience Corporation | Variant nucleic acid libraries for cd3 |
| WO2022266219A1 (en) * | 2021-06-15 | 2022-12-22 | Xencor, Inc. | Heterodimeric antibodies that bind claudin18.2 and cd3 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP7425604B2 (en) | Anti-CTLA4-anti-PD-1 bifunctional antibodies, pharmaceutical compositions and uses thereof | |
| US9574012B2 (en) | AGR2 blocking antibody and use thereof | |
| US20220372112A1 (en) | Tumor therapeutic agent and use thereof | |
| CN111808194B (en) | A humanized antibody that binds claudin for the treatment of cancer | |
| NZ796440A (en) | Anti-TGF-beta antibodies and their use | |
| CA3146341A1 (en) | Bispecific anti lrrc15 and cd3epsilun antibudies | |
| CN114751984B (en) | Monoclonal antibody of targeted human Claudin18.2 protein and application thereof | |
| JP6731933B2 (en) | PCSK9 antibody, pharmaceutical composition and use thereof | |
| JP2024536881A (en) | Anti-LAG3 Antibodies, Pharmaceutical Compositions, and Uses | |
| CN114409797A (en) | Recombinant antibodies and their applications | |
| CA2849746A1 (en) | Anti-icam-1 antibodies to treat multiple-myeloma related disorders | |
| WO2022194201A1 (en) | Cldn18.2-targeting antibody or antigen binding fragment thereof and use thereof | |
| EP4071170A1 (en) | Pharmaceutical composition, preparation method therefor and use thereof | |
| CN102307595A (en) | Antibody and method for treating estrogen receptor-related diseases | |
| WO2025122927A1 (en) | Antibodies directed to claudin 18.2, including bispecific formats thereof | |
| CN117624352A (en) | anti-Tmem 176b antibody, pharmaceutical composition and application | |
| JP7466930B2 (en) | Anti-CD137 Antibodies and Uses Thereof | |
| CN114805581A (en) | Antibodies targeting IL13RA2, chimeric antigen receptors, and uses thereof | |
| US12049502B2 (en) | Antibodies directed to claudin 6, including bispecific formats thereof | |
| EP4549466A1 (en) | Development and use of novel multispecific tumor inhibitor | |
| KR20260017526A (en) | anti-PTK7 chimeric antibodies and anti-PTK7 humanized antibodies and uses thereof | |
| WO2023077068A1 (en) | Novel binding agents related to glycoprotein-a | |
| TW202336032A (en) | Interleukin 2 mutants and complexes containing the same | |
| CN119256013A (en) | Antigen binding protein targeting CD26 and its drug application | |
| WO2024007129A1 (en) | Humanized b7h3 antibody or antigen-binding fragment thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24901650 Country of ref document: EP Kind code of ref document: A1 |