WO2025109176A1 - Optimized sarbecovirus spike s2 subunit binders and compositions comprising the same - Google Patents
Optimized sarbecovirus spike s2 subunit binders and compositions comprising the same Download PDFInfo
- Publication number
- WO2025109176A1 WO2025109176A1 PCT/EP2024/083304 EP2024083304W WO2025109176A1 WO 2025109176 A1 WO2025109176 A1 WO 2025109176A1 EP 2024083304 W EP2024083304 W EP 2024083304W WO 2025109176 A1 WO2025109176 A1 WO 2025109176A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- seq
- sars
- cov
- variant
- binding agent
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C07K16/104—
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K39/395—Antibodies; Immunoglobulins; Immune serum, e.g. antilymphocytic serum
- A61K39/39591—Stabilisation, fragmentation
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P31/00—Antiinfectives, i.e. antibiotics, antiseptics, chemotherapeutics
- A61P31/12—Antivirals
- A61P31/14—Antivirals for RNA viruses
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/505—Medicinal preparations containing antigens or antibodies comprising antibodies
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/20—Immunoglobulins specific features characterized by taxonomic origin
- C07K2317/22—Immunoglobulins specific features characterized by taxonomic origin from camelids, e.g. camel, llama or dromedary
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/20—Immunoglobulins specific features characterized by taxonomic origin
- C07K2317/24—Immunoglobulins specific features characterized by taxonomic origin containing regions, domains or residues from different species, e.g. chimeric, humanized or veneered
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/31—Immunoglobulins specific features characterized by aspects of specificity or valency multispecific
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/52—Constant or Fc region; Isotype
- C07K2317/526—CH3 domain
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/56—Immunoglobulins specific features characterized by immunoglobulin fragments variable (Fv) region, i.e. VH and/or VL
- C07K2317/565—Complementarity determining region [CDR]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/56—Immunoglobulins specific features characterized by immunoglobulin fragments variable (Fv) region, i.e. VH and/or VL
- C07K2317/569—Single domain, e.g. dAb, sdAb, VHH, VNAR or nanobody®
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/72—Increased effector function due to an Fc-modification
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/76—Antagonist effect on antigen, e.g. neutralization or inhibition of binding
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/92—Affinity (KD), association rate (Ka), dissociation rate (Kd) or EC50 value
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/94—Stability, e.g. half-life, pH, temperature or enzyme-resistance
Definitions
- the invention is broadly in the field of binding agents, in particular antibodies. More particularly, the invention pertains to binding agents, in particular antibodies and antigen-binding fragments thereof, binding to the spike protein of a Sarbecovirus , which are capable of potently neutralizing a Sarbecovirus such as SARS-CoV-2, including SARS-CoV-2 variants, and SARS-CoV-1.
- the invention also relates to methods using these binding agents and uses thereof.
- Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the causative agent of COVID- 19 (Zhu et al. 2020, N Engl J Med 382:727-733).
- SARS-CoV-2 infections can be asymptomatic or present with mild to moderately severe symptoms.
- COVID-19 progresses to a more severe stage that is characterized by dyspnoea and hypoxemia, which may progress further to acute respiratory distress requiring often long-term intensive care and causing death in a proportion of patients.
- “Long-COVID” furthermore refers to long-term effects of COVID-19 infection, even when no SARS-CoV-2 virus can be detected anymore.
- the spike of SARS coronaviruses is a major target for neutralizing antibodies.
- This spike protein is a class I fusion protein and is comprised of a membrane distal SI subunit and a membrane proximal S2 subunit.
- the S 1 subunit comprises the receptor-binding domain (RBD) and antibodies directed against this domain can have very strong neutralizing activity (Wheatley et al. 2021. Cell Rep 37: 109822).
- the SI subunit in particular the N-terminal domain and the RBD, can tolerate mutations that result in antigenic variation and immune escape.
- the RBD is also immunodominant (Piccoli et al. 2020.
- the S2 subunit is responsible for the membrane fusion, a process during which S2 undergoes major conformational changes (Dodero- Rojas et al. 2021. eLife 10:e70362).
- the S2 subunit is more conserved and therefore, at least in theory, appears to be an attractive target for the development of neutralizing antibodies with broad anti-Sarbecovirus protective potential.
- VHHs single-domain antibodies
- HR2 domain of the S2 subunit heptad repeat 2 (HR2) domain of the S2 subunit, more particularly within a C-terminal region of the HR2 domain proximal to the viral membrane.
- the antibodies do not only need to exhibit favourable antigenbinding and functional properties, but also be stable, safe, manufacturable, etc.
- nonlimiting factors that affect the potential of antibodies include chemical stability (resistance to e.g., fragmentation, deamidation, oxidation and isomerisation), biophysical stability (e.g., propensity to aggregate), solution properties (e.g., solubility, viscosity).
- Low risk of immunogenicity of an antibody is also a consideration.
- antibodies also need good pharmacokinetic and pharmacodynamic properties, such as slow in vivo clearance rate, long circulation half-life, etc. All these factors can collectively be referred to as development characteristics of an antibody. These are important considerations as these might impact the antibody’s cost of goods, safety profde, dosing schedule, mode of administration, etc. and thus their development potential.
- FIG. 1 Prophylactic treatment with R3_DC23-Fc protects K18-hACE2 mice from lethal SC2 infection.
- K18-hACE2 mice were intraperitoneally injected with 100 pg R3_DC23-Fc or Isotype control antibody (palivizumab) or were left untreated twenty hours prior to intratracheal infection with 3* 10 2 PFU of SARS-CoV-2 D614G variant virus. Animals were monitored on a daily basis by measuring weight change and scoring for humane endpoints.
- mice treated with R3_DC23-Fc displayed significantly lower bodyweight loss as compared to mice treated with palivizumab (p ⁇ 0.05) or untreated mice (p ⁇ 0.01) (mixed-effect analysis with Sidak's multiple comparisons test).
- the graph shows the Kaplan-Meier curve of animal survival portion of the indicated groups. Euthanasia was performed when mice lost more than 25% of their bodyweight as defined on day 0 or when a high score for humane endpoints was reached.
- FIG. 2 LS mutants of humanized R3_DC23-Fc fusions control viral replication in hamsters.
- A-B Treatment of SARS-CoV-2 Wuhan infection by huR3DC23-Fc_LS in Syrian Golden hamsters. Male Syrian Golden hamsters were intranasally infected with SARS-CoV-2 (Wuhan strain) on day 0 and received intraperitoneal treatment with either 10 or 2 mg/kg huR3DC23-Fc_LS, 10 mg/kg Bebtelovimab (positive control) or 10 mg/kg Palivizumab (negative control) 4 hours postinfection.
- FIG. 3 Prophylactic treatment with R3_DC23-Fc protects K18-hACE2 mice against lethal SARS-CoV-2 infection. Twenty hours prior to intratracheal infection with 3* 10 2 PFU of SARS- CoV-2 D614G variant virus, 100 pg R3_DC23-Fc was administered to K18-hACE2 mice and 100 pg of isotype control antibody (palivizumab) was administered to a second group of K18-hACE2 mice and non-permissive wild-type (WT) mice. Animals were monitored on a daily basis by measuring weight change and scoring for humane endpoints. (A) The graph shows the Kaplan- Meier curve of animal survival portion of the indicated groups.
- FIG. 4 Prophylactic treatment with R3_DC23-Fc reduces viral replication of SARS-CoV-2 in the lungs of infected K18-hACE2 mice. Twenty hours prior to intratracheal infection with 3* 10 2 PFU of SARS-CoV-2 D614G variant virus, 100 pg R3_DC23-Fc was administered to K18- hACE2 mice and 100 pg of isotype control antibody (palivizumab) was administered to a second group of K18-hACE2 mice and non-permissive wild-type (WT) mice. Animals were monitored on a daily basis by measuring weight change and scoring for humane endpoints.
- isotype control antibody palivizumab
- FIG. 5 (A) Sequence alignment of the humanized version of the VHH R3 DC23 and modified versions thereof. Sequences of hR3_DC23 and variants are shown: VHH hR3_DC23 (SEQ ID NO:2), VHH RNGL (SEQ ID NO:3), VHH RAGL (SEQ ID NO:4), VHH E2NGL (SEQ ID NO:5), VHH E2AGL (SEQ ID NO:6), VHH QRAGL (SEQ ID NO:8), VHH ERAGL (SEQ ID NO: 10), VHH E10RAGL (SEQ ID NO: 12), VHH E81RAGL (SEQ ID NO: 14), VHH Q75 (SEQ ID NO: 15), VHH E75 (SEQ ID NO: 16), VHH E10E81 (SEQ ID NO: 18), VHH D16 (SEQ ID NO: 19), VHH T19D79 (SEQ ID NO:20), VHH TQD (SEQ ID NO:21), VHH Q75R21Y (
- VHH-Fc multivalent antibodies as described herein consist of two copies of a VHH building block each connected at its C-terminus to an Fc domain as described herein, in particular a human IgGl Fc without (e.g. SEQ ID NO: 82) or with mutations for half-life extension (e.g. SEQ ID NO: 83-86), via a (648)2 linker (SEQ ID NO: 88) at the Fc N-terminus.
- FIG. 6 VHH hR3_DC23 amino acid sequence and illustration of the different CDR annotations as used herein. CDR annotations according to AbM, Chothia, Martin, Kabat, IMGT and MacCallum in grey labeled boxes corresponding to the sequences of VHH hR3_DC23 (SEQ ID NO:2).
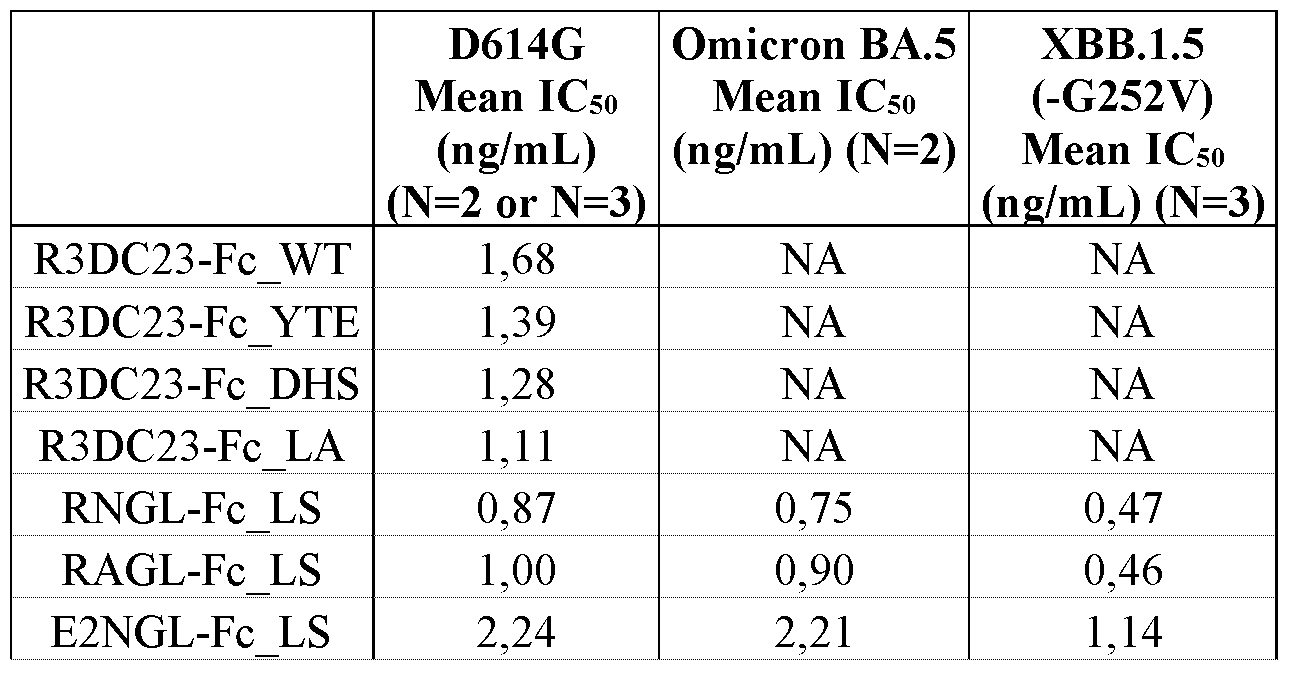
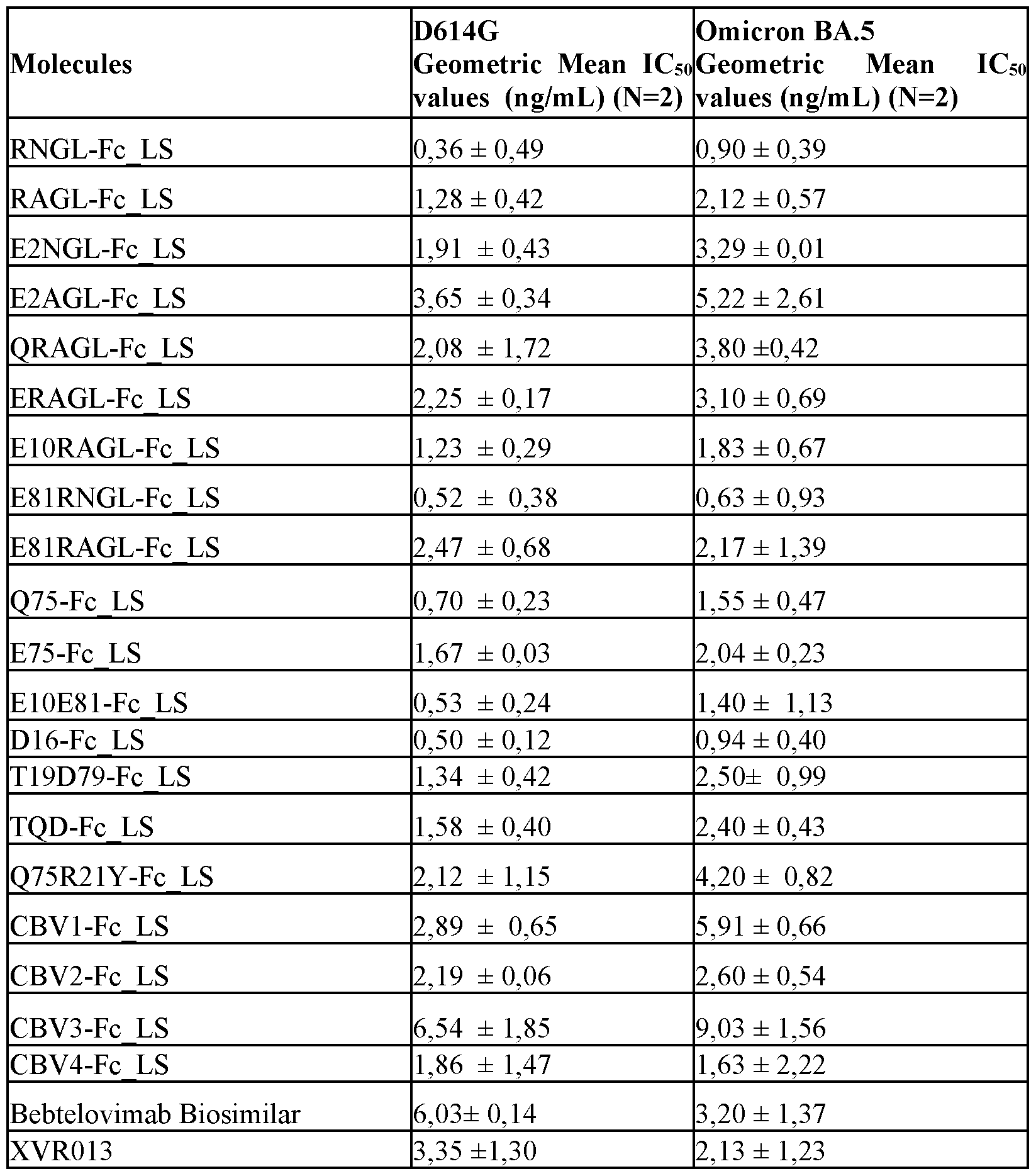
- FIG. 7 S2 targeting molecules neutralize VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 spike proteins.
- Vero E6 cells were transduced with VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 D614G, Omicron BA.5 or XBB.1.5 (-G252V) variant spike protein that had been pre-incubated with different concentrations of the tested S2 targeting molecules or the control antibodies. Sixteen hours later, the GFP fluorescence was measured with a fluorimeter.
- the graphs show the mean GFP fluorescence intensity of the tested molecules for D614G, Omicron BA.5 and XBB1.5 (-G252V) normalized to the GFP fluorescence intensity value of non-infected control cells and of infected cells that were not treated, which were both included in each dilution series.
- the mean IC50 values were calculated by nonlinear regression curve fitting, (log(inhibitor) versus normalized response (four parameters)). Mean values ⁇ SD are shown.
- FIG. 8 Flow cytometric analysis of the binding of the S2 targeting molecules to mammalian (HEK293) cells.
- HEK293 cells were incubated with the indicated S2 targeting molecules or controls (Sotrovimab Biosimilar and XVR013) at 10 or 100 pg/ml for 90 min.
- the graph shows the median fluorescence intensity (MFI) of the AF633-conjugated anti-human IgG to detect binding of the S2 targeting molecules and controls to the HEK293 cells.
- MFI median fluorescence intensity
- FIG. 9 In silica immunogenicity assessment of S2 targeting molecules. An in silica immunogenicity assessment was conducted for the S2 targeting molecules as indicated as described in the examples. (A) Calculated DRB1 scores for the tested S2 targeting molecules and XVR013. (B) The calculated DRB1 scores of the tested S2 targeting molecules and XVR013 were compared with the DRB1 scores of 80 approved antibodies. The variants in each “bin” of the histogram are considered similar in terms of immunogenic risk.
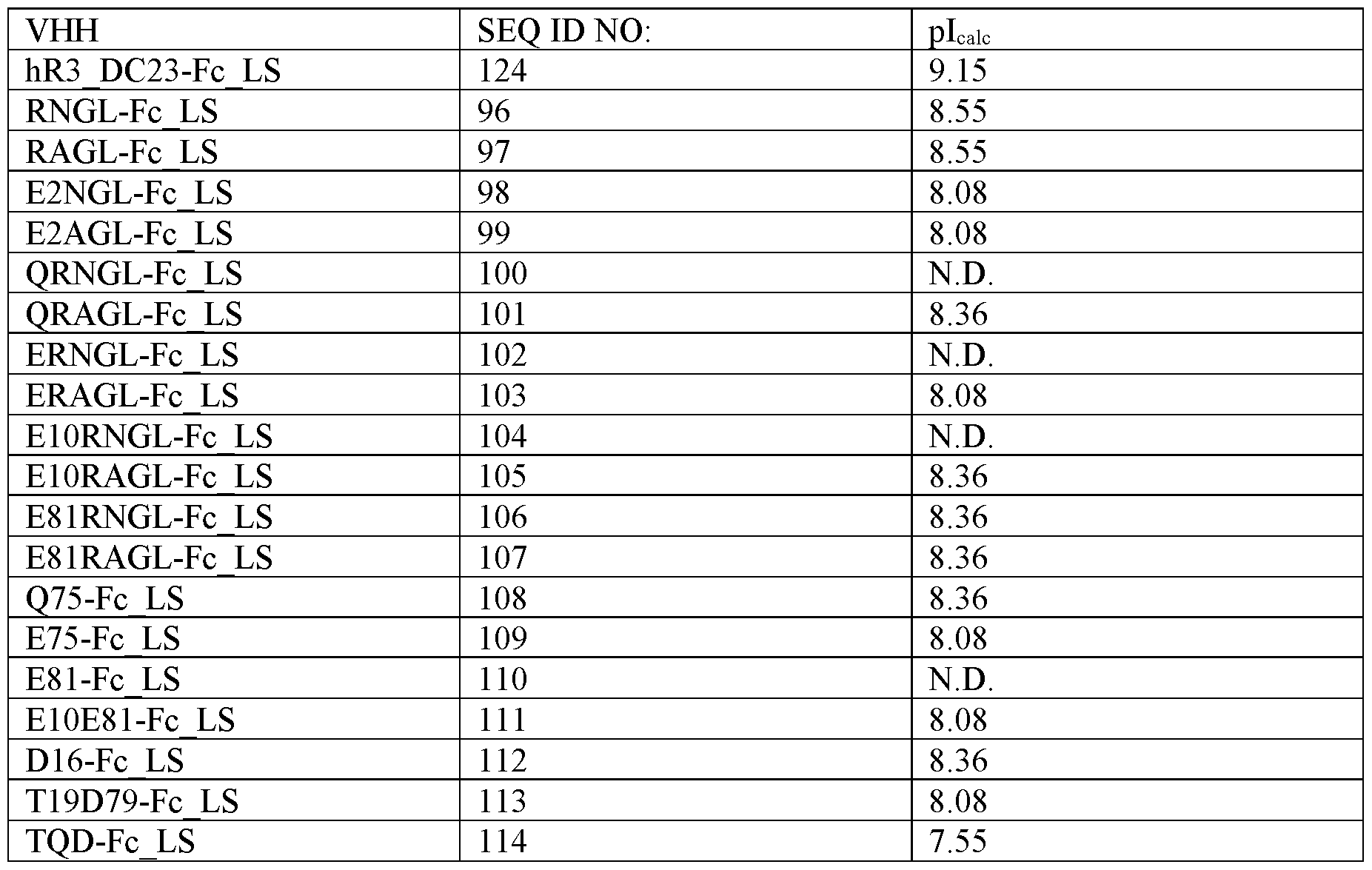
- FIG. 10 Apparent hydrophobicity of the S2 targeting molecules. Retention time of XVR013 and XVR011 (control for hydrophobicity) is indicated for comparison purposes. Note that variants CBV2-Fc_LS (SEQ ID NO: 117) and CBV4-Fc_LS (SEQ ID NO: 119) were analyzed separately: both molecules have lower pl compared to the other tested variants and the pH of the mobile phase used for the HIC assay needed to be adapted accordingly.
- FIG. 11 Strong Cation Exchange (SCX) chromatograms of the S2 targeting molecules and
- FIG. 12. (A) General initial characteristics (top half) and biophysics upon accelerated stress (bottom half) of the indicated S2 targeting molecules, analyzed and classified in color-code categories according to the legend (B).
- FIG. 13 Plasma concentrations of S2 targeting molecules and XVR013.
- An in vivo pharmacokinetic study was performed in human neonatal Fc receptor (hFcRn) tg32 SCID mice for the S2 targeting molecules E2NGL-Fc_LS (SEQ ID NO:98), E75-Fc_LS (SEQ ID NO: 109), CBV2- Fc_LS (SEQ ID NO: 117), CBV3-Fc_LS (SEQ ID NO: 118) and CBV4-Fc_LS (SEQ ID NO: 119), and XVR013.
- Plasma concentrations (Test Article) of the S2 targeting molecules and XVR013 are shown.
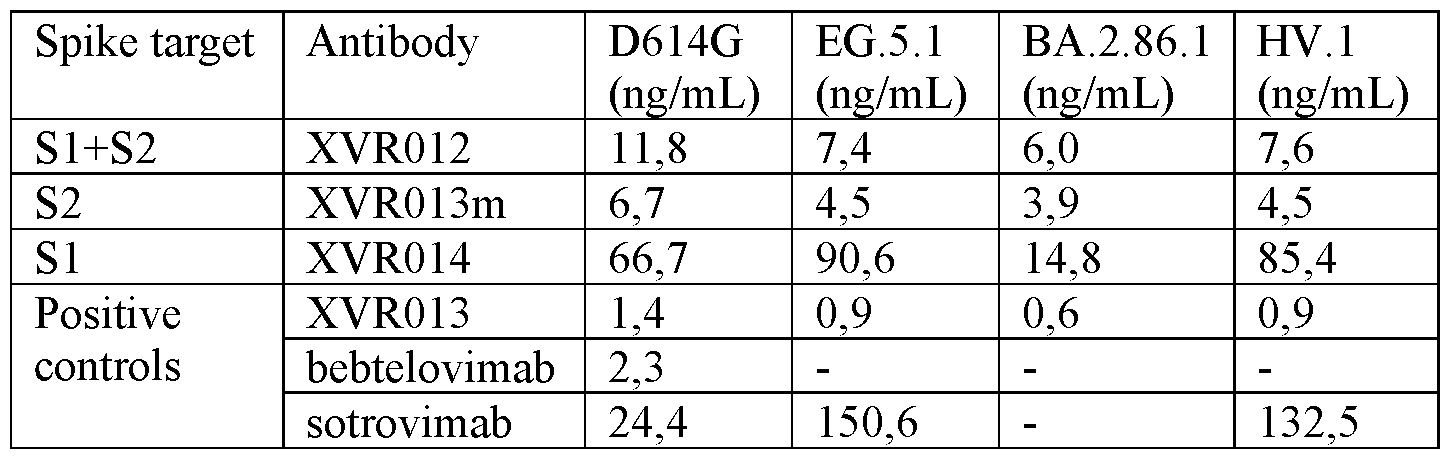
- FIG. 14 Composition of SI and S2 targeting binding agents (XVR012) neutralizes VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 spike proteins.
- Vero E6 cells were transduced with VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 D614G spike protein or with the spike protein of the SARS-CoV-2 variants EG.5.1, BA.2.86.1 and HV.l, which viruses had been preincubated with different concentrations of the XVR012 composition, the constructs XVR013m (CBV3-Fc_LS: SEQ ID NO: 118) or XVR014 (SEQ ID NO: 126), or a control (XVR013, sotrovimab, bebtelovimab or palivizumab).
- the GFP fluorescence was measured with a fluorimeter.
- the mean IC50 values were calculated by nonlinear regression curve fitting, log(inhibitor) versus normalized response (four parameters).
- FIG. 15 Composition of SI and S2 targeting binding agents (XVR012) neutralizes VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 spike proteins.
- Vero E6 cells were transduced with VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 D614G spike protein or with the spike protein of the SARS-CoV-2 variants Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3, Omicron XBB.1.5, Omicron XBB.2.3, Omicron XBB.1.16, Omicron CH.1.1, Omicron EG.5.1, Omicron HK.3, Omicron HV. 1, Omicron BA.2.74, Omicron BA.2.86.1, Omicron FL. 1.5.1, Omicron JN.
- FIG. 16 In vivo efficacy in Syrian Golden Hamster SARS-CoV2 challenge model.
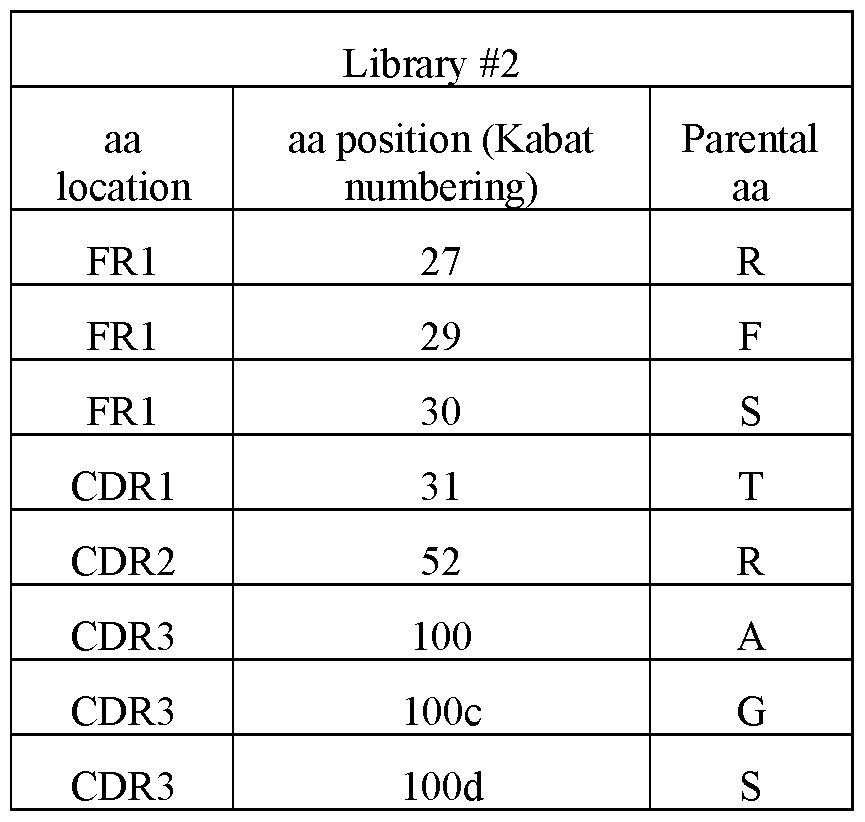
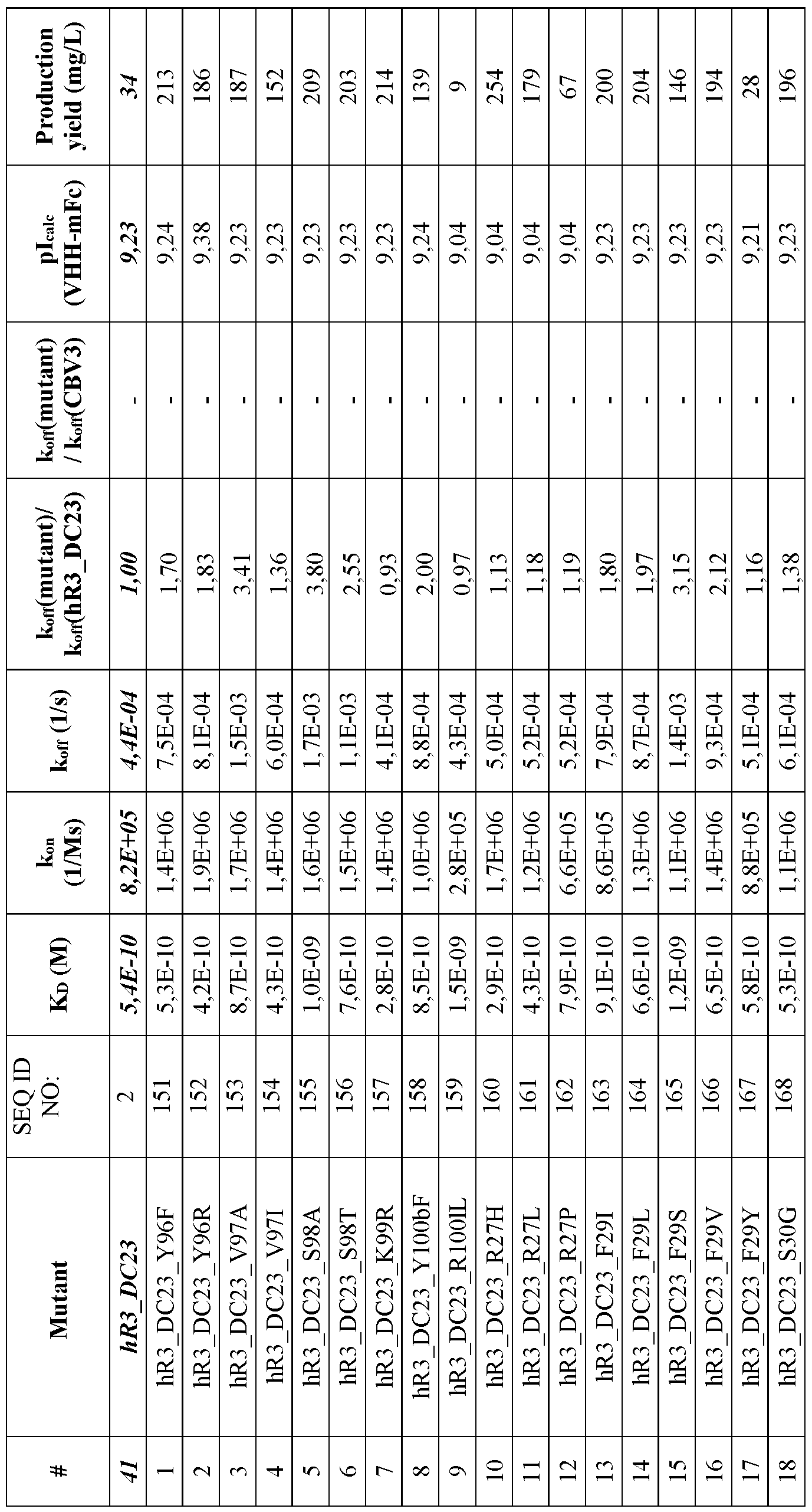
- FIG. 17 Sequence alignments of hR3_DC23 variants (SEQ ID NO:151-182) with hR3_DC23 (SEQ ID NO:2). Numbering ofthe sequences is according to Kabat. CDR1 (positions 31-35), CDR2 (positions 50-65) and CDR3 (positions 95-102) according to Kabat are indicated. Mutations vis-a- vis hR3_DC23 are indicated with grey background. Residues which are believed to constitute the paratope are underlined.
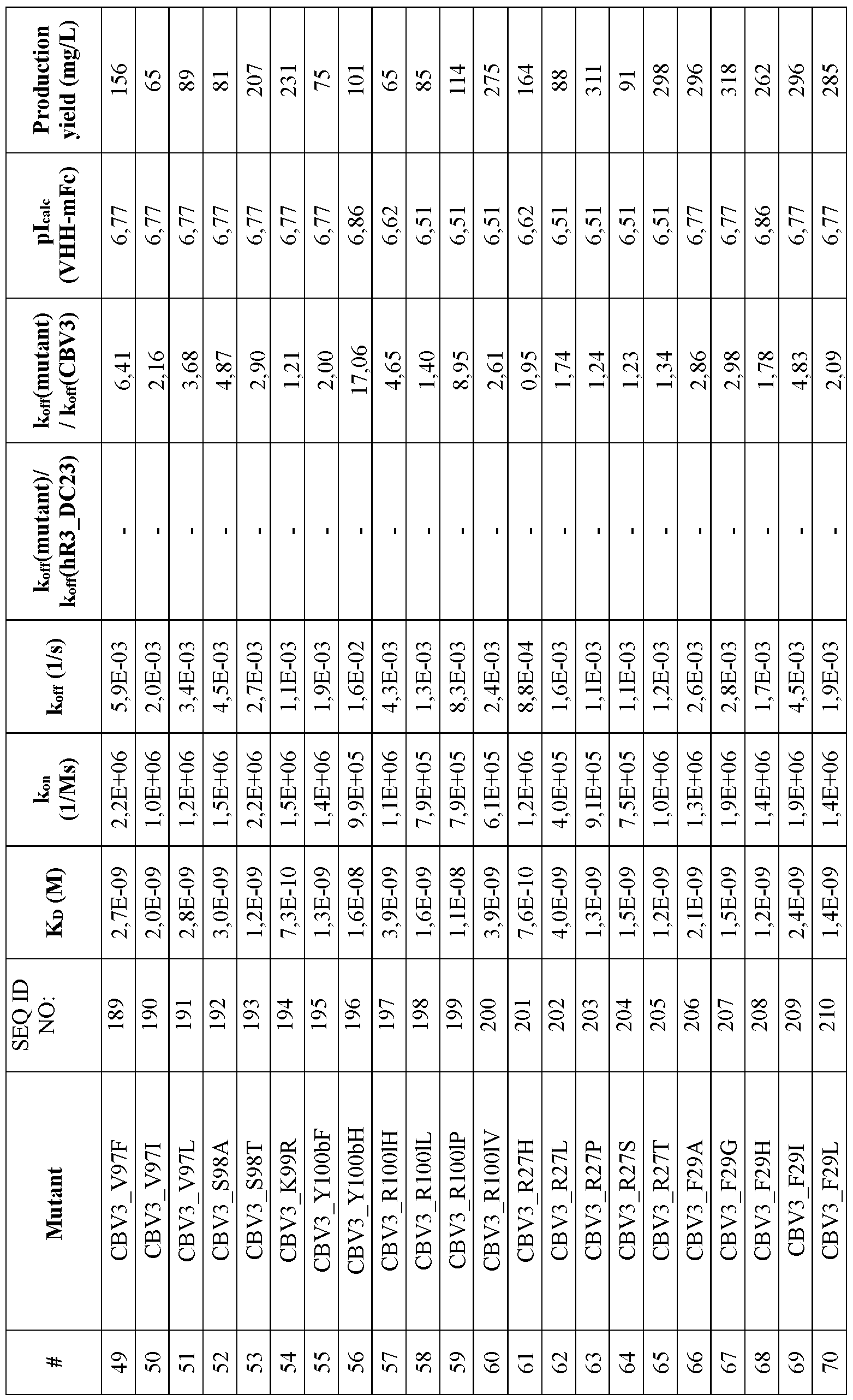
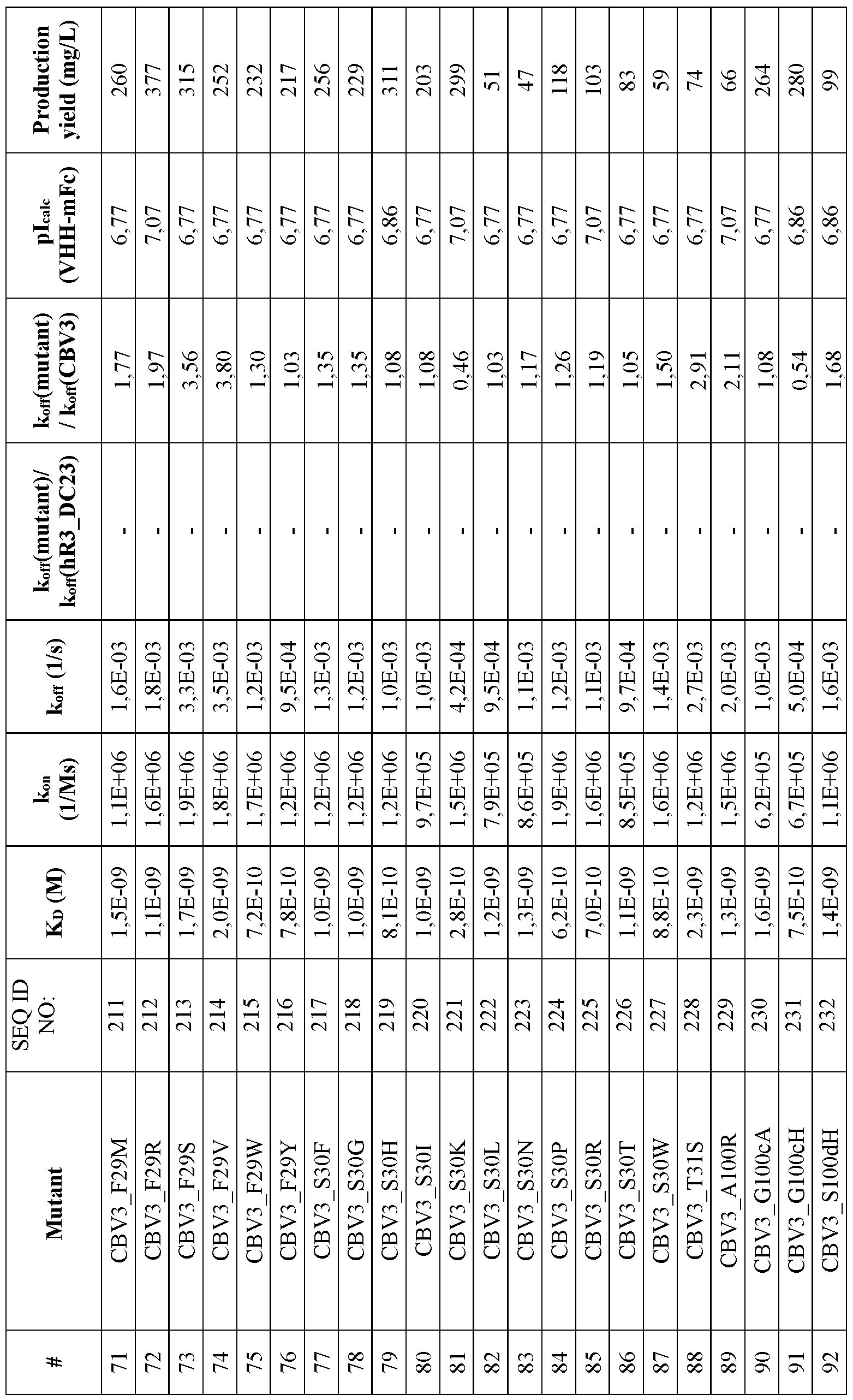
- FIG. 18 Sequence alignments of CBV3 (SEQ ID NO:25) and CBV3 variants (SEQ ID NO:183- 244) with hR3_DC23 (SEQ ID NO:2). Numbering ofthe sequences is according to Kabat. CDR1 (positions 31-35), CDR2(positions 50-65) and CDR3 (positions 95-102) according to Kabat are indicated. Mutations vis-a-vis hR3_DC23 are indicated with grey background. Residues which are believed to constitute the paratope are underlined.
- FIG. 19 Binding of S2 targeting molecules to the spike of Khosta-2 virus, a member of clade-3 of the Sarbecovirus subgenus. Binding of XVR013, XVR013m, XVR014, positive control (sotrovimab) or negative controls (palivizumab, rituximab or an isotype control) to HEK293T cells transiently transfected with expression vectors for GFP fused to the spike of (A) Khosta-2 virus or
- VHHs Sa rhecovn'us -specifc Variable Domains of Heavychain Antibodies
- R3 DC23 and hR3_DC23 said variants exhibiting one or more favourable antibody development characteristics, including, but not limited to, lower pl, reduced non-specific binding, reduced immunogenicity, improved electrostatics (less positive charge patches), increased stability, reduced hydrophobicity, improved pharmacokinetics (PK), or combinations thereof.
- SARS-CoV- 2 variants such as SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta, SARS-CoV-2 Epsilon B. 1.427 variant, SARS-CoV-2 Epsilon B.1.429 variant, SARS-CoV-2 Eta variant, SARS-CoV-2 Iota variant, SARS- CoV-2 Kappa variant, SARS-CoV-2 Mu variant, SARS-CoV-2 Zeta variant, SARS-CoV-2 B. 1.617.3 variant, SARS-CoV-2 Omicron BA.
- SARS-CoV- 2 variants such as SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta, SARS-CoV-2 Epsilon B. 1.427 variant, SARS-CoV
- SARS-CoV-2 Omicron BA.2 variant SARS-CoV-2 Omicron BA.5 variant
- SARS-CoV-2 Omicron XBB.1.5 (-G252V) variant SARS- CoV-2 Omicron XBB.1.5 variant
- SARS-CoV-2 Omicron XBB.2.3 variant SARS-CoV-2 Omicron XBB.1.16 variant
- SARS-CoV-2 Omicron EG.5.1 variant SARS-CoV-2 Omicron
- SARS-CoV-2 Omicron HK.3 variant SARS-CoV-2 Omicron BA.2.74 variant
- SARS-CoV-2 Omicron BA.2.86.1 variant SARS-CoV-2 Omicron HV.
- hR3_DC23 variants thus have advantageously improved antibody development characteristics while maintaining the functional properties such as neutralization of SARS-CoV-2 of hR3_DC23.
- the invention relates to a binding agent capable of neutralizing a Sarbecovirus , characterized in that said binding agent comprises at least one ISVD as described herein, in particular an ISVD comprising a complementarity determining region 1 (CDR1) defined by SEQ ID NO:40, a CDR2 defined by SEQ ID NO:30 and a CDR3 defined by SEQ ID NO:41; or a CDR1 defined by SEQ ID NO:38, a CDR2 defined by SEQ ID NO:39 and a CDR3 defined by SEQ ID NO:41; or an ISVD comprising a CDR1, CDR2 and CDR3, each independently as present in any of SEQ ID NOs: 2 to 26, wherein the CDR1, CDR2 and CDR3 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
- CDR1 complementarity determining region 1
- SEQ ID NO:40 a complementarity determining region 1
- CDR2 defined by
- the invention also relates to affinity matured variants of these binding agents.
- the invention relates to a nucleic acid molecule comprising a polynucleotide sequence encoding the binding agent according to the invention, as well as to a vector comprising such nucleic acid molecule; and a cell comprising such nucleic acid molecule or such vector or a cell expressing the binding agent according to the invention.
- the invention further relates to a pharmaceutical composition comprising the binding agent according to the invention, or the nucleic acid molecule or the vector as described hereinabove; and a pharmaceutically acceptable carrier; as well as to a kit such as a diagnostic kit comprising the binding agent according to the invention.
- a further aspect is directed to the binding agent according to the invention, the nucleic acid molecule or the vector as described hereinabove, the pharmaceutical composition or the kit as described hereinabove for use in medicine such as use in the prevention or treatment of a Sarbecovirus infection in a subject or for use in the diagnosis of a Sarbecovirus infection in a subject.
- the invention further relates to an in vitro or ex vivo method for detecting a Sarbecovirus in a sample, said method comprising:
- one or more or “at least one”, such as one or more members or at least one member of a group of members, is clear per se, by means of further exemplification, the term encompasses inter alia a reference to any one of said members, or to any two or more of said members, such as, e.g., any >3, >4, >5, >6 or >7 etc. of said members, and up to all said members.
- “one or more” or “at least one” may refer to 1, 2, 3, 4, 5, 6, 7 or more.
- the inventors identified novel VHHs that potently neutralize SARS-CoV-2, including SARS-CoV-2 variants such as SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta variant, SARS-CoV- 2 Epsilon B. 1.427 variant, SARS-CoV-2 Epsilon B.
- SARS-CoV-2 variants such as SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta variant, SARS-CoV- 2 Epsilon B.
- SARS-CoV-2 Omicron BA.2 variant SARS-CoV-2 Omicron BA.5 variant
- SARS-CoV-2 Omicron XBB.1.5 (- G252V) variant SARS-CoV-2 Omicron XBB.1.5 variant
- SARS-CoV-2 Omicron XBB.2.3 variant SARS-CoV-2 Omicron XBB.1.16 variant
- SARS-CoV-2 Omicron EG.5.1 variant SARS-CoV-2 Omicron
- SARS-CoV-2 Omicron HK.3 variant SARS-CoV-2 Omicron BA.2.74 variant
- SARS-CoV-2 Omicron BA.2.86.1 variant SARS-CoV-2 Omicron HV.
- binding agents capable of neutralizing a Sarbecovirus , characterized in that said binding agent comprises at least one anti-SARS-CoV-2 spike S2 subunit ISVD having the same or a similar paratope as hR3_DC23 (SEQ ID NO:2).
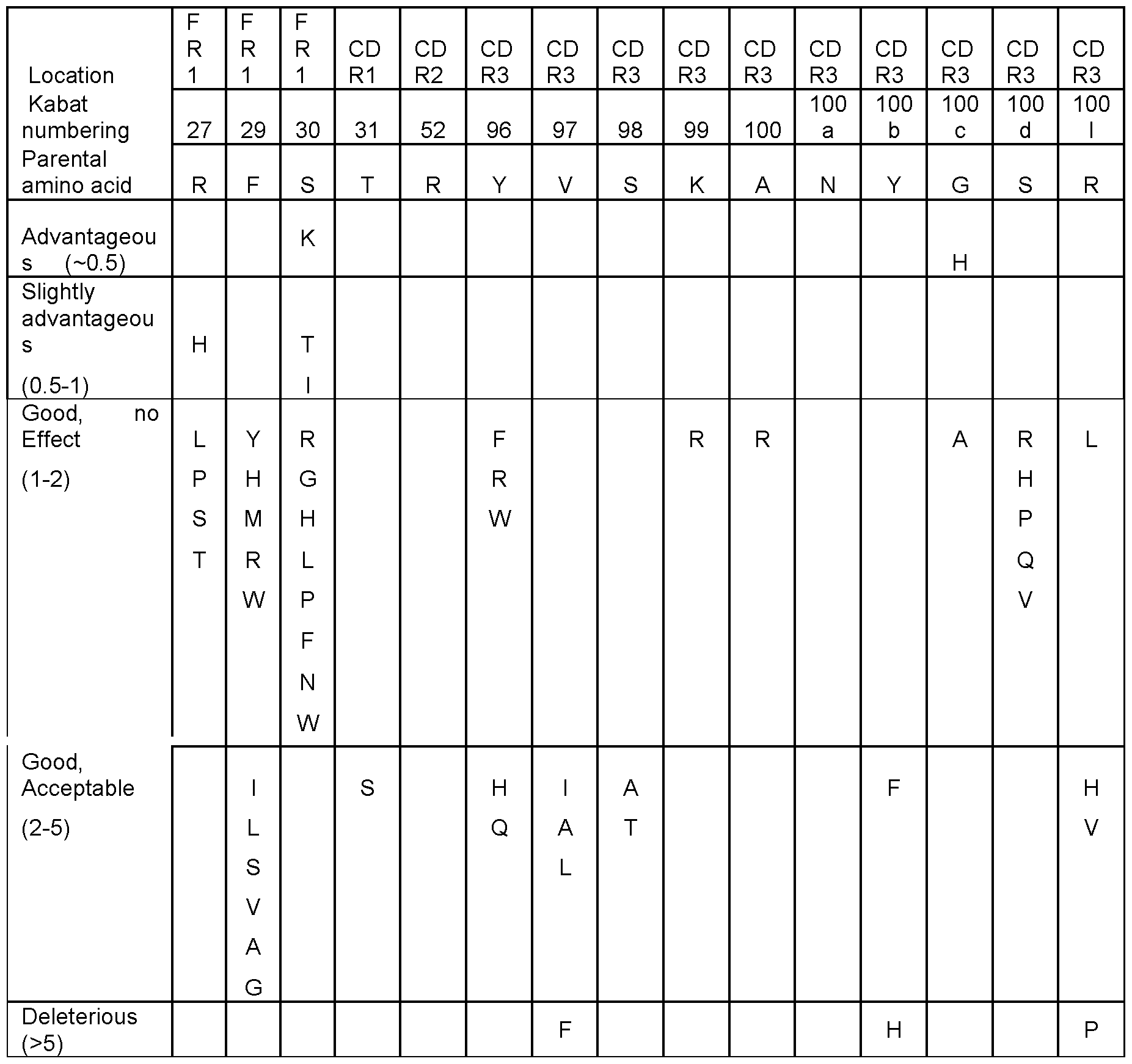
- the anti-SARS-CoV-2 spike S2 subunit ISVD has a paratope comprising or consisting of amino acid residues at positions 96, 97, 98, 99, 100a, 100b and 1001 according to Kabat numbering and/or at positions 27, 29, 30, 31, 52, 100, 100c and lOOd according to Kabat numbering, wherein each independently: the amino acid residue at position 96 is selected from the group consisting of: Y, F, R, W, H and Q; or, with increasing preference, is selected from the group consisting of: Y, F, R and W; or is Y; the amino acid residue at position 97 is selected from the group consisting of: V, I, A, L and F; or, with increasing preference, is selected from the group consisting of: V, I, A and L; or is V; the amino acid residue at position 98 is selected from the group consisting of: S, A and T; or, preferably, is S; the amino acid residue at position 99 is selected from the group consisting
- L, S, V, A and G is selected from the group consisting of: F, Y, H,
- the amino acid residue at position 30 is selected from the group consisting of: S, K, T, I, R, G, H, L, P, F, N and W; or, with increasing preference, is selected from the group consisting of: S, K, T and I; is selected from the group consisting of: K, T and I; or is K; the amino acid residue at position 31 is selected from the group consisting of: T and S; or, preferably is T; the amino acid residue at position 52 is R; the amino acid residue at position 100 is selected from the group consisting of: A and R; or, preferably, is A; the amino acid residue at position 100c is selected from the group consisting of: G, H and A; or, with increasing preference, selected from the group consisting of: G and H; or is H; and the amino acid residue at position lOOd is selected from the group consisting of: S, R, H, P, Q and V; or, preferably, is S.
- the anti-SARS-CoV-2 spike S2 subunit ISVD has a paratope as present in any one of the ISVDs of SEQ ID NO: 151 to 244, 2 or 25, optionally wherein the paratope comprises or consists of the amino acid residue at positions 96, 97, 98, 99, 100a, 100b and 1001 according to Kabat numbering and/or at positions 27, 29, 30, 31, 52, 100, 100c and lOOd according to Kabat numbering of said ISVD.
- the invention relates to a binding agent capable of neutralizing a Sarbecovirus , characterized in that said binding agent comprises at least one ISVD, wherein said ISVD comprises a CDR1 defined by SEQ ID NO:40, a CDR2 defined by SEQ ID NO:30 and a CDR3 defined by SEQ ID NO:41; or a CDR1 defined by SEQ ID NO:38, a CDR2 defined by SEQ ID NO:39 and a CDR3 defined by SEQ ID NO:41; or wherein said ISVD comprises a CDR1, CDR2 and CDR3, each independently as present in any of SEQ ID NOs:2 to 26, wherein the CDR1, CDR2 and CDR3 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
- a “binding agent” generally relates to a molecule that is capable of binding to at least one other molecule, wherein said binding is preferably a specific binding, such as on a defined binding site, pocket or epitope.
- a binding agent may be of any nature or type and is not dependent on its origin.
- a binding agent may be chemically synthesized, naturally occurring, recombinantly produced (and optionally purified), as well as designed and synthetically produced (and optionally purified).
- Said binding agent may hence be, e.g., a small molecule, a chemical, a peptide, a polypeptide, an antibody, or any derivative of any thereof, such as a peptidomimetic, an antibody mimetic, an active fragment, a chemical derivative, among others.
- a functional fragment of a binding agent or a functional part of a binding agent refers to a fragment or part of that binding agent that is functionally equivalent to that binding agent.
- such functional fragment or part of a binding agent as described herein ideally retains one or more of the functional features (1) to (88) of that binding agent as outlined extensively elsewhere herein.
- binding agents described herein are structurally defined as polypeptidic binding agents (i.e. binding agents comprising a peptidic, polypeptidic or proteic moiety, or binding agents comprising a peptide, polypeptide, protein or protein domain) or polypeptide binding agents (i.e. binding agents being peptides, polypeptides or proteins).
- protein polypeptide
- polypeptide are interchangeably used herein to refer to a polymer of amino acid residues and to variants and synthetic analogues of the same; the sequential linear arrangement of the amino acids together resulting in/forming the “amino acid sequence” or “protein sequence”.
- a “peptide” may also be referred to as a partial amino acid sequence derived from its original protein, for instance after enzymatic (e.g. tryptic) digestion. These terms apply to naturally-occurring amino acid polymers as well as to amino acid polymers in which one or more amino acid residues is a synthetic non-naturally occurring amino acid, such as a chemical analogue of a corresponding naturally occurring amino acid.
- proteins comprising one or more posttranslational modifications such as covalent addition of functional groups or proteins (such as glycosylation, phosphorylation, acetylation, ubiquitination, methylation, lipidation and nitrosylation) or such as proteolytic processing.
- functional groups or proteins such as glycosylation, phosphorylation, acetylation, ubiquitination, methylation, lipidation and nitrosylation
- proteolytic processing Based on the amino acid sequence and the modifications, the atomic or molecular mass or weight of a polypeptide is expressed in (kilo)dalton (kDa).
- a further modification of proteins includes addition of a tag, such as a His-tag or sortag.
- a “protein domain” is a distinct functional and/or structural unit in or part of a protein. Usually, a protein domain is responsible for a particular function or interaction, contributing to the overall (biological) role of a protein. Domains may exist in a variety of biological contexts, where similar domains can be found in different proteins with similar or different functions. Protein domains can have a rigid 3D- structure if confined by e.g. a number of intramolecular cysteines (e.g. cysteine- knot proteins) or can, depending on e.g. presence or absence of a bound ligand or e.g. presence or absence of a posttranslational modification, assume different 3D-conformations, or can have a less defined, more fluid 3D-structure.
- cysteines e.g. cysteine- knot proteins
- polypeptidic or polypeptide binding agents described herein are (optionally purified) antibodies or antibody fragments.
- antibody refers to an immunoglobulin (Ig) molecule or a molecule comprising an immunoglobulin (Ig) domain, which specifically binds with an antigen, as well as multimers thereof.
- Antibodies can be intact immunoglobulinsor immunoreactive portions of intact immunoglobulins. The term encompasses naturally, recombinantly, semi-synthetically or synthetically produced antibodies.
- an antibody can be present in or isolated from nature, e.g., produced or expressed natively or endogenously by a cell or tissue and optionally isolated therefrom; or an antibody can be recombinant, i.e., produced by recombinant DNA technology, and/or can be, partly or entirely, chemically or biochemically synthesised.
- an “isolated” or “purified” is meant material that is substantially or essentially free from components that normally accompany it in its native state.
- an “isolated polypeptide” or “purified polypeptide” refers to a polypeptide which has been isolated or purified by any suitable means from a mixture of molecules comprising the to be isolated or to be purified polypeptide of interest.
- An isolated or purified polypeptide of interest can for instance be an immunoglobulin, antibody or nanobody, and the mixture can be a mixture or molecules as present in a cell producing the immunoglobulin, antibody or nanobody, and/or the culture medium into which the immunoglobulin, antibody or nanobody is secreted into (likely together with other molecules secreted by the cell).
- antibody fragment refers to a portion of any antibody that by itself has high affinity for an antigenic determinant, or epitope, and contains one or more complementarity determining regions (CDRs) accounting for such specificity.
- CDRs complementarity determining regions
- antibody fragment and “antigen-binding fragment” and “active antibody fragment” and “functional antibody fragment” as used herein refer to a protein or peptide comprising an immunoglobulin domain or an antigen-binding domain capable of specifically binding to a Sarbecovirus spike protein such as SARS-CoV-2 spike protein, in particular to the S2 subunit of the Sarbecovirus spike protein, more particularly to the HR2 domain of (the S2 subunit of) the Sarbecovirus spike protein.
- Non-limiting examples include immunoglobulin domains, Fab, F(ab)'2, scFv, heavy-light chain dimers, immunoglobulin single variable domains, Nanobodies (or VHH antibodies), domain antibodies, and single chain structures, such as a complete light chain or complete heavy chain.
- immunoglobulin (Ig) domain or more specifically “immunoglobulin variable domain” (abbreviated as “IVD”, also referred to herein as “variable domain”), means an immunoglobulin domain essentially consisting of four “framework regions” which are referred to in the art and herein below as “framework region 1” or “FR1”; as “framework region 2” or “FR2”; as “framework region 3” or “FR3”; and as “framework region 4” or “FR4”, respectively; which framework regions are interrupted by three “complementarity determining regions” or “CDRs”, which are referred to in the art and herein below as “complementarity determining region 1” or “CDR1”; as “complementarity determining region 2” or “CDR2”; and as “complementarity determining region 3” or “CDR3”, respectively.
- an immunoglobulin variable domain can be indicated as follows: FR1 - CDR1 - FR2 - CDR2 - FR3 - CDR3 - FR4. It is the immunoglobulin variable domain(s) (IVDs), and in particular the CDRs therein, even more particularly CDR3 therein, that confer specificity to an antibody for the antigen by carrying the antigen- or epitope-binding site.
- IVDs immunoglobulin variable domain
- a heavy chain variable domain (VH) and a light chain variable domain (VL) interact to form an antigen-binding site.
- CDRs complementarity determining regions
- the antigen-binding domain of a conventional 4-chain antibody such as an IgG, IgM, IgA, IgD or IgE molecule; known in the art
- a conventional 4-chain antibody such as an IgG, IgM, IgA, IgD or IgE molecule; known in the art
- a pair of (associated) immunoglobulin domains such as light and heavy chain variable domains, i.e., by a VH-VL pair of immunoglobulin domains, which jointly bind to an epitope of the respective antigen.
- immunoglobulin single variable domain (abbreviated as "ISVD”), equivalent to the term “single variable domain”, defines molecules wherein the antigen-binding site is present on, and formed by, a single immunoglobulin domain. This sets immunoglobulin single variable domains apart from “conventional” immunoglobulins or their fragments, wherein two immunoglobulin domains, in particular two variable domains, interact to form an antigen-binding site.
- immunoglobulin single variable domain refers to a protein or peptide with an amino acid sequence comprising 4 framework regions (FR) and 3 complementary determining regions (CDR) according to the format of FR1-CDR1-FR2-CDR2-FR3-CDR3-FR4.
- the antigen-binding site of an immunoglobulin single variable domain is formed by a single VH/VHH or VL domain.
- the antigen-binding site of an immunoglobulin single variable domain is formed by no more than three CDRs.
- the single variable domain may be a light chain variable domain sequence (e.g., a VL-sequence) or a suitable fragment thereof; or a heavy chain variable domain sequence (e.g., a VH-sequence or VHH sequence) or a suitable fragment thereof; as long as it is capable of forming a single antigen-binding unit (i.e., a functional antigenbinding unit that essentially consists of the single variable domain, such that the single antigenbinding domain does not need to interact with another variable domain to form a functional antigenbinding unit).
- a light chain variable domain sequence e.g., a VL-sequence
- a heavy chain variable domain sequence e.g., a VH-sequence or VHH sequence
- the immunoglobulin single variable domains are heavy chain variable domain sequences (e.g., a VH-sequence or a VHH-sequence); more specifically, the immunoglobulin single variable domains can be heavy chain variable domain sequences that are derived from a conventional four-chain antibody or heavy chain variable domain sequences that are derived from a heavy chain antibody.
- the immunoglobulin single variable domain may be a (single) domain antibody (or an amino acid sequence that is suitable for use as a (single) domain antibody), a variable domain of a heavy (VH) or light (VL) chain of a conventional antibody (also referred to as a “dAb”) (or an amino acid sequence that is suitable for use as a dAb) or a Nanobody (as defined herein, and including but not limited to a VHH); or any suitable fragment of any one thereof.
- a (single) domain antibody or an amino acid sequence that is suitable for use as a (single) domain antibody
- VH heavy
- VL light
- dAb conventional antibody
- Nanobody as defined herein, and including but not limited to a VHH
- the immunoglobulin single variable domain may be a Nanobody (as defined herein) or a suitable fragment thereof.
- Nanobody®, Nanobodies® and Nanoclone® are registered trademarks of Ablynx N.V. (a Sanofi Company).
- Nanobodies reference is made to the further description below, as well as to the prior art cited herein, such as e.g. described in W02008/020079.
- VHH domains also known as VHHs, VHH domains, VHH antibody fragments, and VHH antibodies, have originally been described as the antigen-binding immunoglobulin (Ig) (variable) domain of “heavy chain antibodies” (i.e., of “antibodies devoid of light chains”; Hamers-Casterman et al. 1993, Nature 363: 446-448).
- the term “VHH domain” has been chosen to distinguish these variable domains from the heavy chain variable domains that are present in conventional 4-chain antibodies (which are referred to herein as “VH domains”) and from the light chain variable domains that are present in conventional 4-chain antibodies (which are referred to herein as “VL domains”).
- the binding agents according to the current invention can be structurally defined as polypeptidic or polypeptide binding agents, in particular antibodies and antibody-fragments, comprising at least one immunoglobulin single variable domain (ISVD) as defined herein.
- the binding agents, in particular antibodies and antibody-fragments, more particularly ISVDs, as described herein comprise at least CDR3 as comprised in an immunoglobulin single variable domain (ISVD) defined by any of SEQ ID NO:2-26, wherein the CDR3 is annotated according to Kabat or Martin.
- the binding agents, in particular the antibodies and antibody fragments, more particularly ISVDs, as described herein comprise at least two of CDR1, CDR2 and CDR3 (e g.
- ISVDs immunoglobulin single variable domain
- Such CDRs may be comprised in any of VHH hR3_DC23 (defined by/set forth in SEQ ID NO:2), VHH RNGL (defined by/set forth in SEQ ID NO:3), VHH RAGL (defined by/set forth in SEQ ID NO:4), VHH E2NGL (defined by/set forth in SEQ ID NO:5), VHH E2AGL (defined by/set forth in SEQ ID NO: 6), VHH QRNGL (defined by/set forth in SEQ ID NO: 7), VHH QRAGL (defined by/set forth in SEQ ID NO: 8), VHH ERNGL (defined by/set forth in SEQ ID NOV), VHH ERAGL (defined by/set forth in SEQ ID NO: 10), VHH E10RNGL (defined by/set forth in SEQ ID NO: 11), VHH E10RAGL (defined by/set forth in SEQ ID NO: 12), VHH E81RNGL (defined by/set forth in SEQ ID NO: 13), VHH E81RAGL (
- VHH RNGL VHH RNGL
- numbering of the amino acid residues of any IVD or ISVD different numbering schemes can be applied. For example, numbering can be performed according to the AHo numbering scheme for all heavy (VH) and light chain variable domains (VL) given by Honegger & Pltickthun (2001. J Mol Biol 309:657-70), as applied to VHH domains from camelids.
- Alternative methods for numbering the amino acid residues of VH domains which can also be applied in an analogous manner to VHH domains, are known in the art.
- the delineation of the FR and CDR sequences can be done by using the Kabat numbering system as applied to VHH domains from camelids by Riechmann & Muyldermans (1999.
- the total number of amino acid residues in each of the CDRs may vary and may not correspond to the total number of amino acid residues indicated by the Kabat numbering (that is, one or more positions according to the Kabat numbering may not be occupied in the actual sequence, or the actual sequence may contain more amino acid residues than the number allowed for by the Kabat numbering).
- the numbering according to Kabat may or may not correspond to the actual numbering of the amino acid residues in the actual sequence.
- the total number of amino acid residues in a VH domain and a VHH domain will usually be in the range of from 110 to 120, often between 112 and 115. It should however be noted that smaller and longer sequences may also be suitable for the purposes described herein.
- CDR regions in an antibody/immunoglobulin sequence generally depends on the algorithm/methodology applied. For example, determination of CDR regions may be done according to the designation based on contact analysis and binding site topography as described in MacCallum et al. (J. Mol. Biol. (1996) 262, 732-745), AbM (AbM is Oxford Molecular Ltd.'s antibody modelling package as described on http://www.bioinf.org.uk/abs/index.html), Chothia (Chothia and Lesk, 1987; Mol Biol. 196:901-17), Martin (Abhinandan, and Martin.
- Fig. 6 illustrates the different annotation -schemes or - methods as applied to the amino acid sequence of VHH hR3_DC23 (SEQ ID NO:2).
- these CDRs can be described as the CDR sequences present in the ISVDs (as described herein) as determined or delineated according to a well-known methodology such as according to any one of the Kabat-, Martin-, Chothia-, aHo, MacCallum et al. 1996, AbM-, or IMGT, numbering scheme or method, such as preferably the Martin numbering scheme or method.
- a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a CDR1 defined by/set forth in any one of SEQ ID NOs: 40 or 38.
- a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a CDR2 defined by/set forth in any one of SEQ ID NOs: 30 or 39.
- a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a CDR3 defined by/set forth in SEQ ID NO: 41.
- the binding agent in particular the antibody or antibody fragment, more particularly the ISVD, as described herein may comprise: - a CDR1 defined by/set forth in SEQ ID NO:40; a CDR2 defined by/set forth in SEQ ID NO:30; and a CDR3 defined by/set forth in SEQ ID NO:41; or
- the binding agent in particular the antibody or antibody fragment, more particularly the ISVD, as described herein does not comprise the sequence set forth in SEQ ID NO: 1 or 2.
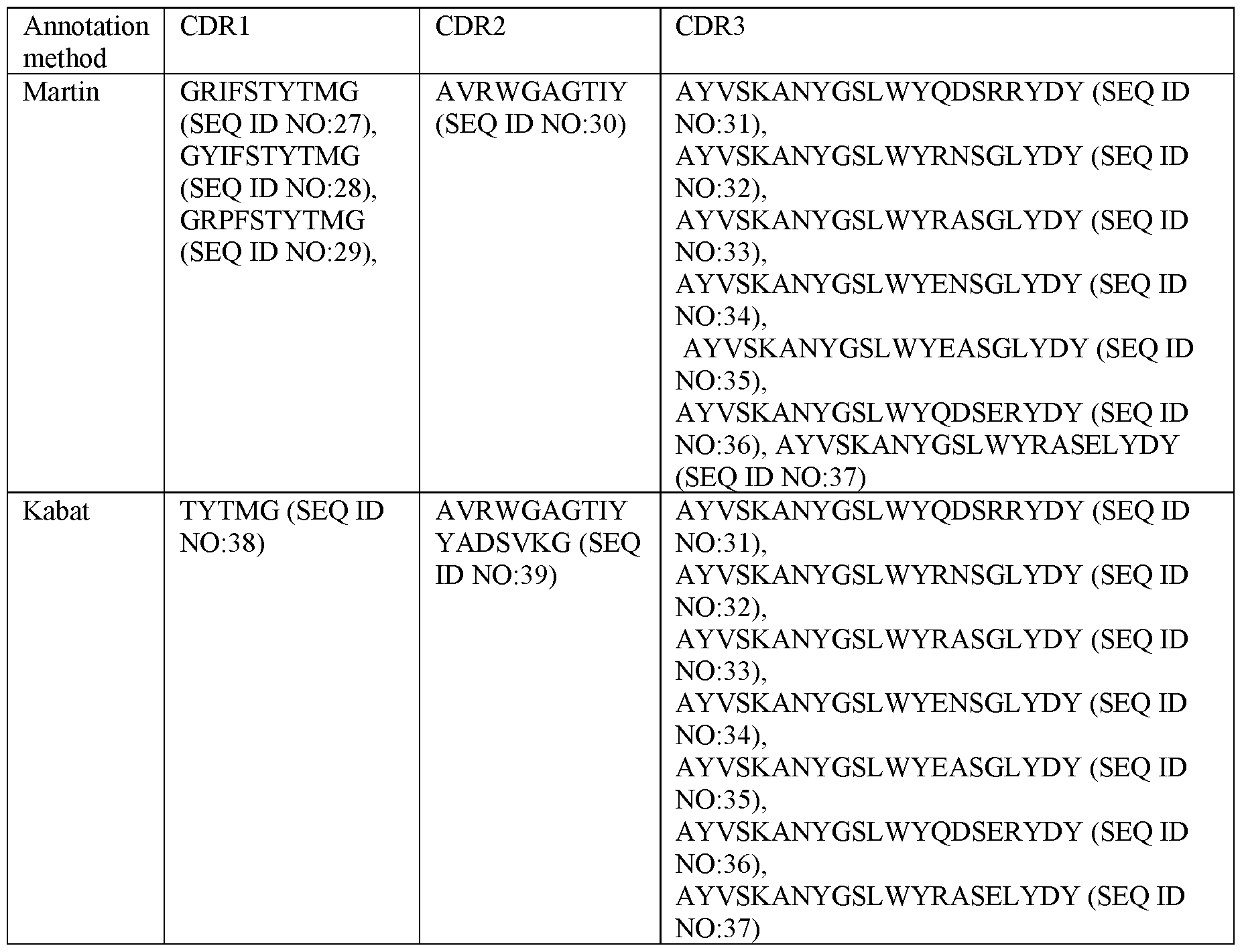
- Table 1 Sequences of CDRs in the VHHs according to certain embodiments:
- a binding agent or Sarbecovirus binding agent in particular an antibody or antibody fragment or Sarbecovirus antibody or antibody-fragments, more particularly an ISVD, as described herein, may be characterized in that it comprises a CDR1 as present in any of SEQ ID NOs:2 to 26, wherein the CDR1 is annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
- a binding agent or Sarbecovirus binding agent in particular an antibody or antibody fragment or Sarbecovirus antibody or antibodyfragments, more particularly an ISVD, as described herein, may be characterized in that it comprises a CDR2 as present in any of SEQ ID NOs:2 to 26, wherein the CDR2 is annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
- a binding agent or Sarbecovirus binding agent in particular an antibody or antibody fragment or Sarbecovirus antibody or antibody-fragments, more particularly an ISVD, as described herein, may be characterized in that it comprises a CDR3 as present in any of SEQ ID NOs:2 to 26, wherein the CDR3 is annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
- a binding agent or Sarbecovirus binding agent in particular an antibody or antibody fragment or Sarbecovirus antibody or antibody-fragments, more particularly an ISVD, as described herein, may be characterized in that it comprises a CDR1, CDR2 and CDR3, each independently as present in any of SEQ ID NOs:2 to 26, wherein the CDR1, CDR2 and CDR3 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
- a binding agent or Sarbecovirus binding agent in particular an antibody or antibody fragment or Sarbecovirus antibody or antibody-fragments, more particularly an ISVD, as described herein, may be characterized in that it comprises a combination of CDR1, CDR2 and CDR3, wherein the CDR1, CDR2 and CDR3 are as present in a particular one of the sequences set forth in SEQ ID NOs:2 to 26, wherein the CDR1, CDR2 and CDR3 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
- a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a CDR1 defined by/set forth in any one of SEQ ID NOs:27, 28, 29, or 38.
- a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a CDR2 defined by/set forth in any one of SEQ ID NOs:30 or 39.
- a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises; and a CDR3 defined by/set forth in any one of SEQ ID NOs:31-37.
- the binding agent, in particular the antibody or antibody fragment, more particularly the ISVD, as described herein may comprise:
- Table 2 Example definitions / sequences of the CDRs in the VHHs of certain embodiments as described herein by employing different annotation methodologies as indicated, in particular CDRs comprised in any of VHH hR3_DC23, VHH RNGL, VHH RAGL, VHH E2NGL, VHH E2AGL, VHH QRNGL, VHH QRAGL, VHH ERNGL, VHH ERAGL, VHH E10RNGL, VHH E10RAGL, VHH E81RNGL, VHH E81RAGL, VHH Q75, VHH E75, VHH E81, VHH E10E81, VHH D16, VHH T19D79, VHH TQD, VHH Q75R21Y, VHH CBV1, VHH CBV2, VHH CBV3, or VHH CBV4, determined according to Kabat or Martin system or method.
- polypeptidic or polypeptide binding agents in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising the complementarity determining regions (CDRs) present in any one of SEQ ID NOs:2-26, wherein the CDRs are defined according to Martin.
- the binding agents, in particular antibodies or antibody fragments, more particularly ISVDs comprise one of following sets of three complementarity determining regions (CDRs):
- -CDR1 defined by/set forth in SEQ ID NO:27, CDR2 defined by/set forth in SEQ ID NO:30, and CDR3 defined by/set forth in SEQ ID NO:34; or -CDR1 defined by/set forth in SEQ ID NO:27, CDR2 defined by/set forth in SEQ ID NO:30, and CDR3 defined by/set forth in SEQ ID NO: 35; or
- CDR1 defined by/set forth in SEQ ID NO:29
- CDR2 defined by/set forth in SEQ ID NO:30
- CDR3 defined by/set forth in SEQ ID NO: 37.
- polypeptidic or polypeptide binding agents in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising the complementarity determining regions (CDRs) present in any one of SEQ ID NOs:2-26, wherein the CDRs are defined according to Kabat.
- the binding agents, in particular antibodies or antibody fragments, more particularly ISVDs comprise one of following sets of three complementarity determining regions (CDRs):
- polypeptidic or polypeptide binding agents in particular the antibodies and antibody fragments, more particularly the ISVDs, according to the current invention can comprise one or more framework regions (FRs) as comprised in any one of SEQ ID NOs:2-26, or variants of such FRs.
- FRs framework regions
- binding agents, antibodies or antibody fragments, or ISVDs may comprise at least one, such as one, two, three or all of an FR1, FR2, FR3, and FR4 region, each independently as comprised in any one of SEQ ID NOs:2-26, or variants of such FRs.
- such binding agents, antibodies or antibody fragment, or ISVDs may comprise an FR1 and FR2 region, an FR1 and FR3 region, an FR1 and FR4 regions, an FR2 and FR3 region, an FR2 and FR4 region, an FR3 and FR4 region, an FR1, FR2 and FR3 region, an FR1, FR2 and FR4 region, an FR2, FR3 and FR4, or an FR1, FR3 and FR4 region as comprised in any one of SEQ ID NOs:2-26, or variants of such FRs.
- any one of the systems or methods for numbering amino acids in immunoglobulin protein sequences as described elsewhere herein and illustrated in Fig. 6 for VHH hR3_DC23, and known to a skilled artisan can be applied.
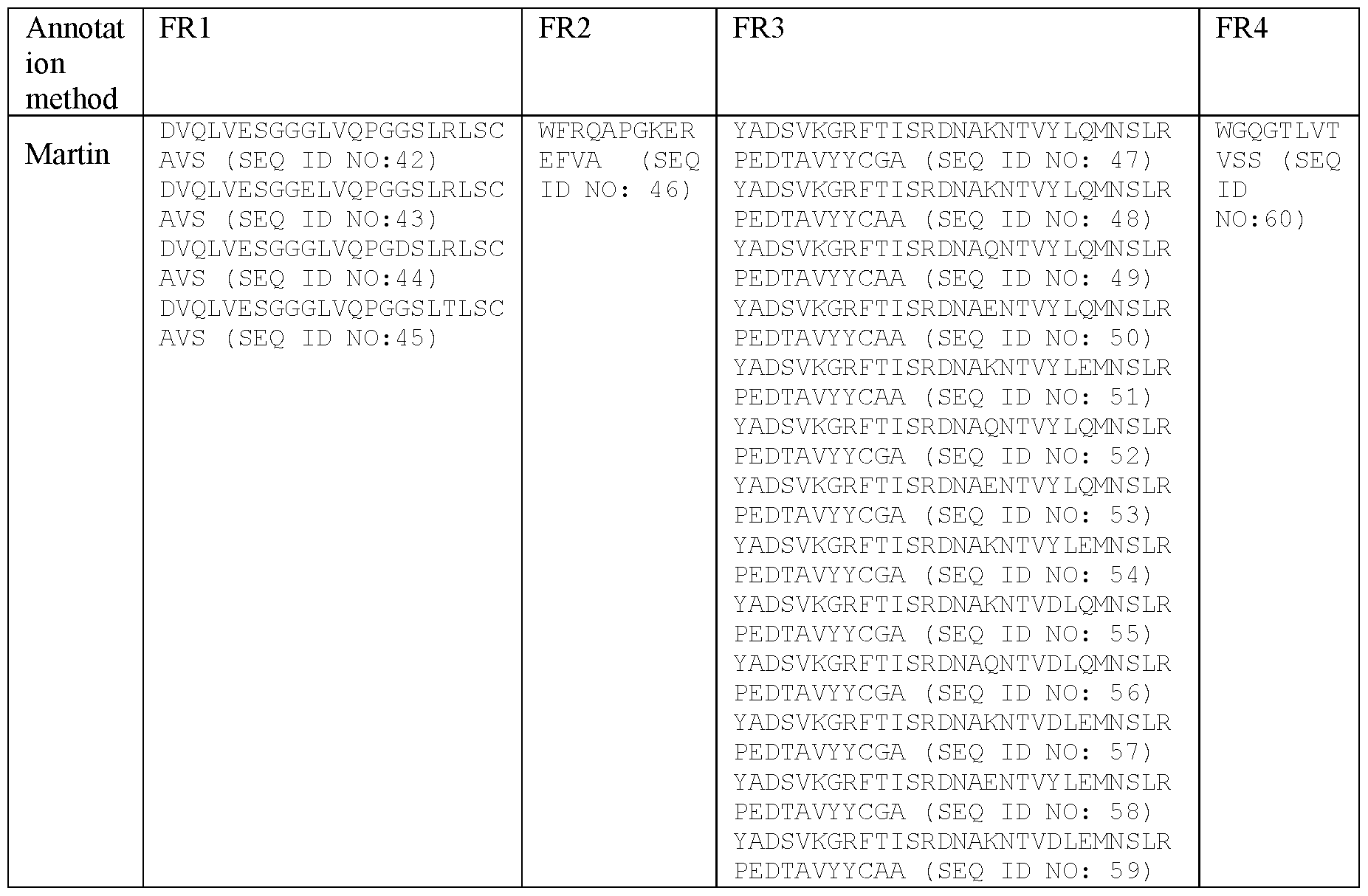
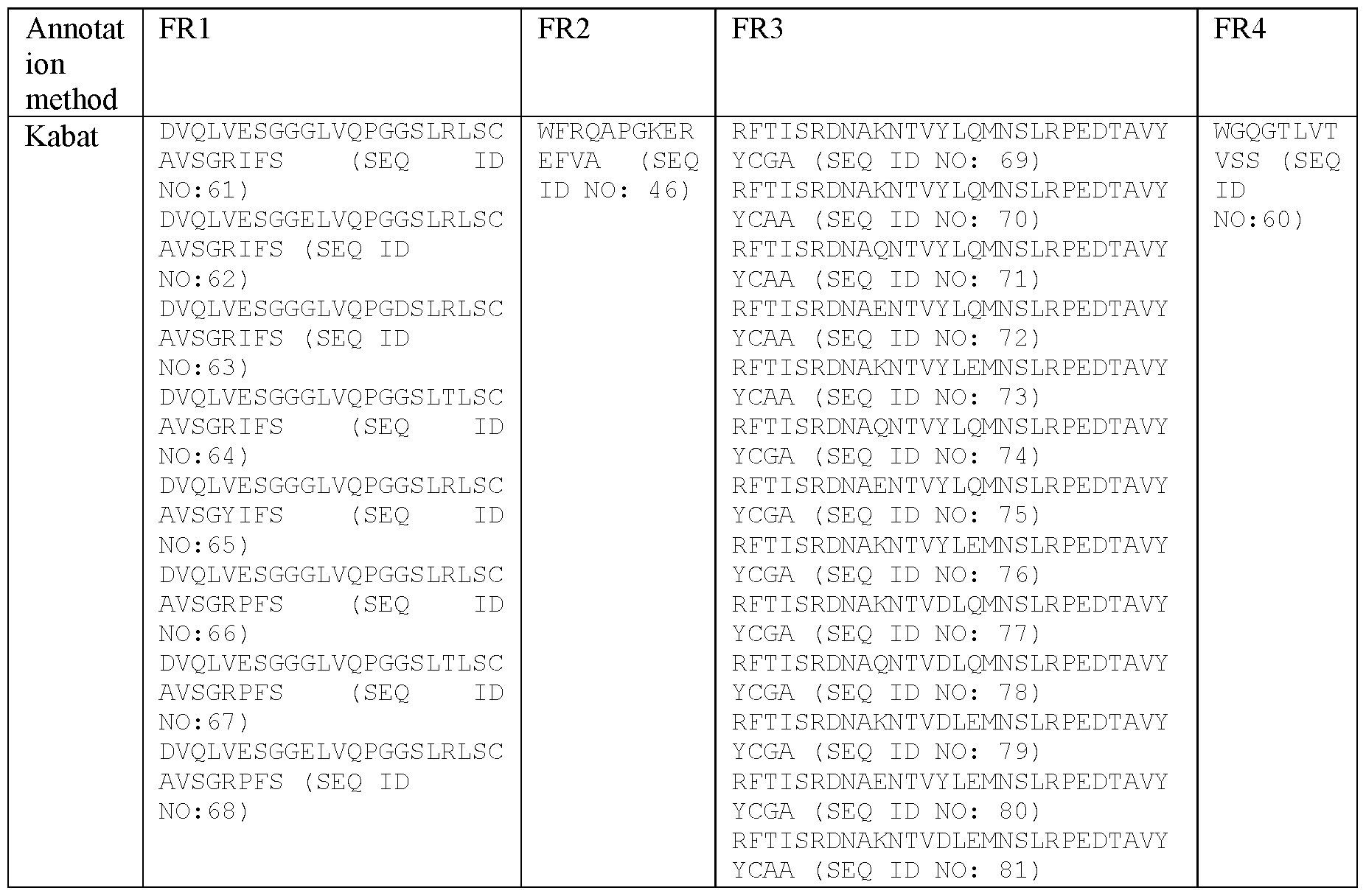
- sequences of the FRs in certain specific VHHs as described herein by employing the Martin or Kabat methodology are shown in Table 3.
- Table 3 Example sequences of the FRs in the VHHs of certain embodiments as described herein by employing the Martin or Kabat methodology.
- a polypeptidic or polypeptide binding agent in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a framework region 1 (FR1) present in any one of SEQ ID NOs:2-26, wherein the FR1 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum, or in that it comprises a variant FR1 which is at least 90% or 95% identical to, or which has at most 3, such as 1, 2 or 3, amino acid substitutions, deletions or additions, such as preferably conservative and/or humanizing substitutions, compared to, a FR1 present in any one of SEQ ID NOs:2-26, wherein the FR1 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum
- a polypeptidic or polypeptide binding agent in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a framework region 2 (FR2) present in any one of SEQ ID NOs:2-26, wherein the FR2 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum, or in that it comprises a variant FR2 which is at least 85% or 90% identical to, or which has at most 2, such as 1 or 2, amino acid substitutions, deletions or additions, such as preferably conservative and/or humanizing substitutions, compared to, a FR2 present in any one of SEQ ID NOs:2-26, wherein the FR2 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
- a polypeptidic or polypeptide binding agent in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a framework region 3 (FR3) present in any one of SEQ ID NOs:2-26, wherein the FR3 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum, or in that it comprises a variant FR3 which is at least 80%, 85%, 90% or 95% identical to, or which has at most 9, such as 1, 2, 3, 4, 5, 6, 7, 8 or 9, amino acid substitutions, deletions or additions, such as preferably conservative and/or humanizing substitutions, compared to, a FR3 present in any one of SEQ ID NOs:2-26, wherein the FR3 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
- FR3 framework region 3
- a polypeptidic or polypeptide binding agent in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a framework region 4 (FR4) present in any one of SEQ ID NOs:2-26, wherein the FR4 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum, or in that it comprises a variant FR4 which is at least 90% identical to, or which has at most 1 amino acid substitution, deletion or addition, such as preferably a conservative and/or humanizing substitution, compared to, a FR4 present in any one of SEQ ID NOs:2-26, wherein the FR4 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
- FR4 framework region 4
- a polypeptidic or polypeptide binding agent in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises, each independently, a FR1 present in any one of SEQ ID Nos:2-26 or a variant FR1 as defined hereinabove; a FR2 present in any one of SEQ ID Nos:2-26 or a variant FR2 as defined hereinabove; a FR3 present in any one of SEQ ID Nos:2-26 or a variant FR3 as defined hereinabove; and a FR4 present in any one of SEQ ID Nos: 2-26 or a variant FR4 as defined hereinabove, wherein the FR1, FR2, FR3 and FR4 are defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
- a polypeptidic or polypeptide binding agent in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises at least one, or the particular combination of two, three or all of the framework regions (FRs) as present in any one of SEQ ID NOs: 2-26, or any variant of said FR or FRs as defined herein above, wherein the FRs are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
- FRs framework regions
- a polypeptidic or polypeptide binding agent in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises at least one, or the particular combination of two, three or all of the framework regions (FRs) present in any one of SEQ ID NOs: 2 to 26, wherein the FRs are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
- FRs framework regions
- polypeptidic or polypeptide binding agents in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising, each independently, a FR1 present in any one of SEQ ID NOs:2-26; a FR2 present in any one of SEQ ID NOs:2-26; a FR3 present in any one of SEQ ID NOs:2-26, and a FR4 present in any one of SEQ ID NOs:2-26, wherein the FR1, FR2, FR3 and FR4 are defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
- a polypeptidic or polypeptide binding agent in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises:
- FR1 defined by any one of SEQ ID NO:42-45, a FR2 defined by SEQ ID NO:46, a FR3 defined by any one of SEQ ID NO:47-59 and a FR4 defined by SEQ ID NO:60; or
- FR1 defined by any one of SEQ ID NO:61-68, a FR2 defined by SEQ ID NO:46, a FR3 defined by any one of SEQ ID NO:69-81 and a FR4 defined by SEQ ID NO:60.
- polypeptidic or polypeptide binding agents in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising a FR1, FR2, FR3 and FR4 as present in the same sequence of any of the sequences shown in SEQ ID NOs:2-26, wherein the FR1, FR2, FR3 and FR4 are defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
- polypeptidic or polypeptide binding agents in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising all four framework regions (FRs) present in any one of SEQ ID NOs:2-26, wherein the FRs are defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
- polypeptidic or polypeptide binding agents in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising the framework regions (FRs) present in any one of SEQ ID NOs:2-26, wherein the FRs are defined according to Martin.
- the binding agents, in particular antibodies or antibody fragments, more particularly ISVDs comprise one of following sets of framework regions (FRs): -FR1 defined by/set forth in SEQ ID NO:42, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:47, and FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:42
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:48
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:42
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:49
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:42
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:50
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:43
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:48
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:42
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:51
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:42, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO: 52, and FR4 defined by/set forth in SEQ ID NO: 60; or
- FR1 defined by/set forth in SEQ ID NO:42
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:53
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:42
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:54
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:43
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:54
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:44
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:47
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:45
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:55
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:45
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:56
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:45
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:59
- FR4 defined by/set forth in SEQ ID NO:60.
- polypeptidic or polypeptide binding agents in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising the framework regions (FRs) present in any one of SEQ ID NOs:2-26, wherein the FRs are defined according to Kabat.
- the binding agents, in particular antibodies or antibody fragments, more particularly ISVDs comprise one of following sets of framework regions (FRs):
- FR1 defined by/set forth in SEQ ID NO:61
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:69
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:61
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:70
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:61
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:71
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:61
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:72
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:62
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:70
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:61
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO: 73
- FR4 defined by/set forth in SEQ ID NO: 60;
- FR1 defined by/set forth in SEQ ID NO:61
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:74
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:61
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:75
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO: 61
- FR2 defined by/set forth in SEQ IDNO:46
- FR3 defined by/set forth in SEQ ID NO:76
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:64
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:77
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:64
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:78
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:65
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:74
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:66
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:76
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:67
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:79
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:68
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:80
- FR4 defined by/set forth in SEQ ID NO:60;
- FR1 defined by/set forth in SEQ ID NO:67
- FR2 defined by/set forth in SEQ ID NO:46
- FR3 defined by/set forth in SEQ ID NO:81
- FR4 defined by/set forth in SEQ ID NO:60.
- polypeptidic or polypeptide binding agents in particular antibodies or antibody fragments, comprise one or more ISVDs individually defined by or set forth in any one of SEQ ID NOs: 2 to 26, or comprise one or more ISVDs comprising or consisting of an amino acid sequence selected from the group of SEQ ID NO: 2 to 26.
- said polypeptidic or polypeptide binding agents in particular antibodies or antibody fragments, more particularly ISVDs, comprise or consist of an amino acid sequence with at least 90% identity to an amino acid sequence selected from the group of SEQ ID NO: 2 to 26, or with at least 95% identity to an amino acid sequence selected from the group of SEQ ID NO: 2 to 26.
- Such non-identity or variability is preferably limited to non-identity or variability in FR amino acid residues.
- Cconservative amino acid substitutions can be introduced in a protein or polypeptide whereby such substitutions have no essential or substantial effect on the protein's activity.
- Preferred conservative substitutions are those fulfilling the criteria defined for an accepted point mutation in Dayhoff et al., Atlas of Protein Sequence and Structure, 5, pp.
- substitutions including but not limited to the following groups: (a) valine, glycine; (b) glycine, alanine; (c) valine, isoleucine, leucine; (d) aspartic acid, glutamic acid; (e) asparagine, glutamine; (f) serine, threonine; (g) lysine, arginine, methionine; and (h) phenylalanine, tyrosine.
- a “homologue”, or “homologues” of a protein of interest encompass(es) proteins having amino acid substitutions, deletions and/or insertions relative to an unmodified protein of interest and having essentially or substantially similar biological and functional activity as the unmodified protein from which it is/they are derived.
- a “percentage (of) sequence identity” is calculated by comparing two optimally aligned (amino acid or nucleic acid) sequences over the window of comparison, determining the number of positions at which the identical amino acid or nucleotide residue occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison (i.e., the window size), and multiplying the result by 100 to yield the percentage of (amino acid or nucleic acid) sequence identity.
- Immunoglobulin single variable domains such as Domain antibodies and Nanobody® (including VHH domains) as described herein can be and/or have been subjected to humanization, i.e. to increase the degree of sequence identity with the closest human germline sequence.
- humanized immunoglobulin single variable domains such as Nanobody® (including VHH domains) are immunoglobulin single variable domains in which at least one amino acid residue is present (and in particular, at least one framework residue) that is and/or that corresponds to a humanizing substitution (as defined further herein).
- Potentially useful humanizing substitutions can be ascertained by comparing the sequence of the framework regions of a VHH sequence with the corresponding framework sequence of one or more closely related human VH sequences, after which one or more of the potentially useful humanizing substitutions (or combinations thereof) thus determined can be introduced into said VHH sequence (in any manner known per se, as further described herein) and the resulting humanized VHH sequences can be tested for affinity for the target, for stability, for ease and level of expression, and/or for other desired properties. In this way, by means of a limited degree of trial and error, other suitable humanizing substitutions (or suitable combinations thereof) can be determined by the skilled person. Also, based on what is described before, (the framework regions of) an immunoglobulin single variable domain, such as a Nanobody® (including VHH domains) may be partially humanized or fully humanized.
- Humanized immunoglobulin single variable domains may have several advantages, such as a reduced immunogenicity, compared to the corresponding non-humanized VHH domains.
- humanized is meant mutated so that immunogenicity upon administration in human patients is minor or non-existent.
- the humanizing substitutions are chosen such that the resulting humanized amino acid sequence and/or ISVD or VHH still retains the favourable properties of the parental (non-humanized) VHH, such as the antigen-binding capacity.
- the skilled person will be able to select humanizing substitutions or suitable combinations of humanizing substitutions which optimize or achieve a desired or suitable balance between the favourable properties provided by the humanizing substitutions on the one hand and the favourable properties of non-humanized VHH domains on the other hand.
- Such methods are known by the skilled addressee.
- a human consensus sequence can be used as target sequence for humanization, but also other means are known in the art.
- One alternative includes a method wherein the skilled person aligns a number of human germline alleles, such as for instance but not limited to the alignment of IGHV3 alleles, and to use said alignment for identification of residues suitable for humanization in the target sequence.
- a subset of human germline alleles most homologous to the target sequence may be aligned as starting point to identify suitable humanisation residues.
- the VHH is analyzed to identify its closest homologue in the human alleles and used for humanisation construct design.
- a humanisation technique applied to Camelidae VHHs may also be performed by a method comprising the replacement of specific amino acids, either alone or in combination. Said replacements may be selected based on what is known from literature, from known humanization efforts, as well as from human consensus sequences compared to the natural VHH sequences, or from the human alleles most similar to the VHH sequence of interest.
- a human-like class of Camelidae single domain antibodies contain the hydrophobic FR2 residues typically found in conventional antibodies of human origin or from other species, but compensating this loss in hydrophilicity by other substitutions at position 103 that substitutes the conserved tryptophan residue present in VH from double-chain antibodies.
- peptides belonging to these two classes show a high amino acid sequence homology to human VH framework regions and said peptides might be administered to a human directly without expectation of an unwanted immune response therefrom, and without the burden of further humanisation.
- some Camelidae VHH sequences display a high sequence homology to human VH framework regions and therefore said VHH might be administered to patients directly without expectation of an immune response therefrom, and without the additional burden or need of humanization.
- Suitable mutations in particular substitutions, can be introduced during humanization to generate a polypeptide with reduced binding to pre-existing antibodies (reference is made for example to WO 2012/175741 and WO2015/173325), for example at least one of the positions: 11, 13, 14, 15, 40, 41, 42, 82, 82a, 82b, 83, 84, 85, 87, 88, 89, 103, or 108.
- the amino acid sequences and/or VHH of the invention may be or have been suitably humanized at any framework residue(s), such as at one or more Hallmark residues (as defined below) or at one or more other framework residues (i.e. nonHallmark residues) or any suitable combination thereof.
- deletions and/or substitutions may also be designed in such a way that one or more sites for posttranslational modification (such as one or more glycosylation sites) are removed, as will be within the ability of the person skilled in the art.
- substitutions or insertions may be designed so as to introduce one or more sites for attachment of functional groups (as described herein), for example to allow site-specific pegylation.
- At least one of the typical Camelidae hallmark residues with hydrophilic characteristics at position 37, 44, 45 and/or 47 is replaced (see Table A-03 of W02008/020079).
- Another example of humanization includes substitution of residues in FR 1, such as position 1, 5, 11, 14, 16, and/or 28; in FR3, such as positions 73, 74, 75, 76, 78, 79, 82b, 83, 84, 93 and/or 94; and in FR4, such as position 10 103, 104, 108 and/or 111 (see Tables A-05 -A08 of W02008/020079; all numbering according to the Kabat-methodology).
- Immunoglobulin single variable domains as described herein can be and/or have been subjected to affinity maturation.
- the ISVD is a variant of an ISVD as described herein, in particular an ISVD of any one of SEQ ID NO:2 to 26, in particular an ISVD of SEQ ID NO:2 or 25, which has an improved affinity for SARS-CoV-2 S2.
- said variant ISVD is obtained by an affinity maturation protocol.
- the ISVD variant may comprise up to 4 such as 1, 2, 3 or 4 substitutions, in particular substitutions of paratope residues, optionally wherein said paratope comprises or consists of residues at positions 96, 97, 98, 99, 100a, 100b and 1001 according to Kabat numbering of the ISVD of any one of SEQ ID NO:2 to 26, in particular an ISVD of SEQ ID NO:2 or 25, and/or at positions 27, 29, 30, 31, 52, 100, 100c and lOOd according to Kabat numbering of the ISVD of any one of SEQ ID NO:2 to 26, in particular an ISVD of SEQ ID NO:2 or 25.
- the ISVD variant may comprise a substitution of a residue at position 96, 97, 98, 99, 100b or 1001 according to Kabat numbering of the ISVD of any one of SEQ ID NO:2 to 26, in particular an ISVD of SEQ ID NO:2 or 25.
- the ISVD may comprise up to 4 such as 1, 2, 3 or 4 substitutions of residues at positions 27, 29, 30, 31, 100, 100c and/or lOOd according to Kabat numbering of the ISVD of any one of SEQ ID NO:2 to 26, in particular an ISVD of SEQ ID NO:2 or 25.
- the residue at position 96 may be substituted with F, R, W, H or Q, preferably with F, R or W.
- the residue at position 97 may be substituted with I, A or L.
- the residue at position 98 may be substituted with A or T.
- the residue at position 99 may be substituted with R.
- the residue at position 100b may be substituted with F.
- the residue at position 1001 may be substituted with L, H or V, more preferably with L.
- the residue at position 27 may be substituted with H, L, P, S or T, preferably with H.
- the residue at position 29 may be substituted with Y, H, M, R, W, I, L, S, V, A or G, preferably with Y, H, M, R or W.
- the residue at position 30 may be substituted with K, T, I, R, G, H, L, P, F, N or W, preferably with K, T or I, more preferably with K.
- the residue at position 31 may be substituted with S.
- the residue at position 100 may be substituted with R.
- the residue at position 100c may be substituted with H or A, preferably with H.
- the residue at position lOOd may be substituted with R, H, P, Q or V.
- the substitution(s) is/are selected from the substitutions set forth in Table 14.
- the ISVD is a variant of hR3_DC23 of SEQ ID NO:2.
- the variant may have an improved affinity for SARS-CoV-2 S2 compared to hR3_DC23.
- the ISVD, in particular the hR3_DC23 variant is an ISVD comprising an amino acid sequence defined by any one of SEQ ID NO: 151-182.
- the ISVD is a variant of CBV3 of SEQ ID NO:25.
- the variant may have an improved affinity for SARS-CoV-2 S2 compared to CBV3.
- the ISVD, in particular the CBV3 variant is an ISVD comprising an amino acid sequence defined by any one of SEQ ID NO: 183-244, preferably an ISVD comprising an amino acid sequence defined by any one of SEQ ID NO: 183-188, 190-195, 197, 198, or 200-244.
- binding agents or Sarbecovirus binding agents can also be described functionally by any individual fimction/embodiment or by any combination of any number of the individual fimctions/embodiments described hereafter and given an arbitrary number “n” between brackets “(n)”.
- the numerical order of these individual functions is random and not imposing any preference on an individual function; similarly, this random numerical order is not imposing any preference on any combination of two or more of the individual functions. Any such combination is furthermore not to be considered as arbitrary as the binding agents or Sarbecovirus binding agents herein exert each of these individual functions.
- the present invention thus provides binding agents, in particular antibodies or antigen-binding fragments thereof, that (1) specifically bind to a Sarbecovirus such as SARS-CoV-2, SARS-CoV-1 and Khosta-2 and may also be referred to herein as Sarbecovirus binding agents or Sarbecovirus antibodies and antibody fragments.
- the binding agents (2) do not bind Middle East respiratory syndrome coronavirus (MERS-CoV).
- Binding means any interaction, be it direct or indirect.
- a direct interaction implies a contact (e.g. physical or chemical) between two binding partners.
- An indirect interaction means any interaction whereby the interaction partners interact in a complex of more than two molecules.
- An interaction can be completely indirect (e.g. two molecules are part of the same complex with the help of one or more bridging molecules but don’t bind in the absence of the bridging molecule(s)).
- An interaction may be partly direct or partly indirect: there is still a direct contact between two interaction partners, but such contact is e.g. not stable, and is stabilized by the interaction with one or more additional molecules.
- Specificity of binding or “binding specificity” or “specifically binding” refers to the situation in which a molecule A is, at a certain concentration (e.g. sufficient to inhibit or neutralize a protein or process of interest) binding to a target of interest (e.g. protein) with higher affinity (e.g. at least 2- fold, 5-fold, orat least 10-fold higher affinity, e.g. at least 20-, 50- or 100-fold or more higher affinity) than the affinity with which it is possibly (if at all) binding to other targets (targets not of interest).
- Specific binding does not mean exclusive binding. However, specific binding does mean that a binder has a certain increased affinity or preference for one or a few of its targets.
- Exclusivity of binding refers to the situation in which a binder is binding only to the target of interest.
- affinity generally refers to the degree to which one molecule (e.g. ligand, chemical, protein or peptide, antibody or antibody fragment) binds to another molecule (e.g. (target) protein or peptide) so as to shift the equilibrium of single molecule monomers towards a complex formed by (specific)(non-covalent) binding of the two molecules.
- Non-covalent interaction or binding between 2 or more binding partners may involve interactions such as van der Waals interaction, hydrogen bonding, and salt bridges.
- the “dissociation constant” or “binding constant” (KD) is commonly used to describe the affinity between the two molecules and it is often calculated by the ratio of the rate constant for the complex formation (referred to as the “k on " value) to the rate constant for dissociation of said complex (the “k O ff” or “kdis” value).
- the measurement of binding affinity of a molecule to another molecule is known to the skilled person and includes, e.g., real-time, label free bio-layer interferometry assay, e.g., an Octet® RED96 system (ForteBio), or surface plasmon resonance (SPR), e.g., BIACORETM, or solution-affinity ELISA.
- real-time, label free bio-layer interferometry assay e.g., an Octet® RED96 system (ForteBio), or surface plasmon resonance (SPR), e.g., BIACORETM, or solution-affinity ELISA.
- coronavirus refers to a family of viruses, which has its name from the large spike protein molecules that are present on the virus surface and give the virions a crown-like shape.
- the Coronoviridae family comprises four genera: Alphacoronavirus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus.
- Coronaviruses represent a diverse family of large enveloped positive-stranded RNA viruses that infect a wide range of animals, a wide variety of vertebrate species, and humans.
- the spike (S) proteins of coronaviruses are essential for host receptor-binding and subsequent fusion of the viral and host cell membrane, effectively resulting in the release of the viral nucleocapsids in the host cell cytoplasm (Letko et al. (2020) Nat Microbiol 5:562-569).
- HCoV-NL63 and HCoV-229E a-coronaviruses
- HCoV-OC43 and HCoV-HKUl P-coronaviruses
- SARS severe acute respiratory syndrome virus
- SARS-CoV-1 SARS-CoV-1
- MERS Middle East respiratory syndrome
- SARS-CoV-2 severe acute respiratory syndrome coronavirus 2
- SARS-CoV severe acute respiratory syndrome coronavirus
- MERS-CoV Middle East respiratory syndrome coronavirus
- SARSr-CoV Severe acute respiratory syndrome-related coronavirus
- SARS coronavirus SARS-CoV-1 and SARS-CoV-2.
- SARS-CoV-2 belongs to the same genus Betacoronavirus as SARS-CoV (lineage B) and MERS-CoV (lineage C), genomic analysis revealed greater similarity between SARS-CoV-2 and SARS-CoV, supporting its classification as a member of lineage B (from the International Committee on Taxonomy of Viruses).
- Betacoronaviruses this virus is characterized by a unique combination of polybasic cleavage sites, a distinctive feature known to increase pathogenicity and transmissibility.
- a bat Sarbecovirus, Bat CoV RaTG13, sampled from a Rhinolophus affinis horseshoe bat was reported to cluster with SARS-CoV-2 in almost all genomic regions with approximately 96% genome sequence identity (and over 93% similarity in the receptor binding domain (RBD) of the spike protein); another mammalian species may have acted as intermediate host.
- One of the suspected intermediate hosts harbours coronaviruses showing high similarity to SARS-CoV-2 in the receptor-binding domain, which contains mutations believed to promote binding to the angiotensinconverting enzyme 2 (ACE2) receptor and demonstrates a 97% amino acid sequence similarity.
- ACE2 angiotensinconverting enzyme 2
- SARS-CoV-1 and -2 both use angiotensin converting enzyme 2 (ACE2) as a receptor on human cells.
- ACE2 angiotensin converting enzyme 2
- SARS-CoV-2 binds ACE2 with a higher affinity than SARS-CoV-1 (Wrapp et al. (2020) Science 367: 1260-1263).
- SARS-CoV-2 differentiates from SARS-CoV-1 and several SARS-related coronaviruses (SARSr-CoVs) as outlined in e.g. Abdelrahman et al. (2020. Front Immunol 11: 552909).
- SARS-CoV-2 refers to the newly-emerged Sarbecovirus which was identified as the cause of a serious and worldwide outbreak of severe acquired pneumonia starting in the city of Wuhan (China).
- multiple mutations in the spike glycoprotein evolved and are evolving, including mutations that are located in the spike SI subunit.
- a skilled person is able to find information about the SARS-CoV-2 variants and their sequences in the relevant literature and databases, such as the SARS-CoV-2 genetic lineage tracing system funded by NIH (https://outbreak.info/).
- a SARS-CoV-2 variant may comprise a mutation at one or more positions selected from N439, K417, S477, L452, T478, E484, P384, N501 and D614 (relative to the SARS-CoV-2 spike amino acid sequence as defined in SEQ ID NO:93).
- SARS-CoV-2 variants include a SARS-CoV-2 variant comprising a mutation at position N501 such as a N501Y variant (e.g. SARS-CoV-2 Alpha variant); a SARS-CoV-2 variant comprising a mutation at positions N501 and E484 such as aN501Y and E484K variant (e.g.
- SARS-CoV-2 Alpha + E484K variant a SARS-CoV-2 variant comprising a mutation at positions K417, E484 and N501 such as a K417N, E484K and N501Y variant
- a SARS-CoV-2 variant comprising a mutation at positions P384, K417, E484 and N501 such as a P384L, K417N, E484K and N501Y variant
- SARS-CoV-2 beta + P384L variant a SARS-CoV-2 variant comprising a mutation at positions L452 and E484 such as a L452R and E484Q variant
- SARS-CoV-2 kappa variant a SARS-CoV-2 variant comprising a mutation at positions L452 and T478 such as a L452R and T478K variant (e.g. SARS-CoV-2 delta variant); a SARS-CoV-2 variant comprising a mutation at position L452 such as a L452R variant (e.g. SARS-CoV-2 epsilon variant); a SARS-CoV-2 variant comprising a mutation at position K417 such as a K417T variant (e.g. SARS-CoV-2 gamma variant); a SARS-CoV-2 variant comprising a mutation at position D614 such as a D614G variant (e.g.
- SARS-CoV-2 D614G variant SARS-CoV- 2 Omicron BA.1 variant or SARS-CoV-2 Omicron BA.2 variant
- a SARS-CoV-2 variant comprising a mutation at positions K147, W152R, F157, 1210, G257, D339, G446 and N460 such as a K147E, W152R, F157L, I210V, G257S, D339H, G446S and N460K variant
- SARS-CoV-2 Omicron BA.4/BA.5 variant a SARS-CoV-2 variant comprising a mutation at positions R346 and N658 such as a R346T and N658S variant
- a SARS-CoV-2 variant comprising a mutation at position R346 such as a R346T variant
- a SARS-CoV-2 variant comprising a mutation at positions R346, K444 and N460 such as a R346T, K444T and N460K variant
- SARS-CoV-2 Omicron BQ.1.1 variant a SARS-CoV-2 variant comprising a mutation at positions V83, Y144, H146, QI 83, V213, R346, L368, V445, G446, N460, F486 and F490 such as a V83A, Y144-, H146Q, Q183E, V213E, R346T, L368I, V445P, G446S, N460K, F486S and F490S variant (e.g.
- SARS-CoV-2 Omicron XBB variant or a V83A, Y144-, H146Q, Q183E, V213E, R346T, L368I, V445P, G446S, N460K, F486P and F490S variant (e.g.
- SARS-CoV-2 Omicron XBB.1.5 (-G252V) variant a SARS-CoV-2 variant comprising a mutation at positions V83, Q52, Y144, H146, F157, Q183, V213, G252, R346, L368, V445, G446, L452, F456, N460, F486 and F490 such as a V83A, Q52H, Y144-, H146Q, F157L, Q183E, V213E, G252V, R346T, L368I, V445P, G446S, L452R, F456L, N460K, F486P and F490S variant (e.g.
- SARS-CoV-2 HV.l a SARS-CoV-2 variant comprising a mutation at positions V83, Q52, Y144, H146, Q183, V213, G252, R346, L368, V445, G446, F456, N460, F486 and F490 such as a V83A, Q52H, Y144-, H146Q, Q183E, V213E, G252V, R346T, L368I, V445P, G446S, F456L, N460K, F486P and F490S variant (e.g.
- SARS-CoV-2 EG5.1 a SARS-CoV-2 variant comprising a mutation at positions T19, R21, L24, P25, P26, A27, S50, H69, V70, V127, G142, Y144, F157, R158, N211, L212, V213, L216, H245, A264, 1332, G339, K356, , S371, S373, S375, T376, R403, D405, R408, K417, N440, V445, G446, N450, L452, N460, S477, T478, N481, V483, E484, F486, Q498, N501, Y505, E554, A570, D614, P621, H655, N679, P681, N764, D796, S939, Q954, N969 and Pl 143 such as a T19I, R21T, L24S, P25-, P26-, A27-, S50L, H69-, V70-, ,
- SARS-CoV-2 BA.2.86.1 The Alpha variant (also known as B.1.1.1.7 lineage) of SARS-CoV-2 was first detected in the UK late 2020 and was one of the first reported variants of concern of SARS-CoV-2; a SARS-CoV-2 variant comprising a mutation at position F456 such as a F456L mutation (e.g.
- the Omicron variant of SARS-CoV-2 was first identified in South Africa and Botswana and was reported to the World Health Organization (WHO) on November 24, 2021, as a novel variant (Fan et al. 2022. Signal Transduct Target Ther. 7: 141).
- the Omicron variant is not a single strain, but evolved into at least three lineages, including BA.1, BA.2, and BA.3. Up to 60 mutations have been identified in the BA. 1 lineage, with as many as 38 of these occurring in the spike (S) protein, one in the envelope (E) protein, two in the membrane (M) protein, and six in the nucleocapsid (N) protein.
- BA.2 lineage possesses 57 mutations, with 31 in the S protein, of which the N-terminus is significantly different from that of BA. l.
- SARS-CoV-2 as used herein covers both the original strain identified in Wuhan as well as variants thereof.
- Khosta-2 refers to Sarbecovirus found in Russian horseshoe bats, which was found to be capable of using human ACE2 to facilitate entry (Seifert et al. 2022 PLoS Pathog 18(9):el010828).
- the binding agents, in particular the antibodies and antibody fragments (3) specifically bind or bind to spike protein of a Sarbecovirus such as SARS-CoV-2 spike protein, SARS-CoV-1 spike protein or Khosta-2 spike protein, in particular the binding agents, in particular the antibodies and antibody fragments, (4) specifically bind or bind to S2 subunit, or to a part of the S2 subunit, of the Sarbecovirus spike protein, more particularly, the binding agents, in particular the antibodies and antibody fragments, (22) specifically bind or bind to or within a region of the S2 subunit located from amino acid E 1188 to amino acid Y 1206, preferably a region located from amino acid N 1192 to amino acid Y1206 or a region located from amino acid El 188 to amino acid L1203, more preferably a region located from amino acid N1192 to amino acid LI 203, even more preferably a region located from amino acid N1194 to amino acid L1203, most preferably a region located from amino acid N1194 to amino acid Q1201 of the S
- the binding agents in particular the antibodies and antibody fragments, (23) specifically bind or bind to or within a region of spike protein of a Sarbecovirus or S2 subunit of the Sarbecovirus spike protein corresponding to the region from amino acid E 1188 to amino acid Y 1206, preferably amino acid N1192 to amino acid Y 1206 or amino acid El 188 to amino acid LI 203, more preferably amino acid N1192 to amino acid L1203, even more preferably amino acid N1194 to amino acid L1203, most preferably amino acid N1194 to amino acid Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93.
- the binding agents in particular the antibodies and antibody fragments, (5) specifically bind or bind to heptad repeat 2 (HR2) domain, or to a part of the HR2 domain, of (the S2 subunit of) the Sarbecovirus spike protein.
- HR2 heptad repeat 2
- the binding agents in particular the antibodies and antibody fragments, (6) specifically bind or bind to or within a region of the HR2 domain proximal to the viral membrane, preferably a region located from amino acid Al 174 to amino acid E1202, more preferably a region located from amino acid 11179 to amino acid El 202, even more preferably a region located from amino acid DI 184 to amino acid El 202, still more preferably a region located from amino acid El 188 to amino acid E1202 or a region located from amino acid VI 189 to amino acid E1202, yet more preferably a region located from amino acid N1194 to amino acid El 202, most preferably a region located from amino acid N1194 to amino acid Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, or (7) specifically bind or bind to a region of the HR2 domain (or of the S2 subunit) corresponding to the region from amino acid El 188 to amino acid Y1206 of the SARS-CoV-2 spike protein as defined in S
- the binding agents in particular the antibodies and antibody fragments, (8) specifically bind or bind to at least one, at least two, at least three, at least four, at least five, at least six, at least seven or all, of the amino acid residues N1192, N1194, SI 196, LI 197, DI 199, L1200, Q1201 and E1202, of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, preferably to at least one, at least two, at least three, at least four or all of the amino acid residues N1194, SI 196, DI 199, Q1201 and E1202 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, more preferably to at least one, at least two, at least three or all of the amino acid residues N1194, S 1196, D1199 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, most preferably to at least one or both of the amino acid residues SI 196 and Q1201
- the binding agents specifically bind or bind to at least one, at least two, at least three, at least four, at least five, at least six, at least seven or all, amino acid residue(s) of spike protein of a Sarbecovirus or S2 subunit or HR2 domain of the Sarbecovirus spike protein corresponding to the amino acid residues N1192, N1194, SI 196, LI 197, DI 199, LI 200, Q1201 and E1202 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, preferably to at least one, at least two, at least three, at least four or all amino acid residue(s) corresponding to the amino acid residues N1194, SI 196, DI 199, Q1201 and E1202 of the SARS-CoV-2 spike protein as defined in SEQ ID NO: 93, more preferably to at least one, at least two, at least three or all amino acid residues(s) corresponding to the amino acid residues N1194, SI 196, DI 199, Q1201 and E1202 of the
- the binding agents specifically bind or bind to the amino acid residues SI 196 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93 or to the amino acid residues of spike protein corresponding to said amino acid residues of SI 196 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, optionally to the amino acid residues N1194, S 1196, D1199 and Q1201ofthe SARS-CoV-2 spike protein as defined in SEQ ID NO:93 or to the amino acid residues of spike protein corresponding to said amino acid residues N1194, SI 196, DI 199 and Q1201of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93.
- (26) at least one, at least two, at least three, at least four, at least five, at least six, at least seven or all, of the amino acid residues N1192, N1194, SI 196, LI 197, DI 199, L1200, and Q1201 and E1202, of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, preferably at least one, at least two, at least three, at least four or all of the amino acid residues N 1194, SI 196, DI 199, Q1201 and E1202 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, more preferably at least one, at least two, at least three or all of the amino acid residues N1194, SI 196, DI 199 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, most preferably at least one or both of the amino acid residues SI 196 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO: 93,
- Assessment of the binding site may be evaluated by determining the crystal structure of a complex of the binding agent, in particular the antibody or antibody fragment, and a spike protein, or an S2 subunit or a peptide comprising a HR2 domain, for example by applying the crystal structure determination method as shown in the examples, and/or by selection and analysis of viral escape variants/mutants, for example by applying the viral escape selection method as shown in the examples, and/or by analysing hydrogen-deuterium exchange on recombinant spike protein (or S2 subunit or HR2 containing peptides) in the presence and absence of the binding agent, for example by applying the hydrogen-deuterium exchange method monitored by mass spectrometry (HDX-MS method) as shown in the examples.
- HDX-MS method mass spectrometry
- these amino acid residues are conserved between different clades of Sarbecoviruses. in particular between clade 1, clade 2, and clade 3 Sarbecoviruses .
- the binding agents, in particular the antibodies or antibody fragments, (9) do not bind to the RBD of the Sarbecovirus spike protein.
- the binding agents, in particular the antibodies and antibody fragments (29) specifically bind or bind to a quaternary epitope of the spike protein.
- the binding agents, in particular the antibodies and antibody fragments (30) specifically bind or bind to a trimeric HR2 domain (or a trimeric S2 subunit or a trimeric spike protein).
- the binding agents, in particular the antibodies and antibody fragments (31) specifically bind or bind to a quaternary epitope within a trimeric HR2 domain (or a trimeric S2 subunit or a trimeric spike protein).
- the binding agents, in particular the antibodies and antibody fragments, (32) specifically bind or bind to a quaternary epitope located within two adjacent HR2 domains or helices.
- the binding agents in particular the antibodies and antibody fragments, (33) specifically bind or bind to a quaternary epitope comprising or consisting of one or more interacting amino acid residues as described herein in one HR2 domain or helix as well as one or more interacting amino acid residues as described herein in an adjacent HR2 domain or helix.
- quaternary epitope refers to a conformational epitope whose structure depends upon or is enhanced by the arrangement of multiple protomers or monomers into a multimeric complex.
- a quaternary epitope may be located in a single protein (or monomer) of a multimeric complex; or it may span multiple protomers, being formed de novo by their interaction.
- Specific binding or binding to a quaternary epitope or a multimeric protein can be assessed by evaluating binding to monomeric and/or (stabilized) multimeric protein by means of an Enzyme Linked Immunosorbent Assay (ELISA) assay, for example by applying the ELISA assay as shown in the examples.
- ELISA Enzyme Linked Immunosorbent Assay
- Stabilization of trimeric spike protein may be achieved by fusing the spike protein to the foldon domain of the trimeric protein fibritin from bacteriophage T4.
- Correlation between binding to the monomeric protein and density of the monomeric protein at elevated densities of the monomeric protein only, such as at a density of 1.0 ng/mm 2 or more, preferably 1.2 ng/mm 2 or more, or 1.5 ng/mm 2 or more, may be indicative for specific binding or binding to a multimeric conformation of the protein.
- enhanced binding to multimeric protein compared to the monomeric protein may indicate specific binding or binding to the multimeric protein.
- the binding agents upon binding to the trimeric spike protein, in particular to a quaternary epitope within the trimeric spike protein, the binding agents, in particular the antibodies and antibody fragments, described herein may stabilize the profusion conformation of the spike protein. More particularly, the binding agent may stabilise or lock the HR2 coiled-coil.
- the binding agents may prevent the unravelling of the HR2 coiled-coil, which is considered a critical early step in the spike-controlled membrane fusion process; or the binding agents may interfere with or block migration of the HR2 alpha helices towards the extended HR1 alpha helices, which is considered a critical step in the refolding of the spike protein from a prehairpin intermediate to a postfusion conformation; and/or the binding agents may prevent the completion of the 6 helix bundle formation, which is considered crucial for the fusion process.
- the binding agents in particular the antibodies and antibody fragments (35) are capable of stabilizing the prefusion conformation of spike protein of a Sarbecovirus.
- the binding agents in particular the antibodies and antibody fragments are (36) capable of stabilizing the HR2 coiled-coil.
- SARS-CoV-2 contains as structural proteins the spike (S) protein, the envelope (E) protein, the membrane (M) protein, and the nucleocapsid (N) protein.
- S spike
- E envelope
- M membrane
- N nucleocapsid
- nspl-16 sixteen nonstructural proteins have been discerned, which are involved in replication and modifying the host defense.
- the Nspl2 protein corresponds to a RNA-dependent RNA polymerase (RdRp).
- spike or S protein which is a transmembrane glycoprotein forming homotrimers protruding from the viral surface and giving the virus a crownlike look.
- the spike protein has two subunits: SI and S2.
- the SI subunit comprises an N-terminal domain (NTD), a receptor binding domain (RBD), and subdomains 1 and 2 (SD1, SD2).
- NTD N-terminal domain
- RBD receptor binding domain
- SD1, SD2 subdomains 1 and 2
- the SI subunit is involved in host receptor binding.
- the spike protein binds to human host cell receptor angiotensin-converting enzyme 2 (ACE2) via the receptor binding domain (RBD) present in the S 1 subunit.
- ACE2 human host cell receptor angiotensin-converting enzyme 2
- the S2 subunit is involved in fusing the membranes of viruses and host cells and viral entry, and comprises multiple domains: an S2’ protease cleavage site (cleavage by a host protease required for fusion), a fusion peptide (FP), a heptad repeat 1 (HR1) domain, a central helix (CH) domain, a connector domain (CD), a heptad repeat 2 (HR2) domain, a transmembrane (TM) domain, and a cytoplasmic tail (CT) domain (Wang et al. (2020). Front Cell Infect Microbiol 10:587269).
- the S protein normally exists in a prefusion conformation.
- SI and S2 cleaved at the S1-S2 furin cleavage site during biosynthesis, remain non-covalently bound to each other - this is different from SARS-CoV in which SI and S2 remain uncleaved.
- PDB: 6VXX the closed state of the S protein
- the 3 RBD domains in the trimer do not protrude from the trimer
- the open state (PDB:6VYB), or “up” conformation, one of the RBD does protrude from the trimer.
- the S-trimer ectodomain with triangular cross-section has a length of approximately 160- Angstrom wherein the S 1 domain adopts a V-shaped form. Sixteen of the 22 N-linked glycosylation sites per protomer appear glycosylated (Walls et al. (2020) Cell 180:281-292).
- the S 1 subunit of the S protein binds with ACE2 through its RBD region to promote the formation of endosomes, which triggers viral fusion activity.
- TMPRSS2 transmembrane protease serine subtype 2
- FP fusion peptide
- the viral membrane is so pulled toward the host cell membrane and tightly binds to it, and the two membranes fuse, resulting in the release of the viral genome into the host cell (Huang et al. (2020) Acta Pharmalogica Sinica 41: 1141-1149).
- spike protein refers to the spike protein of a Sarbecovirus , and can refer to specific S proteins such as SARS-CoV-2 S protein and SARS- CoV-1 S protein.
- spike protein and SARS-CoV-2 spike protein include protein variants of Sarbecovirus or SARS-CoV-2 spike protein isolated from different Sarbecovirus or SARS-CoV-2 isolates, as well as recombinant Sarbecovirus or SARS-CoV-2 spike protein, or a fragment thereof.
- the terms also encompass Sarbecovirus spike protein or SARS-CoV-2 spike protein coupled to, for example, a histidine tag, mouse or human Fc, or a signal sequence.
- SARS-CoV-2 spike protein sequence can be found under/corresponds with or to Genbank Accession: QHQ82464, version QHQ82464.1; and is also defined herein as SEQ ID NO:93:
- SARS-CoV-2 spike protein HR2 domain corresponds with/to amino acids 1169-1202 of SEQ ID NO:93 and as depicted hereafter (SEQ ID NO:94): ISGINASVVNIQKEIDRLNEVAKNLNESLIDLQE (SEQ ID NO:94).
- the SARS-CoV-2 spike protein TM domain corresponds with/to amino acids 1214-1237 of SEQ ID NO:93.
- a region of the HR2 domain “proximal to the viral membrane” refers to a region within the HR2 domain that is within 40 amino acids from the viral membrane.
- the Sars-CoV-1 spike protein sequence can be found under/corresponds with or to GenBank accession NP_828851.1; and is also defined herein as SEQ ID NO:95.
- the SARS-CoV-1 spike protein HR2 domain corresponds with/to amino acids 1151-1184 of SEQ ID NO:95 and as depicted in SEQ ID NO:94.
- the amino acids and amino acid numbering referred to herein is relative to/corresponding to the SARS-CoV-2 spike protein as defined in SEQ ID NO:93; corresponding amino acids in spike proteins or spike protein fragments, domains or regions of other Sarbecoviruses can be easily determined by aligning multiple amino acid sequences.
- Angiotensin converting enzyme 2 refers to mammalian protein belonging to the family of dipeptidyl carboxydipeptidases, and sometimes classified as EC:3.4.17.23.
- the genomic location of the human ACE2 gene is on chrX: 15,561,033- 15,602,158 (GRCh38/hg38; minus strand), or alternatively on chrX: 15,579,156-
- ACE2 acts as a receptor for at least human coronaviruses SARS-CoV and SARS-CoV-2, and NL63/HCoV-NL63 (also known as New Haven coronavirus).
- UniProtKB identifier of human ACE2 protein Q9BYF1.
- Isoform 1 (identifier: Q9BYF1-1) has been chosen as the canonical! sequence.
- a further functional characteristic of the binding agents, in particular the antibodies and antibody fragments, described herein is that they are (10) capable of neutralizing a Sarbecovinis. in particular (11) capable of neutralizing any one or both, preferably both, of SARS-CoV-2 and SARS-CoV-1.
- a "neutralizing binding agent” or a “neutralizing antibody” refers to a binding agent or antibody that binds to a Sarbecovinis. in particular SARS-CoV- 2 and/or SARS-CoV-1, to inhibit or suppress the ability of the Sarbecovinis. or SARS-CoV-2 or SARS-CoV-1, to initiate and/or perpetuate an infection in a host.
- Neutralizing binding agents or antibodies may, for example, interfere with binding of a Sarbecovinis such as SARS-CoV-2 or SARS-CoV-1 to a host receptor, in particular ACE2; and/or with viral entry, e.g. by inducing SI shedding and/or by interfering with viral fusion.
- the binding agents and antibodies according to the current invention are neutralizing, inhibiting, blocking or suppressing a Sarbecovinis infection.
- the binding agents, in particular the antibodies and antibody fragments, described herein (44) do not modulate or interfere with SI shedding.
- the binding agents, in particular the antibodies and antibody fragments, described herein (12) do not induce SI shedding.
- the binding agents, in particular the antibodies and antibody fragments, described herein (45) do not prevent SI shedding.
- the binding agents, in particular the antibodies and antibody fragments, described herein are (13) capable of inhibiting spike-mediated syncytia formation. Consequently, the binding agents, in particular the antibodies and antibody fragments, may be (14) capable of inhibiting viral fusion and, without wishing to be bound by any theory, may as such not allow the Sarbecovinis to complete the infection process into a host cell.
- the binding agents, in particular the antibodies and antibody fragments, described herein (46) do not prevent HR1 unfolding.
- the binding agents in particular the antibodies and antibody fragments, described herein (47) do not prevent folding of HR1 onto HR2 (e.g. as during formation of a S2 6 helix bundle).
- the binding agents, in particular the antibodies and antibody fragments, according to the invention are capable of neutralizing a Sarbecovirus infection potently.
- Neutralizing activity can be measured using a standard neutralization assay as known to one of skill in the art, including, without limitation, a pseudovirus neutralization assay and a plaque reduction test. Exemplary methods for performing such neutralization assays are described herein in the examples. Neutralizing activity can also be evaluated by measuring one or more indicators of a Sarbecovirus , or SARS-CoV-2 or SARS-CoV-1, infection, such as syncytia formation between cells expressing a Sarbecovirus spike protein and cells expressing the Sarbecovirus receptor ACE2 .
- the binding agents are (15) capable of neutralizing a Sarbecovirus, in particular SARS-CoV-2 and/or SARS-CoV-1, with a half maximum inhibitory concentration or 50% inhibitory concentration (IC50) of 100 ng/ml or less, preferably 50 ng/ml or less or 20 ng/ml or less, more preferably 15 ng/ml or less or 10 ng/ml or less, even more preferably 1 ng/ml or less, preferably as determined in a Sarbecovirus spike protein pseudovirus neutralization assay such as a vesicular stomatitis virus JSV)-Sarbecovirus spike protein pseudovirus neutralization assay, more preferably as determined in a SARS-CoV-2 spike protein and/or SARS-CoV-1 spike protein pseudovirus neutralization assay such as a VSV-SARS- CoV-2 spike protein pseudovirus neutralization assay or a VSV-SARS-CoV-1 spike protein pseudovirus neutralization neutral
- the pseudovirus neutralization assay may be based on pseudotyped VSV-delG virus containing the spike protein of a Sarbecovirus such as SARS-CoV-2 spike protein, SARS-CoV-2 variant spike protein or SARS-CoV-1 spike protein.
- a Sarbecovirus such as SARS-CoV-2 spike protein, SARS-CoV-2 variant spike protein or SARS-CoV-1 spike protein.
- half maximum inhibitory concentration or “IC50” refers to a quantity such as a concentration of a binding agent or antibody required for 50% neutralization of the Sarbecovirus.
- the binding agents are (16) capable of neutralizing at least one SARS-CoV-2 variant such as a SARS-CoV-2 variant comprising a mutation at position D614 (relative to the SARS-CoV-2 spike amino acid sequence as defined in SEQ ID NO:93) such as a D614G variant, in particular at least any one or more, preferably all, of SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta variant, SARS-CoV-2 Epsilon B. 1.427 variant, SARS-CoV-2 Epsilon B.
- SARS-CoV-2 variant such as a SARS-CoV-2 variant comprising a mutation at position D614 (relative to the SARS-CoV-2 spike amino acid sequence as defined in SEQ ID NO:93) such as a D614G variant, in particular at least any one or more, preferably all, of SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta
- SARS-CoV-2 Eta variant
- SARS-CoV-2 Iota variant variant
- SARS-CoV-2 Kappa variant variant
- SARS- CoV-2 Mu variant variant
- SARS-CoV-2 Zeta variant SARS-CoV-2 B.1.617.3 variant
- SARS-CoV-2 Omicron BA.l variant SARS-CoV-2 Omicron BA.2 variant
- SARS-CoV-2 Omicron BA.5 variant SARS-CoV-2 Omicron BA.2.75.2 variant
- SARS-CoV-2 Omicron BA.4.6 variant SARS-CoV-2 Omicron BF.7 variant
- SARS-CoV-2 Omicron XBB variant SARS-CoV-2 Omicron XBB. 1.5 (-G252V) variant, SARS-CoV-2 Omicron CH. 1.1 variant, SARS- CoV-2 Omicron EG.5.1 variant, SARS-CoV-2 Omicron BA.2.86.1 variant SARS-CoV-2 Omicron HV. l variant, SARS-CoV-2 Omicron XBB. 1.5 variant, SARS-CoV-2 Omicron FL.
- SARS-CoV-2 Omicron XBB.2.3 variant SARS-CoV-2 Omicron XBB.1.16 variant, SARS-CoV-2 Omicron HK.3 variant, SARS-CoV-2 Omicron BA.2.74 variant, SARS-CoV-2 Omicron JN.1 variant, SARS-CoV-2 Omicron KP.2.3 variant, SARS-CoV-2 Omicron KP.3 variant, SARS-CoV-2 Omicron KP.3.1.1 variant, SARS-CoV-2 Omicron KP.1.1 variant, SARS-CoV-2 Omicron LB.l variant and SARS-CoV-2 Omicron XEC variant.
- the binding agents in particular the antibodies and antibody fragments described herein are characterized in that they are (17) capable of neutralizing SARS-CoV-2 Alpha variant, (78) capable of neutralizing SARS-CoV- 2 Beta variant, (67) capable of neutralizing SARS-CoV-2 Gamma variant, (79) capable of neutralizing SARS-CoV-2 Delta variant, (80) capable of neutralizing SARS-CoV-2 Epsilon B.1.427 variant, (81) capable of neutralizing SARS-CoV-2 Epsilon B.1.429 variant, (68) capable of neutralizing SARS-CoV-2 Eta variant, (69) capable of neutralizing SARS-CoV-2 Iota variant, (70) capable of neutralizing SARS-CoV-2 Zeta variant, (82) capable of neutralizing SARS-CoV-2 Kappa variant, (71) capable of neutralizing SARS-CoV-2 Mu variant, (72) capable of neutralizing SARS- CoV-2 B.1.617.3 variant, (18) capable of neutralizing SARS-CoV-2
- the binding agents are characterized in that they are (86) capable of neutralizing a SARS-CoV-2 variant comprising a F456L mutation, in particular capable of neutralizing one or more, preferably all, of SARS CoV-2 Omicron FL. 1.5.1, SARS CoV-2 Omicron EG.5.1, SARS CoV-2 Omicron HK.3, SARS CoV-2 Omicron HV.l, SARS CoV-2 Omicron KP.1.1, SARS CoV-2 Omicron KP.2.3, SARS CoV-2 Omicron KP.3, SARS CoV-2 Omicron KP.3.1.1, SARS CoV-2 Omicron LB.
- SARS CoV-2 Omicron XEC variants with an IC50 of 100 ng/ml or less, preferably 50 ng/ml or less or 20 ng/ml or less, more preferably 15 ng/ml or less or 10 ng/ml or less, optionally as determined in a SARS-CoV-2 variant spike pseudovirus neutralization assay such as a VSV-SARS-CoV-2 variant spike pseudovirus neutralization assay.
- the binding agents are characterized in that they are (87) capable of neutralizing a SARS-CoV-2 variant comprising a Q493E mutation, in particular capable of neutralizing one or more, preferably all, of SARS CoV-2 Omicron KP.3, SARS CoV-2 Omicron KP.3.1.1, and SARS CoV-2 Omicron XEC variants, with an IC50 of 100 ng/ml or less, preferably 50 ng/ml or less or 20 ng/ml or less, more preferably 15 ng/ml or less or 10 ng/ml or less, optionally as determined in a SARS-CoV-2 variant spike pseudovirus neutralization assay such as a VSV- SARS-CoV-2 variant spike pseudovirus neutralization assay.
- the binding agents are characterized in that they are (88) capable of neutralizing a SARS-CoV-2 variant comprising F456L and Q493E mutations, in particular capable of neutralizing one ore more, preferably all, of SARS CoV-2 Omicron KP.3, SARS CoV-2 Omicron KP.3.1.1, and SARS CoV-2 Omicron XEC variants, with an IC50 of 100 ng/ml or less, preferably 50 ng/ml or less or 20 ng/ml or less, more preferably 15 ng/ml or less or 10 ng/ml or less, optionally as determined in a SARS-CoV-2 variant spike pseudovirus neutralization assay such as a VSV- SARS-CoV-2 variant spike pseudovirus neutralization assay.
- the binding agents, in particular the antibodies and antibody fragments, described herein are further characterized in that they are (14) capable of inhibiting viral fusion.
- the binding agents, in particular the antibodies and antibody fragments, described herein are (13) capable of inhibiting spike-mediated syncytia formation, more particularly they are (20) capable of inhibiting the formation of syncytia between cells expressing a Sarbecovirus spike protein, such as SARS-CoV- 2 and/or SARS-CoV-1 spike protein, and cells expressing the Sarbocovirus host receptor, in particular ACE2 receptor.
- viral fusion refers to fusion of a viral membrane and a host cell membrane.
- Viral fusion assays are well-known to the skilled person and exemplary methods for performing such methods are described herein in the examples.
- complete inhibition is not required and a skilled person is able to identify binding agents, antibodies and antibody fragments that significantly inhibit viral fusion or spike-mediated syncytia formation.
- binding agents, in particular antibodies and antibody fragments, as described herein (21) may induce at least 50% inhibition, preferably at least 60%, at least 70%, at least 80% or at least 90% inhibition.
- a Sarbecovirus binding agent in particular a Sarbecovirus antibody or antibody fragment, as described hereinabove are combined such as to characterize such binding agent, antibody or antibody fragment, e.g. to bind or specifically bind to the Sarbecovirus spike protein HR2 domain and to be capable of neutralizing a Sarbecovirus, in particular at least one or both of SARS-CoV-2 (such as any one or more of SARS-CoV-2 Wuhan strain, SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta variant, SARS-CoV-2 Epsilon B.
- SARS-CoV-2 such as any one or more of SARS-CoV-2 Wuhan strain, SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta variant, SARS-CoV
- SARS-CoV-2 Epsilon B.1.429 variant SARS-CoV-2 Eta variant, SARS-CoV-2 Iota variant, SARS- CoV-2 Zeta variant, SARS-CoV-2 Kappa variant, SARS-CoV-2 Mu variant, SARS-CoV-2 B. 1.617.3 variant, SARS-CoV-2 Omicron BA. l variant, SARS-CoV-2 Omicron BA.2 variant, SARS-CoV-2 Omicron BA.5 variant, SARS-CoV-2 Omicron BA.2.75.2 variant, SARS-CoV-2 Omicron BAA.6 variant, SARS-CoV-2 Omicron BF.7 variant, SARS-CoV-2 Omicron BQ.
- SARS-CoV-2 Omicron XBB variant SARS-CoV-2 Omicron XBB.1.5 variant (-G252V), SARS-CoV-2 Omicron CH. 1.1 variant, SARS-CoV-2 Omicron EG.5.1 variant, SARS-CoV-2 Omicron BA.2.86.1 variant SARS-CoV-2 Omicron HV. l variant, SARS-CoV-2 Omicron XBB.1.5 variant, SARS-CoV-2 Omicron XBB.2.3 variant, SARS-CoV-2 Omicron XBB. 1.16 variant, SARS- CoV-2 Omicron HK.3 variant, SARS-CoV-2 Omicron BA.2.74 variant, SARS-CoV-2 Omicron FL.
- IC50 50% inhibitory concentration
- binding agent, antibody or antibody fragment may further be characterized to be capable of inhibiting spike-mediated syncytia formation between cells expressing the Sarbecovirus spike protein and cells expressing the angiotensin-converting enzyme 2 (ACE2) receptor and/or to be capable of inhibiting viral fusion; and/or by not binding a Middle East respiratory syndrome coronavirus (MERS-CoV).
- ACE2 angiotensin-converting enzyme 2
- the binding agents in particular the antibodies and antibody fragments, described herein are further characterized in that (48) they have an advantageously low isoelectric point, in particular a lower isoelectric point compared with the ISVD defined by SEQ ID NO:2.
- the pl of the binding agent, in particular the antibody or antibody fragment is from 7.5 to 9.5, or from 7.6 to 9.4, or from 7.7 to 9.3, or from 7.8 to 9.3, or from 7.9 to 9.2, preferably from 7.5 to 9.1, or from 7.6 to 9.1, or from 7.7 to 9.1, or from 7.7 to 9.1, or from 7.8 to 9.1, or from 7.9 to 9.1.
- pl refers to the “isoelectric point” of a molecule, in particular a binding agent, antibody or antibody fragment as described herein, and is the pH at which a particular molecule or surface carries no net electrical charge. Antibodies with lower pl values may have longer serum halflives.
- Methods for determining the isoelectric point of a protein are well known to those of one skilled in the art. By way of example, many suitable computer programs for calculating the pl of a protein are generally known in the art, such as EMBOSS iep software, written by Alan Bleasby, available at HGMP-RC, Genome Campus, Hinxton, Cambridge CB10 1SB, UK.
- the binding agents in particular the antibodies and antibody fragments, (50) exhibit advantageously low non-specific binding, in particular reduced non-specific binding compared with the ISVD defined by SEQ ID NO:2, in particular (51) low non-specific binding to mammalian cells, more particularly human cells, such as reduced non-specific binding to mammalian cells, more particularly human cells, compared with the ISVD defined by SEQ ID NO:2.
- non-specific binding or “off-target” of a molecule in particular a binding agent antibody or antibody fragment as described herein, is meant herein the ability of the molecule to bind to a non-specific target.
- a binding agent antibody or antibody fragment as described herein.
- the binding agents in particular the antibodies and antibody fragments, (52) exhibit advantageously low immunogenicity, in particular reduced immunogenicity compared with the ISVD defined by SEQ ID NO:2.
- immunogenicity of a molecule, in particular a binding agent, antibody or antibody fragment as described herein, is meant herein the ability of the molecule to elicit an (undesired) immune response (humoral or cellular) when administered to a recipient. Immunogenicity of a molecule, in particular a binding agent, antibody or antibody fragment, can be determined by in silico analysis as described in the experimental section.
- the binding agents in particular the antibodies and antibody fragments, (53) exhibit advantageously low hydrophobicity, in particular reduced hydrophobicity compared with the ISVD defined by SEQ ID NO:2.
- the binding agents, in particular the antibodies and antibody fragments exhibit advantageously low positive charge patches, in particular reduced positive charge patches compared with the ISVD defined by SEQ ID NO:2.
- the retention time of the binding agents, in particular the antibodies and antibody fragments is 15 min or lower, preferably between 5 and 15 min, or between 6 and 14 min, as determined by strong cation exchange hydrophilic interaction chromatography (SCX), in particular SCX using the conditions specified in the examples section, more particularly SCX using a MAbPac SCX- 10 column, a mobile phase A consisting of 20mM MES pH 5.6 and a mobile phase B consisting of 20 mM HEPES and 120 mM NaCl pH 8.0 and using a linear gradient from 100% A to 100% B in 18 min.
- SCX strong cation exchange hydrophilic interaction chromatography
- the binding agents in particular the antibodies and antibody fragments, are (56) advantageously stable, in particular exhibit (57) improved protein stability compared to the ISVD defined by SEQ ID NO:2.
- the antibody comprises one or more ISVDs as described herein (or variants thereof as described herein) wherein the one or more ISVD (or variant thereof as described herein) is bound or fused to an Fc domain.
- an “Fc domain” as used herein refers to the fragment crystallizable region (Fc region) of a conventional antibody, which is the tail region known to interact with cell surface receptors called Fc receptors and some proteins of the complement system. Said Fc domain is composed of two identical protein fragments, derived from the second and third constant domains of the antibody's two heavy chains. All conventional antibodies comprise an Fc domain, hence, the Fc domain may be an Fc domain derived from or as a variant of the IgG, IgA or IgD antibody Fc regions, even more specifically derived from an IgGl, IgG2 or IgG4 antibody Fc region.
- the hinge region of IgG2 may be replaced by the hinge of human IgGl to generate ISVD fusion constructs, and vice versa.
- Fc variants with known half-life extension may be used such as the M257Y/S259T/T261E variant (also known as YTE), the LS variant (M428L combined with N434S), the DHS variant (comprising L309D, Q311H and N434S substitutions) or the LA variant (comprising M428L and N434A substitutions).
- Fc variants are known to enhance FcRn binding affinity at an endosomal acidic pH and dissociation at physiological pH, thereby prolonging the circulating halflife of the antibody or antibody fragment and mediating half-life extension.
- human Fc domains or humanized Fc domains may be used.
- Humanized forms include but are not limited to the IgG humanization variants known in the art, such as C-terminal deletion of Lysine, alteration or truncation in the hinge region, LALA (L234A and L235A) or LALAPG (L234A, L235A, and P329G) mutations, among other substitutions in the IgG sequence.
- one or more ISVDs (or variants thereof) as described herein are fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NOs: 82-86, as depicted hereafter:
- Fc domain comprising the LS substitutions:
- Fc domain comprising the YTE substitutions:
- Fc domain comprising the DHS substitutions:
- Fc domain comprising the LA substitutions:
- fused to refers in one aspect to “genetic fusion”, e.g., by recombinant DNA technology, as well as to “chemical and/or enzymatic conjugation” resulting in a stable covalent link between two nucleic acid molecules.
- inserted in wherein a fragment of one nucleic acid may be inserted in a second nucleic acid molecule by fusing or ligating the two sequences genetically, enzymatically or chemically.
- Peptides or polypeptides can likewise be fused or connected to one another, such as via peptide bonds or via linking one peptide to a side chain of an amino acid in a second peptide.
- Linkers may be used to fuse an ISVD, such as a herein identified ISVD (or variant thereof as described herein), to an Fc domain such as the human IgGl Fc domain or the LS variant thereof, or the YTE variant thereof, or the DHS variant thereof, or the LA variant thereof or an IgG2 Fc domain, preferably an Fc domain comprising or consisting of the sequence set forth in any of SEQ ID NO: 82- 86.
- the antibody comprising one or more ISVDs as described herein is in a “multivalent” and/or “multispecific” form formed by binding, e.g. chemically or by recombinant DNA techniques, together two or more identical or variant monovalent ISVDs (or variants or humanized forms thereof as described herein).
- Non-limiting examples of multivalent constructs include “bivalent” constructs, “trivalent” constructs, “tetravalent” constructs, and so on, respectively, comprising two, three or four ISVDs.
- the ISVDs comprised within a multivalent construct may be identical or different.
- the term “multispecific antibody” as used herein specifically refers to a multivalent antibody wherein at least one of the two or more ISVDs has a different specificity.
- Non-limiting examples of multi-specific constructs include “bi-specific” constructs, “tri-specific” constructs, “tetra-specific” constructs, and so on.
- any multivalent and multi-specific (as defined herein) antibody of the invention may be directed against two or more different antigens, for example against a Sarbecovirus and one as a half-life extension against Serum Albumin or Staphylococcal protein A (SpA) and/or against two or more different parts of a particular antigen, for example against two or more different parts, regions, subunits or domains of a Sarbecovirus spike protein.
- a Sarbecovirus for example against a Sarbecovirus and one as a half-life extension against Serum Albumin or Staphylococcal protein A (SpA) and/or against two or more different parts of a particular antigen, for example against two or more different parts, regions, subunits or domains of a Sarbecovirus spike protein.
- SpA Staphylococcal protein A
- an antibody in particular a multivalent and/or multispecific antibody, may comprise one or more binding agent, such as ISVD(s), as described herein (or variants thereof as described herein), and one or more binding agents, such as ISVD(s), capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD).
- binding agent such as ISVD(s)
- binding agents such as ISVD(s) capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD).
- ISVDs capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD) are described in WO 2021/156490 Al, WO 2022/167666 Al and WO 2022/238550 Al.
- the combination of at least two ISVDs capable of binding Sarbecovirus spike protein through interaction at 2 different regions of the spike protein, in particular the S2 subunit, more particularly the HR2 domain, and the RBD, in such multivalent and/or multispecific antibody may result in crossreactivity and potent prohibition of infection by Sarbecoviruses. and may further allow for reducing the risk to escape mutant virus emergence.
- the one or more ISVDs capable of binding to a Sarbecovirus spike protein RBD are capable of binding to or competing for the VHH72 epitope (or the epitope specifically bound by VHH72).
- the VHH72 epitope has been described in Wrapp et al. (2020, Cell 184: 1004-1015; WO 2021/156490 Al and WO 2022/238550 Al).
- An ISVD capable of competing for the VHH72 epitope refers to an ISVD that competes with VHH72 for binding to the spike protein as depicted in SEQ ID NO:93, or the RBD.
- an ISVD capable of binding to or competing for the VHH72 epitope may be characterized in that (i) it competes for human receptor (ACE-2 in the case of SARS- CoV-1 and -2) binding upon interaction to the RBD, and/or (ii) is not competing with an ISVD capable of binding to or competing with a VHH3.117 epitope as defined herein.
- Non-limiting examples of ISVDs capable of binding to or competing for the VHH72 epitope include VHH72 family members (including VHH72 (SEQ ID NOVO), VHH2.50, VHH3.17, VHH3.77, VHH3.115, VHH3.144 and VHHBE4), and variants, including VHH72(S56A), and humanized forms thereof; VHH3.83 family members (including VHH3.83 (also referred to as VHH83 herein) (SEQ ID NO:91)) and variants and humanized forms thereof; VHH3.38 family members and variants and humanized forms thereof; VHH3.55 family members and variants and humanized forms thereof; VHH3.36 family members and variants and humanized forms thereof; VHH3.149 family members and variants and humanized forms thereof; and VHH3.29 family members and variants and humanized forms thereof, as described in WO 2021/156490 Al and WO 2022/238550 Al.
- the one or more ISVDs capable of binding to a Sarbecovirus spike protein RBD are capable of binding to or competing for the VHH3.117 epitope (or the epitope specifically bound by VHH3. 117).
- the VHH3. 117 epitope has been described in WO 2022/167666 Al.
- I An ISVD capable of competing for the VHH3.117 epitope refers to an ISVD that competes with VHH3. 117 (SEQ ID NO:92) for binding to the spike protein as depicted in SEQ ID NO:93, or the RBD.
- binding of VHH3.117 to the spike protein as depicted in SEQ ID NO:93 is reduced with at least 30%, or at least 50%, or preferably at least 80% in strength in the presence of an ISVD capable of competing for the VHH3.117 epitope.
- an ISVD capable of binding to or competing for the VHH3.117 epitope may be characterized in that (i) it does not inhibit binding of the RBD with the human receptor (ACE-2 in the case of SARS-CoV-1 and -2), meaning that it allows binding of the receptor and the Sarbecovirus RBD when the ISVD itself is bound to the Sarbecovirus RBD, or alternatively, that the ISVD itself can bind to a Sarbecovirus RBD to which the receptor is bound, and/or (ii) is not competing with an ISVD capable of binding to or competing for the VHH72 epitope as defined herein.
- ACE-2 human receptor
- Non-limiting examples of ISVDs capable of binding to or competing for the VHH3.117 epitope include VHH3.117 family members (including VHH3.117 (SEQ ID NO:92), 3.42, 3.92, 3.94, 3.180) and variants and humanized forms thereof (as described in WO 2022/167666 Al); VHH3.89 family members and variants and humanized forms thereof (as described in WO 2021/156490); VHH3_183 family members and variants and humanized forms thereof; and VHH3C 80 family members and variants and humanized forms thereof (as described in WO 2022/238550 Al).
- the antibody in particular the multivalent and/or multispecific antibody, comprises more than one ISVD capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD), wherein at least one ISVD is capable of binding to or competing for the VHH72 epitope as defined herein, and wherein at least one ISVD is capable of binding to or competing for the VHH3.117 epitope as defined herein.
- RBD Sarbecovirus spike protein receptor binding domain
- the combination of at least two non-competing RBD targeting ISVDs (capable of binding the RBD of the spike protein through interaction at 2 non-competing, different regions of the RBD) and at least one S2 targeting ISVD in the antibody results in cross-reactivity and potent prohibition of infection by Sarbecoviruses , which advantageoulsy allows for further reducing the risk to mutational escape.
- Multivalent antibodies as described herein may be formed e.g. by connecting, such as chemically or by recombinant DNA techniques, the two or more ISVDs directly or via a linker, and/or through fusing (each of) the two or more ISVDs with an Fc domain.
- a single ISVD (or variant thereof) as described herein may be fused e.g. at its C- terminus to an Fc domain, such as an Fc domain comprising or consisting of the sequence set forth in any one of SEQ ID NOs: 82-86, resulting in a Sarbecovirus antibody of bivalent format wherein two of said ISVDs form a heavy chain only antibody-type molecule through disulfide bridges in the hinge region of the Fc part.
- an Fc domain such as an Fc domain comprising or consisting of the sequence set forth in any one of SEQ ID NOs: 82-86
- the binding agent in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in any one of SEQ ID NO:3-26 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO: 83, such as a binding agent, in particular a bivalent antibody, comprising or consisting of the sequence set forth in any one of SEQ ID NO:96-119.
- the binding agent in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in any one of SEQ ID NO:3-26 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in any one of SEQ ID NO: 82, 84, 85, or 86.
- the binding agent in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO: 82.
- the binding agent in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO:83.
- the binding agent in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO: 84.
- the binding agent in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO:85.
- the binding agent in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO: 86.
- one or more ISVDs as described herein are linked, fused or connected directly or via a linker to one or more ISVDs capable of binding to .
- Sa becovirus spike protein RBD as defined herein.
- Such multispecific binding agents may also be referred to herein as “head-to-tail fusions”.
- the C-terminus of a head-to-tail fusion as described herein may be fused, e.g. by a linker, to an Fc domain, which construct upon expression in a host forms a multivalent and/or multispecific antibody through disulfide bridges in the hinge region of the Fc part.
- one or more ISVDs as described herein are linked, fused or connected directly or via a linker to one or more ISVD capable of binding to a Sarbecovirus spike protein RBD to form a multispecific binding agent or construct and said multispecific binding agent or construct is fused to an Fc domain.
- the antibody comprises a trispecific binding agent or construct fused to an Fc domain, wherein said trispecific binding agent or construct comprises one ISVD as described herein (or a variant thereof as described herein), one ISVD capable of binding to or competing with the VHH3.117 epitope as described herein, and one ISVD capable of binding to or competing with the VHH72 epitope as described herein, wherein said ISVDs are linked, fused or connected directly or via a linker to each other, in any order.
- a binding agent in particular a trispecific binding agent, may comprise an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof, an ISVD capable of binding to or competing with the VHH3.
- VHH-VHH-VHH-Fc fusion Such multispecific antibody, in particular trispecific antibody, may also be referred to herein as “VHH-VHH-VHH-Fc fusion”.
- one or more ISVDs as described herein are fused to the N-terminus of an Fc domain, and one or more ISVDs capable of binding to a Sarbecovirus spike protein RBD are fused to the C-terminus of the Fc domain, or one or more ISVDs as described herein (or variants thereof as described herein) are fused to the C- terminus of an Fc domain, and one or more ISVDs capable of binding to a Sarbecovirus spike protein RBD are fused to the N-terminus of the Fc domain.
- the antibody comprises one ISVD as described herein (or a variant thereof as described herein) fused to the N- terminus of an Fc domain and one ISVD capable of binding to a Sarbecovirus spike protein RBD fused to the C-terminus of the Fc domain, or the one ISVD as described herein (or a variant thereof as described herein) is fused to the C-terminus of the Fc domain and the one ISVD capable of binding to a Sarbecovirus spike protein RBD is fused to the N-terminus of the Fc domain.
- Multivalent or multi-specific antibodies as described herein may have (or be engineered and/or selected for) increased avidity and/or improved selectivity for the desired Sarbecovirus interaction, and/or for any other desired property or combination of desired properties that may be obtained by the use of such multivalent and/or multi-specific antibodies.
- the binding agents in particular the multivalent antibodies described herein, more particularly the multivalent antibodies comprising an Fc domain described herein, even more particularly the multivalent antibodies comprising an Fc domain as defined by any one of SEQ ID NO: 83-86, bind FcRn with an affinity (KD) below 300 nM, preferably below 200 nM, more preferably below 100 nM such as from 30 nM to 100 nM or from 40 nM to 100 nM, even more preferably from 30 nM to 80 nM or from 40 nM to 80 nM, most preferably from 30 nM to 70 nM or from 40 nM to 70 nM or from 40 nM to 65 nM, when measured at pH 6.0 by surface plasmon resonance (SPR).
- KD affinity
- the pharmacokinetic properties of binding agents may be influenced by their ability to interact with FcRn.
- the FcRn is expressed on innate immune cells such as macrophages but also on endothelial cells and trophoblasts of the placenta.
- the FcRn binds to IgG Fc domain at the junction of the CH2 and CH3 domains in a pH dependent manner. IgG is endocytosed by cells where it can be shuttled to lysosomes or recycled back to the cell surface.
- IgG to FcRn Binding of IgG to FcRn at low pH (pH ⁇ 6.5) in the endosomes allows the antibody to be trafficked with the FcRn back to the cell surface.
- IgG At the physiologic pH of the extracellular environment IgG has weak affinity for FcRn which results in its release from the FcRn back into circulation.
- the high affinity of IgG to FcRn contributes to the long half-life of antibodies.
- Introduction of specific mutations in IgGl Fc such as LS, LA, DHS and YTE as described elsewhere herein results in a decrease in the dissociation rate and an improvement in binding affinity between Fc and human FcRn at pH6.
- the binding agents in particular the multivalent and/or multi-specific antibodies described herein, more particularly the multivalent and/or multi-specific antibodies comprising an Fc domain described herein, have antibody-dependent cell-mediated cytotoxicity (ADCC) activity. More particularly, the binding agents, in particular the multivalent and/or multispecific antibodies described herein, more particularly the multivalent and/or multi-specific antibodies comprising an Fc domain described herein, are capable of inducing ADCC on target cells expressing a Sarbecovirus spike protein.
- ADCC antibody-dependent cell-mediated cytotoxicity
- ADCC refers to a form of cytotoxicity in which an antibody binds to certain cytotoxic cells (such as NK cells, neutrophils, and macrophages).
- the secretion of Ig on the Fey receptor enables these cytotoxic effector cells to specifically bind to the target cell carrying the antigen, and then kill the target cell using, for example, a cytotoxin.
- an in vitro ADCC assay can be performed.
- nucleic acid sequences encode the polypeptide portion of a polypeptidic or polypeptide Sarbecovirus binding agent, antibody or antibody fragment, or ISVD with all the features and variations described herein, mutatis mutandis.
- the encoded binding agent, antibody or antibody fragment, or ISVD is in essence as described herein.
- Nucleic acid(s)” or “nucleic acid molecule(s)” as used herein refers to a polymeric form of nucleotides of any length, either ribonucleotides or deoxyribonucleotides; the sequential linear arrangement of the nucleotides together resulting in/forming the “nucleotide sequence”, “DNA sequence”, or “RNA sequence”. This term refers only to the primary structure of the molecule. Thus, this term includes double- and single-stranded DNA, and RNA. It also includes known types of modifications, for example, methylation, “caps”, and substitution of one or more of the naturally occurring nucleotides with an analog.
- Modifications to nucleic acids can be introduced at one or more levels: phosphate linkage modification (e.g. introduction of one or more of phosphodiester, phosphoramidate or phosphorothioate bonds), sugar modification (e.g. introduction of one or more of LNA (locked nucleic acids), 2'-O-methyl, 2'-O-methoxy-ethyl, 2’-fluoro, S-constrained ethyl or tricyclo-DNA) and/or non-ribose modifications (e.g. introduction of one or more of phosphorodiamidate morpholines or peptide nucleic acids).
- phosphate linkage modification e.g. introduction of one or more of phosphodiester, phosphoramidate or phosphorothioate bonds
- sugar modification e.g. introduction of one or more of LNA (locked nucleic acids), 2'-O-methyl, 2'-O-methoxy-ethyl, 2’-fluoro, S-constrained
- nucleic acid construct it is meant a nucleic acid molecule that has been constructed in order to comprise one or more functional units not found together in nature, thus having a nucleotide sequence not found in nature (non-native nucleotide sequence).
- examples include circular, linear, doublestranded, extrachromosomal DNA molecules (plasmids), cosmids (plasmids containing COS sequences from lambda phage), viral genomes comprising non-native nucleic acid sequences, RNA or mRNA constructs, and the like.
- a “coding sequence” is a nucleotide sequence that can be transcribed into mRNA and/or translated into a polypeptide when placed under the control of appropriate (gene) regulatory sequences. The boundaries of the coding sequence are determined by a translation start codon at the 5 '-terminus and a translation stop codon at the 3'-terminus.
- a coding sequence can include, but is not limited to mRNA, cDNA, recombinant nucleotide sequences or genomic DNA, while introns may be present as well under certain circumstances.
- RNA or mRNA construct a (poly)peptide coding sequence may be included such that it can be translated by the cellular protein translation machinery.
- a (poly)peptide coding sequence will be typically preceded by an in-frame translation initiation codon and followed by a translation termination codon, to facilitate proper translation.
- the coding sequence in aforementioned constructs is operably linked to a secretion signal sequence.
- an mRNA or mRNA construct encoding the polypeptide portion of a polypeptidic or polypeptide Sarbecovirus binding agent, in particular an antibody or antibody fragment, as identified herein, more particularly an ISVD (or a variant thereof) as described herein, or a functional part thereof.
- the mRNA or mRNA construct comprises (i) a 5'- untranslated region (5’ UTR); (ii) a coding sequence encoding a polypeptidic or polypeptide Sarbecovirus binding agent, an antibody or antibody fragment or an ISVD (or a variant thereof) described herein, or a functional part thereof; (iii) a stop codon; and (iv) a 3 '-untranslated region (3' UTR).
- a secretion signal sequence is operably linked to the coding sequence.
- the mRNA further comprises a 5' terminal cap.
- a 5' cap protects mRNA from RNase degradation.
- the translation system in the cytoplasm recognizes the cap structure through the cap-binding protein (CBP). This helps the small ribosomal subunit bind to mRNA and recognize the start codon to initiate translation.
- CBP cap-binding protein
- introducing a 5’ cap structure at the 5' end of a mRNA or mRNA construct described herein can enhance its translation, improve its stability, reduce its immunogenicity, and/or extend its half-life.
- the 5' caps can be natural or cap analogs. Cap analogs differ from natural (i.e., endogenous, wild-type or physiological) 5'-caps in their chemical structure, while retaining cap function.
- Non-limiting examples of cap structures that can be used herein include a CapO, Capl, Anti -Reverse Cap Analog (ARC A), inosine, N1 -methylguanosine, 2'-fluoro-guanosine, 7-deaza-guanosine, 8-oxo-guanosine, 2-amino-guanosine, LNA- guanosine, 2-azidoguanosine, Cap2, Cap4, 5' methylG cap, or an analog thereof as known to the skilled person.
- Caps may be chemically (i.e., non-enzymatically) or enzymatically synthesized and enzymatically linked to a nucleic acid molecule, in particular an mRNA or mRNA construct as described herein.
- the mRNA or mRNA construct further comprises a polyA tail.
- the addition of a polyA tail to the 3 ’ end of the mRNA or mRNA construct can advantageously increase its stability and/or reduce its immunogenicity.
- the length of the polyA tail can also regulate the halflife of the mRNA or mRNA construct in vivo.
- the polyA tail may be at least about 10, at least about 20, at least about 30, at least about 40, at least about 50, at least about 60, at least about 70, at least about 80, at least about 90 nucleotides in length, or at least about 100 nucleotides in length.
- the polyA tail may be about 10 to about 200, about 20 to about 180, about 50 to about 160, about 70 to about 140, or about 80 to about 120 nucleotides in length.
- a polyA tail may be added to the 3 ’ end of an mRNA or mRNA construct as described herein through polyadenylation directly at the end of the mRNA or mRNA construct by a polyA polymerase, or by RNA ligation, wherein a polyA tail is directly linked to the 3 ’ end of the mRNA or mRNA construct.
- the mRNA or mRNA construct comprises (i) a 5’ cap, (ii) a 5 '-untranslated region (5’ UTR); (iii) a coding sequence encoding a polypeptidic or polypeptide Sarbecovirus binding agent, an antibody or antibody fragment or an ISVD (or a variant thereof) described herein, or a functional part thereof; (iv) a stop codon; (v) a 3 '-untranslated region (3' UTR), and (vi) a polyA tail.
- the mRNA is modified mRNA.
- modified messenger RNA or “modified mRNA” refer to mRNA polynucleotides that include naturally occurring and/or non- naturally occurring modifications, for example, of a sugar, a nucleobase, and/or an intemucleoside linkage (e.g., to a linking phosphate, to a phosphodiester linkage, or to the phosphodiester backbone).
- intemucleoside linkage e.g., to a linking phosphate, to a phosphodiester linkage, or to the phosphodiester backbone.
- sugar modifications and phosphate linkage modifications have been described elsewhere herein.
- modified nucleobases include, but are not limited to, for example, pseudouracil (y), N1 -methylpseudouracil (ml ⁇
- the modification may be introduced with chemical synthesis or with a polymerase enzyme at the terminal of the mRNA or mRNA construct or anywhere else in the mRNA or mRNA construct.
- a “chimeric gene” or “chimeric construct” or “chimeric gene construct” is interchangeably meant a recombinant nucleic acid sequence in which a (gene) promoter or regulatory nucleic acid sequence is operably or operatively linked to, or associated with, a nucleic acid sequence of interest that codes for an RNA (e.g. a coding sequence, an shRNA, etc.), such that the regulatory nucleic acid sequence is able to regulate transcription or expression of the nucleic acid of interest.
- RNA e.g. a coding sequence, an shRNA, etc.
- An "expression cassette” comprises any nucleic acid construct capable of directing the expression of a gene/coding sequence of interest, which is operably linked to a (gene) promoter.
- Expression cassettes are generally DNA constructs preferably including (5’ to 3’ in the direction of transcription): a (gene) promoter region, a polynucleotide sequence of interest with a transcription initiation region, and a termination sequence including a stop signal for RNA polymerase and a polyadenylation signal; all these elements being operably or operatively linked meaning that all of these regions should be capable of operating (being expressed) in a cell, such as prokaryotic (e.g. bacterial) or eukaryotic (e.g.
- the promoter region comprising the transcription initiation region, which preferably includes the RNA polymerase binding site, and the polyadenylation signal may be native to the cell to be transformed, may be derived from an alternative source, or may be synthetic, as long as it is functional in the cell.
- Such expression cassettes can be constructed in e.g. a “vector” or “expression vector” (linear or circular nucleic acids, plasmids, cosmids, viral vectors, phagemids, etc.).
- the nucleic acid molecule may be an mRNA or mRNA construct.
- the present invention also provides a vector including the above-mentioned nucleic acid molecule inserted therein.
- vector refers to a nucleic acid molecule capable of carrying another nucleic acid molecule to which it has been linked.
- Said vectors may include a cloning or expression vector, as well as a delivery vehicle such as a viral, lentiviral or adenoviral vector.
- Expression vectors may comprise plasmids as well as viral vectors and generally contain a desired coding sequence and appropriate DNA sequences necessary for the expression of the operably linked coding sequence in a particular host organism (e.g., bacteria, yeast, plant, insect, or mammal) or in in vitro expression systems.
- an expression vector as described herein may comprise a nucleic acid molecule as described herein comprising a nucleic acid sequence encoding an antibody or an antigen-binding fragment as described herein operably linked to at least one regulatory sequence.
- the vector includes a promoter for driving expression of the nucleic acid of interest, optionally a nucleic acid sequence encoding a signal peptide that secretes the antibody or antigen-binding fragment, and optionally a nucleic acid sequence encoding a terminator.
- the expression vector When the expression vector is manipulated in a production strain or cell line, the vector may or may not be integrated into the genome of the host cell when introduced into the host cell.
- Cloning vectors are generally used to engineer and amplify a certain desired DNA fragment.
- a cloning vectors may contain origin of replication that matches the cell type specified by the cloning vector, and may lack functional sequences needed for expression of the desired DNA fragments.
- the vector contains one or more selection markers. The choice of the selection markers may depend on the host cells of choice, although this is not critical to the present invention as is well known to persons skilled in the art.
- the construction of expression vectors for use in transfecting cells is also well known in the art, and thus can be accomplished via standard techniques (see, for example, Sambrook, Fritsch, and Maniatis, in: Molecular Cloning, A Laboratory Manual, Cold Spring Harbor Laboratory Press, 1989; Gene Transfer and Expression Protocols, pp. 109-128, ed. E. J. Murray, The Humana Press Inc., Clifton, N.J.), and the Ambion 1998 Catalog (Ambion, Austin, Tex.).
- said vector may include any vector known to the skilled person, including any suitable type, but not limited to, for instance, plasmid vectors, cosmid vectors, phage vectors, such as lambda phage, viral vectors, even more particular a lentiviral, adenoviral, AAV or baculoviral vectors, or artificial chromosome vectors such as bacterial artificial chromosomes (BAC), yeast artificial chromosomes (YAC), or Pl artificial chromosomes (PAC).
- BAC bacterial artificial chromosomes
- YAC yeast artificial chromosomes
- PAC Pl artificial chromosomes
- the choice of the vector may bee dependent amongst others on the nature of the host cell of choice.
- One further aspect of the invention provides for a host cell comprising an antibody or antigen-binding fragment thereof, such as an ISVD (or variant thereof) of an antibody or antigen-binding fragment, or part thereof, as described herein.
- the host cell may therefore comprise the nucleic acid molecule encoding said antibody or antigen-binding fragment.
- Host cells can be either prokaryotic or eukaryotic.
- the host cell may also be a recombinant host cell, which involves a cell which has been genetically modified to contain an isolated nucleic acid molecule encoding the antibody or antigenbinding fragment of the invention.
- Representative host cells that may be used to produce said antibodies or antigen-binding fragments such as ISVDs include, but are not limited to, bacterial cells, yeast cells, plant cells and animal cells.
- Bacterial host cells suitable for production of the antibodies or antigen-binding fragment of the invention include Escherichia spp. cells, Bacillus spp. cells, Streptomyces spp. cells, Erwinia spp. cells, Klebsiella spp. cells, Serratia spp. cells, Pseudomonas spp. cells, and Salmonella spp. cells.
- Yeast host cells suitable for use with the invention include species within Saccharomyces, Schizosaccharomyces, Kluyveromyces, Pichia (e.g. Pichia pastoris), Hansenula (e.g. Hansenula polymorpha), Yarowia, Schwaniomyces, Schizosaccharomyces, Zygosaccharomyces and the like. Saccharomyces cerevisiae, S. carlsbergensis and K. lactis are the most commonly used yeast hosts, and are convenient fungal hosts.
- Animal host cells suitable for use with the invention include insect cells and mammalian cells (e.g. derived from Chinese hamster (e.g. CHO), and human cell lines, such as HeLa).
- Exemplary insect cell lines include, but are not limited to, Sf9 cells, baculovirus-insect cell systems (e.g. review Jarvis, Virology Volume 310, Issue 1, 25 May 2003, Pages 1-7).
- the host cells may also be transgenic animals or plants.
- Introduction of a vector in a host cell can be effected by, e.g., calcium phosphate transfection, virus infection, DEAE-dextran-mediated transfection, lipofectamin transfection or electroporation, and any person skilled in the art can select and use an introduction method suitable for the expression vector and host cell used.
- a further aspect of the invention relates to a composition
- a composition comprising a binding agent, such as an antibody or antigen-binding fragment thereof, comprising one or more ISVDs (or variants thereof) as described herein.
- composition refers to a combination of one or more molecules, present in a formulation that retains the binding agents activity, specifically the HR2 (or S2) binding and Sarbecovirus neutralization activity in this case, thus a functional composition.
- the composition thus comprises one or more molecules which constitute one or more binding agents as described herein which specifically bind the Sarbecovirus Spike protein via interaction with its HR2 domain (S2 targeting binding agents or molecules).
- the composition may comprise a bivalent antibody comprising an ISVD (or variant thereof) as described herein fused to an Fc domain.
- Said composition may be a soluble or solid composition.
- composition may further comprise, for instance but not limited to, buffer components, adjuvants, or additional molecules, which may be functional molecules.
- the composition may further comprise one or more binding agents capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD) as described elsewhere herein.
- the composition may further comprise one or more binding agents, such as an antibody or antigen-binding fragment thereof, comprising one or more (such as two, three, four, or more) ISVDs (or variants or humanized forms thereof) capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD) as described elsewhere herein.
- Said composition may thus contain at least two binding agents, characterized in that one binding agent specifically binds the HR2 domain, and the second binding agent specially binds the RBD region, resulting in a composition with at least two binding agents binding in a non-competing manner to the spike protein, possibly simultaneously.
- the binding agent capable of binding to a Sarbecovirus spike protein RBD is capable of binding two non-competing binding sites of the RBD, preferably via two different ISVDs present in said binding agent, wherein said binding agent may be a bispecific binding agent, or multispecific binding agent. More specifically, the binding agent may comprise one or more ISVDs capable of binding to or competing for the VHH72 epitope as defined herein, and one or more ISVDs capable of binding to or competing for the VHH3. 117 epitope as defined herein. Non-limiting examples of binding agents comprising one or more ISVDs capable of binding to or competing for the VHH72 epitope, and one or more ISVDs capable of binding to or competing for the VHH3.
- 117 epitope are described in WO 2022/238550 Al. Also disclosed herein are particular examples of binding agents comprising one or more ISVDs capable of binding to or competing for the VHH72 epitope, and one or more ISVDs capable of binding to or competing for the VHH3.
- 117 epitope in particular antibodies or antigen-binding fragments thereof, comprising i) one or more ISVDs comprising the CDRs present in SEQ ID NO:91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g.
- an ISVD comprising the CDRs present in SEQ ID NO: 92 such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3.117), or a variant or a humanized form thereof, wherein the CDRs are annotated according to Kabat, Martin, MacCallum, IMGT, AbM, or Chothia. More particularly bispecific antibodies comprising i) an ISVD comprising the CDRs present in SEQ ID NO:92, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3.
- the ISVD fused to the C-terminus of the Fc domain may be fused to the C-terminus of the Fc domain, and ii) the ISVD comprising the CDRs present in SEQ ID NO:91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g. VHH83), or a variant or a humanized form thereof, may be fused to the N-terminus of the Fc domain.
- the ISVD fused to the C-terminus of the Fc domain may be modified to reduce reactivity of the ISVD with pre-existing anti -drug antibodies (ADAs).
- said ISVD may comprise a C-terminal modification to reduce reactivity of the ISVD with pre-existing ADAs, such as the C-terminal TVSS sequence (SEQ ID NO: 139) of the ISVD may be replaced by any one of the sequences set forth in SEQ ID NO: 140-150 or the amino acid sequence TVS.
- the same modifications may be provided in any binding agent described herein, in particular a binding agent comprising an ISVD which is not fused at its C-terminus to another moiety such as an Fc moiety, such as more particularly a binding agent comprising an ISVD comprising a C-terminal sequence as set forth in SEQ ID NO: 139.
- the bispecific antibody comprises or consists of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014), or any functional variant thereof, or a variant with at least 90% identity thereof, or a humanized variant thereof, such as a bispecific antibody comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 127-138.
- the composition may comprise (i) a binding agent, in particular an antibody or an antigen-binding fragment thereof, comprising one or more ISVDs comprising the CDRs present in any one of SEQ ID NO: 2 to 26, such as one or more ISVDs comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 2 to 26, or a variant or humanized form thereof; and (ii) a binding agent, in particular an antibody or an antigen-binding fragment thereof, comprising one or more ISVDs comprising the CDRs present in SEQ ID NO:91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g.
- VHH83 or a variant or a humanized form thereof, and an ISVD comprising the CDRs present in SEQ ID NO:92, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3. 117), or a variant or a humanized form thereof, wherein the CDRs are annotated according to Kabat, Martin, MacCallum, IMGT, AbM, or Chothia.
- composition may comprise:
- a binding agent in particular an (optionally bivalent) antibody, comprising an ISVD comprising the CDRs present in any one of SEQ ID NO: 2 to 26, such as an ISVD comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 2 to 26 or a variant or humanized form thereof, fused to an Fc domain, such as an antibody comprising the amino acid sequence set forth in any one of SEQ ID NO: 96 to 119; and
- a binding agent in particular a bispecific antibody, comprising an ISVD comprising the CDRs present in SEQ ID NO:92, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3. 117), or a variant or a humanized form thereof, fused to the N- terminus of an Fc domain as defined herein, and an ISVD comprising the CDRs present in SEQ ID NO:91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g. VHH83), or a variant or a humanized form thereof, fused to the C-terminus of the Fc domain, wherein the CDRs are annotated according to Kabat, Martin, MacCallum, IMGT, AbM, or Chothia.
- the bispecific antibody (ii-a) comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92, fused to the N-terminus of an Fc domain as defined herein, and an ISVD comprising or consisting of the sequence set forth in SEQ ID NO: 91 fused to the C-terminus of the Fc domain.
- the Fc domain comprises or consists of the sequence set forth in any one of SEQ ID NO: 82 to 86.
- the bispecific antibody (ii-a) comprises an ISVD consisting of the sequence set forth in SEQ ID NO:92, fused to the N-terminus of an Fc domain as defined herein, and an ISVD consisting of the sequence set forth in SEQ ID NO: 91 fused to the C-terminus of the Fc domain.
- the Fc domain comprises or consists of the sequence set forth in any one of SEQ ID NO: 82 to 86.
- composition may comprise:
- a binding agent in particular a (optionally bivalent) antibody, comprising an ISVD comprising the CDRs present in any one of SEQ ID NO: 2 to 26, such as an ISVD comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 2 to 26 or a variant or humanized form thereof, fused to an Fc domain, such as an antibody comprising the amino acid sequence set forth in any one of SEQ ID NO: 96 to 119; and
- a binding agent in particular a bispecific antibody, comprising an ISVD comprising the CDRs present in SEQ ID NO:92, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3. 117), or a variant or a humanized form thereof, fused to the C- terminus of an Fc domain as defined herein, and an ISVD comprising the CDRs present in SEQ ID NO: 91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g. VHH83), or a variant or a humanized form thereof, fused to the N-terminus of the Fc domain, wherein the CDRs are annotated according to Kabat, Martin, MacCallum, IMGT, AbM, or Chothia.
- the bispecific antibody (ii-b) comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92, fused to the C-terminus of an Fc domain as defined herein, and an ISVD comprising or consisting of the sequence set forth in SEQ ID NO: 91 fused to the N-terminus of the Fc domain.
- the Fc domain comprises or consists of the sequence set forth in any one of SEQ ID NO: 82 to 86.
- the bispecific antibody (ii-b) comprises an ISVD consisting of the sequence set forth in SEQ ID NO:92, fused to the C-terminus of an Fc domain as defined herein, and an ISVD consisting of the sequence set forth in SEQ ID NO: 91 fused to the N-terminus of the Fc domain.
- the Fc domain comprises or consists of the sequence set forth in any one of SEQ ID NO: 82 to 86.
- composition may comprise:
- a binding agent in particular an (optionally bivalent) antibody, comprising an ISVD comprising the CDRs present in any one of SEQ ID NO: 2 to 26, such as an ISVD comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 2 to 26 or a variant or humanized form thereof, fused to an Fc domain, such as an antibody comprising the amino acid sequence set forth in any one of SEQ ID NO: 96 to 119; and
- a bispecific antibody comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126, or any functional variant thereof, or a variant with at least 90% identity thereof, or a humanized variant thereof.
- composition may comprise:
- a binding agent in particular a (optionally bivalent) antibody, comprising an ISVD comprising the CDRs present in any one of SEQ ID NO: 2 to 26, such as an ISVD comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 2 to 26 or a variant or humanized form thereof, fused to an Fc domain, such as an antibody comprising the amino acid sequence set forth in any one of SEQ ID NO: 96 to 119; and
- a bispecific antibody comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 127-138, or any functional variant thereof, or a variant with at least 90% identity thereof, or a humanized variant thereof.
- the binding agent (i) comprised by any one of the above compositions may be a binding agent, in particular an (optionally bivalent) antibody, comprising an ISVD comprising the CDRs present in SEQ ID NO: 25, such as an ISVD comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 25 (e.g. VHH CBV3) or a variant or humanized form thereof, fused to an Fc domain as defined herein, such as an antibody comprising the amino acid sequence set forth in SEQ ID NO: 118, or a variant or humanized form thereof.
- an antibody comprising the amino acid sequence set forth in SEQ ID NO: 118, or a variant or humanized form thereof.
- the composition may comprise (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof; and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126, or any functional variant thereof, or a variant with at least 90% identity thereof, or a humanized variant thereof, such as a bispecific antibody comprising or consisting of the sequence set forth in any one of SEQ ID NO: 127-138.
- a binding agent in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof
- a binding agent in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126, or any functional variant thereof, or a variant with at least 90% identity thereof, or
- the composition may comprise (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof; and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014).
- a binding agent in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof
- a binding agent in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014).
- composition may comprise (i) a binding agent, in particular a (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof; and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 127.
- a binding agent in particular a (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof
- a binding agent in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 127.
- composition may comprise (i) a binding agent, in particular a (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof; and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 128.
- a binding agent in particular a (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof
- a binding agent in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 128.
- the molecular ratio of the (S2 targeting) binding agent such as an antibody or antigen-binding fragment thereof, comprising one or more ISVDs (or variants thereof), or part thereof, as described herein and the (S 1 targeting) binding agent such as an antibody or antigenbinding fragment thereof, comprising one or more ISVDs (or variants thereof) capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD) in the composition
- RBD Sarbecovirus spike protein receptor binding domain
- the molecular ratio is about 1: 1.
- the mass ratio of the (S2 targeting) binding agent, such as an antibody or antigenbinding fragment thereof, comprising one or more ISVDs (or variants thereof), or part thereof, as described herein and the (SI targeting) binding agent such as an antibody or antigen-binding fragment thereof, comprising one or more ISVDs (or variants thereof) capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD) in the composition may range from 20: 1 to 1:20 such as from 10: 1 to 1: 10, from 9: 1 to 1:9, from 8: 1 to 1:8, from 7: 1 to 1:7, from 6: 1 to 1:6, from 5: 1 to 1:5, from 4: 1 to 1:4 or from 3: 1 to 1:3, preferably from 2: 1 to 1:2, more preferably the mass ratio is about 1: 1.
- An exemplary composition comprises (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof (e.g., XVR013m); and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g.
- An exemplary composition comprises (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof (e.g., XVR013m); and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014) or a variant thereof such as such as a bispecific antibody comprising or consisting of the sequence set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of the binding agent (i) to the binding agent (ii) in the composition is from 10: 1 to 1: 10.
- Another exemplary composition comprises (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof (e.g., XVR013m); and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014) or a variant thereof such as such as a bispecific antibody comprising or consisting of the sequence set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of the binding agent (i) to the binding agent (ii) in the composition is about 1: 10.
- Another exemplary composition comprises (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof (e.g., XVR013m); and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014) or a variant thereof such as such as a bispecific antibody comprising or consisting of the sequence set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of the binding agent (i) to the binding agent (ii) in the composition is about 1: 1.
- a particularly preferred composition comprises XVR013m and XVR014, wherein the mass ratio of XVR013m to XVR014 in the composition is from 20: 1 to 1:20, preferably from 10: 1 to 1: 10 such as from 9: 1 to 1:9, from 8: 1 to 1:8, from 7: 1 to 1:7, from 6: 1 to 1:6, from 5: 1 to 1:5, from 4: 1 to 1:4, from 3: 1 to 1:3, from 2: 1 to 1:2, or about 1: 1.
- the composition comprises XVR013m and XVR014, wherein the mass ratio of XVR013m to XVR014 is about 1: 10.
- the composition comprises XVR013m and XVR014, wherein the mass ratio of XVRO 13m to XVRO 14 is about 1: 1.
- compositions comprise XVRO 13m and a bispecific antibody as set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of XVR013m to the bispecific antibody is from 20: 1 to 1:20, preferably from 10: 1 to 1: 10 such as from 9: 1 to 1:9, from 8: 1 to 1:8, from 7: 1 to 1:7, from 6: 1 to 1:6, from 5: 1 to 1:5, from 4: 1 to 1:4, from 3: 1 to 1:3, from 2: 1 to 1:2, or about 1: 1.
- the composition comprises XVRO 13m and a bispecific antibody as set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio ofXVR013m to the bispecific antibody is about 1: 10. In certain embodiments, the composition comprises XVRO 13m and a bispecific antibody as set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of XVRO 13m to the bispecific antibody is about 1: 1.
- composition may still contain additional binding agent(s) or molecules, which optionally bind further binding regions on the same or different epitopes of the spike protein, or other viral proteins, or may even target totally unrelated target proteins.
- a further aspect of the invention relates to medicaments or pharmaceutical compositions comprising a binding agent, in particular an antibody or antigen-binding fragment, a nucleic acid encoding it, and/or a (recombinant) vector comprising the nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein.
- a pharmaceutical composition is a pharmaceutically acceptable composition; such compositions are preferably further comprising a (pharmaceutically) suitable or acceptable carrier, diluent, adjuvant, excipient, stabilizer, etc.
- pharmaceutically acceptable is meant a material that is not biologically or otherwise undesirable, i.e., the material may be administered to an individual along with the compound, in particular the Sarbecovirus binding agent, more particularly the Sarbecovirus antibody or antigenbinding fragment, without causing any undesirable biological effects or interacting in a deleterious manner with any of the other components of the pharmaceutical composition in which it is contained.
- a pharmaceutically acceptable carrier is preferably a carrier that is relatively non-toxic and innocuous to a patient at concentrations consistent with effective activity of the active ingredient so that any side effects ascribable to the carrier do not vitiate the beneficial effects of the active ingredient.
- Suitable carriers or adj uvantia typically comprise one or more of the compounds included in the following non- exhaustive list: large slowly metabolized macromolecules such as proteins, polysaccharides, polylactic acids, polyglycolic acids, polymeric amino acids, amino acid copolymers and inactive virus particles.
- excipient is intended to include all substances which may be present in a pharmaceutical composition and which are not active ingredients but may contribute to e.g. long-term stability, or therapeutic enhancement on the active ingredient (such as by facilitating drug absorption, reducing viscosity, or enhancing solubility).
- Excipients may include, for example, salts, binders (e.g., lactose, dextrose, sucrose, trehalose, sorbitol, mannitol), lubricants, thickeners, surface active agents, preservatives, emulsifiers, buffer substances, stabilizing agents, flavouring agents or colorants.
- Auxiliary substances such as wetting or emulsifying agents, pH buffering substances, preservatives may be included in such vehicles.
- a pharmaceutically effective amount of binding agents, in particular antibodies or antigen-binding fragments, of the invention is preferably that amount which produces a result or exerts an influence on the particular condition being treated.
- the pharmaceutical composition of this invention can be lyophilized for storage and reconstituted in a suitable carrier prior to use.
- physiologically acceptable carrier, excipient, stabilizer need to be added into the pharmaceutical composition of the invention (Remington's Pharmaceutical Sciences 22nd edition, Ed. Allen, Loyd V, Jr. (2012).
- the preparation containing pharmaceutical composition of this invention should be sterilized before injection. This procedure can be done using sterile filtration membranes before or after lyophilization and reconstitution.
- the pharmaceutical composition can be packaged in a container or vial with sterile access port, such as an i.v. solution bottle with a rubber stopper - the pharmaceutical composition can be present as liquid, or the container or vial is filled with a liquid pharmaceutical composition that is subsequently lyophilized or dried; or can be packaged in a pre-filled syringe.
- compositions of nucleic acid molecules encoding a binding agent as described herein such as an mRNA or an mRNA construct as described herein encoding the polypeptide portion of a polypeptidic or polypeptide Sarbecovirus binding agent, in particular an antibody or antibody fragment, as identified herein, more particularly an ISVD (or a variant thereof) as described herein, or a functional part thereof, may comprise the nucleic acid molecule, mRNA or mRNA construct encapsulated in lipid nanoparticles that protect the nucleic acid molecules, mRNA or mRNA construct and help their absorption into cells.
- lipid nanoparticle or “LNP” refers to a nanosized particle composed of different lipids.
- lipids may be included in such LNP, including, without limitation ionizable lipids, phospholipids (e.g. compounds having a phosphatidylcholine group), sterols (e.g. cholesterol), polyethylene glycol (PEG)-modified lipids.
- phospholipids e.g. compounds having a phosphatidylcholine group
- sterols e.g. cholesterol
- PEG polyethylene glycol
- the lipid nanoparticles contain a cationic lipid, or a lipid species with a net positive charge at a selected pH (e.g., physiological pH), to encapsulate and/or enhance the delivery of mRNA into target cells.
- a selected pH e.g., physiological pH
- the terms “ionizable lipid” or “cationic ionizable lipid” or “cationic lipids” refer to lipids having at least one protonatable or deprotonatable group, such that the lipid is positively charged at a pH at or below physiological pH (e.g., pH 7.4), and neutral at a second pH, preferably at or above physiological pH. It will be understood by one of ordinary skill in the art that the addition or removal of protons as a function of pH is an equilibrium process, and that the reference to a charged or a neutral lipid refers to the nature of the predominant species and does not require that all of the lipid be present in the charged or neutral form.
- the positively charged lipids are able to associate with negatively charged compounds.
- the lipid nanoparticles may further comprise a structural helper lipid.
- Suitable helper lipids are generally known in the art.
- a preferred helper lipid is a steroid or a sterol, more preferably cholesterol.
- Incorporation of a steroid or a sterol (e.g. cholesterol and analogues thereof, ergosterol, phytosterol) in the lipid nanoparticle may help aggregation of other lipids in the lipid nanoparticle.
- Other possible helper lipids are phospholipids including, without limitation, dioleoylphosphatidylethanolamine (DOPE) and l,2-distearoyl-sn-glycero-3 -phosphocholine (DSPC).
- DOPE dioleoylphosphatidylethanolamine
- DSPC l,2-distearoyl-sn-glycero-3 -phosphocholine
- a further aspect of the invention relates to a binding agent, in particular an antibody or antigenbinding fragment, a nucleic acid encoding it as described herein, a vector comprising such nucleic acid, a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein or a pharmaceutical composition comprising a binding agent, in particular an antibody or antigen-binding fragment, nucleic acid encoding it, a (recombinant) vector comprising such nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, for use as a medicine or medicament.
- a binding agent in particular an antibody or antigen-binding fragment, nucleic acid encoding it, a vector comprising such nucleic acid as described herein, or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, or use of a pharmaceutical composition comprising a binding agent, in particular an antibody or antigen-binding fragment, nucleic acid encoding it, a vector comprising such nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, in the manufacture of a medicine or medicament is envisaged.
- the binding agent in particular the antibody or antigen-binding fragment, the nucleic acid encoding it, the vector comprising such nucleic acid or the composition comprising the binding agent, in particular the antibody or antigen-binding fragment, as described herein, or the medicament or pharmaceutical composition comprising a binding agent, in particular an antibody or antigenbinding fragment, a nucleic acid encoding it, a (recombinant) vector comprising such nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, is for use in passive immunisation, for use in treating a subject with a Sarbecovirus infection, for use in preventing infection of a subject with a Sarbecovirus. or for use in protecting a subject from infection with a Sarbecovirus .
- the subject When for use in passive immunisation, the subject may have an infection with a Sarbecovirus (therapeutic passive immunisation) or may not have an infection with a Sarbecovirus (prophylactic passive immunisation).
- a related aspect relates to methods for treating a subject suffering from/having/that has contracted an infection with a Sarbecovirus, the methods comprising administering a binding agent, in particular an antibody or antigen-binding fragment, a nucleic acid encoding it a (recombinant) vector comprising such nucleic acid, or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein to the subject, or comprising administering a medicament or pharmaceutical composition comprising a binding agent , in particular an antibody or antigen-binding fragment, a nucleic acid encoding it, a (recombinant) vector comprising such nucleic acid and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein to the subject.
- a further aspect of the invention relates to methods for protecting a subject from infection with a Sarbecovirus or for preventing infection of a subject with a Sarbecovirus, the methods comprising administering a binding agent, in particular an antibody or antigen-binding fragment, a nucleic acid encoding it, a (recombinant) vector comprising such nucleic acid, or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein to the subject prior to infection, or comprising administering a medicament or pharmaceutical composition as described herein comprising a binding agent, in particular an antibody or antigen-binding fragment, a nucleic acid encoding it, a (recombinant) vector comprising such nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, to the subject prior to infection.
- binding agent in particular an antibody or antigenbinding fragment, nucleic acid encoding it, a vector comprising such nucleic acid as described herein, or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, or use of a pharmaceutical composition comprising a binding agent, in particular an antibody or antigen-binding fragment, nucleic acid encoding it, a vector comprising such nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, in the manufacture of a medicine or medicament for treating a subject with a Sarbecovirus infection, for preventing infection of a subject with a Sarbecovirus. or for protecting a subject from infection with a Sarbecovirus .
- a nucleic acid encoding a binding agent, in particular an antibody or antigen-binding fragment or a (recombinant) vector comprising such nucleic acid as described herein can be used in e.g. gene therapy setting.
- Gene therapy refers to therapy performed by the administration to a subject of an expressed or expressible nucleic acid.
- the nucleic acid molecule or vector as described herein allow for production of the binding agent, antibody or antibody fragment within a cell.
- a large set of methods for gene therapy are available in the art and include, for instance (adeno-associated) virus-mediated gene silencing, or virus-mediated gene therapy (e.g.
- a “therapeutically active agent” generally means any molecule that has or may have a therapeutic effect (i.e. curative or prophylactic effect) in the context of treatment of a disease.
- a therapeutically active agent is a disease-modifying agent, which can be a cytotoxic agent, such as a toxin, or a cytotoxic drug, or an enzyme capable of converting a prodrug into a cytotoxic drug, or a radionuclide, or a cytotoxic cell, or which can be a non-cytotoxic agent.
- a therapeutically active agent has a curative effect on the disease.
- the binding agent in particular the antibody or antibody fragment, or pharmaceutical composition of the invention may act as a therapeutically active agent, when beneficial in treating patients infected with a Sarbecovirus, such SARS-CoV-2 or SARS-CoV-1 or patients suffering from COVID-19.
- the binding agent in particular the antibody or antibody fragment, may comprise a Sarbecovirus-binding ISVD or a variant thereof as described herein, and may contain or be coupled to additional functional groups, advantageous when administrated to a subject.
- Such functional groups can generally comprise all functional groups and techniques mentioned in the art as well as the functional groups and techniques known per se for the modification of pharmaceutical proteins, and in particular for the modification of antibodies or antibody fragments, for which reference is for example made to Remington's Pharmaceutical Sciences, 16th ed., Mack Publishing Co., Easton, PA (1980).
- Such functional groups may for example be linked directly (for example covalently) to the ISVD or active antibody fragment, or optionally via a suitable linker or spacer, as will again be clear to the skilled person.
- One of the most widely used techniques for increasing the half-life and/or reducing immunogenicity of pharmaceutical proteins comprises attachment of a suitable pharmacologically acceptable polymer, such as poly(ethyleneglycol) (PEG) or derivatives thereof (such as methoxypoly(ethyleneglycol) or mPEG).
- a suitable pharmacologically acceptable polymer such as poly(ethyleneglycol) (PEG) or derivatives thereof (such as methoxypoly(ethyleneglycol) or mPEG).
- PEG may be attached to a cysteine residue that naturally occurs in an immunoglobulin single variable domain described herein (or a variant thereof as described herein), an immunoglobulin single variable domain as described herein (or a variant thereof as described herein) may be modified so as to suitably introduce one or more cysteine residues for attachment of PEG, or an amino acid sequence comprising one or more cysteine residues for attachment of PEG may be fused to the N- and/or C-terminus of an ISVD or active antibody fragment as described herein (or a variant thereof as described herein), all using techniques of protein engineering known per se to the skilled person.
- Another, usually less preferred modification comprises N-linked or O-linked glycosylation, usually as part of co-translational and/or post- translational modification, depending on the host cell used for expressing the antibody or active antibody fragment.
- Another technique for increasing the half-life of a binding domain, in particular an antibody or antibody fragment may comprise the engineering into bifunctional or bispecific domains (for example, one ISVD or active antibody fragment against the target Sarbecovirus HR2 domain and one against a serum protein such as albumin or Staphylococcal protein A (SpA) -which is a surface protein abundantly present in the lungs aiding in prolonging half-life)) or into fusions of antibody fragments, in particular immunoglobulin single variable domains, with peptides (for example, a peptide against a serum protein such as albumin).
- an ISVD as described herein can be fused to an immunoglobulin Fc domain as described elsewhere herein. Examples are further shown in the experimental section and are also depicted in the sequence listing.
- the Sarbecovirus is SARS-CoV-2 such as a SARS-CoV-2 variant, or SARS-CoV-1.
- the SARS-CoV-2 variant may be a variant at position N439, K417, S477, L452, T478, E484, P384, N501 and/or D614 (relative to the SARS-CoV-2 spike amino acid sequence as defined in SEQ ID NO:93), more particularly a variant at position N501 such as a N501Y variant (e.g. SARS-CoV-2 Alpha variant), a variant at position N501 and E484 such as a N501Y and E484K variant (e.g. SARS-CoV-2 Alpha + E484K variant), a variant at position K417, E484 and N501 such as a K417N, E484K and N501Y variant (e.g.
- SARS-CoV-2 beta variant a variant at position P384, K417, E484 and N501 such as a P384L, K417N, E484K and N501Y variant (e.g. SARS-CoV-2 beta + P384L variant), a variant at position L452 and E484 such as a L452R and E484Q variant (e.g. SARS-CoV-2 kappa variant), a variant at position L452 and T478 such as a L452R and T478K variant (e.g. SARS-CoV-2 delta variant), a variant at position L452 such as a L452R variant (e.g.
- SARS-CoV-2 epsilon variant a variant at position K417 such as a K417T variant (e.g. SARS-CoV-2 gamma variant), a variant at position D614 such as a D614G variant (e.g. SARS-CoV-2 Omicron variant or SARS-CoV-2 BA. l variant); or a SARS-CoV-2 Omicron EG.5.1 variant, a SARS-CoV-2 Omicron BA.2.86.1 variant, a SARS-CoV-2 Omicron HV.
- K417T variant e.g. SARS-CoV-2 gamma variant
- D614 such as a D614G variant
- SARS-CoV-2 Omicron EG.5.1 variant a SARS-CoV-2 Omicron BA.2.86.1 variant, a SARS-CoV-2 Omicron HV.
- SARS-CoV-2 Omicron XBB.1.5 variant a SARS-CoV-2 Omicron XBB.2.3 variant
- SARS-CoV-2 Omicron XBB.1.16 variant a SARS-CoV-2 Omicron HK.3 variant
- SARS-CoV-2 Omicron BA.2.74 variant a SARS-CoV-2 Omicron FL. 1.5.1 variant, a SARS-CoV-2 Gamma variant, a SARS-CoV-2 Eta variant, a SARS-CoV-2 Iota variant, a SARS- CoV-2 Epsilon B. 1.427 variant, a SARS-CoV-2 Epsilon B.
- a SARS-CoV-2 Mu variant a SARS-CoV-2 Zeta variant, a SARS-CoV-2 B. 1.617.3 variant, a SARS-CoV-2 Omicron CH. 1.1 variant, a SARS-CoV-2 Omicron JN. l variant, a SARS-CoV-2 Omicron KP.2.3 variant, a SARS- CoV-2 Omicron KP.3 variant, a SARS-CoV-2 Omicron KP.3.1.1 variant, a SARS-CoV-2 Omicron KP.1.1 variant, a SARS-CoV-2 Omicron LB. l variant or a SARS-CoV-2 Omicron XEC variant.
- the Sarbecovirus is any one or both of SARS-CoV-2 and SARS-CoV-1.
- SARS-CoV-2 is SARS-CoV-2 Wuhan strain or a SARS-CoV-2 variant, in particular a SARS-CoV-2 variant selected from the group consisting of SARS-CoV-2 Alpha variant, SARS-CoV-2 Omicron BA. l variant and SARS-CoV-2 Omicron BA.2 variant or selected from the group consisting of: SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta variant, SARS-CoV- 2 Epsilon B.
- SARS-CoV-2 Epsilon B. 1.429 variant SARS-CoV-2 Eta variant, SARS- CoV-2 Iota variant, SARS-CoV-2 Kappa variant, SARS-CoV-2 Mu variant, SARS-CoV-2 Zeta variant, SARS-CoV-2 B. 1.617.3 variant, SARS-CoV-2 Omicron BA.5 variant, SARS-CoV-2 Omicron XBB.1.5 (-G252V) variant, SARS-CoV-2 Omicron XBB.1.5 variant, SARS-CoV-2 Omicron XBB.2.3 variant, SARS-CoV-2 Omicron XBB.1.16 variant, SARS-CoV-2 Omicron CH. 1.
- SARS-CoV-2 Omicron EG.5. 1 variant, SARS-CoV-2 Omicron HK.3 variant, SARS- CoV-2 Omicron BA.2.74 variant, SARS-CoV-2 Omicron BA.2.86.
- SARS-CoV-2 Omicron HV. l variant SARS-CoV-2 Omicron FL. 1.5.1 variant, SARS-CoV-2 Omicron JN.l variant, SARS- CoV-2 Omicron KP.2.3 variant, SARS-CoV-2 Omicron KP.3 variant, SARS-CoV-2 Omicron KP.3.1.1 variant, a SARS-CoV-2 Omicron BA.
- SARS-CoV-2 Omicron BA.2 variant a SARS-CoV-2 Omicron KP.1.1 variant
- SARS-CoV-2 Omicron LB. l variant a SARS-CoV-2 Omicron XEC variant.
- the terms “therapy” or “treatment” refer to the alleviation or measurable lessening of one or more symptoms or measurable markers of a pathological condition such as a disease or disorder, in particular a Sarbecovirus infection. Measurable lessening includes any statistically significant decline in a measurable symptom or marker.
- the terms encompass both curative treatments and treatments directed to reduce symptoms and/or slow progression of the disease.
- the terms encompass both the therapeutic treatment of an already developed pathological condition, in particular a Sarbecovirus infection, as well as prophylactic or preventative measures, wherein the aim is to prevent or lessen the chances of incidence of a pathological condition, in particular a Sarbecovirus infection.
- Beneficial or desired clinical results include, but are not limited to, prevention of a disease, reduction of the incidence of a disease, alleviation of symptoms associated with a disease, diminishment of extent of a disease, stabilisation of the disease, delay or slowing of the progression of a disease, amelioration or palliation of a disease, or combinations thereof.
- the terms may relate to therapeutic treatments. In certain other embodiments, the terms may relate to preventative treatments.
- treatment may refer to passive immunisation of a subject having contracted a Sarbecovirus infection (therapeutic treatment).
- Prevention of infection with a Sarbecovirus may be useful in case of e.g. epidemic or pandemic conditions during which subjects known to be most vulnerable to develop severe disease symptoms can be prophylactically treated (preventive or prophylactic immunisation) with a binding agent, in particular an antibody or antigen-binding fragment or a nucleic acid encoding it or a vector comprising such nucleic acid or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment as described herein such as to prevent infection overall, or such as to prevent development or occurrence of severe disease symptoms.
- a binding agent in particular an antibody or antigen-binding fragment or a nucleic acid encoding it or a vector comprising such nucleic acid or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment as described herein such as to prevent infection overall, or such as to prevent development or
- a therapeutically effective amount of a binding in particular an antibody or antigenbinding fragment, nucleic acid, vector or pharmaceutical composition is administered to a subject in need thereof.
- a prophylactically effective amount of a binding agent in particular an antibody or antigen-binding fragment, nucleic acid, vector or pharmaceutical composition is administered to a subject in need thereof.
- a “therapeutically effective amount” or “therapeutically effective dose” indicates an amount of binding agent, in particular antibody or antigen-binding fragment, nucleic acid, vector or pharmaceutical composition that when administered to the subject brings about a clinical positive response with respect to therapeutic treatment of the subject afflicted by a Sarbecovirus infection, such as, e.g. curing infection with a Sarbecovirus .
- a “prophylactically effective amount” or “prophylactically effective dose” refers to an amount of binding agent, in particular antibody or antigen-binding fragment, nucleic acid, vector or pharmaceutical composition that prevents, inhibits or delays the onset of a Sarbecovirus infection and/or prevents or reduces the risk of a clinical manifestation of a Sarbecovirus infection and/or reduces the severity, symptoms and/or duration of a Sarbecovirus infection in the subject.
- the binding agent in particular the antibody or antigen-binding fragment or the nucleic acid encoding it or the vector comprising such nucleic acid or the composition comprising the binding agent, in particular the antibody or antigen-binding fragment as described herein may need to be administered to a subject multiple times, such as with an interval of 1 week or 2 weeks; the interval being dictated by the pharmacokinetic behaviour or characteristics (e.g. half-time or half-life in the subject’s circulation) of the binding agent, in particular the antibody or antigen-binding fragment, nucleic acid or vector.
- a single dose of a binding agent, in particular an antibody or antigen-binding fragment as described herein is administered to the subject is envisaged.
- the single dose may be in the range of 0.5 mg/kg to 25 mg/kg.
- subject relates to any organism such as a vertebrate, particularly any mammal, including both a human and other mammals, for whom diagnosis, therapy or prophylaxis is desired, e.g., an animal such as a rodent, a rabbit, a cow, a sheep, a horse, a dog, a cat, a lama, a pig, or a non-human primate (e.g., a monkey).
- the rodent may be a mouse, rat, hamster, guinea pig, or chinchilla.
- the subject is a human, a rat or a non-human primate.
- the subject is a human.
- a subject is a subject, such as a human subject, with or suspected of having an infection with a Sarbecovirus, also designated ’’patient” or “subject” herein.
- the subject is a mammal susceptible to infection with a Sarbecovirus , such as a human subject that is susceptible to infection with SARS-CoV-2 such as a SARS-CoV-2 variant, or SARS-CoV-1.
- the pharmaceutical composition of the invention can be administered to any patient in accordance with standard techniques.
- the administration can be by any appropriate mode, including oral, parenteral, topical, nasal, ophthalmic, intrathecal, intra-cerebroventricular, sublingual, rectal, vaginal, and the like. Still other techniques of formulation such as nanotechnology and aerosol and inhalant are also within the scope of this invention.
- the dosage and frequency of administration will depend on the age, sex and condition of the patient, concurrent administration of other drugs, counterindications and other parameters to be taken into account by the clinician.
- the binding agent in particular the antibody or antigen-binding fragment, the nucleic acid, the vector or the pharmaceutical composition may be administered to a subject via intravenous injection, subcutaneous injection, or intranasally, or, alternatively via inhalation or pulmonary delivery.
- a further aspect of the invention relates to a binding agent, in particular an antibody or antigenbinding fragment, as described herein for use in diagnosing a Sarbecovirus infection, for use as a diagnostic agent.
- a nucleic acid encoding a Sarbecovirus binding agent, in particular a Sarbecovirus antibody or antigen-binding fragment as described herein, a (recombinant) vector comprising such nucleic acid, or a composition comprising a Sarbecovirus binding agent, in particular a Sarbecovirus antibody or antigen-binding fragment as described herein, can likewise be for use.
- binding agent in particular an antibody or antigen-binding fragment, as described herein in the manufacture of a (in vitro) diagnostic agent or diagnostic kit
- the binding agent, in particular the antibody or antigen-binding fragment, as described herein may be for use in detecting the presence (or absence) of a Sarbecovirus or a part thereof (such as a Sarbecovirus spike protein or a part thereof) in a sample, such as a sample obtained from a subject, such as from a subject suspected to be infected with a Sarbecovirus .
- a nucleic acid encoding a binding agent, in particular an antibody or antigen-binding fragment, as described herein, a (recombinant) vector comprising such nucleic acid or composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein can likewise be used in the manufacture of a diagnostic agent or diagnostic kit, such as an in vitro diagnostic agent or kit.
- a further aspect relates to methods for detecting a Sarbecovirus in a sample, such as a sample obtained from a subject, such as from a subject suspected to be infected with a Sarbecovirus .
- Such methods usually comprise the steps of obtaining a sample, contacting the sample with a binding agent, in particular an antibody or antigen-binding fragment, as described herein, and detecting, determining, assessing, assaying, identifying or measuring binding of the binding agent, in particular the antibody or antigen-binding fragment, with a Sarbecovirus or a part thereof (such as a Sarbecovirus spike protein or a part thereof).
- the Sarbecovirus is selected from the group of clade la, lb, 2 and/or clade 3 Sarbecoviruses.
- SARS-CoV-2 such as SARS-CoV-2, Khosta-2, GD-Pangolin, RaTG13, WIV1, LYRal l, RsSHC014, Rs7327, SARS-CoV-1, Rs4231, Rs4084, Rp3, HKU3-1, or BM48-31 viruses, preferably SARS-CoV-2 such as a SARS-CoV-2 variant or SARS- CoV-1.
- the binding agent in particular the antibody or antibody fragment, as described herein is comprising a detectable moiety fused to it, bound to it, coupled to it, linked to it, complexed to it, or chelated to it.
- a “detectable moiety” in general refers to a moiety that emits a signal or is capable of emitting a signal upon adequate stimulation, or to a moiety that is capable of being detected through binding or interaction with a further molecule (e.g. a tag, such as an affinity tag, that is specifically recognized by a labelled antibody), or is detectable by any means (preferably by a non-invasive means, if detection is in vivo/ inside the human body).
- detectable moiety may allow for computerized composition of an image, as such the detectable moiety may be called an imaging agent.
- Detectable moieties include, without limitation, fluorescence emitters, phosphorescence emitters, positron emitters, radioemitters, etc., enzymes (capable of measurably converting a substrate) and molecular tags.
- radioemitters/radiolabels examples include 68Ga, HOmln, 18F, 45Ti, 44Sc, 47Sc, 61Cu, 60Cu, 62Cu, 66Ga, 64Cu, 55Ca, 72As, 86Y, 90Y, 89Zr, 1251, 74Br, 75Br, 76Br, 77Br, 78Br, U lin, 114mln, 114In, 99mTc, 11C, 32C1, 33C1, 34C1, 1231, 1241, 1311, 186Re, 188Re, 177Lu, 99Tc, 212Bi, 213Bi, 212Pb, 225Ac, 153Sm, and 67Ga.
- Fluorescence emitters include, without limitation, cyanine dyes (e.g. Cy5, Cy5.5, Cy7, Cy7.5), FITC, TRITC, coumarin, indolenine-based dyes, benzoindolenine-based dyes, phenoxazines, BODIPY dyes, rhodamines, Si-rhodamines, Alexa dyes, and derivatives of any thereof.
- molecular tags include affinity tags, such as chitin binding protein (CBP), maltose binding protein (MBP), glutathione-S-transferase (GST), poly(His) (e.g., 6x His or His6), biotin or streptavidin, such as Strep-tag®, Strep-tag II® and Twin- Strep-tag®; solubilizing tags, such as thioredoxin (TRX), poly(NANP) and SUMO; chromatography tags, such as a FLAG-tag; epitope tags, such as V5-tag, myc-tag and HA-tag; fluorescent labels or tags (i.e., fluorochromesZ-phores), such as fluorescent proteins (e.g., GFP, YFP, RFP etc.); luminescent labels or tags, such as luciferase, bioluminescent or chemiluminescent compounds (such as luminal, isoluminol, theromatic
- Binding agents in particular antibodies and antibody fragments, as described herein and comprising a detectable moiety may for example be used for in vitro, in vivo or in situ assays (including immunoassays known per se such as ELISA, RIA, EIA and other "sandwich assays", etc.) as well as in vivo imaging purposes, depending on the choice of the specific label. .
- kits comprising a binding agent, in particular an antibody or antigenbinding fragment, a nucleic acid encoding it, a vector comprising such nucleic acid as described herein, a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, or a pharmaceutical composition comprising a binding agent, in particular an antibody or antigen-binding fragment; a nucleic acid encoding it a vector comprising such nucleic acid or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment as described herein.
- kits may be pharmaceutical kits or medicament kits which are comprising a container or vial (any suitable container or vial, such as a pharmaceutically acceptable container or vial) comprising an amount of binding agent, in particular antibody or antigen-binding fragment, or nucleic acid encoding it or vector comprising such nucleic acid as described herein or composition comprising a binding agent, in particular an antibody or antigen-binding fragment as described herein, and further comprising e.g. a kit insert such as a medical leaflet or package leaflet comprising information on e.g. intended indications (prophylactic or therapeutic treatment of a Sarbecovirus infection) and potential side-effects.
- Pharmaceutical kits or medicament kits may further comprise e.g.
- a syringe for administering the binding agent, in particular the antibody or antigen-binding fragment, nucleic acid encoding it vector comprising such nucleic acid or composition comprising the binding agent, in particular the antibody or antigen-binding fragment as described herein to a subject.
- kits may also be diagnostic kits comprising a container or vial (any suitable container or vial, such as a pharmaceutically acceptable container or vial) comprising an amount of binding agent, in particular antibody or antigen-binding fragment, as described herein, such as a binding agent, in particular an antibody or antigen-binding fragment thereof comprising a detectable moiety.
- diagnostic kits may further comprise e.g. one or more reagents to detect the detectable moiety and/or e.g. instructions on how to use said binding agent, in particular antibody or antigen-binding fragment, for detection of a Sarbecovirus in a sample.
- HEK293-S cells (ThermoFisher Scientific) were cultured in FreeStyle 293 expression media (Life Technologies), at 37°C with 8% CO2 while shaking at 130 rpm.
- HEK293-T cells (ATCC) and Vero E6 cells (ATCC) were cultured at 37°C in the presence of 5% CO2 in DMEM supplemented with 10% heat-inactivated fetal bovine serum (FBS), 1% penicillin, 1% streptomycin, 2 mM 1-glutamine, non-essential amino acids (Invitrogen) and 1 mM sodium pyruvate.
- FBS heat-inactivated fetal bovine serum
- penicillin 1%
- streptomycin 2 mM 1-glutamine
- non-essential amino acids (Invitrogen) and 1 mM sodium pyruvate.
- ExpiCHO-S cells were cultured at 37°C with 8% CO2 while shaking at 130 rpm in ExpiCHO expression media (GIBCO).
- Vero E6-TMPRSS2 cells that stably express human TMPRSS2 (NIBIOHN, JCRB 1819) (Matsuyama et al., PNAS, 2020) were cultured in DMEM containing 10% FBS, penicillin (100 unit/mL), streptomycin (100 ug/mL), Geneticin (G418) (Img/ml). When Vero E6-TMPRSS2 cells were seeded for assays medium without Geneticin was used.
- R3 DC23-Fc(YTE) also referred to herein as huR3DC23-Fc
- XVR013 also referred to herein as R3 DC 23hum-Fc(LS) or huR3DC23-Fc LS or R3DC23-Fc LS
- gBlocks Upon arrival, gBlocks were solubilized in ultraclean water at a concentration of 20 ng/pL. gBlocks were A-tailed using the NEBNext-dA-tailing module (NEB), purified using CleanPCR magnetic beads (CleanNA) and inserted in pcDNA3.4-TOPO vector (ThermoFisher). The ORF of positive clones was fully sequenced, and pDNA of selected clones was prepared using the NucleoBond Xtra Midi kit (Machery-Nagel).
- NEB NEBNext-dA-tailing module
- CleanNA CleanPCR magnetic beads
- pcDNA3.4-TOPO vector ThermoFisher.
- the ORF of positive clones was fully sequenced, and pDNA of selected clones was prepared using the NucleoBond Xtra Midi kit (Machery-Nagel).
- a construct was generated wherein a humanized form of R3 DC23 (SEQ ID NO:2) was fused to a human IgGl Fc containing the LS mutation (SEQ ID NO: 83) via a (648)2 linker (SEQ ID NO: 88) at the Fc N-terminus (R3_DC23hum-Fc(LS) or huR3DC23-Fc_LS or XVR013) (SEQ ID NO: 124).
- the gene encoding huR3DC23-Fc_LS was codon optimized, synthesized, and cloned into the pXLG6 backbone vector at ATUM’s laboratories.
- the R3DC23 DNA sequence was inserted into pXLG6 expression vector and transfected in CHOExpress TM cells at a cell density of 4.00E+6 cells/ml.
- TGE supernatant was harvested by centrifugation and clarified by filtration (0.2pm) after 10 days when cell viability dropped below 10%.
- the protein was further purified by Protein A.
- VHH-Fc_LS constructs SEQ ID NO:96-99, 101,103, 105-109,111-119
- the genes encoding the VHH-Fc_LS constructs were each codon optimized, synthesized and cloned into the manufacturer pcDNA3. 1 backbone vector.
- VHH-Fc_LS proteins SEQ ID NO:96-99, 101, 103, 105-109, 111-119
- the ExpiCHOTM Expression System Thermo Fisher Scientific, Cat. No. A29133.
- the ExpiCHO-S cells were expanded from a working cell bank and passaged in ExpiCHO expression media according to the manufacturer's instructions. The mixtures for transfection were prepared following the protocol instructions described in the ExpiCHO-S system manual. ExpiCHO cells were cultured for 14 days and the harvest was performed by filtration using 0.22 pm filtration units and DE as filter aid (Sartorius, Cat. No. SDLV-0150-05E0-2) after which they were processed immediately.
- the produced antibodies (SEQ ID NO:96-99, 101, 103, 105-109, 111-119) were captured from clarified supernatants using a HiTrap MabSelect PrismA 25 mL column (Cytiva, Cat. nr. 17-5498- 54) on an AKTA Pure 25 FPLC system.
- the column equilibration and protein binding were performed using 20 mM sodium phosphate with 150 mM NaCl pH 7.4 and for protein elution, 100 mM sodium citrate with 150 mM NaCl pH 3.5 was used. After elution, the peak corresponding to affinity purified antibodies was immediately neutralized with 30 % of 1 M Tris pH 8.0.
- protein samples were loaded onto a HiLoad 26/600 Superdex 200 pg (Cytiva, Cat. nr. 28-9893-36) on an AKTA Pure 25 FPLC system.
- the column was equilibrated in 50 mM L-Histidine, 150 mM L-Arginine, 150 mM NaCl pH 7.0 for all constructs except for constructs defined by SEQ NO: 117 and SEQ ID: 119 for which pH 6.0 was used.
- fractions corresponding to the monomeric antibody form were pooled from a 96 deep well plate into a 50 ml falcon tube and filtered through a 0.22 pm PES membrane (Fisher brand, Cat. nr. 15206869, Lot nr. 2103171806) in a laminar flow chamber. Protein samples were transferred to 50 kDa MWCO spin concentrators (Amicon 50K Cat. nr. UFC905024; Lot nr. 0000187574) for concentration and each round of centrifugation was 10 min at 4000 g, and it was repeated until the desired concentration of 2 mg/mL was achieved.
- VHH-Fc samples were characterized by analytical Size Exclusion Chromatography (SEC) to determine the molecular composition of each sample. After rapid thawing in a warm water bath at 25 °C, 10 min centrifugation at 16,000 g and transfer of supernatant to fresh tubes, 5 pl was injected on an AdvanceBioSECs column, 4 (Agilent) with 2.7 pm porous particle size and 300 A pore size, calibrated with PBS. The mobile phase used for was IxPBS with 0.02 sodium azide. The separation was monitored by absorbance at 280 nm with a 16 nm bandwidth, no reference subtraction.
- SEC Size Exclusion Chromatography
- proteins were separated on reducing and non-reducing SDS-PAGE.
- the samples are mixed with 2x concentrate Laemmli buffer containing no DTT (non-reduced conditions) or 200 mM DTT (reducing condition). 3 pg of protein sample are loaded on 12% SDS- PAGE and stain with Coomassie Brilliant Blue.
- HEK293-S cells were transfected with VHH-Fc (S) encoding plasmids using polyethylenimine (PEI). Briefly, suspension-adapted and serum -free HEK293-S cells were seeded at 3 x 10 6 cells/mL in FreeStyle 293 medium (ThermoFisher Scientific). Next, 4.5 pg of pcDNA3.3 -VHH-Fc plasmid DNA was added to the cells and incubated on a shaking platform at 37°C and 8% CO2, for 5 min. Next, 9 pg of PEI was added to the cultures, and cells were further incubated for 5 h, after which an equal culture volume of Ex-Cell-293 (Sigma) was added to the cells.
- PEI polyethylenimine
- huR3DC23-Fc_YTE was expressed in ExpiCHO-STM cells (ThermoFisher Scientific), according to the manufacturer’s protocol. Briefly, a 50 mb culture of 6 x 10 6 cells per mb, grown at 37°C and 8% CO2, was transfected with 40 pg of pcDNA3.3-VHH72-Fc plasmid DNA using ExpiFectamineTM CHO reagent. One day after transfection, 300 pL ExpiCHOTM enhancer and 8 mb ExpiCHOTM feed was added to the cells, and cultures were further incubated at 32°C and 5% CO2. Cells were fed a second time day 5 after transfection. Productions were collected as soon as cell viability dropped below 75%.
- VHH-Fc proteins For purification of the VHH-Fc proteins, supernatants were loaded on a 5 mb MAbSelect SuRe column (GE Healthcare). Unbound proteins were washed away with Mcllvaine buffer pH 7.2, and bound proteins were eluted using Mcllvaine buffer pH 3. Immediately after elution, proteincontaining fractions were neutralized using 30% (v/v) of a saturated Na3PO4 buffer. Next, these fractions were pooled, and loaded on a HiPrep Desalting column for buffer exchange to PBS pH7.4.
- the gene encoding huR3DC23-Fc_LS was codon optimized, synthesized, and cloned into the pXLG6 backbone vector at ATUM’s laboratories.
- parental CHOExpressTM cells were co-transfected with the expression vector and the pXLG5 helper vector.
- the stable pool was generated under 50 mg/L puromycin selective pressure (applied daily) and further expanded.
- the stable pool research cell bank was banked at day 14 when cell viability reached 95%.
- the RCB pool was then expanded for protein production at IL scale and cultured until day 12 (cell density 3.5 x 10 7 cells/mL, cell viability 96%). The supernatant was harvested by centrifugation and clarified by filtration (0.2 pm). The protein was further purified by Protein A using MabSelect SuRe LX resin. Consecutive washed were performed with 20 mM sodium phosphate and 110 mM NaCl at pH 7.2; 100 mM sodium acetate and 500 mM NaCl at pH 5.5; and 20 mM sodium phosphate at pH 7.2. The eluate in 100 mM sodium acetate pH 3.5 was neutralized to pH 7.0 by addition of 1 M Tris pH 11 (10%v/v). After filter sterilization (0.22 pm), the protein was aliquoted at 2 mg/ml.
- K18-hACE2 mice B6.Cg-Tg(K18-ACE2)2Prlmn/J (7-9 weeks of age) were purchased from The Jackson Laboratory and bred in house under specific-pathogen-free (SPF) conditions. SARS-CoV-2 infections were performed under biosafety level 3 (BSL3) conditions. Antibody treatment was performed by intraperitoneal injection using a volume of 100 pl. Animals were anesthetized by isoflurane inhalation and 3xl0 2 PFU of SARS-CoV-2 D614G variant virus (SARS-CoV- 2/human/FRA/702/2020, obtained from the European Virus Archive) was administered by intratracheal instillation.
- SARS-CoV-2 D614G variant virus SARS-CoV- 2/human/FRA/702/2020
- the right lung lobes were stored at -80°C.
- the lung samples were homogenized using a Precellys Evolution tissue homogenizer (Bertin- technologies).
- the lung homogenates were cleared by centrifugation (1,000 x g) for 15 min at 4°C and used to determine the viral titer by plaque assay on VeroE6-TMPRSS2 cells in duplicate using 12-well plates.
- the plates were incubated at 37°C for 2 hours. Subsequently the cells were washed twice and incubated in medium containing Avicel at a concentration of 0.3% (w/v). After 2 days of incubation at 37°C, the overlays were removed, the cells were fixed with 3.7% paraformaldehyde (PFA) and stained with 0.5% crystal -violet dye to visualize the viral plaques.
- PFA paraformaldehyde
- XVR013, at the indicated doses, palivizumab (10 mg/kg), or bebtelovimab (10 mg/kg) were administered by intraperitoneal injection either 4 hours after the SARS-CoV2 challenge (therapeutic setting) or approximately 24 hours prior to infection (prophylactic setting).
- XVR013m, XVR014 or XVR012 (1: 10 ratio of XVR013m:XVR014) were dosed, at the dose levels indicated, by intraperitoneal injection, 4 hours after the SARS-CoV2 challenge (therapeutic setting).
- huR3DC23-Fc_LS used in the hamster study was produced from stable cell pools. Hamsters were monitored daily for behavior, appearance and body weight. On day 4 post-infection, animals were euthanized. At the time of necropsy, gross pathology was performed and abnormalities were recorded. Samples from the right lung lobes were collected and frozen for virological analysis. To determine virus titers, quadruplicate
- the lower limit of detection ranged between 1.1 and 1.3 loglO TCID/g.
- LLOD lower limit of detection
- lung tissue homogenates were used. RNA was isolated and Taqman RT-qPCR was performed. The number of copies (LoglO CP/g) in the different samples was calculated against a standard included in each run.
- the LLOD was 3.5 LoglO CP/g. Blood samples were collected prior to the start of the study on day -2 ( ⁇ 200 pl blood was collected for serum under isoflurane anesthesia) and on day 4 post-infection (p.i.) at time of necropsy for pharmacokinetics analysis.
- Blood samples for serum were immediately transferred to appropriate tubes containing a clot activator. Serum was collected and stored frozen. To inactivate any potential infectious material present and to allow the testing of the serum samples in a BSL-2 environment, serum on day 4 postinfection was heat-treated at 56 °C for 30 minutes.
- the pCGl expression vector for the SARS-CoV-2 spike protein containing the D614G mutation was generated from the pCGl-SARS-2-Sdell8 vector by introducing the specific RBD mutations via QuickChange mutagenesis using appropriate primers, according to the manufacturer’s instructions (Aligent).
- the pCGl expression vector forthe Gamma variant was generated from the pCGl-SARS-2-D614G Sdell8 vector by ordering Gblocks corresponding to the Gamma SARS-CoV-2 amino acid sequence L7-V1189 containing the Gamma mutations (L18F, T20N, P26S, D138Y, R190S, K417T, E484K, N501Y, D614G, H655Y, T1027I, VI 176F), and cloning the Gblocks into the pCGl-SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Eta variant was generated from the pCGl-SARS-2-D614G Sdell8 vector by ordering a Gblock corresponding to the Eta SARS-CoV-2 amino acid sequence S45-P894 containing the Etamutations (Q52R, A67V, H69-, V70-, Y144-, E484K, D614G, Q677H, F888L), and cloning the Gblock into the pCGl-SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Iota variant was generated from the pCGl-SARS-2-D614G Sdell8 vector by ordering a Gblock corresponding to the Iota SARS-CoV-2 amino acid sequence M1-V267 containing the Iota mutations (L5F, T95I, D253G), and cloning the Gblock into the pCGl- SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the B. 1.617.3 variant was generated from the pCGl-SARS-2- D614G Sdel 18 vector by ordering a Gblock corresponding to the B. 1.617.3 SARS-CoV-2 amino acid sequence L10-A688 containing the B. 1.617.3 mutations (T19R, L452R, E484Q, D614G, P681R), and cloning the Gblock into the pCGl-SARS-2D614G Sdel 18 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Mu variant was generated from the pCGl-SARS-2-D614G Sdel 18 vector by ordering a Gblock corresponding to the Mu SARS-CoV-2 amino acid sequence D80-Q965 containing the Mu mutations (T95I, Y144S, Y145N, R346K, E484K, N501Y, D614G, P681H, D950N), and cloning the Gblock into the pCGl-SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Zeta variant was generated from the pCGl-SARS-2-D614G Sdel 18 vector by ordering a Gblock corresponding to the Zeta SARS-CoV-2 amino acid sequence R466-L1193 containing the Zeta mutations (E484K, D614G, VI 176F), and cloning the Gblock into the pCGl-SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Omicron BA.5 variant was generated from the pCGl-SARS-2- BA.2 Sdel 18 vector by introducing the H69-, V70- deletions and the L452R, F486V, R493Q mutations via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
- the pCGl expression vector for the Omicron CH.1.1 variant was generated from the pCGl-SARS- 2-XBB.1.5 Sdell8 vector by ordering a Gblock corresponding to the CH.1.1 SARS-CoV-2 amino acid sequence I65-Q503 containing the CH.1.1 mutations (G142D, K147E, W152R, F157L, I210V, V213G, G257S, Q339H, R346T, S371F, S373P, S375F, T376A, D405N, R408S, K417N, N440K, K444T, G446S, L452R, N460K, S477N, T478K, E484A, F486S), while reverting other XBB. 1.5 specific mutations, and cloning the Gblock into the pCGl-SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Omicron EG.5.1 variant was generated from the pCGl-SARS- 2 -XBB. 1.5 Sdell8 vector by introducing the F456L, Q52H mutations via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
- the pCGl expression vector for the Omicron BA.2.86.1 variant was generated from the pCGl- SARS-2-XBB.1.5 Sdell8 vector by ordering two Gblock’s corresponding to the BA.2.86.1 SARS- CoV-2 amino acid sequence S13-L1144 containing the BA.2.86.1 mutations (T19I, R21T, L24S, P25-, P26-, A27-, S50L, H69-, V70-, , V127F, G142D, Y144-, , F157S, R158G, N211I, L212-, V213G, L216F, H245N, A264D, I332V, G339H, K356T, S371F, S373P, S375F, T376A, R403K, D405N, R408S, K417N, N440K, V445H, G446S, N450D, L452W, N460K, S477N, T4
- the pCGl expression vector for the Omicron HK.3 variant was generated from the pCGl-SARS-2- EG.5.1 Sdell8 vector by introducing the L455F mutation via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
- the pCGl expression vector for the Omicron BA.2.74 variant was generated from the pCGl-SARS- 2-XBB.1.5 Sdell8 vector by ordering a Gblock corresponding to the BA.2.74 SARS-CoV-2 amino acid sequence T70-G501 containing the BA.2.74 mutations (G142D, V213G, G339D, R346T, S371F, S373P, S375F, T376A, D405N, R408S, K417N, N440K, L452M, S477N, T478K, E484A, Q493R), while reverting the XBB.1.5 specific mutations, and cloning the Gblock into the pCGl- SARS-2XBB.1.5 Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Omicron JN. l variant was generated from the pCGl-SARS-2- BA.2.86.1 Sdell8 vector by introducing the L455S mutation via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
- the pCGl expression vector for the Omicron KP.3 variant was generated from the pCGl-SARS-2- JN. l Sdell8 vector by ordering a Gblock corresponding to the KP.3 SARS-CoV-2 amino acid sequence H438-D1110 containing the KP.3 mutations (F456L, Q493E, V1104L), and cloning the Gblock into the pCGl-SARS-2JN. l Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Omicron KP.1.1 variant was generated from the pCGl-SARS- 2 -KP.3 Sdell8 vector by ordering a Gblock corresponding to the KP.1.1 SARS-CoV-2 amino acid sequence P330-S1089 containing the KP.1.1 mutations (R346T and K1086R), while reverting KP.3 specific mutations and cloning the Gblock into the pCGl-SARS-2-KP.3 Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Omicron KP.3. 1. 1 variant was generated from the pCGl-SARS- 2-KP.3 Sdell8 vector by introducing the S31- deletion via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
- the pCGl expression vector for the Omicron KP.2.3 variant was generated from the pCGl-SARS- 2-KP.3 Sdell8 vector by ordering a Gblock corresponding to the KP.2.3 SARS-CoV-2 amino acid sequence L10-V494 containing the KP.2.3 mutations (S31-, H146Q, R346T), while reverting the KP.3 specific mutations, and cloning the Gblock into the pCGl-SARS-2KP.3 Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Omicron LB. 1 variant was generated from the pCGl-SARS-2- JN. l Sdell8 vector by ordering a Gblock corresponding to the LB. l SARS-CoV-2 amino acid sequence V11-A467 containing the LB.l mutations (S31-, Q183H, R346T and F456L) and cloning the Gblock into the pCGl-SARS-2JN. l Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- the pCGl expression vector for the Omicron XEC variant was generated from the pCGl-SARS-2- KP.3 Sdell8 vector by ordering a Gblock corresponding to the XEC SARS-CoV-2 amino acid sequence V3-D75 containing the XEC mutations (T22N and F59S) and cloning the Gblock into the pCGl-SARS-2-KP.3 Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
- VSV vesicular stomatitis virus
- a negative control antibody (palivizumab) or isotype control antibody, and/or a positive control (R3DC23-Fc_LS (XVR013), sotrovimab and/or bebtelovimab) were included in the assay.
- R3DC23-Fc_LS XVR013
- sotrovimab and/or bebtelovimab were included in the assay.
- two or three independent experiments were performed.
- replication-deficient VSV pseudotyped viruses were generated. Briefly, HEK 293T cells, transfected with SARS-CoV-2 Spike were inoculated with a replication deficient VSV vector containing enhanced green fluorescent protein (GFP) and firefly luciferase expression cassettes.
- GFP enhanced green fluorescent protein
- the inoculum was removed, the cells were washed with PBS and incubated in media supplemented with an anti -VSV G monoclonal antibody for 16 hours. Afterwards, pseudoviruses were harvested and clarified by centrifugation. Pseudoviruses were incubated for 1 hour at 37°C with different dilutions of the molecules, the positive control (XVR013, sotrovimab and/or bebtelovimab) and/or the negative control (palivizumab or isotype control) antibody. Next, the incubated pseudoviruses with molecules or controls were added to the subconfluent monolayers of Vero E6 cells for 16 hours at 37°C.
- the cells were lysed using passive lysis buffer.
- the transduction efficiency was quantified by measuring the GFP fluorescence in the prepared cell lysates using a Tecan Infinite 200 Pro plate reader.
- the GFP fluorescence was normalised against the GFP fluorescent of the noninfected cells and the virus infected cells.
- the mean IC50 values were calculated based on one, two or three independent experiments as indicated by nonlinear regression curve fitting (log(inhibitor) versus normalised response (four parameters)).
- SARS-CoV-2 viruses belonging to different lineages D614G, Alpha, Beta, Gamma, Delta, Epsilon B. 1.427, Epsilon B..429, Zeta, Kappa, Omicron BA. 1, Omicron BA.2 and Omicron BA.5 were isolated from nasopharyngeal swabs taken from patients/travelers between January 2020 and July 2022.
- the inoculum was removed and cells were incubated at 37°C with infection medium (up to 18-24 hours post-infection).
- the SARS-CoV2 infected cells were fixed and immunostained with a SARS-CoV nucleocapsid antibody (Sino Biological, Catalogue number: 40143-MM05), followed by HRP -conjugated goat anti-mouse IgG (H+L) secondary antibody (Invitrogen, catalogue number A16072). Spots (infected cells) were counted using Immunospot Image Analyser. For each molecule, the compound concentration showing 50, 80 or 90 % reduction in infection (IC50, ICso or IC90) was calculated based on the Zielinska method. The geometric mean values were calculated based on three independent runs.
- HEK293T cells pre-seeded for 1 day in 6 well plates, were transfected with 800 ng of vector expressing GFP fused to the spike of either Khosta-2 sarbecovirus (Seifert et al. (2022 PLoS Pathog 18(9):e 1010828) or SARS-CoV-2 variant D614G, complexed to polyethylenimine (PEI) in a 1:3 ratio.
- Khosta-2 sarbecovirus Seifert et al. (2022 PLoS Pathog 18(9):e 1010828
- SARS-CoV-2 variant D614G complexed to polyethylenimine (PEI) in a 1:3 ratio.
- PEI polyethylenimine
- SPR surface plasmon resonance
- the non-specific binding of the S2 targeting molecules to the surface of mammalian (HEK293) cells was investigated with a flow cytometry experiment.
- Sotrovimab biosimilar was used as negative control antibody in the assay and XVR013 was used as positive control. All procedures were performed on ice.
- the cells were washed once with PBS and blocked with 1% BSA. Afterwards, the cells were incubated with two concentrations (10 and 100 pg/mL) of the S2 targeting molecules or controls (Sotrovimab Biosimilar and XVR013) for 90 minutes and subsequently washed 3 times with PBS containing 1% BSA. Next, the secondary antibody (anti-human IgG AF633) was incubated for 30 minutes.
- Binding of the molecules or controls was detected using an anti-human IgG AF633 antibody and dead cells were stained using live/dead stain (Invitrogen). Following 3 washes with PBS containing 0.5% BSA, the cells were analyzed by flow cytometry. The median fluorescence intensity of the AF633-conjugated anti-human IgG was measured to detect the binding of the molecules or controls to HEK293 cells.
- the DRB1 score was calculated.
- the DRB 1 score is an approximate score representing a potential immunogenic risk for a peptide and is calculated from the population frequencies of the affected allotypes. Hydrophobic interaction chromatography (HIC) assay
- HIC hydrophobic interaction chromatography
- the mobile phases used for HIC were (A) 1.6 M ammonium sulfate and 50 mM phosphate pH 6.0 (variants CBV2-Fc_LS and CBV4-Fc_LS) or pH 7.0, and (B) 50 mM phosphate pH 6.0 (variants CBV2-Fc_LS and CBV4-Fc_LS) or pH 7.0.
- the S2 targeting molecules were concentration adjusted to Img/mL in 50 mM L-Histidine, 150 mM L-Arginine, pH 6.0 (variants CBV2-Fc_LS and CBV4-Fc_LS) or pH 7.0, and fdtered through a 0.45 pM fdter.
- the proteins (50 pL) were diluted 1: 1 with buffer A to a final ammonium sulfate concentration of 800 mM, mixed and 40 pL of the diluted proteins were injected onto the column. Following a 5 min hold at 50% A and 50% B, bound protein was eluted using a linear gradient from 50 to 100% B in 50 min followed by 5 min hold at 100% B. The column was washed with 100% B, followed by 50 mM ammonium acetate pH 5 and re-equilibration in 50% B for 10 min prior to the next sample. The separation was monitored by absorbance at 214 nm, no reference subtraction.
- the charge proteins variants were assessed using a Strong Cation Exchange Hydrophilic interaction chromatography (SCX) assay employing a MabPac SCX-10 column (Thermo Fisher 063655), containing a stationary phase consisting of an ethylvinylbenzene-divinylbenzene copolymer non- porous beads coated with a layer of highly hydrophilic material consisting sulfonic acid functional groups. All separations were carried out on an Agilent 1100/1260 HPLC equipped with a UV/VIS detector at fixed flow rate of 500 pL/min.
- the mobile phases used for SCX were (A) 20 mM MES pH 5.6, and (B) 20 mM HEPES and 120 mM NaCl pH 8.0.
- the S2 targeting molecules were formulated at Img/mL in 50 mM L-Histidine, 150 mM L-Arginine, 0.02% polysorbate 80, 0.02% sodium azide, pH 6.0 (variants CBV2-Fc_LS and CBV4-Fc_LS) or pH 7.0, and filtered through a 0.45 pM filter.
- the proteins (20 pL) were diluted 1: 1. Proteins (20 pL sample pre-filtered on 0.45 pm filter) were injected onto the column. Following a 2 min hold at 100% A, bound protein was eluted using linear gradient from 100% A to 100% B in 18 min followed by 2 min hold at 100% B, and 4 min hold at 100% A. The separation was monitored by absorbance at 214 nm.
- DSF Di fferential Scanning Fluorimetry
- SLS Static Light Scattering
- PLS Dynamic Light Scatting
- the protein stability was assessed by the determination of their respective melting temperature (Tonset and Tm), aggregation temperatures (Tagg) and particles sizing (AVG-Z, average diameter) using an Unchained Labs UNCLE system running with the Supplier Software v4.01. A temperature gradient of 25 -95 °C was applied and the protein melting temperatures were detected by intrinsic fluorescence (barycentric mean of spectrum, BCM), and protein aggregation propensity by static light scattering with detection at 266 nm (Tagg 266) and 473 nm (Tagg 473).
- Tonset and Tm melting temperature
- Tagg aggregation temperatures
- AVG-Z average diameter
- the S2 targeting molecules candidates were formulated at Img/mL in 50 mM L-Histidine, 150 mM L-Arginine, 0.02% polysorbate 80, 0.02% sodium azide, pH 6.0 (variants CBV2-Fc_LS and CBV4-Fc_LS) or pH 7.0, and filtered through a 0.45 pM filter.
- All proteins were formulated at 1 mg/ml in 50 mM L-His, 150 mM L-Arg at either pH 6 (CBV2- Fc_LS and CBV4-Fc_LS) or pH 7 (RNGL-Fc_LS, RAGL-Fc_LS, E2NGL-Fc_LS, E2AGL-Fc_LS, QRAGL-Fc_LS, ERAGL-Fc_LS, E10RAGL-Fc_LS, E81RNGL-Fc_LS, E81RAGL-Fc_LS, Q75- Fc_LS, E75-Fc_LS, E10E81-Fc_LS, D16-Fc_LS, T19D79-Fc_LS, TQD-Fc_LS, Q75R21Y-Fc_LS, CBVl-Fc_LS and CBV3-Fc_LS), supplemented with 0.02% polysorbate-80, and 0.02% NaN3 to prevent microbial growth.
- each sample was split into two 750 pl aliquots (one for each time point) that were snap-frozen in liquid nitrogen and stored at -80 °C. Samples were rapidly thawed in a water bath at room temperature, followed by centrifugation at 16,000 xg for 10 min to remove potential freeze-thaw aggregates. Supernatant was transferred (only 540 pl to avoid disturbing any freeze-thaw-aggregate pellet) and split into three replicates (120 pl each) in sterile pyrogen-free polypropylene 2 ml screw-cap tubes. Samples were either analyzed immediately afterwards, or used for 14 days accelerated ageing at 37 °C, in a test tube rotatory mixer at 10 rpm, and analyzed subsequently.
- Variants CBV2-Fc_LS and CBV4-Fc_LS were formulated at Img/mL in 50 mM L-Histidine, 150 mM L-Arginine, 0.02% polysorbate-80, 0.02 sodium azide at pH6.0, all other variants were formulated at Img/mL in 50 mM L-Histidine, 150 mM L-Arginine, 0.02% polysorbate-80, 0.02 sodium azide at pH7.0.
- the study samples were prepared in triplicate. 120 pL sample was filled in 2mL screw-cap polypropylene tube and incubated for 2 weeks at 37°C under mild agitation (10 rpm) in turning wheel. All of the variants samples were characterized and analyzed in parallel at time point 0 and after 14 days stress.
- UV/VIS spectrometry performed in an Unchain Labs Lunatic system; total protein quantification at 280 nm complemented by background absorbance at higher wavelengths Analytical SEC to assess degradation/multimerization/aggregation
- XVR014, S2 targeting molecules and a reference molecule (XVR013) were tested in human neonatal Fc receptor (hFcRn) tg32 SCID mice (B6.Cg-FcgrttmlDcr Prkdcscid Tg(FCGRT)32Dcr/DcrJ).
- hFcRn human neonatal Fc receptor
- tg32 SCID mice B6.Cg-FcgrttmlDcr Prkdcscid Tg(FCGRT)32Dcr/DcrJ.
- Six animals per group were included and two dose levels (2 mg/kg and 10 mg/kg) were tested using a single intravenous (IV) administration for evaluating XVR014.
- Five male animals were included per group and one dose level (10 mg/kg) was tested using a single intravenous (IV) administration for testing the S2 targeting molecules and XVR013.
- ADAs pre-existing anti-drug antibodies
- the biosimilar caplacizumab (Proteogenix) was used as negative control and the biosimilar TAS266 (Proteogenix) as a positive control.
- an ELISA method was developed to facilitate the screening of a panel of 70 adult human sera samples (35 male and 35 female of Caucasian, Hispanic, Black or other origin between 18 and 65 years old). Plates were pre-coated with a test compound or a control. Following the addition of human serum, the presence of pre-existing ADAs was detected using an anti-HRP tagged anti-light chain IgG (Fisher Scientific).
- a cut-point was calculated as mean + 1.645*SD (if data were not normally distributed, the cut-point was calculated using the 95th percentile). This cut-point was used to calculate the percentage of healthy adult human donors with pre-existing ADAs against the test compound.
- Saturation mutagenesis primers were designed according to selected mutation sites and custom produced at Genewiz.
- hR3_DC23 SEQ ID NO:2 fragments to be mutated were amplified and ligated by overlap PCR (Q5® Hot Start High-Fidelity 2X Master Mix, New England Biolabs #M0494), followed by gel recovery (QIAquick PCR Purification Kit, Qiagen # 28106).
- Phage vector plasmids and PCR products were digested by Sfi I in rCutSmart® Buffer (New England Biolabs # RO 123 and B7204S) and subsequently purified.
- the purified vector and PCR products were ligated with T4 DNA ligase (New England Biolabs #M02020M and #B0202S), and ligated products were desalted by Gel and PCR Clean-up kit (QIAquick PCR Purification Kit, Qiagen # 28106).
- Phage libraries were pre-incubated with streptavidin (SA)-coated magnetic beads (SA beads, ThermoFisher, 11206) to remove non-specific binders, and then incubated with SA-Bio SARS-Cov2 magnetic beads at room temperature, followed by washing 15 times with 1 x PBST. After washing, 20 ml of MC106F cells were infected at 37°C for 30 minutes. The infected cells were serially diluted and spread on 2xYT plates (containing 100 ug/ml ampicillin) and cultured at 37°C overnight to form colonies. Then 20 microliters of M13K07 helper phage at 5E12/ml were added to the above infected cells and cultured at 37°C for 30 minutes.
- SA streptavidin
- SA beads ThermoFisher, 11206
- the phage pellet obtained after centrifugation was resuspended in 1 ml of 1 x PBS, the debris was discarded by centrifugation at high speed for 5 minutes, and the resulting supernatant was transferred to a new tube and prepared for the second and third rounds of panning.
- the supernatant expressing VHH was tested for binding to biotinylated SARS-CoV-2 spike protein (ACROBiosystems #SPN-C82Ey).
- High-binding ELISA plates were coated with biotinylated SARS- CoV-2 spike protein and additional plates coated with streptavidin protein as a non-specific control, 100 pl/well at 4°C overnight. The next day the plates were washed three times with wash buffer (0.05% PBST). The plates were washed three times after being blocked with 5% milk-PBS for 1 hour. Subsequently, 100 pl of the PPE were added into each well, the plates were incubated at room temperature for 1 hour and washed six times.
- HRP horseshoe radish peroxidase
- anti-Flag-HRP anti-Flag-HRP
- TMB tetramethylbenzidine
- the plasmids encoding full-length heavy-chain-only antibodies were transfected into mammalian host cells (such as Expi293F human cells), and the purified antibodies were obtained by using conventional recombinant protein expression and purification techniques. Specifically, Expi293FTM cells (Gibco, #A 14527) were subcultured and expanded in MetaCell®HEK293-100 expression medium (Cellplus Bio, #L2000-1000). On the day prior to transfection, the viable cell density was adjusted to 3-4 x 10 6 cells/mL and the cells were allowed to grow in 37°C, 8% CO2 shaker overnight.
- mammalian host cells such as Expi293F human cells
- the cell viability and viable cell density reached to 95-99% and approximately 7 x 10 6 cells/mL, respectively. Then the cells were diluted to viable density of 3.6 x 10 6 cells/mL with fresh MetaCell®HEK293-100 expression medium.
- the plasmids mentioned above were transfected into prepared cells with Opti-MEMTM reduced serum medium (Gibco, #31985070) diluted PEI-MAX (Polysciences, #24765). Finally, the transfected cells were cultured in 37°C, 8% CO2 shaker for 5-6 days and harvested when cell viability was decreased to approximately 75%.
- the cell culture media were harvested by centrifugation at 3,300 g for 10 min and filtered by 24-well filter plate (Cytiva, #97032).
- the filtered supernatants were loaded on to pre-equilibrated 24-well purification plate (Cytiva, #7700-9904) containing MabSelect PrismA (Cytiva, #17549802) resin and re-equilibrated by washing with 5-10 column volumes (CV) of PBS.
- the target recombinant antibodies were eluted by acidic buffer (0.1M Citrate buffer, pH3.0), then neutralized with basic buffer (2M Tris-HCl, pH9.0).
- the sample concentration was determined by NanoDrop (Thermo ScientificTM NanoDropTM One) and stored at 4-degree for later use.
- Example 1 Fc fusions of the HR2 targeting VHH.R3 DC23 can protect mice against a lethal viral challenge with SARS-CoV-2
- K18-hACE2 mice that express human ACE2 at the surface of their epithelial cells were treated with 100 pg R3_DC23-Fc (SEQ ID NO: 125) or 100 pg isotype control IgG (palivizumab) 1 day prior to a lethal infection with D614G SARS-CoV-2 virus. Infected mice that were not treated were included as control.
- Figure 1 illustrates that in sharp contrast to mice that were either treated with isotype control antibody or untreated mice, all mice that were treated with R3_DC23-Fc survived the challenge and did not display significant bodyweight loss, indicating that R3_DC23-Fc can protect mice from a lethal viral challenge with SARS-CoV-2.
- Example 2 LS mutants of humanized R3DC23 Fc fusions control viral replication in hamsters
- huR3DC23-Fc The therapeutic potential of huR3DC23-Fc (SEQ ID NO: 124) in the Syrian hamster model was evaluated.
- Hamsters were challenged with an ancestral SARS-CoV-2 isolate (BetaCoV/Munich/BavPatl/2020) and, 4 hours later, treated with either 10 mg/kg or 2 mg/kg huR3DC23-Fc_LS, 10 mg/kg bebtelovivamb (biosimilar) of 10 mg/kg palivizumab (negative control treatment) by intraperitoneal injection.
- At 4 days post infection high levels of huR3DC23-Fc_LS were detected in all hamsters treated with 2 mg/kg and in 4 hamsters treated with 10 mg/kg of this construct (data not shown).
- huR3DC23-Fc_LS In sharp contrast, no or very low levels of huR3DC23-Fc_LS could be detected in the sera of two animals that were treated with 10 mg/kg huR3DC23-Fc_LS. This most likely results from unsuccessful injection, which has been observed by others (Starr et al. 2021 Nature 597:97-102). Apart from these 2 hamsters the lung virus loads, sampled on day 4 after challenge were below the detection limit in the huR3DC23-Fc treated hamsters whereas control treated animals had significantly higher lung virus loads (Fig. 2A). In accordance in the lungs of hamsters treated with either huR3DC23-Fc_LS or bebtelovimab a strong reduction in viral RNA was observed (Fig. 2B). This experiment shows that S2-binding huR3DC23-Fc can strongly restrict SARS-CoV-2 replication in vivo.
- K18-hACE2 mice that express human ACE2 at the surface of their epithelial cells were treated with 100 pg R3_DC23-Fc(YTE) (R3_DC23-Fc; SEQ ID NO: 125) or 100 pg isotype control IgG (palivizumab) via intraperitoneal injection 1 day prior to a lethal infection with SARS-CoV-2 D614G variant virus.
- Palivizumab treated, infected wild type (WT) mice that are nonpermissive for SARS- CoV-2 infection were used as control for protection.
- R3-DC23-Fc protected infected K18-hACE2 mice from bodyweight loss and death, similar to palivizumab treated WT mice. This illustrates that R3-DC23- Fc can protect mice from lethal SARS-CoV-2 infections (Fig. 3A and B).
- R3_DC23-Fc can also control viral replication in the lungs
- K18-hACE2 mice were treated with 100 pg R3_DC23-Fc or 100 pg isotype control IgG (palivizumab) via intraperitoneal injection 1 day prior to infection with SARS-CoV-2 D614G variant virus.
- IgG palivizumab
- R3_DC23-Fc treatment protected K18-hACE2 mice form bodyweight loss and lethality upon viral challenge (Fig. 4A and B).
- high levels of replicating SARS-CoV-2 virus could be isolated from lungs of all K18-hACE2 mice that were treated with palivizumab
- no replicating virus could be detected in the lungs of K18-hACE2 mice treated with R3-DC23-Fc or of WT mice (Fig. 4C).
- no viral RNA could be detected in the lungs of K18-hACE2 mice treated with R3-DC23-Fc or of WT mice (Fig. 4D).
- the molecules RNGL-Fc_LS (SEQ ID NO: 96), RAGL-Fc_LS (SEQ ID NO: 97), E2NGL-Fc_LS (SEQ ID NO:98), E2AGL-Fc_LS (SEQ ID NO:99), QRNGL-Fc_LS (SEQ ID NO: 100), QRAGL- Fc_LS (SEQ ID NO: 101), ERNGL-Fc_LS (SEQ ID NO: 102), ERAGL-Fc_LS (SEQ ID NO: 103), E10RNGL-Fc_LS (SEQ ID NO: 104), E10RAGL-Fc_LS (SEQ ID NO: 105), E81RNGL-Fc_LS (SEQ ID NO: 106), E81RAGL-Fc_LS (SEQ ID NO: 107), Q75-Fc_LS (SEQ ID NO: 108), E75- Fc_LS (SEQ ID NO: 109), E81-Fc_LS (SEQ ID NO: 110
- R3 DC23 (hR3_DC23: SEQ ID NO:2) connected to a human IgGl Fc containing an LS mutation (SEQ ID NO:83) via a (648)2 (SEQ ID NO:88) linker at the Fc N-terminus (Fig. 5B).
- Example 5 S2 targeting molecules neutralize SARS-CoV-2 pseudotyped viruses
- R3DC23-Fc_WT (SEQ ID NO: 120), R3DC23-Fc_YTE (SEQ ID NO: 121), R3DC23- Fc DHS (SEQ ID NO: 122), R3DC23-Fc_LA (SEQ ID NO: 123) and R3DC23-Fc_LS (SEQ ID NO: 124) consist of two copies of the humanized version of the VHH building block R3 DC23 (SEQ ID NO:2) connected to a human IgGl Fc (respectively, comprising SEQ ID NO: 82, 84, 85, 86 and 83) via a (648)2 linker (SEQ ID NO: 88) at their N-terminus.
- the neutralizing potency of the S2 targeting molecules of Example 4 and described above was tested in a neutralization assay using VSV particles pseudotyped with the spike of SARS-CoV-2 (D614G), Omicron variants BA.5 and XBB.1.5 (-G252V).
- Table 5 Neutralization of SARS-CoV-2 variants D614G, Omicron BA.5 and XBB.1.5 (-G252V) determined in a pseudovirus neutralization assay.
- the Table shows the calculated mean IC50 values for the tested molecules.
- N refers to the number of independent experiments. NA: not available.
- Example 6 S2 targeting molecules neutralize authentic SARS-CoV-2 D614G and Omicron BA.5 variants
- SARS-CoV-2 D614G and SARS-CoV-2 Omicron BA.5 variant viruses Dilution series of the molecules or controls were preincubated with a fixed amount of plaque-forming units (PFUs) for both virus variants for 1 hour at 37°C and subsequently used to infect Vero E6-TMPRSS2 cells.
- PFUs plaque-forming units
- the molecules with mutations in the Fc moiety for half-life extension showed improved affinity to human FcRn at pH6.0 compared to the molecule with a wildtype Fc (R3DC23-Fc_WT). As expected, low binding to human FcRn was observed at pH 7.4 for all molecules and the controls. As such, it was not possible to calculate steady state affinity at pH 7.4.
- Example 7 SPR data for human FcRn at pH6.0. The Steady state affinity and 1: 1 binding kinetic parameters for hFcRn at pH6.0 are described in the Table.
- Example 8 Non-specific binding of S2 targeting molecules to the surface of mammalian cells The non-specific binding of the molecules of Example 4 to the surface of mammalian (HEK293) cells was investigated with a flow cytometry experiment. Sotrovimab biosimilar was used as negative control antibody in the assay and XVR013 was used as positive control. As shown in Figure 8, no binding to HEK293 cells was observed for the molecules or the controls at a low concentration (10 pg/mL). At a high concentration (100 pg/mL).
- Example 4 An in silico immunogenicity assessment was performed for the molecules of Example 4 to predict the presence of putative HLA class II restricted epitopes.
- Lonza’s Epibase platform was applied to analyze the HLA binding specificities of all 10-mer peptides derived from the VHH-Fc constructs.
- the DRB1 scores of the tested molecules and XVR013 were similar to the DRB1 scores of humanized antibodies (Fig. 9).
- the DRB1 score is an approximate score representing a potential immunogenic risk for a peptide and is calculated from the population frequencies of the affected allotypes.
- the molecules CBVl-Fc_LS, CBV2-Fc_LS, CBV3-Fc_LS and CBV4-Fc_LS had the lowest DRB 1 score and therefore the lowest immunogenicity risk.
- Example 4 The apparent hydrophobicity of the molecules of Example 4 was assessed by Analytical Hydrophobic Interaction Chromatography (HIC) assay and compared with XVR011 (control for hydrophobicity; XVR011 is a bivalent molecule consisting of two VHH72 building blocks each connected at their C- terminus to an Fc domain with a LALA mutation) and XVR013 (Fig. 10).
- CBV2-Fc_LS SEQ ID NO: 117
- CBV4-Fc_LS SEQ ID NO: 119
- RNGL-Fc_LS, RAGL-Fc_LS, E2NGL-Fc_LS, E2AGL-Fc_LS, QRAGL-Fc_LS, ERAGL-Fc_LS, E10RAGL-Fc_LS, E81RNGL-Fc_LS, E81RAGL-Fc_LS and CBV4-Fc_LS displayed high hydrophobicity with retention time from 23 to 27 min, whereas Q75R21Y-Fc_LS was unusually hydrophobic showing a retention time of 35 min (Fig. 10).
- the hR3_DC23 VHH-Fc_LS variants of Example 4 were analyzed by SCX to detect their charge variants (Fig. 11).
- Example 4 The stability of the S2 targeting molecules of Example 4 was assessed under accelerated stress conditions (ageing study) in order to assess their propensity to degrade in harsh condition, mimicking the worst-case condition a drug candidate may support during manufacturing, shipment and storage operation.
- the samples characteristics are summarized and classified by color-coded categories according to the legend (Fig. 12).
- CBV3-Fc_LS also referred to herein as XVR013m
- XVR013m the animals that had received 10 mg/kg of CBV3-Fc_LS were further followed up until day 95 of the study.
- Serum samples were analysed and the final PK parameters were determined via PK population approach (Table 8). In the Tg32 SCID mouse, a half-life of 9,3 days was modelled. Allometric scaling was then applied to project a terminal half-life of 44 to 47 days in human.
- Table 8 XVR013m and XVR014 PK parameters in the Tg32 SCID mouse as determined using a two-compartment population PK model with linear elimination. Elimination clearance (CL), central and peripheral volume of distribution (VI and V2, respectively), and distribution clearance (Q).
- Example 14 Composition of SI and S2 targeting binding agents (XVR012) neutralizes VSV- GFP reporter viruses pseudotyped with SARS-CoV-2 spike proteins.
- a VHHx-Fc-VHHy construct was generated wherein a humanized form of the SI targeting VHH3. 117 capable of binding to or competing for the VHH3. 117 epitope is fused to a human IgGl Fc containing an LS mutation via a (648)2 linker (SEQ ID NO: 88) at the N-terminus of the Fc domain, and wherein a humanized form of the SI targeting VHH3.83 capable of binding to or competing for the VHH72 epitope is fused to the C-terminus via a (648)2 linker (SEQ ID NO: 88) (XVR014) (SEQ ID NO: 126).
- XVR014 was mixed with CBV3-Fc_LS (also referred to herein as XVR013m, SEQ ID NO: 118) of Example 4 in a ratio 1: 1 to generate a composition or cocktail (XVR012).
- the neutralization potency of XVR012, XVR013m and XVR014 was tested in a pseudovirus neutralization assay using VSV particles pseudotyped with the spike of either SARS-CoV-2 D614G reference strain, or SARS-CoV-2 variants EG.5.1, BA.2.86.1 and HV. l.
- a negative control antibody (palivizumab) and the positive controls XVR013, sotrovimab and bebtelovimab were included in the assay.
- the results ofthe pseudovirus neutralization assay are shown in Fig. 14 and Table 9.
- XVR012, XVR013m and XVR014 were able to neutralize all tested SARS-CoV-2 variants (D614G, EG.5.1, BA.2.86. 1 and HV. 1) with mean IC50 values ranging from 6.0 to 11.8 ng/mL for XVR012, from 3.9 ng/mL to 6.7 ng/mL for XVRO 13m, and from 14.8 to 90,6 ng/mL for XVRO 14 (Table 9) .
- the positive control XVR013 was able to neutralize the tested SARS-CoV-2 variants with IC50 values ranging from 0.6 to 1.4 ng/mL.
- Bebtelovimab neutralized the reference strain D614G (IC50 of 2.3 ng/mL), but could not neutralize the variants EG5.1, BA.2.86.1 and HV. l at the tested concentrations.
- Sotrovimab neutralized the reference strain D614G (IC50 of 24.4 ng/mL), but was less potent against the variants EG5.1 and HV. l variants and could not neutralize the variant BA.2.86.1 at the tested concentrations.
- Table 9 Neutralization of SARS-CoV-2 variants (D614G, EG.5.1, BA.2.86.1 and HV. l) by XVR012, XVR013m, XVR014 and XVR013 as determined in a pseudovirus neutralization assay.
- Vero E6 cells were transduced with VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 D614G spike protein or with the spike protein ofthe SARS-CoV-2 variants EG.5.1, BA.2.86.1 and HV.
- XVR013m CBV3-Fc_LS: SEQ ID NO: 118
- XVR014 SEQ ID NO: 126
- XVR013 SEQ ID NO: 124
- sotrovimab SEQ ID NO: 124
- sotrovimab sotrovimab or bebtelovimab.
- GFP fluorescence was measured with a fluorimeter.
- the mean IC50 values were calculated by nonlinear regression curve fitting, log(inhibitor) versus normalized response (four parameters). indicates that no neutralization was detected.
- the neutralization potency of XVR012, XVR013m and XVR014 was tested in a pseudovirus neutralization assay using VSV particles pseudotyped with the spike of either SARS-CoV-2 D614G reference strain, or SARS-CoV-2 variants Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3, BA.5, EG.5.1, FL.1.5.1, BA.2.86.1, HV. l, XBB.1.5, XBB.2.3, XBB.1.16, CH.1.1, HK.3, BA.2.74, JN. l, KP.2.3, KP.3, KP.3.1.1, KP.1.1, LB. l and XEC.
- a negative control antibody (palivizumab) or an isotype control antibody was included in the assay.
- the results of the pseudovirus neutralization assay are shown in Fig. 15 and Table 10.
- XVR012, XVR013m and XVR014 were able to neutralize all tested SARS-CoV-2 variants (D614G, Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3 and Omicrons BA.5, EG.5.1, FL.1.5.1, BA.2.86.1, HV. l, XBB.1.5, XBB.2.3, XBB.1.16, CH.1.1, HK.3, BA.2.74, JN.
- Table 10 Neutralization of SARS-CoV-2 variants (D614G, Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3, BA.5, EG.5.1, FL.1.5.1, BA.2.86.1, HV. l, XBB.1.5, XBB.2.3, XBB.1.16, CH.1.1, HK.3, BA.2.74, JN. l, KP.2.3, KP.3, KP.3.1.1, KP.1.1, LB.l and XEC) by XVR012, XVR013m and XVR014 as determined in a pseudovirus neutralization assay.
- D614G Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3, BA.5, EG.5.1, FL.1.5.1, BA.2.86.1, HV. l, XBB.1.5, XBB.2.3, XBB.1.16, CH.1.1, HK.3, BA.2.74, JN. l, K
- Vero E6 cells were transduced with VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 D614G spike protein or with the spike protein of the SARS-CoV-2 variants Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3 and Omicrons BA.5, EG.5.1, FL.1.5.1, BA.2.86.1, HV. l, XBB.1.5, XBB.2.3, XBB.1.16, CH.1.1, HK.3, BA.2.74, JN. l, KP.2.3, KP.3, KP.3.1.1, KP.1.1, LB.
- variants were generated of XVR014 (SEQ ID NO: 126) described in Example 14, wherein the C-terminal sequence TVSS (SEQ ID NO: 139) was replaced by the sequence TVSGP (SEQ ID NO: 141) or the sequence TVSAA (SEQ ID NO: 140), thereby obtaining respectively, modified XVRO 14 (TVSGP) consisting of the sequence set forth in SEQ ID NO: 127 and modified XVR01 (TVSAA) consisting of the sequence set forth in SEQ ID NO: 128.
- TVSGP modified XVRO 14
- TVSAA modified XVR01
- the seroprevalence of pre-existing ADAs in drug -naive individuals was assessed against XVRO 13, XVRO 14, the two XVRO 14 variants with C-terminal modifications, the negative control (caplacizumab) and the positive control (TAS266).
- An ELISA based method was used to determine the percentage of healthy adult human donors with pre-existing ADAs against XVR013, XVR014, the XVR014 variants with C-terminal modifications, the negative control and the positive control.
- Pre-existing ADAs were identified in 15%, 29%, 20% and 19% of the healthy donors in XVR013, XVR014, modified XVR014 (TVSGP) and modified XVR014 (TVSAA), respectively (Table 11).
- the C-terminal modifications in XVR014 reduced the percentage of pre-existing ADAs compared to XVR014.
- Example 16 XVR012, XVR013m and XVR014 control viral replication in hamsters
- the therapeutic potential of the cocktail XVR012 in a 1: 10 ratio ofXVR013m and XVR014, or the individual molecules XVR013m and XVR014 in the Syrian hamster challenge model was evaluated.
- Hamsters were challenged with an ancestral SARS-CoV-2 isolate (BetaCoV/Munich/BavPatl/2020) and, 4 hours later, treated with either 2 mg/kg, 1 mg/kg or 0,5 mg/kg XVR013m, 20 mg/kg, 10 mg/kg or 5 mg/kg XVR014, (2+20) mg/kg, (1+10) mg/kg or (0,5+5) mg/kg XVR012 (1: 10 ratio cocktail of XVR013m and XVR014), 10 mg/kg bebtelovivamb (biosimilar, positive control) or 10 mg/kg palivizumab (negative control) by intraperitoneal injection.
- Infectious viral lung loads were completely below detection levels on Day 4 in all animals experimentally confirmed to have been treated with the high, mid and low doses of both the combination, cocktail XVR012, and XVR013m.
- Animals treated with XVR014 displayed a dosedependent effect on lung viral load: respectively, 1 out of 5 and 3 out of 6 animals in the 20 mg/kg and 10 mg/kg dose groups showed a complete reduction in viral load below detection levels, with the other animals in the group showing about 10.000-fold reduction in infectious viral load compared to animals treated with a negative control antibody. In the 5 mg/kg group, none of the animals showed a full reduction in viral load, but still a 1000 to 10.000-fold reduction in infectious viral load compared to the negative control antibody was observed in all but one animal (Fig. 16A).
- Example 17 S2 targeting molecules neutralize authentic SARS-CoV-2 variants
- a microneutralization method was performed using SARS-CoV-2 D614G, Alpha, Beta, Gamma, Delta, Epsilon B. 1.427, Epsilon B. 1.429, Kappa, Zeta, Omicron BA. 1 and Omicron BA.2 variant viruses.
- Eight-point 2-fold dilution series of the molecules or controls were pre-incubated with a fixed amount of plaque-forming units (PFUs) for all virus variants for 1 hour at 37°C.
- PFUs plaque-forming units
- Example 18 Affinity maturation of VHH hR3_DC23 and CBV3
- Table 13 Degenerated libraries constructed. In each library, the aa location, position (Kabat numbering) and parental amino acid is indicated.
- the stringency of selection was increased at each round by lowering the amount of biotinylated SARS- CoV-2 spike used (200, 10 and 1 nM respectively), decreasing the binding time (down to 20 min in R3) and increasing the number of washes.
- Selected phages were used to infect bacteria and single colonies were isolated from the round 2 (R2) and round 3 (R3).
- Periplasmic extract of each single clone (6x 94 clones) was prepared and tested in a binding ELISA. Since almost all the clones showed binding, all the clones were sequenced and the amino acid diversity at each position was analyzed separately.
- VHH- monoFc VHH- monoFc, Nona Biosciences, SEQ ID NO:246
- BLI Bosset Integrity-binding protein
- the monomeric Fc format allows to use protein A for purification, but does not form bivalency to avoid the avidity effect.
- the most prominent amino acids presented at least 5 times
- Table 14 Binding kinetics of mutants selected from phage display to affinity mature hR3_DC23 and/or CBV3. The ratio of off rate of mutant versus its parent was calculated, to evaluate which mutation could be advantageous. Calculated pl and production yield are also displayed.
- Table 15 Classification of the single mutants, from advantageous to deleterious, based on the ratio of off rates compared to their parent, combining results for both hR3_DC23 and CBV3.
- Two mutations S30K and GlOOcH were clearly advantageous, decreasing the off-rate by 2-fold.
- 3 mutations were slightly advantageous (R27H, S30T and S30I). Multiple mutations were acceptable and did not impact the off rate, at positions 27, 29, 30, 96 and lOOd, while only one mutant was acceptable at positions 99, 100, 100c and 1001.
- Example 19 Binding of S2 targeting molecules to the spike of Khosta-2, a clade 3 virus from the Sarbecovirus subgenus
- the transfected cells were incubated with the S2 targeting molecule XVR013m, XVR014, a positive control (XVR013 or sotrovimab) or a negative control (palivizumab or isotype control). Bound molecules were detected by staining for human IgG Fc and flow cytometry analysis. Two independent experiments were performed. As expected, XVR013, XVR013m, XVR014 and sotrovimab highly bound to cells expressing the spike of SARS-CoV-2 D614G variant ( Figure 19). A dose-response was observed for binding to cells expressing the spike of Khosta-2 virus, showing at least partial binding (Figure 19). At 10 pg/ml, XVR013, XVR013m, XVR014 and sotrovimab were able to bind to ⁇ 80, 20, 30 and 20% ofthe cells expressing the Khosta-2 spike, respectively.
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Medicinal Chemistry (AREA)
- Animal Behavior & Ethology (AREA)
- Virology (AREA)
- Veterinary Medicine (AREA)
- Public Health (AREA)
- General Health & Medical Sciences (AREA)
- Pharmacology & Pharmacy (AREA)
- Immunology (AREA)
- Communicable Diseases (AREA)
- Mycology (AREA)
- Microbiology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Molecular Biology (AREA)
- Engineering & Computer Science (AREA)
- Epidemiology (AREA)
- Oncology (AREA)
- Chemical Kinetics & Catalysis (AREA)
- General Chemical & Material Sciences (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Organic Chemistry (AREA)
- Peptides Or Proteins (AREA)
Abstract
This invention relates to Sarbecovirus binding agents, in particular antibodies and antigen-binding fragments thereof, which are capable of potently neutralizing a Sarbecovirus, in particular capable of neutralizing any one or both of SARS-CoV-2, including SARS-CoV-2 variants, and SARS-CoV- 1, and affinity matured variants thereof. The binding agents, in particular the antibodies and antibody fragments, have one or more favourable antibody development characteristics. The invention also relates to methods using these binding agents and uses thereof.
Description
OPTIMIZED SARBECO VIRUS SPIKE S2 SUBUNIT BINDERS AND COMPOSITIONS
COMPRISING THE SAME
FIEED OF THE INVENTION
The invention is broadly in the field of binding agents, in particular antibodies. More particularly, the invention pertains to binding agents, in particular antibodies and antigen-binding fragments thereof, binding to the spike protein of a Sarbecovirus , which are capable of potently neutralizing a Sarbecovirus such as SARS-CoV-2, including SARS-CoV-2 variants, and SARS-CoV-1. The invention also relates to methods using these binding agents and uses thereof.
BACKGROUND OF THE INVENTION
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the causative agent of COVID- 19 (Zhu et al. 2020, N Engl J Med 382:727-733). SARS-CoV-2 infections can be asymptomatic or present with mild to moderately severe symptoms. However, in approximately 10% of patients, COVID-19 progresses to a more severe stage that is characterized by dyspnoea and hypoxemia, which may progress further to acute respiratory distress requiring often long-term intensive care and causing death in a proportion of patients. “Long-COVID” furthermore refers to long-term effects of COVID-19 infection, even when no SARS-CoV-2 virus can be detected anymore.
A particular type of therapeutic approach potentially relies on neutralizing antibodies, i.e. on passive antibody therapy/immunotherapy. The spike of SARS coronaviruses is a major target for neutralizing antibodies. This spike protein is a class I fusion protein and is comprised of a membrane distal SI subunit and a membrane proximal S2 subunit. The S 1 subunit comprises the receptor-binding domain (RBD) and antibodies directed against this domain can have very strong neutralizing activity (Wheatley et al. 2021. Cell Rep 37: 109822). The SI subunit, in particular the N-terminal domain and the RBD, can tolerate mutations that result in antigenic variation and immune escape. The RBD is also immunodominant (Piccoli et al. 2020. Cell 183: 1024-1042). The S2 subunit is responsible for the membrane fusion, a process during which S2 undergoes major conformational changes (Dodero- Rojas et al. 2021. eLife 10:e70362). The S2 subunit is more conserved and therefore, at least in theory, appears to be an attractive target for the development of neutralizing antibodies with broad anti-Sarbecovirus protective potential.
De Cae et al. (2023. bioRxiv. Ultrapotent SARS coronavirus-neutralizing single-domain antibodies that bind a conserved membrane proximal epitope of the spike) describe single-domain antibodies
(VHHs), also known as nanobodies, that can broadly neutralize SARS-CoV-2 with unusually high potency. These VHHs were found to interact with amino acids within the S2 subunit of the spike protein, in particular within the heptad repeat 2 (HR2) domain of the S2 subunit, more particularly within a C-terminal region of the HR2 domain proximal to the viral membrane.
For the development of antibodies, the antibodies do not only need to exhibit favourable antigenbinding and functional properties, but also be stable, safe, manufacturable, etc. For example, nonlimiting factors that affect the potential of antibodies include chemical stability (resistance to e.g., fragmentation, deamidation, oxidation and isomerisation), biophysical stability (e.g., propensity to aggregate), solution properties (e.g., solubility, viscosity). Low risk of immunogenicity of an antibody is also a consideration. Further, antibodies also need good pharmacokinetic and pharmacodynamic properties, such as slow in vivo clearance rate, long circulation half-life, etc. All these factors can collectively be referred to as development characteristics of an antibody. These are important considerations as these might impact the antibody’s cost of goods, safety profde, dosing schedule, mode of administration, etc. and thus their development potential.
There remains a need in the art for potent neutralizing antibodies that target the spike protein of a Sarbecovirus and that exhibit good antibody development characteristics.
DESCRIPTION OF THE DRAWINGS
FIG. 1 Prophylactic treatment with R3_DC23-Fc protects K18-hACE2 mice from lethal SC2 infection. K18-hACE2 mice were intraperitoneally injected with 100 pg R3_DC23-Fc or Isotype control antibody (palivizumab) or were left untreated twenty hours prior to intratracheal infection with 3* 102 PFU of SARS-CoV-2 D614G variant virus. Animals were monitored on a daily basis by measuring weight change and scoring for humane endpoints. (A) The graph shows the mean relative bodyweight change of mice treated with R3_DC23-Fc (n = 3±SEM), palivizumab (n = 2±SEM) or untreated mice (n=6±SEM). Mice treated with R3_DC23-Fc displayed significantly lower bodyweight loss as compared to mice treated with palivizumab (p<0.05) or untreated mice (p<0.01) (mixed-effect analysis with Sidak's multiple comparisons test). (B) The graph shows the Kaplan-Meier curve of animal survival portion of the indicated groups. Euthanasia was performed when mice lost more than 25% of their bodyweight as defined on day 0 or when a high score for humane endpoints was reached.
FIG. 2 LS mutants of humanized R3_DC23-Fc fusions control viral replication in hamsters. (A-B) Treatment of SARS-CoV-2 Wuhan infection by huR3DC23-Fc_LS in Syrian Golden
hamsters. Male Syrian Golden hamsters were intranasally infected with SARS-CoV-2 (Wuhan strain) on day 0 and received intraperitoneal treatment with either 10 or 2 mg/kg huR3DC23-Fc_LS, 10 mg/kg Bebtelovimab (positive control) or 10 mg/kg Palivizumab (negative control) 4 hours postinfection. Animals were euthanised at day 4 and infectious virus (A) and viral RNA (B) were measured in lung tissue on day 4. Horizontal bars indicate the median TCID50/gram (left panel) and RNA copies/gram (right panel) lung tissue. Dotted horizontal lines indicate the LLOD. * 2 animals in high dose (10 mg/kg) group were experimentally confirmed to not have been exposed to R3DC23hum-Fc_LS treatment. Data were analyzed with the one-way ANOVA and Dunn’s multiple comparison test (***P < 0.0001). (#) Data points corresponding to hamsters for which no or very low levels of huR3DC23-Fc_LS was detected in the serum were omitted from statistical analysis.
FIG. 3. Prophylactic treatment with R3_DC23-Fc protects K18-hACE2 mice against lethal SARS-CoV-2 infection. Twenty hours prior to intratracheal infection with 3* 102 PFU of SARS- CoV-2 D614G variant virus, 100 pg R3_DC23-Fc was administered to K18-hACE2 mice and 100 pg of isotype control antibody (palivizumab) was administered to a second group of K18-hACE2 mice and non-permissive wild-type (WT) mice. Animals were monitored on a daily basis by measuring weight change and scoring for humane endpoints. (A) The graph shows the Kaplan- Meier curve of animal survival portion of the indicated groups. Euthanasia was performed when mice lost more than 25% of their bodyweight as defined on day 0 or when a high score for 25 humane endpoints was reached. K18-hACE2 mice treated with R3_DC23-Fc (n=5) were significantly protected from lethality as compared to K18-hACE2 mice (n=5) treated with palivizumab (p = 0.0016, Log-rank, Mantel-cox test). (B) The graph shows the mean relative bodyweight of K18-hACE2 mice treated with R3_DC23-Fc (n = 5 ± SEM) or palivizumab (n = 5 ± SEM) and of WT mice treated with palivizumab (n= 5 ± SEM). From day 4 on mice treated with R3_DC23-Fc displayed significantly lower bodyweight loss as compared to mice treated with palivizumab (p<0.001) (2 -way ANOVA, Tukey's multiple comparisons test).
FIG. 4. Prophylactic treatment with R3_DC23-Fc reduces viral replication of SARS-CoV-2 in the lungs of infected K18-hACE2 mice. Twenty hours prior to intratracheal infection with 3* 102 PFU of SARS-CoV-2 D614G variant virus, 100 pg R3_DC23-Fc was administered to K18- hACE2 mice and 100 pg of isotype control antibody (palivizumab) was administered to a second group of K18-hACE2 mice and non-permissive wild-type (WT) mice. Animals were monitored on a daily basis by measuring weight change and scoring for humane endpoints. (A) The graph shows the Kaplan-Meier curve of animal survival portion of K18-mice treated with R3_DC23-Fc (n=5)
as compared to K18-hACE2 mice (n=4) treated with palivizumab and WT mice treated with palivizumab (n=5). (B) The graph shows the mean relative bodyweight of K18-hACE2 mice treated with R3_DC23-Fc (n = 5 ± SEM) or palivizumab (n = 4 ± SEM) and of WT mice treated with palivizumab (n= 5 ± SEM). From day 5 on K18-hACE2 mice treated with R3_DC23-Fc displayed significantly lower bodyweight loss as compared to K18-hACE2 mice treated with palivizumab (p<0.01) (2 -way ANOVA, Tukey's multiple comparisons test). (C) The graph shows the median and individual viral titer in the lungs of mice from the indicated groups sacrificed at 5 days post infection. K18-hACE2 mice treated with R3_DC23-Fc and WT mice treated with palivizumab had significantly less replicating SARS-CoV-2 virus in their lungs as compared to K18-hACE2 mice treated with palivizumab (p < 0.0005, Kruskal-Wallis test, Dunn's multiple comparisons test). (D) The graph shows the median and individual levels of viral RNA in the lungs of mice from the indicated groups sacrificed at 5 days post infection. K18-hACE2 mice treated with R3_DC23-Fc and WT mice treated with palivizumab had significantly less replicating SARS-CoV-2 virus in their lungs as compared to K18-hACE2 mice treated with palivizumab (p < 0.005, Kruskal-Wallis test, Dunn's multiple comparisons test).
FIG. 5. (A) Sequence alignment of the humanized version of the VHH R3 DC23 and modified versions thereof. Sequences of hR3_DC23 and variants are shown: VHH hR3_DC23 (SEQ ID NO:2), VHH RNGL (SEQ ID NO:3), VHH RAGL (SEQ ID NO:4), VHH E2NGL (SEQ ID NO:5), VHH E2AGL (SEQ ID NO:6), VHH QRAGL (SEQ ID NO:8), VHH ERAGL (SEQ ID NO: 10), VHH E10RAGL (SEQ ID NO: 12), VHH E81RAGL (SEQ ID NO: 14), VHH Q75 (SEQ ID NO: 15), VHH E75 (SEQ ID NO: 16), VHH E10E81 (SEQ ID NO: 18), VHH D16 (SEQ ID NO: 19), VHH T19D79 (SEQ ID NO:20), VHH TQD (SEQ ID NO:21), VHH Q75R21Y (SEQ ID NO:22), VHH CBV1 (SEQ ID NO:23), VHH CBV2 (SEQ ID NO:24), VHH CBV3 (SEQ ID NO:25), or VHH CBV4 (SEQ ID NO:26). Amino acid residue numbering was done according to Kabat numbering. CDR1, 2 and 3 annotated according to Martin are indicated and underlined. (B) Schematic representation of VHH-Fc constructs. VHH-Fc multivalent antibodies as described herein consist of two copies of a VHH building block each connected at its C-terminus to an Fc domain as described herein, in particular a human IgGl Fc without (e.g. SEQ ID NO: 82) or with mutations for half-life extension (e.g. SEQ ID NO: 83-86), via a (648)2 linker (SEQ ID NO: 88) at the Fc N-terminus.
FIG. 6. VHH hR3_DC23 amino acid sequence and illustration of the different CDR annotations as used herein. CDR annotations according to AbM, Chothia, Martin, Kabat, IMGT
and MacCallum in grey labeled boxes corresponding to the sequences of VHH hR3_DC23 (SEQ ID NO:2).
FIG. 7. S2 targeting molecules neutralize VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 spike proteins. Vero E6 cells were transduced with VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 D614G, Omicron BA.5 or XBB.1.5 (-G252V) variant spike protein that had been pre-incubated with different concentrations of the tested S2 targeting molecules or the control antibodies. Sixteen hours later, the GFP fluorescence was measured with a fluorimeter. The graphs show the mean GFP fluorescence intensity of the tested molecules for D614G, Omicron BA.5 and XBB1.5 (-G252V) normalized to the GFP fluorescence intensity value of non-infected control cells and of infected cells that were not treated, which were both included in each dilution series. The mean IC50 values were calculated by nonlinear regression curve fitting, (log(inhibitor) versus normalized response (four parameters)). Mean values ± SD are shown. N=2 or 3 for D614G, N=2 for Omicron BA.5 and N=3 for XBB.1.5(-G252V). N refers to the number of independent experiments.
FIG. 8. Flow cytometric analysis of the binding of the S2 targeting molecules to mammalian (HEK293) cells. HEK293 cells were incubated with the indicated S2 targeting molecules or controls (Sotrovimab Biosimilar and XVR013) at 10 or 100 pg/ml for 90 min. The graph shows the median fluorescence intensity (MFI) of the AF633-conjugated anti-human IgG to detect binding of the S2 targeting molecules and controls to the HEK293 cells.
FIG. 9. In silica immunogenicity assessment of S2 targeting molecules. An in silica immunogenicity assessment was conducted for the S2 targeting molecules as indicated as described in the examples. (A) Calculated DRB1 scores for the tested S2 targeting molecules and XVR013. (B) The calculated DRB1 scores of the tested S2 targeting molecules and XVR013 were compared with the DRB1 scores of 80 approved antibodies. The variants in each “bin” of the histogram are considered similar in terms of immunogenic risk.
FIG. 10. Apparent hydrophobicity of the S2 targeting molecules. Retention time of XVR013 and XVR011 (control for hydrophobicity) is indicated for comparison purposes. Note that variants CBV2-Fc_LS (SEQ ID NO: 117) and CBV4-Fc_LS (SEQ ID NO: 119) were analyzed separately: both molecules have lower pl compared to the other tested variants and the pH of the mobile phase used for the HIC assay needed to be adapted accordingly.
FIG. 11. Strong Cation Exchange (SCX) chromatograms of the S2 targeting molecules and
XVR013 and XVROl l controls.
FIG. 12. (A) General initial characteristics (top half) and biophysics upon accelerated stress (bottom half) of the indicated S2 targeting molecules, analyzed and classified in color-code categories according to the legend (B).
FIG. 13. Plasma concentrations of S2 targeting molecules and XVR013. An in vivo pharmacokinetic study was performed in human neonatal Fc receptor (hFcRn) tg32 SCID mice for the S2 targeting molecules E2NGL-Fc_LS (SEQ ID NO:98), E75-Fc_LS (SEQ ID NO: 109), CBV2- Fc_LS (SEQ ID NO: 117), CBV3-Fc_LS (SEQ ID NO: 118) and CBV4-Fc_LS (SEQ ID NO: 119), and XVR013. Plasma concentrations (Test Article) of the S2 targeting molecules and XVR013 are shown.
FIG. 14. Composition of SI and S2 targeting binding agents (XVR012) neutralizes VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 spike proteins. Vero E6 cells were transduced with VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 D614G spike protein or with the spike protein of the SARS-CoV-2 variants EG.5.1, BA.2.86.1 and HV.l, which viruses had been preincubated with different concentrations of the XVR012 composition, the constructs XVR013m (CBV3-Fc_LS: SEQ ID NO: 118) or XVR014 (SEQ ID NO: 126), or a control (XVR013, sotrovimab, bebtelovimab or palivizumab). Sixteen hours later, the GFP fluorescence was measured with a fluorimeter. The calculated mean IC50 values (ng/ml; N = 1) are shown. The mean IC50 values were calculated by nonlinear regression curve fitting, log(inhibitor) versus normalized response (four parameters).
FIG. 15. Composition of SI and S2 targeting binding agents (XVR012) neutralizes VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 spike proteins. Neutralization potency of (A) XVR013m (neutralization range: 2.3 - 7.7 ng/ml), (B) XVR014 (neutralization range: 11.6 - 461.3 ng/ml) and (C) XVR012 (neutralization range: 4.8 - 12.8 ng/ml) against SARS-CoV-2 variants. Vero E6 cells were transduced with VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 D614G spike protein or with the spike protein of the SARS-CoV-2 variants Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3, Omicron XBB.1.5, Omicron XBB.2.3, Omicron XBB.1.16, Omicron CH.1.1, Omicron EG.5.1, Omicron HK.3, Omicron HV. 1, Omicron BA.2.74, Omicron BA.2.86.1, Omicron FL. 1.5.1, Omicron JN. l, Omicron KP.2.3, Omicron KP.3 and Omicron KP.3.1.1, which viruses had been pre-incubated with different concentrations of the XVR012 composition, the constructs XVR013m (CBV3-Fc_LS: SEQ ID NO: 118) or XVR014 (SEQ ID NO: 126). Sixteen hours later, the
GFP fluorescence was measured with a fluorimeter. The relative percentage of infection compared to the virus control versus antibody concentration (ng/ml) (N = 3, unless indicated by “*” where N = 17 (A), 14 (B) or 12 (C), or by “**” wherein N = 2 or by “***” where N = 1, ) is shown. Each run included three independent repeats. Error bars represent standard deviation. The nonlinear regression curve fittings, log(inhibitor) versus normalized response (four parameters), are shown as well.
FIG. 16. In vivo efficacy in Syrian Golden Hamster SARS-CoV2 challenge model.
(A) Lung infectious viral titers in Syrian golden hamster post-infection challenge model (Wuhan strain) on day 4. (B) Nasal turbinate infectious viral titers in Syrian golden hamster post-infection challenge model (Wuhan strain) on day 4. Mean values + standard error of the mean (SEM) are reported. *Two animals in 2 mg/kg group of XVR013m and one animal in the 20 mg/kg group of XVR014 were experimentally confirmed in PK assays to not have been exposed to the test compound. Dotted lines represent the lower limit of detection (LLOD) range.
FIG. 17. Sequence alignments of hR3_DC23 variants (SEQ ID NO:151-182) with hR3_DC23 (SEQ ID NO:2). Numbering ofthe sequences is according to Kabat. CDR1 (positions 31-35), CDR2 (positions 50-65) and CDR3 (positions 95-102) according to Kabat are indicated. Mutations vis-a- vis hR3_DC23 are indicated with grey background. Residues which are believed to constitute the paratope are underlined.
FIG. 18. Sequence alignments of CBV3 (SEQ ID NO:25) and CBV3 variants (SEQ ID NO:183- 244) with hR3_DC23 (SEQ ID NO:2). Numbering ofthe sequences is according to Kabat. CDR1 (positions 31-35), CDR2(positions 50-65) and CDR3 (positions 95-102) according to Kabat are indicated. Mutations vis-a-vis hR3_DC23 are indicated with grey background. Residues which are believed to constitute the paratope are underlined.
FIG. 19. Binding of S2 targeting molecules to the spike of Khosta-2 virus, a member of clade-3 of the Sarbecovirus subgenus. Binding of XVR013, XVR013m, XVR014, positive control (sotrovimab) or negative controls (palivizumab, rituximab or an isotype control) to HEK293T cells transiently transfected with expression vectors for GFP fused to the spike of (A) Khosta-2 virus or
(B) SARS-CoV-2 D614G variant, or (C) GFP alone, was analyzed by flow cytometry. Different concentrations of antibodies were tested. Two independent experiments were performed. The percentage of bound cells for a representative ofthe 2 experiments is shown.
SUMMARY OF THE INVENTION
As demonstrated in the experimental section, which illustrates certain embodiments of the present invention, the inventors identified variants of Sa rhecovn'us -specifc Variable Domains of Heavychain Antibodies (VHHs) denoted as R3 DC23 and hR3_DC23, said variants exhibiting one or more favourable antibody development characteristics, including, but not limited to, lower pl, reduced non-specific binding, reduced immunogenicity, improved electrostatics (less positive charge patches), increased stability, reduced hydrophobicity, improved pharmacokinetics (PK), or combinations thereof. These variants also potently neutralized SARS-CoV-2, including SARS-CoV- 2 variants such as SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta, SARS-CoV-2 Epsilon B. 1.427 variant, SARS-CoV-2 Epsilon B.1.429 variant, SARS-CoV-2 Eta variant, SARS-CoV-2 Iota variant, SARS- CoV-2 Kappa variant, SARS-CoV-2 Mu variant, SARS-CoV-2 Zeta variant, SARS-CoV-2 B. 1.617.3 variant, SARS-CoV-2 Omicron BA. l variant, SARS-CoV-2 Omicron BA.2 variant, SARS-CoV-2 Omicron BA.5 variant, SARS-CoV-2 Omicron XBB.1.5 (-G252V) variant, SARS- CoV-2 Omicron XBB.1.5 variant, SARS-CoV-2 Omicron XBB.2.3 variant, SARS-CoV-2 Omicron XBB.1.16 variant, SARS-CoV-2 Omicron CH.1.1 variant, SARS-CoV-2 Omicron EG.5.1 variant, SARS-CoV-2 Omicron HK.3 variant, SARS-CoV-2 Omicron BA.2.74 variant, SARS-CoV-2 Omicron BA.2.86.1 variant, SARS-CoV-2 Omicron HV. l variant, SARS-CoV-2 Omicron FL. 1.5.1 variant, SARS-CoV-2 Omicron JN. l variant, SARS-CoV-2 Omicron KP.2.3 variant, SARS-CoV-2 Omicron KP.3 variant, SARS-CoV-2 Omicron KP.3.1.1 variant, SARS-CoV-2 Omicron KP.1.1 variant, SARS-CoV-2 Omicron LB. l variant and SARS-CoV-2 Omicron XEC variant. The hR3_DC23 variants thus have advantageously improved antibody development characteristics while maintaining the functional properties such as neutralization of SARS-CoV-2 of hR3_DC23.
Accordingly, in an aspect the invention relates to a binding agent capable of neutralizing a Sarbecovirus , characterized in that said binding agent comprises at least one ISVD as described herein, in particular an ISVD comprising a complementarity determining region 1 (CDR1) defined by SEQ ID NO:40, a CDR2 defined by SEQ ID NO:30 and a CDR3 defined by SEQ ID NO:41; or a CDR1 defined by SEQ ID NO:38, a CDR2 defined by SEQ ID NO:39 and a CDR3 defined by SEQ ID NO:41; or an ISVD comprising a CDR1, CDR2 and CDR3, each independently as present in any of SEQ ID NOs: 2 to 26, wherein the CDR1, CDR2 and CDR3 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
The invention also relates to affinity matured variants of these binding agents.
In a further aspect, the invention relates to a nucleic acid molecule comprising a polynucleotide sequence encoding the binding agent according to the invention, as well as to a vector comprising such nucleic acid molecule; and a cell comprising such nucleic acid molecule or such vector or a cell expressing the binding agent according to the invention.
The invention further relates to a pharmaceutical composition comprising the binding agent according to the invention, or the nucleic acid molecule or the vector as described hereinabove; and a pharmaceutically acceptable carrier; as well as to a kit such as a diagnostic kit comprising the binding agent according to the invention.
A further aspect is directed to the binding agent according to the invention, the nucleic acid molecule or the vector as described hereinabove, the pharmaceutical composition or the kit as described hereinabove for use in medicine such as use in the prevention or treatment of a Sarbecovirus infection in a subject or for use in the diagnosis of a Sarbecovirus infection in a subject.
The invention further relates to an in vitro or ex vivo method for detecting a Sarbecovirus in a sample, said method comprising:
- contacting the sample with a binding agent according to the invention, and
- determining binding of the binding agent with a Sarbecovirus or a part thereof.
Those skilled in the art will recognize the many other effects and advantages of the present methods, uses or products, and the numerous possibilities for end uses of the present invention from the detailed description and examples provided below.
DETAILED DESCRIPTION OF THE INVENTION
As used herein, the singular forms “a”, “an”, and “the” include both singular and plural referents unless the context clearly dictates otherwise.
The terms “comprising”, “comprises” and “comprised of’ as used herein are synonymous with “including”, “includes” or “containing”, “contains”, and are inclusive or open-ended and do not exclude additional, non-recited members, elements or method steps. The terms also encompass “consisting of’ and “consisting essentially of’, which enjoy well-established meanings in patent terminology.
The recitation of numerical ranges by endpoints includes all numbers and fractions subsumed within the respective ranges, as well as the recited endpoints.
The terms “about” or “approximately” as used herein when referring to a measurable value such as a parameter, an amount, a temporal duration, and the like, are meant to encompass variations of and
from the specified value, such as variations of± 10% or less, preferably ± 5% or less, more preferably ± 1% or less, and still more preferably ± 0.1% or less of and from the specified value, insofar such variations are appropriate to perform in the disclosed invention. It is to be understood that the value to which the modifier “about” refers is itself also specifically, and preferably, disclosed.
Whereas the terms “one or more” or “at least one”, such as one or more members or at least one member of a group of members, is clear per se, by means of further exemplification, the term encompasses inter alia a reference to any one of said members, or to any two or more of said members, such as, e.g., any >3, >4, >5, >6 or >7 etc. of said members, and up to all said members. In another example, “one or more” or “at least one” may refer to 1, 2, 3, 4, 5, 6, 7 or more.
The discussion of the background to the invention herein is included to explain the context of the invention. This is not to be taken as an admission that any of the material referred to was published, known, or part of the common general knowledge in any country as of the priority date of any of the claims.
Throughout this disclosure, various publications, patents and published patent specifications are referenced by an identifying citation. All documents cited in the present specification are hereby incorporated by reference in their entirety. In particular, the teachings or sections of such documents herein specifically referred to are incorporated by reference.
Unless otherwise defined, all terms used in disclosing the invention, including technical and scientific terms, have the meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. By means of further guidance, term definitions are included to better appreciate the teaching of the invention. When specific terms are defined in connection with a particular aspect of the invention or a particular embodiment of the invention, such connotation is meant to apply throughout this specification, i.e., also in the context of other aspects or embodiments of the invention, unless otherwise defined.
In the following passages, different aspects or embodiments of the invention are defined in more detail. Each aspect or embodiment so defined may be combined with any other aspect(s) or embodiment(s) unless clearly indicated to the contrary. In particular, any feature indicated as being preferred or advantageous may be combined with any other feature or features indicated as being preferred or advantageous.
Reference throughout this specification to “one embodiment”, “an embodiment” means that a particular feature, structure or characteristic described in connection with the embodiment is included in at least one embodiment of the present invention. Thus, appearances of the phrases “in one
embodiment” or “in an embodiment” in various places throughout this specification are not necessarily all referring to the same embodiment, but may. Furthermore, the particular features, structures or characteristics may be combined in any suitable manner, as would be apparent to a person skilled in the art from this disclosure, in one or more embodiments. Furthermore, while some embodiments described herein include some but not other features included in other embodiments, combinations of features of different embodiments are meant to be within the scope of the invention, and form different embodiments, as would be understood by those in the art. For example, in the appended claims, any of the claimed embodiments can be used in any combination.
As corroborated by the experimental section, which illustrates certain representative embodiments of the invention, the inventors identified novel VHHs that potently neutralize SARS-CoV-2, including SARS-CoV-2 variants such as SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta variant, SARS-CoV- 2 Epsilon B. 1.427 variant, SARS-CoV-2 Epsilon B. 1.429 variant, SARS-CoV-2 Eta variant, SARS- CoV-2 Iota variant, SARS-CoV-2 Kappa variant, SARS-CoV-2 Mu variant, SARS-CoV-2 Zeta variant, SARS-CoV-2 B. 1.617.3 variant, SARS-CoV-2 Omicron BA. l variant, SARS-CoV-2 Omicron BA.2 variant, SARS-CoV-2 Omicron BA.5 variant, SARS-CoV-2 Omicron XBB.1.5 (- G252V) variant, SARS-CoV-2 Omicron XBB.1.5 variant, SARS-CoV-2 Omicron XBB.2.3 variant, SARS-CoV-2 Omicron XBB.1.16 variant, SARS-CoV-2 Omicron CH. 1.1 variant, SARS-CoV-2 Omicron EG.5.1 variant, SARS-CoV-2 Omicron HK.3 variant, SARS-CoV-2 Omicron BA.2.74 variant, SARS-CoV-2 Omicron BA.2.86.1 variant, SARS-CoV-2 Omicron HV. l variant, SARS- CoV-2 Omicron FL. 1.5.1 variant, SARS-CoV-2 Omicron JN. l variant, SARS-CoV-2 Omicron KP.2.3 variant, SARS-CoV-2 Omicron KP.3 variant, SARS-CoV-2 Omicron KP.3.1.1 variant, SARS-CoV-2 Omicron KP.1.1 variant, SARS-CoV-2 Omicron LB. l variant and SARS-CoV-2 Omicron XEC variant. The novel VHHs were found to also exhibit one or more favourable antibody development characteristics.
Without wishing to be bound by any hypothesis, the inventors’ experiments indicated that the paratope of the hR3_DC23 (SEQ ID NO:2) and CBV3 (SEQ ID NO:25) ISVD is formed or constituted by the amino residues at positions 96, 97, 98, 99, 100a, 100b and 1001 according to Kabat numbering of hR3_DC23 (SEQ ID NO:2) or CBV3 (SEQ ID NO:25) and/or at positions 27, 29, 30, 31, 52, 100, 100c and lOOd according to Kabat numbering of hR3_DC23 (SEQ ID NO:2) or CBV3 (SEQ ID NO:25). Hence, further disclosed herein are binding agents capable of neutralizing a
Sarbecovirus , characterized in that said binding agent comprises at least one anti-SARS-CoV-2 spike S2 subunit ISVD having the same or a similar paratope as hR3_DC23 (SEQ ID NO:2).
In particular embodiments, the anti-SARS-CoV-2 spike S2 subunit ISVD has a paratope comprising or consisting of amino acid residues at positions 96, 97, 98, 99, 100a, 100b and 1001 according to Kabat numbering and/or at positions 27, 29, 30, 31, 52, 100, 100c and lOOd according to Kabat numbering, wherein each independently: the amino acid residue at position 96 is selected from the group consisting of: Y, F, R, W, H and Q; or, with increasing preference, is selected from the group consisting of: Y, F, R and W; or is Y; the amino acid residue at position 97 is selected from the group consisting of: V, I, A, L and F; or, with increasing preference, is selected from the group consisting of: V, I, A and L; or is V; the amino acid residue at position 98 is selected from the group consisting of: S, A and T; or, preferably, is S; the amino acid residue at position 99 is selected from the group consisting of: K and R, or, preferably, is K; the amino acid residue at position 100a is N; the amino acid residue at position 100b is selected from the group consisting of: Y, F and H; or, with increasing preference, is selected from the group consisting of: Y and F; or is Y; the amino acid residue at position 1001 is selected from the group consisting of: R, L, H, V, and P; or, with increasing preference, is selected from the group consisting of: R, L, H and V ; is selected from the group consisting of: R and L; or is R; the amino acid residue at position 27 is selected from the group consisting of: R, H, L, P, S and T; or, with increasing preference, is selected from the group consisting of: R and H; or is H; the amino acid residue at position 29 is selected from the group consisting of: F, Y, H, M, R, W, I,
L, S, V, A and G; or, with increasing preference, is selected from the group consisting of: F, Y, H,
M, R and W; or is F ; the amino acid residue at position 30 is selected from the group consisting of: S, K, T, I, R, G, H, L, P, F, N and W; or, with increasing preference, is selected from the group consisting of: S, K, T and I; is selected from the group consisting of: K, T and I; or is K;
the amino acid residue at position 31 is selected from the group consisting of: T and S; or, preferably is T; the amino acid residue at position 52 is R; the amino acid residue at position 100 is selected from the group consisting of: A and R; or, preferably, is A; the amino acid residue at position 100c is selected from the group consisting of: G, H and A; or, with increasing preference, selected from the group consisting of: G and H; or is H; and the amino acid residue at position lOOd is selected from the group consisting of: S, R, H, P, Q and V; or, preferably, is S.
In particular embodiments, the anti-SARS-CoV-2 spike S2 subunit ISVD has a paratope as present in any one of the ISVDs of SEQ ID NO: 151 to 244, 2 or 25, optionally wherein the paratope comprises or consists of the amino acid residue at positions 96, 97, 98, 99, 100a, 100b and 1001 according to Kabat numbering and/or at positions 27, 29, 30, 31, 52, 100, 100c and lOOd according to Kabat numbering of said ISVD.
In an aspect the invention relates to a binding agent capable of neutralizing a Sarbecovirus , characterized in that said binding agent comprises at least one ISVD, wherein said ISVD comprises a CDR1 defined by SEQ ID NO:40, a CDR2 defined by SEQ ID NO:30 and a CDR3 defined by SEQ ID NO:41; or a CDR1 defined by SEQ ID NO:38, a CDR2 defined by SEQ ID NO:39 and a CDR3 defined by SEQ ID NO:41; or wherein said ISVD comprises a CDR1, CDR2 and CDR3, each independently as present in any of SEQ ID NOs:2 to 26, wherein the CDR1, CDR2 and CDR3 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
A “binding agent” generally relates to a molecule that is capable of binding to at least one other molecule, wherein said binding is preferably a specific binding, such as on a defined binding site, pocket or epitope. A binding agent may be of any nature or type and is not dependent on its origin. A binding agent may be chemically synthesized, naturally occurring, recombinantly produced (and optionally purified), as well as designed and synthetically produced (and optionally purified). Said binding agent may hence be, e.g., a small molecule, a chemical, a peptide, a polypeptide, an antibody, or any derivative of any thereof, such as a peptidomimetic, an antibody mimetic, an active fragment, a chemical derivative, among others. A functional fragment of a binding agent or a functional part of a binding agent refers to a fragment or part of that binding agent that is functionally equivalent to that binding agent. In particular, such functional fragment or part of a binding agent as described
herein ideally retains one or more of the functional features (1) to (88) of that binding agent as outlined extensively elsewhere herein.
The binding agents described herein are structurally defined as polypeptidic binding agents (i.e. binding agents comprising a peptidic, polypeptidic or proteic moiety, or binding agents comprising a peptide, polypeptide, protein or protein domain) or polypeptide binding agents (i.e. binding agents being peptides, polypeptides or proteins).
The terms “protein”, “polypeptide”, and “peptide” are interchangeably used herein to refer to a polymer of amino acid residues and to variants and synthetic analogues of the same; the sequential linear arrangement of the amino acids together resulting in/forming the “amino acid sequence” or “protein sequence”. A “peptide” may also be referred to as a partial amino acid sequence derived from its original protein, for instance after enzymatic (e.g. tryptic) digestion. These terms apply to naturally-occurring amino acid polymers as well as to amino acid polymers in which one or more amino acid residues is a synthetic non-naturally occurring amino acid, such as a chemical analogue of a corresponding naturally occurring amino acid. Also included are proteins comprising one or more posttranslational modifications such as covalent addition of functional groups or proteins (such as glycosylation, phosphorylation, acetylation, ubiquitination, methylation, lipidation and nitrosylation) or such as proteolytic processing. Based on the amino acid sequence and the modifications, the atomic or molecular mass or weight of a polypeptide is expressed in (kilo)dalton (kDa). A further modification of proteins includes addition of a tag, such as a His-tag or sortag. By sortagging (sortase-mediated transpeptidation; Popp et al. 2007, Nat Chem Biol 3:707-708) for instance, a multi-arm PEG nanobody neutralizing SARS-CoV2 was constructed (Moliner-Morro et al. 2020, Biomolecules 10: 1661).
A “protein domain” is a distinct functional and/or structural unit in or part of a protein. Usually, a protein domain is responsible for a particular function or interaction, contributing to the overall (biological) role of a protein. Domains may exist in a variety of biological contexts, where similar domains can be found in different proteins with similar or different functions. Protein domains can have a rigid 3D- structure if confined by e.g. a number of intramolecular cysteines (e.g. cysteine- knot proteins) or can, depending on e.g. presence or absence of a bound ligand or e.g. presence or absence of a posttranslational modification, assume different 3D-conformations, or can have a less defined, more fluid 3D-structure.
Amino acids are presented herein by their 3- or 1 -lettercode nomenclature as defined and provided also in the IUPAC-IUB Joint Commission on Biochemical Nomenclature (Nomenclature and
Symbolism for Amino Acids and Peptides. Eur. J. Biochem. 138: 9-37 (1984)); as follows: Alanine (A or Ala), Cysteine (C or Cys), Aspartic acid (D or Asp), Glutamic acid (E or Glu), Phenylalanine (F or Phe), Glycine (G or Gly), Histidine (H or His), Isoleucine (I or He), Lysine (K or Lys), Leucine (L or Leu), Methionine (M or Met), Asparagine (N or Asn), Proline (P or Pro), Glutamine (Q or Gin), Arginine (R or Arg), Serine (S or Ser), Threonine (T or Thr), Valine (V or Vai), Tryptophan (W or Trp), and Tyrosine (Y or Tyr).
In particular, the polypeptidic or polypeptide binding agents described herein are (optionally purified) antibodies or antibody fragments.
The term “antibody” refers to an immunoglobulin (Ig) molecule or a molecule comprising an immunoglobulin (Ig) domain, which specifically binds with an antigen, as well as multimers thereof. “Antibodies” can be intact immunoglobulinsor immunoreactive portions of intact immunoglobulins. The term encompasses naturally, recombinantly, semi-synthetically or synthetically produced antibodies. Hence, for example, an antibody can be present in or isolated from nature, e.g., produced or expressed natively or endogenously by a cell or tissue and optionally isolated therefrom; or an antibody can be recombinant, i.e., produced by recombinant DNA technology, and/or can be, partly or entirely, chemically or biochemically synthesised.
By “isolated” or “purified” is meant material that is substantially or essentially free from components that normally accompany it in its native state. For example, an “isolated polypeptide” or “purified polypeptide” refers to a polypeptide which has been isolated or purified by any suitable means from a mixture of molecules comprising the to be isolated or to be purified polypeptide of interest. An isolated or purified polypeptide of interest can for instance be an immunoglobulin, antibody or nanobody, and the mixture can be a mixture or molecules as present in a cell producing the immunoglobulin, antibody or nanobody, and/or the culture medium into which the immunoglobulin, antibody or nanobody is secreted into (likely together with other molecules secreted by the cell).
The terms “antibody fragment”, “antigen-binding fragment”, “functional antibody fragment” and "active antibody fragment" refer to a portion of any antibody that by itself has high affinity for an antigenic determinant, or epitope, and contains one or more complementarity determining regions (CDRs) accounting for such specificity. The terms “antibody fragment” and “antigen-binding fragment” and “active antibody fragment” and “functional antibody fragment” as used herein refer to a protein or peptide comprising an immunoglobulin domain or an antigen-binding domain capable of specifically binding to a Sarbecovirus spike protein such as SARS-CoV-2 spike protein, in particular to the S2 subunit of the Sarbecovirus spike protein, more particularly to the HR2 domain
of (the S2 subunit of) the Sarbecovirus spike protein. Non-limiting examples include immunoglobulin domains, Fab, F(ab)'2, scFv, heavy-light chain dimers, immunoglobulin single variable domains, Nanobodies (or VHH antibodies), domain antibodies, and single chain structures, such as a complete light chain or complete heavy chain.
The term “immunoglobulin (Ig) domain”, or more specifically “immunoglobulin variable domain” (abbreviated as “IVD”, also referred to herein as “variable domain”), means an immunoglobulin domain essentially consisting of four “framework regions” which are referred to in the art and herein below as “framework region 1” or “FR1”; as “framework region 2” or “FR2”; as “framework region 3” or “FR3”; and as “framework region 4” or “FR4”, respectively; which framework regions are interrupted by three “complementarity determining regions” or “CDRs”, which are referred to in the art and herein below as “complementarity determining region 1” or “CDR1”; as “complementarity determining region 2” or “CDR2”; and as “complementarity determining region 3” or “CDR3”, respectively. Thus, the general structure or sequence of an immunoglobulin variable domain can be indicated as follows: FR1 - CDR1 - FR2 - CDR2 - FR3 - CDR3 - FR4. It is the immunoglobulin variable domain(s) (IVDs), and in particular the CDRs therein, even more particularly CDR3 therein, that confer specificity to an antibody for the antigen by carrying the antigen- or epitope-binding site. Typically, in conventional immunoglobulins, a heavy chain variable domain (VH) and a light chain variable domain (VL) interact to form an antigen-binding site. In this case, the complementarity determining regions (CDRs) of both VH and VL contribute (although not necessarily evenly) to the antigen-binding site, i.e. atotal of 6 CDRs will be involved in antigen-binding site formation. In view of the above definition, the antigen-binding domain of a conventional 4-chain antibody (such as an IgG, IgM, IgA, IgD or IgE molecule; known in the art) or of a Fab fragment, a F(ab')2 fragment, an Fv fragment such as a disulphide linked Fv or a scFv fragment, or a diabody (all known in the art) derived from such conventional 4-chain antibody, will bind to the respective epitope of an antigen by a pair of (associated) immunoglobulin domains such as light and heavy chain variable domains, i.e., by a VH-VL pair of immunoglobulin domains, which jointly bind to an epitope of the respective antigen.
An “immunoglobulin single variable domain” (abbreviated as "ISVD"), equivalent to the term “single variable domain”, defines molecules wherein the antigen-binding site is present on, and formed by, a single immunoglobulin domain. This sets immunoglobulin single variable domains apart from “conventional” immunoglobulins or their fragments, wherein two immunoglobulin domains, in particular two variable domains, interact to form an antigen-binding site. An
“immunoglobulin single variable domain” (or “ISVD”) as used herein, refers to a protein or peptide with an amino acid sequence comprising 4 framework regions (FR) and 3 complementary determining regions (CDR) according to the format of FR1-CDR1-FR2-CDR2-FR3-CDR3-FR4. The antigen-binding site of an immunoglobulin single variable domain is formed by a single VH/VHH or VL domain. Hence, the antigen-binding site of an immunoglobulin single variable domain is formed by no more than three CDRs. As such, the single variable domain may be a light chain variable domain sequence (e.g., a VL-sequence) or a suitable fragment thereof; or a heavy chain variable domain sequence (e.g., a VH-sequence or VHH sequence) or a suitable fragment thereof; as long as it is capable of forming a single antigen-binding unit (i.e., a functional antigenbinding unit that essentially consists of the single variable domain, such that the single antigenbinding domain does not need to interact with another variable domain to form a functional antigenbinding unit). In certain embodiments, the immunoglobulin single variable domains are heavy chain variable domain sequences (e.g., a VH-sequence or a VHH-sequence); more specifically, the immunoglobulin single variable domains can be heavy chain variable domain sequences that are derived from a conventional four-chain antibody or heavy chain variable domain sequences that are derived from a heavy chain antibody. For example, the immunoglobulin single variable domain may be a (single) domain antibody (or an amino acid sequence that is suitable for use as a (single) domain antibody), a variable domain of a heavy (VH) or light (VL) chain of a conventional antibody (also referred to as a “dAb”) (or an amino acid sequence that is suitable for use as a dAb) or a Nanobody (as defined herein, and including but not limited to a VHH); or any suitable fragment of any one thereof.
In embodiments, the immunoglobulin single variable domain may be a Nanobody (as defined herein) or a suitable fragment thereof. Note: Nanobody®, Nanobodies® and Nanoclone® are registered trademarks of Ablynx N.V. (a Sanofi Company). For a general description of Nanobodies, reference is made to the further description below, as well as to the prior art cited herein, such as e.g. described in W02008/020079. “VHH domains”, also known as VHHs, VHH domains, VHH antibody fragments, and VHH antibodies, have originally been described as the antigen-binding immunoglobulin (Ig) (variable) domain of “heavy chain antibodies” (i.e., of “antibodies devoid of light chains”; Hamers-Casterman et al. 1993, Nature 363: 446-448). The term “VHH domain” has been chosen to distinguish these variable domains from the heavy chain variable domains that are present in conventional 4-chain antibodies (which are referred to herein as “VH domains”) and from the light chain variable domains that are present in conventional 4-chain antibodies (which are referred to herein as “VL domains”). For a further description of VHHs and Nanobody, reference is
made to the review article by Muyldermans (2001. Rev Mol Biotechnol 74: 277-302), as well as to the following patent applications, which are mentioned as general background art: WO 94/04678, WO 95/04079, WO 96/34103, WO 94/25591, WO 99/37681, WO 00/40968, WO 00/43507, WO 00/65057, WO 01/40310, WO 01/44301, EP 1134231, WO 02/48193, WO 97/49805, WO 01/21817, WO 03/035694, WO 03/054016, WO 03/055527, WO 03/050531, WO 01/90190, WO 03/025020 (= EP 1433793), WO 04/041867, WO 04/041862, WO 04/041865, WO 04/041863, WO 04/062551, WO 05/044858, WO 06/40153, WO 06/079372, WO 06/122786, WO 06/122787 and WO 06/122825. As described in these references, Nanobody (in particular VHH sequences and partially humanized Nanobody) can in particular be characterized by the presence of one or more “hallmark residues” in one or more of the framework sequences.
The binding agents according to the current invention can be structurally defined as polypeptidic or polypeptide binding agents, in particular antibodies and antibody-fragments, comprising at least one immunoglobulin single variable domain (ISVD) as defined herein. In certain embodiments, the binding agents, in particular antibodies and antibody-fragments, more particularly ISVDs, as described herein comprise at least CDR3 as comprised in an immunoglobulin single variable domain (ISVD) defined by any of SEQ ID NO:2-26, wherein the CDR3 is annotated according to Kabat or Martin. In other embodiments, the binding agents, in particular the antibodies and antibody fragments, more particularly ISVDs, as described herein comprise at least two of CDR1, CDR2 and CDR3 (e g. CDR1 and CDR3, CDR2 and CDR3, CDR1 and CDR2), or all three of CDR1, CDR2 and CDR3, as comprised in an immunoglobulin single variable domain (ISVDs) defined by any of SEQ ID NO:2-26, wherein the CDR1, CDR2 and CDR3 are annotated according to Kabat or Martin. Such CDRs may be comprised in any of VHH hR3_DC23 (defined by/set forth in SEQ ID NO:2), VHH RNGL (defined by/set forth in SEQ ID NO:3), VHH RAGL (defined by/set forth in SEQ ID NO:4), VHH E2NGL (defined by/set forth in SEQ ID NO:5), VHH E2AGL (defined by/set forth in SEQ ID NO: 6), VHH QRNGL (defined by/set forth in SEQ ID NO: 7), VHH QRAGL (defined by/set forth in SEQ ID NO: 8), VHH ERNGL (defined by/set forth in SEQ ID NOV), VHH ERAGL (defined by/set forth in SEQ ID NO: 10), VHH E10RNGL (defined by/set forth in SEQ ID NO: 11), VHH E10RAGL (defined by/set forth in SEQ ID NO: 12), VHH E81RNGL (defined by/set forth in SEQ ID NO: 13), VHH E81RAGL (defined by/set forth in SEQ ID NO: 14), VHH Q75 (defined by/set forth in SEQ ID NO: 15), VHH E75 (defined by/set forth in SEQ ID NO: 16), VHH E81 (defined by/set forth in SEQ ID NO: 17), VHH E10E81 (defined by/set forth in SEQ ID NO: 18), VHH D16 (defined by/set forth in SEQ ID NO: 19), VHH T19D79 (defined by/set forth in SEQ ID NO:20), VHH TQD (defined by/set forth in SEQ ID NO:21), VHH Q75R21Y (defined by/set forth in SEQ
ID NO:22), VHH CBV1 (defined by/set forth in SEQ ID NO:23), VHH CBV2 (defined by/set forth in SEQ ID NO:24), VHH CBV3 (defined by/set forth in SEQ ID NO:25), or VHH CBV4 (defined by/set forth in SEQ ID NO:26), as depicted hereafter:
VHH hR3_DC23
DVQLVESGGGLVQPGGSLRLSCAVSGRI FSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLQMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSRRYDYWGQGTLVTVSS
( SEQ ID NO : 2 )
VHH RNGL
DVQLVESGGGLVQPGGSLRLSCAVSGRI FSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLQMNSLRPEDTAVYYCAAAYVSKANYGSLWYRNSGLYDYWGQGTLVTVSS
( SEQ ID NO : 3 )
VHH RAGE
DVQLVESGGGLVQPGGSLRLSCAVSGRI FSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLQMNSLRPEDTAVYYCAAAYVSKANYGSLWYRASGLYDYWGQGTLVTVSS ( SEQ ID NO : 4 )
VHH E2NGL
DVQLVESGGGLVQPGGSLRLSCAVSGRI FSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLQMNSLRPEDTAVYYCAAAYVSKANYGSLWYENSGLYDYWGQGTLVTVSS
( SEQ ID NO : 5 )
VHH E2AGL
DVQLVESGGGLVQPGGSLRLSCAVSGRI FSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLQMNSLRPEDTAVYYCAAAYVSKANYGSLWYEASGLYDYWGQGTLVTVSS ( SEQ ID NO : 6 )
VHH QRNGL
DVQLVESGGGLVQPGGSLRLSCAVSGRI FSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAQNTVYLQMNSLRPEDTAVYYCAAAYVSKANYGSLWYRNSGLYDYWGQGTLVTVSS ( SEQ ID NO : 7 )
VHH QRAGL
DVQLVESGGGLVQPGGSLRLSCAVSGRI FSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG RFTISRDNAQNTVYLQMNSLRPEDTAVYYCAAAYVSKANYGSLWYRASGLYDYWGQGTLVTVSS ( SEQ ID NO : 8 )
VHH ERNGL
DVQLVESGGGLVQPGGSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAENTVYLQMNSLRPEDTAVYYCAAAYVSKANYGSLWYRNSGLYDYWGQGTLVTVSS (SEQ ID NO: 9)
VHH ERAGL
DVQLVESGGGLVQPGGSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAENTVYLQMNSLRPEDTAVYYCAAAYVSKANYGSLWYRASGLYDYWGQGTLVTVSS (SEQ ID NO:10)
VHHE10RNGL
DVQLVESGGELVQPGGSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLQMNSLRPEDTAVYYCAAAYVSKANYGSLWYRNSGLYDYWGQGTLVTVSS (SEQ ID NO:11)
VHHE10RAGL
DVQLVESGGELVQPGGSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLQMNSLRPEDTAVYYCAAAYVSKANYGSLWYRASGLYDYWGQGTLVTVSS (SEQ ID N0:12)
VHHE81RNGL
DVQLVESGGGLVQPGGSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLEMNSLRPEDTAVYYCAAAYVSKANYGSLWYRNSGLYDYWGQGTLVTVSS (SEQ ID N0:13)
VHHE81RAGL
DVQLVESGGGLVQPGGSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLEMNSLRPEDTAVYYCAAAYVSKANYGSLWYRASGLYDYWGQGTLVTVSS (SEQ ID N0:14)
VHHQ75
DVQLVESGGGLVQPGGSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAQNTVYLQMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSRRYDYWGQGTLVTVSS (SEQ ID N0:15)
VHHE75
DVQLVESGGGLVQPGGSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAENTVYLQMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSRRYDYWGQGTLVTVSS (SEQ ID N0:16)
VHHE81
DVQLVESGGGLVQPGGSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLEMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSRRYDYWGQGTLVTVSS (SEQ ID NO:17)
VHHE10E81
DVQLVESGGELVQPGGSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLEMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSRRYDYWGQGTLVTVSS (SEQ ID NO:18)
VHHD16
DVQLVESGGGLVQPGDSLRLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLQMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSRRYDYWGQGTLVTVSS (SEQ ID NO:19)
VHHT19D79
DVQLVESGGGLVQPGGSLTLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVDLQMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSRRYDYWGQGTLVTVSS (SEQ ID NQ:20)
VHHTQD
DVQLVESGGGLVQPGGSLTLSCAVSGRIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAQNTVDLQMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSRRYDYWGQGTLVTVSS (SEQ ID NO:21)
VHHQ75R21Y
DVQLVESGGGLVQPGGSLRLSCAVSGYIFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAQNTVYLQMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSRRYDYWGQGTLVTVSS (SEQ ID NO:22)
VHH CBV1
DVQLVESGGGLVQPGGSLRLSCAVSGRPFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVYLEMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSERYDYWGQGTLVTVSS (SEQ ID NO:23)
VHH CBV2
DVQLVESGGGLVQPGGSLTLSCAVSGRPFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG
RFTISRDNAKNTVDLEMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSERYDYWGQGTLVTVSS (SEQ ID NO:24)
VHH CBV3
DVQLVESGGELVQPGGSLRLSCAVSGRPFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG RFTISRDNAENTVYLEMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSERYDYWGQGTLVTVSS ( SEQ ID NO : 25 )
VHH CBV4
DVQLVESGGGLVQPGGSLTLSCAVSGRPFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG RFTISRDNAKNTVDLEMNSLRPEDTAVYYCAAAYVSKANYGSLWYRASELYDYWGQGTLVTVSS ( SEQ ID NO : 26 )
For numbering of the amino acid residues of any IVD or ISVD different numbering schemes can be applied. For example, numbering can be performed according to the AHo numbering scheme for all heavy (VH) and light chain variable domains (VL) given by Honegger & Pltickthun (2001. J Mol Biol 309:657-70), as applied to VHH domains from camelids. Alternative methods for numbering the amino acid residues of VH domains, which can also be applied in an analogous manner to VHH domains, are known in the art. For example, the delineation of the FR and CDR sequences can be done by using the Kabat numbering system as applied to VHH domains from camelids by Riechmann & Muyldermans (1999. J Immunol Methods 231:25-38). It should be noted that - as is well known in the art for VH domains and for VHH domains - the total number of amino acid residues in each of the CDRs may vary and may not correspond to the total number of amino acid residues indicated by the Kabat numbering (that is, one or more positions according to the Kabat numbering may not be occupied in the actual sequence, or the actual sequence may contain more amino acid residues than the number allowed for by the Kabat numbering). This means that, generally, the numbering according to Kabat may or may not correspond to the actual numbering of the amino acid residues in the actual sequence. The total number of amino acid residues in a VH domain and a VHH domain will usually be in the range of from 110 to 120, often between 112 and 115. It should however be noted that smaller and longer sequences may also be suitable for the purposes described herein.
The determination of the CDR regions in an antibody/immunoglobulin sequence generally depends on the algorithm/methodology applied. For example, determination of CDR regions may be done according to the designation based on contact analysis and binding site topography as described in MacCallum et al. (J. Mol. Biol. (1996) 262, 732-745), AbM (AbM is Oxford Molecular Ltd.'s antibody modelling package as described on http://www.bioinf.org.uk/abs/index.html), Chothia (Chothia and Lesk, 1987; Mol Biol. 196:901-17), Martin (Abhinandan, and Martin. Molecular Immunology 45 (2008) 3832-3839; as shown in http://bioinf.org.uk/abs/info.html), Kabat (Kabat et
al., 1991; 5th edition, NIH publication 91-3242), or IMGT (LeFranc, 2014; Frontiers in Immunology. 5 (22): 1-22). Said annotations further include delineation of CDRs and framework regions (FRs) in immunoglobulin-domain-containing proteins, and are known methods and systems to a skilled artisan who thus can apply these annotations onto any antibody/immunoglobulin protein sequences without undue burden. As an example, Fig. 6 illustrates the different annotation -schemes or - methods as applied to the amino acid sequence of VHH hR3_DC23 (SEQ ID NO:2).
Applying different methods to the same antibody/immunoglobulin sequence may give rise to different CDR amino acid sequences wherein the differences may reside in CDR sequence length and/or -delineation within the antibody/immunoglobulin/IVD sequence (as illustrated in Fig. 6 for VHH hR3_DC23). The CDRs of the ISVD binding agents, in particular antibodies and antibody fragments as described herein can therefore be described as the CDR sequences as present in the ISVDs characterized herein. Alternatively, these CDRs can be described as the CDR sequences present in the ISVDs (as described herein) as determined or delineated according to a well-known methodology such as according to any one of the Kabat-, Martin-, Chothia-, aHo, MacCallum et al. 1996, AbM-, or IMGT, numbering scheme or method, such as preferably the Martin numbering scheme or method.
As outlined above, many systems or methods (Kabat, MacCallum, IMGT, AbM, Chothia, Martin) exist for numbering amino acids in immunoglobulin protein sequences, including for delineation of CDRs and framework regions (FRs) in these protein sequences. These systems or methods are known to a skilled artisan who thus can apply these systems or methods on any immunoglobulin protein sequences without undue burden (as illustrated in Fig. 6 for VHH hR3_DC23).
In certain embodiments, a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a CDR1 defined by/set forth in any one of SEQ ID NOs: 40 or 38. In certain embodiments, a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a CDR2 defined by/set forth in any one of SEQ ID NOs: 30 or 39. In certain embodiments, a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a CDR3 defined by/set forth in SEQ ID NO: 41.
In certain embodiments, the binding agent, in particular the antibody or antibody fragment, more particularly the ISVD, as described herein may comprise:
- a CDR1 defined by/set forth in SEQ ID NO:40; a CDR2 defined by/set forth in SEQ ID NO:30; and a CDR3 defined by/set forth in SEQ ID NO:41; or
- a CDR1 defined by/set forth in SEQ ID NO:38; a CDR2 defined by/set forth in SEQ ID NO:39; and a CDR3 defined by/set forth in SEQ ID NO:41.
In particular embodiments, the binding agent, in particular the antibody or antibody fragment, more particularly the ISVD, as described herein does not comprise the sequence set forth in SEQ ID NO: 1 or 2.
In certain preferred embodiments, a binding agent or Sarbecovirus binding agent, in particular an antibody or antibody fragment or Sarbecovirus antibody or antibody-fragments, more particularly an ISVD, as described herein, may be characterized in that it comprises a CDR1 as present in any of SEQ ID NOs:2 to 26, wherein the CDR1 is annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia. In certain preferred embodiments, a binding agent or Sarbecovirus binding agent, in particular an antibody or antibody fragment or Sarbecovirus antibody or antibodyfragments, more particularly an ISVD, as described herein, may be characterized in that it comprises a CDR2 as present in any of SEQ ID NOs:2 to 26, wherein the CDR2 is annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia. In certain preferred embodiments, a binding agent or Sarbecovirus binding agent, in particular an antibody or antibody fragment or Sarbecovirus antibody or antibody-fragments, more particularly an ISVD, as described herein, may be characterized in that it comprises a CDR3 as present in any of SEQ ID NOs:2 to 26, wherein the CDR3 is annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia. In certain preferred embodiments, a binding agent or Sarbecovirus binding agent, in particular an antibody or antibody fragment or Sarbecovirus antibody or antibody-fragments, more particularly an ISVD, as described herein, may be characterized in that it comprises a CDR1, CDR2 and CDR3,
each independently as present in any of SEQ ID NOs:2 to 26, wherein the CDR1, CDR2 and CDR3 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
In certain preferred embodiments, a binding agent or Sarbecovirus binding agent, in particular an antibody or antibody fragment or Sarbecovirus antibody or antibody-fragments, more particularly an ISVD, as described herein, may be characterized in that it comprises a combination of CDR1, CDR2 and CDR3, wherein the CDR1, CDR2 and CDR3 are as present in a particular one of the sequences set forth in SEQ ID NOs:2 to 26, wherein the CDR1, CDR2 and CDR3 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
In certain embodiments, a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a CDR1 defined by/set forth in any one of SEQ ID NOs:27, 28, 29, or 38. In certain embodiments, a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a CDR2 defined by/set forth in any one of SEQ ID NOs:30 or 39. In certain embodiments, a binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises; and a CDR3 defined by/set forth in any one of SEQ ID NOs:31-37. In certain embodiments, the binding agent, in particular the antibody or antibody fragment, more particularly the ISVD, as described herein may comprise:
- a CDR1 defined by/set forth in any one of SEQ ID NO:27-29; a CDR2 defined by/set forth in SEQ ID NO:30; and a CDR3 defined by/set forth in any one of SEQ ID NO:31-37; or
- a CDR1 defined by/set forth in SEQ ID NO:38; a CDR2 defined by/set forth in SEQ ID NO:39; and a CDR3 defined by/set forth in any one of SEQ ID NO:31-37.
Table 2: Example definitions / sequences of the CDRs in the VHHs of certain embodiments as described herein by employing different annotation methodologies as indicated, in particular CDRs comprised in any of VHH hR3_DC23, VHH RNGL, VHH RAGL, VHH E2NGL, VHH E2AGL, VHH QRNGL, VHH QRAGL, VHH ERNGL, VHH ERAGL, VHH E10RNGL, VHH E10RAGL, VHH E81RNGL, VHH E81RAGL, VHH Q75, VHH E75, VHH E81, VHH E10E81, VHH D16, VHH T19D79, VHH TQD, VHH Q75R21Y, VHH CBV1, VHH CBV2, VHH CBV3, or VHH CBV4, determined according to Kabat or Martin system or method.
For example, polypeptidic or polypeptide binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising the complementarity determining regions (CDRs) present in any one of SEQ ID NOs:2-26, wherein the CDRs are defined according to Martin. In certain embodiments, the binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, comprise one of following sets of three complementarity determining regions (CDRs):
-CDR1 defined by/set forth in SEQ ID NO:27, CDR2 defined by/set forth in SEQ ID NO:30, and CDR3 defined by/set forth in SEQ ID NO: 31; or
-CDR1 defined by/set forth in SEQ ID NO:27, CDR2 defined by/set forth in SEQ ID NO:30, and CDR3 defined by/set forth in SEQ ID NO:32; or
-CDR1 defined by/set forth in SEQ ID NO:27, CDR2 defined by/set forth in SEQ ID NO:30, and
CDR3 defined by/set forth in SEQ ID NO: 33; or
-CDR1 defined by/set forth in SEQ ID NO:27, CDR2 defined by/set forth in SEQ ID NO:30, and CDR3 defined by/set forth in SEQ ID NO:34; or
-CDR1 defined by/set forth in SEQ ID NO:27, CDR2 defined by/set forth in SEQ ID NO:30, and CDR3 defined by/set forth in SEQ ID NO: 35; or
-CDR1 defined by/set forth in SEQ ID NO:28, CDR2 defined by/set forth in SEQ ID NO:30, and CDR3 defined by/set forth in SEQ ID NO: 31; or
-CDR1 defined by/set forth in SEQ ID NO:29, CDR2 defined by/set forth in SEQ ID NO:30, and CDR3 defined by/set forth in SEQ ID NO:36; or
-CDR1 defined by/set forth in SEQ ID NO:29, CDR2 defined by/set forth in SEQ ID NO:30, and CDR3 defined by/set forth in SEQ ID NO: 37.
In particular embodiments, polypeptidic or polypeptide binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising the complementarity determining regions (CDRs) present in any one of SEQ ID NOs:2-26, wherein the CDRs are defined according to Kabat. In certain embodiments, the binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, comprise one of following sets of three complementarity determining regions (CDRs):
-CDR1 defined by/set forth in SEQ ID NO:38, CDR2 defined by/set forth in SEQ ID NO:39, and CDR3 defined by/set forth in SEQ ID NO: 31; or
-CDR1 defined by/set forth in SEQ ID NO:38, CDR2 defined by/set forth in SEQ ID NO:39, and CDR3 defined by/set forth in SEQ ID NO:32; or
-CDR1 defined by/set forth in SEQ ID NO:38, CDR2 defined by/set forth in SEQ ID NO:39, and CDR3 defined by/set forth in SEQ ID NO: 33; or
-CDR1 defined by/set forth in SEQ ID NO:38, CDR2 defined by/set forth in SEQ ID NO:39, and CDR3 defined by/set forth in SEQ ID NO:34; or
-CDR1 defined by/set forth in SEQ ID NO:38, CDR2 defined by/set forth in SEQ ID NO:39, and CDR3 defined by/set forth in SEQ ID NO: 35; or
-CDR1 defined by/set forth in SEQ ID NO:38, CDR2 defined by/set forth in SEQ ID NO:39, and CDR3 defined by/set forth in SEQ ID NO:36; or
-CDR1 defined by/set forth in SEQ ID NO:38, CDR2 defined by/set forth in SEQ ID NO:39, and CDR3 defined by/set forth in SEQ ID NO: 37.
In further embodiments, the polypeptidic or polypeptide binding agents, in particular the antibodies and antibody fragments, more particularly the ISVDs, according to the current invention can comprise one or more framework regions (FRs) as comprised in any one of SEQ ID NOs:2-26, or variants of such FRs. More in particular, such binding agents, antibodies or antibody fragments, or ISVDs may comprise at least one, such as one, two, three or all of an FR1, FR2, FR3, and FR4 region, each independently as comprised in any one of SEQ ID NOs:2-26, or variants of such FRs. For example, such binding agents, antibodies or antibody fragment, or ISVDs, may comprise an FR1 and FR2 region, an FR1 and FR3 region, an FR1 and FR4 regions, an FR2 and FR3 region, an FR2 and FR4 region, an FR3 and FR4 region, an FR1, FR2 and FR3 region, an FR1, FR2 and FR4 region, an FR2, FR3 and FR4, or an FR1, FR3 and FR4 region as comprised in any one of SEQ ID NOs:2-26, or variants of such FRs. For delineation of FRs in these protein sequences, any one of the systems or methods for numbering amino acids in immunoglobulin protein sequences as described elsewhere herein and illustrated in Fig. 6 for VHH hR3_DC23, and known to a skilled artisan can be applied. By means of an example, sequences of the FRs in certain specific VHHs as described herein by employing the Martin or Kabat methodology are shown in Table 3.
Table 3 : Example sequences of the FRs in the VHHs of certain embodiments as described herein by employing the Martin or Kabat methodology.
A polypeptidic or polypeptide binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a framework region 1 (FR1) present in any one of SEQ ID NOs:2-26, wherein the FR1 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum, or in that it comprises a variant FR1 which is at least 90% or 95% identical to, or which has at most 3, such as 1, 2 or 3, amino acid substitutions, deletions or additions, such as preferably conservative and/or humanizing substitutions, compared to, a FR1 present in any one of SEQ ID NOs:2-26, wherein the FR1 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum
A polypeptidic or polypeptide binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a framework region 2 (FR2) present in any one of SEQ ID NOs:2-26, wherein the FR2 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum, or in that it comprises a variant FR2 which is at least 85% or 90% identical to, or which has at most 2, such as 1 or 2, amino acid substitutions, deletions or additions, such as preferably conservative and/or humanizing substitutions, compared to, a FR2 present in any one of SEQ ID NOs:2-26, wherein the FR2 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
A polypeptidic or polypeptide binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a framework region 3 (FR3) present in any one of SEQ ID NOs:2-26, wherein the FR3 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum, or in that it comprises a variant FR3 which is at least 80%, 85%, 90% or 95% identical to, or which has at most 9, such as 1, 2, 3, 4, 5, 6, 7, 8 or 9, amino acid substitutions, deletions or additions, such as preferably conservative and/or humanizing substitutions, compared to, a FR3 present in any one of SEQ ID NOs:2-26, wherein the FR3 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
A polypeptidic or polypeptide binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises a framework region 4 (FR4) present in any one of SEQ ID NOs:2-26, wherein the FR4 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum, or in that it comprises a variant FR4 which is at least 90% identical to, or which has at most 1 amino acid substitution, deletion or addition, such as preferably a conservative and/or humanizing substitution, compared to, a FR4 present in any one of SEQ ID NOs:2-26, wherein the FR4 is defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
In further embodiments, a polypeptidic or polypeptide binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises, each independently, a FR1 present in any one of SEQ ID Nos:2-26 or a variant FR1 as defined hereinabove; a FR2 present in any one of SEQ ID Nos:2-26 or a variant FR2 as defined hereinabove; a FR3 present in any one of SEQ ID Nos:2-26 or a variant FR3 as defined hereinabove; and a FR4 present in any one of SEQ ID Nos: 2-26 or a variant FR4 as defined hereinabove, wherein the FR1, FR2, FR3 and FR4 are defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
In particular embodiments, a polypeptidic or polypeptide binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises at least one, or the particular combination of two, three or all of the framework regions (FRs) as present in any one of SEQ ID NOs: 2-26, or any variant of said FR or FRs as defined herein above, wherein the FRs are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
In particular embodiments, a polypeptidic or polypeptide binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it
comprises at least one, or the particular combination of two, three or all of the framework regions (FRs) present in any one of SEQ ID NOs: 2 to 26, wherein the FRs are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
In further particular embodiments, polypeptidic or polypeptide binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising, each independently, a FR1 present in any one of SEQ ID NOs:2-26; a FR2 present in any one of SEQ ID NOs:2-26; a FR3 present in any one of SEQ ID NOs:2-26, and a FR4 present in any one of SEQ ID NOs:2-26, wherein the FR1, FR2, FR3 and FR4 are defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
In particular embodiments, a polypeptidic or polypeptide binding agent, in particular an antibody or antibody fragment, more particularly an ISVD, as described herein may be characterized in that it comprises:
- a FR1 defined by any one of SEQ ID NO:42-45, a FR2 defined by SEQ ID NO:46, a FR3 defined by any one of SEQ ID NO:47-59 and a FR4 defined by SEQ ID NO:60; or
- a FR1 defined by any one of SEQ ID NO:61-68, a FR2 defined by SEQ ID NO:46, a FR3 defined by any one of SEQ ID NO:69-81 and a FR4 defined by SEQ ID NO:60.
In further particular embodiments, polypeptidic or polypeptide binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising a FR1, FR2, FR3 and FR4 as present in the same sequence of any of the sequences shown in SEQ ID NOs:2-26, wherein the FR1, FR2, FR3 and FR4 are defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
In yet further particular embodiments, polypeptidic or polypeptide binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising all four framework regions (FRs) present in any one of SEQ ID NOs:2-26, wherein the FRs are defined according to any one of AbM, Chothia, Martin, Kabat, IMGT or MacCallum.
For example, polypeptidic or polypeptide binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising the framework regions (FRs) present in any one of SEQ ID NOs:2-26, wherein the FRs are defined according to Martin. In certain embodiments, the binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, comprise one of following sets of framework regions (FRs):
-FR1 defined by/set forth in SEQ ID NO:42, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:47, and FR4 defined by/set forth in SEQ ID NO:60;
-FR1 defined by/set forth in SEQ ID NO:42, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:48, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:42, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:49, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:42, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:50, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:43, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:48, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:42, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:51, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:42, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO: 52, and FR4 defined by/set forth in SEQ ID NO: 60; or
-FR1 defined by/set forth in SEQ ID NO:42, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:53, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:42, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:54, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:43, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:54, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:44, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:47, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:45, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:55, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:45, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:56, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:45, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:57, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:43, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:58, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:45, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:59, and FR4 defined by/set forth in SEQ ID NO:60.
For example, polypeptidic or polypeptide binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, as described herein can be defined as comprising the framework regions (FRs) present in any one of SEQ ID NOs:2-26, wherein the FRs are defined according to Kabat. In certain embodiments, the binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, comprise one of following sets of framework regions (FRs):
-FR1 defined by/set forth in SEQ ID NO:61, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:69, and FR4 defined by/set forth in SEQ ID NO:60;
-FR1 defined by/set forth in SEQ ID NO:61, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:70, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:61, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:71, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:61, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:72, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:62, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:70, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:61, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO: 73, and FR4 defined by/set forth in SEQ ID NO: 60; or
-FR1 defined by/set forth in SEQ ID NO:61, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:74, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:61, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:75, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO: 61, FR2 defined by/set forth in SEQ IDNO:46, FR3 defined by/set forth in SEQ ID NO:76, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:62, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:76, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:63, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:69, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:64, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:77, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:64, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:78, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:65, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:74, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:66, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:76, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:67, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:79, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:68, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:80, and FR4 defined by/set forth in SEQ ID NO:60; or
-FR1 defined by/set forth in SEQ ID NO:67, FR2 defined by/set forth in SEQ ID NO:46, FR3 defined by/set forth in SEQ ID NO:81, and FR4 defined by/set forth in SEQ ID NO:60.
In particular embodiments, the polypeptidic or polypeptide binding agents, in particular antibodies or antibody fragments, comprise one or more ISVDs individually defined by or set forth in any one of SEQ ID NOs: 2 to 26, or comprise one or more ISVDs comprising or consisting of an amino acid sequence selected from the group of SEQ ID NO: 2 to 26.
In further embodiments, said polypeptidic or polypeptide binding agents, in particular antibodies or antibody fragments, more particularly ISVDs, comprise or consist of an amino acid sequence with at least 90% identity to an amino acid sequence selected from the group of SEQ ID NO: 2 to 26, or with at least 95% identity to an amino acid sequence selected from the group of SEQ ID NO: 2 to 26. Such non-identity or variability, is preferably limited to non-identity or variability in FR amino acid residues. Cconservative amino acid substitutions can be introduced in a protein or polypeptide whereby such substitutions have no essential or substantial effect on the protein's activity. Preferred conservative substitutions are those fulfilling the criteria defined for an accepted point mutation in Dayhoff et al., Atlas of Protein Sequence and Structure, 5, pp. 345-352 (1978 & Supp.), which is incorporated herein by reference. Examples of conservative substitutions are substitutions including
but not limited to the following groups: (a) valine, glycine; (b) glycine, alanine; (c) valine, isoleucine, leucine; (d) aspartic acid, glutamic acid; (e) asparagine, glutamine; (f) serine, threonine; (g) lysine, arginine, methionine; and (h) phenylalanine, tyrosine. A “homologue”, or “homologues” of a protein of interest encompass(es) proteins having amino acid substitutions, deletions and/or insertions relative to an unmodified protein of interest and having essentially or substantially similar biological and functional activity as the unmodified protein from which it is/they are derived.
A “percentage (of) sequence identity” is calculated by comparing two optimally aligned (amino acid or nucleic acid) sequences over the window of comparison, determining the number of positions at which the identical amino acid or nucleotide residue occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison (i.e., the window size), and multiplying the result by 100 to yield the percentage of (amino acid or nucleic acid) sequence identity.
Immunoglobulin single variable domains such as Domain antibodies and Nanobody® (including VHH domains) as described herein can be and/or have been subjected to humanization, i.e. to increase the degree of sequence identity with the closest human germline sequence. In particular, humanized immunoglobulin single variable domains, such as Nanobody® (including VHH domains) are immunoglobulin single variable domains in which at least one amino acid residue is present (and in particular, at least one framework residue) that is and/or that corresponds to a humanizing substitution (as defined further herein). Potentially useful humanizing substitutions can be ascertained by comparing the sequence of the framework regions of a VHH sequence with the corresponding framework sequence of one or more closely related human VH sequences, after which one or more of the potentially useful humanizing substitutions (or combinations thereof) thus determined can be introduced into said VHH sequence (in any manner known per se, as further described herein) and the resulting humanized VHH sequences can be tested for affinity for the target, for stability, for ease and level of expression, and/or for other desired properties. In this way, by means of a limited degree of trial and error, other suitable humanizing substitutions (or suitable combinations thereof) can be determined by the skilled person. Also, based on what is described before, (the framework regions of) an immunoglobulin single variable domain, such as a Nanobody® (including VHH domains) may be partially humanized or fully humanized.
Humanized immunoglobulin single variable domains, in particular Nanobody®, may have several advantages, such as a reduced immunogenicity, compared to the corresponding non-humanized VHH domains. By humanized is meant mutated so that immunogenicity upon administration in human
patients is minor or non-existent. The humanizing substitutions are chosen such that the resulting humanized amino acid sequence and/or ISVD or VHH still retains the favourable properties of the parental (non-humanized) VHH, such as the antigen-binding capacity. Based on the description provided herein, the skilled person will be able to select humanizing substitutions or suitable combinations of humanizing substitutions which optimize or achieve a desired or suitable balance between the favourable properties provided by the humanizing substitutions on the one hand and the favourable properties of non-humanized VHH domains on the other hand. Such methods are known by the skilled addressee. A human consensus sequence can be used as target sequence for humanization, but also other means are known in the art. One alternative includes a method wherein the skilled person aligns a number of human germline alleles, such as for instance but not limited to the alignment of IGHV3 alleles, and to use said alignment for identification of residues suitable for humanization in the target sequence. Also a subset of human germline alleles most homologous to the target sequence may be aligned as starting point to identify suitable humanisation residues. Alternatively, the VHH is analyzed to identify its closest homologue in the human alleles and used for humanisation construct design. A humanisation technique applied to Camelidae VHHs may also be performed by a method comprising the replacement of specific amino acids, either alone or in combination. Said replacements may be selected based on what is known from literature, from known humanization efforts, as well as from human consensus sequences compared to the natural VHH sequences, or from the human alleles most similar to the VHH sequence of interest. As can be seen from the data on the VHH entropy and VHH variability given in Tables A-5-A-8 of WO 08/020079, some amino acid residues in the framework regions are more conserved between human and Camelidae than others. Generally, although the invention in its broadest sense is not limited thereto, any substitutions, deletions or insertions (or additions) are preferably made at positions that are less conserved. Also, generally, amino acid substitutions are preferred over amino acid deletions or insertions. For instance, a human-like class of Camelidae single domain antibodies contain the hydrophobic FR2 residues typically found in conventional antibodies of human origin or from other species, but compensating this loss in hydrophilicity by other substitutions at position 103 that substitutes the conserved tryptophan residue present in VH from double-chain antibodies. As such, peptides belonging to these two classes show a high amino acid sequence homology to human VH framework regions and said peptides might be administered to a human directly without expectation of an unwanted immune response therefrom, and without the burden of further humanisation. Indeed, some Camelidae VHH sequences display a high sequence homology to human VH framework
regions and therefore said VHH might be administered to patients directly without expectation of an immune response therefrom, and without the additional burden or need of humanization.
Suitable mutations, in particular substitutions, can be introduced during humanization to generate a polypeptide with reduced binding to pre-existing antibodies (reference is made for example to WO 2012/175741 and WO2015/173325), for example at least one of the positions: 11, 13, 14, 15, 40, 41, 42, 82, 82a, 82b, 83, 84, 85, 87, 88, 89, 103, or 108. The amino acid sequences and/or VHH of the invention may be or have been suitably humanized at any framework residue(s), such as at one or more Hallmark residues (as defined below) or at one or more other framework residues (i.e. nonHallmark residues) or any suitable combination thereof.
Depending on the host organism used to express the amino acid sequence, ISVD or VHH or polypeptide as described herein, such deletions and/or substitutions may also be designed in such a way that one or more sites for posttranslational modification (such as one or more glycosylation sites) are removed, as will be within the ability of the person skilled in the art. Alternatively, substitutions or insertions may be designed so as to introduce one or more sites for attachment of functional groups (as described herein), for example to allow site-specific pegylation.
In some cases, at least one of the typical Camelidae hallmark residues with hydrophilic characteristics at position 37, 44, 45 and/or 47 is replaced (see Table A-03 of W02008/020079). Another example of humanization includes substitution of residues in FR 1, such as position 1, 5, 11, 14, 16, and/or 28; in FR3, such as positions 73, 74, 75, 76, 78, 79, 82b, 83, 84, 93 and/or 94; and in FR4, such as position 10 103, 104, 108 and/or 111 (see Tables A-05 -A08 of W02008/020079; all numbering according to the Kabat-methodology).
Immunoglobulin single variable domains as described herein can be and/or have been subjected to affinity maturation. In embodiments, the ISVD is a variant of an ISVD as described herein, in particular an ISVD of any one of SEQ ID NO:2 to 26, in particular an ISVD of SEQ ID NO:2 or 25, which has an improved affinity for SARS-CoV-2 S2. In embodiments, said variant ISVD is obtained by an affinity maturation protocol. In embodiments, the ISVD variant may comprise up to 4 such as 1, 2, 3 or 4 substitutions, in particular substitutions of paratope residues, optionally wherein said paratope comprises or consists of residues at positions 96, 97, 98, 99, 100a, 100b and 1001 according to Kabat numbering of the ISVD of any one of SEQ ID NO:2 to 26, in particular an ISVD of SEQ ID NO:2 or 25, and/or at positions 27, 29, 30, 31, 52, 100, 100c and lOOd according to Kabat numbering of the ISVD of any one of SEQ ID NO:2 to 26, in particular an ISVD of SEQ ID NO:2 or 25. In embodiments, the ISVD variant may comprise a substitution of a residue at position 96, 97,
98, 99, 100b or 1001 according to Kabat numbering of the ISVD of any one of SEQ ID NO:2 to 26, in particular an ISVD of SEQ ID NO:2 or 25. In embodiments, the ISVD may comprise up to 4 such as 1, 2, 3 or 4 substitutions of residues at positions 27, 29, 30, 31, 100, 100c and/or lOOd according to Kabat numbering of the ISVD of any one of SEQ ID NO:2 to 26, in particular an ISVD of SEQ ID NO:2 or 25. In certain embodiments, the residue at position 96 may be substituted with F, R, W, H or Q, preferably with F, R or W. In certain embodiments, the residue at position 97 may be substituted with I, A or L. In certain embodiments, the residue at position 98 may be substituted with A or T. In certain embodiments, the residue at position 99 may be substituted with R. In certain embodiments, the residue at position 100b may be substituted with F. In certain embodiments, the residue at position 1001 may be substituted with L, H or V, more preferably with L. In certain embodiments, the residue at position 27 may be substituted with H, L, P, S or T, preferably with H. In certain embodiments, the residue at position 29 may be substituted with Y, H, M, R, W, I, L, S, V, A or G, preferably with Y, H, M, R or W. In certain embodiments, the residue at position 30 may be substituted with K, T, I, R, G, H, L, P, F, N or W, preferably with K, T or I, more preferably with K. In certain embodiments, the residue at position 31 may be substituted with S. In certain embodiments, the residue at position 100 may be substituted with R. In certain embodiments, the residue at position 100c may be substituted with H or A, preferably with H. In certain embodiments, the residue at position lOOd may be substituted with R, H, P, Q or V. Preferably, the substitution(s) is/are selected from the substitutions set forth in Table 14.
In particular embodiments, the ISVD is a variant of hR3_DC23 of SEQ ID NO:2. Advantageously, the variant may have an improved affinity for SARS-CoV-2 S2 compared to hR3_DC23. In certain embodiments, the ISVD, in particular the hR3_DC23 variant, is an ISVD comprising an amino acid sequence defined by any one of SEQ ID NO: 151-182.
In particular embodiments, the ISVD is a variant of CBV3 of SEQ ID NO:25. Advantageously, the variant may have an improved affinity for SARS-CoV-2 S2 compared to CBV3. In certain embodiments, the ISVD, in particular the CBV3 variant, is an ISVD comprising an amino acid sequence defined by any one of SEQ ID NO: 183-244, preferably an ISVD comprising an amino acid sequence defined by any one of SEQ ID NO: 183-188, 190-195, 197, 198, or 200-244.
The binding agents or Sarbecovirus binding agents (can be used interchangeably) according to the current invention can also be described functionally by any individual fimction/embodiment or by any combination of any number of the individual fimctions/embodiments described hereafter and given an arbitrary number “n” between brackets “(n)”. The numerical order of these individual
functions is random and not imposing any preference on an individual function; similarly, this random numerical order is not imposing any preference on any combination of two or more of the individual functions. Any such combination is furthermore not to be considered as arbitrary as the binding agents or Sarbecovirus binding agents herein exert each of these individual functions.
The present invention thus provides binding agents, in particular antibodies or antigen-binding fragments thereof, that (1) specifically bind to a Sarbecovirus such as SARS-CoV-2, SARS-CoV-1 and Khosta-2 and may also be referred to herein as Sarbecovirus binding agents or Sarbecovirus antibodies and antibody fragments. In certain embodiments, the binding agents (2) do not bind Middle East respiratory syndrome coronavirus (MERS-CoV).
“Binding” means any interaction, be it direct or indirect. A direct interaction implies a contact (e.g. physical or chemical) between two binding partners. An indirect interaction means any interaction whereby the interaction partners interact in a complex of more than two molecules. An interaction can be completely indirect (e.g. two molecules are part of the same complex with the help of one or more bridging molecules but don’t bind in the absence of the bridging molecule(s)). An interaction may be partly direct or partly indirect: there is still a direct contact between two interaction partners, but such contact is e.g. not stable, and is stabilized by the interaction with one or more additional molecules.
“Specificity of binding” or “binding specificity” or “specifically binding” refers to the situation in which a molecule A is, at a certain concentration (e.g. sufficient to inhibit or neutralize a protein or process of interest) binding to a target of interest (e.g. protein) with higher affinity (e.g. at least 2- fold, 5-fold, orat least 10-fold higher affinity, e.g. at least 20-, 50- or 100-fold or more higher affinity) than the affinity with which it is possibly (if at all) binding to other targets (targets not of interest). Specific binding does not mean exclusive binding. However, specific binding does mean that a binder has a certain increased affinity or preference for one or a few of its targets. Exclusivity of binding refers to the situation in which a binder is binding only to the target of interest. The term "affinity", as used herein, generally refers to the degree to which one molecule (e.g. ligand, chemical, protein or peptide, antibody or antibody fragment) binds to another molecule (e.g. (target) protein or peptide) so as to shift the equilibrium of single molecule monomers towards a complex formed by (specific)(non-covalent) binding of the two molecules. Non-covalent interaction or binding between 2 or more binding partners may involve interactions such as van der Waals interaction, hydrogen bonding, and salt bridges. The “dissociation constant” or “binding constant” (KD) is commonly used to describe the affinity between the two molecules and it is often calculated by the ratio of the rate
constant for the complex formation (referred to as the "kon" value) to the rate constant for dissociation of said complex (the "kOff" or "kdis" value). The measurement of binding affinity of a molecule to another molecule, such as an antibody or antibody-fragment to an antigen, or a ligand to a receptor, is known to the skilled person and includes, e.g., real-time, label free bio-layer interferometry assay, e.g., an Octet® RED96 system (ForteBio), or surface plasmon resonance (SPR), e.g., BIACORE™, or solution-affinity ELISA.
The terms "('oronaviridae" and the more common name “coronavirus” refer to a family of viruses, which has its name from the large spike protein molecules that are present on the virus surface and give the virions a crown-like shape. The Coronoviridae family comprises four genera: Alphacoronavirus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus. Coronaviruses represent a diverse family of large enveloped positive-stranded RNA viruses that infect a wide range of animals, a wide variety of vertebrate species, and humans. The spike (S) proteins of coronaviruses are essential for host receptor-binding and subsequent fusion of the viral and host cell membrane, effectively resulting in the release of the viral nucleocapsids in the host cell cytoplasm (Letko et al. (2020) Nat Microbiol 5:562-569).
Four coronaviruses, presumably from a zoonotic origin, are endemic in humans: HCoV-NL63 and HCoV-229E (a-coronaviruses) and HCoV-OC43 and HCoV-HKUl (P-coronaviruses). In addition, 3 episodes of severe respiratory disease caused by -coronaviruses have occurred since 2000: severe acute respiratory syndrome virus (SARS), caused by SARS-CoV-1, emerged from a zoonotic origin (bats via civet cats as an intermediate species) and disappeared in 2004 (Drosten et al. 2003, N Engl J Med 348: 1967-1976). Over 8000 SARS cases were reported with a mortality rate of approximately 10%. In 2012, Middle East respiratory syndrome (MERS) emerged in the Arabian Peninsula. MERS is caused by MERS-CoV, has been confirmed in over 2500 cases and has a case fatality rate of 34% (de Groot et al. 2013, N Engl J Virol 87:7790-7792). Starting at the end of 2019, the third zoonotic human coronavirus emerged with cases of severe acquired pneumonia reported in the city of Wuhan (China) being caused by a new P-coronavirus, now known as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), given its genetic relationship with SARS-CoV-1 (Chen et al. (2020) Lancet 395:507-513). Similar to severe acute respiratory syndrome coronavirus (SARS-CoV) and Middle East respiratory syndrome coronavirus (MERS-CoV) infections, patients exhibited symptoms of viral pneumonia including fever, difficult breathing, and bilateral lung infiltration in the most severe cases (Gralinski et al. (2020) Viruses 12: 135).
The term “Sarbecovirus'” as used herein refers to a subgenus within the genus Betacoronavirus and includes the species Severe acute respiratory syndrome-related coronavirus (SARSr-CoV or SARS- CoV, also known as SARS coronavirus, SARS-related coronavirus, and Severe acute respiratory syndrome coronavirus, which are used as synonyms herein). Non-limiting examples of strains belonging to the SARS-CoV species include SARS-CoV-1 and SARS-CoV-2.
The first available genome sequence placed the novel human pathogen SARS-CoV-2 in the Sarbecovirus subgenus of Coronaviridae, the same subgenus as the SARS virus. Although SARS- CoV-2 belongs to the same genus Betacoronavirus as SARS-CoV (lineage B) and MERS-CoV (lineage C), genomic analysis revealed greater similarity between SARS-CoV-2 and SARS-CoV, supporting its classification as a member of lineage B (from the International Committee on Taxonomy of Viruses).
Among other Betacoronaviruses, this virus is characterized by a unique combination of polybasic cleavage sites, a distinctive feature known to increase pathogenicity and transmissibility. A bat Sarbecovirus, Bat CoV RaTG13, sampled from a Rhinolophus affinis horseshoe bat was reported to cluster with SARS-CoV-2 in almost all genomic regions with approximately 96% genome sequence identity (and over 93% similarity in the receptor binding domain (RBD) of the spike protein); another mammalian species may have acted as intermediate host. One of the suspected intermediate hosts, the Malayan pangolin, harbours coronaviruses showing high similarity to SARS-CoV-2 in the receptor-binding domain, which contains mutations believed to promote binding to the angiotensinconverting enzyme 2 (ACE2) receptor and demonstrates a 97% amino acid sequence similarity.
SARS-CoV-1 and -2 both use angiotensin converting enzyme 2 (ACE2) as a receptor on human cells. SARS-CoV-2 binds ACE2 with a higher affinity than SARS-CoV-1 (Wrapp et al. (2020) Science 367: 1260-1263). SARS-CoV-2 differentiates from SARS-CoV-1 and several SARS-related coronaviruses (SARSr-CoVs) as outlined in e.g. Abdelrahman et al. (2020. Front Immunol 11: 552909).
SARS-CoV-2 refers to the newly-emerged Sarbecovirus which was identified as the cause of a serious and worldwide outbreak of severe acquired pneumonia starting in the city of Wuhan (China). The long-term global spread of SARS-CoV-2, together with selective pressure for immune escape, led to adaptation of the virus to the host and generation of new SARS-CoV-2 variants. Specifically, multiple mutations in the spike glycoprotein evolved and are evolving, including mutations that are located in the spike SI subunit. A skilled person is able to find information about the SARS-CoV-2 variants and their sequences in the relevant literature and databases, such as the SARS-CoV-2 genetic
lineage tracing system funded by NIH (https://outbreak.info/). For example, a SARS-CoV-2 variant may comprise a mutation at one or more positions selected from N439, K417, S477, L452, T478, E484, P384, N501 and D614 (relative to the SARS-CoV-2 spike amino acid sequence as defined in SEQ ID NO:93). Further non-limiting examples of SARS-CoV-2 variants include a SARS-CoV-2 variant comprising a mutation at position N501 such as a N501Y variant (e.g. SARS-CoV-2 Alpha variant); a SARS-CoV-2 variant comprising a mutation at positions N501 and E484 such as aN501Y and E484K variant (e.g. SARS-CoV-2 Alpha + E484K variant); a SARS-CoV-2 variant comprising a mutation at positions K417, E484 and N501 such as a K417N, E484K and N501Y variant (e.g. SARS-CoV-2 beta variant); a SARS-CoV-2 variant comprising a mutation at positions P384, K417, E484 and N501 such as a P384L, K417N, E484K and N501Y variant (e.g. SARS-CoV-2 beta + P384L variant); a SARS-CoV-2 variant comprising a mutation at positions L452 and E484 such as a L452R and E484Q variant (e.g. SARS-CoV-2 kappa variant); a SARS-CoV-2 variant comprising a mutation at positions L452 and T478 such as a L452R and T478K variant (e.g. SARS-CoV-2 delta variant); a SARS-CoV-2 variant comprising a mutation at position L452 such as a L452R variant (e.g. SARS-CoV-2 epsilon variant); a SARS-CoV-2 variant comprising a mutation at position K417 such as a K417T variant (e.g. SARS-CoV-2 gamma variant); a SARS-CoV-2 variant comprising a mutation at position D614 such as a D614G variant (e.g. SARS-CoV-2 D614G variant, SARS-CoV- 2 Omicron BA.1 variant or SARS-CoV-2 Omicron BA.2 variant); a SARS-CoV-2 variant comprising a mutation at positions K147, W152R, F157, 1210, G257, D339, G446 and N460 such as a K147E, W152R, F157L, I210V, G257S, D339H, G446S and N460K variant (e.g. SARS-CoV-2 Omicron BA.2.75 variant, SARS-CoV-2 Omicron BA.2.75.2 variant); a SARS-CoV-2 variant comprising a mutation at positions R346, F486 and DI 199 such as a R346T, F486S and D1199N variant (e.g. SARS-CoV-2 Omicron BA.2.75.2 variant); a SARS-CoV-2 variant comprising a mutation at positions H69, V70, L452 and F486 such as a H69-, V70-, L452R and F486V variant (e.g. SARS- CoV-2 Omicron BA.4/BA.5 variant); a SARS-CoV-2 variant comprising a mutation at positions R346 and N658 such as a R346T and N658S variant (e.g. SARS-CoV-2 Omicron BA.4.6 variant); a SARS-CoV-2 variant comprising a mutation at position R346 such as a R346T variant (e.g. SARS- CoV-2 Omicron BF.7 variant); a SARS-CoV-2 variant comprising a mutation at positions R346, K444 and N460 such as a R346T, K444T and N460K variant (e.g. SARS-CoV-2 Omicron BQ.1.1 variant); a SARS-CoV-2 variant comprising a mutation at positions V83, Y144, H146, QI 83, V213, R346, L368, V445, G446, N460, F486 and F490 such as a V83A, Y144-, H146Q, Q183E, V213E, R346T, L368I, V445P, G446S, N460K, F486S and F490S variant (e.g. SARS-CoV-2 Omicron XBB variant) or a V83A, Y144-, H146Q, Q183E, V213E, R346T, L368I, V445P, G446S, N460K, F486P
and F490S variant (e.g. SARS-CoV-2 Omicron XBB.1.5 (-G252V) variant); a SARS-CoV-2 variant comprising a mutation at positions V83, Q52, Y144, H146, F157, Q183, V213, G252, R346, L368, V445, G446, L452, F456, N460, F486 and F490 such as a V83A, Q52H, Y144-, H146Q, F157L, Q183E, V213E, G252V, R346T, L368I, V445P, G446S, L452R, F456L, N460K, F486P and F490S variant (e.g. SARS-CoV-2 HV.l); a SARS-CoV-2 variant comprising a mutation at positions V83, Q52, Y144, H146, Q183, V213, G252, R346, L368, V445, G446, F456, N460, F486 and F490 such as a V83A, Q52H, Y144-, H146Q, Q183E, V213E, G252V, R346T, L368I, V445P, G446S, F456L, N460K, F486P and F490S variant (e.g. SARS-CoV-2 EG5.1); a SARS-CoV-2 variant comprising a mutation at positions T19, R21, L24, P25, P26, A27, S50, H69, V70, V127, G142, Y144, F157, R158, N211, L212, V213, L216, H245, A264, 1332, G339, K356, , S371, S373, S375, T376, R403, D405, R408, K417, N440, V445, G446, N450, L452, N460, S477, T478, N481, V483, E484, F486, Q498, N501, Y505, E554, A570, D614, P621, H655, N679, P681, N764, D796, S939, Q954, N969 and Pl 143 such as a T19I, R21T, L24S, P25-, P26-, A27-, S50L, H69-, V70-, , V127F, G142D, Y144-, , F157S, R158G, N211I, L212-, V213G, L216F, H245N, A264D, I332V, G339H, K356T, S371F, S373P, S375F, T376A, R403K, D405N, R408S, K417N, N440K, V445H, G446S, N450D, L452W, N460K, S477N, T478K, N481K, V483-, E484K, F486P, Q498R, N501Y, Y505H, E554K, A570V, D614G, P621S, H655Y, N679K, P681R, N764K, D796Y, S939F, Q954H, N969K and P1143L variant (e.g. SARS-CoV-2 BA.2.86.1). The Alpha variant (also known as B.1.1.1.7 lineage) of SARS-CoV-2 was first detected in the UK late 2020 and was one of the first reported variants of concern of SARS-CoV-2; a SARS-CoV-2 variant comprising a mutation at position F456 such as a F456L mutation (e.g. SARS CoV-2 Omicron FL.1.5.1, SARS CoV-2 Omicron EG.5.1, SARS CoV- 2 Omicron HK.3, SARS CoV-2 Omicron HV.l, SARS CoV-2 Omicron KP.1.1, SARS CoV-2 Omicron KP.2.3, SARS CoV-2 Omicron KP.3, SARS CoV-2 Omicron KP.3.1.1, SARS CoV-2 Omicron LB.l, and SARS CoV-2 Omicron XEC); a SARS-CoV-2 variant comprising a mutation at position Q493 such as a Q493E mutation (e.g. SARS CoV-2 Omicron KP.3, SARS CoV-2 Omicron KP.3.1.1, and SARS CoV-2 Omicron XEC); or a SARS-CoV-2 variant comprising a mutation at position F456 and a mutation at position Q493 such as F456L and Q493E mutations (e.g. SARS CoV-2 Omicron KP.3, SARS CoV-2 Omicron KP.3.1.1, and SARS CoV-2 Omicron XEC). It contained several mutations in the spike protein, including N501Y mutation and D614G mutation. The Omicron variant of SARS-CoV-2 was first identified in South Africa and Botswana and was reported to the World Health Organization (WHO) on November 24, 2021, as a novel variant (Fan et al. 2022. Signal Transduct Target Ther. 7: 141). The Omicron variant is not a single strain, but evolved into at least three lineages, including BA.1, BA.2, and BA.3. Up to 60 mutations have been
identified in the BA. 1 lineage, with as many as 38 of these occurring in the spike (S) protein, one in the envelope (E) protein, two in the membrane (M) protein, and six in the nucleocapsid (N) protein. BA.2 lineage possesses 57 mutations, with 31 in the S protein, of which the N-terminus is significantly different from that of BA. l. The term “SARS-CoV-2” as used herein covers both the original strain identified in Wuhan as well as variants thereof.
Khosta-2 as used herein refers to Sarbecovirus found in Russian horseshoe bats, which was found to be capable of using human ACE2 to facilitate entry (Seifert et al. 2022 PLoS Pathog 18(9):el010828).
The binding agents, in particular the antibodies and antibody fragments (3) specifically bind or bind to spike protein of a Sarbecovirus such as SARS-CoV-2 spike protein, SARS-CoV-1 spike protein or Khosta-2 spike protein, in particular the binding agents, in particular the antibodies and antibody fragments, (4) specifically bind or bind to S2 subunit, or to a part of the S2 subunit, of the Sarbecovirus spike protein, more particularly, the binding agents, in particular the antibodies and antibody fragments, (22) specifically bind or bind to or within a region of the S2 subunit located from amino acid E 1188 to amino acid Y 1206, preferably a region located from amino acid N 1192 to amino acid Y1206 or a region located from amino acid El 188 to amino acid L1203, more preferably a region located from amino acid N1192 to amino acid LI 203, even more preferably a region located from amino acid N1194 to amino acid L1203, most preferably a region located from amino acid N1194 to amino acid Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93. In certain embodiments, the binding agents, in particular the antibodies and antibody fragments, (23) specifically bind or bind to or within a region of spike protein of a Sarbecovirus or S2 subunit of the Sarbecovirus spike protein corresponding to the region from amino acid E 1188 to amino acid Y 1206, preferably amino acid N1192 to amino acid Y 1206 or amino acid El 188 to amino acid LI 203, more preferably amino acid N1192 to amino acid L1203, even more preferably amino acid N1194 to amino acid L1203, most preferably amino acid N1194 to amino acid Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93. More particularly, the binding agents, in particular the antibodies and antibody fragments, (5) specifically bind or bind to heptad repeat 2 (HR2) domain, or to a part of the HR2 domain, of (the S2 subunit of) the Sarbecovirus spike protein. In certain embodiments, the binding agents, in particular the antibodies and antibody fragments, (6) specifically bind or bind to or within a region of the HR2 domain proximal to the viral membrane, preferably a region located from amino acid Al 174 to amino acid E1202, more preferably a region located from amino acid 11179 to amino acid El 202, even more preferably a region located from amino acid DI 184 to amino acid El 202, still more preferably a region located from amino acid El 188 to amino
acid E1202 or a region located from amino acid VI 189 to amino acid E1202, yet more preferably a region located from amino acid N1194 to amino acid El 202, most preferably a region located from amino acid N1194 to amino acid Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, or (7) specifically bind or bind to a region of the HR2 domain (or of the S2 subunit) corresponding to the region from amino acid El 188 to amino acid Y1206 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, preferably a region of the HR2 domain (or of the S2 subunit) corresponding to the region from amino acid El 188 to amino acid Y1203 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, more preferably a region of the HR2 domain (or of the S2 subunit) corresponding to the region from amino acid Al 190 to amino acid L1203 of the SARS- CoV-2 spike protein as defined in SEQ ID NO:93, such as a region of the HR2 domain (or of the S2 subunit) corresponding to the region from amino acid KI 191 to amino acid E1202 of the SARS- CoV-2 spike protein as defined in SEQ ID NO:93, or a region of the HR2 domain (or of the S2 subunit) corresponding to the region from amino acid N1192 to amino acid Q1201 of the SARS- CoV-2 spike protein as defined in SEQ ID NO:93 (such as the region from amino acid N1192 to amino acid Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93), even more preferably a region of the HR2 domain (or of the S2 subunit) corresponding to the region from amino acid N1194 to amino acid L1203 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, most preferably a region of the HR2 domain (or of the S2 subunit) corresponding to the region from amino acid N1194 to amino acid Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93 such as a region of the HR2 domain (or of the S2 subunit) corresponding to the region from amino acid SI 196 to amino acid Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93. In particular embodiments, the binding agents, in particular the antibodies and antibody fragments, (8) specifically bind or bind to at least one, at least two, at least three, at least four, at least five, at least six, at least seven or all, of the amino acid residues N1192, N1194, SI 196, LI 197, DI 199, L1200, Q1201 and E1202, of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, preferably to at least one, at least two, at least three, at least four or all of the amino acid residues N1194, SI 196, DI 199, Q1201 and E1202 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, more preferably to at least one, at least two, at least three or all of the amino acid residues N1194, S 1196, D1199 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, most preferably to at least one or both of the amino acid residues SI 196 and Q1201 of the SARS- CoV-2 spike protein as defined in SEQ ID NO:93. In embodiments, the binding agents, in particular the antibodies and antibody fragments, (24) specifically bind or bind to at least one, at least two, at least three, at least four, at least five, at least six, at least seven or all, amino acid residue(s) of spike
protein of a Sarbecovirus or S2 subunit or HR2 domain of the Sarbecovirus spike protein corresponding to the amino acid residues N1192, N1194, SI 196, LI 197, DI 199, LI 200, Q1201 and E1202 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, preferably to at least one, at least two, at least three, at least four or all amino acid residue(s) corresponding to the amino acid residues N1194, SI 196, DI 199, Q1201 and E1202 of the SARS-CoV-2 spike protein as defined in SEQ ID NO: 93, more preferably to at least one, at least two, at least three or all amino acid residues(s) corresponding to the amino acid residues N1194, SI 196, DI 199 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, most preferably to at least one or both amino acid residue(s) corresponding to the amino acid residues SI 196 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93. In particular embodiments, the binding agents, in particular the antibodies and antibody fragments, (25) specifically bind or bind to the amino acid residues SI 196 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93 or to the amino acid residues of spike protein corresponding to said amino acid residues of SI 196 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, optionally to the amino acid residues N1194, S 1196, D1199 and Q1201ofthe SARS-CoV-2 spike protein as defined in SEQ ID NO:93 or to the amino acid residues of spike protein corresponding to said amino acid residues N1194, SI 196, DI 199 and Q1201of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93.
In particular embodiments, (26) at least one, at least two, at least three, at least four, at least five, at least six, at least seven or all, of the amino acid residues N1192, N1194, SI 196, LI 197, DI 199, L1200, and Q1201 and E1202, of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, preferably at least one, at least two, at least three, at least four or all of the amino acid residues N 1194, SI 196, DI 199, Q1201 and E1202 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, more preferably at least one, at least two, at least three or all of the amino acid residues N1194, SI 196, DI 199 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, most preferably at least one or both of the amino acid residues SI 196 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO: 93, are indispensable for binding of the binding agents, in particular the antibodies and antibody fragments, to spike protein. In embodiments, (27) at least one, at least two, at least three, at least four, at least five, at least six, at least seven or all, amino acid residue(s) of spike protein of a Sarbecovirus or S2 subunit or HR2 domain of the Sarbecovirus spike protein corresponding to the amino acid residues N1192, N1194, SI 196, LI 197, DI 199, LI 200, Q1201 and E1202 ofthe SARS-CoV-2 spike protein as defined in SEQ ID NO:93, preferably at least one, at least two, at least three, at least four or all amino acid residue(s) corresponding to the amino acid residues N1194, SI 196, DI 199, Q1201 and E1202 ofthe SARS-CoV-2 spike protein as defined
in SEQ ID NO: 93, more preferably at least one, at least two, at least three or all amino acid residues(s) corresponding to the amino acid residues N1194, SI 196, DI 199 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, most preferably at least one or both amino acid residue(s) corresponding to the amino acid residues S 1196 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, are indispensable for binding of the binding agents, in particular the antibodies and antibody fragments, to spike protein. In particular embodiments, (28) the amino acid residues SI 196 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93 or the amino acid residues of spike protein corresponding to said amino acid residues of SI 196 and Q1201 of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, optionally the amino acid residues N1194, S 1196, D1199 and Q1201ofthe SARS-CoV-2 spike protein as defined in SEQ ID NO:93 or the amino acid residues of spike protein corresponding to said amino acid residues N1194, SI 196, DI 199 and Q1201of the SARS-CoV-2 spike protein as defined in SEQ ID NO:93, are indispensable for binding of the binding agents, in particular the antibodies and antibody fragments, to spike protein.
Assessment of the binding site may be evaluated by determining the crystal structure of a complex of the binding agent, in particular the antibody or antibody fragment, and a spike protein, or an S2 subunit or a peptide comprising a HR2 domain, for example by applying the crystal structure determination method as shown in the examples, and/or by selection and analysis of viral escape variants/mutants, for example by applying the viral escape selection method as shown in the examples, and/or by analysing hydrogen-deuterium exchange on recombinant spike protein (or S2 subunit or HR2 containing peptides) in the presence and absence of the binding agent, for example by applying the hydrogen-deuterium exchange method monitored by mass spectrometry (HDX-MS method) as shown in the examples.
Advantageously, these amino acid residues are conserved between different clades of Sarbecoviruses. in particular between clade 1, clade 2, and clade 3 Sarbecoviruses . In preferred embodiments, the binding agents, in particular the antibodies or antibody fragments, (9) do not bind to the RBD of the Sarbecovirus spike protein.
The binding agents, in particular the antibodies and antibody fragments (29) specifically bind or bind to a quaternary epitope of the spike protein. In particular, the binding agents, in particular the antibodies and antibody fragments (30) specifically bind or bind to a trimeric HR2 domain (or a trimeric S2 subunit or a trimeric spike protein). In particular, the binding agents, in particular the antibodies and antibody fragments (31) specifically bind or bind to a quaternary epitope within a
trimeric HR2 domain (or a trimeric S2 subunit or a trimeric spike protein). More particularly, the binding agents, in particular the antibodies and antibody fragments, (32) specifically bind or bind to a quaternary epitope located within two adjacent HR2 domains or helices. In particular embodiments, the binding agents, in particular the antibodies and antibody fragments, (33) specifically bind or bind to a quaternary epitope comprising or consisting of one or more interacting amino acid residues as described herein in one HR2 domain or helix as well as one or more interacting amino acid residues as described herein in an adjacent HR2 domain or helix. In particular embodiments, the binding agents, in particular the antibodies and antibody fragments, (34) specifically bind or bind to a quaternary epitope within a trimeric spike protein, wherein amino acid residues, particularly one or more interacting amino acid residues as described herein, from at least two such as two monomers of the trimeric spike protein contribute to said quaternary epitope.
As used herein, the term “quaternary epitope” refers to a conformational epitope whose structure depends upon or is enhanced by the arrangement of multiple protomers or monomers into a multimeric complex. A quaternary epitope may be located in a single protein (or monomer) of a multimeric complex; or it may span multiple protomers, being formed de novo by their interaction.
Specific binding or binding to a quaternary epitope or a multimeric protein can be assessed by evaluating binding to monomeric and/or (stabilized) multimeric protein by means of an Enzyme Linked Immunosorbent Assay (ELISA) assay, for example by applying the ELISA assay as shown in the examples. Stabilization of trimeric spike protein may be achieved by fusing the spike protein to the foldon domain of the trimeric protein fibritin from bacteriophage T4. Correlation between binding to the monomeric protein and density of the monomeric protein at elevated densities of the monomeric protein only, such as at a density of 1.0 ng/mm2 or more, preferably 1.2 ng/mm2 or more, or 1.5 ng/mm2 or more, may be indicative for specific binding or binding to a multimeric conformation of the protein. For a given density of monomeric and multimeric protein, enhanced binding to multimeric protein compared to the monomeric protein may indicate specific binding or binding to the multimeric protein.
Without wishing to be bound by any theory, upon binding to the trimeric spike protein, in particular to a quaternary epitope within the trimeric spike protein, the binding agents, in particular the antibodies and antibody fragments, described herein may stabilize the profusion conformation of the spike protein. More particularly, the binding agent may stabilise or lock the HR2 coiled-coil. As such, the binding agents may prevent the unravelling of the HR2 coiled-coil, which is considered a critical early step in the spike-controlled membrane fusion process; or the binding agents may
interfere with or block migration of the HR2 alpha helices towards the extended HR1 alpha helices, which is considered a critical step in the refolding of the spike protein from a prehairpin intermediate to a postfusion conformation; and/or the binding agents may prevent the completion of the 6 helix bundle formation, which is considered crucial for the fusion process. In embodiments, the binding agents, in particular the antibodies and antibody fragments (35) are capable of stabilizing the prefusion conformation of spike protein of a Sarbecovirus. In embodiments, the binding agents, in particular the antibodies and antibody fragments are (36) capable of stabilizing the HR2 coiled-coil. SARS-CoV-2 contains as structural proteins the spike (S) protein, the envelope (E) protein, the membrane (M) protein, and the nucleocapsid (N) protein. Furthermore, sixteen nonstructural proteins (nspl-16) have been discerned, which are involved in replication and modifying the host defense. The Nspl2 protein corresponds to a RNA-dependent RNA polymerase (RdRp).
Of specific interest in the current invention is the spike or S protein, which is a transmembrane glycoprotein forming homotrimers protruding from the viral surface and giving the virus a crownlike look. The spike protein has two subunits: SI and S2.
The SI subunit comprises an N-terminal domain (NTD), a receptor binding domain (RBD), and subdomains 1 and 2 (SD1, SD2). The SI subunit is involved in host receptor binding. The spike protein binds to human host cell receptor angiotensin-converting enzyme 2 (ACE2) via the receptor binding domain (RBD) present in the S 1 subunit.
The S2 subunit is involved in fusing the membranes of viruses and host cells and viral entry, and comprises multiple domains: an S2’ protease cleavage site (cleavage by a host protease required for fusion), a fusion peptide (FP), a heptad repeat 1 (HR1) domain, a central helix (CH) domain, a connector domain (CD), a heptad repeat 2 (HR2) domain, a transmembrane (TM) domain, and a cytoplasmic tail (CT) domain (Wang et al. (2020). Front Cell Infect Microbiol 10:587269).
The S protein normally exists in a prefusion conformation. In said prefusion conformation, SI and S2, cleaved at the S1-S2 furin cleavage site during biosynthesis, remain non-covalently bound to each other - this is different from SARS-CoV in which SI and S2 remain uncleaved. In the closed state of the S protein (PDB: 6VXX), the 3 RBD domains in the trimer do not protrude from the trimer whereas in the open state (PDB:6VYB), or “up” conformation, one of the RBD does protrude from the trimer. The S-trimer ectodomain with triangular cross-section has a length of approximately 160- Angstrom wherein the S 1 domain adopts a V-shaped form. Sixteen of the 22 N-linked glycosylation sites per protomer appear glycosylated (Walls et al. (2020) Cell 180:281-292).
The S 1 subunit of the S protein binds with ACE2 through its RBD region to promote the formation of endosomes, which triggers viral fusion activity. After S1-ACE2 binding, S is cleaved by cellular proteases, such as transmembrane protease serine subtype 2 (TMPRSS2) or endosomal cathepsins, which exposes the fusion peptide (FP) that is located in the S2 subunit. The FP inserts into the host cell membrane, thereby shortening the distance between the viral membrane and host cell membrane, the HR1 domain of the S protein is in close proximity to the host cell membrane, whereas the HR2 domain is closer to the viral membrane side. Then, HR2 folds back to HR1, whereby the two HR domains form a six-helix structure in an antiparallel format of the fusion core. The viral membrane is so pulled toward the host cell membrane and tightly binds to it, and the two membranes fuse, resulting in the release of the viral genome into the host cell (Huang et al. (2020) Acta Pharmalogica Sinica 41: 1141-1149).
The terms “spike protein”, “S” or “S protein” as used herein as synonyms refer to the spike protein of a Sarbecovirus , and can refer to specific S proteins such as SARS-CoV-2 S protein and SARS- CoV-1 S protein. The terms “spike protein” and “SARS-CoV-2 spike protein” include protein variants of Sarbecovirus or SARS-CoV-2 spike protein isolated from different Sarbecovirus or SARS-CoV-2 isolates, as well as recombinant Sarbecovirus or SARS-CoV-2 spike protein, or a fragment thereof. The terms also encompass Sarbecovirus spike protein or SARS-CoV-2 spike protein coupled to, for example, a histidine tag, mouse or human Fc, or a signal sequence.
The SARS-CoV-2 spike protein sequence can be found under/corresponds with or to Genbank Accession: QHQ82464, version QHQ82464.1; and is also defined herein as SEQ ID NO:93:
MFVFLVLLPLVSSQCVNLTTRTQLPPAYTNSFTRGVYYPDKVFRSSVLHSTQDLFLPFFSNVTWFH AIHVSGTNGTKRFDNPVLPFNDGVYFASTEKSNI IRGWI FGTTLDSKTQSLLIVNNATNVVIKVCE FQFCNDPFLGVYYHKNNKSWMESEFRVYSSANNCTFEYVSQPFLMDLEGKQGNFKNLREFVFKNID GYFKIYSKHTPINLVRDLPQGFSALEPLVDLPIGINITRFQTLLALHRSYLTPGDSSSGWTAGAAA YYVGYLQPRTFLLKYNENGTITDAVDCALDPLSETKCTLKSFTVEKGIYQTSNFRVQPTESIVRFP NITNLCPFGEVFNATRFASVYAWNRKRISNCVADYSVLYNSASFSTFKCYGVSPTKLNDLCFTNVY ADSFVIRGDEVRQIAPGQTGKIADYNYKLPDDFTGCVIAWNSNNLDSKVGGNYNYLYRLFRKSNLK PFERDISTEIYQAGSTPCNGVEGFNCYFPLQSYGFQPTNGVGYQPYRVVVLSFELLHAPATVCGPK KSTNLVKNKCVNFNFNGLTGTGVLTESNKKFLPFQQFGRDIADTTDAVRDPQTLEILDITPCSFGG VSVITPGTNTSNQVAVLYQDVNCTEVPVAIHADQLTPTWRVYSTGSNVFQTRAGCLIGAEHVNNSY ECDIPIGAGICASYQTQTNSPRRARSVASQSI IAYTMSLGAENSVAYSNNSIAIPTNFTISVTTEI LPVSMTKTSVDCTMYICGDSTECSNLLLQYGSFCTQLNRALTGIAVEQDKNTQEVFAQVKQIYKTP
PIKDFGGFNFSQILPDPSKPSKRSFIEDLLFNKVTLADAGFIKQYGDCLGDIAARDLICAQKFNGL TVLPPLLTDEMIAQYTSALLAGTITSGWTFGAGAALQIPFAMQMAYRFNGIGVTQNVLYENQKLIA NQFNSAIGKIQDSLSSTASALGKLQDVVNQNAQALNTLVKQLSSNFGAISSVLNDILSRLDKVEAE VQIDRLITGRLQSLQTYVTQQLIRAAEIRASANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQSA PHGVVFLHVTYVPAQEKNFTTAPAICHDGKAHFPREGVFVSNGTHWFVTQRNFYEPQIITTDNTFV SGNCDVVIGIVNNTVYDPLQPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRLNE VAKNLNESLIDLQELGKYEQYIKWPWYIWLGFIAGLIAIVMVTIMLCCMTSCCSCLKGCCSCGSCC KFDEDDSEPVLKGVKLHYT
Herein, the SARS-CoV-2 spike protein HR2 domain corresponds with/to amino acids 1169-1202 of SEQ ID NO:93 and as depicted hereafter (SEQ ID NO:94): ISGINASVVNIQKEIDRLNEVAKNLNESLIDLQE (SEQ ID NO:94).
Herein, the SARS-CoV-2 spike protein TM domain corresponds with/to amino acids 1214-1237 of SEQ ID NO:93.
As used herein a region of the HR2 domain “proximal to the viral membrane” refers to a region within the HR2 domain that is within 40 amino acids from the viral membrane.
The Sars-CoV-1 spike protein sequence can be found under/corresponds with or to GenBank accession NP_828851.1; and is also defined herein as SEQ ID NO:95. Herein, the SARS-CoV-1 spike protein HR2 domain corresponds with/to amino acids 1151-1184 of SEQ ID NO:95 and as depicted in SEQ ID NO:94. The amino acids and amino acid numbering referred to herein is relative to/corresponding to the SARS-CoV-2 spike protein as defined in SEQ ID NO:93; corresponding amino acids in spike proteins or spike protein fragments, domains or regions of other Sarbecoviruses can be easily determined by aligning multiple amino acid sequences.
“Angiotensin converting enzyme 2”, “ACE2”, or “ACE-2” as used herein interchangeably refers to mammalian protein belonging to the family of dipeptidyl carboxydipeptidases, and sometimes classified as EC:3.4.17.23. The genomic location of the human ACE2 gene is on chrX: 15,561,033- 15,602,158 (GRCh38/hg38; minus strand), or alternatively on chrX: 15,579,156-
15,620,271(GRCh37/hgl9; minus strand). ACE2 acts as a receptor for at least human coronaviruses SARS-CoV and SARS-CoV-2, and NL63/HCoV-NL63 (also known as New Haven coronavirus). UniProtKB identifier of human ACE2 protein: Q9BYF1. Isoform 1 (identifier: Q9BYF1-1) has been chosen as the canonical! sequence. Reference DNA sequence of the human ACE2 gene in GenBank: NC_000023.11. Reference mRNA sequences of human ACE2 in GenBank NM_001371415.1 and
NM_021804.3.
A further functional characteristic of the binding agents, in particular the antibodies and antibody fragments, described herein is that they are (10) capable of neutralizing a Sarbecovinis. in particular (11) capable of neutralizing any one or both, preferably both, of SARS-CoV-2 and SARS-CoV-1.
As used herein, a "neutralizing binding agent" or a “neutralizing antibody” (or "binding agent or antibody that is capable of neutralizing a Sarbecovinis. in particular SARS-CoV-2 and/or SARS- CoV-1") refers to a binding agent or antibody that binds to a Sarbecovinis. in particular SARS-CoV- 2 and/or SARS-CoV-1, to inhibit or suppress the ability of the Sarbecovinis. or SARS-CoV-2 or SARS-CoV-1, to initiate and/or perpetuate an infection in a host. Neutralizing binding agents or antibodies may, for example, interfere with binding of a Sarbecovinis such as SARS-CoV-2 or SARS-CoV-1 to a host receptor, in particular ACE2; and/or with viral entry, e.g. by inducing SI shedding and/or by interfering with viral fusion. At present it is not fully clear how the binding agents and antibodies according to the current invention are neutralizing, inhibiting, blocking or suppressing a Sarbecovinis infection. In certain embodiments, the binding agents, in particular the antibodies and antibody fragments, described herein (44) do not modulate or interfere with SI shedding. In certain embodiments, the binding agents, in particular the antibodies and antibody fragments, described herein (12) do not induce SI shedding. In certain embodiments, the binding agents, in particular the antibodies and antibody fragments, described herein (45) do not prevent SI shedding. In certain embodiments, the binding agents, in particular the antibodies and antibody fragments, described herein are (13) capable of inhibiting spike-mediated syncytia formation. Consequently, the binding agents, in particular the antibodies and antibody fragments, may be (14) capable of inhibiting viral fusion and, without wishing to be bound by any theory, may as such not allow the Sarbecovinis to complete the infection process into a host cell. In certain embodiments, the binding agents, in particular the antibodies and antibody fragments, described herein (46) do not prevent HR1 unfolding. In certain embodiments, the binding agents, in particular the antibodies and antibody fragments, described herein (47) do not prevent folding of HR1 onto HR2 (e.g. as during formation of a S2 6 helix bundle). Independent of their mechanism of action, the binding agents, in particular the antibodies and antibody fragments, according to the invention are capable of neutralizing a Sarbecovirus infection potently.
Neutralizing activity can be measured using a standard neutralization assay as known to one of skill in the art, including, without limitation, a pseudovirus neutralization assay and a plaque reduction test. Exemplary methods for performing such neutralization assays are described herein in the
examples. Neutralizing activity can also be evaluated by measuring one or more indicators of a Sarbecovirus , or SARS-CoV-2 or SARS-CoV-1, infection, such as syncytia formation between cells expressing a Sarbecovirus spike protein and cells expressing the Sarbecovirus receptor ACE2 .
In particular embodiments, the binding agents, in particular the antibodies and antibody fragments, are (15) capable of neutralizing a Sarbecovirus, in particular SARS-CoV-2 and/or SARS-CoV-1, with a half maximum inhibitory concentration or 50% inhibitory concentration (IC50) of 100 ng/ml or less, preferably 50 ng/ml or less or 20 ng/ml or less, more preferably 15 ng/ml or less or 10 ng/ml or less, even more preferably 1 ng/ml or less, preferably as determined in a Sarbecovirus spike protein pseudovirus neutralization assay such as a vesicular stomatitis virus JSV)-Sarbecovirus spike protein pseudovirus neutralization assay, more preferably as determined in a SARS-CoV-2 spike protein and/or SARS-CoV-1 spike protein pseudovirus neutralization assay such as a VSV-SARS- CoV-2 spike protein pseudovirus neutralization assay or a VSV-SARS-CoV-1 spike protein pseudovirus neutralization assay. In particular, the pseudovirus neutralization assay may be based on pseudotyped VSV-delG virus containing the spike protein of a Sarbecovirus such as SARS-CoV-2 spike protein, SARS-CoV-2 variant spike protein or SARS-CoV-1 spike protein. As used herein in connection with the neutralizing activity of a binding agent or antibody, “half maximum inhibitory concentration” or “IC50” refers to a quantity such as a concentration of a binding agent or antibody required for 50% neutralization of the Sarbecovirus.
In particular embodiments, the binding agents, in particular the antibodies and antibody fragments, are (16) capable of neutralizing at least one SARS-CoV-2 variant such as a SARS-CoV-2 variant comprising a mutation at position D614 (relative to the SARS-CoV-2 spike amino acid sequence as defined in SEQ ID NO:93) such as a D614G variant, in particular at least any one or more, preferably all, of SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta variant, SARS-CoV-2 Epsilon B. 1.427 variant, SARS-CoV-2 Epsilon B. 1.429 variant, SARS-CoV-2 Eta, variant, SARS-CoV-2 Iota variant, SARS-CoV-2 Kappa variant, SARS- CoV-2 Mu variant, SARS-CoV-2 Zeta variant, SARS-CoV-2 B.1.617.3 variant, SARS-CoV-2 Omicron BA.l variant SARS-CoV-2 Omicron BA.2 variant, SARS-CoV-2 Omicron BA.5 variant, SARS-CoV-2 Omicron BA.2.75.2 variant, SARS-CoV-2 Omicron BA.4.6 variant, SARS-CoV-2 Omicron BF.7 variant, SARS-CoV-2 Omicron BQ. 1. 1 variant, SARS-CoV-2 Omicron XBB variant, SARS-CoV-2 Omicron XBB. 1.5 (-G252V) variant, SARS-CoV-2 Omicron CH. 1.1 variant, SARS- CoV-2 Omicron EG.5.1 variant, SARS-CoV-2 Omicron BA.2.86.1 variant SARS-CoV-2 Omicron HV. l variant, SARS-CoV-2 Omicron XBB. 1.5 variant, SARS-CoV-2 Omicron FL. 1.5.1 variant,
SARS-CoV-2 Omicron XBB.2.3 variant, SARS-CoV-2 Omicron XBB.1.16 variant, SARS-CoV-2 Omicron HK.3 variant, SARS-CoV-2 Omicron BA.2.74 variant, SARS-CoV-2 Omicron JN.1 variant, SARS-CoV-2 Omicron KP.2.3 variant, SARS-CoV-2 Omicron KP.3 variant, SARS-CoV-2 Omicron KP.3.1.1 variant, SARS-CoV-2 Omicron KP.1.1 variant, SARS-CoV-2 Omicron LB.l variant and SARS-CoV-2 Omicron XEC variant. In particular embodiments, the binding agents, in particular the antibodies and antibody fragments described herein are characterized in that they are (17) capable of neutralizing SARS-CoV-2 Alpha variant, (78) capable of neutralizing SARS-CoV- 2 Beta variant, (67) capable of neutralizing SARS-CoV-2 Gamma variant, (79) capable of neutralizing SARS-CoV-2 Delta variant, (80) capable of neutralizing SARS-CoV-2 Epsilon B.1.427 variant, (81) capable of neutralizing SARS-CoV-2 Epsilon B.1.429 variant, (68) capable of neutralizing SARS-CoV-2 Eta variant, (69) capable of neutralizing SARS-CoV-2 Iota variant, (70) capable of neutralizing SARS-CoV-2 Zeta variant, (82) capable of neutralizing SARS-CoV-2 Kappa variant, (71) capable of neutralizing SARS-CoV-2 Mu variant, (72) capable of neutralizing SARS- CoV-2 B.1.617.3 variant, (18) capable of neutralizing SARS-CoV-2 Omicron BA.l variant, (19) capable of neutralizing SARS-CoV-2 Omicron BA.2 variant, (37) capable of neutralizing SARS- CoV-2 Omicron BA.5 variant, (38) capable of neutralizing SARS-CoV-2 Omicron BA.2.75.2 variant, (39) capable of neutralizing SARS-CoV-2 Omicron BA.4.6 variant, (40) capable of neutralizing SARS-CoV-2 Omicron BF.7 variant, (41) capable of neutralizing SARS-CoV-2 Omicron BQ.1.1 variant, (42) capable of neutralizing SARS-CoV-2 Omicron XBB variant, (43) capable of neutralizing SARS-CoV-2 Omicron XBB.1.5 (-G252V) variant, (73) capable of neutralizing SARS-CoV-2 Omicron CH.1.1 variant, (58) capable of neutralizing SARS-CoV-2 Omicron EG.5.1 variant, (59) capable of neutralizing SARS-CoV-2 Omicron BA.2.86.1 variant, (60) capable of neutralizing SARS-CoV-2 Omicron HV.l variant, (61) capable of neutralizing SARS- CoV-2 Omicron XBB.1.5 variant, (62) capable of neutralizing SARS-CoV-2 Omicron XBB.2.3 variant, (63) capable of neutralizing SARS-CoV-2 Omicron XBB.1.16 variant, (64) capable of neutralizing SARS-CoV-2 Omicron HK.3 variant, (65) capable of neutralizing SARS-CoV-2 Omicron BA.2.74 variant, (66) capable of neutralizing SARS-CoV-2 Omicron FL.1.5.1 variant, (74) capable of neutralizing SARS-CoV-2 Omicron JN.l variant, (75) capable of neutralizing SARS- CoV-2 Omicron KP.2.3 variant, (76) capable of neutralizing SARS-CoV-2 Omicron KP.3 variant, (77) capable of neutralizing SARS-CoV-2 Omicron KP.3.1.1 variant, (83) capable of neutralizing SARS-CoV-2 Omicron KP.1.1 variant, (84) capable of neutralizing SARS-CoV-2 Omicron LB.l variant and/or (85) capable of neutralizing SARS-CoV-2 Omicron XEC variant with an IC50 of 100 ng/ml or less, preferably 50 ng/ml or less or 20 ng/ml or less, more preferably 15 ng/ml or less or 10
ng/ml or less, even more preferably 1 ng/ml or less, preferably as determined in a SARS-CoV-2 variant spike pseudovirus neutralization assay such as a VSV-SARS-CoV-2 variant spike pseudovirus neutralization assay. In particular embodiments, the binding agents, in particular the antibodies and antibody fragments described herein, are characterized in that they are (86) capable of neutralizing a SARS-CoV-2 variant comprising a F456L mutation, in particular capable of neutralizing one or more, preferably all, of SARS CoV-2 Omicron FL. 1.5.1, SARS CoV-2 Omicron EG.5.1, SARS CoV-2 Omicron HK.3, SARS CoV-2 Omicron HV.l, SARS CoV-2 Omicron KP.1.1, SARS CoV-2 Omicron KP.2.3, SARS CoV-2 Omicron KP.3, SARS CoV-2 Omicron KP.3.1.1, SARS CoV-2 Omicron LB. l, and SARS CoV-2 Omicron XEC variants, with an IC50 of 100 ng/ml or less, preferably 50 ng/ml or less or 20 ng/ml or less, more preferably 15 ng/ml or less or 10 ng/ml or less, optionally as determined in a SARS-CoV-2 variant spike pseudovirus neutralization assay such as a VSV-SARS-CoV-2 variant spike pseudovirus neutralization assay. In particular embodiments, the binding agents, in particular the antibodies and antibody fragments described herein, are characterized in that they are (87) capable of neutralizing a SARS-CoV-2 variant comprising a Q493E mutation, in particular capable of neutralizing one or more, preferably all, of SARS CoV-2 Omicron KP.3, SARS CoV-2 Omicron KP.3.1.1, and SARS CoV-2 Omicron XEC variants, with an IC50 of 100 ng/ml or less, preferably 50 ng/ml or less or 20 ng/ml or less, more preferably 15 ng/ml or less or 10 ng/ml or less, optionally as determined in a SARS-CoV-2 variant spike pseudovirus neutralization assay such as a VSV- SARS-CoV-2 variant spike pseudovirus neutralization assay. In particular embodiments, the binding agents, in particular the antibodies and antibody fragments described herein, are characterized in that they are (88) capable of neutralizing a SARS-CoV-2 variant comprising F456L and Q493E mutations, in particular capable of neutralizing one ore more, preferably all, of SARS CoV-2 Omicron KP.3, SARS CoV-2 Omicron KP.3.1.1, and SARS CoV-2 Omicron XEC variants, with an IC50 of 100 ng/ml or less, preferably 50 ng/ml or less or 20 ng/ml or less, more preferably 15 ng/ml or less or 10 ng/ml or less, optionally as determined in a SARS-CoV-2 variant spike pseudovirus neutralization assay such as a VSV- SARS-CoV-2 variant spike pseudovirus neutralization assay.
The binding agents, in particular the antibodies and antibody fragments, described herein are further characterized in that they are (14) capable of inhibiting viral fusion. In particular embodiments, the binding agents, in particular the antibodies and antibody fragments, described herein are (13) capable of inhibiting spike-mediated syncytia formation, more particularly they are (20) capable of inhibiting the formation of syncytia between cells expressing a Sarbecovirus spike protein, such as SARS-CoV-
2 and/or SARS-CoV-1 spike protein, and cells expressing the Sarbocovirus host receptor, in particular ACE2 receptor.
As used herein “viral fusion” refers to fusion of a viral membrane and a host cell membrane. Viral fusion assays are well-known to the skilled person and exemplary methods for performing such methods are described herein in the examples. As will be clear to a skilled person, complete inhibition is not required and a skilled person is able to identify binding agents, antibodies and antibody fragments that significantly inhibit viral fusion or spike-mediated syncytia formation. Preferably, binding agents, in particular antibodies and antibody fragments, as described herein (21) may induce at least 50% inhibition, preferably at least 60%, at least 70%, at least 80% or at least 90% inhibition. In particular embodiments, some of the functional characteristics of a Sarbecovirus binding agent, in particular a Sarbecovirus antibody or antibody fragment, as described hereinabove are combined such as to characterize such binding agent, antibody or antibody fragment, e.g. to bind or specifically bind to the Sarbecovirus spike protein HR2 domain and to be capable of neutralizing a Sarbecovirus, in particular at least one or both of SARS-CoV-2 (such as any one or more of SARS-CoV-2 Wuhan strain, SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta variant, SARS-CoV-2 Epsilon B. 1.427 variant, SARS-CoV-2 Epsilon B.1.429 variant, SARS-CoV-2 Eta variant, SARS-CoV-2 Iota variant, SARS- CoV-2 Zeta variant, SARS-CoV-2 Kappa variant, SARS-CoV-2 Mu variant, SARS-CoV-2 B. 1.617.3 variant, SARS-CoV-2 Omicron BA. l variant, SARS-CoV-2 Omicron BA.2 variant, SARS-CoV-2 Omicron BA.5 variant, SARS-CoV-2 Omicron BA.2.75.2 variant, SARS-CoV-2 Omicron BAA.6 variant, SARS-CoV-2 Omicron BF.7 variant, SARS-CoV-2 Omicron BQ. 1.1 variant, SARS-CoV-2 Omicron XBB variant, SARS-CoV-2 Omicron XBB.1.5 variant (-G252V), SARS-CoV-2 Omicron CH. 1.1 variant, SARS-CoV-2 Omicron EG.5.1 variant, SARS-CoV-2 Omicron BA.2.86.1 variant SARS-CoV-2 Omicron HV. l variant, SARS-CoV-2 Omicron XBB.1.5 variant, SARS-CoV-2 Omicron XBB.2.3 variant, SARS-CoV-2 Omicron XBB. 1.16 variant, SARS- CoV-2 Omicron HK.3 variant, SARS-CoV-2 Omicron BA.2.74 variant, SARS-CoV-2 Omicron FL. 1.5.1 variant, SARS-CoV-2 Omicron JN. 1 variant, SARS-CoV-2 Omicron KP.2.3 variant, SARS-CoV-2 Omicron KP.3 variant, SARS-CoV-2 Omicron KP.3.1.1 variant, SARS-CoV-2 Omicron KP.1.1 variant, SARS-CoV-2 Omicron LB.l variant and SARS-CoV-2 Omicron XEC variant) and SARS-CoV-1, preferably to be capable of neutralizing the Sarbecovirus with a 50% inhibitory concentration (IC50) of 100 ng/ml or less, preferably 50 ng/ml or less or 20 ng/ml or less, more preferably 15 ng/ml or less or 10 ng/ml or less, even more preferably 1 ng/ml or less, as determined in a vesicular stomatitis virus JSV)-Sarbecovirus spike protein pseudovirus
neutralization assay. Such binding agent, antibody or antibody fragment may further be characterized to be capable of inhibiting spike-mediated syncytia formation between cells expressing the Sarbecovirus spike protein and cells expressing the angiotensin-converting enzyme 2 (ACE2) receptor and/or to be capable of inhibiting viral fusion; and/or by not binding a Middle East respiratory syndrome coronavirus (MERS-CoV).
The binding agents, in particular the antibodies and antibody fragments, described herein are further characterized in that (48) they have an advantageously low isoelectric point, in particular a lower isoelectric point compared with the ISVD defined by SEQ ID NO:2. In particular embodiments, (49) the pl of the binding agent, in particular the antibody or antibody fragment, is from 7.5 to 9.5, or from 7.6 to 9.4, or from 7.7 to 9.3, or from 7.8 to 9.3, or from 7.9 to 9.2, preferably from 7.5 to 9.1, or from 7.6 to 9.1, or from 7.7 to 9.1, or from 7.7 to 9.1, or from 7.8 to 9.1, or from 7.9 to 9.1.
As used herein, “pl” refers to the “isoelectric point” of a molecule, in particular a binding agent, antibody or antibody fragment as described herein, and is the pH at which a particular molecule or surface carries no net electrical charge. Antibodies with lower pl values may have longer serum halflives. Methods for determining the isoelectric point of a protein are well known to those of one skilled in the art. By way of example, many suitable computer programs for calculating the pl of a protein are generally known in the art, such as EMBOSS iep software, written by Alan Bleasby, available at HGMP-RC, Genome Campus, Hinxton, Cambridge CB10 1SB, UK.
In embodiments, the binding agents, in particular the antibodies and antibody fragments, (50) exhibit advantageously low non-specific binding, in particular reduced non-specific binding compared with the ISVD defined by SEQ ID NO:2, in particular (51) low non-specific binding to mammalian cells, more particularly human cells, such as reduced non-specific binding to mammalian cells, more particularly human cells, compared with the ISVD defined by SEQ ID NO:2.
As used herein, with “non-specific” binding or “off-target” of a molecule, in particular a binding agent antibody or antibody fragment as described herein, is meant herein the ability of the molecule to bind to a non-specific target. There are several tests that can be performed to determine if an antibody has any cross-reactivity with other antigens, such as, for example but without limitation, the binding to HEK293 cells using flow cytometry as described in the experimental section.
In embodiments, the binding agents, in particular the antibodies and antibody fragments, (52) exhibit advantageously low immunogenicity, in particular reduced immunogenicity compared with the ISVD defined by SEQ ID NO:2.
With “immunogenicity” of a molecule, in particular a binding agent, antibody or antibody fragment as described herein, is meant herein the ability of the molecule to elicit an (undesired) immune response (humoral or cellular) when administered to a recipient. Immunogenicity of a molecule, in particular a binding agent, antibody or antibody fragment, can be determined by in silico analysis as described in the experimental section.
In embodiments, the binding agents, in particular the antibodies and antibody fragments, (53) exhibit advantageously low hydrophobicity, in particular reduced hydrophobicity compared with the ISVD defined by SEQ ID NO:2.
In embodiments, the binding agents, in particular the antibodies and antibody fragments, (54) exhibit advantageously low positive charge patches, in particular reduced positive charge patches compared with the ISVD defined by SEQ ID NO:2. In particular embodiments, (55) the retention time of the binding agents, in particular the antibodies and antibody fragments, is 15 min or lower, preferably between 5 and 15 min, or between 6 and 14 min, as determined by strong cation exchange hydrophilic interaction chromatography (SCX), in particular SCX using the conditions specified in the examples section, more particularly SCX using a MAbPac SCX- 10 column, a mobile phase A consisting of 20mM MES pH 5.6 and a mobile phase B consisting of 20 mM HEPES and 120 mM NaCl pH 8.0 and using a linear gradient from 100% A to 100% B in 18 min.
In embodiments, the binding agents, in particular the antibodies and antibody fragments, are (56) advantageously stable, in particular exhibit (57) improved protein stability compared to the ISVD defined by SEQ ID NO:2.
In certain embodiments, the antibody comprises one or more ISVDs as described herein (or variants thereof as described herein) wherein the one or more ISVD (or variant thereof as described herein) is bound or fused to an Fc domain.
An “Fc domain” as used herein refers to the fragment crystallizable region (Fc region) of a conventional antibody, which is the tail region known to interact with cell surface receptors called Fc receptors and some proteins of the complement system. Said Fc domain is composed of two identical protein fragments, derived from the second and third constant domains of the antibody's two heavy chains. All conventional antibodies comprise an Fc domain, hence, the Fc domain may be an Fc domain derived from or as a variant of the IgG, IgA or IgD antibody Fc regions, even more specifically derived from an IgGl, IgG2 or IgG4 antibody Fc region. For example, the hinge region of IgG2, may be replaced by the hinge of human IgGl to generate ISVD fusion constructs, and vice versa. In addition, Fc variants with known half-life extension may be used such as the
M257Y/S259T/T261E variant (also known as YTE), the LS variant (M428L combined with N434S), the DHS variant (comprising L309D, Q311H and N434S substitutions) or the LA variant (comprising M428L and N434A substitutions). These Fc variants are known to enhance FcRn binding affinity at an endosomal acidic pH and dissociation at physiological pH, thereby prolonging the circulating halflife of the antibody or antibody fragment and mediating half-life extension. Preferably, human Fc domains or humanized Fc domains may be used. Humanized forms, include but are not limited to the IgG humanization variants known in the art, such as C-terminal deletion of Lysine, alteration or truncation in the hinge region, LALA (L234A and L235A) or LALAPG (L234A, L235A, and P329G) mutations, among other substitutions in the IgG sequence.
In particular embodiments, one or more ISVDs (or variants thereof) as described herein are fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NOs: 82-86, as depicted hereafter:
WT Fc domain:
DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLP PSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVMHEALHNHYTQKSLSLSPG ( SEQ ID NO : 82 )
Fc domain comprising the LS substitutions:
DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLP PSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVLHEALHSHYTQKSLSLSPG ( SEQ ID NO : 83 )
Fc domain comprising the YTE substitutions:
DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLYITREPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLP PSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVMHEALHNHYTQKSLSLSPG ( SEQ ID NO : 84 )
Fc domain comprising the DHS substitutions:
DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVDHHDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLP
PSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ
QGNVFSCSVMHEALHSHYTQKSLSLSPG ( SEQ ID NO : 85 )
Fc domain comprising the LA substitutions:
DKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLP PSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVLHEALHAHYTQKSLSLSPG ( SEQ ID NO : 86 )
The term “fused to”, as used herein interchangeably with “connected to”, “conjugated to”, “ligated to” refers in one aspect to “genetic fusion”, e.g., by recombinant DNA technology, as well as to “chemical and/or enzymatic conjugation” resulting in a stable covalent link between two nucleic acid molecules. The same applies for the term “inserted in”, wherein a fragment of one nucleic acid may be inserted in a second nucleic acid molecule by fusing or ligating the two sequences genetically, enzymatically or chemically. Peptides or polypeptides can likewise be fused or connected to one another, such as via peptide bonds or via linking one peptide to a side chain of an amino acid in a second peptide.
Linkers may be used to fuse an ISVD, such as a herein identified ISVD (or variant thereof as described herein), to an Fc domain such as the human IgGl Fc domain or the LS variant thereof, or the YTE variant thereof, or the DHS variant thereof, or the LA variant thereof or an IgG2 Fc domain, preferably an Fc domain comprising or consisting of the sequence set forth in any of SEQ ID NO: 82- 86. A non-limiting example of a linker comprises a Gly-Ser linker such as (G4S)n, with n=l-6 (SEQ ID NO: 87), preferably 2-3 (SEQ ID NO: 88-89).
In certain embodiments, the antibody comprising one or more ISVDs as described herein (or variants thereof as described herein) is in a “multivalent” and/or “multispecific” form formed by binding, e.g. chemically or by recombinant DNA techniques, together two or more identical or variant monovalent ISVDs (or variants or humanized forms thereof as described herein).
Non-limiting examples of multivalent constructs include “bivalent” constructs, “trivalent” constructs, “tetravalent” constructs, and so on, respectively, comprising two, three or four ISVDs. The ISVDs comprised within a multivalent construct may be identical or different. The term “multispecific antibody” as used herein specifically refers to a multivalent antibody wherein at least one of the two or more ISVDs has a different specificity. Non-limiting examples of multi-specific constructs include “bi-specific” constructs, “tri-specific” constructs, “tetra-specific” constructs, and
so on. To illustrate this further, any multivalent and multi-specific (as defined herein) antibody of the invention may be directed against two or more different antigens, for example against a Sarbecovirus and one as a half-life extension against Serum Albumin or Staphylococcal protein A (SpA) and/or against two or more different parts of a particular antigen, for example against two or more different parts, regions, subunits or domains of a Sarbecovirus spike protein.
In particular embodiments, an antibody, in particular a multivalent and/or multispecific antibody, may comprise one or more binding agent, such as ISVD(s), as described herein (or variants thereof as described herein), and one or more binding agents, such as ISVD(s), capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD). Non-limiting examples of ISVDs capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD) are described in WO 2021/156490 Al, WO 2022/167666 Al and WO 2022/238550 Al. Advantageously, the combination of at least two ISVDs capable of binding Sarbecovirus spike protein through interaction at 2 different regions of the spike protein, in particular the S2 subunit, more particularly the HR2 domain, and the RBD, in such multivalent and/or multispecific antibody may result in crossreactivity and potent prohibition of infection by Sarbecoviruses. and may further allow for reducing the risk to escape mutant virus emergence.
In certain further embodiments, the one or more ISVDs capable of binding to a Sarbecovirus spike protein RBD are capable of binding to or competing for the VHH72 epitope (or the epitope specifically bound by VHH72). The VHH72 epitope has been described in Wrapp et al. (2020, Cell 184: 1004-1015; WO 2021/156490 Al and WO 2022/238550 Al). An ISVD capable of competing for the VHH72 epitope refers to an ISVD that competes with VHH72 for binding to the spike protein as depicted in SEQ ID NO:93, or the RBD. With ‘competing’ is meant that the binding ofVHH72 to the spike protein as depicted in SEQ ID NO:93, or the RBD, is reduced with at least 30%, or at least 50%, or preferably at least 80% in strength in the presence of an ISVD capable of competing for the VHH72 epitope. In embodiments, an ISVD capable of binding to or competing for the VHH72 epitope may be characterized in that (i) it competes for human receptor (ACE-2 in the case of SARS- CoV-1 and -2) binding upon interaction to the RBD, and/or (ii) is not competing with an ISVD capable of binding to or competing with a VHH3.117 epitope as defined herein. Non-limiting examples of ISVDs capable of binding to or competing for the VHH72 epitope include VHH72 family members (including VHH72 (SEQ ID NOVO), VHH2.50, VHH3.17, VHH3.77, VHH3.115, VHH3.144 and VHHBE4), and variants, including VHH72(S56A), and humanized forms thereof; VHH3.83 family members (including VHH3.83 (also referred to as VHH83 herein) (SEQ ID
NO:91)) and variants and humanized forms thereof; VHH3.38 family members and variants and humanized forms thereof; VHH3.55 family members and variants and humanized forms thereof; VHH3.36 family members and variants and humanized forms thereof; VHH3.149 family members and variants and humanized forms thereof; and VHH3.29 family members and variants and humanized forms thereof, as described in WO 2021/156490 Al and WO 2022/238550 Al.
In certain further embodiments, the one or more ISVDs capable of binding to a Sarbecovirus spike protein RBD are capable of binding to or competing for the VHH3.117 epitope (or the epitope specifically bound by VHH3. 117). The VHH3. 117 epitope has been described in WO 2022/167666 Al. I An ISVD capable of competing for the VHH3.117 epitope refers to an ISVD that competes with VHH3. 117 (SEQ ID NO:92) for binding to the spike protein as depicted in SEQ ID NO:93, or the RBD. With ‘competing’ is meant that the binding of VHH3.117 to the spike protein as depicted in SEQ ID NO:93 is reduced with at least 30%, or at least 50%, or preferably at least 80% in strength in the presence of an ISVD capable of competing for the VHH3.117 epitope. In embodiments, an ISVD capable of binding to or competing for the VHH3.117 epitope may be characterized in that (i) it does not inhibit binding of the RBD with the human receptor (ACE-2 in the case of SARS-CoV-1 and -2), meaning that it allows binding of the receptor and the Sarbecovirus RBD when the ISVD itself is bound to the Sarbecovirus RBD, or alternatively, that the ISVD itself can bind to a Sarbecovirus RBD to which the receptor is bound, and/or (ii) is not competing with an ISVD capable of binding to or competing for the VHH72 epitope as defined herein. Non-limiting examples of ISVDs capable of binding to or competing for the VHH3.117 epitope include VHH3.117 family members (including VHH3.117 (SEQ ID NO:92), 3.42, 3.92, 3.94, 3.180) and variants and humanized forms thereof (as described in WO 2022/167666 Al); VHH3.89 family members and variants and humanized forms thereof (as described in WO 2021/156490); VHH3_183 family members and variants and humanized forms thereof; and VHH3C 80 family members and variants and humanized forms thereof (as described in WO 2022/238550 Al).
In yet other further embodiments, the antibody, in particular the multivalent and/or multispecific antibody, comprises more than one ISVD capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD), wherein at least one ISVD is capable of binding to or competing for the VHH72 epitope as defined herein, and wherein at least one ISVD is capable of binding to or competing for the VHH3.117 epitope as defined herein. Advantageously, the combination of at least two non-competing RBD targeting ISVDs (capable of binding the RBD of the spike protein through interaction at 2 non-competing, different regions of the RBD) and at least one S2 targeting ISVD in
the antibody results in cross-reactivity and potent prohibition of infection by Sarbecoviruses , which advantageoulsy allows for further reducing the risk to mutational escape.
Multivalent antibodies as described herein may be formed e.g. by connecting, such as chemically or by recombinant DNA techniques, the two or more ISVDs directly or via a linker, and/or through fusing (each of) the two or more ISVDs with an Fc domain.
For example, a single ISVD (or variant thereof) as described herein may be fused e.g. at its C- terminus to an Fc domain, such as an Fc domain comprising or consisting of the sequence set forth in any one of SEQ ID NOs: 82-86, resulting in a Sarbecovirus antibody of bivalent format wherein two of said ISVDs form a heavy chain only antibody-type molecule through disulfide bridges in the hinge region of the Fc part.
In particular embodiments, the binding agent, in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in any one of SEQ ID NO:3-26 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO: 83, such as a binding agent, in particular a bivalent antibody, comprising or consisting of the sequence set forth in any one of SEQ ID NO:96-119. In further embodiments, the binding agent, in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in any one of SEQ ID NO:3-26 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in any one of SEQ ID NO: 82, 84, 85, or 86.
In particular embodiments, the binding agent, in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO: 82.
In particular embodiments, the binding agent, in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO:83.
In particular embodiments, the binding agent, in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO: 84.
In particular embodiments, the binding agent, in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO:85.
In particular embodiments, the binding agent, in particular a bivalent antibody, comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof fused to an Fc domain comprising or consisting of the sequence set forth in SEQ ID NO: 86.
In particular embodiments, one or more ISVDs as described herein (or variants thereof as described herein) are linked, fused or connected directly or via a linker to one or more ISVDs capable of binding to . Sa becovirus spike protein RBD as defined herein. Non-limiting examples of suitable linkers for linking the ISVDs include peptide linkers such as a (G4S)n, wherein n=l, 2, 3, 4, 5 or 6. Such multispecific binding agents may also be referred to herein as “head-to-tail fusions”.
In further embodiments, the C-terminus of a head-to-tail fusion as described herein may be fused, e.g. by a linker, to an Fc domain, which construct upon expression in a host forms a multivalent and/or multispecific antibody through disulfide bridges in the hinge region of the Fc part. Accordingly, in particular embodiments, one or more ISVDs as described herein (or variants thereof as described herein) are linked, fused or connected directly or via a linker to one or more ISVD capable of binding to a Sarbecovirus spike protein RBD to form a multispecific binding agent or construct and said multispecific binding agent or construct is fused to an Fc domain. In preferred embodiments, the antibody comprises a bispecific binding agent or construct fused to an Fc domain, wherein said bispecific binding agent or construct comprises one ISVD as described herein (or a variant thereof as described herein) linked, fused or connected directly or via a linker to one ISVD capable of binding to a Sarbecovirus spike protein RBD as described herein. For example, a binding agent, in particular a bispecific antibody, may comprise an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof, linked, fused or connected directly or via a linker to an ISVD capable of binding to a Sarbecovirus spike protein RBD as described herein. Such multispecific antibody, in particular bispecific antibody, may also be referred to herein as “VHH- VHH-Fc fusion”.
In particular embodiments, the antibody comprises a trispecific binding agent or construct fused to an Fc domain, wherein said trispecific binding agent or construct comprises one ISVD as described herein (or a variant thereof as described herein), one ISVD capable of binding to or competing with the VHH3.117 epitope as described herein, and one ISVD capable of binding to or competing with the VHH72 epitope as described herein, wherein said ISVDs are linked, fused or connected directly or via a linker to each other, in any order. For example, a binding agent, in particular a trispecific binding agent, may comprise an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof, an ISVD capable of binding to or competing with the VHH3. 117 epitope
as described herein, and an ISVD capable of binding to or competing with the VHH72 epitope as described herein, wherein said ISVDs are linked, fused or connected directly or via a linker to each other, in any order. Such multispecific antibody, in particular trispecific antibody, may also be referred to herein as “VHH-VHH-VHH-Fc fusion”.
In other particular embodiments, one or more ISVDs as described herein (or variants thereof as described herein) are fused to the N-terminus of an Fc domain, and one or more ISVDs capable of binding to a Sarbecovirus spike protein RBD are fused to the C-terminus of the Fc domain, or one or more ISVDs as described herein (or variants thereof as described herein) are fused to the C- terminus of an Fc domain, and one or more ISVDs capable of binding to a Sarbecovirus spike protein RBD are fused to the N-terminus of the Fc domain. In particular embodiments, the antibody comprises one ISVD as described herein (or a variant thereof as described herein) fused to the N- terminus of an Fc domain and one ISVD capable of binding to a Sarbecovirus spike protein RBD fused to the C-terminus of the Fc domain, or the one ISVD as described herein (or a variant thereof as described herein) is fused to the C-terminus of the Fc domain and the one ISVD capable of binding to a Sarbecovirus spike protein RBD is fused to the N-terminus of the Fc domain. For example, a binding agent, in particular a bispecific or multispecific antibody, may comprise an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof, fused to the N-terminus of an Fc domain as described herein and an ISVD capable of binding to a Sarbecovirus spike protein RBD fused to the C-terminus of the Fc domain; or the ISVD comprising or consisting of the sequence set forth in SEQ ID NO:25 or a variant thereof may be fused to the C-terminus of the Fc domain and the ISVD capable of binding to a Sarbecovirus spike protein RBD fused to the N-terminus of the Fc domain. Such multispecific antibody may also be referred to herein as “VHH-Fc-VHH fusions” or “moonlander”.
Multivalent or multi-specific antibodies as described herein may have (or be engineered and/or selected for) increased avidity and/or improved selectivity for the desired Sarbecovirus interaction, and/or for any other desired property or combination of desired properties that may be obtained by the use of such multivalent and/or multi-specific antibodies.
In particular embodiments, the binding agents, in particular the multivalent antibodies described herein, more particularly the multivalent antibodies comprising an Fc domain described herein, even more particularly the multivalent antibodies comprising an Fc domain as defined by any one of SEQ ID NO: 83-86, bind FcRn with an affinity (KD) below 300 nM, preferably below 200 nM, more preferably below 100 nM such as from 30 nM to 100 nM or from 40 nM to 100 nM, even more
preferably from 30 nM to 80 nM or from 40 nM to 80 nM, most preferably from 30 nM to 70 nM or from 40 nM to 70 nM or from 40 nM to 65 nM, when measured at pH 6.0 by surface plasmon resonance (SPR).
The pharmacokinetic properties of binding agents, in particular multivalent antibodies described herein, more particularly multivalent antibodies comprising an Fc domain described herein, may be influenced by their ability to interact with FcRn. The FcRn is expressed on innate immune cells such as macrophages but also on endothelial cells and trophoblasts of the placenta. The FcRn binds to IgG Fc domain at the junction of the CH2 and CH3 domains in a pH dependent manner. IgG is endocytosed by cells where it can be shuttled to lysosomes or recycled back to the cell surface. Binding of IgG to FcRn at low pH (pH < 6.5) in the endosomes allows the antibody to be trafficked with the FcRn back to the cell surface. At the physiologic pH of the extracellular environment IgG has weak affinity for FcRn which results in its release from the FcRn back into circulation. The high affinity of IgG to FcRn contributes to the long half-life of antibodies. Introduction of specific mutations in IgGl Fc such as LS, LA, DHS and YTE as described elsewhere herein results in a decrease in the dissociation rate and an improvement in binding affinity between Fc and human FcRn at pH6.
In particular embodiments, the binding agents, in particular the multivalent and/or multi-specific antibodies described herein, more particularly the multivalent and/or multi-specific antibodies comprising an Fc domain described herein, have antibody-dependent cell-mediated cytotoxicity (ADCC) activity. More particularly, the binding agents, in particular the multivalent and/or multispecific antibodies described herein, more particularly the multivalent and/or multi-specific antibodies comprising an Fc domain described herein, are capable of inducing ADCC on target cells expressing a Sarbecovirus spike protein. "Antibody-dependent cell-mediated cytotoxicity" or "ADCC" refers to a form of cytotoxicity in which an antibody binds to certain cytotoxic cells (such as NK cells, neutrophils, and macrophages). The secretion of Ig on the Fey receptor enables these cytotoxic effector cells to specifically bind to the target cell carrying the antigen, and then kill the target cell using, for example, a cytotoxin. In order to evaluate the ADCC activity of an antibody of interest, an in vitro ADCC assay can be performed.
In yet another aspect, the invention provides nucleic acid molecules such as isolated nucleic acids, (isolated) chimeric gene constructs, expression cassettes, and RNA-based constructs comprising a polynucleotide sequence, such as a coding sequence, that is encoding the polypeptide portion of a polypeptidic or polypeptide Sarbecovirus binding agent, in particular an antibody or antibody
fragment, as identified herein, more particularly an ISVD (or a variant thereof) as described herein, or a functional part thereof. It is particularly envisaged that the nucleic acid sequences encode the polypeptide portion of a polypeptidic or polypeptide Sarbecovirus binding agent, antibody or antibody fragment, or ISVD with all the features and variations described herein, mutatis mutandis. Thus, the encoded binding agent, antibody or antibody fragment, or ISVD, is in essence as described herein.
“Nucleic acid(s)” or “nucleic acid molecule(s)” as used herein refers to a polymeric form of nucleotides of any length, either ribonucleotides or deoxyribonucleotides; the sequential linear arrangement of the nucleotides together resulting in/forming the “nucleotide sequence”, “DNA sequence”, or “RNA sequence”. This term refers only to the primary structure of the molecule. Thus, this term includes double- and single-stranded DNA, and RNA. It also includes known types of modifications, for example, methylation, “caps”, and substitution of one or more of the naturally occurring nucleotides with an analog. Modifications to nucleic acids can be introduced at one or more levels: phosphate linkage modification (e.g. introduction of one or more of phosphodiester, phosphoramidate or phosphorothioate bonds), sugar modification (e.g. introduction of one or more of LNA (locked nucleic acids), 2'-O-methyl, 2'-O-methoxy-ethyl, 2’-fluoro, S-constrained ethyl or tricyclo-DNA) and/or non-ribose modifications (e.g. introduction of one or more of phosphorodiamidate morpholines or peptide nucleic acids).
By "nucleic acid construct" it is meant a nucleic acid molecule that has been constructed in order to comprise one or more functional units not found together in nature, thus having a nucleotide sequence not found in nature (non-native nucleotide sequence). Examples include circular, linear, doublestranded, extrachromosomal DNA molecules (plasmids), cosmids (plasmids containing COS sequences from lambda phage), viral genomes comprising non-native nucleic acid sequences, RNA or mRNA constructs, and the like.
A “coding sequence” is a nucleotide sequence that can be transcribed into mRNA and/or translated into a polypeptide when placed under the control of appropriate (gene) regulatory sequences. The boundaries of the coding sequence are determined by a translation start codon at the 5 '-terminus and a translation stop codon at the 3'-terminus. A coding sequence can include, but is not limited to mRNA, cDNA, recombinant nucleotide sequences or genomic DNA, while introns may be present as well under certain circumstances.
In an RNA or mRNA construct a (poly)peptide coding sequence may be included such that it can be translated by the cellular protein translation machinery. In aforementioned constructs a (poly)peptide
coding sequence will be typically preceded by an in-frame translation initiation codon and followed by a translation termination codon, to facilitate proper translation. Preferably, the coding sequence in aforementioned constructs is operably linked to a secretion signal sequence.
Particularly disclosed herein is an mRNA or mRNA construct encoding the polypeptide portion of a polypeptidic or polypeptide Sarbecovirus binding agent, in particular an antibody or antibody fragment, as identified herein, more particularly an ISVD (or a variant thereof) as described herein, or a functional part thereof. In embodiments, the mRNA or mRNA construct comprises (i) a 5'- untranslated region (5’ UTR); (ii) a coding sequence encoding a polypeptidic or polypeptide Sarbecovirus binding agent, an antibody or antibody fragment or an ISVD (or a variant thereof) described herein, or a functional part thereof; (iii) a stop codon; and (iv) a 3 '-untranslated region (3' UTR). In preferred embodiments, a secretion signal sequence is operably linked to the coding sequence.
In particular embodiments, the mRNA further comprises a 5' terminal cap. A 5' cap protects mRNA from RNase degradation. Moreover, the translation system in the cytoplasm recognizes the cap structure through the cap-binding protein (CBP). This helps the small ribosomal subunit bind to mRNA and recognize the start codon to initiate translation. Advantageously, introducing a 5’ cap structure at the 5' end of a mRNA or mRNA construct described herein can enhance its translation, improve its stability, reduce its immunogenicity, and/or extend its half-life. The 5' caps can be natural or cap analogs. Cap analogs differ from natural (i.e., endogenous, wild-type or physiological) 5'-caps in their chemical structure, while retaining cap function. Non-limiting examples of cap structures that can be used herein include a CapO, Capl, Anti -Reverse Cap Analog (ARC A), inosine, N1 -methylguanosine, 2'-fluoro-guanosine, 7-deaza-guanosine, 8-oxo-guanosine, 2-amino-guanosine, LNA- guanosine, 2-azidoguanosine, Cap2, Cap4, 5' methylG cap, or an analog thereof as known to the skilled person. Caps may be chemically (i.e., non-enzymatically) or enzymatically synthesized and enzymatically linked to a nucleic acid molecule, in particular an mRNA or mRNA construct as described herein.
In particular embodiments, the mRNA or mRNA construct further comprises a polyA tail. The addition of a polyA tail to the 3 ’ end of the mRNA or mRNA construct can advantageously increase its stability and/or reduce its immunogenicity. The length of the polyA tail can also regulate the halflife of the mRNA or mRNA construct in vivo. In certain embodiments, the polyA tail may be at least about 10, at least about 20, at least about 30, at least about 40, at least about 50, at least about 60, at least about 70, at least about 80, at least about 90 nucleotides in length, or at least about 100
nucleotides in length. For example, the polyA tail may be about 10 to about 200, about 20 to about 180, about 50 to about 160, about 70 to about 140, or about 80 to about 120 nucleotides in length. A polyA tail may be added to the 3 ’ end of an mRNA or mRNA construct as described herein through polyadenylation directly at the end of the mRNA or mRNA construct by a polyA polymerase, or by RNA ligation, wherein a polyA tail is directly linked to the 3 ’ end of the mRNA or mRNA construct.
In further embodiments, the mRNA or mRNA construct comprises (i) a 5’ cap, (ii) a 5 '-untranslated region (5’ UTR); (iii) a coding sequence encoding a polypeptidic or polypeptide Sarbecovirus binding agent, an antibody or antibody fragment or an ISVD (or a variant thereof) described herein, or a functional part thereof; (iv) a stop codon; (v) a 3 '-untranslated region (3' UTR), and (vi) a polyA tail.
Preferably, the mRNA is modified mRNA. As used herein, the terms “modified messenger RNA” or “modified mRNA” refer to mRNA polynucleotides that include naturally occurring and/or non- naturally occurring modifications, for example, of a sugar, a nucleobase, and/or an intemucleoside linkage (e.g., to a linking phosphate, to a phosphodiester linkage, or to the phosphodiester backbone). Non-limiting examples of sugar modifications and phosphate linkage modifications have been described elsewhere herein. Commonly used modified nucleobases include, but are not limited to, for example, pseudouracil (y), N1 -methylpseudouracil (ml\|/), 1 -ethylpseudouracil, 2-thiouracil (s2U), 4 ’-thiouracil, 5 -methylcytosine, 5 -methyluracil, 5 -methoxyuracil, and any combination thereof. The modification may be introduced with chemical synthesis or with a polymerase enzyme at the terminal of the mRNA or mRNA construct or anywhere else in the mRNA or mRNA construct.
With a “chimeric gene” or “chimeric construct” or “chimeric gene construct” is interchangeably meant a recombinant nucleic acid sequence in which a (gene) promoter or regulatory nucleic acid sequence is operably or operatively linked to, or associated with, a nucleic acid sequence of interest that codes for an RNA (e.g. a coding sequence, an shRNA, etc.), such that the regulatory nucleic acid sequence is able to regulate transcription or expression of the nucleic acid of interest. The operable or operative linkage in a chimeric gene between the regulatory nucleic acid sequence and the nucleic acid sequence of interest is not found in nature.
An "expression cassette" comprises any nucleic acid construct capable of directing the expression of a gene/coding sequence of interest, which is operably linked to a (gene) promoter. Expression cassettes are generally DNA constructs preferably including (5’ to 3’ in the direction of transcription): a (gene) promoter region, a polynucleotide sequence of interest with a transcription initiation region, and a termination sequence including a stop signal for RNA polymerase and a
polyadenylation signal; all these elements being operably or operatively linked meaning that all of these regions should be capable of operating (being expressed) in a cell, such as prokaryotic (e.g. bacterial) or eukaryotic (e.g. mammalian, yeast, insect, fungal, plant, algal) cells, when transformed into that cell. The promoter region comprising the transcription initiation region, which preferably includes the RNA polymerase binding site, and the polyadenylation signal may be native to the cell to be transformed, may be derived from an alternative source, or may be synthetic, as long as it is functional in the cell. Such expression cassettes can be constructed in e.g. a “vector” or “expression vector” (linear or circular nucleic acids, plasmids, cosmids, viral vectors, phagemids, etc.).
In particular embodiments, the nucleic acid molecule may be an mRNA or mRNA construct.
The present invention also provides a vector including the above-mentioned nucleic acid molecule inserted therein.
The term “vector”, “vector construct”, “expression vector”, “recombinant vector” or “gene transfer vector”, as used herein, is intended to refer to a nucleic acid molecule capable of carrying another nucleic acid molecule to which it has been linked.
Said vectors may include a cloning or expression vector, as well as a delivery vehicle such as a viral, lentiviral or adenoviral vector. Expression vectors may comprise plasmids as well as viral vectors and generally contain a desired coding sequence and appropriate DNA sequences necessary for the expression of the operably linked coding sequence in a particular host organism (e.g., bacteria, yeast, plant, insect, or mammal) or in in vitro expression systems. In particular, an expression vector as described herein may comprise a nucleic acid molecule as described herein comprising a nucleic acid sequence encoding an antibody or an antigen-binding fragment as described herein operably linked to at least one regulatory sequence. Regulatory sequences are selected to direct the expression of the protein of interest, in particular the antibody or antigen-binding fragment, in a suitable host cell, and include promoters, enhancers, and other expression control elements as known to the skilled person. Hence, in embodiments, the vector includes a promoter for driving expression of the nucleic acid of interest, optionally a nucleic acid sequence encoding a signal peptide that secretes the antibody or antigen-binding fragment, and optionally a nucleic acid sequence encoding a terminator. When the expression vector is manipulated in a production strain or cell line, the vector may or may not be integrated into the genome of the host cell when introduced into the host cell. Cloning vectors are generally used to engineer and amplify a certain desired DNA fragment. Thus, a cloning vectors may contain origin of replication that matches the cell type specified by the cloning vector, and may lack functional sequences needed for expression of the desired DNA fragments. Preferably, the vector
contains one or more selection markers. The choice of the selection markers may depend on the host cells of choice, although this is not critical to the present invention as is well known to persons skilled in the art. The construction of expression vectors for use in transfecting cells is also well known in the art, and thus can be accomplished via standard techniques (see, for example, Sambrook, Fritsch, and Maniatis, in: Molecular Cloning, A Laboratory Manual, Cold Spring Harbor Laboratory Press, 1989; Gene Transfer and Expression Protocols, pp. 109-128, ed. E. J. Murray, The Humana Press Inc., Clifton, N.J.), and the Ambion 1998 Catalog (Ambion, Austin, Tex.).
More particular, said vector may include any vector known to the skilled person, including any suitable type, but not limited to, for instance, plasmid vectors, cosmid vectors, phage vectors, such as lambda phage, viral vectors, even more particular a lentiviral, adenoviral, AAV or baculoviral vectors, or artificial chromosome vectors such as bacterial artificial chromosomes (BAC), yeast artificial chromosomes (YAC), or Pl artificial chromosomes (PAC). The choice of the vector may bee dependent amongst others on the nature of the host cell of choice.
One further aspect of the invention provides for a host cell comprising an antibody or antigen-binding fragment thereof, such as an ISVD (or variant thereof) of an antibody or antigen-binding fragment, or part thereof, as described herein. The host cell may therefore comprise the nucleic acid molecule encoding said antibody or antigen-binding fragment. Host cells can be either prokaryotic or eukaryotic. The host cell may also be a recombinant host cell, which involves a cell which has been genetically modified to contain an isolated nucleic acid molecule encoding the antibody or antigenbinding fragment of the invention. Representative host cells that may be used to produce said antibodies or antigen-binding fragments such as ISVDs, include, but are not limited to, bacterial cells, yeast cells, plant cells and animal cells. Bacterial host cells suitable for production of the antibodies or antigen-binding fragment of the invention include Escherichia spp. cells, Bacillus spp. cells, Streptomyces spp. cells, Erwinia spp. cells, Klebsiella spp. cells, Serratia spp. cells, Pseudomonas spp. cells, and Salmonella spp. cells. Yeast host cells suitable for use with the invention include species within Saccharomyces, Schizosaccharomyces, Kluyveromyces, Pichia (e.g. Pichia pastoris), Hansenula (e.g. Hansenula polymorpha), Yarowia, Schwaniomyces, Schizosaccharomyces, Zygosaccharomyces and the like. Saccharomyces cerevisiae, S. carlsbergensis and K. lactis are the most commonly used yeast hosts, and are convenient fungal hosts. Animal host cells suitable for use with the invention include insect cells and mammalian cells (e.g. derived from Chinese hamster (e.g. CHO), and human cell lines, such as HeLa). Exemplary insect cell lines include, but are not limited to, Sf9 cells, baculovirus-insect cell systems (e.g. review Jarvis, Virology Volume 310, Issue 1, 25
May 2003, Pages 1-7). Alternatively, the host cells may also be transgenic animals or plants.
Introduction of a vector in a host cell can be effected by, e.g., calcium phosphate transfection, virus infection, DEAE-dextran-mediated transfection, lipofectamin transfection or electroporation, and any person skilled in the art can select and use an introduction method suitable for the expression vector and host cell used.
A further aspect of the invention relates to a composition comprising a binding agent, such as an antibody or antigen-binding fragment thereof, comprising one or more ISVDs (or variants thereof) as described herein.
A ‘composition’, as used herein, refers to a combination of one or more molecules, present in a formulation that retains the binding agents activity, specifically the HR2 (or S2) binding and Sarbecovirus neutralization activity in this case, thus a functional composition. The composition thus comprises one or more molecules which constitute one or more binding agents as described herein which specifically bind the Sarbecovirus Spike protein via interaction with its HR2 domain (S2 targeting binding agents or molecules). In particular embodiments, the composition may comprise a bivalent antibody comprising an ISVD (or variant thereof) as described herein fused to an Fc domain. Said composition may be a soluble or solid composition.
In addition to said S2 targeting binding agent molecule(s), the composition may further comprise, for instance but not limited to, buffer components, adjuvants, or additional molecules, which may be functional molecules.
In particular embodiments, the composition may further comprise one or more binding agents capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD) as described elsewhere herein. In particular embodiments, the composition may further comprise one or more binding agents, such as an antibody or antigen-binding fragment thereof, comprising one or more (such as two, three, four, or more) ISVDs (or variants or humanized forms thereof) capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD) as described elsewhere herein. Said composition may thus contain at least two binding agents, characterized in that one binding agent specifically binds the HR2 domain, and the second binding agent specially binds the RBD region, resulting in a composition with at least two binding agents binding in a non-competing manner to the spike protein, possibly simultaneously.
In preferred embodiments, the binding agent capable of binding to a Sarbecovirus spike protein RBD is capable of binding two non-competing binding sites of the RBD, preferably via two different
ISVDs present in said binding agent, wherein said binding agent may be a bispecific binding agent, or multispecific binding agent. More specifically, the binding agent may comprise one or more ISVDs capable of binding to or competing for the VHH72 epitope as defined herein, and one or more ISVDs capable of binding to or competing for the VHH3. 117 epitope as defined herein. Non-limiting examples of binding agents comprising one or more ISVDs capable of binding to or competing for the VHH72 epitope, and one or more ISVDs capable of binding to or competing for the VHH3. 117 epitope are described in WO 2022/238550 Al. Also disclosed herein are particular examples of binding agents comprising one or more ISVDs capable of binding to or competing for the VHH72 epitope, and one or more ISVDs capable of binding to or competing for the VHH3. 117 epitope , in particular antibodies or antigen-binding fragments thereof, comprising i) one or more ISVDs comprising the CDRs present in SEQ ID NO:91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g. VHH83), or a variant or a humanized form thereof, and ii) an ISVD comprising the CDRs present in SEQ ID NO: 92, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3.117), or a variant or a humanized form thereof, wherein the CDRs are annotated according to Kabat, Martin, MacCallum, IMGT, AbM, or Chothia. More particularly bispecific antibodies comprising i) an ISVD comprising the CDRs present in SEQ ID NO:92, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3. 117), or a variant or a humanized form thereof, fused to the N-terminus of an Fc domain as defined herein, and ii) an ISVD comprising the CDRs present in SEQ ID NO:91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g. VHH83), or a variant or a humanized form thereof, fused to the C-terminus of the Fc domain. Or i) the ISVD comprising the CDRs present in SEQ ID NO: 92, such as the ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3. 117), or a variant or a humanized form thereof, may be fused to the C-terminus of the Fc domain, and ii) the ISVD comprising the CDRs present in SEQ ID NO:91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g. VHH83), or a variant or a humanized form thereof, may be fused to the N-terminus of the Fc domain. In embodiments, the ISVD fused to the C-terminus of the Fc domain may be modified to reduce reactivity of the ISVD with pre-existing anti -drug antibodies (ADAs). In particular said ISVD may comprise a C-terminal modification to reduce reactivity of the ISVD with pre-existing ADAs, such as the C-terminal TVSS sequence (SEQ ID NO: 139) of the ISVD may be replaced by any one of the sequences set forth in SEQ ID NO: 140-150 or the amino acid sequence TVS. The same modifications may be provided in any binding agent described herein, in particular a binding agent comprising an ISVD which is not fused at its C-terminus to another moiety such as an Fc moiety,
such as more particularly a binding agent comprising an ISVD comprising a C-terminal sequence as set forth in SEQ ID NO: 139.
In further particular embodiments, the bispecific antibody comprises or consists of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014), or any functional variant thereof, or a variant with at least 90% identity thereof, or a humanized variant thereof, such as a bispecific antibody comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 127-138.
In particular embodiments, the composition may comprise (i) a binding agent, in particular an antibody or an antigen-binding fragment thereof, comprising one or more ISVDs comprising the CDRs present in any one of SEQ ID NO: 2 to 26, such as one or more ISVDs comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 2 to 26, or a variant or humanized form thereof; and (ii) a binding agent, in particular an antibody or an antigen-binding fragment thereof, comprising one or more ISVDs comprising the CDRs present in SEQ ID NO:91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g. VHH83), or a variant or a humanized form thereof, and an ISVD comprising the CDRs present in SEQ ID NO:92, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3. 117), or a variant or a humanized form thereof, wherein the CDRs are annotated according to Kabat, Martin, MacCallum, IMGT, AbM, or Chothia.
In further particular embodiments, the composition may comprise:
(i) a binding agent, in particular an (optionally bivalent) antibody, comprising an ISVD comprising the CDRs present in any one of SEQ ID NO: 2 to 26, such as an ISVD comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 2 to 26 or a variant or humanized form thereof, fused to an Fc domain, such as an antibody comprising the amino acid sequence set forth in any one of SEQ ID NO: 96 to 119; and
(ii-a) a binding agent, in particular a bispecific antibody, comprising an ISVD comprising the CDRs present in SEQ ID NO:92, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3. 117), or a variant or a humanized form thereof, fused to the N- terminus of an Fc domain as defined herein, and an ISVD comprising the CDRs present in SEQ ID NO:91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g. VHH83), or a variant or a humanized form thereof, fused to the C-terminus of the Fc domain, wherein the CDRs are annotated according to Kabat, Martin, MacCallum, IMGT, AbM, or Chothia.
In certain embodiments, the bispecific antibody (ii-a) comprises an ISVD comprising or consisting
of the sequence set forth in SEQ ID NO:92, fused to the N-terminus of an Fc domain as defined herein, and an ISVD comprising or consisting of the sequence set forth in SEQ ID NO: 91 fused to the C-terminus of the Fc domain. Optionally, the Fc domain comprises or consists of the sequence set forth in any one of SEQ ID NO: 82 to 86.
In certain embodiments, the bispecific antibody (ii-a) comprises an ISVD consisting of the sequence set forth in SEQ ID NO:92, fused to the N-terminus of an Fc domain as defined herein, and an ISVD consisting of the sequence set forth in SEQ ID NO: 91 fused to the C-terminus of the Fc domain. Optionally, the Fc domain comprises or consists of the sequence set forth in any one of SEQ ID NO: 82 to 86.
In further particular embodiments, the composition may comprise:
(i) a binding agent, in particular a (optionally bivalent) antibody, comprising an ISVD comprising the CDRs present in any one of SEQ ID NO: 2 to 26, such as an ISVD comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 2 to 26 or a variant or humanized form thereof, fused to an Fc domain, such as an antibody comprising the amino acid sequence set forth in any one of SEQ ID NO: 96 to 119; and
(ii-b) a binding agent, in particular a bispecific antibody, comprising an ISVD comprising the CDRs present in SEQ ID NO:92, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92 (e.g. VHH3. 117), or a variant or a humanized form thereof, fused to the C- terminus of an Fc domain as defined herein, and an ISVD comprising the CDRs present in SEQ ID NO: 91, such as an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:91 (e.g. VHH83), or a variant or a humanized form thereof, fused to the N-terminus of the Fc domain, wherein the CDRs are annotated according to Kabat, Martin, MacCallum, IMGT, AbM, or Chothia.
In certain embodiments, the bispecific antibody (ii-b) comprises an ISVD comprising or consisting of the sequence set forth in SEQ ID NO:92, fused to the C-terminus of an Fc domain as defined herein, and an ISVD comprising or consisting of the sequence set forth in SEQ ID NO: 91 fused to the N-terminus of the Fc domain. Optionally, the Fc domain comprises or consists of the sequence set forth in any one of SEQ ID NO: 82 to 86.
In certain embodiments, the bispecific antibody (ii-b) comprises an ISVD consisting of the sequence set forth in SEQ ID NO:92, fused to the C-terminus of an Fc domain as defined herein, and an ISVD consisting of the sequence set forth in SEQ ID NO: 91 fused to the N-terminus of the Fc domain. Optionally, the Fc domain comprises or consists of the sequence set forth in any one of SEQ ID NO:
82 to 86.
In further particular embodiments, the composition may comprise:
(i) a binding agent, in particular an (optionally bivalent) antibody, comprising an ISVD comprising the CDRs present in any one of SEQ ID NO: 2 to 26, such as an ISVD comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 2 to 26 or a variant or humanized form thereof, fused to an Fc domain, such as an antibody comprising the amino acid sequence set forth in any one of SEQ ID NO: 96 to 119; and
(ii) a bispecific antibody comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126, or any functional variant thereof, or a variant with at least 90% identity thereof, or a humanized variant thereof.
In further particular embodiments, the composition may comprise:
(i) a binding agent, in particular a (optionally bivalent) antibody, comprising an ISVD comprising the CDRs present in any one of SEQ ID NO: 2 to 26, such as an ISVD comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 2 to 26 or a variant or humanized form thereof, fused to an Fc domain, such as an antibody comprising the amino acid sequence set forth in any one of SEQ ID NO: 96 to 119; and
(ii) a bispecific antibody comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 127-138, or any functional variant thereof, or a variant with at least 90% identity thereof, or a humanized variant thereof.
In particular embodiments, the binding agent (i) comprised by any one of the above compositions may be a binding agent, in particular an (optionally bivalent) antibody, comprising an ISVD comprising the CDRs present in SEQ ID NO: 25, such as an ISVD comprising or consisting of the amino acid sequence set forth in any one of SEQ ID NO: 25 (e.g. VHH CBV3) or a variant or humanized form thereof, fused to an Fc domain as defined herein, such as an antibody comprising the amino acid sequence set forth in SEQ ID NO: 118, or a variant or humanized form thereof.
In yet further particular embodiments, the composition may comprise (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof; and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126, or any functional variant thereof, or a variant with at least 90% identity thereof, or a humanized
variant thereof, such as a bispecific antibody comprising or consisting of the sequence set forth in any one of SEQ ID NO: 127-138.
For example, the composition may comprise (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof; and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014).
In another example the composition may comprise (i) a binding agent, in particular a (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof; and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 127.
In another example the composition may comprise (i) a binding agent, in particular a (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof; and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 128.
In embodiments, the molecular ratio of the (S2 targeting) binding agent, such as an antibody or antigen-binding fragment thereof, comprising one or more ISVDs (or variants thereof), or part thereof, as described herein and the (S 1 targeting) binding agent such as an antibody or antigenbinding fragment thereof, comprising one or more ISVDs (or variants thereof) capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD) in the composition may range from 20: 1 to 1:20 such as from 10: 1 to 1: 10, from 9: 1 to 1:9, from 8: 1 to 1:8, from 7: 1 to 1:7, from 6: 1 to 1:6, from 5: 1 to 1:5, from 4: 1 to 1:4 or from 3: 1 to 1:3, preferably from 2: 1 to 1:2, more preferably the molecular ratio is about 1: 1.
In embodiments, the mass ratio of the (S2 targeting) binding agent, such as an antibody or antigenbinding fragment thereof, comprising one or more ISVDs (or variants thereof), or part thereof, as described herein and the (SI targeting) binding agent such as an antibody or antigen-binding fragment thereof, comprising one or more ISVDs (or variants thereof) capable of binding to a Sarbecovirus spike protein receptor binding domain (RBD) in the composition may range from 20: 1 to 1:20 such as from 10: 1 to 1: 10, from 9: 1 to 1:9, from 8: 1 to 1:8, from 7: 1 to 1:7, from 6: 1 to 1:6, from 5: 1 to 1:5, from 4: 1 to 1:4 or from 3: 1 to 1:3, preferably from 2: 1 to 1:2, more preferably the mass ratio is about 1: 1.
An exemplary composition comprises (i) a binding agent, in particular an (optionally bivalent)
antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof (e.g., XVR013m); and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014) or a variant thereof such as such as a bispecific antibody comprising or consisting of the sequence set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of the binding agent (i) to the binding agent (ii) in the composition is from 20 : 1 to 1:20, preferably from 10 : 1 to 1: 10 such as from 9: 1 to 1:9, from 8: 1 to 1:8, from 7: 1 to 1:7, from 6: 1 to 1:6, from 5: 1 to 1:5, from 4: 1 to 1:4, from 3: 1 to 1:3, from 2: 1 to 1:2, or about 1: 1.
An exemplary composition comprises (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof (e.g., XVR013m); and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014) or a variant thereof such as such as a bispecific antibody comprising or consisting of the sequence set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of the binding agent (i) to the binding agent (ii) in the composition is from 10: 1 to 1: 10.
Another exemplary composition comprises (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof (e.g., XVR013m); and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014) or a variant thereof such as such as a bispecific antibody comprising or consisting of the sequence set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of the binding agent (i) to the binding agent (ii) in the composition is about 1: 10.
Another exemplary composition comprises (i) a binding agent, in particular an (optionally bivalent) antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 118, or a variant or a humanized form thereof (e.g., XVR013m); and (ii) a binding agent, in particular a bispecific antibody, comprising or consisting of the amino acid sequence set forth in SEQ ID NO: 126 (e.g. XVR014) or a variant thereof such as such as a bispecific antibody comprising or consisting of the sequence set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of the binding agent (i) to the binding agent (ii) in the composition is about 1: 1.
A particularly preferred composition comprises XVR013m and XVR014, wherein the mass ratio of XVR013m to XVR014 in the composition is from 20: 1 to 1:20, preferably from 10: 1 to 1: 10 such as from 9: 1 to 1:9, from 8: 1 to 1:8, from 7: 1 to 1:7, from 6: 1 to 1:6, from 5: 1 to 1:5, from 4: 1 to 1:4,
from 3: 1 to 1:3, from 2: 1 to 1:2, or about 1: 1. In certain embodiments, the composition comprises XVR013m and XVR014, wherein the mass ratio of XVR013m to XVR014 is about 1: 10. In certain embodiments, the composition comprises XVR013m and XVR014, wherein the mass ratio of XVRO 13m to XVRO 14 is about 1: 1.
Further particularly preferred compositions comprise XVRO 13m and a bispecific antibody as set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of XVR013m to the bispecific antibody is from 20: 1 to 1:20, preferably from 10: 1 to 1: 10 such as from 9: 1 to 1:9, from 8: 1 to 1:8, from 7: 1 to 1:7, from 6: 1 to 1:6, from 5: 1 to 1:5, from 4: 1 to 1:4, from 3: 1 to 1:3, from 2: 1 to 1:2, or about 1: 1. In certain embodiments, the composition comprises XVRO 13m and a bispecific antibody as set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio ofXVR013m to the bispecific antibody is about 1: 10. In certain embodiments, the composition comprises XVRO 13m and a bispecific antibody as set forth in any one of SEQ ID NO: 127-138, wherein the mass ratio of XVRO 13m to the bispecific antibody is about 1: 1.
Moreover said composition may still contain additional binding agent(s) or molecules, which optionally bind further binding regions on the same or different epitopes of the spike protein, or other viral proteins, or may even target totally unrelated target proteins.
A further aspect of the invention relates to medicaments or pharmaceutical compositions comprising a binding agent, in particular an antibody or antigen-binding fragment, a nucleic acid encoding it, and/or a (recombinant) vector comprising the nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein. In particular, a pharmaceutical composition is a pharmaceutically acceptable composition; such compositions are preferably further comprising a (pharmaceutically) suitable or acceptable carrier, diluent, adjuvant, excipient, stabilizer, etc.
By “pharmaceutically acceptable” is meant a material that is not biologically or otherwise undesirable, i.e., the material may be administered to an individual along with the compound, in particular the Sarbecovirus binding agent, more particularly the Sarbecovirus antibody or antigenbinding fragment, without causing any undesirable biological effects or interacting in a deleterious manner with any of the other components of the pharmaceutical composition in which it is contained. A pharmaceutically acceptable carrier is preferably a carrier that is relatively non-toxic and innocuous to a patient at concentrations consistent with effective activity of the active ingredient so that any side effects ascribable to the carrier do not vitiate the beneficial effects of the active ingredient. Suitable carriers or adj uvantia typically comprise one or more of the compounds included
in the following non- exhaustive list: large slowly metabolized macromolecules such as proteins, polysaccharides, polylactic acids, polyglycolic acids, polymeric amino acids, amino acid copolymers and inactive virus particles. The term “excipient”, as used herein, is intended to include all substances which may be present in a pharmaceutical composition and which are not active ingredients but may contribute to e.g. long-term stability, or therapeutic enhancement on the active ingredient (such as by facilitating drug absorption, reducing viscosity, or enhancing solubility). Excipients may include, for example, salts, binders (e.g., lactose, dextrose, sucrose, trehalose, sorbitol, mannitol), lubricants, thickeners, surface active agents, preservatives, emulsifiers, buffer substances, stabilizing agents, flavouring agents or colorants. A “diluent”, such as in particular a “pharmaceutically acceptable diluent”, includes vehicles such as water, saline, physiological salt solutions, glycerol, ethanol, etc. Auxiliary substances such as wetting or emulsifying agents, pH buffering substances, preservatives may be included in such vehicles.
A pharmaceutically effective amount of binding agents, in particular antibodies or antigen-binding fragments, of the invention is preferably that amount which produces a result or exerts an influence on the particular condition being treated.
The pharmaceutical composition of this invention can be lyophilized for storage and reconstituted in a suitable carrier prior to use. When prepared as lyophilization or liquid, physiologically acceptable carrier, excipient, stabilizer need to be added into the pharmaceutical composition of the invention (Remington's Pharmaceutical Sciences 22nd edition, Ed. Allen, Loyd V, Jr. (2012). The preparation containing pharmaceutical composition of this invention should be sterilized before injection. This procedure can be done using sterile filtration membranes before or after lyophilization and reconstitution. The pharmaceutical composition can be packaged in a container or vial with sterile access port, such as an i.v. solution bottle with a rubber stopper - the pharmaceutical composition can be present as liquid, or the container or vial is filled with a liquid pharmaceutical composition that is subsequently lyophilized or dried; or can be packaged in a pre-filled syringe.
Pharmaceutical compositions of nucleic acid molecules encoding a binding agent as described herein, such as an mRNA or an mRNA construct as described herein encoding the polypeptide portion of a polypeptidic or polypeptide Sarbecovirus binding agent, in particular an antibody or antibody fragment, as identified herein, more particularly an ISVD (or a variant thereof) as described herein, or a functional part thereof, may comprise the nucleic acid molecule, mRNA or mRNA construct encapsulated in lipid nanoparticles that protect the nucleic acid molecules, mRNA or mRNA construct and help their absorption into cells. As used herein, the term “lipid nanoparticle” or “LNP”
refers to a nanosized particle composed of different lipids. Many different types of lipids may be included in such LNP, including, without limitation ionizable lipids, phospholipids (e.g. compounds having a phosphatidylcholine group), sterols (e.g. cholesterol), polyethylene glycol (PEG)-modified lipids. Preferably, the lipid nanoparticles contain a cationic lipid, or a lipid species with a net positive charge at a selected pH (e.g., physiological pH), to encapsulate and/or enhance the delivery of mRNA into target cells. As used herein, the terms “ionizable lipid” or “cationic ionizable lipid” or “cationic lipids” refer to lipids having at least one protonatable or deprotonatable group, such that the lipid is positively charged at a pH at or below physiological pH (e.g., pH 7.4), and neutral at a second pH, preferably at or above physiological pH. It will be understood by one of ordinary skill in the art that the addition or removal of protons as a function of pH is an equilibrium process, and that the reference to a charged or a neutral lipid refers to the nature of the predominant species and does not require that all of the lipid be present in the charged or neutral form. Advantageously, the positively charged lipids are able to associate with negatively charged compounds. Also advantageously, a positive charge on the LNP may promote association with the negatively charged cell membrane to enhance cellular uptake. In addition to the ionizable lipids, in particular cationic ionizable lipids, the lipid nanoparticles may further comprise a structural helper lipid. Suitable helper lipids are generally known in the art. A preferred helper lipid is a steroid or a sterol, more preferably cholesterol. Incorporation of a steroid or a sterol (e.g. cholesterol and analogues thereof, ergosterol, phytosterol) in the lipid nanoparticle may help aggregation of other lipids in the lipid nanoparticle. Other possible helper lipids are phospholipids including, without limitation, dioleoylphosphatidylethanolamine (DOPE) and l,2-distearoyl-sn-glycero-3 -phosphocholine (DSPC).
A further aspect of the invention relates to a binding agent, in particular an antibody or antigenbinding fragment, a nucleic acid encoding it as described herein, a vector comprising such nucleic acid, a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein or a pharmaceutical composition comprising a binding agent, in particular an antibody or antigen-binding fragment, nucleic acid encoding it, a (recombinant) vector comprising such nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, for use as a medicine or medicament. Alternatively, use of a binding agent, in particular an antibody or antigen-binding fragment, nucleic acid encoding it, a vector comprising such nucleic acid as described herein, or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, or use of a pharmaceutical composition comprising a binding agent, in particular an antibody or antigen-binding fragment, nucleic acid encoding it, a vector comprising such nucleic acid, and/or
a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, in the manufacture of a medicine or medicament is envisaged.
In particular, the binding agent, in particular the antibody or antigen-binding fragment, the nucleic acid encoding it, the vector comprising such nucleic acid or the composition comprising the binding agent, in particular the antibody or antigen-binding fragment, as described herein, or the medicament or pharmaceutical composition comprising a binding agent, in particular an antibody or antigenbinding fragment, a nucleic acid encoding it, a (recombinant) vector comprising such nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, is for use in passive immunisation, for use in treating a subject with a Sarbecovirus infection, for use in preventing infection of a subject with a Sarbecovirus. or for use in protecting a subject from infection with a Sarbecovirus .
When for use in passive immunisation, the subject may have an infection with a Sarbecovirus (therapeutic passive immunisation) or may not have an infection with a Sarbecovirus (prophylactic passive immunisation).
A related aspect relates to methods for treating a subject suffering from/having/that has contracted an infection with a Sarbecovirus, the methods comprising administering a binding agent, in particular an antibody or antigen-binding fragment, a nucleic acid encoding it a (recombinant) vector comprising such nucleic acid, or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein to the subject, or comprising administering a medicament or pharmaceutical composition comprising a binding agent , in particular an antibody or antigen-binding fragment, a nucleic acid encoding it, a (recombinant) vector comprising such nucleic acid and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein to the subject.
A further aspect of the invention relates to methods for protecting a subject from infection with a Sarbecovirus or for preventing infection of a subject with a Sarbecovirus, the methods comprising administering a binding agent, in particular an antibody or antigen-binding fragment, a nucleic acid encoding it, a (recombinant) vector comprising such nucleic acid, or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein to the subject prior to infection, or comprising administering a medicament or pharmaceutical composition as described herein comprising a binding agent, in particular an antibody or antigen-binding fragment, a nucleic acid encoding it, a (recombinant) vector comprising such nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, to
the subject prior to infection.
Further related aspects are directed to use of a binding agent, in particular an antibody or antigenbinding fragment, nucleic acid encoding it, a vector comprising such nucleic acid as described herein, or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, or use of a pharmaceutical composition comprising a binding agent, in particular an antibody or antigen-binding fragment, nucleic acid encoding it, a vector comprising such nucleic acid, and/or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein, in the manufacture of a medicine or medicament for treating a subject with a Sarbecovirus infection, for preventing infection of a subject with a Sarbecovirus. or for protecting a subject from infection with a Sarbecovirus .
In the above medical aspects, a nucleic acid encoding a binding agent, in particular an antibody or antigen-binding fragment or a (recombinant) vector comprising such nucleic acid as described herein can be used in e.g. gene therapy setting. “Gene therapy” as used herein refers to therapy performed by the administration to a subject of an expressed or expressible nucleic acid. For such applications, the nucleic acid molecule or vector as described herein allow for production of the binding agent, antibody or antibody fragment within a cell. A large set of methods for gene therapy are available in the art and include, for instance (adeno-associated) virus-mediated gene silencing, or virus-mediated gene therapy (e.g. US 20040023390; Mendell et al 2017, N Eng J Med 377: 1713-1722). A plethora of delivery methods are well known to those of skill in the art and include but are not limited to viral delivery systems, microinjection of DNA plasmids, biolistics of naked nucleic acids, use of a liposome or an artificial exosome, administration of the nucleic acid or vector formulated in a nanoparticle or lipid or lipid-comprising particle. In vivo delivery by administration to an individual patient occurs typically by systemic administration (e.g., intravenous, intraperitoneal infusion or brain injection; e.g. Mendell et al 2017, N Eng J Med 377: 1713-1722).
A “therapeutically active agent” generally means any molecule that has or may have a therapeutic effect (i.e. curative or prophylactic effect) in the context of treatment of a disease. Preferably, a therapeutically active agent is a disease-modifying agent, which can be a cytotoxic agent, such as a toxin, or a cytotoxic drug, or an enzyme capable of converting a prodrug into a cytotoxic drug, or a radionuclide, or a cytotoxic cell, or which can be a non-cytotoxic agent. Even more preferably, a therapeutically active agent has a curative effect on the disease. The binding agent, in particular the antibody or antibody fragment, or pharmaceutical composition of the invention may act as a therapeutically active agent, when beneficial in treating patients infected with a Sarbecovirus, such
SARS-CoV-2 or SARS-CoV-1 or patients suffering from COVID-19. The binding agent, in particular the antibody or antibody fragment, may comprise a Sarbecovirus-binding ISVD or a variant thereof as described herein, and may contain or be coupled to additional functional groups, advantageous when administrated to a subject. Examples of such functional groups and of techniques for introducing them will be clear to the skilled person, and can generally comprise all functional groups and techniques mentioned in the art as well as the functional groups and techniques known per se for the modification of pharmaceutical proteins, and in particular for the modification of antibodies or antibody fragments, for which reference is for example made to Remington's Pharmaceutical Sciences, 16th ed., Mack Publishing Co., Easton, PA (1980). Such functional groups may for example be linked directly (for example covalently) to the ISVD or active antibody fragment, or optionally via a suitable linker or spacer, as will again be clear to the skilled person. One of the most widely used techniques for increasing the half-life and/or reducing immunogenicity of pharmaceutical proteins comprises attachment of a suitable pharmacologically acceptable polymer, such as poly(ethyleneglycol) (PEG) or derivatives thereof (such as methoxypoly(ethyleneglycol) or mPEG). For example, for this purpose, PEG may be attached to a cysteine residue that naturally occurs in an immunoglobulin single variable domain described herein (or a variant thereof as described herein), an immunoglobulin single variable domain as described herein (or a variant thereof as described herein) may be modified so as to suitably introduce one or more cysteine residues for attachment of PEG, or an amino acid sequence comprising one or more cysteine residues for attachment of PEG may be fused to the N- and/or C-terminus of an ISVD or active antibody fragment as described herein (or a variant thereof as described herein), all using techniques of protein engineering known per se to the skilled person. Another, usually less preferred modification comprises N-linked or O-linked glycosylation, usually as part of co-translational and/or post- translational modification, depending on the host cell used for expressing the antibody or active antibody fragment. Another technique for increasing the half-life of a binding domain, in particular an antibody or antibody fragment, may comprise the engineering into bifunctional or bispecific domains (for example, one ISVD or active antibody fragment against the target Sarbecovirus HR2 domain and one against a serum protein such as albumin or Staphylococcal protein A (SpA) -which is a surface protein abundantly present in the lungs aiding in prolonging half-life)) or into fusions of antibody fragments, in particular immunoglobulin single variable domains, with peptides (for example, a peptide against a serum protein such as albumin). In yet another example, an ISVD as described herein (or a variant thereof as described herein) can be fused to an immunoglobulin Fc domain as described elsewhere herein. Examples are further shown in the experimental section and
are also depicted in the sequence listing. In embodiments, in the above medical aspects, the Sarbecovirus is SARS-CoV-2 such as a SARS-CoV-2 variant, or SARS-CoV-1. The SARS-CoV-2 variant may be a variant at position N439, K417, S477, L452, T478, E484, P384, N501 and/or D614 (relative to the SARS-CoV-2 spike amino acid sequence as defined in SEQ ID NO:93), more particularly a variant at position N501 such as a N501Y variant (e.g. SARS-CoV-2 Alpha variant), a variant at position N501 and E484 such as a N501Y and E484K variant (e.g. SARS-CoV-2 Alpha + E484K variant), a variant at position K417, E484 and N501 such as a K417N, E484K and N501Y variant (e.g. SARS-CoV-2 beta variant), a variant at position P384, K417, E484 and N501 such as a P384L, K417N, E484K and N501Y variant (e.g. SARS-CoV-2 beta + P384L variant), a variant at position L452 and E484 such as a L452R and E484Q variant (e.g. SARS-CoV-2 kappa variant), a variant at position L452 and T478 such as a L452R and T478K variant (e.g. SARS-CoV-2 delta variant), a variant at position L452 such as a L452R variant (e.g. SARS-CoV-2 epsilon variant), a variant at position K417 such as a K417T variant (e.g. SARS-CoV-2 gamma variant), a variant at position D614 such as a D614G variant (e.g. SARS-CoV-2 Omicron variant or SARS-CoV-2 BA. l variant); or a SARS-CoV-2 Omicron EG.5.1 variant, a SARS-CoV-2 Omicron BA.2.86.1 variant, a SARS-CoV-2 Omicron HV. l variant, a SARS-CoV-2 Omicron XBB.1.5 variant, a SARS-CoV-2 Omicron XBB.2.3 variant, a SARS-CoV-2 Omicron XBB.1.16 variant, a SARS-CoV-2 Omicron HK.3 variant, SARS-CoV-2 Omicron BA.2.74 variant, a SARS-CoV-2 Omicron FL. 1.5.1 variant, a SARS-CoV-2 Gamma variant, a SARS-CoV-2 Eta variant, a SARS-CoV-2 Iota variant, a SARS- CoV-2 Epsilon B. 1.427 variant, a SARS-CoV-2 Epsilon B. 1.429 variant, a SARS-CoV-2 Mu variant, a SARS-CoV-2 Zeta variant, a SARS-CoV-2 B. 1.617.3 variant, a SARS-CoV-2 Omicron CH. 1.1 variant, a SARS-CoV-2 Omicron JN. l variant, a SARS-CoV-2 Omicron KP.2.3 variant, a SARS- CoV-2 Omicron KP.3 variant, a SARS-CoV-2 Omicron KP.3.1.1 variant, a SARS-CoV-2 Omicron KP.1.1 variant, a SARS-CoV-2 Omicron LB. l variant or a SARS-CoV-2 Omicron XEC variant. In particular embodiments, the Sarbecovirus is any one or both of SARS-CoV-2 and SARS-CoV-1. In further particular embodiments, SARS-CoV-2 is SARS-CoV-2 Wuhan strain or a SARS-CoV-2 variant, in particular a SARS-CoV-2 variant selected from the group consisting of SARS-CoV-2 Alpha variant, SARS-CoV-2 Omicron BA. l variant and SARS-CoV-2 Omicron BA.2 variant or selected from the group consisting of: SARS-CoV-2 D614G variant, SARS-CoV-2 Alpha variant, SARS-CoV-2 Beta variant, SARS-CoV-2 Gamma variant, SARS-CoV-2 Delta variant, SARS-CoV- 2 Epsilon B. 1.427 variant, SARS-CoV-2 Epsilon B. 1.429 variant, SARS-CoV-2 Eta variant, SARS- CoV-2 Iota variant, SARS-CoV-2 Kappa variant, SARS-CoV-2 Mu variant, SARS-CoV-2 Zeta variant, SARS-CoV-2 B. 1.617.3 variant, SARS-CoV-2 Omicron BA.5 variant, SARS-CoV-2
Omicron XBB.1.5 (-G252V) variant, SARS-CoV-2 Omicron XBB.1.5 variant, SARS-CoV-2 Omicron XBB.2.3 variant, SARS-CoV-2 Omicron XBB.1.16 variant, SARS-CoV-2 Omicron CH. 1. 1 variant, SARS-CoV-2 Omicron EG.5. 1 variant, SARS-CoV-2 Omicron HK.3 variant, SARS- CoV-2 Omicron BA.2.74 variant, SARS-CoV-2 Omicron BA.2.86. 1 variant, SARS-CoV-2 Omicron HV. l variant, SARS-CoV-2 Omicron FL. 1.5.1 variant, SARS-CoV-2 Omicron JN.l variant, SARS- CoV-2 Omicron KP.2.3 variant, SARS-CoV-2 Omicron KP.3 variant, SARS-CoV-2 Omicron KP.3.1.1 variant, a SARS-CoV-2 Omicron BA. l variant, a SARS-CoV-2 Omicron BA.2 variant, a SARS-CoV-2 Omicron KP.1.1 variant, a SARS-CoV-2 Omicron LB. l variant and a SARS-CoV-2 Omicron XEC variant.
As used herein, the terms “therapy” or “treatment” refer to the alleviation or measurable lessening of one or more symptoms or measurable markers of a pathological condition such as a disease or disorder, in particular a Sarbecovirus infection. Measurable lessening includes any statistically significant decline in a measurable symptom or marker. Generally, the terms encompass both curative treatments and treatments directed to reduce symptoms and/or slow progression of the disease. The terms encompass both the therapeutic treatment of an already developed pathological condition, in particular a Sarbecovirus infection, as well as prophylactic or preventative measures, wherein the aim is to prevent or lessen the chances of incidence of a pathological condition, in particular a Sarbecovirus infection. Beneficial or desired clinical results include, but are not limited to, prevention of a disease, reduction of the incidence of a disease, alleviation of symptoms associated with a disease, diminishment of extent of a disease, stabilisation of the disease, delay or slowing of the progression of a disease, amelioration or palliation of a disease, or combinations thereof. In certain embodiments, the terms may relate to therapeutic treatments. In certain other embodiments, the terms may relate to preventative treatments.
For example, treatment may refer to passive immunisation of a subject having contracted a Sarbecovirus infection (therapeutic treatment). Prevention of infection with a Sarbecovirus may be useful in case of e.g. epidemic or pandemic conditions during which subjects known to be most vulnerable to develop severe disease symptoms can be prophylactically treated (preventive or prophylactic immunisation) with a binding agent, in particular an antibody or antigen-binding fragment or a nucleic acid encoding it or a vector comprising such nucleic acid or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment as described herein such as to prevent infection overall, or such as to prevent development or occurrence of severe disease symptoms.
In embodiments, a therapeutically effective amount of a binding, in particular an antibody or antigenbinding fragment, nucleic acid, vector or pharmaceutical composition is administered to a subject in need thereof. In other embodiments, a prophylactically effective amount of a binding agent, in particular an antibody or antigen-binding fragment, nucleic acid, vector or pharmaceutical composition is administered to a subject in need thereof. A “therapeutically effective amount” or “therapeutically effective dose” indicates an amount of binding agent, in particular antibody or antigen-binding fragment, nucleic acid, vector or pharmaceutical composition that when administered to the subject brings about a clinical positive response with respect to therapeutic treatment of the subject afflicted by a Sarbecovirus infection, such as, e.g. curing infection with a Sarbecovirus . Similarly, a “prophylactically effective amount” or “prophylactically effective dose” refers to an amount of binding agent, in particular antibody or antigen-binding fragment, nucleic acid, vector or pharmaceutical composition that prevents, inhibits or delays the onset of a Sarbecovirus infection and/or prevents or reduces the risk of a clinical manifestation of a Sarbecovirus infection and/or reduces the severity, symptoms and/or duration of a Sarbecovirus infection in the subject. In order to achieve the therapeutic effect or the preventive or prophylactic effect, the binding agent, in particular the antibody or antigen-binding fragment or the nucleic acid encoding it or the vector comprising such nucleic acid or the composition comprising the binding agent, in particular the antibody or antigen-binding fragment as described herein may need to be administered to a subject multiple times, such as with an interval of 1 week or 2 weeks; the interval being dictated by the pharmacokinetic behaviour or characteristics (e.g. half-time or half-life in the subject’s circulation) of the binding agent, in particular the antibody or antigen-binding fragment, nucleic acid or vector. Alternatively, therapeutic treatments and prophylactic treatments in which a single dose of a binding agent, in particular an antibody or antigen-binding fragment as described herein is administered to the subject is envisaged. The single dose may be in the range of 0.5 mg/kg to 25 mg/kg.
The term “subject”, “individual” or “patient”, used interchangeably herein, relates to any organism such as a vertebrate, particularly any mammal, including both a human and other mammals, for whom diagnosis, therapy or prophylaxis is desired, e.g., an animal such as a rodent, a rabbit, a cow, a sheep, a horse, a dog, a cat, a lama, a pig, or a non-human primate (e.g., a monkey). The rodent may be a mouse, rat, hamster, guinea pig, or chinchilla. In one embodiment, the subject is a human, a rat or a non-human primate. Preferably, the subject is a human. In particular embodiments, a subject is a subject, such as a human subject, with or suspected of having an infection with a Sarbecovirus, also designated ’’patient” or “subject” herein. However, it will be understood that the aforementioned
terms do not imply that symptoms are present. In particular embodiments, the subject is a mammal susceptible to infection with a Sarbecovirus , such as a human subject that is susceptible to infection with SARS-CoV-2 such as a SARS-CoV-2 variant, or SARS-CoV-1.
The pharmaceutical composition of the invention can be administered to any patient in accordance with standard techniques. The administration can be by any appropriate mode, including oral, parenteral, topical, nasal, ophthalmic, intrathecal, intra-cerebroventricular, sublingual, rectal, vaginal, and the like. Still other techniques of formulation such as nanotechnology and aerosol and inhalant are also within the scope of this invention. The dosage and frequency of administration will depend on the age, sex and condition of the patient, concurrent administration of other drugs, counterindications and other parameters to be taken into account by the clinician.
In particular embodiments of the herein described medical aspects, the binding agent, in particular the antibody or antigen-binding fragment, the nucleic acid, the vector or the pharmaceutical composition may be administered to a subject via intravenous injection, subcutaneous injection, or intranasally, or, alternatively via inhalation or pulmonary delivery.
A further aspect of the invention relates to a binding agent, in particular an antibody or antigenbinding fragment, as described herein for use in diagnosing a Sarbecovirus infection, for use as a diagnostic agent. A nucleic acid encoding a Sarbecovirus binding agent, in particular a Sarbecovirus antibody or antigen-binding fragment as described herein, a (recombinant) vector comprising such nucleic acid, or a composition comprising a Sarbecovirus binding agent, in particular a Sarbecovirus antibody or antigen-binding fragment as described herein, can likewise be for use.
Use of a binding agent, in particular an antibody or antigen-binding fragment, as described herein in the manufacture of a (in vitro) diagnostic agent or diagnostic kit is also envisaged. In particular, the binding agent, in particular the antibody or antigen-binding fragment, as described herein may be for use in detecting the presence (or absence) of a Sarbecovirus or a part thereof (such as a Sarbecovirus spike protein or a part thereof) in a sample, such as a sample obtained from a subject, such as from a subject suspected to be infected with a Sarbecovirus . A nucleic acid encoding a binding agent, in particular an antibody or antigen-binding fragment, as described herein, a (recombinant) vector comprising such nucleic acid or composition comprising a binding agent, in particular an antibody or antigen-binding fragment, as described herein can likewise be used in the manufacture of a diagnostic agent or diagnostic kit, such as an in vitro diagnostic agent or kit.
A further aspect relates to methods for detecting a Sarbecovirus in a sample, such as a sample
obtained from a subject, such as from a subject suspected to be infected with a Sarbecovirus . Such methods usually comprise the steps of obtaining a sample, contacting the sample with a binding agent, in particular an antibody or antigen-binding fragment, as described herein, and detecting, determining, assessing, assaying, identifying or measuring binding of the binding agent, in particular the antibody or antigen-binding fragment, with a Sarbecovirus or a part thereof (such as a Sarbecovirus spike protein or a part thereof).
In particular embodiments of the herein described diagnostic aspects, the Sarbecovirus is selected from the group of clade la, lb, 2 and/or clade 3 Sarbecoviruses. such as SARS-CoV-2, Khosta-2, GD-Pangolin, RaTG13, WIV1, LYRal l, RsSHC014, Rs7327, SARS-CoV-1, Rs4231, Rs4084, Rp3, HKU3-1, or BM48-31 viruses, preferably SARS-CoV-2 such as a SARS-CoV-2 variant or SARS- CoV-1.
In embodiments of the herein described diagnostic aspects, the binding agent, in particular the antibody or antibody fragment, as described herein is comprising a detectable moiety fused to it, bound to it, coupled to it, linked to it, complexed to it, or chelated to it. A “detectable moiety” in general refers to a moiety that emits a signal or is capable of emitting a signal upon adequate stimulation, or to a moiety that is capable of being detected through binding or interaction with a further molecule (e.g. a tag, such as an affinity tag, that is specifically recognized by a labelled antibody), or is detectable by any means (preferably by a non-invasive means, if detection is in vivo/ inside the human body). Furthermore, the detectable moiety may allow for computerized composition of an image, as such the detectable moiety may be called an imaging agent. Detectable moieties include, without limitation, fluorescence emitters, phosphorescence emitters, positron emitters, radioemitters, etc., enzymes (capable of measurably converting a substrate) and molecular tags. Examples of radioemitters/radiolabels include 68Ga, HOmln, 18F, 45Ti, 44Sc, 47Sc, 61Cu, 60Cu, 62Cu, 66Ga, 64Cu, 55Ca, 72As, 86Y, 90Y, 89Zr, 1251, 74Br, 75Br, 76Br, 77Br, 78Br, U lin, 114mln, 114In, 99mTc, 11C, 32C1, 33C1, 34C1, 1231, 1241, 1311, 186Re, 188Re, 177Lu, 99Tc, 212Bi, 213Bi, 212Pb, 225Ac, 153Sm, and 67Ga. Fluorescence emitters include, without limitation, cyanine dyes (e.g. Cy5, Cy5.5, Cy7, Cy7.5), FITC, TRITC, coumarin, indolenine-based dyes, benzoindolenine-based dyes, phenoxazines, BODIPY dyes, rhodamines, Si-rhodamines, Alexa dyes, and derivatives of any thereof. Non-limiting examples of molecular tags include affinity tags, such as chitin binding protein (CBP), maltose binding protein (MBP), glutathione-S-transferase (GST), poly(His) (e.g., 6x His or His6), biotin or streptavidin, such as Strep-tag®, Strep-tag II® and Twin- Strep-tag®; solubilizing tags, such as thioredoxin (TRX), poly(NANP) and SUMO; chromatography
tags, such as a FLAG-tag; epitope tags, such as V5-tag, myc-tag and HA-tag; fluorescent labels or tags (i.e., fluorochromesZ-phores), such as fluorescent proteins (e.g., GFP, YFP, RFP etc.); luminescent labels or tags, such as luciferase, bioluminescent or chemiluminescent compounds (such as luminal, isoluminol, theromatic acridinium ester, imidazole, acridinium salts, oxalate ester, dioxetane or GFP and its analogs); phosphorescent labels; a metal chelator; and (other) enzymatic labels (e.g., peroxidase, alkaline phosphatase, beta-galactosidase, urease or glucose oxidase).
Binding agents, in particular antibodies and antibody fragments, as described herein and comprising a detectable moiety may for example be used for in vitro, in vivo or in situ assays (including immunoassays known per se such as ELISA, RIA, EIA and other "sandwich assays", etc.) as well as in vivo imaging purposes, depending on the choice of the specific label. .
A further aspect relates to kits comprising a binding agent, in particular an antibody or antigenbinding fragment, a nucleic acid encoding it, a vector comprising such nucleic acid as described herein, a composition comprising a binding agent, in particular an antibody or antigen-binding fragment, or a pharmaceutical composition comprising a binding agent, in particular an antibody or antigen-binding fragment; a nucleic acid encoding it a vector comprising such nucleic acid or a composition comprising a binding agent, in particular an antibody or antigen-binding fragment as described herein.
Such kits may be pharmaceutical kits or medicament kits which are comprising a container or vial (any suitable container or vial, such as a pharmaceutically acceptable container or vial) comprising an amount of binding agent, in particular antibody or antigen-binding fragment, or nucleic acid encoding it or vector comprising such nucleic acid as described herein or composition comprising a binding agent, in particular an antibody or antigen-binding fragment as described herein, and further comprising e.g. a kit insert such as a medical leaflet or package leaflet comprising information on e.g. intended indications (prophylactic or therapeutic treatment of a Sarbecovirus infection) and potential side-effects. Pharmaceutical kits or medicament kits may further comprise e.g. a syringe for administering the binding agent, in particular the antibody or antigen-binding fragment, nucleic acid encoding it vector comprising such nucleic acid or composition comprising the binding agent, in particular the antibody or antigen-binding fragment as described herein to a subject.
Such kits may also be diagnostic kits comprising a container or vial (any suitable container or vial, such as a pharmaceutically acceptable container or vial) comprising an amount of binding agent, in particular antibody or antigen-binding fragment, as described herein, such as a binding agent, in particular an antibody or antigen-binding fragment thereof comprising a detectable moiety. Such
diagnostic kits may further comprise e.g. one or more reagents to detect the detectable moiety and/or e.g. instructions on how to use said binding agent, in particular antibody or antigen-binding fragment, for detection of a Sarbecovirus in a sample.
While the invention has been described in conjunction with specific embodiments thereof, it is evident that many alternatives, modifications, and variations will be apparent to those skilled in the art in light of the foregoing description. Accordingly, it is intended to embrace all such alternatives, modifications, and variations as follows in the spirit and broad scope of the appended claims.
The herein disclosed aspects and embodiments of the invention are further supported by the following non-limiting examples.
EXAMPLES
Material and methods
Cell Lines
HEK293-S cells (ThermoFisher Scientific) were cultured in FreeStyle 293 expression media (Life Technologies), at 37°C with 8% CO2 while shaking at 130 rpm. HEK293-T cells (ATCC) and Vero E6 cells (ATCC) were cultured at 37°C in the presence of 5% CO2 in DMEM supplemented with 10% heat-inactivated fetal bovine serum (FBS), 1% penicillin, 1% streptomycin, 2 mM 1-glutamine, non-essential amino acids (Invitrogen) and 1 mM sodium pyruvate. ExpiCHO-S cells (GIBCO) were cultured at 37°C with 8% CO2 while shaking at 130 rpm in ExpiCHO expression media (GIBCO). Vero E6-TMPRSS2 cells that stably express human TMPRSS2 (NIBIOHN, JCRB 1819) (Matsuyama et al., PNAS, 2020) were cultured in DMEM containing 10% FBS, penicillin (100 unit/mL), streptomycin (100 ug/mL), Geneticin (G418) (Img/ml). When Vero E6-TMPRSS2 cells were seeded for assays medium without Geneticin was used.
Sotrovimab, bebtelovimab and palivizumab
Bebtelovimab Biosimilar (PX-TA1750) and sotrovimab Biosimilar (PX-TA1637) were commercially purchased from Proteogenix. Clinical grade Palivizumab was obtained from the Ghent University hospital.
Generation of S2 targeting molecules
Constructs (SEQ ID NO:96-119) were generated wherein a modified version (SEQ ID NO:3-26; Fig. 5 A) of the humanized version of the VHH building block R3 DC23 (hR3_DC23: SEQ ID NO:2) was connected to a human IgGl Fc containing an LS mutation (SEQ ID NO: 83) via a (648)2 (SEQ
ID NO:88) linker at the Fc N-terminus (Fig. 5B).
Generation of R3 DC23-Fc(YTE) (also referred to herein as huR3DC23-Fc) (SEQ ID NO: 125) and XVR013 (also referred to herein as R3 DC 23hum-Fc(LS) or huR3DC23-Fc LS or R3DC23-Fc LS) (SEQ ID NO: 124)
A humanized (Q1D, Q5V, A14P, D16G, T19R, M63V, S73N, D79Y, T82cL, K83R and Q108L, according to Kabat numbering; the T82cL modification in particular serves to inactivate the glycosylation of the N82a position, and can be useful in both humanized and non-humanized version, for expression in mammalian cells) version of R3_DC23 was fused via a (648)2 linker to a human IgGl Fc (EPKSCdel_YTE_K447del) ordered synthetically at IDT as gBlocks. Upon arrival, gBlocks were solubilized in ultraclean water at a concentration of 20 ng/pL. gBlocks were A-tailed using the NEBNext-dA-tailing module (NEB), purified using CleanPCR magnetic beads (CleanNA) and inserted in pcDNA3.4-TOPO vector (ThermoFisher). The ORF of positive clones was fully sequenced, and pDNA of selected clones was prepared using the NucleoBond Xtra Midi kit (Machery-Nagel).
A construct was generated wherein a humanized form of R3 DC23 (SEQ ID NO:2) was fused to a human IgGl Fc containing the LS mutation (SEQ ID NO: 83) via a (648)2 linker (SEQ ID NO: 88) at the Fc N-terminus (R3_DC23hum-Fc(LS) or huR3DC23-Fc_LS or XVR013) (SEQ ID NO: 124).
R3_DC23hum-Fc(LS) or huR3DC23-Fc_LS or XVR013:
DVQLVESGGGLVQPGGSLRLSCAVSGRI FSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG RFTISRDNAKNTVYLQMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSRRYDYWGQGTLVTVSSGG GGSGGGGSDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR EPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKL TVDKSRWQQGNVFSCSVLHEALHSHYTQKSLSLSPG ( SEQ ID NO : 124 )
Transient production of huR3DC23-Fc LS (also referred to herein as R3 DC 23hum-Fc(LS) or XVR013) (SEQ ID NO: 124)
The gene encoding huR3DC23-Fc_LS was codon optimized, synthesized, and cloned into the pXLG6 backbone vector at ATUM’s laboratories. Upon gene and codon optimization the R3DC23 DNA sequence was inserted into pXLG6 expression vector and transfected in CHOExpress ™ cells at a cell density of 4.00E+6 cells/ml. TGE supernatant was harvested by centrifugation and clarified by filtration (0.2pm) after 10 days when cell viability dropped below 10%. The protein was further
purified by Protein A.
Transient production of the S2 targeting molecules
The genes encoding the VHH-Fc_LS constructs (SEQ ID NO:96-99, 101,103, 105-109,111-119) were each codon optimized, synthesized and cloned into the manufacturer pcDNA3. 1 backbone vector.
The expression of the VHH-Fc_LS proteins (SEQ ID NO:96-99, 101, 103, 105-109, 111-119) was performed using the ExpiCHO™ Expression System (Thermo Fisher Scientific, Cat. No. A29133). The ExpiCHO-S cells were expanded from a working cell bank and passaged in ExpiCHO expression media according to the manufacturer's instructions. The mixtures for transfection were prepared following the protocol instructions described in the ExpiCHO-S system manual. ExpiCHO cells were cultured for 14 days and the harvest was performed by filtration using 0.22 pm filtration units and DE as filter aid (Sartorius, Cat. No. SDLV-0150-05E0-2) after which they were processed immediately.
The produced antibodies (SEQ ID NO:96-99, 101, 103, 105-109, 111-119) were captured from clarified supernatants using a HiTrap MabSelect PrismA 25 mL column (Cytiva, Cat. nr. 17-5498- 54) on an AKTA Pure 25 FPLC system. The column equilibration and protein binding were performed using 20 mM sodium phosphate with 150 mM NaCl pH 7.4 and for protein elution, 100 mM sodium citrate with 150 mM NaCl pH 3.5 was used. After elution, the peak corresponding to affinity purified antibodies was immediately neutralized with 30 % of 1 M Tris pH 8.0. To polish the samples and to achieve a high purity > 95 % monomeric form, protein samples were loaded onto a HiLoad 26/600 Superdex 200 pg (Cytiva, Cat. nr. 28-9893-36) on an AKTA Pure 25 FPLC system. The column was equilibrated in 50 mM L-Histidine, 150 mM L-Arginine, 150 mM NaCl pH 7.0 for all constructs except for constructs defined by SEQ NO: 117 and SEQ ID: 119 for which pH 6.0 was used. The fractions corresponding to the monomeric antibody form were pooled from a 96 deep well plate into a 50 ml falcon tube and filtered through a 0.22 pm PES membrane (Fisher brand, Cat. nr. 15206869, Lot nr. 2103171806) in a laminar flow chamber. Protein samples were transferred to 50 kDa MWCO spin concentrators (Amicon 50K Cat. nr. UFC905024; Lot nr. 0000187574) for concentration and each round of centrifugation was 10 min at 4000 g, and it was repeated until the desired concentration of 2 mg/mL was achieved.
Characterization of the S2 targeting molecules
Purified VHH-Fc samples were characterized by analytical Size Exclusion Chromatography (SEC) to determine the molecular composition of each sample. After rapid thawing in a warm water bath at
25 °C, 10 min centrifugation at 16,000 g and transfer of supernatant to fresh tubes, 5 pl was injected on an AdvanceBioSECs column, 4 (Agilent) with 2.7 pm porous particle size and 300 A pore size, calibrated with PBS. The mobile phase used for was IxPBS with 0.02 sodium azide. The separation was monitored by absorbance at 280 nm with a 16 nm bandwidth, no reference subtraction.
For additional quality control, proteins were separated on reducing and non-reducing SDS-PAGE. The samples are mixed with 2x concentrate Laemmli buffer containing no DTT (non-reduced conditions) or 200 mM DTT (reducing condition). 3 pg of protein sample are loaded on 12% SDS- PAGE and stain with Coomassie Brilliant Blue.
HEK S transfection and protein purification protocol; Production of YTE variants of VHH-Fc in mammalian cells
HEK293-S cells were transfected with VHH-Fc (S) encoding plasmids using polyethylenimine (PEI). Briefly, suspension-adapted and serum -free HEK293-S cells were seeded at 3 x 106 cells/mL in FreeStyle 293 medium (ThermoFisher Scientific). Next, 4.5 pg of pcDNA3.3 -VHH-Fc plasmid DNA was added to the cells and incubated on a shaking platform at 37°C and 8% CO2, for 5 min. Next, 9 pg of PEI was added to the cultures, and cells were further incubated for 5 h, after which an equal culture volume of Ex-Cell-293 (Sigma) was added to the cells. Transfections were incubated for 4 days, after which cells were pelleted (10’, 300 g) and supernatants were filtered before further use. For purification of the VHH-Fc proteins, supernatants were loaded on a 5 mb MAbSelect SuRe column (GE Healthcare). Unbound proteins were washed away with Mcllvaine buffer, pH 7.2, and bound proteins were eluted using Mcllvaine buffer pH 3. Immediately after elution, proteincontaining fractions were neutralized using 30% (v/v) of a saturated NasPCfi buffer. Next, these fractions were pooled, and loaded on a HiPrep Desalting column for buffer exchange to PBS, pH 7.4. Additionally, huR3DC23-Fc_YTE was expressed in ExpiCHO-S™ cells (ThermoFisher Scientific), according to the manufacturer’s protocol. Briefly, a 50 mb culture of 6 x 106 cells per mb, grown at 37°C and 8% CO2, was transfected with 40 pg of pcDNA3.3-VHH72-Fc plasmid DNA using ExpiFectamine™ CHO reagent. One day after transfection, 300 pL ExpiCHO™ enhancer and 8 mb ExpiCHOTM feed was added to the cells, and cultures were further incubated at 32°C and 5% CO2. Cells were fed a second time day 5 after transfection. Productions were collected as soon as cell viability dropped below 75%.
For purification of the VHH-Fc proteins, supernatants were loaded on a 5 mb MAbSelect SuRe column (GE Healthcare). Unbound proteins were washed away with Mcllvaine buffer pH 7.2, and
bound proteins were eluted using Mcllvaine buffer pH 3. Immediately after elution, proteincontaining fractions were neutralized using 30% (v/v) of a saturated Na3PO4 buffer. Next, these fractions were pooled, and loaded on a HiPrep Desalting column for buffer exchange to PBS pH7.4.
Fed batch production of huR3DC23-Fc LS from stable pool at IL scale
The gene encoding huR3DC23-Fc_LS was codon optimized, synthesized, and cloned into the pXLG6 backbone vector at ATUM’s laboratories. Upon expansion to a density of about 4 x 106 cells/ml, parental CHOExpress™ cells were co-transfected with the expression vector and the pXLG5 helper vector. The stable pool was generated under 50 mg/L puromycin selective pressure (applied daily) and further expanded. The stable pool research cell bank was banked at day 14 when cell viability reached 95%.
The RCB pool was then expanded for protein production at IL scale and cultured until day 12 (cell density 3.5 x 107 cells/mL, cell viability 96%). The supernatant was harvested by centrifugation and clarified by filtration (0.2 pm). The protein was further purified by Protein A using MabSelect SuRe LX resin. Consecutive washed were performed with 20 mM sodium phosphate and 110 mM NaCl at pH 7.2; 100 mM sodium acetate and 500 mM NaCl at pH 5.5; and 20 mM sodium phosphate at pH 7.2. The eluate in 100 mM sodium acetate pH 3.5 was neutralized to pH 7.0 by addition of 1 M Tris pH 11 (10%v/v). After filter sterilization (0.22 pm), the protein was aliquoted at 2 mg/ml.
In vivo viral challenge in K18-hACE2 mice
K18-hACE2 mice: B6.Cg-Tg(K18-ACE2)2Prlmn/J (7-9 weeks of age) were purchased from The Jackson Laboratory and bred in house under specific-pathogen-free (SPF) conditions. SARS-CoV-2 infections were performed under biosafety level 3 (BSL3) conditions. Antibody treatment was performed by intraperitoneal injection using a volume of 100 pl. Animals were anesthetized by isoflurane inhalation and 3xl02 PFU of SARS-CoV-2 D614G variant virus (SARS-CoV- 2/human/FRA/702/2020, obtained from the European Virus Archive) was administered by intratracheal instillation. Animals were monitored on a daily base by a blind observer who measured weight change and scored for humane endpoints: Hunchback (1 point) Piloerection (1 point), less movement upon opening cage (1 point), motionless upon touching (2 points), neurological symptoms (shaking, balance, 3 points), heavy breathing (3 points). Mice that lost more than 25% of their initial body weight or reached a humane endpoint with a score of 5 points were euthanized.
Titration of SARS-CoV-2 viral titer in lung homogenates of mice
After dissection, the right lung lobes were stored at -80°C. For the quantification of the lung viral
titer the lung samples were homogenized using a Precellys Evolution tissue homogenizer (Bertin- technologies). The lung homogenates were cleared by centrifugation (1,000 x g) for 15 min at 4°C and used to determine the viral titer by plaque assay on VeroE6-TMPRSS2 cells in duplicate using 12-well plates. After addition of the lung homogenate dilution series to the cells, the plates were incubated at 37°C for 2 hours. Subsequently the cells were washed twice and incubated in medium containing Avicel at a concentration of 0.3% (w/v). After 2 days of incubation at 37°C, the overlays were removed, the cells were fixed with 3.7% paraformaldehyde (PFA) and stained with 0.5% crystal -violet dye to visualize the viral plaques.
To quantify the SARS-CoV-2 viral RNA levels, RNA was prepared from the lung homogenates and analyzed by qRT-PCR. cDNA was prepared using the iScript™ cDNA Synthesis Kit and random hexamer primers. QPCR was performed using the SARS-CoV-2 Research Use Only (RUO) qPCR Primers & Probes kit (IDT) according to the manufacturer’s instructions.
Syrian Golden hamster challenge model
9-to 10-weeks-old male Syrian golden hamsters (Mesocricetus auratus) weighing 89.8 g to 132.3 g were obtained from Janvier (France). Six (example 2) or five to eight (example 2) hamsters per group were infected intranasally with 10A2.0 50% tissue culture infectious dose (TCID50)/dose SARS-CoV-2 (BetaCoV/Munich/BavPat 1/2020 of Wuhan lineage) in a total dose volume of 100 pl, divided equally over both nostrils. In a first study (example 2), XVR013, at the indicated doses, palivizumab (10 mg/kg), or bebtelovimab (10 mg/kg) were administered by intraperitoneal injection either 4 hours after the SARS-CoV2 challenge (therapeutic setting) or approximately 24 hours prior to infection (prophylactic setting). In a 2ndsecond study (example 16), XVR013m, XVR014 or XVR012 (1: 10 ratio of XVR013m:XVR014) were dosed, at the dose levels indicated, by intraperitoneal injection, 4 hours after the SARS-CoV2 challenge (therapeutic setting). In both studies, the irrelevant antibody Palivizumab (Synagis, anti-RSV antibody) was used as a negative control, while bebtelovimab was used as a positive control. The huR3DC23-Fc_LS used in the hamster study was produced from stable cell pools. Hamsters were monitored daily for behavior, appearance and body weight. On day 4 post-infection, animals were euthanized. At the time of necropsy, gross pathology was performed and abnormalities were recorded. Samples from the right lung lobes were collected and frozen for virological analysis. To determine virus titers, quadruplicate
10-fold serial dilutions were used in confluent layers of Vero E6 cells. To this end, serial dilutions of the samples (lung tissue homogenates) were incubated on Vero E6 monolayers for 1 hour at 37°C. Vero E6 monolayers were then washed and incubated for 5 or 6 days at 37°C, after which plates were
stained and scored based on cytopathic effect (CPE) by using the vitality marker WST8 (colorimetric readout). Viral titers (LoglO TCID50/g) were calculated using the Spearman-Karber method (Karber. 1931 Archiv F. Experiment. Pathol, u. Pharmakoi. 162:480-483). For the viral titration, the lower limit of detection (LLOD) ranged between 1.1 and 1.3 loglO TCID/g. To detect viral RNA, lung tissue homogenates were used. RNA was isolated and Taqman RT-qPCR was performed. The number of copies (LoglO CP/g) in the different samples was calculated against a standard included in each run. For the viral RNA in lung tissue, the LLOD was 3.5 LoglO CP/g. Blood samples were collected prior to the start of the study on day -2 (~ 200 pl blood was collected for serum under isoflurane anesthesia) and on day 4 post-infection (p.i.) at time of necropsy for pharmacokinetics analysis. Blood samples for serum were immediately transferred to appropriate tubes containing a clot activator. Serum was collected and stored frozen. To inactivate any potential infectious material present and to allow the testing of the serum samples in a BSL-2 environment, serum on day 4 postinfection was heat-treated at 56 °C for 30 minutes.
Generation of spike protein expression vectors for the production of VSVdelG pseudovirus particles expressing spike proteins containing RBD mutations ofSARS-CoV-2 variants
The pCGl expression vector for the SARS-CoV-2 spike protein containing the D614G mutation was generated from the pCGl-SARS-2-Sdell8 vector by introducing the specific RBD mutations via QuickChange mutagenesis using appropriate primers, according to the manufacturer’s instructions (Aligent).
The pCGl expression vector forthe Gamma variant was generated from the pCGl-SARS-2-D614G Sdell8 vector by ordering Gblocks corresponding to the Gamma SARS-CoV-2 amino acid sequence L7-V1189 containing the Gamma mutations (L18F, T20N, P26S, D138Y, R190S, K417T, E484K, N501Y, D614G, H655Y, T1027I, VI 176F), and cloning the Gblocks into the pCGl-SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Eta variant was generated from the pCGl-SARS-2-D614G Sdell8 vector by ordering a Gblock corresponding to the Eta SARS-CoV-2 amino acid sequence S45-P894 containing the Etamutations (Q52R, A67V, H69-, V70-, Y144-, E484K, D614G, Q677H, F888L), and cloning the Gblock into the pCGl-SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Iota variant was generated from the pCGl-SARS-2-D614G Sdell8 vector by ordering a Gblock corresponding to the Iota SARS-CoV-2 amino acid sequence M1-V267 containing the Iota mutations (L5F, T95I, D253G), and cloning the Gblock into the pCGl- SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the B. 1.617.3 variant was generated from the pCGl-SARS-2- D614G Sdel 18 vector by ordering a Gblock corresponding to the B. 1.617.3 SARS-CoV-2 amino acid sequence L10-A688 containing the B. 1.617.3 mutations (T19R, L452R, E484Q, D614G, P681R), and cloning the Gblock into the pCGl-SARS-2D614G Sdel 18 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Mu variant was generated from the pCGl-SARS-2-D614G Sdel 18 vector by ordering a Gblock corresponding to the Mu SARS-CoV-2 amino acid sequence D80-Q965 containing the Mu mutations (T95I, Y144S, Y145N, R346K, E484K, N501Y, D614G, P681H, D950N), and cloning the Gblock into the pCGl-SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Zeta variant was generated from the pCGl-SARS-2-D614G Sdel 18 vector by ordering a Gblock corresponding to the Zeta SARS-CoV-2 amino acid sequence R466-L1193 containing the Zeta mutations (E484K, D614G, VI 176F), and cloning the Gblock into the pCGl-SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Omicron BA.5 variant was generated from the pCGl-SARS-2- BA.2 Sdel 18 vector by introducing the H69-, V70- deletions and the L452R, F486V, R493Q mutations via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
The pCGl expression vector for the Omicron XBB.1.5 (-G252V) variant was generated from the pCGl-SARS-2-XBB Sdell8 vector by introducing the F486P mutation via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent). The spike protein sequence of the XBB.1.5 (-G252V) variant was based on the sequence published on the European Centre for Disease Prevention and Control (ECDC) website. This XBB 1.5 spike variant sequence lacked the G252V mutation, present in approximately 80% of the circulating XBB.1.5 viruses.
The pCGl expression vector for the Omicron XBB. 1.5 variant (including the G252V mutation) was generated from the pCGl-SARS-2-XBB.1.5 (-G252V) Sdell8 vector by introducing the G252V mutation via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
The pCGl expression vector for the Omicron CH.1.1 variant was generated from the pCGl-SARS- 2-XBB.1.5 Sdell8 vector by ordering a Gblock corresponding to the CH.1.1 SARS-CoV-2 amino acid sequence I65-Q503 containing the CH.1.1 mutations (G142D, K147E, W152R, F157L, I210V, V213G, G257S, Q339H, R346T, S371F, S373P, S375F, T376A, D405N, R408S, K417N, N440K, K444T, G446S, L452R, N460K, S477N, T478K, E484A, F486S), while reverting other XBB. 1.5 specific mutations, and cloning the Gblock into the pCGl-SARS-2D614G Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Omicron EG.5.1 variant was generated from the pCGl-SARS- 2 -XBB. 1.5 Sdell8 vector by introducing the F456L, Q52H mutations via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
The pCGl expression vector for the Omicron HV.l variant was generated from the pCGl-SARS-2- XBB.1.5 Sdell8 vector by ordering a Gblock corresponding to the HV. l SARS-CoV-2 amino acid sequence V39-F460 containing the HV. 1 mutations (Q52H, F157L, L452R, F456L) and all XBB. 1.5 specific mutations, and cloning the Gblock into the pCGl -SARS-2XBB .1.5 Sdel 18 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Omicron BA.2.86.1 variant was generated from the pCGl- SARS-2-XBB.1.5 Sdell8 vector by ordering two Gblock’s corresponding to the BA.2.86.1 SARS- CoV-2 amino acid sequence S13-L1144 containing the BA.2.86.1 mutations (T19I, R21T, L24S, P25-, P26-, A27-, S50L, H69-, V70-, , V127F, G142D, Y144-, , F157S, R158G, N211I, L212-, V213G, L216F, H245N, A264D, I332V, G339H, K356T, S371F, S373P, S375F, T376A, R403K, D405N, R408S, K417N, N440K, V445H, G446S, N450D, L452W, N460K, S477N, T478K, N481K, V483-, E484K, F486P, Q498R, N501Y, Y505H, E554K, A570V, D614G, P621S, H655Y, N679K, P681R, N764K, D796Y, S939F, Q954H, N969K, P1143L), while reverting the XBB. 1.5 specific mutations and cloning the Gblock’s into the pCGl-SARS-2XBB.1.5 Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Omicron XBB.2.3 variant was generated from the pCGl-SARS- 2 -XBB. 1.5 (-G252V) Sdell8 vector by introducing the D253G, P521S mutations via QuickChange
mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
The pCGl expression vector for the Omicron XBB. 1.16 variant was generated from the pCGl- SARS-2-XBB.1.5 Sdell8 vector by introducing the E180V, T478R mutations via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
The pCGl expression vector for the Omicron HK.3 variant was generated from the pCGl-SARS-2- EG.5.1 Sdell8 vector by introducing the L455F mutation via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
The pCGl expression vector for the Omicron FL. 1.5. 1 variant was generated from the pCGl-SARS- 2-XBB.1.5 Sdell8 vector by introducing the F456L, T478R, A701V mutations via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
The pCGl expression vector for the Omicron BA.2.74 variant was generated from the pCGl-SARS- 2-XBB.1.5 Sdell8 vector by ordering a Gblock corresponding to the BA.2.74 SARS-CoV-2 amino acid sequence T70-G501 containing the BA.2.74 mutations (G142D, V213G, G339D, R346T, S371F, S373P, S375F, T376A, D405N, R408S, K417N, N440K, L452M, S477N, T478K, E484A, Q493R), while reverting the XBB.1.5 specific mutations, and cloning the Gblock into the pCGl- SARS-2XBB.1.5 Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Omicron JN. l variant was generated from the pCGl-SARS-2- BA.2.86.1 Sdell8 vector by introducing the L455S mutation via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
The pCGl expression vector for the Omicron KP.3 variant was generated from the pCGl-SARS-2- JN. l Sdell8 vector by ordering a Gblock corresponding to the KP.3 SARS-CoV-2 amino acid sequence H438-D1110 containing the KP.3 mutations (F456L, Q493E, V1104L), and cloning the Gblock into the pCGl-SARS-2JN. l Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Omicron KP.1.1 variant was generated from the pCGl-SARS- 2 -KP.3 Sdell8 vector by ordering a Gblock corresponding to the KP.1.1 SARS-CoV-2 amino acid sequence P330-S1089 containing the KP.1.1 mutations (R346T and K1086R), while reverting KP.3
specific mutations and cloning the Gblock into the pCGl-SARS-2-KP.3 Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Omicron KP.3. 1. 1 variant was generated from the pCGl-SARS- 2-KP.3 Sdell8 vector by introducing the S31- deletion via QuickChange mutagenesis using appropriate primers (ordered at IDT), according to the manufacturer’s instructions (Agilent).
The pCGl expression vector for the Omicron KP.2.3 variant was generated from the pCGl-SARS- 2-KP.3 Sdell8 vector by ordering a Gblock corresponding to the KP.2.3 SARS-CoV-2 amino acid sequence L10-V494 containing the KP.2.3 mutations (S31-, H146Q, R346T), while reverting the KP.3 specific mutations, and cloning the Gblock into the pCGl-SARS-2KP.3 Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Omicron LB. 1 variant was generated from the pCGl-SARS-2- JN. l Sdell8 vector by ordering a Gblock corresponding to the LB. l SARS-CoV-2 amino acid sequence V11-A467 containing the LB.l mutations (S31-, Q183H, R346T and F456L) and cloning the Gblock into the pCGl-SARS-2JN. l Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
The pCGl expression vector for the Omicron XEC variant was generated from the pCGl-SARS-2- KP.3 Sdell8 vector by ordering a Gblock corresponding to the XEC SARS-CoV-2 amino acid sequence V3-D75 containing the XEC mutations (T22N and F59S) and cloning the Gblock into the pCGl-SARS-2-KP.3 Sdell8 vector via Gibson assembly according to the manufacturer’s instructions (New England Biolabs).
After sequencing, clones containing the correct spike coding sequence were prepared using the Qiagen plasmid kit. Before usage the spike coding sequence of the prepared pCGl vectors was confirmed by Sanger sequencing.
Pseudovirus neutralization assay
Pseudovirus neutralization assays were conducted using vesicular stomatitis virus (VSV) particles pseudotyped with the spike of SARS-CoV-2 of either the D614G reference strain, Pre-Omicron variants Gamma, Eta, Iota, Mu, Zeta, B.1.617.3 or Omicron variants BA.5, XBB.1.5 (-G252V), XBB.1.5, XBB.2.3, XBB.1.16, CH.1.1, EG.5.1, HK.3, HV. l, BA.2.74, BA.2.86.1, FL1.5.1, JN. l, KP.2.3, KP.3, KP.3.1.1, KP.1.1, LB. l or XEC. A negative control antibody (palivizumab) or isotype control antibody, and/or a positive control (R3DC23-Fc_LS (XVR013), sotrovimab and/or bebtelovimab) were included in the assay. For each variant one, two or three independent
experiments were performed. First, replication-deficient VSV pseudotyped viruses were generated. Briefly, HEK 293T cells, transfected with SARS-CoV-2 Spike were inoculated with a replication deficient VSV vector containing enhanced green fluorescent protein (GFP) and firefly luciferase expression cassettes. After a 2 h incubation at 37°C, the inoculum was removed, the cells were washed with PBS and incubated in media supplemented with an anti -VSV G monoclonal antibody for 16 hours. Afterwards, pseudoviruses were harvested and clarified by centrifugation. Pseudoviruses were incubated for 1 hour at 37°C with different dilutions of the molecules, the positive control (XVR013, sotrovimab and/or bebtelovimab) and/or the negative control (palivizumab or isotype control) antibody. Next, the incubated pseudoviruses with molecules or controls were added to the subconfluent monolayers of Vero E6 cells for 16 hours at 37°C. Afterwards, the cells were lysed using passive lysis buffer. The transduction efficiency was quantified by measuring the GFP fluorescence in the prepared cell lysates using a Tecan Infinite 200 Pro plate reader. The GFP fluorescence was normalised against the GFP fluorescent of the noninfected cells and the virus infected cells. The mean IC50 values were calculated based on one, two or three independent experiments as indicated by nonlinear regression curve fitting (log(inhibitor) versus normalised response (four parameters)).
Live virus assay
The neutralization potency of the S2 targeting molecules was tested in a live virus assay (microneutralization method). SARS-CoV-2 viruses belonging to different lineages (D614G, Alpha, Beta, Gamma, Delta, Epsilon B. 1.427, Epsilon B..429, Zeta, Kappa, Omicron BA. 1, Omicron BA.2 and Omicron BA.5) were isolated from nasopharyngeal swabs taken from patients/travelers between January 2020 and July 2022. More specifically, the following clinical isolates were used: SARS- CoV-2 Isolate BavPatl/2020/Germany (09 Feb 2020), Alpha variant hCoV- 19/USA/CA_CDC_5574/2020, Beta variant hCoV-19/South Africa/KRISP-K005325/2020, Gamma variant hCoV-19/Japan/TY7-503/2021, Delta variant hCoV-19/USA/MD-HP05647/2021, Epsilon B. 1.427 variant hCoV-19/USA/CA/VRLC009/2021, Epsilon B.1.429 variant hCoV- 19/USA/CA/VRLCO 14/2021, Zeta variant hCoV-19/USA/CA/VRLC012/2021, Kappa variant hCoV-19/USA/CA-Stanford-15_S02/2021, Omicron BA. l variant hCoV-19/Netherlands/NH- RIVM-72291/2021, Omicron BA.2 variant hCoV 19/USA/CO-CDPHE-2102544747/2021 and Omicron BA.5 variant Clinical isolate hCovl9/NL/VCB-20220714-2/2022. Dose-dependent neutralization of the molecules, the positive controls (bebtelovimab biosimilar, sotrovimab
biosimilar or XVR013) or a negative control (palivizumab or isotype control) were assessed in an authentic virus neutralization assay.
Briefly, 2 or 5 -fold dilutions of the molecules were incubated for 1 hour at room temperature with the following amount of plaque-forming units (PFUs) of the virus: 118, 89, 103, 56, 87, 132, 247, 129, 185, 108, 143 and 103 PFU/well for D614G, Alpha, Beta, Gamma, Delta, Epsilon B. 1.427, Epsilon B. 1.429, Zeta, Kappa, Omicron BA. 1, Omicron BA.2 and Omicron BA.5, respectively. Afterwards, the Vero-E6 cell monolayer (25000 cells pre-seeded 24 hours in advance) was inoculated with virus antibody mixtures for 1 hour at 37°C. In a next step, the inoculum was removed and cells were incubated at 37°C with infection medium (up to 18-24 hours post-infection). Afterwards, the SARS-CoV2 infected cells were fixed and immunostained with a SARS-CoV nucleocapsid antibody (Sino Biological, Catalogue number: 40143-MM05), followed by HRP -conjugated goat anti-mouse IgG (H+L) secondary antibody (Invitrogen, catalogue number A16072). Spots (infected cells) were counted using Immunospot Image Analyser. For each molecule, the compound concentration showing 50, 80 or 90 % reduction in infection (IC50, ICso or IC90) was calculated based on the Zielinska method. The geometric mean values were calculated based on three independent runs.
Binding of S2 targeting molecules to the spike of Khosta-2, a clade 3 virus from the Sarbecovirus subgenus
2.105 HEK293T cells, pre-seeded for 1 day in 6 well plates, were transfected with 800 ng of vector expressing GFP fused to the spike of either Khosta-2 sarbecovirus (Seifert et al. (2022 PLoS Pathog 18(9):e 1010828) or SARS-CoV-2 variant D614G, complexed to polyethylenimine (PEI) in a 1:3 ratio. As controls, cells were either transfected with a vector expressing GFP only, or left untransfected. After 2 days of incubation at 37°C and 5% CO2, cells were harvested, blocked with PBS + 1% BSA, then incubated for Ih at 4°C with XVR013, XVR013m, XVR014, positive control (sotrovimab) or negative control (palivizumab, rituximab or an isotype control PCRV). To detect binding of the S 1 and/or S2 targeting molecules or controls, cells were stained with a phycoerythrin (PE) anti-human IgG Fc detection antibody (1/200) and eFluor780 (1/400) fixable viability dye. Cells were fixed and analyzed by flow cytometry. GPF-positive cells were considered as transfected.
Affinity assessment of S2 targeting molecules on FcRn
To determine the binding kinetics for FcRn, surface plasmon resonance (SPR) analysis of the S2 targeting molecules binding to purified recombinant human FcRn/FCGRT-B2M protein was performed on a Biacore 8 K+ instrument. Sotrovimab biosimilar and XVR013 were used as controls
in the assay. In brief, the experimental set-up was as follows: the S2 targeting molecules or control antibodies were immobilized at low density on a CM5 sensor chip (Cytiva) and the human FcRn at pH6.0 or pH7.4 was injected in solution in 2-fold serial dilutions in the range of 1000 to 0.98 nM. Analyte injections were performed in running buffer during 2 minutes at 30 pl/min; assay runs with 0 nM concentration were included as blank reference. SPR running buffer consisted of lx HBS-EP at pH 6.0 or 7.4. A multicycle kinetics protocol was applied (off-rate measurement 90 seconds). Data was analysed using the Biacore Insight Evaluation Software. Kinetic parameters were calculated using the 1: 1 stoichiometry Langmuir binding model and steady state affinity binding model.
Non-specific binding of S2 targeting molecules to the surface of mammalian cells
The non-specific binding of the S2 targeting molecules to the surface of mammalian (HEK293) cells was investigated with a flow cytometry experiment. Sotrovimab biosimilar was used as negative control antibody in the assay and XVR013 was used as positive control. All procedures were performed on ice. The cells were washed once with PBS and blocked with 1% BSA. Afterwards, the cells were incubated with two concentrations (10 and 100 pg/mL) of the S2 targeting molecules or controls (Sotrovimab Biosimilar and XVR013) for 90 minutes and subsequently washed 3 times with PBS containing 1% BSA. Next, the secondary antibody (anti-human IgG AF633) was incubated for 30 minutes. Binding of the molecules or controls was detected using an anti-human IgG AF633 antibody and dead cells were stained using live/dead stain (Invitrogen). Following 3 washes with PBS containing 0.5% BSA, the cells were analyzed by flow cytometry. The median fluorescence intensity of the AF633-conjugated anti-human IgG was measured to detect the binding of the molecules or controls to HEK293 cells.
In silico immunogenicity analysis
An in silico immunogenicity assessment has been performed to predict the presence of putative HLA class II restricted epitopes, a.k.a. T helper (Th) cell epitopes (Lonza Biologies, UK). Lonza’s Epibase platform was applied to analyze the HLA binding specificities of all 10-mer peptides derived from a molecule. For molecules RNGL-Fc_LS, RAGL-Fc_LS, E2NGL-Fc_LS, E2AGL-Fc_LS, QRAGL- Fc_LS, ERAGL-Fc_LS, E10RAGL-Fc_LS, E81RNGL-Fc_LS, E81RAGL-Fc_LS, Q75-Fc_LS, E75-Fc_LS, E10E81-Fc_LS, D16-Fc_LS, T19D79-Fc_LS, TQD-Fc_LS, Q75R21Y-Fc_LS, CBV1- Fc_LS, CBV2-Fc_LS, CBV3-Fc_LS, CBV4-Fc_LS and XVR013, the DRB1 score was calculated. The DRB 1 score is an approximate score representing a potential immunogenic risk for a peptide and is calculated from the population frequencies of the affected allotypes.
Hydrophobic interaction chromatography (HIC) assay
Apparent hydrophobicity was assessed using a hydrophobic interaction chromatography (HIC) assay employing a Dionex ProPac HIC-10 column, 100 mm x 4.6 mm (Thermo Fisher 063655), containing a stationary phase consisting of a mixed population of ethyl and amide functional groups bonded to silica. All separations were carried out on an Agilent 1100/1260 HPLC equipped with a UV/VIS detector. The column temperature was maintained at 25 °C throughout the run and the flow rate was 800 pL/min. The mobile phases used for HIC were (A) 1.6 M ammonium sulfate and 50 mM phosphate pH 6.0 (variants CBV2-Fc_LS and CBV4-Fc_LS) or pH 7.0, and (B) 50 mM phosphate pH 6.0 (variants CBV2-Fc_LS and CBV4-Fc_LS) or pH 7.0.
Prior to HIC analysis, the S2 targeting molecules were concentration adjusted to Img/mL in 50 mM L-Histidine, 150 mM L-Arginine, pH 6.0 (variants CBV2-Fc_LS and CBV4-Fc_LS) or pH 7.0, and fdtered through a 0.45 pM fdter. The proteins (50 pL) were diluted 1: 1 with buffer A to a final ammonium sulfate concentration of 800 mM, mixed and 40 pL of the diluted proteins were injected onto the column. Following a 5 min hold at 50% A and 50% B, bound protein was eluted using a linear gradient from 50 to 100% B in 50 min followed by 5 min hold at 100% B. The column was washed with 100% B, followed by 50 mM ammonium acetate pH 5 and re-equilibration in 50% B for 10 min prior to the next sample. The separation was monitored by absorbance at 214 nm, no reference subtraction.
Strong Cation Exchange, hydrophilic interaction chromatography (SCX) assay
The charge proteins variants were assessed using a Strong Cation Exchange Hydrophilic interaction chromatography (SCX) assay employing a MabPac SCX-10 column (Thermo Fisher 063655), containing a stationary phase consisting of an ethylvinylbenzene-divinylbenzene copolymer non- porous beads coated with a layer of highly hydrophilic material consisting sulfonic acid functional groups. All separations were carried out on an Agilent 1100/1260 HPLC equipped with a UV/VIS detector at fixed flow rate of 500 pL/min. The mobile phases used for SCX were (A) 20 mM MES pH 5.6, and (B) 20 mM HEPES and 120 mM NaCl pH 8.0.
Prior to SCX analysis, the S2 targeting molecules were formulated at Img/mL in 50 mM L-Histidine, 150 mM L-Arginine, 0.02% polysorbate 80, 0.02% sodium azide, pH 6.0 (variants CBV2-Fc_LS and CBV4-Fc_LS) or pH 7.0, and filtered through a 0.45 pM filter. The proteins (20 pL) were diluted 1: 1. Proteins (20 pL sample pre-filtered on 0.45 pm filter) were injected onto the column. Following a 2 min hold at 100% A, bound protein was eluted using linear gradient from 100% A to 100% B in
18 min followed by 2 min hold at 100% B, and 4 min hold at 100% A. The separation was monitored by absorbance at 214 nm.
Di fferential Scanning Fluorimetry (DSF), Static Light Scattering (SLS) and Dynamic Light Scatting (PLS) assay
The protein stability was assessed by the determination of their respective melting temperature (Tonset and Tm), aggregation temperatures (Tagg) and particles sizing (AVG-Z, average diameter) using an Unchained Labs UNCLE system running with the Supplier Software v4.01. A temperature gradient of 25 -95 °C was applied and the protein melting temperatures were detected by intrinsic fluorescence (barycentric mean of spectrum, BCM), and protein aggregation propensity by static light scattering with detection at 266 nm (Tagg 266) and 473 nm (Tagg 473). Prior loading in the system, the S2 targeting molecules candidates were formulated at Img/mL in 50 mM L-Histidine, 150 mM L-Arginine, 0.02% polysorbate 80, 0.02% sodium azide, pH 6.0 (variants CBV2-Fc_LS and CBV4-Fc_LS) or pH 7.0, and filtered through a 0.45 pM filter.
Ageing study
To assess the developability of the S2 targeting molecules, the stability of each variant was evaluated in an accelerated stress study.
All proteins were formulated at 1 mg/ml in 50 mM L-His, 150 mM L-Arg at either pH 6 (CBV2- Fc_LS and CBV4-Fc_LS) or pH 7 (RNGL-Fc_LS, RAGL-Fc_LS, E2NGL-Fc_LS, E2AGL-Fc_LS, QRAGL-Fc_LS, ERAGL-Fc_LS, E10RAGL-Fc_LS, E81RNGL-Fc_LS, E81RAGL-Fc_LS, Q75- Fc_LS, E75-Fc_LS, E10E81-Fc_LS, D16-Fc_LS, T19D79-Fc_LS, TQD-Fc_LS, Q75R21Y-Fc_LS, CBVl-Fc_LS and CBV3-Fc_LS), supplemented with 0.02% polysorbate-80, and 0.02% NaN3 to prevent microbial growth. After dilution and filtering (0.22 pm), each sample was split into two 750 pl aliquots (one for each time point) that were snap-frozen in liquid nitrogen and stored at -80 °C. Samples were rapidly thawed in a water bath at room temperature, followed by centrifugation at 16,000 xg for 10 min to remove potential freeze-thaw aggregates. Supernatant was transferred (only 540 pl to avoid disturbing any freeze-thaw-aggregate pellet) and split into three replicates (120 pl each) in sterile pyrogen-free polypropylene 2 ml screw-cap tubes. Samples were either analyzed immediately afterwards, or used for 14 days accelerated ageing at 37 °C, in a test tube rotatory mixer at 10 rpm, and analyzed subsequently.
Variants CBV2-Fc_LS and CBV4-Fc_LS were formulated at Img/mL in 50 mM L-Histidine, 150 mM L-Arginine, 0.02% polysorbate-80, 0.02 sodium azide at pH6.0, all other variants were
formulated at Img/mL in 50 mM L-Histidine, 150 mM L-Arginine, 0.02% polysorbate-80, 0.02 sodium azide at pH7.0. The study samples were prepared in triplicate. 120 pL sample was filled in 2mL screw-cap polypropylene tube and incubated for 2 weeks at 37°C under mild agitation (10 rpm) in turning wheel. All of the variants samples were characterized and analyzed in parallel at time point 0 and after 14 days stress. By
Visual inspection to assess presence of visible particles
UV/VIS spectrometry performed in an Unchain Labs Lunatic system; total protein quantification at 280 nm complemented by background absorbance at higher wavelengths Analytical SEC to assess degradation/multimerization/aggregation
Analytical SCX on hydrophiic resin to detect charges variants
SDS-PAGE in non-reduced and reduced condition to assess degradation/isomerization/aggregation
Pharmacokinetic (PK) study
XVR014, S2 targeting molecules and a reference molecule (XVR013) were tested in human neonatal Fc receptor (hFcRn) tg32 SCID mice (B6.Cg-FcgrttmlDcr Prkdcscid Tg(FCGRT)32Dcr/DcrJ). Six animals per group were included and two dose levels (2 mg/kg and 10 mg/kg) were tested using a single intravenous (IV) administration for evaluating XVR014. Five male animals were included per group and one dose level (10 mg/kg) was tested using a single intravenous (IV) administration for testing the S2 targeting molecules and XVR013.
Serial blood samples were collected, processed to plasma and then analyzed in a qualified human IgG Fc ELISA assay. Individual PK profiles were fed into a two-compartment population PK model with linear elimination. Performance was checked based on visual checks (individual predictions, observations vs individual predictions, residuals vs time...) and on the quality of the model parameters estimates (small residual error). Allometric scaling was performed to project human PK parameters. Clearance (CL) and volume of distribution of the peripheral compartment (V2) were used to compute the half-life beta (terminal phase).
Evaluation of pre-existing anti-drug-antibodies
The seroprevalence of pre-existing anti-drug antibodies (ADAs) in drug-naive individuals against a test compound was evaluated by an ELISA method. The biosimilar caplacizumab (Proteogenix) was used as negative control and the biosimilar TAS266 (Proteogenix) as a positive control. Briefly, an ELISA method was developed to facilitate the screening of a panel of 70 adult human sera samples
(35 male and 35 female of Caucasian, Hispanic, Black or other origin between 18 and 65 years old). Plates were pre-coated with a test compound or a control. Following the addition of human serum, the presence of pre-existing ADAs was detected using an anti-HRP tagged anti-light chain IgG (Fisher Scientific). For each dataset, a cut-point was calculated as mean + 1.645*SD (if data were not normally distributed, the cut-point was calculated using the 95th percentile). This cut-point was used to calculate the percentage of healthy adult human donors with pre-existing ADAs against the test compound.
Phage mutagenesis library construction
Saturation mutagenesis primers were designed according to selected mutation sites and custom produced at Genewiz. hR3_DC23 (SEQ ID NO:2) fragments to be mutated were amplified and ligated by overlap PCR (Q5® Hot Start High-Fidelity 2X Master Mix, New England Biolabs #M0494), followed by gel recovery (QIAquick PCR Purification Kit, Qiagen # 28106). Phage vector plasmids and PCR products were digested by Sfi I in rCutSmart® Buffer (New England Biolabs # RO 123 and B7204S) and subsequently purified. The purified vector and PCR products were ligated with T4 DNA ligase (New England Biolabs #M02020M and #B0202S), and ligated products were desalted by Gel and PCR Clean-up kit (QIAquick PCR Purification Kit, Qiagen # 28106).
Ligated products were electroporated into E.coli SS320 (MC1061F’) competent cells (Lucigen® Corporation #60512-1), and library size was calculated one day later by counting the colonies spread on ampicillin-containing plates (Amp plates).
Phages were rescued by infecting the culture of transformed MC1061F’ cells. Library bacteria were inoculated in 2x Yeast Extract Tryptone (YT) (containing ampicillin, 2% glucose) medium and cultured at 37°C until an OD value of 0.6. Then helper phage was added, and the mixtures were incubated at 37°C for 1 hour. Induction was performed overnight at 30°C by the addition of 0.5 mM isopropyl P-D-l -thiogalactopyranoside (IPTG) and 50 pg/ml kanamycin. The next day, after centrifugation, the library phages were precipitated with 20% PEG8000/ 2.5 M NaCl. The phages were resuspended in 1 X PBS, and the concentration was determined. The resuspended phages were used as input for the first round of panning
Phage Panning
Phage libraries were pre-incubated with streptavidin (SA)-coated magnetic beads (SA beads, ThermoFisher, 11206) to remove non-specific binders, and then incubated with SA-Bio SARS-Cov2 magnetic beads at room temperature, followed by washing 15 times with 1 x PBST. After washing,
20 ml of MC106F cells were infected at 37°C for 30 minutes. The infected cells were serially diluted and spread on 2xYT plates (containing 100 ug/ml ampicillin) and cultured at 37°C overnight to form colonies. Then 20 microliters of M13K07 helper phage at 5E12/ml were added to the above infected cells and cultured at 37°C for 30 minutes. Finally, 80 ml of 2xYT (containing 100 ug/ml ampicillin, 50 ug/ml kanamycin, and 1 mM IPTG) was added and incubated overnight at 30 °C. 100 ml of the overnight culture was centrifuged, fdtered through a 0.22 pm filter, and 20 ml of 20% PEG8000/2.5M NaCl was added to the supernatant, followed by centrifugation at 4°C for 30 minutes. The phage pellet obtained after centrifugation was resuspended in 1 ml of 1 x PBS, the debris was discarded by centrifugation at high speed for 5 minutes, and the resulting supernatant was transferred to a new tube and prepared for the second and third rounds of panning.
The processes of the second and third rounds of panning were the same as the previous steps except specific modification of certain conditions as indicated.
Screening ELISA
Overnight colonies were picked into 96 deep-well plates with 2xYT medium (containing ampicillin, 2% glucose) and grown for 6 hours at 37°C. The supernatant was discarded by centrifugation, the pellet bacteria were resuspended in new 2xYT medium (containing ampicillin, ImM IPTG) to induce antibody expression overnight at 30°C, and the supernatant (bacterial periplasm extracts, PPE) was harvested for ELISA experiments after centrifugation.
The supernatant expressing VHH was tested for binding to biotinylated SARS-CoV-2 spike protein (ACROBiosystems #SPN-C82Ey). High-binding ELISA plates were coated with biotinylated SARS- CoV-2 spike protein and additional plates coated with streptavidin protein as a non-specific control, 100 pl/well at 4°C overnight. The next day the plates were washed three times with wash buffer (0.05% PBST). The plates were washed three times after being blocked with 5% milk-PBS for 1 hour. Subsequently, 100 pl of the PPE were added into each well, the plates were incubated at room temperature for 1 hour and washed six times. Next, 100 pl of horseshoe radish peroxidase (HRP)- conjugated secondary antibody (anti-Flag-HRP, 1:5000) was added to each well, the plates were incubated at room temperature for 1 hour and were washed six times. Color development was performed by adding 100 pl of tetramethylbenzidine (TMB) for 10 minutes, and then terminated by ELISA stop solution. The absorbance at 450 nm OD450 was read on a SpectraMax instrument (Molecular Devices).
Sequencing and production ofVHH-monoFc HCAbs
Sanger sequencing of the VHH genes of the selected clones was performed by Genewiz Inc., part of Azenta Life Sciences (https://www.genewiz.com/).
The sequences encoding variable domains of llama/alpaca were synthesized and cloned into pTT5 vector (licensed from National Research Council of Canada, NRC) encoding engineered monomeric human IgGl Fc region (SEQ ID NO:246) to generate the plasmids encoding full-length heavy-chain- only antibodies by using conventional recombinant DNA technology (Tsingke Biotechnologies).
The plasmids encoding full-length heavy-chain-only antibodies were transfected into mammalian host cells (such as Expi293F human cells), and the purified antibodies were obtained by using conventional recombinant protein expression and purification techniques. Specifically, Expi293F™ cells (Gibco, #A 14527) were subcultured and expanded in MetaCell®HEK293-100 expression medium (Cellplus Bio, #L2000-1000). On the day prior to transfection, the viable cell density was adjusted to 3-4 x 106 cells/mL and the cells were allowed to grow in 37°C, 8% CO2 shaker overnight. On the day of transfection, the cell viability and viable cell density reached to 95-99% and approximately 7 x 106 cells/mL, respectively. Then the cells were diluted to viable density of 3.6 x 106 cells/mL with fresh MetaCell®HEK293-100 expression medium. The plasmids mentioned above were transfected into prepared cells with Opti-MEM™ reduced serum medium (Gibco, #31985070) diluted PEI-MAX (Polysciences, #24765). Finally, the transfected cells were cultured in 37°C, 8% CO2 shaker for 5-6 days and harvested when cell viability was decreased to approximately 75%.
The cell culture media were harvested by centrifugation at 3,300 g for 10 min and filtered by 24-well filter plate (Cytiva, #97032). The filtered supernatants were loaded on to pre-equilibrated 24-well purification plate (Cytiva, #7700-9904) containing MabSelect PrismA (Cytiva, #17549802) resin and re-equilibrated by washing with 5-10 column volumes (CV) of PBS. The target recombinant antibodies were eluted by acidic buffer (0.1M Citrate buffer, pH3.0), then neutralized with basic buffer (2M Tris-HCl, pH9.0). The sample concentration was determined by NanoDrop (Thermo Scientific™ NanoDrop™ One) and stored at 4-degree for later use.
Bio-layer interferometry
To determine the binding kinetics of 96 VHH-monoFc HCAbs with SARS-CoV-2 Spike trimer protein, single-concentration binding assays were conducted using Bio-Layer Interferometry (BLI) on an Octet RED96e instrument (Sartorius). Biotinylated SARS-CoV-2 Spike trimer protein (Aero, Cat#SPN-C82Ey) was captured onto streptavidin (SA) biosensors (Sartorius, Cat#18-5019) as ligand with a loading level of 1.2- 1.4 nm. 96 VHH-monoFc HCAbs were used as analyte in a concentration of 30 nM. All samples were diluted with freshly prepared l x kinetic buffer (10x Kinetic Buffer (Sartorius, Cat#18-1105) diluted with PBS (BBI Life Sciences, Cat#E607016-0500)). Binding activities were performed at a shake speed of 1000 rpm at 25°C with an association time of 180 seconds and a dissociation time of 400 seconds. Obtained binding curves were reference signal
I l l subtracted and were fitted with a simple 1 : 1 binding model to generate kinetic data using Octet data analysis software (Sartorius, version 12.0). The fitted association rate (kon), dissociation rate (kOfr), equilibrium dissociation constant (KD) and binding graphs are shown.
Calculation of pl
An application internally developed by Nona Biosciences, with the algorithm and pKa parameters disclosed in EMBOSS (http://emboss.open-bio.org/rel/rel6/apps/iep.html). was used to predict pl.
Example 1: Fc fusions of the HR2 targeting VHH.R3 DC23 can protect mice against a lethal viral challenge with SARS-CoV-2
K18-hACE2 mice that express human ACE2 at the surface of their epithelial cells were treated with 100 pg R3_DC23-Fc (SEQ ID NO: 125) or 100 pg isotype control IgG (palivizumab) 1 day prior to a lethal infection with D614G SARS-CoV-2 virus. Infected mice that were not treated were included as control. Figure 1 illustrates that in sharp contrast to mice that were either treated with isotype control antibody or untreated mice, all mice that were treated with R3_DC23-Fc survived the challenge and did not display significant bodyweight loss, indicating that R3_DC23-Fc can protect mice from a lethal viral challenge with SARS-CoV-2.
Example 2: LS mutants of humanized R3DC23 Fc fusions control viral replication in hamsters
The therapeutic potential of huR3DC23-Fc (SEQ ID NO: 124) in the Syrian hamster model was evaluated. Hamsters were challenged with an ancestral SARS-CoV-2 isolate (BetaCoV/Munich/BavPatl/2020) and, 4 hours later, treated with either 10 mg/kg or 2 mg/kg huR3DC23-Fc_LS, 10 mg/kg bebtelovivamb (biosimilar) of 10 mg/kg palivizumab (negative control treatment) by intraperitoneal injection. At 4 days post infection high levels of huR3DC23-Fc_LS were detected in all hamsters treated with 2 mg/kg and in 4 hamsters treated with 10 mg/kg of this construct (data not shown). In sharp contrast, no or very low levels of huR3DC23-Fc_LS could be detected in the sera of two animals that were treated with 10 mg/kg huR3DC23-Fc_LS. This most likely results from unsuccessful injection, which has been observed by others (Starr et al. 2021 Nature 597:97-102). Apart from these 2 hamsters the lung virus loads, sampled on day 4 after challenge were below the detection limit in the huR3DC23-Fc treated hamsters whereas control treated animals had significantly higher lung virus loads (Fig. 2A). In accordance in the lungs of hamsters treated with either huR3DC23-Fc_LS or bebtelovimab a strong reduction in viral RNA was observed (Fig. 2B). This experiment shows that S2-binding huR3DC23-Fc can strongly restrict SARS-CoV-2
replication in vivo.
Example 3: Prophylactic treatment with R3_DC23-Fc protects K18-hACE2 mice against SARS-CoV-2 infection
K18-hACE2 mice that express human ACE2 at the surface of their epithelial cells were treated with 100 pg R3_DC23-Fc(YTE) (R3_DC23-Fc; SEQ ID NO: 125) or 100 pg isotype control IgG (palivizumab) via intraperitoneal injection 1 day prior to a lethal infection with SARS-CoV-2 D614G variant virus. Palivizumab treated, infected wild type (WT) mice that are nonpermissive for SARS- CoV-2 infection were used as control for protection.
As expected, all palivizumab treated K18-hACE2 displayed marked bodyweight loss and succumbed from the viral challenge. In sharp contrast, R3-DC23-Fc protected infected K18-hACE2 mice from bodyweight loss and death, similar to palivizumab treated WT mice. This illustrates that R3-DC23- Fc can protect mice from lethal SARS-CoV-2 infections (Fig. 3A and B).
To test if R3_DC23-Fc can also control viral replication in the lungs, K18-hACE2 mice were treated with 100 pg R3_DC23-Fc or 100 pg isotype control IgG (palivizumab) via intraperitoneal injection 1 day prior to infection with SARS-CoV-2 D614G variant virus. At five days post-infection part of the mice were sacrificed to isolate the lungs for quantification of viral replication by plaque assay and qPCR and part of the mice were used to monitor the bodyweight and mortality.
R3_DC23-Fc treatment protected K18-hACE2 mice form bodyweight loss and lethality upon viral challenge (Fig. 4A and B). Whereas high levels of replicating SARS-CoV-2 virus could be isolated from lungs of all K18-hACE2 mice that were treated with palivizumab, no replicating virus could be detected in the lungs of K18-hACE2 mice treated with R3-DC23-Fc or of WT mice (Fig. 4C). Similarly, in contrast to palivizumab-treated K18-hACE2 mice, no viral RNA could be detected in the lungs of K18-hACE2 mice treated with R3-DC23-Fc or of WT mice (Fig. 4D). These data illustrate that prophylactic treatment with R3-DC23-Fc can prevent or strongly reduce viral replication in the highly permissive K18-hACE2 mice.
Example 4: Variants of hR3_DC23 with reduced pl
The molecules RNGL-Fc_LS (SEQ ID NO: 96), RAGL-Fc_LS (SEQ ID NO: 97), E2NGL-Fc_LS (SEQ ID NO:98), E2AGL-Fc_LS (SEQ ID NO:99), QRNGL-Fc_LS (SEQ ID NO: 100), QRAGL- Fc_LS (SEQ ID NO: 101), ERNGL-Fc_LS (SEQ ID NO: 102), ERAGL-Fc_LS (SEQ ID NO: 103), E10RNGL-Fc_LS (SEQ ID NO: 104), E10RAGL-Fc_LS (SEQ ID NO: 105), E81RNGL-Fc_LS (SEQ ID NO: 106), E81RAGL-Fc_LS (SEQ ID NO: 107), Q75-Fc_LS (SEQ ID NO: 108), E75-
Fc_LS (SEQ ID NO: 109), E81-Fc_LS (SEQ ID NO: 110), E10E81-Fc_LS (SEQ ID NO: 111), DI 6- Fc_LS (SEQ ID NO: 112), T19D79-Fc_LS (SEQ ID NO: 113), TQD-Fc_LS (SEQ ID NO: 114), Q75R21Y-Fc_LS (SEQ ID NO: 115), CBVl-Fc LS (SEQ ID NO: 116), CBV2-Fc-LS (SEQ ID NO: 117), CBV3-Fc_LS (SEQ ID NO: 118) and CBV4-Fc_LS (SEQ ID NO: 119) consist of two copies of a modified version (SEQ ID NO: 3 -26) of the humanized version of the VHH building block
R3 DC23 (hR3_DC23: SEQ ID NO:2) connected to a human IgGl Fc containing an LS mutation (SEQ ID NO:83) via a (648)2 (SEQ ID NO:88) linker at the Fc N-terminus (Fig. 5B).
CBV3-Fc_LS or XVR013m:
DVQLVESGGELVQPGGSLRLSCAVSGRPFSTYTMGWFRQAPGKEREFVAAVRWGAGTIYYADSVKG RFTISRDNAENTVYLEMNSLRPEDTAVYYCGAAYVSKANYGSLWYQDSERYDYWGQGTLVTVSSGG
GGSGGGGSDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWY
VDGVEVHNAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPR
EPQVYTLPPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKL TVDKSRWQQGNVFSCSVLHEALHSHYTQKSLSLSPG ( SEQ ID NO : 118 ) The theoretical pl of the constructs is shown in Table 4. The variant S2 targeting molecules exhibited lower pls than XVR013. The theoretical pl was calculated based on the amino acid sequence.
Example 5: S2 targeting molecules neutralize SARS-CoV-2 pseudotyped viruses
Molecules R3DC23-Fc_WT (SEQ ID NO: 120), R3DC23-Fc_YTE (SEQ ID NO: 121), R3DC23- Fc DHS (SEQ ID NO: 122), R3DC23-Fc_LA (SEQ ID NO: 123) and R3DC23-Fc_LS (SEQ ID NO: 124) consist of two copies of the humanized version of the VHH building block R3 DC23 (SEQ ID NO:2) connected to a human IgGl Fc (respectively, comprising SEQ ID NO: 82, 84, 85, 86 and 83) via a (648)2 linker (SEQ ID NO: 88) at their N-terminus. Molecule R3DC23-Fc_WT has a wild type human IgGl Fc (comprising SEQ ID NO: 82), while molecules R3DC23-Fc_YTE (SEQ ID NO: 121), R3DC23-Fc_DHS (SEQ ID NO: 121), R3DC23-Fc_LA (SEQ ID NO: 121) and R3DC23- Fc_LS (SEQ ID NO: 124) have mutations in the Fc part of the human IgGl Fc for half-life extension (Fig. 5B).
The neutralizing potency of the S2 targeting molecules of Example 4 and described above was tested in a neutralization assay using VSV particles pseudotyped with the spike of SARS-CoV-2 (D614G), Omicron variants BA.5 and XBB.1.5 (-G252V). Dose-dependent neutralization of the molecules, the positive control (XVR013 = hR3_DC23-Fc_LS), and anegative control (palivizumab) were assessed in a pseudovirus neutralization assay. All molecules were able to neutralize all tested variants with mean IC50 values ranging from 0.87 to 5.41 ng/mL for the D614G variant, from 0.75 ng/mL to 4.01 ng/mL for Omicron BA.5, and from 0.41 to 2.52 ng/mL for XBB1.5 (-G252V) (Fig. 7 and Table 5).
Table 5: Neutralization of SARS-CoV-2 variants D614G, Omicron BA.5 and XBB.1.5 (-G252V) determined in a pseudovirus neutralization assay. The Table shows the calculated mean IC50 values for the tested molecules. N refers to the number of independent experiments. NA: not available.
Example 6: S2 targeting molecules neutralize authentic SARS-CoV-2 D614G and Omicron BA.5 variants
To test if the S2 targeting molecules of Examples 4 and 5 can also potently neutralize authentic SARS-CoV-2 virus, a microneutralization method was performed using SARS-CoV-2 D614G and SARS-CoV-2 Omicron BA.5 variant viruses. Dilution series of the molecules or controls were preincubated with a fixed amount of plaque-forming units (PFUs) for both virus variants for 1 hour at 37°C and subsequently used to infect Vero E6-TMPRSS2 cells. Dose-dependent neutralization of the molecules, the positive controls (bebtelovimab biosimilar and XVR013), and a negative control (palivizumab) were assessed in an authentic virus neutralization assay. Table 6 shows that all molecules were able to neutralize both variants with geometric mean IC50 values (N=2) ranging from 0.36 to 6.54 ng/mL for D614G variant and from 0.63 ng/mLto 9.03 ng/mL for Omicron BA.5 variant.
Table 6. Neutralization of authentic SARS-CoV-2 virus D614G and Omicron BA.5 determined by microneutralization method. Geometric mean values ± SD are reported. N refers to the number of independent experiments.
Example 7. Affinity assessment of S2 targeting molecules on FcRn
To determine the binding kinetics of the molecules for human hFcRn (at pH6.0 and pH7.4), the molecules of Examples 4 and 5 and controls (sotrovimab biosimilar and XVR013) were immobilized on CM5 chips at low density and recombinant human FcRn-p2m was injected in-solution. The affinity for human FcRn at pH6.0 was similar between the two fitting models. The calculated KDS at pH6.0 ranged between 42.9 and 99.5 nM for all molecules and the controls expect for R3DC23- Fc_WT, which had a KD of 395 nM (Table 7). The molecules with mutations in the Fc moiety for half-life extension showed improved affinity to human FcRn at pH6.0 compared to the molecule with a wildtype Fc (R3DC23-Fc_WT). As expected, low binding to human FcRn was observed at pH 7.4
for all molecules and the controls. As such, it was not possible to calculate steady state affinity at pH 7.4.
Table 7. SPR data for human FcRn at pH6.0. The Steady state affinity and 1: 1 binding kinetic parameters for hFcRn at pH6.0 are described in the Table.
Example 8. Non-specific binding of S2 targeting molecules to the surface of mammalian cells The non-specific binding of the molecules of Example 4 to the surface of mammalian (HEK293) cells was investigated with a flow cytometry experiment. Sotrovimab biosimilar was used as negative control antibody in the assay and XVR013 was used as positive control. As shown in Figure 8, no binding to HEK293 cells was observed for the molecules or the controls at a low concentration (10 pg/mL). At a high concentration (100 pg/mL). non-specific binding to HEK293 cells was observed for the molecules RNGL-Fc_LS, RAGL-Fc_LS, E10RAGL-Fc_LS, E81RNGL-Fc_LS, E81RAGL- Fc_LS and the positive control XVR013.
Example 9. Immunogenicity risk of S2 targeting molecules
An in silico immunogenicity assessment was performed for the molecules of Example 4 to predict the presence of putative HLA class II restricted epitopes. Lonza’s Epibase platform was applied to analyze the HLA binding specificities of all 10-mer peptides derived from the VHH-Fc constructs. The DRB1 scores of the tested molecules and XVR013 were similar to the DRB1 scores of humanized antibodies (Fig. 9). The DRB1 score is an approximate score representing a potential immunogenic risk for a peptide and is calculated from the population frequencies of the affected allotypes. The molecules CBVl-Fc_LS, CBV2-Fc_LS, CBV3-Fc_LS and CBV4-Fc_LS had the lowest DRB 1 score and therefore the lowest immunogenicity risk.
Example 10. Apparent hydrophobicity of S2 targeting molecules
The apparent hydrophobicity of the molecules of Example 4 was assessed by Analytical Hydrophobic Interaction Chromatography (HIC) assay and compared with XVR011 (control for hydrophobicity; XVR011 is a bivalent molecule consisting of two VHH72 building blocks each connected at their C- terminus to an Fc domain with a LALA mutation) and XVR013 (Fig. 10). CBV2-Fc_LS (SEQ ID NO: 117) and CBV4-Fc_LS (SEQ ID NO: 119) were analysed separately: both molecules have lower pl compared to the other tested molecules and the pH of the mobile phase used for the HIC assay needed to be adapted accordingly.
RNGL-Fc_LS, RAGL-Fc_LS, E2NGL-Fc_LS, E2AGL-Fc_LS, QRAGL-Fc_LS, ERAGL-Fc_LS, E10RAGL-Fc_LS, E81RNGL-Fc_LS, E81RAGL-Fc_LS and CBV4-Fc_LS displayed high hydrophobicity with retention time from 23 to 27 min, whereas Q75R21Y-Fc_LS was unusually hydrophobic showing a retention time of 35 min (Fig. 10). Q75-Fc_LS, E75-Fc_LS, E10E81-Fc_LS, D16-Fc_LS, T19D79-Fc_LS, TQD-Fc_LS, CBVl-Fc_LS, CBV2-Fc_LS and CBV3-Fc_LS were remarkedly more hydrophilic with retention times below the retention time of XVR013.
Example 11. SCX analysis of S2 targeting molecules
The hR3_DC23 VHH-Fc_LS variants of Example 4 were analyzed by SCX to detect their charge variants (Fig. 11).
The retention time of almost all variants was lower compared to XVR013 indicating lower positive charge patches on the protein surface compared to XVR013.
It should be noted that all variants eluted in 2 peaks indicating the presence of a charge variant comprising approximately 15% of the protein material. Without wishing to be bound by any theory this may be due to the transient production process used to generate the S2 targeting molecules, which was not optimized.
Example 12. Developability assessment of S2 targeting molecules
The stability of the S2 targeting molecules of Example 4 was assessed under accelerated stress conditions (ageing study) in order to assess their propensity to degrade in harsh condition, mimicking the worst-case condition a drug candidate may support during manufacturing, shipment and storage operation. The samples characteristics are summarized and classified by color-coded categories according to the legend (Fig. 12).
Example 13. Pharmacokinetic (PK) study in tg32 SCID mice
In order to study the pharmacokinetics of XVRO 14 and the S2 targeting molecules, in vivo PK studies were performed in human neonatal Fc receptor (hFcRn) tg32 SCID mice. This in vivo pre-clinical model was chosen given its proven utility for PK characterisation of engineered monoclonal antibodies with enhanced affinity for hFcRn (and avoiding inaccurate PK-estimation due to speciesspecific immunogenic responses). The pharmacokinetic properties of selected pre-lead candidates were compared with those of XVRO 13.
An in vivo PK study was performed in hFcRn tg32 SCID mice for E2NGL-Fc_LS, E75-Fc_LS, CBV2-Fc_LS, CBV3-Fc_LS, CBV4-Fc_LS and XVR013. Five male animals were included per group. One dose level (10 mg/kg) was tested using a single intravenous (IV) administration.
An interim read-out in the PK assay was performed at day 14. Serum samples were analysed and the interim PK parameters of the variants were compared to those of XVRO 13.
The PK results indicate that the exposure in plasma of the selected S2 targeting molecules was improved compared to XVR013; best plasma exposure was detected for CBV3-Fc_LS (Fig. 13).
CBV3-Fc_LS (also referred to herein as XVR013m) was selected for further follow up and the animals that had received 10 mg/kg of CBV3-Fc_LS were further followed up until day 95 of the study. Serum samples were analysed and the final PK parameters were determined via PK population approach (Table 8). In the Tg32 SCID mouse, a half-life of 9,3 days was modelled. Allometric scaling was then applied to project a terminal half-life of 44 to 47 days in human.
An in vivo PK study was also performed for XVR014 in hFcRn tg32 SCID mice. Six male animals were included per group. Two dose levels were tested, 10 mg/kg and a 2 mg/kg, using a single intravenous (IV) administration. Animals were followed up until day 95 of the study. Serum samples were analysed and the final PK parameters were determined via PK population approach (Table 8). In the Tg32 SCID mouse, a half-life of 16,7 days (2 mg/kg dose group) to 18,3 days (10 mg/kg dose group) was modelled. Allometric scaling was then applied to project a terminal half-life of 73 to 77 days in human.
Table 8. XVR013m and XVR014 PK parameters in the Tg32 SCID mouse as determined using a two-compartment population PK model with linear elimination. Elimination clearance (CL), central and peripheral volume of distribution (VI and V2, respectively), and distribution clearance (Q).
Example 14. Composition of SI and S2 targeting binding agents (XVR012) neutralizes VSV- GFP reporter viruses pseudotyped with SARS-CoV-2 spike proteins.
A VHHx-Fc-VHHy construct was generated wherein a humanized form of the SI targeting VHH3. 117 capable of binding to or competing for the VHH3. 117 epitope is fused to a human IgGl Fc containing an LS mutation via a (648)2 linker (SEQ ID NO: 88) at the N-terminus of the Fc domain, and wherein a humanized form of the SI targeting VHH3.83 capable of binding to or competing for the VHH72 epitope is fused to the C-terminus via a (648)2 linker (SEQ ID NO: 88) (XVR014) (SEQ ID NO: 126).
XVR014:
DVQLVESGGGLVQPGGSLRLSCAASGKAVSISDMGWYRQPPGKQRELVATITKTGSTNYADSVKGR FTISRDNTKNTVYLEMNSLRPEDTAVYYCNAWLPYGLGPDYYGLELWGQGTLVTVSSGGGGSGGGG
SDKTHTCPPCPAPELLGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSHEDPEVKFNWYVDGVEVH
NAKTKPREEQYNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKALPAPIEKTISKAKGQPREPQVYTL PPSRDELTKNQVSLTCLVKGFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRW QQGNVFSCSVLHEALHSHYTQKSLSLSPGGGGGSGGGGSDVQLVESGGGLVQPGDSLRLSCVLSGG VFTSYAMGWFRQAPGKEREFLAAITFNSDATYYADSVKGRFTISRDNAKNTAYLQMNSLRPEDTAV YSCAAGGNHYNPQYYHDYDKYDHWGQGTLVTVSS ( SEQ ID NO : 126 )
XVR014 was mixed with CBV3-Fc_LS (also referred to herein as XVR013m, SEQ ID NO: 118) of Example 4 in a ratio 1: 1 to generate a composition or cocktail (XVR012).
The neutralization potency of XVR012, XVR013m and XVR014 was tested in a pseudovirus neutralization assay using VSV particles pseudotyped with the spike of either SARS-CoV-2 D614G reference strain, or SARS-CoV-2 variants EG.5.1, BA.2.86.1 and HV. l. A negative control antibody (palivizumab) and the positive controls XVR013, sotrovimab and bebtelovimab were included in the assay. The results ofthe pseudovirus neutralization assay are shown in Fig. 14 and Table 9. XVR012, XVR013m and XVR014 were able to neutralize all tested SARS-CoV-2 variants (D614G, EG.5.1, BA.2.86. 1 and HV. 1) with mean IC50 values ranging from 6.0 to 11.8 ng/mL for XVR012, from 3.9 ng/mL to 6.7 ng/mL for XVRO 13m, and from 14.8 to 90,6 ng/mL for XVRO 14 (Table 9) . The positive control XVR013 was able to neutralize the tested SARS-CoV-2 variants with IC50 values ranging from 0.6 to 1.4 ng/mL. Bebtelovimab neutralized the reference strain D614G (IC50 of 2.3 ng/mL), but could not neutralize the variants EG5.1, BA.2.86.1 and HV. l at the tested concentrations. Sotrovimab neutralized the reference strain D614G (IC50 of 24.4 ng/mL), but was less potent against the variants EG5.1 and HV. l variants and could not neutralize the variant BA.2.86.1 at the tested concentrations.
Table 9: Neutralization of SARS-CoV-2 variants (D614G, EG.5.1, BA.2.86.1 and HV. l) by XVR012, XVR013m, XVR014 and XVR013 as determined in a pseudovirus neutralization assay. Vero E6 cells were transduced with VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 D614G spike protein or with the spike protein ofthe SARS-CoV-2 variants EG.5.1, BA.2.86.1 and HV. l, which viruses had been pre-incubated with different concentrations of the XVR012 composition, the constructs XVR013m (CBV3-Fc_LS: SEQ ID NO: 118) or XVR014 (SEQ ID NO: 126), XVR013 (SEQ ID NO: 124), sotrovimab or bebtelovimab. Sixteen hours later, the GFP fluorescence was measured with a fluorimeter. The calculated mean IC50 values (ng/ml; N = 1) are shown. The mean IC50 values were calculated by nonlinear regression curve fitting, log(inhibitor) versus normalized response (four parameters). indicates that no neutralization was detected.
The neutralization potency of XVR012, XVR013m and XVR014 was tested in a pseudovirus neutralization assay using VSV particles pseudotyped with the spike of either SARS-CoV-2 D614G reference strain, or SARS-CoV-2 variants Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3, BA.5, EG.5.1, FL.1.5.1, BA.2.86.1, HV. l, XBB.1.5, XBB.2.3, XBB.1.16, CH.1.1, HK.3, BA.2.74, JN. l, KP.2.3, KP.3, KP.3.1.1, KP.1.1, LB. l and XEC. A negative control antibody (palivizumab) or an isotype control antibody was included in the assay. The results of the pseudovirus neutralization assay are shown in Fig. 15 and Table 10. XVR012, XVR013m and XVR014 were able to neutralize all tested SARS-CoV-2 variants (D614G, Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3 and Omicrons BA.5, EG.5.1, FL.1.5.1, BA.2.86.1, HV. l, XBB.1.5, XBB.2.3, XBB.1.16, CH.1.1, HK.3, BA.2.74, JN. l, KP.2.3, KP.3, KP.3.1.1, KP.1.1, LB. l and XEC) with mean IC50 values ranging from 4.8 to 12.8 ng/mL for XVR012, from 2.3 ng/mL to 7.7 ng/mL for XVR013m, and from 11.6 to 461.3 ng/mL for XVR014 (Table 10).
Table 10: Neutralization of SARS-CoV-2 variants (D614G, Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3, BA.5, EG.5.1, FL.1.5.1, BA.2.86.1, HV. l, XBB.1.5, XBB.2.3, XBB.1.16, CH.1.1, HK.3, BA.2.74, JN. l, KP.2.3, KP.3, KP.3.1.1, KP.1.1, LB.l and XEC) by XVR012, XVR013m and XVR014 as determined in a pseudovirus neutralization assay. Vero E6 cells were transduced with VSV-GFP reporter viruses pseudotyped with SARS-CoV-2 D614G spike protein or with the spike protein of the SARS-CoV-2 variants Gamma, Eta, Iota, Mu, Zeta, B. 1.617.3 and Omicrons BA.5, EG.5.1, FL.1.5.1, BA.2.86.1, HV. l, XBB.1.5, XBB.2.3, XBB.1.16, CH.1.1, HK.3, BA.2.74, JN. l, KP.2.3, KP.3, KP.3.1.1, KP.1.1, LB. l and XEC which viruses had been pre-incubated with different concentrations of the XVR012 composition, the constructs XVR013m (CBV3-Fc_LS: SEQ ID NO: 118) or XVR014 (SEQ ID NO: 126). Sixteen hours later, the GFP fluorescence was measured with a fluorimeter. The calculated mean IC50 values (ng/ml; N = 3, unless indicated by “*” where N = 1, by ‘**’ where N = 2, by '***' where N = 12, by ‘****’ where N = 17, or by ‘*****’ where N = 14) are shown. The mean IC50 values were calculated by nonlinear regression curve fitting, log(inhibitor) versus normalized response (four parameters), nt: not tested.
Example 15. Evaluation of pre-existing ADAs
In an attempt to reduce pre-existing anti-drug antibodies (ADAs), variants were generated of XVR014 (SEQ ID NO: 126) described in Example 14, wherein the C-terminal sequence TVSS (SEQ ID NO: 139) was replaced by the sequence TVSGP (SEQ ID NO: 141) or the sequence TVSAA (SEQ ID NO: 140), thereby obtaining respectively, modified XVRO 14 (TVSGP) consisting of the sequence set forth in SEQ ID NO: 127 and modified XVR01 (TVSAA) consisting of the sequence set forth in SEQ ID NO: 128.
The seroprevalence of pre-existing ADAs in drug -naive individuals was assessed against XVRO 13, XVRO 14, the two XVRO 14 variants with C-terminal modifications, the negative control (caplacizumab) and the positive control (TAS266).
An ELISA based method was used to determine the percentage of healthy adult human donors with pre-existing ADAs against XVR013, XVR014, the XVR014 variants with C-terminal modifications, the negative control and the positive control. Pre-existing ADAs were identified in 15%, 29%, 20% and 19% of the healthy donors in XVR013, XVR014, modified XVR014 (TVSGP) and modified XVR014 (TVSAA), respectively (Table 11). The C-terminal modifications in XVR014 reduced the percentage of pre-existing ADAs compared to XVR014.
Table 11. Evaluation of pre-existing ADAs in drug -naive individuals against XVR013, XVR014, two modified XVR014 variants with C-terminal modifications (modified XVR014 (TVSGP) and modified XVR014 (TVSAA)), caplacizumab (negative control) and TAS266 (positive control).
Example 16: XVR012, XVR013m and XVR014 control viral replication in hamsters
The therapeutic potential of the cocktail XVR012 in a 1: 10 ratio ofXVR013m and XVR014, or the individual molecules XVR013m and XVR014 in the Syrian hamster challenge model was evaluated. Hamsters were challenged with an ancestral SARS-CoV-2 isolate (BetaCoV/Munich/BavPatl/2020) and, 4 hours later, treated with either 2 mg/kg, 1 mg/kg or 0,5 mg/kg XVR013m, 20 mg/kg, 10 mg/kg or 5 mg/kg XVR014, (2+20) mg/kg, (1+10) mg/kg or (0,5+5) mg/kg XVR012 (1: 10 ratio cocktail of XVR013m and XVR014), 10 mg/kg bebtelovivamb (biosimilar, positive control) or 10 mg/kg palivizumab (negative control) by intraperitoneal injection.
At 4 days post infection expected levels ofXVR012 were detected in all hamsters treated with (2+20) mg/kg, (1+10) mg/kg and (0,5+5) mg/kg of XVR012, in 6 out of 8 hamsters treated with 2 mg/kg of XVR013m and in all hamsters treated with 1 mg/kg and 0,5 mg/kg of XVR013m, in 5 out of 6 animals treated with 20 mg/kg of XVR014 and in all animals treated with 10 mg/kg and 5 mg/kg of XVR014 (data not shown). In sharp contrast, no or very low levels of XVR013m or XVR014 could be detected in the sera of respectively two animals that were treated with 2 mg/kg XVR013m and two animals that were treated with 20 mg/kg XVR014. This most likely results from unsuccessful injection, which has been observed by others (Starr et al. 2021 Nature 597:97-102).
Infectious viral lung loads were completely below detection levels on Day 4 in all animals
experimentally confirmed to have been treated with the high, mid and low doses of both the combination, cocktail XVR012, and XVR013m. Animals treated with XVR014 displayed a dosedependent effect on lung viral load: respectively, 1 out of 5 and 3 out of 6 animals in the 20 mg/kg and 10 mg/kg dose groups showed a complete reduction in viral load below detection levels, with the other animals in the group showing about 10.000-fold reduction in infectious viral load compared to animals treated with a negative control antibody. In the 5 mg/kg group, none of the animals showed a full reduction in viral load, but still a 1000 to 10.000-fold reduction in infectious viral load compared to the negative control antibody was observed in all but one animal (Fig. 16A).
A dose-dependent effect was observed on the viral load detected in the nasal turbinates at Day 4 for the combination XVR012 and the individual components XVR013m and XVR014. A clear synergistic effect was observed with the combination XVR012 demonstrating substantially lower levels of viral load at all dose levels compared to the individual components XVR013m and XVR014 (Fig. 16B).
Example 17: S2 targeting molecules neutralize authentic SARS-CoV-2 variants
To test the therapeutic potential of the cocktail XVR012 in a 1: 1 weight ratio of XVR013m and XVR014, and of the individual molecules XVR013m and XVR014, a microneutralization method was performed using SARS-CoV-2 D614G, Alpha, Beta, Gamma, Delta, Epsilon B. 1.427, Epsilon B. 1.429, Kappa, Zeta, Omicron BA. 1 and Omicron BA.2 variant viruses. Eight-point 2-fold dilution series of the molecules or controls were pre-incubated with a fixed amount of plaque-forming units (PFUs) for all virus variants for 1 hour at 37°C. The mixtures were then transferred to Vero E6- TMPRSS2 cells and incubated for 1 hour after which the inoculum was removed and replaced with fresh medium containing carboxymethylcellulose. Then the plates were incubated overnight after which the cells were fixed and immunostained to visualize virus infected cells. Virus control wells (cells + virus) and cell control wells (cells only) were included. A positive control (Sotrovimab biosimilar), and a negative control (isotype control) were also assessed. The ICso/ICso/ICgo titers were calculated according to the method described by Zielinski et al. (2005, Virology Journal 2:84). Table 12 shows that all molecules were able to neutralize both variants with MN50 values (N=l) ranging from 11.96 to 33.04 ng/ml for the cocktail XVR012, 11.69 to 40.30 ng/ml for XVR013m and 20.46 to 191.86 ng/ml for XVR014.
Table 12. Neutralization of authentic SARS-CoV-2 virus D614G, Alpha, Beta, Gamma, Delta, Epsilon B. 1.427, Epsilon B. 1.429, Kappa, Zeta, Omicron BA.l and Omicron BA.2, determined by microneutralization method. IC50, ICso and IC90 are shown. 1 independent experiment was performed.
Example 18: Affinity maturation of VHH hR3_DC23 and CBV3
In order to identify variants of VHH hR3_DC23 (SEQ ID N0:2) and of VHH CBV3 (SEQ ID NO:25) that improve and/or conserve most of the binding activity, site targeted random mutagenesis was done. In short, the amino acids in close proximity of the HR2 epitope were identified and divided into 2 groups, each group of amino acids binding to separate helices of the HR2 (Table 13).
Table 13: Degenerated libraries constructed. In each library, the aa location, position (Kabat numbering) and parental amino acid is indicated.
Phage display libraries of VHH hR3_DC23 (SEQ ID NO:2) were generated with codons NNK (where N = A/C/G/T, and K = G/T) at each of the indicated position to fully degenerate the amino acids. These degenerated libraries were used into 3 rounds of phage display selection using biotinylated SARS-CoV-2 spike protein (full spike of BQ1.1 variant, AA Vai 16 - Pro 1213, Accession # QHD43416.1; SEQ ID NO:245). Shortly, the biotinylated SARS-CoV-2 spike protein was captured on streptavidin beads in round 1 and round 2 and on ELISA plates in round 3. The stringency of selection was increased at each round by lowering the amount of biotinylated SARS- CoV-2 spike used (200, 10 and 1 nM respectively), decreasing the binding time (down to 20 min in R3) and increasing the number of washes. Selected phages were used to infect bacteria and single colonies were isolated from the round 2 (R2) and round 3 (R3). Periplasmic extract of each single
clone (6x 94 clones) was prepared and tested in a binding ELISA. Since almost all the clones showed binding, all the clones were sequenced and the amino acid diversity at each position was analyzed separately.
A selection of single mutants was produced and tested for their binding kinetics to SARS-CoV-2 spike. The mutations were selected based on their frequency after the phage display: each amino acid present at least twice at a particular position after phage display was introduced in the VHH CBV3 (SEQ ID NO:25) and produced by mammalian cells in a monovalent VHH-humanFc format (VHH- monoFc, Nona Biosciences, SEQ ID NO:246), purified using protein A and the off rate was measured using BLI (Octet) using a single concentration. The monomeric Fc format allows to use protein A for purification, but does not form bivalency to avoid the avidity effect. In addition, the most prominent amino acids (present at least 5 times) were also introduced in the VHH hR3_DC23 (SEQ ID NO:2), produced, purified and tested as above. Finally, some combinations of mutations were tested as well.
The on rate (kon), off rate (kOff) and affinity (KD) measured are in Table 14, as well as the ratio of the off rate of each mutant compared to its parent (hR3_DC23 (SEQ ID NO:2) = mutant #41, or CBV3 (SEQ ID NO:25) = mutant #42). Note that the same can be done for kon and KD. For simplicity and in vivo relevance, kofrwas selected.
Table 14: Binding kinetics of mutants selected from phage display to affinity mature hR3_DC23 and/or CBV3. The ratio of off rate of mutant versus its parent was calculated, to evaluate which mutation could be advantageous. Calculated pl and production yield are also displayed.
Analyzing the ratio of off rates for both hR3_DC23 and CBV3 mutants, the single mutants were classified as indicated in Table 15.
Table 15: Classification of the single mutants, from advantageous to deleterious, based on the ratio of off rates compared to their parent, combining results for both hR3_DC23 and CBV3.
Two mutations (S30K and GlOOcH) were clearly advantageous, decreasing the off-rate by 2-fold. 3 mutations were slightly advantageous (R27H, S30T and S30I). Multiple mutations were acceptable and did not impact the off rate, at positions 27, 29, 30, 96 and lOOd, while only one mutant was acceptable at positions 99, 100, 100c and 1001. Multiple mutations were still acceptable, impacting minimally the very good off rate of hR3_DC23 and CBV3 (positions 29, 96, 97, 98 and 1001), while
only one mutant was acceptable at positions 31 and 100b. Finally, no mutants or deleterious mutants could be selected for positions 52, 97, 100a, 100b and 1001.
The calculated pl of each VHH-monoFc was also analyzed in Table 14. All hR3_DC23 mutants had a pl in the same range as their parent. Similarly, all CBV3 mutants had similar pl as their parent, indicating that the mutations did not trigger a change in pl and are thus not expected to trigger a change in PK properties. The production yield (mg of antibody produced / L of cellular culture) was also reviewed. All mutants produced well, except one (#9, hR3_DC23_R1001L).
Example 19: Binding of S2 targeting molecules to the spike of Khosta-2, a clade 3 virus from the Sarbecovirus subgenus
The breadth of binding of the S2 targeting molecules beyond SARS-CoV-2, among members of the Sarbecovirus subgenus, was evaluated. For this, the ability to bind to the spike of the distantly related bat virus Khosta-2, a member of the clade 3 sarbecoviruses able to use human ACE2 as a receptor to infect human cells (Seifert et al. 2022 PLoS Pathog 18(9):el010828), was evaluated by flow cytometry. Briefly, cells were transfected with vectors expressing GFP fused to the spike of Khosta- 2 or SARS-CoV-2 variant D614G, or expressing GFP alone. The transfected cells were incubated with the S2 targeting molecule XVR013m, XVR014, a positive control (XVR013 or sotrovimab) or a negative control (palivizumab or isotype control). Bound molecules were detected by staining for human IgG Fc and flow cytometry analysis. Two independent experiments were performed. As expected, XVR013, XVR013m, XVR014 and sotrovimab highly bound to cells expressing the spike of SARS-CoV-2 D614G variant (Figure 19). A dose-response was observed for binding to cells expressing the spike of Khosta-2 virus, showing at least partial binding (Figure 19). At 10 pg/ml, XVR013, XVR013m, XVR014 and sotrovimab were able to bind to ~80, 20, 30 and 20% ofthe cells expressing the Khosta-2 spike, respectively.
Claims
1. A binding agent capable of neutralizing a Sarbecovirus , characterized in that said binding agent comprises at least one immunoglobulin single variable domain (ISVD), wherein the ISVD comprises a complementarity determining region 1 (CDR1) defined by SEQ ID NO:40, a CDR2 defined by SEQ ID NO:30 and a CDR3 defined by SEQ ID NO:41; or a CDR1 defined by SEQ ID NO:38, a CDR2 defined by SEQ ID NO:39 and a CDR3 defined by SEQ ID NO:41 .
2. A binding agent capable of neutralizing a Sarbecovirus, characterized in that said binding agent comprises at least one immunoglobulin single variable domain (ISVD), wherein the ISVD comprises a CDR1, CDR2 and CDR3, each independently as present in any of SEQ ID NOs: 2 to 26, wherein the CDR1, CDR2 and CDR3 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
3. The binding agent according to claim 1 or 2, wherein the ISVD comprises a CDR1 defined by any one of SEQ ID NO:27-29, a CDR2 defined by SEQ ID NO:30; and a CDR3 defined by any one of SEQ ID NO:31-37; or a CDR1 defined by SEQ ID NO:38, a CDR2 defined by SEQ ID NO:39; and a CDR3 defined by any one of SEQ ID NO:31-37.
4. The binding agent according to claim 2, wherein the ISVD comprises a combination of CDR1, CDR2 and CDR3, wherein the CDR1, CDR2 and CDR3 are as present in a particular one of the sequences defined by SEQ ID NOs: 2 to 26, wherein the CDR1, CDR2 and CDR3 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
5. The binding agent according to any one of claims 1 to 4, wherein the ISVD further comprises a combination of framework region 1 (FR1), FR2, FR3 and FR4 each independently as present in any of SEQ ID NOs: 2 to 26, wherein the FR1, FR2, FR3 and FR4 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia; or a FR1 defined by any one of SEQ ID NO:42- 45, a FR2 defined by SEQ ID NO:46, a FR3 defined by any one of SEQ ID NO:47-59 and a FR4 defined by SEQ ID NO:60; or a FR1 defined by any one of SEQ ID NO:61-68, a FR2 defined by SEQ ID NO:46, a FR3 defined by any one of SEQ ID NO:69-81 and a FR4 defined by SEQ ID NO:60.
6. The binding agent according to any one of claims 1 to 5, wherein the ISVD further comprises a combination of FR1, FR2, FR3 and FR4, wherein the FR1, FR2, FR3 and FR4 are as present in a particular one of the sequences defined by SEQ ID NOs: 2 to 26, wherein the FR1, FR2, FR3 and FR4 are annotated according to any one of Kabat, MacCallum, IMGT, AbM, Martin or Chothia.
7. The binding agent according to any one of claims 1 to 6. wherein the ISVD comprises an amino acid sequence with at least 90% identity to any one of the sequences defined by SEQ ID NO:2-26.
8. The binding agent according to any one of claims 1 to 7, wherein the ISVD does not comprise an amino acid sequence defined by SEQ ID NO: 1 or 2.
9. The binding agent according to any one of claims 1 to 7, wherein the ISVD comprises an amino acid sequence defined by any one of SEQ ID NO:3-26.
10. A binding agent capable of neutralizing a Sarbecovirus , characterized in that said binding agent comprises at least one immunoglobulin single variable domain (ISVD), wherein the ISVD is an affinity matured variant of an ISVD as defined in any one of claims 1 to 9, optionally wherein said ISVD comprises an amino acid sequence defined by any one of SEQ ID NO: 183-188, 190-195, 197, 198, 200-244, or 151-182.
11. The binding agent according to any one of claims 1 to 10, which is in a multivalent form.
12. The binding agent according to any one of claims 1 to 11, wherein the ISVD is fused to an Fc domain.
13. The binding agent according to any one of claims 1 to 12, wherein the ISVD is fused to an Fc domain comprising a sequence defined by any one of SEQ ID NO: 82-86.
14. A nucleic acid molecule comprising a polynucleotide sequence encoding the binding agent according to any one of claims 1 to 13, optionally wherein said nucleic acid molecule is mRNA.
15. A vector comprising the nucleic acid molecule according to claim 14.
16. A cell expressing the binding agent according to any one of claims 1 to 13 or comprising the nucleic acid molecule according to claim 14 or the vector according to claim 15.
17. A composition comprising the binding agent as defined in any one of claims 1 to 13.
18. The composition according to claim 17, further comprising a binding agent which specifically binds to a Sarbecovirus spike protein receptor binding domain (RBD).
19. The composition according to claim 18, wherein the binding agent specifically binding to the RBD comprises at least one ISVD capable of binding to or competing for the epitope specifically bound by VHH72 defined by SEQ ID NO:90 and at least one ISVD capable of binding to or competing for the epitope specifically bound by VHH3.117 defined by SEQ ID NO:92, such as a binding agent defined by any one of SEQ ID NO: 126-138.
20. The composition according to claim 18 or 19 comprising the binding agent defined by SEQ ID NO: 118, and the binding agent defined by SEQ ID NO: 126 or a variant thereof such as a binding agent defined by any one of SEQ ID NO: 127-138.
21. A pharmaceutical composition comprising the binding agent according to any one of claims 1 to 13, the nucleic acid molecule according to claim 14, the vector according to claim 15, or the composition according to any one of claims 17 to 20, and a pharmaceutically acceptable carrier.
22. A kit such as a diagnostic kit comprising the binding agent according to any one of claims 1 to 13, or the composition according to any one of claims 17 to 20.
23. The binding agent according to any one of claims 1 to 13, the nucleic acid molecule according to claim 14, the vector according to claim 15, the composition according to any one of claims 17 to 20, the pharmaceutical composition according to claim 21, or the kit according to claim 22 for use in medicine.
24. The binding agent according to any one of claims 1 to 13, the nucleic acid molecule according to claim 14, the vector according to claim 15, the composition according to any one of claims 17 to 20, the pharmaceutical composition according to claim 21, or the kit according to claim 22 for use in the prevention or treatment of a Sarbecovirus infection in a subject.
25. The binding agent according to any one of claims 1 to 13, the nucleic acid molecule according to claim 14, the vector according to claim 15, the composition according to any one of claims 17 to 20, the pharmaceutical composition according to claim 21, or the kit according to claim 22 for use in the diagnosis of a Sarbecovirus infection in a subject.
26. An in vitro or ex vivo method for detecting a Sarbecovirus in a sample comprising:
- contacting the sample with a binding agent according to any one of claims 1 to 13, and
- determining binding of the binding agent with a Sarbecovirus or a part thereof.
Applications Claiming Priority (8)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP23211603.8 | 2023-11-22 | ||
| EP23211603 | 2023-11-22 | ||
| EP24151698.8 | 2024-01-12 | ||
| EP24151698 | 2024-01-12 | ||
| EP24172808 | 2024-04-26 | ||
| EP24172808.8 | 2024-04-26 | ||
| EP24206518 | 2024-10-14 | ||
| EP24206518.3 | 2024-10-14 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025109176A1 true WO2025109176A1 (en) | 2025-05-30 |
Family
ID=93650489
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/EP2024/083304 Pending WO2025109176A1 (en) | 2023-11-22 | 2024-11-22 | Optimized sarbecovirus spike s2 subunit binders and compositions comprising the same |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025109176A1 (en) |
Citations (38)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1994004678A1 (en) | 1992-08-21 | 1994-03-03 | Casterman Cecile | Immunoglobulins devoid of light chains |
| WO1994025591A1 (en) | 1993-04-29 | 1994-11-10 | Unilever N.V. | PRODUCTION OF ANTIBODIES OR (FUNCTIONALIZED) FRAGMENTS THEREOF DERIVED FROM HEAVY CHAIN IMMUNOGLOBULINS OF $i(CAMELIDAE) |
| WO1995004079A1 (en) | 1993-08-02 | 1995-02-09 | Raymond Hamers | Recombinant vector containing a lipoprotein gene sequence for expressing nucleotide sequences |
| WO1996034103A1 (en) | 1995-04-25 | 1996-10-31 | Vrije Universiteit Brussel | Variable fragments of immunoglobulins - use for therapeutic or veterinary purposes |
| WO1997049805A2 (en) | 1996-06-27 | 1997-12-31 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Recognition molecules interacting specifically with the active site or cleft of a target molecule |
| WO1999037681A2 (en) | 1998-01-26 | 1999-07-29 | Unilever Plc | Method for producing antibody fragments |
| WO2000040968A1 (en) | 1999-01-05 | 2000-07-13 | Unilever Plc | Binding of antibody fragments to solid supports |
| WO2000043507A1 (en) | 1999-01-19 | 2000-07-27 | Unilever Plc | Method for producing antibody fragments |
| WO2000065057A1 (en) | 1999-04-22 | 2000-11-02 | Unilever Plc | Inhibition of viral infection using monovalent antigen-binding proteins |
| WO2001021817A1 (en) | 1999-09-24 | 2001-03-29 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Recombinant phages capable of entering host cells via specific interaction with an artificial receptor |
| WO2001040310A2 (en) | 1999-11-29 | 2001-06-07 | Unilever Plc | Immobilisation of proteins using a polypeptide segment |
| WO2001044301A1 (en) | 1999-11-29 | 2001-06-21 | Unilever Plc | Immobilized single domain antigen-binding molecules |
| EP1134231A1 (en) | 2000-03-14 | 2001-09-19 | Unilever N.V. | Antibody heavy chain variable domains against human dietary enzymes, and their uses |
| WO2001090190A2 (en) | 2000-05-26 | 2001-11-29 | National Research Council Of Canada | Single-domain antigen-binding antibody fragments derived from llama antibodies |
| WO2002048193A2 (en) | 2000-12-13 | 2002-06-20 | Unilever N.V. | Camelidae antibody arrays |
| WO2003025020A1 (en) | 2001-09-13 | 2003-03-27 | Institute For Antibodies Co., Ltd. | Method of constructing camel antibody library |
| WO2003035694A2 (en) | 2001-10-24 | 2003-05-01 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Functional heavy chain antibodies, fragments thereof, library thereof and methods of production thereof |
| WO2003050531A2 (en) | 2001-12-11 | 2003-06-19 | Algonomics N.V. | Method for displaying loops from immunoglobulin domains in different contexts |
| WO2003054016A2 (en) | 2001-12-21 | 2003-07-03 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Method for cloning of variable domain sequences |
| WO2003055527A2 (en) | 2002-01-03 | 2003-07-10 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Immunoconjugates useful for treatment of tumours |
| US20040023390A1 (en) | 2002-08-05 | 2004-02-05 | Davidson Beverly L. | SiRNA-mediated gene silencing with viral vectors |
| WO2004041865A2 (en) | 2002-11-08 | 2004-05-21 | Ablynx N.V. | Stabilized single domain antibodies |
| WO2004041867A2 (en) | 2002-11-08 | 2004-05-21 | Ablynx N.V. | Camelidae antibodies against imminoglobulin e and use thereof for the treatment of allergic disorders |
| WO2004062551A2 (en) | 2003-01-10 | 2004-07-29 | Ablynx N.V. | RECOMBINANT VHH SINGLE DOMAIN ANTIBODY FROM CAMELIDAE AGAINST VON WILLEBRAND FACTOR (vWF) OR AGAINST COLLAGEN |
| WO2005044858A1 (en) | 2003-11-07 | 2005-05-19 | Ablynx N.V. | Camelidae single domain antibodies vhh directed against epidermal growth factor receptor and uses therefor |
| WO2006040153A2 (en) | 2004-10-13 | 2006-04-20 | Ablynx N.V. | Single domain camelide anti -amyloid beta antibodies and polypeptides comprising the same for the treatment and diagnosis of degenarative neural diseases such as alzheimer's disease |
| WO2006079372A1 (en) | 2005-01-31 | 2006-08-03 | Ablynx N.V. | Method for generating variable domain sequences of heavy chain antibodies |
| WO2006122786A2 (en) | 2005-05-18 | 2006-11-23 | Ablynx Nv | Improved nanobodies™ against tumor necrosis factor-alpha |
| WO2006122825A2 (en) | 2005-05-20 | 2006-11-23 | Ablynx Nv | Single domain vhh antibodies against von willebrand factor |
| WO2008020079A1 (en) | 2006-08-18 | 2008-02-21 | Ablynx N.V. | Amino acid sequences directed against il-6r and polypeptides comprising the same for the treatment of deseases and disorders associated with il-6-mediated signalling |
| WO2012175741A2 (en) | 2011-06-23 | 2012-12-27 | Ablynx Nv | Techniques for predicting, detecting and reducing aspecific protein interference in assays involving immunoglobulin single variable domains |
| WO2015173325A2 (en) | 2014-05-16 | 2015-11-19 | Ablynx Nv | Improved immunoglobulin variable domains |
| WO2021156490A2 (en) | 2020-02-06 | 2021-08-12 | Vib Vzw | Corona virus binders |
| WO2022167666A1 (en) | 2021-02-05 | 2022-08-11 | Vib Vzw | Sarbecovirus binders |
| WO2022238550A1 (en) | 2021-05-12 | 2022-11-17 | Vib Vzw | Pan-specific corona virus binders |
| WO2023035016A1 (en) * | 2021-09-03 | 2023-03-09 | The Uab Research Foundation | Human neutralizing antibodies against sars-cov-2 spike s2 domain and uses thereof |
| WO2023076881A1 (en) * | 2021-10-26 | 2023-05-04 | The United States Of America, As Represented By The Secretary, Department Of Health And Human Services | Single domain antibodies targeting the s2 subunit of sars-cov-2 spike protein |
| WO2023222825A1 (en) * | 2022-05-18 | 2023-11-23 | Vib Vzw | Sarbecovirus spike s2 subunit binders |
-
2024
- 2024-11-22 WO PCT/EP2024/083304 patent/WO2025109176A1/en active Pending
Patent Citations (42)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1994004678A1 (en) | 1992-08-21 | 1994-03-03 | Casterman Cecile | Immunoglobulins devoid of light chains |
| WO1994025591A1 (en) | 1993-04-29 | 1994-11-10 | Unilever N.V. | PRODUCTION OF ANTIBODIES OR (FUNCTIONALIZED) FRAGMENTS THEREOF DERIVED FROM HEAVY CHAIN IMMUNOGLOBULINS OF $i(CAMELIDAE) |
| WO1995004079A1 (en) | 1993-08-02 | 1995-02-09 | Raymond Hamers | Recombinant vector containing a lipoprotein gene sequence for expressing nucleotide sequences |
| WO1996034103A1 (en) | 1995-04-25 | 1996-10-31 | Vrije Universiteit Brussel | Variable fragments of immunoglobulins - use for therapeutic or veterinary purposes |
| WO1997049805A2 (en) | 1996-06-27 | 1997-12-31 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Recognition molecules interacting specifically with the active site or cleft of a target molecule |
| WO1999037681A2 (en) | 1998-01-26 | 1999-07-29 | Unilever Plc | Method for producing antibody fragments |
| WO2000040968A1 (en) | 1999-01-05 | 2000-07-13 | Unilever Plc | Binding of antibody fragments to solid supports |
| WO2000043507A1 (en) | 1999-01-19 | 2000-07-27 | Unilever Plc | Method for producing antibody fragments |
| WO2000065057A1 (en) | 1999-04-22 | 2000-11-02 | Unilever Plc | Inhibition of viral infection using monovalent antigen-binding proteins |
| WO2001021817A1 (en) | 1999-09-24 | 2001-03-29 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Recombinant phages capable of entering host cells via specific interaction with an artificial receptor |
| WO2001040310A2 (en) | 1999-11-29 | 2001-06-07 | Unilever Plc | Immobilisation of proteins using a polypeptide segment |
| WO2001044301A1 (en) | 1999-11-29 | 2001-06-21 | Unilever Plc | Immobilized single domain antigen-binding molecules |
| EP1134231A1 (en) | 2000-03-14 | 2001-09-19 | Unilever N.V. | Antibody heavy chain variable domains against human dietary enzymes, and their uses |
| WO2001090190A2 (en) | 2000-05-26 | 2001-11-29 | National Research Council Of Canada | Single-domain antigen-binding antibody fragments derived from llama antibodies |
| WO2002048193A2 (en) | 2000-12-13 | 2002-06-20 | Unilever N.V. | Camelidae antibody arrays |
| WO2003025020A1 (en) | 2001-09-13 | 2003-03-27 | Institute For Antibodies Co., Ltd. | Method of constructing camel antibody library |
| EP1433793A1 (en) | 2001-09-13 | 2004-06-30 | Institute for Antibodies Co., Ltd. | Method of constructing camel antibody library |
| WO2003035694A2 (en) | 2001-10-24 | 2003-05-01 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Functional heavy chain antibodies, fragments thereof, library thereof and methods of production thereof |
| WO2003050531A2 (en) | 2001-12-11 | 2003-06-19 | Algonomics N.V. | Method for displaying loops from immunoglobulin domains in different contexts |
| WO2003054016A2 (en) | 2001-12-21 | 2003-07-03 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Method for cloning of variable domain sequences |
| WO2003055527A2 (en) | 2002-01-03 | 2003-07-10 | Vlaams Interuniversitair Instituut Voor Biotechnologie Vzw | Immunoconjugates useful for treatment of tumours |
| US20040023390A1 (en) | 2002-08-05 | 2004-02-05 | Davidson Beverly L. | SiRNA-mediated gene silencing with viral vectors |
| WO2004041863A2 (en) | 2002-11-08 | 2004-05-21 | Ablynx N.V. | Single domain antibodies directed against interferon- gamma and uses therefor |
| WO2004041862A2 (en) | 2002-11-08 | 2004-05-21 | Ablynx N.V. | Single domain antibodies directed against tumour necrosis factor-alpha and uses therefor |
| WO2004041867A2 (en) | 2002-11-08 | 2004-05-21 | Ablynx N.V. | Camelidae antibodies against imminoglobulin e and use thereof for the treatment of allergic disorders |
| WO2004041865A2 (en) | 2002-11-08 | 2004-05-21 | Ablynx N.V. | Stabilized single domain antibodies |
| WO2004062551A2 (en) | 2003-01-10 | 2004-07-29 | Ablynx N.V. | RECOMBINANT VHH SINGLE DOMAIN ANTIBODY FROM CAMELIDAE AGAINST VON WILLEBRAND FACTOR (vWF) OR AGAINST COLLAGEN |
| WO2005044858A1 (en) | 2003-11-07 | 2005-05-19 | Ablynx N.V. | Camelidae single domain antibodies vhh directed against epidermal growth factor receptor and uses therefor |
| WO2006040153A2 (en) | 2004-10-13 | 2006-04-20 | Ablynx N.V. | Single domain camelide anti -amyloid beta antibodies and polypeptides comprising the same for the treatment and diagnosis of degenarative neural diseases such as alzheimer's disease |
| WO2006079372A1 (en) | 2005-01-31 | 2006-08-03 | Ablynx N.V. | Method for generating variable domain sequences of heavy chain antibodies |
| WO2006122787A1 (en) | 2005-05-18 | 2006-11-23 | Ablynx Nv | Serum albumin binding proteins |
| WO2006122786A2 (en) | 2005-05-18 | 2006-11-23 | Ablynx Nv | Improved nanobodies™ against tumor necrosis factor-alpha |
| WO2006122825A2 (en) | 2005-05-20 | 2006-11-23 | Ablynx Nv | Single domain vhh antibodies against von willebrand factor |
| WO2008020079A1 (en) | 2006-08-18 | 2008-02-21 | Ablynx N.V. | Amino acid sequences directed against il-6r and polypeptides comprising the same for the treatment of deseases and disorders associated with il-6-mediated signalling |
| WO2012175741A2 (en) | 2011-06-23 | 2012-12-27 | Ablynx Nv | Techniques for predicting, detecting and reducing aspecific protein interference in assays involving immunoglobulin single variable domains |
| WO2015173325A2 (en) | 2014-05-16 | 2015-11-19 | Ablynx Nv | Improved immunoglobulin variable domains |
| WO2021156490A2 (en) | 2020-02-06 | 2021-08-12 | Vib Vzw | Corona virus binders |
| WO2022167666A1 (en) | 2021-02-05 | 2022-08-11 | Vib Vzw | Sarbecovirus binders |
| WO2022238550A1 (en) | 2021-05-12 | 2022-11-17 | Vib Vzw | Pan-specific corona virus binders |
| WO2023035016A1 (en) * | 2021-09-03 | 2023-03-09 | The Uab Research Foundation | Human neutralizing antibodies against sars-cov-2 spike s2 domain and uses thereof |
| WO2023076881A1 (en) * | 2021-10-26 | 2023-05-04 | The United States Of America, As Represented By The Secretary, Department Of Health And Human Services | Single domain antibodies targeting the s2 subunit of sars-cov-2 spike protein |
| WO2023222825A1 (en) * | 2022-05-18 | 2023-11-23 | Vib Vzw | Sarbecovirus spike s2 subunit binders |
Non-Patent Citations (41)
| Title |
|---|
| "GenBank", Database accession no. NM_001371415.1 |
| "Genbank", Database accession no. QHQ82464 |
| "Nomenclature and Symbolism for Amino Acids and Peptides", EUR. J. BIOCHEM., vol. 138, 1984, pages 9 - 37 |
| "Remington's Pharmaceutical Sciences", 1980, MACK PUBLISHING CO. |
| "UniProtKB", Database accession no. Q9BYF1 |
| ABDELRAHMAN ET AL., FRONT IMMUNOL, vol. 11, 2020, pages 552909 |
| ABHINANDANMARTIN, MOLECULAR IMMUNOLOGY, vol. 45, 2008, pages 3832 - 3839 |
| CHEN ET AL., LANCET, vol. 395, 2020, pages 507 - 513 |
| CHOTHIALESK, MOL BIOL., vol. 196, 1987, pages 901 - 17 |
| DAYHOFF ET AL.: "Atlas of Protein Sequence and Structure", vol. 5, 1978, pages: 345 - 352 |
| DE CAE ET AL.: "Ultrapotent SARS coronavirus-neutralizing single-domain antibodies that bind a conserved membrane proximal epitope of the spike", BIORXIV, 2023 |
| DE CAE SIEGLINDE ET AL: "Ultrapotent SARS coronavirus-neutralizing single-domain antibodies that bind a conserved membrane proximal epitope of the spike", BIORXIV, 1 March 2023 (2023-03-01), XP093070242, Retrieved from the Internet <URL:www.biorxiv.org/content/10.1101/2023.03.10.531533v1.full.pdf> [retrieved on 20230803], DOI: 10.1101/2023.03.10.531533 * |
| DE GROOT ET AL., N ENGL J VIROL, vol. 87, 2013, pages 7790 - 7792 |
| DODERO-ROJAS ET AL., ELIFE, vol. 10, 2021, pages e70362 |
| DROSTEN ET AL., N ENGL J MED, vol. 348, 2003, pages 1967 - 1976 |
| FAN ET AL., SIGNAL TRANSDUCT TARGET THER., vol. 7, 2022, pages 141 |
| GRALINSKI ET AL., VIRUSES, vol. 12, 2020, pages 135 |
| HAMERS-CASTERMAN ET AL., NATURE, vol. 363, 1993, pages 446 - 448 |
| HONEGGERPLÜCKTHUN, J MOL BIOL, vol. 309, 2001, pages 657 - 70 |
| HUANG ET AL., ACTA PHARMALOGICA SINICA, vol. 41, 2020, pages 1141 - 1149 |
| JARVIS, VIROLOGY, vol. 310, no. 1, 25 May 2003 (2003-05-25), pages 109 - 128 |
| KABAT ET AL.: "NIH publication 91-3242", 1991 |
| KÄRBER, ARCHIV F. EXPERIMENT. PATHOL. U. PHARMAKOL., vol. 162, 1931, pages 480 - 483 |
| LEFRANC, FRONTIERS IN IMMUNOLOGY, vol. 5, no. 22, 2014, pages 1 - 22 |
| LETKO, NAT MICROBIOL, vol. 5, 2020, pages 562 - 569 |
| MACCALLUM ET AL., J. MOL. BIOL., vol. 262, 1996, pages 732 - 745 |
| MATSUYAMA ET AL., PNAS, 2020 |
| MENDELL ET AL., N ENG J MED, vol. 377, 2017, pages 1713 - 1722 |
| MOLINER-MORRO ET AL., BIOMOLECULES, vol. 10, 2020, pages 1661 |
| MUYLDERMANS, REV MOL BIOTECHNOL, vol. 74, 2001, pages 277 - 302 |
| PICCOLI ET AL., CELL, vol. 184, 2020, pages 1004 - 1015 |
| POPP ET AL., NAT CHEM BIOL, vol. 3, 2007, pages 707 - 708 |
| RIECHMANNMUYLDERMANS, J IMMUNOL METHODS, vol. 231, 1999, pages 25 - 38 |
| SAMBROOKFRITSCHMANIATIS: "Molecular Cloning, A Laboratory Manual", 1989, COLD SPRING HARBOR LABORATORY PRESS |
| SEIFERT ET AL., PLOS PATHOG, vol. 18, no. 9, 2022, pages e1010828 |
| STARR ET AL., NATURE, vol. 597, 2021, pages 97 - 102 |
| WANG ET AL., FRONT CELL INFECT MICROBIOL, vol. 10, 2020, pages 587269 |
| WHEATLEY ET AL., CELL REP, vol. 37, 2021, pages 109822 |
| WRAPP ET AL., SCIENCE, vol. 367, 2020, pages 1260 - 1263 |
| ZHU ET AL., N ENGL J MED, vol. 382, 2020, pages 727 - 733 |
| ZIELINSKI ET AL., VIROLOGY JOURNAL, vol. 2, 2005, pages 84 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| WO2021183947A1 (en) | MOLECULES THAT BIND TO SARS-CoV-2 | |
| US20250313611A1 (en) | Sarbecovirus spike s2 subunit binders | |
| AU2014255864A1 (en) | Human antibodies binding to RSV G protein | |
| US20240228596A1 (en) | Pan-specific corona virus binders | |
| EP2986638A1 (en) | Human antibodies binding to rsv g proteins | |
| US11440952B2 (en) | Compositions for preventing or treating viral and other microbial infections | |
| JP2023516280A (en) | coronavirus binder | |
| US20240101647A1 (en) | Sarbecovirus binders | |
| US20240398911A1 (en) | Engineered receptors and monoclonal antibodies for cronaviruses and uses thereof | |
| WO2025109176A1 (en) | Optimized sarbecovirus spike s2 subunit binders and compositions comprising the same | |
| US20220340658A1 (en) | NEUTRALIZING ANTI-SARS-CoV-2 ANTIBODIES AND USE THEREOF | |
| IL302460A (en) | Anti-sars-cov-2 antigen antibodies and related compositions and methods | |
| US20250296987A1 (en) | Rsv/hmpv cross-reactive antibodies | |
| WO2024068744A1 (en) | Antivirals against human parainfluenza virus | |
| CA3267635A1 (en) | Antivirals against human parainfluenza virus |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24812491 Country of ref document: EP Kind code of ref document: A1 |