WO2025038492A1 - Anti-ctla-4 antibodies and related binding molecules and methods and uses thereof - Google Patents
Anti-ctla-4 antibodies and related binding molecules and methods and uses thereof Download PDFInfo
- Publication number
- WO2025038492A1 WO2025038492A1 PCT/US2024/041822 US2024041822W WO2025038492A1 WO 2025038492 A1 WO2025038492 A1 WO 2025038492A1 US 2024041822 W US2024041822 W US 2024041822W WO 2025038492 A1 WO2025038492 A1 WO 2025038492A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- cdr
- seq
- region
- set forth
- sequence set
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/2803—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily
- C07K16/2818—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily against CD28 or CD152
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/505—Medicinal preparations containing antigens or antibodies comprising antibodies
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/76—Antagonist effect on antigen, e.g. neutralization or inhibition of binding
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/92—Affinity (KD), association rate (Ka), dissociation rate (Kd) or EC50 value
Definitions
- CTLA-4 protein cytotoxic T-lymphocyte-associated protein 4
- CTLA-4 is a cell surface protein predominantly expressed on T cells and plays a crucial role in regulating immune responses.
- the novel antibodies or antibody fragments described may provide potent moieties for modulation of human immune function with potential therapeutic applications in various diseases, including cancer.
- Cancer therapies comprise a wide range of therapeutic approaches, including surgery, radiation, and chemotherapy. While the various approaches offer a broad selection of options for treating cancer, many therapeutics suffer from disadvantages, such as a lack of selectivity of targeting cancer cells over normal, healthy cells, and the development of resistance by the cancer to the treatment.
- Immunotherapy targeting checkpoint molecules has emerged as one of the most compelling approaches for treating patients with cancers, due to promising results from preclinical and clinical trials. Despite this promise, certain immunotherapies, including CTLA-4 targeting antibodies, have shown limited efficacy in a substantial number of patients and have been associated with unwanted toxicities, including immune-related adverse events. Thus there exists a need for providing improved immunotherapies. The disclosure provided herein addresses this need.
- an anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) antibody or antigen-binding fragment thereof comprising a heavy chain variable (VH) region and a light chain variable (VL) region
- the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1), a heavy chain complementarity determining region 2 (CDR-H2), and a heavy chain complementarity determining region 3 (CDR-H3) contained within any one of SEQ ID NOs: 1-27 and 112
- the VL region comprises a light chain complementarity determining region 1 (CDR-L1), a light chain complementarity determining region 2 (CDR-L2) and a light chain complementarity determining region 3 (CDR-L3) contained within any one of SEQ ID NOs: 28-48 and 118, wherein the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR- H3 contained within SEQ ID NO: 1
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118
- the VH region comprises a CDR-H1, a CDR- H2, and a CDR-H3 contained within SEQ ID NO: 2
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118
- the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 3
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained
- an anti-CTLA-4 antibody or antigenbinding fragment thereof comprising a heavy chain variable (VH) region and a light chain variable (VL) region
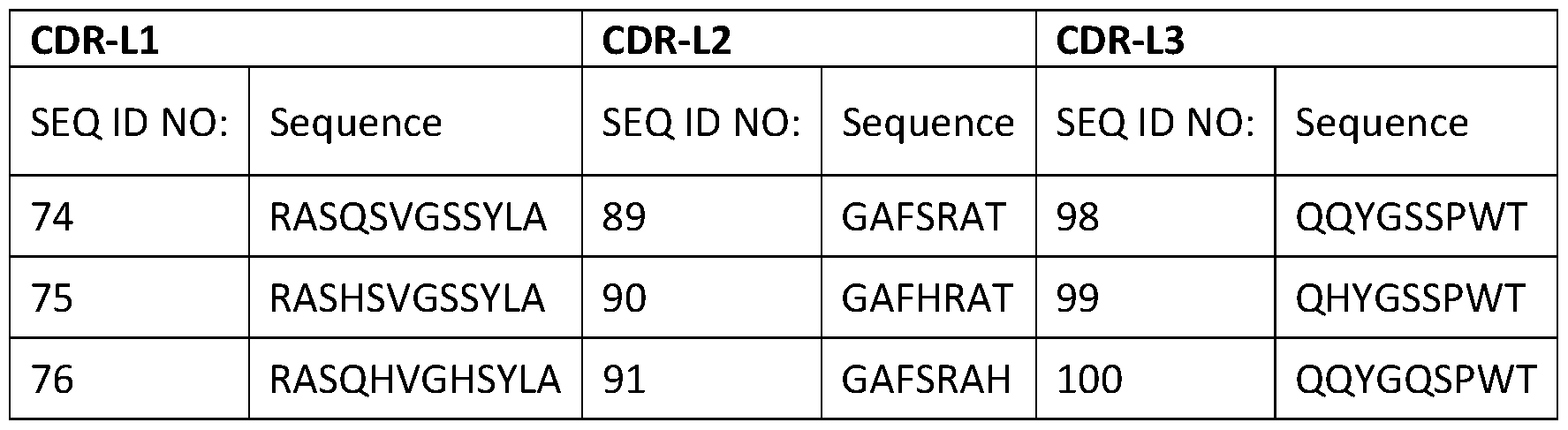
- the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1) comprising the sequence set forth in any one of SEQ ID NOs: 49-56, a heavy chain complementarity determining region 2 (CDR-H2) comprising the sequence set forth in any one of SEQ ID NOs: 57-67, and a heavy chain complementarity determining region 3 (CDR-H3) comprising the sequence set forth in any one of SEQ ID NOs: 68-73; and the VL region comprises a light chain complementarity determining region 1 (CDR- Ll) comprising the sequence set forth in any one of SEQ ID NOs: 74-88, a light chain complementarity determining region 2 (CDR-L2) comprising the sequence set forth in any one of SEQ ID NOs: 89-
- the VH region can comprise a heavy chain complementarity determining region 1 (CDR-H1) comprising the sequence set forth in any one of SEQ ID NOs: 50-56, a heavy chain complementarity determining region 2 (CDR-H2) comprising the sequence set forth in any one of SEQ ID NOs: 58-67, and a heavy chain complementarity determining region 3 (CDR-H3) comprising the sequence set forth in any one of SEQ ID NOs: 69-73.
- CDR-H1 heavy chain complementarity determining region 1
- CDR-H2 heavy chain complementarity determining region 2
- CDR-H3 heavy chain complementarity determining region 3
- the VL region can comprise a light chain complementarity determining region 1 (CDR-L1) comprising the sequence set forth in any one of SEQ ID NOs: 75-88, a light chain complementarity determining region 2 (CDR-L2) comprising the sequence set forth in any one of SEQ ID NOs: 90-97, and a light chain complementarity determining region 3 (CDR-L3) comprising the sequence set forth in any one of SEQ ID NOs: 99-100.
- CDR-L1 comprising the sequence set forth in any one of SEQ ID NOs: 75-88
- CDR-L2 comprising the sequence set forth in any one of SEQ ID NOs: 90-97
- CDR-L3 comprising the sequence set forth in any one of SEQ ID NOs: 99-100.
- the VH region can comprise a CDR-H1, a CDR- H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 58, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively;
- the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 69, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively;
- the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID
- the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 1
- the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118
- the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 2
- the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 9
- the VH region and the VL region can be or can comprise: the sequence set forth in SEQ ID NO: 1 and 118, respectively; the sequence set forth in SEQ ID NO: 2 and 118, respectively; the sequence set forth in SEQ ID NO: 3 and 118, respectively; the sequence set forth in SEQ ID NO: 4 and 118, respectively; the sequence set forth in SEQ ID NO: 5 and 118, respectively; the sequence set forth in SEQ ID NO: 6 and 118, respectively; the sequence set forth in SEQ ID NO: 7 and 118, respectively; the sequence set forth in SEQ ID NO: 8 and 118, respectively; the sequence set forth in SEQ ID NO: 9 and 118, respectively; the sequence set forth in SEQ ID NO: 10 and 118, respectively; the sequence set forth in SEQ ID NO: 11 and 118, respectively; the sequence set forth in SEQ ID NO: 12 and 118, respectively; the sequence set forth in SEQ ID NO: 13 and 118, respectively; the sequence set forth in SEQ ID NO: 1 and 118, respectively; the
- an anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) antibody or antigen-binding fragment thereof comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein the VH region and the VL region can be or can comprise: the sequence set forth in SEQ ID NO: 1 and 118, respectively; the sequence set forth in SEQ ID NO: 2 and 118, respectively; the sequence set forth in SEQ ID NO: 3 and 118, respectively; the sequence set forth in SEQ ID NO: 4 and 118, respectively; the sequence set forth in SEQ ID NO: 5 and 118, respectively; the sequence set forth in SEQ ID NO: 6 and 118, respectively; the sequence set forth in SEQ ID NO: 7 and 118, respectively; the sequence set forth in SEQ ID NO: 8 and 118, respectively; the sequence set forth in SEQ ID NO: 9 and 118, respectively; the sequence set forth in SEQ ID NO: 10 and 118, respectively; the sequence set forth in SEQ ID NO: 1 and 118,
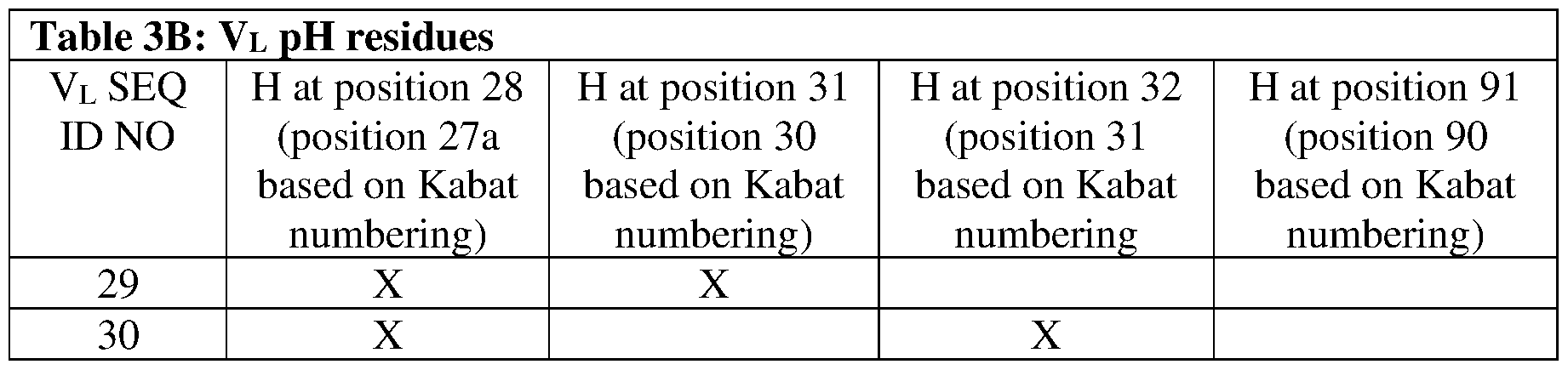
- the VH region comprises one or more amino acid residues selected from the group consisting of: a histidine (H) at position 98, an arginine (R) at position 31, a glutamic acid (E) or a histidine (H) at position 53, a histidine (H) at position 58, and a histidine (H) at position 97, or any combination thereof, and/or the VL region comprises one or more amino acid residues selected from the group consisting of: a histidine (H) at position 27a, a histidine (H) at position 30, a histidine (H) at position 31, and a histidine (H) at position 90, or any combination thereof, where numbering of amino acid residues is by Kabat numbering.
- the amino acid residues are mutations (amino acid substitutions) compared to a reference sequence.
- the VH region can comprise one or more mutations selected from the group consisting of: S31R, D53H, D53E, Y58H, and W97H
- the VL region comprises one or more mutations selected from the group consisting of: S28H, S31H, S32H, and Q91H, wherein numbering of amino acid residues is by Kabat numbering.
- the reference sequence is a VH sequence set forth in SEQ ID NO: 112 and a VL sequence set forth in SEQ ID NO: 118.
- the VH region can comprise L102H (corresponding to histidine at position 98 by Kabat numbering).
- the VH region can comprise a CDR-H1, a CDR- H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively
- the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
- the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively
- the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
- the VH region can comprise a CDR- Hl, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively
- the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
- the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively
- the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively
- the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively
- the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- the antibody can be a full length antibody.
- the full length antibody comprises a constant region (Fc) from an IgAl, IgA2, IgD, IgE, IgGl, IgG2, IgG3, IgG4, or IgM.
- the constant region is a human IgGl heavy chain constant region.
- the human IgGl heavy chain constant region comprises one or more of mutations selected from the group consisting of G236A, S239D, M252Y, S254T, T256E, A330L, I332E, M428L, and N434S, numbered 1 according to the EU numbering system.
- the human IgGl heavy chain constant region can comprise S239D/A330L/I332E mutations, numbered according to the EU numbering system.
- the antibody region can comprise a human IgGl heavy chain constant region comprising S239D/I332E mutations, numbered according to the EU numbering system.
- the antibody region can comprise a human IgGl heavy chain constant region comprising G236A/S239D/A330L/I332E mutations, numbered according to the EU numbering system.
- the antibody region can comprise a human IgGl heavy chain constant region comprising G236A/A330L/I332E mutations, numbered according to the EU numbering system. In any of the embodiments herein, the antibody region can comprise a human IgGl heavy chain constant region comprising M252Y/S254T/T256E mutations, numbered according to the EU numbering system. In any of the embodiments herein, the antibody region can comprise a human IgGl heavy chain constant region comprising M428L/N434S mutations, numbered according to the EU numbering system.
- the antibody can be an antigen-binding fragment.
- the antigen-binding fragment is selected from the group consisting of a single domain antibody, a single chain antibody, an unibody, a single chain variable fragment (scFv), a Fab fragment, and a F(ab')2 fragment.
- the anti-CTLA-4 antibody or antigen-binding fragment thereof can be recombinant.
- the VH region and the VL region can be human or can be from a human protein.
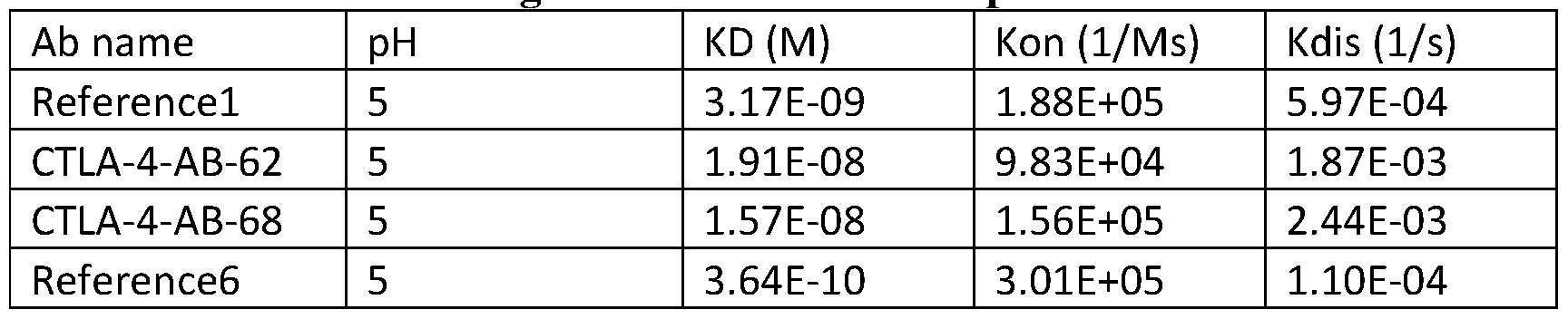
- the ratio of dissociation constant for binding at acidic pH (e.g., pH 5.0 to 6.0) to the dissociation constant for binding at physiological pH (e.g., pH 7.35 to 7.45) to human CTLA-4 protein can be greater than 2.
- the antibody or antigen-binding fragment can exhibit a ratio of a dissociation constant for binding at acidic pH (e.g., pH 5.0 to 6.0) to dissociation constant for binding at physiological pH (e.g., pH 7.35 to 7.45) of more than 7.
- the ratio of EC50 value of the antibody or antigen-binding fragment thereof for human CTLA-4 at physiological pH (e.g., pH 7.35 to 7.45) to EC50 value of the antibody or antigen-binding fragment thereof for human CTLA-4 at acidic pH (e.g., pH 5.0 to 6.0) can be greater than 2.
- the antibody or antigen-binding fragment thereof can possess pH dependent binding properties to human CTLA-4 that facilitates the dissociation of antibody from the antibody :CTLA-4 complex upon internalization into a cell, allowing either the antibody, CTLA-4, or both to be recycled instead of being degraded in the lysosome.
- the antibody or antigen-binding fragment thereof can preserve high affinity for CTLA-4 at pH>7, lower affinity at pH 5.5-7, and significantly reduced affinity for CTLA-4 at pH ⁇ 5.5.
- the antibody or antigen-binding fragment thereof can induce strong deletion of Treg and local T cell activation in tumor microenvironment but minimal systemic T cell activation.
- the antibody or antigen-binding fragment thereof can induce a favorable ratio of Treg cells to T effector cells in the tumor microenvironment.
- the antibody or antigen-binding fragment thereof can induce a favorable ratio of immune cell subsets in the tumor or blood.
- the antibody or antigen-binding fragment can induce a favorable ratio of CTLA-4 expression on immune cell subsets in the tumor or blood.
- the immune cell subsets comprise myeloid cells, T cells, CD4 + T cells, CD8 + T cells, and/or Treg cells.
- a conjugate comprising the anti-CTLA- 4 antibody or antigen-binding fragment thereof of any of the embodiments herein and a heterologous molecule or moiety.
- the heterologous molecule or moiety is a cytotoxic agent, a drug, enzymatically active toxin or fragment thereof, or a radioactive atom.
- a bispecific antibody comprising the anti- CTLA-4 antibody or antigen-binding fragment thereof of any of the embodiments herein.
- a polynucleotide comprising a nucleic acid encoding the anti-CTLA-4 antibody or antigen-binding domain thereof of any of the embodiments herein.
- a vector comprising the polynucleotide of any of the embodiments herein.
- the vector is a viral vector.
- the viral vector is a retroviral vector or a lentiviral vector.
- a cell comprising the polynucleotide of any of the embodiments herein, or the vector of any of the embodiments herein.
- the cell is a mammalian cell.
- provided herein is a method of producing an antibody comprising culturing the cell of any of the embodiments herein under a condition that produces the antibody. In some embodiments, the method further comprises recovering the antibody produced by the cell.
- an antibody or antigen-binding fragment thereof produced by the method of any of the embodiments herein.
- composition comprising the anti-CTLA- 4 antibody or antigen-binding fragment thereof of any of the embodiments herein, the conjugate of any of the embodiments herein, the bispecific antibody of any of the embodiments herein.
- the composition further comprises a pharmaceutically acceptable excipient.
- a method of stimulating an immune response in a subject comprising administering to a subject in need thereof the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of the embodiments herein, the conjugate of any of the embodiments herein, the bispecific antibody of any of the embodiments herein or the composition of any of the embodiments herein.
- stimulating an immune response treats a disease or condition in the subject.
- the disease or condition is a cancer.
- a method of cancer immunotherapy comprising administering to a subject in need thereof a therapeutically effective amount of the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of the embodiments herein, the conjugate of any of the embodiments herein, the bispecific antibody of any of the embodiments herein or the composition of any of the embodiments herein.
- the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of the embodiments herein, the conjugate of any of the embodiments herein, the bispecific antibody of any of the embodiments herein or the composition of any of the embodiments herein is administered to the subject in need thereof at about 0.01 pg/kg body weight to about 20 mg/kg body weight.
- the method further comprises administering to the subject an effective amount of at least one additional therapeutic agent.
- the at least one additional therapeutic agent is selected from the group consisting of: viral gene therapy, immune checkpoint inhibitors, target therapies, radiation therapies, vaccination therapies, and chemotherapies.
- the at least one additional therapeutic agent is selected from the group consisting of: pomalyst, revlimid, lenalidomide, pomalidomide, thalidomide, a DNA-alkylating platinum-containing derivative, cisplatin, 5- fluorouracil, cyclophosphamide, an anti-CD137 antibody, an anti-PD-1 antibody, an anti-PD- L1 antibody, an anti-CD20 antibody, an anti-CD40 antibody, an anti-DR5 antibody, an antiCD Id antibody, an anti-TIM3 antibody, an anti-SLAMF7 antibody, an anti-KIR receptor antibody, an anti-OX40 antibody, an anti-HER2 antibody, an anti-ErbB-2 antibody, an anti- EGFR antibody, cetuximab, rituximab, trastuzumab, pembrolizumab, an antibody drug conjugation, radiotherapy, single dose radiation, fractionated radiation, focal radiation, whole organ radiation, IL- 12, IFNa, GM
- the method comprises administering to the subject an effective amount of the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of the embodiments herein, the conjugate of any of the embodiments herein, the bispecific antibody of any of the embodiments herein or the composition of any of the embodiments herein prior to a surgery or after a surgery to remove a solid tumor in the subject.
- FIGs. 1A-S show results of binding tests of exemplary provided antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. 1A shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-7, CTLA-4- AB-9, CTLA-4- AB- 10, and CTLA-4- AB- 11) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. IB shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB- 13, CTLA-4- AB- 14, CTLA-4-AB-15, and CTLA-4-AB-16) and reference antibodies for binding to human CTLA- 4 at pH 5 and at pH 7.4.
- FIG. 1C shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB- 17, CTLA-4-AB-18, CTLA-4-AB-19, and CTLA-4-AB-20) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. ID shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB -21, CTLA-4- AB-22, CTLA-4-AB-23, and CTLA-4- AB-24) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. IE shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-25, CTLA-4-AB-26, CTLA-4- AB-27, and CTLA-4- AB-28) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. IF shows results of binding tests of anti-CTLA- 4 antibodies (CTLA-4- AB-29, CTLA-4- AB-30, CTLA-4-AB-31, and CTLA-4- AB-32) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. 1G shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-34, CTLA-4- AB-35, CTLA- 4-AB-36, CTLA-4-AB-37, CTLA-4-AB-38 and CTLA-4-AB-39) for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. 1H shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB -42, CTLA-4-AB-43, CTLA-4-AB-44, and CTLA-4-AB-45) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. II shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-8, CTLA-4- AB- 12, CTLA-4- AB-33, and CTLA-4- AB-46) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. 1J shows results of binding tests of anti-CTLA-4 antibody CTLA-4- AB- 47 and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. IK shows results of binding tests of anti-CTLA-4 antibody CTLA-4-AB-48 and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. IL shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-49, CTLA-4- AB-50, and CTLA-4- AB- 51) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. IM shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-52, CTLA-4- AB-53, and CTLA-4- AB-54) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. IN shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4-AB-55, CTLA-4- AB-56, and CTLA-4- AB-57) and reference antibodies for binding to human CTLA- 4 at pH 5 and at pH 7.4.
- FIG. IO shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-58, CTLA-4- AB-59, and CTLA-4- AB-60) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. IP shows results of binding tests of anti-CTLA- 4 antibodies and reference antibodies (CTLA-4- AB -61, CTLA-4- AB-62, and CTLA-4- AB-63) for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. IQ shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB -64, CTLA-4-AB-65, and CTLA-4-AB-66) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. 1R shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4-AB-67, CTLA-4-AB-68, and CTLA-4- AB -69) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4.
- FIG. IS shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB -70, CTLA-4- AB -71, and CTLA-4- AB -72) and reference antibodies for binding to human CTLA- 4 at pH 5 and at pH 7.4.
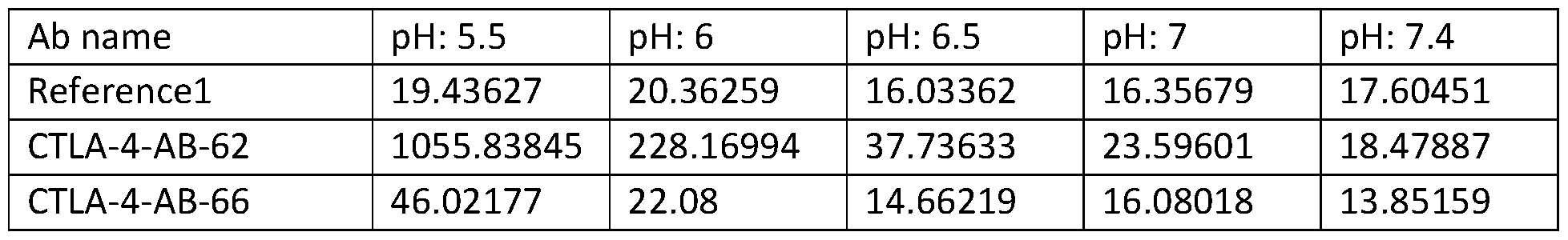
- FIGs. 2A-2C show results of binding tests of exemplary provided antibodies for binding to human CTLA-4 at different pH’s.
- FIG. 2A shows results of binding tests of a reference antibody for binding to human CTLA-4 at pH 5.5, pH 6, pH 6.5, pH 7 and pH 7.4.
- FIG. 2B shows results of binding tests of CTLA-4-AB-62 for binding to human CTLA-4 at pH 5.5, pH 6, pH 6.5, pH 7 and pH 7.4.
- FIG. 2C shows results of binding tests of CTLA-4- AB-68 for binding to human CTLA-4 at pH 5.5, pH 6, pH 6.5, pH 7 and pH 7.4.
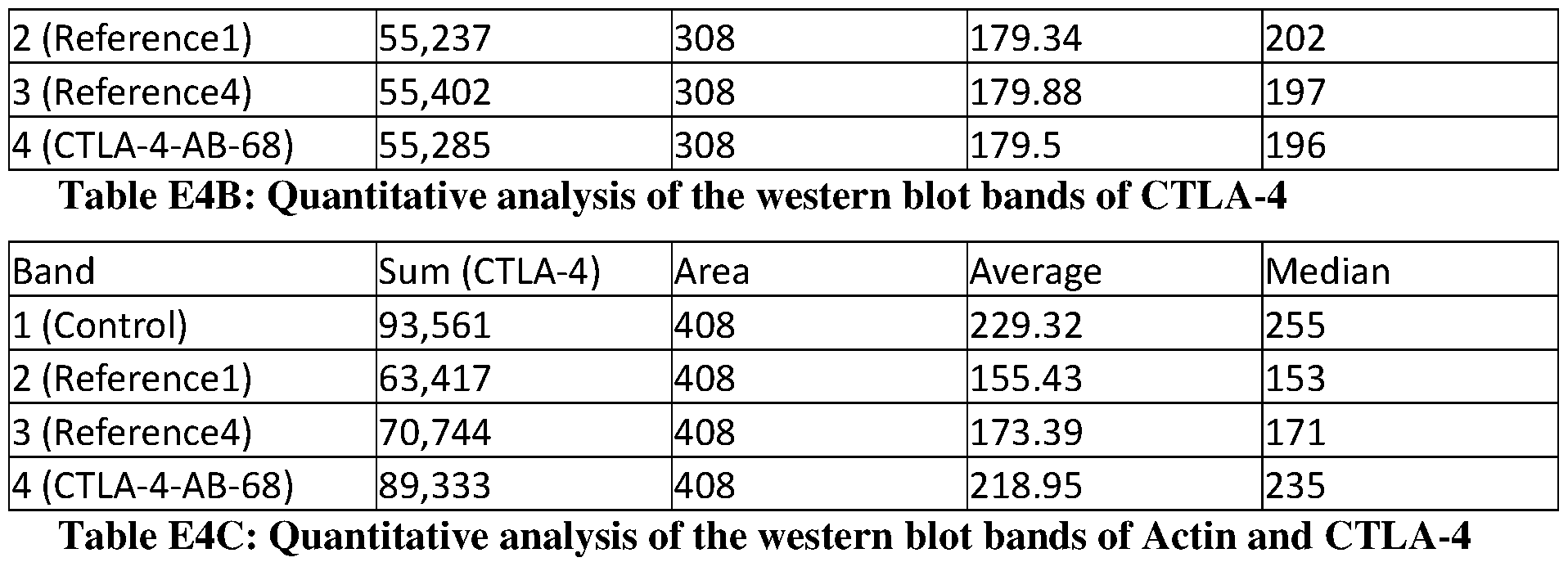
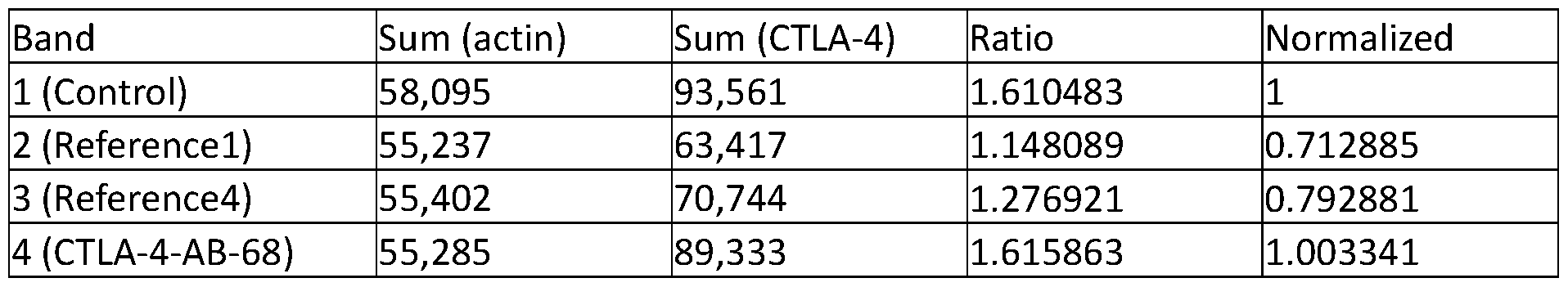
- FIGs. 3A-3C show effects of anti-CTLA-4 antibodies on CTLA-4 expression in treated human-CTLA-4 expressing Cells.
- FIG. 3A shows hCTLA-4 expression levels in cells treated with anti-CTLA-4 antibodies.
- FIG. 3B shows a schematic drawing of the workflow of confocal microscopy.
- FIG. 3C shows results of confocal microscopy.
- HEK293-hCTLA-4 cells were incubated with lOpg/mL antibody
- FIGs. 4A-4B show effect of anti-CTLA-4 antibodies on antibody dependent cellular cytotoxicity (ADCC).
- FIG. 4A shows selection of NK cells.
- FIG. 4B shows target cell lysis was evaluated at various concentrations of four different antibodies and EC50 values calculated from dose-response curves.
- FIGs. 5A-5E show in vivo anti-tumor efficacy of anti-CTLA-4 antibodies.
- FIG. 5A shows a schematic drawing of the experimental procedure.
- FIGs. 6A-6E show effect of anti-CTLA-4 antibodies on immune cell subsets in the tumor and blood of hCTLA4 knock-in mice.
- FIG. 6A shows gating strategy for tumor samples to analyze This panel was designed to analyze absolute cell numbers of CD45+, Myeloid , T cells, CD4 T, CD8 T and Treg in tumor of CT26 model in hCTLA4 knock-in mice.
- FIG. 6A shows gating strategy for tumor samples to analyze This panel was designed to analyze absolute cell numbers of CD45+, Myeloid , T cells, CD4 T, CD8 T and Treg in tumor of CT26 model in hCTLA4 knock-in mice.
- FIG. 6B shows gating strategy for blood samples to analyze absolute cell numbers of CD45+, Myeloid, T cells, CD4 T, CD8 T and Treg in blood of CT26 model in hCTLA4 knock-in mice, and the expression level of hCTLA-4 on Treg cells in the blood of CT26 model in hCTLA4 knock-in mice.
- FIG. 6C shows results of the FCM analysis for tumor samples. One-way ANOVA was used for statistical analysis. When a significant F-statistics (a ratio of treatment variance to the error variance) was obtained, comparisons between group 1 and other groups were carried out with Games-Howell (equal variances not assumed) or Tukey (equal variances assumed) test. * p ⁇ 0.05; ** p ⁇ 0.01; *** p ⁇ 0.001.
- FIG. 6D shows results of the FCM analysis for blood samples showing percentages of myeloid, T cells and T cell subpopulations in CD45+ cells.
- FIG. 6E shows results of the FCM analysis for blood samples showing percentages of T cell subpopulations in T cells.
- FIG. 6F shows results of the FCM analysis for blood samples showing cell numbers of CD45+, myeloid, T cells and T cell subpopulations per 100 pL blood.
- FIG. 6G shows results of the FCM analysis for blood samples showing ratios between T cell subpopulations.
- CD4Teff refers to CD4T cells other than Treg.
- FIG. 6H shows MFI of hCTLA4 of Treg cells.
- the present disclosure concerns anti-CTLA-4 antibodies or antibody fragments (i.e. antigen binding fragment that is able to bind CTLA-4) suitable for therapeutic use. Relative to existing known therapies, some of these anti-CTLA-4 antibodies or antibody fragments confer specific binding properties to human CTLA-4 under different pH conditions. In some embodiments, the disclosed anti-CTLA-4 antibodies or antibody fragments also exhibit physiochemical or biological properties that lead to improved efficacy and safety to anti- CTLA-4 antibodies known in the art. In some embodiments, the provided anti-CTLA-4 antibody, including any antigen binding antibody fragments, can be used to treat cancer.
- Cancer is a disease state of genomic instability caused by the accumulation of point mutations, resulting in behavioral and structural alterations to disaffected cells. Cancer presents in the form of abnormal growths defined by the proliferation of morphologically ambiguous cells that have the potential to invade both proximal and distant tissues, usually in the form of solid state tumors (Hassanpour SH, Dehghani M. Review of cancer from perspective of molecular. Journal of Cancer Research and. Practice. July 2017. doi:10.1016/j.jcrpr.2017.07.001).
- the immune system is responsible for defending the body against foreign substances, both internal and external , such as pathogens and abnormal cells (Daeron M. The immune system as a system of relations. Front Immunol. 2022; 13:984678). Composed of innate and adaptive immunity, the immune system has impressive flexibility when identifying disease-causing agents. Innate immune cells provide preconfigured responses to broad situations and stimuli (Kaur BP, Secord E. Innate Immunity. Immunol Allergy Clin North Am.
- adaptive immune cells provide more tailored responses to each stimulus through learning how to distinguish between previously encountered and new stimuli (Geckin B, Konstantin Fdhse F, Dominguez-Andres J, Netea MG. Trained immunity: implications for vaccination. Curr Opin Immunol. 2022;77: 102190). Because of its capacity to be “trained” to eliminate entities deemed harmful, the adaptive immune system has potential to be utilized for targeted cancer therapy, and the emergence of therapies such as CAR-T serves as compelling evidence of this strategy (Baulu E, Gardet C, Chuvin N, Depil S. TCR- engineered T cell therapy in solid tumors: State of the art and perspectives. Sci Adv. 2023;9(7):eadf3700).
- T cells Central to the immune response are T cells, a type of white blood cell, which act as key orchestrators of immune reactions (Kumar BV, Connors TJ, Farber DL. Human T Cell Development, Localization, and Function throughout Life. Immunity. 2018;48(2):202-213).
- T cells Starting as a hematopoietic stem cell, T cells mature into a variety of differentiated forms that can directly kill threats, recruit other immune cells to assist in the containment and eradication of threats, or mitigate excessive and indiscriminate immune activity that targets otherwise healthy and harmless cells (Shah K, Al-Haidari A, Sun J, Kazi JU. T cell receptor (TCR) signaling in health and disease. Signal Transduct Target Ther. 2021;6(l):412).
- T cells A unique feature of T cells is their ability to discriminate between healthy and abnormal surface markers, with abnormal marker detection eliciting activation of T cells (Feinerman O, Germain RN, Altan-Bonnet G. Quantitative challenges in understanding ligand discrimination by alphabeta T cells. Mol Immunol. 2008;45(3):619-631).
- T cell activation is tightly regulated to prevent excessive immune responses that could lead to tissue damage and autoimmune disorders.
- CTLA-4 a protein predominantly expressed on the surface of activated T cells, serves as an important checkpoint molecule for downregulating immune responses.
- CTLA-4 acts by binding to its ligands, CD80 (B7-1) and CD86 (B7-2), on antigen- presenting cells (APCs) (Berg M, Zavazava N. Regulation of CD28 expression on CD8+ T cells by CTLA-4. J Leukoc Biol. 2008;83(4):853-863), thereby attenuating T cell activation and proliferation.
- APCs antigen- presenting cells
- the engagement of CTLA-4 with its ligands delivers inhibitory signals to T cells, leading to suppression of immune reactions (Alegre ML, Frauwirth KA, Thompson CB. T-cell regulation by CD28 and CTLA-4. Nat Rev Immunol. 2001;l(3):220-228).
- CTLA-4 competes with CD28 to bind at CD80/86, with CTLA-4 having preferential binding with CD80/86 relative to CD28 (Krummel MF, Allison JP. CD28 and CTLA-4 have opposing effects on the response of T cells to stimulation. J Exp Med. 1995;182(2):459-465).
- TCR T cell receptor
- MHC-II major histocompability complex II
- CTLA-4 Upon TCR stimulation, CTLA-4 circulates to the cell surface, but is then internalized by unphosphorylated cytoplasmic domains and either recycled to the plasma membrane or subjected to lysosomal degradation (Qureshi OS, Kaur S, Hou TZ, et al. Constitutive clathrin-mediated endocytosis of CTLA-4 persists during T cell activation. J Biol Chem. 2012;287( 12):9429-9440).
- ICIs immune checkpoint inhibitors
- they serve to induce an antitumor immune response by blocking immune checkpoints that downregulate intratumoral T-cell responses
- ICIs immune checkpoint inhibitors
- they serve to induce an antitumor immune response by blocking immune checkpoints that downregulate intratumoral T-cell responses
- proof-of-concept experiments supporting anti- CTLA-4 therapy were initially described by James Allison’s research group, which discovered that blockade of CTLA-4 on effector and Treg compartments led to tumor size reduction (Peggs KS, Quezada SA, Chambers CA, Korman AJ, Allison JP.
- CTLA4 particularly sCTLA4’s link to improved growth/survival of cells grown in the presence of primary blood mononuclear cells, suggesting an immunomodulatory effect of CTLA-4 (Kennedy PT, Saulters EL, Duckworth AD, et al. Soluble CTLA-4 attenuates T cell activation and modulates anti-tumor immunity. Mol Ther. 2024;32(2):457-468). [0048] Based on promising preclinical tumor model studies, the clinical potential of antibodies against CTLA-4 has been explored in different human malignancies.
- CTLA-4 anti-CTLA-4
- Tremelimumab marketed as Imjudo
- Treatment and targeting of CTLA-4 is associated with autoimmune like toxicities (Sharma P, Goswami S, Raychaudhuri D, et al. Immune checkpoint therapy-current perspectives and future directions. Cell. 2023; 186(8): 1652- 1669).
- irAEs immune-related adverse events
- the most common irAEs are skin rash, hepatitis, colitis and endocrinopathies, particularly hypopituitarism (Liu Y, Zheng P. Preserving the CTLA-4 Checkpoint for Safer and More Effective Cancer Immunotherapy. Trends Pharmacol Sci. 2020;41(l):4-12).
- CTLA-4 is recycled between the plasma membrane and endosomes by binding to lipopolysaccharide-responsive and beige-like anchor protein (LRBA).
- LRBA lipopolysaccharide-responsive and beige-like anchor protein
- Antibodies that bind CTLA-4 disrupt the recycling process. Consequently, CTLA-4 is systemically directed to lysosomal degradation, and autoimmunity-related adverse effects are developed due to the unstoppable action of cytotoxic T cells.
- complete loss of CTLA-4 recycling leads to increased irAEs, as the immune system engages in off-target actions that disaffect otherwise healthy cells.
- Antibodies are central components of the immune system that help prevent and control pathogen spread while providing long-term protection from reinfection. They are composed of two identical fragment antigen binding (Fab) domains that are coupled through a hinge to a fragment crystallizable (Fc) region.
- Fab fragment antigen binding
- the Fabs provide specificity and mediate target antigen binding, while the Fc region interacts with Fcgamma Receptors (FcgR), a class of receptors that deliver either inhibitory or activatory signals to modulate immune cell responses to regulate immune cell activity (Liu R, Oldham RJ, Teal E, Beers SA, Cragg MS. Fc-Engineering for Modulated Effector Functions-Improving Antibodies for Cancer Treatment. Antibodies (Basel). 2020;9(4)).
- FcgR Fcgamma Receptors
- ADCC antibody-dependent cellular cytotoxicity
- Fc enhancements are afucosylation, a process in which oligosaccharides of the Fc region are removed of focuses - a sugar molecule found in the Fc region of antibodies - that enhances antibody Fc-FcgR interactions, improving T-cell signaling and function (Shinkawa T, Nakamura K, Yamane N, et al. The absence of fucose but not the presence of galactose or bisecting N- acetylgluco s amine of human IgGl complextype oligosaccharides shows the critical role of enhancing antibody-dependent cellular cytotoxicity. J Biol Chem. 2003;278(5):3466-3473).
- CTLA-4 trafficking plays an essential role in supporting the inhibitory signaling required to maintain tolerance, and that it may be possible to exploit the specificity of its intracellular routes to reshape the whole signaling network.
- the present disclosure concerns novel anti-CTLA-4 antibodies or antibody fragments suitable for therapeutic use. Some of these antibodies have been engineered using innovative techniques to confer specific binding properties to human CTLA-4 under different pH conditions.
- the pH sensitivity of the disclosed antibodies may enable them to selectively engage with CTLA-4 in acidic or neutral pH environments, such as within a tumor microenvironment or endosome of a cell.
- the pH sensitivity of the disclosed antibodies may also enable them to dissociate from the target under acidic pH in endosomes, allow physiological CTLA-4 recycling, and reduce adverse effects.
- the disclosed antibodies may provide options for more targeted therapy with reduced irAEs.
- the disclosed anti-CTLA-4 antibodies or antibody fragments may exhibit distinct physiochemical or biological properties that lead to significant improvements in efficacy and safety relative to anti-CTLA-4 antibodies known in the art.
- provided anti-CTLA-4 antibodies or antibody fragments thereof possess pH dependent binding properties to human CTLA-4 that facilitates the dissociation of antibody from the antibody:CTLA-4 complex upon internalization into a cell, allowing either the antibody, CTLA-4, or both to be recycled instead of being degraded in the lysosome.
- the antibody or antigen-binding fragment thereof preserves high affinity for CTLA-4 in neutral pH environments (such as at pH>7), lower affinity in acidic environments (such as at pH 5.5-7 found in early endosomes), and significantly reduced affinity for CTLA-4 at more acidic environments (such as at pH ⁇ 5.5 found in late endosomes and lysosomes).
- provided anti-CTLA-4 antibodies or antibody fragments thereof have a lower binding affinity (i.e. greater dissociation constant) to CTLA-4 at an acidic pH as present in endosomes of a cell in comparison with the binding affinity of CTLA-4 present in normal tissue.
- the dissociation constant (KD) for binding at acidic pH e.g., pH 5.0 to 6.0
- the dissociation constant for binding at physiological pH e.g., e.g. pH 7.35 to 7.45.
- these anti-CTLA-4 antibodies or antibody fragments have reduced side-effects, in comparison with monoclonal anti-CTLA-4 antibodies known in the art. This may permit use of higher dosages of these anti-CTLA-4 antibodies or antibody fragments, thus providing a more effective therapeutic option without a corresponding significant increase in side effects.
- the provided antibodies can be used to block inhibitory activity of CTLA-4 by CTLA-4-expressing cells including regulatory T cells (Tregs), and in some cases may also mediate depletion of such cells by an ADCC mechanism.
- Tregs regulatory T cells
- depletion of Tregs in the tumor contributes to anti-tumor efficacy of the provided antibodies.
- provided antibodies reduce CTLA-4 mediated signaling in effector T cells and exhibit activity to deplete Tregs in the tumor microenvironment that might otherwise suppress an anti-tumor immune response.
- the antibody or antigen-binding fragment thereof or a pharmaceutical composition comprising the antibody or antigen-binding fragment thereof induces strong deletion of Treg and local T cell activation in tumor microenvironment but minimal systemic T cell activation.
- the antibody or antigen-binding fragment thereof or a pharmaceutical composition comprising the antibody or antigen-binding fragment thereof induces a favorable ratio of Treg cells to T effector cells in the tumor microenvironment.
- anti-CTLA-4 antibodies including full-length antibodies or antigen-binding fragments, that contain a heavy chain variable region (VH) sequence and a light chain variable region (VL) sequence as described, or a sufficient antigen-binding portion thereof.
- VH is the region of the anti-CTLA-4 antibody that comprises the three heavy chain complementarity determining regions (CDRs)
- VL chain is the region of the anti-CTLA-4 antibody that comprises the three light chain CDRs.
- provided anti-CTLA-4 antibodies including full-length antibodies or antigen-binding fragments, are antibodies that contains a VH region sequence that contains a CDR-H1, a CDR- H2 and a CDR-H3 as described and contains a VL region sequence that contains a CDR-L1, a CDR-L2 and a CDR-L3 as described.
- the antibodies include antibodies that specifically bind to CTLA-4, e.g., human CTLA-4.
- the provided anti-CTLA-4 antibodies are human antibodies, or antibodies that are modified from or variant of human antibodies.
- the antibodies include isolated antibodies.
- CTLA-4-binding molecules containing such antibodies, such as single-chain proteins, fusion proteins, conjugates and/or recombinant receptors such as chimeric antigen receptors.
- an CTLA-4-targeted antibody-drug conjugate comprising an anti-CTLA-4 antibody or antigen binding fragment thereof described herein.
- a bispecific immune cell engager that is a fusion protein comprising the anti-CTLA-4 antibody or antigen binding fragment thereof described herein.
- the bispecific immune cell engager is a bispecific T cell engager.
- the CTLA-4-binding molecules include isolated molecules.
- polynucleotides containing nucleic acids sequences encoding all or a portion of such antibodies, including an antigen-binding fragment, or binding molecule can be incorporated into constructs, such as deoxyribonucleic acid (DNA) or RNA constructs, such as those that can be introduced into cells for expression of the encoded anti-CTLA-4 antibodies or binding molecules.
- DNA deoxyribonucleic acid
- RNA constructs such as those that can be introduced into cells for expression of the encoded anti-CTLA-4 antibodies or binding molecules.
- antibody herein is used in the broadest sense and includes polyclonal and monoclonal antibodies, including intact antibodies and functional (antigen-binding) antibody fragments, including fragment antigen binding (Fab) fragments, F(ab’)2 fragments, Fab’ fragments, Fv fragments, recombinant IgG (rlgG) fragments, heavy chain variable (VH) regions capable of specifically binding the antigen, single chain antibody fragments, including single chain variable fragments (scFv), and single domain antibodies e.g., sdAb, sdFv, nanobody, VHH) fragments.
- Fab fragment antigen binding
- rlgG fragment antigen binding
- VH heavy chain variable
- immunoglobulins such as intrabodies, peptibodies, chimeric antibodies, fully human antibodies, humanized antibodies, and heteroconjugate antibodies, multispecific, e.g., bispecific or trispecific, antibodies, diabodies, triabodies, and tetrabodies, tandem di-scFv, tandem tri-scFv.
- antibody should be understood to encompass functional antibody fragments thereof also referred to herein as “antigen-binding fragments.”
- the term also encompasses intact or full-length antibodies, including antibodies of any class or sub-class, including IgG and sub-classes thereof, IgM, IgE, IgA, and IgD.
- CDR complementarity determining region
- HVR hypervariable region
- CDR-H1, CDR-H2, CDR-H3 three CDRs in each heavy chain variable region
- CDR-L1, CDR-L2, CDR-L3 three CDRs in each light chain variable region
- “Framework regions” and “FR” are known to refer to the non-CDR portions of the variable regions of the heavy and light chains.
- FR-H1, FR-H2, FR-H3, and FR-H4 there are four FRs in each full-length heavy chain variable region (FR-H1, FR-H2, FR-H3, and FR-H4), and four FRs in each full-length light chain variable region (FR-L1, FR-L2, FR-L3, and FR-L4).
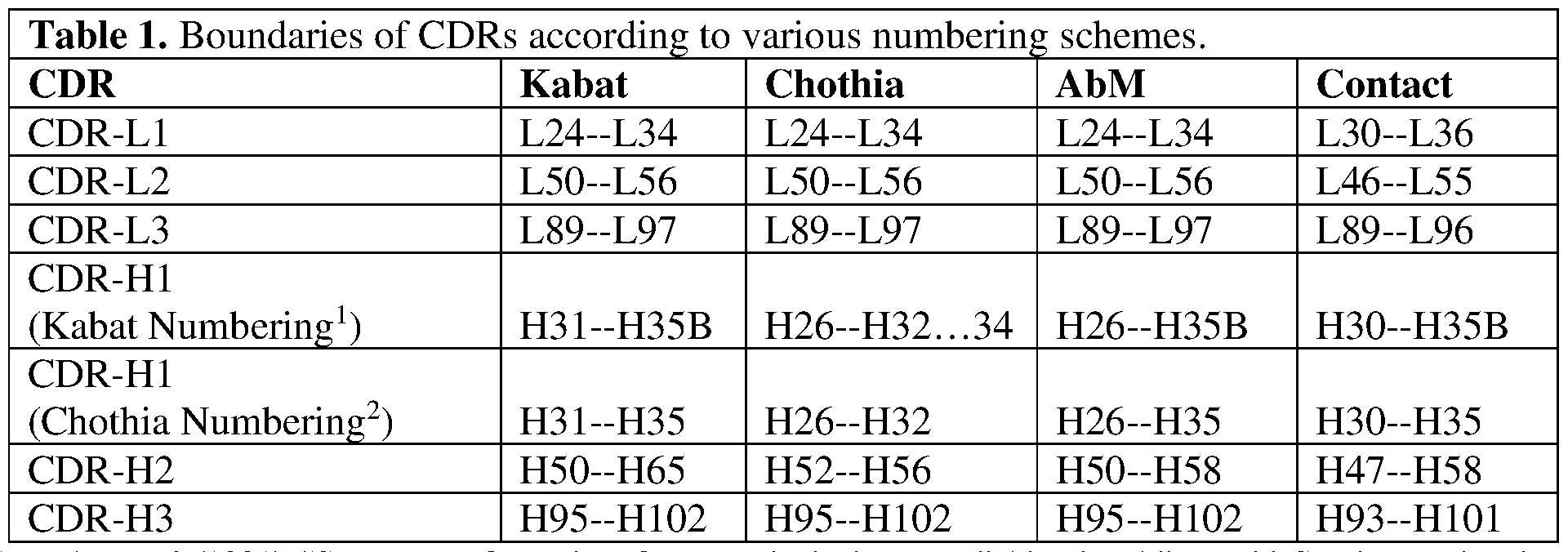
- the boundaries of a given CDR or FR may vary depending on the scheme used for identification.
- the Kabat scheme is based on structural alignments
- the Chothia scheme is based on structural information. Numbering for both the Kabat and Chothia schemes is based upon the most common antibody region sequence lengths, in some cases with insertions. Insertions in the sequence relative to the standard numbering scheme are indicated using insertion letter codes. For example, residues that are inserted between residues L30 and L31 are indicated as L31A, L31B, etc. Deletions in the sequence relative to the standard scheme are accommodated by skipping numbers. The two schemes place certain insertions and deletions (“indels”) at different positions, resulting in differential numbering.
- the Chothia numbering scheme is nearly identical to the Kabat numbering scheme, except that insertions are placed at structural positions and topologically equivalents residues do get assigned the same numbers.
- the Contact scheme is based on analysis of complex crystal structures and is similar in many respects to the Chothia numbering scheme.
- the AbM scheme is a compromise between Kabat and Chothia definitions based on that used by Oxford Molecular’s AbM antibody modeling software.
- the IgBLAST scheme is based on matching to germline V, D and J genes, and can be determined using National Center for Biotechnology Information (NCBI)’s IgBLAST tool.
- Kabat numbering can be determined by known sequence rules as described in, for example, Kabat et al. (1991), “Sequences of Proteins of Immunological Interest,” 5th Ed. Public Health Service, National Institutes of Health, Bethesda, MD.
- the Kabat numbering scheme in some aspects can include any of the following rules to designate CDRs: CDR-L1 starts at approximately residue 24 of the light chain, always has a preceding C residue, and always has a following W residue; the end of CDR-L1 is defined by a stretch of 3 residues, where the W residue can be followed by Y, L, or F, followed by Q or L; CDR-1 has a length of 10 to 17 residues; CDR-L2 always starts 16 residues after the end of CDR-L1; the two residues before CDR-L2 are I and Y but can also be V and Y, I and K, or I and F; CDR-L2 is always 7 residues long; CDR-L3 always starts 33 residues after the end of CDR-L2, always has a preceding C residue, and is strictly followed by a F-G-X-G sequence motif, where X is any amino acid; CDR-L3 has a length of 7 to 11 residues; CDR-H1
- boundary positions of certain CDRs can differ based on different definitions for the CDRs (See e.g., Martin, ACR, Antibody Information: How to identify the CDRs by looking at a sequence [online] bioinf.org.uk/abs/info.html).
- the boundary positions for CDR-F1 according to Chothia numbering can be F26— F32 (Chothia et al., Science, 1986; 233(4765):755-8 and Chothia C. and Fesk A.M. J Mol Biol, 1987; 196(4):901-17).
- the boundary positions for CDR-F1 can be E25--E32 (Al-Fazikani et al., J Mol Biol, 1997; 273(4):927-48).
- the boundary positions for CDR-F2 can be F50— F52 and for CDR-F3 can be F91— F96 (Chothia et al., Science, 1986; 233(4765):755-8; Chothia C. and Fesk A.M. J Mol Biol, 1987; 196(4):901-17; and Al-Fazikani et al., J Mol Biol, 1997; 273(4):927-48).
- the boundary positions for CDR-H1 according to Chothia numbering can be H26— H32 (Chothia et al., Science, 1986; 233(4765):755-8; Chothia C. and Fesk A.M. J Mol Biol, 1987; 196(4):901-17; and Al-Fazikani et al., J Mol Biol, 1997; 273(4):927-48).
- the boundary positions for CDR-H2 can be H53— H55 (Chothia et al., Science, 1986; 233(4765):755-8 and Chothia C. and Fesk A.M.
- the boundary positions for CDR-H3 can be H96— H101 (Chothia et al., Science, 1986; 233(4765):755-8 and Chothia C. and Fesk A.M. J Mol Biol., 1987; 196(4):901-17).
- the boundary positions for CDR-H3 can be H92— H104 (Morea et al., Biophys Chem, 1997; 68(1-3): 9-16 and Morea et al., J Mol Biol., 1998; 275(2): 269-94).
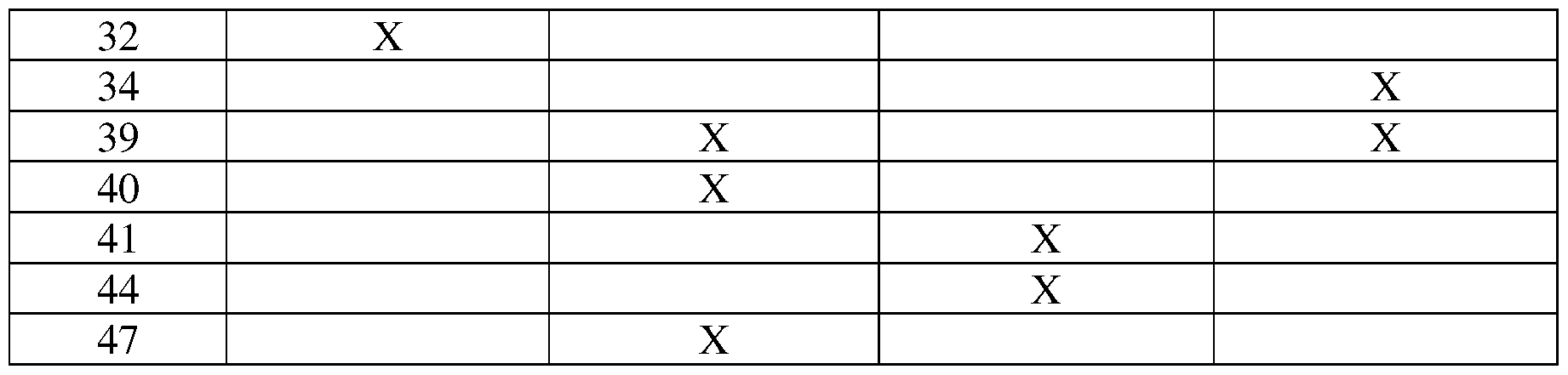
- Table 1 exemplifies exemplary numbering and lists exemplary position boundaries of CDR-L1, CDR-L2, CDR-L3 and CDR-H1, CDR-H2, CDR-H3 as identified by Kabat, Chothia, AbM, and Contact schemes, respectively.
- residue numbering is listed using both the Kabat and Chothia numbering schemes.
- FRs are located between CDRs, for example, with FR-L1 located before CDR-L1, FR-L2 located between CDR-L1 and CDR- L2, FR-L3 located between CDR-L2 and CDR-L3 and so forth. It is noted that because the shown Kabat numbering scheme places insertions at H35A and H35B, the end of the Chothia CDR-H1 loop when numbered using the shown Kabat numbering convention varies between
- CDR complementary determining region
- individual specified CDRs e.g., CDR-H1, CDR-H2, CDR-H3
- CDR-H1, CDR-H2, CDR-H3 individual specified CDRs
- a particular CDR e.g., a CDR-H3
- a CDR-H3 contains the amino acid sequence of a corresponding CDR in a given VH or VL region amino acid sequence
- a CDR has a sequence of the corresponding CDR (e.g., CDR-H3) within the variable region, as defined by any of the aforementioned schemes, or other known schemes.
- an antibody or antigen-binding fragment thereof comprises a CDR-H1, a CDR-H2, and a CDR-H3 as contained within a given VH region amino acid sequence and a CDR-L1, a CDR- L2, and a CDR-L3 as contained within a given VL region amino acid sequence
- the CDRs can be defined by any of the aforementioned schemes, such as Kabat, Chothia, AbM, IgBLAST, IMGT, or Contact method, or other known scheme.
- specific CDR sequences are specified. Exemplary CDR sequences of provided antibodies are described using various numbering schemes, although it is understood that a provided antibody can include CDRs as described according to any of the other aforementioned numbering schemes or other known numbering schemes.
- FR or individual specified FR(s) e.g., FR- Hl, FR-H2, FR-H3, FR-H4, FR-L1, FR-L2, FR-L3, and/or FR-L4
- FR- Hl FR- Hl
- FR- Hl FR- Hl, FR-H2, FR-H3, FR-H4, FR-L1, FR-L2, FR-L3, and/or FR-L4
- the scheme for identification of a particular CDR, FR, or FRs or CDRs is specified, such as the CDR as defined by the Kabat, Chothia, AbM, IgBLAST, IMGT, or Contact method, or other known schemes.
- the particular amino acid sequence of a CDR or FR is given.
- an antibody or antigen-binding fragment thereof comprises a FR-H1, a FR-H2, a FR-H3, and a FR-H4 as contained within a given VH region amino acid sequence and a FR-L1, a FR-L2, a FR-L3, and a FR-L4 as contained within a given VL region amino acid sequence
- the FRs can be defined by any of the aforementioned schemes, such as Kabat, Chothia, AbM, IgBLAST, IMGT, or Contact method, or other known scheme.
- variable region refers to the domain of an antibody heavy or light chain that is involved in binding the antibody to antigen.
- the variable regions of the heavy chain and light chain (VH and VL, respectively) of a native antibody generally have similar structures, with each domain comprising four conserved framework regions (FRs) and three CDRs (See, e.g., Kindt et al. Kuby Immunology, 6th ed., W.H. Freeman and Co., page 91 (2007)).
- FRs conserved framework regions
- antibodies that bind a particular antigen may be isolated using a VH or VL domain from an antibody that binds the antigen to screen a library of complementary VL or VH domains, respectively. See, e.g., Portolano et al., J. Immunol. 150:880-887 (1993); Clarkson et al., Nature 352:624-628 (1991).
- an “antibody fragment” or “antigen-binding fragment” refers to a molecule other than an intact antibody that comprises a portion of an intact antibody that binds the antigen to which the intact antibody binds.
- an antigen binding fragment includes all CDRs of a variable heavy chain (VH) and variable light chain (VL) sequence from antibodies that bind CTLA-4 set forth herein.
- antibody fragments include but are not limited to Fv, Fab, Fab’, Fab’-SH, F(ab’)2; diabodies; linear antibodies; heavy chain variable (VH) regions, single-chain antibody molecules such as scFvs and single-domain antibodies comprising only the VH region; and multispecific antibodies formed from antibody fragments.
- Papain digestion of antibodies produce two identical antigen-binding fragments, called “Fab” fragments, and a residual “Fc” fragment, a designation reflecting the ability to crystallize readily.
- the Fab fragment is composed of an entire E chain along with the variable region domain of the H chain (VH), and the first constant domain of one heavy chain (CHI).
- Each Fab fragment is monovalent with respect to antigen binding, i.e., it has a single antigen-binding site.
- Pepsin treatment of an antibody yields a single large F(ab’)2 fragment which roughly corresponds to two disulfide linked Fab fragments having different antigen-binding activity and is still capable of crosslinking antigen.
- Fab’ fragments differ from Fab fragments by having a few additional residues at the carboxy terminus of the CHI domain including one or more cysteines from the antibody hinge region.
- Fab’-SH is the designation herein for Fab’ in which the cysteine residue(s) of the constant domains bear a free thiol group.
- the antibody is or comprises an antibody fragment comprising a variable heavy chain (VH) and a variable light chain (VL) region.
- the antibodies are single-chain antibody fragments comprising a heavy chain variable (VH) region and/or a light chain variable (VL) region, such as scFvs.
- Antibody fragments can be made by various techniques, including but not limited to proteolytic digestion of an intact antibody as well as production by recombinant host cells.
- the antibodies are recombinantly -produced fragments, such as fragments comprising arrangements that do not occur naturally, such as those with two or more antibody regions or chains joined by synthetic linkers, e.g. , peptide linkers, and/or that are may not be produced by enzyme digestion of a naturally-occurring intact antibody.
- the antibody fragments are scFvs.
- F(ab) refers to two of the protein fragments resulting from proteolytic cleavage of immunoglobulin G (IgG) molecules by the enzyme papain. Each F(ab) comprises a covalent heterodimer of the VH chain and VL chain and includes an intact antigen-binding site.
- F(ab)2 refers to a protein fragment of IgG generated by proteolytic cleavage by the enzyme pepsin. Each F(ab’)2 fragment comprises two F(ab) fragments, thus comprising both antigen-binding sites.
- An “Fv fragment” for use according to certain embodiments of the present invention can be produced by preferential proteolytic cleavage of an IgM, and on rare occasions of an IgG or IgA immunoglobulin molecule. Fv fragments are, however, more commonly derived using recombinant techniques known in the art.
- the Fv fragment includes a non-covalent VH"VL heterodimer including an antigen-binding site which retains much of the antigen recognition and binding capabilities of the native antibody molecule, but lacking the CHI and CL domains contained within a Fab.
- single chain Fv (scFv) antibodies are contemplated and may be prepared using standard molecular biology techniques following the teachings of the present application with regard to selecting antibodies having the desired specificity.
- chimeric antibodies may be made.
- a chimeric antibody may comprise CDRs and framework regions from different antibodies. These antibodies may be produced through recombinant molecular biological techniques or may be physically conjugated together.
- a scFv polypeptide is a covalently linked VH"VL heterodimer which is expressed from a gene fusion including VH- and Vi.-cncoding genes linked by a peptide-encoding linker.
- a number of methods have been described to discern chemical structures for converting the naturally aggregated — but chemically separated — light and heavy polypeptide chains from an antibody V region into an scFv molecule which will fold into a three dimensional structure substantially similar to the structure of an antigen-binding site. See, e.g., U.S. Pat. Nos. 5,091,513 and 5,132,405, to Huston et al. and U.S. Pat. No. 4,946,778, to Eadner et al.
- the antibodies described herein may be provided in the form of a UniBody®.
- a UniBody® is an IgG4 antibody with the hinge region removed (see GenMab Utrecht, The Netherlands; see also, e.g., US20090226421). This antibody technology creates a stable, smaller antibody format with an anticipated longer therapeutic window than current small antibody formats. IgG4 antibodies are considered inert and thus do not interact with the immune system. Fully human IgG4 antibodies may be modified by eliminating the hinge region of the antibody to obtain half-molecule fragments having distinct stability properties relative to the corresponding intact IgG4 (GenMab, Utrecht).
- the UniBody® Halving the IgG4 molecule leaves only one area on the UniBody® that can bind to cognate antigens (e.g., disease targets) and the UniBody® therefore binds univalently to only one site on target cells. For certain cancer cell surface antigens, this univalent binding may not stimulate the cancer cells to grow as may be seen using bivalent antibodies having the same antigen specificity, and hence UniBody® technology may afford treatment options for some types of cancer that may be refractory to treatment with conventional antibodies.
- the small size of the UniBody® can be a great benefit when treating some forms of cancer, allowing for better distribution of the molecule over larger solid tumors and potentially increasing efficacy.
- the antibodies of the present disclosure may be chimeric antibodies.
- a chimeric antibody is comprised of an antigen-binding fragment of an anti-CTLA-4 antibody operably linked or otherwise fused to a heterologous Fc portion of a different antibody.
- the heterologous Fc domain is of human origin.
- the heterologous Fc domain may be from a different Ig class from the parent antibody, including IgA (including subclasses IgAl and IgA2), IgD, IgE, IgG (including subclasses IgGl, IgG2, IgG3, and IgG4), and IgM.
- the heterologous Fc domain may be comprised of CH2 and CH3 domains from one or more of the different Ig classes.
- the antibodies of the present disclosure may be “non- naturally occurring” antibodies.
- Non-naturally occurring antibodies can refer to antibodies that comprise one or more amino acid modifications, such that the resultant antibody is substantially non-naturally occurring (e.g., does not exists in nature). These amino acid modifications can include point mutations, wherein a naturally occurring amino acid is substituted for another naturally occurring amino acid. In some embodiments, the amino acid modifications can include point mutations wherein a non-naturally occurring amino acid is substituted for a naturally occurring amino acid.
- Non-naturally occurring antibodies can also refer to antibodies that are conjugated to a heterologous protein or compound, such as a detectable marker. [0083] Among the provided antibodies are human antibodies.
- a “human antibody” is an antibody with an amino acid sequence corresponding to that of an antibody produced by a human or a human cell, or non-human source that utilizes human antibody repertoires or other human antibody-encoding sequences, including human antibody libraries.
- the term excludes humanized forms of non-human antibodies comprising non-human antigen-binding regions, such as those in which all or substantially all CDRs are non-human.
- the term includes antigenbinding fragments of human antibodies.
- Human antibodies may be prepared by administering an immunogen to a transgenic animal that has been modified to produce intact human antibodies or intact antibodies with human variable regions in response to antigenic challenge. Such animals typically contain all or a portion of the human immunoglobulin loci, which replace the endogenous immunoglobulin loci, or which are present extrachromosomally or integrated randomly into the animal’s chromosomes. In such transgenic animals, the endogenous immunoglobulin loci have generally been inactivated. Human antibodies also may be derived from human antibody libraries, including phage display and cell-free libraries, containing antibody-encoding sequences derived from a human repertoire.
- monoclonal antibodies including monoclonal antibody fragments.
- the term “monoclonal antibody” as used herein refers to an antibody obtained from or within a population of substantially homogeneous antibodies, i.e., the individual antibodies comprising the population are identical, except for possible variants containing naturally occurring mutations or arising during production of a monoclonal antibody preparation, such variants generally being present in minor amounts.
- polyclonal antibody preparations which typically include different antibodies directed against different epitopes
- each monoclonal antibody of a monoclonal antibody preparation is directed against a single epitope on an antigen.
- the term is not to be construed as requiring production of the antibody by any particular method.
- a monoclonal antibody may be made by a variety of techniques, including but not limited to generation from a hybridoma, recombinant DNA methods, phage-display and other antibody display methods.
- the antibody or the antigen-binding fragments of the antibody is isolated.
- variants of the antibodies disclosed herein contemplates variants of the antibodies disclosed herein.
- such variant antibodies or antigen-binding fragments, or CDRs thereof bind to CTLA-4 at least about 50%, at least about 70%, at least about 80%, at least about 85%, at least about 90% and in certain embodiments, at least about 95% as well as an antibody sequence specifically set forth herein.
- such variant antibodies or antigen-binding fragments, or CDRs thereof bind to CTLA-4 with greater affinity than the antibodies set forth herein, for example, that bind quantitatively at least about 105%, 106%, 107%, 108%, 109%, or 110% as well as an antibody sequence specifically set forth herein.
- a subject antibody may have: a) a heavy chain variable region having an amino acid sequence that is at least 80% identical, at least 85% identical, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, or 99% or 100% identical, to a heavy chain variable region of an anti-CTLA-4 antibody described herein; and b) a light chain variable region having an amino acid sequence that is at least 80% identical, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, or 99% or 100% identical, to a light chain variable region of an anti-CTLA-4 antibody described herein.
- the anti-CTLA-4 antibody contains a VH region and a VL region containing a combination of six CDRs as described below.
- the anti-CTLA-4 antibody contains a VH region and a VL region as described below.
- a VH region sequence can be any of the VH region sequence described herein.
- a VL region sequence can be any of the VL region sequence described herein.
- any of the VH region sequence and any of the VL region sequence described herein can be used in combination.
- the antibody is an antigen-binding fragment, such as a Fab or an scFv.
- the antibody or antigen-binding fragment further comprises at least a portion of an immunoglobulin constant region or a variant thereof.
- the antibody is a full-length antibody that also contains a constant region.
- an anti- CTLA-4 antibody comprises at least one heavy chain comprising a VH region and at least a portion of a heavy chain constant region, and at least one light chain comprising a VL region and at least a portion of a light chain constant region.
- an anti-CTLA-4 antibody comprises two heavy chains, wherein each heavy chain comprises a VH region and at least a portion of a heavy chain constant region, and two light chains, wherein each light chain comprises a VL region and at least a portion of a light chain constant region.
- an anti-CTLA-4 antibody comprises two heavy chains, wherein each heavy chain comprises a VH region and a heavy chain constant region, and two light chains, wherein each light chain comprises a VL region and a light chain constant region.
- Anti-CTLA-4 antibodies include any combination of the heavy chain and light chain complementarity-determining regions (CDRs) discussed herein.
- the anti-CTLA-4 antibody or antigen-binding fragment thereof comprises any one of the CDR-H1 as described herein, any one of the CDR- H2 as described herein, any one of the CDR-H3 as described herein, any one of the CDR-L1 as described herein, any one of the CDR-L2 as described herein and any one of the CDR-L3 as described herein.
- any one or more of the CDR-H1, the CDR-H2 and the CDR-H3 sequences described herein, and any one or more of the CDR-L1, the CDR-L2 and the CDR-L3 sequences described herein can be used in combination.
- a provided anti-CTLA-4 antibody or an antigen-binding fragment thereof has a CDR-H1, a CDR-H2 and a CDR-H3 present in a VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112.
- a provided anti-CTLA-4 antibody or an antigen-binding fragment thereof has a CDR-L1, a CDR-L2 and a CDR-L3 present in a VL region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118.
- the provided anti- CTLA-4 antibody contains a combination of any of such six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) described above).
- a provided anti-CTLA-4 antibody or an antigen-binding fragment thereof has a CDR-H1, a CDR- H2 and a CDR-H3 present in a VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112, and a CDR-L1, a CDR-L2 and a CDR-L3 present in a VL region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118, wherein the CDR-H1, CDR- H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3 are not set forth in SEQ ID NOS: 49, 57, 68, 74, 89 and 98, respectively.
- the combination of six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) is according to Kabat numbering. In some embodiments, the combination of six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) is according to Chothia numbering.
- the combination of six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) is according to AbM numbering, In some embodiments, the combination of six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) is according to IMGT numbering. In some embodiments, the combination of six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) is according to IgBLAST numbering.
- the VH region contains a CDR-H1 set forth in any one of SEQ ID NOs: 49-56, a CDR-H2 set forth in any one of SEQ ID NOs: 57- 67, and a CDR-H3 set forth in any one of SEQ ID NOs: 68-73; and the VL region contains a CDR-L1 set forth in any one of SEQ ID NOs: 74-88, a CDR-L2 set forth in any one of SEQ ID NOs: 89-97, and a CDR-L3 set forth in any one of SEQ ID NOs: 98-100.
- the anti-CTLA-4 antibody or antigen-binding fragment does not contain the combination of CDRs set forth in SEQ ID NO: 49, SEQ ID NO: 57, SEQ ID NO: 68, SEQ ID NO: 74, SEQ ID NO: 89, and SEQ ID NO: 98.
- the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3 are not set forth in SEQ ID NOS: 49, 57, 68, 74, 89 and 98, respectively.
- the VH region comprises a CDR-H1 comprising the sequence set forth in in any one of SEQ ID NOs: 49-56, a CDR-H2 comprising the sequence set forth in any one of SEQ ID NOs: 57-67, and a CDR-H3 comprising the sequence set forth in any one of SEQ ID NOs: 68-73
- the VL region comprises a CDR-L1 comprising the sequence set forth in any one of SEQ ID NOs: 75-88, a CDR-L2 comprising the sequence set forth in any one of SEQ ID NOs: 90-97; and a CDR-L3 comprising the sequence set forth in any one of SEQ ID NOs: 99 and 100.
- the VH region comprises a CDR- H1 comprising the sequence set forth in any one of SEQ ID NOs: 50-56, a CDR-H2 comprising the sequence set forth in any one of SEQ ID NOs: 58-67, and a CDR-H3 comprising the sequence set forth in any one of SEQ ID NOs: 69-73
- the VL region comprises a CDR-L1 comprising the sequence set forth in any one of SEQ ID NOs: 74-88, a CDR-L2 comprising the sequence set forth in any one of SEQ ID NOs: 89-97; and a CDR-L3 comprising the sequence set forth in any one of SEQ ID NOs: 98-100.
- antibodies are those having sequences at least at or about 90%, at or about 91%, at or about 92%, at or about 93%, at or about 94%, at or about 95%, at or about 96%, at or about 97%, at or about 98%, or at or about 99% identical to any such CDR sequence, e.g., any of the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2, CDR-L3.
- among the antibodies are those in which a CDR contained therein has no more than 2 amino acid difference compared to any such above CDR sequence, e.g., any of the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2, CDR-L3. In some embodiments, among the antibodies are those in which a CDR contained therein has no more than 1 amino acid difference compared to any such above CDR sequence, e.g., any of the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2, CDR-L3.
- the VH region contains a CDR-H1 set forth in any one of SEQ ID NOs: 50-56, a CDR-H2 set forth in any one of SEQ ID NOs: 58- 67, and a CDR-H3 set forth in any one of SEQ ID NOs: 69-73; and the VL region contains a CDR-L1 set forth in any one of SEQ ID NOs: 75-88, a CDR-L2 set forth in any one of SEQ ID NOs: 90-97, and a CDR-L3 set forth in any one of SEQ ID NOs: 99-100.
- the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 1. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 2. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 3.
- the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 4. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 5. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 6.
- the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 7. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 8. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 9.
- the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 10. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 11. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 12.
- the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 13. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 14. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 15.
- the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 16. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 17. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 18.
- the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 19. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 20. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 21.
- the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 22. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 23. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 24.
- the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 25. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 26. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 27.
- the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 112.
- the CDR sequence is according to Kabat numbering.
- the CDR sequence is according to Chothia numbering.
- the CDR sequence is according to AbM numbering.
- the CDR sequence is according to IMGT numbering.
- the CDR sequence is according to IgBLAST numbering.
- the antibody or antigen -binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 28. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR- Ll, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 29. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR- L3 present in SEQ ID NO: 30.
- the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 31. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 32. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 33.
- the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 34. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 35. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR- Ll, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 36.
- the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR- L3 present in SEQ ID NO: 37. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 38. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 39.
- the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 40. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 41. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 42.
- the antibody or antigen-binding fragment has a VL region that contains a CDR- Ll, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 43. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR- L3 present in SEQ ID NO: 44. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 46.
- the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 46. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 47. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ
- the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 118.
- the CDR sequence is according to Kabat numbering.
- the CDR sequence is according to Chothia numbering.
- the CDR sequence is according to AbM numbering.
- the CDR sequence is according to IMGT numbering.
- the CDR sequence is according to IgBLAST numbering.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 58, and 68, respectively; SEQ ID NOS: 49, 57, and 69, respectively; SEQ ID NOS: 51, 57, and 68, respectively; SEQ ID NOS: 50, 59, and 68, respectively; SEQ ID NOS: 52, 60, and 68, respectively; SEQ ID NOS: 52, 61, and 68, respectively; SEQ ID NOS: 49, 49, 58, and 68, respectively; SEQ ID NOS: 49, 57, and 69, respectively; SEQ ID NOS: 51, 57, and 68, respectively; SEQ ID NOS: 50, 59, and 68, respectively; SEQ ID NOS: 52, 60, and 68, respectively; SEQ ID NOS: 52, 61, and 68, respectively; SEQ ID NOS: 49, 49, 58, and 68
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 75, 89, and 98, respectively; SEQ ID NOS: 76, 89, and 98, respectively; SEQ ID NOS: 77, 89, and 98, respectively; SEQ ID NOS: 78, 90, and 98, respectively; SEQ ID NOS: 79, 90, and 98, respectively; SEQ ID NOS: 80, 91, and 98, respectively; SEQ ID NOS: 78, 89, and 99, respectively; SEQ ID NOS: 81, 92, and 98, respectively; SEQ ID NOS: 82, 89, and 98, respectively; SEQ ID NOS: 82, 93, and 98, respectively; SEQ ID NOS: 83, 92, and 98, respectively; SEQ ID NOS: 83, 92, and 98, respectively; SEQ ID NOS: 83,
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 58, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 69, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 51, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 59, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 61, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 51, 62, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 63, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 57, and 69, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 53, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 53, 57, and 70, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 54, 57, and 71, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 55, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 56, 57, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 64, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 60, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 66, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 71, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 73, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 75, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 77, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 78, 90, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 79, 90, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 80, 91, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 78, 89, and 99, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 81, 92, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 82, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 82, 93, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 83, 92, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 100, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 86, 94, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 87, 95, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 96, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 88, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 97, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 88, 89, and 100, respectively.
- the provided antibody is a human antibody or antigenbinding fragment thereof that contains a framework region that contains human germline gene segment sequences.
- the human antibody contains a VH region in which the framework region, e.g.FRl, FR2, FR3 and FR4, has at least 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a framework region encoded by a human germline antibody segment, such as a V segment and/or J segment.
- the human antibody contains a VL region in which the framework region e.g.FRl, FR2, FR3 and FR4, has at least 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a framework region encoded by a human germline antibody segment, such as a V segment and/or J segment.
- the framework region sequence contained within the VH region and/or VL region differs by no more than 10 amino acids, such as no more than 9, 8, 7, 6, 5, 4, 3, 2 or 1 amino acid, compared to the framework region sequence encoded by a human germline antibody segment.
- the VH region comprises any of the CDR-H1, CDR-H2 and CDR-H3 as described and comprises a framework region 1 (FR1), a FR2, a FR3 and/or a FR4 having at least at or about 95%, at or about 96%, at or about 97%, at or about 98%, or at or about 99% sequence identity, respectively, to a FR1, a FR2, a FR3 and/or a FR4 contained within the VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112.
- FR1 framework region 1
- FR2 a FR2
- FR3 and/or a FR4 having at least at or about 95%, at or about 96%, at or about 97%, at or about 98%, or at or about 99% sequence identity, respectively, to a FR1, a FR2, a FR3 and/or a FR4 contained within the VH region amino acid sequence set forth in any one of SEQ ID
- the VH region comprises any of the CDR-H1, CDR-H2 and CDR-H3 as described and comprises a framework region 1 (FR1), a FR2, a FR3 and/or a FR4 contained within the VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112.
- the VH region comprises any of the CDR-L1, CDR-L2 and CDR-L3 as described and comprises a framework region 1 (FR1), a FR2, a FR3 and/or a FR4 having at least at or about 95%, at or about 96%, at or about 97%, at or about 98%, or at or about 99% sequence identity, respectively, to a FR1, a FR2, a FR3 and/or a FR4 contained within the VL region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118.
- FR1 framework region 1
- FR2 a FR3 and/or a FR4 having at least at or about 95%, at or about 96%, at or about 97%, at or about 98%, or at or about 99% sequence identity, respectively, to a FR1, a FR2, a FR3 and/or a FR4 contained within the VL region amino acid sequence set forth in any one of SEQ ID NOs:
- the VL region comprises any of the CDR-H1, CDR-H2 and CDR-H3 as described and comprises a framework region 1 (FR1), a FR2, a FR3 and/or a FR4 contained within the VH region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118.
- FR1 framework region 1
- FR2 framework region 2
- FR3 framework region 3
- FR4 contained within the VH region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118.
- any of the provided anti-CTLA-4 antibodies or antigen binding fragments has a VH region having the amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112, and has a VL region having the amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 1, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 1.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 2, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 2.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 3, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 3.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 4, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 4.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 5, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 5.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 6, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO:6.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 7, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 7.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 8, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 8.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 9, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 9.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 10, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 10.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 11, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 11.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 12, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 12.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 13, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 13.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 14, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 14.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 15, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 15.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 16, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 16.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 17, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 17.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 18, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 18.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 19, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 19.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 20, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 20.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 21, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 21.
- the antibody or antigen-binding fragment has has a VH region having the amino acid sequence set forth in SEQ ID NO: 22, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 22.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 23, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 23.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 24, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 24.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 25, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 25.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 26, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 26.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 27, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 27.
- the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 112, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 112. In some embodiments, the antibody or antigen-binding fragment does not have a VH region having the amino acid sequence set forth in SEQ ID NO: 112.
- the antibody or antigen -binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 28, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 28.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 29, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 29.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 30, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 30.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 31, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 31.
- a VL region having the amino acid sequence set forth in SEQ ID NO: 32 or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 32.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 33, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 33.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 34, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 34.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 35, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 35.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 36, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 36.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 37, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 37.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 38, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 38.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 39, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 39.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 40, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 40.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 41, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 41.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 42, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 42.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 43, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 43.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 44, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 44.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 45, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 45.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 46, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 46.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 47, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 47.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 48, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 48.
- the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 118, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 118. In some embodiments, the antibody or antigen-binding fragment does not have a VL region having the amino acid sequence set forth in SEQ ID NO: 118.
- the antibody or antigen-binding fragment has a VH region having an amino acid sequence that is less than 100% identical to SEQ ID NO: 112 and has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 112.
- the antibody or antigen-binding fragment has a VL region having an amino acid sequence that is less than 100% identical to SEQ ID NO: 118 and has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 118.
- the VH region does not have the amino acid sequence set forth in SEQ ID NO: 112.
- the VL region does not have the amino acid sequence set forth in SEQ ID NO: 118.
- the VH region and the VL region of the antibody are not set forth SEQ ID NO: 112 and 118, respectively.
- the antibody or antigen-binding fragment has a VH region having an amino acid sequence that is less than 100% identical to SEQ ID NO: 112 and has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 112, and a VL region having an amino acid sequence set forth in SEQ ID NO: 118.
- the antibody or antigen-binding fragment has a VH region having an amino acid sequence set forth in SEQ ID NO: 112, and a VL region having an amino acid sequence that is less than 100% identical to SEQ ID NO: 118 and has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in set forth in SEQ ID NO: 118.
- the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 1
- the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118
- the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 2
- the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%,
- the VH region of the antibody or antigen-binding fragment thereof comprises the amino acid sequence of any one of SEQ ID NOs: 1-27 and 112
- the VL region of the antibody or antigen-binding fragment comprises the amino acid sequence of any one of SEQ ID NOs: 28-48 and 118.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 1 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 2 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 3 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 4 and 118, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 5 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 6 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 7 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 8 and 118, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 9 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 10 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 11 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 12 and 118, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 13 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 14 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 15 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 16 and 118, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 17 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 18 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 19 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 20 and 118, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 21 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 22 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 28, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 29, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 30, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 31, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 32, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 33, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 34, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 35, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 36, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 37, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 38, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 40, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 41, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 42, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 43, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 44, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 45, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 46, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 23 and 41, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 24 and 41, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 25 and 41, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 26 and 41, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 22 and 41, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 27 and 41, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 23 and 47, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 24 and 47, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 25 and 47, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 26 and 47, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 22 and 47, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 27 and 47, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 23 and 29, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 24 and 29, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 25 and 29, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 26 and 29, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 22 and 29, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 27 and 29, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 23 and 39, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 24 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 25 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 26 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 22 and 39, respectively.
- the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 27 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 5 and 48, respectively.
- Table El provides the SEQ ID NOS: of exemplary provided antibody or antigenbinding fragment thereof.
- the CTLA-4-binding antibody or fragment thereof comprises a VH region that comprises the CDR-H1, the CDR-H2 and the CDR-H3 sequence and a VL region that comprises the CDR-L1, the CDR-L2 and the CDR-L3 sequence set forth in the SEQ ID NOS: listed in each row of Table El below (by Kabat numbering scheme).
- the CTLA-4-binding antibody or fragment thereof comprises a VH region sequence and a VL region sequence set forth in the SEQ ID NOS: listed in each row of Table El below, or an antibody comprising a VH region and a VL region amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region sequence and the VL region sequence set forth in the SEQ ID NOS: listed in each row of Table El below.
- the CTLA-4-binding antibody or fragment thereof comprises a VH region sequence and a VL region sequence set forth in the SEQ ID NOS: listed in each row of Table El below.
- the provided antibody or antigen-binding fragment thereof comprises a VH region and a VL region, wherein the VH region of the antibody or antigen- binding fragment thereof can contain a combination of any of the CDR-H1, the CDR-H2 and the CDR-H3 amino acid sequences set forth in Table El, and the VL region of the antibody or antigen-binding fragment thereof can contain a combination of any of the CDR-L1, the CDR- L2 and the CDR-L3 amino acid sequences set forth in Table El.
- the provided antibody or antigen-binding fragment thereof comprises a VH region and/or a VL region set forth in Table El, in any combination, orientation or containing a different linker.
- the antibody or antigen-binding fragment thereof comprises a VH region described in Table El.
- the antibody or antigen-binding fragment thereof comprises a VL region described in Table El.
- provided antibodies include those that include amino acid residues (such as due to mutation compared to a reference sequence) that alter pH binding such that the antibodies exhibit better binding at neutral pH (e.g., pH 7.4) compared to acidic pH (e.g., pH 5.0).
- neutral pH e.g., pH 7.4
- acidic pH e.g., pH 5.0
- provided antibodies, including antibody fragments, herein contain one or more amino acid mutation in the VH region that is S31H, D54H, D54E, Y59H, W101H, or L102H, which correspond to S31R, D53H, D53E, Y58H, W97H, or L98H based on Kabat numbering, respectively, or any combination thereof, compared to the VH region set forth in SEQ ID NO: 112; and/or one or more amino acid mutation in the VL region that is S28H, S31H, S32H, or Q91H, which correspond to S27aH, S30H, S31H, or Q90H based on Kabat numbering, respectively, or any combination thereof, compared to the VL region set forth in SEQ ID NO: 118.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises one or more mutations selected from the group consisting of: S31R, D54H, D54E, Y59H, W101H, and L102H (S31R, D53H, D53E, Y58H, W97H, and L98H, respectively, based on Kabat numbering) in the VH region, such as compared to a reference anti-CTLA-4 antibody VH, e.g. set forth in SEQ ID NO: 112.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprise a histidine (H) at position 98 based on Kabat numbering (e.g., L102H mutation compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering) (e.g., S31R mutation compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a glutamic acid (E) at position 53 based on Kabat numbering) (e.g., D54E mutation compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 53 based on Kabat numbering (e.g., D54H mutation compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 58 based on Kabat numbering (e.g., Y59H mutation compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 97 based on Kabat numbering (e.g., W101H mutation compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering and a arginine (R) at position 31 based on Kabat numbering (e.g., L102H and S31R mutation compared to SEQ ID NO: 112) in the VH region.
- the provided anti- CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering and a glutamic acid (E) at position 53 based on Kabat numbering (e.g., L102H and D54E mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering and a histidine (H) at position 53 based on Kabat numbering (e.g., L102H and D54H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H and W 101H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering) and a glutamic acid (E) at position 53 based on Kabat numbering (e.g., S31R and D54E mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 53 based on Kabat numbering (e.g., S31R and D54H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., S31R and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., S31R and W 101H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a glutamic acid (E) at position 53 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., D54E and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a glutamic acid (E) at position 53 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., D54E and W 101H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 53 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., D54H and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 53 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., D54H and W 101H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 58 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., Y59H and W 101H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering and a glutamic acid (E) at position 53 based on Kabat numbering (e.g., L102H, S31R, and D54E mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- R arginine
- E glutamic acid
- the provided anti-CTLA-4 antibody or antigenbinding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 53 based on Kabat numbering (e.g., L102H, S31R, and D54H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- R arginine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H, S31R, and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- R arginine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- R arginine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H, D54E, and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- E glutamic acid
- H histidine
- the provided anti- CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, D54E, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- E glutamic acid
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a a histidine (H) at position 53 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H, D54H, and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a histidine (H) at position 53 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, D54H, and W 101H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, Y59H, and W 101H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering), a glutamic acid (E) at position based on Kabat numbering, and a histidine (H) at position 58 based on Kabat numbering (e.g., S31R, D54E, and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- R arginine
- E glutamic acid
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering), a glutamic acid (E) at position 53 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., S31R, D54E, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- R arginine
- E glutamic acid
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering), a histidine (H) at position 53 based on Kabat numbering, and a histidine (H) at position 58 based on Kabat numbering (e.g., S31R, D54H, and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- R arginine
- H histidine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering), a histidine (H) at position 53 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., S31R, D54H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- R arginine
- H histidine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering), a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., S31R, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- R arginine
- H histidine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a glutamic acid (E) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., D54E, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- E glutamic acid
- H histidine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., D54H, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering, and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H, S31R, D54E, and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, D54E, and W 101H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a histidine (H) at position 53 based on Kabat numbering, and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H, S31R, D54H, and Y59H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti- CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a histidine (H) at position 53 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, D54H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, D54E, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- E glutamic acid
- H histidine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a histidine (H) at position 53, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, D54H, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., S31R, D54E, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- R arginine
- E glutamic acid
- H histidine
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, D54E, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a histidine (H) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, D54H, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises one or more mutations selected from the group consisting of: S28H, S31H, S32H, and Q91H (S27aH, S30H, S31H, and Q90H, respectively, based on Kabat numbering) in the VL region, such as compared to a reference anti-CTLA-4 antibody VL, e.g. set forth in SEQ ID NO: 118.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering (e.g., S28H mutation compared to SEQ ID NO: 118) in the VL region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 30 based on Kabat numbering (e.g., S31H mutation compared to SEQ ID NO: 118) in the VL region.
- the provided anti- CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 31 based on Kabat numbering (e.g., S32H mutation compared to SEQ ID NO: 118) in the VL region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 90 based on Kabat numbering (e.g., Q91H mutation compared to SEQ ID NO: 118) in the VL region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering and a histidine (H) at position 30 based on Kabat numbering (e.g., S28H and S31H mutations compared to SEQ ID NO: 118) in the VL region.
- the provided anti- CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position27a based on Kabat numbering and a histidine (H) at position 31 based on Kabat numbering (e.g., S28H and S32H mutations compared to SEQ ID NO: 118) in the VL region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering and a histidine (H) at position 90 based on Kabat numbering (e.g., S28H and Q91H mutations compared to SEQ ID NO: 118) in the VL region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 30 based on Kabat numbering and a histidine (H) at position 31 based on Kabat numbering (e.g., S31H and S32H mutations compared to SEQ ID NO: 118) in the VL region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 30 based on Kabat numbering and a histidine (H) at position 90 based on Kabat numbering (e.g., S31H and Q91H mutations compared to SEQ ID NO: 118) in the VL region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 31 based on Kabat numbering and a histidine (H) at position 90 based on Kabat numbering (e.g., S32H and Q91H mutations compared to SEQ ID NO: 118) in the VL region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering, a histidine (H) at position 30 based on Kabat numbering, and a histidine (H) at position 31 based on Kabat numbering) (e.g., S28H, S31H, S32H mutations compared to SEQ ID NO: 118 (S27aH, S30H, S31H mutations compared to SEQ ID NO: 118, based on Kabat numbering)) in the VL region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 28 (position 27a based on Kabat numbering), a histidine (H) at position 30 based on Kabat numbering, and a histidine (H) at position 90 based on Kabat numbering (e.g., S28H, S31H, Q91H mutations compared to SEQ ID NO: 118) in the VL region.
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering, a histidine (H) at position 31 based on Kabat numbering, and a histidine (H) at position 90 based on Kabat numbering (e.g., S28H, S32H, Q91H mutations compared to SEQ ID NO: 118) in the VL region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 30 based on Kabat numbering, a histidine (H) at position 31 based on Kabat numbering, and a histidine (H) at position 90 based on Kabat numbering (e.g., S31H, S32H, Q91H mutations compared to SEQ ID NO: 118) in the VL region.
- H histidine
- the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering, a histidine (H) at position 30 based on Kabat numbering, a histidine (H) at position 31 based on Kabat numbering, and a histidine (H) at position 91 (position 90 based on Kabat numbering) (e.g., S28H, S31H, S32H, Q91H mutations compared to SEQ ID NO: 118) in the VL region.
- the VH region comprises an arginine (R) at position 31, glutamic acid (E) or histidine (H) at position 53, histidine (H) at position 58, histidine (H) at position 97, or histidine (H) at position 98, respectively, based on Kabat numbering, or any combination thereof (e.g., S31R, D54H, D54E, Y59H, W101H, or L102H mutations, or any combination thereof, compared to SEQ ID NO:112), and the VL region comprises a histidine (H) at position 27a, histidine (H) at position 30, histidine (H) at position 31, or histidine (H) at position 90, respectively, based on Kabat numbering), or any combination thereof (e.g., S28H, S31H, S32H, or Q91H mutations, or any combination thereof, compared to SEQ ID NO: 118).
- R arginine
- E glutamic acid
- H hist
- Exemplary antibody VH and VL regions with one or more of the above residues are set forth in Tables 3A and 3B.
- any antibody or antigen binding fragment comprising at least one of such VH or VL region set forth in Table 3A or Table 3B, paired with any respective other VH and VL described herein.
- antibodies or antigen binding fragments containing a paired combination of any VH set forth in Table 3A and any VL set forth in Table 3B are also provided.
- the antibody or antigen binding fragment comprises a VH set forth in any one of SEQ ID NOs: 5, 6, 14, 17-19, and 21-27, and a VL set forth in any one of SEQ ID NOs: 29, 30, 32, 34, 39-41, 44, and 47.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 29.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 30.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 40.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 29.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 39.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 47.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 34.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 44.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 32.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 41.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 30.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 40.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 29.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 39.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 47.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 34.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 44.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 32.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 41.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 30.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 40.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 29.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 39.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 47.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 34.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 44.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 32.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 41.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 30.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 40.
- the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 47.
- the above examples are non-limiting and any combinations are contemplated and provided by the present disclosure.
- antibodies that contain a VH region sequence that exhibits a sequence identity of at least 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% to any of such VH region sequences in Table 3A, in which one of the pH residues in Table 3A is present.
- antibodies that contain a VL region sequence that exhibits a sequence identity of at least 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% to any of such V L region sequences in Table 3B, in which one of the pH residues in Table 3B is present.
- antibodies that contain a VH region sequence that exhibits a sequence identity of at least 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% to any of such VH region sequences in Table 3A, in which one of the pH residues in Table 3A is present; and a VL region sequence that exhibits a sequence identity of at least 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% to any of such VL region sequences in Table 3B, in which one of the pH residues in Table 3B is present.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89 and 98, respectively.
- the VH region comprises the sequence set forth in SEQ ID NO: 25 and the VL comprises the sequence set forth in SEQ ID NO: 29.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67 and 68, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89 and 99, respectively.
- the VH region comprises the sequence set forth in SEQ ID NO: 23 and the VL comprises the sequence set forth in SEQ ID NO: 39.
- the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, 72, respectively
- the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89 and 99, respectively.
- the VH region comprises the sequence set forth in SEQ ID NO: 25 and the VL comprises the sequence set forth in SEQ ID NO: 39.
- antibody variable domains as described are fused to immunoglobulin constant domain sequences.
- the fusion is with an Ig heavy chain constant domain, comprising at least part of the hinge, CH2, and CH3 regions.
- the antibody further comprises a heavy chain constant domain and/or a light chain constant domain.
- the heavy chain and/or light constant domain is murine or human.
- a “constant region” or “constant domain” is a domain in an antibody heavy or light chain that contains a sequence of amino acids that is comparatively more conserved among antibodies than the variable region domain.
- the constant regions of immunoglobulins show less sequence diversity than the variable regions, and are responsible for binding a number of natural proteins to elicit important biochemical events.
- Each light chain has a single light chain constant region (CL) domain, which is either of the kappa or lambda type.
- Each heavy chain contains one or more heavy chain constant region (CH) domains that can be assigned to different classes or isotypes.
- IgA, IgD, IgE, IgG and IgM having heavy chains designated ex, 8, e, y and p., respectively.
- Full-length IgA, IgD and IgG isotypes contain CHI, CH2, CH3 and a hinge region, while IgE and IgM contain CHI, CH2, CH3 and CH4.
- the y and ex classes are further divided into subclasses on the basis of relatively minor differences in the CH sequence and function, e.g., humans express the following subclasses: IgGl, IgG2, IgG3, IgG4, IgAl and IgA2.
- IgGl antibodies can exist in multiple polymorphic variants termed allotypes (reviewed in Jefferis and Lefranc 2009. mAbs Vol 1 Issue 4 1-7) any of which are suitable. In humans there are five different classes of antibodies including IgA (which includes subclasses IgAl and IgA2), IgD, IgE, IgG (which includes subclasses IgGl, IgG2, IgG3, and IgG4), and IgM. The distinguishing features between these antibody classes are their constant, Fc regions, although subtler differences may exist in the V region. An antibody constant region can include an Fc portion.
- the human IgG heavy-chain Fc region usually includes the region containing the CH2 and CH3 domains and the hinge region, such as an amino acid residue at position Cys226, or from Pro230, to the carboxyl-terminus thereof.
- a provided antibody contains an Ig heavy chain constant domain comprising at least part of the hinge, CH2, and CH3 regions. Certain embodiments have the first heavy-chain constant region (CHI) containing the site necessary for light chain bonding, present in at least one of the fusions. DNAs encoding the immunoglobulin heavy chain fusions and, if desired, the immunoglobulin light chain, are inserted into separate expression vectors, and are co-transfected into a suitable host cell. This provides for greater flexibility in adjusting the mutual proportions of the three polypeptide fragments in embodiments when unequal ratios of the three polypeptide chains used in the construction provide the optimum yield of the desired fusion antibody.
- CHI first heavy-chain constant region
- the antibody or antigen-binding fragment thereof may contain at least a portion of an immunoglobulin constant region, such as one or more constant region domain.
- the constant regions include a light chain constant region and/or a heavy chain constant region 1 (CHI).
- the antibody includes at least a portion of a hinge region or a variant thereof.
- the antibody includes a CH2 and/or CH3 domain, such as an Fc region.
- the Fc region is an Fc region from an IgAl, IgA2, IgD, IgE, IgGl, IgG2, IgG3, IgG4, or IgM.
- the heavy chain includes a variable heavy chain as described that is joined to a human constant region.
- the human constant region includes the CH1-CH2-CH3 constant domains.
- the human constant region is human IgGl or is derived from human IgGl, or is or is derived from a naturally occurring variant thereof.
- the naturally occurring variant is an IgG allotype for instance with the K97R, D239E, or L241M mutation.
- the heavy chain constant region is mutated or modified, i.e., is a modified constant region.
- the heavy chain constant region is a modified human IgG heavy chain constant region.
- the mutations include one or more amino acid substitutions in the heavy chain constant region, such as the heavy chain constant region of IgGl, to increase effector activity of the heavy chain constant region.
- increased effector activity is an enhanced antibody dependent cell cytotoxicity (ADCC), such as mediated by FcyR-expressing cells.
- ADCC antibody dependent cell cytotoxicity
- the one or more amino acid substitutions results in binding to FcyRI.
- FcyRIIa or FcyRIIIa that is enhanced compared to an antibody in which the heavy chain constant region does not contain the one or more amino acid substitutions.
- the enhanced binding to the Fcyl, Fcylla and/or Fey Illa receptor results in increased ADCC.
- an activity of the provided antibodies is to mediate depletion of CTLA- 4-expressing cells, such as Tregs, via ADCC, enhanced binding to the FcyR (e.g, FcyRI. FcyRIIa or FcyRIIIa) mediates a reduction of Tregs at the tumor site.
- FcyR e.g, FcyRI. FcyRIIa or FcyRIIIa
- I l l substitutions in the heavy chain constant region to enhance FcR binding and/or ADCC can include any as described in U.S. Patent No. US7317091 or U.S. Patent No. 10196445, or any combination of any such described amino acid substitutions.
- the heavy chain constant region contains at least one amino acid substitution selected from the group consisting of P230A, E233D, L234E, L234Y, L234I, L235D, L235S, L235Y, L235I, S239D, S239E, S239N, S239Q, S239T, V240I, V240M, F243L, V264I, V264T, V264Y, V266I, E272Y, K274T, K274E, K274R, K274L, K274Y, F275W, N276L, Y278T, V302I, E318R, S324D, S324I, S324V, N325T, K326I, K326T, L328M, L328I, L328Q, L328D, L328V, L328T, A330Y, A330L, A330I, I332D, I332E, I3
- the heavy chain constant region contains at amino acid substitutions V264I, F243E/V264I, E328M, I332E, E328M/I332E, V264VI332E, S298A/I332E, S239E/I332E, S239Q/I332E, S239E, A330Y, I332D, E328VI332E, E328Q/I332E, V264T, V240I, V266I, S239D, S239D/I332D, S239D/I332E, S239D/I332N, S239D/I332Q, S239E/I332D, S239E/I332N, S239E/I332Q, S239N/I332D, S239N/I332E, S239Q/I332D, A330Y/I332E, V264VA330Y/I332E, V264VA330Y/I3
- the mutation in the IgG (e.g., IgGl) heavy chain to enhance binding to FcyR-expressing cells is selected from one or more mutations G236A, S239D, S298A, A330E, I332E, E333A and K334A, wherein the numbering of the residues in the Fc region is that of the EU index as in Kabat.
- the mutation in the IgG heavy chain to enhance binding to FcyR-expressing cells is selected from mutations S239D and I332E.
- the mutation in the IgG heavy chain to enhance binding to FcyR- expressing cells is selected from mutations S239D, A330L and I332E.
- the mutation in the IgG heavy chain to enhance binding to FcyR-expressing cells is selected from mutations G236A, A330L and I332E. In some embodiments, the mutation in the IgG heavy chain to enhance binding to FcyR-expressing cells is selected from mutations G236A, S239D, A330E and I332E. In some embodiments, the mutation in the IgG heavy chain to enhance binding to FcyR-expressing cells is selected from mutations S298A, E333A and K334A.
- the mutation in the IgG heavy chain increase the affinity of IgG for the neonatal Fc receptor, FcRn, which prolongs the in vivo half-life of antibodies and results in increased anti-tumor activity.
- the mutations in the IgG is M428E and/or N434S.
- the mutations include one or more amino acid substitutions to reduce effector activity of the heavy chain constant region.
- the heavy chain constant region is modified to reduce effector activity of the antibody.
- the modified human IgG heavy chain constant region has reduced effector activity.
- effector activity is reduced compared to a wild-type human IgG heavy chain constant region.
- the modified human IgG heavy chain constant region is modified compared to the constant region of wildtype human IgGl.
- the modified human IgG heavy chain constant region has a sequence that is modified by one or more amino acid substitutions compared to SEQ ID NO: 196 (or a naturally occurring variant thereof, e.g.
- Various examples of mutations to heavy chain constant regions to alter, such as reduce, effector function are known, including any as described below.
- reference to amino acid substitutions in a heavy chain constant region is by EU numbering by Kabat (also called Kabat numbering) unless described with reference to a specific SEQ ID NO.
- EU numbering is known and is according to the most recently updated IMGT Scientific Chart (IMGT®, the international ImMunoGeneTics information system®, http://www.imgt.org/IMGTScientificChart/Numbering/Hu_IGHGnber.html (created: 17 May 2001, last updated: 10 Jan 2013) and the EU index as reported in Kabat, E.A. et al. Sequences of Proteins of Immunological interest. 5th ed. US Department of Health and Human Services, NIH publication No. 91-3242 (1991).
- the heavy chain constant region is modified to alter antibody-dependent cellular cytotoxicity (ADCC) and/or complement-dependent cytotoxicity (CDC), e.g., the amino acid modifications described in Natsume et al., 2008 Cancer Res, 68(10): 3863-72; Idusogie et al., 2001 J Immunol, 166(4): 2571-5; Moore et al., 2010 mAbs, 2(2): 181-189; Lazar et al., 2006 PNAS, 103(11): 4005-4010, Shields et al., 2001 JBC, 276(9): 6591-6604; Stavenhagen et al., 2007 Cancer Res, 67(18): 8882-8890; Stavenhagen et al., 2008 Advan. Enzyme Regul., 48: 152-164; Alegre et al, 1992 J Immunol, 148: 3461-3468; Reviewed in Kaneko and Niwa
- the heavy chain constant region is altered at one or more of the following positions to reduce Fc receptor binding: Leu 234 (L234), Leu235 (L235), Asp265 (D265), Asp270 (D270), Ser298 (S298), Asn297 (N297), Asn325 (N325), Ala327 (A327) or Pro329 (P329).
- Leu 234Ala (L234A), Leu235Ala (L235A), Leu235Glu (L235E), Asp265Asn (D265N), Asp265Ala (D265A), Asp270Asn (D270N), Ser298Asn (S298N), Asn297Ala (N297A), Pro329Ala (P329A) or Pro239Gly (P329G), Asn325Glu (N325E) orAla327Ser (A327S).
- modifications within the heavy chain constant region reduce binding to Fc-receptor-gamma receptors while have minimal impact on binding to the neonatal Fc receptor (FcRn).
- the heavy chain constant region is modified at amino acid Asn297 (Kabat Numbering) to prevent glycosylation of the chimeric antibody, e.g., Asn297Ala (N297A) or Asn297Asp (N297D).
- the heavy chain constant region is modified at amino acid Leu235 (Kabat Numbering) to alter Fc receptor interactions, e.g., Leu235Glu (L235E) or Leu235Ala (L235A).
- the heavy chain constant region of the chimeric antibody is modified at amino acid Leu234 (Kabat Numbering) to alter Fc receptor interactions, e.g., Leu234Ala (L234A).
- the heavy chain constant region of the chimeric antibody is modified at amino acid Leu234 (Kabat Numbering) to alter Fc receptor interactions, e.g., Leu235Glu (L235E).
- the heavy chain constant region of the chimeric antibody is altered at both amino acids 234 and 235, e.g., Leu234Ala and Leu235Ala (L234A/L235A) or Leu234Val and Leu235Ala (L234V/L235A).
- the modified heavy chain constant region comprises Leu234Ala and Leu235Ala (L234A/L235A) mutations.
- the heavy chain constant region of the chimeric antibody is altered at amino acids 234, 235, and 297, e.g., Leu234Ala, Leu235Ala, Asn297Ala (L234A/L235A/N297A). In some embodiments, the heavy chain constant region of the chimeric antibody is altered at amino acids at 234, 235, and 329, e.g., Leu234Ala, Leu235Ala, Pro239Ala (L234A/L235A/P329A). In some embodiments, the heavy chain constant region of the chimeric antibody is modified at amino acid Asp265 (Kabat Numbering) to alter Fc receptor interactions, e.g.
- the heavy chain constant region of the chimeric antibody is modified at amino acid Pro329 (Kabat Numbering) to alter Fc receptor interactions, e.g. Pro329Ala (P329A) or Pro329Gly (P329G).
- the heavy chain constant region of the chimeric antibody is altered at both amino acids 265 and 329, e.g., Asp265Ala and Pro329Ala (D265A/P329A) or Asp265Ala and Pro329Gly (D265A/P329G).
- the heavy chain constant region of the chimeric antibody is altered at amino acids at 234, 235, and 265, e.g., Leu234Ala, Leu235Ala, Asp265Ala (L234A/L235A/D265A). In some embodiments, the heavy chain constant region of the chimeric antibody is altered at amino acids at 234, 235, and 329, e.g., Leu234Ala, Leu235Ala, Pro329Gly (L234A/L235A/P329G).
- the heavy chain constant region of the chimeric antibody is altered at amino acids at 234, 235, 265 and 329, e.g., Leu234Ala, Leu235Ala, Asp265Ala, Pro329Gly
- the heavy chain constant region of the chimeric antibody is altered at Gly235 to reduce Fc receptor binding. For example, wherein Gly235 is deleted from the heavy chain constant region of the chimeric antibody.
- the heavy chain constant region of the chimeric antibody is modified at amino acid Gly236 to enhance the interaction with CD32A, e.g., Gly236Ala (G236A).
- the heavy chain constant region of the chimeric antibody lacks Lys447 (EU index of Kabat et al 1991 Sequences of Proteins of Immunological Interest).
- the heavy chain constant region of the chimeric antibody is lacking an amino acid at one or more of the following positions to reduce Fc receptor binding: Glu233 (E233), Leu234 (L234), or Leu235 (L235).
- the heavy chain constant region of the chimeric antibody is lacking an amino acid at one or more of the following positions Glu233 (E233), Leu234 (L234), or Leu235 (L235), and is modified at one or more of Asp265 (D265), Asn297 (N297), or Pro329 (P329), to reduce Fc receptor binding.
- the heavy chain constant region of the chimeric antibody comprises a three amino acid deletion in the lower hinge corresponding to IgGl E233, L234, and L235.
- such heavy chain constant regions s do not engage FcyRs and thus are referred to as “effector silent” or “effector null.”
- the modified heavy chain constant region includes Leu234Ala and Leu235Ala (L234A/L235A) mutations.
- the heavy chain constant sequence includes an IgG4 containing the S228P mutation, which has been shown to prevent recombination between a therapeutic antibody and an endogenous IgG4 by Fab-arm exchange (see e.g. Fabrijin et al. (2009) Nat. Biotechnol., 27(8): 767-71.).
- the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 102.
- the heavy chain constant domain is derived from an IgGl.
- the heavy chain constant domain comprises one or more of the mutations selected from the group consisting of G236A, S239D, M252Y, S254T, T256E, A330E, I332E, M428E, and N434S.
- the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 101.
- the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 106. In some embodiments, the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 107.
- the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 108. In some embodiments, the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 109.
- the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 110. In some embodiments, the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 111. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 101.
- the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 106. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 107. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 108. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 109. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 110. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 111.
- the light chain constant domain comprises the sequence set forth in SEQ ID NO: 103.
- the heavy chain further comprises a signal peptide.
- the signal peptide can be cleaved upon expression from a cell.
- the heavy chain signal peptide is set forth in SEQ ID NO: 104.
- the light chain signal peptide is set forth in SEQ ID NO: 105.
- the light chain constant region is a kappa light chain.
- the kappa light chain comprises the sequence set forth in SEQ ID NO: 103.
- the light chain further comprises a signal peptide.
- the signal peptide can be cleaved upon expression from a cell.
- the signal peptide is set forth in SEQ ID NO: 105.
- the antibody or antigen-binding fragment thereof is a singlechain antibody fragment, such as a single chain variable fragment (scFv).
- the antibody or antigen binding fragment is a multi-domain antibody, such as an scFv comprising a heavy chain variable (VH) region and a light chain variable (VL) region.
- the single-chain antibody fragment e.g., scFv
- the linker typically is a peptide linker, e.g., a flexible and/or soluble peptide linker.
- the linkers are those rich in glycine and serine and/or in some cases threonine.
- the linkers further include charged residues such as lysine and/or glutamate, which can improve solubility.
- the linkers further include one or more proline.
- the linkers rich in glycine and serine (and/or threonine) include at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% such amino acid(s). In some embodiments, they include at least at or about 50%, 55%, 60%, 70%, or 75%, glycine, serine, and/or threonine. In some embodiments, the linker is comprised substantially entirely of glycine, serine, and/or threonine.
- the linkers generally are between about 5 and about 50 amino acids in length, typically between at or about 10 and at or about 30, e.g., 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30, and in some examples between 10 and 25 amino acids in length.
- Exemplary linkers include linkers having various numbers of repeats of the sequence GGGGS (4GS; SEQ ID NO: 124) or GGGS (3GS; SEQ ID NO: 125), such as between 2, 3, 4 and 5 repeats of such a sequence.
- Exemplary linkers include those having or consisting of an sequence set forth in SEQ ID NO: 126 (GGGGSGGGGSGGGGS).
- Exemplary linkers further include those having or consisting of the sequence set forth in SEQ ID NO: 127 (GSTSGSGKPGSGEGSTKG). Exemplary linkers further include those having or consisting of the sequence set forth in SEQ ID NO: 128 (SRGGGGSGGGGSGGGGSLEMA). An exemplary linker includes those having or consisting of the sequence set forth in SEQ ID NO: 129 (GSRGGGGSGGGGSGGGGSLEMA).
- the provided antibodies have one or more specified functional features, such as binding properties, including binding to particular epitopes or exhibiting lower or reduced binding to a related but non-specific antigen.
- the provided antibodies can bind to an epitope that is similar to or overlaps with epitopes of other antibodies, such as reference antibodies, and/or exhibit particular binding affinities.
- the provided antibodies can bind to an epitope that is different from epitopes of other antibodies, e.g., binding a conformational epitope.
- the provided antibodies or antigen-binding fragment thereof specifically bind to a cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) protein.
- CTLA-4 refers to human CTLA-4.
- the provided antibodies and antibody fragments specifically bind to human CTLA-4.
- a molecule is said to exhibit "specific binding" or “preferential binding” if it reacts or associates more frequently, more rapidly, with greater duration and/or with greater affinity with a particular cell or substance (e.g. a cell expressing a CTLA-4 antigen or a recombinant CTLA-4 antigen) than it does with alternative cells or substances.
- An antibody specifically binds or preferentially binds to a target (i.e. CTLA-4) if it binds with greater affinity, avidity, more readily, and/or with greater duration than it binds to other substances.
- a target i.e. CTLA-4
- an antibody that specifically or preferentially binds to CTLA-4 is an antibody that binds CTLA-4 with greater affinity, avidity, more readily, and/or with greater duration than it binds to a different protein antigen. It is also understood by reading this definition that specific binding or preferential binding does not necessarily require (although it can include) exclusive binding. Methods to determine such specific or preferential binding are also well known in the art, e.g., an immunoassay.
- an antibody or other binding molecule binds to CTLA-4 or specifically binds to CTLA-4 does not necessarily mean that it binds to CTLA-4 from every species.
- features of binding to CTLA-4 such as the ability to specifically bind thereto and/or to compete for binding thereto with a reference antibody, and/or to bind with a particular affinity or compete to a particular degree, in some embodiments, refers to the ability with respect to a human CTLA-4 protein and the antibody may not have this feature with respect to a CTLA-4 of another species such as mouse.
- the provided antibodies are capable of binding CTLA-4, such as human CTLA-4, with at least a certain affinity, as measured by any of a number of known methods.
- the affinity is represented by absorbance.
- the affinity is represented by equilibrium dissociation constant (KD).
- ELISA enzyme linked immunosorbent assays
- ELISA enzyme linked immunosorbent assays
- the antigen target macromolecule
- a solid surface microplate
- Detection is accomplished by measuring the activity of the reporter enzyme via incubation with the appropriate substrate to produce a measurable product.
- a variety of assays are known for assessing binding affinity, equilibrium dissociation constant (KD), equilibrium association constant (KA), ECSO, on-rate (association rate constant; k on or k a ; units of 1/Ms or M ⁇ s' 1 ) and the off-rate (dissociation rate constant; k o ff or ka; units of 1/s or s' 1 ) and/or determining whether a binding molecule (e.g., an antibody or fragment thereof) specifically binds to a particular ligand (e.g., an antigen, such as a CTLA-4 protein).
- KD equilibrium dissociation constant
- KA equilibrium association constant
- ECSO on-rate (association rate constant; k on or k a ; units of 1/Ms or M ⁇ s' 1 ) and the off-rate (dissociation rate constant; k o ff or ka; units of 1/s or s' 1 ) and/or determining whether a binding molecule
- a BIAcore® instrument can be used to determine the binding kinetics and constants of a complex between two proteins (e.g., an antibody or fragment thereof, and an antigen, such as a CTLA-4 protein), using surface plasmon resonance (SPR) analysis (see, e.g., Scatchard et al., Ann. N.Y. Acad. Sci. 51:660, 1949; Wilson, Science 295:2103, 2002; Wolff et al., Cancer Res. 53:2560, 1993; and U.S. Patent Nos. 5,283,173, 5,468,614, or the equivalent).
- SPR surface plasmon resonance
- SPR measures changes in the concentration of molecules at a sensor surface as molecules bind to or dissociate from the surface.
- the change in the SPR signal is directly proportional to the change in mass concentration close to the surface, thereby allowing measurement of binding kinetics between two molecules.
- the dissociation rate constant (k o ff or kd), the association rate constant (k on or k a ) and/or equilibrium dissociation constant (KD) and/or equilibrium association constant (KA) for the complex can be determined by monitoring changes in the refractive index with respect to time as buffer is passed over the chip.
- suitable assays for measuring the binding of one protein to another include, for example, immunoassays such as enzyme linked immunosorbent assays (ELISA) and radioimmunoassays (RIA), or determination of binding by monitoring the change in the spectroscopic or optical properties of the proteins through fluorescence, UV absorption, circular dichroism, or nuclear magnetic resonance (NMR).
- immunoassays such as enzyme linked immunosorbent assays (ELISA) and radioimmunoassays (RIA), or determination of binding by monitoring the change in the spectroscopic or optical properties of the proteins through fluorescence, UV absorption, circular dichroism, or nuclear magnetic resonance (NMR).
- exemplary assays include, but are not limited to, Western blot, analytical ultracentrifugation, spectroscopy, flow cytometry, sequencing, genetic reporter assays, flow cytometry, and other methods for detection of expressed nucleic acids or binding of proteins.
- binding can be determined under conditions that mimic those that exist in natural environments in the body of a subject. For instance, acidic pH such as pH about 5.0 mimics more closely the endosomes in a cell. In contrast, physiological pH is a more neutral pH 7.35 to 7.45, such as pH about 7.4 and more closely mimics the environment of normal tissues or cells in the blood.
- acidic pH such as pH about 5.0 mimics more closely the endosomes in a cell.
- physiological pH is a more neutral pH 7.35 to 7.45, such as pH about 7.4 and more closely mimics the environment of normal tissues or cells in the blood.
- the antibody or fragment exhibits a similar CTLA-4 binding at an acidic pH as present in the microenvironment of tumor (such as at pH 5.0 to 6.8, e.g. pH 6.0 or pH 5.0) compared to CTLA-4 binding at normal pH as present in blood or a normal tissue (such as at pH 7.35 to 7.45, e.g. pH 7.4).
- the KD at an acidic pH as present in the microenvironment of tumor (such as at pH 5.0 to 6.8, e.g. pH 6.0 or pH 5.0) is about the same (e.g. 0.7-fold to 1.3-fold) the KD at pH of a non-tumor microenvironment (such as at pH 7.35 to 7.45, e.g. 7.4) as present at normal pH of blood or a normal tissue.
- the provided antibodies are described in relation to properties observed for another antibody, e.g., a reference antibody.
- the reference antibody is a human or humanized anti-CTLA-4 antibody.
- Exemplary reference anti-CTLA-4 antibodies include ipilimumab (Vn and VL set forth in SEQ ID Nos: 112 and 118, respectively), tremelimumab (Vn and VL set forth in SEQ ID NOs: 113 and 119, respectively), botensilimab (Vn and VL set forth in SEQ ID NOs: 114 and 120, respectively), gotistobart (Vn and VL set forth in SEQ ID NOs: 115 and 121, respectively), quavonlimab (Vn and VL set forth in SEQ ID NOs: 116 and 122, respectively), ipi.101 (Vn and VL set forth in SEQ ID NOs: 117 and 123, respectively).
- the provided antibody contains heavy
- a provided antibody or fragment exhibits a lower or similar binding affinity (e.g. lower absorbance) than the reference antibody to CTLA-4 at an acidic pH as present in the endosomes of a cell (such as at pH 5.0).
- a provided antibody or fragment exhibits a higher CTLA-4 binding compared to a reference antibody to CTLA-4 at a neutral pH (such as at pH 7.4) as present at normal pH of blood or a normal tissue.
- the CTLA-4 binding of a provided antibody or fragment at a neutral pH as present in the blood or a normal tissue is higher by at least or 1.2, 1.3, 1.4, 1.5, 1.6, 1.7, 1.8, 1.9, 2.0, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 30, 40, 50 or more compared to the reference antibody for CTLA-4 binding at the same pH or condition.
- the CTLA-4 binding is higher by at least 5%, 10%, 15%, 20%, 25%, 35%, 50%, 100%, 2-fold, 5-fold, 10- fold, 20-fold or more.
- the absorbance of a provided antibody for CTLA- 4 binding in an ELISA or other binding assay at a neutral pH as present in the blood or a normal tissue (such as at pH 7.4) is about 1.5-fold, 2-fold, 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold, 10-fold, 20-fold, 25-fold, 30-fold, 40-fold, 50-fold or 100-fold higher than the absorbance of the reference antibody for CTLA-4 binding at the same pH or condition.
- the EC50 of a provided antibody for CTLA-4 binding at an acidic pH as present within a tumor microenvironment or endosome of a cell is about 1.5-fold, 2-fold, 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold, 10-fold, 20-fold, 25-fold, 30-fold, 40-fold, 50-fold or 100-fold higher than the EC50 of the reference antibody for CTLA-4 binding at the same pH or condition.
- the KD of provided antibody for CTLA-4 binding at an acidic pH as present within a tumor microenvironment or endosome of a cell is about 1.5-fold, 2-fold, 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold, 10-fold, 20-fold, 25-fold, 30-fold, 40-fold, 50-fold or 100-fold higher than the K D of the reference antibody for CTLA-4 binding at the same pH or condition.
- the provided antibody or antigen-binding fragment thereof can impact safety and efficacy through involvement with the broader immune system.
- the provided antibody or antigen-binding fragment thereof induces a favorable ratio of immune cell subsets in the tumor or blood, such as T cells, CD4 + T cells, CD8 + T cells, and/or Treg cells.
- the provided antibody or antigen-binding fragment induces a favorable ratio of CTLA-4 expression on immune cell subsets in the tumor or blood, such as T cells, CD4 + T cells, CD8 + T cells, and/or Treg cells.
- amino acid sequence modification(s) of the antibodies described herein are contemplated. Lor example, it may be desirable to improve the binding affinity and/or other biological properties of the antibody.
- amino acid sequence variants of an antibody may be prepared by introducing appropriate nucleotide changes into a polynucleotide that encodes the antibody, or a chain thereof, or by peptide synthesis. Such modifications include, for example, deletions from, and/or insertions into and/or substitutions of, residues within the amino acid sequences of the antibody. Any combination of deletion, insertion, and substitution may be made to arrive at the final antibody, provided that the final construct possesses the desired characteristics (e.g., high affinity binding to CTLA-4).
- the amino acid changes may alter post-translational processes of the antibody, such as changing the number or position of glycosylation sites. Any of the variations and modifications described above for polypeptides of the present invention may be included in antibodies of the present invention.
- the antibodies include one or more amino acid variations, e.g., substitutions, deletions, insertions, and/or mutations, compared to the sequence of an antibody described herein.
- Exemplary variants include those designed to improve the binding affinity and/or other biological properties of the antibody.
- Amino acid sequence variants of an antibody may be prepared by introducing appropriate modifications into the nucleotide sequence encoding the antibody, or by peptide synthesis. Such modifications include, for example, deletions from, and/or insertions into and/or substitutions of residues within the amino acid sequences of the antibody. Any combination of deletion, insertion, and substitution can be made to arrive at the final construct, provided that the final construct possesses the desired characteristics, e.g., antigen-binding.
- the antibodies include one or more amino acid substitutions, e.g., as compared to an antibody sequence described herein and/or compared to a sequence of a natural repertoire, e.g., human repertoire.
- Sites of interest for substitutional mutagenesis include the CDRs and FRs.
- Amino acid substitutions may be introduced into an antibody of interest and the products screened for a desired activity, e.g., retained/improved antigen binding, decreased immunogenicity, improved half-life, and/or improved effector function, such as the ability to promote antibody-dependent cellular cytotoxicity (ADCC) or complement-dependent cytotoxicity (CDC).
- ADCC antibody-dependent cellular cytotoxicity
- CDC complement-dependent cytotoxicity
- one or more residues within a CDR of a parent antibody is/are substituted.
- the substitution is made to revert a sequence or position in the sequence to a germline sequence, such as an antibody sequence found in the germline (e.g., human germline), for example, to reduce the likelihood of immunogenicity, e.g., upon administration to a human subject.
- alterations are made in CDR “hotspots,” residues encoded by codons that undergo mutation at high frequency during the somatic maturation process (see, e.g., Chowdhury, Methods Mol. Biol. 207:179-196 (2008)), and/or residues that contact antigen, with the resulting variant VH or VL being tested for binding affinity.
- Affinity maturation by constructing and reselecting from secondary libraries has been described, e.g., in Hoogenboom et al. in Methods in Molecular Biology 178:1-37 (O’Brien et al., ed., Human Press, Totowa, NJ, (2001)).
- affinity maturation diversity is introduced into the variable genes chosen for maturation by any of a variety of methods (e.g., error-prone PCR, chain shuffling, or oligonucleotide-directed mutagenesis).
- a secondary library is then created. The library is then screened to identify any antibody variants with the desired affinity.
- Another method to introduce diversity involves CDR-directed approaches, in which several CDR residues (e.g., 4-6 residues at a time) are randomized.
- CDR residues involved in antigen binding may be specifically identified, e.g., using alanine scanning mutagenesis or modeling.
- CDR-H3 and CDR-L3 in particular are often targeted.
- substitutions, insertions, or deletions may occur within one or more CDRs so long as such alterations do not substantially reduce the ability of the antibody to bind antigen.
- conservative alterations e.g., conservative substitutions as provided herein
- Such alterations may, for example, be outside of antigen contacting residues in the CDRs.
- each CDR either is unaltered, or contains no more than one, two or three amino acid substitutions.
- Amino acid sequence insertions include amino- and/or carboxyl-terminal fusions ranging in length from one residue to polypeptides containing a hundred or more residues, as well as intrasequence insertions of single or multiple amino acid residues.
- terminal insertions include an antibody with an N-terminal methionyl residue.
- Other insertional variants of the antibody molecule include the fusion to the N- or C-terminus of the antibody to an enzyme or a polypeptide which increases the serum half-life of the antibody.
- the antibody is altered to increase or decrease the extent to which the antibody is glycosylated, for example, by removing or inserting one or more glycosylation sites by altering the amino acid sequence and/or by modifying the oligosaccharide(s) attached to the glycosylation sites, e.g., using certain cell lines.
- an N-linked glycosylation which is a glycosylation site that occurs at asparagines in the consensus sequence -Asn-Xaa-Ser/Thr is removed or inserted. In some embodiments, one or more re replaced with another amino acid to remove the glycosylation site.
- modified antibodies are those having one or more amino acid modifications in the Fc region, such as those having a human Fc region sequence or other portion of a constant region (e.g., a human IgGl, IgG2, IgG3 or IgG4 Fc region) comprising an amino acid modification (e.g. a substitution) at one or more amino acid positions.
- a human Fc region sequence or other portion of a constant region e.g., a human IgGl, IgG2, IgG3 or IgG4 Fc region
- an amino acid modification e.g. a substitution
- Such modifications can be made, e.g., to improve half-life, alter binding to one or more types of Fc receptors, and/or alter effector functions.
- cysteine engineered antibodies such as “thioMAbs” and other cysteine engineered variants, in which one or more residues of an antibody are substituted with cysteine residues, in order to generate reactive thiol groups at accessible sites, e.g., for use in conjugation of agents and linker-agents, to produce immunoconjugates.
- Cysteine engineered antibodies are described, e.g., in U.S. Patent Nos. 7,855,275 and 7,521,541.
- the antibodies are modified to contain additional nonproteinaceous moieties, including water soluble polymers.
- exemplary polymers include, but are not limited to, polyethylene glycol (PEG), copolymers of ethylene glycol/propylene glycol, carboxymethylcellulose, dextran, polyvinyl alcohol, polyvinyl pyrrolidone, poly-1, 3- dioxolane, poly-l,3,6-trioxane, ethylene/maleic anhydride copolymer, polyaminoacids (either homopolymers or random copolymers), and dextran or poly(n-vinyl pyrrolidone)poly ethylene glycol, propropylene glycol homopolymers, prolypropylene oxide/ethylene oxide copolymers, polyoxyethylated polyols (e.g., glycerol), polyvinyl alcohol, and mixtures thereof.
- PEG polyethylene glycol
- Polyethylene glycol propionaldehyde may have advantages in manufacturing due to its stability in water.
- the polymer may be of any molecular weight, and may be branched or unbranched.
- the number of polymers attached to the antibody may vary, and if more than one polymer is attached, they can be the same or different molecules. In general, the number and/or type of polymers used for derivatization can be determined based on considerations including, but not limited to, the particular properties or functions of the antibody to be improved, whether the antibody derivative will be used in a therapy under defined conditions, etc.
- Antibodies, and antigen-binding fragments and variants thereof, of the present invention may also be modified to include a detectable label, e.g., an epitope tag or label, e.g., for use in purification or diagnostic applications. These may be conjugated to the antibody as a fusion protein or conjugate, e.g., using a linker or linking group.
- a linker or linking group There are many linking groups known in the art for making antibody conjugates, including, for example, those disclosed in U.S. Pat. No. 5,208,020 or EP Patent 0 425 235 Bl, and Chari et al., Cancer Research 52: 127-131 (1992).
- Linking groups include disulfide groups, thioester groups, acid labile groups, photolabile groups, peptidase labile groups, or esterase labile groups, as disclosed in the above-identified patents.
- tags and/or labels can include, but are not limited to, FLAG tags, poly-histidine tags (e.g. 6xHis), cMyc tags, glutathione- S- transferase tags, avidin, fluorescent labels, polymer particles, metal particles, haptens, enzyme labels, luminescent labels, electrochemiluminescent labels, bioluminescent labels, radioisotopes, or oligonucleotides.
- binding molecules that comprise any of the provided anti-CTLA-4 antibodies or antigen binding fragments.
- the binding molecules can include conjugates with another moiety, such as a chemical or other protein molecule.
- the other moiety is a protein molecule and the conjugate is a fusion protein.
- binding molecules that are multispecific antibodies in which at least one antibody binding arm is a provided anti-CTLA-4 antibody or antigen binding fragment.
- conjugates containing an anti-CTLA-4 antibody or antigenbinding fragment provided herein and one or more further moiety can be an effector moiety, which provides at least one additional function or property of the conjugate as compared to the unconjugated antibody.
- an effector moiety can be any substance having biological or detectable activity, for example, therapeutic agents, detectable labels, binding agents, or prodrugs, which are metabolized to an active agent in vivo.
- the moiety can be a targeting moiety, a small molecule drug (nonpolypeptide drug of less than 500 Daltons molar mass), a toxin, a cytostatic agent, a cytotoxic agent, an immunosuppressive agent, a radioactive agent suitable for diagnostic purposes, a radioactive metal ion for therapeutic purposes, a prodrug-activating enzyme, an agent that increases biological half-life, or a diagnostic or detectable agent.
- the effector moiety is a protein, peptide or a nucleic acid molecule, which can be synthesized or produced by recombinant means.
- the effector moiety is a drug moiety, which may be synthesized artificially or purified from a natural source.
- the drug moiety has an intracellular activity.
- the anti-CTLA-4 antibody or antigen-binding fragment of the conjugate is internalized and the drug moiety is a cytotoxin that blocks the protein synthesis of the cell, therein leading to cell death.
- the conjugate can be used for inhibiting the multiplication of a tumor cell or cancer cell, causing apoptosis in a tumor or cancer cell, or for treating cancer in a patient.
- the conjugate can be used accordingly in a variety of settings for the treatment of animal cancers.
- the conjugate can be used to deliver a drug to a tumor cell or cancer cell.
- the conjugate and/or drug upon binding to CTLA-4 on a tumor cell, the conjugate and/or drug can be taken up inside a tumor cell or cancer cell through receptor-mediated endocytosis.
- the conjugate is an antibody drug conjugate (ADC, also called immunoconjugates) containing an anti-CTLA-4 antibody or antigen-binding fragment provided herein conjugated to a drug moiety that acts as a therapeutic agent, which is either cytotoxic, cytostatic or otherwise provides some therapeutic benefit.
- the cytotoxic agent is a chemotherapeutic agent, a drug, a growth inhibitory agent, a toxin (e.g., an enzymatically active toxin of bacterial, fungal, plant, or animal origin, or fragments thereof), or a radioactive isotope (i.e., a radioconjugate).
- provided antibody drug conjugates of the present disclosure allow targeted-delivery of the drug moiety to tumors. In some cases, this can result in targeted killing of the tumor cell.
- the drug moiety is an auristatin, such as auristatin E (also known in the art as a derivative of dolastatin-10) or a derivative thereof.
- the auristatin can be, for example, an ester formed between auristatin E and a keto acid.
- auristatin E can be reacted with paraacetyl benzoic acid or benzoylvaleric acid to produce AEB and AEVB, respectively.
- Other typical auristatins include AFP, MMAF, and MMAE.
- the synthesis and structure of exemplary auristatins are described in U.S. Pat. Nos. 6,884,869, 7,098,308, 7,256,257, 7,423,116, 7,498,298 and 7,745,394, each of which is incorporated by reference herein in its entirety and for all purposes.
- Auristatins have been shown to interfere with microtubule dynamics and nuclear and cellular division and have anticancer activity.
- Auristatins of the present invention bind tubulin and can exert a cytotoxic or cytostatic effect on a 5T4 expressing cell or cell line.
- Assays known in the art, that can be used for determining whether an auristatin or resultant antibody-drug conjugate exerts a cytostatic or cytotoxic effect on a desired cell or cell line.
- Methods for determining whether a compound binds tubulin are known in the art. See, for example, Muller et al., Anal. Chem 2006, 78, 4390-4397; Hamel et al., Molecular Pharmacology, 1995 47: 965-976; and Hamel et al., The Journal of Biological Chemistry, 1990 265:28, 17141-17149.
- Examples of drugs or pay loads are selected from the group consisting of DM1 (maytansine, N2'-deacetyl-N2'-(3-mercapto-l -oxopropyl)- or N2'-deacetyl-N2'-(3-mercapto- l-oxopropyl)-maytansine), mc-MMAD (6-maleimidocaproyl-monomethylauristatin-D or N- methyl-L-valyl-N-[(lS,2R)-2-methoxy-4-[(2S)-2-[(lR,2R)-l-methoxy-2-methyl-3-oxo-3- [[(lS)-2-phenyl-l-(2-thiazolyl)ethyl]amino]propyl]-l-pyrrolidinyl]-l-[(lS)-l-methylpropyl]- 4-oxobutyl]-N-methyl
- DM1 is a derivative of the tubulin inhibitor maytansine while MMAD, MMAE, and MMAF are auristatin derivatives.
- the preferred payloads of the present invention are selected from the group consisting of mc- MMAF and mc-Val-Cit-PABA-MMAE.
- the drug moiety is an anti-cancer agent.
- anticancer agents include, but are not limited to, cytostatics, enzyme inhibitors, gene regulators, cytotoxic nucleosides, tubulin binding agents or tubulin inhibitors, proteasome inhibitors, hormones and hormone antagonists, anti-angiogenesis agents, and the like.
- cytostatic anti-cancer agents include alkylating agents such as the anthracycline family of drugs (e.g.
- adriamycin carminomycin, cyclosporin- A, chloroquine, methopterin, mithramycin, porfiromycin, streptonigrin, porfiromycin, anthracenediones, and aziridines).
- cytostatic anti-cancer agents include DNA synthesis inhibitors (e.g., methotrexate and dichloromethotrexate, 3-amino-l,2,4-benzotriazine 1,4-dioxide, aminopterin, cytosine P-D- arabinofuranoside, 5-fluoro-5'-deoxyuridine, 5-fluorouracil, ganciclovir, hydroxyurea, actinomycin-D, and mitomycin C), DNA-intercalators or cross-linkers (e.g., bleomycin, carboplatin, carmustine, chlorambucil, cyclophosphamide, cis-diammineplatinum(II) dichloride (cisplatin), melphalan, mitoxantrone, and oxaliplatin), and DNA-RNA transcription regulators (e.g., actinomycin D, daunorubicin, doxorubicin, homoharringtonine, and idarubi
- quinonoid derivatives e.g. quinolones, genistein, bactacyclin
- busulfan ifosfamide, mechlorethamine
- triaziquone diaziquone
- carbazilquinone indoloquinone EO9
- diaziridinyl-benzoquinone methyl DZQ triethylenephospho
- a conjugate comprising anti-CTLA-4 antibody or antigen-binding fragment provided herein conjugated with a toxin.
- the therapeutic agent is a cytotoxin comprising a polypeptide having ribosomeinactivating activity including, for example, gelonin, bouganin, saporin, ricin, ricin A chain, bryodin, diphtheria toxin, restrictocin, Pseudomonas exotoxin A and variants thereof.
- the effector moiety is a radionuclide (e.g., a peptide receptor radionuclide) or a radiolabel.
- the effector moiety is a radionuclide or radiolabel with high-energy ionizing radiation that are capable of causing multiple strand breaks in nuclear DNA, leading to cell death.
- high-energy radionuclides include: 90Y, 1251, 1311, 1231, U lin, 105Rh, 153Sm, 67Cu, 67Ga, 166Ho, 177Lu, 186Re and 188Re. These isotopes typically produce high energy a- or P-particles which have a short path length.
- radionuclides kill cells to which they are in close proximity, for example neoplastic cells to which the conjugate has attached or has entered. They have little or no effect on nonlocalized cells and are essentially non-immunogenic.
- high-energy isotopes may be generated by thermal irradiation of an otherwise stable isotope, for example as in boron neutron-capture therapy (Guan et al., PNAS, 95: 13206-10, 1998).
- conjugate comprising an anti-CTLA-4 antibody or antigen-binding fragment provided herein conjugated with a label, which can generate a detectable signal, indirectly or directly.
- conjugates can be used for research or diagnostic applications, such as for the in vivo detection of cancer.
- the label is preferably capable of producing, either directly or indirectly, a detectable signal.
- the label may be radio-opaque or a radioisotope, such as 3H, 14C, 32P, 35S, 1231, 1251, 1311; a fluorescent (fluorophore) or chemiluminescent (chromophore) compound, such as fluorescein isothiocyanate, rhodamine or luciferin; an enzyme, such as alkaline phosphatase, P- galactosidase or horseradish peroxidase; an imaging agent; or a metal ion.
- a radioisotope such as 3H, 14C, 32P, 35S, 1231, 1251, 1311
- a fluorescent (fluorophore) or chemiluminescent (chromophore) compound such as fluorescein isothiocyanate, rhodamine or luciferin
- an enzyme such as alkaline phosphatase, P- galactosidase or horseradish peroxidase
- the label is a radioactive atom for scintigraphic studies, for example 99Tc or 1231, or a spin label for nuclear magnetic resonance (NMR) imaging (also known as magnetic resonance imaging, MRI), such as zirconium-89, iodine-123, iodine-131, indium-111, fluorine- 19, carbon- 13, nitrogen- 15, oxygen- 17, gadolinium, manganese or iron.
- NMR nuclear magnetic resonance
- zirconium-89 may be complexed to various metal chelating agents and conjugated to antibodies, e.g., for PET imaging (WO 2011/056983).
- the conjugates may be prepared using any methods known in the art. See, e.g., WO 2009/067800, WO 2011/133886, and U.S. Patent Application Publication No. 2014322129, incorporated by reference herein in their entirety.
- the linkage of an antibody or antigen binding fragment to a moiety is a direct or indirect linkage.
- the attachment can be covalent or non-covalent, e.g., via a biotinstreptavidin non-covalent interaction.
- a moiety can be attached by alkylation (e.g., at the epsilon-amino group lysines or the N-terminus of antibodies), reductive amination of oxidized carbohydrate, transesterification between hydroxyl and carboxyl groups, amidation at amino groups or carboxyl groups, and conjugation to thiols.
- the attachment of a moiety can be by chemical conjugation and linkage methods known in the art.
- the moiety can be linked to an antibody by a linker.
- linkers such as peptide linkers, cleavable linkers, non-cleavable linkers or linkers that aid in the conjugation reaction, can be used to link or conjugate the effector moieties to the antibody or antigen-binding fragment. Attachment of a linker to an antibody can be accomplished in a variety of ways, such as through surface lysines, reductive- coupling to oxidized carbohydrates, and through cysteine residues liberated by reducing interchain disulfide linkages. A variety of antibody drug conjugate linkage systems are known in the art, including hydrazone-, disulfide- and peptide-based linkages.
- an anti-CTLA-4 antibody or antigen-binding fragment provided herein is conjugated to one or more moieties, e.g. about 1 to about 20 drug moieties, through a linker (L).
- the conjugate comprises the following components: (antibody or antigen-binding fragment), (L) q and (moiety) m , wherein the antibody or antigen-binding fragment is any of the described capable of specifically binding CTLA-4 as described; L is a linker for linking the protein or polypeptide to the moiety; m is at least 1; q is 0 or more; and the resulting conjugate binds to CTLA-4.
- m is 1 to 4 and q is 0 to 8.
- the linker may be composed of one or more linker components.
- the linker typically has two reactive functional groups, i.e. bivalency in a reactive sense.
- Bivalent linker reagents which are useful to attach two or more functional or biologically active moieties, such as peptides, nucleic acids, drugs, toxins, antibodies, haptens, and reporter groups are known, and methods have been described their resulting conjugates (Hermanson, G. T. (1996) Bioconjugate Techniques; Academic Press: New York, p 234-242).
- the linker may comprise amino acid residues.
- Exemplary amino acid linker components include a dipeptide, a tripeptide, a tetrapeptide or a pentapeptide.
- Exemplary dipeptides include: valine-citrulline (vc or val-cit), alanine-phenylalanine (af or ala- phe).
- Exemplary tripeptides include: glycine-valine-citrulline (gly-val-cit) and glycine- glycine-glycine (gly-gly-gly).
- Amino acid residues which comprise an amino acid linker component include those occurring naturally, as well as minor amino acids and non-naturally occurring amino acid analogs, such as citrulline.
- Amino acid linker components can be designed and optimized in their selectivity for enzymatic cleavage by a particular enzymes, for example, a tumor-associated protease, cathepsin B, C and D, at a plasmin protease.
- Exemplary linker components include 6-maleimidocaproyl (“MC”), maleimidopropanoyl (“MP”), valine-citrulline (“val-cit”), a alanine-phenylalanine (“ala-phe”), p-aminobenzyloxycarbonyl (“PAB”), N-Succinimidyl 4-(2-pyridylthio)pentanoate (“SPP”), N-Succinimidyl 4-(N-maleimidomethyl)cyclohexane-I carboxylate (“SMCC”), and N- Succinimidyl (4-iodo-acetyl)aminobenzoate (“SLAB”).
- MC 6-maleimidocaproyl
- MP maleimidopropanoyl
- val-cit valine-citrulline
- ala-phe alanine-phenylalanine
- PAB p-aminobenzyloxycarbonyl
- Conjugates of an anti-CTLA-4 antibody or antigen-binding fragment provided herein and cytotoxic agent can be made using a variety of bifunctional protein-coupling agents such as N-succinimidyl-3-(2-pyridyldithiol) propionate (SPDP), iminothiolane (IT), bifunctional derivatives of imidoesters (such as dimethyl adipimidate HC1), active esters (such as disuccinimidyl substrate), aldehydes (such as glutaraldehyde), bis-azido compounds (such as bis(p-azidobenzoyl) hexanediamine), bis-diazonium derivatives (such as bis-(p- diazoniumbenzoyl)-ethylenediamine), diisocyanates (such as toluene 2,6-diisocyanate), and bis-active fluorine compounds (such as l,5-difluoro-2,4-di
- the antibody drug conjugate can be prepared by a variety of methods, such as organic chemistry reactions, conditions, and reagents known to those skilled in the art.
- a fusion protein containing an antibody or antigen-binding fragment and cytotoxic agent may be made, e.g., by recombinant techniques or peptide synthesis.
- the length of DNA may comprise respective regions encoding the two portions of the conjugate either adjacent one another or separated by a region encoding a linker peptide which does not destroy the desired properties of the conjugate.
- CTLA-4-binding polypeptides that are multispecific containing at least one antibody or antigen-binding fragment that binds CTLA-4 and one or more additional binding domains.
- the one or more additional domains bind to a second antigen or protein other than CTLA-4.
- the further antigen or protein may be an antigen expressed on a tumor, a molecule or receptor expressed on an immune cell, such as a T cell, e.g. a CD3, or an additional inhibitory receptor (e.g. PD-1, LAG3, TIM3, VISTA, TIGIT, SIRPa, NKG2A, B7H3, B7H4) or an activating receptor (e.g.
- the one or more additional domain is an antibody or antigenbinding fragment specific for the second antigen or protein.
- the present disclosure provides a nucleic acid or nucleic acids encoding an antibody or antigen-binding fragment thereof.
- the nucleic acid(s) of the present disclosure may comprise a polynucleotide sequence encoding any one of the anti-CTLA-4 antibodies or antigen-binding fragments thereof disclosed herein.
- the polypeptide sequences may be used to determine appropriate nucleic acid sequences encoding the particular antibody disclosed thereby.
- the nucleic acid sequence may be optimized to reflect particular codon "preferences" for various expression systems according to standard methods well known to those of skill in the art.
- nucleic acid molecules are those that comprise polynucleotides that encode one or more polypeptides (e.g., an immunoglobulin heavy chain or light chain) of a provided anti-CTLA-4 antibody.
- a nucleic acid molecule comprises a polynucleotide that encodes a heavy chain or a light chain of an anti-CTLA-4 antibody.
- a nucleic acid molecule comprises both a polynucleotide that encodes a heavy chain and a polynucleotide that encodes a light chain of an anti-CTLA-4 antibody.
- a first nucleic acid molecule comprises a first polynucleotide that encodes a heavy chain and a second nucleic acid molecule comprises a second polynucleotide that encodes a light chain.
- nucleic acid molecules comprising polynucleotides that encode one or more polypeptides of a binding molecule containing any of the provided anti-CTLA-4 antibodies, such as a CTLA-4-targeted antibody-drug conjugate (ADC) or a bispecific immune cell engager comprising the anti-CTLA-4 antibody or antigen binding fragment thereof as described herein.
- ADC CTLA-4-targeted antibody-drug conjugate
- bispecific immune cell engager comprising the anti-CTLA-4 antibody or antigen binding fragment thereof as described herein.
- the nucleic acid(s) is an isolated nucleic acid(s).
- the polynucleotides may include those encompassing natural and/or non-naturally occurring nucleotides and bases, e.g., including those with backbone modifications.
- the terms “nucleic acid molecule”, “nucleic acid”, “sequence of nucleotides”, and “polynucleotide” may be used interchangeably, and refer to a polymer of nucleotides. Such polymers of nucleotides may contain natural and/or non-natural nucleotides, and include, but are not limited to, DNA, RNA, and PNA.
- Nucleic acid sequence refers to the linear sequence of nucleotides that comprise the nucleic acid molecule or polynucleotide.
- the heavy chain and the light chain are expressed from one nucleic acid molecule, or from two separate nucleic acid molecules, as two separate polypeptides.
- a single polynucleotide encodes a single polypeptide comprising both a heavy chain variable domain fragment and a light chain variable domain fragment linked together.
- a nucleic acid of the present disclosure may be operably linked to a transcriptional control element, e.g., a promoter, and enhancer, etc.
- a transcriptional control element e.g., a promoter, and enhancer, etc.
- Suitable promoter and enhancer elements are known to those of skill in the art.
- a polynucleotide encoding a heavy chain or light chain of an anti-CTLA-4 antibody comprises a nucleotide sequence that encodes a leader sequence, which, when translated, is located at the N terminus of the heavy chain or light chain.
- the leader sequence may be the native heavy or light chain leader sequence, or may be another heterologous leader sequence.
- Nucleic acid molecules may be constructed using conventional recombinant DNA techniques well-known in the art.
- a nucleic acid molecule is an expression vector that is suitable for expression in a selected host cell.
- Vectors comprising polynucleotides that encode anti-CTLA-4 heavy chains and/or anti-CTLA-4 light chains are provided herein. Vectors comprising polynucleotides that encode anti-CTLA-4 heavy chains and/or anti-CTLA-4 light chains are also provided herein.
- the vector is selected from the group consisting of a DNA vector, an RNA vector, a plasmid, a lentiviral vector, an adenoviral vector, an adeno-associated viral vector, and a retroviral vector.
- the vector is an expression vector.
- a nucleic acid of the present disclosure may be present within an expression vector and/or a cloning vector.
- An expression vector can include a selectable marker, an origin of replication, and other features that provide for replication and/or maintenance of the vector.
- Suitable expression vectors include, e.g., plasmids, viral vectors, and the like. Large numbers of suitable vectors and promoters are known to those of skill in the art; many are commercially available for generating a subject recombinant construct.
- Bacterial pBs, phagescript, PsiX174, pBluescript SK, pBs KS, pNH8a, pNH16a, pNH18a, pNH46a (Stratagene, La Jolla, Calif., USA); pTrc99A, pKK223-3, pKK233-3, pDR540, and pRIT5 (Pharmacia, Uppsala, Sweden).
- Eukaryotic pWLneo, pSV2cat, pOG44, PXR1, pSG (Stratagene) pSVK3, pBPV, pMSG and pSVL (Pharmacia).
- Expression vectors generally have convenient restriction sites located near the promoter sequence to provide for the insertion of nucleic acid sequences encoding heterologous proteins.
- a selectable marker operative in the expression host may be present.
- Suitable expression vectors include, but are not limited to, viral vectors (e.g. viral vectors based on vaccinia virus; poliovirus; adenovirus (see, e.g., Li et al., Invest. Opthalmol. Vis. Sci. (1994) 35: 2543-2549; Borras et al., Gene Ther. (1999) 6: 515-524; Li and Davidson, Proc. Natl. Acad. Sci.
- viral vectors e.g. viral vectors based on vaccinia virus; poliovirus; adenovirus (see, e.g., Li et al., Invest. Opthalmol. Vis. Sci. (1994) 35: 2543-2549; Borras et al., Gene
- a retroviral vector e.g., Murine Leukemia Virus, spleen necrosis virus, and vectors derived from retroviruses such as Rous Sarcoma Virus, Harvey Sarcoma Virus, avian leukosis virus, human immunodeficiency virus, myeloproliferative sarcoma virus, and mammary tumor virus; and the like.
- Additional expression vectors suitable for use are, e.g., without limitation, a lentivirus vector, a gamma retrovirus vector, a foamy virus vector, an adeno-associated virus vector, an adenovirus vector, a pox virus vector, a herpes virus vector, an engineered hybrid virus vector, a transposon mediated vector, and the like.
- Viral vector technology is well known in the art and is described, for example, in Sambrook et al., 2012, Molecular Cloning: A Laboratory Manual, volumes 1-4, Cold Spring Harbor Press, NY), and in other virology and molecular biology manuals.
- Viruses, which are useful as vectors include, but are not limited to, retroviruses, adenoviruses, adeno- associated viruses, herpes viruses, and lentiviruses.
- a suitable vector contains an origin of replication functional in at least one organism, a promoter sequence, convenient restriction endonuclease sites, and one or more selectable markers, (e.g., WO 01/96584; WO 01/29058; and U.S. Pat. No. 6,326,193).
- a vector comprises a first polynucleotide sequence encoding a heavy chain and a second polynucleotide sequence encoding a light chain.
- the heavy chain and light chain are expressed from the vector as two separate polypeptides.
- the heavy chain variable domain fragment and light chain variable domain fragment are expressed as part of a single polypeptide, such as, for example, when the antibody is an scFv.
- a first vector comprises a polynucleotide that encodes a heavy chain and a second vector comprises a polynucleotide that encodes a light chain.
- the first vector and second vector are transfected into host cells in similar amounts (such as similar molar amounts or similar mass amounts).
- a mole- or mass-ratio of between 5:1 and 1:5 of the first vector and the second vector is transfected into host cells.
- a mass ratio of between 1:1 and 1:5 for the vector encoding the heavy chain and the vector encoding the light chain is used.
- a mass ratio of 1:2 for the vector encoding the heavy chain and the vector encoding the light chain is used.
- a host cell comprising any of the vectors or nucleic acids disclosed herein.
- the host cell may be of eukaryotic, prokaryotic, mammalian, or bacterial origin.
- anti-CTLA-4 heavy chains and/or anti-CTLA-4 light chains may be expressed in prokaryotic cells, such as bacterial cells; or in eukaryotic cells, such as fungal cells (such as yeast), plant cells, insect cells, and mammalian cells. Such expression may be carried out, for example, according to procedures known in the art.
- Exemplary eukaryotic cells that may be used to express polypeptides include, but are not limited to, COS cells, including COS 7 cells; HEK293 cells, including HEK293-6E cells; CHO cells, including CHO-S, DG44. Lecl3 CHO cells, and FUT8 CHO cells; PER.C6® cells (Crucell); and NSO cells.
- anti-CTLA-4 heavy chains and/or anti-CTLA-4 light chains may be expressed in yeast. See, e.g., U.S. Publication No. US 2006/0270045 Al.
- a particular eukaryotic host cell is selected based on its ability to make desired post-translational modifications to the anti-CTLA-4 heavy chains and/or anti-CTLA-4 light chains.
- CHO cells produce polypeptides that have a higher level of sialylation than the same polypeptide produced in HEK293 cells.
- nucleic acids may be transiently or stably transfected in the desired host cells, according to any suitable method.
- host cells comprising any of the polynucleotides or vectors described herein.
- host cells are capable of over-expressing heterologous DNAs can be used for the purpose of isolating the genes encoding the antibody, polypeptide or protein of interest.
- mammalian host cells include but not limited to COS, HeLa, and CHO cells. See also PCT Publication No. WO 87/04462.
- Suitable nonmammalian host cells include prokaryotes (such as E. coli or B. subtilis) and yeast (such as .S'. cerevisiae, S. pombe; or K. laclis).
- a method of producing an antibody or antigen-binding fragment thereof that binds to CTLA-4 comprises culturing the host cell.
- the nucleic acids encoding a subject antibody are introduced directly into a host cell, and the cell incubated under conditions sufficient to induce expression of the encoded antibody. Expression may conveniently be achieved by culturing under appropriate conditions recombinant host cells containing the nucleic acid.
- an antibody or antigen-binding fragment thereof may be isolated and/or purified using any suitable technique, and then used as desired.
- Anti-CTLA-4 antibodies may be purified by any suitable method. Such methods include, but are not limited to, the use of affinity matrices or hydrophobic interaction chromatography. Suitable affinity ligands such as ligands that bind antibody constant regions. For example, a Protein A, Protein G, Protein A/G, or an antibody affinity column may be used to bind the constant region and to purify an anti-CTEA-4 antibody. Hydrophobic interactive chromatography, for example, a butyl or phenyl column, may also suitable for purifying some polypeptides such as antibodies. Ion exchange chromatography (e.g., anion exchange chromatography and/or cation exchange chromatography) may also suitable for purifying some polypeptides such as antibodies.
- affinity ligands such as ligands that bind antibody constant regions.
- Hydrophobic interactive chromatography for example, a butyl or phenyl column, may also suitable for purifying some polypeptides such as antibodies.
- Ion exchange chromatography
- Mixed-mode chromatography e.g., reversed phase/anion exchange, reversed phase/cation exchange, hydrophilic interaction/anion exchange, hydrophilic interaction/cation exchange, etc.
- Antibodies or antigen-binding fragments thereof as provided herein, and encoding nucleic acid molecules and vectors may be isolated and/or purified, e.g. from their natural environment, in substantially pure or homogeneous form, or, in the case of nucleic acid, free or substantially free of nucleic acid or genes of origin other than the sequence encoding a polypeptide with the desired function.
- the pharmaceutical composition can further comprise a pharmaceutically acceptable excipient.
- the pharmaceutical composition can contain one or more excipients for modifying, maintaining or preserving, for example, the pH, osmolarity, viscosity, clarity, color, isotonicity, odor, sterility, stability, rate of dissolution or release, adsorption, or penetration of the composition.
- compositions may comprise buffers such as neutral buffered saline, phosphate buffered saline and the like; carbohydrates such as glucose, mannose, sucrose or dextrans, mannitol; proteins; polypeptides or amino acids such as glycine; antioxidants; chelating agents such as EDTA or glutathione; adjuvants (e.g., aluminum hydroxide); and preservatives.
- buffers such as neutral buffered saline, phosphate buffered saline and the like
- carbohydrates such as glucose, mannose, sucrose or dextrans, mannitol
- proteins such as glucose, mannose, sucrose or dextrans, mannitol
- proteins such as glucose, mannose, sucrose or dextrans, mannitol
- proteins such as glucose, mannose, sucrose or dextrans, mannitol
- proteins such as glucose, mannose, sucrose or dextrans, mannitol
- proteins such as glucose, mannose
- the pharmaceutical composition is a solid, such as a powder, capsule, or tablet.
- the components of the pharmaceutical composition can be lyophilized.
- the solid pharmaceutical composition is reconstituted or dissolved in a liquid prior to administration.
- the pharmaceutical composition is a liquid, for example a provided protein dissolved in an aqueous solution (such as physiological saline or Ringer’s solution).
- the pH of the pharmaceutical composition is between about 4.0 and about 8.5 (such as between about 4.0 and about 5.0, between about 4.5 and about 5.5, between about 5.0 and about 6.0, between about 5.5 and about 6.5, between about 6.0 and about 7.0, between about 6.5 and about 7.5, between about 7.0 and about 8.0, or between about 7.5 and about 8.5).
- the pharmaceutical composition comprises a pharmaceutically-acceptable excipient, for example a filler, binder, coating, preservative, lubricant, flavoring agent, sweetening agent, coloring agent, a solvent, a buffering agent, a chelating agent, or stabilizer.
- a pharmaceutically-acceptable excipient for example a filler, binder, coating, preservative, lubricant, flavoring agent, sweetening agent, coloring agent, a solvent, a buffering agent, a chelating agent, or stabilizer.
- pharmaceutically-acceptable fillers include cellulose, dibasic calcium phosphate, calcium carbonate, microcrystalline cellulose, sucrose, lactose, glucose, mannitol, sorbitol, maltol, pregelatinized starch, com starch, or potato starch.
- Examples of pharmaceutically-acceptable binders include polyvinylpyrrolidone, starch, lactose, xylitol, sorbitol, maltitol, gelatin, sucrose, polyethylene glycol, methyl cellulose, or cellulose.
- Examples of pharmaceutically-acceptable coatings include hydroxypropyl methylcellulose (HPMC), shellac, com protein zein, or gelatin.
- Examples of pharmaceutically- acceptable disintegrants include polyvinylpyrrolidone, carboxymethyl cellulose, or sodium starch glycolate.
- Examples of pharmaceutically-acceptable lubricants include polyethylene glycol, magnesium stearate, or stearic acid.
- Examples of pharmaceutically-acceptable preservatives include methyl parabens, ethyl parabens, propyl paraben, benzoic acid, or sorbic acid.
- Examples of pharmaceutically-acceptable sweetening agents include sucrose, saccharine, aspartame, or sorbitol.
- Examples of pharmaceutically-acceptable buffering agents include carbonates, citrates, gluconates, acetates, phosphates, or tartrates.
- the pharmaceutical composition further comprises an agent for the controlled or sustained release of the product, such as injectable microspheres, bio- erodible particles, polymeric compounds (polylactic acid, polyglycolic acid), beads, or liposomes.
- an agent for the controlled or sustained release of the product such as injectable microspheres, bio- erodible particles, polymeric compounds (polylactic acid, polyglycolic acid), beads, or liposomes.
- the pharmaceutical composition is sterile. Sterilization may be accomplished by filtration through sterile filtration membranes or radiation. Where the composition is lyophilized, sterilization using this method may be conducted either prior to or following lyophilization and reconstitution.
- the composition for parenteral administration may be stored in lyophilized form or in solution.
- parenteral compositions generally are placed into a container having a sterile access port, for example, an intravenous solution bag or vial having a stopper pierceable by a hypodermic injection needle.
- a pharmaceutically acceptable carrier may be a pharmaceutically acceptable material, composition, or vehicle.
- the carrier may be a liquid or solid filler, diluent, excipient, solvent, or encapsulating material, or some combination thereof.
- Each component of the carrier must be “pharmaceutically acceptable” in that it must be compatible with the other ingredients of the formulation. It also must be suitable for contact with any tissue, organ, or portion of the body that it may encounter, meaning that it must not carry a risk of toxicity, irritation, allergic response, immunogenicity, or any other complication that excessively outweighs its therapeutic benefits.
- the pharmaceutical composition is for administration to a subject.
- dosages and routes of administration of the pharmaceutical composition are determined according to the size and condition of the subject, according to standard pharmaceutical practice.
- the therapeutically effective dose can be estimated initially either in cell culture assays or in animal models such as mice, rats, rabbits, dogs, pigs, or monkeys. An animal model may also be used to determine the appropriate concentration range and route of administration. Such information can then be used to determine useful doses and routes for administration in humans. The exact dosage will be determined in light of factors related to the subject requiring treatment. Dosage and administration are adjusted to provide sufficient levels of the active compound or to maintain the desired effect. Factors that may be taken into account include the severity of the disease state, the general health of the subject, the age, weight, and gender of the subject, time and frequency of administration, drug combination(s), reaction sensitivities, and response to therapy.
- compositions may be administered every 3 to 4 days, every week, or biweekly depending on the half-life and clearance rate of the particular formulation. The frequency of dosing will depend upon the pharmacokinetic parameters of the molecule in the formulation used. Typically, a composition is administered until a dosage is reached that achieves the desired effect. The composition may therefore be administered as a single dose, or as multiple doses (at the same or different concentrations/dosages) over time, or as a continuous infusion. Further refinement of the appropriate dosage is routinely made. Appropriate dosages may be ascertained through use of appropriate dose-response data.
- the pharmaceutical composition is for administration to a subject through any route, including orally, transdermally, by inhalation, intravenously, intraarterially, intramuscularly, direct application to a wound site, application to a surgical site, intraperitoneally, by suppository, subcutaneously, intradermally, transcutaneously, by nebulization, intrapleurally, intraventricularly, intra-articularly, intraocularly, or intraspinally.
- a provided pharmaceutical formulation may, for example, be in a form suitable for intravenous infusion.
- the dosage of the pharmaceutical composition is a single dose or a repeated dose.
- the doses are given to a subject once per day, twice per day, three times per day, or four or more times per day.
- about 1 or more (such as about 2 or more, about 3 or more, about 4 or more, about 5 or more, about 6 or more, or about 7 or more) doses are given in a week.
- multiple doses are given over the course of days, weeks, months, or years.
- a course of treatment is about 1 or more doses (such as about 2 or more does, about 3 or more doses, about 4 or more doses, about 5 or more doses, about 7 or more doses, about 10 or more doses, about 15 or more doses, about 25 or more doses, about 40 or more doses, about 50 or more doses, or about 100 or more doses).
- an administered dose of the pharmaceutical composition is about 1 pg of protein per kg subject body mass or more (such as about 2 pg of protein per kg subject body mass or more, about 5 pg of protein per kg subject body mass or more, about 10 pg of protein per kg subject body mass or more, about 25 pg of protein per kg subject body mass or more, about 50 pg of protein per kg subject body mass or more, about 100 pg of protein per kg subject body mass or more, about 250 pg of protein per kg subject body mass or more, about 500 pg of protein per kg subject body mass or more, about 1 mg of protein per kg subject body mass or more, about 2 mg of protein per kg subject body mass or more, or about 5 mg of protein per kg subject body mass or more).
- the antibodies and antigen-binding fragments or CTLA-4 binding molecules containing the same described herein, including pharmaceutical compositions thereof, can be used in a variety of therapeutic applications, such as the treatment of a disease.
- Such methods and uses include therapeutic methods and uses, for example, involving administration of the molecules or compositions containing the same, to a subject having a disease or disorder.
- the molecule or engineered cell is administered in an effective amount to effect treatment of the disease or disorder.
- Uses include uses of the antibodies and antigenbinding fragments or CTLA-4 binding molecules containing the same described herein, including pharmaceutical compositions thereof, in such methods and treatments, and in the preparation of a medicament in order to carry out such therapeutic methods.
- the methods are carried out by administering the antibodies and antigen-binding fragments or CTLA-4 binding molecules containing the same, including pharmaceutical compositions thereof, or compositions comprising the same, to the subject having or suspected of having the disease or disorder. In some embodiments, the methods thereby treat the disease or disorder in the subject. Also provided are any methods or uses involving administering a provided engineered cell as described, or pharmaceutical compositions containing the same, for treating a disease or disorder.
- the disease or disorder is a tumor or cancer.
- cancer refers to a neoplasm or tumor resulting from abnormal uncontrolled growth of cells. “Cancer” explicitly includes leukemias and lymphomas.
- carcinoma also refers to a disease involving cells that have the potential to metastasize to distal sites.
- the disease or disorder is carcinoma, including that of the bladder, breast, colon, kidney, liver, lung, ovary, pancreas, stomach, cervix, thyroid and skin; including squamous cell carcinoma; hematopoietic tumors of lymphoid lineage, including leukemia, acute lymphocytic leukemia, acute lymphoblastic leukemia, B-cell lymphoma, T-cell lymphoma, Berketts lymphoma; hematopoietic tumors of myeloid lineage, including acute and chronic myelogenous leukemias and promyelocytic leukemia; tumors of mesenchymal origin, including fibrosarcoma and rhabdomyoscarcoma; other tumors, including melanoma, seminoma, tetratocarcinoma, neuroblastoma and gliom
- cancers caused by aberrations in apoptosis would also be treated by the methods and compositions of the invention.
- Such cancers may include, but are not be limited to, follicular lymphomas, carcinomas with p53 mutations, hormone dependent tumors of the breast, prostate and ovary, and precancerous lesions such as familial adenomatous polyposis, and myelodysplastic syndromes.
- Illustrative subjects include mammalian subjects, such as farm animals, domestic animals, and human patients.
- the subject is a human subject.
- subject and patient are used interchangeably herein.
- a subject is selected that is known, suspected or that has been identified as having T cells expressing CTLA-4.
- a provided antibody or antigen-binding fragment, CTLA-4 binding molecule or engineered cell as described herein, or pharmaceutical composition comprising the same, is administered to the subject.
- the administration to the subject will generally have an effect due to its binding with the CTLA-4 target.
- alleviation or treatment of a disease or disorder involves the lessening of one or more symptoms or medical problems associated with the disease or disorder.
- the therapeutically effective amount of the drug can accomplish one or a combination of the following: reduce the number of cancer cells; reduce the tumor size; inhibit (i.e., to decrease to some extent and/or stop) cancer cell infiltration into peripheral organs; inhibit tumor metastasis; inhibit, to some extent, tumor growth; and/or relieve to some extent one or more of the symptoms associated with the cancer.
- a composition of this disclosure can be used to prevent the onset or reoccurrence of the disease or disorder in the subject.
- the antibody or antigen-binding fragment, CTLA-4 binding molecule or engineered cell as described herein, or pharmaceutical composition comprising the same can be used to inhibit growth of mammalian cancer cells (such as human cancer cells).
- a method of treating cancer can include administering an effective amount of any of the pharmaceutical compositions described herein to a subject with cancer.
- the effective amount of the pharmaceutical composition can be administered to inhibit, halt, or reverse progression of cancers.
- Human cancer cells can be treated in vivo, or ex vivo. In ex vivo treatment of a human patient, tissue or fluids containing cancer cells are treated outside the body and then the tissue or fluids are reintroduced back into the patient.
- the cancer is treated in a human patient in vivo by administration of the therapeutic composition into the patient.
- the antibody or antigen-binding fragment, CTLA-4 binding molecule or engineered cell as described herein, or pharmaceutical composition comprising the same are useful in treating, alleviating a symptom of, ameliorating and/or delaying the progression of a cancer or other neoplastic condition.
- the cancer is bladder cancer, breast cancer, uterine/cervical cancer, ovarian cancer, prostate cancer, testicular cancer, esophageal cancer, gastrointestinal cancer, pancreatic cancer, colorectal cancer, colon cancer, kidney cancer, head and neck cancer, lung cancer, stomach cancer, germ cell cancer, bone cancer, liver cancer, thyroid cancer, skin cancer, neoplasm of the central nervous system, lymphoma, leukemia, myeloma, sarcoma, and virus-related cancer.
- the cancer is a metastatic cancer, refractory cancer, or recurrent cancer.
- compositions of the invention can be administered in dosages and routes and at times to be determined in appropriate pre-clinical and clinical experimentation and trials. Compositions may be administered multiple times at dosages within these ranges. Administration of the compositions may be combined with other methods useful to treat the desired disease or condition as determined by those of skill in the art. Typically, precise amount of the compositions of the present disclosure to be administered can be determined by a physician with consideration of individual differences in age, weight, tumor size, extent of infection or metastasis, and condition of the patient (subject).
- a therapeutically effective dose may be, by way of nonlimiting example, from about 0.01 pg/kg body weight to about 20 mg/kg body weight. In some embodiments, the therapeutically effective dose may be, by way of nonlimiting example, from about 0.01 mg/kg body weight to about 5-20 mg/kg body weight. Intervals between single doses can be, for example, daily, weekly, monthly, every three months or yearly.
- An exemplary treatment regimen entails administration once per week, once every two weeks, once every three weeks, once every four weeks, once a month, once every three months or once every three to six months. Common dosing frequencies may range, for example, from twice daily to once a week.
- the antibody or antigen-binding fragment, CTLA-4 binding molecule or pharmaceutical composition comprising the same, of the present disclosure can be administered alone or in combination with other modes of treatment, such as other anti-cancer agents. They can be provided before, substantially contemporaneous with, or after other modes of treatment (i.e., concurrently or sequentially).
- the method of treatment described herein can further include administering: radiation therapy, chemotherapy, vaccination, targeted tumor therapy, CAR-T therapy, oncolytic virus therapy, cancer immunotherapy, cytokine therapy, surgical resection, chromatin modification, ablation, cryotherapy, an antisense agent against a tumor target, a siRNA agent against a tumor target, a microRNA agent against a tumor target or an anti-cancer/tumor agent, or a biologic, such as an antibody, cytokine, or receptor extracellular domain-Fc fusion.
- the antibody or antigen-binding fragment, CTLA-4 binding molecule or pharmaceutical composition comprising the same, of the present disclosure can be administered in combination with a molecule that disrupts or enhances alternative immunomodulatory pathways (such as TIM3, TIM4, 0X40, CD40, GITR, 4-1-BB, B7-H1, PD- 1, B7-H3, B7-H4, LIGHT, BTLA, ICOS, CD27 or LAG3) or modulates the activity of effecter molecules such as cytokines (e.g., IL-4, IL-7, IL-10, IL-12, IL-15, IL-17, GF-beta, IFNg, Flt3, BLys) and chemokines (e.g., CCL21) in order to enhance the immunomodulatory effects.
- cytokines e.g., IL-4, IL-7, IL-10, IL-12, IL-15, IL-17, GF-beta, IFNg, Flt3, BLys
- Specific embodiments include a bispecific antibody comprising an anti-CTLA-4 antibody described herein or antigen binding fragment thereof, in combination with anti-PD-1 (pembrolizumab (Keytruda) or Nivolumab (Opdivo)), anti-B7-Hl (atezolizumab (Tecentriq) or durvalumab (Imfinzi)), anti-B7-H3, anti-B7-H4, anti-LIGHT, anti-LAG3, anti-TIM3, anti- TIM4 anti-CD40, anti-OX40, anti-GITR, anti- BTLA, anti-CD27, anti-ICOS or anti-4-lBB.
- an antibody of the invention or antigen binding fragment thereof is administered in combination with a molecule that activates different stages or aspects of the immune response in order to achieve a broader immune response, such as IDO inhibitors.
- the antibody or antigen-binding fragment, CTLA-4 binding molecule or pharmaceutical composition comprising the same, of the present disclosure can be administered in combination with at least one additional therapeutic agent selected from the group consisting of: pomalyst, revlimid, lenalidomide, pomalidomide, thalidomide, a DNA- alkylating platinum-containing derivative, cisplatin, 5-fluorouracil, cyclophosphamide, an anti-CD137 antibody, an anti-PD-1 antibody, an anti-PD-Ll antibody, an anti-CD20 antibody, an anti-CD40 antibody, an anti-DR5 antibody, an anti-CDld antibody, an anti-TIM3 antibody, an anti-SLAMF7 antibody, an anti-KIR receptor antibody, an anti-OX40 antibody, an anti- HER2 antibody, an anti-ErbB-2 antibody, an anti-EGFR antibody, cetuximab, rituximab, trastuzumab, pembrolizum
- the antibody or antigen-binding fragment, CTLA-4 binding molecule or pharmaceutical composition comprising the same, of the present disclosure can be administered prior to a surgery to remove a solid tumor in the subject. In some embodiments, the antibody or antigen-binding fragment, CTLA-4 binding molecule or pharmaceutical composition comprising the same, of the present disclosure can be administered after a surgery to remove a solid tumor in the subject.
- kits comprising the provided compositions.
- the kit comprises any of the provided compositions and instructions for administering the composition to a subject.
- Kits can optionally include one or more components such as instructions for use, devices and additional reagents (e.g., sterilized water or saline solutions for dilution of the compositions and/or reconstitution of lyophilized protein), and components, such as tubes, containers and syringes for practice of the methods.
- additional reagents e.g., sterilized water or saline solutions for dilution of the compositions and/or reconstitution of lyophilized protein
- components such as tubes, containers and syringes for practice of the methods.
- kits can further contain reagents for collection of samples, preparation and processing of samples, and/or reagents for quantitating the amount of one or more surface markers in a sample, such as, but not limited to, detection reagents, such as antibodies, buffers, substrates for enzymatic staining, chromagens or other materials, such as slides, containers, microtiter plates, and optionally, instructions for performing the methods.
- detection reagents such as antibodies, buffers, substrates for enzymatic staining, chromagens or other materials, such as slides, containers, microtiter plates, and optionally, instructions for performing the methods.
- kits can be provided as articles of manufacture that include packing materials for the packaging of the cells, antibodies or reagents, or compositions thereof, or one or more other components.
- the kits can contain containers, bottles, tubes, vial and any packaging material suitable for separating or organizing the components of the kit.
- the one or more containers may be formed from a variety of materials such as glass or plastic.
- the one or more containers hold a composition comprising cells or an antibody or other reagents for use in the methods.
- the article of manufacture or kit herein may comprise the cells, antibodies or reagents in separate containers or in the same container.
- the one or more containers holding the composition may be a single-use vial or a multi-use vial, which, in some cases, may allow for repeat use of the composition.
- the article of manufacture or kit may further comprise a second container comprising a suitable diluent.
- the article of manufacture or kit may further include other materials desirable from a commercial, therapeutic, and user standpoint, including other buffers, diluents, filters, needles, syringes, therapeutic agents and/or package inserts with instructions for use.
- the kit can, optionally, include instructions. Instructions typically include a tangible expression describing the cell composition, reagents and/or antibodies and, optionally, other components included in the kit, and methods for using such. In some embodiments, the instructions indicate methods for using the cell compositions and antibodies for administration to a subject for treating a disease or condition, such as in accord with any of the provided embodiments. In some embodiments, the instructions are provided as a label or a package insert, which is on or associated with the container. In some embodiments, the instructions may indicate directions for reconstitution and/or use of the composition.
- an optionally substituted group means that the group is unsubstituted or is substituted.
- nucleic acid molecule refers to a polymer of nucleotides.
- polymers of nucleotides may contain natural and/or non-natural nucleotides, and include, but are not limited to, DNA, RNA, and PNA.
- Nucleic acid sequence refers to the linear sequence of nucleotides comprised in the nucleic acid molecule or polynucleotide.
- isolated refers to a molecule that has been separated from at least some of the components with which it is typically found in nature or produced.
- a polypeptide is referred to as “isolated” when it is separated from at least some of the components of the cell in which it was produced.
- a polypeptide is secreted by a cell after expression, physically separating the supernatant containing the polypeptide from the cell that produced it is considered to be “isolating” the polypeptide.
- a polynucleotide is referred to as “isolated” when it is not part of the larger polynucleotide (such as, for example, genomic DNA or mitochondrial DNA, in the case of a DNA polynucleotide) in which it is typically found in nature, or is separated from at least some of the components of the cell in which it was produced, for example, in the case of an RNA polynucleotide.
- a DNA polynucleotide that is contained in a vector inside a host cell may be referred to as “isolated”.
- polypeptide and “protein” are used interchangeably to refer to a polymer of amino acid residues, and are not limited to a minimum length. Such polymers of amino acid residues may contain natural or non-natural amino acid residues, and include, but are not limited to, peptides, oligopeptides, dimers, trimers, and multimers of amino acid residues. Both full-length proteins and fragments thereof are encompassed by the definition.
- the terms also include post-expression modifications of the polypeptide, for example, glycosylation, sialylation, acetylation, phosphorylation, and the like.
- polypeptide refers to a protein which includes modifications, such as deletions, additions, and substitutions (generally conservative in nature), to the native sequence, as long as the protein maintains the desired activity. These modifications may be deliberate, as through site-directed mutagenesis, or may be accidental, such as through mutations of hosts which produce the proteins or errors due to PCR amplification.
- isolated protein means that a subject protein (1) is free of at least some other proteins with which it would typically be found in nature, (2) is essentially free of other proteins from the same source, e.g., from the same species, (3) is expressed by a cell from a different species, (4) has been separated from at least about 50 percent of polynucleotides, lipids, carbohydrates, or other materials with which it is associated in nature, (5) is not associated (by covalent or noncovalent interaction) with portions of a protein with which the "isolated protein" is associated in nature, (6) is operably associated (by covalent or noncovalent interaction) with a polypeptide with which it is not associated in nature, or (7) does not occur in nature.
- Such an isolated protein can be encoded by genomic DNA, cDNA, mRNA or other RNA, of may be of synthetic origin, or any combination thereof.
- the isolated protein is substantially pure or substantially free from proteins or polypeptides or other contaminants that are found in its natural environment that would interfere with its use (therapeutic, diagnostic, prophylactic, research or otherwise).
- nucleic acids or proteins generally denotes a nucleic acid or polypeptide that is substantially free from other components as determined by analytical techniques well known in the art (e.g., a purified polypeptide or polynucleotide forms a discrete band in an electrophoretic gel, chromatographic eluate, and/or a media subjected to density gradient centrifugation).
- nucleic acid or polypeptide that gives rise to essentially one band in an electrophoretic gel is “purified.”
- a purified nucleic acid or protein is at least about 50% pure, usually at least about 75%, 80%, 85%, 90%, 95%, 96%, 99% or more pure (e.g., percent by weight or on a molar basis).
- recombinant indicates that the material (e.g., a nucleic acid or a polypeptide) has been artificially (i.e., non-naturally) altered by human intervention. The alteration can be performed on the material within, or removed from, its natural environment or state.
- a “recombinant nucleic acid” is one that is made by recombining nucleic acids, e.g., during cloning, affinity modification, DNA shuffling or other well-known molecular biological procedures.
- a “recombinant DNA molecule,” is comprised of segments of DNA joined together by means of such molecular biological techniques.
- recombinant protein or “recombinant polypeptide” as used herein refers to a protein molecule which is expressed using a recombinant DNA molecule.
- a “recombinant host cell” is a cell that contains and/or expresses a recombinant nucleic acid or that is otherwise altered by genetic engineering, such as by introducing into the cell a nucleic acid molecule encoding a recombinant protein, such as a immunomodulatory protein provided herein.
- Transcriptional control signals in eukaryotes comprise “promoter” and “enhancer” elements. Promoters and enhancers consist of short arrays of DNA sequences that interact specifically with cellular proteins involved in transcription.
- Promoter and enhancer elements have been isolated from a variety of eukaryotic sources including genes in yeast, insect and mammalian cells and viruses (analogous control elements, i.e., promoters, are also found in prokaryotes). The selection of a particular promoter and enhancer depends on what cell type is to be used to express the protein of interest.
- substantially pure means an object species is the predominant species present i.e., on a molar basis it is more abundant than any other individual species in the composition), and a substantially purified fraction is a composition wherein the object species comprises at least about 50 percent (on a molar basis) of all macromolecular species present. Generally, a substantially pure composition will comprise more than about 80 percent of all macromolecular species present in the composition, for example, in some embodiments, more than about 85%, 90%, 95%, and 99%. In some embodiments, the object species is purified to essential homogeneity (contaminant species cannot be detected in the composition by conventional detection methods) wherein the composition consists essentially of a single macromolecular species.
- substantially similar denotes a sufficiently high degree of similarity between two or more numeric values such that one of skill in the art would consider the difference between the two or more values to be of little or no biological and/or statistical significance within the context of the biological characteristic measured by said value.
- the two or more substantially similar values differ by no more than about any one of 5%, 10%, 15%, 20%, 25%, or 50%.
- a polypeptide “variant” means a biologically active polypeptide having at least about 80% amino acid sequence identity with the native sequence polypeptide after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity.
- Such variants include, for instance, polypeptides wherein one or more amino acid residues are added, or deleted, at the N- or C-terminus of the polypeptide.
- a variant will have at least about 80% amino acid sequence identity.
- a variant will have at least about 90% amino acid sequence identity.
- a variant will have at least about 95% amino acid sequence identity with the native sequence polypeptide.
- percent (%) amino acid sequence identity and “homology” with respect to a peptide, polypeptide or antibody sequence are defined as the percentage of amino acid residues in a candidate sequence that are identical with the amino acid residues in the specific peptide or polypeptide sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity. Alignment for purposes of determining percent amino acid sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST, BLAST-2, ALIGN or MEGALIGNTM (DNASTAR) software. Those skilled in the art can determine appropriate parameters for measuring alignment, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared.
- amino acid substitution may include but are not limited to the replacement of one amino acid in a polypeptide with another amino acid. Exemplary substitutions are shown in Table 4. Amino acid substitutions may be introduced into an antibody of interest and the products screened for a desired activity, for example, retained/improved antigen binding, decreased immunogenicity, or improved ADCC or CDC.
- Amino acids may be grouped according to common side-chain properties:
- Non-conservative substitutions will entail exchanging a member of one of these classes for another class.
- conservative amino acid substitution as used herein means an amino acid substitution in which an amino acid residue is substituted by another amino acid residue having a side chain R group with similar chemical properties (e.g., charge or hydrophobicity).
- Examples of groups of amino acids that have side chains with similar chemical properties include 1) aliphatic side chains: glycine, alanine, valine, leucine, and isoleucine; 2) aliphatic - hydroxyl side chains: serine and threonine; 3) amide-containing side chains: asparagine and glutamine; 4) aromatic side chains: phenylalanine, tyrosine, and tryptophan; 5) basic side chains: lysine, arginine, and histidine; 6) acidic side chains: aspartic acid and glutamic acid; and 7) sulfur-containing side chains: cysteine and methionine.
- Conservative amino acids substitution groups are: valine-leucine-isoleucine, phenylalanine-tyrosine, lysine-arginine, alanine-valine, glutamate-aspartate, and asparagine-glutamine.
- nucleotides or amino acid positions “correspond to” nucleotides or amino acid positions in a disclosed sequence refers to nucleotides or amino acid positions identified upon alignment with the disclosed sequence based on structural sequence alignment or using a standard alignment algorithm, such as the GAP algorithm. By aligning the sequences, one skilled in the art can identify corresponding residues, for example, using conserved and identical amino acid residues as guides.
- the term “specifically binds” as used herein means the ability of a protein, under specific binding conditions, to bind to a target protein such that its affinity or avidity is at least 10 times as great, but optionally 50, 100, 250 or 500 times as great, or even at least 1000 times as great as the average affinity or avidity of the same protein to a collection of random peptides or polypeptides of sufficient statistical size.
- a specifically binding protein need not bind exclusively to a single target molecule but may specifically bind to more than one target molecule. In some cases, a specifically binding protein may bind to a protein that has similarity in structural conformation with the target protein (e.g., paralogs or orthologs).
- a protein of the invention may specifically bind to more than one distinct species of target molecule due to cross-reactivity.
- Solid-phase ELISA immunoassays, ForteBio Octet or Biacore measurements can be used to determine specific binding between two proteins.
- interactions between two binding proteins have dissociation constants (Kd) less than about IxlO' 5 M, and often as low as about 1 x 10' 12 M.
- dissociation constants Kd
- interactions between two binding proteins have dissociation constants of less than about IxlO' 6 M, IxlO' 7 M, IxlO' 8 M, IxlO' 9 M, IxlO' 10 M, or IxlO' 11 M or less.
- affinity refers to the strength of the sum total of noncovalent interactions between a single binding site of a molecule (for example, an antibody) and its binding partner (for example, an antigen).
- the affinity or the apparent affinity of a molecule X for its partner Y can generally be represented by the dissociation constant (KD) or the Ko-apparent, respectively.
- KD dissociation constant
- Affinity can be measured by common methods known in the art (such as, for example, ELISA KD, KinExA, flow cytometry, and/or surface plasmon resonance devices), including those described herein. Such methods include, but are not limited to, methods involving BIAcore®, Octet®, or flow cytometry.
- the KD of the antigen-binding molecule is measured by flow cytometry using an antigen-expressing cell line and fitting the mean fluorescence measured at each antibody concentration to a non-linear one-site binding equation (Prism Software graphpad).
- vector is used to describe a polynucleotide that can be engineered to contain a cloned polynucleotide or polynucleotides that can be propagated in a host cell.
- a vector can include one or more of the following elements: an origin of replication, one or more regulatory sequences (such as, for example, promoters and/or enhancers) that regulate the expression of the polypeptide of interest, and/or one or more selectable marker genes (such as, for example, antibiotic resistance genes and genes that can be used in colorimetric assays, for example, P-galactosidase).
- expression vector refers to a vector that is used to express a polypeptide of interest in a host cell.
- a “host cell” refers to a cell that may be or has been a recipient of a vector or isolated polynucleotide.
- Host cells may be prokaryotic cells or eukaryotic cells.
- Exemplary eukaryotic cells include mammalian cells, such as primate or non-primate animal cells; fungal cells, such as yeast; plant cells; and insect cells.
- Nonlimiting exemplary mammalian cells include, but are not limited to, NSO cells, PER.C6® cells (Crucell), and 293 and CHO cells, and their derivatives, such as 293-6E, CHO-DG44, CHO-K1, CHO-S, and CHO-DS cells.
- Host cells include progeny of a single host cell, and the progeny may not necessarily be completely identical (in morphology or in genomic DNA complement) to the original parent cell due to natural, accidental, or deliberate mutation.
- a host cell includes cells transfected in vivo with a polynucleotide(s) a provided herein.
- expression refers to the process by which a polynucleotide is transcribed from a DNA template (such as into an mRNA or other RNA transcript) and/or the process by which a transcribed mRNA is subsequently translated into peptide, polypeptides or proteins.
- Transcripts and encoded polypeptides may be collectively referred to as “gene product.” If the polynucleotide is derived from genomic DNA, expression may include splicing of the mRNA in a eukaryotic cell.
- tumor microenvironment refers to the environment in which a tumor exists, which is the non-cellular area within the tumor and the area directly outside the tumorous tissue but does not pertain to the intracellular compartment of the cancer cell itself.
- the tumor and the tumor microenvironment are closely related and interact constantly.
- a tumor can change its microenvironment, and the microenvironment can affect how a tumor grows and spreads.
- the tumor microenvironment has a low pH in the range of 5.6 to 6.8, more commonly in the range of 6.0 to 6.8, in the range of 6.2-6.8.
- a normal physiological pH is in the range of 7.2-7.8, more commonly 7.35 to 7.45, such as about 7.4.
- the tumor microenvironment is also known to have lower concentration of glucose and other nutrients, but higher concentration of lactic acid, in comparison with blood plasma. Furthermore, the tumor microenvironment can have a temperature that is 0.3 to 1° C. higher than the normal physiological temperature.
- the tumor microenvironment has been discussed in Gillies et al., “MRI of the Tumor Microenvironment,” Journal of Magnetic Resonance Imaging, vol. 16, pp. 430-450, 2002, hereby incorporated by reference herein its entirety.
- the term “non-tumor microenvironment” refers to a microenvironment at a site other than a tumor.
- the terms “individual” and “subject” are used interchangeably herein to refer to an animal; for example a mammal.
- patient includes human and veterinary subjects.
- methods of treating mammals including, but not limited to, humans, rodents, simians, felines, canines, equines, bovines, porcines, ovines, caprines, mammalian laboratory animals, mammalian farm animals, mammalian sport animals, and mammalian pets, are provided.
- the subject can be male or female and can be any suitable age, including infant, juvenile, adolescent, adult, and geriatric subjects.
- the subject to receive the treatment can be a patient, designating the fact that the subject has been identified as having a disorder of relevance to the treatment, or being at adequate risk of contracting the disorder.
- the subject is a human, such as a human patient.
- composition refers to any mixture of two or more products, substances, or compounds, including cells or antibodies. It may be a solution, a suspension, liquid, powder, a paste, aqueous, non-aqueous or any combination thereof.
- the preparation is generally in such form as to permit the biological activity of the active ingredient (e.g. antibody) to be effective.
- a “pharmaceutically acceptable carrier” refers to an ingredient in a pharmaceutical formulation, other than an active ingredient, which is nontoxic to a subject.
- a pharmaceutically acceptable carrier includes, but is not limited to, a buffer, excipient, stabilizer, or preservative.
- treatment refers to clinical intervention designed to alter the natural course of the individual or cell being treated during the course of clinical pathology. Desirable effects of treatment include decreasing the rate of disease progression, ameliorating or palliating the disease state, and remission or improved prognosis.
- An individual is successfully “treated”, for example, if one or more symptoms associated with a disorder (e.g., an eosinophil-mediated disease) are mitigated or eliminated.
- an individual is successfully “treated” if treatment results in increasing the quality of life of those suffering from a disease, decreasing the dose of other medications required for treating the disease, reducing the frequency of recurrence of the disease, lessening severity of the disease, delaying the development or progression of the disease, and/or prolonging survival of individuals.
- an “effective amount” refers to at least an amount effective, at dosages and for periods of time necessary, to achieve the desired or indicated effect, including a therapeutic or prophylactic result.
- An effective amount can be provided in one or more administrations.
- a “therapeutically effective amount” is at least the minimum dose of cells required to effect a measurable improvement of a particular disorder.
- a therapeutically effective amount is the amount of a composition that reduces the severity, the duration and/or the symptoms associated with cancer, viral infection, microbial infection, or septic shock in an animal.
- a therapeutically effective amount herein may vary according to factors such as the disease state, age, sex, and weight of the patient.
- a therapeutically effective amount may also be one in which any toxic or detrimental effects of the antibody are outweighed by the therapeutically beneficial effects.
- a “prophylactically effective amount” refers to an amount effective, at the dosages and for periods of time necessary, to achieve the desired prophylactic result. Typically but not necessarily, since a prophylactic dose is used in subjects prior to or at the earlier stage of disease, the prophylactically effective amount can be less than the therapeutically effective amount.
- a “disease” or “disorder” as used herein refers to a condition where treatment is needed and/or desired.
- tumor cell refers to a cell (or cells) exhibiting an uncontrolled growth and/or abnormal increased cell survival and/or inhibition of apoptosis which interferes with the normal functioning of bodily organs and systems. Included in this definition are benign and malignant cancers, polyps, hyperplasia, as well as dormant tumors or micrometastases .
- cancer and “tumor” encompass solid and hematological/lymphatic cancers and also encompass malignant, pre-malignant, and benign growth, such as dysplasia. Also, included in this definition are cells having abnormal proliferation that is not impeded (e.g. immune evasion and immune escape mechanisms) by the immune system (e.g. virus infected cells).
- immune evasion and immune escape mechanisms e.g. virus infected cells.
- Exemplary cancers include, but are not limited to: carcinoma, including that of the bladder, breast, colon, kidney, liver, lung, ovary, pancreas, stomach, cervix, thyroid and skin; including squamous cell carcinoma; hematopoietic tumors of lymphoid lineage, including leukemia, acute lymphocytic leukemia, acute lymphoblastic leukemia, B-cell lymphoma, T- cell lymphoma, Berketts lymphoma; hematopoietic tumors of myeloid lineage, including acute and chronic myelogenous leukemias and promyelocytic leukemia; tumors of mesenchymal origin, including fibrosarcoma and rhabdomyoscarcoma; other tumors, including melanoma, seminoma, tetratocarcinoma, neuroblastoma and glioma; tumors of the central and peripheral nervous system, including astrocytoma, neuroblastoma,
- cancers caused by aberrations in apoptosis would also be treated by the methods and compositions of the invention.
- Such cancers may include, but are not be limited to, follicular lymphomas, carcinomas with p53 mutations, hormone dependent tumors of the breast, prostate and ovary, and precancerous lesions such as familial adenomatous polyposis, and myelodysplastic syndromes, n specific
- malignancy or dysproliferative changes are treated or prevented by the methods and compositions of the invention in the ovary, bladder, breast, colon, lung, skin, pancreas, or uterus.
- sarcoma, melanoma, or leukemia is treated or prevented by the methods and compositions of the invention.
- combination refers to any association between or among two or more items.
- the combination can be two or more separate items, such as two compositions or two collections, can be a mixture thereof, such as a single mixture of the two or more items, or any variation thereof.
- the elements of a combination are generally functionally associated or related.
- kits are packaged combinations that optionally includes other elements, such as additional agents and instructions for use of the combination or elements thereof, for a purpose including, but not limited to, therapeutic uses.
- wild-type or “natural” or “native,” which are used interchangeably, as used herein is used in connection with biological materials such as nucleic acid molecules, proteins, host cells, and the like, that are found in nature and not modified by human intervention.
- Embodiment 1 An anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) antibody or antigen -binding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein: the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1), a heavy chain complementarity determining region 2 (CDR-H2), and a heavy chain complementarity determining region 3 (CDR-H3) contained within any one of SEQ ID NOs: 1-27 and 112, and the VL region comprises a light chain complementarity determining region 1 (CDR-L1), a light chain complementarity determining region 2 (CDR-L2) and a light chain complementarity determining region 3 (CDR- L3) contained within any one of SEQ ID NOs: 28-48 and 118, wherein the CDR-H1, CDR- H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3 are not set
- Embodiment 2 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 1, wherein: the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 1, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 3, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises
- Embodiment 3 An anti-CTLA-4 antibody or antigen-binding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein: the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1) comprising the sequence set forth in any one of SEQ ID NOs: 49-56, a heavy chain complementarity determining region 2 (CDR-H2) comprising the sequence set forth in any one of SEQ ID NOs: 57-67, and a heavy chain complementarity determining region 3 (CDR-H3) comprising the sequence set forth in any one of SEQ ID NOs: 68-73; and the VL region comprises a light chain complementarity determining region 1 (CDR-L1) comprising the sequence set forth in any one of SEQ ID NOs: 74-88, a light chain complementarity determining region 2 (CDR-L2) comprising the sequence set forth in any one of SEQ ID NOs: 89-97, and
- Embodiment 4 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-3, wherein the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1) comprising the sequence set forth in any one of SEQ ID NOs: 50-56, a heavy chain complementarity determining region 2 (CDR-H2) comprising the sequence set forth in any one of SEQ ID NOs: 58-67, and a heavy chain complementarity determining region 3 (CDR-H3) comprising the sequence set forth in any one of SEQ ID NOs: 69-73.
- CDR-H1 heavy chain complementarity determining region 1
- CDR-H2 heavy chain complementarity determining region 2
- CDR-H3 heavy chain complementarity determining region 3
- Embodiment 5 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiments 1-4, wherein the VL region comprises a light chain complementarity determining region 1 (CDR-L1) comprising the sequence set forth in any one of SEQ ID NOs: 75-88, a light chain complementarity determining region 2 (CDR-L2) comprising the sequence set forth in any one of SEQ ID NOs: 90-97, and a light chain complementarity determining region 3 (CDR-L3) comprising the sequence set forth in any one of SEQ ID NOs: 99-100.
- CDR-L1 light chain complementarity determining region 1
- CDR-L2 light chain complementarity determining region 2
- CDR-L3 light chain complementarity determining region 3
- Embodiment 6 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-5, wherein: the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 58, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 69, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1,
- Embodiment 7 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-6, wherein: the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 1, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 2, and the VL region is or comprises an amino acid
- Embodiment 8 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-7, wherein the VH region and the VL region are or comprise: the sequence set forth in SEQ ID NO: 1 and 118, respectively; the sequence set forth in SEQ ID NO: 2 and 118, respectively; the sequence set forth in SEQ ID NO: 3 and 118, respectively; the sequence set forth in SEQ ID NO: 4 and 118, respectively; the sequence set forth in SEQ ID NO: 5 and 118, respectively; the sequence set forth in SEQ ID NO: 6 and 118, respectively; the sequence set forth in SEQ ID NO: 7 and 118, respectively; the sequence set forth in SEQ ID NO: 8 and 118, respectively; the sequence set forth in SEQ ID NO: 9 and 118, respectively; the sequence set forth in SEQ ID NO: 10 and 118, respectively; the sequence set forth in SEQ ID NO: 11 and 118, respectively; the sequence set forth in SEQ ID NO: 12 and 118, respectively; the sequence set forth in
- Embodiment 9 An anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) antibody or antigen -binding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein the VH region and the VL region are or comprise: the sequence set forth in SEQ ID NO: 1 and 118, respectively; the sequence set forth in SEQ ID NO: 2 and 118, respectively; the sequence set forth in SEQ ID NO: 3 and 118, respectively; the sequence set forth in SEQ ID NO: 4 and 118, respectively; the sequence set forth in SEQ ID NO: 5 and 118, respectively; the sequence set forth in SEQ ID NO: 6 and 118, respectively; the sequence set forth in SEQ ID NO: 7 and 118, respectively; the sequence set forth in SEQ ID NO: 8 and 118, respectively; the sequence set forth in SEQ ID NO: 9 and 118, respectively; the sequence set forth in SEQ ID NO: 10 and 118, respectively; the sequence set forth in SEQ ID NO:
- Embodiment 10 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-9, wherein the VH region comprises L102H (histidine at position 98 by Kabat numbering).
- Embodiment 11 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10 wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
- the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10 wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
- Embodiment 13 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
- Embodiment 14 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- Embodiment 15 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- Embodiment 16 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
- Embodiment 17 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-16, wherein the antibody is a full length antibody.
- Embodiment 18 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 17, wherein the full length antibody comprises a constant region (Fc) from an IgAl, IgA2, IgD, IgE, IgGl, IgG2, IgG3, IgG4, or IgM.
- Fc constant region
- Embodiment 19 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 18, wherein the constant region is a human IgGl heavy chain constant region.
- Embodiment 20 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19, wherein the human IgGl heavy chain constant region comprises one or more of mutations selected from the group consisting of G236A, S239D, M252Y, S254T, T256E, A330L, I332E, M428L, and N434S, numbered according to the EU numbering system.
- Embodiment 21 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the human IgGl heavy chain constant region comprises S239D/A330L/I332E mutations, numbered according to the EU numbering system.
- Embodiment 22 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising S239D/I332E mutations, numbered according to the EU numbering system.
- Embodiment 23 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising G236A/S239D/A330L/I332E mutations, numbered according to the EU numbering system.
- Embodiment 24 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising G236A/A330L/I332E mutations, numbered according to the EU numbering system.
- Embodiment 26 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising M428L/N434S mutations, numbered according to the EU numbering system.
- Embodiment 27 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-16, wherein the antibody is an antigen-binding fragment.
- Embodiment 28 The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 27, wherein the antigen-binding fragment is selected from the group consisting of a single domain antibody, a single chain antibody, an unibody, a single chain variable fragment (scFv), a Fab fragment, and a F(ab')2 fragment.
- Embodiment 29 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-16, wherein the anti-CTLA-4 antibody or antigen-binding fragment thereof is recombinant.
- Embodiment 30 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-29, wherein the VH region and the VL region is human or is from a human protein.
- Embodiment 31 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-30, wherein the ratio of dissociation constant for binding at acidic pH (e.g., pH 5.0 to 6.0) to the dissociation constant for binding at physiological pH (e.g., pH 7.35 to 7.45) to human CTLA-4 protein is greater than 2.
- acidic pH e.g., pH 5.0 to 6.0
- physiological pH e.g., pH 7.35 to 7.45
- Embodiment 32 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-31, wherein the antibody or antigen-binding fragment exhibits a ratio of a dissociation constant for binding at acidic pH (e.g., pH 5.0 to 6.0) to dissociation constant for binding at physiological pH (e.g., pH 7.35 to 7.45) of more than 7.
- acidic pH e.g., pH 5.0 to 6.0
- physiological pH e.g., pH 7.35 to 7.45
- Embodiment 33 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-32, wherein the ratio of EC50 value of the antibody or antigenbinding fragment thereof for human CTLA-4 at physiological pH (e.g., pH 7.35 to 7.45) to EC50 value of the antibody or antigen-binding fragment thereof for human CTLA-4 at acidic pH (e.g., pH 5.0 to 6.0) is greater than 2.
- physiological pH e.g., pH 7.35 to 7.45
- acidic pH e.g., pH 5.0 to 6.0
- Embodiment 34 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-33, wherein the antibody or antigen-binding fragment thereof possesses pH dependent binding properties to human CTLA-4 that facilitates the dissociation of antibody from the antibody: CTLA-4 complex upon internalization into a cell, allowing either the antibody, CTLA-4, or both to be recycled instead of being degraded in the lysosome.
- Embodiment 35 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-34, wherein the antibody or antigen-binding fragment thereof preserves high affinity for CTLA-4 at pH>7, lower affinity at pH 5.5-7, and significantly reduced affinity for CTLA-4 at pH ⁇ 5.5.
- Embodiment 36 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-35, wherein the antibody or antigen-binding fragment thereof induces strong deletion of Treg and local T cell activation in tumor microenvironment but minimal systemic T cell activation.
- Embodiment 37 The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-36, wherein the antibody or antigen-binding fragment thereof induces a favorable ratio of Treg cells to T effector cells in the tumor microenvironment.
- Embodiment 38 A conjugate, comprising the anti-CTLA-4 antibody or antigenbinding fragment thereof of any of Embodiments 1-37 and a heterologous molecule or moiety.
- Embodiment 39 The conjugate of Embodiment 38, wherein the heterologous molecule or moiety is a cytotoxic agent, a drug, enzymatically active toxin or fragment thereof, or a radioactive atom.
- Embodiment 40 A bispecific antibody, comprising the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-37.
- Embodiment 41 A polynucleotide comprising a nucleic acid encoding the anti- CTLA-4 antibody or antigen-binding domain thereof of any of Embodiments 1-37.
- Embodiment 42 A vector, comprising the polynucleotide of Embodiment 41.
- Embodiment 43 The vector of Embodiment 42, wherein the vector is a viral vector.
- Embodiment 44 The vector of Embodiment 43, wherein the viral vector is a retroviral vector or a lentiviral vector.
- Embodiment 45 A cell comprising the polynucleotide of Embodiment 41, or the vector of any of Embodiments 42-44.
- Embodiment 46 The cell of Embodiment 45, wherein the cell is a mammalian cell.
- Embodiment 47 A method of producing an antibody comprising culturing the cell of Embodiment 45 or 46 under a condition that produces the antibody.
- Embodiment 48 The method of Embodiment 47, further comprising recovering the antibody produced by the cell.
- [0401] 49 An antibody or antigen-binding fragment thereof produced by the method of Embodiment 47 or Embodiment 48.
- Embodiment 50 A composition comprising the anti-CTLA-4 antibody or antigenbinding fragment thereof of any of Embodiments 1-37 and 49, the conjugate of Embodiment 38 or 39, the bispecific antibody of Embodiment 40.
- Embodiment 51 The composition of Embodiment 50, further comprising a pharmaceutically acceptable excipient.
- Embodiment 52 A method of stimulating an immune response in a subject, the method comprising administering to a subject in need thereof the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-37 and 49, the conjugate of Embodiment 38 or 39, the bispecific antibody of Embodiment 40 or the composition of Embodiment 50 or 51.
- Embodiment 53 The method of Embodiment 52, wherein stimulating an immune response treats a disease or condition in the subject.
- Embodiment 54 The method of Embodiment 53, wherein the disease or condition is a cancer.
- Embodiment 55 A method of cancer immunotherapy, the method comprising administering to a subject in need thereof a therapeutically effective amount of the anti-CTLA- 4 antibody or antigen-binding fragment thereof of any of Embodiments 1-37 and 49, the conjugate of Embodiment 38 or 39, the bispecific antibody of Embodiment 40 or the composition of Embodiment 50 or 51.
- Anti-CTLA-4 antibodies were computationally designed via a series of steps to identify novel antibodies with potentially desirable properties. The process included:
- Coding sequences for the computationally designed antibodies were recombinantly expressed in HEK293 cells. Briefly, the gene encoding the protein-of-interest was synthesized via homologous recombination amplification, and the target gene was inserted into the an expression vector via restriction enzyme digestion and ligation with nucleic acid encoding a constant heavy chain (SEQ ID NO: 101 or SEQ ID NO: 106) with a heavy chain signal peptide (SEQ ID NO: 104), and nucleic acid encoding a constant light chain (SEQ ID NO: 103) with a light chain signal peptide (SEQ ID NO: 105). The vectors were sequenced to confirm the correct insertion of the target gene.
- the vectors were transiently transfected into host cells and after culture, protein was purified from the culture supernatant via affinity chromatography or a combination of chromatography methods.
- the final purified product was subjected to buffer exchange into the storage buffer. Methods such as SDS-PAGE, UV (OD280), and SEC were used to assess the purity and quantity of the crude samples after each purification step as well as the final protein product.
- Table El lists sequence identifiers (SEQ ID NO:) corresponding to amino acid (aa) sequences of the variable heavy (Vn) chain and variable light (VL) chain of the designed antibodies, including complementarity determining regions (CDRs, by Kabat numbering) of each chain.
- the tested concentrations of antibodies ranged from 0.1152 ng/mL to 1800 ng/mL and results were compared to reference antibodies (e.g., Referencel, ipilimumab; Reference2, tremelimumab; Referenced, gotistobart; and Reference6, ipi.101).
- reference antibodies e.g., Referencel, ipilimumab; Reference2, tremelimumab; Referenced, gotistobart; and Reference6, ipi.101.
- FIGs. 1A-1S the computationally designed antibodies exhibited differences in binding relative to the reference antibodies and to each other at pH 5.0 and pH 7.4.
- the antibodies are antibodies that exhibit better binding at neutral pH than acidic pH.
- the ratio of binding at neutral pH (e.g., pH 7.4) than acidic pH (e.g., pH 5.0) was substantially increased compared to reference antibodies, while retaining similar or better binding at pH 7.4 than reference antibodies.
- antibodies with such a pH-dependent binding feature are enriched in antibodies that contain one or more of the following amino acids in the VH region: a histidine (H) at position 98, an arginine (R) at position 31, a glutamic acid (E) or a histidine (H) at position 53, a histidine (H) at position 58, and a histidine (H) at position 97, or any combination thereof; and/or in the VL region: a histidine (H) at position 27a, a histidine (H) at position 30, a histidine (H) at position 31, and a histidine (H) at position 90, or any combination thereof, each based on Kabat numbering.
- an antibody with higher activity at neutral pH (e.g., pH 7.4) than at acidic pH (e.g., pH 5.0) will exhibit less risk of toxic effect because the antibody can dissociate from CTLA-4 upon internalization of the antibody - CTLA-4 complex, thereby allowing CTLA-4 and the anti-CTLA-4 antibody to recycle to the cell surface.
- neutral pH e.g., pH 7.4
- acidic pH e.g., pH 5.0
- the antibodies are antibodies that exhibit better binding to the EcGRIIIA receptor than antibodies comprising an IgGl constant region domain.
- a significant contributor to the anti-tumor response of anti- CTLA-4 antibodies in the clinic involves depletion of Tregs via Fc gamma receptor engagement and mediated antibody-dependent cellular cytotoxicity & phagocytosis. Priming of the immune response via FcR engagement is also a suggested contributor to the mechanism of anti-CTLA4 antibodies in cancer treatment.
- the provided antibodies are expected to alter the ratio of effector T cells to regulatory T cells via Fc gamma receptor interaction in the tumor microenvironment in a favorable way.
- BLI Biolayer interferometry
- KD equilibrium dissociation constant
- k on association rate constant
- kdis dissociation rate constant
- BLI is an optical analytical technique that analyzes the interference pattern of white light reflected from two surfaces: a layer of immobilized protein on the biosensor tip, and an internal reference layer. Any change in the number of molecules bound to the biosensor tip causes a shift in the interference pattern that can be measured in real-time.
- Table E3A and Table E3B the computationally designed antibodies exhibited differences in kinetics relative to the reference antibody and to each other.
- ADCC antibody dependent cellular cytotoxicity
- the ratios tested included 0:1, 1:1, 2:1, 4:1, and 8:1. Based on the percentage of target cell lysis, the optimal E:T ratio of 2:1 was selected for further characterization. Purified NK cells and labeled target cells were co-incubated for 6 hours at the optimized E:T ratio of 2:1. Target cell lysis was measured by the reduction in CellTrace Far-Red fluorescence. The percentage of target cell lysis was evaluated at various concentrations of four different antibodies, with EC50 values calculated from dose-response curves. As depicted in FIG.
- CTLA-4- AB-68 (IgGl) showed slightly higher ADCC compared to Reference 1 (IgGl), and the isotype control antibody (IgGl) exhibited minimal ADCC activity.
- CTLA-4- AB-68.1 (IgGl S239D/A330L/I332E), an Fc variant of CTLA-4-AB-68 (IgGl), demonstrated substantially greater ADCC compared to the other molecules tested.
- anti-CTLA-4 antibodies were assessed using a syngeneic mouse model for colon cancer (CT26 tumor-bearing mice). This model was specifically selected because reference anti-human-CTLA-4 antibodies do not cross-react with murine CTLA-4, necessitating the use of human CTLA-4 (hCTLA-4) knock-in mice to accurately evaluate the therapeutic potential of the antibodies. These mice were developed on a BALB/c background, providing an immune-competent, genetically consistent environment that closely mimics the human immune response. The study utilized female hCTLA-4 knock- in BALB/c mice, aged 6-8 weeks and weighing approximately 18-22g.
- CT-26 tumor cells were cultured in vitro as a monolayer in RPML1640 medium supplemented with 10% heat-inactivated fetal bovine serum, 100 U/ml penicillin, and 100 pg/ml streptomycin. Cultures were maintained at 37°C in a 5% CO2 atmosphere and subcultured twice weekly using trypsin-EDTA treatment. Cells in the exponential growth phase were harvested and counted for tumor inoculation.
- each mouse was inoculated subcutaneously on the right lower flank with 3 x 10 A 5 CT-26 tumor cells in 0.1 mL of serum-free PBS. To ensure accurate subcutaneous delivery, the skin fold was lifted with one hand while the cells were injected with the other. To minimize tumor ulceration, a new syringe and needle were used for each mouse, and any droplets of cells were wiped from the skin with KimWipes postinjection. The animals were randomized, and treatment commenced when the average tumor volume reached 80 mm A 3. Treatment was administered intraperitoneally at a fixed mg/kg dose, with the dosing volume adjusted based on body weight (10 pL/g). Treatment was delivered bi-weekly for 3 weeks, totaling six doses, as shown in FIG. 5A.
- FIGs. 5B-5E Tumor volume results are summarized in FIGs. 5B-5E. While tumor growth progressed in all vehicle control-treated mice, treatment with CTLA-4- AB-68.1 led to complete tumor rejection in several mice and significantly slowed tumor growth in others. Notably, mice treated with CTLA-4- AB-68.1 appeared to demonstrate improved tumor rejection at increased doses and superior tumor rejection compared to those treated with Reference 1 at a low (1 mg/kg) dose.
- hCTLA4 knock-in mice bearing CT26 colorectal carcinoma tumors were prepared as described in Example 6.
- Multi-parameter flow cytometry assays were designed to evaluate the absolute cell numbers of various immune cell subsets in tumor and blood samples 5 days after a single 5 mg/kg dose treatment of Vehicle control, CTLA-4- AB-68.1, or Referencel. Each group included four mice. The animals were randomized, and treatment commenced when the average tumor volume reached 80 mm A 3. Treatment was administered intraperitoneally .
- FIG. 6C The results of the FCM analysis for tumor are presented in FIG. 6C, and the results of the FCM analysis for blood samples are shown in FIGs. 6D-6H, respectively.
- FIG. 6C cell numbers of CD45+, myeloid, T cells and T cell subpopulations per 100 mg tumor are shown.
- CD4Teff refers to CD4T cells other than Treg.
- CTLA-4- AB-68.1 treatment increased the ratio of CD4Teff/TReg and the ratio of CD8T/TReg cells in the tumor relative to the vehicle control and Reference 1. Additionally, while treatment with Reference 1 resulted in depleted levels of CTLA-4 expression on TReg cells in the blood compared to the vehicle control, treatment with CTLA- 4- AB-68.1 did not deplete CTLA-4 expression on TReg cells in the blood.
- the present invention is not intended to be limited in scope to the particular disclosed embodiments, which are provided, for example, to illustrate various aspects of the invention. Various modifications to the compositions and methods described will become apparent from the description and teachings herein. Such variations may be practiced without departing from the true scope and spirit of the disclosure and are intended to fall within the scope of the present disclosure.
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Immunology (AREA)
- Organic Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- General Health & Medical Sciences (AREA)
- Medicinal Chemistry (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Biochemistry (AREA)
- Animal Behavior & Ethology (AREA)
- General Chemical & Material Sciences (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Pharmacology & Pharmacy (AREA)
- Biophysics (AREA)
- Genetics & Genomics (AREA)
- Molecular Biology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Peptides Or Proteins (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
Abstract
Provided are novel monoclonal antibodies or antibody fragments against protein cytotoxic T-lymphocyte- associated protein 4 (CTLA-4). The antibodies or antibody fragments disclosed can be used to provide potent agents for the treatment of cancers via the modulation of human immune function.
Description
ANTI-CTLA-4 ANTIBODIES AND RELATED BINDING MOLECULES AND METHODS AND USES THEREOF
Cross-Reference to Related Applications
[0001] This application claims priority from U.S. provisional application No. 63/532,336 filed August 11, 2023, entitled, “ANTI-CTLA-4 ANTIBODIES AND RELATED BINDING MOLECULES AND METHODS AND USES THEREOF”, the contents of which is incorporated by reference in its entireties.
Incorporation by Reference of Sequence Listing
[0002] The present application is being filed along with a Sequence Listing in electronic format. The Sequence Listing is provided as a file entitled 281632000240.XML, created August 8, 2024, which is 169,088 bytes in size. The information in the electronic format of the Sequence Listing is incorporated by reference in its entirety.
Field
[0003] The present disclosure provides novel monoclonal antibodies or antibody fragments against protein cytotoxic T-lymphocyte- associated protein 4 (CTLA-4). CTLA-4 is a cell surface protein predominantly expressed on T cells and plays a crucial role in regulating immune responses. The novel antibodies or antibody fragments described may provide potent moieties for modulation of human immune function with potential therapeutic applications in various diseases, including cancer.
Background
[0004] Cancer therapies comprise a wide range of therapeutic approaches, including surgery, radiation, and chemotherapy. While the various approaches offer a broad selection of options for treating cancer, many therapeutics suffer from disadvantages, such as a lack of selectivity of targeting cancer cells over normal, healthy cells, and the development of resistance by the cancer to the treatment. Immunotherapy targeting checkpoint molecules has emerged as one of the most compelling approaches for treating patients with cancers, due to promising results from preclinical and clinical trials. Despite this promise, certain
immunotherapies, including CTLA-4 targeting antibodies, have shown limited efficacy in a substantial number of patients and have been associated with unwanted toxicities, including immune-related adverse events. Thus there exists a need for providing improved immunotherapies. The disclosure provided herein addresses this need.
Summary
[0005] In some embodiments, provided herein is an anti-cytotoxic T-lymphocyte- associated protein 4 (CTLA-4) antibody or antigen-binding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein: the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1), a heavy chain complementarity determining region 2 (CDR-H2), and a heavy chain complementarity determining region 3 (CDR-H3) contained within any one of SEQ ID NOs: 1-27 and 112, and the VL region comprises a light chain complementarity determining region 1 (CDR-L1), a light chain complementarity determining region 2 (CDR-L2) and a light chain complementarity determining region 3 (CDR-L3) contained within any one of SEQ ID NOs: 28-48 and 118, wherein the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3 are not set forth in SEQ ID NOS: 49, 57, 68, 74, 89 and 98, respectively.
[0006] In some embodiments, the VH region comprises a CDR-H1, a CDR-H2, and a CDR- H3 contained within SEQ ID NO: 1, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR- H2, and a CDR-H3 contained within SEQ ID NO: 2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 3, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:4, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:5, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:6, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:7, and the VL region comprises a CDR-L1, a CDR-L2,
and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR- H2, and a CDR-H3 contained within SEQ ID NO:8, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:9, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 10, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 11, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 12, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 13, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 14, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 15, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 16, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 17, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 18, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 19, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:20, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:21, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained
within SEQ ID NO:22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 contained within SEQ ID NO: 28; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 30; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 31; the Vnregion comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 32; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 33; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 34; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 35; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 36; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 37; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 38; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 40; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region
comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 42; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 43; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 44; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 45; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 46; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 23, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 24, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 25, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 26, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 27, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 23, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 24, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 25, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 26, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained
within SEQ ID NO: 47; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 27, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 23, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 24, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 25, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 26, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 27, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 23, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 24, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 25, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 26, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 27, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; or the VH region comprises a CDR-
Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 5, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 48.
[0007] In some embodiments, provided herein is an anti-CTLA-4 antibody or antigenbinding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein: the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1) comprising the sequence set forth in any one of SEQ ID NOs: 49-56, a heavy chain complementarity determining region 2 (CDR-H2) comprising the sequence set forth in any one of SEQ ID NOs: 57-67, and a heavy chain complementarity determining region 3 (CDR-H3) comprising the sequence set forth in any one of SEQ ID NOs: 68-73; and the VL region comprises a light chain complementarity determining region 1 (CDR- Ll) comprising the sequence set forth in any one of SEQ ID NOs: 74-88, a light chain complementarity determining region 2 (CDR-L2) comprising the sequence set forth in any one of SEQ ID NOs: 89-97, and a light chain complementarity determining region 3 (CDR-L3) comprising the sequence set forth in any one of SEQ ID NOs: 98-100, wherein the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3 are not set forth in SEQ ID NOS: 49, 57, 68, 74, 89 and 98, respectively.
[0008] In any of the embodiments herein, the VH region can comprise a heavy chain complementarity determining region 1 (CDR-H1) comprising the sequence set forth in any one of SEQ ID NOs: 50-56, a heavy chain complementarity determining region 2 (CDR-H2) comprising the sequence set forth in any one of SEQ ID NOs: 58-67, and a heavy chain complementarity determining region 3 (CDR-H3) comprising the sequence set forth in any one of SEQ ID NOs: 69-73.
[0009] In any of the embodiments herein, the VL region can comprise a light chain complementarity determining region 1 (CDR-L1) comprising the sequence set forth in any one of SEQ ID NOs: 75-88, a light chain complementarity determining region 2 (CDR-L2) comprising the sequence set forth in any one of SEQ ID NOs: 90-97, and a light chain complementarity determining region 3 (CDR-L3) comprising the sequence set forth in any one of SEQ ID NOs: 99-100.
[0010] In any of the embodiments herein, the VH region can comprise a CDR-H1, a CDR- H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 58, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise
a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 69, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 51, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 59, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 61, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 51, 62, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 63, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 57, and 69, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 53, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 53, 57, and 70, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise
a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 54, 57, and 71, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 55, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 56, 57, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 64, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 60, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 66, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 71, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 73, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region can comprise
a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 75, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 77, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 78, 90, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 79, 90, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 80, 91, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 78, 89, and 99, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 81, 92, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 82, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 82, 93, and 98, respectively; the VH region can comprise
a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 83, 92, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 100, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 86, 94, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 87, 95, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 96, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 88, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 97, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region can comprise
a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region can comprise
a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region can comprise
a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR- L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; or the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 88, 89, and 100, respectively.
[0011] In any of the embodiments herein, the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 1, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 2, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 3, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 4, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at
or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 5, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 6, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 7, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 8, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 9, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 10, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 11, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%,
91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 12, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 13, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 14, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 15, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 16, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 17, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 18, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or
99% identity to SEQ ID NO: 19, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 20, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 21, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 28; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 30; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or
can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 31; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 32; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 33; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 34; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 35; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 36; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 37; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%,
90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 38; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 40; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 42; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 43; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 44; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 45; the VH
region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 46; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%,
90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region can be or
can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%,
93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; or the VH region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 5, and the VL region can be or can comprise an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 48.
[0012] In any of the embodiments herein, the VH region and the VL region can be or can comprise: the sequence set forth in SEQ ID NO: 1 and 118, respectively; the sequence set forth in SEQ ID NO: 2 and 118, respectively; the sequence set forth in SEQ ID NO: 3 and 118, respectively; the sequence set forth in SEQ ID NO: 4 and 118, respectively; the sequence set forth in SEQ ID NO: 5 and 118, respectively; the sequence set forth in SEQ ID NO: 6 and 118, respectively; the sequence set forth in SEQ ID NO: 7 and 118, respectively; the sequence set forth in SEQ ID NO: 8 and 118, respectively; the sequence set forth in SEQ ID NO: 9 and 118, respectively; the sequence set forth in SEQ ID NO: 10 and 118, respectively; the sequence set forth in SEQ ID NO: 11 and 118, respectively; the sequence set forth in SEQ ID NO: 12 and 118, respectively; the sequence set forth in SEQ ID NO: 13 and 118, respectively; the sequence set forth in SEQ ID NO: 14 and 118, respectively; the sequence set forth in SEQ ID NO: 15 and 118, respectively; the sequence set forth in SEQ ID NO: 16 and 118, respectively; the sequence set forth in SEQ ID NO: 17 and 118, respectively; the sequence set forth in SEQ ID
NO: 18 and 118, respectively; the sequence set forth in SEQ ID NO: 19 and 118, respectively; the sequence set forth in SEQ ID NO: 20 and 118, respectively; the sequence set forth in SEQ ID NO: 21 and 118, respectively; the sequence set forth in SEQ ID NO: 22 and 118, respectively; the sequence set forth in SEQ ID NO: 112 and 28, respectively; the sequence set forth in SEQ ID NO: 112 and 29, respectively; the sequence set forth in SEQ ID NO: 112 and 30, respectively; the sequence set forth in SEQ ID NO: 112 and 31, respectively; the sequence set forth in SEQ ID NO: 112 and 32, respectively; the sequence set forth in SEQ ID NO: 112 and 33, respectively; the sequence set forth in SEQ ID NO: 112 and 34, respectively; the sequence set forth in SEQ ID NO: 112 and 35, respectively; the sequence set forth in SEQ ID NO: 112 and 36, respectively; the sequence set forth in SEQ ID NO: 112 and 37, respectively; the sequence set forth in SEQ ID NO: 112 and 38, respectively; the sequence set forth in SEQ ID NO: 112 and 39, respectively; the sequence set forth in SEQ ID NO: 112 and 40, respectively; the sequence set forth in SEQ ID NO: 112 and 41, respectively; the sequence set forth in SEQ ID NO: 112 and 42, respectively; the sequence set forth in SEQ ID NO: 112 and 43, respectively; the sequence set forth in SEQ ID NO: 112 and 44, respectively; the sequence set forth in SEQ ID NO: 112 and 45, respectively; the sequence set forth in SEQ ID NO: 112 and 46, respectively; the sequence set forth in SEQ ID NO: 23 and 41, respectively; the sequence set forth in SEQ ID NO: 24 and 41, respectively; the sequence set forth in SEQ ID NO: 25 and 41, respectively; the sequence set forth in SEQ ID NO: 26 and 41, respectively; the sequence set forth in SEQ ID NO: 22 and 41, respectively; the sequence set forth in SEQ ID NO: 27 and 41, respectively; the sequence set forth in SEQ ID NO: 23 and 47, respectively; the sequence set forth in SEQ ID NO: 24 and 47, respectively; the sequence set forth in SEQ ID NO: 25 and 47, respectively; the sequence set forth in SEQ ID NO: 26 and 47, respectively; the sequence set forth in SEQ ID NO: 22 and 47, respectively; the sequence set forth in SEQ ID NO: 27 and 47, respectively; the sequence set forth in SEQ ID NO: 23 and 29, respectively; the sequence set forth in SEQ ID NO: 24 and 29, respectively; the sequence set forth in SEQ ID NO: 25 and 29, respectively; the sequence set forth in SEQ ID NO: 26 and 29, respectively; the sequence set forth in SEQ ID NO: 22 and 29, respectively; the sequence set forth in SEQ ID NO: 27 and 29, respectively; the sequence set forth in SEQ ID NO: 23 and 39, respectively; the sequence set forth in SEQ ID NO: 24 and 39, respectively; the sequence set forth in SEQ ID NO: 25 and 39, respectively; the sequence set forth in SEQ ID NO: 26 and 39, respectively;
the sequence set forth in SEQ ID NO: 22 and 39, respectively; the sequence set forth in SEQ ID NO: 27 and 39, respectively; or the sequence set forth in SEQ ID NO: 5 and 48, respectively.
[0013] In some embodiments, provided herein is an anti-cytotoxic T-lymphocyte- associated protein 4 (CTLA-4) antibody or antigen-binding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein the VH region and the VL region can be or can comprise: the sequence set forth in SEQ ID NO: 1 and 118, respectively; the sequence set forth in SEQ ID NO: 2 and 118, respectively; the sequence set forth in SEQ ID NO: 3 and 118, respectively; the sequence set forth in SEQ ID NO: 4 and 118, respectively; the sequence set forth in SEQ ID NO: 5 and 118, respectively; the sequence set forth in SEQ ID NO: 6 and 118, respectively; the sequence set forth in SEQ ID NO: 7 and 118, respectively; the sequence set forth in SEQ ID NO: 8 and 118, respectively; the sequence set forth in SEQ ID NO: 9 and 118, respectively; the sequence set forth in SEQ ID NO: 10 and 118, respectively; the sequence set forth in SEQ ID NO: 11 and 118, respectively; the sequence set forth in SEQ ID NO: 12 and 118, respectively; the sequence set forth in SEQ ID NO: 13 and 118, respectively; the sequence set forth in SEQ ID NO: 14 and 118, respectively; the sequence set forth in SEQ ID NO: 15 and 118, respectively; the sequence set forth in SEQ ID NO: 16 and 118, respectively; the sequence set forth in SEQ ID NO: 17 and 118, respectively; the sequence set forth in SEQ ID NO: 18 and 118, respectively; the sequence set forth in SEQ ID NO: 19 and 118, respectively; the sequence set forth in SEQ ID NO: 20 and 118, respectively; the sequence set forth in SEQ ID NO: 21 and 118, respectively; the sequence set forth in SEQ ID NO: 22 and 118, respectively; the sequence set forth in SEQ ID NO: 112 and 28, respectively; the sequence set forth in SEQ ID NO: 112 and 29, respectively; the sequence set forth in SEQ ID NO: 112 and 30, respectively; the sequence set forth in SEQ ID NO: 112 and 31, respectively; the sequence set forth in SEQ ID NO: 112 and 32, respectively; the sequence set forth in SEQ ID NO: 112 and 33, respectively; the sequence set forth in SEQ ID NO: 112 and 34, respectively; the sequence set forth in SEQ ID NO: 112 and 35, respectively; the sequence set forth in SEQ ID NO: 112 and 36, respectively; the sequence set forth in SEQ ID NO: 112 and 37, respectively; the sequence set forth in SEQ ID NO: 112 and 38, respectively; the sequence set forth in SEQ ID NO: 112 and 39, respectively; the sequence set forth in SEQ ID NO: 112 and 40, respectively; the sequence set forth in SEQ ID NO: 112 and 41, respectively; the sequence set forth in SEQ ID NO: 112 and 42, respectively; the sequence set forth in SEQ ID NO: 112 and 43, respectively; the sequence set forth in SEQ ID NO: 112
and 44, respectively; the sequence set forth in SEQ ID NO: 112 and 45, respectively; the sequence set forth in SEQ ID NO: 112 and 46, respectively; the sequence set forth in SEQ ID NO: 23 and 41, respectively; the sequence set forth in SEQ ID NO: 24 and 41, respectively; the sequence set forth in SEQ ID NO: 25 and 41, respectively; the sequence set forth in SEQ ID NO: 26 and 41, respectively; the sequence set forth in SEQ ID NO: 22 and 41, respectively; the sequence set forth in SEQ ID NO: 27 and 41, respectively; the sequence set forth in SEQ ID NO: 23 and 47, respectively; the sequence set forth in SEQ ID NO: 24 and 47, respectively; the sequence set forth in SEQ ID NO: 25 and 47, respectively; the sequence set forth in SEQ ID NO: 26 and 47, respectively; the sequence set forth in SEQ ID NO: 22 and 47, respectively; the sequence set forth in SEQ ID NO: 27 and 47, respectively; the sequence set forth in SEQ ID NO: 23 and 29, respectively; the sequence set forth in SEQ ID NO: 24 and 29, respectively; the sequence set forth in SEQ ID NO: 25 and 29, respectively; the sequence set forth in SEQ ID NO: 26 and 29, respectively; the sequence set forth in SEQ ID NO: 22 and 29, respectively; the sequence set forth in SEQ ID NO: 27 and 29, respectively; the sequence set forth in SEQ ID NO: 23 and 39, respectively; the sequence set forth in SEQ ID NO: 24 and 39, respectively; the sequence set forth in SEQ ID NO: 25 and 39, respectively; the sequence set forth in SEQ ID NO: 26 and 39, respectively; the sequence set forth in SEQ ID NO: 22 and 39, respectively; the sequence set forth in SEQ ID NO: 27 and 39, respectively; or the sequence set forth in SEQ ID NO: 5 and 48, respectively.
[0014] In some embodiments, the VH region comprises one or more amino acid residues selected from the group consisting of: a histidine (H) at position 98, an arginine (R) at position 31, a glutamic acid (E) or a histidine (H) at position 53, a histidine (H) at position 58, and a histidine (H) at position 97, or any combination thereof, and/or the VL region comprises one or more amino acid residues selected from the group consisting of: a histidine (H) at position 27a, a histidine (H) at position 30, a histidine (H) at position 31, and a histidine (H) at position 90, or any combination thereof, where numbering of amino acid residues is by Kabat numbering. In some embodiments, the amino acid residues are mutations (amino acid substitutions) compared to a reference sequence. In any of the embodiments herein, the VH region can comprise one or more mutations selected from the group consisting of: S31R, D53H, D53E, Y58H, and W97H, and/or the VL region comprises one or more mutations selected from the group consisting of: S28H, S31H, S32H, and Q91H, wherein numbering of amino acid residues
is by Kabat numbering. In some embodiments, the reference sequence is a VH sequence set forth in SEQ ID NO: 112 and a VL sequence set forth in SEQ ID NO: 118.
[0015] In any of the some embodiments herein, In any of the embodiments herein, the VH region can comprise L102H (corresponding to histidine at position 98 by Kabat numbering).
[0016] In any of the embodiments herein, the VH region can comprise a CDR-H1, a CDR- H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively. In any of the embodiments herein, the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively. In any of the embodiments herein, the VH region can comprise a CDR- Hl, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively. In any of the embodiments herein, the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively. In any of the embodiments herein, the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively. In any of the embodiments herein, the VH region can comprise a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region can comprise a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0017] In any of the embodiments herein, the antibody can be a full length antibody. In some embodiments, the full length antibody comprises a constant region (Fc) from an IgAl, IgA2, IgD, IgE, IgGl, IgG2, IgG3, IgG4, or IgM. In some embodiments, the constant region is a human IgGl heavy chain constant region. In some embodiments, the human IgGl heavy chain constant region comprises one or more of mutations selected from the group consisting of G236A, S239D, M252Y, S254T, T256E, A330L, I332E, M428L, and N434S, numbered 1
according to the EU numbering system. In any of the embodiments herein, the human IgGl heavy chain constant region can comprise S239D/A330L/I332E mutations, numbered according to the EU numbering system. In any of the embodiments herein, the antibody region can comprise a human IgGl heavy chain constant region comprising S239D/I332E mutations, numbered according to the EU numbering system. In any of the embodiments herein, the antibody region can comprise a human IgGl heavy chain constant region comprising G236A/S239D/A330L/I332E mutations, numbered according to the EU numbering system. In any of the embodiments herein, the antibody region can comprise a human IgGl heavy chain constant region comprising G236A/A330L/I332E mutations, numbered according to the EU numbering system. In any of the embodiments herein, the antibody region can comprise a human IgGl heavy chain constant region comprising M252Y/S254T/T256E mutations, numbered according to the EU numbering system. In any of the embodiments herein, the antibody region can comprise a human IgGl heavy chain constant region comprising M428L/N434S mutations, numbered according to the EU numbering system.
[0018] In any of the embodiments herein, the antibody can be an antigen-binding fragment. In some embodiments, the antigen-binding fragment is selected from the group consisting of a single domain antibody, a single chain antibody, an unibody, a single chain variable fragment (scFv), a Fab fragment, and a F(ab')2 fragment.
[0019] In any of the embodiments herein, the anti-CTLA-4 antibody or antigen-binding fragment thereof can be recombinant. In any of the embodiments herein, the VH region and the VL region can be human or can be from a human protein.
[0020] In any of the embodiments herein, the ratio of dissociation constant for binding at acidic pH (e.g., pH 5.0 to 6.0) to the dissociation constant for binding at physiological pH (e.g., pH 7.35 to 7.45) to human CTLA-4 protein can be greater than 2. In any of the embodiments herein, the antibody or antigen-binding fragment can exhibit a ratio of a dissociation constant for binding at acidic pH (e.g., pH 5.0 to 6.0) to dissociation constant for binding at physiological pH (e.g., pH 7.35 to 7.45) of more than 7. In any of the embodiments herein, the ratio of EC50 value of the antibody or antigen-binding fragment thereof for human CTLA-4 at physiological pH (e.g., pH 7.35 to 7.45) to EC50 value of the antibody or antigen-binding fragment thereof for human CTLA-4 at acidic pH (e.g., pH 5.0 to 6.0) can be greater than 2. In any of the embodiments herein, the antibody or antigen-binding fragment thereof can possess pH dependent binding properties to human CTLA-4 that facilitates the dissociation of antibody
from the antibody :CTLA-4 complex upon internalization into a cell, allowing either the antibody, CTLA-4, or both to be recycled instead of being degraded in the lysosome. In any of the embodiments herein, the antibody or antigen-binding fragment thereof can preserve high affinity for CTLA-4 at pH>7, lower affinity at pH 5.5-7, and significantly reduced affinity for CTLA-4 at pH<5.5. In any of the embodiments herein, the antibody or antigen-binding fragment thereof can induce strong deletion of Treg and local T cell activation in tumor microenvironment but minimal systemic T cell activation. In any of the embodiments herein, the antibody or antigen-binding fragment thereof can induce a favorable ratio of Treg cells to T effector cells in the tumor microenvironment. In any of the embodiments herein, the antibody or antigen-binding fragment thereof can induce a favorable ratio of immune cell subsets in the tumor or blood. In any of the embodiments herein, the antibody or antigen-binding fragment can induce a favorable ratio of CTLA-4 expression on immune cell subsets in the tumor or blood. In some embodiments, the immune cell subsets comprise myeloid cells, T cells, CD4+ T cells, CD8+ T cells, and/or Treg cells.
[0021] In some embodiments, provided herein is a conjugate, comprising the anti-CTLA- 4 antibody or antigen-binding fragment thereof of any of the embodiments herein and a heterologous molecule or moiety. In some embodiments, the heterologous molecule or moiety is a cytotoxic agent, a drug, enzymatically active toxin or fragment thereof, or a radioactive atom.
[0022] In some embodiments, provided herein is a bispecific antibody, comprising the anti- CTLA-4 antibody or antigen-binding fragment thereof of any of the embodiments herein.
[0023] In some embodiments, provided herein is a polynucleotide comprising a nucleic acid encoding the anti-CTLA-4 antibody or antigen-binding domain thereof of any of the embodiments herein.
[0024] In some embodiments, provided herein is a vector, comprising the polynucleotide of any of the embodiments herein. In some embodiments, the vector is a viral vector. In some embodiments, the viral vector is a retroviral vector or a lentiviral vector.
[0025] In some embodiments, provided herein is a cell comprising the polynucleotide of any of the embodiments herein, or the vector of any of the embodiments herein. In some embodiments, the cell is a mammalian cell.
[0026] In some embodiments, provided herein is a method of producing an antibody comprising culturing the cell of any of the embodiments herein under a condition that produces
the antibody. In some embodiments, the method further comprises recovering the antibody produced by the cell.
[0027] In some embodiments, provided herein is an antibody or antigen-binding fragment thereof produced by the method of any of the embodiments herein.
[0028] In some embodiments, provided herein is a composition comprising the anti-CTLA- 4 antibody or antigen-binding fragment thereof of any of the embodiments herein, the conjugate of any of the embodiments herein, the bispecific antibody of any of the embodiments herein. In some embodiments, the composition further comprises a pharmaceutically acceptable excipient.
[0029] In some embodiments, provided herein is a method of stimulating an immune response in a subject, the method comprising administering to a subject in need thereof the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of the embodiments herein, the conjugate of any of the embodiments herein, the bispecific antibody of any of the embodiments herein or the composition of any of the embodiments herein. In some embodiments, stimulating an immune response treats a disease or condition in the subject.
[0030] In some embodiments, the disease or condition is a cancer.
[0031] In some embodiments, provided herein is a method of cancer immunotherapy, the method comprising administering to a subject in need thereof a therapeutically effective amount of the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of the embodiments herein, the conjugate of any of the embodiments herein, the bispecific antibody of any of the embodiments herein or the composition of any of the embodiments herein. In some embodiments, the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of the embodiments herein, the conjugate of any of the embodiments herein, the bispecific antibody of any of the embodiments herein or the composition of any of the embodiments herein is administered to the subject in need thereof at about 0.01 pg/kg body weight to about 20 mg/kg body weight.
[0032] In some embodiments, the method further comprises administering to the subject an effective amount of at least one additional therapeutic agent. In some embodiments, the at least one additional therapeutic agent is selected from the group consisting of: viral gene therapy, immune checkpoint inhibitors, target therapies, radiation therapies, vaccination therapies, and chemotherapies. In some embodiments, the at least one additional therapeutic agent is selected from the group consisting of: pomalyst, revlimid, lenalidomide,
pomalidomide, thalidomide, a DNA-alkylating platinum-containing derivative, cisplatin, 5- fluorouracil, cyclophosphamide, an anti-CD137 antibody, an anti-PD-1 antibody, an anti-PD- L1 antibody, an anti-CD20 antibody, an anti-CD40 antibody, an anti-DR5 antibody, an antiCD Id antibody, an anti-TIM3 antibody, an anti-SLAMF7 antibody, an anti-KIR receptor antibody, an anti-OX40 antibody, an anti-HER2 antibody, an anti-ErbB-2 antibody, an anti- EGFR antibody, cetuximab, rituximab, trastuzumab, pembrolizumab, an antibody drug conjugation, radiotherapy, single dose radiation, fractionated radiation, focal radiation, whole organ radiation, IL- 12, IFNa, GM-CSF, a chimeric antigen receptor, adoptively transferred T cells, an anti-cancer vaccine, and an oncolytic virus. In some embodiments, the method comprises administering to the subject an effective amount of the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of the embodiments herein, the conjugate of any of the embodiments herein, the bispecific antibody of any of the embodiments herein or the composition of any of the embodiments herein prior to a surgery or after a surgery to remove a solid tumor in the subject.
Brief Description of the Drawings
[0033] FIGs. 1A-S show results of binding tests of exemplary provided antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. 1A shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-7, CTLA-4- AB-9, CTLA-4- AB- 10, and CTLA-4- AB- 11) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. IB shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB- 13, CTLA-4- AB- 14, CTLA-4-AB-15, and CTLA-4-AB-16) and reference antibodies for binding to human CTLA- 4 at pH 5 and at pH 7.4. FIG. 1C shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB- 17, CTLA-4-AB-18, CTLA-4-AB-19, and CTLA-4-AB-20) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. ID shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB -21, CTLA-4- AB-22, CTLA-4-AB-23, and CTLA-4- AB-24) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. IE shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-25, CTLA-4-AB-26, CTLA-4- AB-27, and CTLA-4- AB-28) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. IF shows results of binding tests of anti-CTLA- 4 antibodies (CTLA-4- AB-29, CTLA-4- AB-30, CTLA-4-AB-31, and CTLA-4- AB-32) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. 1G shows
results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-34, CTLA-4- AB-35, CTLA- 4-AB-36, CTLA-4-AB-37, CTLA-4-AB-38 and CTLA-4-AB-39) for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. 1H shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB -42, CTLA-4-AB-43, CTLA-4-AB-44, and CTLA-4-AB-45) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. II shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-8, CTLA-4- AB- 12, CTLA-4- AB-33, and CTLA-4- AB-46) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. 1J shows results of binding tests of anti-CTLA-4 antibody CTLA-4- AB- 47 and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. IK shows results of binding tests of anti-CTLA-4 antibody CTLA-4-AB-48 and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. IL shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-49, CTLA-4- AB-50, and CTLA-4- AB- 51) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. IM shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-52, CTLA-4- AB-53, and CTLA-4- AB-54) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. IN shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4-AB-55, CTLA-4- AB-56, and CTLA-4- AB-57) and reference antibodies for binding to human CTLA- 4 at pH 5 and at pH 7.4. FIG. IO shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB-58, CTLA-4- AB-59, and CTLA-4- AB-60) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. IP shows results of binding tests of anti-CTLA- 4 antibodies and reference antibodies (CTLA-4- AB -61, CTLA-4- AB-62, and CTLA-4- AB-63) for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. IQ shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB -64, CTLA-4-AB-65, and CTLA-4-AB-66) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. 1R shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4-AB-67, CTLA-4-AB-68, and CTLA-4- AB -69) and reference antibodies for binding to human CTLA-4 at pH 5 and at pH 7.4. FIG. IS shows results of binding tests of anti-CTLA-4 antibodies (CTLA-4- AB -70, CTLA-4- AB -71, and CTLA-4- AB -72) and reference antibodies for binding to human CTLA- 4 at pH 5 and at pH 7.4.
[0034] FIGs. 2A-2C show results of binding tests of exemplary provided antibodies for binding to human CTLA-4 at different pH’s. FIG. 2A shows results of binding tests of a reference antibody for binding to human CTLA-4 at pH 5.5, pH 6, pH 6.5, pH 7 and pH 7.4.
FIG. 2B shows results of binding tests of CTLA-4-AB-62 for binding to human CTLA-4 at pH 5.5, pH 6, pH 6.5, pH 7 and pH 7.4. FIG. 2C shows results of binding tests of CTLA-4- AB-68 for binding to human CTLA-4 at pH 5.5, pH 6, pH 6.5, pH 7 and pH 7.4.
[0035] FIGs. 3A-3C show effects of anti-CTLA-4 antibodies on CTLA-4 expression in treated human-CTLA-4 expressing Cells. FIG. 3A shows hCTLA-4 expression levels in cells treated with anti-CTLA-4 antibodies. FIG. 3B shows a schematic drawing of the workflow of confocal microscopy. FIG. 3C shows results of confocal microscopy. HEK293-hCTLA-4 cells were incubated with lOpg/mL antibody
[0036] FIGs. 4A-4B show effect of anti-CTLA-4 antibodies on antibody dependent cellular cytotoxicity (ADCC). FIG. 4A shows selection of NK cells. FIG. 4B shows target cell lysis was evaluated at various concentrations of four different antibodies and EC50 values calculated from dose-response curves.
[0037] FIGs. 5A-5E show in vivo anti-tumor efficacy of anti-CTLA-4 antibodies. FIG. 5A shows a schematic drawing of the experimental procedure. FIG. 5B shows tumor volume results of the vehicle, biweekly for 3 weeks, n=8. FIG. 5C shows tumor volume results of reference 1 at 1 mg/kg, biweekly for 3 weeks, n=8. FIG. 5D shows tumor volume results of CTLA-4- AB-68.1 at 1 mg/kg, biweekly for 3 weeks, n=8. FIG. 5E shows tumor volume results of CTLA-4- AB-68.1 at 10 mg/kg, biweekly for 3 weeks, n=8.
[0038] FIGs. 6A-6E show effect of anti-CTLA-4 antibodies on immune cell subsets in the tumor and blood of hCTLA4 knock-in mice. FIG. 6A shows gating strategy for tumor samples to analyze This panel was designed to analyze absolute cell numbers of CD45+, Myeloid , T cells, CD4 T, CD8 T and Treg in tumor of CT26 model in hCTLA4 knock-in mice. FIG. 6B shows gating strategy for blood samples to analyze absolute cell numbers of CD45+, Myeloid, T cells, CD4 T, CD8 T and Treg in blood of CT26 model in hCTLA4 knock-in mice, and the expression level of hCTLA-4 on Treg cells in the blood of CT26 model in hCTLA4 knock-in mice. FIG. 6C shows results of the FCM analysis for tumor samples. One-way ANOVA was used for statistical analysis. When a significant F-statistics (a ratio of treatment variance to the error variance) was obtained, comparisons between group 1 and other groups were carried out with Games-Howell (equal variances not assumed) or Tukey (equal variances assumed) test. * p<0.05; ** p<0.01; *** p<0.001. Error bars represented Standard Error of Mean (SEM). FIG. 6D shows results of the FCM analysis for blood samples showing percentages of myeloid, T cells and T cell subpopulations in CD45+ cells. FIG. 6E shows results of the FCM analysis for
blood samples showing percentages of T cell subpopulations in T cells. FIG. 6F shows results of the FCM analysis for blood samples showing cell numbers of CD45+, myeloid, T cells and T cell subpopulations per 100 pL blood. FIG. 6G shows results of the FCM analysis for blood samples showing ratios between T cell subpopulations. CD4Teff refers to CD4T cells other than Treg. FIG. 6H shows MFI of hCTLA4 of Treg cells. One-way ANOVA was used for statistical analysis. When a significant F-statistics (a ratio of treatment variance to the error variance) was obtained, comparisons between group 1 and other groups were carried out with Games-Howell (equal variances not assumed) or Tukey (equal variances assumed) test. * p<0.05; ** p<0.01; *** p<0.001. Error bars represented Standard Error of Mean (SEM).
Detailed Description
[0039] The present disclosure concerns anti-CTLA-4 antibodies or antibody fragments (i.e. antigen binding fragment that is able to bind CTLA-4) suitable for therapeutic use. Relative to existing known therapies, some of these anti-CTLA-4 antibodies or antibody fragments confer specific binding properties to human CTLA-4 under different pH conditions. In some embodiments, the disclosed anti-CTLA-4 antibodies or antibody fragments also exhibit physiochemical or biological properties that lead to improved efficacy and safety to anti- CTLA-4 antibodies known in the art. In some embodiments, the provided anti-CTLA-4 antibody, including any antigen binding antibody fragments, can be used to treat cancer.
[0040] Cancer is a disease state of genomic instability caused by the accumulation of point mutations, resulting in behavioral and structural alterations to disaffected cells. Cancer presents in the form of abnormal growths defined by the proliferation of morphologically ambiguous cells that have the potential to invade both proximal and distant tissues, usually in the form of solid state tumors (Hassanpour SH, Dehghani M. Review of cancer from perspective of molecular. Journal of Cancer Research and. Practice. July 2017. doi:10.1016/j.jcrpr.2017.07.001). Consequently, these tumors severely impact local and global systems in the body, leading to premature death and severe decrease in quality of life, and the increasing prevalence of cancer has spurred significant interest in identifying treatments to reduce or eliminate tumors (Global Burden of Disease 2019 Cancer Collaboration, Kocamik JM, Compton K, et al. Cancer Incidence, Mortality, Years of Life Lost, Years Lived With Disability, and Disability- Adjusted Life Years for 29 Cancer Groups
From 2010 to 2019: A Systematic Analysis for the Global Burden of Disease Study 2019. JAMA Oncol. 2022;8(3):420-444.).
[0041] Historically, tumors have been treated with a plethora of therapeutic approaches, including chemotherapy, radiation, and surgery (Knezevic CE, Clarke W. Cancer chemotherapy: the case for therapeutic drug monitoring. Ther Drug Monit. 2020;42(l):6-19; Baskar R, Lee KA, Yeo R, Yeoh K-W. Cancer and radiation therapy: current advances and future directions. Int J Med Sci. 2012;9(3): 193-199; Sanai N, Berger MS. Surgical oncology for gliomas: the state of the art. Nat Rev Clin Oncol. 2018; 15(2): 112- 125). While their efficacy has improved over time, these strategies have limitations with respect to lack of specificity for targeting cancer cells, development of resistance to treatment, and debilitating side-effects that significantly impact the patient’s quality of life (Gyanani V, Haley JC, Goswami R. Challenges of Current Anticancer Treatment Approaches with Focus on Liposomal Drug Delivery Systems. Pharmaceuticals (Basel). 2021; 14(9)). Hence, there is considerable interest in developing mechanisms for targeted therapy, in which cancer cells can be targeted with minimal collateral damage to healthy cells.
[0042] The immune system is responsible for defending the body against foreign substances, both internal and external , such as pathogens and abnormal cells (Daeron M. The immune system as a system of relations. Front Immunol. 2022; 13:984678). Composed of innate and adaptive immunity, the immune system has impressive flexibility when identifying disease-causing agents. Innate immune cells provide preconfigured responses to broad situations and stimuli (Kaur BP, Secord E. Innate Immunity. Immunol Allergy Clin North Am. 2021;41(4):535-541), and adaptive immune cells provide more tailored responses to each stimulus through learning how to distinguish between previously encountered and new stimuli (Geckin B, Konstantin Fdhse F, Dominguez-Andres J, Netea MG. Trained immunity: implications for vaccination. Curr Opin Immunol. 2022;77: 102190). Because of its capacity to be “trained” to eliminate entities deemed harmful, the adaptive immune system has potential to be utilized for targeted cancer therapy, and the emergence of therapies such as CAR-T serves as compelling evidence of this strategy (Baulu E, Gardet C, Chuvin N, Depil S. TCR- engineered T cell therapy in solid tumors: State of the art and perspectives. Sci Adv. 2023;9(7):eadf3700).
[0043] Central to the immune response are T cells, a type of white blood cell, which act as key orchestrators of immune reactions (Kumar BV, Connors TJ, Farber DL. Human T Cell
Development, Localization, and Function throughout Life. Immunity. 2018;48(2):202-213). Starting as a hematopoietic stem cell, T cells mature into a variety of differentiated forms that can directly kill threats, recruit other immune cells to assist in the containment and eradication of threats, or mitigate excessive and indiscriminate immune activity that targets otherwise healthy and harmless cells (Shah K, Al-Haidari A, Sun J, Kazi JU. T cell receptor (TCR) signaling in health and disease. Signal Transduct Target Ther. 2021;6(l):412). A unique feature of T cells is their ability to discriminate between healthy and abnormal surface markers, with abnormal marker detection eliciting activation of T cells (Feinerman O, Germain RN, Altan-Bonnet G. Quantitative challenges in understanding ligand discrimination by alphabeta T cells. Mol Immunol. 2008;45(3):619-631).
[0044] T cell activation is tightly regulated to prevent excessive immune responses that could lead to tissue damage and autoimmune disorders. CTLA-4, a protein predominantly expressed on the surface of activated T cells, serves as an important checkpoint molecule for downregulating immune responses.
[0045] CTLA-4 acts by binding to its ligands, CD80 (B7-1) and CD86 (B7-2), on antigen- presenting cells (APCs) (Berg M, Zavazava N. Regulation of CD28 expression on CD8+ T cells by CTLA-4. J Leukoc Biol. 2008;83(4):853-863), thereby attenuating T cell activation and proliferation. The engagement of CTLA-4 with its ligands delivers inhibitory signals to T cells, leading to suppression of immune reactions (Alegre ML, Frauwirth KA, Thompson CB. T-cell regulation by CD28 and CTLA-4. Nat Rev Immunol. 2001;l(3):220-228). Dysregulation of the CTLA-4 pathway has been implicated in several pathological conditions, including autoimmune diseases as well as cancer, where tumor cells can exploit the immune system’s regulatory mechanisms to evade destruction (Hossen MM, Ma Y, Yin Z, et al. Current understanding of CTLA-4: from mechanism to autoimmune diseases. Front Immunol. 2023; 14: 1198365).
[0046] Mechanistically, CTLA-4 competes with CD28 to bind at CD80/86, with CTLA-4 having preferential binding with CD80/86 relative to CD28 (Krummel MF, Allison JP. CD28 and CTLA-4 have opposing effects on the response of T cells to stimulation. J Exp Med. 1995;182(2):459-465). However, under conditions favoring CD28 binding to CD80/86, a cascade of signaling events leads to the activation and proliferation of naive T-cells in conjunction with stimulation of T cell receptor (TCR) signaling by major histocompability complex II (MHC-II) on APCs, resulting in biological events at the immunological synapse,
downstream phosphorylation and post-translational modifications, transcriptional changes, and cytoskeletal remodeling (Frauwirth KA, Riley JL, Harris MH, et al. The CD28 signaling pathway regulates glucose metabolism. Immunity. 2002;16(6):769-777). Upon TCR stimulation, CTLA-4 circulates to the cell surface, but is then internalized by unphosphorylated cytoplasmic domains and either recycled to the plasma membrane or subjected to lysosomal degradation (Qureshi OS, Kaur S, Hou TZ, et al. Constitutive clathrin-mediated endocytosis of CTLA-4 persists during T cell activation. J Biol Chem. 2012;287( 12):9429-9440). Taken together, these findings support a balance between achieving sufficient effector T cell activation to achieve optimal cancer immunotherapeutic effect (CITE) while maintaining sufficient regulatory T cell (Treg) function to mitigate excessive effector T cell activity, which leads to a side effect known as an immune-related adverse event (irAE) (Du X, Liu M, Su J, et al. Uncoupling therapeutic from immunotherapy-related adverse effects for safer and effective anti-CTLA-4 antibodies in CTLA4 humanized mice. Cell Res. 2018;28(4):433-447; Ramos- Casals M, Brahmer JR, Callahan MK, et al. Immune-related adverse events of checkpoint inhibitors. Nat Rev Dis Primers. 2020;6(l):38).
[0047] Cancer immunotherapies utilizing surface proteins like CTLA-4 are known as immune checkpoint inhibitors (ICIs), and they serve to induce an antitumor immune response by blocking immune checkpoints that downregulate intratumoral T-cell responses (Ramos- Casals M, Brahmer JR, Callahan MK, et al. Immune-related adverse events of checkpoint inhibitors. Nat Rev Dis Primers. 2020;6(l ):38). Proof-of-concept experiments supporting anti- CTLA-4 therapy were initially described by James Allison’s research group, which discovered that blockade of CTLA-4 on effector and Treg compartments led to tumor size reduction (Peggs KS, Quezada SA, Chambers CA, Korman AJ, Allison JP. Blockade of CTLA-4 on both effector and regulatory T cell compartments contributes to the antitumor activity of anti-CTLA-4 antibodies. J Exp Med. 2009;206(8):1717-1725). In recent years, studies have highlighted the role that soluble CTLA-4 (sCTLA4) plays in cancer progression (Ward EJ, Dahal LN, Wijesekera SK, et al. The soluble isoform of CTLA-4 as a regulator of T-cell responses. Eur J Immunol. 2013;43(5): 1274-1285), particularly sCTLA4’s link to improved growth/survival of cells grown in the presence of primary blood mononuclear cells, suggesting an immunomodulatory effect of CTLA-4 (Kennedy PT, Saulters EL, Duckworth AD, et al. Soluble CTLA-4 attenuates T cell activation and modulates anti-tumor immunity. Mol Ther. 2024;32(2):457-468).
[0048] Based on promising preclinical tumor model studies, the clinical potential of antibodies against CTLA-4 has been explored in different human malignancies. Although anti- CTLA-4 (Ipilimumab, marketed as Yervoy) and Tremelimumab (marketed as Imjudo) have demonstrated efficacy in treating solid tumors such as melanoma, treatment and targeting of CTLA-4 is associated with autoimmune like toxicities (Sharma P, Goswami S, Raychaudhuri D, et al. Immune checkpoint therapy-current perspectives and future directions. Cell. 2023; 186(8): 1652- 1669). Characteristic side effects from inhibition of CTLA-4 are generally called immune-related adverse events (irAEs) and the most common irAEs are skin rash, hepatitis, colitis and endocrinopathies, particularly hypopituitarism (Liu Y, Zheng P. Preserving the CTLA-4 Checkpoint for Safer and More Effective Cancer Immunotherapy. Trends Pharmacol Sci. 2020;41(l):4-12).
[0049] A key challenge in developing therapeutic antibodies against CTLA-4 is to uncouple their desirable effects from their unwanted autoimmune side effects. Without wishing to be bound by any theory, CTLA-4 is recycled between the plasma membrane and endosomes by binding to lipopolysaccharide-responsive and beige-like anchor protein (LRBA). Antibodies that bind CTLA-4 disrupt the recycling process. Consequently, CTLA-4 is systemically directed to lysosomal degradation, and autoimmunity-related adverse effects are developed due to the unstoppable action of cytotoxic T cells. Increasingly, it is recognized that complete loss of CTLA-4 recycling leads to increased irAEs, as the immune system engages in off-target actions that disaffect otherwise healthy cells. Furthermore, current anti-CTLA antibodies fail to sufficiently engage with signaling pathways that enhance effector T cell function, leading to decreased immune response to cancer cells. Strategies to improve CITE while reducing irAEs point to designing therapies that improve both CTLA-4 recycling and antibody affinity for receptors that enhance effector T cell activation. Therefore, there is a desire to improve the therapeutic potential of anti-CTLA-4 antibodies by increasing efficacy while reducing the associated irAEs.
[0050] Recently, engineered next-generation anti-CTLA-4 antibodies have been reported with improved therapeutic characteristics (Zhang Y, Du X, Liu M, et al. Hijacking antibody- induced CTLA-4 lysosomal degradation for safer and more effective cancer immunotherapy. Cell Res. 2019;29(8):609-627). Antibodies are central components of the immune system that help prevent and control pathogen spread while providing long-term protection from reinfection. They are composed of two identical fragment antigen binding (Fab) domains that
are coupled through a hinge to a fragment crystallizable (Fc) region. The Fabs provide specificity and mediate target antigen binding, while the Fc region interacts with Fcgamma Receptors (FcgR), a class of receptors that deliver either inhibitory or activatory signals to modulate immune cell responses to regulate immune cell activity (Liu R, Oldham RJ, Teal E, Beers SA, Cragg MS. Fc-Engineering for Modulated Effector Functions-Improving Antibodies for Cancer Treatment. Antibodies (Basel). 2020;9(4)). Engagement of these receptors drives antibody effector functions including antibody-dependent cellular cytotoxicity (ADCC), a critical feature underlying the efficacy of anti-CTLA-4 in solid tumors that involves the release of cytotoxic granules by natural killer cells, granulocytes, and myeloid cells to kill target cells (Galvez-Cancino F, Simpson AP, Costoya C, et al. Fey receptors and immunomodulatory antibodies in cancer. Nat Rev Cancer. 2024;24(l):51-71). An example of these beneficial Fc enhancements is afucosylation, a process in which oligosaccharides of the Fc region are removed of focuses - a sugar molecule found in the Fc region of antibodies - that enhances antibody Fc-FcgR interactions, improving T-cell signaling and function (Shinkawa T, Nakamura K, Yamane N, et al. The absence of fucose but not the presence of galactose or bisecting N- acetylgluco s amine of human IgGl complextype oligosaccharides shows the critical role of enhancing antibody-dependent cellular cytotoxicity. J Biol Chem. 2003;278(5):3466-3473). Supporting this finding, a recent study showed that modification of anti-CTLA-4 antibody Fc regions led to improved CITE, resulting in improved T cell-mediated targeting of tumor cells independent of Treg depletion (Waight JD, Chand D, Dietrich S, et al. Selective FcyR Co-engagement on APCs Modulates the Activity of Therapeutic Antibodies Targeting T Cell Antigens. Cancer Cell.
2018;33(6): 1033-1047. e5). Together, these findings support improved FcgR engagement as a potent mechanism of action for improving CITE.
[0051] Given the importance of maintaining global Treg function to reduce irAEs, additional studies have identified means to retain Treg function while inhibiting intratumoral Treg-mediated suppression by pH-dependent disassociation of CTLA-4, allowing for successful recycling of CTLA-4 that consequently reduces irAE occurrence (Zhang Y, Du X, Liu M, et al. Hijacking antibody-induced CTLA-4 lysosomal degradation for safer and more effective cancer immunotherapy. Cell Res. 2019;29(8):609-627; Altman A, Kong K-F. pH- sensitive anti-CTLA4 antibodies: yes to efficacy, no to toxicity. Cell Res. 2019;29(8):601- 602). These molecules provide crucial insights that update and broaden our understanding of
the signaling dynamics of immune checkpoints. They demonstrate that CTLA-4 trafficking plays an essential role in supporting the inhibitory signaling required to maintain tolerance, and that it may be possible to exploit the specificity of its intracellular routes to reshape the whole signaling network.
[0052] The present disclosure concerns novel anti-CTLA-4 antibodies or antibody fragments suitable for therapeutic use. Some of these antibodies have been engineered using innovative techniques to confer specific binding properties to human CTLA-4 under different pH conditions. In some embodiments, the pH sensitivity of the disclosed antibodies may enable them to selectively engage with CTLA-4 in acidic or neutral pH environments, such as within a tumor microenvironment or endosome of a cell. In some embodiments, the pH sensitivity of the disclosed antibodies may also enable them to dissociate from the target under acidic pH in endosomes, allow physiological CTLA-4 recycling, and reduce adverse effects. By leveraging pH differences between the environments where an antibody may interact with CTLA-4, the disclosed antibodies may provide options for more targeted therapy with reduced irAEs. The disclosed anti-CTLA-4 antibodies or antibody fragments may exhibit distinct physiochemical or biological properties that lead to significant improvements in efficacy and safety relative to anti-CTLA-4 antibodies known in the art.
[0053] In some embodiments, provided anti-CTLA-4 antibodies or antibody fragments thereof possess pH dependent binding properties to human CTLA-4 that facilitates the dissociation of antibody from the antibody:CTLA-4 complex upon internalization into a cell, allowing either the antibody, CTLA-4, or both to be recycled instead of being degraded in the lysosome.
[0054] In some embodiments, the antibody or antigen-binding fragment thereof preserves high affinity for CTLA-4 in neutral pH environments (such as at pH>7), lower affinity in acidic environments (such as at pH 5.5-7 found in early endosomes), and significantly reduced affinity for CTLA-4 at more acidic environments (such as at pH<5.5 found in late endosomes and lysosomes).
[0055] In some embodiments, provided anti-CTLA-4 antibodies or antibody fragments thereof have a lower binding affinity (i.e. greater dissociation constant) to CTLA-4 at an acidic pH as present in endosomes of a cell in comparison with the binding affinity of CTLA-4 present in normal tissue. In some embodiments, the dissociation constant (KD) for binding at acidic pH (e.g., pH 5.0 to 6.0) is higher than the dissociation constant for binding at physiological pH
(e.g., e.g. pH 7.35 to 7.45). In some such embodiments, these anti-CTLA-4 antibodies or antibody fragments have reduced side-effects, in comparison with monoclonal anti-CTLA-4 antibodies known in the art. This may permit use of higher dosages of these anti-CTLA-4 antibodies or antibody fragments, thus providing a more effective therapeutic option without a corresponding significant increase in side effects.
[0056] In some embodiments, the provided antibodies can be used to block inhibitory activity of CTLA-4 by CTLA-4-expressing cells including regulatory T cells (Tregs), and in some cases may also mediate depletion of such cells by an ADCC mechanism. In some embodiments, depletion of Tregs in the tumor contributes to anti-tumor efficacy of the provided antibodies. In some embodiments, provided antibodies reduce CTLA-4 mediated signaling in effector T cells and exhibit activity to deplete Tregs in the tumor microenvironment that might otherwise suppress an anti-tumor immune response.
[0057] In some embodiments, the antibody or antigen-binding fragment thereof or a pharmaceutical composition comprising the antibody or antigen-binding fragment thereof induces strong deletion of Treg and local T cell activation in tumor microenvironment but minimal systemic T cell activation. In some embodiments, the antibody or antigen-binding fragment thereof or a pharmaceutical composition comprising the antibody or antigen-binding fragment thereof induces a favorable ratio of Treg cells to T effector cells in the tumor microenvironment.
[0058] All publications, including patent documents, scientific articles and databases, referred to in this application are incorporated by reference in their entirety for all purposes to the same extent as if each individual publication were individually incorporated by reference. If a definition set forth herein is contrary to or otherwise inconsistent with a definition set forth in the patents, applications, published applications and other publications that are herein incorporated by reference, the definition set forth herein prevails over the definition that is incorporated herein by reference.
[0059] The section headings used herein are for organizational purposes only and are not to be construed as limiting the subject matter described.
I. ANTI-CTLA-4 ANTIBODIES AND ANTIGEN-BINDING FRAGMENTS AND BINDING MOLECULES THEREOF
[0060] Provided herein are anti-CTLA-4 antibodies, including full-length antibodies or antigen-binding fragments, that contain a heavy chain variable region (VH) sequence and a light
chain variable region (VL) sequence as described, or a sufficient antigen-binding portion thereof. In some such embodiments, the VH is the region of the anti-CTLA-4 antibody that comprises the three heavy chain complementarity determining regions (CDRs) and the VL chain is the region of the anti-CTLA-4 antibody that comprises the three light chain CDRs. Among provided anti-CTLA-4 antibodies, including full-length antibodies or antigen-binding fragments, are antibodies that contains a VH region sequence that contains a CDR-H1, a CDR- H2 and a CDR-H3 as described and contains a VL region sequence that contains a CDR-L1, a CDR-L2 and a CDR-L3 as described. In some embodiments, the antibodies include antibodies that specifically bind to CTLA-4, e.g., human CTLA-4. Among the provided anti-CTLA-4 antibodies are human antibodies, or antibodies that are modified from or variant of human antibodies. The antibodies include isolated antibodies. Also provided are CTLA-4-binding molecules containing such antibodies, such as single-chain proteins, fusion proteins, conjugates and/or recombinant receptors such as chimeric antigen receptors. In some embodiments, provided herein is an CTLA-4-targeted antibody-drug conjugate (ADC) comprising an anti-CTLA-4 antibody or antigen binding fragment thereof described herein. In some embodiments, provided herein is a bispecific immune cell engager that is a fusion protein comprising the anti-CTLA-4 antibody or antigen binding fragment thereof described herein. In some embodiments, the bispecific immune cell engager is a bispecific T cell engager. In some aspects, the CTLA-4-binding molecules include isolated molecules.
[0061] Also provided are polynucleotides containing nucleic acids sequences encoding all or a portion of such antibodies, including an antigen-binding fragment, or binding molecule. The provided polynucleotides can be incorporated into constructs, such as deoxyribonucleic acid (DNA) or RNA constructs, such as those that can be introduced into cells for expression of the encoded anti-CTLA-4 antibodies or binding molecules.
[0062] The term “antibody” herein is used in the broadest sense and includes polyclonal and monoclonal antibodies, including intact antibodies and functional (antigen-binding) antibody fragments, including fragment antigen binding (Fab) fragments, F(ab’)2 fragments, Fab’ fragments, Fv fragments, recombinant IgG (rlgG) fragments, heavy chain variable (VH) regions capable of specifically binding the antigen, single chain antibody fragments, including single chain variable fragments (scFv), and single domain antibodies e.g., sdAb, sdFv, nanobody, VHH) fragments. The term encompasses genetically engineered and/or otherwise modified forms of immunoglobulins, such as intrabodies, peptibodies, chimeric antibodies,
fully human antibodies, humanized antibodies, and heteroconjugate antibodies, multispecific, e.g., bispecific or trispecific, antibodies, diabodies, triabodies, and tetrabodies, tandem di-scFv, tandem tri-scFv. Unless otherwise stated, the term “antibody” should be understood to encompass functional antibody fragments thereof also referred to herein as “antigen-binding fragments.” The term also encompasses intact or full-length antibodies, including antibodies of any class or sub-class, including IgG and sub-classes thereof, IgM, IgE, IgA, and IgD.
[0063] The terms “complementarity determining region,” and “CDR,” synonymous with “hypervariable region” or “HVR,” are known to refer to non-contiguous sequences of amino acids within antibody variable regions, which confer antigen specificity and/or binding affinity. In general, there are three CDRs in each heavy chain variable region (CDR-H1, CDR-H2, CDR-H3) and three CDRs in each light chain variable region (CDR-L1, CDR-L2, CDR-L3). “Framework regions” and “FR” are known to refer to the non-CDR portions of the variable regions of the heavy and light chains. In general, there are four FRs in each full-length heavy chain variable region (FR-H1, FR-H2, FR-H3, and FR-H4), and four FRs in each full-length light chain variable region (FR-L1, FR-L2, FR-L3, and FR-L4).
[0064] The precise amino acid sequence boundaries of a given CDR or FR can be readily determined using any of a number of well-known schemes, including those described by Kabat et al. (1991), “Sequences of Proteins of Immunological Interest,” 5th Ed. Public Health Service, National Institutes of Health, Bethesda, MD (“Kabat” numbering scheme); Al-Lazikani et al., J Mol Biol, 1997; 273(4):927-48 (“Chothia” numbering scheme); MacCallum et al., J. Mol. Biol, 1996; 262:732-745.” (“Contact” numbering scheme); Lefranc MP et al., Dev Comp Immunol, 2003; 27(l):55-77 (“IMGT” numbering scheme); Honegger A and Pliickthun A, J Mol Biol, 2001; 309(3):657-70, (“Aho” numbering scheme); Martin et al., PNAS, 1989; 86(23):9268-9272, (“AbM” numbering scheme); and Ye et al., Nucleic Acids Res. 2013; 41(Web Server issue):W34-40, (“IgBLAST numbering scheme). Details regarding various numbering schemes are also described in, for example, Jarasch et al., Proteins, 2017; 85( 1):65- 71; Martin et al., Bioinformatics tools for antibody engineering. In: Diibel, S. (editor) Handbook of Therapeutic Antibodies, Vol. 1. Wiley-VCH, Weinheim, Germany; Martin, A.C.R. (2010). Protein Sequence and Structure Analysis of Antibody Variable Domains. In: Kontermann, R., Diibel, S. (eds) Antibody Engineering. Springer Protocols Handbooks. Springer, Berlin, Heidelberg; and Martin, ACR, Antibody Information: How to identify the CDRs by looking at a sequence [online] bioinf.org.uk/abs/info.html, all of which are
incorporated by reference in their entireties. Various prediction algorithm tools are available and known for numbering antibody residues and CDRs (e.g., AbYsis, Abnum, AbYmod, AbRSA, IgBLAST, IMGT, or ANARCI).
[0065] The boundaries of a given CDR or FR may vary depending on the scheme used for identification. For example, the Kabat scheme is based on structural alignments, while the Chothia scheme is based on structural information. Numbering for both the Kabat and Chothia schemes is based upon the most common antibody region sequence lengths, in some cases with insertions. Insertions in the sequence relative to the standard numbering scheme are indicated using insertion letter codes. For example, residues that are inserted between residues L30 and L31 are indicated as L31A, L31B, etc. Deletions in the sequence relative to the standard scheme are accommodated by skipping numbers. The two schemes place certain insertions and deletions (“indels”) at different positions, resulting in differential numbering. For instance, the Chothia numbering scheme is nearly identical to the Kabat numbering scheme, except that insertions are placed at structural positions and topologically equivalents residues do get assigned the same numbers. The Contact scheme is based on analysis of complex crystal structures and is similar in many respects to the Chothia numbering scheme. The AbM scheme is a compromise between Kabat and Chothia definitions based on that used by Oxford Molecular’s AbM antibody modeling software. The IgBLAST scheme is based on matching to germline V, D and J genes, and can be determined using National Center for Biotechnology Information (NCBI)’s IgBLAST tool.
[0066] In some embodiments, Kabat numbering can be determined by known sequence rules as described in, for example, Kabat et al. (1991), “Sequences of Proteins of Immunological Interest,” 5th Ed. Public Health Service, National Institutes of Health, Bethesda, MD. In some embodiments, the Kabat numbering scheme in some aspects can include any of the following rules to designate CDRs: CDR-L1 starts at approximately residue 24 of the light chain, always has a preceding C residue, and always has a following W residue; the end of CDR-L1 is defined by a stretch of 3 residues, where the W residue can be followed by Y, L, or F, followed by Q or L; CDR-1 has a length of 10 to 17 residues; CDR-L2 always starts 16 residues after the end of CDR-L1; the two residues before CDR-L2 are I and Y but can also be V and Y, I and K, or I and F; CDR-L2 is always 7 residues long; CDR-L3 always starts 33 residues after the end of CDR-L2, always has a preceding C residue, and is strictly followed by a F-G-X-G sequence motif, where X is any amino acid; CDR-L3 has a length of
7 to 11 residues; CDR-H1 starts at approximately position 26 of the heavy chain; the first amino acid in CDR-H1 is always 9 residues after a conserved C residue; CDR-H1 is followed by an invariant W residue followed by typically V, but also can be I or A; CDR-H1 has a length of 5 to 7 residues; CDR-H2 always starts at 15 residues after the end of CDR-H1; the first residue in CDR-H2 is usually preceded by the sequence motif L-E-W-I-G but a number of variations exist; the end of CDR-H2 is defined by a motif of 3 residues - the first residue of the motif of 3 residues can be either K or R, the second residue of the motif of 3 residues can be L, I, V, F, T, or A, the third residue of the motif of 3 residues can be T, S, I, or A; CDR-H2 has a length of 16 to 19 residues; CDR-H3 always starts 33 residues after the end of CDR-H2 and is always 3 residues after a C residue - the first residue of CDR-H3 is preceded by the conserved C residue followed by two residues, which are usually A-R; the residues following CDR-H3 is strictly followed by a W-G-X-G sequence motif, where the X is any amino acid; CDR-H3 typically has a length of 3 to 25 residues; CDR-H3 can be much longer than 25 residues.
[0067] In some cases, according to the Chothia numbering scheme, exact boundary positions of certain CDRs can differ based on different definitions for the CDRs (See e.g., Martin, ACR, Antibody Information: How to identify the CDRs by looking at a sequence [online] bioinf.org.uk/abs/info.html). For example, in some instances, the boundary positions for CDR-F1 according to Chothia numbering can be F26— F32 (Chothia et al., Science, 1986; 233(4765):755-8 and Chothia C. and Fesk A.M. J Mol Biol, 1987; 196(4):901-17). In some instances, the boundary positions for CDR-F1 can be E25--E32 (Al-Fazikani et al., J Mol Biol, 1997; 273(4):927-48). In some instances, the boundary positions for CDR-F2 can be F50— F52 and for CDR-F3 can be F91— F96 (Chothia et al., Science, 1986; 233(4765):755-8; Chothia C. and Fesk A.M. J Mol Biol, 1987; 196(4):901-17; and Al-Fazikani et al., J Mol Biol, 1997; 273(4):927-48). In some instances, the boundary positions for CDR-H1 according to Chothia numbering can be H26— H32 (Chothia et al., Science, 1986; 233(4765):755-8; Chothia C. and Fesk A.M. J Mol Biol, 1987; 196(4):901-17; and Al-Fazikani et al., J Mol Biol, 1997; 273(4):927-48). In some instances, the boundary positions for CDR-H2 can be H53— H55 (Chothia et al., Science, 1986; 233(4765):755-8 and Chothia C. and Fesk A.M. J Mol Biol, 1987, 196(4):901-17); H52a-H55 (Tramontano et al., J Mol Biol, 1990, 215(1): 175-82), or H52— H56 (Al-Fazikani et al., J Mol Biol., 1997; 273(4):927-48). In some instances, the boundary positions for CDR-H3 can be H96— H101 (Chothia et al., Science, 1986; 233(4765):755-8 and Chothia C. and Fesk A.M. J Mol Biol., 1987; 196(4):901-17). In some
instances, the boundary positions for CDR-H3 can be H92— H104 (Morea et al., Biophys Chem, 1997; 68(1-3): 9-16 and Morea et al., J Mol Biol., 1998; 275(2): 269-94).
[0068] Table 1, below, exemplifies exemplary numbering and lists exemplary position boundaries of CDR-L1, CDR-L2, CDR-L3 and CDR-H1, CDR-H2, CDR-H3 as identified by Kabat, Chothia, AbM, and Contact schemes, respectively. For CDR-H1, residue numbering is listed using both the Kabat and Chothia numbering schemes. FRs are located between CDRs, for example, with FR-L1 located before CDR-L1, FR-L2 located between CDR-L1 and CDR- L2, FR-L3 located between CDR-L2 and CDR-L3 and so forth. It is noted that because the shown Kabat numbering scheme places insertions at H35A and H35B, the end of the Chothia CDR-H1 loop when numbered using the shown Kabat numbering convention varies between
1 - Kabat etal. (1991), “Sequences of Proteins of Immunological Interest,” 5th Ed. Public Health Service, National Institutes of Health, Bethesda, MD
2 - Al-Lazikani et al., J Mol Biol., 1997; 273(4):927-48).
[0069] Thus, unless otherwise specified, a “CDR” or “complementary determining region,” or individual specified CDRs (e.g., CDR-H1, CDR-H2, CDR-H3), of a given antibody or region thereof, such as a variable region thereof, should be understood to encompass a (or the specific) complementary determining region as defined by any of the aforementioned schemes, or other known schemes. For example, where it is stated that a particular CDR (e.g., a CDR-H3) contains the amino acid sequence of a corresponding CDR in a given VH or VL region amino acid sequence, it is understood that such a CDR has a sequence of the corresponding CDR (e.g., CDR-H3) within the variable region, as defined by any of the aforementioned schemes, or other known schemes. In some embodiments, where it is stated that an antibody or antigen-binding fragment thereof comprises a CDR-H1, a CDR-H2, and a CDR-H3 as contained within a given VH region amino acid sequence and a CDR-L1, a CDR- L2, and a CDR-L3 as contained within a given VL region amino acid sequence, the CDRs can
be defined by any of the aforementioned schemes, such as Kabat, Chothia, AbM, IgBLAST, IMGT, or Contact method, or other known scheme. In some embodiments, specific CDR sequences are specified. Exemplary CDR sequences of provided antibodies are described using various numbering schemes, although it is understood that a provided antibody can include CDRs as described according to any of the other aforementioned numbering schemes or other known numbering schemes.
[0070] Likewise, unless otherwise specified, a FR or individual specified FR(s) (e.g., FR- Hl, FR-H2, FR-H3, FR-H4, FR-L1, FR-L2, FR-L3, and/or FR-L4), of a given antibody or region thereof, such as a variable region thereof, should be understood to encompass a (or the specific) framework region as defined by any of the known schemes. In some instances, the scheme for identification of a particular CDR, FR, or FRs or CDRs is specified, such as the CDR as defined by the Kabat, Chothia, AbM, IgBLAST, IMGT, or Contact method, or other known schemes. In other cases, the particular amino acid sequence of a CDR or FR is given. In some embodiments, where it is stated that an antibody or antigen-binding fragment thereof comprises a FR-H1, a FR-H2, a FR-H3, and a FR-H4 as contained within a given VH region amino acid sequence and a FR-L1, a FR-L2, a FR-L3, and a FR-L4 as contained within a given VL region amino acid sequence, the FRs can be defined by any of the aforementioned schemes, such as Kabat, Chothia, AbM, IgBLAST, IMGT, or Contact method, or other known scheme.
[0071] The term “variable region” or “variable domain” refers to the domain of an antibody heavy or light chain that is involved in binding the antibody to antigen. The variable regions of the heavy chain and light chain (VH and VL, respectively) of a native antibody generally have similar structures, with each domain comprising four conserved framework regions (FRs) and three CDRs (See, e.g., Kindt et al. Kuby Immunology, 6th ed., W.H. Freeman and Co., page 91 (2007)). A single VH or VL domain may be sufficient to confer antigen-binding specificity. Furthermore, antibodies that bind a particular antigen may be isolated using a VH or VL domain from an antibody that binds the antigen to screen a library of complementary VL or VH domains, respectively. See, e.g., Portolano et al., J. Immunol. 150:880-887 (1993); Clarkson et al., Nature 352:624-628 (1991).
[0072] Among the provided antibodies are antibody fragments. An “antibody fragment” or “antigen-binding fragment” refers to a molecule other than an intact antibody that comprises a portion of an intact antibody that binds the antigen to which the intact antibody binds. Typically, an antigen binding fragment includes all CDRs of a variable heavy chain (VH) and
variable light chain (VL) sequence from antibodies that bind CTLA-4 set forth herein. Examples of antibody fragments include but are not limited to Fv, Fab, Fab’, Fab’-SH, F(ab’)2; diabodies; linear antibodies; heavy chain variable (VH) regions, single-chain antibody molecules such as scFvs and single-domain antibodies comprising only the VH region; and multispecific antibodies formed from antibody fragments. Papain digestion of antibodies produce two identical antigen-binding fragments, called “Fab” fragments, and a residual “Fc” fragment, a designation reflecting the ability to crystallize readily. The Fab fragment is composed of an entire E chain along with the variable region domain of the H chain (VH), and the first constant domain of one heavy chain (CHI). Each Fab fragment is monovalent with respect to antigen binding, i.e., it has a single antigen-binding site. Pepsin treatment of an antibody yields a single large F(ab’)2 fragment which roughly corresponds to two disulfide linked Fab fragments having different antigen-binding activity and is still capable of crosslinking antigen. Fab’ fragments differ from Fab fragments by having a few additional residues at the carboxy terminus of the CHI domain including one or more cysteines from the antibody hinge region. Fab’-SH is the designation herein for Fab’ in which the cysteine residue(s) of the constant domains bear a free thiol group. F(ab’)2 antibody fragments originally were produced as pairs of Fab’ fragments which have hinge cysteines between them. Other chemical couplings of antibody fragments are also known. In some embodiments, the antibody is or comprises an antibody fragment comprising a variable heavy chain (VH) and a variable light chain (VL) region. In particular embodiments, the antibodies are single-chain antibody fragments comprising a heavy chain variable (VH) region and/or a light chain variable (VL) region, such as scFvs.
[0073] Antibody fragments can be made by various techniques, including but not limited to proteolytic digestion of an intact antibody as well as production by recombinant host cells. In some embodiments, the antibodies are recombinantly -produced fragments, such as fragments comprising arrangements that do not occur naturally, such as those with two or more antibody regions or chains joined by synthetic linkers, e.g. , peptide linkers, and/or that are may not be produced by enzyme digestion of a naturally-occurring intact antibody. In some aspects, the antibody fragments are scFvs.
[0074] The term “F(ab)” refers to two of the protein fragments resulting from proteolytic cleavage of immunoglobulin G (IgG) molecules by the enzyme papain. Each F(ab) comprises
a covalent heterodimer of the VH chain and VL chain and includes an intact antigen-binding site.
[0075] The term “F(ab)2” refers to a protein fragment of IgG generated by proteolytic cleavage by the enzyme pepsin. Each F(ab’)2 fragment comprises two F(ab) fragments, thus comprising both antigen-binding sites.
[0076] An “Fv fragment” for use according to certain embodiments of the present invention can be produced by preferential proteolytic cleavage of an IgM, and on rare occasions of an IgG or IgA immunoglobulin molecule. Fv fragments are, however, more commonly derived using recombinant techniques known in the art. The Fv fragment includes a non-covalent VH"VL heterodimer including an antigen-binding site which retains much of the antigen recognition and binding capabilities of the native antibody molecule, but lacking the CHI and CL domains contained within a Fab. Inbar et al. (1972) Proc. Nat. Acad. Sci. USA 69:2659- 2662; Hochman et al. (1976) Biochem 75:2706-2710; and Ehrlich et al. (1980) Biochem 79:4091-4096.
[0077] In certain embodiments, single chain Fv (scFv) antibodies are contemplated and may be prepared using standard molecular biology techniques following the teachings of the present application with regard to selecting antibodies having the desired specificity.
[0078] In still further embodiments, chimeric antibodies may be made. For example, a chimeric antibody may comprise CDRs and framework regions from different antibodies. These antibodies may be produced through recombinant molecular biological techniques or may be physically conjugated together.
[0079] A scFv polypeptide is a covalently linked VH"VL heterodimer which is expressed from a gene fusion including VH- and Vi.-cncoding genes linked by a peptide-encoding linker. Huston et al. (1988) Proc. Nat. Acad. Sci. USA S5(16):5879-5883. A number of methods have been described to discern chemical structures for converting the naturally aggregated — but chemically separated — light and heavy polypeptide chains from an antibody V region into an scFv molecule which will fold into a three dimensional structure substantially similar to the structure of an antigen-binding site. See, e.g., U.S. Pat. Nos. 5,091,513 and 5,132,405, to Huston et al. and U.S. Pat. No. 4,946,778, to Eadner et al.
[0080] In certain embodiments, the antibodies described herein may be provided in the form of a UniBody®. A UniBody® is an IgG4 antibody with the hinge region removed (see GenMab Utrecht, The Netherlands; see also, e.g., US20090226421). This antibody technology
creates a stable, smaller antibody format with an anticipated longer therapeutic window than current small antibody formats. IgG4 antibodies are considered inert and thus do not interact with the immune system. Fully human IgG4 antibodies may be modified by eliminating the hinge region of the antibody to obtain half-molecule fragments having distinct stability properties relative to the corresponding intact IgG4 (GenMab, Utrecht). Halving the IgG4 molecule leaves only one area on the UniBody® that can bind to cognate antigens (e.g., disease targets) and the UniBody® therefore binds univalently to only one site on target cells. For certain cancer cell surface antigens, this univalent binding may not stimulate the cancer cells to grow as may be seen using bivalent antibodies having the same antigen specificity, and hence UniBody® technology may afford treatment options for some types of cancer that may be refractory to treatment with conventional antibodies. The small size of the UniBody® can be a great benefit when treating some forms of cancer, allowing for better distribution of the molecule over larger solid tumors and potentially increasing efficacy.
[0081] In certain embodiments, the antibodies of the present disclosure may be chimeric antibodies. In this regard, a chimeric antibody is comprised of an antigen-binding fragment of an anti-CTLA-4 antibody operably linked or otherwise fused to a heterologous Fc portion of a different antibody. In certain embodiments, the heterologous Fc domain is of human origin. In further embodiments, the heterologous Fc domain may be from a different Ig class from the parent antibody, including IgA (including subclasses IgAl and IgA2), IgD, IgE, IgG (including subclasses IgGl, IgG2, IgG3, and IgG4), and IgM. In further embodiments, the heterologous Fc domain may be comprised of CH2 and CH3 domains from one or more of the different Ig classes.
[0082] In certain embodiments, the antibodies of the present disclosure may be “non- naturally occurring” antibodies. Non-naturally occurring antibodies can refer to antibodies that comprise one or more amino acid modifications, such that the resultant antibody is substantially non-naturally occurring (e.g., does not exists in nature). These amino acid modifications can include point mutations, wherein a naturally occurring amino acid is substituted for another naturally occurring amino acid. In some embodiments, the amino acid modifications can include point mutations wherein a non-naturally occurring amino acid is substituted for a naturally occurring amino acid. Non-naturally occurring antibodies can also refer to antibodies that are conjugated to a heterologous protein or compound, such as a detectable marker.
[0083] Among the provided antibodies are human antibodies. A “human antibody” is an antibody with an amino acid sequence corresponding to that of an antibody produced by a human or a human cell, or non-human source that utilizes human antibody repertoires or other human antibody-encoding sequences, including human antibody libraries. The term excludes humanized forms of non-human antibodies comprising non-human antigen-binding regions, such as those in which all or substantially all CDRs are non-human. The term includes antigenbinding fragments of human antibodies.
[0084] Human antibodies may be prepared by administering an immunogen to a transgenic animal that has been modified to produce intact human antibodies or intact antibodies with human variable regions in response to antigenic challenge. Such animals typically contain all or a portion of the human immunoglobulin loci, which replace the endogenous immunoglobulin loci, or which are present extrachromosomally or integrated randomly into the animal’s chromosomes. In such transgenic animals, the endogenous immunoglobulin loci have generally been inactivated. Human antibodies also may be derived from human antibody libraries, including phage display and cell-free libraries, containing antibody-encoding sequences derived from a human repertoire.
[0085] Among the provided antibodies are monoclonal antibodies, including monoclonal antibody fragments. The term “monoclonal antibody” as used herein refers to an antibody obtained from or within a population of substantially homogeneous antibodies, i.e., the individual antibodies comprising the population are identical, except for possible variants containing naturally occurring mutations or arising during production of a monoclonal antibody preparation, such variants generally being present in minor amounts. In contrast to polyclonal antibody preparations, which typically include different antibodies directed against different epitopes, each monoclonal antibody of a monoclonal antibody preparation is directed against a single epitope on an antigen. The term is not to be construed as requiring production of the antibody by any particular method. A monoclonal antibody may be made by a variety of techniques, including but not limited to generation from a hybridoma, recombinant DNA methods, phage-display and other antibody display methods.
[0086] In some aspects, the antibody or the antigen-binding fragments of the antibody is isolated.
[0087] The present disclosure contemplates variants of the antibodies disclosed herein. In certain embodiments, such variant antibodies or antigen-binding fragments, or CDRs thereof,
bind to CTLA-4 at least about 50%, at least about 70%, at least about 80%, at least about 85%, at least about 90% and in certain embodiments, at least about 95% as well as an antibody sequence specifically set forth herein. In further embodiments, such variant antibodies or antigen-binding fragments, or CDRs thereof, bind to CTLA-4 with greater affinity than the antibodies set forth herein, for example, that bind quantitatively at least about 105%, 106%, 107%, 108%, 109%, or 110% as well as an antibody sequence specifically set forth herein.
[0088] In particular embodiments, a subject antibody may have: a) a heavy chain variable region having an amino acid sequence that is at least 80% identical, at least 85% identical, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, or 99% or 100% identical, to a heavy chain variable region of an anti-CTLA-4 antibody described herein; and b) a light chain variable region having an amino acid sequence that is at least 80% identical, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, or 99% or 100% identical, to a light chain variable region of an anti-CTLA-4 antibody described herein.
A. Exemplary Antibodies
[0089] In some embodiments, the anti-CTLA-4 antibody contains a VH region and a VL region containing a combination of six CDRs as described below. In some embodiments, the anti-CTLA-4 antibody contains a VH region and a VL region as described below. In any such embodiments, a VH region sequence can be any of the VH region sequence described herein. In any such embodiments, a VL region sequence can be any of the VL region sequence described herein. In any such embodiments, any of the VH region sequence and any of the VL region sequence described herein can be used in combination. In some such embodiments, the antibody is an antigen-binding fragment, such as a Fab or an scFv. In some embodiments, the antibody or antigen-binding fragment further comprises at least a portion of an immunoglobulin constant region or a variant thereof. In some such embodiments, the antibody is a full-length antibody that also contains a constant region. In some embodiments, an anti- CTLA-4 antibody comprises at least one heavy chain comprising a VH region and at least a portion of a heavy chain constant region, and at least one light chain comprising a VL region and at least a portion of a light chain constant region. In some embodiments, an anti-CTLA-4 antibody comprises two heavy chains, wherein each heavy chain comprises a VH region and at least a portion of a heavy chain constant region, and two light chains, wherein each light chain comprises a VL region and at least a portion of a light chain constant region. In some embodiments, an anti-CTLA-4 antibody comprises two heavy chains, wherein each heavy
chain comprises a VH region and a heavy chain constant region, and two light chains, wherein each light chain comprises a VL region and a light chain constant region.
[0090] Anti-CTLA-4 antibodies, including antigen-binding fragments thereof, include any combination of the heavy chain and light chain complementarity-determining regions (CDRs) discussed herein. In some embodiments, the anti-CTLA-4 antibody or antigen-binding fragment thereof comprises any one of the CDR-H1 as described herein, any one of the CDR- H2 as described herein, any one of the CDR-H3 as described herein, any one of the CDR-L1 as described herein, any one of the CDR-L2 as described herein and any one of the CDR-L3 as described herein. In some of any such embodiments, any one or more of the CDR-H1, the CDR-H2 and the CDR-H3 sequences described herein, and any one or more of the CDR-L1, the CDR-L2 and the CDR-L3 sequences described herein can be used in combination.
[0091] In some embodiments, a provided anti-CTLA-4 antibody or an antigen-binding fragment thereof has a CDR-H1, a CDR-H2 and a CDR-H3 present in a VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112. In some embodiments, a provided anti-CTLA-4 antibody or an antigen-binding fragment thereof has a CDR-L1, a CDR-L2 and a CDR-L3 present in a VL region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118. In some embodiments, the provided anti- CTLA-4 antibody contains a combination of any of such six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) described above). In some embodiments, a provided anti-CTLA-4 antibody or an antigen-binding fragment thereof has a CDR-H1, a CDR- H2 and a CDR-H3 present in a VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112, and a CDR-L1, a CDR-L2 and a CDR-L3 present in a VL region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118, wherein the CDR-H1, CDR- H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3 are not set forth in SEQ ID NOS: 49, 57, 68, 74, 89 and 98, respectively. In some embodiments, the combination of six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) is according to Kabat numbering. In some embodiments, the combination of six CDRs (a CDR-H1, a CDR-H2, a
CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) is according to Chothia numbering. In some embodiments, the combination of six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) is according to AbM numbering, In some embodiments, the combination of six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) is according to IMGT numbering. In some embodiments, the combination of six CDRs (a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2 and a CDR-L3) is according to IgBLAST numbering.
[0092] Exemplary heavy and light chain CDR sequences of the anti-CTLA-4 antibodies and antigen-binding fragments thereof are provided in Tables 2A and 2B.
[0093] In some of any of the provided embodiments, the VH region contains a CDR-H1 set forth in any one of SEQ ID NOs: 49-56, a CDR-H2 set forth in any one of SEQ ID NOs: 57- 67, and a CDR-H3 set forth in any one of SEQ ID NOs: 68-73; and the VL region contains a CDR-L1 set forth in any one of SEQ ID NOs: 74-88, a CDR-L2 set forth in any one of SEQ ID NOs: 89-97, and a CDR-L3 set forth in any one of SEQ ID NOs: 98-100. In some embodiments, the anti-CTLA-4 antibody or antigen-binding fragment does not contain the combination of CDRs set forth in SEQ ID NO: 49, SEQ ID NO: 57, SEQ ID NO: 68, SEQ ID NO: 74, SEQ ID NO: 89, and SEQ ID NO: 98. In some embodiments, the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3 are not set forth in SEQ ID NOS: 49, 57, 68, 74, 89 and 98, respectively. In some embodiments, the VH region comprises a CDR-H1 comprising the sequence set forth in in any one of SEQ ID NOs: 49-56, a CDR-H2 comprising the sequence set forth in any one of SEQ ID NOs: 57-67, and a CDR-H3 comprising the sequence set forth in any one of SEQ ID NOs: 68-73, and the VL region comprises a CDR-L1 comprising the sequence set forth in any one of SEQ ID NOs: 75-88, a CDR-L2 comprising the sequence set forth in any one of SEQ ID NOs: 90-97; and a CDR-L3 comprising the sequence set forth in any one of SEQ ID NOs: 99 and 100. In some embodiments, the VH region comprises a CDR- H1 comprising the sequence set forth in any one of SEQ ID NOs: 50-56, a CDR-H2 comprising the sequence set forth in any one of SEQ ID NOs: 58-67, and a CDR-H3 comprising the
sequence set forth in any one of SEQ ID NOs: 69-73, and the VL region comprises a CDR-L1 comprising the sequence set forth in any one of SEQ ID NOs: 74-88, a CDR-L2 comprising the sequence set forth in any one of SEQ ID NOs: 89-97; and a CDR-L3 comprising the sequence set forth in any one of SEQ ID NOs: 98-100. Also among the antibodies are those having sequences at least at or about 90%, at or about 91%, at or about 92%, at or about 93%, at or about 94%, at or about 95%, at or about 96%, at or about 97%, at or about 98%, or at or about 99% identical to any such CDR sequence, e.g., any of the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2, CDR-L3. In some embodiments, among the antibodies are those in which a CDR contained therein has no more than 2 amino acid difference compared to any such above CDR sequence, e.g., any of the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2, CDR-L3. In some embodiments, among the antibodies are those in which a CDR contained therein has no more than 1 amino acid difference compared to any such above CDR sequence, e.g., any of the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2, CDR-L3.
[0094] In some of any of the provided embodiments, the VH region contains a CDR-H1 set forth in any one of SEQ ID NOs: 50-56, a CDR-H2 set forth in any one of SEQ ID NOs: 58- 67, and a CDR-H3 set forth in any one of SEQ ID NOs: 69-73; and the VL region contains a CDR-L1 set forth in any one of SEQ ID NOs: 75-88, a CDR-L2 set forth in any one of SEQ ID NOs: 90-97, and a CDR-L3 set forth in any one of SEQ ID NOs: 99-100.
[0095] In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 1. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 2. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 3. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 4. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 5. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 6. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 7. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 8. In some
embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 9. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 10. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 11. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 12. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 13. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 14. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 15. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 16. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 17. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 18. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 19. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 20. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 21. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 22. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 23. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 24. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR- Hl, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 25. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR- H3 present in SEQ ID NO: 26. In some embodiments, the antibody or antigen-binding
fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 27. In some embodiments, the antibody or antigen-binding fragment has a VH region that contains a CDR-H1, a CDR-H2 and a CDR-H3 present in SEQ ID NO: 112. In some embodiments, the CDR sequence is according to Kabat numbering. In some embodiments, the CDR sequence is according to Chothia numbering. In some embodiments, the CDR sequence is according to AbM numbering. In some embodiments, the CDR sequence is according to IMGT numbering. In some embodiments, the CDR sequence is according to IgBLAST numbering.
[0096] In some embodiments, the antibody or antigen -binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 28. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR- Ll, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 29. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR- L3 present in SEQ ID NO: 30. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 31. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 32. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 33. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 34. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 35. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR- Ll, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 36. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR- L3 present in SEQ ID NO: 37. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 38. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 39. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 40. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ
ID NO: 41. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 42. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR- Ll, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 43. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR- L3 present in SEQ ID NO: 44. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 46. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 46. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 47. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ
ID NO: 48. In some embodiments, the antibody or antigen-binding fragment has a VL region that contains a CDR-L1, a CDR-L2 and a CDR-L3 present in SEQ ID NO: 118. In some embodiments, the CDR sequence is according to Kabat numbering. In some embodiments, the CDR sequence is according to Chothia numbering. In some embodiments, the CDR sequence is according to AbM numbering. In some embodiments, the CDR sequence is according to IMGT numbering. In some embodiments, the CDR sequence is according to IgBLAST numbering.
[0097] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 58, and 68, respectively; SEQ ID NOS: 49, 57, and 69, respectively; SEQ ID NOS: 51, 57, and 68, respectively; SEQ ID NOS: 50, 59, and 68, respectively; SEQ ID NOS: 52, 60, and 68, respectively; SEQ ID NOS: 52, 61, and 68, respectively; SEQ ID
NOS: 51, 62, and 68, respectively; SEQ ID NOS: 50, 63, and 68, respectively; SEQ ID NOS:
50, 57, and 69, respectively; SEQ ID NOS: 53, 57, and 68, respectively; SEQ ID NOS: 53, 57, and 70, respectively; SEQ ID NOS: 54, 57, and 71, respectively; SEQ ID NOS: 55, 57, and 68, respectively; SEQ ID NOS: 56, 57, and 72, respectively; SEQ ID NOS: 49, 64, and 68, respectively; SEQ ID NOS: 49, 60, and 68, respectively; SEQ ID NOS: 49, 65, and 68, respectively; SEQ ID NOS: 49, 66, and 68, respectively; SEQ ID NOS: 49, 57, and 71, respectively; SEQ ID NOS: 49, 57, and 73, respectively; SEQ ID NOS: 49, 57, and 72, respectively; SEQ ID NOS: 49, 57, and 68, respectively; SEQ ID NOS: 49, 67, and 68,
respectively; SEQ ID NOS: 49, 65, and 73, respectively; SEQ ID NOS: 52, 60, and 72, respectively; SEQ ID NOS: 52, 67, and 68, respectively; SEQ ID NOS: 49, 57, and 72, respectively; or SEQ ID NOS: 49, 65, and 72, respectively.
[0098] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 75, 89, and 98, respectively; SEQ ID NOS: 76, 89, and 98, respectively; SEQ ID NOS: 77, 89, and 98, respectively; SEQ ID NOS: 78, 90, and 98, respectively; SEQ ID NOS: 79, 90, and 98, respectively; SEQ ID NOS: 80, 91, and 98, respectively; SEQ ID NOS: 78, 89, and 99, respectively; SEQ ID NOS: 81, 92, and 98, respectively; SEQ ID NOS: 82, 89, and 98, respectively; SEQ ID NOS: 82, 93, and 98, respectively; SEQ ID NOS: 83, 92, and 98, respectively; SEQ ID NOS: 84, 89, and 99, respectively; SEQ ID NOS: 84, 89, and 100, respectively; SEQ ID NOS: 85, 89, and 98, respectively; SEQ ID NOS: 86, 94, and 98, respectively; SEQ ID NOS: 87, 95, and 98, respectively; SEQ ID NOS: 85, 96, and 98, respectively; SEQ ID NOS: 88, 89, and 98, respectively; SEQ ID NOS: 74, 97, and 98, respectively; SEQ ID NOS: 74, 89, and 98, respectively; SEQ ID NOS: 84, 89, and 98, respectively; or SEQ ID NOS: 88, 89, and 100, respectively.
[0099] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 58, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0100] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 69, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0101] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 51, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0102] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 59, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0103] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0104] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 61, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0105] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 51, 62, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0106] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 63, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0107] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 57, and 69, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0108] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set
forth in SEQ ID NOS: 53, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0109] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 53, 57, and 70, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0110] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 54, 57, and 71, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0111] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 55, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0112] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 56, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0113] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 64, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0114] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 60, and 68, respectively, and the VL region comprises a CDR-L1, a
CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0115] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0116] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 66, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0117] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 71, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0118] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 73, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0119] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively.
[0120] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 75, 89, and 98, respectively.
[0121] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
[0122] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 77, 89, and 98, respectively.
[0123] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 78, 90, and 98, respectively.
[0124] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 79, 90, and 98, respectively.
[0125] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 80, 91, and 98, respectively.
[0126] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 78, 89, and 99, respectively.
[0127] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set
forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 81, 92, and 98, respectively.
[0128] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 82, 89, and 98, respectively.
[0129] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 82, 93, and 98, respectively.
[0130] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 83, 92, and 98, respectively.
[0131] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0132] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 100, respectively.
[0133] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a
CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
[0134] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 86, 94, and 98, respectively.
[0135] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 87, 95, and 98, respectively.
[0136] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 96, and 98, respectively.
[0137] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 88, 89, and 98, respectively.
[0138] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 97, and 98, respectively.
[0139] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
[0140] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
[0141] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
[0142] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
[0143] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
[0144] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
[0145] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
[0146] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set
forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
[0147] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
[0148] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2 and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2 and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
[0149] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
[0150] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
[0151] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
[0152] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a
CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
[0153] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
[0154] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
[0155] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
[0156] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
[0157] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0158] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0159] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0160] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0161] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0162] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0163] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 88, 89, and 100, respectively.
[0164] In some embodiments, the provided antibody is a human antibody or antigenbinding fragment thereof that contains a framework region that contains human germline gene segment sequences. For example, in some embodiments, the human antibody contains a VH region in which the framework region, e.g.FRl, FR2, FR3 and FR4, has at least 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a framework region encoded by a human germline antibody segment, such as a V segment and/or J segment. In some embodiments, the human antibody contains a VL region in which the framework region e.g.FRl, FR2, FR3 and
FR4, has at least 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a framework region encoded by a human germline antibody segment, such as a V segment and/or J segment. For example, in some such embodiments, the framework region sequence contained within the VH region and/or VL region differs by no more than 10 amino acids, such as no more than 9, 8, 7, 6, 5, 4, 3, 2 or 1 amino acid, compared to the framework region sequence encoded by a human germline antibody segment.
[0165] In some embodiments of the antibody or antigen-binding fragment thereof provided herein, the VH region comprises any of the CDR-H1, CDR-H2 and CDR-H3 as described and comprises a framework region 1 (FR1), a FR2, a FR3 and/or a FR4 having at least at or about 95%, at or about 96%, at or about 97%, at or about 98%, or at or about 99% sequence identity, respectively, to a FR1, a FR2, a FR3 and/or a FR4 contained within the VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112. In some embodiments of the antibody or antigen-binding fragment thereof provided herein, the VH region comprises any of the CDR-H1, CDR-H2 and CDR-H3 as described and comprises a framework region 1 (FR1), a FR2, a FR3 and/or a FR4 contained within the VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112.
[0166] In some embodiments of the antibody or antigen-binding fragment thereof provided herein, the VH region comprises any of the CDR-L1, CDR-L2 and CDR-L3 as described and comprises a framework region 1 (FR1), a FR2, a FR3 and/or a FR4 having at least at or about 95%, at or about 96%, at or about 97%, at or about 98%, or at or about 99% sequence identity, respectively, to a FR1, a FR2, a FR3 and/or a FR4 contained within the VL region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118. In some embodiments of the antibody or antigen-binding fragment thereof provided herein, the VL region comprises any of the CDR-H1, CDR-H2 and CDR-H3 as described and comprises a framework region 1 (FR1), a FR2, a FR3 and/or a FR4 contained within the VH region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118.
[0167] In some embodiments, any of the provided anti-CTLA-4 antibodies or antigen binding fragments has a VH region having the amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in any one of SEQ ID NOs: 1-27 and 112, and has a VL region having the amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and
118, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in any one of SEQ ID NOs: 28-48 and 118.
[0168] In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 1, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 1. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 2, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 2. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 3, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 3. In some embodiment, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 4, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 4. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 5, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 5. In some embodiment, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 6, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO:6. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 7, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 7. In some embodiment, the antibody or antigen-binding fragment has a VH region having the amino acid
sequence set forth in SEQ ID NO: 8, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 8. In some embodiment, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 9, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 9. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 10, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 10. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 11, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 11. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 12, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 12. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 13, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 13. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 14, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 14. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 15, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 15. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid
sequence set forth in SEQ ID NO: 16, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 16. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 17, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 17. In some embodiment, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 18, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 18. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 19, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 19. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 20, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 20. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 21, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 21. In some embodiments, the antibody or antigen-binding fragment has has a VH region having the amino acid sequence set forth in SEQ ID NO: 22, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 22. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 23, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 23. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid
sequence set forth in SEQ ID NO: 24, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 24. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 25, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 25. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 26, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 26. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 27, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 27. In some embodiments, the antibody or antigen-binding fragment has a VH region having the amino acid sequence set forth in SEQ ID NO: 112, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 112. In some embodiments, the antibody or antigen-binding fragment does not have a VH region having the amino acid sequence set forth in SEQ ID NO: 112.
[0169] In some embodiments, the antibody or antigen -binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 28, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 28. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 29, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 30, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%,
97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 31, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 31. In some embodiments, a VL region having the amino acid sequence set forth in SEQ ID NO: 32, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 33, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 33. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 34, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 35, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 35. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 36, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 36. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 37, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 37. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 38, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region
amino acid sequence set forth in SEQ ID NO: 38. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 39, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 40, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 41, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 42, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 42. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 43, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 43. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 44, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 44. In some embodiment, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 45, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 45. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 46, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region
amino acid sequence set forth in SEQ ID NO: 46. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 47, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 48, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 48. In some embodiments, the antibody or antigen-binding fragment has a VL region having the amino acid sequence set forth in SEQ ID NO: 118, or an amino acid sequence that has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 118. In some embodiments, the antibody or antigen-binding fragment does not have a VL region having the amino acid sequence set forth in SEQ ID NO: 118.
[0170] In some embodiments, the antibody or antigen-binding fragment has a VH region having an amino acid sequence that is less than 100% identical to SEQ ID NO: 112 and has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 112. In some embodiments, the antibody or antigen-binding fragment has a VL region having an amino acid sequence that is less than 100% identical to SEQ ID NO: 118 and has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in SEQ ID NO: 118. In some embodiments, the VH region does not have the amino acid sequence set forth in SEQ ID NO: 112. In some embodiments, the VL region does not have the amino acid sequence set forth in SEQ ID NO: 118. In some embodiments, the VH region and the VL region of the antibody are not set forth SEQ ID NO: 112 and 118, respectively. In some embodiments, the antibody or antigen-binding fragment has a VH region having an amino acid sequence that is less than 100% identical to SEQ ID NO: 112 and has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region amino acid sequence set forth in SEQ ID NO: 112, and a VL region having an amino acid sequence set forth in SEQ ID NO: 118. In some embodiments, the antibody or antigen-binding
fragment has a VH region having an amino acid sequence set forth in SEQ ID NO: 112, and a VL region having an amino acid sequence that is less than 100% identical to SEQ ID NO: 118 and has at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VL region amino acid sequence set forth in set forth in SEQ ID NO: 118.
[0171] In some embodiments, the antibody or antigen-binding fragment provided herein, the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 1, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 3, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 4, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 5, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 6, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an
amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 7, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 8, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 9, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 10, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 11, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 12, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 13, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 14, and the VL region is or comprises an amino acid
sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 15, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 16, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 17, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 18, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 19, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 20, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 21, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence
having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 28; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 30; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 31; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 32; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 33; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having
at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 34; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 35; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 36; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 37; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 38; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 40; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or
about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 42; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 43; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 44; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 45; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 46; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%,
88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%,
90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%,
91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; or the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 5, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 48.
[0172] In some embodiments, the VH region of the antibody or antigen-binding fragment thereof comprises the amino acid sequence of any one of SEQ ID NOs: 1-27 and 112, and the VL region of the antibody or antigen-binding fragment comprises the amino acid sequence of any one of SEQ ID NOs: 28-48 and 118.
[0173] In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 1 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 2 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 3 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 4 and 118, respectively. In some embodiments of the antibody or antigenbinding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 5 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 6 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 7 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 8 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 9 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 10 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 11 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 12 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 13 and 118, respectively. In some embodiments of the
antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 14 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 15 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 16 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 17 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 18 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 19 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 20 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 21 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 22 and 118, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 28, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 29, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 30, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 31, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 32, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 33, respectively. In some embodiments of the antibody or
antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 34, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 35, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 36, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 37, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 38, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 40, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 41, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 42, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 43, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 44, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 45, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 112 and 46, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 23 and 41, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 24 and 41, respectively. In some embodiments of the antibody or antigen-binding
fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 25 and 41, respectively. In some embodiments of the antibody or antigenbinding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 26 and 41, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 22 and 41, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 27 and 41, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 23 and 47, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 24 and 47, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 25 and 47, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 26 and 47, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 22 and 47, respectively. In some embodiments of the antibody or antigenbinding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 27 and 47, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 23 and 29, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 24 and 29, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 25 and 29, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 26 and 29, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 22 and 29, respectively. In some embodiments of the antibody or antigen-binding fragment
provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 27 and 29, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 23 and 39, respectively. In some embodiments of the antibody or antigenbinding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 24 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 25 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 26 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 22 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 27 and 39, respectively. In some embodiments of the antibody or antigen-binding fragment provided herein, the VH region and the VL region are or comprise the sequence set forth in SEQ ID NO: 5 and 48, respectively.
[0174] Table El provides the SEQ ID NOS: of exemplary provided antibody or antigenbinding fragment thereof. In some embodiments, the CTLA-4-binding antibody or fragment thereof comprises a VH region that comprises the CDR-H1, the CDR-H2 and the CDR-H3 sequence and a VL region that comprises the CDR-L1, the CDR-L2 and the CDR-L3 sequence set forth in the SEQ ID NOS: listed in each row of Table El below (by Kabat numbering scheme). In some embodiments, the CTLA-4-binding antibody or fragment thereof comprises a VH region sequence and a VL region sequence set forth in the SEQ ID NOS: listed in each row of Table El below, or an antibody comprising a VH region and a VL region amino acid sequence that has at least 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to the VH region sequence and the VL region sequence set forth in the SEQ ID NOS: listed in each row of Table El below. In some embodiments, the CTLA-4-binding antibody or fragment thereof comprises a VH region sequence and a VL region sequence set forth in the SEQ ID NOS: listed in each row of Table El below.
[0175] In some embodiments, the provided antibody or antigen-binding fragment thereof comprises a VH region and a VL region, wherein the VH region of the antibody or antigen-
binding fragment thereof can contain a combination of any of the CDR-H1, the CDR-H2 and the CDR-H3 amino acid sequences set forth in Table El, and the VL region of the antibody or antigen-binding fragment thereof can contain a combination of any of the CDR-L1, the CDR- L2 and the CDR-L3 amino acid sequences set forth in Table El. In some embodiments, the provided antibody or antigen-binding fragment thereof comprises a VH region and/or a VL region set forth in Table El, in any combination, orientation or containing a different linker. In some aspects, the antibody or antigen-binding fragment thereof comprises a VH region described in Table El. In some aspects, the antibody or antigen-binding fragment thereof comprises a VL region described in Table El.
[0176] In some embodiments, provided antibodies include those that include amino acid residues (such as due to mutation compared to a reference sequence) that alter pH binding such that the antibodies exhibit better binding at neutral pH (e.g., pH 7.4) compared to acidic pH (e.g., pH 5.0). Results herein surprisingly found enrichment of a number of antibodies, including antibody fragments, with such activity that contain one or more of the following amino acids in the VH region: arginine (R) at position 31, glutamic acid (E) or histidine (H) at position 54, histidine (H) at position 59, histidine (H) at position 101, or histidine (H) at position 102 (corresponding to arginine (R) at position 31, glutamic acid (E) or histidine (H) at position 53, histidine (H) at position 58, histidine (H) at position 97, or histidine (H) at position 98, respectively, based on Kabat numbering), or any combination thereof; and/or in the VL region: histidine (H) at position 28, histidine (H) at position 31, histidine (H) at position 32, or a histidine (H) at position 91 (corresponding to histidine (H) at position 27a, histidine (H) at position 30, histidine (H) at position 31, or histidine (H) at position 90, respectively, based on Kabat numbering), or any combination thereof. In some embodiments, provided antibodies, including antibody fragments, herein contain one or more amino acid mutation in the VH region that is S31H, D54H, D54E, Y59H, W101H, or L102H, which correspond to S31R, D53H, D53E, Y58H, W97H, or L98H based on Kabat numbering, respectively, or any combination thereof, compared to the VH region set forth in SEQ ID NO: 112; and/or one or more amino acid mutation in the VL region that is S28H, S31H, S32H, or Q91H, which correspond to S27aH, S30H, S31H, or Q90H based on Kabat numbering, respectively, or any combination thereof, compared to the VL region set forth in SEQ ID NO: 118.
[0177] In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises one or more mutations selected from the group consisting of:
S31R, D54H, D54E, Y59H, W101H, and L102H (S31R, D53H, D53E, Y58H, W97H, and L98H, respectively, based on Kabat numbering) in the VH region, such as compared to a reference anti-CTLA-4 antibody VH, e.g. set forth in SEQ ID NO: 112.
[0178] In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprise a histidine (H) at position 98 based on Kabat numbering (e.g., L102H mutation compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering) (e.g., S31R mutation compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a glutamic acid (E) at position 53 based on Kabat numbering) (e.g., D54E mutation compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 53 based on Kabat numbering (e.g., D54H mutation compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 58 based on Kabat numbering (e.g., Y59H mutation compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 97 based on Kabat numbering (e.g., W101H mutation compared to SEQ ID NO: 112) in the VH region.
[0179] In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering and a arginine (R) at position 31 based on Kabat numbering (e.g., L102H and S31R mutation compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti- CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering and a glutamic acid (E) at position 53 based on Kabat numbering (e.g., L102H and D54E mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering and a histidine (H) at position 53 based on Kabat numbering (e.g., L102H and D54H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H and Y59H
mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H and W 101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering) and a glutamic acid (E) at position 53 based on Kabat numbering (e.g., S31R and D54E mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 53 based on Kabat numbering (e.g., S31R and D54H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., S31R and Y59H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., S31R and W 101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a glutamic acid (E) at position 53 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., D54E and Y59H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a glutamic acid (E) at position 53 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., D54E and W 101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 53 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., D54H and Y59H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 53 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., D54H and W 101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H)
at position 58 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., Y59H and W 101H mutations compared to SEQ ID NO: 112) in the VH region.
[0180] In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering and a glutamic acid (E) at position 53 based on Kabat numbering (e.g., L102H, S31R, and D54E mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigenbinding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 53 based on Kabat numbering (e.g., L102H, S31R, and D54H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H, S31R, and Y59H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H, D54E, and Y59H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti- CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, D54E, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a a histidine (H) at position 53 based on Kabat numbering and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H, D54H, and Y59H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine
(H) at position 98 based on Kabat numbering, a histidine (H) at position 53 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, D54H, and W 101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, Y59H, and W 101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering), a glutamic acid (E) at position based on Kabat numbering, and a histidine (H) at position 58 based on Kabat numbering (e.g., S31R, D54E, and Y59H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering), a glutamic acid (E) at position 53 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., S31R, D54E, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering), a histidine (H) at position 53 based on Kabat numbering, and a histidine (H) at position 58 based on Kabat numbering (e.g., S31R, D54H, and Y59H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering), a histidine (H) at position 53 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., S31R, D54H, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering), a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., S31R, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a glutamic acid (E) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., D54E, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding
fragment thereof comprises a histidine (H) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., D54H, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
[0181] In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering, and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H, S31R, D54E, and Y59H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, D54E, and W 101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a histidine (H) at position 53 based on Kabat numbering, and a histidine (H) at position 58 based on Kabat numbering (e.g., L102H, S31R, D54H, and Y59H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti- CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a histidine (H) at position 53 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, D54H, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, D54E,
Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, a histidine (H) at position 53, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, D54H, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises an arginine (R) at position 31 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., S31R, D54E, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a glutamic acid (E) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, D54E, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 98 based on Kabat numbering, an arginine (R) at position 31 based on Kabat numbering, a histidine (H) at position 53 based on Kabat numbering, a histidine (H) at position 58 based on Kabat numbering, and a histidine (H) at position 97 based on Kabat numbering (e.g., L102H, S31R, D54H, Y59H, and W101H mutations compared to SEQ ID NO: 112) in the VH region.
[0182] In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises one or more mutations selected from the group consisting of: S28H, S31H, S32H, and Q91H (S27aH, S30H, S31H, and Q90H, respectively, based on Kabat numbering) in the VL region, such as compared to a reference anti-CTLA-4 antibody VL, e.g. set forth in SEQ ID NO: 118. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering (e.g., S28H mutation compared to SEQ ID NO: 118) in the VL region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 30 based on Kabat numbering (e.g., S31H mutation compared to SEQ ID NO: 118) in the VL region. In some embodiments, the provided anti-
CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 31 based on Kabat numbering (e.g., S32H mutation compared to SEQ ID NO: 118) in the VL region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 90 based on Kabat numbering (e.g., Q91H mutation compared to SEQ ID NO: 118) in the VL region.
[0183] In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering and a histidine (H) at position 30 based on Kabat numbering (e.g., S28H and S31H mutations compared to SEQ ID NO: 118) in the VL region. In some embodiments, the provided anti- CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position27a based on Kabat numbering and a histidine (H) at position 31 based on Kabat numbering (e.g., S28H and S32H mutations compared to SEQ ID NO: 118) in the VL region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering and a histidine (H) at position 90 based on Kabat numbering (e.g., S28H and Q91H mutations compared to SEQ ID NO: 118) in the VL region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 30 based on Kabat numbering and a histidine (H) at position 31 based on Kabat numbering (e.g., S31H and S32H mutations compared to SEQ ID NO: 118) in the VL region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 30 based on Kabat numbering and a histidine (H) at position 90 based on Kabat numbering (e.g., S31H and Q91H mutations compared to SEQ ID NO: 118) in the VL region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 31 based on Kabat numbering and a histidine (H) at position 90 based on Kabat numbering (e.g., S32H and Q91H mutations compared to SEQ ID NO: 118) in the VL region.
[0184] In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering, a histidine (H) at position 30 based on Kabat numbering, and a histidine (H) at position 31 based on Kabat numbering) (e.g., S28H, S31H, S32H mutations compared to SEQ ID NO: 118 (S27aH, S30H, S31H mutations compared to SEQ ID NO: 118, based on Kabat numbering)) in the VL region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding
fragment thereof comprises a histidine (H) at position 28 (position 27a based on Kabat numbering), a histidine (H) at position 30 based on Kabat numbering, and a histidine (H) at position 90 based on Kabat numbering (e.g., S28H, S31H, Q91H mutations compared to SEQ ID NO: 118) in the VL region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering, a histidine (H) at position 31 based on Kabat numbering, and a histidine (H) at position 90 based on Kabat numbering (e.g., S28H, S32H, Q91H mutations compared to SEQ ID NO: 118) in the VL region. In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 30 based on Kabat numbering, a histidine (H) at position 31 based on Kabat numbering, and a histidine (H) at position 90 based on Kabat numbering (e.g., S31H, S32H, Q91H mutations compared to SEQ ID NO: 118) in the VL region.
[0185] In some embodiments, the provided anti-CTLA-4 antibody or antigen-binding fragment thereof comprises a histidine (H) at position 27a based on Kabat numbering, a histidine (H) at position 30 based on Kabat numbering, a histidine (H) at position 31 based on Kabat numbering, and a histidine (H) at position 91 (position 90 based on Kabat numbering) (e.g., S28H, S31H, S32H, Q91H mutations compared to SEQ ID NO: 118) in the VL region.
[0186] In some embodiments, the VH region comprises an arginine (R) at position 31, glutamic acid (E) or histidine (H) at position 53, histidine (H) at position 58, histidine (H) at position 97, or histidine (H) at position 98, respectively, based on Kabat numbering, or any combination thereof (e.g., S31R, D54H, D54E, Y59H, W101H, or L102H mutations, or any combination thereof, compared to SEQ ID NO:112), and the VL region comprises a histidine (H) at position 27a, histidine (H) at position 30, histidine (H) at position 31, or histidine (H) at position 90, respectively, based on Kabat numbering), or any combination thereof (e.g., S28H, S31H, S32H, or Q91H mutations, or any combination thereof, compared to SEQ ID NO: 118).
[0187] Exemplary antibody VH and VL regions with one or more of the above residues are set forth in Tables 3A and 3B. Provided herein are any antibody or antigen binding fragment comprising at least one of such VH or VL region set forth in Table 3A or Table 3B, paired with any respective other VH and VL described herein. Also provided are antibodies or antigen binding fragments containing a paired combination of any VH set forth in Table 3A and any VL set forth in Table 3B. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in any one of SEQ ID NOs: 5, 6, 14, 17-19, and 21-27, and a VL set
forth in any one of SEQ ID NOs: 29, 30, 32, 34, 39-41, 44, and 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 5 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 6 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in
SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 14 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 17 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID
NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 18 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 19 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID
NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 21 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 22 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID
NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 23 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 24 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 25 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID
NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 26 and a VL set forth in SEQ ID NO: 47. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 29. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 30. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 32. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 34. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 39. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 40. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 41. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 44. In some embodiments, the antibody or antigen binding fragment comprises a VH set forth in SEQ ID NO: 27 and a VL set forth in SEQ ID NO: 47. The above examples are non-limiting and any combinations are contemplated and provided by the present disclosure.
[0188] Also provided herein are antibodies that contain a VH region sequence that exhibits a sequence identity of at least 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% to any of such VH region sequences in Table 3A, in which one
of the pH residues in Table 3A is present. Also provided herein are antibodies that contain a VL region sequence that exhibits a sequence identity of at least 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% to any of such VL region sequences in Table 3B, in which one of the pH residues in Table 3B is present. Also provided herein are antibodies that contain a VH region sequence that exhibits a sequence identity of at least 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% to any of such VH region sequences in Table 3A, in which one of the pH residues in Table 3A is present; and a VL region sequence that exhibits a sequence identity of at least 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% or 100% to any of such VL region sequences in Table 3B, in which one of the pH residues in Table 3B is present.
[0189] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89 and 98, respectively. In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises the sequence set forth in SEQ ID NO: 25 and the VL comprises the sequence set forth in SEQ ID NO: 29.
[0190] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67 and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89 and 99, respectively. In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises the sequence set forth in SEQ ID NO: 23 and the VL comprises the sequence set forth in SEQ ID NO: 39.
[0191] In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89 and 99, respectively. In some embodiments of the anti-CTLA-4 antibody or antigen-binding fragment, the VH region comprises the sequence set forth in SEQ ID NO: 25 and the VL comprises the sequence set forth in SEQ ID NO: 39.
[0192] In certain embodiments, antibody variable domains as described are fused to immunoglobulin constant domain sequences. In certain embodiments, the fusion is with an Ig heavy chain constant domain, comprising at least part of the hinge, CH2, and CH3 regions. In some of any of the embodiments herein, the antibody further comprises a heavy chain constant
domain and/or a light chain constant domain. In some embodiments, the heavy chain and/or light constant domain is murine or human.
[0193] A “constant region” or “constant domain” is a domain in an antibody heavy or light chain that contains a sequence of amino acids that is comparatively more conserved among antibodies than the variable region domain. The constant regions of immunoglobulins show less sequence diversity than the variable regions, and are responsible for binding a number of natural proteins to elicit important biochemical events. Each light chain has a single light chain constant region (CL) domain, which is either of the kappa or lambda type. Each heavy chain contains one or more heavy chain constant region (CH) domains that can be assigned to different classes or isotypes. There are five classes of immunoglobulins: IgA, IgD, IgE, IgG and IgM, having heavy chains designated ex, 8, e, y and p., respectively. Full-length IgA, IgD and IgG isotypes contain CHI, CH2, CH3 and a hinge region, while IgE and IgM contain CHI, CH2, CH3 and CH4. The y and ex classes are further divided into subclasses on the basis of relatively minor differences in the CH sequence and function, e.g., humans express the following subclasses: IgGl, IgG2, IgG3, IgG4, IgAl and IgA2. IgGl antibodies can exist in multiple polymorphic variants termed allotypes (reviewed in Jefferis and Lefranc 2009. mAbs Vol 1 Issue 4 1-7) any of which are suitable. In humans there are five different classes of antibodies including IgA (which includes subclasses IgAl and IgA2), IgD, IgE, IgG (which includes subclasses IgGl, IgG2, IgG3, and IgG4), and IgM. The distinguishing features between these antibody classes are their constant, Fc regions, although subtler differences may exist in the V region. An antibody constant region can include an Fc portion. Although the boundaries of the Fc region of an immunoglobulin heavy chain might vary, the human IgG heavy-chain Fc region usually includes the region containing the CH2 and CH3 domains and the hinge region, such as an amino acid residue at position Cys226, or from Pro230, to the carboxyl-terminus thereof.
[0194] In certain embodiments, a provided antibody contains an Ig heavy chain constant domain comprising at least part of the hinge, CH2, and CH3 regions. Certain embodiments have the first heavy-chain constant region (CHI) containing the site necessary for light chain bonding, present in at least one of the fusions. DNAs encoding the immunoglobulin heavy chain fusions and, if desired, the immunoglobulin light chain, are inserted into separate expression vectors, and are co-transfected into a suitable host cell. This provides for greater flexibility in adjusting the mutual proportions of the three polypeptide fragments in
embodiments when unequal ratios of the three polypeptide chains used in the construction provide the optimum yield of the desired fusion antibody. It is, however, possible to insert the coding sequences for two or all three polypeptide chains into a single expression vector when the expression of at least two polypeptide chains in equal ratios results in high yields or when the ratios have no significant effect on the yield of the desired chain combination.
[0195] In some embodiments, the antibody or antigen-binding fragment thereof, may contain at least a portion of an immunoglobulin constant region, such as one or more constant region domain. In some embodiments, the constant regions include a light chain constant region and/or a heavy chain constant region 1 (CHI). In some embodiments, the antibody includes at least a portion of a hinge region or a variant thereof. In some embodiments, the antibody includes a CH2 and/or CH3 domain, such as an Fc region. In some embodiments, the Fc region is an Fc region from an IgAl, IgA2, IgD, IgE, IgGl, IgG2, IgG3, IgG4, or IgM.
[0196] In some embodiments, the heavy chain includes a variable heavy chain as described that is joined to a human constant region. In some embodiments, the human constant region includes the CH1-CH2-CH3 constant domains. In some embodiments, the human constant region is human IgGl or is derived from human IgGl, or is or is derived from a naturally occurring variant thereof. In some embodiments, the naturally occurring variant is an IgG allotype for instance with the K97R, D239E, or L241M mutation.
[0197] In some embodiments, the heavy chain constant region is mutated or modified, i.e., is a modified constant region. In some embodiments, the heavy chain constant region is a modified human IgG heavy chain constant region.
[0198] In some cases, the mutations include one or more amino acid substitutions in the heavy chain constant region, such as the heavy chain constant region of IgGl, to increase effector activity of the heavy chain constant region. In some embodiments, increased effector activity is an enhanced antibody dependent cell cytotoxicity (ADCC), such as mediated by FcyR-expressing cells. In some embodiments, the one or more amino acid substitutions results in binding to FcyRI. FcyRIIa or FcyRIIIa that is enhanced compared to an antibody in which the heavy chain constant region does not contain the one or more amino acid substitutions. In some embodiments, the enhanced binding to the Fcyl, Fcylla and/or Fey Illa receptor results in increased ADCC. Since an activity of the provided antibodies is to mediate depletion of CTLA- 4-expressing cells, such as Tregs, via ADCC, enhanced binding to the FcyR (e.g, FcyRI. FcyRIIa or FcyRIIIa) mediates a reduction of Tregs at the tumor site. Exemplary amino acid
I l l
substitutions in the heavy chain constant region to enhance FcR binding and/or ADCC can include any as described in U.S. Patent No. US7317091 or U.S. Patent No. 10196445, or any combination of any such described amino acid substitutions.
[0199] In in some embodiments, the heavy chain constant region contains at least one amino acid substitution selected from the group consisting of P230A, E233D, L234E, L234Y, L234I, L235D, L235S, L235Y, L235I, S239D, S239E, S239N, S239Q, S239T, V240I, V240M, F243L, V264I, V264T, V264Y, V266I, E272Y, K274T, K274E, K274R, K274L, K274Y, F275W, N276L, Y278T, V302I, E318R, S324D, S324I, S324V, N325T, K326I, K326T, L328M, L328I, L328Q, L328D, L328V, L328T, A330Y, A330L, A330I, I332D, I332E, I332N, I332Q, T335D, T335R, and T335Y, wherein the numbering of the residues in the Fc region is that of the EU index as in Kabat. In some embodiments, the heavy chain constant region contains at amino acid substitutions V264I, F243E/V264I, E328M, I332E, E328M/I332E, V264VI332E, S298A/I332E, S239E/I332E, S239Q/I332E, S239E, A330Y, I332D, E328VI332E, E328Q/I332E, V264T, V240I, V266I, S239D, S239D/I332D, S239D/I332E, S239D/I332N, S239D/I332Q, S239E/I332D, S239E/I332N, S239E/I332Q, S239N/I332D, S239N/I332E, S239Q/I332D, A330Y/I332E, V264VA330Y/I332E,
A330E/I332E, V264VA330E/I332E, E234E, E234Y, E234I, E235D, E235S, E235Y, E235I, S239T, V240M, V264Y, A330I, N325T, E328D/I332E, E328V/I332E, E328T/I332E, E328VI332E, S239E/V264VI332E, S239Q/V264VI332E, S239E/V264VA330Y/I332E, S239D/A330Y/I332E, S239N/A330Y/I332E, S239D/A330E/I332E, S239N/A330E/I332E, V264VS298A/I332E, S239D/S298A/I332E, S239N/S298A/I332E, S239D/V264VI332E, S239D/V264VS298A/I332E, S239D/V264VA330E/I332E, S239D/I332E/A330I, P230A, P230A/E233D/I332E, E272Y, K274T, K274E, K274R, K274E, K274Y, F275W, N276E, Y278T, V302I, E318R, S324D, S324I, S324V, K326I, K326T, T335D, T335R, T335Y, V240VV266I, S239D/A330Y/I332E/E234I, S239D/A330Y/I332E/E235D,
S239D/A330Y/I332E/V240I, S239D/A330Y/I332E/V264T, S239D/A330Y/I332E/K326E, or S239D/A330Y/I332E/K326T, wherein the numbering of the residues in the Fc region is that of the EU index as in Kabat.
[0200] In some embodiments, the mutation in the IgG (e.g., IgGl) heavy chain to enhance binding to FcyR-expressing cells is selected from one or more mutations G236A, S239D, S298A, A330E, I332E, E333A and K334A, wherein the numbering of the residues in the Fc region is that of the EU index as in Kabat. In some embodiments, the mutation in the IgG heavy
chain to enhance binding to FcyR-expressing cells is selected from mutations S239D and I332E. In some embodiments, the mutation in the IgG heavy chain to enhance binding to FcyR- expressing cells is selected from mutations S239D, A330L and I332E. In some embodiments, the mutation in the IgG heavy chain to enhance binding to FcyR-expressing cells is selected from mutations G236A, A330L and I332E. In some embodiments, the mutation in the IgG heavy chain to enhance binding to FcyR-expressing cells is selected from mutations G236A, S239D, A330E and I332E. In some embodiments, the mutation in the IgG heavy chain to enhance binding to FcyR-expressing cells is selected from mutations S298A, E333A and K334A.
[0201] In some embodiments, the mutation in the IgG heavy chain increase the affinity of IgG for the neonatal Fc receptor, FcRn, which prolongs the in vivo half-life of antibodies and results in increased anti-tumor activity. In some embodiments, the mutations in the IgG is M428E and/or N434S.
[0202] In some cases, the mutations include one or more amino acid substitutions to reduce effector activity of the heavy chain constant region. In some embodiments, the heavy chain constant region is modified to reduce effector activity of the antibody. In some embodiments, the modified human IgG heavy chain constant region has reduced effector activity. In some embodiments, effector activity is reduced compared to a wild-type human IgG heavy chain constant region. In some of any of the described embodiments, the modified human IgG heavy chain constant region is modified compared to the constant region of wildtype human IgGl. In some embodiments, the modified human IgG heavy chain constant region has a sequence that is modified by one or more amino acid substitutions compared to SEQ ID NO: 196 (or a naturally occurring variant thereof, e.g. with K97R, D239E or E241M mutations) and that exhibits at least 85%, at least 90%, at least 95% or at least 98% sequence identity to SEQ ID NO: 196 or the natural variant thereof and contains the one or more amino acid substitutions, for example to reduce effector activity of the heavy chain constant region.
[0203] Various examples of mutations to heavy chain constant regions to alter, such as reduce, effector function are known, including any as described below. In some embodiments, reference to amino acid substitutions in a heavy chain constant region is by EU numbering by Kabat (also called Kabat numbering) unless described with reference to a specific SEQ ID NO. EU numbering is known and is according to the most recently updated IMGT Scientific Chart (IMGT®, the international ImMunoGeneTics information system®,
http://www.imgt.org/IMGTScientificChart/Numbering/Hu_IGHGnber.html (created: 17 May 2001, last updated: 10 Jan 2013) and the EU index as reported in Kabat, E.A. et al. Sequences of Proteins of Immunological interest. 5th ed. US Department of Health and Human Services, NIH publication No. 91-3242 (1991).
[0204] In some embodiments, the heavy chain constant region is modified to alter antibody-dependent cellular cytotoxicity (ADCC) and/or complement-dependent cytotoxicity (CDC), e.g., the amino acid modifications described in Natsume et al., 2008 Cancer Res, 68(10): 3863-72; Idusogie et al., 2001 J Immunol, 166(4): 2571-5; Moore et al., 2010 mAbs, 2(2): 181-189; Lazar et al., 2006 PNAS, 103(11): 4005-4010, Shields et al., 2001 JBC, 276(9): 6591-6604; Stavenhagen et al., 2007 Cancer Res, 67(18): 8882-8890; Stavenhagen et al., 2008 Advan. Enzyme Regul., 48: 152-164; Alegre et al, 1992 J Immunol, 148: 3461-3468; Reviewed in Kaneko and Niwa, 2011 Biodrugs, 25(1): 1-11.
[0205] In some embodiments, the heavy chain constant region is altered at one or more of the following positions to reduce Fc receptor binding: Leu 234 (L234), Leu235 (L235), Asp265 (D265), Asp270 (D270), Ser298 (S298), Asn297 (N297), Asn325 (N325), Ala327 (A327) or Pro329 (P329). For example, Leu 234Ala (L234A), Leu235Ala (L235A), Leu235Glu (L235E), Asp265Asn (D265N), Asp265Ala (D265A), Asp270Asn (D270N), Ser298Asn (S298N), Asn297Ala (N297A), Pro329Ala (P329A) or Pro239Gly (P329G), Asn325Glu (N325E) orAla327Ser (A327S). In some embodiments, modifications within the heavy chain constant region reduce binding to Fc-receptor-gamma receptors while have minimal impact on binding to the neonatal Fc receptor (FcRn).
[0206] In some embodiments, the heavy chain constant region is modified at amino acid Asn297 (Kabat Numbering) to prevent glycosylation of the chimeric antibody, e.g., Asn297Ala (N297A) or Asn297Asp (N297D). In some embodiments, the heavy chain constant region is modified at amino acid Leu235 (Kabat Numbering) to alter Fc receptor interactions, e.g., Leu235Glu (L235E) or Leu235Ala (L235A). In some embodiments, the heavy chain constant region of the chimeric antibody is modified at amino acid Leu234 (Kabat Numbering) to alter Fc receptor interactions, e.g., Leu234Ala (L234A). In some embodiments, the heavy chain constant region of the chimeric antibody is modified at amino acid Leu234 (Kabat Numbering) to alter Fc receptor interactions, e.g., Leu235Glu (L235E). In some embodiments, the heavy chain constant region of the chimeric antibody is altered at both amino acids 234 and 235, e.g., Leu234Ala and Leu235Ala (L234A/L235A) or Leu234Val and Leu235Ala (L234V/L235A).
In some embodiments, the modified heavy chain constant region comprises Leu234Ala and Leu235Ala (L234A/L235A) mutations. In some embodiments, the heavy chain constant region of the chimeric antibody is altered at amino acids 234, 235, and 297, e.g., Leu234Ala, Leu235Ala, Asn297Ala (L234A/L235A/N297A). In some embodiments, the heavy chain constant region of the chimeric antibody is altered at amino acids at 234, 235, and 329, e.g., Leu234Ala, Leu235Ala, Pro239Ala (L234A/L235A/P329A). In some embodiments, the heavy chain constant region of the chimeric antibody is modified at amino acid Asp265 (Kabat Numbering) to alter Fc receptor interactions, e.g. Asp265Ala (D265A). In some embodiments, the heavy chain constant region of the chimeric antibody is modified at amino acid Pro329 (Kabat Numbering) to alter Fc receptor interactions, e.g. Pro329Ala (P329A) or Pro329Gly (P329G). In some embodiments, the heavy chain constant region of the chimeric antibody is altered at both amino acids 265 and 329, e.g., Asp265Ala and Pro329Ala (D265A/P329A) or Asp265Ala and Pro329Gly (D265A/P329G). In some embodiments, the heavy chain constant region of the chimeric antibody is altered at amino acids at 234, 235, and 265, e.g., Leu234Ala, Leu235Ala, Asp265Ala (L234A/L235A/D265A). In some embodiments, the heavy chain constant region of the chimeric antibody is altered at amino acids at 234, 235, and 329, e.g., Leu234Ala, Leu235Ala, Pro329Gly (L234A/L235A/P329G). In some embodiments, the heavy chain constant region of the chimeric antibody is altered at amino acids at 234, 235, 265 and 329, e.g., Leu234Ala, Leu235Ala, Asp265Ala, Pro329Gly
(L234A/L235A/D265A/P329G). In some embodiments, the heavy chain constant region of the chimeric antibody is altered at Gly235 to reduce Fc receptor binding. For example, wherein Gly235 is deleted from the heavy chain constant region of the chimeric antibody. In some embodiments, the heavy chain constant region of the chimeric antibody is modified at amino acid Gly236 to enhance the interaction with CD32A, e.g., Gly236Ala (G236A). In some embodiments, the heavy chain constant region of the chimeric antibody lacks Lys447 (EU index of Kabat et al 1991 Sequences of Proteins of Immunological Interest).
[0207] In some embodiments, the heavy chain constant region of the chimeric antibody is lacking an amino acid at one or more of the following positions to reduce Fc receptor binding: Glu233 (E233), Leu234 (L234), or Leu235 (L235). In some embodiments, the heavy chain constant region of the chimeric antibody is lacking an amino acid at one or more of the following positions Glu233 (E233), Leu234 (L234), or Leu235 (L235), and is modified at one or more of Asp265 (D265), Asn297 (N297), or Pro329 (P329), to reduce Fc receptor binding.
In some embodiments, the heavy chain constant region of the chimeric antibody comprises a three amino acid deletion in the lower hinge corresponding to IgGl E233, L234, and L235. In some aspects, such heavy chain constant regions s do not engage FcyRs and thus are referred to as “effector silent” or “effector null.”
[0208] In some embodiments, the modified heavy chain constant region includes Leu234Ala and Leu235Ala (L234A/L235A) mutations.
[0209] In some embodiments, the heavy chain constant sequence includes an IgG4 containing the S228P mutation, which has been shown to prevent recombination between a therapeutic antibody and an endogenous IgG4 by Fab-arm exchange (see e.g. Fabrijin et al. (2009) Nat. Biotechnol., 27(8): 767-71.). In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 102.
[0210] In some embodiments, the heavy chain constant domain is derived from an IgGl. In some embodiments, the heavy chain constant domain comprises one or more of the mutations selected from the group consisting of G236A, S239D, M252Y, S254T, T256E, A330E, I332E, M428E, and N434S. In some embodiments, the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 101. In some embodiments, the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 106. In some embodiments, the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 107. In some embodiments, the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 108. In some embodiments, the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 109. In some embodiments, the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 110. In some embodiments, the heavy chain constant domain comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 111.
In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 101. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 106. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 107. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 108. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 109. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 110. In some embodiments, the heavy chain constant domain comprises the sequence set forth in SEQ ID NO: 111. Mutations or modifications in the constant heavy chain constant domain could affect the function of the antibody or antigen-binding fragment thereof, including ADCC, ADCP, and CDC (See, e.g., US10196445B1, US7317091B2, US10912831B1, each of which is incorporated herein in its entirety by reference).
[0211] In some embodiments, the light chain constant domain comprises the sequence set forth in SEQ ID NO: 103. In some embodiments, the heavy chain further comprises a signal peptide. In some embodiments, the signal peptide can be cleaved upon expression from a cell. In some embodiments, the heavy chain signal peptide is set forth in SEQ ID NO: 104. In some embodiments, the light chain signal peptide is set forth in SEQ ID NO: 105.
[0212] In some embodiments, the light chain constant region is a kappa light chain. In some embodiments, the kappa light chain comprises the sequence set forth in SEQ ID NO: 103. In some embodiments, the light chain further comprises a signal peptide. In some embodiments, the signal peptide can be cleaved upon expression from a cell. In some embodiments, the signal peptide is set forth in SEQ ID NO: 105.
[0213] In some embodiments, the antibody or antigen-binding fragment thereof is a singlechain antibody fragment, such as a single chain variable fragment (scFv). In some embodiments, the antibody or antigen binding fragment is a multi-domain antibody, such as an scFv comprising a heavy chain variable (VH) region and a light chain variable (VL) region. In some embodiments, the single-chain antibody fragment e.g., scFv) includes one or more linkers joining two antibody domains or regions, such as a heavy chain variable (VH) region and a light chain variable (VL) region. The linker typically is a peptide linker, e.g., a flexible and/or soluble peptide linker. Among the linkers are those rich in glycine and serine and/or in some cases threonine. In some embodiments, the linkers further include charged residues such
as lysine and/or glutamate, which can improve solubility. In some embodiments, the linkers further include one or more proline.
[0214] In some aspects, the linkers rich in glycine and serine (and/or threonine) include at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% such amino acid(s). In some embodiments, they include at least at or about 50%, 55%, 60%, 70%, or 75%, glycine, serine, and/or threonine. In some embodiments, the linker is comprised substantially entirely of glycine, serine, and/or threonine. The linkers generally are between about 5 and about 50 amino acids in length, typically between at or about 10 and at or about 30, e.g., 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, or 30, and in some examples between 10 and 25 amino acids in length. Exemplary linkers include linkers having various numbers of repeats of the sequence GGGGS (4GS; SEQ ID NO: 124) or GGGS (3GS; SEQ ID NO: 125), such as between 2, 3, 4 and 5 repeats of such a sequence. Exemplary linkers include those having or consisting of an sequence set forth in SEQ ID NO: 126 (GGGGSGGGGSGGGGS). Exemplary linkers further include those having or consisting of the sequence set forth in SEQ ID NO: 127 (GSTSGSGKPGSGEGSTKG). Exemplary linkers further include those having or consisting of the sequence set forth in SEQ ID NO: 128 (SRGGGGSGGGGSGGGGSLEMA). An exemplary linker includes those having or consisting of the sequence set forth in SEQ ID NO: 129 (GSRGGGGSGGGGSGGGGSLEMA).
B. Exemplary Features
[0215] In some aspects, the provided antibodies have one or more specified functional features, such as binding properties, including binding to particular epitopes or exhibiting lower or reduced binding to a related but non-specific antigen. In some aspects, the provided antibodies can bind to an epitope that is similar to or overlaps with epitopes of other antibodies, such as reference antibodies, and/or exhibit particular binding affinities. In some aspects, the provided antibodies can bind to an epitope that is different from epitopes of other antibodies, e.g., binding a conformational epitope.
[0216] In some embodiments, the provided antibodies or antigen-binding fragment thereof specifically bind to a cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) protein. In some of any of the embodiments provided herein, CTLA-4 refers to human CTLA-4. In particular embodiments, the provided antibodies and antibody fragments specifically bind to human CTLA-4. A molecule is said to exhibit "specific binding" or "preferential binding" if it reacts
or associates more frequently, more rapidly, with greater duration and/or with greater affinity with a particular cell or substance (e.g. a cell expressing a CTLA-4 antigen or a recombinant CTLA-4 antigen) than it does with alternative cells or substances. An antibody specifically binds or preferentially binds to a target (i.e. CTLA-4) if it binds with greater affinity, avidity, more readily, and/or with greater duration than it binds to other substances. For example, an antibody that specifically or preferentially binds to CTLA-4 is an antibody that binds CTLA-4 with greater affinity, avidity, more readily, and/or with greater duration than it binds to a different protein antigen. It is also understood by reading this definition that specific binding or preferential binding does not necessarily require (although it can include) exclusive binding. Methods to determine such specific or preferential binding are also well known in the art, e.g., an immunoassay.
[0217] The observation that an antibody or other binding molecule binds to CTLA-4 or specifically binds to CTLA-4 does not necessarily mean that it binds to CTLA-4 from every species. For example, in some embodiments, features of binding to CTLA-4, such as the ability to specifically bind thereto and/or to compete for binding thereto with a reference antibody, and/or to bind with a particular affinity or compete to a particular degree, in some embodiments, refers to the ability with respect to a human CTLA-4 protein and the antibody may not have this feature with respect to a CTLA-4 of another species such as mouse.
[0218] In some embodiments, the provided antibodies are capable of binding CTLA-4, such as human CTLA-4, with at least a certain affinity, as measured by any of a number of known methods. In some embodiments, the affinity is represented by absorbance. In some embodiments, the affinity is represented by equilibrium dissociation constant (KD).
[0219] A variety of assays can be used for assessing absorbance. For example, enzyme linked immunosorbent assays (ELISA) is a plate-based assay technique designed for detecting and quantifying soluble substances. In an ELISA, the antigen (target macromolecule) is immobilized on a solid surface (microplate) and then complexed with an antibody that is linked to a reporter enzyme. Detection is accomplished by measuring the activity of the reporter enzyme via incubation with the appropriate substrate to produce a measurable product.
[0220] A variety of assays are known for assessing binding affinity, equilibrium dissociation constant (KD), equilibrium association constant (KA), ECSO, on-rate (association rate constant; kon or ka; units of 1/Ms or M^s'1) and the off-rate (dissociation rate constant; koff or ka; units of 1/s or s'1) and/or determining whether a binding molecule (e.g., an antibody or
fragment thereof) specifically binds to a particular ligand (e.g., an antigen, such as a CTLA-4 protein). One can determine the binding affinity of a binding molecule, e.g., an antibody or an antigen-binding fragment thereof, for an antigen, e.g., CTLA-4, such as human CTLA-4, such as by using any of a number of binding assays that are well known. For example, in some embodiments, a BIAcore® instrument can be used to determine the binding kinetics and constants of a complex between two proteins (e.g., an antibody or fragment thereof, and an antigen, such as a CTLA-4 protein), using surface plasmon resonance (SPR) analysis (see, e.g., Scatchard et al., Ann. N.Y. Acad. Sci. 51:660, 1949; Wilson, Science 295:2103, 2002; Wolff et al., Cancer Res. 53:2560, 1993; and U.S. Patent Nos. 5,283,173, 5,468,614, or the equivalent).
[0221] SPR measures changes in the concentration of molecules at a sensor surface as molecules bind to or dissociate from the surface. The change in the SPR signal is directly proportional to the change in mass concentration close to the surface, thereby allowing measurement of binding kinetics between two molecules. The dissociation rate constant (koff or kd), the association rate constant (kon or ka) and/or equilibrium dissociation constant (KD) and/or equilibrium association constant (KA) for the complex can be determined by monitoring changes in the refractive index with respect to time as buffer is passed over the chip. Other suitable assays for measuring the binding of one protein to another include, for example, immunoassays such as enzyme linked immunosorbent assays (ELISA) and radioimmunoassays (RIA), or determination of binding by monitoring the change in the spectroscopic or optical properties of the proteins through fluorescence, UV absorption, circular dichroism, or nuclear magnetic resonance (NMR). Other exemplary assays include, but are not limited to, Western blot, analytical ultracentrifugation, spectroscopy, flow cytometry, sequencing, genetic reporter assays, flow cytometry, and other methods for detection of expressed nucleic acids or binding of proteins.
[0222] In some embodiments, binding can be determined under conditions that mimic those that exist in natural environments in the body of a subject. For instance, acidic pH such as pH about 5.0 mimics more closely the endosomes in a cell. In contrast, physiological pH is a more neutral pH 7.35 to 7.45, such as pH about 7.4 and more closely mimics the environment of normal tissues or cells in the blood.
[0223] In some embodiments, the antibody or fragment exhibits a similar CTLA-4 binding at an acidic pH as present in the microenvironment of tumor (such as at pH 5.0 to 6.8, e.g. pH 6.0 or pH 5.0) compared to CTLA-4 binding at normal pH as present in blood or a normal
tissue (such as at pH 7.35 to 7.45, e.g. pH 7.4). In some embodiments, the KD at an acidic pH as present in the microenvironment of tumor (such as at pH 5.0 to 6.8, e.g. pH 6.0 or pH 5.0) is about the same (e.g. 0.7-fold to 1.3-fold) the KD at pH of a non-tumor microenvironment (such as at pH 7.35 to 7.45, e.g. 7.4) as present at normal pH of blood or a normal tissue.
[0224] In some embodiments, properties or features of the provided antibodies are described in relation to properties observed for another antibody, e.g., a reference antibody. In some embodiments, the reference antibody is a human or humanized anti-CTLA-4 antibody. Exemplary reference anti-CTLA-4 antibodies include ipilimumab (Vn and VL set forth in SEQ ID Nos: 112 and 118, respectively), tremelimumab (Vn and VL set forth in SEQ ID NOs: 113 and 119, respectively), botensilimab (Vn and VL set forth in SEQ ID NOs: 114 and 120, respectively), gotistobart (Vn and VL set forth in SEQ ID NOs: 115 and 121, respectively), quavonlimab (Vn and VL set forth in SEQ ID NOs: 116 and 122, respectively), ipi.101 (Vn and VL set forth in SEQ ID NOs: 117 and 123, respectively). In some embodiments, the provided antibody contains heavy and light chain CDRs that are distinct from the CDRs present in the reference antibody or antibodies.
[0225] In some embodiments, a provided antibody or fragment exhibits a lower or similar binding affinity (e.g. lower absorbance) than the reference antibody to CTLA-4 at an acidic pH as present in the endosomes of a cell (such as at pH 5.0).
[0226] In some embodiments, a provided antibody or fragment exhibits a higher CTLA-4 binding compared to a reference antibody to CTLA-4 at a neutral pH (such as at pH 7.4) as present at normal pH of blood or a normal tissue. In some embodiments, the CTLA-4 binding of a provided antibody or fragment at a neutral pH as present in the blood or a normal tissue (such as at pH 7.4) is higher by at least or 1.2, 1.3, 1.4, 1.5, 1.6, 1.7, 1.8, 1.9, 2.0, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 30, 40, 50 or more compared to the reference antibody for CTLA-4 binding at the same pH or condition. In some examples, the CTLA-4 binding is higher by at least 5%, 10%, 15%, 20%, 25%, 35%, 50%, 100%, 2-fold, 5-fold, 10- fold, 20-fold or more. In some embodiments, the absorbance of a provided antibody for CTLA- 4 binding in an ELISA or other binding assay at a neutral pH as present in the blood or a normal tissue (such as at pH 7.4) is about 1.5-fold, 2-fold, 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold, 10-fold, 20-fold, 25-fold, 30-fold, 40-fold, 50-fold or 100-fold higher than the absorbance of the reference antibody for CTLA-4 binding at the same pH or condition. In some embodiments, the EC50 of a provided antibody for CTLA-4 binding at an acidic pH as present
within a tumor microenvironment or endosome of a cell (such as at pH 5.0) is about 1.5-fold, 2-fold, 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold, 10-fold, 20-fold, 25-fold, 30-fold, 40-fold, 50-fold or 100-fold higher than the EC50 of the reference antibody for CTLA-4 binding at the same pH or condition. In some embodiments, the KD of provided antibody for CTLA-4 binding at an acidic pH as present within a tumor microenvironment or endosome of a cell (such as at pH 5.0) is about 1.5-fold, 2-fold, 3-fold, 4-fold, 5-fold, 6-fold, 7-fold, 8-fold, 9-fold, 10-fold, 20-fold, 25-fold, 30-fold, 40-fold, 50-fold or 100-fold higher than the KD of the reference antibody for CTLA-4 binding at the same pH or condition.
[0227] In some embodiments, the provided antibody or antigen-binding fragment thereof can impact safety and efficacy through involvement with the broader immune system. In some embodiments, the provided antibody or antigen-binding fragment thereof induces a favorable ratio of immune cell subsets in the tumor or blood, such as T cells, CD4+ T cells, CD8+ T cells, and/or Treg cells. In some embodiments, the provided antibody or antigen-binding fragment induces a favorable ratio of CTLA-4 expression on immune cell subsets in the tumor or blood, such as T cells, CD4+ T cells, CD8+ T cells, and/or Treg cells.
C. Variants and Modifications
[0228] Amino acid sequence modification(s) of the antibodies described herein are contemplated. Lor example, it may be desirable to improve the binding affinity and/or other biological properties of the antibody. Lor example, amino acid sequence variants of an antibody may be prepared by introducing appropriate nucleotide changes into a polynucleotide that encodes the antibody, or a chain thereof, or by peptide synthesis. Such modifications include, for example, deletions from, and/or insertions into and/or substitutions of, residues within the amino acid sequences of the antibody. Any combination of deletion, insertion, and substitution may be made to arrive at the final antibody, provided that the final construct possesses the desired characteristics (e.g., high affinity binding to CTLA-4). The amino acid changes may alter post-translational processes of the antibody, such as changing the number or position of glycosylation sites. Any of the variations and modifications described above for polypeptides of the present invention may be included in antibodies of the present invention.
[0229] In certain embodiments, the antibodies include one or more amino acid variations, e.g., substitutions, deletions, insertions, and/or mutations, compared to the sequence of an antibody described herein. Exemplary variants include those designed to improve the binding affinity and/or other biological properties of the antibody. Amino acid sequence variants of an
antibody may be prepared by introducing appropriate modifications into the nucleotide sequence encoding the antibody, or by peptide synthesis. Such modifications include, for example, deletions from, and/or insertions into and/or substitutions of residues within the amino acid sequences of the antibody. Any combination of deletion, insertion, and substitution can be made to arrive at the final construct, provided that the final construct possesses the desired characteristics, e.g., antigen-binding.
[0230] In certain embodiments, the antibodies include one or more amino acid substitutions, e.g., as compared to an antibody sequence described herein and/or compared to a sequence of a natural repertoire, e.g., human repertoire. Sites of interest for substitutional mutagenesis include the CDRs and FRs. Amino acid substitutions may be introduced into an antibody of interest and the products screened for a desired activity, e.g., retained/improved antigen binding, decreased immunogenicity, improved half-life, and/or improved effector function, such as the ability to promote antibody-dependent cellular cytotoxicity (ADCC) or complement-dependent cytotoxicity (CDC).
[0231] In some embodiments, one or more residues within a CDR of a parent antibody (e.g. a humanized or human antibody) is/are substituted. In some embodiments, the substitution is made to revert a sequence or position in the sequence to a germline sequence, such as an antibody sequence found in the germline (e.g., human germline), for example, to reduce the likelihood of immunogenicity, e.g., upon administration to a human subject.
[0232] In some embodiments, alterations are made in CDR “hotspots,” residues encoded by codons that undergo mutation at high frequency during the somatic maturation process (see, e.g., Chowdhury, Methods Mol. Biol. 207:179-196 (2008)), and/or residues that contact antigen, with the resulting variant VH or VL being tested for binding affinity. Affinity maturation by constructing and reselecting from secondary libraries has been described, e.g., in Hoogenboom et al. in Methods in Molecular Biology 178:1-37 (O’Brien et al., ed., Human Press, Totowa, NJ, (2001)). In some embodiments of affinity maturation, diversity is introduced into the variable genes chosen for maturation by any of a variety of methods (e.g., error-prone PCR, chain shuffling, or oligonucleotide-directed mutagenesis). A secondary library is then created. The library is then screened to identify any antibody variants with the desired affinity. Another method to introduce diversity involves CDR-directed approaches, in which several CDR residues (e.g., 4-6 residues at a time) are randomized. CDR residues
involved in antigen binding may be specifically identified, e.g., using alanine scanning mutagenesis or modeling. CDR-H3 and CDR-L3 in particular are often targeted.
[0233] In certain embodiments, substitutions, insertions, or deletions may occur within one or more CDRs so long as such alterations do not substantially reduce the ability of the antibody to bind antigen. For example, conservative alterations (e.g., conservative substitutions as provided herein) that do not substantially reduce binding affinity may be made in CDRs. Such alterations may, for example, be outside of antigen contacting residues in the CDRs. In certain embodiments of the variant VH and VL sequences provided above, each CDR either is unaltered, or contains no more than one, two or three amino acid substitutions.
[0234] Amino acid sequence insertions include amino- and/or carboxyl-terminal fusions ranging in length from one residue to polypeptides containing a hundred or more residues, as well as intrasequence insertions of single or multiple amino acid residues. Examples of terminal insertions include an antibody with an N-terminal methionyl residue. Other insertional variants of the antibody molecule include the fusion to the N- or C-terminus of the antibody to an enzyme or a polypeptide which increases the serum half-life of the antibody.
[0235] In certain embodiments, the antibody is altered to increase or decrease the extent to which the antibody is glycosylated, for example, by removing or inserting one or more glycosylation sites by altering the amino acid sequence and/or by modifying the oligosaccharide(s) attached to the glycosylation sites, e.g., using certain cell lines.
[0236] In some embodiments, an N-linked glycosylation, which is a glycosylation site that occurs at asparagines in the consensus sequence -Asn-Xaa-Ser/Thr is removed or inserted. In some embodiments, one or more re replaced with another amino acid to remove the glycosylation site.
[0237] Exemplary modifications, variants, and cell lines are described, e.g., in Patent Publication Nos. US 2003/0157108, US 2004/0093621, US 2003/0157108; WO 2000/61739; WO 2001/29246; US 2003/0115614; US 2002/0164328; US 2004/0093621; US 2004/0132140; US 2004/0110704; US 2004/0110282; US 2004/0109865; WO 2003/085119; WO 2003/084570; WO 2005/035586; WO 2005/035778; W02005/053742; W02002/031140; Okazaki et al. J. Mol. Biol. 336:1239-1249 (2004); Yamane-Ohnuki et al. Biotech. Bioeng. 87: 614 (2004). Ripka et al. Arch. Biochem. Biophys. 249:533-545 (1986); US Pat Appl No US 2003/0157108 Al, Presta, L; and WO 2004/056312 Al, Yamane-Ohnuki et al. Biotech. Bioeng. 87: 614 (2004); Kanda, Y. et al., Biotechnol. Bioeng., 94(4):680-688 (2006); and
W02003/085107); WO 2003/011878 (Jean-Mairet et al.); US Patent No. 6,602,684 (Umana et al.); and US 2005/0123546 (Umana et al.); WO 1997/30087 (Patel et al.); WO 1998/58964 (Raju, S.); and WO 1999/22764 (Raju, S.).
[0238] Among the modified antibodies are those having one or more amino acid modifications in the Fc region, such as those having a human Fc region sequence or other portion of a constant region (e.g., a human IgGl, IgG2, IgG3 or IgG4 Fc region) comprising an amino acid modification (e.g. a substitution) at one or more amino acid positions.
[0239] Such modifications can be made, e.g., to improve half-life, alter binding to one or more types of Fc receptors, and/or alter effector functions.
[0240] Also among the variants are cysteine engineered antibodies such as “thioMAbs” and other cysteine engineered variants, in which one or more residues of an antibody are substituted with cysteine residues, in order to generate reactive thiol groups at accessible sites, e.g., for use in conjugation of agents and linker-agents, to produce immunoconjugates. Cysteine engineered antibodies are described, e.g., in U.S. Patent Nos. 7,855,275 and 7,521,541.
[0241] In some embodiments, the antibodies are modified to contain additional nonproteinaceous moieties, including water soluble polymers. Exemplary polymers include, but are not limited to, polyethylene glycol (PEG), copolymers of ethylene glycol/propylene glycol, carboxymethylcellulose, dextran, polyvinyl alcohol, polyvinyl pyrrolidone, poly-1, 3- dioxolane, poly-l,3,6-trioxane, ethylene/maleic anhydride copolymer, polyaminoacids (either homopolymers or random copolymers), and dextran or poly(n-vinyl pyrrolidone)poly ethylene glycol, propropylene glycol homopolymers, prolypropylene oxide/ethylene oxide copolymers, polyoxyethylated polyols (e.g., glycerol), polyvinyl alcohol, and mixtures thereof. Polyethylene glycol propionaldehyde may have advantages in manufacturing due to its stability in water. The polymer may be of any molecular weight, and may be branched or unbranched. The number of polymers attached to the antibody may vary, and if more than one polymer is attached, they can be the same or different molecules. In general, the number and/or type of polymers used for derivatization can be determined based on considerations including, but not limited to, the particular properties or functions of the antibody to be improved, whether the antibody derivative will be used in a therapy under defined conditions, etc.
[0242] Antibodies, and antigen-binding fragments and variants thereof, of the present invention may also be modified to include a detectable label, e.g., an epitope tag or label, e.g.,
for use in purification or diagnostic applications. These may be conjugated to the antibody as a fusion protein or conjugate, e.g., using a linker or linking group. There are many linking groups known in the art for making antibody conjugates, including, for example, those disclosed in U.S. Pat. No. 5,208,020 or EP Patent 0 425 235 Bl, and Chari et al., Cancer Research 52: 127-131 (1992). Linking groups include disulfide groups, thioester groups, acid labile groups, photolabile groups, peptidase labile groups, or esterase labile groups, as disclosed in the above-identified patents. Examples of tags and/or labels can include, but are not limited to, FLAG tags, poly-histidine tags (e.g. 6xHis), cMyc tags, glutathione- S- transferase tags, avidin, fluorescent labels, polymer particles, metal particles, haptens, enzyme labels, luminescent labels, electrochemiluminescent labels, bioluminescent labels, radioisotopes, or oligonucleotides.
D. Binding Molecules
[0243] Also provided herein are binding molecules that comprise any of the provided anti-CTLA-4 antibodies or antigen binding fragments. The binding molecules can include conjugates with another moiety, such as a chemical or other protein molecule. In some embodiments, the other moiety is a protein molecule and the conjugate is a fusion protein. In some embodiments, provided herein are binding molecules that are multispecific antibodies in which at least one antibody binding arm is a provided anti-CTLA-4 antibody or antigen binding fragment.
/. Conjugates
[0244] Provided herein are conjugates containing an anti-CTLA-4 antibody or antigenbinding fragment provided herein and one or more further moiety. The further moiety can be an effector moiety, which provides at least one additional function or property of the conjugate as compared to the unconjugated antibody. In some embodiments, an effector moiety can be any substance having biological or detectable activity, for example, therapeutic agents, detectable labels, binding agents, or prodrugs, which are metabolized to an active agent in vivo. In some embodiments, the moiety can be a targeting moiety, a small molecule drug (nonpolypeptide drug of less than 500 Daltons molar mass), a toxin, a cytostatic agent, a cytotoxic agent, an immunosuppressive agent, a radioactive agent suitable for diagnostic purposes, a radioactive metal ion for therapeutic purposes, a prodrug-activating enzyme, an agent that increases biological half-life, or a diagnostic or detectable agent. In some embodiments, the
effector moiety is a protein, peptide or a nucleic acid molecule, which can be synthesized or produced by recombinant means. In some embodiments, the effector moiety is a drug moiety, which may be synthesized artificially or purified from a natural source.
[0245] In some embodiments, the drug moiety has an intracellular activity. In some embodiments, the anti-CTLA-4 antibody or antigen-binding fragment of the conjugate is internalized and the drug moiety is a cytotoxin that blocks the protein synthesis of the cell, therein leading to cell death. The conjugate can be used for inhibiting the multiplication of a tumor cell or cancer cell, causing apoptosis in a tumor or cancer cell, or for treating cancer in a patient. The conjugate can be used accordingly in a variety of settings for the treatment of animal cancers. The conjugate can be used to deliver a drug to a tumor cell or cancer cell. In some embodiments, upon binding to CTLA-4 on a tumor cell, the conjugate and/or drug can be taken up inside a tumor cell or cancer cell through receptor-mediated endocytosis.
[0246] In some embodiments, the conjugate is an antibody drug conjugate (ADC, also called immunoconjugates) containing an anti-CTLA-4 antibody or antigen-binding fragment provided herein conjugated to a drug moiety that acts as a therapeutic agent, which is either cytotoxic, cytostatic or otherwise provides some therapeutic benefit. In some embodiments, the cytotoxic agent is a chemotherapeutic agent, a drug, a growth inhibitory agent, a toxin (e.g., an enzymatically active toxin of bacterial, fungal, plant, or animal origin, or fragments thereof), or a radioactive isotope (i.e., a radioconjugate). In some embodiments, provided antibody drug conjugates of the present disclosure allow targeted-delivery of the drug moiety to tumors. In some cases, this can result in targeted killing of the tumor cell.
[0247] In some embodiments, the drug moiety is an auristatin, such as auristatin E (also known in the art as a derivative of dolastatin-10) or a derivative thereof. The auristatin can be, for example, an ester formed between auristatin E and a keto acid. For example, auristatin E can be reacted with paraacetyl benzoic acid or benzoylvaleric acid to produce AEB and AEVB, respectively. Other typical auristatins include AFP, MMAF, and MMAE. The synthesis and structure of exemplary auristatins are described in U.S. Pat. Nos. 6,884,869, 7,098,308, 7,256,257, 7,423,116, 7,498,298 and 7,745,394, each of which is incorporated by reference herein in its entirety and for all purposes.
[0248] Auristatins have been shown to interfere with microtubule dynamics and nuclear and cellular division and have anticancer activity. Auristatins of the present invention bind tubulin and can exert a cytotoxic or cytostatic effect on a 5T4 expressing cell or cell line. There
are a number of different assays, known in the art, that can be used for determining whether an auristatin or resultant antibody-drug conjugate exerts a cytostatic or cytotoxic effect on a desired cell or cell line. Methods for determining whether a compound binds tubulin are known in the art. See, for example, Muller et al., Anal. Chem 2006, 78, 4390-4397; Hamel et al., Molecular Pharmacology, 1995 47: 965-976; and Hamel et al., The Journal of Biological Chemistry, 1990 265:28, 17141-17149.
[0249] Examples of drugs or pay loads are selected from the group consisting of DM1 (maytansine, N2'-deacetyl-N2'-(3-mercapto-l -oxopropyl)- or N2'-deacetyl-N2'-(3-mercapto- l-oxopropyl)-maytansine), mc-MMAD (6-maleimidocaproyl-monomethylauristatin-D or N- methyl-L-valyl-N-[(lS,2R)-2-methoxy-4-[(2S)-2-[(lR,2R)-l-methoxy-2-methyl-3-oxo-3- [[(lS)-2-phenyl-l-(2-thiazolyl)ethyl]amino]propyl]-l-pyrrolidinyl]-l-[(lS)-l-methylpropyl]- 4-oxobutyl]-N-methyl-(9Cl)-L-valinamide), mc-MMAF (maleimidocaproyl- monomethylauristatin F or N-[6-(2,5-dihydro-2,5-dioxo-lH-pyrrol-l-yl)-l-oxohexyl]-N- methyl-E-valyl-E-valyl-(3R,4S,5S)-3-methoxy-5-methyl-4-(methylamino)heptanoyl- (aR,pR,2S)-P-methoxy-a-methyl-2-pyrrolidinepropanoyl-E-phenylalanine) and mc-Val-Cit- PABA-MMAE (6-maleimidocaproyl-ValcCit-(p-aminobenzyloxycarbonyl)- monomethylauristatin E or N-[[[4-[[N-[6-(2,5-dihydro-2,5-dioxo-lH-pyrrol-l-yl)-l- oxohexyl]-L-valyl-N5-(aminocarbonyl)-L-omithyl]amino]phenyl]methoxy]carbonyl]-N- methyl-L-valyl-N-[(lS,2R)-4-[(2S)-2-[(lR,2R)-3-[[(lR,2S)-2-hydroxy-l-methyl-2- phenylethyl] amino] - 1 -methoxy-2-methyl-3-oxopropyl] - 1 -pyrrolidinyl] -2-methoxy- 1 - [( 1 S)- 1 - methylpropyl]-4-oxobutyl]-N-methyl-L-valinamide). DM1 is a derivative of the tubulin inhibitor maytansine while MMAD, MMAE, and MMAF are auristatin derivatives. The preferred payloads of the present invention are selected from the group consisting of mc- MMAF and mc-Val-Cit-PABA-MMAE.
[0250] In a further embodiment, the drug moiety is an anti-cancer agent. Exemplary anticancer agents include, but are not limited to, cytostatics, enzyme inhibitors, gene regulators, cytotoxic nucleosides, tubulin binding agents or tubulin inhibitors, proteasome inhibitors, hormones and hormone antagonists, anti-angiogenesis agents, and the like. Exemplary cytostatic anti-cancer agents include alkylating agents such as the anthracycline family of drugs (e.g. adriamycin, carminomycin, cyclosporin- A, chloroquine, methopterin, mithramycin, porfiromycin, streptonigrin, porfiromycin, anthracenediones, and aziridines). Other cytostatic anti-cancer agents include DNA synthesis inhibitors (e.g., methotrexate and
dichloromethotrexate, 3-amino-l,2,4-benzotriazine 1,4-dioxide, aminopterin, cytosine P-D- arabinofuranoside, 5-fluoro-5'-deoxyuridine, 5-fluorouracil, ganciclovir, hydroxyurea, actinomycin-D, and mitomycin C), DNA-intercalators or cross-linkers (e.g., bleomycin, carboplatin, carmustine, chlorambucil, cyclophosphamide, cis-diammineplatinum(II) dichloride (cisplatin), melphalan, mitoxantrone, and oxaliplatin), and DNA-RNA transcription regulators (e.g., actinomycin D, daunorubicin, doxorubicin, homoharringtonine, and idarubicin). Other exemplary cytostatic agents that are compatible with the present disclosure include ansamycin benzoquinones, quinonoid derivatives (e.g. quinolones, genistein, bactacyclin), busulfan, ifosfamide, mechlorethamine, triaziquone, diaziquone, carbazilquinone, indoloquinone EO9, diaziridinyl-benzoquinone methyl DZQ, triethylenephosphoramide, and nitrosourea compounds (e.g. carmustine, lomustine, semustine).
[0251] In some embodiments, there is provided a conjugate comprising anti-CTLA-4 antibody or antigen-binding fragment provided herein conjugated with a toxin. In some embodiments, the therapeutic agent is a cytotoxin comprising a polypeptide having ribosomeinactivating activity including, for example, gelonin, bouganin, saporin, ricin, ricin A chain, bryodin, diphtheria toxin, restrictocin, Pseudomonas exotoxin A and variants thereof.
[0252] In some embodiments, the effector moiety is a radionuclide (e.g., a peptide receptor radionuclide) or a radiolabel. In some embodiments, the effector moiety is a radionuclide or radiolabel with high-energy ionizing radiation that are capable of causing multiple strand breaks in nuclear DNA, leading to cell death. Exemplary high-energy radionuclides include: 90Y, 1251, 1311, 1231, U lin, 105Rh, 153Sm, 67Cu, 67Ga, 166Ho, 177Lu, 186Re and 188Re. These isotopes typically produce high energy a- or P-particles which have a short path length. Such radionuclides kill cells to which they are in close proximity, for example neoplastic cells to which the conjugate has attached or has entered. They have little or no effect on nonlocalized cells and are essentially non-immunogenic. Alternatively, high-energy isotopes may be generated by thermal irradiation of an otherwise stable isotope, for example as in boron neutron-capture therapy (Guan et al., PNAS, 95: 13206-10, 1998).
[0253] In some embodiments, there is provided a conjugate comprising an anti-CTLA-4 antibody or antigen-binding fragment provided herein conjugated with a label, which can generate a detectable signal, indirectly or directly. These conjugates can be used for research or diagnostic applications, such as for the in vivo detection of cancer. The label is preferably
capable of producing, either directly or indirectly, a detectable signal. For example, the label may be radio-opaque or a radioisotope, such as 3H, 14C, 32P, 35S, 1231, 1251, 1311; a fluorescent (fluorophore) or chemiluminescent (chromophore) compound, such as fluorescein isothiocyanate, rhodamine or luciferin; an enzyme, such as alkaline phosphatase, P- galactosidase or horseradish peroxidase; an imaging agent; or a metal ion. In some embodiments, the label is a radioactive atom for scintigraphic studies, for example 99Tc or 1231, or a spin label for nuclear magnetic resonance (NMR) imaging (also known as magnetic resonance imaging, MRI), such as zirconium-89, iodine-123, iodine-131, indium-111, fluorine- 19, carbon- 13, nitrogen- 15, oxygen- 17, gadolinium, manganese or iron. Zirconium-89 may be complexed to various metal chelating agents and conjugated to antibodies, e.g., for PET imaging (WO 2011/056983).
[0254] The conjugates may be prepared using any methods known in the art. See, e.g., WO 2009/067800, WO 2011/133886, and U.S. Patent Application Publication No. 2014322129, incorporated by reference herein in their entirety.
[0255] In some embodiments, the linkage of an antibody or antigen binding fragment to a moiety, such as a drug or toxin or other heterologous moiety, is a direct or indirect linkage. In some embodiments, the attachment can be covalent or non-covalent, e.g., via a biotinstreptavidin non-covalent interaction. For example, a moiety can be attached by alkylation (e.g., at the epsilon-amino group lysines or the N-terminus of antibodies), reductive amination of oxidized carbohydrate, transesterification between hydroxyl and carboxyl groups, amidation at amino groups or carboxyl groups, and conjugation to thiols. In some embodiments, the attachment of a moiety, such as a therapeutic agent, can be by chemical conjugation and linkage methods known in the art. In some embodiments, the moiety can be linked to an antibody by a linker. In some embodiments, linkers such as peptide linkers, cleavable linkers, non-cleavable linkers or linkers that aid in the conjugation reaction, can be used to link or conjugate the effector moieties to the antibody or antigen-binding fragment. Attachment of a linker to an antibody can be accomplished in a variety of ways, such as through surface lysines, reductive- coupling to oxidized carbohydrates, and through cysteine residues liberated by reducing interchain disulfide linkages. A variety of antibody drug conjugate linkage systems are known in the art, including hydrazone-, disulfide- and peptide-based linkages.
[0256] In some embodiments, an anti-CTLA-4 antibody or antigen-binding fragment provided herein is conjugated to one or more moieties, e.g. about 1 to about 20 drug moieties,
through a linker (L). In some embodiments, the conjugate comprises the following components: (antibody or antigen-binding fragment), (L)q and (moiety)m, wherein the antibody or antigen-binding fragment is any of the described capable of specifically binding CTLA-4 as described; L is a linker for linking the protein or polypeptide to the moiety; m is at least 1; q is 0 or more; and the resulting conjugate binds to CTLA-4. In particular embodiments, m is 1 to 4 and q is 0 to 8.
[0257] The linker may be composed of one or more linker components. For covalent attachment of the antibody and the drug moiety the linker typically has two reactive functional groups, i.e. bivalency in a reactive sense. Bivalent linker reagents which are useful to attach two or more functional or biologically active moieties, such as peptides, nucleic acids, drugs, toxins, antibodies, haptens, and reporter groups are known, and methods have been described their resulting conjugates (Hermanson, G. T. (1996) Bioconjugate Techniques; Academic Press: New York, p 234-242).
[0258] In some embodiments, the linker may comprise amino acid residues. Exemplary amino acid linker components include a dipeptide, a tripeptide, a tetrapeptide or a pentapeptide. Exemplary dipeptides include: valine-citrulline (vc or val-cit), alanine-phenylalanine (af or ala- phe). Exemplary tripeptides include: glycine-valine-citrulline (gly-val-cit) and glycine- glycine-glycine (gly-gly-gly). Amino acid residues which comprise an amino acid linker component include those occurring naturally, as well as minor amino acids and non-naturally occurring amino acid analogs, such as citrulline. Amino acid linker components can be designed and optimized in their selectivity for enzymatic cleavage by a particular enzymes, for example, a tumor-associated protease, cathepsin B, C and D, at a plasmin protease.
[0259] Exemplary linker components include 6-maleimidocaproyl (“MC”), maleimidopropanoyl (“MP”), valine-citrulline (“val-cit”), a alanine-phenylalanine (“ala-phe”), p-aminobenzyloxycarbonyl (“PAB”), N-Succinimidyl 4-(2-pyridylthio)pentanoate (“SPP”), N-Succinimidyl 4-(N-maleimidomethyl)cyclohexane-I carboxylate (“SMCC”), and N- Succinimidyl (4-iodo-acetyl)aminobenzoate (“SLAB”).
[0260] Conjugates of an anti-CTLA-4 antibody or antigen-binding fragment provided herein and cytotoxic agent can be made using a variety of bifunctional protein-coupling agents such as N-succinimidyl-3-(2-pyridyldithiol) propionate (SPDP), iminothiolane (IT), bifunctional derivatives of imidoesters (such as dimethyl adipimidate HC1), active esters (such as disuccinimidyl substrate), aldehydes (such as glutaraldehyde), bis-azido compounds (such
as bis(p-azidobenzoyl) hexanediamine), bis-diazonium derivatives (such as bis-(p- diazoniumbenzoyl)-ethylenediamine), diisocyanates (such as toluene 2,6-diisocyanate), and bis-active fluorine compounds (such as l,5-difluoro-2,4-dinitrobenzene).
[0261] The antibody drug conjugate can be prepared by a variety of methods, such as organic chemistry reactions, conditions, and reagents known to those skilled in the art. Alternatively, a fusion protein containing an antibody or antigen-binding fragment and cytotoxic agent may be made, e.g., by recombinant techniques or peptide synthesis. The length of DNA may comprise respective regions encoding the two portions of the conjugate either adjacent one another or separated by a region encoding a linker peptide which does not destroy the desired properties of the conjugate.
2. Multispecific Antibodies
[0262] Provided herein are CTLA-4-binding polypeptides that are multispecific containing at least one antibody or antigen-binding fragment that binds CTLA-4 and one or more additional binding domains. Typically, the one or more additional domains bind to a second antigen or protein other than CTLA-4. In some aspects, the further antigen or protein may be an antigen expressed on a tumor, a molecule or receptor expressed on an immune cell, such as a T cell, e.g. a CD3, or an additional inhibitory receptor (e.g. PD-1, LAG3, TIM3, VISTA, TIGIT, SIRPa, NKG2A, B7H3, B7H4) or an activating receptor (e.g. 0X40, GITR, 41BB, CD40, CD27, CD28 or ICOS) or to confer an additional specificity to a target cell (e.g. CD8 or CD4). In some embodiments, the one or more additional domain is an antibody or antigenbinding fragment specific for the second antigen or protein.
E. Nucleic Acids and Expression Vectors and Methods of Producing Antibodies
[0263] The present disclosure provides a nucleic acid or nucleic acids encoding an antibody or antigen-binding fragment thereof. The nucleic acid(s) of the present disclosure may comprise a polynucleotide sequence encoding any one of the anti-CTLA-4 antibodies or antigen-binding fragments thereof disclosed herein. The polypeptide sequences may be used to determine appropriate nucleic acid sequences encoding the particular antibody disclosed thereby. The nucleic acid sequence may be optimized to reflect particular codon "preferences" for various expression systems according to standard methods well known to those of skill in the art.
[0264] Among the provided nucleic acid molecules are those that comprise polynucleotides that encode one or more polypeptides (e.g., an immunoglobulin heavy chain or light chain) of a provided anti-CTLA-4 antibody. In some embodiments, a nucleic acid molecule comprises a polynucleotide that encodes a heavy chain or a light chain of an anti-CTLA-4 antibody. In some embodiments, a nucleic acid molecule comprises both a polynucleotide that encodes a heavy chain and a polynucleotide that encodes a light chain of an anti-CTLA-4 antibody. In some embodiments, a first nucleic acid molecule comprises a first polynucleotide that encodes a heavy chain and a second nucleic acid molecule comprises a second polynucleotide that encodes a light chain. Also provided herein are nucleic acid molecules comprising polynucleotides that encode one or more polypeptides of a binding molecule containing any of the provided anti-CTLA-4 antibodies, such as a CTLA-4-targeted antibody-drug conjugate (ADC) or a bispecific immune cell engager comprising the anti-CTLA-4 antibody or antigen binding fragment thereof as described herein.
[0265] In some embodiments, the nucleic acid(s) is an isolated nucleic acid(s). The polynucleotides may include those encompassing natural and/or non-naturally occurring nucleotides and bases, e.g., including those with backbone modifications. The terms “nucleic acid molecule”, “nucleic acid”, “sequence of nucleotides”, and “polynucleotide” may be used interchangeably, and refer to a polymer of nucleotides. Such polymers of nucleotides may contain natural and/or non-natural nucleotides, and include, but are not limited to, DNA, RNA, and PNA. “Nucleic acid sequence” refers to the linear sequence of nucleotides that comprise the nucleic acid molecule or polynucleotide.
[0266] In some such embodiments, the heavy chain and the light chain are expressed from one nucleic acid molecule, or from two separate nucleic acid molecules, as two separate polypeptides. In some embodiments, such as when an antibody is an scFv, a single polynucleotide encodes a single polypeptide comprising both a heavy chain variable domain fragment and a light chain variable domain fragment linked together.
[0267] This disclosure is not intended to be limited with regard to the source of the antibody or the manner in which it is made. Various procedures known within the art may be used for the production of monoclonal antibodies directed against CTLA-4 (See, for example, Antibodies: A Laboratory Manual, Harlow E, and Lane D, 1988, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, incorporated herein by reference). In some
embodiments, the anti-CTLA-4 antibodies, and antigen binding fragments thereof, are recombinantly expressed and produced.
[0268] In some embodiments, a nucleic acid of the present disclosure may be operably linked to a transcriptional control element, e.g., a promoter, and enhancer, etc. Suitable promoter and enhancer elements are known to those of skill in the art.
[0269] In some embodiments, a polynucleotide encoding a heavy chain or light chain of an anti-CTLA-4 antibody comprises a nucleotide sequence that encodes a leader sequence, which, when translated, is located at the N terminus of the heavy chain or light chain. The leader sequence may be the native heavy or light chain leader sequence, or may be another heterologous leader sequence.
[0270] Nucleic acid molecules may be constructed using conventional recombinant DNA techniques well-known in the art. In some embodiments, a nucleic acid molecule is an expression vector that is suitable for expression in a selected host cell.
[0271] Vectors comprising polynucleotides that encode anti-CTLA-4 heavy chains and/or anti-CTLA-4 light chains are provided herein. Vectors comprising polynucleotides that encode anti-CTLA-4 heavy chains and/or anti-CTLA-4 light chains are also provided herein. In certain embodiments, the vector is selected from the group consisting of a DNA vector, an RNA vector, a plasmid, a lentiviral vector, an adenoviral vector, an adeno-associated viral vector, and a retroviral vector. In certain embodiments, the vector is an expression vector. A nucleic acid of the present disclosure may be present within an expression vector and/or a cloning vector. An expression vector can include a selectable marker, an origin of replication, and other features that provide for replication and/or maintenance of the vector. Suitable expression vectors include, e.g., plasmids, viral vectors, and the like. Large numbers of suitable vectors and promoters are known to those of skill in the art; many are commercially available for generating a subject recombinant construct. The following vectors are provided by way of example and should not be construed in anyway as limiting: Bacterial: pBs, phagescript, PsiX174, pBluescript SK, pBs KS, pNH8a, pNH16a, pNH18a, pNH46a (Stratagene, La Jolla, Calif., USA); pTrc99A, pKK223-3, pKK233-3, pDR540, and pRIT5 (Pharmacia, Uppsala, Sweden). Eukaryotic: pWLneo, pSV2cat, pOG44, PXR1, pSG (Stratagene) pSVK3, pBPV, pMSG and pSVL (Pharmacia).
[0272] Expression vectors generally have convenient restriction sites located near the promoter sequence to provide for the insertion of nucleic acid sequences encoding heterologous
proteins. A selectable marker operative in the expression host may be present. Suitable expression vectors include, but are not limited to, viral vectors (e.g. viral vectors based on vaccinia virus; poliovirus; adenovirus (see, e.g., Li et al., Invest. Opthalmol. Vis. Sci. (1994) 35: 2543-2549; Borras et al., Gene Ther. (1999) 6: 515-524; Li and Davidson, Proc. Natl. Acad. Sci. USA (1995) 92: 7700-7704; Sakamoto et al., H. Gene Ther. (1999) 5: 1088-1097; WO 94/12649, WO 93/03769; WO 93/19191; WO 94/28938; WO 95/11984 and WO 95/00655); adeno-associated virus (see, e.g., Ali et al., Hum. Gene Ther. (1998) 9: 81-86, Flannery et al., Proc. Natl. Acad. Sci. USA (1997) 94: 6916-6921; Bennett et al., Invest. Opthalmol. Vis. Sci. (1997) 38: 2857-2863; Jomary et al., Gene Ther. (1997) 4:683 690, Rolling et al., Hum. Gene Ther. (1999) 10: 641-648; Ali et al., Hum. Mol. Genet. (1996) 5: 591-594; Srivastava in WO 93/09239, Samulski et al., J. Vir. (1989) 63: 3822-3828; Mendelson et al., Virol. (1988) 166: 154-165; and Flotte et al., Proc. Natl. Acad. Sci. USA (1993) 90: 10613-10617); SV40; herpes simplex virus; human immunodeficiency virus (see, e.g., Miyoshi et al., Proc. Natl. Acad. Sci. USA (1997) 94: 10319-23; Takahashi et al., J. Virol. (1999) 73: 7812-7816); a retroviral vector (e.g., Murine Leukemia Virus, spleen necrosis virus, and vectors derived from retroviruses such as Rous Sarcoma Virus, Harvey Sarcoma Virus, avian leukosis virus, human immunodeficiency virus, myeloproliferative sarcoma virus, and mammary tumor virus); and the like.
[0273] Additional expression vectors suitable for use are, e.g., without limitation, a lentivirus vector, a gamma retrovirus vector, a foamy virus vector, an adeno-associated virus vector, an adenovirus vector, a pox virus vector, a herpes virus vector, an engineered hybrid virus vector, a transposon mediated vector, and the like. Viral vector technology is well known in the art and is described, for example, in Sambrook et al., 2012, Molecular Cloning: A Laboratory Manual, volumes 1-4, Cold Spring Harbor Press, NY), and in other virology and molecular biology manuals. Viruses, which are useful as vectors include, but are not limited to, retroviruses, adenoviruses, adeno- associated viruses, herpes viruses, and lentiviruses.
[0274] In general, a suitable vector contains an origin of replication functional in at least one organism, a promoter sequence, convenient restriction endonuclease sites, and one or more selectable markers, (e.g., WO 01/96584; WO 01/29058; and U.S. Pat. No. 6,326,193).
[0275] In some embodiments, a vector comprises a first polynucleotide sequence encoding a heavy chain and a second polynucleotide sequence encoding a light chain. In some embodiments, the heavy chain and light chain are expressed from the vector as two separate
polypeptides. In some embodiments, the heavy chain variable domain fragment and light chain variable domain fragment are expressed as part of a single polypeptide, such as, for example, when the antibody is an scFv.
[0276] In some embodiments, a first vector comprises a polynucleotide that encodes a heavy chain and a second vector comprises a polynucleotide that encodes a light chain. In some embodiments, the first vector and second vector are transfected into host cells in similar amounts (such as similar molar amounts or similar mass amounts). In some embodiments, a mole- or mass-ratio of between 5:1 and 1:5 of the first vector and the second vector is transfected into host cells. In some embodiments, a mass ratio of between 1:1 and 1:5 for the vector encoding the heavy chain and the vector encoding the light chain is used. In some embodiments, a mass ratio of 1:2 for the vector encoding the heavy chain and the vector encoding the light chain is used.
[0277] Also provided is a host cell comprising any of the vectors or nucleic acids disclosed herein. The host cell may be of eukaryotic, prokaryotic, mammalian, or bacterial origin. In various embodiments, anti-CTLA-4 heavy chains and/or anti-CTLA-4 light chains may be expressed in prokaryotic cells, such as bacterial cells; or in eukaryotic cells, such as fungal cells (such as yeast), plant cells, insect cells, and mammalian cells. Such expression may be carried out, for example, according to procedures known in the art. Exemplary eukaryotic cells that may be used to express polypeptides include, but are not limited to, COS cells, including COS 7 cells; HEK293 cells, including HEK293-6E cells; CHO cells, including CHO-S, DG44. Lecl3 CHO cells, and FUT8 CHO cells; PER.C6® cells (Crucell); and NSO cells. In some embodiments, anti-CTLA-4 heavy chains and/or anti-CTLA-4 light chains may be expressed in yeast. See, e.g., U.S. Publication No. US 2006/0270045 Al. In some embodiments, a particular eukaryotic host cell is selected based on its ability to make desired post-translational modifications to the anti-CTLA-4 heavy chains and/or anti-CTLA-4 light chains. For example, in some embodiments, CHO cells produce polypeptides that have a higher level of sialylation than the same polypeptide produced in HEK293 cells.
[0278] Introduction of one or more nucleic acids into a desired host cell may be accomplished by any method, including but not limited to, calcium phosphate transfection, DEAE-dextran mediated transfection, cationic lipid-mediated transfection, electroporation, transduction, infection, etc. Nonlimiting exemplary methods are described, e.g., in Sambrook et al., Molecular Cloning, A Laboratory Manual, 3rd ed. Cold Spring Harbor Laboratory Press
(2001). Nucleic acids may be transiently or stably transfected in the desired host cells, according to any suitable method.
[0279] Also provided herein are host cells comprising any of the polynucleotides or vectors described herein. In some embodiments host cells are capable of over-expressing heterologous DNAs can be used for the purpose of isolating the genes encoding the antibody, polypeptide or protein of interest. Non-limiting examples of mammalian host cells include but not limited to COS, HeLa, and CHO cells. See also PCT Publication No. WO 87/04462. Suitable nonmammalian host cells include prokaryotes (such as E. coli or B. subtilis) and yeast (such as .S'. cerevisiae, S. pombe; or K. laclis).
[0280] A method of producing an antibody or antigen-binding fragment thereof that binds to CTLA-4 is also provided herein, wherein the method comprises culturing the host cell. In some embodiments, the nucleic acids encoding a subject antibody are introduced directly into a host cell, and the cell incubated under conditions sufficient to induce expression of the encoded antibody. Expression may conveniently be achieved by culturing under appropriate conditions recombinant host cells containing the nucleic acid. Following production by expression, an antibody or antigen-binding fragment thereof, may be isolated and/or purified using any suitable technique, and then used as desired.
[0281] Anti-CTLA-4 antibodies may be purified by any suitable method. Such methods include, but are not limited to, the use of affinity matrices or hydrophobic interaction chromatography. Suitable affinity ligands such as ligands that bind antibody constant regions. For example, a Protein A, Protein G, Protein A/G, or an antibody affinity column may be used to bind the constant region and to purify an anti-CTEA-4 antibody. Hydrophobic interactive chromatography, for example, a butyl or phenyl column, may also suitable for purifying some polypeptides such as antibodies. Ion exchange chromatography (e.g., anion exchange chromatography and/or cation exchange chromatography) may also suitable for purifying some polypeptides such as antibodies. Mixed-mode chromatography (e.g., reversed phase/anion exchange, reversed phase/cation exchange, hydrophilic interaction/anion exchange, hydrophilic interaction/cation exchange, etc.) may also suitable for purifying some polypeptides such as antibodies. Many methods of purifying polypeptides are known in the art.
[0282] Antibodies or antigen-binding fragments thereof as provided herein, and encoding nucleic acid molecules and vectors, may be isolated and/or purified, e.g. from their natural
environment, in substantially pure or homogeneous form, or, in the case of nucleic acid, free or substantially free of nucleic acid or genes of origin other than the sequence encoding a polypeptide with the desired function.
IL PHARMACEUTICAL COMPOSITIONS
[0283] Provided herein are compositions containing any of the provided antibodies or antigen-binding fragments or binding molecules containing same as described herein. The pharmaceutical composition can further comprise a pharmaceutically acceptable excipient. For example, the pharmaceutical composition can contain one or more excipients for modifying, maintaining or preserving, for example, the pH, osmolarity, viscosity, clarity, color, isotonicity, odor, sterility, stability, rate of dissolution or release, adsorption, or penetration of the composition. Such compositions may comprise buffers such as neutral buffered saline, phosphate buffered saline and the like; carbohydrates such as glucose, mannose, sucrose or dextrans, mannitol; proteins; polypeptides or amino acids such as glycine; antioxidants; chelating agents such as EDTA or glutathione; adjuvants (e.g., aluminum hydroxide); and preservatives.
[0284] In some embodiments, the pharmaceutical composition is a solid, such as a powder, capsule, or tablet. For example, the components of the pharmaceutical composition can be lyophilized. In some embodiments, the solid pharmaceutical composition is reconstituted or dissolved in a liquid prior to administration.
[0285] In some embodiments, the pharmaceutical composition is a liquid, for example a provided protein dissolved in an aqueous solution (such as physiological saline or Ringer’s solution). In some embodiments, the pH of the pharmaceutical composition is between about 4.0 and about 8.5 (such as between about 4.0 and about 5.0, between about 4.5 and about 5.5, between about 5.0 and about 6.0, between about 5.5 and about 6.5, between about 6.0 and about 7.0, between about 6.5 and about 7.5, between about 7.0 and about 8.0, or between about 7.5 and about 8.5).
[0286] In some embodiments, the pharmaceutical composition comprises a pharmaceutically-acceptable excipient, for example a filler, binder, coating, preservative, lubricant, flavoring agent, sweetening agent, coloring agent, a solvent, a buffering agent, a chelating agent, or stabilizer. Examples of pharmaceutically-acceptable fillers include cellulose, dibasic calcium phosphate, calcium carbonate, microcrystalline cellulose, sucrose, lactose, glucose, mannitol, sorbitol, maltol, pregelatinized starch, com starch, or potato starch.
Examples of pharmaceutically-acceptable binders include polyvinylpyrrolidone, starch, lactose, xylitol, sorbitol, maltitol, gelatin, sucrose, polyethylene glycol, methyl cellulose, or cellulose. Examples of pharmaceutically-acceptable coatings include hydroxypropyl methylcellulose (HPMC), shellac, com protein zein, or gelatin. Examples of pharmaceutically- acceptable disintegrants include polyvinylpyrrolidone, carboxymethyl cellulose, or sodium starch glycolate. Examples of pharmaceutically-acceptable lubricants include polyethylene glycol, magnesium stearate, or stearic acid. Examples of pharmaceutically-acceptable preservatives include methyl parabens, ethyl parabens, propyl paraben, benzoic acid, or sorbic acid. Examples of pharmaceutically-acceptable sweetening agents include sucrose, saccharine, aspartame, or sorbitol. Examples of pharmaceutically-acceptable buffering agents include carbonates, citrates, gluconates, acetates, phosphates, or tartrates.
[0287] In some embodiments, the pharmaceutical composition further comprises an agent for the controlled or sustained release of the product, such as injectable microspheres, bio- erodible particles, polymeric compounds (polylactic acid, polyglycolic acid), beads, or liposomes.
[0288] In some embodiments, the pharmaceutical composition is sterile. Sterilization may be accomplished by filtration through sterile filtration membranes or radiation. Where the composition is lyophilized, sterilization using this method may be conducted either prior to or following lyophilization and reconstitution. The composition for parenteral administration may be stored in lyophilized form or in solution. In addition, parenteral compositions generally are placed into a container having a sterile access port, for example, an intravenous solution bag or vial having a stopper pierceable by a hypodermic injection needle.
[0289] A pharmaceutically acceptable carrier may be a pharmaceutically acceptable material, composition, or vehicle. For example, the carrier may be a liquid or solid filler, diluent, excipient, solvent, or encapsulating material, or some combination thereof. Each component of the carrier must be “pharmaceutically acceptable” in that it must be compatible with the other ingredients of the formulation. It also must be suitable for contact with any tissue, organ, or portion of the body that it may encounter, meaning that it must not carry a risk of toxicity, irritation, allergic response, immunogenicity, or any other complication that excessively outweighs its therapeutic benefits.
[0290] In some embodiments, the pharmaceutical composition is for administration to a subject. Generally, dosages and routes of administration of the pharmaceutical composition are
determined according to the size and condition of the subject, according to standard pharmaceutical practice. For example, the therapeutically effective dose can be estimated initially either in cell culture assays or in animal models such as mice, rats, rabbits, dogs, pigs, or monkeys. An animal model may also be used to determine the appropriate concentration range and route of administration. Such information can then be used to determine useful doses and routes for administration in humans. The exact dosage will be determined in light of factors related to the subject requiring treatment. Dosage and administration are adjusted to provide sufficient levels of the active compound or to maintain the desired effect. Factors that may be taken into account include the severity of the disease state, the general health of the subject, the age, weight, and gender of the subject, time and frequency of administration, drug combination(s), reaction sensitivities, and response to therapy.
[0291] Long-acting pharmaceutical compositions may be administered every 3 to 4 days, every week, or biweekly depending on the half-life and clearance rate of the particular formulation. The frequency of dosing will depend upon the pharmacokinetic parameters of the molecule in the formulation used. Typically, a composition is administered until a dosage is reached that achieves the desired effect. The composition may therefore be administered as a single dose, or as multiple doses (at the same or different concentrations/dosages) over time, or as a continuous infusion. Further refinement of the appropriate dosage is routinely made. Appropriate dosages may be ascertained through use of appropriate dose-response data.
[0292] In some embodiments, the pharmaceutical composition is for administration to a subject through any route, including orally, transdermally, by inhalation, intravenously, intraarterially, intramuscularly, direct application to a wound site, application to a surgical site, intraperitoneally, by suppository, subcutaneously, intradermally, transcutaneously, by nebulization, intrapleurally, intraventricularly, intra-articularly, intraocularly, or intraspinally.
[0293] A provided pharmaceutical formulation may, for example, be in a form suitable for intravenous infusion.
[0294] In some embodiments, the dosage of the pharmaceutical composition is a single dose or a repeated dose. In some embodiments, the doses are given to a subject once per day, twice per day, three times per day, or four or more times per day. In some embodiments, about 1 or more (such as about 2 or more, about 3 or more, about 4 or more, about 5 or more, about 6 or more, or about 7 or more) doses are given in a week. In some embodiments, multiple doses are given over the course of days, weeks, months, or years. In some embodiments, a
course of treatment is about 1 or more doses (such as about 2 or more does, about 3 or more doses, about 4 or more doses, about 5 or more doses, about 7 or more doses, about 10 or more doses, about 15 or more doses, about 25 or more doses, about 40 or more doses, about 50 or more doses, or about 100 or more doses).
[0295] In some embodiments, an administered dose of the pharmaceutical composition is about 1 pg of protein per kg subject body mass or more (such as about 2 pg of protein per kg subject body mass or more, about 5 pg of protein per kg subject body mass or more, about 10 pg of protein per kg subject body mass or more, about 25 pg of protein per kg subject body mass or more, about 50 pg of protein per kg subject body mass or more, about 100 pg of protein per kg subject body mass or more, about 250 pg of protein per kg subject body mass or more, about 500 pg of protein per kg subject body mass or more, about 1 mg of protein per kg subject body mass or more, about 2 mg of protein per kg subject body mass or more, or about 5 mg of protein per kg subject body mass or more).
III. METHODS OF USE
[0296] The antibodies and antigen-binding fragments or CTLA-4 binding molecules containing the same described herein, including pharmaceutical compositions thereof, can be used in a variety of therapeutic applications, such as the treatment of a disease. Such methods and uses include therapeutic methods and uses, for example, involving administration of the molecules or compositions containing the same, to a subject having a disease or disorder. In some embodiments, the molecule or engineered cell is administered in an effective amount to effect treatment of the disease or disorder. Uses include uses of the antibodies and antigenbinding fragments or CTLA-4 binding molecules containing the same described herein, including pharmaceutical compositions thereof, in such methods and treatments, and in the preparation of a medicament in order to carry out such therapeutic methods. In some embodiments, the methods are carried out by administering the antibodies and antigen-binding fragments or CTLA-4 binding molecules containing the same, including pharmaceutical compositions thereof, or compositions comprising the same, to the subject having or suspected of having the disease or disorder. In some embodiments, the methods thereby treat the disease or disorder in the subject. Also provided are any methods or uses involving administering a provided engineered cell as described, or pharmaceutical compositions containing the same, for treating a disease or disorder.
[0297] In some embodiments, the disease or disorder is a tumor or cancer. As used herein, the term "cancer" refers to a neoplasm or tumor resulting from abnormal uncontrolled growth of cells. “Cancer” explicitly includes leukemias and lymphomas. The term “cancer” also refers to a disease involving cells that have the potential to metastasize to distal sites. In some embodiments, the disease or disorder is carcinoma, including that of the bladder, breast, colon, kidney, liver, lung, ovary, pancreas, stomach, cervix, thyroid and skin; including squamous cell carcinoma; hematopoietic tumors of lymphoid lineage, including leukemia, acute lymphocytic leukemia, acute lymphoblastic leukemia, B-cell lymphoma, T-cell lymphoma, Berketts lymphoma; hematopoietic tumors of myeloid lineage, including acute and chronic myelogenous leukemias and promyelocytic leukemia; tumors of mesenchymal origin, including fibrosarcoma and rhabdomyoscarcoma; other tumors, including melanoma, seminoma, tetratocarcinoma, neuroblastoma and glioma; tumors of the central and peripheral nervous system, including astrocytoma, neuroblastoma, glioma, and schwannomas; tumors of mesenchymal origin, including fibrosarcoma, rhabdomyosarcoma, and osteosarcoma; and other tumors, including melanoma, xenoderma pegmentosum, keratoactanthoma, seminoma, thyroid follicular cancer and teratocarcinoma. It is also contemplated that cancers caused by aberrations in apoptosis would also be treated by the methods and compositions of the invention. Such cancers may include, but are not be limited to, follicular lymphomas, carcinomas with p53 mutations, hormone dependent tumors of the breast, prostate and ovary, and precancerous lesions such as familial adenomatous polyposis, and myelodysplastic syndromes.
[0298] Illustrative subjects include mammalian subjects, such as farm animals, domestic animals, and human patients. In particular embodiments, the subject is a human subject. The terms subject and patient are used interchangeably herein.
[0299] In some cases, a subject is selected that is known, suspected or that has been identified as having T cells expressing CTLA-4. A provided antibody or antigen-binding fragment, CTLA-4 binding molecule or engineered cell as described herein, or pharmaceutical composition comprising the same, is administered to the subject. The administration to the subject will generally have an effect due to its binding with the CTLA-4 target.
[0300] Generally, alleviation or treatment of a disease or disorder involves the lessening of one or more symptoms or medical problems associated with the disease or disorder. For example, in the case of cancer, the therapeutically effective amount of the drug can accomplish
one or a combination of the following: reduce the number of cancer cells; reduce the tumor size; inhibit (i.e., to decrease to some extent and/or stop) cancer cell infiltration into peripheral organs; inhibit tumor metastasis; inhibit, to some extent, tumor growth; and/or relieve to some extent one or more of the symptoms associated with the cancer. In some embodiments, a composition of this disclosure can be used to prevent the onset or reoccurrence of the disease or disorder in the subject.
[0301] In some embodiments, the antibody or antigen-binding fragment, CTLA-4 binding molecule or engineered cell as described herein, or pharmaceutical composition comprising the same, can be used to inhibit growth of mammalian cancer cells (such as human cancer cells). A method of treating cancer can include administering an effective amount of any of the pharmaceutical compositions described herein to a subject with cancer. The effective amount of the pharmaceutical composition can be administered to inhibit, halt, or reverse progression of cancers. Human cancer cells can be treated in vivo, or ex vivo. In ex vivo treatment of a human patient, tissue or fluids containing cancer cells are treated outside the body and then the tissue or fluids are reintroduced back into the patient. In some embodiments, the cancer is treated in a human patient in vivo by administration of the therapeutic composition into the patient.
[0302] In some embodiments, the antibody or antigen-binding fragment, CTLA-4 binding molecule or engineered cell as described herein, or pharmaceutical composition comprising the same, are useful in treating, alleviating a symptom of, ameliorating and/or delaying the progression of a cancer or other neoplastic condition. In some embodiments, the cancer is bladder cancer, breast cancer, uterine/cervical cancer, ovarian cancer, prostate cancer, testicular cancer, esophageal cancer, gastrointestinal cancer, pancreatic cancer, colorectal cancer, colon cancer, kidney cancer, head and neck cancer, lung cancer, stomach cancer, germ cell cancer, bone cancer, liver cancer, thyroid cancer, skin cancer, neoplasm of the central nervous system, lymphoma, leukemia, myeloma, sarcoma, and virus-related cancer. In certain embodiments, the cancer is a metastatic cancer, refractory cancer, or recurrent cancer.
[0303] Compositions of the invention can be administered in dosages and routes and at times to be determined in appropriate pre-clinical and clinical experimentation and trials. Compositions may be administered multiple times at dosages within these ranges. Administration of the compositions may be combined with other methods useful to treat the desired disease or condition as determined by those of skill in the art. Typically, precise amount
of the compositions of the present disclosure to be administered can be determined by a physician with consideration of individual differences in age, weight, tumor size, extent of infection or metastasis, and condition of the patient (subject).
[0304] In some embodiments, a therapeutically effective dose may be, by way of nonlimiting example, from about 0.01 pg/kg body weight to about 20 mg/kg body weight. In some embodiments, the therapeutically effective dose may be, by way of nonlimiting example, from about 0.01 mg/kg body weight to about 5-20 mg/kg body weight. Intervals between single doses can be, for example, daily, weekly, monthly, every three months or yearly. An exemplary treatment regimen entails administration once per week, once every two weeks, once every three weeks, once every four weeks, once a month, once every three months or once every three to six months. Common dosing frequencies may range, for example, from twice daily to once a week.
[0305] The antibody or antigen-binding fragment, CTLA-4 binding molecule or pharmaceutical composition comprising the same, of the present disclosure can be administered alone or in combination with other modes of treatment, such as other anti-cancer agents. They can be provided before, substantially contemporaneous with, or after other modes of treatment (i.e., concurrently or sequentially). In some embodiments, the method of treatment described herein can further include administering: radiation therapy, chemotherapy, vaccination, targeted tumor therapy, CAR-T therapy, oncolytic virus therapy, cancer immunotherapy, cytokine therapy, surgical resection, chromatin modification, ablation, cryotherapy, an antisense agent against a tumor target, a siRNA agent against a tumor target, a microRNA agent against a tumor target or an anti-cancer/tumor agent, or a biologic, such as an antibody, cytokine, or receptor extracellular domain-Fc fusion.
[0306] In some embodiments, the antibody or antigen-binding fragment, CTLA-4 binding molecule or pharmaceutical composition comprising the same, of the present disclosure can be administered in combination with a molecule that disrupts or enhances alternative immunomodulatory pathways (such as TIM3, TIM4, 0X40, CD40, GITR, 4-1-BB, B7-H1, PD- 1, B7-H3, B7-H4, LIGHT, BTLA, ICOS, CD27 or LAG3) or modulates the activity of effecter molecules such as cytokines (e.g., IL-4, IL-7, IL-10, IL-12, IL-15, IL-17, GF-beta, IFNg, Flt3, BLys) and chemokines (e.g., CCL21) in order to enhance the immunomodulatory effects. Specific embodiments include a bispecific antibody comprising an anti-CTLA-4 antibody described herein or antigen binding fragment thereof, in combination with anti-PD-1
(pembrolizumab (Keytruda) or Nivolumab (Opdivo)), anti-B7-Hl (atezolizumab (Tecentriq) or durvalumab (Imfinzi)), anti-B7-H3, anti-B7-H4, anti-LIGHT, anti-LAG3, anti-TIM3, anti- TIM4 anti-CD40, anti-OX40, anti-GITR, anti- BTLA, anti-CD27, anti-ICOS or anti-4-lBB. In yet another embodiment, an antibody of the invention or antigen binding fragment thereof is administered in combination with a molecule that activates different stages or aspects of the immune response in order to achieve a broader immune response, such as IDO inhibitors.
[0307] In some embodiments, the antibody or antigen-binding fragment, CTLA-4 binding molecule or pharmaceutical composition comprising the same, of the present disclosure can be administered in combination with at least one additional therapeutic agent selected from the group consisting of: pomalyst, revlimid, lenalidomide, pomalidomide, thalidomide, a DNA- alkylating platinum-containing derivative, cisplatin, 5-fluorouracil, cyclophosphamide, an anti-CD137 antibody, an anti-PD-1 antibody, an anti-PD-Ll antibody, an anti-CD20 antibody, an anti-CD40 antibody, an anti-DR5 antibody, an anti-CDld antibody, an anti-TIM3 antibody, an anti-SLAMF7 antibody, an anti-KIR receptor antibody, an anti-OX40 antibody, an anti- HER2 antibody, an anti-ErbB-2 antibody, an anti-EGFR antibody, cetuximab, rituximab, trastuzumab, pembrolizumab, an antibody drug conjugation, radiotherapy, single dose radiation, fractionated radiation, focal radiation, whole organ radiation, IL- 12, IFNa, GM-CSF, a chimeric antigen receptor, adoptively transferred T cells, an anti-cancer vaccine, and an oncolytic virus.
[0308] In some embodiments, the antibody or antigen-binding fragment, CTLA-4 binding molecule or pharmaceutical composition comprising the same, of the present disclosure can be administered prior to a surgery to remove a solid tumor in the subject. In some embodiments, the antibody or antigen-binding fragment, CTLA-4 binding molecule or pharmaceutical composition comprising the same, of the present disclosure can be administered after a surgery to remove a solid tumor in the subject.
IV. ARTICLES OF MANUFACTURE OR KITS
[0309] Provided herein are articles of manufacture and kits comprising the provided compositions. In some embodiments, the kit comprises any of the provided compositions and instructions for administering the composition to a subject.
[0310] Kits can optionally include one or more components such as instructions for use, devices and additional reagents (e.g., sterilized water or saline solutions for dilution of the compositions and/or reconstitution of lyophilized protein), and components, such as tubes,
containers and syringes for practice of the methods. In some embodiments, the kits can further contain reagents for collection of samples, preparation and processing of samples, and/or reagents for quantitating the amount of one or more surface markers in a sample, such as, but not limited to, detection reagents, such as antibodies, buffers, substrates for enzymatic staining, chromagens or other materials, such as slides, containers, microtiter plates, and optionally, instructions for performing the methods. Those of skill in the art will recognize many other possible containers and plates and reagents that can be used in accord with the provided methods.
[0311] In some embodiments, the kits can be provided as articles of manufacture that include packing materials for the packaging of the cells, antibodies or reagents, or compositions thereof, or one or more other components. For example, the kits can contain containers, bottles, tubes, vial and any packaging material suitable for separating or organizing the components of the kit. The one or more containers may be formed from a variety of materials such as glass or plastic. In some embodiments, the one or more containers hold a composition comprising cells or an antibody or other reagents for use in the methods. The article of manufacture or kit herein may comprise the cells, antibodies or reagents in separate containers or in the same container.
[0312] In some embodiments, the one or more containers holding the composition may be a single-use vial or a multi-use vial, which, in some cases, may allow for repeat use of the composition. In some embodiments, the article of manufacture or kit may further comprise a second container comprising a suitable diluent. The article of manufacture or kit may further include other materials desirable from a commercial, therapeutic, and user standpoint, including other buffers, diluents, filters, needles, syringes, therapeutic agents and/or package inserts with instructions for use.
[0313] In some embodiments, the kit can, optionally, include instructions. Instructions typically include a tangible expression describing the cell composition, reagents and/or antibodies and, optionally, other components included in the kit, and methods for using such. In some embodiments, the instructions indicate methods for using the cell compositions and antibodies for administration to a subject for treating a disease or condition, such as in accord with any of the provided embodiments. In some embodiments, the instructions are provided as a label or a package insert, which is on or associated with the container. In some
embodiments, the instructions may indicate directions for reconstitution and/or use of the composition.
V. DEFINITIONS
[0314] Unless defined otherwise, all terms of art, notations and other technical and scientific terms or terminology used herein are intended to have the same meaning as is commonly understood by one of ordinary skill in the art to which the claimed subject matter pertains. In some cases, terms with commonly understood meanings are defined herein for clarity and/or for ready reference, and the inclusion of such definitions herein should not necessarily be construed to represent a substantial difference over what is generally understood in the art.
[0315] It is understood that embodiments of the invention described herein include “consisting” and/or “consisting essentially of’ embodiments. As used herein, the singular form “a”, “an”, and “the” includes plural references unless indicated otherwise. Use of the term “or” herein is not meant to imply that alternatives are mutually exclusive.
[0316] In this application, the use of “or” means “and/or” unless expressly stated or understood by one skilled in the art. In the context of a multiple dependent claim, the use of “or” refers back to more than one preceding independent or dependent claim.
[0317] The term “about” as used herein refers to the usual error range for the respective value readily known to the skilled person in this technical field. Reference to “about” a value or parameter herein includes (and describes) embodiments that are directed to that value or parameter per se. For example, description referring to “about X” includes description of “X”.
[0318] As used herein, "optional" or "optionally" means that the subsequently described event or circumstance does or does not occur, and that the description includes instances where said event or circumstance occurs and instances where it does not. For example, an optionally substituted group means that the group is unsubstituted or is substituted.
[0319] The terms “nucleic acid molecule”, “nucleic acid” and “polynucleotide” may be used interchangeably, and refer to a polymer of nucleotides. Such polymers of nucleotides may contain natural and/or non-natural nucleotides, and include, but are not limited to, DNA, RNA, and PNA. “Nucleic acid sequence” refers to the linear sequence of nucleotides comprised in the nucleic acid molecule or polynucleotide.
[0320] The term “isolated” as used herein refers to a molecule that has been separated from at least some of the components with which it is typically found in nature or produced.
For example, a polypeptide is referred to as “isolated” when it is separated from at least some of the components of the cell in which it was produced. Where a polypeptide is secreted by a cell after expression, physically separating the supernatant containing the polypeptide from the cell that produced it is considered to be “isolating” the polypeptide. Similarly, a polynucleotide is referred to as “isolated” when it is not part of the larger polynucleotide (such as, for example, genomic DNA or mitochondrial DNA, in the case of a DNA polynucleotide) in which it is typically found in nature, or is separated from at least some of the components of the cell in which it was produced, for example, in the case of an RNA polynucleotide. Thus, a DNA polynucleotide that is contained in a vector inside a host cell may be referred to as “isolated”.
[0321] The terms “polypeptide” and “protein” are used interchangeably to refer to a polymer of amino acid residues, and are not limited to a minimum length. Such polymers of amino acid residues may contain natural or non-natural amino acid residues, and include, but are not limited to, peptides, oligopeptides, dimers, trimers, and multimers of amino acid residues. Both full-length proteins and fragments thereof are encompassed by the definition. The terms also include post-expression modifications of the polypeptide, for example, glycosylation, sialylation, acetylation, phosphorylation, and the like. Furthermore, for purposes of the present disclosure, a “polypeptide” refers to a protein which includes modifications, such as deletions, additions, and substitutions (generally conservative in nature), to the native sequence, as long as the protein maintains the desired activity. These modifications may be deliberate, as through site-directed mutagenesis, or may be accidental, such as through mutations of hosts which produce the proteins or errors due to PCR amplification.
[0322] The term "isolated protein" referred to herein means that a subject protein (1) is free of at least some other proteins with which it would typically be found in nature, (2) is essentially free of other proteins from the same source, e.g., from the same species, (3) is expressed by a cell from a different species, (4) has been separated from at least about 50 percent of polynucleotides, lipids, carbohydrates, or other materials with which it is associated in nature, (5) is not associated (by covalent or noncovalent interaction) with portions of a protein with which the "isolated protein" is associated in nature, (6) is operably associated (by covalent or noncovalent interaction) with a polypeptide with which it is not associated in nature, or (7) does not occur in nature. Such an isolated protein can be encoded by genomic DNA, cDNA, mRNA or other RNA, of may be of synthetic origin, or any combination thereof. In certain embodiments, the isolated protein is substantially pure or substantially free from
proteins or polypeptides or other contaminants that are found in its natural environment that would interfere with its use (therapeutic, diagnostic, prophylactic, research or otherwise).
[0323] The term “purified” as applied to nucleic acids or proteins generally denotes a nucleic acid or polypeptide that is substantially free from other components as determined by analytical techniques well known in the art (e.g., a purified polypeptide or polynucleotide forms a discrete band in an electrophoretic gel, chromatographic eluate, and/or a media subjected to density gradient centrifugation). For example, a nucleic acid or polypeptide that gives rise to essentially one band in an electrophoretic gel is “purified.” A purified nucleic acid or protein is at least about 50% pure, usually at least about 75%, 80%, 85%, 90%, 95%, 96%, 99% or more pure (e.g., percent by weight or on a molar basis).
[0324] The term “recombinant” indicates that the material (e.g., a nucleic acid or a polypeptide) has been artificially (i.e., non-naturally) altered by human intervention. The alteration can be performed on the material within, or removed from, its natural environment or state. For example, a “recombinant nucleic acid” is one that is made by recombining nucleic acids, e.g., during cloning, affinity modification, DNA shuffling or other well-known molecular biological procedures. A “recombinant DNA molecule,” is comprised of segments of DNA joined together by means of such molecular biological techniques. The term “recombinant protein” or “recombinant polypeptide” as used herein refers to a protein molecule which is expressed using a recombinant DNA molecule. A “recombinant host cell” is a cell that contains and/or expresses a recombinant nucleic acid or that is otherwise altered by genetic engineering, such as by introducing into the cell a nucleic acid molecule encoding a recombinant protein, such as a immunomodulatory protein provided herein. Transcriptional control signals in eukaryotes comprise “promoter” and “enhancer” elements. Promoters and enhancers consist of short arrays of DNA sequences that interact specifically with cellular proteins involved in transcription. Promoter and enhancer elements have been isolated from a variety of eukaryotic sources including genes in yeast, insect and mammalian cells and viruses (analogous control elements, i.e., promoters, are also found in prokaryotes). The selection of a particular promoter and enhancer depends on what cell type is to be used to express the protein of interest.
[0325] As used herein, “substantially pure” means an object species is the predominant species present i.e., on a molar basis it is more abundant than any other individual species in the composition), and a substantially purified fraction is a composition wherein the object
species comprises at least about 50 percent (on a molar basis) of all macromolecular species present. Generally, a substantially pure composition will comprise more than about 80 percent of all macromolecular species present in the composition, for example, in some embodiments, more than about 85%, 90%, 95%, and 99%. In some embodiments, the object species is purified to essential homogeneity (contaminant species cannot be detected in the composition by conventional detection methods) wherein the composition consists essentially of a single macromolecular species.
[0326] The term “substantially similar” or “substantially the same,” as used herein, denotes a sufficiently high degree of similarity between two or more numeric values such that one of skill in the art would consider the difference between the two or more values to be of little or no biological and/or statistical significance within the context of the biological characteristic measured by said value. In some embodiments the two or more substantially similar values differ by no more than about any one of 5%, 10%, 15%, 20%, 25%, or 50%.
[0327] A polypeptide “variant” means a biologically active polypeptide having at least about 80% amino acid sequence identity with the native sequence polypeptide after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity. Such variants include, for instance, polypeptides wherein one or more amino acid residues are added, or deleted, at the N- or C-terminus of the polypeptide. In some embodiments, a variant will have at least about 80% amino acid sequence identity. In some embodiments, a variant will have at least about 90% amino acid sequence identity. In some embodiments, a variant will have at least about 95% amino acid sequence identity with the native sequence polypeptide.
[0328] As used herein, “percent (%) amino acid sequence identity” and “homology” with respect to a peptide, polypeptide or antibody sequence are defined as the percentage of amino acid residues in a candidate sequence that are identical with the amino acid residues in the specific peptide or polypeptide sequence, after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent sequence identity, and not considering any conservative substitutions as part of the sequence identity. Alignment for purposes of determining percent amino acid sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST, BLAST-2, ALIGN or MEGALIGNTM (DNASTAR) software. Those skilled in the
art can determine appropriate parameters for measuring alignment, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared.
[0329] An amino acid substitution may include but are not limited to the replacement of one amino acid in a polypeptide with another amino acid. Exemplary substitutions are shown in Table 4. Amino acid substitutions may be introduced into an antibody of interest and the products screened for a desired activity, for example, retained/improved antigen binding, decreased immunogenicity, or improved ADCC or CDC.
[0330] Amino acids may be grouped according to common side-chain properties:
(1) hydrophobic: Norleucine, Met, Ala, Vai, Leu, He;
(2) neutral hydrophilic: Cys, Ser, Thr, Asn, Gin;
(3) acidic: Asp, Glu;
(4) basic: His, Lys, Arg;
(5) residues that influence chain orientation: Gly, Pro;
(6) aromatic: Trp, Tyr, Phe.
[0331] Non-conservative substitutions will entail exchanging a member of one of these classes for another class.
[0332] The term “conservative amino acid substitution” as used herein means an amino acid substitution in which an amino acid residue is substituted by another amino acid residue having a side chain R group with similar chemical properties (e.g., charge or hydrophobicity). Examples of groups of amino acids that have side chains with similar chemical properties include 1) aliphatic side chains: glycine, alanine, valine, leucine, and isoleucine; 2) aliphatic - hydroxyl side chains: serine and threonine; 3) amide-containing side chains: asparagine and glutamine; 4) aromatic side chains: phenylalanine, tyrosine, and tryptophan; 5) basic side chains: lysine, arginine, and histidine; 6) acidic side chains: aspartic acid and glutamic acid; and 7) sulfur-containing side chains: cysteine and methionine. Conservative amino acids substitution groups are: valine-leucine-isoleucine, phenylalanine-tyrosine, lysine-arginine, alanine-valine, glutamate-aspartate, and asparagine-glutamine.
[0333] The term, “corresponding to” with reference to positions of a protein, such as recitation that nucleotides or amino acid positions “correspond to” nucleotides or amino acid positions in a disclosed sequence, such as set forth in the Sequence Listing, refers to nucleotides or amino acid positions identified upon alignment with the disclosed sequence based on structural sequence alignment or using a standard alignment algorithm, such as the GAP algorithm. By aligning the sequences, one skilled in the art can identify corresponding residues, for example, using conserved and identical amino acid residues as guides.
[0334] The term “specifically binds” as used herein means the ability of a protein, under specific binding conditions, to bind to a target protein such that its affinity or avidity is at least 10 times as great, but optionally 50, 100, 250 or 500 times as great, or even at least 1000 times as great as the average affinity or avidity of the same protein to a collection of random peptides or polypeptides of sufficient statistical size. A specifically binding protein need not bind exclusively to a single target molecule but may specifically bind to more than one target molecule. In some cases, a specifically binding protein may bind to a protein that has similarity in structural conformation with the target protein (e.g., paralogs or orthologs). Those of skill will recognize that specific binding to a molecule having the same function in a different species of animal (i.e., ortholog) or to a molecule having a substantially similar epitope as the target molecule (e.g., paralog) is possible and does not detract from the specificity of binding which is determined relative to a statistically valid collection of unique non-targets (e.g., random polypeptides). Thus, a protein of the invention may specifically bind to more than one distinct species of target molecule due to cross-reactivity. Solid-phase ELISA immunoassays,
ForteBio Octet or Biacore measurements can be used to determine specific binding between two proteins. Generally, interactions between two binding proteins have dissociation constants (Kd) less than about IxlO'5 M, and often as low as about 1 x 10'12 M. In certain aspects of the present disclosure, interactions between two binding proteins have dissociation constants of less than about IxlO'6 M, IxlO'7 M, IxlO'8 M, IxlO'9 M, IxlO'10 M, or IxlO'11 M or less.
[0335] The term “affinity” with reference to binding refers to the strength of the sum total of noncovalent interactions between a single binding site of a molecule (for example, an antibody) and its binding partner (for example, an antigen). The affinity or the apparent affinity of a molecule X for its partner Y can generally be represented by the dissociation constant (KD) or the Ko-apparent, respectively. Affinity can be measured by common methods known in the art (such as, for example, ELISA KD, KinExA, flow cytometry, and/or surface plasmon resonance devices), including those described herein. Such methods include, but are not limited to, methods involving BIAcore®, Octet®, or flow cytometry. In some embodiments, the KD of the antigen-binding molecule is measured by flow cytometry using an antigen-expressing cell line and fitting the mean fluorescence measured at each antibody concentration to a non-linear one-site binding equation (Prism Software graphpad).
[0336] The term “vector” is used to describe a polynucleotide that can be engineered to contain a cloned polynucleotide or polynucleotides that can be propagated in a host cell. A vector can include one or more of the following elements: an origin of replication, one or more regulatory sequences (such as, for example, promoters and/or enhancers) that regulate the expression of the polypeptide of interest, and/or one or more selectable marker genes (such as, for example, antibiotic resistance genes and genes that can be used in colorimetric assays, for example, P-galactosidase). The term “expression vector” refers to a vector that is used to express a polypeptide of interest in a host cell.
[0337] A “host cell” refers to a cell that may be or has been a recipient of a vector or isolated polynucleotide. Host cells may be prokaryotic cells or eukaryotic cells. Exemplary eukaryotic cells include mammalian cells, such as primate or non-primate animal cells; fungal cells, such as yeast; plant cells; and insect cells. Nonlimiting exemplary mammalian cells include, but are not limited to, NSO cells, PER.C6® cells (Crucell), and 293 and CHO cells, and their derivatives, such as 293-6E, CHO-DG44, CHO-K1, CHO-S, and CHO-DS cells. Host cells include progeny of a single host cell, and the progeny may not necessarily be completely identical (in morphology or in genomic DNA complement) to the original parent
cell due to natural, accidental, or deliberate mutation. A host cell includes cells transfected in vivo with a polynucleotide(s) a provided herein.
[0338] The term “expression” refers to the process by which a polynucleotide is transcribed from a DNA template (such as into an mRNA or other RNA transcript) and/or the process by which a transcribed mRNA is subsequently translated into peptide, polypeptides or proteins. Transcripts and encoded polypeptides may be collectively referred to as “gene product.” If the polynucleotide is derived from genomic DNA, expression may include splicing of the mRNA in a eukaryotic cell.
[0339] The term “tumor microenvironment” as used herein refers to the environment in which a tumor exists, which is the non-cellular area within the tumor and the area directly outside the tumorous tissue but does not pertain to the intracellular compartment of the cancer cell itself. The tumor and the tumor microenvironment are closely related and interact constantly. A tumor can change its microenvironment, and the microenvironment can affect how a tumor grows and spreads. Typically, the tumor microenvironment has a low pH in the range of 5.6 to 6.8, more commonly in the range of 6.0 to 6.8, in the range of 6.2-6.8. On the other hand, a normal physiological pH is in the range of 7.2-7.8, more commonly 7.35 to 7.45, such as about 7.4. The tumor microenvironment is also known to have lower concentration of glucose and other nutrients, but higher concentration of lactic acid, in comparison with blood plasma. Furthermore, the tumor microenvironment can have a temperature that is 0.3 to 1° C. higher than the normal physiological temperature. The tumor microenvironment has been discussed in Gillies et al., “MRI of the Tumor Microenvironment,” Journal of Magnetic Resonance Imaging, vol. 16, pp. 430-450, 2002, hereby incorporated by reference herein its entirety. The term “non-tumor microenvironment” refers to a microenvironment at a site other than a tumor.
[0340] The terms “individual” and “subject” are used interchangeably herein to refer to an animal; for example a mammal. The term patient includes human and veterinary subjects. In some embodiments, methods of treating mammals, including, but not limited to, humans, rodents, simians, felines, canines, equines, bovines, porcines, ovines, caprines, mammalian laboratory animals, mammalian farm animals, mammalian sport animals, and mammalian pets, are provided. The subject can be male or female and can be any suitable age, including infant, juvenile, adolescent, adult, and geriatric subjects. In some embodiments, the subject to receive the treatment can be a patient, designating the fact that the subject has been identified as having
a disorder of relevance to the treatment, or being at adequate risk of contracting the disorder. In particular embodiments, the subject is a human, such as a human patient.
[0341] The term “composition” refers to any mixture of two or more products, substances, or compounds, including cells or antibodies. It may be a solution, a suspension, liquid, powder, a paste, aqueous, non-aqueous or any combination thereof. The preparation is generally in such form as to permit the biological activity of the active ingredient (e.g. antibody) to be effective.
[0342] A “pharmaceutically acceptable carrier” refers to an ingredient in a pharmaceutical formulation, other than an active ingredient, which is nontoxic to a subject. A pharmaceutically acceptable carrier includes, but is not limited to, a buffer, excipient, stabilizer, or preservative.
[0343] As used herein, the term “treatment” or “treating” refers to clinical intervention designed to alter the natural course of the individual or cell being treated during the course of clinical pathology. Desirable effects of treatment include decreasing the rate of disease progression, ameliorating or palliating the disease state, and remission or improved prognosis. An individual is successfully “treated”, for example, if one or more symptoms associated with a disorder (e.g., an eosinophil-mediated disease) are mitigated or eliminated. For example, an individual is successfully “treated” if treatment results in increasing the quality of life of those suffering from a disease, decreasing the dose of other medications required for treating the disease, reducing the frequency of recurrence of the disease, lessening severity of the disease, delaying the development or progression of the disease, and/or prolonging survival of individuals.
[0344] An “effective amount” refers to at least an amount effective, at dosages and for periods of time necessary, to achieve the desired or indicated effect, including a therapeutic or prophylactic result. An effective amount can be provided in one or more administrations. A “therapeutically effective amount” is at least the minimum dose of cells required to effect a measurable improvement of a particular disorder. In some embodiments, a therapeutically effective amount is the amount of a composition that reduces the severity, the duration and/or the symptoms associated with cancer, viral infection, microbial infection, or septic shock in an animal. A therapeutically effective amount herein may vary according to factors such as the disease state, age, sex, and weight of the patient. A therapeutically effective amount may also be one in which any toxic or detrimental effects of the antibody are outweighed by the therapeutically beneficial effects. A “prophylactically effective amount” refers to an amount
effective, at the dosages and for periods of time necessary, to achieve the desired prophylactic result. Typically but not necessarily, since a prophylactic dose is used in subjects prior to or at the earlier stage of disease, the prophylactically effective amount can be less than the therapeutically effective amount.
[0345] A “disease” or “disorder” as used herein refers to a condition where treatment is needed and/or desired.
[0346] The term “tumor cell”, “cancer cell”, “cancer”, “tumor”, and/or “neoplasm”, unless otherwise designated, are used herein interchangeably and refer to a cell (or cells) exhibiting an uncontrolled growth and/or abnormal increased cell survival and/or inhibition of apoptosis which interferes with the normal functioning of bodily organs and systems. Included in this definition are benign and malignant cancers, polyps, hyperplasia, as well as dormant tumors or micrometastases .
[0347] The terms “cancer” and “tumor” encompass solid and hematological/lymphatic cancers and also encompass malignant, pre-malignant, and benign growth, such as dysplasia. Also, included in this definition are cells having abnormal proliferation that is not impeded (e.g. immune evasion and immune escape mechanisms) by the immune system (e.g. virus infected cells). Exemplary cancers include, but are not limited to: carcinoma, including that of the bladder, breast, colon, kidney, liver, lung, ovary, pancreas, stomach, cervix, thyroid and skin; including squamous cell carcinoma; hematopoietic tumors of lymphoid lineage, including leukemia, acute lymphocytic leukemia, acute lymphoblastic leukemia, B-cell lymphoma, T- cell lymphoma, Berketts lymphoma; hematopoietic tumors of myeloid lineage, including acute and chronic myelogenous leukemias and promyelocytic leukemia; tumors of mesenchymal origin, including fibrosarcoma and rhabdomyoscarcoma; other tumors, including melanoma, seminoma, tetratocarcinoma, neuroblastoma and glioma; tumors of the central and peripheral nervous system, including astrocytoma, neuroblastoma, glioma, and schwannomas; tumors of mesenchymal origin, including fibrosarcoma, rhabdomyosarcoma, and osteosarcoma; and other tumors, including melanoma, xenoderma pegmentosum, keratoactanthoma, seminoma, thyroid follicular cancer and teratocarcinoma. It is also contemplated that cancers caused by aberrations in apoptosis would also be treated by the methods and compositions of the invention. Such cancers may include, but are not be limited to, follicular lymphomas, carcinomas with p53 mutations, hormone dependent tumors of the breast, prostate and ovary,
and precancerous lesions such as familial adenomatous polyposis, and myelodysplastic syndromes, n specific
[0348] embodiments, malignancy or dysproliferative changes (such as metaplasias and dysplasias), or hyperproliferative disorders, are treated or prevented by the methods and compositions of the invention in the ovary, bladder, breast, colon, lung, skin, pancreas, or uterus. In other specific embodiments, sarcoma, melanoma, or leukemia is treated or prevented by the methods and compositions of the invention.
[0349] As used herein, combination refers to any association between or among two or more items. The combination can be two or more separate items, such as two compositions or two collections, can be a mixture thereof, such as a single mixture of the two or more items, or any variation thereof. The elements of a combination are generally functionally associated or related.
[0350] As used herein, a kit is a packaged combination that optionally includes other elements, such as additional agents and instructions for use of the combination or elements thereof, for a purpose including, but not limited to, therapeutic uses.
[0351] The term “wild-type” or “natural” or “native,” which are used interchangeably, as used herein is used in connection with biological materials such as nucleic acid molecules, proteins, host cells, and the like, that are found in nature and not modified by human intervention.
VI. EXEMPLARY EMBODIMENTS
[0352] Among the provided embodiments are:
[0353] Embodiment 1. An anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) antibody or antigen -binding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein: the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1), a heavy chain complementarity determining region 2 (CDR-H2), and a heavy chain complementarity determining region 3 (CDR-H3) contained within any one of SEQ ID NOs: 1-27 and 112, and the VL region comprises a light chain complementarity determining region 1 (CDR-L1), a light chain complementarity determining region 2 (CDR-L2) and a light chain complementarity determining region 3 (CDR- L3) contained within any one of SEQ ID NOs: 28-48 and 118, wherein the CDR-H1, CDR-
H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3 are not set forth in SEQ ID NOS: 49, 57, 68, 74, 89 and 98, respectively.
[0354] Embodiment 2. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 1, wherein: the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 1, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 3, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:4, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:5, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:6, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:7, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR- H2, and a CDR-H3 contained within SEQ ID NO:8, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:9, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 10, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 11, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 12, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 13, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a
CDR-H2, and a CDR-H3 contained within SEQ ID NO: 14, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 15, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 16, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 17, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 18, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 19, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:20, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:21, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 contained within SEQ ID NO: 28; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 30; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 31; the Vnregion comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 32; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a
CDR-L3 contained within SEQ ID NO: 33; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 34; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 35; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 36; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 37; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 38; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 40; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 42; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 43; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 44; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 45; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 46; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 23, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 24, and the
VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 25, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 26, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 27, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 41; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 23, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 24, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 25, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 26, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 27, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 47; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 23, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 24, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 25, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 26, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3
contained within SEQ ID NO: 22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 27, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 23, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 24, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 25, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 26, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 27, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; or the VH region comprises a CDR- Hl, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 5, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 48.
[0355] Embodiment 3. An anti-CTLA-4 antibody or antigen-binding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein: the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1) comprising the sequence set forth in any one of SEQ ID NOs: 49-56, a heavy chain complementarity determining region 2 (CDR-H2) comprising the sequence set forth in any one of SEQ ID NOs: 57-67, and a heavy chain complementarity determining region 3 (CDR-H3) comprising the sequence set forth in any one of SEQ ID NOs: 68-73; and the VL region comprises a light chain complementarity determining region 1 (CDR-L1) comprising the sequence set forth in any one of SEQ ID NOs: 74-88, a light chain complementarity determining region 2 (CDR-L2) comprising the sequence set forth in any one of SEQ ID NOs: 89-97, and a light chain complementarity determining region 3 (CDR-L3) comprising the sequence set forth in any one of SEQ ID NOs: 98-100, wherein the CDR-H1, CDR-H2, CDR-
H3, CDR-L1, CDR-L2 and CDR-L3 are not set forth in SEQ ID NOS: 49, 57, 68, 74, 89 and 98, respectively.
[0356] Embodiment 4. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-3, wherein the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1) comprising the sequence set forth in any one of SEQ ID NOs: 50-56, a heavy chain complementarity determining region 2 (CDR-H2) comprising the sequence set forth in any one of SEQ ID NOs: 58-67, and a heavy chain complementarity determining region 3 (CDR-H3) comprising the sequence set forth in any one of SEQ ID NOs: 69-73.
[0357] Embodiment 5. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiments 1-4, wherein the VL region comprises a light chain complementarity determining region 1 (CDR-L1) comprising the sequence set forth in any one of SEQ ID NOs: 75-88, a light chain complementarity determining region 2 (CDR-L2) comprising the sequence set forth in any one of SEQ ID NOs: 90-97, and a light chain complementarity determining region 3 (CDR-L3) comprising the sequence set forth in any one of SEQ ID NOs: 99-100.
[0358] Embodiment 6. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-5, wherein: the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 58, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 69, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 51, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 59, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98,
respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 61, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 51, 62, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 63, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 57, and 69, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 53, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 53, 57, and 70, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 54, 57, and 71, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 55, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 56, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 64, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a
CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 60, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 66, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 71, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 73, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 75, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 77, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 78, 90, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth
in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 79, 90, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 80, 91, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 78, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 81, 92, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 82, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 82, 93, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 83, 92, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 100, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively,
and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 86, 94, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 87, 95, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 96, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 88, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 97, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and
a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in
SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; or the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 88, 89, and 100, respectively.
[0359] Embodiment 7. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-6, wherein: the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%,
97%, 98% or 99% identity to SEQ ID NO: 1, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 3, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 4, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 5, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 6, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 7, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 8, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%,
98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 9, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 10, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 11, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 12, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 13, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 14, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 15, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%,
97%, 98% or 99% identity to SEQ ID NO: 16, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 17, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 18, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 19, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 20, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 21, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%,
98% or 99% identity to SEQ ID NO: 28; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 30; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 31; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 32; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 33; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 34; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 35; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99%
identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 36; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 37; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 38; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 40; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 42; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to
SEQ ID NO: 43; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 44; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 45; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 112, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 46; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the
VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 41; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 47; the VH region
is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 29; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 23, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 24, and the VL region is or
comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 25, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 26, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; or the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 5, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 48.
[0360] Embodiment 8. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-7, wherein the VH region and the VL region are or comprise: the sequence set forth in SEQ ID NO: 1 and 118, respectively; the sequence set forth in SEQ ID NO: 2 and 118, respectively; the sequence set forth in SEQ ID NO: 3 and 118, respectively; the sequence set forth in SEQ ID NO: 4 and 118, respectively; the sequence set forth in SEQ ID NO: 5 and 118, respectively; the sequence set forth in SEQ ID NO: 6 and 118, respectively; the sequence set forth in SEQ ID NO: 7 and 118, respectively; the sequence set forth in SEQ ID NO: 8 and 118, respectively; the sequence set forth in SEQ ID NO: 9 and 118, respectively;
the sequence set forth in SEQ ID NO: 10 and 118, respectively; the sequence set forth in SEQ ID NO: 11 and 118, respectively; the sequence set forth in SEQ ID NO: 12 and 118, respectively; the sequence set forth in SEQ ID NO: 13 and 118, respectively; the sequence set forth in SEQ ID NO: 14 and 118, respectively; the sequence set forth in SEQ ID NO: 15 and 118, respectively; the sequence set forth in SEQ ID NO: 16 and 118, respectively; the sequence set forth in SEQ ID NO: 17 and 118, respectively; the sequence set forth in SEQ ID NO: 18 and 118, respectively; the sequence set forth in SEQ ID NO: 19 and 118, respectively; the sequence set forth in SEQ ID NO: 20 and 118, respectively; the sequence set forth in SEQ ID NO: 21 and 118, respectively; the sequence set forth in SEQ ID NO: 22 and 118, respectively; the sequence set forth in SEQ ID NO: 112 and 28, respectively; the sequence set forth in SEQ ID NO: 112 and 29, respectively; the sequence set forth in SEQ ID NO: 112 and 30, respectively; the sequence set forth in SEQ ID NO: 112 and 31, respectively; the sequence set forth in SEQ ID NO: 112 and 32, respectively; the sequence set forth in SEQ ID NO: 112 and 33, respectively; the sequence set forth in SEQ ID NO: 112 and 34, respectively; the sequence set forth in SEQ ID NO: 112 and 35, respectively; the sequence set forth in SEQ ID NO: 112 and 36, respectively; the sequence set forth in SEQ ID NO: 112 and 37, respectively; the sequence set forth in SEQ ID NO: 112 and 38, respectively; the sequence set forth in SEQ ID NO: 112 and 39, respectively; the sequence set forth in SEQ ID NO: 112 and 40, respectively; the sequence set forth in SEQ ID NO: 112 and 41, respectively; the sequence set forth in SEQ ID NO: 112 and 42, respectively; the sequence set forth in SEQ ID NO: 112 and 43, respectively; the sequence set forth in SEQ ID NO: 112 and 44, respectively; the sequence set forth in SEQ ID NO: 112 and 45, respectively; the sequence set forth in SEQ ID NO: 112 and
46, respectively; the sequence set forth in SEQ ID NO: 23 and 41, respectively; the sequence set forth in SEQ ID NO: 24 and 41, respectively; the sequence set forth in SEQ ID NO: 25 and 41, respectively; the sequence set forth in SEQ ID NO: 26 and 41, respectively; the sequence set forth in SEQ ID NO: 22 and 41, respectively; the sequence set forth in SEQ ID NO: 27 and 41, respectively; the sequence set forth in SEQ ID NO: 23 and 47, respectively; the sequence set forth in SEQ ID NO: 24 and 47, respectively; the sequence set forth in SEQ ID NO: 25 and
47, respectively; the sequence set forth in SEQ ID NO: 26 and 47, respectively; the sequence set forth in SEQ ID NO: 22 and 47, respectively; the sequence set forth in SEQ ID NO: 27 and 47, respectively; the sequence set forth in SEQ ID NO: 23 and 29, respectively; the sequence set forth in SEQ ID NO: 24 and 29, respectively; the sequence set forth in SEQ ID NO: 25 and
29, respectively; the sequence set forth in SEQ ID NO: 26 and 29, respectively; the sequence set forth in SEQ ID NO: 22 and 29, respectively; the sequence set forth in SEQ ID NO: 27 and 29, respectively; the sequence set forth in SEQ ID NO: 23 and 39, respectively; the sequence set forth in SEQ ID NO: 24 and 39, respectively; the sequence set forth in SEQ ID NO: 25 and 39, respectively; the sequence set forth in SEQ ID NO: 26 and 39, respectively; the sequence set forth in SEQ ID NO: 22 and 39, respectively; the sequence set forth in SEQ ID NO: 27 and 39, respectively; or the sequence set forth in SEQ ID NO: 5 and 48, respectively.
[0361] Embodiment 9. An anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) antibody or antigen -binding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein the VH region and the VL region are or comprise: the sequence set forth in SEQ ID NO: 1 and 118, respectively; the sequence set forth in SEQ ID NO: 2 and 118, respectively; the sequence set forth in SEQ ID NO: 3 and 118, respectively; the sequence set forth in SEQ ID NO: 4 and 118, respectively; the sequence set forth in SEQ ID NO: 5 and 118, respectively; the sequence set forth in SEQ ID NO: 6 and 118, respectively; the sequence set forth in SEQ ID NO: 7 and 118, respectively; the sequence set forth in SEQ ID NO: 8 and 118, respectively; the sequence set forth in SEQ ID NO: 9 and 118, respectively; the sequence set forth in SEQ ID NO: 10 and 118, respectively; the sequence set forth in SEQ ID NO: 11 and 118, respectively; the sequence set forth in SEQ ID NO: 12 and 118, respectively; the sequence set forth in SEQ ID NO: 13 and 118, respectively; the sequence set forth in SEQ ID NO: 14 and 118, respectively; the sequence set forth in SEQ ID NO: 15 and 118, respectively; the sequence set forth in SEQ ID NO: 16 and 118, respectively; the sequence set forth in SEQ ID NO: 17 and 118, respectively; the sequence set forth in SEQ ID NO: 18 and 118, respectively; the sequence set forth in SEQ ID NO: 19 and 118, respectively; the sequence set forth in SEQ ID NO: 20 and 118, respectively; the sequence set forth in SEQ ID NO: 21 and 118, respectively; the sequence set forth in SEQ ID NO: 22 and 118, respectively; the sequence set forth in SEQ ID NO: 112 and 28, respectively; the sequence set forth in SEQ ID NO: 112 and 29, respectively; the sequence set forth in SEQ ID NO: 112 and 30, respectively; the sequence set forth in SEQ ID NO: 112 and 31, respectively; the sequence set forth in SEQ ID NO: 112 and 32, respectively; the sequence set forth in SEQ ID NO: 112 and 33, respectively; the sequence set forth in SEQ ID NO: 112 and 34, respectively; the sequence set forth in SEQ ID NO: 112 and 35, respectively; the sequence set forth in SEQ ID NO: 112 and 36, respectively; the sequence set forth in SEQ ID NO: 112 and 37, respectively; the
sequence set forth in SEQ ID NO: 112 and 38, respectively; the sequence set forth in SEQ ID NO: 112 and 39, respectively; the sequence set forth in SEQ ID NO: 112 and 40, respectively; the sequence set forth in SEQ ID NO: 112 and 41, respectively; the sequence set forth in SEQ ID NO: 112 and 42, respectively; the sequence set forth in SEQ ID NO: 112 and 43, respectively; the sequence set forth in SEQ ID NO: 112 and 44, respectively; the sequence set forth in SEQ ID NO: 112 and 45, respectively; the sequence set forth in SEQ ID NO: 112 and
46, respectively; the sequence set forth in SEQ ID NO: 23 and 41, respectively; the sequence set forth in SEQ ID NO: 24 and 41, respectively; the sequence set forth in SEQ ID NO: 25 and 41, respectively; the sequence set forth in SEQ ID NO: 26 and 41, respectively; the sequence set forth in SEQ ID NO: 22 and 41, respectively; the sequence set forth in SEQ ID NO: 27 and 41, respectively; the sequence set forth in SEQ ID NO: 23 and 47, respectively; the sequence set forth in SEQ ID NO: 24 and 47, respectively; the sequence set forth in SEQ ID NO: 25 and
47, respectively; the sequence set forth in SEQ ID NO: 26 and 47, respectively; the sequence set forth in SEQ ID NO: 22 and 47, respectively; the sequence set forth in SEQ ID NO: 27 and 47, respectively; the sequence set forth in SEQ ID NO: 23 and 29, respectively; the sequence set forth in SEQ ID NO: 24 and 29, respectively; the sequence set forth in SEQ ID NO: 25 and 29, respectively; the sequence set forth in SEQ ID NO: 26 and 29, respectively; the sequence set forth in SEQ ID NO: 22 and 29, respectively; the sequence set forth in SEQ ID NO: 27 and 29, respectively; the sequence set forth in SEQ ID NO: 23 and 39, respectively; the sequence set forth in SEQ ID NO: 24 and 39, respectively; the sequence set forth in SEQ ID NO: 25 and 39, respectively; the sequence set forth in SEQ ID NO: 26 and 39, respectively; the sequence set forth in SEQ ID NO: 22 and 39, respectively; the sequence set forth in SEQ ID NO: 27 and 39, respectively; or the sequence set forth in SEQ ID NO: 5 and 48, respectively.
[0362] Embodiment 10. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-9, wherein the VH region comprises L102H (histidine at position 98 by Kabat numbering).
[0363] Embodiment 11. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10 wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively.
[0364] Embodiment 12. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively.
[0365] Embodiment 13. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively.
[0366] Embodiment 14. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0367] Embodiment 15. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0368] Embodiment 16. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively.
[0369] Embodiment 17. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-16, wherein the antibody is a full length antibody.
[0370] Embodiment 18. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 17, wherein the full length antibody comprises a constant region (Fc) from an IgAl, IgA2, IgD, IgE, IgGl, IgG2, IgG3, IgG4, or IgM.
[0371] Embodiment 19. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 18, wherein the constant region is a human IgGl heavy chain constant region.
[0372] Embodiment 20. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19, wherein the human IgGl heavy chain constant region comprises one or more of mutations selected from the group consisting of G236A, S239D, M252Y, S254T, T256E, A330L, I332E, M428L, and N434S, numbered according to the EU numbering system.
[0373] Embodiment 21. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the human IgGl heavy chain constant region comprises S239D/A330L/I332E mutations, numbered according to the EU numbering system.
[0374] Embodiment 22. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising S239D/I332E mutations, numbered according to the EU numbering system.
[0375] Embodiment 23. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising G236A/S239D/A330L/I332E mutations, numbered according to the EU numbering system.
[0376] Embodiment 24. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising G236A/A330L/I332E mutations, numbered according to the EU numbering system.
[0377] 25. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising M252Y/S254T/T256E mutations, numbered according to the EU numbering system.
[0378] Embodiment 26. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising M428L/N434S mutations, numbered according to the EU numbering system.
[0379] Embodiment 27. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-16, wherein the antibody is an antigen-binding fragment.
[0380] Embodiment 28. The anti-CTLA-4 antibody or antigen-binding fragment thereof of Embodiment 27, wherein the antigen-binding fragment is selected from the group consisting of a single domain antibody, a single chain antibody, an unibody, a single chain variable fragment (scFv), a Fab fragment, and a F(ab')2 fragment.
[0381] Embodiment 29. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-16, wherein the anti-CTLA-4 antibody or antigen-binding fragment thereof is recombinant.
[0382] Embodiment 30. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-29, wherein the VH region and the VL region is human or is from a human protein.
[0383] Embodiment 31. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-30, wherein the ratio of dissociation constant for binding at acidic pH (e.g., pH 5.0 to 6.0) to the dissociation constant for binding at physiological pH (e.g., pH 7.35 to 7.45) to human CTLA-4 protein is greater than 2.
[0384] Embodiment 32. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-31, wherein the antibody or antigen-binding fragment exhibits a ratio of a dissociation constant for binding at acidic pH (e.g., pH 5.0 to 6.0) to dissociation constant for binding at physiological pH (e.g., pH 7.35 to 7.45) of more than 7.
[0385] Embodiment 33. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-32, wherein the ratio of EC50 value of the antibody or antigenbinding fragment thereof for human CTLA-4 at physiological pH (e.g., pH 7.35 to 7.45) to EC50 value of the antibody or antigen-binding fragment thereof for human CTLA-4 at acidic pH (e.g., pH 5.0 to 6.0) is greater than 2.
[0386] Embodiment 34. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-33, wherein the antibody or antigen-binding fragment thereof possesses pH dependent binding properties to human CTLA-4 that facilitates the dissociation of antibody from the antibody: CTLA-4 complex upon internalization into a cell, allowing either the antibody, CTLA-4, or both to be recycled instead of being degraded in the lysosome.
[0387] Embodiment 35. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-34, wherein the antibody or antigen-binding fragment thereof preserves high affinity for CTLA-4 at pH>7, lower affinity at pH 5.5-7, and significantly reduced affinity for CTLA-4 at pH<5.5.
[0388] Embodiment 36. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-35, wherein the antibody or antigen-binding fragment thereof induces strong deletion of Treg and local T cell activation in tumor microenvironment but minimal systemic T cell activation.
[0389] Embodiment 37. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-36, wherein the antibody or antigen-binding fragment thereof induces a favorable ratio of Treg cells to T effector cells in the tumor microenvironment.
[0390] Embodiment 38. A conjugate, comprising the anti-CTLA-4 antibody or antigenbinding fragment thereof of any of Embodiments 1-37 and a heterologous molecule or moiety.
[0391] Embodiment 39. The conjugate of Embodiment 38, wherein the heterologous molecule or moiety is a cytotoxic agent, a drug, enzymatically active toxin or fragment thereof, or a radioactive atom.
[0392] Embodiment 40. A bispecific antibody, comprising the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-37.
[0393] Embodiment 41. A polynucleotide comprising a nucleic acid encoding the anti- CTLA-4 antibody or antigen-binding domain thereof of any of Embodiments 1-37.
[0394] Embodiment 42. A vector, comprising the polynucleotide of Embodiment 41.
[0395] Embodiment 43. The vector of Embodiment 42, wherein the vector is a viral vector.
[0396] Embodiment 44. The vector of Embodiment 43, wherein the viral vector is a retroviral vector or a lentiviral vector.
[0397] Embodiment 45. A cell comprising the polynucleotide of Embodiment 41, or the vector of any of Embodiments 42-44.
[0398] Embodiment 46. The cell of Embodiment 45, wherein the cell is a mammalian cell.
[0399] Embodiment 47. A method of producing an antibody comprising culturing the cell of Embodiment 45 or 46 under a condition that produces the antibody.
[0400] Embodiment 48. The method of Embodiment 47, further comprising recovering the antibody produced by the cell.
[0401] 49. An antibody or antigen-binding fragment thereof produced by the method of Embodiment 47 or Embodiment 48.
[0402] Embodiment 50. A composition comprising the anti-CTLA-4 antibody or antigenbinding fragment thereof of any of Embodiments 1-37 and 49, the conjugate of Embodiment 38 or 39, the bispecific antibody of Embodiment 40.
[0403] Embodiment 51. The composition of Embodiment 50, further comprising a pharmaceutically acceptable excipient.
[0404] Embodiment 52. A method of stimulating an immune response in a subject, the method comprising administering to a subject in need thereof the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of Embodiments 1-37 and 49, the conjugate of Embodiment 38 or 39, the bispecific antibody of Embodiment 40 or the composition of Embodiment 50 or 51.
[0405] Embodiment 53. The method of Embodiment 52, wherein stimulating an immune response treats a disease or condition in the subject.
[0406] Embodiment 54. The method of Embodiment 53, wherein the disease or condition is a cancer.
[0407] Embodiment 55. A method of cancer immunotherapy, the method comprising administering to a subject in need thereof a therapeutically effective amount of the anti-CTLA- 4 antibody or antigen-binding fragment thereof of any of Embodiments 1-37 and 49, the conjugate of Embodiment 38 or 39, the bispecific antibody of Embodiment 40 or the composition of Embodiment 50 or 51.
VII. EXAMPLES
[0408] The following examples are included for illustrative purposes only and are not intended to limit the scope of the invention.
Example 1 : Generation of Anti-CTLA-4 Antibodies
[0409] Anti-CTLA-4 antibodies were computationally designed via a series of steps to identify novel antibodies with potentially desirable properties. The process included:
1) Analyzing the structure, function, and relevant biological context of the CTLA-4 antigen and existing antibodies that may have potential for CTLA-4 targeting.
2) Generating in silico models of candidate antibodies using molecular modeling techniques and predicting their properties, such as binding affinity and stability, using computational methods, such as molecular dynamics simulations.
3) Using protein engineering algorithms to introduce mutations, deletions, or insertions in the antibody sequences to optimize their properties, such as affinity, specificity, stability, immunogenicity, and isoelectric point.
4) Assessing the designed antibody sequences using computational methods, such as molecular docking and molecular dynamics simulations, to validate their binding interactions with CTLA-4 and stability in different environments.
5) Refining and optimizing the antibody designs iteratively based on the computational and experimental results, using a feedback loop between computational predictions and experimental validation.
[0410] Coding sequences for the computationally designed antibodies were recombinantly expressed in HEK293 cells. Briefly, the gene encoding the protein-of-interest was synthesized via homologous recombination amplification, and the target gene was inserted into the an expression vector via restriction enzyme digestion and ligation with nucleic acid encoding a constant heavy chain (SEQ ID NO: 101 or SEQ ID NO: 106) with a heavy chain signal peptide (SEQ ID NO: 104), and nucleic acid encoding a constant light chain (SEQ ID NO: 103) with a light chain signal peptide (SEQ ID NO: 105). The vectors were sequenced to confirm the correct insertion of the target gene. The vectors were transiently transfected into host cells and after culture, protein was purified from the culture supernatant via affinity chromatography or a combination of chromatography methods. The final purified product was subjected to buffer exchange into the storage buffer. Methods such as SDS-PAGE, UV (OD280), and SEC were used to assess the purity and quantity of the crude samples after each purification step as well as the final protein product.
[0411] Table El lists sequence identifiers (SEQ ID NO:) corresponding to amino acid (aa) sequences of the variable heavy (Vn) chain and variable light (VL) chain of the designed antibodies, including complementarity determining regions (CDRs, by Kabat numbering) of each chain.
Example 2 : Functional Analyses of Anti-CTLA-4 Antibodies
[0412] The engineered antibodies generated in HEK293 cells as described in Example 1 were tested for binding to recombinant human CTLA-4 using Enzyme-Linked ImmunoSorbent Assay (ELISA) at different pH’s.
[0413] The tested concentrations of antibodies ranged from 0.1152 ng/mL to 1800 ng/mL and results were compared to reference antibodies (e.g., Referencel, ipilimumab; Reference2, tremelimumab; Referenced, gotistobart; and Reference6, ipi.101). As shown in FIGs. 1A-1S, the computationally designed antibodies exhibited differences in binding relative to the reference antibodies and to each other at pH 5.0 and pH 7.4.
[0414] Additionally, antibodies were tested for binding to human CTLA-4 using ELISA at different pH conditions, and EC50 and absorbance were measured. As shown in FIGs. 2A-2C and Table E2A, the computationally designed antibodies exhibited differences in binding relative to the reference antibodies and to each other at different pH’s.
[0415] Among the antibodies are antibodies that exhibit better binding at neutral pH than acidic pH. In many cases, the ratio of binding at neutral pH (e.g., pH 7.4) than acidic pH (e.g., pH 5.0) was substantially increased compared to reference antibodies, while retaining similar or better binding at pH 7.4 than reference antibodies. In particular, it is found herein that antibodies with such a pH-dependent binding feature are enriched in antibodies that contain one or more of the following amino acids in the VH region: a histidine (H) at position 98, an arginine (R) at position 31, a glutamic acid (E) or a histidine (H) at position 53, a histidine (H) at position 58, and a histidine (H) at position 97, or any combination thereof; and/or in the VL region: a histidine (H) at position 27a, a histidine (H) at position 30, a histidine (H) at position 31, and a histidine (H) at position 90, or any combination thereof, each based on Kabat numbering. Without wishing to be bound by theory, it is believed that an antibody with higher activity at neutral pH (e.g., pH 7.4) than at acidic pH (e.g., pH 5.0) will exhibit less risk of toxic effect because the antibody can dissociate from CTLA-4 upon internalization of the antibody - CTLA-4 complex, thereby allowing CTLA-4 and the anti-CTLA-4 antibody to recycle to the cell surface. The concept that recycling of CTLA-4 is important in human immune function is supported by the observation that defects in this pathway due to lipopolysaccharide (LPS)- responsive and beige-like anchor protein (LRBA) deficiency are associated with autoimmunity. Immunotherapy-related adverse effect (irAE) can also be caused by loss of CTLA-4 at the cell surface. Lor anti-CTLA-4 antibodies that display pH sensitive binding to CTLA-4, as the pH drops in early endosomes, it is more likely the anti-CTLA4 antibodies will dissociate from CTLA-4, enabling CTLA-4 and the anti-CTLA-4 antibodies to traffic back to the cell surface via interaction with LBRA and the neonatal Ec receptor (EcRn) respectively. Contrarily, CTLA-4 interaction with anti-CTLA4 antibodies that bind better at a lower pH can cause depletion of both CTLA-4 and the anti-CTLA-4 antibody via lysosomal degradation. This property is critical for proper CTLA-4 function and balance between immune system activation and self-immune tolerance. Thus, the provided antibodies are expected to maintain therapeutic efficacy while also being safe due to critical properties enabling proper CTLA-4 function and balance between immune system activation and self-immune tolerance.
[0416] Among the antibodies are antibodies that exhibit better binding to the EcGRIIIA receptor than antibodies comprising an IgGl constant region domain. Without wishing to be
bound by theory, it is believed that a significant contributor to the anti-tumor response of anti- CTLA-4 antibodies in the clinic involves depletion of Tregs via Fc gamma receptor engagement and mediated antibody-dependent cellular cytotoxicity & phagocytosis. Priming of the immune response via FcR engagement is also a suggested contributor to the mechanism of anti-CTLA4 antibodies in cancer treatment. Thus the provided antibodies are expected to alter the ratio of effector T cells to regulatory T cells via Fc gamma receptor interaction in the tumor microenvironment in a favorable way.
Example 3 Binding Kinetics to Human CTLA-4 at Different pH Conditions
[0417] The engineered antibodies generated in HEK293 cells as described in Example 1 were measured for binding kinetics to human CTLA-4 at different pH conditions. Biolayer interferometry (BLI) was used to measure equilibrium dissociation constant (KD), association rate constant (kon), and dissociation rate constant (kdis) of the interaction between the anti- CTLA-4 antibodies as described above and CTLA-4 at pH 5.0 or pH 7.4, and compared to a reference antibody. Briefly, BLI is an optical analytical technique that analyzes the interference pattern of white light reflected from two surfaces: a layer of immobilized protein on the biosensor tip, and an internal reference layer. Any change in the number of molecules bound to the biosensor tip causes a shift in the interference pattern that can be measured in real-time. As shown in Table E3A and Table E3B the computationally designed antibodies exhibited differences in kinetics relative to the reference antibody and to each other.
Table E3B: In vitro binding kinetics to CTLA-4 at pH 5.0
Example 4 Effect of Anti-CTLA-4 Antibodies on CTLA-4 Expression in Treated Human-CTLA-4 Expressing Cells
[0418] Functional experiments were conducted to evaluate the impact of anti-CTLA-4 antibodies on CTLA-4 expression levels in CTLA-4 expressing cells. Human embryonic kidney (HEK) 293T cells were transiently transfected with CTLA-4 cDNA and incubated for four hours with various antibodies, including a control hlgG-Fc, Reference 1, Referenced, and CTLA-4- AB-68. The CTLA-4 protein levels were assessed using western blot analysis, with ACTB (beta-actin) serving as the loading control.
[0419] Quantitative analysis of the western blot bands was performed using image analysis software, which calculated the sum, area, average, and median pixel intensities for each band. The CTLA-4 levels were normalized to the beta-actin levels to account for variations in sample loading. As depicted in FIG. 3A and shown in Tables E4A-E4C, cells treated with CTLA-4-AB-68 exhibited hCTLA-4 levels similar to those of the negative control. In contrast, cells treated with Reference 1, known for depleting CTLA-4, showed reduced hCTLA-4 levels. Referenced, an anti-CTLA-4 antibody with a pH-dependent binding profile reported to reduce CTLA-4 degradation, also displayed reduced hCTLA-4 levels but to a lesser extent than Referencel. These findings suggest that Referencel promotes the depletion of CTLA-4 via trafficking to the lysosome and subsequent degradation, while Referenced, despite its pH-dependent binding, still leads to some CTLA-4 degradation. On the other hand, CTLA-4- AB-68 effectively maintained CTLA-4 levels, likely due to its unique pH-depending binding properties that facilitate CTLA-4 recycling.
[0420] Additionally, confocal microscopy experiments were performed to visualize the trafficking of CTLA-4 and the antibodies. Although not a quantitative readout, this qualitative analysis provided visual evidence supporting the western blot findings, demonstrating the distinct intracellular localization patterns of both CTLA-4 and anti-CTLA- 4 antibodies upon treatment with CTLA-4- AB-68 vs. Referencel. A simplified workflow is provided in FIG. 3B. HEK293-hCTLA-4 cells were incubated with lOpg/mL antibody. Results in FIG. 3C further underscores the unique property of CTLA-4- AB-68 in preserving CTLA-4 recycling to reduce CTLA-4 depletion.
Example 5 Effect of Anti-CTLA-4 Antibodies on Antibody Dependent Cellular Cytotoxicity
[0421] The ability of anti-CTLA-4 antibodies to induce antibody dependent cellular cytotoxicity (ADCC) was assessed by co-culturing human CTLA-4-expressing target cells with NK cells isolated from the peripheral blood mononuclear cells PBMCs of healthy donors. Briefly, NK cells were purified using negative selection techniques and verified by flow cytometry, gating out CD3-positive cells and selecting CD56-positive cells as shown in FIG. 4A. HEK293 cells engineered to overexpress human CTLA-4 (293T-hCTLA-4) were used as target cells. These cells were labeled with 50 nM CellTrace Far-Red for identification. Multiple Effector: Target cell ratios were tested to identify the optimal condition for differentiating ADCC activity. The ratios tested included 0:1, 1:1, 2:1, 4:1, and 8:1. Based on the percentage of target cell lysis, the optimal E:T ratio of 2:1 was selected for further characterization. Purified NK cells and labeled target cells were co-incubated for 6 hours at the optimized E:T ratio of 2:1. Target cell lysis was measured by the reduction in CellTrace Far-Red fluorescence. The percentage of target cell lysis was evaluated at various concentrations of four different antibodies, with EC50 values calculated from dose-response curves. As depicted in FIG. 4B and shown in Table E5A, CTLA-4- AB-68 (IgGl) showed slightly higher ADCC compared to Reference 1 (IgGl), and the isotype control antibody (IgGl) exhibited minimal ADCC activity. Notably, CTLA-4- AB-68.1 (IgGl
S239D/A330L/I332E), an Fc variant of CTLA-4-AB-68 (IgGl), demonstrated substantially greater ADCC compared to the other molecules tested.
Example 6 Evaluation of In Vivo Anti-Tumor Efficacy of Anti-CTLA-4 Antibodies in the Subcutaneous CT-26 Murine Colorectal Carcinoma Syngeneic Model Using hCTLA-4 Knock-In Mice
[0422] In this study, the antitumor activity of anti-CTLA-4 antibodies was assessed using a syngeneic mouse model for colon cancer (CT26 tumor-bearing mice). This model was specifically selected because reference anti-human-CTLA-4 antibodies do not cross-react with murine CTLA-4, necessitating the use of human CTLA-4 (hCTLA-4) knock-in mice to accurately evaluate the therapeutic potential of the antibodies. These mice were developed on a BALB/c background, providing an immune-competent, genetically consistent environment that closely mimics the human immune response. The study utilized female hCTLA-4 knock- in BALB/c mice, aged 6-8 weeks and weighing approximately 18-22g.
[0423] CT-26 tumor cells were cultured in vitro as a monolayer in RPML1640 medium supplemented with 10% heat-inactivated fetal bovine serum, 100 U/ml penicillin, and 100 pg/ml streptomycin. Cultures were maintained at 37°C in a 5% CO2 atmosphere and subcultured twice weekly using trypsin-EDTA treatment. Cells in the exponential growth phase were harvested and counted for tumor inoculation.
[0424] For tumor development, each mouse was inoculated subcutaneously on the right lower flank with 3 x 10A5 CT-26 tumor cells in 0.1 mL of serum-free PBS. To ensure accurate subcutaneous delivery, the skin fold was lifted with one hand while the cells were injected with the other. To minimize tumor ulceration, a new syringe and needle were used for each mouse, and any droplets of cells were wiped from the skin with KimWipes postinjection. The animals were randomized, and treatment commenced when the average tumor volume reached 80 mmA3. Treatment was administered intraperitoneally at a fixed mg/kg dose, with the dosing volume adjusted based on body weight (10 pL/g). Treatment was delivered bi-weekly for 3 weeks, totaling six doses, as shown in FIG. 5A.
[0425] Tumor dimensions were measured three times per week during both the treatment and observation periods. Measurements were taken in two dimensions using calipers, and
tumor volume was calculated using the formula: V = 0.5 * a * bA2, where “a” and “b” represent the long and short diameters of the tumor, respectively.
[0426] Tumor volume results are summarized in FIGs. 5B-5E. While tumor growth progressed in all vehicle control-treated mice, treatment with CTLA-4- AB-68.1 led to complete tumor rejection in several mice and significantly slowed tumor growth in others. Notably, mice treated with CTLA-4- AB-68.1 appeared to demonstrate improved tumor rejection at increased doses and superior tumor rejection compared to those treated with Reference 1 at a low (1 mg/kg) dose.
Example 7 Effect of Anti-CTLA-4 Antibodies on Immune Cell Subsets in the Tumor and Blood of hCTLA4 Knock-In Mice
[0427] hCTLA4 knock-in mice bearing CT26 colorectal carcinoma tumors were prepared as described in Example 6. Multi-parameter flow cytometry assays were designed to evaluate the absolute cell numbers of various immune cell subsets in tumor and blood samples 5 days after a single 5 mg/kg dose treatment of Vehicle control, CTLA-4- AB-68.1, or Referencel. Each group included four mice. The animals were randomized, and treatment commenced when the average tumor volume reached 80 mmA3. Treatment was administered intraperitoneally .
[0428] Five days after administration of the single dose, tumor and blood samples were collected from the mice, and single-cell suspensions were prepared. Red blood cells were lysed from the blood samples to isolate leukocytes. The flow cytometry panels with fluorophore-conjugated-antibodies were designed as shown in Table E7A and Table E7B for tumor and blood samples respectively. Cells were stained with antibodies according to the manufacturer's instructions. After staining, cells were washed and resuspended in an appropriate buffer for flow cytometry analysis.
[0429] Data acquisition was performed using multi-parameter flow cytometry, incorporating compensation controls and single- stained controls to ensure accuracy. The flow cytometry data were analyzed to gate and quantify the following cell populations: CD45+ leukocytes, CD4 T cells (CD3+, CD4+), CD8 T cells (CD3+, CD8+), regulatory T cells (CD4+, CD25+, FoxP3+), and myeloid cells (CDl lb+).
[0430] The gating strategy for tumor and blood samples are detailed in FIG. 6A and FIG. 6B, respectively. Absolute cell numbers were calculated per 100 mg of tumor tissue and per 100 microliters of blood.
[0431] The results of the FCM analysis for tumor are presented in FIG. 6C, and the results of the FCM analysis for blood samples are shown in FIGs. 6D-6H, respectively. In FIG. 6C, cell numbers of CD45+, myeloid, T cells and T cell subpopulations per 100 mg tumor are shown. CD4Teff refers to CD4T cells other than Treg. Treatment of hCTLA4 knock-in mice bearing CT26 colorectal carcinoma tumors with CTLA-4- AB-68.1 led to significant differences in immune cell subsets in both the tumor and the blood compared to the vehicle control and Reference 1.
[0432] Notably, CTLA-4- AB-68.1 treatment increased the ratio of CD4Teff/TReg and the ratio of CD8T/TReg cells in the tumor relative to the vehicle control and Reference 1. Additionally, while treatment with Reference 1 resulted in depleted levels of CTLA-4 expression on TReg cells in the blood compared to the vehicle control, treatment with CTLA- 4- AB-68.1 did not deplete CTLA-4 expression on TReg cells in the blood.
[0433] The present invention is not intended to be limited in scope to the particular disclosed embodiments, which are provided, for example, to illustrate various aspects of the invention. Various modifications to the compositions and methods described will become apparent from the description and teachings herein. Such variations may be practiced without departing from the true scope and spirit of the disclosure and are intended to fall within the scope of the present disclosure.
Claims
1. An anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) antibody or antigenbinding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein: the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1), a heavy chain complementarity determining region 2 (CDR-H2), and a heavy chain complementarity determining region 3 (CDR-H3) contained within any one of SEQ ID NOs: 1-27 and 112, and the VL region comprises a light chain complementarity determining region 1 (CDR-L1), a light chain complementarity determining region 2 (CDR-L2) and a light chain complementarity determining region 3 (CDR-L3) contained within any one of SEQ ID NOs: 28-48 and 118, wherein the CDR-H1, CDR- H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3 are not set forth in SEQ ID NOS: 49, 57, 68, 74, 89 and 98, respectively.
2. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 1, wherein: the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:
1, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:
2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:
3, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:4, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:5, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:6, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:7, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:8, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:9, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 10, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 11, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 12, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:13, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 14, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 15, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:16, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 17, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 18, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 19, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:20, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:21, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 118; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 28; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 29; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 30; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 31; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 32; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 33; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 112, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 34;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:2, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO:;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:
26, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 22, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO:
27, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 39; or the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 contained within SEQ ID NO: 5, and the VL region comprises a CDR-L1, a CDR-L2, and a CDR-L3 contained within SEQ ID NO: 48.
3. An anti-CTLA-4 antibody or antigen-binding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein: the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1) comprising the sequence set forth in any one of SEQ ID NOs: 49-56, a heavy chain complementarity determining region 2 (CDR-H2) comprising the sequence set forth in any one of SEQ ID NOs: 57-67, and a heavy chain complementarity determining region 3 (CDR-H3) comprising the sequence set forth in any one of SEQ ID NOs: 68-73; and the VL region comprises a light chain complementarity determining region 1 (CDR-L1) comprising the sequence set forth in any one of SEQ ID NOs: 74-88, a light chain complementarity determining region 2 (CDR-L2) comprising the sequence set forth in any one of SEQ ID NOs: 89-97, and a light chain complementarity determining region 3 (CDR-L3) comprising the sequence set forth in any one of SEQ ID NOs: 98-100, wherein the CDR-H1, CDR-H2, CDR-H3, CDR-L1, CDR-L2 and CDR-L3 are not set forth in SEQ ID NOS: 49, 57, 68, 74, 89 and 98, respectively.
4. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-3, wherein the VH region comprises a heavy chain complementarity determining region 1 (CDR-H1) comprising the sequence set forth in any one of SEQ ID NOs: 50-56, a heavy chain complementarity determining region 2 (CDR-H2) comprising the sequence set forth in any one of SEQ ID NOs: 58-67, and a heavy chain complementarity determining region 3 (CDR-H3) comprising the sequence set forth in any one of SEQ ID NOs: 69-73.
5. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claims 1-4, wherein the VL region comprises a light chain complementarity determining region 1 (CDR-L1) comprising the sequence set forth in any one of SEQ ID NOs: 75-88, a light chain complementarity determining region 2 (CDR-L2) comprising the sequence set forth in any one of SEQ ID NOs: 90-97, and a light chain complementarity determining region 3 (CDR-L3) comprising the sequence set forth in any one of SEQ ID NOs: 99-100.
6. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-5, wherein: the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 58, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 69, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 51, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 59, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 61, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 51, 62, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 63, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 50, 57, and 69, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 53, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 53, 57, and 70, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 54, 57, and 71, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 55, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 56, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 64, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 60, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 66, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 71, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 73, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 75, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 77, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 78, 90, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 79, 90, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 80, 91, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 78, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 81, 92, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 82, 89, and 98, respectively;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 82, 93, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 83, 92, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 100, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 86, 94, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 87, 95, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 96, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 88, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 74, 97, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 73, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 57, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively;
the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 65, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively; or the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 88, 89, and 100, respectively.
7. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-6, wherein: the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:
1, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:
2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:
3, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:
4, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:
5, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:
6, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:
, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%,8%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%,8%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 118; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:18; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:0, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:18; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:1, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:18; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:18; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:3, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:18; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,7%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:4, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,
%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:8; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:8; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:8; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:8; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:8; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:8; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:8; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,
%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:8; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:8; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,
%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,
%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:2, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,
%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,
%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,
%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%,%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO:, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%,
87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 22, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 27, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 39; or the VH region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 5, and the VL region is or comprises an amino acid sequence having at least at or about 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% identity to SEQ ID NO: 48.
8. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-7, wherein the VH region and the VL region are or comprise: the sequence set forth in SEQ ID NO: 1 and 118, respectively; the sequence set forth in SEQ ID NO: 2 and 118, respectively; the sequence set forth in SEQ ID NO: 3 and 118, respectively; the sequence set forth in SEQ ID NO: 4 and 118, respectively; the sequence set forth in SEQ ID NO: 5 and 118, respectively; the sequence set forth in SEQ ID NO: 6 and 118, respectively; the sequence set forth in SEQ ID NO: 7 and 118, respectively; the sequence set forth in SEQ ID NO: 8 and 118, respectively; the sequence set forth in SEQ ID NO: 9 and 118, respectively; the sequence set forth in SEQ ID NO: 10 and 118, respectively; the sequence set forth in SEQ ID NO: 11 and 118, respectively; the sequence set forth in SEQ ID NO: 12 and 118, respectively; the sequence set forth in SEQ ID NO: 13 and 118, respectively; the sequence set forth in SEQ ID NO: 14 and 118, respectively; the sequence set forth in SEQ ID NO: 15 and 118, respectively; the sequence set forth in SEQ ID NO: 16 and 118, respectively; the sequence set forth in SEQ ID NO: 17 and 118, respectively; the sequence set forth in SEQ ID NO: 18 and 118, respectively; the sequence set forth in SEQ ID NO: 19 and 118, respectively; the sequence set forth in SEQ ID NO: 20 and 118, respectively; the sequence set forth in SEQ ID NO: 21 and 118, respectively; the sequence set forth in SEQ ID NO: 22 and 118, respectively; the sequence set forth in SEQ ID NO: 112 and 28, respectively; the sequence set forth in SEQ ID NO: 112 and 29, respectively; the sequence set forth in SEQ ID NO: 112 and 30, respectively; the sequence set forth in SEQ ID NO: 112 and 31, respectively; the sequence set forth in
SEQ ID NO: 112 and 32, respectively; the sequence set forth in SEQ ID NO: 112 and 33, respectively; the sequence set forth in SEQ ID NO: 112 and 34, respectively; the sequence set forth in SEQ ID NO: 112 and 35, respectively; the sequence set forth in SEQ ID NO: 112 and 36, respectively; the sequence set forth in SEQ ID NO: 112 and 37, respectively; the sequence set forth in SEQ ID NO: 112 and 38, respectively; the sequence set forth in SEQ ID NO: 112 and 39, respectively; the sequence set forth in SEQ ID NO: 112 and 40, respectively; the sequence set forth in SEQ ID NO: 112 and 41, respectively; the sequence set forth in SEQ ID NO: 112 and 42, respectively; the sequence set forth in SEQ ID NO: 112 and 43, respectively; the sequence set forth in SEQ ID NO: 112 and 44, respectively; the sequence set forth in SEQ ID NO: 112 and 45, respectively; the sequence set forth in SEQ ID NO: 112 and 46, respectively; the sequence set forth in SEQ ID NO: 23 and 41, respectively; the sequence set forth in SEQ ID NO: 24 and 41, respectively; the sequence set forth in SEQ ID NO: 25 and 41, respectively; the sequence set forth in SEQ ID NO: 26 and 41, respectively; the sequence set forth in SEQ ID NO:
22 and 41, respectively; the sequence set forth in SEQ ID NO: 27 and 41, respectively; the sequence set forth in SEQ ID NO: 23 and 47, respectively; the sequence set forth in SEQ ID NO: 24 and 47, respectively; the sequence set forth in SEQ ID NO: 25 and 47, respectively; the sequence set forth in SEQ ID NO: 26 and 47, respectively; the sequence set forth in SEQ ID NO: 22 and 47, respectively; the sequence set forth in SEQ ID NO: 27 and 47, respectively; the sequence set forth in SEQ ID NO:
23 and 29, respectively; the sequence set forth in SEQ ID NO: 24 and 29, respectively; the sequence set forth in SEQ ID NO: 25 and 29, respectively; the sequence set forth in SEQ ID NO: 26 and 29, respectively; the sequence set forth in SEQ ID NO: 22 and 29, respectively; the sequence set forth in SEQ ID NO: 27 and 29, respectively; the sequence set forth in SEQ ID NO: 23 and 39, respectively; the sequence set forth in SEQ ID NO: 24 and 39, respectively; the sequence set forth in SEQ ID NO: 25 and 39, respectively; the sequence set forth in SEQ ID NO: 26 and 39, respectively; the sequence set forth in SEQ ID NO: 22 and 39, respectively; the sequence set forth in SEQ ID NO: 27 and 39, respectively; or the sequence set forth in SEQ ID NO: 5 and 48, respectively.
9. An anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) antibody or antigenbinding fragment thereof, comprising a heavy chain variable (VH) region and a light chain variable (VL) region, wherein the VH region and the VL region are or comprise: the sequence set forth in SEQ ID NO: 1 and 118, respectively; the sequence set forth in SEQ ID NO: 2 and 118, respectively; the sequence set forth in SEQ ID NO: 3 and 118, respectively; the sequence set forth in SEQ ID NO: 4 and 118, respectively; the sequence set forth in SEQ ID NO: 5 and 118, respectively; the sequence set forth in SEQ ID NO: 6 and 118, respectively; the sequence set forth in SEQ ID NO: 7 and 118, respectively; the sequence set forth in SEQ ID NO: 8 and 118, respectively; the sequence set forth in SEQ ID NO: 9 and 118, respectively; the sequence set forth in SEQ ID NO: 10 and 118, respectively; the sequence set
forth in SEQ ID NO: 11 and 118, respectively; the sequence set forth in SEQ ID NO: 12 and 118, respectively; the sequence set forth in SEQ ID NO: 13 and 118, respectively; the sequence set forth in SEQ ID NO: 14 and 118, respectively; the sequence set forth in SEQ ID NO: 15 and 118, respectively; the sequence set forth in SEQ ID NO: 16 and 118, respectively; the sequence set forth in SEQ ID NO: 17 and 118, respectively; the sequence set forth in SEQ ID NO: 18 and 118, respectively; the sequence set forth in SEQ ID NO: 19 and 118, respectively; the sequence set forth in SEQ ID NO: 20 and 118, respectively; the sequence set forth in SEQ ID NO: 21 and 118, respectively; the sequence set forth in SEQ ID NO: 22 and 118, respectively; the sequence set forth in SEQ ID NO: 112 and 28, respectively; the sequence set forth in SEQ ID NO: 112 and 29, respectively; the sequence set forth in SEQ ID NO: 112 and 30, respectively; the sequence set forth in SEQ ID NO: 112 and 31, respectively; the sequence set forth in SEQ ID NO: 112 and 32, respectively; the sequence set forth in SEQ ID NO: 112 and 33, respectively; the sequence set forth in SEQ ID NO: 112 and 34, respectively; the sequence set forth in SEQ ID NO: 112 and 35, respectively; the sequence set forth in SEQ ID NO: 112 and 36, respectively; the sequence set forth in SEQ ID NO: 112 and 37, respectively; the sequence set forth in SEQ ID NO: 112 and 38, respectively; the sequence set forth in SEQ ID NO: 112 and 39, respectively; the sequence set forth in SEQ ID NO: 112 and 40, respectively; the sequence set forth in SEQ ID NO: 112 and 41, respectively; the sequence set forth in SEQ ID NO: 112 and 42, respectively; the sequence set forth in SEQ ID NO: 112 and 43, respectively; the sequence set forth in SEQ ID NO: 112 and 44, respectively; the sequence set forth in SEQ ID NO: 112 and 45, respectively; the sequence set forth in SEQ ID NO: 112 and 46, respectively; the sequence set forth in SEQ ID NO: 23 and 41, respectively; the sequence set forth in SEQ ID NO: 24 and 41, respectively; the sequence set forth in SEQ ID NO: 25 and 41, respectively; the sequence set forth in SEQ ID NO: 26 and 41, respectively; the sequence set forth in SEQ ID NO: 22 and 41, respectively; the sequence set forth in SEQ ID NO: 27 and 41, respectively; the sequence set forth in SEQ ID NO: 23 and 47, respectively; the sequence set forth in SEQ ID NO: 24 and 47, respectively; the sequence set forth in SEQ ID NO: 25 and 47, respectively; the sequence set forth in SEQ ID NO: 26 and 47, respectively; the sequence set forth in SEQ ID NO: 22 and 47, respectively; the sequence set forth in SEQ ID NO: 27 and 47, respectively; the sequence set forth in SEQ ID NO: 23 and 29, respectively; the sequence set forth in SEQ ID NO: 24 and 29, respectively; the sequence set forth in SEQ ID NO: 25 and 29, respectively; the sequence set forth in SEQ ID NO:
26 and 29, respectively; the sequence set forth in SEQ ID NO: 22 and 29, respectively; the sequence set forth in SEQ ID NO: 27 and 29, respectively; the sequence set forth in SEQ ID NO: 23 and 39, respectively; the sequence set forth in SEQ ID NO: 24 and 39, respectively; the sequence set forth in SEQ ID NO: 25 and 39, respectively; the sequence set forth in SEQ ID NO: 26 and 39, respectively; the sequence set forth in SEQ ID NO: 22 and 39, respectively; the sequence set forth in SEQ ID NO:
27 and 39, respectively; or the sequence set forth in SEQ ID NO: 5 and 48, respectively.
10. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-9, wherein the VH region comprises one or more amino acid residues selected from the group consisting of: a histidine (H) at position 98, an arginine (R) at position 31, a glutamic acid (E) or a histidine (H) at position 53, a histidine (H) at position 58, and a histidine (H) at position 97, or any combination thereof, and/or the VL region comprises one or more amino acid residues selected from the group consisting of: a histidine (H) at position 27a, a histidine (H) at position 30, a histidine (H) at position 31 , and a histidine (H) at position 90, or any combination thereof, wherein numbering of amino acid residues is by Kabat numbering.
11. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-10 wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 85, 89, and 98, respectively, optionally wherein the VH region comprises the sequence of amino acids set forth in SEQ ID NO: 25 and the VL region comprises the sequence of amino acids set forth in SEQ ID NO: 41.
12. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 98, respectively, optionally wherein the VH region comprises the sequence of amino acids set forth in SEQ ID NO: 23 and the VL region comprises the sequence of amino acids set forth in SEQ ID NO: 47.
13. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 76, 89, and 98, respectively, optionally wherein the VH region comprises the sequence of amino acids set forth in SEQ ID NO: 25 and the VL region comprises the sequence of amino acids set forth in SEQ ID NO: 29.
14. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 49, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively,
optionally wherein the VH region comprises the sequence of amino acids set forth in SEQ ID NO: 23 and the VL region comprises the sequence of amino acids set forth in SEQ ID NO: 39.
15. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 60, and 72, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively, optionally wherein the VH region comprises the sequence of amino acids set forth in SEQ ID NO: 25 and the VL region comprises the sequence of amino acids set forth in SEQ ID NO: 39.
16. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-10, wherein the VH region comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising the sequence set forth in SEQ ID NOS: 52, 67, and 68, respectively, and the VL region comprises a CDR-L1, a CDR- L2, and a CDR-L3 comprising the sequence set forth in SEQ ID NOS: 84, 89, and 99, respectively, optionally wherein the VH region comprises the sequence of amino acids set forth in SEQ ID NO: 26 and the VL region comprises the sequence of amino acids set forth in SEQ ID NO: 39.
17. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-16, wherein the antibody is a full length antibody.
18. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 17, wherein the full length antibody comprises a constant region (Fc) from an IgAl, IgA2, IgD, IgE, IgGl, IgG2, IgG3, IgG4, or IgM.
19. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 18, wherein the constant region is a human IgGl heavy chain constant region.
20. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 19, wherein the human IgGl heavy chain constant region comprises one or more of mutations selected from the group consisting of G236A, S239D, M252Y, S254T, T256E, A330L, I332E, M428L, and N434S, numbered according to the EU numbering system.
21. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 19 or 20, wherein the human IgGl heavy chain constant region comprises S239D/A330L/I332E mutations, numbered according to the EU numbering system.
22. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising S239D/I332E mutations, numbered according to the EU numbering system.
23. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising G236A/S239D/A330L/I332E mutations, numbered according to the EU numbering system.
24. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising G236A/A330L/I332E mutations, numbered according to the EU numbering system.
25. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising M252Y/S254T/T256E mutations, numbered according to the EU numbering system.
26. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 19 or 20, wherein the antibody comprises a human IgGl heavy chain constant region comprising M428L/N434S mutations, numbered according to the EU numbering system.
27. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-16, wherein the antibody is an antigen-binding fragment.
28. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 27, wherein the antigen-binding fragment is selected from the group consisting of a single domain antibody, a single chain antibody, an unibody, a single chain variable fragment (scFv), a Fab fragment, and a F(ab')2 fragment.
29. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-16, wherein the anti-CTLA-4 antibody or antigen-binding fragment thereof is recombinant.
30. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-29, wherein the VH region and the VL region is human or is from a human protein.
31. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-30, wherein the ratio of dissociation constant for binding at acidic pH (e.g., pH 5.0 to 6.0) to the dissociation constant for binding at physiological pH (e.g., pH 7.35 to 7.45) to human CTLA-4 protein is greater than 2.
32. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-31, wherein the antibody or antigen-binding fragment exhibits a ratio of a dissociation constant for binding at acidic pH (e.g., pH 5.0 to 6.0) to dissociation constant for binding at physiological pH (e.g., pH 7.35 to 7.45) of more than 7.
33. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-32, wherein the ratio of EC50 value of the antibody or antigen-binding fragment thereof for human CTLA- 4 at physiological pH (e.g., pH 7.35 to 7.45) to EC50 value of the antibody or antigen-binding fragment thereof for human CTLA-4 at acidic pH (e.g., pH 5.0 to 6.0) is greater than 2.
34. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-33, wherein the antibody or antigen-binding fragment thereof possesses pH dependent binding properties to human CTLA-4 that facilitates the dissociation of antibody from the antibody: CTLA-4 complex upon internalization into a cell, allowing either the antibody, CTLA-4, or both to be recycled instead of being degraded in the lysosome.
35. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-34, wherein the antibody or antigen-binding fragment thereof preserves high affinity for CTLA-4 at pH>7, lower affinity at pH 5.5-7, and significantly reduced affinity for CTLA-4 at pH<5.5.
36. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-35, wherein the antibody or antigen-binding fragment thereof induces strong deletion of Treg and local T cell activation in tumor microenvironment but minimal systemic T cell activation.
37. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-36, wherein the antibody or antigen-binding fragment thereof induces a favorable ratio of regulatory T cells (Treg cells) to T effector cells in the tumor microenvironment.
38. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-37, wherein the antibody or antigen-binding fragment thereof induces a favorable ratio of immune cell subsets in the tumor or blood.
39. The anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-38, wherein the antibody or antigen-binding fragment induces a favorable ratio of CTLA-4 expression on immune cell subsets in the tumor or blood.
40. The anti-CTLA-4 antibody or antigen-binding fragment thereof of claim 38 or 39, wherein the immune cell subsets comprise myeloid cells, T cells, CD4+ T cells, CD8+ T cells, and/or Treg cells.
41. A conjugate, comprising the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-40 and a heterologous molecule or moiety.
42. The conjugate of claim 41, wherein the heterologous molecule or moiety is a cytotoxic agent, a drug, enzymatically active toxin or fragment thereof, or a radioactive atom.
43. A bispecific antibody, comprising the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-40.
44. A polynucleotide comprising a nucleic acid encoding the anti-CTLA-4 antibody or antigen-binding domain thereof of any of claims 1-40.
45. A vector, comprising the polynucleotide of claim 44.
46. The vector of claim 45, wherein the vector is a viral vector.
47. The vector of claim 46, wherein the viral vector is a retroviral vector or a lentiviral vector.
48. A cell comprising the polynucleotide of claim 41, or the vector of any of claims 45-47.
49. The cell of claim 48, wherein the cell is a mammalian cell.
50. A method of producing an antibody comprising culturing the cell of claim 48 or 49 under a condition that produces the antibody.
51. The method of claim 50, further comprising recovering the antibody produced by the cell.
52. An antibody or antigen-binding fragment thereof produced by the method of claim 50 or claim 51.
53. A composition comprising the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-40 and 52, the conjugate of claim 41 or 42, the bispecific antibody of claim 43.
54. The composition of claim 53, further comprising a pharmaceutically acceptable excipient.
55. A method of stimulating an immune response in a subject, the method comprising administering to a subject in need thereof the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-40 and 52, the conjugate of claim 41 or 42, the bispecific antibody of claim 43 or the composition of claim 53 or 54.
56. The method of claim 55, wherein stimulating an immune response treats a disease or condition in the subject.
57. The method of claim 56, wherein the disease or condition is a cancer.
58. A method of cancer immunotherapy, the method comprising administering to a subject in need thereof a therapeutically effective amount of the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-40 and 52, the conjugate of claim 41 or 42, the bispecific antibody of claim 43 or the composition of claim 53 or 54.
59. The method of claim 58, wherein the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-40 and 52, the conjugate of claim 41 or 42, the bispecific antibody of claim 43 or the composition of claim 53 or 54 is administered to the subject in need thereof at about 0.01 pg/kg body weight to about 20 mg/kg body weight.
60. The method of claim 58 or 59, further comprising administering to the subject an effective amount of at least one additional therapeutic agent.
61. The method of claim 60, wherein the at least one additional therapeutic agent is selected from the group consisting of: viral gene therapy, immune checkpoint inhibitors, target therapies, radiation therapies, vaccination therapies, and chemotherapies.
62. The method of claim 61 , wherein the at least one additional therapeutic agent is selected from the group consisting of: pomalyst, revlimid, lenalidomide, pomalidomide, thalidomide, a DNA- alkylating platinum-containing derivative, cisplatin, 5-fluorouracil, cyclophosphamide, an anti-CD137 antibody, an anti-PD-1 antibody, an anti-PD-Ll antibody, an anti-CD20 antibody, an anti-CD40 antibody, an anti-DR5 antibody, an anti-CDld antibody, an anti-TIM3 antibody, an anti-SLAMF7 antibody, an anti-KIR receptor antibody, an anti-OX40 antibody, an anti-HER2 antibody, an anti-ErbB- 2 antibody, an anti-EGFR antibody, cetuximab, rituximab, trastuzumab, pembrolizumab, an antibody drug conjugation, radiotherapy, single dose radiation, fractionated radiation, focal radiation, whole organ radiation, IL-12, IFNa, GM-CSF, a chimeric antigen receptor, adoptively transferred T cells, an anti-cancer vaccine, and an oncolytic virus.
63. The method of any of claims 58-62, wherein the method comprises administering to the subject an effective amount of the anti-CTLA-4 antibody or antigen-binding fragment thereof of any of claims 1-40 and 52, the conjugate of claim 41 or 42, the bispecific antibody of claim 43 or the composition of claim 53 or 54 prior to a surgery or after a surgery to remove a solid tumor in the subject.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363532336P | 2023-08-11 | 2023-08-11 | |
| US63/532,336 | 2023-08-11 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025038492A1 true WO2025038492A1 (en) | 2025-02-20 |
Family
ID=92543311
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/041822 Pending WO2025038492A1 (en) | 2023-08-11 | 2024-08-09 | Anti-ctla-4 antibodies and related binding molecules and methods and uses thereof |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025038492A1 (en) |
Citations (55)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1987004462A1 (en) | 1986-01-23 | 1987-07-30 | Celltech Limited | Recombinant dna sequences, vectors containing them and method for the use thereof |
| US4946778A (en) | 1987-09-21 | 1990-08-07 | Genex Corporation | Single polypeptide chain binding molecules |
| US5091513A (en) | 1987-05-21 | 1992-02-25 | Creative Biomolecules, Inc. | Biosynthetic antibody binding sites |
| US5132405A (en) | 1987-05-21 | 1992-07-21 | Creative Biomolecules, Inc. | Biosynthetic antibody binding sites |
| WO1993003769A1 (en) | 1991-08-20 | 1993-03-04 | THE UNITED STATES OF AMERICA, represented by THE SECRETARY, DEPARTEMENT OF HEALTH AND HUMAN SERVICES | Adenovirus mediated transfer of genes to the gastrointestinal tract |
| US5208020A (en) | 1989-10-25 | 1993-05-04 | Immunogen Inc. | Cytotoxic agents comprising maytansinoids and their therapeutic use |
| WO1993009239A1 (en) | 1991-11-08 | 1993-05-13 | Research Corporation Technologies, Inc. | Adeno-associated virus-2 basal vectors |
| WO1993019191A1 (en) | 1992-03-16 | 1993-09-30 | Centre National De La Recherche Scientifique | Defective recombinant adenoviruses expressing cytokines for use in antitumoral treatment |
| US5283173A (en) | 1990-01-24 | 1994-02-01 | The Research Foundation Of State University Of New York | System to detect protein-protein interactions |
| WO1994012649A2 (en) | 1992-12-03 | 1994-06-09 | Genzyme Corporation | Gene therapy for cystic fibrosis |
| WO1994028938A1 (en) | 1993-06-07 | 1994-12-22 | The Regents Of The University Of Michigan | Adenovirus vectors for gene therapy sponsorship |
| WO1995000655A1 (en) | 1993-06-24 | 1995-01-05 | Mc Master University | Adenovirus vectors for gene therapy |
| WO1995011984A2 (en) | 1993-10-25 | 1995-05-04 | Canji, Inc. | Recombinant adenoviral vector and methods of use |
| EP0425235B1 (en) | 1989-10-25 | 1996-09-25 | Immunogen Inc | Cytotoxic agents comprising maytansinoids and their therapeutic use |
| WO1997030087A1 (en) | 1996-02-16 | 1997-08-21 | Glaxo Group Limited | Preparation of glycosylated antibodies |
| WO1998058964A1 (en) | 1997-06-24 | 1998-12-30 | Genentech, Inc. | Methods and compositions for galactosylated glycoproteins |
| WO1999022764A1 (en) | 1997-10-31 | 1999-05-14 | Genentech, Inc. | Methods and compositions comprising glycoprotein glycoforms |
| WO2000061739A1 (en) | 1999-04-09 | 2000-10-19 | Kyowa Hakko Kogyo Co., Ltd. | Method for controlling the activity of immunologically functional molecule |
| WO2001029058A1 (en) | 1999-10-15 | 2001-04-26 | University Of Massachusetts | Rna interference pathway genes as tools for targeted genetic interference |
| WO2001029246A1 (en) | 1999-10-19 | 2001-04-26 | Kyowa Hakko Kogyo Co., Ltd. | Process for producing polypeptide |
| US6326193B1 (en) | 1999-11-05 | 2001-12-04 | Cambria Biosciences, Llc | Insect control agent |
| WO2001096584A2 (en) | 2000-06-12 | 2001-12-20 | Akkadix Corporation | Materials and methods for the control of nematodes |
| WO2002031140A1 (en) | 2000-10-06 | 2002-04-18 | Kyowa Hakko Kogyo Co., Ltd. | Cells producing antibody compositions |
| US20020164328A1 (en) | 2000-10-06 | 2002-11-07 | Toyohide Shinkawa | Process for purifying antibody |
| WO2003011878A2 (en) | 2001-08-03 | 2003-02-13 | Glycart Biotechnology Ag | Antibody glycosylation variants having increased antibody-dependent cellular cytotoxicity |
| US20030115614A1 (en) | 2000-10-06 | 2003-06-19 | Yutaka Kanda | Antibody composition-producing cell |
| US6602684B1 (en) | 1998-04-20 | 2003-08-05 | Glycart Biotechnology Ag | Glycosylation engineering of antibodies for improving antibody-dependent cellular cytotoxicity |
| US20030157108A1 (en) | 2001-10-25 | 2003-08-21 | Genentech, Inc. | Glycoprotein compositions |
| WO2003085107A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | Cells with modified genome |
| WO2003084570A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | DRUG CONTAINING ANTIBODY COMPOSITION APPROPRIATE FOR PATIENT SUFFERING FROM FcϜRIIIa POLYMORPHISM |
| WO2003085119A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | METHOD OF ENHANCING ACTIVITY OF ANTIBODY COMPOSITION OF BINDING TO FcϜ RECEPTOR IIIa |
| US20040093621A1 (en) | 2001-12-25 | 2004-05-13 | Kyowa Hakko Kogyo Co., Ltd | Antibody composition which specifically binds to CD20 |
| US20040110282A1 (en) | 2002-04-09 | 2004-06-10 | Kyowa Hakko Kogyo Co., Ltd. | Cells in which activity of the protein involved in transportation of GDP-fucose is reduced or lost |
| US20040109865A1 (en) | 2002-04-09 | 2004-06-10 | Kyowa Hakko Kogyo Co., Ltd. | Antibody composition-containing medicament |
| WO2004056312A2 (en) | 2002-12-16 | 2004-07-08 | Genentech, Inc. | Immunoglobulin variants and uses thereof |
| US20040132140A1 (en) | 2002-04-09 | 2004-07-08 | Kyowa Hakko Kogyo Co., Ltd. | Production process for antibody composition |
| WO2005035778A1 (en) | 2003-10-09 | 2005-04-21 | Kyowa Hakko Kogyo Co., Ltd. | PROCESS FOR PRODUCING ANTIBODY COMPOSITION BY USING RNA INHIBITING THE FUNCTION OF α1,6-FUCOSYLTRANSFERASE |
| WO2005035586A1 (en) | 2003-10-08 | 2005-04-21 | Kyowa Hakko Kogyo Co., Ltd. | Fused protein composition |
| US6884869B2 (en) | 2001-04-30 | 2005-04-26 | Seattle Genetics, Inc. | Pentapeptide compounds and uses related thereto |
| US20050123546A1 (en) | 2003-11-05 | 2005-06-09 | Glycart Biotechnology Ag | Antigen binding molecules with increased Fc receptor binding affinity and effector function |
| WO2005053742A1 (en) | 2003-12-04 | 2005-06-16 | Kyowa Hakko Kogyo Co., Ltd. | Medicine containing antibody composition |
| US20060270045A1 (en) | 2003-10-22 | 2006-11-30 | Keck Graduate Institute | Methods of synthesizing heteromultimeric polypeptides in yeast using a haploid mating strategy |
| US7256257B2 (en) | 2001-04-30 | 2007-08-14 | Seattle Genetics, Inc. | Pentapeptide compounds and uses related thereto |
| US7317091B2 (en) | 2002-03-01 | 2008-01-08 | Xencor, Inc. | Optimized Fc variants |
| US7498298B2 (en) | 2003-11-06 | 2009-03-03 | Seattle Genetics, Inc. | Monomethylvaline compounds capable of conjugation to ligands |
| US7521541B2 (en) | 2004-09-23 | 2009-04-21 | Genetech Inc. | Cysteine engineered antibodies and conjugates |
| WO2009067800A1 (en) | 2007-11-27 | 2009-06-04 | Viventia Biotech Inc. | Antibodies against a cancer-associated epitope of variant nfkbib and uses thereof |
| US20090226421A1 (en) | 2005-11-28 | 2009-09-10 | Genmab A/S | Recombinant monovalent antibodies and methods for production thereof |
| WO2009125825A1 (en) * | 2008-04-11 | 2009-10-15 | 中外製薬株式会社 | Antigen-binding molecule capable of binding to two or more antigen molecules repeatedly |
| WO2011056983A1 (en) | 2009-11-05 | 2011-05-12 | Genentech, Inc. | Zirconium-radiolabeled, cysteine engineered antibody conjugates |
| WO2011133886A2 (en) | 2010-04-23 | 2011-10-27 | Genentech, Inc. | Production of heteromultimeric proteins |
| US20140322129A1 (en) | 2013-03-14 | 2014-10-30 | Genentech, Inc. | Anti-b7-h4 antibodies and immunoconjugates |
| US10196445B1 (en) | 2015-03-17 | 2019-02-05 | Bristol-Myers Squibb Company | Ipilimumab variant with enhanced ADCC |
| WO2020214748A1 (en) * | 2019-04-18 | 2020-10-22 | Bristol-Myers Squibb Company | Ipilimumab variants with enhanced specificity for binding at low ph |
| US10912831B1 (en) | 2016-12-07 | 2021-02-09 | Agenus Inc. | Anti-CTLA-4 antibodies and methods of use thereof |
-
2024
- 2024-08-09 WO PCT/US2024/041822 patent/WO2025038492A1/en active Pending
Patent Citations (61)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1987004462A1 (en) | 1986-01-23 | 1987-07-30 | Celltech Limited | Recombinant dna sequences, vectors containing them and method for the use thereof |
| US5091513A (en) | 1987-05-21 | 1992-02-25 | Creative Biomolecules, Inc. | Biosynthetic antibody binding sites |
| US5132405A (en) | 1987-05-21 | 1992-07-21 | Creative Biomolecules, Inc. | Biosynthetic antibody binding sites |
| US4946778A (en) | 1987-09-21 | 1990-08-07 | Genex Corporation | Single polypeptide chain binding molecules |
| US5208020A (en) | 1989-10-25 | 1993-05-04 | Immunogen Inc. | Cytotoxic agents comprising maytansinoids and their therapeutic use |
| EP0425235B1 (en) | 1989-10-25 | 1996-09-25 | Immunogen Inc | Cytotoxic agents comprising maytansinoids and their therapeutic use |
| US5283173A (en) | 1990-01-24 | 1994-02-01 | The Research Foundation Of State University Of New York | System to detect protein-protein interactions |
| US5468614A (en) | 1990-01-24 | 1995-11-21 | The Research Foundation Of State University Of New York | System to detect protein-protein interactions |
| WO1993003769A1 (en) | 1991-08-20 | 1993-03-04 | THE UNITED STATES OF AMERICA, represented by THE SECRETARY, DEPARTEMENT OF HEALTH AND HUMAN SERVICES | Adenovirus mediated transfer of genes to the gastrointestinal tract |
| WO1993009239A1 (en) | 1991-11-08 | 1993-05-13 | Research Corporation Technologies, Inc. | Adeno-associated virus-2 basal vectors |
| WO1993019191A1 (en) | 1992-03-16 | 1993-09-30 | Centre National De La Recherche Scientifique | Defective recombinant adenoviruses expressing cytokines for use in antitumoral treatment |
| WO1994012649A2 (en) | 1992-12-03 | 1994-06-09 | Genzyme Corporation | Gene therapy for cystic fibrosis |
| WO1994028938A1 (en) | 1993-06-07 | 1994-12-22 | The Regents Of The University Of Michigan | Adenovirus vectors for gene therapy sponsorship |
| WO1995000655A1 (en) | 1993-06-24 | 1995-01-05 | Mc Master University | Adenovirus vectors for gene therapy |
| WO1995011984A2 (en) | 1993-10-25 | 1995-05-04 | Canji, Inc. | Recombinant adenoviral vector and methods of use |
| WO1997030087A1 (en) | 1996-02-16 | 1997-08-21 | Glaxo Group Limited | Preparation of glycosylated antibodies |
| WO1998058964A1 (en) | 1997-06-24 | 1998-12-30 | Genentech, Inc. | Methods and compositions for galactosylated glycoproteins |
| WO1999022764A1 (en) | 1997-10-31 | 1999-05-14 | Genentech, Inc. | Methods and compositions comprising glycoprotein glycoforms |
| US6602684B1 (en) | 1998-04-20 | 2003-08-05 | Glycart Biotechnology Ag | Glycosylation engineering of antibodies for improving antibody-dependent cellular cytotoxicity |
| WO2000061739A1 (en) | 1999-04-09 | 2000-10-19 | Kyowa Hakko Kogyo Co., Ltd. | Method for controlling the activity of immunologically functional molecule |
| WO2001029058A1 (en) | 1999-10-15 | 2001-04-26 | University Of Massachusetts | Rna interference pathway genes as tools for targeted genetic interference |
| WO2001029246A1 (en) | 1999-10-19 | 2001-04-26 | Kyowa Hakko Kogyo Co., Ltd. | Process for producing polypeptide |
| US6326193B1 (en) | 1999-11-05 | 2001-12-04 | Cambria Biosciences, Llc | Insect control agent |
| WO2001096584A2 (en) | 2000-06-12 | 2001-12-20 | Akkadix Corporation | Materials and methods for the control of nematodes |
| WO2002031140A1 (en) | 2000-10-06 | 2002-04-18 | Kyowa Hakko Kogyo Co., Ltd. | Cells producing antibody compositions |
| US20020164328A1 (en) | 2000-10-06 | 2002-11-07 | Toyohide Shinkawa | Process for purifying antibody |
| US20030115614A1 (en) | 2000-10-06 | 2003-06-19 | Yutaka Kanda | Antibody composition-producing cell |
| US7423116B2 (en) | 2001-04-30 | 2008-09-09 | Seattle Genetics Inc. | Pentapeptide compounds and uses related thereto |
| US6884869B2 (en) | 2001-04-30 | 2005-04-26 | Seattle Genetics, Inc. | Pentapeptide compounds and uses related thereto |
| US7256257B2 (en) | 2001-04-30 | 2007-08-14 | Seattle Genetics, Inc. | Pentapeptide compounds and uses related thereto |
| US7098308B2 (en) | 2001-04-30 | 2006-08-29 | Seattle Genetics, Inc. | Pentapeptide compounds and uses related thereto |
| WO2003011878A2 (en) | 2001-08-03 | 2003-02-13 | Glycart Biotechnology Ag | Antibody glycosylation variants having increased antibody-dependent cellular cytotoxicity |
| US20030157108A1 (en) | 2001-10-25 | 2003-08-21 | Genentech, Inc. | Glycoprotein compositions |
| US20040093621A1 (en) | 2001-12-25 | 2004-05-13 | Kyowa Hakko Kogyo Co., Ltd | Antibody composition which specifically binds to CD20 |
| US7317091B2 (en) | 2002-03-01 | 2008-01-08 | Xencor, Inc. | Optimized Fc variants |
| WO2003084570A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | DRUG CONTAINING ANTIBODY COMPOSITION APPROPRIATE FOR PATIENT SUFFERING FROM FcϜRIIIa POLYMORPHISM |
| US20040132140A1 (en) | 2002-04-09 | 2004-07-08 | Kyowa Hakko Kogyo Co., Ltd. | Production process for antibody composition |
| US20040110704A1 (en) | 2002-04-09 | 2004-06-10 | Kyowa Hakko Kogyo Co., Ltd. | Cells of which genome is modified |
| US20040109865A1 (en) | 2002-04-09 | 2004-06-10 | Kyowa Hakko Kogyo Co., Ltd. | Antibody composition-containing medicament |
| WO2003085107A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | Cells with modified genome |
| US20040110282A1 (en) | 2002-04-09 | 2004-06-10 | Kyowa Hakko Kogyo Co., Ltd. | Cells in which activity of the protein involved in transportation of GDP-fucose is reduced or lost |
| WO2003085119A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | METHOD OF ENHANCING ACTIVITY OF ANTIBODY COMPOSITION OF BINDING TO FcϜ RECEPTOR IIIa |
| WO2004056312A2 (en) | 2002-12-16 | 2004-07-08 | Genentech, Inc. | Immunoglobulin variants and uses thereof |
| WO2005035586A1 (en) | 2003-10-08 | 2005-04-21 | Kyowa Hakko Kogyo Co., Ltd. | Fused protein composition |
| WO2005035778A1 (en) | 2003-10-09 | 2005-04-21 | Kyowa Hakko Kogyo Co., Ltd. | PROCESS FOR PRODUCING ANTIBODY COMPOSITION BY USING RNA INHIBITING THE FUNCTION OF α1,6-FUCOSYLTRANSFERASE |
| US20060270045A1 (en) | 2003-10-22 | 2006-11-30 | Keck Graduate Institute | Methods of synthesizing heteromultimeric polypeptides in yeast using a haploid mating strategy |
| US20050123546A1 (en) | 2003-11-05 | 2005-06-09 | Glycart Biotechnology Ag | Antigen binding molecules with increased Fc receptor binding affinity and effector function |
| US7745394B2 (en) | 2003-11-06 | 2010-06-29 | Seattle Genetics, Inc. | Monomethylvaline compounds capable of conjugation to ligands |
| US7498298B2 (en) | 2003-11-06 | 2009-03-03 | Seattle Genetics, Inc. | Monomethylvaline compounds capable of conjugation to ligands |
| WO2005053742A1 (en) | 2003-12-04 | 2005-06-16 | Kyowa Hakko Kogyo Co., Ltd. | Medicine containing antibody composition |
| US7521541B2 (en) | 2004-09-23 | 2009-04-21 | Genetech Inc. | Cysteine engineered antibodies and conjugates |
| US7855275B2 (en) | 2004-09-23 | 2010-12-21 | Genentech, Inc. | Cysteine engineered antibodies and conjugates |
| US20090226421A1 (en) | 2005-11-28 | 2009-09-10 | Genmab A/S | Recombinant monovalent antibodies and methods for production thereof |
| WO2009067800A1 (en) | 2007-11-27 | 2009-06-04 | Viventia Biotech Inc. | Antibodies against a cancer-associated epitope of variant nfkbib and uses thereof |
| WO2009125825A1 (en) * | 2008-04-11 | 2009-10-15 | 中外製薬株式会社 | Antigen-binding molecule capable of binding to two or more antigen molecules repeatedly |
| WO2011056983A1 (en) | 2009-11-05 | 2011-05-12 | Genentech, Inc. | Zirconium-radiolabeled, cysteine engineered antibody conjugates |
| WO2011133886A2 (en) | 2010-04-23 | 2011-10-27 | Genentech, Inc. | Production of heteromultimeric proteins |
| US20140322129A1 (en) | 2013-03-14 | 2014-10-30 | Genentech, Inc. | Anti-b7-h4 antibodies and immunoconjugates |
| US10196445B1 (en) | 2015-03-17 | 2019-02-05 | Bristol-Myers Squibb Company | Ipilimumab variant with enhanced ADCC |
| US10912831B1 (en) | 2016-12-07 | 2021-02-09 | Agenus Inc. | Anti-CTLA-4 antibodies and methods of use thereof |
| WO2020214748A1 (en) * | 2019-04-18 | 2020-10-22 | Bristol-Myers Squibb Company | Ipilimumab variants with enhanced specificity for binding at low ph |
Non-Patent Citations (93)
| Title |
|---|
| ALEGRE ET AL., J IMMUNOL, vol. 148, 1992, pages 3461 - 3468 |
| ALEGRE MLFRAUWIRTH KATHOMPSON CB: "T-cell regulation by CD28 and CTLA-4", NAT REV IMMUNOL., vol. 1, no. 3, 2001, pages 220 - 228, XP055345954, DOI: 10.1038/35105024 |
| ALI ET AL., HUM. GENE THER., vol. 9, 1998, pages 81 - 86 |
| ALI ET AL., HUM. MOL. GENET., vol. 5, 1996, pages 591 - 594 |
| AL-LAZIKANI ET AL., J MOL BIOL, vol. 273, no. 4, 1997, pages 927 - 48 |
| AL-LAZIKANI ET AL., J MOL BIOL., vol. 273, no. 4, 1997, pages 927 - 48 |
| ALTMAN AKONG K-F: "pH-sensitive anti-CTLA4 antibodies: yes to efficacy, no to toxicity", CELL RES., vol. 29, no. 8, 2019, pages 601 - 602 |
| BASKAR RLEE KAYEO RYEOH K-W: "Cancer and radiation therapy: current advances and future directions", INT J MED SCI., vol. 9, no. 3, 2012, pages 193 - 199, XP055668349, DOI: 10.7150/ijms.3635 |
| BAULU EGARDET CCHUVIN NDEPIL S: "TCR-engineered T cell therapy in solid tumors: State of the art and perspectives", SCI ADV., vol. 9, no. 7, 2023, pages eadf3700, XP093068209, DOI: 10.1126/sciadv.adf3700 |
| BENNETT ET AL., INVEST. OPTHALMOL. VIS. SCI., vol. 38, 1997, pages 2857 - 2863 |
| BERG MZAVAZAVA N.: "Regulation of CD28 expression on CD8+ T cells by CTLA-4", J LEUKOC BIOL., vol. 83, no. 4, 2008, pages 853 - 863 |
| BORRAS ET AL., GENE THER., vol. 6, 1999, pages 515 - 524 |
| CHARI ET AL., CANCER RESEARCH, vol. 52, 1992, pages 127 - 131 |
| CHOTHIA C.LESK A.M., J MOL BIOL, vol. 196, no. 4, 1987, pages 901 - 17 |
| CHOTHIA C.LESK A.M., J MOL BIOL., vol. 196, no. 4, 1987, pages 901 - 17 |
| CHOTHIA ET AL., SCIENCE, vol. 233, no. 4765, 1986, pages 755 - 8 |
| CHOWDHURY, METHODS MOL. BIOL., vol. 207, 2008, pages 179 - 196 |
| CLARKSON ET AL., NATURE, vol. 352, 1991, pages 624 - 628 |
| COMPTON K ET AL.: "Cancer Incidence, Mortality, Years of Life Lost, Years Lived With Disability, and Disability-Adjusted Life Years for 29 Cancer Groups From 2010 to 2019: A Systematic Analysis for the Global Burden of Disease Study 2019", JAMA ONCOL., vol. 8, no. 3, 2022, pages 420 - 444 |
| DU XLIU MSU J ET AL.: "Uncoupling therapeutic from immunotherapy-related adverse effects for safer and effective anti-CTLA-4 antibodies in CTLA4 humanized mice", CELL RES., vol. 28, no. 4, 2018, pages 433 - 447 |
| EHRLICH ET AL., BIOCHEM, vol. 19, 1980, pages 4091 - 4096 |
| FEINERMAN OGERMAIN RNALTAN-BONNET G.: "Quantitative challenges in understanding ligand discrimination by alphabeta T cells", MOL IMMUNOL., vol. 45, no. 3, 2008, pages 619 - 631, XP022287698, DOI: 10.1016/j.molimm.2007.03.028 |
| FLANNERY ET AL., PROC. NATL. ACAD. SCI. USA, vol. 94, 1997, pages 10319 - 6921 |
| FLOTTE ET AL., PROC. NATL. ACAD. SCI. USA, vol. 90, 1993, pages 10613 - 10617 |
| FRAUWIRTH KARILEY JLHARRIS MH ET AL.: "The CD28 signaling pathway regulates glucose metabolism", IMMUNITY, vol. 16, no. 6, 2002, pages 769 - 777 |
| GALVEZ-CANCINO FSIMPSON APCOSTOYA C ET AL.: "Fcy receptors and immunomodulatory antibodies in cancer", NAT REV CANCER., vol. 24, no. 1, 2024, pages 51 - 71 |
| GECKIN BKONSTANTIN FOHSE FDOMÍNGUEZ-ANDRÉS JNETEA MG: "Trained immunity: implications for vaccination", CURR OPIN IMMUNOL., vol. 77, 2022, pages 102190, XP087129969, DOI: 10.1016/j.coi.2022.102190 |
| GILLIES ET AL.: "MRI of the Tumor Microenvironment", JOURNAL OF MAGNETIC RESONANCE IMAGING, vol. 16, 2002, pages 430 - 450, XP055045243, DOI: 10.1002/jmri.10181 |
| GUAN ET AL., PNAS, vol. 95, 1998, pages 13206 - 10 |
| GYANANI VHALEY JCGOSWAMI R.: "Challenges of Current Anticancer Treatment Approaches with Focus on Liposomal Drug Delivery Systems", PHARMACEUTICALS (BASEL), vol. 14, no. 9, 2021 |
| HAMEL ET AL., MOLECULAR PHARMACOLOGY, vol. 47, 1995, pages 965 - 976 |
| HAMEL ET AL., THE JOURNAL OF BIOLOGICAL CHEMISTRY, vol. 265, no. 28, 1990, pages 17141 - 17149 |
| HASSANPOUR SHDEHGHANI M.: "Review of cancer from perspective of molecular", JOURNAL OF CANCER RESEARCH AND PRACTICE., July 2017 (2017-07-01) |
| HOCHMAN ET AL., BIOCHEM, vol. 15, 1976, pages 2706 - 2710 |
| HONEGGER APLIICKTHUN A, J MOL BIOL, vol. 309, no. 3, 2001, pages 657 - 70 |
| HOOGENBOOM ET AL.: "Molecular Cloning, A Laboratory Manual", vol. 178, 2001, COLD SPRING HARBOR LABORATORY PRESS, pages: 1 - 37 |
| HOSSEN MMMA YYIN Z ET AL.: "Current understanding of CTLA-4: from mechanism to autoimmune diseases", FRONT IMMUNOL., vol. 14, 2023, pages 1198365 |
| HUSTON ET AL., PROC. NAT. ACAD. SCI. USA, vol. 85, no. 16, 1988, pages 5879 - 5883 |
| IDUSOGIE ET AL., J IMMUNOL, vol. 166, no. 4, 2001, pages 2571 - 5 |
| INBAR ET AL., PROC. NAT. ACAD. SCI. USA, vol. 69, 1972, pages 2659 - 2662 |
| JARASCH ET AL., PROTEINS, vol. 85, no. 1, 2017, pages 65 - 71 |
| JOMARY ET AL., GENE THER., vol. 4, 1997, pages 683 690 |
| KABAT, E.A. ET AL.: "Sequences of Proteins of Immunological Interest", 1991, US DEPARTMENT OF HEALTH AND HUMAN SERVICES, NIH PUBLICATION |
| KANDA, Y. ET AL., BIOTECHNOL. BIOENG., vol. 94, no. 4, 2006, pages 680 - 688 |
| KANEKONIWA, BIODRUGS, vol. 25, no. 1, 2011, pages 1 - 11 |
| KAUR BPSECORD E.: "Innate Immunity", IMMUNOL ALLERGY CLIN NORTH AM., vol. 41, no. 4, 2021, pages 535 - 541 |
| KENNEDY PTSAULTERS ELDUCKWORTH AD ET AL.: "Soluble CTLA-4 attenuates T cell activation and modulates anti-tumor immunity", MOL THER., vol. 32, no. 2, 2024, pages 457 - 468 |
| KNEZEVIC CECLARKE W.: "Cancer chemotherapy: the case for therapeutic drug monitoring", THER DRUG MONIT., vol. 42, no. 1, 2020, pages 6 - 19 |
| KRUMMEL MFALLISON JP: "CD28 and CTLA-4 have opposing effects on the response of T cells to stimulation", J EXP MED., vol. 182, no. 2, 1995, pages 459 - 465, XP001031165, DOI: 10.1084/jem.182.2.459 |
| KUMAR BVCONNORS TJFARBER DL: "Human T Cell Development, Localization, and Function throughout Life", IMMUNITY, vol. 48, no. 2, 2018, pages 202 - 213 |
| LABRIJIN, NAT. BIOTECHNOL., vol. 27, no. 8, 2009, pages 767 - 71 |
| LAZAR ET AL., PNAS, vol. 103, no. 11, 2006, pages 4005 - 4010 |
| LEFRANC MP ET AL., DEV COMP IMMUNOL, vol. 27, no. 1, 2003, pages 55 - 77 |
| LI ET AL., INVEST. OPTHALMOL. VIS. SCI., vol. 35, 1994, pages 2543 - 2549 |
| LIDAVIDSON, PROC. NATL. ACAD. SCI. USA, vol. 92, 1995, pages 7700 - 7704 |
| LIU YZHENG P: "Preserving the CTLA-4 Checkpoint for Safer and More Effective Cancer Immunotherapy", TRENDS PHARMACOL SCI., vol. 41, no. 1, 2020, pages 4 - 12, XP085972925, DOI: 10.1016/j.tips.2019.11.003 |
| MACCALLUM ET AL., J. MOL. BIOL, vol. 262, 1996, pages 732 - 745 |
| MARTIN ET AL., PNAS, vol. 86, no. 23, 1989, pages 9268 - 9272 |
| MARTIN ET AL.: "Handbook of Therapeutic Antibodies", vol. 1, WILEY-VCH, article "Bioinformatics tools for antibody engineering" |
| MARTIN, A.C.R.: "Antibody Engineering", 2010, SPRINGER PROTOCOLS HANDBOOKS. SPRINGER, article "Protein Sequence and Structure Analysis of Antibody Variable Domains" |
| MENDELSON ET AL., VIROL., vol. 166, 1988, pages 154 - 165 |
| MOORE ET AL., MABS, vol. 2, no. 2, 2010, pages 181 - 189 |
| MOREA ET AL., BIOPHYS CHEM, vol. 68, no. 1-3, 1997, pages 9 - 16 |
| MOREA ET AL., J MOL BIOL., vol. 275, no. 2, 1998, pages 269 - 94 |
| MULLER ET AL., ANAL. CHEM, vol. 78, 2006, pages 4390 - 4397 |
| NATSUME ET AL., CANCER RES, vol. 68, no. 10, 2008, pages 3863 - 72 |
| OKAZAKI ET AL., J. MOL. BIOL., vol. 336, 2004, pages 1239 - 1249 |
| PEGGS KSQUEZADA SACHAMBERS CAKORMAN AJALLISON JP: "Blockade of CTLA-4 on both effector and regulatory T cell compartments contributes to the antitumor activity of anti-CTLA-4 antibodies", J EXP MED., vol. 206, no. 8, 2009, pages 1717 - 1725 |
| PORTOLANO ET AL., J. IMMUNOL., vol. 150, 1993, pages 880 - 887 |
| QURESHI OSKAUR SHOU TZ ET AL.: "Constitutive clathrin-mediated endocytosis of CTLA-4 persists during T cell activation", J BIOL CHEM., vol. 287, no. 12, 2012, pages 9429 - 9440, XP055850230, DOI: 10.1074/jbc.M111.304329 |
| RAMOS-CASALS MBRAHMER JRCALLAHAN MK ET AL.: "Immune-related adverse events of checkpoint inhibitors", NAT REV DIS PRIMERS., vol. 6, no. 1, 2020, pages 38, XP037131257, DOI: 10.1038/s41572-020-0160-6 |
| RIPKA ET AL., ARCH. BIOCHEM. BIOPHYS., vol. 249, 1986, pages 533 - 545 |
| ROLLING ET AL., HUM. GENE THER., vol. 10, 1999, pages 641 - 648 |
| SAKAMOTO ET AL., H. GENE THER., vol. 5, 1999, pages 1088 - 1097 |
| SAMBROOK ET AL.: "Molecular Cloning: A Laboratory Manual", vol. 1-4, 2012, COLD SPRING HARBOR PRESS |
| SAMULSKI ET AL., J. VIR., vol. 63, 1989, pages 3822 - 3828 |
| SANAI NBERGER MS: "Surgical oncology for gliomas: the state of the art", NAT REV CLIN ONCOL., vol. 15, no. 2, 2018, pages 112 - 125 |
| SCATCHARD ET AL., ANN. N.Y. ACAD. SCI., vol. 51, 1949, pages 660 |
| SHAH KAL-HAIDARI ASUN JKAZI JU: "T cell receptor (TCR) signaling in health and disease", SIGNAL TRANSDUCT TARGET THER., vol. 6, no. 1, 2021, pages 412 |
| SHARMA PGOSWAMI SRAYCHAUDHURI D ET AL.: "Immune checkpoint therapy-current perspectives and future directions", CELL, vol. 186, no. 8, 2023, pages 1652 - 1669, XP087297532, DOI: 10.1016/j.cell.2023.03.006 |
| SHIELDS ET AL., JBC, vol. 276, no. 9, 2001, pages 6591 - 6604 |
| SHINKAWA TNAKAMURA KYAMANE N ET AL.: "The absence of fucose but not the presence of galactose or bisecting N-acetylglucosamine of human IgG1 complex-type oligo saccharides shows the critical role of enhancing antibody-dependent cellular cytotoxicity", J BIOL CHEM., vol. 278, no. 5, 2003, pages 3466 - 3473 |
| STAVENHAGEN ET AL., ADVAN. ENZYME REGUL., vol. 48, 2008, pages 152 - 164 |
| STAVENHAGEN ET AL., CANCER RES, vol. 67, no. 18, 2007, pages 8882 - 8890 |
| TAKAHASHI ET AL., J. VIROL., vol. 73, 1999, pages 7812 - 7816 |
| TRAMONTANO ET AL., J MOL BIOL, vol. 215, no. 1, 1990, pages 175 - 82 |
| WAIGHT JDCHAND DDIETRICH S ET AL.: "Selective FcγR Co-engagement on APCs Modulates the Activity of Therapeutic Antibodies Targeting T Cell Antigens", CANCER CELL, vol. 33, no. 6, 2018, pages 1033 - 1047, XP055713036, DOI: 10.1016/j.ccell.2018.05.005 |
| WARD FJDAHAL LNWIJESEKERA SK ET AL.: "The soluble isoform of CTLA-4 as a regulator of T-cell responses", EUR J IMMUNOL., vol. 43, no. 5, 2013, pages 1274 - 1285 |
| WILSON, SCIENCE, vol. 295, 2002, pages 2103 |
| WOLFF ET AL., CANCER RES., vol. 53, 1993, pages 2560 |
| YAMANE-OHNUKI ET AL., BIOTECH. BIOENG., vol. 87, 2004, pages 614 |
| YE ET AL., NUCLEIC ACIDS RES., 2013 |
| ZHANG YDU XLIU M ET AL.: "Hijacking antibody-induced CTLA-4 lysosomal degradation for safer and more effective cancer immunotherapy", CELL RES., vol. 29, no. 8, 2019, pages 609 - 627, XP036917063, DOI: 10.1038/s41422-019-0184-1 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US20230272084A1 (en) | B7-h3 directed antibody drug conjugates | |
| US20230272063A1 (en) | Anti-claudin 18 antibodies and methods of use thereof | |
| CN112513085B (en) | PSMA binders and their uses | |
| KR102584675B1 (en) | Antibodies specific for GUCY2c and their uses | |
| TWI870335B (en) | Variant chimeric 4d5 antibodies and uses thereof in combination with anti-pd-1 antibodies for the treatment of cancer | |
| US10160812B2 (en) | Bispecific HER2 antibodies | |
| CN113227142B (en) | anti-PD-1 antibodies and uses thereof | |
| KR102608723B1 (en) | Anti-PD-1 antibodies and uses thereof | |
| US20230112085A1 (en) | Anti-b7-h4 constructs and uses thereof | |
| KR20210062026A (en) | Anti-TNFRSF9 antibody and uses thereof | |
| WO2020153467A1 (en) | Novel cancer antigens and antibodies of said antigens | |
| JP2023524102A (en) | Antibodies specific for ABCB5 and uses thereof | |
| WO2025038492A1 (en) | Anti-ctla-4 antibodies and related binding molecules and methods and uses thereof | |
| CN117355331A (en) | Anti-Siglec compositions and uses thereof | |
| US12521446B2 (en) | Anti-CEACAM5 antibody drug conjugates | |
| WO2023109900A1 (en) | Anti-ox40 antibodies, multispecific antibodies and methods of use | |
| RU2800779C2 (en) | Bispecific antigen-binding molecules and methods of their use | |
| WO2025240670A2 (en) | Anti-pd-1 antibodies and related binding molecules and methods and uses thereof | |
| WO2025199352A2 (en) | Antibodies specific for solute carrier family 34 member 2 (slc34a2) | |
| WO2024147097A1 (en) | Her3-binding antibody-drug conjugate | |
| JP2024528217A (en) | Bispecific antibodies and methods of use | |
| HK40050386A (en) | Anti-pd-1 antibodies and uses thereof | |
| HK40050386B (en) | Anti-pd-1 antibodies and uses thereof | |
| EA048967B1 (en) | CLAUDINE 18 ANTIBODIES AND METHODS OF THEIR APPLICATION | |
| HK40049718A (en) | Anti-tnfrsf9 antibodies and uses thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24762190 Country of ref document: EP Kind code of ref document: A1 |