WO2024098023A2 - Interferon alpha polypeptides and conjugates - Google Patents
Interferon alpha polypeptides and conjugates Download PDFInfo
- Publication number
- WO2024098023A2 WO2024098023A2 PCT/US2023/078726 US2023078726W WO2024098023A2 WO 2024098023 A2 WO2024098023 A2 WO 2024098023A2 US 2023078726 W US2023078726 W US 2023078726W WO 2024098023 A2 WO2024098023 A2 WO 2024098023A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- ifnα
- conjugate
- polypeptide
- amino acid
- certain embodiments
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/52—Cytokines; Lymphokines; Interferons
- C07K14/555—Interferons [IFN]
- C07K14/56—IFN-alpha
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K45/00—Medicinal preparations containing active ingredients not provided for in groups A61K31/00 - A61K41/00
- A61K45/06—Mixtures of active ingredients without chemical characterisation, e.g. antiphlogistics and cardiaca
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K47/00—Medicinal preparations characterised by the non-active ingredients used, e.g. carriers or inert additives; Targeting or modifying agents chemically bound to the active ingredient
- A61K47/50—Medicinal preparations characterised by the non-active ingredients used, e.g. carriers or inert additives; Targeting or modifying agents chemically bound to the active ingredient the non-active ingredient being chemically bound to the active ingredient, e.g. polymer-drug conjugates
- A61K47/51—Medicinal preparations characterised by the non-active ingredients used, e.g. carriers or inert additives; Targeting or modifying agents chemically bound to the active ingredient the non-active ingredient being chemically bound to the active ingredient, e.g. polymer-drug conjugates the non-active ingredient being a modifying agent
- A61K47/56—Medicinal preparations characterised by the non-active ingredients used, e.g. carriers or inert additives; Targeting or modifying agents chemically bound to the active ingredient the non-active ingredient being chemically bound to the active ingredient, e.g. polymer-drug conjugates the non-active ingredient being a modifying agent the modifying agent being an organic macromolecular compound, e.g. an oligomeric, polymeric or dendrimeric molecule
- A61K47/59—Medicinal preparations characterised by the non-active ingredients used, e.g. carriers or inert additives; Targeting or modifying agents chemically bound to the active ingredient the non-active ingredient being chemically bound to the active ingredient, e.g. polymer-drug conjugates the non-active ingredient being a modifying agent the modifying agent being an organic macromolecular compound, e.g. an oligomeric, polymeric or dendrimeric molecule obtained otherwise than by reactions only involving carbon-to-carbon unsaturated bonds, e.g. polyureas or polyurethanes
- A61K47/60—Medicinal preparations characterised by the non-active ingredients used, e.g. carriers or inert additives; Targeting or modifying agents chemically bound to the active ingredient the non-active ingredient being chemically bound to the active ingredient, e.g. polymer-drug conjugates the non-active ingredient being a modifying agent the modifying agent being an organic macromolecular compound, e.g. an oligomeric, polymeric or dendrimeric molecule obtained otherwise than by reactions only involving carbon-to-carbon unsaturated bonds, e.g. polyureas or polyurethanes the organic macromolecular compound being a polyoxyalkylene oligomer, polymer or dendrimer, e.g. PEG, PPG, PEO or polyglycerol
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K47/00—Medicinal preparations characterised by the non-active ingredients used, e.g. carriers or inert additives; Targeting or modifying agents chemically bound to the active ingredient
- A61K47/50—Medicinal preparations characterised by the non-active ingredients used, e.g. carriers or inert additives; Targeting or modifying agents chemically bound to the active ingredient the non-active ingredient being chemically bound to the active ingredient, e.g. polymer-drug conjugates
- A61K47/51—Medicinal preparations characterised by the non-active ingredients used, e.g. carriers or inert additives; Targeting or modifying agents chemically bound to the active ingredient the non-active ingredient being chemically bound to the active ingredient, e.g. polymer-drug conjugates the non-active ingredient being a modifying agent
- A61K47/62—Medicinal preparations characterised by the non-active ingredients used, e.g. carriers or inert additives; Targeting or modifying agents chemically bound to the active ingredient the non-active ingredient being chemically bound to the active ingredient, e.g. polymer-drug conjugates the non-active ingredient being a modifying agent the modifying agent being a protein, peptide or polyamino acid
- A61K47/65—Peptidic linkers, binders or spacers, e.g. peptidic enzyme-labile linkers
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K38/00—Medicinal preparations containing peptides
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/50—Fusion polypeptide containing protease site
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/70—Fusion polypeptide containing domain for protein-protein interaction
Definitions
- the present disclosure generally relates to interferon alpha (IFN ⁇ ) polypeptides comprising one or more non-natural amino acid or modified amino acid substitution mutations.
- IFN ⁇ interferon alpha
- the present disclosure is also directed to IFN ⁇ conjugates comprising IFN ⁇ polypeptides site- specifically linked to at least one masking moiety, optionally via a linker.
- Type I interferons are cytokines that play an important role in modulating the innate and adaptive immune response.
- type I interferons are also now known to suppress the proliferation of cancer cells.
- the IFN-1 family comprises 17 functional genes on chromosome 9 that encode 16 proteins, IFN ⁇ , IFN ⁇ , IFN ⁇ , IFN ⁇ , and 12 subtypes of IFN ⁇ .
- IFN ⁇ is generally produced by leukocytes, while IFN ⁇ is generally produced by fibroblasts, but both IFN ⁇ / ⁇ elicit an immune response by binding to the IFN- ⁇ receptor (IFNAR).
- IFNAR IFN- ⁇ receptor
- the 12 subtypes of IFN ⁇ exhibit slightly different binding affinities and subtle differences, the first and most widely studied of the IFN ⁇ subtypes is IFN ⁇ 2.
- IFN ⁇ 2a and IFN ⁇ 2b which differ only at residue 23 (lysine or arginine, respectively).
- IFN ⁇ 2b IFN ⁇ 2b developed by Merck Sharp & Dohme was the first IFN-1 to gain approval in the United States. Intron A ® is approved as a treatment for hairy cell leukemia, malignant melanoma, follicular lymphoma, condylomata 1
- IFN ⁇ 2a Recombinant IFN ⁇ 2a was developed by Hoffmann-La Roche Inc. as Roferon-A for various types of cancer, AIDS-related sarcoma, and hepatitis.
- Various pegylated forms of IFN ⁇ 2a and IFN ⁇ 2b have also been developed, including Pegasys (Peg IFN ⁇ 2a) for chronic hepatitis B and C, Pegintron (Peg IFN ⁇ 2b) for hepatitis C, and Sylatron (Peg IFN ⁇ 2b) for melanoma.
- IFN ⁇ 2a and IFN ⁇ 2b exhibit minimal attenuation of activity and are non-tumor specific. Thus, their clinical use is limited due to poor systemic tolerability. In fact, the use of IFN ⁇ recombinant and pegylated forms can be accompanied by severe adverse side effects. Most patients experience acute toxicity in the form of flu-like symptoms, including fever, chills, myalgia, headache, and nausea. Hematological side effects, including decreases in blood counts, are also commonly observed. Hepatic, gastrointestinal, neurological, cardiovascular, and endocrine side effects have also been reported (Sleijfer, S. et al., 2005, Pharmacy World and Science, 27, pages 423–431).
- IFN ⁇ polypeptides and conjugates that are characterized by limited systemic activation and improved tumor-selectivity that provide effective and tolerable IFN ⁇ therapy for the treatment of cancer and other diseases.
- the IFN ⁇ polypeptides comprise at least one non-natural amino acid or modified amino acid. In one embodiment, the IFN ⁇ polypeptides comprise at least one non-natural amino acid. In one embodiment, the IFN ⁇ polypeptides comprise at least one modified amino acid.
- the IFN ⁇ conjugates comprise IFN ⁇ polypeptides site-specifically conjugated to at least one masking moiety, optionally via a linker. In certain embodiments, while the IFN ⁇ conjugates are in systemic circulation, the masking moiety acts as a mask to attenuate IFN ⁇ activity and/or extend the half-life of the conjugate.
- the conjugate comes in contact with the tumor microenvironment (TME)
- TME tumor microenvironment
- the conjugate is cleaved from the masking moiety either preferentially by tumor-selective proteases enriched in the TME or the acidic nature of the TME. This restores the activity of the IFN ⁇ 2
- the IFN ⁇ conjugates and polypeptides described herein exhibit reduced toxicity, for example, systemic toxicity, compared to wild- type IFN ⁇ . In other embodiments, the IFN ⁇ conjugates and polypeptides described herein exhibit increased stability, for example in serum, compared to wild-type IFN ⁇ .
- the IFN ⁇ conjugates and polypeptides described herein exhibit reduced toxicity, for example, systemic toxicity, and increased stability, for example in serum, compared to wild- type IFN ⁇ .
- the disclosure provides IFN ⁇ polypeptides capable of binding the IFN ⁇ receptor ⁇ IFNAR).
- the IFN ⁇ polypeptides comprise at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
- the IFN ⁇ polypeptides comprise at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156.
- the non-natural amino acid or modified amino acid is a non-natural amino acid. In one embodiment, the non-natural amino acid or modified amino acid is a modified amino acid. In certain embodiments, the positions are in reference to human wild- type IFN ⁇ (SEQ ID NO: 33). In certain embodiments, the positions are in reference to mice wild-type IFN ⁇ (SEQ ID NO: 38). In certain embodiments, the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 33. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 33. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 33.
- the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 38. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 38. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 38. [00010]
- IFN ⁇ conjugates comprising an IFN ⁇ polypeptide and a masking moiety wherein the IFN ⁇ polypeptide is site-specifically linked to the masking moiety, optionally via a linker. In certain embodiments, the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide comprising at least one non-natural amino acid or modified 3
- amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156 wherein the at least one non-natural amino acid or modified amino acid is linked to a masking moiety optionally via a linker.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide comprising at least one non- natural amino acid or modified amino acid at a position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 wherein the at least one non-natural amino acid or modified amino acid is linked to a masking moiety optionally via a linker.
- the non- natural amino acid or modified amino acid is a non-natural amino acid.
- the non-natural amino acid or modified amino acid is a modified amino acid.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide and at least one masking moiety wherein the IFN ⁇ polypeptide is site-specifically linked to the at least one masking moiety via a protease cleavable linker and the water masking moiety is a water-soluble polymer or carbohydrate.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide and at least one masking moiety wherein the IFN ⁇ polypeptide is site-specifically linked to the at least one masking moiety via a cathepsin B cleavable linker.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide and at least one masking moiety wherein the IFN ⁇ polypeptide is site-specifically linked to the at least one masking moiety via a pH- sensitive linker.
- the IFN ⁇ conjugate comprises at least two masking moieties. In certain embodiments, the IFN ⁇ conjugate comprises at least three masking moieties. In certain embodiments, the IFN ⁇ conjugate comprises at least four masking moieties. In certain embodiments, the IFN ⁇ conjugate comprises at least five masking moieties. In certain embodiments, the IFN ⁇ conjugate comprises at least six or more masking moieties.
- the masking moiety is attached the IFN ⁇ polypeptide via a linker as described herein.
- polynucleotides encoding the IFN ⁇ polypeptides and/or fusion constructs described herein.
- expression vectors comprising the polynucleotides.
- cells comprising the polynucleotides or expression vectors.
- the cells are selected from bacterial cells, fungal cells, and mammalian cells.
- the cells are selected from E. coli cells, Saccharomyces cerevisiae cells, and CHO cells.
- methods of making the IFN ⁇ polypeptides for instance using the polynucleotides, expression vectors, and/or cells described herein. 4
- the method includes administering to the subject an effective amount of the IFN ⁇ polypeptide, IFN ⁇ conjugate, or fusion construct of any of the foregoing embodiments, or a composition or a pharmaceutical composition containing the same.
- the disease or condition is selected from a cancer, an autoimmune disease, an inflammatory disease, and an infection.
- the effective amount is a therapeutically effective amount.
- the disease or condition is cancer.
- Anti-tumor immune memory is the immune system’s ability to recognize, or memorize, a previously encountered tumor antigen.
- a IFN ⁇ polypeptide, IFN ⁇ conjugate, or fusion construct described herein or a pharmaceutical composition thereof treats the disease or condition, for example, cancer, by activating anti-tumor immunity.
- a IFN ⁇ polypeptide, IFN ⁇ conjugate, or fusion construct described herein or a pharmaceutical composition thereof treats the disease or condition, for example, cancer, by inducing or enhancing anti-tumor immune memory.
- Embodiments disclosed herein are also directed to the use of the IFN ⁇ polypeptide, IFN ⁇ conjugates, or fusion constructs of any of the foregoing embodiments for treating, preventing, or diagnosing a disease or condition in a subject in need thereof.
- Embodiments disclosed herein are also directed to the IFN ⁇ polypeptides, IFN ⁇ conjugates, or fusion constructs of any of the foregoing embodiments for use in treating, preventing, or diagnosing a disease or condition in a subject in need thereof.
- Embodiments disclosed herein are also directed to the IFN ⁇ polypeptides, IFN ⁇ conjugates, or fusion constructs of any of the foregoing embodiments for use in the manufacture of a medicament for treating, preventing, or diagnosing a disease or condition in a subject in need thereof.
- the disease or condition is selected from a cancer, an autoimmune disease, an inflammatory disease, and an infection.
- FIG.1A is a graph of the effect of a 3 mg/kg dose of Conjugate 1, Conjugate 16 and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 41 post treatment.
- FIG.1B is a graph of the effect of a 10 mg/kg dose of Conjugate 1, Conjugate 16 and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 41 post treatment.
- FIG.1C is a graph of the effect of a 3 mg/kg dose of Conjugate 1, Conjugate 16 and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 44 post treatment.
- FIG.1D is a graph of the effect of a 15 mg/kg dose of Conjugate 1, Conjugate 16 and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 44 post treatment.
- FIG.1E is a graph of the effect of Conjugate 1, Conjugate 16 and Conjugate 18 (dosed at 3 mg/kg and 15 mg/kg) on MDA-MB-231 tumor size up until the end of the study at day 44 post treatment.
- FIG.2A is a graph of the effect of a 3 mg/kg dose of Conjugate 1, Conjugate 2, Conjugate 13, and Conjugate 18 on MDA-MB-231 tumor growth up.
- FIG.2B is a graph of the effect of a 1 mg/kg dose of Conjugate 1, Conjugate 2, Conjugate 13, and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 32 post treatment.
- FIG.2C is a graph of the effect of a 0.1 mg/kg dose of Conjugate 1, Conjugate 2, Conjugate 13, and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 32 post treatment.
- FIG.2D is a graph of the effect of a 0.05 mg/kg dose of Conjugate 1, Conjugate 2, Conjugate 13, and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 32 post treatment.
- FIG.2E is a graph of the effect of Conjugate 1, Conjugate 2, Conjugate 13, and Conjugate 18 (dosed at 0.05 mg/kg, 0.01 mg/kg, 1 mg/kg and 3 mg/kg) on MDA-MB-231 tumor size up until the end of the study at day 44 post treatment as described in Example 9.
- FIG.3A is a graph illustrating the serum concentrations of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 following intravenous administration at doses ranging from 3 mg/kg to 45 mg/kg. 6
- FIG. 3B is a graph showing the PK profile of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 following an intravenous dose (15 mg/kg).
- FIG.3C is a graph showing percent body weight changes in hamsters following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg.
- FIG.3D is a graph showing percent body weight changes in hamsters following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at a dose of 15 mg/kg.
- FIG.3E is a graph illustrating ALT fold change as measured on day 2 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg.
- FIG.3F is a graph illustrating AST fold change as measured on day 2 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg.
- FIG.3G is a graph illustrating ALP fold change as measured on day 2 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg.
- FIG.3H is a graph illustrating ALT fold change as measured on day 7 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg.
- FIG.3I is a graph illustrating AST fold change as measured on day 7 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg.
- FIG.3J is a graph illustrating ALP fold change as measured on day 7 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg.
- FIG.3K and FIG.3L are graphs illustrating tolerability of HLE-Interferon variants in golden Syrian hamsters.
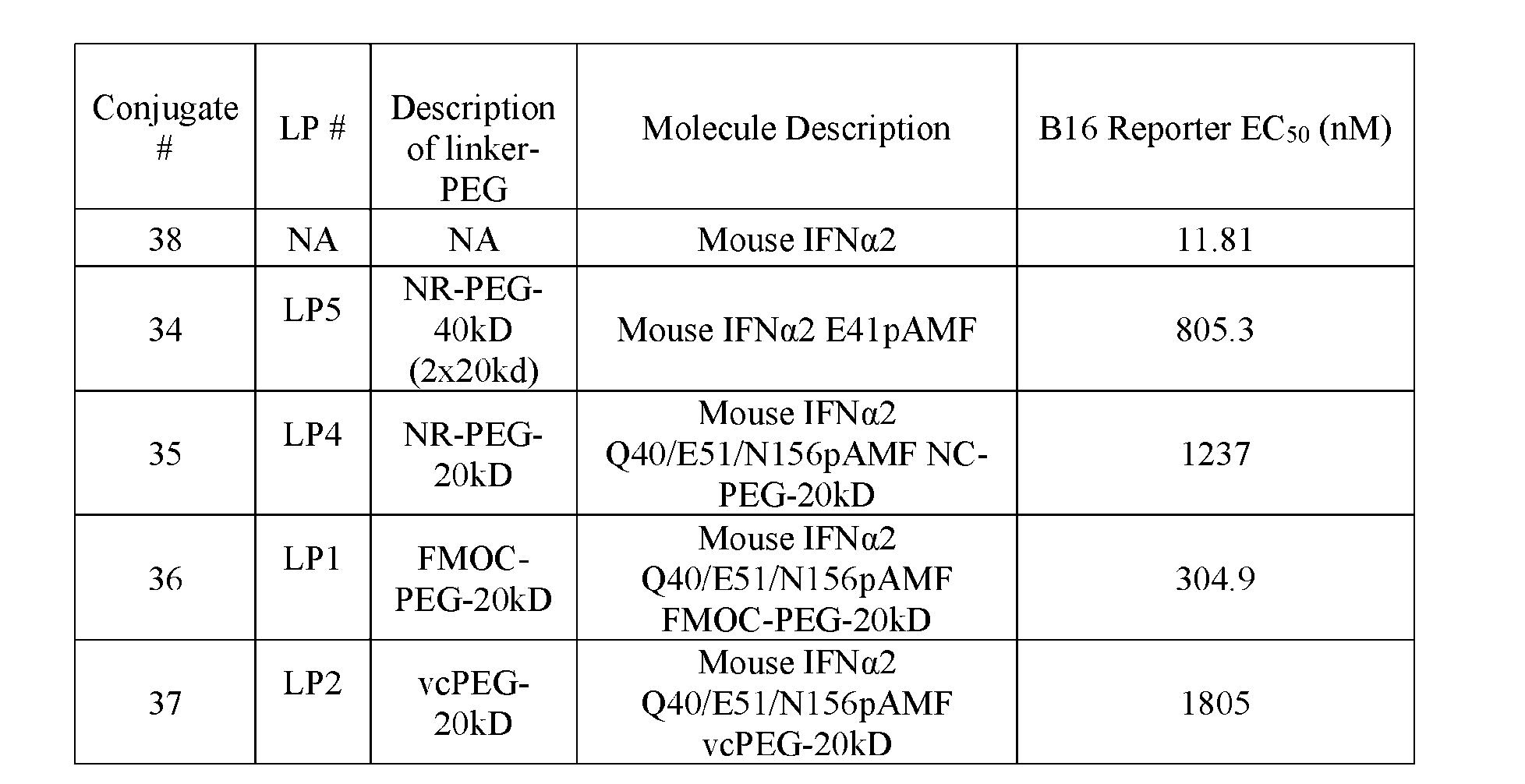
- FIG.4A is a graph of the effect of a 3 mg/kg dose of Conjugate 34, Conjugate 35, Conjugate 36, and Conjugate 37 on B16F10 tumor growth.
- FIG.4B is a graph of the effect of a 1 mg/kg dose of Conjugate 34 and Conjugate 36 on B16F10 tumor growth.
- FIG.4C is a graph of the effect of Conjugate 35 and Conjugate 37 (dosed with 3 mg/kg and 10 mg/kg) on B16F10 tumor growth. 7
- FIG. 4D is a graph of the effect of Conjugate 35 and Conjugate 37 (dosed at 3 mg/kg and 10 mg/kg) on B16F10 tumor size.
- FIG.4E is a graph of the effect of Conjugate 37 administered at 3 mg/kg compared to the vehicle on the production of CD45 positive cells.
- FIG.4F is a graph of the effect of Conjugate 37 administered at 3 mg/kg compared to the vehicle on the level of Granzyme B secreted from CD8 T-cells.
- FIG.4G is a graph of the effect of Conjugate 37 administered at 3 mg/kg compared to the vehicle on the level of Granzyme B secreted from NK cells.
- FIG. 4G is a graph of the effect of Conjugate 37 administered at 3 mg/kg compared to the vehicle on the level of Granzyme B secreted from NK cells.
- FIG. 4H and FIG. 4I are graphs showing mouse surrogate efficacy and PD in MC38-HCEA.
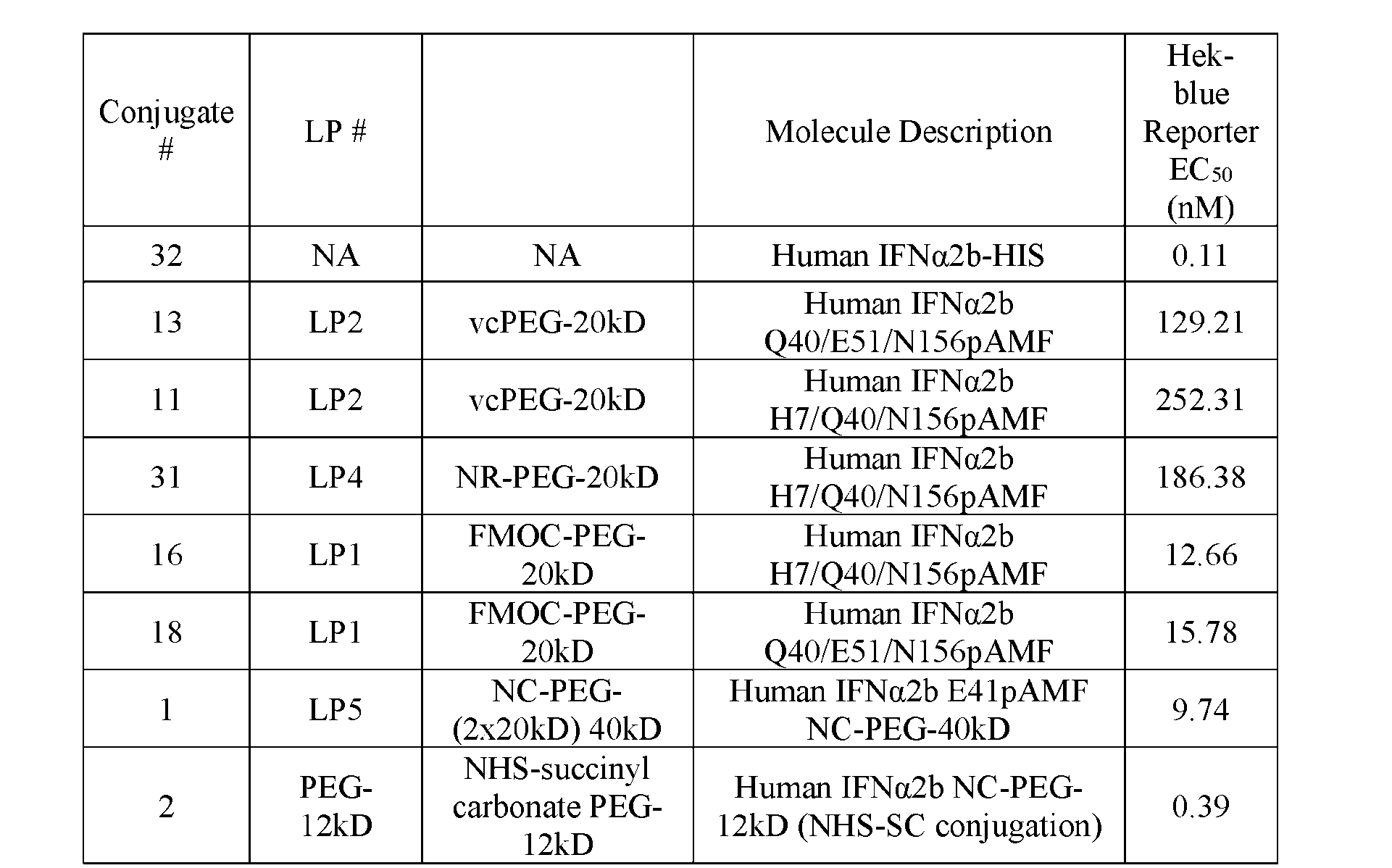
- FIG.5A is a graph of dose response curves for human IFN ⁇ Conjugate 1, Conjugate 2, Conjugate 11, Conjugate 13, Conjugate 16, Conjugate 18, and Conjugate 31 compared to wildtype human IFN ⁇ (Conjugate 32) in HEK-blue human IFN ⁇ / ⁇ Reporter Assay.
- FIG. 5B is a graph of dose response curves for mouse IFN ⁇ Conjugate 34, Conjugate 35, Conjugate 36, and Conjugate 37 compared to wildtype mouse IFN ⁇ (Conjugate 38) in B16-Blue mouse IFN ⁇ / ⁇ Reporter Assay.
- FIG.6 is a graph showing the PK of IFN ⁇ Conjugate 1, Conjugate 13, Conjugate 18, and Conjugate 33, in C57B1/6 mice.
- FIG. 7A, FIG. 7B, and FIG. 7C show the combination of Conjugate 37 with a checkpoint inhibitor in B16F10 synergist mouse tumor model.
- FIG.8 is an image showing how the IFN ⁇ conjugates of the present description are masked in circulation, but unmasked in the tumor microenvironment (TME).
- FIG. 9A shows the SDS-PAGE gel showing expression and purification of 3XnnAA-IFN ⁇ .
- FIG. 9B is the analytical SEC chromatogram showing purity of final PEGylated 3XnnAA-IFN ⁇ .
- FIG. 9C is a Hek-Blue assay of in vitro cell activation with IFN ⁇ samples.
- FIG.9D is a table summarizing strains tested, fermentation format and scales, titers, and product quality metrics for 3XnnAA-IFN ⁇ proteins. Strains with a product plasmid bearing an extra copy of AS tRNA are denoted as “+ AS tRNA” while strains bearing a product plasmid with only the 3XnnAA-IFN ⁇ coding sequence are denoted as “No AS tRNA”. The % full length 3XnnAA-IFN ⁇ was calculated by intact LC-MS analysis, purity % was assessed by analytical SEC, and PEG-to-protein ratio was calculated using SDS-PAGE gel densitometry. [00054] FIG. 10A, FIG. 10B, and FIG.
- FIG. 10C show expression plasmids and shake flask analysis of 3x-nnAA-IFN ⁇ .
- the high copy product plasmid used for the initial expression of 3X-nnAA-IFN ⁇ in shake flasks is shown in FIG.10A.
- the high copy product plasmid (+ AS tRNA) used to increase tRNA concentration/amber suppression for the expression of 3X- nnAA-IFN ⁇ . This is the final plasmid used in high density fermentations is shown in FIG.10B.
- the SDS-PAGE gel analysis of HisSUMO-IFN ⁇ and 3X-nnAA-IFN ⁇ in shake flasks is shown in FIG.10C.
- the arrow denotes the presence of the SUMO-tagged IFN ⁇ band.
- FIG. 11C show intact LC-MS analysis of 3X-nnAA- IFN ⁇ proteins expressed with either product plasmid and include the deconvoluted LC-MS spectrum of purified and Ulp1-cleaved IFN ⁇ that has been conjugated with a small molecule DBCO-amine.
- FIG. 11A shows 3X-nnAA-IFN ⁇ produced in shake flasks with a product plasmid bearing an extra copy of the AS tRNA (product plasmid + AS tRNA, see FIG.10B).
- FIG.11B shows 3X-nnAA-IFN ⁇ produced in shake flasks with a product plasmid with no extra copy of the AS tRNA (product plasmid, see FIG.10A).
- FIG.11C shows 3X-nnAA-IFN ⁇ produced in high density fermentation with a product plasmid bearing an extra copy of the AS tRNA (product plasmid + AS tRNA, see FIG.10B). Individual peaks in each LC-MS spectrum are labeled.
- FIG. 12A is a graph showing increased Granzyme B levels in tumor infiltrating CD8 T-cells harvested from MC38-hTrop2 tumor bearing mice following treatment with a single IV dose of 1 mg/kg of Conjugate 37 or Conjugate 34 compared to the vehicle.
- FIG.12B is a graph showing increased Granzyme B levels in tumor infiltrating NK cells harvested from MC38-hTrop2 tumor bearing mice following treatment with a single IV dose of 1 mg/kg of Conjugate 37 or Conjugate 34 compared to the vehicle. 9
- FIG. 12C is a graph showing increased activation of monocytes following treatment with a single IV dose of 1 mg/kg of Conjugate 37 or Conjugate 34 compared to the vehicle.
- FIG. 12D is a graph showing increased activation of dendritic cells following treatment with a single IV dose of 1 mg/kg of Conjugate 37 or Conjugate 34 compared to the vehicle.
- FIG.12E is a graph showing increased activation of plasmacytoid dendritic cells following treatment with a single IV dose of 1 mg/kg of Conjugate 37 or Conjugate 34 compared to the vehicle.
- FIG.12F is a lymph node analysis showing that treatment with Conjugate 34 results in similar increase in levels of GranzymeB in CD8 T-cells from both tumor-draining and non- draining lymph node, while treatment with Conjugate 37 results in a greater increase in GranzymeB levels in CD8 T-cells from tumor-draining lymph node compared to non-draining lymph nodes.
- FIG. 13 is a showing the tumor size of complete responder mice obtained from Conjugate 37 treatment that were treated with 300 ⁇ g anti-CD8 antibody or Isotype antibody and rechallenged with 5x10 6 MC38-hCEA cells on D0.
- IFN ⁇ polypeptides When rechallenged, Conjugate 37- treated complete responder mice that received Isotype control antibody demonstrated no recurrence of tumors, while CD8 depletion ablated Conjugate 37-induced anti-tumor immune memory.
- the IFN ⁇ polypeptides comprise at least one non-natural amino acid or modified amino acid substitution.
- the IFN ⁇ conjugates comprise an IFN ⁇ polypeptide site-specifically linked to a masking moiety, optionally via a linker. As disclosed herein, the site-specific linkage to the masking moiety provides an extended half-life of the conjugates.
- the masking moiety acts as a mask to attenuate IFN ⁇ activity while the IFN ⁇ conjugate is in systemic circulation but once the masked-IFN ⁇ conjugate comes in contact with the tumor microenvironment (TME), tumor-selective proteases or the acidic environment of the TME cleave the masking moiety to activate the IFN ⁇ polypeptide for immune cell activation and tumor cell killing. Therefore, the IFN ⁇ conjugates bonded to a masking moiety using site-specific conjugation are advantageous for the delivery of IFN ⁇ because the 10
- polypeptides exhibit less systemic toxicity, while simultaneously exhibiting enhanced and selective tumor cell killing. In certain embodiments, these IFN ⁇ polypeptides exhibit improved characteristics, for example reduced toxicity and increased stability, relative to a wild type (i.e., parent) IFN ⁇ .
- methods of treating, preventing, or diagnosing a disease or condition, for example, cancer, in a subject in need thereof comprising administering to the subject an effective amount of an IFN ⁇ polypeptide or IFN ⁇ conjugate described herein or a pharmaceutical composition thereof.
- a IFN ⁇ polypeptide or a IFN ⁇ conjugate described herein or a pharmaceutical composition thereof treats the disease or condition, for example, cancer, by activating anti-tumor immunity.
- IFN ⁇ conjugates described herein activated the innate and adaptive immune system when administered to tumor-bearing mice; analysis of the TME cell types showed that administration of the IFN ⁇ conjugates led to increased levels of Granzyme B levels in tumor infiltrating CD8 T-cells and NK cells and increased activation of multiple innate immune cells in the TME including monocytes, dendritic cells and plasmacytoid dendritic cells.
- a IFN ⁇ polypeptide or a IFN ⁇ conjugate described herein or a pharmaceutical composition thereof treats the disease or conditions, for example, cancer, by inducing or enhancing anti-tumor immune memory via T cell activation and proliferation.
- kits and reagents are generally carried out in accordance with manufacturer-defined protocols and conditions unless otherwise noted.
- the singular forms “a,” “an,” and “the,” include the plural referents unless the context clearly indicates otherwise.
- the term “about,” as used herein, indicates and encompasses an indicated value and a range above and below that value. In certain embodiments, the term “about” indicates the designated value ⁇ 10%, ⁇ 5%, or ⁇ 1%. In certain embodiments, the term “about” indicates the designated value ⁇ one standard deviation of that value.
- the term “combinations thereof,” as used herein, includes every possible combination of elements to which the term refers to.
- Ranges throughout this disclosure, various aspects of the disclosure can be presented in a range format. It should be understood that the description in range format is merely for convenience and brevity and should not be construed as an inflexible limitation on the scope of the disclosure. Accordingly, the description of a range should be considered to have specifically disclosed all the possible subranges as well as individual numerical values within that range. For example, description of a range such as from 1 to 6 should be considered to have specifically disclosed subranges such as from 1 to 3, from 1 to 4, from 1 to 5, from 2 to 4, from 2 to 6, from 3 to 6 etc., as well as individual numbers within that range, for example, 1, 2, 2.7, 3, 4, 5, 5.3, and 6. This applies regardless of the breadth of the range.
- IFN ⁇ is a key part of the innate immune response with potent antiviral, antiproliferative and immunomodulatory properties. IFN ⁇ , like other type I interferons, binds a plasma membrane receptor made of IFNAR1 and IFNAR2 that is ubiquitously expressed in the human body. In humans, IFN ⁇ gene is located on chromosome 9. There are 12 subtypes of IFN ⁇ .
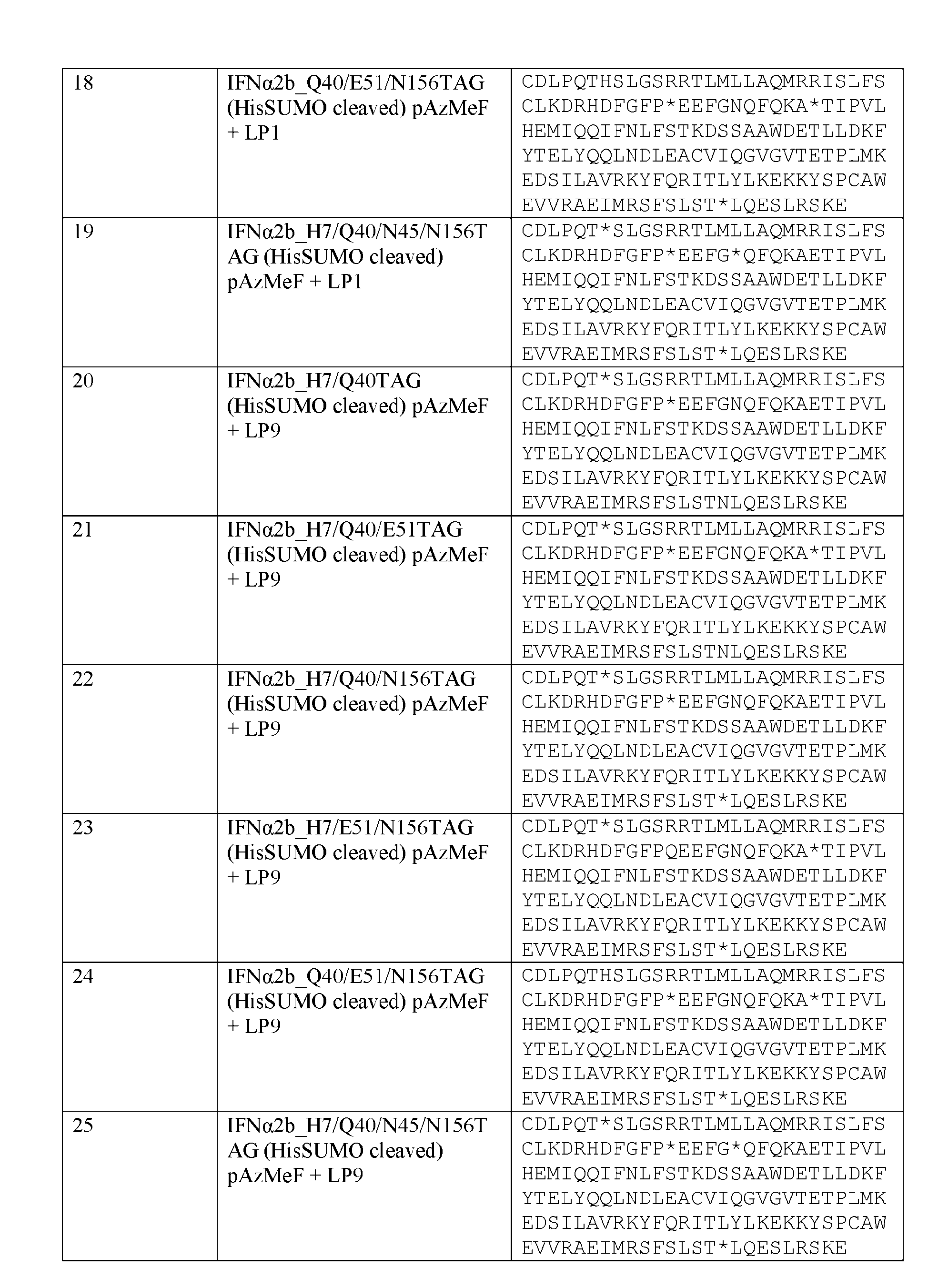
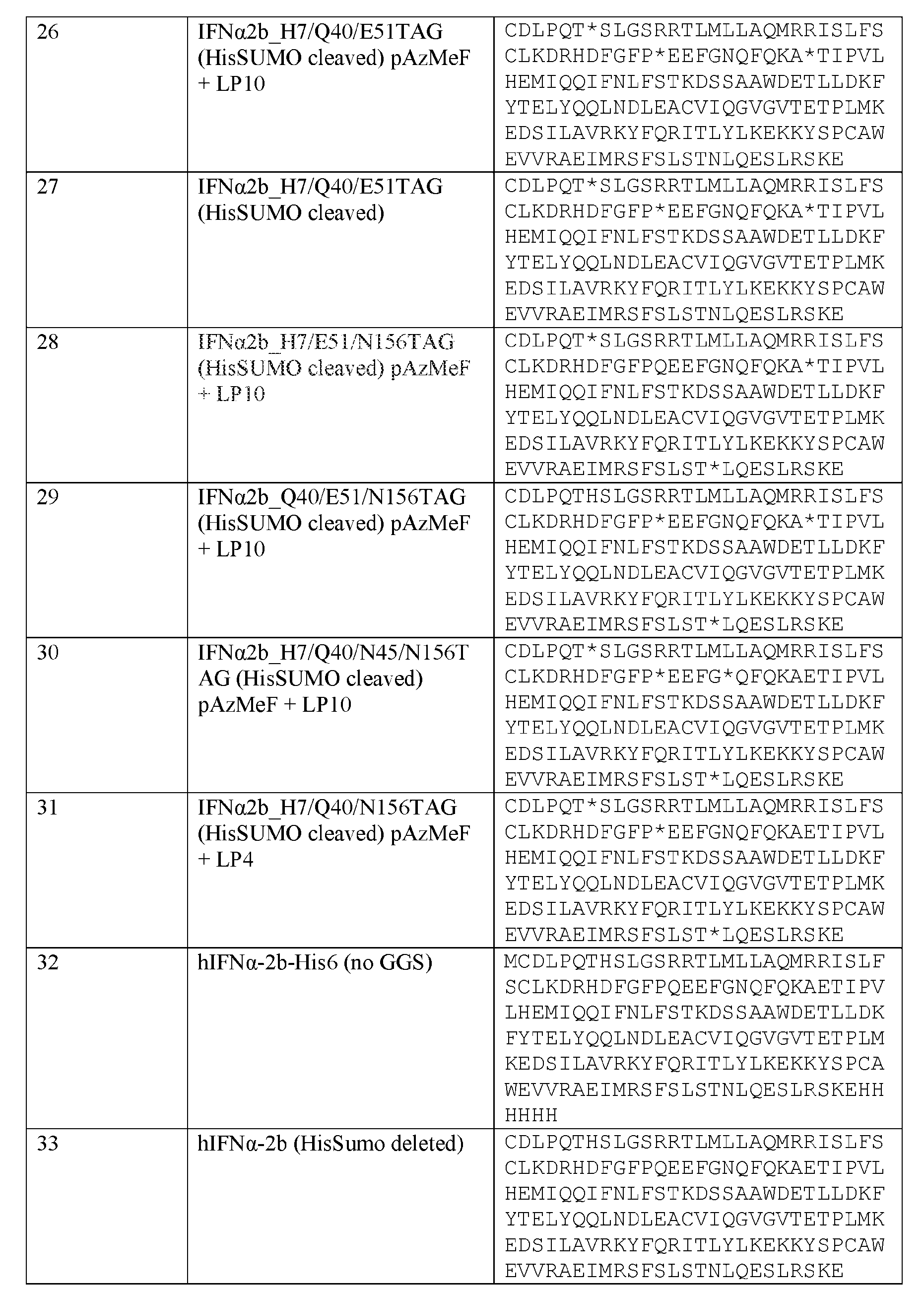
- IFN ⁇ 2 sequence is provided by SEQ ID NO: 33: CDLPQTHSLG SRRTLMLLAQMRRISLFSCLKDRHDFGFPQEEFGNQFQKA ETIPVLHEMI QQIFNLFSTKDSSAAWDETLLDKFYTELYQQLNDLEACVI QGVGVTETPL MKEDSILAVRKYFQRITLYLKEKKYSPCAWEVVRAEIMRS FSLSTNLQES LRSKE.
- the IFNAR is composed of two subunits: IFNAR1 and IFNAR2. Binding to either the IFNAR1 or IFNAR2 subunit by an IFN-1, including IFN ⁇ , brings the two subunits close 12
- human interferon alpha/beta receptor 1 or “human IFN-R-1” as used herein, refers to the IFNAR1 subunit of the receptor for IFN ⁇ encoded by the IFNAR1 gene. Representative IFN-R-1 sequences are provided by UniProt. Accession No. P17181.
- human interferon alpha/beta receptor 2 or “human IFN-R-2” as used herein, refers to the IFNAR2 subunit of the receptor for IFN ⁇ encoded by the IFNAR2 gene. Representative IFN-R-2 sequences are provided by UniProt.
- operable-linked refers to a functional linkage between two elements, regardless of orientation or distance between the two elements, such that the function of one element is controlled or affected by the other element.
- operable linkage with reference to a promoter and heterologous coding sequence means that the transcription of the heterologous coding sequence is under the control of, or driven by, the promoter.
- operable linkage with reference to an enhancer and promoter means that the enhancer increases the level of transcription driven by a promoter.
- isolated refers to a substance that has been separated and/or recovered from its natural environment.
- a naturally occurring polynucleotide or polypeptide present in a living animal is not isolated, but the same polynucleotide or polypeptide, separated from some or all of the co-existing materials in the natural system, is isolated.
- Such polynucleotide could be part of a vector and/or such polynucleotide or polypeptide could be part of a composition (e.g., a cell lysate), and still be isolated in that such vector or composition is not part of the natural environment for the nucleic acid or polypeptide.
- An “isolated IFN ⁇ variant” or “isolated IFN ⁇ polypeptide” is one that has been separated and/or recovered from a component of its natural environment.
- Components of the natural environment may include enzymes, hormones, and other proteinaceous or nonproteinaceous materials.
- an isolated IFN ⁇ polypeptide is purified to a degree sufficient to obtain at least 15 residues of N-terminal or internal amino acid sequence, for example by use of a spinning cup sequenator.
- an isolated IFN ⁇ polypeptide is purified to homogeneity by gel electrophoresis (e.g., SDS-PAGE) under reducing or nonreducing conditions, with detection by Coomassie blue or silver stain.
- an isolated IFN ⁇ polypeptide is prepared by at least one purification step.
- the term “substantially pure” with respect to a composition comprising a polypeptide IFN ⁇ refers to a composition that includes at least 80%, 85%, 90% or 95% by 13
- an isolated IFN ⁇ polypeptide is purified to at least 80%, 85%, 90%, 95%, or 99% by weight. In some embodiments, an isolated IFN ⁇ polypeptide is purified to at least 80%, 85%, 90%, 95%, or 99% by volume.
- an isolated IFN ⁇ polypeptide is provided as a solution comprising at least 85%, 90%, 95%, 98%, 99% to 100% by weight. In some embodiments, an isolated IFN ⁇ polypeptide is provided as a solution comprising at least 85%, 90%, 95%, 98%, 99% to 100% by volume.
- Affinity refers to the strength of the sum total of non-covalent interactions between a single binding site of a molecule (e.g., IFN ⁇ ) and its binding partner (e.g., IFNAR).
- binding affinity refers to intrinsic binding affinity, which reflects a 1:1 interaction between members of a binding pair (e.g., IFN ⁇ and IFNAR, subunit IFNAR1 or IFNAR2).
- the affinity of a molecule X for its partner Y can be represented by the dissociation constant (K D ).
- Affinity can be measured by common methods known in the art, including those described herein. Affinity can be determined, for example, using surface plasmon resonance (SPR) technology, such as a Biacore ® instrument. In some embodiments, the affinity is determined at 25°C.
- the terms “specific,” “specific binding,” “specifically binds to,” “specific for,” “selectively binds,” or “selective for,” as used herein, refers to a particular receptor or ligand that exhibits binding that is measurably different from a non-specific or non-selective interaction.
- Specific binding can be measured, for example, by determining binding of a molecule compared to binding of a control molecule.
- Specific binding can also be determined by competition with a control molecule that competes with the ligand for binding to the receptor. In that case, specific binding is indicated if the binding of the ligand to the receptor is competitively inhibited by the control molecule.
- k d (sec -1 ), as used herein, refers to the dissociation rate constant of a particular receptor-ligand interaction. This value is also referred to as the koff value.
- k a (M -1 ⁇ sec -1 ), as used herein, refers to the association rate constant of a particular receptor-ligand interaction. This value is also referred to as the kon value. 14
- K D K D
- M dissociation equilibrium constant of a particular receptor-ligand interaction.
- K D k d /k a .
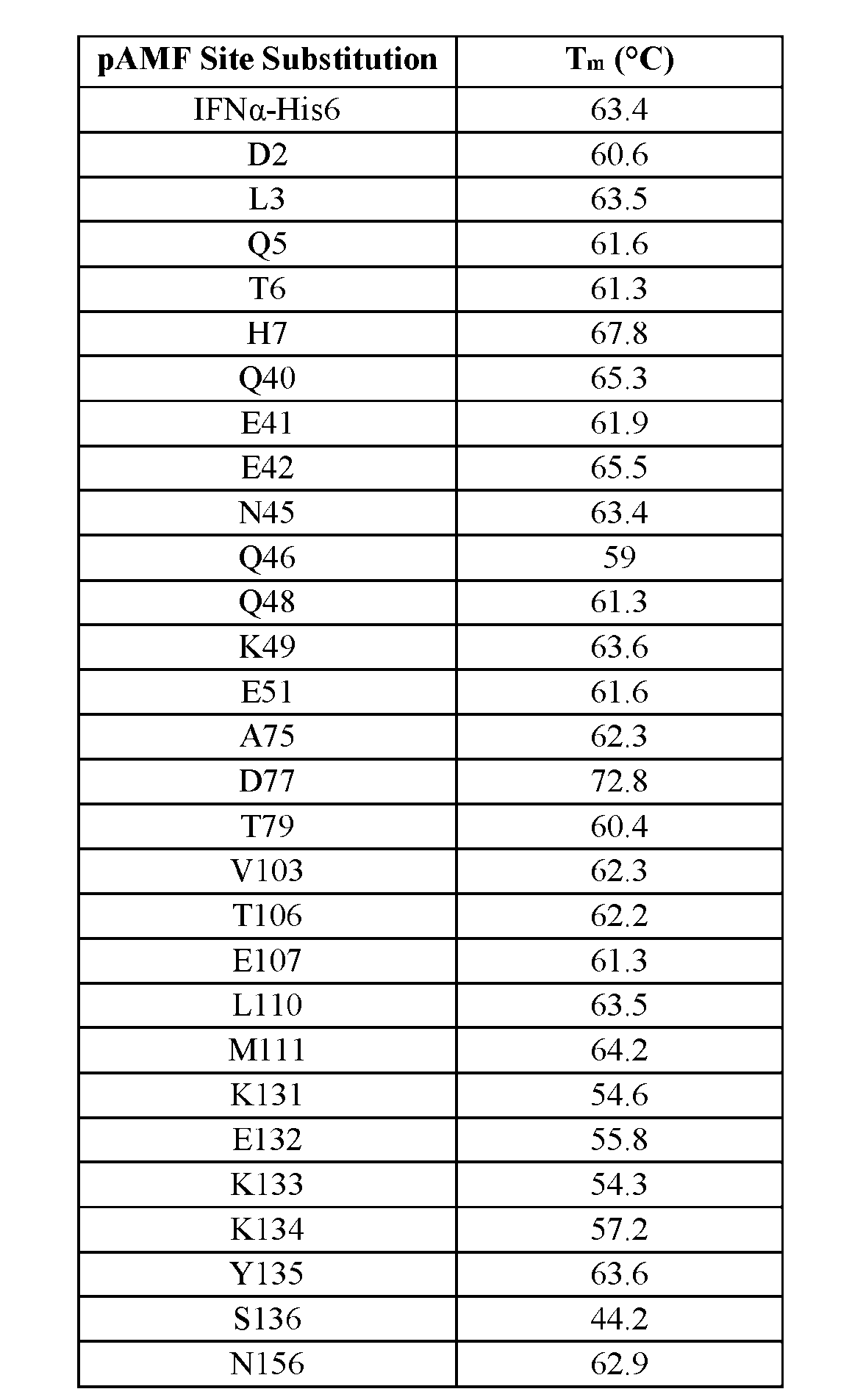
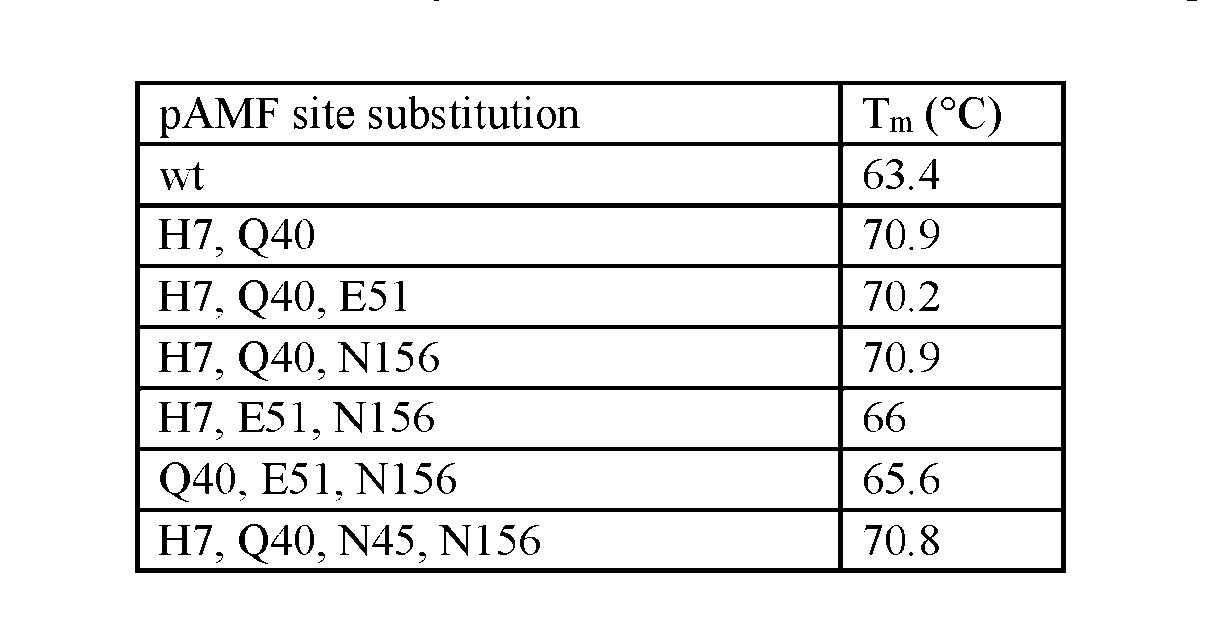
- Tm Tm as used herein, has the meaning commonly understood in the art and refers to is the temperature at which the equilibrium between folded and unfolded forms of the enzyme is at its mid-point.
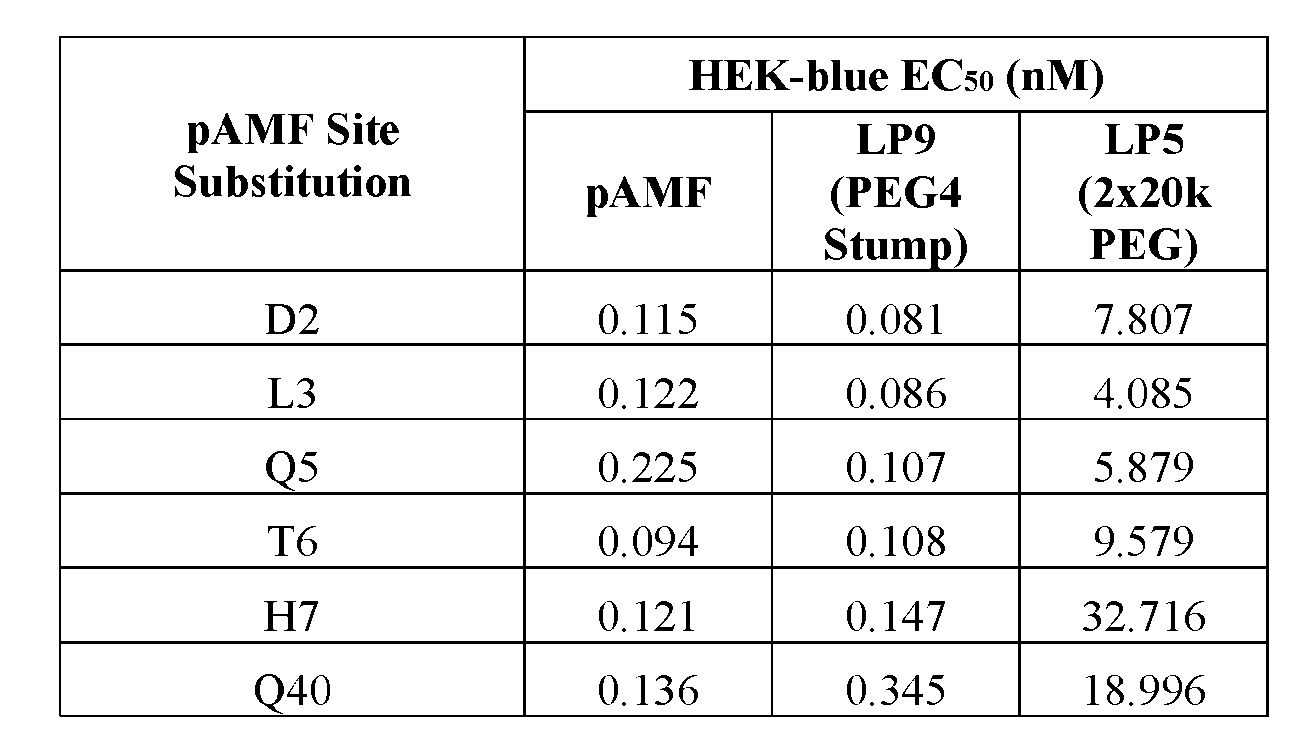
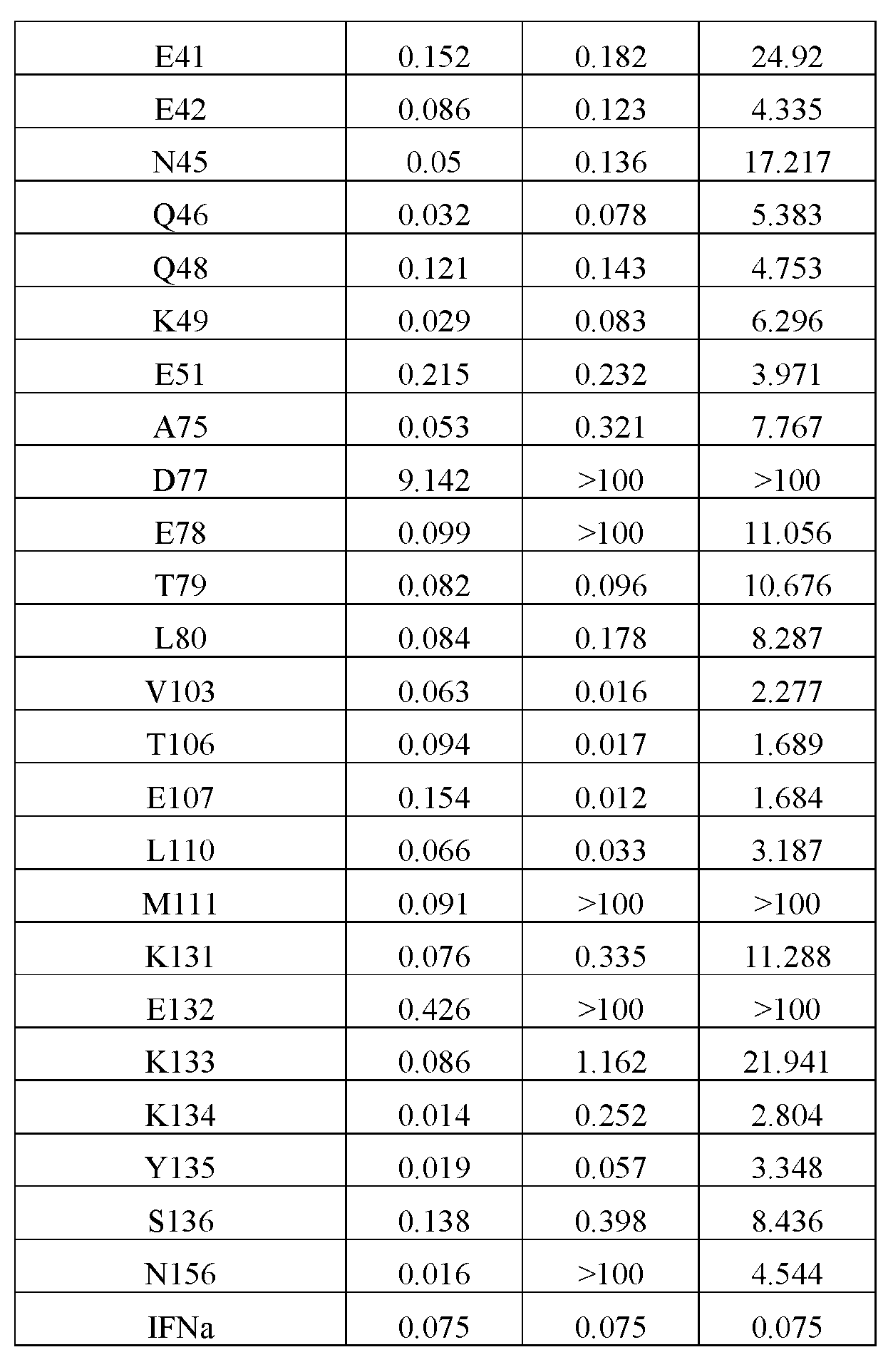
- EC 50 half maximal effective concentration
- EC50 has the meaning commonly understood in the art and refers to the concentration of a substance e.g., a drug, e.g., an IFN ⁇ polypeptide, which induces a response halfway between the baseline and maximum after a specified exposure time.
- EC50 can be defined as the concentration required to obtain a 50% effect and represents the concentration of a compound where 50% of its maximal effect is observed.
- half-life” or t1/2 refers to the amount of time required for the drug concentration measured in a sample to be reduced to half of its starting concentration or amount.
- terminal t1/2 refers to the amount of time required for the drug concentration measured in a sample to be reduced to half of its pseudo- equilibrium concentration or amount.
- C 0 has the meaning commonly understood in the art and refers to the plasma concentration at the time of dosing (time 0).
- AUC as used herein, has the meaning commonly understood in the art of pharmacokinetics, and refers to the area under the plasma drug concentration-time curve (AUC) and reflects the measure of how much drug reaches an individual’s bloodstream in a given period of time after a dose is given. AUC is dependent on the rate of elimination of the drug from the body and the dose administered.
- AUC is directly proportional to the dose when the drug follows linear kinetics and is inversely proportional to the clearance of the drug.
- AUC 0-last has the meaning commonly understood in the art of pharmacokinetics, and refers to the AUC from dosing (time 0) to the last measurable concentration.
- AUC 0-inf has the meaning commonly understood in the art of pharmacokinetics, and refers to the AUC from dosing (time 0) extrapolated to infinity.
- steady state as used herein has the meaning commonly understood in the art of pharmacokinetics, and refers to the condition when the administration of a drug and the clearance are balanced, creating a plasma concentration that is unchanged by time.
- protein refers to a polymer of amino acid residues linked together by peptide (amide) bonds.
- amide peptide bonds
- a peptide will be at least three amino acids long and equal to or less than about 10 amino acids in length.
- a polypeptide is typically greater than 10 amino acids in length.
- a protein, peptide, or polypeptide may refer to an individual protein or a collection of proteins.
- One or more of the amino acids in a protein, peptide, or polypeptide may be modified, for example, by the addition of a chemical entity such as a carbohydrate group, a hydroxyl group, a phosphate group, a farnesyl group, an isofarnesyl group, a fatty acid group, a linker for conjugation, functionalization, or other modification, etc., or may be substituted with a non- natural amino acid.
- a protein, peptide, or polypeptide may also be a single molecule or may be a multi-molecular complex.
- a protein, peptide, or polypeptide may be just a fragment of a naturally occurring protein or peptide.
- a protein, peptide, or polypeptide may be naturally occurring, recombinant, or synthetic, or any combination thereof.
- a protein may comprise different domains, for example, a protein binding domain and a catalytic domain.
- Any of the proteins provided herein may be produced by any method known in the art.
- the proteins provided herein may be produced via recombinant protein expression and purification. Methods for recombinant protein expression and purification are well known, and include those described by Green and Sambrook, Molecular Cloning: A Laboratory Manual (4 th ed., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. (2012)), the entire contents of which are incorporated herein by reference.
- a “mutation” as used herein refers to a change in nucleic acid or polypeptide sequence relative to a reference sequence.
- a mutation may comprise a substitution of a residue within a sequence, e.g., a nucleic acid or amino acid sequence, with another residue, or a 16
- a mutation can be a “substitution” mutation wherein the amino acid, or nucleotide at a particular position in a reference sequence is substituted with a different amino acid or nucleotide at that position in the amino acid or nucleic acid sequence.
- a substitution replaces one amino acid at a specific location in a polypeptide or protein sequence for a different amino acid in that position of the polypeptide or protein sequence.
- a “substitution” replaces a natural amino acid at a specific location in a polypeptide or protein sequence for a non-natural amino acid or modified amino acid in that position of the polypeptide or protein sequence.
- substitution refers to as “substitution” mutation as disclosed herein above.
- reversion mutation refers to a particular type of substitution mutation wherein a polypeptide or nucleic acid sequence having a substitution mutation at a specific position in the sequence, acquires a mutation at that specific position that restores the original sequence.
- a polypeptide sequence having a mutation at a specific position in the polypeptide sequence acquires a mutation that restores the amino acid at that specific position to the amino acid found in the reference sequence e.g., restores the wild-type sequence).
- wild-type or “parent” refers to a naturally occurring gene or protein. These include a naturally occurring IFN ⁇ gene or protein.
- variant refers to a nucleic acid or polypeptide sequence having at least one mutation relative to a reference sequence. Accordingly, a “variant” or “mutant” typically has less than 100% sequence identity to a reference sequence.
- identity in the context of two or more polypeptide or nucleic acid molecule sequences, means two or more sequences or subsequences that are the same or have a specified percentage of amino acid residues or nucleotides that are the same over a specified region (e.g., 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% identity), when compared and aligned for maximum correspondence over a comparison window, or designated region, as measured using 17
- a sequence comparison algorithm by manual alignment, or by visual inspection. Alignment for purposes of determining percent amino acid sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST (Altschul et al. Nucleic Acids Res.2007, 25, 3389-3402), BLAST-2, ALIGN, MEGALIGN (DNASTAR), CLUSTALW, CLUSTAL OMEGA, or MUSCLE software. Those skilled in the art can determine appropriate parameters for aligning sequences, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared.
- sequence analysis software is used for analysis, the results of the analysis are based on the “default values” of the program referenced. “Default values” mean any set of values or parameters which originally load with the software when first initialized.
- Default values mean any set of values or parameters which originally load with the software when first initialized.
- the term “substantial similarity” or “substantially similar” means that two peptide sequences, when optimally aligned, such as by the programs GAP or BESTFIT using default gap weights, share at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.5%, or 100% sequence identity.
- amino acid refers to a D- or L-natural or non-naturally occurring amino acid, including, but not limited to, the twenty common naturally occurring amino acids.
- Naturally occurring amino acids include alanine (Ala; A), arginine (Arg; R), asparagine (Asn; N), aspartic acid (Asp; D), cysteine (Cys; C); glutamic acid (Glu; E), glutamine (Gln; Q), Glycine (Gly; G); histidine (His; H), isoleucine (Ile; I), leucine (Leu; L), lysine (Lys; K), methionine (Met; M), phenylalanine (Phe; F), proline (Pro; P), serine (Ser; S), threonine (Thr; T), tryptophan (Trp; W), tyrosine (Tyr; Y), and valine (Val; V).
- amino acids include, but are not limited to, alanine, ⁇ -alanine, arginine, asparagine, aspartic acid, cysteine, cystine, glutamic acid, glutamine, glycine, phenylalanine, histidine, isoleucine, lysine, leucine, methionine, proline, serine, threonine, valine, tryptophan, or tyrosine, among others.
- amino acid also includes “non-natural amino acids” (nnAA) and “modified amino acids.”
- nAA non-natural amino acids
- modified amino acids are used herein interchangeably.
- Naturally encoded amino acids are the proteinogenic amino acids known to those of skill in the art. They include the 20 common amino acids (alanine, arginine, asparagine, 18 aspartic acid, cysteine, glutamine, glutamic acid, glycine, histidine, isoleucine, leucine, lysine, methionine, phenylalanine, proline, serine, threonine, tryptophan, tyrosine, and valine) and the less common pyrrolysine and selenocysteine.
- 20 common amino acids alanine, arginine, asparagine, 18 aspartic acid, cysteine, glutamine, glutamic acid, glycine, histidine, isoleucine, leucine, lysine, methionine, phenylalanine, proline, serine, threonine, tryptophan, tyrosine, and valine
- Naturally encoded amino acids include post- translational variants of the 22 naturally occurring amino acids such as prenylated amino acids, isoprenylated amino acids, myristoylated amino acids, palmitoylated amino acids, N-linked glycosylated amino acids, O-linked glycosylated amino acids, phosphorylated amino acids and acylated amino acids.
- a “conservative substitution,” or a “conservative amino acid substitution,” as used herein, refers to the substitution of an amino acid with a chemically or functionally similar amino acid. Conservative substitution tables providing similar amino acids are well known in the art.
- polypeptide sequences having such substitutions are known as “conservatively modified variants.” Such conservatively modified variants are in addition to and do not exclude polymorphic variants, interspecies homologs, and alleles.
- the groups of amino acids provided in Tables 1-3 are, in some embodiments, considered conservative substitutions for one another. Table 1. Selected groups of amino acids that are considered conservative substitutions 19 Table 2. Additional selected groups of amino acids that are considered conservative substitutions for one another, in certain embodiments. Group 1 A, S, and T Group 2 D and E Group 3 N and Q Group 4 R and K Group 5 I, L, and M Group 6 F, Y, and W Table 3. Further selected groups of amino acids that are considered conservative substitutions for one another, in certain embodiments.
- modified amino acid refers to an amino acid that is not a proteinogenic amino acid, or a post-translationally modified variant thereof.
- non-natural amino acid refers to an amino acid that is not one of the 20 common amino acids or pyrrolysine or selenocysteine, or post-translationally modified variants thereof.
- modified amino acids include e.g., p-acetylphenylalanine (pAcF), azido-lysine (AzK), and p- azidomethyl-L-phenylalanine (pAMF).
- Additional non-limiting examples include O-methyl- L-tyrosine, 3-methyl-phenylalanine, O-4-allyl-L-tyrosine, 4-propyl-L-tyrosine, fluorinated phenylalanine, isopropyl-L-phenylalanine, p-azido-L-phenylalanine, p-acyl-L-phenylalanine, p-A benzoyl-L-phenylalanine, p-iodo-phenylalanine, p-bromophenylalanine, p-amino-L- phenylalanine, isopropyl-L-phenylalanine, and p-propargyloxy-pheny
- Modified amino acids such as pAcF, AzK, and pAMF, provide side chains to which various secondary molecules e.g., polyethyleneglycol (PEG) can be conjugated/bound.
- PEG polyethyleneglycol
- a modified amino acid is pAMF.
- pAMF is typically incorporated into proteins at the TAG 20 amber codon using method known in the art (see e.g., Zimmerman, E. S. et al. Bioconjug. Chem. 25, 351–361 (2014)).
- pAMF incorporation provides an efficient approach for site- specific modification of the proteins and subsequent conjugation-site specific modification.
- non-natural amino acid or “unnatural amino acid”) or “synthetic amino acids” are ⁇ , ⁇ , ⁇ , or ⁇ amino acids, also includes but is not limited to, amino acids found in proteins, i.e., glycine, alanine, valine, leucine, isoleucine, methionine, phenylalanine, tryptophan, proline, serine, threonine, cysteine, tyrosine, asparagine, glutamine, aspartate, glutamate, lysine, arginine and histidine.
- the amino acid is in the L- configuration.
- the amino acid can be a derivative of alanyl, valinyl, leucinyl, isoleucinyl, prolinyl, phenylalaninyl, tryptophanyl, methioninyl, glycinyl, serinyl, threoninyl, cysteinyl, tyrosinyl, asparaginyl, glutaminyl, aspartoyl, glutaroyl, lysinyl, argininyl, histidinyl, ⁇ -alanyl, ⁇ -valinyl, ⁇ -leucinyl, ⁇ -isoleuccinyl, ⁇ -prolinyl, ⁇ -phenylalaninyl, ⁇ -tryptophanyl, ⁇ -methioninyl, ⁇ -glycinyl, ⁇ -serinyl, ⁇ -threoninyl, ⁇ -cysteinyl
- Unnatural amino acids are not proteinogenic amino acids, or post-translationally modified variants thereof that either occur naturally or are chemically synthesized.
- unnatural amino acid refers to an amino acid that is not one of the 20 common amino acids or pyrrolysine or selenocysteine, or post-translationally modified variants thereof.
- Non-limiting examples of unnatural amino acids include sulfoalanine, hydroxyproline (Hyp), beta-alanine, citrulline (Cit), ornithine (Orn), norleucine (Nle), 3-nitrotyrosine, nitroarginine, pyroglutamic acid (Pyr), naphtylalanine (Nal), 2,4-diaminobutyric acid (DAB), methionine sulfoxide, and methionine sulfone.
- the term “disease,” or “disease or disorder” as used herein, refers any condition or disorder that damages or interferes with the normal function of a cell, tissue, or organ.
- treating or “treatment” of any disease or disorder refers, in certain embodiments, to ameliorating a disease or disorder that exists in a subject. “Treating” or “treatment” includes ameliorating at least one physical parameter, which may be indiscernible by the subject. In yet another embodiment, “treating” or “treatment” includes modulating the disease or disorder, either physically (e.g., stabilization of a discernible symptom) or physiologically (e.g., stabilization of a physical parameter) or both. In yet another embodiment, “treating” or “treatment” includes delaying or preventing the onset of the disease or disorder. For example, in an exemplary embodiment, the phrase “treating cancer” refers to inhibition of 21
- treatment includes preventing or delaying the recurrence of the disease, delaying or slowing the progression of the disease, ameliorating the disease state, providing a remission (partial or total) of the disease, decreasing the dose of one or more other medications required to treat the disease, delaying the progression of the disease, increasing or improving the quality of life, increasing weight gain, and/or prolonging survival. Also encompassed by “treatment” is a reduction of pathological consequence of cancer.
- anti-tumor immune memory refers to the immune system’s ability to recognize a previously encountered tumor antigen, and through T cell activation and proliferation, mount a stronger and faster response to the tumor antigen compared to the first encounter based on the memory of the first encounter.
- therapeutically effective amount or “effective amount” refers to an amount of a substance e.g., an IFN ⁇ polypeptide or IFN ⁇ conjugate disclosed herein, or a composition comprising a substance, that when administered to a subject is effective to treat a disease or disorder.
- the phrase “effective amount” is used interchangeably with “therapeutically effective amount” or “therapeutically effective dose” and the like, and means an amount of a therapeutic agent that is effective to prevent or ameliorate a disease or the progression of the disease e.g., cancer, or result in amelioration of symptoms.
- Effective amounts of the compositions provided herein may vary according to factors such as the disease state, age, sex, weight of the animal or human.
- the term “subject,” as used herein, refers to a mammalian subject. Exemplary subjects include, but are not limited to humans, monkeys, dogs, cats, mice, rats, cows, pigs, horses, camels, avians, goats, and sheep.
- the subject is a human.
- the subject has a disease that can be treated with an IFN ⁇ polypeptide or IFN ⁇ conjugate provided herein.
- the term “therapeutically effective amount,” or “effective amount” as used herein, refers to the amount of the subject compound or composition that will elicit the biological, physiologic, clinical or medical response of a cell, tissue, organ, system, or subject that is being sought by the researcher, veterinarian, medical doctor or other clinician.
- terapéuticaally effective amount refers to an amount of a compound e.g., an IFN ⁇ polypeptide or IFN ⁇ conjugate, or composition that, when administered, is sufficient to prevent development of, or treat at least to some extent, one or more of the signs or symptoms of the 22
- a pharmaceutical composition refers to a composition that can be administrated to a subject in the context of treatment of a disease or disorder.
- a pharmaceutical composition comprises an active ingredient, e.g., an IFN ⁇ polypeptide as disclosed herein, and a pharmaceutically acceptable excipient.
- IFN ⁇ Polypeptides Provided herein are IFN ⁇ polypeptides that comprise at least one non-natural amino acid or modified amino acid substitution compared to a wild type IFN ⁇ .
- the IFN ⁇ can be any IFN ⁇ known to the person of skill. In certain embodiments, the IFN ⁇ is any IFN ⁇ subtype. In certain embodiments, the IFN ⁇ is IFN ⁇ 2. In some embodiments, the IFN ⁇ polypeptides comprise at least two non-natural amino acid or modified amino acid substitutions. In some embodiments, the IFN ⁇ polypeptides comprise at least three, four, five, six, or more non- natural amino acid or modified amino acid substitutions. [000116] The at least one non-natural amino acid or modified amino acid substitution can be made by standard techniques. In certain embodiments, the substitution is made by one or more mutations in the genetic sequence encoding the IFN ⁇ polypeptide.
- the IFN ⁇ polypeptide comprises at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
- the IFN ⁇ polypeptide comprises at least two non-natural amino acids or modified amino acids at positions selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
- the IFN ⁇ polypeptide comprises at least three non-natural amino acids or modified amino acids at positions selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
- the IFN ⁇ polypeptide comprises at least four non-natural amino acids or modified amino acids at positions selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
- the IFN ⁇ polypeptide comprises at least five or more non-natural amino acids or modified amino acids at positions selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
- the IFN ⁇ polypeptide comprises a non-natural amino acid or modified amino acid in at least one amino acid position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156.
- the IFN ⁇ polypeptide comprises at least two non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFN ⁇ polypeptide comprises at least three non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFN ⁇ polypeptide comprises at least four non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156.
- the IFN ⁇ polypeptide comprises at least five non- natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. [000119] In some embodiments, the IFN ⁇ polypeptide comprises a non-natural amino acid or modified amino acid at H7. In some embodiments, the IFN ⁇ polypeptide comprises a non- natural amino acid or modified amino acid at Q40. In some embodiments, the IFN ⁇ polypeptide comprises a non-natural amino acid or modified amino acid at E41. In some embodiments, the IFN ⁇ polypeptide comprises a non-natural amino acid or modified amino acid at N45.
- the IFN ⁇ polypeptide comprises a non-natural amino acid or modified amino acid at E51. In some embodiments, the IFN ⁇ polypeptide comprises a non-natural amino acid or modified amino acid at N156. [000120] In some embodiments, the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at H7 and Q40. In some embodiments, the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at H7 and Q40 and further comprises at least one additional non-natural amino acid or modified amino acid at a position selected from E51, N45, and N156. In some embodiments, the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at H7, Q40 and E51.
- the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at H7, Q40 and N156. In some embodiments, the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at H7, Q40, N45, and N156. In some embodiments, the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at H7 and E51. In some 24
- the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at H7, E51 and N156. [000121] In some embodiments, the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at Q40 and E51. In some embodiments, the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at Q40 and N156. In some embodiments, the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at E51 and N156. In some embodiments, the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at Q40, E51, and N156. [000122] In some embodiments, the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at Q40 and N156.
- the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids at Q40 and N156 and further comprises at least one non-natural amino acids or modified amino acids at a position selected from H7 and E51.
- the IFN ⁇ polypeptide comprises at least one non-natural amino acid or modified amino acid in a position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprises at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136.
- the IFN ⁇ polypeptide comprises at least two non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprises at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136.
- the IFN ⁇ polypeptide comprises at least three non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprises at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136.
- the IFN ⁇ polypeptide comprises non-natural amino acids or modified amino acids in positions Q40 and N156 and further comprises at least one non-natural amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, 25
- the non-natural amino acid or modified amino acid is a non-natural amino acid.
- the non-natural amino acid or modified amino acid is a modified amino acid.
- the non-natural amino acid or modified amino acid comprises a residue of a moiety selected from the group consisting of amino, carboxy, acetyl, hydrazino, hydrazido, semicarbazido, sulfanyl, azido and alkynyl.
- the modified amino acid is selected from the group consisting of p-acetyl-L-phenylalanine, O-methyl-L-tyrosine, 3-methyl-phenylalanine, O-4-allyl-L-tyrosine, 4-propyl-L-tyrosine, fluorinated phenylalanine, isopropyl-L- phenylalanine, p-azido-L-phenylalanine, p-acyl-L-phenylalanine, p-benzoyl-L-phenylalanine, p-iodo-phenylalanine, p-bromophenylalanine, p-amino-L-phenylalanine, isopropyl-L- phenylalanine, p-propargyloxy-phenylalanine, and p-azidomethyl-L-phenylalanine residues.
- the non-natural amino acid is a p-azidomethyl-L- phenylalanine residue.
- the IFN ⁇ polypeptide comprises a p-azidomethyl-L- phenylalanine residue in at least one amino acid position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
- the IFN ⁇ polypeptide comprises a p-azidomethyl-L- phenylalanine residue in at least one amino acid position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFN ⁇ polypeptide comprises a p-azidomethyl-L-phenylalanine residue in at least two amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156.
- the IFN ⁇ polypeptide comprises a p-azidomethyl-L-phenylalanine residue in at least three amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFN ⁇ polypeptide comprises a p-azidomethyl-L-phenylalanine residue in at least four amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156.
- the IFN ⁇ polypeptide comprises a p-azidomethyl-L- phenylalanine residue in at least five amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFN ⁇ polypeptide comprises a p-azidomethyl-L-phenylalanine residue in positions H7, Q40, E41, N45, E51, and N156. 26
- the IFN ⁇ polypeptide comprises a p-azidomethyl-L- phenylalanine residue at H7. In some embodiments, the IFN ⁇ polypeptide comprises a p- azidomethyl-L-phenylalanine residue at Q40. In some embodiments, the IFN ⁇ polypeptide comprises a p-azidomethyl-L-phenylalanine residue at E41. In some embodiments, the IFN ⁇ polypeptide comprises a p-azidomethyl-L-phenylalanine residue at N45. In some embodiments, the IFN ⁇ polypeptide comprises p-azidomethyl-L-phenylalanine residue at E51.
- the IFN ⁇ polypeptide comprises a p-azidomethyl-L-phenylalanine residue at N156. [000133] In some embodiments, the IFN ⁇ polypeptide comprises p-azidomethyl-L- phenylalanine residues at H7 and Q40. In some embodiments, the IFN ⁇ polypeptide comprises p-azidomethyl-L-phenylalanine residues at H7 and Q40 and further comprises at least one additional p-azidomethyl-L-phenylalanine residue at a position selected from E51, N45, and N156.
- the IFN ⁇ polypeptide comprises p-azidomethyl-L-phenylalanine residues at H7, Q40 and E51. In some embodiments, the IFN ⁇ polypeptide comprises p- azidomethyl-L-phenylalanine residues at H7, Q40 and N156. In some embodiments, the IFN ⁇ polypeptide comprises p-azidomethyl-L-phenylalanine residues at H7, Q40, N45, and N156. In some embodiments, the IFN ⁇ polypeptide comprises p-azidomethyl-L-phenylalanine residues at H7 and E51.
- the IFN ⁇ polypeptide comprises p-azidomethyl- L-phenylalanine residues at H7, E51 and N156. [000134] In some embodiments, the IFN ⁇ polypeptide comprises p-azidomethyl-L- phenylalanine residues at Q40 and E51. In some embodiments, the IFN ⁇ polypeptide comprises p-azidomethyl-L-phenylalanine residues at Q40 and N156. In some embodiments, the IFN ⁇ polypeptide comprises p-azidomethyl-L-phenylalanine residues at E51 and N156.
- the IFN ⁇ polypeptide comprises p-azidomethyl-L-phenylalanine residues at Q40, E51, and N156. [000135] In some embodiments, the IFN ⁇ polypeptide comprises p-azidomethyl-L- phenylalanine residues at Q40 and N156. In some embodiments, the IFN ⁇ polypeptide comprises p-azidomethyl-L-phenylalanine residues at Q40 and N156 and further comprises at least one p-azidomethyl-L-phenylalanine residue at a position selected from H7 and E51. [000136] In some embodiments, the amino acid substitution position is according to the sequence of wild-type IFN ⁇ .
- the amino acid substitution is with reference to SEQ ID NO: 33.
- the IFN ⁇ polypeptide comprises an amino acid sequence having at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 33.
- the IFN ⁇ polypeptide comprises an amino acid sequence according to: SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, or SEQ ID NO: 8.
- SEQ ID NO: 3 amino acid sequence according to: SEQ ID NO: 4
- SEQ ID NO: 5 amino acid sequence according to: SEQ ID NO: 6
- SEQ ID NO: 7 amino acid sequence according to: SEQ ID NO: 8.
- SEQ ID NO: 3 amino acid sequence according to: SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, or SEQ ID NO: 8.
- post-translationally modified variants of the IFN ⁇ polypeptides disclosed herein Any of the IFN ⁇ polypeptides provided herein can be post- translationally modified in any manner recognized by those of skill in the art. Typical post- translational modifications for IFN ⁇ polypeptides include interchain disulfide bonding and glycosylation. The post-translational modification
- the post-translational modification can be an intentional modification by a practitioner, for instance, using the methods provided herein.
- IFN ⁇ polypeptides fused to further peptides or polypeptides include, but are not limited to, e.g., a methionyl IFN ⁇ polypeptide in which a methionine is linked to the N-terminus of the IFN ⁇ polypeptide resulting from recombinant expression, fusions for the purpose of purification (including but not limited to, to poly-histidine or affinity epitopes), fusions with serum albumin binding peptides, and fusions with serum proteins such as serum albumin.
- the IFN ⁇ polypeptides may comprise protease cleavage sequences, IFN ⁇ polypeptide-binding domains (including but not limited to, FLAG or poly-His) or other affinity-based sequences (including but not limited to, FLAG, poly-His, GST, etc.).
- the IFN ⁇ polypeptides may also comprise linked molecules that improve detection (including, but not limited to, GFP), purification, or other features of the IFN ⁇ polypeptide.
- the IFN ⁇ polypeptides comprise a C-terminal affinity sequence that facilitates purification of full length IFN ⁇ polypeptides.
- such C-terminal affinity sequence is a poly-His sequence, e.g., a 6-His sequence.
- the IFN ⁇ polypeptides comprise an N-terminal affinity sequence that facilitates purification of full length IFN ⁇ polypeptides.
- such N- terminal affinity sequence is a poly-His sequence, e.g., a 6-His sequence.
- the IFN ⁇ polypeptides are fused to a polypeptide sequence that facilitates expression or purification.
- the fusion polypeptide sequence is a small ubiquitin modifying protein (SUMO; Butt et al., 2009, Protein Expr Purif. 43(1): 1–9).
- the fusion protein can be cleaved from the IFN ⁇ polypeptide during or after expression or purification.
- the fused peptide or polypeptide specifically binds to a target molecule other than the target molecule bound by the IFN ⁇ polypeptide.
- the at least one non-natural amino acid or modified amino acid substitution provides an IFN ⁇ polypeptide that has reduced IFNAR binding compared to wild-type IFN ⁇ . In some embodiments, the at least one non-natural amino acid substitution or modified amino acid provides an IFN ⁇ polypeptide that has reduced toxicity, for example, systemic toxicity, compared to wild-type IFN ⁇ . In some embodiments, the at least one non- natural amino acid or modified amino acid substitution provides an IFN ⁇ polypeptide that has increased stability, for example, increased stability in serum, compared to wild-type IFN ⁇ .
- the at least one non-natural amino acid or modified amino acid substitution provides an IFN ⁇ polypeptide that has a longer half-life in serum compared to wild-type IFN ⁇ . In some embodiments, the at least one non-natural amino acid or modified amino acid substitution provides an IFN ⁇ polypeptide that has reduced toxicity and increased stability compared to wild-type IFN ⁇ . [000141] In certain embodiments, the IFN ⁇ polypeptide has increased affinity for IFNAR. In certain embodiments, the at least one non-natural amino acid or modified amino acid is on an IFNAR receptor contacting surface of the IFN ⁇ polypeptide.
- the at least one non-natural amino acid or modified amino acid in the IFN ⁇ polypeptide is located at an amino acid position that contacts IFNAR through hydrogen bonds and/or ionic bonds. In certain embodiments, the at least one non-natural amino acid or modified amino acid in the IFN ⁇ polypeptide is at a position that contacts IFNAR through ionic bonds. In certain embodiments, one or more non-natural amino acids or modified amino acids increase binding of IFN ⁇ polypeptide to IFNAR relative to an IFN ⁇ of the same sequence, other than the one or more non-natural amino acids or modified amino acids.
- one or more non-natural amino acids or modified amino acids increase binding of IFN ⁇ polypeptide to IFNAR by 10%, 20%, 25%, 50%, 75%, 100%, 125%, 150%, 200%, 250%, 300%, 400%, 500%, 1000%, 2000%, 3000%, or more.
- the one or more non-natural amino acids or modified amino acid increase the stability of the IFN ⁇ polypeptide.
- the one or more non-natural amino acids or modified amino acids increase the serum half-life of the IFN ⁇ polypeptide.
- the one or more non-natural amino acids or modified amino acids increase the serum half-life of the IFN ⁇ polypeptide relative to wild-type IFN ⁇ .
- the one or more non-natural amino acids or modified amino acids increase the serum half-life of the IFN ⁇ polypeptide relative to an IFN ⁇ of the same sequence, other than the one or more non-natural amino acids or modified amino acid. In certain embodiments, the one or more non-natural amino acids increase the serum half-life of the IFN ⁇ 29
- the non-natural amino acid or modified amino acid is a non-natural amino acid.
- the non-natural amino acid or modified amino acid is a modified amino acid.
- IFN ⁇ Conjugates comprising an IFN ⁇ polypeptide and a masking moiety, wherein the IFN ⁇ polypeptide is linked to the masking moiety, optionally via a linker.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide as described herein with at least one non-natural amino acid or modified amino acid wherein the at least one non-natural amino acid or modified amino acid is linked to a masking moiety, optionally via a linker.
- the masking moiety is a water-soluble polymer, a carbohydrate, or a peptide.
- the non-natural amino acid or modified amino acid is a non-natural amino acid.
- the non-natural amino acid or modified amino acid is a modified amino acid.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide site- specifically linked to a masking moiety via a protease cleavable linker wherein the masking moiety is a water-soluble polymer or carbohydrate.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide site- specifically linked to a masking moiety via a pH-sensitive linker.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide site-specifically linked to a masking moiety via a cathepsin B cleavable linker.
- the masking moiety is a water- soluble polymer, a carbohydrate, or a peptide.
- the IFN ⁇ conjugates described herein can be linked to one, two, three, four, five, six, or more masking moieties optionally via linker(s).
- the linker can be any linker capable of forming at least one bond to the IFN ⁇ polypeptide and at least one bond to a masking moiety. Useful linkers are described the sections and examples below.
- the linkers are protease cleavable or pH-sensitive.
- the conjugate can be formed from an IFN ⁇ polypeptide that comprises one or more reactive groups.
- the conjugate can be formed from an IFN ⁇ polypeptide comprising all naturally encoded amino acids. Those of skill in the art will recognize that several naturally encoded amino acids include reactive groups capable of conjugation to a masking moiety or to a linker. These reactive groups include 30
- the conjugate can comprise a masking moiety or linker linked to the residue of a reactive group.
- the masking moiety precursor or linker precursor comprises a reactive group capable of forming a bond with a reactive group.
- Typical reactive groups include maleimide groups, activated carbonates (including but not limited to, p-nitrophenyl ester), activated esters (including but not limited to, N-hydroxysuccinimide, p-nitrophenyl ester, and aldehydes).
- Particularly useful reactive groups include maleimide and succinimide, for instance N-hydroxysuccinimide, for forming bonds to cysteine and lysine side chains.
- the IFN ⁇ polypeptide comprises one or more modified amino acids having a reactive group, as described herein.
- the modified amino acid is not a naturally encoded amino acid.
- These modified amino acids can comprise a reactive group useful for forming a covalent bond to a masking moiety precursor or to a payload precursor.
- One of skill in the art can use the reactive group to link the IFN ⁇ polypeptide to any molecular entity capable of forming a covalent bond to the modified amino acid.
- conjugates comprising an IFN ⁇ polypeptide comprising a modified amino acid residue linked to a payload directly or indirectly via a linker.
- modified amino acids are described in the sections below.
- the modified amino acids have reactive groups capable of forming bonds to linkers or payloads with complementary reactive groups.
- the non-natural amino acids or modified amino acids are positioned at select locations in a polypeptide chain of the IFN ⁇ polypeptide. These locations were identified as providing optimum sites for substitution with the non-natural amino acids or modified amino acids. Each site is capable of bearing a non-natural amino acid or modified amino acid with optimum structure, function and/or methods for producing the IFN ⁇ polypeptide.
- a site-specific position for substitution provides an IFN ⁇ polypeptide or conjugate that is stable. Stability can be measured by any technique apparent to those of skill in the art.
- a site-specific position for substitution provides an IFN ⁇ polypeptide or conjugate that has optimal functional properties. For instance, the IFN ⁇ polypeptide or conjugate can show little or no loss of binding affinity for its target antigen compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. In certain embodiments, the IFN ⁇ polypeptide or conjugate can show 31
- a site-specific position for substitution provides an IFN ⁇ polypeptide or conjugate that can be made advantageously.
- the IFN ⁇ polypeptide or conjugate shows advantageous properties in its methods of synthesis, discussed below.
- the IFN ⁇ polypeptide or conjugate can show little or no loss in yield in production compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid.
- the IFN ⁇ polypeptide or conjugate can show enhanced yield in production compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. In certain embodiments, the IFN ⁇ polypeptide or conjugate can show little or no loss of tRNA suppression compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. In certain embodiments, the IFN ⁇ polypeptide or conjugate can show enhanced tRNA suppression in production compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. [000154] In certain embodiments, a site-specific position for substitution provides an IFN ⁇ polypeptide or conjugate that has advantageous solubility.
- the IFN ⁇ polypeptide or conjugate can show little or no loss in solubility compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. In certain embodiments, the IFN ⁇ polypeptide or conjugate can show enhanced solubility compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. [000155] In certain embodiments, a site-specific position for substitution provides an IFN ⁇ polypeptide or conjugate that has advantageous expression. In certain embodiments, the IFN ⁇ polypeptide or conjugate can show little or no loss in expression compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid.
- the IFN ⁇ polypeptide or conjugate can show enhanced expression compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid.
- a site-specific position for substitution provides an IFN ⁇ polypeptide or conjugate that has advantageous folding.
- the IFN ⁇ polypeptide or conjugate can show little or no loss in proper folding compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino 32
- the IFN ⁇ polypeptide or conjugate can show enhanced folding compared to an IFN ⁇ polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid.
- a site-specific position for substitution provides an IFN ⁇ polypeptide that is capable of advantageous conjugation.
- several nonnatural amino acids have side chains or functional groups that facilitate conjugation of the IFN ⁇ polypeptide to a second agent, either directly or via a linker.
- the IFN ⁇ polypeptide can show enhanced conjugation efficiency compared to an IFN ⁇ polypeptide without the same or other non-natural amino acids or modified amino acids at other positions.
- the IFN ⁇ polypeptide can show enhanced conjugation yield compared to an IFN ⁇ polypeptide without the same or other non-natural amino acids or modified amino acids at other positions. In certain embodiments, the IFN ⁇ polypeptide can show enhanced conjugation specificity compared to an IFN ⁇ polypeptide without the same or other non-natural amino acids or modified amino acids at other positions. [000158] The one or more non-natural amino acids or modified amino acids are located at selected site-specific positions in at least one polypeptide chain of the IFN ⁇ conjugate.
- the polypeptide chain can be any polypeptide chain of the IFN ⁇ polypeptide without limitation.
- the IFN ⁇ polypeptides or conjugate provided herein comprise one non-natural amino acid or modified amino acid at a site-specific position. In certain embodiments, the IFN ⁇ polypeptide or conjugate provided herein comprise two non- natural amino acids or modified amino acids at site-specific positions. In certain embodiments, the IFN ⁇ polypeptide or conjugate provided herein comprise three non-natural amino acids or modified amino acids at site-specific positions. In certain embodiments, the IFN ⁇ polypeptide or conjugate provided herein comprise more than three non-natural amino acids or modified amino acids at site-specific positions. In certain embodiments, the IFN ⁇ polypeptide or conjugate provided herein comprise four non-natural amino acids or modified amino acids at site-specific positions.
- the non-natural or modified amino acid is a non-natural amino acid. In certain embodiments, the non-natural or modified amino acid a modified amino acid. [000160] In certain embodiments, the IFN ⁇ conjugate is of Formula (I): 33
- COMP is an IFN ⁇ polypeptide
- L is a linker, for example, a linker that comprises a protease cleavable linker or a pH-sensitive linker
- MM is a masking moiety
- x is an integer selected from 0 and 1
- y is an integer between 1 and 30.
- IFN ⁇ is an IFN ⁇ polypeptide that comprises at least one non-natural amino acid or modified amino acid selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
- IFN ⁇ is an IFN ⁇ polypeptide that comprises at least one non-natural amino acid or modified amino acid selected from the group consisting of: H7, Q40, E41, N45, E51, and N156.
- linker L is a cathepsin B cleavable linker.
- the masking moiety is a water-soluble polymer, a carbohydrate, or a peptide.
- IFN ⁇ is an IFN ⁇ polypeptide
- L is a protease cleavable linker
- MM is a water soluble polymer or a carbohydrate.
- IFN ⁇ is an IFN ⁇ polypeptide and L is a pH-sensitive linker.
- IFN ⁇ is an IFN ⁇ polypeptide and L is a cathepsin B cleavable linker.
- x is 0. In certain embodiments, x is 1. In certain embodiments, y is 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10. In certain embodiments, y is 1, 2, 3, 4, 5, or 6. In certain embodiments, y is 1, 2, 3, or 4. In certain embodiments, y is 1. In certain embodiments, y is 2. In certain embodiments, y is 3. In certain embodiments, y is 4. [000168] In certain embodiments, x is 0 and y is 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10. In certain embodiments, x is 1 and y is 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10. In certain embodiments, x is 0 and y is 1, 2, 3, 4, 5, or 6.
- x is 1 and y is 1, 2, 3, 4, 5, or 6. In certain embodiments, x is 0 and y is 1. In certain embodiments, x is 0 and y is 2. In certain embodiments, x is 0 and y is 3. In certain embodiments, x is 0 and y is 4. In certain embodiments, x is 0 and y is 5. In certain embodiments, x is 0 and y is 6. In certain embodiments, x is 1 and y is 1. In certain embodiments, x is 1 and y is 2. In certain 34
- x is 1 and y is 3. In certain embodiments, x is 1 and y is 4. In certain embodiments, x is 1 and y is 5. In certain embodiments, x is 1 and y is 6. 1.3.1.
- the masking moiety can be any macromolecule deemed suitable by the person of skill in the art. In certain embodiments, the masking moiety is a protein, peptide, antibody or antigen binding fragment thereof, nucleic acid, carbohydrate, or other large molecule composed of polymerized monomers. In certain embodiments, the masking moiety is a protein.
- the masking moiety is an antibody or an antigen binding fragment thereof. In some embodiments, the masking moiety is a residue of a polypeptide. In some embodiments, the masking moiety is a residue of an antibody. In some embodiments, the masking moiety is a residue of an antibody chain. [000171] In certain embodiments, the masking moiety is a polymer, denotated as POLY herein, for example a water-soluble polymer. These polymers can be linked to the polypeptide via a naturally encoded amino acid, via a non-naturally encoded amino acid, or any functional substituent of a natural or modified amino acid, or any substituent or functional group added to a natural or modified amino acid.
- the polymer can also be linked to the polypeptide via a linker as described herein.
- the molecular weight of the polymer may be of a wide range, including but not limited to, between about 100 Da and about 100,000 Da or more.
- the polymer selected may be water soluble so that a protein to which it is attached does not precipitate in an aqueous environment, such as a physiological environment.
- the polymer may be branched or unbranched.
- the polymer will be pharmaceutically acceptable.
- the water-soluble polymer may be any structural form including, but not limited to linear, forked or branched.
- the water soluble polymer is a poly(alkylene glycol), such as poly(ethylene glycol) (PEG), but other water soluble polymers can also be employed.
- PEG poly(ethylene glycol)
- Alternative examples of polymers include, but are not limited to, other poly(alkylene glycols) such as poly(propylene glycol) (“PPG”), copolymers of ethylene glycol and propylene glycol and the like, poly(oxyethylated polyol), poly(olefinic alcohol), poly(vinylpyrrolidone), poly(hydroxyalkylmethacrylamide), poly(hydroxy- alkylmethacrylate), poly(saccharides), poly( ⁇ -hydroxy acid), poly(vinyl alcohol), 35
- PPG poly(propylene glycol)
- PPG poly(propylene glycol)
- copolymers of ethylene glycol and propylene glycol and the like poly(oxyethylated polyol), poly(olefinic alcohol), poly(viny
- PEG polyphosphazene, polyoxazolines (“POZ”) (which are described in WO 2008/106186), poly(N-acryloylmorpholine), and combinations of any of the foregoing.
- PEG is a well-known, water-soluble polymer that is commercially available or can be prepared by ring-opening polymerization of ethylene glycol according to methods well known in the art (Sandler and Karo, Polymer Synthesis, Academic Press, New York, Vol.3, pages 138-161).
- PEG is used broadly to encompass any polyethylene glycol molecule, without regard to size or to modification at an end of the PEG, and can be represented as linked to a polypeptide by the formula: XO–(CH 2 CH 2 O) n –CH 2 CH 2 –Y where n is 2 to 10,000,X is H or a terminal modification, including but not limited to, a C1-4 alkyl, and Y is the attachment point to the polypeptide.
- a PEG terminates on one end with hydroxy or methoxy, i.e., X is H or CH3 (“methoxy PEG” or “mPEG”).
- the PEG can terminate with a reactive group, thereby forming a bifunctional polymer.
- Typical reactive groups can include those reactive groups that are commonly used to react with the functional groups found in the 20 common amino acids (including but not limited to, maleimide groups, activated carbonates (including but not limited to, p-nitrophenyl ester), activated esters (including but not limited to, N hydroxysuccinimide, p-nitrophenyl ester, and aldehydes) as well as functional groups that are inert to the 20 common amino acids but that react specifically with complementary functional groups present in non-naturally encoded or modified amino acids (including but not limited to, azide groups, alkyne groups).
- functional groups found in the 20 common amino acids including but not limited to, maleimide groups, activated carbonates (including but not limited to, p-nitrophenyl ester), activated esters (including but not limited to, N hydroxysuccinimide, p-nitrophenyl ester, and aldehydes) as well as functional groups that are inert to the 20 common amino acids but that react specifically with complementary functional groups present
- Y may be an amide, carbamate or urea linkage to an amine group (including but not limited to, the epsilon amine of lysine or the N-terminus) of the polypeptide.
- Y may be a maleimide linkage to a thiol group (including but not limited to, the thiol group of cysteine).
- Y may be a linkage to a residue not commonly accessible via the 20 common amino acids.
- an azide group on the PEG can be reacted with an alkyne group on the polypeptide to form a Huisgen [3+2] cycloaddition product.
- an alkyne group on the PEG can be reacted with an azide group present in a non-naturally encoded or modified amino acid, such as the modified amino acids described herein, to form a similar product.
- a strong nucleophile including but not limited to, hydrazine, hydrazide, hydroxylamine, semicarbazide
- a strong nucleophile can be reacted with an aldehyde or ketone group present in a non-naturally encoded or modified amino acid to form a hydrazone, oxime or semicarbazone, as applicable, which in some cases can be further reduced by treatment with an appropriate reducing agent.
- the strong nucleophile can be incorporated into the polypeptide via a non- naturally encoded or modified amino acid and used to react preferentially with a ketone or aldehyde group present in the water-soluble polymer.
- the proportion of polyethylene glycol molecules to polypeptide molecules will vary, as will their concentrations in the reaction mixture.
- the optimum ratio in terms of efficiency of reaction in that there is minimal excess unreacted protein or polymer may be determined by the molecular weight of the polyethylene glycol selected and on the number of available reactive groups available. As it relates to molecular weight, typically the higher the molecular weight of the polymer, the fewer number of polymer molecules which may be attached to the protein.
- any molecular mass for a PEG can be used as practically desired, including but not limited to, from about 100 Daltons (Da) to 100,000 Da or more as desired (including but not limited to, sometimes 0.1-50 kDa or 10-40 kDa).
- Branched chain PEGs including but not limited to, PEG molecules with each chain having a MW ranging from 1-100 kDa (including but not limited to, 150 kDa or 5-20 kDa) can also be used.
- PEG molecules are described in, including but not limited to, the Shearwater Polymers, Inc. catalog, and the Nektar Therapeutics catalog, incorporated herein by reference.
- at least one terminus of the PEG molecule is available for reaction with the IFN ⁇ polypeptide or a linker of the IFN ⁇ polypeptide.
- PEG derivatives bearing alkyne and azide moieties for reaction with amino acid side chains can be used to attach PEG to non-naturally encoded or modified amino acids as described herein.
- the PEG will typically contain either an alkyne moiety to effect formation of the [3+2] cycloaddition product or an activated PEG species (i.e., ester, carbonate) containing a phosphine group to effect formation of the amide linkage.
- the PEG will typically contain an azide moiety to effect formation of the [3+2] Huisgen cycloaddition product.
- the PEG will typically comprise a potent nucleophile (including but not limited to, a hydrazide, hydrazine, hydroxylamine, or semicarbazide functionality) in order to effect formation of corresponding hydrazone, oxime, and semicarbazone linkages, respectively.
- a potent nucleophile including but not limited to, a hydrazide, hydrazine, hydroxylamine, or semicarbazide functionality
- the PEG molecule contains a chemical functionality that is reactive with the chemical functionality present on the side chain of the non-naturally encoded or modified amino acid.
- the PEG molecule terminates in an amine which is available for reaction with a linker attached to the polypeptide.
- the linker may terminate in an electrophile, for example, a carboxylic acid, and the amine of the PEG molecule is a nucleophile to form an amide bond.
- the PEG molecule is a mPEG- NHS reagent, including mPEG-succinimidyl ester.
- the NHS ester can react with an amine group, for example on the polypeptide or the linker, at a pH of 7-8.5 to form a stable amide bond.
- the masking moiety is an azide- or acetylene-containing polymer comprising a water-soluble polymer backbone having an average molecular weight from about 800 Da to about 100,000 Da.
- the masking moiety is an amine- or N-hydroxysuccinimide-containing polymer comprising a water-soluble polymer backbone having an average molecular weight from about 800 Da to about 100,000 Da.
- the polymer backbone of the water-soluble polymer can be poly(ethylene glycol).
- water soluble polymers including but not limited to poly(ethylene)glycol and other related polymers, including poly(dextran) and poly(propylene glycol), are also suitable for use and that the use of the term PEG or poly(ethylene glycol) is intended to encompass and include all such molecules.
- PEG includes, but is not limited to, poly(ethylene glycol) in any of its forms, including bifunctional PEG, multiarmed PEG, derivatized PEG, forked PEG, branched PEG, pendent PEG (i.e.
- the polymer backbone can be linear or branched. Branched polymer backbones are generally known in the art. Typically, a branched polymer has a central branch core moiety and a plurality of linear polymer chains linked to the central branch core. PEG is commonly used in branched forms that can be prepared by addition of ethylene oxide to various polyols, such as glycerol, glycerol oligomers, pentaerythritol and sorbitol. The central branch moiety can also be derived from several amino acids, such as lysine.
- the branched poly(ethylene glycol) can be represented in general form as R(-PEG-OH)m in which R is derived from a core moiety, such as glycerol, glycerol oligomers, or pentaerythritol, and m represents the number of arms.
- Multi-armed PEG molecules such as those described in U.S. Pat. Nos.5,932,4625,643,575; 5,229,490; 4,289,872; U.S. Pat. Appl.2003/0143596; WO 96/21469; and WO 93/21259, each of which is incorporated by reference herein in its entirety, can also be used as the polymer backbone.
- Branched PEG can also be in the form of a forked PEG represented by PEG(YCHZ 2 ) n , where Y is a linking group and Z is an activated terminal group linked to CH by a chain of atoms of defined length.
- the pendant PEG has reactive groups, such as carboxyl, along the PEG backbone rather than at the end of PEG chains.
- the polymer can also be prepared with weak or degradable linkages in the backbone.
- PEG can be prepared with ester linkages in the polymer backbone that are subject to hydrolysis.
- poly(ethylene glycol) or PEG represents or includes all the forms known in the art including but not limited to those disclosed herein.
- polymer backbones that are water-soluble, with from 2 to about 300 termini, are particularly suitable.
- suitable polymers include, but are not limited to, other poly(alkylene glycols), such as poly(propylene glycol) (“PPG”), copolymers thereof (including but not limited to copolymers of ethylene glycol and propylene glycol), terpolymers thereof, mixtures thereof, and the like.
- PPG poly(propylene glycol)
- copolymers thereof including but not limited to copolymers of ethylene glycol and propylene glycol
- terpolymers thereof mixtures thereof, and the like.
- the molecular weight of each chain of the polymer backbone can vary, it is typically in the range of from about 800 Da to about 100,000 Da, often from about 6,000 Da to about 80,000 Da.
- substantially water-soluble backbones is by no means exhaustive and is merely illustrative, and that all polymeric materials having the qualities described herein are contemplated as being suitable for use.
- the polymer derivatives are “multi-functional”, meaning that the polymer backbone has at least two termini, and possibly as many as about 300 termini, functionalized or activated with a functional group.
- Multifunctional polymer derivatives include, but are not limited to, linear polymers having two termini, each terminus being bonded to a functional group which may be the same or different.
- POLY is polyethylene glycol (PEG), methoxypolyethylene glycol (mPEG), poly(propylene glycol) (PPG), copolymers of ethylene glycol and propylene glycol, poly(oxyethylated polyol), poly(olefinic alcohol), poly(vinylpyrrolidone), poly(hydroxyalkylmethacrylamide), poly(hydroxyalkylmethacrylate), poly(saccharides), poly( ⁇ -hydroxy acid), poly(vinyl alcohol), polyphosphazene, polyoxazolines (POZ), poly(N- acryloylmorpholine), polysarcosine, or a combination thereof.
- PEG polyethylene glycol
- mPEG methoxypolyethylene glycol
- PPG poly(propylene glycol)
- copolymers of ethylene glycol and propylene glycol poly(oxyethylated polyol), poly(olefinic alcohol), poly(vinylpyrrolidone), poly(hydroxy
- POLY is polyethylene glycol (PEG). In some embodiments, POLY is methoxypolyethylene glycol (mPEG). In some embodiments, POLY is poly(propylene glycol) (PPG). In some embodiments, POLY is copolymers of ethylene glycol and propylene glycol. In some embodiments, POLY is poly(oxyethylated polyol). In some embodiments, POLY is poly(olefinic alcohol). In some embodiments, POLY is poly(vinylpyrrolidone). In some embodiments, POLY is poly(hydroxyalkylmethacrylamide). In some embodiments, POLY is poly(hydroxyalkylmethacrylate). In some embodiments, POLY is poly(saccharides).
- POLY is poly( ⁇ -hydroxy acid). In some embodiments, POLY is poly(vinyl alcohol). In some embodiments, POLY is polyphosphazene. In some embodiments, POLY is polyoxazolines (POZ). In some embodiments, POLY is poly(N-acryloylmorpholine). In some embodiments, POLY is polysarcosine. In some embodiments, POLY is a nonpeptidic, water- soluble polymer. In certain embodiments, POLY includes a polyethylene glycol (PEG) or methoxypolyethylene glycol (mPEG). In certain embodiments, POLY is , wherein represents attachment to the remainder of the compound, and wherein n1 is an integer from 1 to 10,000.
- n1 is an integer from 1 to 5,000. In certain embodiments, n1 is an integer from 1 to 2,500. In certain embodiments, n1 is an integer from 1 to 2,000. In certain embodiments, n1 is an integer from 1 to 1,000. In certain embodiments, n1 is an integer from 100 to 1,000. In certain embodiments, n1 is an integer from 100 to 700. In certain embodiments, n1 is an integer from 300 to 700. In certain embodiments, n1 is an integer from 400 to 500. In certain embodiments, n1 is an integer from 600 to 700. In certain embodiments, n1 is an integer from 100 to 500.
- POLY is a residue of a nonpeptidic, hydrophilic polymer.
- POLY is a residue of polyethylene glycol (PEG), methoxypolyethylene glycol (mPEG), poly(propylene glycol) (PPG), copolymers of ethylene glycol and propylene glycol, poly(oxyethylated polyol), poly(olefinic alcohol), poly(vinylpyrrolidone), poly(hydroxyalkylmethacrylamide), 40
- POLY is a residue of polyethylene glycol (PEG), methoxypolyethylene glycol (mPEG), poly(propylene glycol) (PPG), or a copolymer of ethylene glycol and propylene glycol.
- POLY is a residue of methoxypolyethylene glycol (mPEG).
- POLY is a residue of polyethylene glycol (PEG). In certain embodiments, including any of the foregoing, POLY is a residue of poly(propylene glycol) (PPG). In certain embodiments, including any of the foregoing, POLY is a residue of copolymers of ethylene glycol and propylene glycol. In certain embodiments, including any of the foregoing, POLY is a residue of poly(oxyethylated polyol). In certain embodiments, including any of the foregoing, POLY is a residue of poly(olefinic alcohol). In certain embodiments, including any of the foregoing, POLY is a residue of poly(vinylpyrrolidone).
- POLY is a residue of poly(hydroxyalkylmethacrylamide). In certain embodiments, including any of the foregoing, POLY is a residue of poly(hydroxyalkylmethacrylate). In certain embodiments, including any of the foregoing, POLY is a residue of poly(saccharides). In certain embodiments, including any of the foregoing, POLY is a residue of poly( ⁇ -hydroxy acid). In certain embodiments, including any of the foregoing, POLY is a residue of poly(vinyl alcohol). In certain embodiments, including any of the foregoing, POLY is a residue of polyphosphazene.
- POLY is a residue of polyoxazolines (POZ). In certain embodiments, including any of the foregoing, POLY is a residue of poly(N-acryloylmorpholine). In certain embodiments, including any of the foregoing, POLY is a residue of polysarcosine. [000193] In certain embodiments, including any of the foregoing, POLY is , wherein R 1 is hydrogen or methyl, n1 is an integer from 1 to 10,000, inclusive, and represents attachment to the remainder of the compound or conjugate. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 5000. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 2500. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 1500. In certain embodiments, 41
- n1 is an integer between 100 to 1000. In certain embodiments, n1 is an integer from 100 to 700. In certain embodiments, n1 is an integer from 300 to 700. In certain embodiments, n1 is an integer from 400 to 500. In certain embodiments, n1 is an integer from 600 to 700. In certain embodiments, including any of the foregoing, n1 is an integer between 100 to 500. [000194] In other embodiments, the making group is a carbohydrate. In certain embodiments, the IFN ⁇ may be altered to increase, decrease, or eliminate the extent to which it is glycosylated.
- N-linked glycosylation refers to the attachment of a carbohydrate moiety to the side chain of an asparagine residue.
- the tripeptide sequences asparagine-X-serine and asparagine-X-threonine, where X is any amino acid except proline, are the recognition sequences for enzymatic attachment of the carbohydrate moiety to the asparagine side chain.
- X is any amino acid except proline
- O-linked glycosylation refers to the attachment of one of the sugars Nacetylgalactosamine, galactose, or xylose to a hydroxyamino acid, most commonly serine or threonine, although 5-hydroxyproline or 5-hydroxylysine may also be used.
- Addition of N-linked glycosylation sites to the protein may be accomplished by altering the amino acid sequence such that one or more of the above-described tripeptide sequences is created.
- Addition of O-linked glycosylation sites may be accomplished by addition or substitution of one or more serine or threonine residues in or to (as the case may be) the sequence of a protein. 1.3.2.
- the IFN ⁇ conjugates described herein comprise an IFN ⁇ polypeptide and a masking moiety wherein the IFN ⁇ conjugate is site-specifically linked to the masking moiety, optionally via a linker.
- the linker is cleavable.
- the linker is cleavable in vivo.
- the linker is cleavable by a protease.
- the linker is cleavable by a cathepsin.
- the linker is cleavable by cathepsin B.
- the linker is a pH-sensitive linker.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide site- specifically linked to a masking moiety via a protease cleavable linker wherein the masking moiety is a water-soluble polymer or carbohydrate.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide site-specifically linked to a masking moiety via a cathepsin B 42
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide site-specifically linked to a masking moiety via a pH-sensitive linker.
- the IFN ⁇ conjugate comprises an IFN ⁇ polypeptide site-specifically linked to a masking moiety via a pH-sensitive linker.
- an IFN ⁇ conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof comprising a linker of Formula (L1): wherein RG is a reactive group residue; W 1 and W 2 are independently absent or a divalent attaching group; L 1 is absent, a protease cleavable linker, or a pH-sensitive linker; SG 1 is a divalent spacer group; is a bond to the IFN ⁇ polypeptide; and is a bond to the masking moiety.
- an IFN ⁇ conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof comprising a linker of Formula (L2): wherein RG is a reactive group residue; W 1 and W 2 are independently absent or a divalent attaching group; L 1 is absent, a protease cleavable linker, or a pH-sensitive linker; SG 2 is a trivalent spacer group; is a bond to the IFN ⁇ polypeptide; and is a bond to the masking moiety.
- an IFN ⁇ conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof comprising a linker of Formula (L1) wherein W 1 and W 2 are both divalent attaching groups, L 1 is a protease cleavable linker 43
- the protease cleavable linker is a cathepsin B cleavable linker.
- an IFN ⁇ conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof comprising a linker of Formula (L1) wherein W 1 and W 2 are both divalent attaching groups, L 1 is absent, and RG and SG 1 are as defined herein.
- an IFN ⁇ conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof comprising a linker of Formula (L2) wherein W 1 and W 2 are both divalent attaching groups, L 1 is absent, and RG and SG 2 are as defined herein.
- L2 linker of Formula
- W 1 and W 2 are both divalent attaching groups
- L 1 is absent
- RG and SG 2 are as defined herein.
- Reactive Group Residues [000205] Reactive groups (or conjugating groups) facilitate conjugation of the masking moiety described herein to a second moiety, such as an FN ⁇ described herein.
- the reactive group is designated R herein. Reactive groups can react via any suitable reaction mechanism known to those of skill in the art.
- a reactive group reacts through a [3+2] alkyne-azide cycloaddition reaction, inverse-electron demand Diels-Alder ligation reaction, thiol-electrophile reaction, or carbonyl-oxyamine reaction, as described in detail herein.
- the reactive group comprises an alkyne, strained alkyne, tetrazine, thiol, para-acetyl-phenylalanine residue, oxyamine, maleimide, or azide.
- the reactive group is: , , , , , , , , , , , , , , , , , –N3, methylcyclopropene (e.g. ), 44
- R 201 is lower alkyl.
- the reactive group is: or .
- R 201 is methyl, ethyl, or propyl.
- R 201 is methyl. Additional conjugating groups are described in, for example, U.S. Patent Publication No. 2014/0356385, U.S. Patent Publication No. 2013/0189287, U.S. Patent Publication No.2013/0251783, U.S. Patent No. 8,703,936, U.S. Patent No. 9,145,361, U.S. Patent No.9,222,940, and U.S. Patent No.8,431,558.
- a divalent residue of the reactive group is formed and is bonded to the residue of an IFN ⁇ polypeptide.
- the structure of the divalent residue is determined by the type of conjugation reaction employed to form the conjugate.
- the divalent residue RG comprises a triazole ring or fused cyclic group comprising a triazole ring.
- the divalent residue RG is: and/or .
- the divalent residue RG when a conjugate is formed through a tetrazine inverse electron demand Diels-Alder ligation reaction, the divalent residue RG comprises a fused bicyclic ring having at least two adjacent nitrogen atoms in the ring. In certain embodiments when a conjugate is formed through a tetrazine inverse electron demand Diels-Alder ligation reaction, the divalent residue RG is: or . [000210] In certain embodiments when a conjugate is formed through a thiol-maleimide reaction, the divalent residue RG comprises succinimidylene and a sulfur linkage. In certain 45
- the divalent residue RG when a conjugate is formed through a thiol-maleimide reaction, the divalent residue RG is: , or . [000211] In certain embodiments, a conjugate is formed through a thiol-N- hydroxysuccinimide reaction using the following group: . [000212] The reaction involved for formation of the conjugate comprises the following step: , [000213] and the resulting divalent residue RG is: . [000214] In certain embodiments when a conjugate is formed through a carbonyl-oxyamine reaction, the divalent residue RG comprises a divalent residue of a modified amino acid. In certain embodiments when a conjugate is formed through a carbonyl-oxyamine reaction, the divalent residue RG is: 46
- the divalent residue of RG comprises an oxime linkage. In certain embodiments when a conjugate is formed through a carbonyl-oxyamine reaction, the divalent residue of RG is: .
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises a triazole ring.
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer; wherein RG is a triazole ring or fused cyclic group comprising a triazole ring.
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: or .
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises a fused bicyclic ring having at least two adjacent nitrogen atoms in the ring.
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: 47
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises a sulfur linkage.
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: , , or .
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises a divalent residue of a modified amino acid.
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: or .
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises an oxime linkage.
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: 48
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises an oxime linkage.
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: .
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer or regioisomer thereof; wherein RG is: , , , , , , , , or .
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer or regioisomer thereof; wherein RG is: 49
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W 1 is a divalent attaching group selected from -C(O)-C1-6alkylene-, -C(O)(C1-6alkylene)NR 4 -, -C(O)(C1-6alkylene)O-, and -C(O)(C1- 6 alkylene)S- wherein R 4 is independently hydrogen or C 1-6 alkyl, RG is connected to W 1 at -C(O)-, and the C1-6alkylene is optionally substituted with one, two, or three substituents selected from halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloal
- W 1 is a divalent attaching group of the formula -C(O)- C1-6alkylene-, -C(O)-C1-5alkylene-, -C(O)-C1-4alkylene-, -C(O)-C4-6alkylene-, or -C(O)-C2- 5alkylene-.
- W 1 is a divalent attaching group of the formula -C(O)-C 6 alkylene-, -C(O)-C 5 alkylene-, -C(O)-C 4 alkylene-, -C(O)-C3alkylene-, -C(O)-C2alkylene-, or -C(O)-CH2alkylene.
- W 1 is a divalent attaching group of -C(O)-C 4 alkylene-.
- W 1 is a divalent attaching group of the formula -C(O)(C 1 - 6 alkylene)NH-, C(O)(C 1 - 5 alkylene)NH-, C(O)(C 1 - 4 alkylene)NH-, C(O)(C 1 - 3alkylene)NH-, or C(O)(C2-5alkylene)NH-.
- W 1 is a divalent attaching group of the formula -C(O)(C6alkylene)NH-, C(O)(C5alkylene)NH-, C(O)(C4alkylene)NH-, C(O)(C3 alkylene)NH-, C(O)(C2alkylene)NH- or C(O)(CH2alkylene)NH-.
- W 1 is a divalent attaching group of the formula -C(O)(C 2 alkylene)NH-.
- W 1 is a divalent attaching group of the formula -C(O)(C 1 - 6 alkylene)O-, C(O)(C 1 - 5 alkylene)O-, C(O)(C 1 - 4 alkylene)O-, C(O)(C 1 - 3 alkylene)O-, or C(O)(C2-5alkylene)O-.
- W 1 is a divalent attaching group of the formula -C(O)(C6alkylene)O-, C(O)(C5alkylene)O-, C(O)(C4alkylene)O-, C(O)(C3alkylene)O-, C(O)(C 2 alkylene)O- or C(O)(CH 2 alkylene)O-.
- W 1 is a divalent attaching group of the formula -C(O)(C2alkylene)O-. 50
- W 1 is a divalent attaching group of the formula -C(O)(C 1 - 6 alkylene)S-, C(O)(C 1 - 5 alkylene)S-, C(O)(C 1 - 4 alkylene)S-, C(O)(C 1 - 3 alkylene)O-, or C(O)(C2-5alkylene)S-.
- W 1 is a divalent attaching group of the formula -C(O)(C6alkylene)S-, C(O)(C5alkylene)S-, C(O)(C4alkylene)S-, C(O)(C3alkylene)S-, C(O)(C 2 alkylene)S- or C(O)(CH 2 alkylene)S-.
- W 1 is a divalent attaching group of the formula -C(O)(C2alkylene)S-.
- W 1 is a divalent attaching group of the formula -C(O)(C2alkyl)NH- or -C(O)-C4alkyl-.
- the C 1-6 alkylene of a W 1 divalent attaching group is unsubstituted.
- the C1-6alkylene of a W 1 divalent attaching group is optionally substituted with one, two, or three substituents selected from halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer or regioisomer thereof; wherein W 2 is wherein X 1 is absent, a divalent water-soluble polymer, -C1-6alkylene-, -NR 4 (C1- 6alkylene)-, or -O(C1-6alkylene)-; X 2 is absent or -C1-6alkylene-; X 3 is absent, -NR 4 -, or -O-; R 4 is independently hydrogen or C1-6alkyl; and wherein the C 1 - 6 alkylene of X 1 or X 2 is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl,
- a halogen e.g., flu
- R 1 is hydrogen or methyl and n2 is an integer between 1 and 50, inclusive.
- n2 is an integer between 10 and 40.
- n2 is an integer between 20 and 50.
- n2 is an integer between 1 and 20.
- n2 is an integer between 1 and 15.
- n2 is an integer between 1 and 10.
- n2 is an integer between 10 and 20.
- wherein n2 is 20.
- wherein n2 is 10.
- n2 is an integer between 1 and 6.
- n2 is 4, 5, or 6. In certain embodiments, wherein n2 is 4.
- X 1 is unsubstituted -(C1-6alkylene)-.
- X 1 is -(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- X 1 is -(C1-3alkylene)- wherein the C1-3alkylene is unsubstituted.
- X 1 is -(C 1 - 3 alkylene)- wherein the C 1 - 3 alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- X 1 is -(C3-6alkylene)- wherein the C3-6alkylene is unsubstituted.
- X 1 is -(C 3 - 6 alkylene)- wherein the C 1 - 3 alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 1 is -(CH2)-, -(C2alkylene)-, -(C3alkylene)-, -(C4alkylene)-, -(C5alkylene)-, or -(C6alkylene)- wherein the alkylene is unsubstituted. In certain embodiments, X 1 is unsubstituted -(CH2)-.
- X 1 is -(CH 2 )-, -(C 2 alkylene)-, -(C 3 alkylene)-, -(C 4 alkylene)-, -(C 5 alkylene)-, or -(C 6 alkylene)- wherein the alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 1 is -O(C 1 - 6 alkylene)- wherein the C 1 - 6 alkylene is unsubstituted.
- X 1 is -O(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and 52
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 1 is -O(C 1 - 3 alkylene)- wherein the C 1 - 3 alkylene is unsubstituted. In certain embodiments, X 1 is -O(C 1 - 3 alkylene)- wherein the C 1 - 3 alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 1 is -O(C3-6alkylene)- wherein the C3-6alkylene is unsubstituted.
- X 1 is -O(C 3 - 6 alkylene)- wherein the C 1 - 3 alkylene is optionally substituted with one, two, or three substituents selected a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 1 is -O(CH 2 )-, -O(C 2 alkylene)-, -O(C 3 alkylene)-, -O(C 4 alkylene)-, -O(C 5 alkylene)-, or -O(C6alkylene)- wherein the alkylene is unsubstituted.
- X 1 is -O(CH2)- wherein the CH2 group is unsubstituted.
- X 1 is -O(CH2)-, -O(C2alkylene)-, -O(C3alkylene)-, -O(C4alkylene)-, -O(C5alkylene)-, or -O(C6alkylene)- wherein the alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 1 is -NR 4 (C1-6alkylene)- wherein the C1-6alkylene is unsubstituted.
- X 1 is -NR 4 (C 1 - 6 alkylene)- wherein the C 1 - 6 alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 1 is -NR 4 (C1-3alkylene)- wherein the C1-3alkylene is unsubstituted.
- X 1 is -NR 4 (C 1 - 3 alkylene)- wherein the C 1 - 3 alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 1 is -O(C3-6alkylene)- wherein the C3-6alkylene is unsubstituted.
- X 1 is -NR 4 (C3-6alkylene)- wherein the C1-3alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 1 is -NR 4 (CH 2 )-, -NR 4 (C 2 alkylene)-, -NR 4 (C 3 alkylene)-, -NR 4 (C 4 alkylene)-, -NR 4 (C 5 alkylene)- , or -NR 4 (C6alkylene)- wherein the alkylene is unsubstituted.
- X 1 is -NR 4 (C 2 alkylene)- wherein the C 2 alkylene is unsubstituted.
- X 1 is 53
- X 1 is -NR 4 (CH 2 )- wherein the CH 2 group is unsubstituted. In certain embodiments, X 1 is - NH(CH2)- wherein the CH2 group is unsubstituted.
- X 1 is -NR 4 (CH2)-, -NR 4 (C2alkylene)-, -NR 4 (C3alkylene)-, -NR 4 (C4alkylene)-, - NR 4 (C5alkylene) -, or -NR 4 (C 6 alkylene)- wherein the alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- R 4 is independently hydrogen. In any of the foregoing embodiments, R 4 is independently C 1-6 alkyl. In any of the foregoing embodiments, R 4 is independently methyl. [000246] In certain embodiments, X 2 is absent. In certain embodiments, X 2 is unsubstituted -(C1-6alkylene)-.
- X 2 is -(C1-6alkylene)- wherein the C1- 6alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy e.g., X 2 is -(C1-3alkylene)- wherein the C1- 3 alkylene is unsubstituted.
- X 2 is -(C 1 - 3 alkylene)- wherein the C 1 - 3alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- X 2 is -(C3-6alkylene)- wherein the C3- 6 alkylene is unsubstituted.
- X 2 is -(C 3 - 6 alkylene)- wherein the C 1 - 3alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 2 is -(CH2)-, -(C2alkylene)-, -(C3alkylene)-, -(C4alkylene)-, -(C5alkylene)-, or -(C6alkylene)- wherein the alkylene is unsubstituted. In certain embodiments, X 2 is unsubstituted -(C 2 alkylene)-.
- X 2 is -(CH 2 )-, -(C 2 alkylene)-, -(C 3 alkylene)-, -(C 4 alkylene)-, -(C 5 alkylene)-, or -(C 6 alkylene)- wherein the alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 3 is absent. In certain embodiments, X 3 is -NR 4 -. In certain embodiments, X 3 is -NH-. In any of the foregoing embodiments, X 3 is -N(CH 3 )-. In certain embodiments, X 3 is -O-. [000250] In certain embodiments, X 1 , X 2 , and X 3 are absent. In certain embodiments, X 1 is absent. In certain embodiments, X 2 is absent. In certain embodiments, X 3 is absent. In certain embodiments, X 2 and X 3 are absent. In certain embodiments, X 1 and X 3 are absent. In certain embodiments, X 1 and X 2 are absent.
- Non-limiting examples of W 2 include , , , , , , , , , , and .
- W 2 is selected from , , , , , , , and . 1.3.2.3.
- Protease Cleavable Linker or pH-sensitive Linker (L 1 ) [000253]
- a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein the linker is cleavable.
- the linker is cleavable in vivo.
- the linker is cleavable by a protease.
- the linker is cleavable by a cathepsin. In certain embodiments, the linker is cleavable by cathepsin B. In certain embodiments, the linker is a pH-sensitive linker.
- L 1 comprises a compound of the formula: wherein R 5 is hydrogen, an electron donating group, or an electron withdrawing group; X 4 is -O- or -NR 6 -; X 5 is a linker; R 6 is hydrogen or an electron withdrawing group; and m is an integer selected from 1 to 4. [000255] In certain embodiments, X 5 includes an ester wherein the carbonyl carbon of the ester functional group is covalently bound to the fluorene.
- X 5 includes an amide wherein the carbonyl carbon of the amide functional group is covalently bound to the fluorene. In certain embodiments, X 5 includes a ketone wherein the carbonyl carbon of the ketone functional group is covalently bound to the fluorene. In certain embodiments, X 5 includes an anhydride wherein one of the carbonyls of the anhydride functional group is covalently bound to the fluorene. In certain embodiments, X 5 includes a sulfonyl wherein the sulfur of the sulfonyl functional group is covalently bound to the fluorene.
- X 5 includes an ammonium where the positively charged nitrogen of the ammonium functional group is covalently bound to the fluorene.
- X 5 is -C(O)-, -C(O)NR 4 , -C(O)-(C1-6alkylene)-, -C(O)-O-(C 1-6 alkylene)-, -C(O)-NR 4 -(C 1-6 alkylene)-, or -C(O)-S-(C 1-6 alkylene)- wherein the -C(O)- is bound to the fluorene and the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., flu
- X 5 is -C(O)-. In certain embodiments, X 5 is -C(O)NR 4 . In certain embodiments, X 5 is -C(O)NH-. In certain embodiments, X 5 is - C(O)N(CH3)-. 56
- X 5 is -C(O)-(C 1-6 alkylene)- wherein the C 1-6 alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 5 is-C(O)-(C 1-3 alkylene)- wherein the C 1- 6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 5 is-C(O)-(C3-6alkylene)- wherein the C 1-6 alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 5 is -C(O)-O-(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 5 is -C(O)-O-(C1-3alkylene)- wherein the C1- 6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 5 is -C(O)-O-(C 3-6 alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 5 is -C(O)-NR 4 (C 1-6 alkylene)- wherein the C 1-6 alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 5 is -C(O)-NR 4 (C1-3alkylene)- wherein the C 1-6 alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 5 is -C(O)-NR 4 (C3-6alkylene)- wherein the C 1-6 alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 5 is -C(O)-S-(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- X 5 is -C(O)-S-(C 1-3 alkylene)- wherein the C 1- 57
- 6 alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- a halogen e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)
- X 5 is -C(O)-S-(C3-6alkylene)- wherein the C 1-6 alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
- each R 5 is hydrogen, an electron donating group, or an electron withdrawing group.
- each R 5 is hydrogen.
- each R 5 is an electron donating group.
- each R 5 is an electron withdrawing group.
- each R 5 is independently selected from the group consisting of hydrogen, haloalkyl, halogen, -CN, -SO3H, -C(O)R 3 , -C(O)OR 3 , -OR 3 , -N(H)C(O)R 3 , -N(H)CO2R 3 , and -N(H)C(O)C(H)(R 3 )CO2H wherein each R 3 is independently alkyl, cycloalkyl, heteroalkyl, heterocycloalkyl, aryl, or heteroaryl.
- each R 5 is independently selected from the group consisting of haloalkyl, halogen, -CN, -SO 3 H, -C(O)R 3 , -C(O)OR 3 , -OR 3 , -N(H)C(O)R 3 , -N(H)CO2R 3 , and -N(H)C(O)C(H)(R 3 )CO2H wherein each R 3 is independently alkyl, cycloalkyl, heteroalkyl, heterocycloalkyl, aryl, or heteroaryl.
- each R 5 is independently selected from the group consisting of -H, -CF3, -Br, -Cl, -F, -CN, - SO 3 H, -C(O)Me, -CO2Me, -Ome, -N(H)C(O)Me, -N(H)CO2Me, and N(H)C(O)C(H)(Me)CO2H.
- each R 5 is independently selected from the group consisting of -CF 3 , - Br, -Cl, -F, -CN, -SO 3 H, -C(O)Me, -CO 2 Me, -OMe, -N(H)C(O)Me, -N(H)CO 2 Me, and N(H)C(O)C(H)(Me)CO2H.
- R 5 is hydrogen.
- R 5 is -Br.
- R 5 is -Cl.
- R 5 is -F.
- R 5 is -CN.
- R 5 is -SO 3 H.
- R 5 is -C(O)Me. In certain embodiments, R 5 is -OMe.
- X 4 is -O-. In certain embodiments, X 4 is -NR 6 - wherein R 6 is hydrogen or an electron withdrawing group. In certain embodiments, R 6 is hydrogen. In certain embodiments, R 6 is an electron withdrawing group. The electron withdrawing group can be any electron withdrawing group deemed suitable to the person of skill in the art. In certain embodiments, R 6 is independently selected from the group consisting of -C(O)R 3 , - 58
- R 3 is independently alkyl, cycloalkyl, heteroalkyl, heterocycloalkyl, aryl, or heteroaryl.
- R 6 is independently selected from the group consisting of -C(O)R 3 , C(O)OR 3 , and -S(O) 2 R 3 wherein each R 3 is independently alkyl, cycloalkyl, heteroalkyl, heterocycloalkyl, aryl, or heteroaryl.
- R 6 is independently selected from the group consisting of hydrogen, -CF 3 , -C(O)Me, -CO 2 Me, and -S(O) 2 CH 3 . In certain embodiments, R 6 is independently selected from the group consisting of -C(O)Me, and -S(O) 2 R 3 . In certain embodiments, R 6 is -C(O)Me. In certain embodiments, R 6 is -S(O) 2 CH 3 . [000264] In certain embodiments, m is 1. In certain embodiments, m is 2. In certain embodiments, m is 3. In certain embodiments, m is 4. [000265] Non-limiting examples of include: X 5 R 5 O , , , , , , , and . [000266] In certain embodiments, is . 59
- L 1 comprises a peptide. In certain embodiments, L 1 comprises a dipeptide. In certain embodiments, L 1 comprises a tripeptide or a tetrapeptide. In certain embodiments, L 1 comprises natural and non-natural amino acids.
- L 1 comprises at least one natural amino acid selected from alanine, ⁇ -alanine, arginine, asparagine, aspartic acid, cysteine, cystine, glutamic acid, glutamine, glycine, phenylalanine, histidine, isoleucine, lysine, leucine, methionine, proline, serine, threonine, valine, tryptophan, and tyrosine.
- L 1 comprises at least one non-natural amino acid selected from sulfoalanine, hydroxyproline (Hyp), beta-alanine, citrulline (Cit), ornithine (Orn), norleucine (Nle), 3-nitrotyrosine, nitroarginine, pyroglutamic acid (Pyr), naphtylalanine (Nal), 2,4-diaminobutyric acid (DAB), methionine sulfoxide, and methionine sulfone.
- non-natural amino acid selected from sulfoalanine, hydroxyproline (Hyp), beta-alanine, citrulline (Cit), ornithine (Orn), norleucine (Nle), 3-nitrotyrosine, nitroarginine, pyroglutamic acid (Pyr), naphtylalanine (Nal), 2,4-diaminobutyric acid (DAB), methionine sulfoxide, and
- L 1 comprises at least one natural amino acid selected from alanine, ⁇ -alanine, arginine, asparagine, aspartic acid, cysteine, cystine, glutamic acid, glutamine, glycine, phenylalanine, histidine, isoleucine, lysine, leucine, methionine, proline, serine, threonine, valine, tryptophan, and tyrosine and further comprises at least one non- natural amino acid from sulfoalanine, hydroxyproline (Hyp), beta-alanine, citrulline (Cit), ornithine (Orn), norleucine (Nle), 3-nitrotyrosine, nitroarginine, pyroglutamic acid (Pyr), naphtylalanine (Nal), 2,4-diaminobutyric acid (DAB), methionine sulfoxide, and methionine sulfone.
- alanine
- L 1 comprises a dipeptide selected from the group consisting of -Phe-Lys-, -Val-Ala-, -Val-Lys-, -Ala-Lys-, -Val-Cit-, -Phe-Cit-, -Leu-Cit-, -Ile- Cit-, -Phe-Arg-, and -Trp-Cit-.
- L 1 comprises -Val-Ala-.
- L 1 comprises -Val-Cit-.
- L 1 comprises a peptide-self immolative group selected from the group consisting of -Phe-Lys-PABC-, -Val-Ala-PABC-, -Val-Lys-PABC-, -Ala-Lys- PABC-, -Val-Cit-PABC-, -Phe-Cit-PABC-, -Leu-Cit-PABC-, -Ile-Cit-PABC-, -Phe-Arg- PABC-, -Trp-Cit-PABC-, and Val-Glu-PABC.
- L 1 comprises -Val- Ala-PABC-.
- L 1 comprises -Val-Cit-PABC-.
- the peptide and/or self immolative group can be in either orientation with respect to W 2 or SG 1 or (SG 2 ).
- PABC refers 60
- L 1 is -Val-Cit-.
- L 1 is -Val-Cit- PABC- wherein the -PABC- is covalently bound to W 2 and -Val- is covalently bound to SG 1 or SG 2 .
- L 1 is absent.
- Spacer Groups facilitate spacing of the conjugating group from the other groups of the compounds described herein. This spacing can lead to more efficient conjugation. The spacer group can also stabilize the conjugating group and lead to improved overall IFN ⁇ conjugate properties.
- a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein, SG 1 is any divalent spacer group.
- SG 1 is , , , , , , , , , or ; wherein a and c are an integer independently selected from 0, 1, 2, 3, 4, 5, and 6; and b is an integer selected from 1, 2, 3, 4, 5, and 6.
- SG 1 is , , , , , , , or ; wherein a is an integer selected from 0, 1, 2, 3, 4, 5, and 6; and b is an integer selected from 1, 2, 3, 4, 5, and 6.
- Non-limiting examples of SG 1 include , , , , , and .
- provided herein is a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein, SG 2 is any trivalent spacer group.
- SG 2 is or ; wherein a is independently an integer selected from 0, 1, 2, 3, 4, 5, and 6. [000280] In certain embodiments, SG 2 is . [000281] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W 1 is -C(O)CH2CH2NH-, W 2 is , and RG, L 1 , and 62
- W 1 is -C(O)CH 2 CH 2 NH-, W 2 is , and RG, L 1 , and SG 1 are as defined herein.
- W 1 is -C(O)CH2CH2NH-, W 2 is , RG is or , and SG 1 is or , and L 1 is as defined herein.
- W 1 is -C(O)CH2CH2NH-, W 2 is , RG is or , L 1 is a pH-sensitive linker, and SG 1 is as defined herein.
- W 1 is -C(O)CH2CH2NH-, W 2 is , RG is or , L 1 is , and SG 1 is as defined herein.
- W 1 is -C(O)CH2CH2NH-, W 2 is , RG is or , L 1 is , and SG 1 is as defined herein.
- W 1 is -C(O)CH 2 CH 2 NH-, W 2 is , RG is or , L 1 is , and SG 1 is .
- W 1 is -C(O)CH2CH2NH-, W 2 is , RG is or , L 1 is , and SG 1 is .
- a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W 1 is -C(O)CH 2 CH 2 NH-, W 2 is , and RG, L 1 , and SG 1 are as defined herein.
- W 1 is -C(O)CH2CH2NH-, W 2 is , and RG, L 1 , and SG 1 are as defined herein. In certain embodiments, W 1 is -C(O)CH2CH2NH-, W 2 is selected from , , and , and RG, L 1 , and SG 1 are as defined herein. In certain embodiments, W 1 is -C(O)CH 2 CH 2 NH-, W 2 is selected from 64
- W 1 is -C(O)CH2CH2NH-
- W 2 is selected from , , and , RG is or , L 1 is a cathepsin B cleavable linker, and SG 1 are as defined herein.
- W 1 is -C(O)CH 2 CH 2 NH-
- W 2 is selected from , , and , RG is or , L 1 is comprises a dipeptide, and SG 1 are as defined herein.
- W 1 is -C(O)CH 2 CH 2 NH-
- W 2 is selected from , , and , RG is or , L 1 is comprises -Val-Cit-, and SG 1 is as defined herein.
- W 1 is -C(O)CH 2 CH 2 NH-
- W 2 is selected from 65
- RG is or , L 1 comprises a peptide-self immolative group, and SG 1 is .
- W 1 is -C(O)(CH2CH2)NH-
- W 2 is selected from , , and , RG is or , L 1 is -Val-Cit-PABC-, and SG 1 is .
- a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W 1 is -C(O)CH 2 CH 2 CH 2 CH 2 NH-, W 2 is absent, and RG, L 1 , and SG 1 are as defined herein.
- W 1 is -C(O)CH2CH2CH2CH2NH-, W 2 is absent, RG is or , L 1 is a protease cleavable linker, and SG 1 is as defined herein.
- W 1 is -C(O)CH 2 CH 2 CH 2 CH 2 NH-, W 2 is absent, RG is or , L 1 is a cathepsin B cleavable linker, and SG 1 is as defined herein. In certain embodiments, W 1 is -C(O)CH 2 CH 2 CH 2 CH 2 NH-, W 2 is absent, RG 66
- W 1 is -C(O)CH2CH2CH2CH2NH-
- W 2 is absent
- RG is or
- L 1 is comprises -Val-Cit-
- SG 1 is as defined herein.
- W 1 is -C(O)CH2CH2CH2CH2NH-
- W 2 is absent
- RG is or
- L 1 comprises a peptide-self immolative group
- SG 1 is .
- W 1 is -C(O)CH 2 CH 2 CH 2 CH 2 NH-
- W 2 is absent, RG is or
- L 1 is -Val-Cit-PABC-
- SG 1 is .
- a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W 1 is -C(O)(C 1 - 6 alkylene)NH-, W 2 is , and RG, L 1 , and SG 1 are as defined herein.
- W 1 is -C(O)(C 1 - 6 alkylene)NH- and W 2 is . In certain embodiments, W 1 is -C(O)CH2CH2NH- and W 2 is . In certain embodiments, W 1 is -C(O)CH 2 CH 2 NH- and W 2 is . In certain 67
- W 1 is -C(O)CH 2 CH 2 NH- and W 2 is . In certain embodiments, W 1 is -C(O)CH 2 CH 2 NH- and W 2 is . In certain embodiments, W 1 is -C(O)CH2CH2NH- and W 2 is . In certain embodiments, W 1 is -C(O)CH 2 CH 2 NH- and W 2 is . In certain embodiments, W 1 is -C(O)(CH 2 CH 2 )NH-, W 2 is , RG is or , SG is , and L 1 is absent. In certain embodiments, W 1 is -C(O)CH 2 CH 2 NH-, W 2 is , RG is or , SG is . and L 1 is absent. In certain embodiments, W 1 is -C(O)CH2CH2NH-, W 2 is , RG is or , SG is , and L 1 is absent. In certain embodiments, W 1 is -C(O)CH2CH2NH
- W 1 is -C(O)CH 2 CH 2 NH-
- W 2 is , RG is or , SG is , and L 1 is absent.
- L1 is a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W 1 is -C(O)(C 1 - 6 alkylene)-, W 2 is , and RG, L 1 , and SG 1 are as defined herein.
- W 1 is -C(O)CH2CH2CH2CH2-, W 2 is , and RG, L 1 , and SG 1 are as defined herein.
- W 1 O N 1-6 is -C(O)CH 2 CH 2 CH 2 CH 2 -, W 2 is R4 , and RG, L 1 , and SG 1 are as defined herein.
- W 1 is -C(O)CH 2 CH 2 CH 2 CH 2 -, W 2 is , and RG, L 1 , and SG 1 are as defined herein.
- W 1 is -C(O)CH2CH2CH2CH2-, W 2 is , RG is or , and SG 1 is , and L 1 is as defined herein. In certain embodiments, W 1 is -C(O)CH2CH2CH2CH2-, W 2 is , RG is or , L 1 is absent, and SG 1 is . In certain 69
- W 1 is -C(O)CH 2 CH 2 CH 2 CH 2 -, W 2 is , RG is or , L 1 is absent, and SG 1 is .
- a conjugate comprising a linker according to Formula (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W 1 is -C(O)(C 1 - 6 alkylene)-, W 2 is , and RG, L 1 , and SG 2 are as defined herein.
- W 1 is -C(O)(C1-6alkylene)-, W 2 is , and RG, L 1 , and SG 2 are as defined herein.
- W 1 is -C(O)(C 1 - 6 alkylene)-, W 2 is , and RG, L 1 , and SG 2 are as defined herein.
- W 1 is -C(O)(C1-6alkylene)-, W 2 is , and RG is or , L 1 , and SG 2 is as defined herein.
- W 1 is -C(O)(C1-6alkylene)-, W 2 is , and RG is or 70
- W 1 is -C(O)(C1-6alkylene)-, W 2 is , and RG is or , L 1 is absent, and SG 2 is .
- W 1 is -C(O)(C 1 - 6 alkylene)-, W 2 is , and RG is or , L 1 is absent, and SG 2 is .
- 1.3.3. Conjugating Groups and Residues Thereof [000287]
- a conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof according to any of the following formulas: 71
- MM is a polymer (POLY), for example a water-soluble polymer.
- POLY is , wherein n1 is an integer from 1 to 10,000, inclusive, and represents attachment to the remainder of the compound or conjugate. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 5000. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 2500. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 1500. In certain embodiments, including any of the foregoing, n1 is an integer between 100 to 1000.
- n1 is an integer from 100 to 700. In certain embodiments, n1 is an integer from 300 to 700. In certain embodiments, n1 is an integer from 400 to 500. In certain embodiments, n1 is an integer from 600 to 700. In certain embodiments, including any of the foregoing, n1 is an integer between 100 to 500. [000289]
- the present disclosure encompasses each and every regioisomer of the conjugate structures depicted below: 75
- n1 is an integer between 300 and 800, inclusive. In any of the foregoing embodiments, n1 is an integer between 400 and 600, inclusive. In any of the foregoing embodiments, n1 is an integer between 400 and 500, inclusive. [000291] In any of the foregoing embodiments, n is an integer from 1 to 8. In any of the foregoing embodiments, n is 1. In any of the foregoing embodiments, n is 2. In any of the 77
- n is 3. In any of the foregoing embodiments, n is 4. In any of the foregoing embodiments, n is 6. In any of the foregoing embodiments, n is 8. [000292] The present disclosure encompasses the conjugate structures depicted below: 78
- the bracketed structure can be covalently bonded to one or more non-natural amino acids or modified amino acids of the IFN ⁇ polypeptide, wherein the one or more non-natural amino acids or modified amino acids are located at sites selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156 of SEQ ID NO: 33.
- the one or more non-natural amino acid or modified amino acid is two non-natural amino acids or modified amino acids, respectively. In the foregoing embodiment, the one or more non-natural amino acid or modified amino acid is three non-natural amino acids or modified amino acids, respectively. In the foregoing embodiment, the one or more non-natural amino acid or modified amino acid is four non-natural amino acids or modified amino acids, respectively. In the foregoing embodiment, the one or more non-natural amino acid or modified amino acid is five or more non-natural amino acids or modified amino acids, respectively.
- the bracketed structure can be covalently bonded to one or more non-natural amino acids or modified amino acids of the IFN ⁇ polypeptide, wherein the one or more non-natural amino acids or modified amino acids are located at sites selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33.
- the bracketed structure can be covalently bonded to two non-natural amino acids or modified amino acids at sites selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33.
- the bracketed structure can be covalently bonded to three non- natural amino acids or modified amino acids at sites selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33.
- the bracketed structure can be covalently bonded to four non-natural amino acids or modified amino acids at sites selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33.
- the bracketed structure can be covalently bonded to five or more non-natural amino acids or modified amino acids at sites selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33.
- the conjugate comprises a non-natural amino acid or modified amino acid located at amino acid position H7.
- the conjugate comprises a non-natural amino acid or modified amino acid located at amino acid position Q40.
- the conjugate comprises a non-natural amino acid or modified amino 83
- the conjugate comprises a non-natural amino acid or modified amino acid located at amino acid position N45. In certain embodiments, the conjugate comprises a non-natural amino acid or modified amino acid located at amino acid position E51. In certain embodiments, the conjugate comprises a non- natural amino acid or modified amino acid located at amino acid position N156. [000296] In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at H7 and Q40.
- the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at H7 and Q40 and further comprising at least one additional non-natural amino acid or modified amino acid at a position selected from E51, N45, and N156.
- the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at H7, Q40 and E51.
- the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at H7, Q40 and N156.
- the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at H7, Q40, N45, and N156.
- the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at H7 and E51. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at H7, E51 and N156. [000297] In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at Q40 and E51. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at Q40 and N156. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at E51 and N156.
- the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at Q40, E51, and N156. [000298] In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at Q40 and N156. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids at Q40 and N156 and further comprising at least one non-natural amino acid or modified amino acid at a position selected from H7 and E51.
- the conjugate comprises an IFN ⁇ polypeptide comprising at least one non-natural amino acid or modified amino acid in a position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprising at least one non- 84
- natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136.
- the conjugate comprises an IFN ⁇ polypeptide comprising at least two non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprising at least one non- natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136.
- the conjugate comprises an IFN ⁇ polypeptide comprising at least three non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprising at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136.
- the conjugate comprises an IFN ⁇ polypeptide comprising non-natural amino acids or modified amino acids in positions Q40 and N156 and further comprising at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136.
- the non-natural or modified amino acid is a non-natural amino acid.
- the non-natural or modified amino acid is a modified amino acid.
- one or more linkers and/or masking moieties are conjugated to the one or more non-natural amino acids or modified amino acids.
- the non-natural amino acid residue or modified amino acid residue comprises a residue of a moiety selected from the group consisting of amino, carboxy, acetyl, hydrazino, hydrazido, semicarbazido, sulfanyl, azido and alkynyl.
- the modified amino acid residue is selected from the group consisting of: p-acetyl-L-phenylalanine, O- methyl-L-tyrosine, 3-methyl-phenylalanine, O-4-allyl-L-tyrosine, 4-propyl-L-tyrosine, fluorinated phenylalanine, isopropyl-L-phenylalanine, p-azido-L-phenylalanine, p-acetyl-L- phenylalanine, p-benzoyl-L-phenylalanine, p-iodo-phenylalanine, p-bromophenylalanine, p- amino-L-phenylalanine, isopropyl-L-phenylalanine, p-propargyloxy-phenylalanine, and p- azidomethyl-L-phenylalanine residues.
- the modified amino acid 85 is selected from the group consisting of:
- the modified amino acid is para-azidomethyl-L-phenylalanine (pAMF).
- the modified amino acid is para-azidomethyl-L-phenylalanine (pAMF) and is located at an amino acid position selected from amino acid positions: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156 of SEQ ID NO: 33.
- the modified amino acid is para- azidomethyl-L-phenylalanine (pAMF) and is located at an amino acid position selected from amino acid positions: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33.
- pAMF para- azidomethyl-L-phenylalanine
- the conjugate comprises an IFN ⁇ polypeptide comprising a p-azidomethyl-L-phenylalanine residue in at least one amino acid position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
- the conjugate comprises an IFN ⁇ polypeptide comprising a p-azidomethyl-L-phenylalanine residue in at least one amino acid position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising a p-azidomethyl-L-phenylalanine residue in at least two amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156.
- the conjugate comprises an IFN ⁇ polypeptide comprising a p- azidomethyl-L-phenylalanine residue in at least three amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising a p-azidomethyl-L-phenylalanine residue in at least four amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156.
- the conjugate comprises an IFN ⁇ polypeptide comprising a p- azidomethyl-L-phenylalanine residue in at least five amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising a p-azidomethyl-L-phenylalanine residue in positions H7, Q40, E41, N45, E51, and N156. [000307] In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising a p-azidomethyl-L-phenylalanine residue at H7.
- the conjugate comprises an IFN ⁇ polypeptide comprising a p-azidomethyl-L-phenylalanine residue at Q40. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising a p-azidomethyl-L- phenylalanine residue at E41. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising a p-azidomethyl-L-phenylalanine residue at N45. In some 86
- the conjugate comprises an IFN ⁇ polypeptide comprising p-azidomethyl-L- phenylalanine residue at E51. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising a p-azidomethyl-L-phenylalanine residue at N156. [000308] In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising p- azidomethyl-L-phenylalanine residues at H7 and Q40.
- the conjugate comprises an IFN ⁇ polypeptide comprising p-azidomethyl-L-phenylalanine residues at H7 and Q40 and further comprising at least one additional p-azidomethyl-L-phenylalanine residue at a position selected from E51, N45, and N156.
- the conjugate comprises an IFN ⁇ polypeptide comprising p-azidomethyl-L-phenylalanine residues at H7, Q40 and E51.
- the conjugate comprises an IFN ⁇ polypeptide comprising p- azidomethyl-L-phenylalanine residues at H7, Q40 and N156.
- the conjugate comprises an IFN ⁇ polypeptide comprising p-azidomethyl-L-phenylalanine residues at H7, Q40, N45, and N156. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising p-azidomethyl-L-phenylalanine residues at H7 and E51. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising p-azidomethyl-L- phenylalanine residues at H7, E51 and N156. [000309] In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising p- azidomethyl-L-phenylalanine residues at Q40 and E51.
- the conjugate comprises an IFN ⁇ polypeptide comprising p-azidomethyl-L-phenylalanine residues at Q40 and N156. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising p- azidomethyl-L-phenylalanine residues at E51 and N156. In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising p-azidomethyl-L-phenylalanine residues at Q40, E51, and N156. [000310] In some embodiments, the conjugate comprises an IFN ⁇ polypeptide comprising p- azidomethyl-L-phenylalanine residues at Q40 and N156.
- the conjugate comprises an IFN ⁇ polypeptide comprising p-azidomethyl-L-phenylalanine residues at Q40 and N156 and further comprising at least one p-azidomethyl-L-phenylalanine residues at a position selected from H7 and E51.
- the masking moiety is a water soluble polymer (POLY) selected from the group consisting of is polyethylene glycol (PEG), poly(propylene glycol) (PPG), copolymers of ethylene glycol and propylene glycol, poly(oxyethylated polyol), poly(olefinic alcohol), poly(vinylpyrrolidone), poly(hydroxyalkylmethacrylamide), poly(hydroxyalkylmethacrylate), poly(saccharides), poly(a-hydroxy acid), poly(vinyl alcohol), polyphosphazene, polyoxazolines (POZ), poly(N-acryloylmorpholine), and combinations 87
- POLY water soluble polymer
- the water soluble polymer is PEG.
- the PEG has an average molecular weight of between about 5KDa and about 50 KDa.
- the PEG is selected from the group consisting of a linear or branched PEG molecule having an average molecular weight of 10Kda, 20Kda, 30Kda, or 40Kda.
- the PEG has an average molecular weight of 30Kda.
- the PEG has an average molecular weight of 40Kda.
- the conjugate has an extended half-life compared to an identical polypeptide lacking the water-soluble polymer.
- IFN ⁇ polypeptides comprising one or more non-natural amino acids or modified amino acids.
- These non-natural amino acids or modified amino acids can facilitate conjugation to a masking moiety or linker to form conjugates.
- the non-natural amino acid or modified amino acid is at a position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
- the non-natural amino acid or modified amino acid is at a position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156.
- a polynucleotide is provided, encoding one of these IFN ⁇ polypeptides.
- the polynucleotide encodes a TAG codon to facilitate incorporation of a non-natural amino acid or modified amino acid according to the expression techniques described herein. Any non-natural amino acid or modified amino acid can be incorporated at the TAG position.
- the non-natural amino acid or modified amino acid is one described herein.
- the non-natural or modified amino acid is a modified amino acid and the modified amino acid is p- azidomethylphenylalanine.
- the IFN ⁇ polypeptide has an amino acid sequence according to SEQ ID NO: 1, SEQ ID NO: 2, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22, or SEQ ID NO: 23, SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 29, SEQ ID NO: 30, SEQ ID NO: 31, SEQ ID NO: 35, SEQ ID NO: 36, and SEQ ID NO: 37.
- the IFN ⁇ polypeptide has an amino acid sequence selected from SEQ ID NO: 11, SEQ ID NO: 13, SEQ ID NO: 16, and SEQ ID NO: 18.
- the IFN ⁇ polypeptide conjugate comprises a PEG having an average molecular weight of 20Kda, 30Kda or 40Kda.
- the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 11. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 11. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 11. [000316] In certain embodiments, the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 16. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 16. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 16.
- the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 18. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 18. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 18. [000318] In certain embodiments, the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 33. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 33. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 33.
- the TAG (*) position of the above amino acid sequences indicates a non-natural amino acid or modified amino acid.
- the non- natural amino acid or modified amino acid is one described herein.
- the non-natural or modified amino acid is a modified amino acid and the modified amino acid is p-azidomethylphenylalanine.
- Vectors, Host Cells, and Recombinant Methods [000320] Also provided are isolated nucleic acids encoding IFN ⁇ polypeptide, vectors and host cells comprising the nucleic acids, and recombinant techniques for the production of the IFN ⁇ polypeptide and cytokines.
- the nucleic acid encoding it may be isolated and inserted into a replicable vector for further cloning (i.e., amplification of the DNA) or expression.
- the nucleic acid may be produced by homologous recombination, for example as described in U.S. Patent No.5,204,244.
- Many different vectors are known in the art.
- the vector components generally include, but are not limited to, one or more of the following: a signal sequence, an origin of replication, one or more marker genes, an enhancer element, a promoter, and a transcription termination sequence, for example as described in U.S. Patent No.5,534,615. 89
- Suitable host cells include any prokaryotic (e.g., bacterial), lower eukaryotic (e.g., yeast), or higher eukaryotic (e.g., mammalian) cells.
- Suitable prokaryotes include eubacteria, such as Gram-negative or Gram-positive organisms, for example, Enterobacteriaceae such as Escherichia (E. coli), Enterobacter, Erwinia, Klebsiella, Proteus, Salmonella (S. typhimurium), Serratia (S. marcescans), Shigella, Bacilli (B.
- E. coli 294 One useful E. coli cloning host is E. coli 294, although other strains such as E. coli B, E. coli X1776, and E. coli W3110 are suitable.
- eukaryotic microbes such as filamentous fungi or yeast are also suitable cloning or expression hosts for IFN ⁇ polypeptide-encoding vectors. Saccharomyces cerevisiae, or common baker’s yeast, is a commonly used lower eukaryotic host microorganism.
- Schizosaccharomyces pombe Kluyveromyces (K. lactis, K. fragilis, K. bulgaricus K. wickeramii, K. waltii, K. drosophilarum, K. thermotolerans, and K. marxianus), Yarrowia, Pichia pastoris, Candida (C. albicans), Trichoderma reesia, Neurospora crassa, Schwanniomyces (S. occidentalis), and filamentous fungi such as, for example Penicillium, Tolypocladium, and Aspergillus (A. nidulans and A.
- Useful mammalian host cells include COS-7 cells, HEK293 cells; baby hamster kidney (BHK) cells; Chinese hamster ovary (CHO); mouse cells; African green monkey kidney cells (VERO-76), and the like.
- the host cells used to produce the IFN ⁇ polypeptides may be cultured in a variety of media.
- Commercially available media such as, for example, Ham’s F10, Minimal Essential Medium (MEM), RPMI-1640, and Dulbecco’s Modified Eagle’s Medium (DMEM) are suitable for culturing the host cells.
- any of these media may be supplemented as necessary with hormones and/or other growth factors (such as insulin, transferrin, or epidermal growth factor), salts (such as sodium chloride, calcium, magnesium, and phosphate), buffers (such as HEPES), nucleotides (such as adenosine and thymidine), antibiotics, trace elements (defined as inorganic compounds usually present at final concentrations in the micromolar range), and glucose or an equivalent energy 90
- growth factors such as insulin, transferrin, or epidermal growth factor
- salts such as sodium chloride, calcium, magnesium, and phosphate
- buffers such as HEPES
- nucleotides such as adenosine and thymidine
- antibiotics such as adenosine and thymidine
- trace elements defined as inorganic compounds usually present at final concentrations in the micromolar range
- a IFN ⁇ polypeptide is produced by a method comprising the step of culturing a host cell described herein.
- the host cell comprises a nucleic acid, vector, or expression vector described herein for producing the IFN ⁇ polypeptide.
- the IFN ⁇ polypeptide variant comprises one or more non- natural amino acids or modified amino acids as described herein.
- the host cell further comprises a nucleic acid, vector, or expression vector encoding an aminoacyl tRNA synthetase (RS) specific for the non-natural amino acid or modified amino acid.
- the host cell further comprises a nucleic acid, vector, or expression vector encoding a tRNA specific for the non-natural amino acid or modified amino acid.
- any or each nucleic acid, vector, or expression vector is codon optimized for the host cell.
- the non-natural or modified amino acid is a modified amino acid and the modified amino acid is p-azidomethylphenylalanine.
- the host cell is E. coli. In certain embodiments, the host cells, for instance E.
- the IFN ⁇ polypeptides can be produced intracellularly, in the periplasmic space, or directly secreted into the medium. If the IFN ⁇ polypeptide is produced intracellularly, as a first step, the particulate debris, either host cells or lysed fragments, is removed, for example, by centrifugation or ultrafiltration. For example, Carter et al. (Bio/Technology, 1992, 10:163-167) describes a procedure for isolating polypeptides which are secreted to the periplasmic space of E. coli.
- the IFN ⁇ polypeptide is produced in a cell-free system.
- the cell-free system is an in vitro transcription and translation system as described in Yin et al., mAbs, 2012, 4:217-225, incorporated by reference in its entirety.
- the cell-free system utilizes a cell-free extract from a eukaryotic cell or from a prokaryotic cell.
- the prokaryotic cell is E. coli.
- Cell-free expression of the IFN ⁇ polypeptide may be useful, for example, where the IFN ⁇ polypeptide accumulates in a cell as an insoluble aggregate, or where yields from periplasmic expression are low.
- IFN ⁇ polypeptide is secreted into the medium
- supernatants from such expression systems are generally first concentrated using a commercially available protein concentration filter, for example, an Amicon ® or Millipore ® Pellcon ® ultrafiltration unit.
- a protease inhibitor such as PMSF may be included in any of the foregoing steps to inhibit proteolysis and antibiotics may be included to prevent the growth of adventitious contaminants.
- the IFN ⁇ polypeptide composition prepared from the cells can be purified using, for example, hydroxyapatite chromatography, gel electrophoresis, dialysis, and affinity chromatography, with affinity chromatography being a particularly useful purification technique.
- the matrix to which the affinity ligand is attached is most often agarose, but other matrices are available.
- Mechanically stable matrices such as controlled pore glass or poly(styrenedivinyl)benzene allow for faster flow rates and shorter processing times than can be achieved with agarose.
- Other techniques for protein purification such as fractionation on an ion-exchange column, ethanol precipitation, Reverse Phase HPLC, chromatography on silica, chromatography on heparin Sepharose ® , chromatofocusing, SDS-PAGE, and ammonium sulfate precipitation are also available, and can be applied by one of skill in the art.
- the mixture comprising the IFN ⁇ polypeptide of interest and contaminants may be subjected to low pH hydrophobic interaction chromatography using an elution buffer at a pH between about 2.5-4.5, generally performed at low salt concentrations (e.g., from about 0-0.25 M salt).
- the IFN ⁇ polypeptide is conjugated, for instance as described below. 1.5. Conjugation [000339]
- the conjugates can be prepared by standard techniques.
- an IFN ⁇ is contacted with a masking moiety or linker precursor under conditions suitable for forming a bond from the IFN ⁇ polypeptide to the masking moiety to form an IFN ⁇ -masking moiety conjugate.
- an IFN ⁇ polypeptide is contacted with a linker precursor under conditions suitable for forming a bond from the IFN ⁇ polypeptide to the linker.
- the resulting IFN ⁇ polypeptide-linker is contacted with a masking moiety precursor under conditions suitable for forming a bond from the IFN ⁇ -linker to the masking moiety to form an IFN ⁇ -linker-masking moiety conjugate.
- a masking moiety precursor is contacted with a linker precursor under conditions suitable for forming a bond from the 92
- an IFN ⁇ conjugate is prepared by contacting an IFN ⁇ as disclosed herein with a linker precursor having a structure selected from: , , , 93
- n1 is an integer between 300 and 800, inclusive. In any of the foregoing embodiments, n1 is an integer between 400 and 600, inclusive. In any of the foregoing embodiments, n1 is an integer between 400 and 500, inclusive.
- an IFN ⁇ conjugate is prepared by contacting an IFN ⁇ as disclosed herein with a linker precursor having a structure selected from: 94
- the IFN ⁇ polypeptides or conjugates provided herein can be formulated into pharmaceutical compositions using methods available in the art and those disclosed herein. Any of the IFN ⁇ polypeptides or conjugates provided herein can be provided in the appropriate pharmaceutical composition and be administered by a suitable route of administration. [000344]
- the methods provided herein encompass administering pharmaceutical compositions comprising at least one IFN ⁇ polypeptides or conjugates provided herein and one or more compatible and pharmaceutically acceptable carriers.
- pharmaceutically acceptable means approved by a regulatory agency of the Federal or a state government or listed in the U.S. Pharmacopeia or other generally recognized pharmacopeia for 96
- carrier includes a diluent, excipient, or vehicle with which the therapeutic is administered.
- Such pharmaceutical carriers can be sterile liquids, such as water and oils, including those of petroleum, animal, vegetable or synthetic origin, such as peanut oil, soybean oil, mineral oil, sesame oil and the like. Water can be used as a carrier when the pharmaceutical composition is administered intravenously. Saline solutions and aqueous dextrose and glycerol solutions can also be employed as liquid carriers, particularly for injectable solutions. Examples of suitable pharmaceutical carriers are described in Martin, E.W., Remington’s Pharmaceutical Sciences.
- compositions or IFN ⁇ polypeptide or conjugate provided herein may be administered by any route known in the art.
- a pharmaceutical composition or IFN ⁇ polypeptide or conjugate provided herein is administered parenterally.
- the compositions for parenteral administration can be emulsions or sterile solutions.
- Parenteral compositions may include, for example, propylene glycol, polyethylene glycol, vegetable oils, and injectable organic esters (e.g., ethyl oleate). These compositions can also contain wetting, isotonizing, emulsifying, dispersing and stabilizing agents.
- a composition provided herein is a pharmaceutical composition or a single unit dosage form.
- Pharmaceutical compositions and single unit dosage forms provided herein comprise a prophylactically or therapeutically effective amount of one or more prophylactic or therapeutic IFN ⁇ polypeptide or conjugate.
- Typical pharmaceutical compositions and dosage forms comprise one or more excipients.
- Suitable excipients are well-known to those skilled in the art of pharmacy, and non- limiting examples of suitable excipients include starch, glucose, lactose, sucrose, gelatin, malt, rice, flour, chalk, silica gel, sodium stearate, glycerol monostearate, talc, sodium chloride, dried skim milk, glycerol, propylene, glycol, water, ethanol and the like. Whether a particular excipient is suitable for incorporation into a pharmaceutical composition or dosage form depends on a variety of factors well known in the art including, but not limited to, the way in which the dosage form will be administered to a subject and the specific IFN ⁇ polypeptide or conjugate in the dosage form.
- the composition or single unit dosage form if desired, can also contain minor amounts of wetting or emulsifying agents, or pH buffering agents. 97
- Lactose free compositions can comprise excipients that are well known in the art and are listed, for example, in the U.S. Pharmocopeia (USP) SP (XXI)/NF (XVI).
- lactose free compositions comprise an active ingredient, a binder/filler, and a lubricant in pharmaceutically compatible and pharmaceutically acceptable amounts.
- Exemplary lactose free dosage forms comprise an active ingredient, microcrystalline cellulose, pre gelatinized starch, and magnesium stearate.
- Components of the pharmaceutical composition can be supplied either separately or mixed together in unit dosage form, for example, as a dry lyophilized powder or water free concentrate.
- the composition is to be administered by infusion, it can be dispensed with an infusion bottle containing sterile pharmaceutical grade water or saline. Where the composition is administered by injection, an ample of sterile water for injection or saline can be provided so that the ingredients may be mixed prior to administration.
- the pharmaceutical composition is supplied as a dry sterilized lyophilized powder that is capable of being reconstituted to the appropriate concentration for administration to a subject.
- IFN ⁇ polypeptides or conjugates are supplied as a water free concentrate.
- the pharmaceutical composition is supplied in liquid form.
- the pharmaceutical composition is provided in liquid form and is substantially free of surfactants and/or inorganic salts.
- the pharmaceutical composition is formulated as a salt form.
- Pharmaceutically acceptable salts include those formed with anions such as those derived from hydrochloric, phosphoric, acetic, oxalic, tartaric acids, etc., and those formed with cations such as those derived from sodium, potassium, ammonium, calcium, ferric hydroxides, isopropylamine, triethylamine, 2-ethylamino ethanol, histidine, procaine, etc.
- anhydrous pharmaceutical compositions and dosage forms comprising an IFN ⁇ polypeptide or conjugate, since water can facilitate the degradation of some IFN ⁇ polypeptide or conjugate.
- Anhydrous pharmaceutical compositions and dosage forms provided herein can be prepared using anhydrous or low moisture containing ingredients and low moisture or low humidity conditions.
- Pharmaceutical compositions and dosage forms that comprise lactose and at least one active ingredient that comprises a primary or secondary amine can be anhydrous if substantial contact with moisture and/or humidity during manufacturing, packaging, and/or storage is expected.
- anhydrous pharmaceutical composition should be prepared and stored such that its anhydrous nature is maintained. Accordingly, anhydrous compositions can be packaged using materials known to prevent exposure to water such that they can be included in suitable formulary kits. Examples of suitable packaging include, but are not limited to, hermetically sealed foils, plastics, unit dose containers (e.g., vials), blister packs, and strip packs. [000357] Further provided are pharmaceutical compositions and dosage forms that comprise one or more excipients that reduce the rate by which an IFN ⁇ polypeptide or conjugate will decompose. Such excipients, which are referred to herein as “stabilizers,” include, but are not limited to, antioxidants such as ascorbic acid, pH buffers, or salt buffers.
- parenteral dosage forms can be administered to subjects by various routes including, but not limited to, subcutaneous, intravenous (including bolus injection), intramuscular, and intraarterial. Because their administration typically bypasses subjects’ natural defenses against contaminants, parenteral dosage forms are typically, sterile or capable of being sterilized prior to administration to a subject. Examples of parenteral dosage forms include, but are not limited to, solutions ready for injection, dry products ready to be dissolved or suspended in a pharmaceutically acceptable vehicle for injection, suspensions ready for injection, and emulsions.
- Suitable vehicles that can be used to provide parenteral dosage forms are well known to those skilled in the art. Examples include, but are not limited to: Water for Injection USP; aqueous vehicles such as, but not limited to, Sodium Chloride Injection, Ringer’s Injection, Dextrose Injection, Dextrose and Sodium Chloride Injection, and Lactated Ringer’s Injection; water miscible vehicles such as, but not limited to, ethyl alcohol, polyethylene glycol, and polypropylene glycol; and non-aqueous vehicles such as, but not limited to, corn oil, cottonseed oil, peanut oil, sesame oil, ethyl oleate, isopropyl myristate, and benzyl benzoate.
- aqueous vehicles such as, but not limited to, Sodium Chloride Injection, Ringer’s Injection, Dextrose Injection, Dextrose and Sodium Chloride Injection, and Lactated Ringer’s Injection
- the amount of the IFN ⁇ polypeptide or conjugate or composition which will be effective in the prevention or treatment of a disorder or one or more symptoms thereof will vary with the nature and severity of the disease or condition, and the route by which the IFN ⁇ polypeptide or conjugate is administered.
- the frequency and dosage will also vary according to factors specific for each subject depending on the specific therapy (e.g., therapeutic or prophylactic agents) administered, the severity of the disorder, disease, or condition, the route of administration, as well as age, body, weight, response, and the past medical history of the subject.
- Effective doses may be extrapolated from dose-response curves derived from in vitro or animal model test systems.
- the dose can be administered according to a suitable schedule, for example, once, two times, three times, or for times weekly. It may be necessary to use dosages of the IFN ⁇ polypeptide or conjugate outside the ranges disclosed herein in some cases, as will be apparent to those of ordinary skill in the art. Furthermore, it is noted that the clinician or treating physician will know how and when to interrupt, adjust, or terminate therapy in conjunction with subject response. [000365] Different therapeutically effective amounts may be applicable for different diseases and conditions, as will be readily known by those of ordinary skill in the art.
- amounts sufficient to prevent, manage, treat or ameliorate such disorders, but insufficient to cause, or sufficient to reduce, adverse effects associated with the IFN ⁇ polypeptide or conjugate provided herein are also encompassed by the herein described dosage amounts and dose frequency schedules.
- the dosage administered to the subject may be increased to improve the prophylactic or therapeutic effect of the composition or it may be decreased to reduce one or more side effects that a particular subject is experiencing.
- treatment or prevention can be initiated with one or more loading doses of an IFN ⁇ polypeptide or conjugate or composition provided herein followed by one or more maintenance doses.
- a dose of an IFN ⁇ polypeptide or conjugate or composition provided herein can be administered to achieve a steady-state concentration of the IFN ⁇ polypeptide or conjugate in blood or serum of the subject.
- the steady-state concentration can be determined by measurement according to techniques available to those of skill or can be based on the physical characteristics of the subject such as height, weight and age.
- IFN ⁇ polypeptides or conjugates disclosed herein are administered to a mammal, generally a human, in a pharmaceutically acceptable dosage form such as those known in the art and those discussed above.
- the IFN ⁇ polypeptides or conjugates disclosed herein may be administered to a human intravenously as a bolus or by continuous infusion over a period of time, by intravenous, intramuscular, intraperitoneal, intra- cerebrospinal, subcutaneous, intra-articular, intrasynovial, intrathecal, or intratumoral routes.
- the IFN ⁇ polypeptides or conjugates can also be suitably administered by peritumoral, intralesional, or perilesional routes, to exert local as well as systemic therapeutic effects.
- a therapeutically effective amount of the IFN ⁇ polypeptide or conjugate or composition is an amount that is effective to reduce the severity, the duration and/or the symptoms of a particular disease or condition.
- the amount of the IFN ⁇ polypeptide or conjugate or composition that will be therapeutically effective in the prevention, management, treatment and/or amelioration of a particular disease can be determined by standard clinical techniques.
- the precise amount of the IFN ⁇ polypeptide or conjugate or composition to be administered will depend, in part, on the route of administration, the seriousness of the particular disease or condition, and should be decided according to the judgment of the practitioner and each subject’s circumstances. 1.8. Methods of Treatment [000371]
- the IFN ⁇ polypeptides and conjugates provided herein can be administered to a mammal, generally a human, for the treatment of any disease, disorder, or condition that would benefit from the stimulation of amplification of the immune response.
- the disease or condition is abnormal cellular proliferation.
- the disease, disorder, or condition is cancer.
- Any suitable cancer may be treated with the IFN ⁇ polypeptides and conjugates provided herein.
- Illustrative suitable cancers include, for example, acute lymphoblastic leukemia (ALL), acute myeloid leukemia (AML), adrenocortical carcinoma, anal cancer, appendix cancer, astrocytoma, basal cell carcinoma, brain tumor, bile duct cancer, bladder cancer, bone cancer, breast cancer (including triple-negative breast cancer, or TNBC), bronchial tumor, carcinoma of unknown primary origin, cardiac tumor, cervical cancer, chordoma, colon cancer, colorectal cancer, craniopharyngioma, ductal carcinoma, embryonal tumor, endometrial cancer, ependymoma, esophageal cancer, esthesioneuroblastoma, fallopian tube carcinoma, fibrous histiocytoma, Ewing sarcoma, eye cancer, germ cell tumor, gallbladder cancer, gastric cancer, gastrointestinal carcinoid tumor, gastrointestinal stromal tumor, 101
- ALL acute lymphoblastic leukemia
- AML acute my
- gestational trophoblastic disease glioma, head and neck cancer, hepatocellular cancer, histiocytosis, Hodgkin lymphoma, hypopharyngeal cancer, intraocular melanoma, islet cell tumor, Kaposi sarcoma, kidney cancer, Langerhans cell histiocytosis, laryngeal cancer, lip and oral cavity cancer, liver cancer, lobular carcinoma in situ, lung cancer, macroglobulinemia, malignant fibrous histiocytoma, melanoma, Merkel cell carcinoma, mesothelioma, metastatic squamous neck cancer with occult primary, midline tract carcinoma involving NUT gene, mouth cancer, multiple endocrine neoplasia syndrome, multiple myeloma, mycosis fungoides, myelodysplastic syndrome, myelodysplastic/myeloproliferative neoplasm, nasal cavity and par nasal sinus cancer, nasophary
- the disease to be treated with the IFN ⁇ polypeptides and conjugates provided herein is melanoma, gastric cancer, colorectal cancer, renal cell carcinoma, cervical cancer, non-small cell lung carcinoma, ovarian cancer, uterine cancer, fallopian tube carcinoma, primary peritoneal carcinoma, uterine corpus carcinoma, endometrial carcinoma, prostate cancer, and breast cancer.
- the disease to be treated is breast cancer.
- the disease to be treated is melanoma.
- a IFN ⁇ polypeptide or IFN ⁇ conjugates described herein or a pharmaceutical composition thereof treats the disease or condition, for example, cancer, by activating anti-tumor immunity.
- a IFN ⁇ polypeptide or IFN ⁇ conjugate described herein or a pharmaceutical composition thereof treats the disease or condition, for example, cancer, by inducing or enhancing anti-tumor immune memory.
- the disease or condition is a viral infection, for example hepatitis B (HBV) or hepatitis C (HCV).
- the viral infection is hepatitis C, including drug resistant and multidrug resistant forms of HCV or HBV and related disease states, conditions, or complications of an HCV or HBV infection, including cirrhosis and related hepatotoxicities,
- the HBV or HCV is chronic.
- a INF ⁇ polypeptide or conjugate as described herein is administered with a second active agent, for example, for the treatment of cancer.
- the second active agent is an immune checkpoint inhibitor, including but not limited to, a PD-1 inhibitor, a PD-L1 inhibitor, a PD-L2 inhibitor, a CTLA-4 inhibitor, or a LAG-3 inhibitor.
- the immune checkpoint inhibitor is a PD-1 inhibitor.
- the immune checkpoint inhibitor is a PD-1 immune checkpoint inhibitor selected from, but not limited to, nivolumab (Opdivo), pembrolizumab (Keytruda), and cemiplimab (Libtayo). In certain embodiments, the PD-1 immune checkpoint inhibitor is pembrolizumab (Keytruda). [000378] In certain embodiments, the immune checkpoint inhibitor is a PD-L1 inhibitor. In certain embodiments, the immune checkpoint inhibitor is a PD-1 immune checkpoint inhibitor selected from, but not limited to, atezolizumab (Texentriq), durvalumab (Imfinzi), and Avelumab (Bavencio).
- the immune checkpoint inhibitor is a CTLA-4 immune checkpoint inhibitor.
- the immune checkpoint inhibitor is a PD-1 immune checkpoint inhibitor selected from, but not limited to, ipilimumab (Yervoy).
- the immune checkpoint inhibitor is a LAG-3 immune checkpoint inhibitor, for example, Relatlimab.
- a INF ⁇ polypeptide or conjugate as described herein is administered with an immune checkpoint inhibitor selected from a PD-1 inhibitor, a PD-L1 inhibitor, a PD-L2 inhibitor, a CTLA-4 inhibitor, or a LAG-3 inhibitor for the treatment of melanoma.
- the immune checkpoint inhibitor is a PD-1 inhibitor, for example, pembrolizumab (Keytruda).
- a IFN ⁇ polypeptide or conjugate as described herein is administered in combination with a second active agent for the treatment of hepatitis B or hepatitis C, including, but not limited to ribavirin.
- a IFN ⁇ polypeptide or conjugate as described herein is administered in combination ribavirin for the treatment of hepatitis C.
- Additional second active agents that can be administered in combination with a IFN ⁇ polypeptide or conjugate as described herein for the treatment of hepatitis C include, but are not limited to, a protease inhibitor (such as telaprevir (Incivek), boceprevir (Victrelis), and 103
- a protease inhibitor such as telaprevir (Incivek), boceprevir (Victrelis), and 103
- simeprevir (Olysio); a NS5A inhibitor (such as daclatasvir (Daklinza) and velpatasvir (Epclusa); a NS5B inhibitor (such as dasabuvir (Exviera) and sofosbuvir (Sovaldi)); or a combination drug (such as Harvoni (ledipasvir/sofosbuvir), Viekira Pak (ombitasvir/paritaprevir/ritonavir/dasabuvir), Viekirax (ombitasvir/paritaprevir/ritonavir), Mavyret (paritaprevir and glecaprevir), Technivie (ombitasvir/paritaprevir/ritonavir), Epclusa (sofosbuvir/velpatasvir) and Zepatier (elbasvir and grazoprevir)).
- Harvoni ledipasvir/sofosbuvir
- Additional second active agents that can be administered in combination with a IFN ⁇ polypeptide or conjugate as described herein for the treatment of hepatitis B include, but are not limited to, a nucleoside reverse transcriptase inhibitor, for example, Epivir (Lamivudine), Hepsera (Adefovir dipivoxil), Baraclude (Entecavir), Tyzeka (Telbivudine), Viread (Tenofovir), Vemlidy (tenofovir alfenamide), and Levovir (Cledvudine). 1.10. Diagnostic Applications [000385] In some embodiments, the IFN ⁇ polypeptides or conjugates provided herein are used in diagnostic applications.
- an IFN ⁇ polypeptide or conjugate disclosed herein that is specific for a given receptor may be useful in assays for the given receptor.
- the IFN ⁇ polypeptide or conjugate can be used to detect the expression of the given receptor in various cells and tissues. These assays may be useful, for example, diagnosing cancer, infection and autoimmune disease.
- the formation of a complex between the IFN ⁇ polypeptide or conjugate and receptor can be detected by any method known to those of skill in the art. Examples include assays that use secondary reagents for detection, ELISA’s and immunoprecipitation and agglutination assays.
- the IFN ⁇ polypeptide or conjugate may be administered to a subject by methods known in the art such as, for example, intravenous, intranasal, intraperitoneal, intracerebral, intraarterial injection such that a specific binding between the IFN ⁇ polypeptide or conjugate and receptor may occur.
- the IFN ⁇ polypeptide or conjugate/receptor complex may conveniently be detected through a label attached to the IFN ⁇ polypeptide or conjugate or any other art-known method of detection.
- the IFN ⁇ polypeptide or conjugate may be labeled with a detectable moiety. Suitable detectable moieties include, but are not limited to radioisotopes, fluorescent labels, and enzyme-substrate labels. 1.11. Kits [000389] In some embodiments, an IFN ⁇ polypeptide or conjugate as described herein can be provided in a kit, i.e., a packaged combination of reagents in predetermined amounts with instructions for performing a procedure. In some embodiments, the procedure is a diagnostic assay. In other embodiments, the procedure is a therapeutic procedure.
- IFN ⁇ variants were expressed in a cell-free protein synthesis reaction as described in Zawada et al. Biotechnol. Bioeng., 2011, 108:1570-1578. Briefly, cell-free extracts were added to a premix containing cell-free reaction components (Groff et al., mAbs, 2014, 6:671- 678) and 10ug/mL plasmid DNA template.
- IFN ⁇ -His6 variants were quantified via high throughput capillary electrophoresis using the LabChip GXII ® (Perkin Elmer) against an IFN ⁇ standard curve, according to the manufacturer's instructions.
- Thermal Stability of pAMF substituted variants by Differential Scanning Fluorimetry was determined by differential scanning fluorimetry (DSF) as previously described in He et al. J Pharm Sci, 2010, 99:1707-1720. Briefly, in a CFX384 (Bio-Rad Laboratories), variants were heated in the presence of Sypro Orange protein stain (Millipore). Fluorescence was monitored and Tm was determined by the minimum of the negative derivative. The thermal stability of selected single-site IFN ⁇ variants is provided in Table 4. 105 [000393] Table 4.
- IFN ⁇ variants were bound to IFNAR2 at concentrations from 0.8 nM to 100 nM.
- the data was fit with the Biacore T200 Evaluation software using a 1:1 Langmuir binding model.
- the binding affinity of selected single-site IFN ⁇ variants to hIFNAR1 and hIFNAR2 are provided in Table 5. [000395] Table 5. Binding affinity of selected single-site IFN ⁇ variants to hIFNAR1 and hIFNAR2 107
- HEK-blue IFN ⁇ / ⁇ Reporter Assay for human IFN ⁇ [000396] HEK-Blue IFN ⁇ / ⁇ Reporter Cells (Invivogen, Cat# hkb-ifnab) were maintained in complete DMEM/F-12 Media (Corning) with 100IU Penicillin/100 ⁇ g/mL Streptomycin (Corning), 2mM GlutaMax (Gibco), 10% h.i. FBS (Sigma), 100 ⁇ g/mL Normocin (Invivogen), and HEK-Blue Selection antibiotics mix (Invivogen).
- IFN ⁇ samples in Table 6 were conjugated with LP5 or PEG (2x20 kD) or LP9 (DBCO-PEG 4 -amine), a residual linker remaining after the PEG mask is released through protease cleavage.
- the potency of LP5-conjugated IFN ⁇ variants was attenuated compared to unconjugated IFN ⁇ variants and an IFN ⁇ control without pAMF incorporation.
- LP9-conjugated IFN ⁇ variants had in most cases similar potency as un-conjugated IFN ⁇ variants.
- the structure of LP5 and LP9 are provided below and the conjugation was performed as described herein.
- IFN ⁇ samples in Table 7 were conjugated with LP6 (20kD PEG) or LP11, a residual linker remaining after the PEG mask is released through protease cleavage.
- the potency of LP6-conjugated IFN ⁇ variants was attenuated compared to unconjugated IFN ⁇ variants and an IFN ⁇ control without pAMF incorporation.
- LP11-conjugated IFN ⁇ variants had in most cases similar potency as un-conjugated IFN ⁇ variants.
- the structure of LP6 and LP11 are provided below and the conjugation was performed as described herein. 108
- Recombinant human cathepsin B (catB, 953- CY-010, R&D Systems) was pre-activated with 5 mM DTT in 10 mM MES pH 5.0 at room temperature for 15 min.
- In vitro catB PEG release assays were performed with 20 uM conjugated IFNa and 240 nM activated catB in 50 mM sodium phosphate, pH 6.0 at 40 °C for 16 hours.
- Table 8 shows the %PEG release calculated by SDS-PAGE gel densitometry of IFN ⁇ variants conjugated to DBCO-valcit-pAB-PEGs with varying linker lengths.
- the structures of LP2, LP6, LP7, and LP8 are provided below.
- cell free reactions were prepared by the addition of 37.5% v/v S30 extract, 3 ug/mL plasmid encoding IFN variants and a supermix containing amino-acids, NMPs and small molecules for energy generation (Cai, Q. et al. Biotechnol. Prog.31, 823–831 (2015)).
- Four macromolecular reagents were individually over-expressed in E. coli and added to the XpressCF+ ® reaction as reagent lysates at ⁇ 1% v/v each: T7 RNA polymerase, E. coli peptide deformylase, and the orthogonal tRNA synthetase / tRNA pair from M.
- the digested reaction was analyzed by 4-12% SDS-PAGE to verify full cleavage of the His SUMO tag prior to HiPrep Desalting column with Sephadex G-25 resin for rapid buffer exchange into 20 mM Tris-acetate, 150mM NaCl, 1mM DTT, pH 7.5.
- IMAC affinity polish and buffer exchange [000407]
- the desalted IFN variants were applied to a HisTrap Exel affinity column equilibrated with 15 mM Tris-acetate, 150mM NaCl, 1mM DTT pH 7.5 as a flow through chromatography process.
- the target IFN ⁇ variants were eluted from the column without adsorption whereas the remaining contaminants were strongly bound.
- DBCO-Fmoc-mPEG (20 kDa) (LP1) [000410] DBCO-Fmoc-mPEG (20 kDa) (LP1) was synthesized as shown below in Scheme 1.
- Scheme 1 [000411] Step 1: In a 1000 mL flask equipped with a magnetic stir bar, mPEG-NH 2 (20,000 kDa) (23.98 g, 1.2 mmol) in anhydrous toluene (250 mL) was added. The mixture was azeotropically dried under reduced pressure at 45 °C on a rotary evaporator, lyophilized overnight, and then dissolved in anhydrous DCM (200 mL).
- Step 2 To an oven-dried 250 mL flask equipped with a magnetic stir bar, Fmoc PEGylated amide compound 4 (11.4 g, 0.52 mmol) (azeotropically dried with 100 mL toluene removed at 50° C under vacuum prior to use) and anhydrous DCM (70 mL) were added.
- mPEG (20kDa)-valcit-pAB-(PEG) 4 -DBCO (LP2) [000413] mPEG (20 kDa)-valcit-pAB-(PEG)4-DBCO (LP2) was synthesized as shown below in Scheme 2.
- DBCO-C 6 -NHS ester (in about 10% excess) and mPEG-amine was dissolved in anhydrous DCM and the reaction was stirred for 24 hours while monitoring the consumption of DBCO-C 6 -NHS ester.
- the compounds were purified by repeated crystallization from MTBE until no DBCO-C6-NHS ester was detected by HPLC.
- DBCO PEG compounds LP4 and LP5 were confirmed by 1 H NMR (CDCl3), MALDI-TOF, and analytical ELSD-HPLC.
- PEG density analysis using gel densitometry was used to estimate PEG density.1-4 ug of PEGylated IFN ⁇ was loaded on 4-12% Bis-tris SDS-PAGE (NuPAGETM Invitrogen). The gel ran in 1x NuPAGETM MES SDS Running Buffer (Invitrogen) with constant voltage at 400 volts for 35 minutes. The gel image was scanned using Bio-Rad Gel DOC EZ Imager and exported for densitometry analysis using ImageQuant TL 7.0 (GE Health). The PEGylated IFN ⁇ migrated slower than unpegylated IFN ⁇ .
- B16-Blue IFN ⁇ / ⁇ Reporter Cells (Invivogen, Cat# hkb-ifnab) were maintained in complete DMEM/F-12 Media (Corning) with 100IU Penicillin/100 ⁇ g/mL Streptomycin (Corning), 2mM GlutaMax (Gibco), 10% h.i. FBS (Sigma), 100 ⁇ g/mL Normocin (Invivogen), and HEK-Blue selection antibiotics mix (Invivogen).
- cells were harvested with Accutase, counted and resuspend at 0.5 x 106 cells/mL 25 ⁇ L of cells were seeded per well in 384-well clear-bottom plate.
- FIG. 6 illustrates plasma concentrations of hIFNa2b from each test article.
- Conjugate 33 had the fastest plasma clearance and was not detected 4 hours post dosing.
- Conjugate 1 and Conjugate 13 had similar half-life extension, whereas Conjugate 18 had a faster clearance (FIG.6).
- FIG. 1A and FIG. 1B illustrate the effects of Conjugate 1, Conjugate 16 and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 41 post 125
- FIG.1A The effect of a 3 mg/kg dose is shown in FIG.1A and the effect of a 10 mg/kg dose is shown in FIG.1B.
- Analysis of tumor sizes was done on day 41, when the mean of vehicle- treated tumors reached the study endpoint (>1,500 mm 3 ).
- Conjugate 1 induced greater tumor growth suppression (47% TGI) compared to Conjugate 18 (34% TGI) and Conjugate 16 (36% TGI) (FIG. 1A).
- Conjugate 1, Conjugate 18, and Conjugate 16 showed similar tumor growth suppression (51% TGI and 51% TGI and 45% TGI, respectively) (FIG.1B).
- FIGS.1C, 1D, and 1E illustrate the effects of Conjugate 1, Conjugate 13, Conjugate 11, and Conjugate 31 on MDA-MB-231 tumor growth up until the end of the study at day 44 post treatment.
- FIG. 1C and FIG. 1D show the effect of a 3 mg/kg and 15 mg/kg dose, respectively.
- FIG.1E is a graph showing the tumor size on day 44. Analysis of tumor sizes was done on day 44, when the mean of vehicle-treated tumors reached >1,200 mm 3 .
- Overall Conjugate 11, Conjugate 13, and Conjugate 31 exhibited trends of dose-dependent anti-tumor activity.
- Conjugate 1 induced greater tumor growth suppression (50% TGI) compared to Conjugate 11 (10% TGI), Conjugate 13 (37% TGI) (FIG. 1C).
- Conjugate 11 and Conjugate 13 induced similar activity (38% TGI and 46% TGI, respectively) (FIG.1D).
- Conjugate 31, which has a non-releasable PEG did not demonstrate significant effect when dosed at 3 mg/kg and 15 mg/kg (0% and 12% TGI) (FIG.1E) compared to other IFN ⁇ -variants, indicating that cleavage of PEG is essential to confer anti-tumor activity.
- mice were engrafted intraperitoneally with 5x10 6 human PBMCs and implanted with 5x10 6 MDA-MB-231 tumor cells in the mammary fat pad. Mice were randomized and enrolled into the study 7 days post implant, with tumor sizes around 100 mm 3 - 150 mm 3 . Tumor-bearing mice from both studies were administered three weekly doses (qwx3) of the test articles at doses ranging from 0.05 mg/kg to 3 mg/kg. All treatments were well tolerated with normal body weight gain throughout the course of the study.
- FIGS. 2A-2E summarize results illustrating dose-dependent effects of different IFN ⁇ variants on growth of MDA-MB-231 tumors up in a mouse model engrafted with human PBMCs. At equivalent doses of 3 mg/kg, all test articles demonstrated prolonged tumor stasis 126
- Conjugate 1 induced mildly greater tumor growth suppression (104% TGI) compared to Conjugate 18 (91% TGI), Conjugate 13 (83% TGI) and Conjugate 2 (82% TGI) (FIG.2A).
- activity of Conjugate 1 and Conjugate 18 was the greatest and nearly comparable (95% and 92% TGI, respectively) followed by Conjugate 13 (70% TGI) and Conjugate 2 (51% TGI) (FIG.2B).
- Conjugate 1 Treatment with Conjugate 1 demonstrated potent and dose-dependent tumor growth suppression at 0.1 mg/kg and 0.05 mg/kg doses (81% TGI at 0.1 mg/kg and 60% TGI at 0.05 mg/kg) (FIGS. 2C and 2D).
- Conjugate 18 demonstrated comparable anti-tumor activity at 0.1 mg/kg and 0.05 mg/kg dose (59% and 52% TGI, respectively) (FIG.2C and 2D).
- a similar trend was seen with Conjugate 13, inducing similar effect at 0.1 mg/kg and 0.05 mg/kg dose (32% and 40% TGI, respectively) (FIG.2C and 2D).
- Activity of Conjugate 2 was the least, with 23% TGI at 0.1 mg/kg dose (FIG. 2C and 2E).
- Conjugate 1 demonstrated the greatest TGI amongst all IFN ⁇ variants tested at equivalent doses (FIG.2E).
- EXAMPLE 11 TOLERABILITY OF HLE-INTERFERON VARIANTS IN GOLDEN SYRIAN HAMSTERS [000437] In vivo tolerability of HLE-Interferon variants was tested in naive Golden Syrian hamsters in a single dose study. Briefly Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 were dosed IV at doses ranging from 3 mg/kg to 45 mg/kg. Animal body weights were monitored regularly for 7 days throughout the course of the study. Serum samples were collected at different timepoints to assess plasma levels of each test article.
- FIG. 3A illustrates serum concentrations of each test article.
- Conjugate 13, Conjugate 18, Conjugate 1 was detectable until Day 7 post treatment.
- Treatment with Conjugate 18 and Conjugate 13 followed linear dose-dependent pharmacokinetics (FIG. 3A).
- Conjugate 13 and Conjugate 1 showed similar PK profile and had greater half-life extension compared to Conjugate 18 and Conjugate 2 (FIG. 3B).
- Overall Conjugate 2 had the fastest clearance, with total interferon levels undetectable in 2 out of 3 animals by day 3 of study (FIG.3B).
- hamsters treated with Conjugate 18 showed dose dependent body weight loss (-7% at 3 mg/kg on Day 7, -10.4% loss at 15 mg/kg on Day 7, -20.6% at 45mg/kg on Day 6).
- a single dose of 45 mg/kg of Conjugate 18 was not tolerated (FIG. 3C).
- a single dose of 15 mg/kg Conjugate 1 also induced 12.4% reduction in hamster body weight on Day 7 post treatment (FIG.3D).
- a single dose of Conjugate 13 did not result in any hamster body weight loss up to a dose of 45 mg/kg (FIG.3C).
- FIGS.3E-J illustrate effects of all test articles on induction of liver enzymes AST, ALT, and ALP compared to the vehicle-treated serum on 2 days and 7 days of the study. Analysis of serum revealed treatment with Conjugate 18 at all doses resulted in early induction of ALT (8.5-20 fold) on day 2 of the study that returned to baseline levels by day 7 (FIG.3E and 3H).
- Conjugate 13 resultsed in ⁇ 5 fold increase in AST levels on day 7 of study (FIG.3I).
- Treatment with 15 mg/kg of Conjugate 1 resulted in a time-dependent increase in levels of AST (4 fold on day 2, 9 fold on day 7) and ALP (1.5 fold on day 2, 4 fold on day 7) (FIG.3F, 3G, 3I, and 3J).
- Conjugate 1 Similar to Conjugate 13 and Conjugate 18, Conjugate 1 also caused initial induction of ALT on day 2 ( ⁇ 10.5 fold) that returned to baseline by day 7 (FIG.3E and 3H).
- FIGS.3K-3L demonstrate the effect of all test articles on platelet and reticulocyte counts. Analysis of whole blood was done on Day 7 post treatment. Analysis revealed that treatment with Conjugate 18 resulted in greater than 50% reduction in reticulocyte count at the lowest dose 3 mg/kg. At doses greater than 3 mg/kg, Conjugate 18 showed dose-dependent decrease in reticulocyte count (92% reduction at 15 mg/kg and 99% reduction at 45 mg/kg dose) (FIG.3K). Treatment with Conjugate 1 resulted in ⁇ 97% decrease in reticulocyte count. Treatment with Conjugate 13 showed evidence of dose-dependent loss in reticulocyte count 128
- FIG.3K illustrates the effect of all IFNa variants on platelet count on Day 7.
- Treatment with Conjugate 18 showed reduction in platelet counts at all doses (40-55% reduction).
- Treatment with Conjugate 13 and Conjugate 2 did not result in platelet loss.
- Conjugate 1 at 15 mg/kg induced ⁇ 60% reduction in platelet count (FIG.3L).
- EXAMPLE 12 IN VITRO ACTIVITY OF MOUSE SURROGATE
- Mouse IFNa molecules with pAMF incorporated at the same sites as in human IFNa were made in order to study the efficacy of the IFNa variants in mouse models with intact immune system.
- the in vitro activity of mouse IFNa molecules were evaluated using a B16-Blue IFN ⁇ / ⁇ Reporter assay.
- B16-Blue IFN ⁇ / ⁇ Reporter Cells (Invivogen, Cat# hkb-ifnab) were maintained in complete DMEM/F-12 Media (Corning) with 100IU Penicillin/100 ⁇ g/mL Streptomycin (Corning), 2mM GlutaMax (Gibco), 10% FBS (Sigma), 100 ⁇ g/mL Normocin (Invivogen), and HEK-Blue selection antibiotics mix (Invivogen).
- cells were harvested with Accutase, counted and resuspend at 0.5 x 106 cells/mL 25 ⁇ L of cells were seeded per well in 384-well clear-bottom plate.
- FIG.4A-4G illustrates the effects of different test articles on B16F10 tumor growth. Analysis of tumor sizes was done on study day 10, when the mean of vehicle-treated tumors reached the study endpoint. Overall treatment with Conjugate 36 resulted in the most potent anti-tumor activity compared to all other test articles (FIG. 4D).
- Conjugate 36 showed significantly greater TGI (90%) followed by Conjugate 37 (73%), Conjugate 34 (65%) and Conjugate 35 (25%) (FIG.4A).
- Conjugate 36 induced greater tumor growth suppression compared to Conjugate 34 (79% and 44% 130
- Conjugate 37 showed significantly greater TGI compared to Conjugate 35: 73% and 25% TGI, respectively at 3 mg/kg; 81% and 35% TGI respectively, at 10 mg/kg, indicating that cleavage of PEG is essential to confer anti-tumor activity (FIG.4C).
- Anti-tumor activity of Conjugate 37 was also evaluated in MC38 cells expressing a human tumor associated antigen. Briefly, C57/Bl6 mice were implanted subcutaneously with 1x10 6 MC38-hCEA cells and enrolled into the study with tumor size around 150 mm 3 .
- FIG.4H illustrates that treatment with Conjugate 37 elicits potent dose-dependent anti-tumor activity and results in high complete response rates in this model.
- Analysis of tumor sizes done on study day 17 revealed 48% TGI at 0.1 mg/kg, 96% TGI at 1 mg/kg.
- 12.5% mice in the 1 mg/kg dose group and 100% mice in the 3 mg/kg dose group were tumor free.
- FIGS.4E-4G illustrates the effect of Conjugate 37 on immune activation in TME.
- mice were implanted subcutaneously with 1x10 6 B16F10 cells and enrolled into the study with tumor size around 70 mm 3 .
- Tumor bearing mice were administered 3 weekly (qwx3) doses of Conjugate 37 at 3 mg/kg as a single-agent or in combination with 10 mg/kg anti-PD-1.
- Mouse 131
- FIG. 7A illustrates single-agent and combination anti-tumor activity of Conjugate 37 and anti-PD-1 on B16F10 tumor growth. Analysis of tumors on study day 10 when the vehicle group tumor volume reached ⁇ 1,000 mm 3 revealed anti-PD-1 treatment alone did not induce anti-tumor activity (FIG.7A). Analysis of tumors on study day 24 revealed single agent treatment with Conjugate 37 revealed potent anti- tumor activity. Conjugate 37 combined with anti-PD-1 exhibited trends of greater anti-tumor activity compared to treatment with Conjugate 37 alone (FIG. 7A and FIG.
- FIG.7C demonstrates mice treated with combination of Conjugate 37 and anti-PD-1 exhibit longer time to reach tumor volume of 500 mm 3 compared to mice treated with Conjugate 37 alone.
- EXAMPLE 15 PRODUCTION OF INTERFERON ALPHA CONTAINING 3 PAMF AMINO ACIDS IN E. COLI CELLS WITH HIGH DENSITY FERMENTATION
- IFN ⁇ Interferon alpha
- NNAAs non-natural amino acids
- CDS coding sequence for an aminoacyl tRNA synthetase (RS) specific for para-azidomethylphenylalanine
- pAMF para-azidomethylphenylalanine
- pAMF para- azidomethylphenylalanine
- pAMF RS CDS Three copies of a tRNA specific for para- azidomethylphenylalanine (pAMF) were cloned behind the pAMF RS CDS, with 23 nucleotide non-coding DNA spacers before each tRNA sequence.
- IFN ⁇ human interferon alpha
- HisSUMO-IFN ⁇ Q40/E51/N156 TAG was codon optimized for E. coli.
- the construct was cloned behind a T7p and strong RBS into a high copy (pUC origin) plasmid with a kanamycin (Kan) selection cassette.
- the E. coli strain for expression of IFN ⁇ was generated by transforming the E. coli Snuggle strain with both the RS plasmid and product plasmid. Transformations were plated on LB agar containing 50 ⁇ g/mL kanamycin and 100 ⁇ g/mL carbenicillin. Single colonies were picked and transferred into culture tubes with 3 mL of TB media containing 50 ⁇ g/mL kanamycin and 100 ⁇ g/mL carbenicillin for overnight growth at 37° C.
- the culture tube was used to inoculate a shake flask with I17-SF shake flask media containing 50 ⁇ g/mL of kanamycin and 100 ⁇ g/mL of carbenicillin at 8% (v/v) seeding density.
- the shake flask was harvested once the culture achieved an OD 595 nm greater than 3.
- Glycerol was added to the shake flask to a final concentration of 16-20% (v/v).
- the cell bank was collected and aliquoted into 2 mL vials, flash frozen in liquid nitrogen and stored at -80°C.
- the fermentation process began by taking a 2 mL vial of the cell bank and inoculating a shake flask with I17-SF shake flask media (as described in Hanson, J.; Groff, D.; Carlos, A.; Usman, H.; Fong, K.; Yu, A.; Armstrong, S.; Dwyer, A.; Masikat, M.R.; Yuan, D.; et al. An Integrated In Vivo/In Vitro Protein Production Platform for Site- Specific Antibody Drug Conjugates.
- the shake flask culture was used to inoculate a 1 L bioreactor at a seeding density of 6% (v/v) in batched media.
- the batched media consisted of 2.4% (v/v) 5x I17 Media in DI H2O, 50 ⁇ g/mL of kanamycin, 100 ⁇ g/mL of carbenicillin, and 0.1% (v/v) A204 antifoam.
- the bioreactor temperature, dissolved oxygen and pH setpoints at inoculation were 37° C, 30% and 7, respectively.
- the fed batch phase began by feeding 5x I17 media at an exponential rate of 0.15 h-1. After 21 hours in the fed batch phase, the temperature decreased to 25° C, and the exponential feed rate decreased to 0.02 h-1.
- the induction phase began by adding pAMF to a target concentration of 4 mM and L-Arabinose to a target concentration of 4 g/L based on the culture volume in the bioreactor prior to induction. The induction phase took 24 hours before the bioreactor was harvested.
- the culture was collected and centrifuged at 18,592 xG and 2-8° C for 15 min in a floor centrifuge. The supernatant was discarded, and the cell pellets were resuspended with DPBS at a concentration of 16.67% (w/w).
- the cell resuspension 133 was then passed twice through an Avestin Homogenizer (EmulsiFlex-C5) at 17,000 Psi to disrupt the cells and generate the crude lysate.
- the crude lysate was clarified by centrifuging at 18,000-20,000 xG and 2-8° C for 30 minutes in a floor centrifuge.
- the supernatant (clarified lysate) was collected and aliquoted, flash frozen in liquid nitrogen and stored at -80°C.
- Lysate supernatants were applied to Ni-NTA resin that had been pre-equilibrated with PBS. After application of the supernatant, the resin was washed with PBS containing 10 mM imidazole before the protein was eluted across several fractions with PBS containing 200 mM imidazole. The purest fractions were identified by analysis via SDS-PAGE then pooled and concentrated in 10 kDa MWCO Amicon centrifuge filters.
- Samples were quantified by adjusting the absorbance at 280 nm according to the calculated molar absorbance of the protein and considering the % purity calculated by gel densitometry analysis from an SDS-PAGE gel. [000458] The presence of full-length protein was verified using intact LC-MS analysis. Protein samples were digested with Ulp1 (1:20 w/w ratio) for 1 hour at 22 o C, after which 10- 15 pmol of each protein sample was injected onto a reverse phase column via the autosampler of an Agilent 1200 series HPLC.
- This vector has a kanamycin resistance marker and a pUC high copy origin of replication, and the expression cassette has a T7 promoter for high level transcription. Plasmid sequence was verified by sequencing. [000460] For the incorporation of nnAAs into proteins of interest, the selected codons in product genes where nnAAs would be incorporated were substituted with the amber codon “TAG.” The gene for 3XnnAA-IFN ⁇ was mutated to contain 3 TAG codons at the positions coding for amino acids 40, 51, and N156 as described in Example 15. The coding sequence for the pAMF RS was cloned into a medium copy pJ434 plasmid behind a constitutive Pc0 promoter.
- Proteins were first buffer exchanged into 20 mM Tris, 300 mM sodium chloride, pH 7.5 with Cytiva Sephadex G-25 fine resin. Then they were applied back onto Cytiva Ni Sepharose excel resin and the flowthrough contained the target protein. The final pool was concentrated and buffer exchanged into PBS, 9% sucrose, pH 6 with Amicon centrifuge filters (10 kDa MWCO) for conjugation. [000466] For the PEG conjugation step, the conjugation reaction was diluted with water, adjusted to pH 5 using 1 M acetic acid, and applied to Cytiva Capto SP ImpRes resin.
- Mobile phase A consisted of 10 mM citric acid, pH 5 and mobile phase B consisted of 10 mM citric acid, 300 mM sodium chloride, pH 5. Protein was eluted using a linear gradient from 0% to 100% mobile phase B. Targeted fractions were collected and buffer exchanged into PBS, 9% sucrose, pH 6 with Amicon centrifuge filters (10 kDa MWCO) for further testing.
- Peaks were filtered by setting a signal-to-noise ratio of > 30.0. Top 90% of the peak height was used to calculate average mass.
- protein concentrations were brought to 1 mg/mL in DPBS.
- the DBCO-amine was added at a drug to pAMF ratio of 3:1, and 500 mM NaCl was added to the reaction to improve DBCO-amine solubility.
- the conjugation reaction was incubated overnight at 30°C prior to LC-MS analysis.
- proteins were dialyzed into 1x DPBS + 9% Sucrose prior to conjugation.
- the PEG of interest was prepared in water as a 5mM stock solution.
- the protein was formulated in DPBS buffer, and 3 molar equivalents of PEG were added per mole of pAMF. Conjugation reactions were incubated at 25°C (3XnnAA-IFN ⁇ ) overnight in a Thermomixer (Fisher scientific, Allentown Pennsylvania) with agitation at 450rpm.. [000472] After conjugate cleanup, PEGylated proteins were analyzed via SDS-PAGE. PEG- to-protein ratios were calculated by gel densitometry analysis using the Lane and Bands image analysis tools in the Image Lab software (version 5.2.1, Bio-Rad).
- IFNa/b HEK-Blue cells were obtained from Invivogen (catalog number hkb-ifnab) and were plated in 384-well plates at a density of 12,500 cells/well in HEK-Blue detection media (Invivogen, catalog number hb-det2). The cells were treated with various concentrations of test articles and incubated at 37°C with 5% CO2 137
- Interferon ⁇ is a cytokine protein that has successfully been used for the treatment of ulcerative melanoma and renal cell carcinoma and has shown promise as a treatment for leukemias and solid tumors.
- Conditional activation of cytokines such as IFN ⁇ through PEGylation can reduce toxicity and extend drug half-life while maintaining anti-tumor efficacy.
- This strategy requires the attachment of multiple, releasable PEGs that initially mask receptor binding in a prodrug form.
- Incorporation of multiple nnAAs into a single protein chain is known to reduce the amount of full length protein produced, but this effect can be overcome by increasing the cellular level of AS tRNA which helps overcome translational termination at TAG codons.
- a construct was designed that would result in the incorporation of pAMF at 3 solvent-exposed sites on IFN ⁇ . Because the variant contained several nnAA sites, the product plasmid was co-transformed into E.
- coli SBDG419 with an RS plasmid containing 3 copies of the pAMF tRNA in order to increase intracellular AS tRNA levels and amber suppression efficiency (FIGS.10A-10B).
- Initial test expressions revealed that the nnAA-containing IFN ⁇ variant (3XnnAA-IFN ⁇ ) expressed at levels comparable to the wild-type protein (FIG.10C).
- analysis of the unlabeled protein revealed significant truncation (18%) at the C- terminal pAMF site (FIGS. 11A-11C). Therefore, an additional copy of the AS tRNA was cloned at the 3’ end of the IFN ⁇ coding sequence of the high copy product plasmid (FIG.10B).
- the 3XnnAA-IFN ⁇ protein that had been expressed in shake flasks using this construct was purified and analyzed the sample via LC-MS, which revealed that the addition of tRNA to the product plasmid improve amber suppression efficiency and significantly reduced the truncation observed at the C-terminal pAMF site (FIG.11C).
- a conjugation reaction with a small molecule DBCO- amine was performed. Analysis by intact LC-MS showed that >96% of each sample had been labeled with three DBCO-amine molecules (FIGS. 11A-11C), confirming the successful incorporation of three nnAAs.
- the deconvoluted spectra predominantly consisted of a main peak, with the presence of 1% truncated product (FIGS.11A-11C).
- the 3XnnAA- IFN ⁇ was produced at a titer of approximately 540 mg/L (FIG.9D).
- Fed-batch fermentation of this strain was scaled to 500 mL to produce sufficient material for downstream processing and activity analysis. Growth rates, OD595 and product titers compared well with previous fermentations.
- the 3XnnAA-IFN ⁇ protein was expressed at titers of 600 mg/L as calculated by purification of the protein over IMAC resin (FIG.9D).
- the PEGylated 3XnnAA-IFN ⁇ showed no cell activation activity, demonstrating effective activity attenuation (expected activity of a prodrug) through site-specific PEGylation at multiple sites.
- the successful activity attenuation of 3XnnAA-IFN ⁇ through site specific PEGylation suggests that the E. coli SBDG419 production strain could feasibly be used to generate mutants of other therapeutic proteins containing multiple conjugatable pAMF handles.
- FIGS.12A-F illustrate the effect of Conjugate 37 and Conjugate 34 on different cell types in the TME. Analysis revealed both HLE-Interferon variants increased Granzyme B levels in tumor infiltrating CD8 T-cells and NK cells compared to vehicle-treated tumors (FIGS.12A-12B). Additionally Conjugate 37 and Conjugate 34 treatment increased activation of multiple innate immune cells in TME including monocytes, dendritic cells and plasmacytoid dendritic cells (FIGS. 12C-12E).
- Lymph node analysis revealed that treatment with Conjugate 34 resulted in a similar increase in levels of GranzymeB in CD8 T-cells from both tumor-draining and non-draining lymph node.
- Conjugate 37 treatment resulted in a greater increase in GranzymeB levels in CD8 T-cells from tumor-draining lymph node compared to a non-draining lymph node (FIG 12F).
- EXAMPLE 18 EVALUATION OF FUNCTIONAL ROLE OF CD8 T-CELLS IN FORMATION OF HLE- INTERFERON VARIANT INDUCED ANTI-TUMOR IMMUNE MEMORY [000481] The functional role of CD8 T-cells in formation of HLE-Interferon variant induced anti-tumor immune memory was evaluated in tumor free (complete responder) mice obtained from Conjugate 37 treatment. Complete responder mice were treated with 300 ⁇ g anti-CD8 antibody or Isotype antibody and rechallenged with 5x10 6 MC38-hCEA cells on D0. CD8 depletion was conducted prior to rechallenge and maintained throughout the course of the study.
- Conjugate 37-treated complete responder mice that received Isotype control antibody demonstrated no recurrence of tumors when rechallenged with MC38-hCEA cells.
- CD8 depletion ablated Conjugate 37-induced anti-tumor immune memory in complete responder mice as seen by formation of tumors when rechallenged with MC38-hCEA cells (FIG 13).
- HLE-Interferon variants are tested in a non-human primate system, such as cynomolgus 140 monkey.
- HLE-Interferon variants are dosed to cynomolgus monkeys by slow intravenous infusion and subcutaneous injection. The monkeys are dosed at different dose levels (e.g., 6.75 mg/kg or 20.25 mg/kg) to understand the toxicity and PK-PD relationship. Blood is drawn at various timepoints to monitor drug exposure and proof of mechanism.
- HLE- Interferon variants PK profile is aligned with observed toxicity and markers of mechanism to generate a better understanding of the drug safety profile.
- PK profile of the drugs is evaluated by measuring concentration of total Interferon and PEGylated Interferon. Presence of higher concentrations of non-PEGylated Interferon can contribute to interferon-mediated toxicity.
- Proof of mechanism and PD profile for HLE- Interferon variants is evaluated by measuring levels of different mechanism of action and inflammatory cytokines in the circulation. Blood clinical chemistry is evaluated to monitor for signs of liver or kidney toxicity. Blood hematology, such as various red and white blood cells, is also monitored to detect hematological toxicities.
- HLE-Interferon variants can be inferred by the PK-PD profile. Additionally, safety of the HLE-Interferon variants can be partially represented by the drug’s Highest Non-Severely Toxic Dose (HNSTD), or dose at which no or limited toxicities are observed. The higher the HNSTD, the higher probability of the drug safety.
- HNSTD Highest Non-Severely Toxic Dose
- EXAMPLE 20 SEQUENCES Table 13 provides sequences referred to herein. [000485] In certain embodiments, SEQ ID NO.2-31 and 33-38 are preceded by a methionine amino acid. In Table 13, the sequence of the indicated HisSUMO fusions (SEQ. ID NO: 39) are not shown for clarity.
- the cleavable HisSUMO fusion facilitates expression and purification.
- IFN ⁇ polypeptides according to any of SEQ ID NOS. 2-38 In certain embodiments, provided herein are IFN ⁇ polypeptides according to any of SEQ ID NOS: 2-38 fused to HisSUMO SEQ ID NO: 39. [000486] Table 13. Sequence table Conjugate No. Name Amino acid sequence
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Medicinal Chemistry (AREA)
- General Health & Medical Sciences (AREA)
- Pharmacology & Pharmacy (AREA)
- Epidemiology (AREA)
- Animal Behavior & Ethology (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Engineering & Computer Science (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Organic Chemistry (AREA)
- Zoology (AREA)
- Gastroenterology & Hepatology (AREA)
- Biochemistry (AREA)
- Biophysics (AREA)
- Genetics & Genomics (AREA)
- Molecular Biology (AREA)
- Toxicology (AREA)
- Peptides Or Proteins (AREA)
- Medicinal Preparation (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
Abstract
The present disclosure generally relates to interferon alpha (IFNα) polypeptides comprising one or more non-natural amino acid or modified amino acid substitution mutations. The present disclosure is also directed to IFNα conjugates comprising IFNα polypeptides site-specifically linked to at least one masking moiety, optionally via a linker. Also provided are pharmaceutical compositions, diagnostic compositions and kits containing the polypeptides and conjugates disclosed herein, nucleic acids and expression vectors encoding the polypeptides disclosed herein, cells comprising the same, and methods of using the polypeptides, conjugates, nucleic acids, expression vectors, and cells for therapeutic, and diagnostic purposes.
Description
INTERFERON ALPHA POLYPEPTIDES AND CONJUGATES CROSS-REFERENCE TO RELATED APPLICATIONS [0001] This application claims the benefit of U.S. Provisional Application No.63/422,795 filed November 4, 2022, and U.S. Provisional Application No. 63/493,439 filed March 31, 2023. Each of these applications is incorporated for all purposes in their entireties. FIELD [0002] The present disclosure generally relates to interferon alpha (IFNα) polypeptides comprising one or more non-natural amino acid or modified amino acid substitution mutations. The present disclosure is also directed to IFNα conjugates comprising IFNα polypeptides site- specifically linked to at least one masking moiety, optionally via a linker. Also provided are pharmaceutical compositions, diagnostic compositions and kits containing the polypeptides and conjugates disclosed herein, nucleic acids and expression vectors encoding the polypeptides disclosed herein, cells comprising the same, and methods of using the polypeptides, conjugates, nucleic acids, expression vectors, and cells for therapeutic, and diagnostic purposes. BACKGROUND [0003] Type I interferons (IFN-1) are cytokines that play an important role in modulating the innate and adaptive immune response. First discovered as anti-viral agents, type I interferons are also now known to suppress the proliferation of cancer cells. The IFN-1 family comprises 17 functional genes on chromosome 9 that encode 16 proteins, IFNβ, IFNε, IFNκ, IFNω, and 12 subtypes of IFNα. IFNα is generally produced by leukocytes, while IFNβ is generally produced by fibroblasts, but both IFNα/β elicit an immune response by binding to the IFN-α receptor (IFNAR). [0004] While the 12 subtypes of IFNα exhibit slightly different binding affinities and subtle differences, the first and most widely studied of the IFNα subtypes is IFNα2. Several allelic variants of IFNα2 have been identified, including IFNα2a and IFNα2b, which differ only at residue 23 (lysine or arginine, respectively). [0005] Various forms of recombinant and pegylated type I interferons have been developed and approved as therapeutics. In 1986, recombinant IFNα2b (Intron A®) developed by Merck Sharp & Dohme was the first IFN-1 to gain approval in the United States. Intron A® is approved as a treatment for hairy cell leukemia, malignant melanoma, follicular lymphoma, condylomata 1
acuminate, AIDS-related Kaposi’s sarcoma, and chronic hepatitis B and C. Recombinant IFNα2a was developed by Hoffmann-La Roche Inc. as Roferon-A for various types of cancer, AIDS-related sarcoma, and hepatitis. Various pegylated forms of IFNα2a and IFNα2b have also been developed, including Pegasys (Peg IFNα2a) for chronic hepatitis B and C, Pegintron (Peg IFNα2b) for hepatitis C, and Sylatron (Peg IFNα2b) for melanoma. [0006] However, these currently marketed pegylated forms of IFNα2a and IFNα2b exhibit minimal attenuation of activity and are non-tumor specific. Thus, their clinical use is limited due to poor systemic tolerability. In fact, the use of IFNα recombinant and pegylated forms can be accompanied by severe adverse side effects. Most patients experience acute toxicity in the form of flu-like symptoms, including fever, chills, myalgia, headache, and nausea. Hematological side effects, including decreases in blood counts, are also commonly observed. Hepatic, gastrointestinal, neurological, cardiovascular, and endocrine side effects have also been reported (Sleijfer, S. et al., 2005, Pharmacy World and Science, 27, pages 423–431). Most significantly, FDA-approved alpha interferons come with a black box warning for potential development of life-threatening neuropsychiatric, autoimmune, ischemic, and infectious disorders. Peg IFNα2b Sylatron warns of serious depression and suicidal ideation. [0007] These harmful side effects can result in the administration of lower, less effective doses, and even the discontinuation of the therapy. Therefore, what are needed in the art are IFNα polypeptides and conjugates that are characterized by limited systemic activation and improved tumor-selectivity that provide effective and tolerable IFNα therapy for the treatment of cancer and other diseases. SUMMARY [0008] Provided herein are IFNα polypeptides and conjugates. The IFNα polypeptides comprise at least one non-natural amino acid or modified amino acid. In one embodiment, the IFNα polypeptides comprise at least one non-natural amino acid. In one embodiment, the IFNα polypeptides comprise at least one modified amino acid. The IFNα conjugates comprise IFNα polypeptides site-specifically conjugated to at least one masking moiety, optionally via a linker. In certain embodiments, while the IFNα conjugates are in systemic circulation, the masking moiety acts as a mask to attenuate IFNα activity and/or extend the half-life of the conjugate. Once the IFNα conjugate comes in contact with the tumor microenvironment (TME), the conjugate is cleaved from the masking moiety either preferentially by tumor-selective proteases enriched in the TME or the acidic nature of the TME. This restores the activity of the IFNα 2
polypeptide to participate in immune cell activation and tumor cell killing. The use of the masking moiety is advantageous for the delivery of IFNα and widens the therapeutic window because systemic activity and toxicity of the IFNα is limited, but tumor cell killing is selective and enhanced. Therefore, in certain embodiments, the IFNα conjugates and polypeptides described herein exhibit reduced toxicity, for example, systemic toxicity, compared to wild- type IFNα. In other embodiments, the IFNα conjugates and polypeptides described herein exhibit increased stability, for example in serum, compared to wild-type IFNα. In certain embodiments, the IFNα conjugates and polypeptides described herein exhibit reduced toxicity, for example, systemic toxicity, and increased stability, for example in serum, compared to wild- type IFNα. [0009] In one aspect, the disclosure provides IFNα polypeptides capable of binding the IFNα receptor ^IFNAR). In certain embodiments, the IFNα polypeptides comprise at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156. In certain embodiments, the IFNα polypeptides comprise at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In one embodiment, the non-natural amino acid or modified amino acid is a non-natural amino acid. In one embodiment, the non-natural amino acid or modified amino acid is a modified amino acid. In certain embodiments, the positions are in reference to human wild- type IFNα (SEQ ID NO: 33). In certain embodiments, the positions are in reference to mice wild-type IFNα (SEQ ID NO: 38). In certain embodiments, the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 33. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 33. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 33. In certain embodiments, the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 38. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 38. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 38. [00010] In another aspect, provided herein are IFNα conjugates comprising an IFNα polypeptide and a masking moiety wherein the IFNα polypeptide is site-specifically linked to the masking moiety, optionally via a linker. In certain embodiments, the IFNα conjugate comprises an IFNα polypeptide comprising at least one non-natural amino acid or modified 3
amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156 wherein the at least one non-natural amino acid or modified amino acid is linked to a masking moiety optionally via a linker. In certain embodiments, the IFNα conjugate comprises an IFNα polypeptide comprising at least one non- natural amino acid or modified amino acid at a position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 wherein the at least one non-natural amino acid or modified amino acid is linked to a masking moiety optionally via a linker. In one embodiment, the non- natural amino acid or modified amino acid is a non-natural amino acid. In one embodiment, the non-natural amino acid or modified amino acid is a modified amino acid. In another embodiment, the IFNα conjugate comprises an IFNα polypeptide and at least one masking moiety wherein the IFNα polypeptide is site-specifically linked to the at least one masking moiety via a protease cleavable linker and the water masking moiety is a water-soluble polymer or carbohydrate. In another embodiment, the IFNα conjugate comprises an IFNα polypeptide and at least one masking moiety wherein the IFNα polypeptide is site-specifically linked to the at least one masking moiety via a cathepsin B cleavable linker. In another embodiment, the IFNα conjugate comprises an IFNα polypeptide and at least one masking moiety wherein the IFNα polypeptide is site-specifically linked to the at least one masking moiety via a pH- sensitive linker. [00011] In certain embodiments, the IFNα conjugate comprises at least two masking moieties. In certain embodiments, the IFNα conjugate comprises at least three masking moieties. In certain embodiments, the IFNα conjugate comprises at least four masking moieties. In certain embodiments, the IFNα conjugate comprises at least five masking moieties. In certain embodiments, the IFNα conjugate comprises at least six or more masking moieties. In any of the foregoing embodiments, the masking moiety is attached the IFNα polypeptide via a linker as described herein. [00012] In another aspect, provided herein are polynucleotides encoding the IFNα polypeptides and/or fusion constructs described herein. In a further aspect, provided herein are expression vectors comprising the polynucleotides. In a further aspect, provided herein are cells comprising the polynucleotides or expression vectors. In some embodiments, the cells are selected from bacterial cells, fungal cells, and mammalian cells. In some embodiments, the cells are selected from E. coli cells, Saccharomyces cerevisiae cells, and CHO cells. In another aspect, provided herein are methods of making the IFNα polypeptides, for instance using the polynucleotides, expression vectors, and/or cells described herein. 4
[00013] In another aspect, provided herein are methods of treating, preventing, or diagnosing a disease or condition in a subject in need thereof, wherein the method includes administering to the subject an effective amount of the IFNα polypeptide, IFNα conjugate, or fusion construct of any of the foregoing embodiments, or a composition or a pharmaceutical composition containing the same. In some embodiments, the disease or condition is selected from a cancer, an autoimmune disease, an inflammatory disease, and an infection. In some embodiments, the effective amount is a therapeutically effective amount. In certain embodiments, the disease or condition is cancer. Anti-tumor immune memory is the immune system’s ability to recognize, or memorize, a previously encountered tumor antigen. If the previously encountered tumor antigen is recognized, the immune system can respond more quickly and aggressively (through T cell activation and proliferation) than the first response based on the memory of the initial encounter. In certain embodiments, a IFNα polypeptide, IFNα conjugate, or fusion construct described herein or a pharmaceutical composition thereof treats the disease or condition, for example, cancer, by activating anti-tumor immunity. In certain embodiments, a IFNα polypeptide, IFNα conjugate, or fusion construct described herein or a pharmaceutical composition thereof treats the disease or condition, for example, cancer, by inducing or enhancing anti-tumor immune memory. [00014] Embodiments disclosed herein are also directed to the use of the IFNα polypeptide, IFNα conjugates, or fusion constructs of any of the foregoing embodiments for treating, preventing, or diagnosing a disease or condition in a subject in need thereof. Embodiments disclosed herein are also directed to the IFNα polypeptides, IFNα conjugates, or fusion constructs of any of the foregoing embodiments for use in treating, preventing, or diagnosing a disease or condition in a subject in need thereof. Embodiments disclosed herein are also directed to the IFNα polypeptides, IFNα conjugates, or fusion constructs of any of the foregoing embodiments for use in the manufacture of a medicament for treating, preventing, or diagnosing a disease or condition in a subject in need thereof. In some embodiments, the disease or condition is selected from a cancer, an autoimmune disease, an inflammatory disease, and an infection. [00015] These and other embodiments along with many of its features are described in more detail in conjunction with the text below and attached figures. Other features, objects, and advantages will be apparent from the disclosure that follows. 5
BRIEF DESCRIPTION OF THE DRAWINGS [00016] FIG.1A is a graph of the effect of a 3 mg/kg dose of Conjugate 1, Conjugate 16 and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 41 post treatment. [00017] FIG.1B is a graph of the effect of a 10 mg/kg dose of Conjugate 1, Conjugate 16 and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 41 post treatment. [00018] FIG.1C is a graph of the effect of a 3 mg/kg dose of Conjugate 1, Conjugate 16 and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 44 post treatment. [00019] FIG.1D is a graph of the effect of a 15 mg/kg dose of Conjugate 1, Conjugate 16 and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 44 post treatment. [00020] FIG.1E is a graph of the effect of Conjugate 1, Conjugate 16 and Conjugate 18 (dosed at 3 mg/kg and 15 mg/kg) on MDA-MB-231 tumor size up until the end of the study at day 44 post treatment. [00021] FIG.2A is a graph of the effect of a 3 mg/kg dose of Conjugate 1, Conjugate 2, Conjugate 13, and Conjugate 18 on MDA-MB-231 tumor growth up. [00022] FIG.2B is a graph of the effect of a 1 mg/kg dose of Conjugate 1, Conjugate 2, Conjugate 13, and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 32 post treatment. [00023] FIG.2C is a graph of the effect of a 0.1 mg/kg dose of Conjugate 1, Conjugate 2, Conjugate 13, and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 32 post treatment. [00024] FIG.2D is a graph of the effect of a 0.05 mg/kg dose of Conjugate 1, Conjugate 2, Conjugate 13, and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 32 post treatment. [00025] FIG.2E is a graph of the effect of Conjugate 1, Conjugate 2, Conjugate 13, and Conjugate 18 (dosed at 0.05 mg/kg, 0.01 mg/kg, 1 mg/kg and 3 mg/kg) on MDA-MB-231 tumor size up until the end of the study at day 44 post treatment as described in Example 9. [00026] FIG.3A is a graph illustrating the serum concentrations of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 following intravenous administration at doses ranging from 3 mg/kg to 45 mg/kg. 6
[00027] FIG. 3B is a graph showing the PK profile of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 following an intravenous dose (15 mg/kg). [00028] FIG.3C is a graph showing percent body weight changes in hamsters following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg. [00029] FIG.3D is a graph showing percent body weight changes in hamsters following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at a dose of 15 mg/kg. [00030] FIG.3E is a graph illustrating ALT fold change as measured on day 2 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg. [00031] FIG.3F is a graph illustrating AST fold change as measured on day 2 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg. [00032] FIG.3G is a graph illustrating ALP fold change as measured on day 2 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg. [00033] FIG.3H is a graph illustrating ALT fold change as measured on day 7 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg. [00034] FIG.3I is a graph illustrating AST fold change as measured on day 7 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg. [00035] FIG.3J is a graph illustrating ALP fold change as measured on day 7 of the study following intravenous administration of Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 at doses ranging from 3 mg/kg to 45 mg/kg. [00036] FIG.3K and FIG.3L are graphs illustrating tolerability of HLE-Interferon variants in golden Syrian hamsters. [00037] FIG.4A is a graph of the effect of a 3 mg/kg dose of Conjugate 34, Conjugate 35, Conjugate 36, and Conjugate 37 on B16F10 tumor growth. [00038] FIG.4B is a graph of the effect of a 1 mg/kg dose of Conjugate 34 and Conjugate 36 on B16F10 tumor growth. [00039] FIG.4C is a graph of the effect of Conjugate 35 and Conjugate 37 (dosed with 3 mg/kg and 10 mg/kg) on B16F10 tumor growth. 7
[00040] FIG. 4D is a graph of the effect of Conjugate 35 and Conjugate 37 (dosed at 3 mg/kg and 10 mg/kg) on B16F10 tumor size. [00041] FIG.4E is a graph of the effect of Conjugate 37 administered at 3 mg/kg compared to the vehicle on the production of CD45 positive cells. [00042] FIG.4F is a graph of the effect of Conjugate 37 administered at 3 mg/kg compared to the vehicle on the level of Granzyme B secreted from CD8 T-cells. [00043] FIG.4G is a graph of the effect of Conjugate 37 administered at 3 mg/kg compared to the vehicle on the level of Granzyme B secreted from NK cells. [00044] FIG. 4H and FIG. 4I are graphs showing mouse surrogate efficacy and PD in MC38-HCEA. [00045] FIG.5A is a graph of dose response curves for human IFNα Conjugate 1, Conjugate 2, Conjugate 11, Conjugate 13, Conjugate 16, Conjugate 18, and Conjugate 31 compared to wildtype human IFNα (Conjugate 32) in HEK-blue human IFNα/β Reporter Assay. [00046] FIG. 5B is a graph of dose response curves for mouse IFN ^^ Conjugate 34, Conjugate 35, Conjugate 36, and Conjugate 37 compared to wildtype mouse IFNα (Conjugate 38) in B16-Blue mouse IFNα/β Reporter Assay. [00047] FIG.6 is a graph showing the PK of IFNα Conjugate 1, Conjugate 13, Conjugate 18, and Conjugate 33, in C57B1/6 mice. [00048] FIG. 7A, FIG. 7B, and FIG. 7C show the combination of Conjugate 37 with a checkpoint inhibitor in B16F10 synergist mouse tumor model. [00049] FIG.8 is an image showing how the IFNα conjugates of the present description are masked in circulation, but unmasked in the tumor microenvironment (TME). [00050] FIG. 9A shows the SDS-PAGE gel showing expression and purification of 3XnnAA-IFNα. From left to right: Soluble lysate from 200 mL fermenter expression of 3XnnAA-IFNα, soluble lysate from 200 mL fermenter (+ AS tRNA = using a product plasmid with an extra copy of the AS tRNA) expression of 3XnnAA-IFNα, representative sample of purified HisSUMO-3XnnAA-IFNα (non-reducing), representative sample of purified HisSUMO-3XnnAA-IFNα (reducing). [00051] FIG. 9B is the analytical SEC chromatogram showing purity of final PEGylated 3XnnAA-IFNα. [00052] FIG. 9C is a Hek-Blue assay of in vitro cell activation with IFNα samples. A commercial WT IFNα standard (orange diamonds) was compared to 3XnnAA-IFNα (green triangles) and PEGylated 3XnnAA-IFNα (red squares). For this assay, increases in OD630 corresponds to cell activation. 8
[00053] FIG.9D is a table summarizing strains tested, fermentation format and scales, titers, and product quality metrics for 3XnnAA-IFNα proteins. Strains with a product plasmid bearing an extra copy of AS tRNA are denoted as “+ AS tRNA” while strains bearing a product plasmid with only the 3XnnAA-IFNα coding sequence are denoted as “No AS tRNA”. The % full length 3XnnAA-IFNα was calculated by intact LC-MS analysis, purity % was assessed by analytical SEC, and PEG-to-protein ratio was calculated using SDS-PAGE gel densitometry. [00054] FIG. 10A, FIG. 10B, and FIG. 10C show expression plasmids and shake flask analysis of 3x-nnAA-IFNα. The high copy product plasmid used for the initial expression of 3X-nnAA-IFNα in shake flasks is shown in FIG.10A. The high copy product plasmid (+ AS tRNA) used to increase tRNA concentration/amber suppression for the expression of 3X- nnAA-IFNα. This is the final plasmid used in high density fermentations is shown in FIG.10B. The SDS-PAGE gel analysis of HisSUMO-IFNα and 3X-nnAA-IFNα in shake flasks is shown in FIG.10C. Lanes from left to right: 1) Uninduced lysate, 2) wild type HisSUMO-IFNα, 3) 3X-nnAA-IFNα expressed using plasmid A (without extra AS tRNA on the product plasmid), 4) 3X-nnAA-IFNα expressed using plasmid B (+ AS tRNA on the product plasmid). The arrow denotes the presence of the SUMO-tagged IFNα band. [00055] FIG. 11A, FIG. 11B, and FIG. 11C show intact LC-MS analysis of 3X-nnAA- IFNα proteins expressed with either product plasmid and include the deconvoluted LC-MS spectrum of purified and Ulp1-cleaved IFNα that has been conjugated with a small molecule DBCO-amine. FIG. 11A shows 3X-nnAA-IFNα produced in shake flasks with a product plasmid bearing an extra copy of the AS tRNA (product plasmid + AS tRNA, see FIG.10B). FIG.11B shows 3X-nnAA-IFNα produced in shake flasks with a product plasmid with no extra copy of the AS tRNA (product plasmid, see FIG.10A). FIG.11C shows 3X-nnAA-IFNα produced in high density fermentation with a product plasmid bearing an extra copy of the AS tRNA (product plasmid + AS tRNA, see FIG.10B). Individual peaks in each LC-MS spectrum are labeled. [00056] FIG. 12A is a graph showing increased Granzyme B levels in tumor infiltrating CD8 T-cells harvested from MC38-hTrop2 tumor bearing mice following treatment with a single IV dose of 1 mg/kg of Conjugate 37 or Conjugate 34 compared to the vehicle. [00057] FIG.12B is a graph showing increased Granzyme B levels in tumor infiltrating NK cells harvested from MC38-hTrop2 tumor bearing mice following treatment with a single IV dose of 1 mg/kg of Conjugate 37 or Conjugate 34 compared to the vehicle. 9
[00058] FIG. 12C is a graph showing increased activation of monocytes following treatment with a single IV dose of 1 mg/kg of Conjugate 37 or Conjugate 34 compared to the vehicle. [00059] FIG. 12D is a graph showing increased activation of dendritic cells following treatment with a single IV dose of 1 mg/kg of Conjugate 37 or Conjugate 34 compared to the vehicle. [00060] FIG.12E is a graph showing increased activation of plasmacytoid dendritic cells following treatment with a single IV dose of 1 mg/kg of Conjugate 37 or Conjugate 34 compared to the vehicle. [00061] FIG.12F is a lymph node analysis showing that treatment with Conjugate 34 results in similar increase in levels of GranzymeB in CD8 T-cells from both tumor-draining and non- draining lymph node, while treatment with Conjugate 37 results in a greater increase in GranzymeB levels in CD8 T-cells from tumor-draining lymph node compared to non-draining lymph nodes. [00062] FIG. 13 is a showing the tumor size of complete responder mice obtained from Conjugate 37 treatment that were treated with 300 ^g anti-CD8 antibody or Isotype antibody and rechallenged with 5x106 MC38-hCEA cells on D0. When rechallenged, Conjugate 37- treated complete responder mice that received Isotype control antibody demonstrated no recurrence of tumors, while CD8 depletion ablated Conjugate 37-induced anti-tumor immune memory. DETAILED DESCRIPTION [00063] Provided herein are IFNα polypeptides, IFNα conjugates, fusion constructs and compositions comprising the same. The IFNα polypeptides comprise at least one non-natural amino acid or modified amino acid substitution. The IFNα conjugates comprise an IFNα polypeptide site-specifically linked to a masking moiety, optionally via a linker. As disclosed herein, the site-specific linkage to the masking moiety provides an extended half-life of the conjugates. The masking moiety acts as a mask to attenuate IFNα activity while the IFNα conjugate is in systemic circulation but once the masked-IFNα conjugate comes in contact with the tumor microenvironment (TME), tumor-selective proteases or the acidic environment of the TME cleave the masking moiety to activate the IFNα polypeptide for immune cell activation and tumor cell killing. Therefore, the IFNα conjugates bonded to a masking moiety using site-specific conjugation are advantageous for the delivery of IFNα because the 10
polypeptides exhibit less systemic toxicity, while simultaneously exhibiting enhanced and selective tumor cell killing. In certain embodiments, these IFNα polypeptides exhibit improved characteristics, for example reduced toxicity and increased stability, relative to a wild type (i.e., parent) IFNα. [00064] Also provided herein are methods of treating, preventing, or diagnosing a disease or condition, for example, cancer, in a subject in need thereof, comprising administering to the subject an effective amount of an IFNα polypeptide or IFNα conjugate described herein or a pharmaceutical composition thereof. In certain embodiments, a IFNα polypeptide or a IFNα conjugate described herein or a pharmaceutical composition thereof treats the disease or condition, for example, cancer, by activating anti-tumor immunity. For example, as described in Example 17, certain IFNα conjugates described herein activated the innate and adaptive immune system when administered to tumor-bearing mice; analysis of the TME cell types showed that administration of the IFNα conjugates led to increased levels of Granzyme B levels in tumor infiltrating CD8 T-cells and NK cells and increased activation of multiple innate immune cells in the TME including monocytes, dendritic cells and plasmacytoid dendritic cells. Further, in certain embodiments, a IFNα polypeptide or a IFNα conjugate described herein or a pharmaceutical composition thereof treats the disease or conditions, for example, cancer, by inducing or enhancing anti-tumor immune memory via T cell activation and proliferation. As described in Example 18, tumor free and complete responder mice obtained from previous treatment with an IFNα conjugate described herein demonstrated no reoccurrence of tumors after receiving an isotype antibody when rechallenged with the tumors. 1.1. Definitions [00065] Unless otherwise defined, all terms of art, notations and other scientific terminology used herein are intended to have the meanings commonly understood by those of skill in the art to which this invention pertains. In some cases, terms with commonly understood meanings are defined herein for clarity and/or for ready reference, and the inclusion of such definitions herein should not necessarily be construed to represent a difference over what is generally understood in the art. The techniques and procedures described or referenced herein are generally well understood and commonly employed using conventional methodologies by those skilled in the art, such as, for example, the widely utilized molecular cloning methodologies described in Green & Sambrook, Molecular Cloning: A Laboratory Manual 4th ed. (2012) Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, or in Current Protocols in Molecular Biology (2022), Wiley Periodicals LLC. As appropriate, procedures 11
involving the use of commercially available kits and reagents are generally carried out in accordance with manufacturer-defined protocols and conditions unless otherwise noted. [00066] As used herein, the singular forms “a,” “an,” and “the,” include the plural referents unless the context clearly indicates otherwise. [00067] The term “about,” as used herein, indicates and encompasses an indicated value and a range above and below that value. In certain embodiments, the term “about” indicates the designated value ± 10%, ± 5%, or ± 1%. In certain embodiments, the term “about” indicates the designated value ± one standard deviation of that value. [00068] The term “combinations thereof,” as used herein, includes every possible combination of elements to which the term refers to. [00069] Ranges: throughout this disclosure, various aspects of the disclosure can be presented in a range format. It should be understood that the description in range format is merely for convenience and brevity and should not be construed as an inflexible limitation on the scope of the disclosure. Accordingly, the description of a range should be considered to have specifically disclosed all the possible subranges as well as individual numerical values within that range. For example, description of a range such as from 1 to 6 should be considered to have specifically disclosed subranges such as from 1 to 3, from 1 to 4, from 1 to 5, from 2 to 4, from 2 to 6, from 3 to 6 etc., as well as individual numbers within that range, for example, 1, 2, 2.7, 3, 4, 5, 5.3, and 6. This applies regardless of the breadth of the range. [00070] IFNα is a key part of the innate immune response with potent antiviral, antiproliferative and immunomodulatory properties. IFNα, like other type I interferons, binds a plasma membrane receptor made of IFNAR1 and IFNAR2 that is ubiquitously expressed in the human body. In humans, IFNα gene is located on chromosome 9. There are 12 subtypes of IFNα. In certain embodiments, the term “human interferon-alpha,” or “human IFNα,” or “hIFNα,” as used herein, refers to IFNα2 having an amino acid sequence according to amino acids 24-188 of UniProt Accession No. P01563 (SEQ ID NO: 33). A representative active form IFNα2 sequence is provided by SEQ ID NO: 33: CDLPQTHSLG SRRTLMLLAQMRRISLFSCLKDRHDFGFPQEEFGNQFQKA ETIPVLHEMI QQIFNLFSTKDSSAAWDETLLDKFYTELYQQLNDLEACVI QGVGVTETPL MKEDSILAVRKYFQRITLYLKEKKYSPCAWEVVRAEIMRS FSLSTNLQES LRSKE. [00071] The IFNAR is composed of two subunits: IFNAR1 and IFNAR2. Binding to either the IFNAR1 or IFNAR2 subunit by an IFN-1, including IFNα, brings the two subunits close 12
together and activates the JAK-STAT signaling cascade. This results in the activation of interferon-regulated genes that induce an interferon response. The term “human interferon alpha/beta receptor 1” or “human IFN-R-1” as used herein, refers to the IFNAR1 subunit of the receptor for IFNα encoded by the IFNAR1 gene. Representative IFN-R-1 sequences are provided by UniProt. Accession No. P17181. [00072] The term “human interferon alpha/beta receptor 2” or “human IFN-R-2” as used herein, refers to the IFNAR2 subunit of the receptor for IFNα encoded by the IFNAR2 gene. Representative IFN-R-2 sequences are provided by UniProt. Accession No. P48551. [00073] The term “operably-linked,” as used herein, refers to a functional linkage between two elements, regardless of orientation or distance between the two elements, such that the function of one element is controlled or affected by the other element. For example, operable linkage with reference to a promoter and heterologous coding sequence means that the transcription of the heterologous coding sequence is under the control of, or driven by, the promoter. In another example, operable linkage with reference to an enhancer and promoter means that the enhancer increases the level of transcription driven by a promoter. [00074] The term “isolated,” as used herein, refers to a substance that has been separated and/or recovered from its natural environment. For example, a naturally occurring polynucleotide or polypeptide present in a living animal is not isolated, but the same polynucleotide or polypeptide, separated from some or all of the co-existing materials in the natural system, is isolated. Such polynucleotide could be part of a vector and/or such polynucleotide or polypeptide could be part of a composition (e.g., a cell lysate), and still be isolated in that such vector or composition is not part of the natural environment for the nucleic acid or polypeptide. [00075] An “isolated IFNα variant” or “isolated IFNα polypeptide” is one that has been separated and/or recovered from a component of its natural environment. Components of the natural environment may include enzymes, hormones, and other proteinaceous or nonproteinaceous materials. In some embodiments, an isolated IFNα polypeptide is purified to a degree sufficient to obtain at least 15 residues of N-terminal or internal amino acid sequence, for example by use of a spinning cup sequenator. In some embodiments, an isolated IFNα polypeptide is purified to homogeneity by gel electrophoresis (e.g., SDS-PAGE) under reducing or nonreducing conditions, with detection by Coomassie blue or silver stain. In some aspects, an isolated IFNα polypeptide is prepared by at least one purification step. [00076] The term “substantially pure” with respect to a composition comprising a polypeptide IFNα refers to a composition that includes at least 80%, 85%, 90% or 95% by 13
weight or, in certain embodiments, 95%, 98%, 99% or 100% by weight, e.g., dry weight, of the IFNα polypeptide relative to the remaining portion of the composition. The weight percentage can be relative to the total weight of protein in the composition or relative to the total weight of IFNα polypeptide in the composition. Purity can be determined by techniques apparent to those of skill in the art, for instance SDS-PAGE. [00077] In some embodiments, an isolated IFNα polypeptide is purified to at least 80%, 85%, 90%, 95%, or 99% by weight. In some embodiments, an isolated IFNα polypeptide is purified to at least 80%, 85%, 90%, 95%, or 99% by volume. In some embodiments, an isolated IFNα polypeptide is provided as a solution comprising at least 85%, 90%, 95%, 98%, 99% to 100% by weight. In some embodiments, an isolated IFNα polypeptide is provided as a solution comprising at least 85%, 90%, 95%, 98%, 99% to 100% by volume. [00078] “Affinity” refers to the strength of the sum total of non-covalent interactions between a single binding site of a molecule (e.g., IFNα) and its binding partner (e.g., IFNAR). Unless indicated otherwise, as used herein, “binding affinity” refers to intrinsic binding affinity, which reflects a 1:1 interaction between members of a binding pair (e.g., IFNα and IFNAR, subunit IFNAR1 or IFNAR2). The affinity of a molecule X for its partner Y can be represented by the dissociation constant (KD). Affinity can be measured by common methods known in the art, including those described herein. Affinity can be determined, for example, using surface plasmon resonance (SPR) technology, such as a Biacore® instrument. In some embodiments, the affinity is determined at 25°C. [00079] With regard to the binding of receptor to a target molecule (ligand), the terms “specific,” “specific binding,” “specifically binds to,” “specific for,” “selectively binds,” or “selective for,” as used herein, refers to a particular receptor or ligand that exhibits binding that is measurably different from a non-specific or non-selective interaction. Specific binding can be measured, for example, by determining binding of a molecule compared to binding of a control molecule. Specific binding can also be determined by competition with a control molecule that competes with the ligand for binding to the receptor. In that case, specific binding is indicated if the binding of the ligand to the receptor is competitively inhibited by the control molecule. [00080] The term “kd” (sec-1), as used herein, refers to the dissociation rate constant of a particular receptor-ligand interaction. This value is also referred to as the koff value. [00081] The term “ka” (M-1×sec-1), as used herein, refers to the association rate constant of a particular receptor-ligand interaction. This value is also referred to as the kon value. 14
[00082] The term “KD” (M), as used herein, refers to the dissociation equilibrium constant of a particular receptor-ligand interaction. KD = kd/ka. [00083] The term “KA” (M-1), as used herein, refers to the association equilibrium constant of a particular receptor-ligand interaction. KA = ka/kd. [00084] The term “Tm” as used herein, has the meaning commonly understood in the art and refers to is the temperature at which the equilibrium between folded and unfolded forms of the enzyme is at its mid-point. [00085] The term “EC50” or “half maximal effective concentration” as used herein, has the meaning commonly understood in the art and refers to the concentration of a substance e.g., a drug, e.g., an IFNα polypeptide, which induces a response halfway between the baseline and maximum after a specified exposure time. Thus, EC50 can be defined as the concentration required to obtain a 50% effect and represents the concentration of a compound where 50% of its maximal effect is observed. [00086] The term “half-life” or “t1/2” as used herein refers to the amount of time required for the drug concentration measured in a sample to be reduced to half of its starting concentration or amount. The term “terminal t1/2” as used herein, refers to the amount of time required for the drug concentration measured in a sample to be reduced to half of its pseudo- equilibrium concentration or amount. [00087] The term “C0” as used herein, has the meaning commonly understood in the art and refers to the plasma concentration at the time of dosing (time 0). [00088] The term “AUC” as used herein, has the meaning commonly understood in the art of pharmacokinetics, and refers to the area under the plasma drug concentration-time curve (AUC) and reflects the measure of how much drug reaches an individual’s bloodstream in a given period of time after a dose is given. AUC is dependent on the rate of elimination of the drug from the body and the dose administered. AUC is directly proportional to the dose when the drug follows linear kinetics and is inversely proportional to the clearance of the drug. [00089] The term “AUC0-last” as used herein, has the meaning commonly understood in the art of pharmacokinetics, and refers to the AUC from dosing (time 0) to the last measurable concentration. [00090] The term “AUC0-inf” as used herein, has the meaning commonly understood in the art of pharmacokinetics, and refers to the AUC from dosing (time 0) extrapolated to infinity. 15
[00091] The term “clearance” as used herein, refers to the rate at which an active drug e.g., an IFNα polypeptide as disclosed herein, is removed from the body. “Clearance” is typically reported as the ratio of the elimination rate of a drug to the plasma drug concentration [00092] The term “Vss” as used herein, has the meaning commonly understood in the art and refers to the apparent volume of distribution at steady state which describes the physiological distribution of the drug candidate. [00093] The term “steady state” as used herein has the meaning commonly understood in the art of pharmacokinetics, and refers to the condition when the administration of a drug and the clearance are balanced, creating a plasma concentration that is unchanged by time. [00094] The terms “protein,” “peptide,” and “polypeptide” are used interchangeably herein, and refer to a polymer of amino acid residues linked together by peptide (amide) bonds. Although the terms refer to a protein, peptide, or polypeptide of any size, structure, or function. Typically, a peptide will be at least three amino acids long and equal to or less than about 10 amino acids in length. A polypeptide is typically greater than 10 amino acids in length. A protein, peptide, or polypeptide may refer to an individual protein or a collection of proteins. One or more of the amino acids in a protein, peptide, or polypeptide may be modified, for example, by the addition of a chemical entity such as a carbohydrate group, a hydroxyl group, a phosphate group, a farnesyl group, an isofarnesyl group, a fatty acid group, a linker for conjugation, functionalization, or other modification, etc., or may be substituted with a non- natural amino acid. A protein, peptide, or polypeptide may also be a single molecule or may be a multi-molecular complex. A protein, peptide, or polypeptide may be just a fragment of a naturally occurring protein or peptide. A protein, peptide, or polypeptide may be naturally occurring, recombinant, or synthetic, or any combination thereof. A protein may comprise different domains, for example, a protein binding domain and a catalytic domain. Any of the proteins provided herein may be produced by any method known in the art. For example, the proteins provided herein may be produced via recombinant protein expression and purification. Methods for recombinant protein expression and purification are well known, and include those described by Green and Sambrook, Molecular Cloning: A Laboratory Manual (4th ed., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. (2012)), the entire contents of which are incorporated herein by reference. [00095] A “mutation” as used herein, refers to a change in nucleic acid or polypeptide sequence relative to a reference sequence. A mutation may comprise a substitution of a residue within a sequence, e.g., a nucleic acid or amino acid sequence, with another residue, or a 16
deletion or insertion of one or more residues within a sequence. Mutations are typically described herein by identifying the original residue followed by the position of the residue within the sequence and by the identity of the newly substituted residue. Various methods for making the amino acid substitutions (mutations) provided herein are well known in the art, and are provided by, for example, Green and Sambrook, Molecular Cloning: A Laboratory Manual (4th ed., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. (2012)). [00096] As noted above, a mutation can be a “substitution” mutation wherein the amino acid, or nucleotide at a particular position in a reference sequence is substituted with a different amino acid or nucleotide at that position in the amino acid or nucleic acid sequence. In some embodiments, a substitution replaces one amino acid at a specific location in a polypeptide or protein sequence for a different amino acid in that position of the polypeptide or protein sequence. In some embodiments, a “substitution” replaces a natural amino acid at a specific location in a polypeptide or protein sequence for a non-natural amino acid or modified amino acid in that position of the polypeptide or protein sequence. Thus, the term “substitution” as used herein, refers to as “substitution” mutation as disclosed herein above. [00097] The term “reversion mutation,” or “reversion” as used herein, refers to a particular type of substitution mutation wherein a polypeptide or nucleic acid sequence having a substitution mutation at a specific position in the sequence, acquires a mutation at that specific position that restores the original sequence. Thus, in some embodiments, a polypeptide sequence having a mutation at a specific position in the polypeptide sequence acquires a mutation that restores the amino acid at that specific position to the amino acid found in the reference sequence e.g., restores the wild-type sequence). [00098] The term “wild-type” or “parent” refers to a naturally occurring gene or protein. These include a naturally occurring IFNα gene or protein. [00099] The term “variant” or “mutant” as used herein, refers to a nucleic acid or polypeptide sequence having at least one mutation relative to a reference sequence. Accordingly, a “variant” or “mutant” typically has less than 100% sequence identity to a reference sequence. [000100] The terms “identical,” or “percent identity,” in the context of two or more polypeptide or nucleic acid molecule sequences, means two or more sequences or subsequences that are the same or have a specified percentage of amino acid residues or nucleotides that are the same over a specified region (e.g., 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% identity), when compared and aligned for maximum correspondence over a comparison window, or designated region, as measured using 17
methods known in the art, such as a sequence comparison algorithm, by manual alignment, or by visual inspection. Alignment for purposes of determining percent amino acid sequence identity can be achieved in various ways that are within the skill in the art, for instance, using publicly available computer software such as BLAST (Altschul et al. Nucleic Acids Res.2007, 25, 3389-3402), BLAST-2, ALIGN, MEGALIGN (DNASTAR), CLUSTALW, CLUSTAL OMEGA, or MUSCLE software. Those skilled in the art can determine appropriate parameters for aligning sequences, including any algorithms needed to achieve maximal alignment over the full length of the sequences being compared. Within the context of this disclosure, it is understood that where sequence analysis software is used for analysis, the results of the analysis are based on the “default values” of the program referenced. “Default values” mean any set of values or parameters which originally load with the software when first initialized. [000101] As applied to polypeptides, the term “substantial similarity” or “substantially similar” means that two peptide sequences, when optimally aligned, such as by the programs GAP or BESTFIT using default gap weights, share at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.5%, or 100% sequence identity. In some aspects, residue positions, which are not identical, differ by conservative amino acid substitutions. [000102] The term “amino acid,” as used herein, refers to a D- or L-natural or non-naturally occurring amino acid, including, but not limited to, the twenty common naturally occurring amino acids. Naturally occurring amino acids include alanine (Ala; A), arginine (Arg; R), asparagine (Asn; N), aspartic acid (Asp; D), cysteine (Cys; C); glutamic acid (Glu; E), glutamine (Gln; Q), Glycine (Gly; G); histidine (His; H), isoleucine (Ile; I), leucine (Leu; L), lysine (Lys; K), methionine (Met; M), phenylalanine (Phe; F), proline (Pro; P), serine (Ser; S), threonine (Thr; T), tryptophan (Trp; W), tyrosine (Tyr; Y), and valine (Val; V). Representative amino acids include, but are not limited to, alanine, β-alanine, arginine, asparagine, aspartic acid, cysteine, cystine, glutamic acid, glutamine, glycine, phenylalanine, histidine, isoleucine, lysine, leucine, methionine, proline, serine, threonine, valine, tryptophan, or tyrosine, among others. The term “amino acid” also includes “non-natural amino acids” (nnAA) and “modified amino acids.” The terms “non-natural amino acids” and “modified amino acids” are used herein interchangeably. [000103] Naturally encoded amino acids are the proteinogenic amino acids known to those of skill in the art. They include the 20 common amino acids (alanine, arginine, asparagine, 18
aspartic acid, cysteine, glutamine, glutamic acid, glycine, histidine, isoleucine, leucine, lysine, methionine, phenylalanine, proline, serine, threonine, tryptophan, tyrosine, and valine) and the less common pyrrolysine and selenocysteine. Naturally encoded amino acids include post- translational variants of the 22 naturally occurring amino acids such as prenylated amino acids, isoprenylated amino acids, myristoylated amino acids, palmitoylated amino acids, N-linked glycosylated amino acids, O-linked glycosylated amino acids, phosphorylated amino acids and acylated amino acids. [000104] A “conservative substitution,” or a “conservative amino acid substitution,” as used herein, refers to the substitution of an amino acid with a chemically or functionally similar amino acid. Conservative substitution tables providing similar amino acids are well known in the art. Polypeptide sequences having such substitutions are known as “conservatively modified variants.” Such conservatively modified variants are in addition to and do not exclude polymorphic variants, interspecies homologs, and alleles. By way of example, the groups of amino acids provided in Tables 1-3 are, in some embodiments, considered conservative substitutions for one another. Table 1. Selected groups of amino acids that are considered conservative substitutions
19
Table 2. Additional selected groups of amino acids that are considered conservative substitutions for one another, in certain embodiments. Group 1 A, S, and T Group 2 D and E Group 3 N and Q Group 4 R and K Group 5 I, L, and M Group 6 F, Y, and W Table 3. Further selected groups of amino acids that are considered conservative substitutions for one another, in certain embodiments. Group A A and G Group B D and E Group C N and Q Group D R, K, and H Group E I, L, M, V Group F F, Y, and W Group G S and T Group H C and M [000105] Additional conservative substitutions may be found, for example, in Creighton, Proteins: Structures and Molecular Properties 2nd ed. (1993) W. H. Freeman & Co., New York, NY. A polypeptide or protein generated by making one or more conservative substitutions of amino acid residues in a parent polypeptide or protein is referred to as a “conservatively modified variant.” [000106] The term “modified amino acid” (or “non-natural amino acid”) refers to an amino acid that is not a proteinogenic amino acid, or a post-translationally modified variant thereof. In particular, the term refers to an amino acid that is not one of the 20 common amino acids or pyrrolysine or selenocysteine, or post-translationally modified variants thereof. Exemplary modified amino acids include e.g., p-acetylphenylalanine (pAcF), azido-lysine (AzK), and p- azidomethyl-L-phenylalanine (pAMF). Additional non-limiting examples include O-methyl- L-tyrosine, 3-methyl-phenylalanine, O-4-allyl-L-tyrosine, 4-propyl-L-tyrosine, fluorinated phenylalanine, isopropyl-L-phenylalanine, p-azido-L-phenylalanine, p-acyl-L-phenylalanine, p-A benzoyl-L-phenylalanine, p-iodo-phenylalanine, p-bromophenylalanine, p-amino-L- phenylalanine, isopropyl-L-phenylalanine, and p-propargyloxy-phenylalanine. Modified amino acids, such as pAcF, AzK, and pAMF, provide side chains to which various secondary molecules e.g., polyethyleneglycol (PEG) can be conjugated/bound. In preferred embodiments, a modified amino acid is pAMF. pAMF is typically incorporated into proteins at the TAG 20
amber codon using method known in the art (see e.g., Zimmerman, E. S. et al. Bioconjug. Chem. 25, 351–361 (2014)). pAMF incorporation provides an efficient approach for site- specific modification of the proteins and subsequent conjugation-site specific modification. [000107] The term “non-natural amino acid” (or “unnatural amino acid”) or “synthetic amino acids” are α, β, γ, or δ amino acids, also includes but is not limited to, amino acids found in proteins, i.e., glycine, alanine, valine, leucine, isoleucine, methionine, phenylalanine, tryptophan, proline, serine, threonine, cysteine, tyrosine, asparagine, glutamine, aspartate, glutamate, lysine, arginine and histidine. In certain embodiments, the amino acid is in the L- configuration. Alternatively, the amino acid can be a derivative of alanyl, valinyl, leucinyl, isoleucinyl, prolinyl, phenylalaninyl, tryptophanyl, methioninyl, glycinyl, serinyl, threoninyl, cysteinyl, tyrosinyl, asparaginyl, glutaminyl, aspartoyl, glutaroyl, lysinyl, argininyl, histidinyl, β-alanyl, β-valinyl, β-leucinyl, β-isoleuccinyl, β-prolinyl, β-phenylalaninyl, β-tryptophanyl, β-methioninyl, β-glycinyl, β-serinyl, β-threoninyl, β-cysteinyl, β-tyrosinyl, β-asparaginyl, β- glutaminyl, β-aspartoyl, β-glutaroyl, β-lysinyl, β-argininyl or β-histidinyl. Unnatural amino acids are not proteinogenic amino acids, or post-translationally modified variants thereof that either occur naturally or are chemically synthesized. In particular, the term unnatural amino acid refers to an amino acid that is not one of the 20 common amino acids or pyrrolysine or selenocysteine, or post-translationally modified variants thereof. Non-limiting examples of unnatural amino acids include sulfoalanine, hydroxyproline (Hyp), beta-alanine, citrulline (Cit), ornithine (Orn), norleucine (Nle), 3-nitrotyrosine, nitroarginine, pyroglutamic acid (Pyr), naphtylalanine (Nal), 2,4-diaminobutyric acid (DAB), methionine sulfoxide, and methionine sulfone. [000108] The term “disease,” or “disease or disorder” as used herein, refers any condition or disorder that damages or interferes with the normal function of a cell, tissue, or organ. Examples of diseases include neoplasia, pathogen infection of cell, etc. [000109] As used herein, “treating” or “treatment” of any disease or disorder refers, in certain embodiments, to ameliorating a disease or disorder that exists in a subject. “Treating” or “treatment” includes ameliorating at least one physical parameter, which may be indiscernible by the subject. In yet another embodiment, “treating” or “treatment” includes modulating the disease or disorder, either physically (e.g., stabilization of a discernible symptom) or physiologically (e.g., stabilization of a physical parameter) or both. In yet another embodiment, “treating” or “treatment” includes delaying or preventing the onset of the disease or disorder. For example, in an exemplary embodiment, the phrase “treating cancer” refers to inhibition of 21
cancer cell proliferation, inhibition of cancer spread (metastasis), inhibition of tumor growth, reduction of cancer cell number or tumor growth, decrease in the malignant grade of a cancer (e.g., increased differentiation), or improved cancer-related symptoms. Further, as used herein, “treatment” includes preventing or delaying the recurrence of the disease, delaying or slowing the progression of the disease, ameliorating the disease state, providing a remission (partial or total) of the disease, decreasing the dose of one or more other medications required to treat the disease, delaying the progression of the disease, increasing or improving the quality of life, increasing weight gain, and/or prolonging survival. Also encompassed by “treatment” is a reduction of pathological consequence of cancer. [000110] As used herein, the term “anti-tumor immune memory” refers to the immune system’s ability to recognize a previously encountered tumor antigen, and through T cell activation and proliferation, mount a stronger and faster response to the tumor antigen compared to the first encounter based on the memory of the first encounter. [000111] As used herein, the term “therapeutically effective amount” or “effective amount” refers to an amount of a substance e.g., an IFNα polypeptide or IFNα conjugate disclosed herein, or a composition comprising a substance, that when administered to a subject is effective to treat a disease or disorder. For example, in an exemplary embodiment, the phrase “effective amount” is used interchangeably with “therapeutically effective amount” or “therapeutically effective dose” and the like, and means an amount of a therapeutic agent that is effective to prevent or ameliorate a disease or the progression of the disease e.g., cancer, or result in amelioration of symptoms. Effective amounts of the compositions provided herein may vary according to factors such as the disease state, age, sex, weight of the animal or human. [000112] The term “subject,” as used herein, refers to a mammalian subject. Exemplary subjects include, but are not limited to humans, monkeys, dogs, cats, mice, rats, cows, pigs, horses, camels, avians, goats, and sheep. In certain embodiments, the subject is a human. In some embodiments, the subject has a disease that can be treated with an IFNα polypeptide or IFNα conjugate provided herein. [000113] The term “therapeutically effective amount,” or “effective amount” as used herein, refers to the amount of the subject compound or composition that will elicit the biological, physiologic, clinical or medical response of a cell, tissue, organ, system, or subject that is being sought by the researcher, veterinarian, medical doctor or other clinician. The term “therapeutically effective amount” refers to an amount of a compound e.g., an IFNα polypeptide or IFNα conjugate, or composition that, when administered, is sufficient to prevent development of, or treat at least to some extent, one or more of the signs or symptoms of the 22
disorder or disease being treated. The therapeutically effective amount will vary depending on the compound or composition, the disease and its severity and the age, weight, etc., of the subject to be treated. [000114] The term “pharmaceutical composition,” as used herein, refers to a composition that can be administrated to a subject in the context of treatment of a disease or disorder. In some embodiments, a pharmaceutical composition comprises an active ingredient, e.g., an IFNα polypeptide as disclosed herein, and a pharmaceutically acceptable excipient. 1.2. IFNα Polypeptides [000115] Provided herein are IFNα polypeptides that comprise at least one non-natural amino acid or modified amino acid substitution compared to a wild type IFNα. The IFNα can be any IFNα known to the person of skill. In certain embodiments, the IFNα is any IFNα subtype. In certain embodiments, the IFNα is IFNα2. In some embodiments, the IFNα polypeptides comprise at least two non-natural amino acid or modified amino acid substitutions. In some embodiments, the IFNα polypeptides comprise at least three, four, five, six, or more non- natural amino acid or modified amino acid substitutions. [000116] The at least one non-natural amino acid or modified amino acid substitution can be made by standard techniques. In certain embodiments, the substitution is made by one or more mutations in the genetic sequence encoding the IFNα polypeptide. [000117] In certain embodiments, the IFNα polypeptide comprises at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156. In certain embodiments, the IFNα polypeptide comprises at least two non-natural amino acids or modified amino acids at positions selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156. In certain embodiments, the IFNα polypeptide comprises at least three non-natural amino acids or modified amino acids at positions selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156. In certain embodiments, the IFNα polypeptide comprises at least four non-natural amino acids or modified amino acids at positions selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156. In certain embodiments, 23
the IFNα polypeptide comprises at least five or more non-natural amino acids or modified amino acids at positions selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156. [000118] In some embodiments, the IFNα polypeptide comprises a non-natural amino acid or modified amino acid in at least one amino acid position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFNα polypeptide comprises at least two non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFNα polypeptide comprises at least three non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFNα polypeptide comprises at least four non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFNα polypeptide comprises at least five non- natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. [000119] In some embodiments, the IFNα polypeptide comprises a non-natural amino acid or modified amino acid at H7. In some embodiments, the IFNα polypeptide comprises a non- natural amino acid or modified amino acid at Q40. In some embodiments, the IFNα polypeptide comprises a non-natural amino acid or modified amino acid at E41. In some embodiments, the IFNα polypeptide comprises a non-natural amino acid or modified amino acid at N45. In some embodiments, the IFNα polypeptide comprises a non-natural amino acid or modified amino acid at E51. In some embodiments, the IFNα polypeptide comprises a non-natural amino acid or modified amino acid at N156. [000120] In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at H7 and Q40. In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at H7 and Q40 and further comprises at least one additional non-natural amino acid or modified amino acid at a position selected from E51, N45, and N156. In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at H7, Q40 and E51. In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at H7, Q40 and N156. In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at H7, Q40, N45, and N156. In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at H7 and E51. In some 24
embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at H7, E51 and N156. [000121] In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at Q40 and E51. In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at Q40 and N156. In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at E51 and N156. In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at Q40, E51, and N156. [000122] In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at Q40 and N156. In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids at Q40 and N156 and further comprises at least one non-natural amino acids or modified amino acids at a position selected from H7 and E51. [000123] In some embodiments, the IFNα polypeptide comprises at least one non-natural amino acid or modified amino acid in a position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprises at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136. [000124] In some embodiments, the IFNα polypeptide comprises at least two non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprises at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136. [000125] In some embodiments, the IFNα polypeptide comprises at least three non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprises at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136. [000126] In some embodiments, the IFNα polypeptide comprises non-natural amino acids or modified amino acids in positions Q40 and N156 and further comprises at least one non-natural amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, 25
K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136. [000127] In any of the foregoing embodiments, the non-natural amino acid or modified amino acid is a non-natural amino acid. In any of the foregoing embodiments, the non-natural amino acid or modified amino acid is a modified amino acid. [000128] In any of the foregoing embodiments, the non-natural amino acid or modified amino acid comprises a residue of a moiety selected from the group consisting of amino, carboxy, acetyl, hydrazino, hydrazido, semicarbazido, sulfanyl, azido and alkynyl. [000129] In any of the foregoing embodiments, the modified amino acid is selected from the group consisting of p-acetyl-L-phenylalanine, O-methyl-L-tyrosine, 3-methyl-phenylalanine, O-4-allyl-L-tyrosine, 4-propyl-L-tyrosine, fluorinated phenylalanine, isopropyl-L- phenylalanine, p-azido-L-phenylalanine, p-acyl-L-phenylalanine, p-benzoyl-L-phenylalanine, p-iodo-phenylalanine, p-bromophenylalanine, p-amino-L-phenylalanine, isopropyl-L- phenylalanine, p-propargyloxy-phenylalanine, and p-azidomethyl-L-phenylalanine residues. In any of the foregoing embodiments, the non-natural amino acid is a p-azidomethyl-L- phenylalanine residue. [000130] In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L- phenylalanine residue in at least one amino acid position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156. [000131] In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L- phenylalanine residue in at least one amino acid position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L-phenylalanine residue in at least two amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L-phenylalanine residue in at least three amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L-phenylalanine residue in at least four amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L- phenylalanine residue in at least five amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L-phenylalanine residue in positions H7, Q40, E41, N45, E51, and N156. 26
[000132] In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L- phenylalanine residue at H7. In some embodiments, the IFNα polypeptide comprises a p- azidomethyl-L-phenylalanine residue at Q40. In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L-phenylalanine residue at E41. In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L-phenylalanine residue at N45. In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L-phenylalanine residue at E51. In some embodiments, the IFNα polypeptide comprises a p-azidomethyl-L-phenylalanine residue at N156. [000133] In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L- phenylalanine residues at H7 and Q40. In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L-phenylalanine residues at H7 and Q40 and further comprises at least one additional p-azidomethyl-L-phenylalanine residue at a position selected from E51, N45, and N156. In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L-phenylalanine residues at H7, Q40 and E51. In some embodiments, the IFNα polypeptide comprises p- azidomethyl-L-phenylalanine residues at H7, Q40 and N156. In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L-phenylalanine residues at H7, Q40, N45, and N156. In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L-phenylalanine residues at H7 and E51. In some embodiments, the IFNα polypeptide comprises p-azidomethyl- L-phenylalanine residues at H7, E51 and N156. [000134] In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L- phenylalanine residues at Q40 and E51. In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L-phenylalanine residues at Q40 and N156. In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L-phenylalanine residues at E51 and N156. In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L-phenylalanine residues at Q40, E51, and N156. [000135] In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L- phenylalanine residues at Q40 and N156. In some embodiments, the IFNα polypeptide comprises p-azidomethyl-L-phenylalanine residues at Q40 and N156 and further comprises at least one p-azidomethyl-L-phenylalanine residue at a position selected from H7 and E51. [000136] In some embodiments, the amino acid substitution position is according to the sequence of wild-type IFNα. In some embodiments, the amino acid substitution is with reference to SEQ ID NO: 33. In some embodiments, the IFNα polypeptide comprises an amino acid sequence having at least 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to SEQ ID NO: 33. 27
[000137] In some embodiments, the IFNα polypeptide comprises an amino acid sequence according to: SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, or SEQ ID NO: 8. [000138] Also within the scope are post-translationally modified variants of the IFNα polypeptides disclosed herein. Any of the IFNα polypeptides provided herein can be post- translationally modified in any manner recognized by those of skill in the art. Typical post- translational modifications for IFNα polypeptides include interchain disulfide bonding and glycosylation. The post-translational modification can occur during production, in vivo, in vitro or otherwise. In some embodiments, the post-translational modification can be an intentional modification by a practitioner, for instance, using the methods provided herein. [000139] Further included within the scope are IFNα polypeptides fused to further peptides or polypeptides. Exemplary fusions include, but are not limited to, e.g., a methionyl IFNα polypeptide in which a methionine is linked to the N-terminus of the IFNα polypeptide resulting from recombinant expression, fusions for the purpose of purification (including but not limited to, to poly-histidine or affinity epitopes), fusions with serum albumin binding peptides, and fusions with serum proteins such as serum albumin. The IFNα polypeptides may comprise protease cleavage sequences, IFNα polypeptide-binding domains (including but not limited to, FLAG or poly-His) or other affinity-based sequences (including but not limited to, FLAG, poly-His, GST, etc.). The IFNα polypeptides may also comprise linked molecules that improve detection (including, but not limited to, GFP), purification, or other features of the IFNα polypeptide. In certain embodiments, the IFNα polypeptides comprise a C-terminal affinity sequence that facilitates purification of full length IFNα polypeptides. In certain embodiments, such C-terminal affinity sequence is a poly-His sequence, e.g., a 6-His sequence. In certain embodiments, the IFNα polypeptides comprise an N-terminal affinity sequence that facilitates purification of full length IFNα polypeptides. In certain embodiments, such N- terminal affinity sequence is a poly-His sequence, e.g., a 6-His sequence. In certain embodiments, the IFNα polypeptides are fused to a polypeptide sequence that facilitates expression or purification. In certain embodiments, the fusion polypeptide sequence is a small ubiquitin modifying protein (SUMO; Butt et al., 2009, Protein Expr Purif. 43(1): 1–9). In advantageous embodiments, the fusion protein can be cleaved from the IFNα polypeptide during or after expression or purification. In some embodiments, the fused peptide or polypeptide specifically binds to a target molecule other than the target molecule bound by the IFNα polypeptide. 28
[000140] In some embodiments, the at least one non-natural amino acid or modified amino acid substitution provides an IFNα polypeptide that has reduced IFNAR binding compared to wild-type IFNα. In some embodiments, the at least one non-natural amino acid substitution or modified amino acid provides an IFNα polypeptide that has reduced toxicity, for example, systemic toxicity, compared to wild-type IFNα. In some embodiments, the at least one non- natural amino acid or modified amino acid substitution provides an IFNα polypeptide that has increased stability, for example, increased stability in serum, compared to wild-type IFNα. In some embodiments, the at least one non-natural amino acid or modified amino acid substitution provides an IFNα polypeptide that has a longer half-life in serum compared to wild-type IFNα. In some embodiments, the at least one non-natural amino acid or modified amino acid substitution provides an IFNα polypeptide that has reduced toxicity and increased stability compared to wild-type IFNα. [000141] In certain embodiments, the IFNα polypeptide has increased affinity for IFNAR. In certain embodiments, the at least one non-natural amino acid or modified amino acid is on an IFNAR receptor contacting surface of the IFNα polypeptide. In certain embodiments, the at least one non-natural amino acid or modified amino acid in the IFNα polypeptide is located at an amino acid position that contacts IFNAR through hydrogen bonds and/or ionic bonds. In certain embodiments, the at least one non-natural amino acid or modified amino acid in the IFNα polypeptide is at a position that contacts IFNAR through ionic bonds. In certain embodiments, one or more non-natural amino acids or modified amino acids increase binding of IFNα polypeptide to IFNAR relative to an IFNα of the same sequence, other than the one or more non-natural amino acids or modified amino acids. In certain embodiments, one or more non-natural amino acids or modified amino acids increase binding of IFNα polypeptide to IFNAR by 10%, 20%, 25%, 50%, 75%, 100%, 125%, 150%, 200%, 250%, 300%, 400%, 500%, 1000%, 2000%, 3000%, or more. [000142] In certain embodiments, the one or more non-natural amino acids or modified amino acid increase the stability of the IFNα polypeptide. In certain embodiments, the one or more non-natural amino acids or modified amino acids increase the serum half-life of the IFNα polypeptide. In certain embodiments, the one or more non-natural amino acids or modified amino acids increase the serum half-life of the IFNα polypeptide relative to wild-type IFNα. In certain embodiments, the one or more non-natural amino acids or modified amino acids increase the serum half-life of the IFNα polypeptide relative to an IFNα of the same sequence, other than the one or more non-natural amino acids or modified amino acid. In certain embodiments, the one or more non-natural amino acids increase the serum half-life of the IFNα 29
polypeptide by 10%, 20%, 25%, 50%, 75%, 100%, 125%, 150%, 200%, 250%, 300%, 400%, 500%, 1000%, 2000%, 3000%, or more. [000143] In any of the foregoing embodiments, the non-natural amino acid or modified amino acid is a non-natural amino acid. In any of the foregoing embodiments, the non-natural amino acid or modified amino acid is a modified amino acid. 1.3. IFNα Conjugates [000144] In another aspect, provided herein are IFNα conjugates comprising an IFNα polypeptide and a masking moiety, wherein the IFNα polypeptide is linked to the masking moiety, optionally via a linker. In certain embodiments, the IFNα conjugate comprises an IFNα polypeptide as described herein with at least one non-natural amino acid or modified amino acid wherein the at least one non-natural amino acid or modified amino acid is linked to a masking moiety, optionally via a linker. In a further embodiment, the masking moiety is a water-soluble polymer, a carbohydrate, or a peptide. In certain embodiments, the non-natural amino acid or modified amino acid is a non-natural amino acid. In certain embodiments, the non-natural amino acid or modified amino acid is a modified amino acid. [000145] In another embodiment, the IFNα conjugate comprises an IFNα polypeptide site- specifically linked to a masking moiety via a protease cleavable linker wherein the masking moiety is a water-soluble polymer or carbohydrate. [000146] In another embodiment, the IFNα conjugate comprises an IFNα polypeptide site- specifically linked to a masking moiety via a pH-sensitive linker. In another embodiment, the IFNα conjugate comprises an IFNα polypeptide site-specifically linked to a masking moiety via a cathepsin B cleavable linker. In a further embodiment, the masking moiety is a water- soluble polymer, a carbohydrate, or a peptide. [000147] The IFNα conjugates described herein can be linked to one, two, three, four, five, six, or more masking moieties optionally via linker(s). The linker can be any linker capable of forming at least one bond to the IFNα polypeptide and at least one bond to a masking moiety. Useful linkers are described the sections and examples below. In certain embodiments, the linkers are protease cleavable or pH-sensitive. [000148] In certain embodiments, the conjugate can be formed from an IFNα polypeptide that comprises one or more reactive groups. In certain embodiments, the conjugate can be formed from an IFNα polypeptide comprising all naturally encoded amino acids. Those of skill in the art will recognize that several naturally encoded amino acids include reactive groups capable of conjugation to a masking moiety or to a linker. These reactive groups include 30
cysteine side chains, lysine side chains, and amino-terminal groups. In these embodiments, the conjugate can comprise a masking moiety or linker linked to the residue of a reactive group. In these embodiments, the masking moiety precursor or linker precursor comprises a reactive group capable of forming a bond with a reactive group. Typical reactive groups include maleimide groups, activated carbonates (including but not limited to, p-nitrophenyl ester), activated esters (including but not limited to, N-hydroxysuccinimide, p-nitrophenyl ester, and aldehydes). Particularly useful reactive groups include maleimide and succinimide, for instance N-hydroxysuccinimide, for forming bonds to cysteine and lysine side chains. Additional reactive groups include alkynes, for example strained alkynes, and azides, for forming bonds to non-natural amino acids or modified amino acids incorporated in IFNα polypeptide chains. Additional reactive groups are described in the sections and examples below. [000149] In certain embodiments, the IFNα polypeptide comprises one or more modified amino acids having a reactive group, as described herein. Typically, the modified amino acid is not a naturally encoded amino acid. These modified amino acids can comprise a reactive group useful for forming a covalent bond to a masking moiety precursor or to a payload precursor. One of skill in the art can use the reactive group to link the IFNα polypeptide to any molecular entity capable of forming a covalent bond to the modified amino acid. Thus, provided herein are conjugates comprising an IFNα polypeptide comprising a modified amino acid residue linked to a payload directly or indirectly via a linker. Exemplary modified amino acids are described in the sections below. Generally, the modified amino acids have reactive groups capable of forming bonds to linkers or payloads with complementary reactive groups. [000150] The non-natural amino acids or modified amino acids are positioned at select locations in a polypeptide chain of the IFNα polypeptide. These locations were identified as providing optimum sites for substitution with the non-natural amino acids or modified amino acids. Each site is capable of bearing a non-natural amino acid or modified amino acid with optimum structure, function and/or methods for producing the IFNα polypeptide. [000151] In certain embodiments, a site-specific position for substitution provides an IFNα polypeptide or conjugate that is stable. Stability can be measured by any technique apparent to those of skill in the art. [000152] In certain embodiments, a site-specific position for substitution provides an IFNα polypeptide or conjugate that has optimal functional properties. For instance, the IFNα polypeptide or conjugate can show little or no loss of binding affinity for its target antigen compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. In certain embodiments, the IFNα polypeptide or conjugate can show 31
enhanced binding compared to an IFNα polypeptide without the site-specific non-natural amino acid or modified amino acid. [000153] In certain embodiments, a site-specific position for substitution provides an IFNα polypeptide or conjugate that can be made advantageously. For instance, in certain embodiments, the IFNα polypeptide or conjugate shows advantageous properties in its methods of synthesis, discussed below. In certain embodiments, the IFNα polypeptide or conjugate can show little or no loss in yield in production compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. In certain embodiments, the IFNα polypeptide or conjugate can show enhanced yield in production compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. In certain embodiments, the IFNα polypeptide or conjugate can show little or no loss of tRNA suppression compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. In certain embodiments, the IFNα polypeptide or conjugate can show enhanced tRNA suppression in production compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. [000154] In certain embodiments, a site-specific position for substitution provides an IFNα polypeptide or conjugate that has advantageous solubility. In certain embodiments, the IFNα polypeptide or conjugate can show little or no loss in solubility compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. In certain embodiments, the IFNα polypeptide or conjugate can show enhanced solubility compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. [000155] In certain embodiments, a site-specific position for substitution provides an IFNα polypeptide or conjugate that has advantageous expression. In certain embodiments, the IFNα polypeptide or conjugate can show little or no loss in expression compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. In certain embodiments, the IFNα polypeptide or conjugate can show enhanced expression compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. [000156] In certain embodiments, a site-specific position for substitution provides an IFNα polypeptide or conjugate that has advantageous folding. In certain embodiments, the IFNα polypeptide or conjugate can show little or no loss in proper folding compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino 32
acid. In certain embodiments, the IFNα polypeptide or conjugate can show enhanced folding compared to an IFNα polypeptide or conjugate without the site-specific non-natural amino acid or modified amino acid. [000157] In certain embodiments, a site-specific position for substitution provides an IFNα polypeptide that is capable of advantageous conjugation. As described below, several nonnatural amino acids have side chains or functional groups that facilitate conjugation of the IFNα polypeptide to a second agent, either directly or via a linker. In certain embodiments, the IFNα polypeptide can show enhanced conjugation efficiency compared to an IFNα polypeptide without the same or other non-natural amino acids or modified amino acids at other positions. In certain embodiments, the IFNα polypeptide can show enhanced conjugation yield compared to an IFNα polypeptide without the same or other non-natural amino acids or modified amino acids at other positions. In certain embodiments, the IFNα polypeptide can show enhanced conjugation specificity compared to an IFNα polypeptide without the same or other non-natural amino acids or modified amino acids at other positions. [000158] The one or more non-natural amino acids or modified amino acids are located at selected site-specific positions in at least one polypeptide chain of the IFNα conjugate. The polypeptide chain can be any polypeptide chain of the IFNα polypeptide without limitation. [000159] In certain embodiments, the IFNα polypeptides or conjugate provided herein comprise one non-natural amino acid or modified amino acid at a site-specific position. In certain embodiments, the IFNα polypeptide or conjugate provided herein comprise two non- natural amino acids or modified amino acids at site-specific positions. In certain embodiments, the IFNα polypeptide or conjugate provided herein comprise three non-natural amino acids or modified amino acids at site-specific positions. In certain embodiments, the IFNα polypeptide or conjugate provided herein comprise more than three non-natural amino acids or modified amino acids at site-specific positions. In certain embodiments, the IFNα polypeptide or conjugate provided herein comprise four non-natural amino acids or modified amino acids at site-specific positions. In certain embodiments, the non-natural or modified amino acid is a non-natural amino acid. In certain embodiments, the non-natural or modified amino acid a modified amino acid. [000160] In certain embodiments, the IFNα conjugate is of Formula (I): 33
or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein COMP is an IFNα polypeptide; L is a linker, for example, a linker that comprises a protease cleavable linker or a pH-sensitive linker; MM is a masking moiety; x is an integer selected from 0 and 1; and y is an integer between 1 and 30. [000161] In certain embodiments, IFNα is an IFNα polypeptide that comprises at least one non-natural amino acid or modified amino acid selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156. In certain embodiments, IFNα is an IFNα polypeptide that comprises at least one non-natural amino acid or modified amino acid selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. [000162] In certain embodiments, linker L is a cathepsin B cleavable linker. [000163] In certain embodiments, the masking moiety is a water-soluble polymer, a carbohydrate, or a peptide. [000164] In certain embodiments, IFNα is an IFNα polypeptide, L is a protease cleavable linker, and MM is a water soluble polymer or a carbohydrate. [000165] In certain embodiments, IFNα is an IFNα polypeptide and L is a pH-sensitive linker. [000166] In certain embodiments, IFNα is an IFNα polypeptide and L is a cathepsin B cleavable linker. [000167] In certain embodiments, x is 0. In certain embodiments, x is 1. In certain embodiments, y is 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10. In certain embodiments, y is 1, 2, 3, 4, 5, or 6. In certain embodiments, y is 1, 2, 3, or 4. In certain embodiments, y is 1. In certain embodiments, y is 2. In certain embodiments, y is 3. In certain embodiments, y is 4. [000168] In certain embodiments, x is 0 and y is 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10. In certain embodiments, x is 1 and y is 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10. In certain embodiments, x is 0 and y is 1, 2, 3, 4, 5, or 6. In certain embodiments, x is 1 and y is 1, 2, 3, 4, 5, or 6. In certain embodiments, x is 0 and y is 1. In certain embodiments, x is 0 and y is 2. In certain embodiments, x is 0 and y is 3. In certain embodiments, x is 0 and y is 4. In certain embodiments, x is 0 and y is 5. In certain embodiments, x is 0 and y is 6. In certain embodiments, x is 1 and y is 1. In certain embodiments, x is 1 and y is 2. In certain 34
embodiments, x is 1 and y is 3. In certain embodiments, x is 1 and y is 4. In certain embodiments, x is 1 and y is 5. In certain embodiments, x is 1 and y is 6. 1.3.1. Masking Groups of IFNα Conjugates [000169] The IFNα conjugates described herein comprise an IFNα polypeptide and at least one masking moiety. [000170] The masking moiety can be any macromolecule deemed suitable by the person of skill in the art. In certain embodiments, the masking moiety is a protein, peptide, antibody or antigen binding fragment thereof, nucleic acid, carbohydrate, or other large molecule composed of polymerized monomers. In certain embodiments, the masking moiety is a protein. In certain embodiments, the masking moiety is an antibody or an antigen binding fragment thereof. In some embodiments, the masking moiety is a residue of a polypeptide. In some embodiments, the masking moiety is a residue of an antibody. In some embodiments, the masking moiety is a residue of an antibody chain. [000171] In certain embodiments, the masking moiety is a polymer, denotated as POLY herein, for example a water-soluble polymer. These polymers can be linked to the polypeptide via a naturally encoded amino acid, via a non-naturally encoded amino acid, or any functional substituent of a natural or modified amino acid, or any substituent or functional group added to a natural or modified amino acid. The polymer can also be linked to the polypeptide via a linker as described herein. The molecular weight of the polymer may be of a wide range, including but not limited to, between about 100 Da and about 100,000 Da or more. [000172] The polymer selected may be water soluble so that a protein to which it is attached does not precipitate in an aqueous environment, such as a physiological environment. The polymer may be branched or unbranched. Preferably, for therapeutic use of the end-product preparation, the polymer will be pharmaceutically acceptable. [000173] The water-soluble polymer may be any structural form including, but not limited to linear, forked or branched. Typically, the water soluble polymer is a poly(alkylene glycol), such as poly(ethylene glycol) (PEG), but other water soluble polymers can also be employed. By way of example, PEG is used to describe certain embodiments. [000174] Alternative examples of polymers include, but are not limited to, other poly(alkylene glycols) such as poly(propylene glycol) (“PPG”), copolymers of ethylene glycol and propylene glycol and the like, poly(oxyethylated polyol), poly(olefinic alcohol), poly(vinylpyrrolidone), poly(hydroxyalkylmethacrylamide), poly(hydroxy- alkylmethacrylate), poly(saccharides), poly(α-hydroxy acid), poly(vinyl alcohol), 35
polyphosphazene, polyoxazolines (“POZ”) (which are described in WO 2008/106186), poly(N-acryloylmorpholine), and combinations of any of the foregoing. [000175] PEG is a well-known, water-soluble polymer that is commercially available or can be prepared by ring-opening polymerization of ethylene glycol according to methods well known in the art (Sandler and Karo, Polymer Synthesis, Academic Press, New York, Vol.3, pages 138-161). The term “PEG” is used broadly to encompass any polyethylene glycol molecule, without regard to size or to modification at an end of the PEG, and can be represented as linked to a polypeptide by the formula: XO–(CH2CH2O)n–CH2CH2–Y where n is 2 to 10,000,X is H or a terminal modification, including but not limited to, a C1-4 alkyl, and Y is the attachment point to the polypeptide. [000176] In some cases, a PEG terminates on one end with hydroxy or methoxy, i.e., X is H or CH3 (“methoxy PEG” or “mPEG”). Alternatively, the PEG can terminate with a reactive group, thereby forming a bifunctional polymer. Typical reactive groups can include those reactive groups that are commonly used to react with the functional groups found in the 20 common amino acids (including but not limited to, maleimide groups, activated carbonates (including but not limited to, p-nitrophenyl ester), activated esters (including but not limited to, N hydroxysuccinimide, p-nitrophenyl ester, and aldehydes) as well as functional groups that are inert to the 20 common amino acids but that react specifically with complementary functional groups present in non-naturally encoded or modified amino acids (including but not limited to, azide groups, alkyne groups). It is noted that the other end of the PEG, which is shown in the above formula by Y, will attach either directly or indirectly to a polypeptide via a naturally-occurring, non-naturally encoded, or modified amino acid. For instance, Y may be an amide, carbamate or urea linkage to an amine group (including but not limited to, the epsilon amine of lysine or the N-terminus) of the polypeptide. Alternatively, Y may be a maleimide linkage to a thiol group (including but not limited to, the thiol group of cysteine). Alternatively, Y may be a linkage to a residue not commonly accessible via the 20 common amino acids. For example, an azide group on the PEG can be reacted with an alkyne group on the polypeptide to form a Huisgen [3+2] cycloaddition product. Alternatively, an alkyne group on the PEG can be reacted with an azide group present in a non-naturally encoded or modified amino acid, such as the modified amino acids described herein, to form a similar product. In some embodiments, a strong nucleophile (including but not limited to, hydrazine, hydrazide, hydroxylamine, semicarbazide) can be reacted with an aldehyde or ketone group present in a non-naturally encoded or modified amino acid to form a hydrazone, oxime or semicarbazone, as applicable, which in some cases can be further reduced by treatment with an appropriate reducing agent. 36
Alternatively, the strong nucleophile can be incorporated into the polypeptide via a non- naturally encoded or modified amino acid and used to react preferentially with a ketone or aldehyde group present in the water-soluble polymer. [000177] In certain embodiments, the proportion of polyethylene glycol molecules to polypeptide molecules will vary, as will their concentrations in the reaction mixture. In general, the optimum ratio (in terms of efficiency of reaction in that there is minimal excess unreacted protein or polymer) may be determined by the molecular weight of the polyethylene glycol selected and on the number of available reactive groups available. As it relates to molecular weight, typically the higher the molecular weight of the polymer, the fewer number of polymer molecules which may be attached to the protein. Similarly, branching of the polymer should be taken into account when optimizing these parameters. Generally, the higher the molecular weight (or the more branches) the higher the polymer:protein ratio. [000178] Any molecular mass for a PEG can be used as practically desired, including but not limited to, from about 100 Daltons (Da) to 100,000 Da or more as desired (including but not limited to, sometimes 0.1-50 kDa or 10-40 kDa). Branched chain PEGs, including but not limited to, PEG molecules with each chain having a MW ranging from 1-100 kDa (including but not limited to, 150 kDa or 5-20 kDa) can also be used. A wide range of PEG molecules are described in, including but not limited to, the Shearwater Polymers, Inc. catalog, and the Nektar Therapeutics catalog, incorporated herein by reference. [000179] Generally, at least one terminus of the PEG molecule is available for reaction with the IFNα polypeptide or a linker of the IFNα polypeptide. For example, PEG derivatives bearing alkyne and azide moieties for reaction with amino acid side chains can be used to attach PEG to non-naturally encoded or modified amino acids as described herein. If the non-naturally encoded or modified amino acid comprises an azide, then the PEG will typically contain either an alkyne moiety to effect formation of the [3+2] cycloaddition product or an activated PEG species (i.e., ester, carbonate) containing a phosphine group to effect formation of the amide linkage. Alternatively, if the non-naturally encoded amino acid or modified comprises an alkyne, then the PEG will typically contain an azide moiety to effect formation of the [3+2] Huisgen cycloaddition product. If the non-naturally encoded or modified amino acid comprises a carbonyl group, the PEG will typically comprise a potent nucleophile (including but not limited to, a hydrazide, hydrazine, hydroxylamine, or semicarbazide functionality) in order to effect formation of corresponding hydrazone, oxime, and semicarbazone linkages, respectively. In other alternatives, a reverse of the orientation of the reactive groups described 37
herein can be used, i.e., an azide moiety in the non-naturally encoded or modified amino acid can be reacted with a PEG derivative containing an alkyne. [000180] In some embodiments, the PEG molecule contains a chemical functionality that is reactive with the chemical functionality present on the side chain of the non-naturally encoded or modified amino acid. [000181] In alternative embodiments, the PEG molecule terminates in an amine which is available for reaction with a linker attached to the polypeptide. For example, the linker may terminate in an electrophile, for example, a carboxylic acid, and the amine of the PEG molecule is a nucleophile to form an amide bond. In other embodiments, the PEG molecule is a mPEG- NHS reagent, including mPEG-succinimidyl ester. The NHS ester can react with an amine group, for example on the polypeptide or the linker, at a pH of 7-8.5 to form a stable amide bond. [000182] In certain embodiments, the masking moiety is an azide- or acetylene-containing polymer comprising a water-soluble polymer backbone having an average molecular weight from about 800 Da to about 100,000 Da. In other embodiments, the masking moiety is an amine- or N-hydroxysuccinimide-containing polymer comprising a water-soluble polymer backbone having an average molecular weight from about 800 Da to about 100,000 Da. The polymer backbone of the water-soluble polymer can be poly(ethylene glycol). However, it should be understood that a wide variety of water soluble polymers including but not limited to poly(ethylene)glycol and other related polymers, including poly(dextran) and poly(propylene glycol), are also suitable for use and that the use of the term PEG or poly(ethylene glycol) is intended to encompass and include all such molecules. The term PEG includes, but is not limited to, poly(ethylene glycol) in any of its forms, including bifunctional PEG, multiarmed PEG, derivatized PEG, forked PEG, branched PEG, pendent PEG (i.e. PEG or related polymers having one or more functional groups pendent to the polymer backbone), or PEG with degradable linkages therein. [000183] The polymer backbone can be linear or branched. Branched polymer backbones are generally known in the art. Typically, a branched polymer has a central branch core moiety and a plurality of linear polymer chains linked to the central branch core. PEG is commonly used in branched forms that can be prepared by addition of ethylene oxide to various polyols, such as glycerol, glycerol oligomers, pentaerythritol and sorbitol. The central branch moiety can also be derived from several amino acids, such as lysine. The branched poly(ethylene glycol) can be represented in general form as R(-PEG-OH)m in which R is derived from a core moiety, such as glycerol, glycerol oligomers, or pentaerythritol, and m represents the number of arms. 38
Multi-armed PEG molecules, such as those described in U.S. Pat. Nos.5,932,4625,643,575; 5,229,490; 4,289,872; U.S. Pat. Appl.2003/0143596; WO 96/21469; and WO 93/21259, each of which is incorporated by reference herein in its entirety, can also be used as the polymer backbone. [000184] Branched PEG can also be in the form of a forked PEG represented by PEG(YCHZ2)n, where Y is a linking group and Z is an activated terminal group linked to CH by a chain of atoms of defined length. [000185] Yet another branched form, the pendant PEG, has reactive groups, such as carboxyl, along the PEG backbone rather than at the end of PEG chains. [000186] In addition to these forms of PEG, the polymer can also be prepared with weak or degradable linkages in the backbone. For example, PEG can be prepared with ester linkages in the polymer backbone that are subject to hydrolysis. As shown herein, this hydrolysis results in cleavage of the polymer into fragments of lower molecular weight: PEG-CO2-PEG- + H2O → PEG-CO2H + HO-PEG- It is understood by those skilled in the art that the term poly(ethylene glycol) or PEG represents or includes all the forms known in the art including but not limited to those disclosed herein. [000187] Many other polymers are also suitable for use. In some embodiments, polymer backbones that are water-soluble, with from 2 to about 300 termini, are particularly suitable. Examples of suitable polymers include, but are not limited to, other poly(alkylene glycols), such as poly(propylene glycol) (“PPG”), copolymers thereof (including but not limited to copolymers of ethylene glycol and propylene glycol), terpolymers thereof, mixtures thereof, and the like. Although the molecular weight of each chain of the polymer backbone can vary, it is typically in the range of from about 800 Da to about 100,000 Da, often from about 6,000 Da to about 80,000 Da. [000188] Those of ordinary skill in the art will recognize that the foregoing list for substantially water-soluble backbones is by no means exhaustive and is merely illustrative, and that all polymeric materials having the qualities described herein are contemplated as being suitable for use. [000189] In some embodiments the polymer derivatives are “multi-functional”, meaning that the polymer backbone has at least two termini, and possibly as many as about 300 termini, functionalized or activated with a functional group. Multifunctional polymer derivatives include, but are not limited to, linear polymers having two termini, each terminus being bonded to a functional group which may be the same or different. 39
[000190] In certain embodiments, POLY is polyethylene glycol (PEG), methoxypolyethylene glycol (mPEG), poly(propylene glycol) (PPG), copolymers of ethylene glycol and propylene glycol, poly(oxyethylated polyol), poly(olefinic alcohol), poly(vinylpyrrolidone), poly(hydroxyalkylmethacrylamide), poly(hydroxyalkylmethacrylate), poly(saccharides), poly( β-hydroxy acid), poly(vinyl alcohol), polyphosphazene, polyoxazolines (POZ), poly(N- acryloylmorpholine), polysarcosine, or a combination thereof. In some embodiments, POLY is polyethylene glycol (PEG). In some embodiments, POLY is methoxypolyethylene glycol (mPEG). In some embodiments, POLY is poly(propylene glycol) (PPG). In some embodiments, POLY is copolymers of ethylene glycol and propylene glycol. In some embodiments, POLY is poly(oxyethylated polyol). In some embodiments, POLY is poly(olefinic alcohol). In some embodiments, POLY is poly(vinylpyrrolidone). In some embodiments, POLY is poly(hydroxyalkylmethacrylamide). In some embodiments, POLY is poly(hydroxyalkylmethacrylate). In some embodiments, POLY is poly(saccharides). In some embodiments, POLY is poly( β-hydroxy acid). In some embodiments, POLY is poly(vinyl alcohol). In some embodiments, POLY is polyphosphazene. In some embodiments, POLY is polyoxazolines (POZ). In some embodiments, POLY is poly(N-acryloylmorpholine). In some embodiments, POLY is polysarcosine. In some embodiments, POLY is a nonpeptidic, water- soluble polymer. In certain embodiments, POLY includes a polyethylene glycol (PEG) or methoxypolyethylene glycol (mPEG). In certain embodiments, POLY is , wherein represents attachment to the remainder of the compound, and wherein n1 is an integer from 1 to 10,000. In certain embodiments, n1 is an integer from 1 to 5,000. In certain embodiments, n1 is an integer from 1 to 2,500. In certain embodiments, n1 is an integer from 1 to 2,000. In certain embodiments, n1 is an integer from 1 to 1,000. In certain embodiments, n1 is an integer from 100 to 1,000. In certain embodiments, n1 is an integer from 100 to 700. In certain embodiments, n1 is an integer from 300 to 700. In certain embodiments, n1 is an integer from 400 to 500. In certain embodiments, n1 is an integer from 600 to 700. In certain embodiments, n1 is an integer from 100 to 500. [000191] In certain embodiments, including any of the foregoing, POLY is a residue of a nonpeptidic, hydrophilic polymer. In certain embodiments, POLY is a residue of polyethylene glycol (PEG), methoxypolyethylene glycol (mPEG), poly(propylene glycol) (PPG), copolymers of ethylene glycol and propylene glycol, poly(oxyethylated polyol), poly(olefinic alcohol), poly(vinylpyrrolidone), poly(hydroxyalkylmethacrylamide), 40
poly(hydroxyalkylmethacrylate), poly(saccharides), poly( β-hydroxy acid), poly(vinyl alcohol), polyphosphazene, polyoxazolines (POZ), poly(N-acryloylmorpholine), polysarcosine, or a combination thereof. In certain embodiments, including any of the foregoing, POLY is a residue of polyethylene glycol (PEG), methoxypolyethylene glycol (mPEG), poly(propylene glycol) (PPG), or a copolymer of ethylene glycol and propylene glycol. In certain embodiments, including any of the foregoing, POLY is a residue of methoxypolyethylene glycol (mPEG). [000192] In certain embodiments, including any of the foregoing, POLY is a residue of polyethylene glycol (PEG). In certain embodiments, including any of the foregoing, POLY is a residue of poly(propylene glycol) (PPG). In certain embodiments, including any of the foregoing, POLY is a residue of copolymers of ethylene glycol and propylene glycol. In certain embodiments, including any of the foregoing, POLY is a residue of poly(oxyethylated polyol). In certain embodiments, including any of the foregoing, POLY is a residue of poly(olefinic alcohol). In certain embodiments, including any of the foregoing, POLY is a residue of poly(vinylpyrrolidone). In certain embodiments, including any of the foregoing, POLY is a residue of poly(hydroxyalkylmethacrylamide). In certain embodiments, including any of the foregoing, POLY is a residue of poly(hydroxyalkylmethacrylate). In certain embodiments, including any of the foregoing, POLY is a residue of poly(saccharides). In certain embodiments, including any of the foregoing, POLY is a residue of poly( β-hydroxy acid). In certain embodiments, including any of the foregoing, POLY is a residue of poly(vinyl alcohol). In certain embodiments, including any of the foregoing, POLY is a residue of polyphosphazene. In certain embodiments, including any of the foregoing, POLY is a residue of polyoxazolines (POZ). In certain embodiments, including any of the foregoing, POLY is a residue of poly(N-acryloylmorpholine). In certain embodiments, including any of the foregoing, POLY is a residue of polysarcosine. [000193] In certain embodiments, including any of the foregoing, POLY is , wherein R1 is hydrogen or methyl, n1 is an integer from 1 to 10,000, inclusive, and represents attachment to the remainder of the compound or conjugate. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 5000. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 2500. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 1500. In certain embodiments, 41
including any of the foregoing, n1 is an integer between 100 to 1000. In certain embodiments, n1 is an integer from 100 to 700. In certain embodiments, n1 is an integer from 300 to 700. In certain embodiments, n1 is an integer from 400 to 500. In certain embodiments, n1 is an integer from 600 to 700. In certain embodiments, including any of the foregoing, n1 is an integer between 100 to 500. [000194] In other embodiments, the making group is a carbohydrate. In certain embodiments, the IFNα may be altered to increase, decrease, or eliminate the extent to which it is glycosylated. Glycosylation of polypeptides is typically either “N-linked” or “O-linked.” [000195] “N-linked” glycosylation refers to the attachment of a carbohydrate moiety to the side chain of an asparagine residue. The tripeptide sequences asparagine-X-serine and asparagine-X-threonine, where X is any amino acid except proline, are the recognition sequences for enzymatic attachment of the carbohydrate moiety to the asparagine side chain. Thus, the presence of either of these tripeptide sequences in a polypeptide creates a potential glycosylation site. [000196] “O-linked” glycosylation refers to the attachment of one of the sugars Nacetylgalactosamine, galactose, or xylose to a hydroxyamino acid, most commonly serine or threonine, although 5-hydroxyproline or 5-hydroxylysine may also be used. [000197] Addition of N-linked glycosylation sites to the protein may be accomplished by altering the amino acid sequence such that one or more of the above-described tripeptide sequences is created. Addition of O-linked glycosylation sites may be accomplished by addition or substitution of one or more serine or threonine residues in or to (as the case may be) the sequence of a protein. 1.3.2. Linkers of IFNα Conjugates [000198] The IFNα conjugates described herein comprise an IFNα polypeptide and a masking moiety wherein the IFNα conjugate is site-specifically linked to the masking moiety, optionally via a linker. In particular embodiments, the linker is cleavable. In certain embodiments, the linker is cleavable in vivo. In certain embodiments, the linker is cleavable by a protease. In certain embodiments, the linker is cleavable by a cathepsin. In certain embodiments, the linker is cleavable by cathepsin B. In certain embodiments, the linker is a pH-sensitive linker. [000199] In another embodiment, the IFNα conjugate comprises an IFNα polypeptide site- specifically linked to a masking moiety via a protease cleavable linker wherein the masking moiety is a water-soluble polymer or carbohydrate. In another embodiment, the IFNα conjugate comprises an IFNα polypeptide site-specifically linked to a masking moiety via a cathepsin B 42
cleavable linker. In another embodiment, the IFNα conjugate comprises an IFNα polypeptide site-specifically linked to a masking moiety via a pH-sensitive linker. [000200] In certain embodiments, provided herein is an IFNα conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof comprising a linker of Formula (L1): wherein RG is a reactive group residue; W1 and W2 are independently absent or a divalent attaching group; L1 is absent, a protease cleavable linker, or a pH-sensitive linker; SG1 is a divalent spacer group; is a bond to the IFNα polypeptide; and is a bond to the masking moiety. [000201] In certain embodiments, provided herein is an IFNα conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof comprising a linker of Formula (L2): wherein RG is a reactive group residue; W1 and W2 are independently absent or a divalent attaching group; L1 is absent, a protease cleavable linker, or a pH-sensitive linker; SG2 is a trivalent spacer group; is a bond to the IFNα polypeptide; and is a bond to the masking moiety. [000202] In certain embodiments, provided herein is an IFNα conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof comprising a linker of Formula (L1) wherein W1 and W2 are both divalent attaching groups, L1 is a protease cleavable linker 43
or a pH-sensitive linker, and RG and SG1 are as defined herein. In some embodiments, the protease cleavable linker is a cathepsin B cleavable linker. [000203] In certain embodiments, provided herein is an IFNα conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof comprising a linker of Formula (L1) wherein W1 and W2 are both divalent attaching groups, L1 is absent, and RG and SG1 are as defined herein. [000204] In certain embodiments, provided herein is an IFNα conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof comprising a linker of Formula (L2) wherein W1 and W2 are both divalent attaching groups, L1 is absent, and RG and SG2 are as defined herein. 1.3.2.1. Reactive Group Residues [000205] Reactive groups (or conjugating groups) facilitate conjugation of the masking moiety described herein to a second moiety, such as an FNα described herein. In certain embodiments, the reactive group is designated R herein. Reactive groups can react via any suitable reaction mechanism known to those of skill in the art. In certain embodiments, a reactive group reacts through a [3+2] alkyne-azide cycloaddition reaction, inverse-electron demand Diels-Alder ligation reaction, thiol-electrophile reaction, or carbonyl-oxyamine reaction, as described in detail herein. In certain embodiments, the reactive group comprises an alkyne, strained alkyne, tetrazine, thiol, para-acetyl-phenylalanine residue, oxyamine, maleimide, or azide. In certain embodiments, the reactive group is: , , , , , , , , , , –N3, methylcyclopropene (e.g. ), 44
or –SH; wherein R201 is lower alkyl. In certain embodiments, the reactive group is: or . [000206] In an embodiment, R201 is methyl, ethyl, or propyl. In an embodiment, R201 is methyl. Additional conjugating groups are described in, for example, U.S. Patent Publication No. 2014/0356385, U.S. Patent Publication No. 2013/0189287, U.S. Patent Publication No.2013/0251783, U.S. Patent No. 8,703,936, U.S. Patent No. 9,145,361, U.S. Patent No.9,222,940, and U.S. Patent No.8,431,558. [000207] After conjugation, a divalent residue of the reactive group, designated as RG, is formed and is bonded to the residue of an IFNα polypeptide. The structure of the divalent residue is determined by the type of conjugation reaction employed to form the conjugate. [000208] In certain embodiments when a conjugate is formed through a [3+2] alkyne-azide cycloaddition reaction, the divalent residue RG comprises a triazole ring or fused cyclic group comprising a triazole ring. In certain embodiment when a conjugate is formed through a strain- promoted [3+2] alkyne-azide cycloaddition (SPAAC) reaction, the divalent residue RG is: and/or . [000209] In certain embodiments when a conjugate is formed through a tetrazine inverse electron demand Diels-Alder ligation reaction, the divalent residue RG comprises a fused bicyclic ring having at least two adjacent nitrogen atoms in the ring. In certain embodiments when a conjugate is formed through a tetrazine inverse electron demand Diels-Alder ligation reaction, the divalent residue RG is: or . [000210] In certain embodiments when a conjugate is formed through a thiol-maleimide reaction, the divalent residue RG comprises succinimidylene and a sulfur linkage. In certain 45
embodiments when a conjugate is formed through a thiol-maleimide reaction, the divalent residue RG is: , or . [000211] In certain embodiments, a conjugate is formed through a thiol-N- hydroxysuccinimide reaction using the following group: . [000212] The reaction involved for formation of the conjugate comprises the following step: , [000213] and the resulting divalent residue RG is: . [000214] In certain embodiments when a conjugate is formed through a carbonyl-oxyamine reaction, the divalent residue RG comprises a divalent residue of a modified amino acid. In certain embodiments when a conjugate is formed through a carbonyl-oxyamine reaction, the divalent residue RG is: 46
or . [000215] In certain embodiments when a conjugate is formed through a carbonyl-oxyamine reaction, the divalent residue of RG comprises an oxime linkage. In certain embodiments when a conjugate is formed through a carbonyl-oxyamine reaction, the divalent residue of RG is: . [000216] In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises a triazole ring. In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer; wherein RG is a triazole ring or fused cyclic group comprising a triazole ring. In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: or . [000217] In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises a fused bicyclic ring having at least two adjacent nitrogen atoms in the ring. In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: 47
or . [000218] In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises a sulfur linkage. In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: , , or . [000219] In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises a divalent residue of a modified amino acid. In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: or . [000220] In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises an oxime linkage. In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: 48
. [000221] In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG comprises an oxime linkage. In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein RG is: . [000222] In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer or regioisomer thereof; wherein RG is: , , , , , , , , , , or . [000223] In an embodiment, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer or regioisomer thereof; wherein RG is: 49
or . 1.3.2.2. Divalent Attaching Groups [000224] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W1 is a divalent attaching group selected from -C(O)-C1-6alkylene-, -C(O)(C1-6alkylene)NR4-, -C(O)(C1-6alkylene)O-, and -C(O)(C1- 6alkylene)S- wherein R4 is independently hydrogen or C1-6alkyl, RG is connected to W1 at -C(O)-, and the C1-6alkylene is optionally substituted with one, two, or three substituents selected from halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000225] In certain embodiments, W1 is a divalent attaching group of the formula -C(O)- C1-6alkylene-, -C(O)-C1-5alkylene-, -C(O)-C1-4alkylene-, -C(O)-C4-6alkylene-, or -C(O)-C2- 5alkylene-. In certain embodiments of formula (L1) or (L2), W1 is a divalent attaching group of the formula -C(O)-C6alkylene-, -C(O)-C5alkylene-, -C(O)-C4alkylene-, -C(O)-C3alkylene-, -C(O)-C2alkylene-, or -C(O)-CH2alkylene. In certain embodiments, W1 is a divalent attaching group of -C(O)-C4alkylene-. [000226] In certain embodiments, W1 is a divalent attaching group of the formula -C(O)(C1-6alkylene)NH-, C(O)(C1-5alkylene)NH-, C(O)(C1-4alkylene)NH-, C(O)(C1- 3alkylene)NH-, or C(O)(C2-5alkylene)NH-. [000227] In certain embodiments, W1 is a divalent attaching group of the formula -C(O)(C6alkylene)NH-, C(O)(C5alkylene)NH-, C(O)(C4alkylene)NH-, C(O)(C3 alkylene)NH-, C(O)(C2alkylene)NH- or C(O)(CH2alkylene)NH-. In certain embodiments, W1 is a divalent attaching group of the formula -C(O)(C2alkylene)NH-. [000228] In certain embodiments, W1 is a divalent attaching group of the formula -C(O)(C1-6alkylene)O-, C(O)(C1-5alkylene)O-, C(O)(C1-4alkylene)O-, C(O)(C1-3alkylene)O-, or C(O)(C2-5alkylene)O-. [000229] In certain embodiments, W1 is a divalent attaching group of the formula -C(O)(C6alkylene)O-, C(O)(C5alkylene)O-, C(O)(C4alkylene)O-, C(O)(C3alkylene)O-, C(O)(C2alkylene)O- or C(O)(CH2alkylene)O-. In certain embodiments, W1 is a divalent attaching group of the formula -C(O)(C2alkylene)O-. 50
[000230] In certain embodiments, W1 is a divalent attaching group of the formula -C(O)(C1-6alkylene)S-, C(O)(C1-5alkylene)S-, C(O)(C1-4alkylene)S-, C(O)(C1-3alkylene)O-, or C(O)(C2-5alkylene)S-. [000231] In certain embodiments, W1 is a divalent attaching group of the formula -C(O)(C6alkylene)S-, C(O)(C5alkylene)S-, C(O)(C4alkylene)S-, C(O)(C3alkylene)S-, C(O)(C2alkylene)S- or C(O)(CH2alkylene)S-. In certain embodiments, W1 is a divalent attaching group of the formula -C(O)(C2alkylene)S-. [000232] In certain embodiments, W1 is a divalent attaching group of the formula -C(O)(C2alkyl)NH- or -C(O)-C4alkyl-. [000233] In any of the foregoing embodiments, the C1-6alkylene of a W1 divalent attaching group is unsubstituted. In any of the foregoing embodiments, the C1-6alkylene of a W1 divalent attaching group is optionally substituted with one, two, or three substituents selected from halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000234] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer or regioisomer thereof; wherein W2 is wherein X1 is absent, a divalent water-soluble polymer, -C1-6alkylene-, -NR4(C1- 6alkylene)-, or -O(C1-6alkylene)-; X2 is absent or -C1-6alkylene-; X3 is absent, -NR4-, or -O-; R4 is independently hydrogen or C1-6alkyl; and wherein the C1-6alkylene of X1 or X2 is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy; and wherein the carbonyl is attached to W1. [000235] In certain embodiments, X1 is absent. In certain embodiments, X1 is a divalent water-soluble polymer. In certain embodiments, the divalent water-soluble polymer is 51
wherein R1 is hydrogen or methyl and n2 is an integer between 1 and 50, inclusive. In certain embodiments, n2 is an integer between 10 and 40. In certain embodiments, n2 is an integer between 20 and 50. In certain embodiments, n2 is an integer between 1 and 20. In certain embodiments, n2 is an integer between 1 and 15. In certain embodiments, n2 is an integer between 1 and 10. In certain embodiments, n2 is an integer between 10 and 20. In certain embodiments, wherein n2 is 20. In certain embodiments, wherein n2 is 10. In certain embodiments, n2 is an integer between 1 and 6. In certain embodiments, n2 is 4, 5, or 6. In certain embodiments, wherein n2 is 4. [000236] In certain embodiments, X1 is unsubstituted -(C1-6alkylene)-. In certain embodiments, X1 is -(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X1 is -(C1-3alkylene)- wherein the C1-3alkylene is unsubstituted. In certain embodiments, X1 is -(C1-3alkylene)- wherein the C1-3alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X1 is -(C3-6alkylene)- wherein the C3-6alkylene is unsubstituted. In certain embodiments, X1 is -(C3-6alkylene)- wherein the C1-3alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000237] In certain embodiments, X1 is -(CH2)-, -(C2alkylene)-, -(C3alkylene)-, -(C4alkylene)-, -(C5alkylene)-, or -(C6alkylene)- wherein the alkylene is unsubstituted. In certain embodiments, X1 is unsubstituted -(CH2)-. [000238] In certain embodiments, X1 is -(CH2)-, -(C2alkylene)-, -(C3alkylene)-, -(C4alkylene)-, -(C5alkylene)-, or -(C6alkylene)- wherein the alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000239] In certain embodiments, X1 is -O(C1-6alkylene)- wherein the C1-6alkylene is unsubstituted. In certain embodiments, X1 is -O(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and 52
alkoxy. In certain embodiments, X1 is -O(C1-3alkylene)- wherein the C1-3alkylene is unsubstituted. In certain embodiments, X1 is -O(C1-3alkylene)- wherein the C1-3alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X1 is -O(C3-6alkylene)- wherein the C3-6alkylene is unsubstituted. In certain embodiments, X1 is -O(C3-6alkylene)- wherein the C1-3alkylene is optionally substituted with one, two, or three substituents selected a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000240] In certain embodiments, X1 is -O(CH2)-, -O(C2alkylene)-, -O(C3alkylene)-, -O(C4alkylene)-, -O(C5alkylene)-, or -O(C6alkylene)- wherein the alkylene is unsubstituted. In certain embodiments, X1 is -O(CH2)- wherein the CH2 group is unsubstituted. [000241] In certain embodiments, X1 is -O(CH2)-, -O(C2alkylene)-, -O(C3alkylene)-, -O(C4alkylene)-, -O(C5alkylene)-, or -O(C6alkylene)- wherein the alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000242] In certain embodiments, X1 is -NR4(C1-6alkylene)- wherein the C1-6alkylene is unsubstituted. In certain embodiments, X1 is -NR4(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X1 is -NR4(C1-3alkylene)- wherein the C1-3alkylene is unsubstituted. In certain embodiments, X1 is -NR4(C1-3alkylene)- wherein the C1-3alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X1 is -O(C3-6alkylene)- wherein the C3-6alkylene is unsubstituted. In certain embodiments, X1 is -NR4(C3-6alkylene)- wherein the C1-3alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000243] In certain embodiments, X1 is -NR4(CH2)-, -NR4(C2alkylene)-, -NR4(C3alkylene)-, -NR4(C4alkylene)-, -NR4(C5alkylene)- , or -NR4(C6alkylene)- wherein the alkylene is unsubstituted. In certain embodiments, X1 is -NR4(C2alkylene)- wherein the C2alkylene is unsubstituted. In certain embodiments, X1 is 53
-NH(C2alkylene)- wherein the C2alkylene is unsubstituted. In certain embodiments, X1 is -NR4(CH2)- wherein the CH2 group is unsubstituted. In certain embodiments, X1 is - NH(CH2)- wherein the CH2 group is unsubstituted. [000244] In certain embodiments, X1 is -NR4(CH2)-, -NR4(C2alkylene)-, -NR4(C3alkylene)-, -NR4(C4alkylene)-, - NR4(C5alkylene) -, or -NR4(C6alkylene)- wherein the alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000245] In any of the foregoing embodiments, R4 is independently hydrogen. In any of the foregoing embodiments, R4 is independently C1-6alkyl. In any of the foregoing embodiments, R4 is independently methyl. [000246] In certain embodiments, X2 is absent. In certain embodiments, X2 is unsubstituted -(C1-6alkylene)-. In certain embodiments, X2 is -(C1-6alkylene)- wherein the C1- 6alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X2 is -(C1-3alkylene)- wherein the C1- 3alkylene is unsubstituted. In certain embodiments, X2 is -(C1-3alkylene)- wherein the C1- 3alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X2 is -(C3-6alkylene)- wherein the C3- 6alkylene is unsubstituted. In certain embodiments, X2 is -(C3-6alkylene)- wherein the C1- 3alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000247] In certain embodiments, X2 is -(CH2)-, -(C2alkylene)-, -(C3alkylene)-, -(C4alkylene)-, -(C5alkylene)-, or -(C6alkylene)- wherein the alkylene is unsubstituted. In certain embodiments, X2 is unsubstituted -(C2alkylene)-. [000248] In certain embodiments, X2 is -(CH2)-, -(C2alkylene)-, -(C3alkylene)-, -(C4alkylene)-, -(C5alkylene)-, or -(C6alkylene)- wherein the alkylene is optionally substituted with one, two, or three substituents selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. 54
[000249] In certain embodiments, X3 is absent. In certain embodiments, X3 is -NR4-. In certain embodiments, X3 is -NH-. In any of the foregoing embodiments, X3 is -N(CH3)-. In certain embodiments, X3 is -O-. [000250] In certain embodiments, X1, X2, and X3 are absent. In certain embodiments, X1 is absent. In certain embodiments, X2 is absent. In certain embodiments, X3 is absent. In certain embodiments, X2 and X3 are absent. In certain embodiments, X1 and X3 are absent. In certain embodiments, X1 and X2 are absent. [000251] Non-limiting examples of W2 include , , , , , , , , , , , , , and . [000252] In certain embodiments, W2 is selected from , , , , , , , and . 1.3.2.3. Protease Cleavable Linker or pH-sensitive Linker (L1) [000253] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1) or (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein the linker is cleavable. In certain embodiments, the linker is cleavable in vivo. In certain embodiments, the linker is cleavable by a protease. In certain 55
embodiments, the linker is cleavable by a cathepsin. In certain embodiments, the linker is cleavable by cathepsin B. In certain embodiments, the linker is a pH-sensitive linker. [000254] In certain embodiments, L1 comprises a compound of the formula: wherein R5 is hydrogen, an electron donating group, or an electron withdrawing group; X4 is -O- or -NR6-; X5 is a linker; R6 is hydrogen or an electron withdrawing group; and m is an integer selected from 1 to 4. [000255] In certain embodiments, X5 includes an ester wherein the carbonyl carbon of the ester functional group is covalently bound to the fluorene. In certain embodiments, X5 includes an amide wherein the carbonyl carbon of the amide functional group is covalently bound to the fluorene. In certain embodiments, X5 includes a ketone wherein the carbonyl carbon of the ketone functional group is covalently bound to the fluorene. In certain embodiments, X5 includes an anhydride wherein one of the carbonyls of the anhydride functional group is covalently bound to the fluorene. In certain embodiments, X5 includes a sulfonyl wherein the sulfur of the sulfonyl functional group is covalently bound to the fluorene. In certain embodiments, X5 includes an ammonium where the positively charged nitrogen of the ammonium functional group is covalently bound to the fluorene. [000256] In certain embodiments, X5 is -C(O)-, -C(O)NR4, -C(O)-(C1-6alkylene)-, -C(O)-O-(C1-6alkylene)-, -C(O)-NR4-(C1-6alkylene)-, or -C(O)-S-(C1-6alkylene)- wherein the -C(O)- is bound to the fluorene and the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000257] In certain embodiments, X5 is -C(O)-. In certain embodiments, X5 is -C(O)NR4. In certain embodiments, X5 is -C(O)NH-. In certain embodiments, X5 is - C(O)N(CH3)-. 56
[000258] In certain embodiments, X5 is -C(O)-(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X5 is-C(O)-(C1-3alkylene)- wherein the C1- 6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X5 is-C(O)-(C3-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000259] In certain embodiments, X5 is -C(O)-O-(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X5 is -C(O)-O-(C1-3alkylene)- wherein the C1- 6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X5 is -C(O)-O-(C3-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000260] In certain embodiments, X5 is -C(O)-NR4(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X5 is -C(O)-NR4(C1-3alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X5 is -C(O)-NR4(C3-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000261] In certain embodiments, X5 is -C(O)-S-(C1-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X5 is -C(O)-S-(C1-3alkylene)- wherein the C1- 57
6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. In certain embodiments, X5 is -C(O)-S-(C3-6alkylene)- wherein the C1-6alkylene is optionally substituted with one, two, or three groups independently selected from a halogen (e.g., fluoro (F), chloro (Cl), bromo (Br), or iodo (I)), alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy. [000262] In certain embodiments, each R5 is hydrogen, an electron donating group, or an electron withdrawing group. In certain embodiments, each R5 is hydrogen. In certain embodiments, each R5 is an electron donating group. In certain embodiments, each R5 is an electron withdrawing group. The electron donating group can be any electron donating group deemed suitable to the person of skill in the art. The electron withdrawing group can be any electron withdrawing group deemed suitable to the person of skill in the art. In certain embodiments each R5 is independently selected from the group consisting of hydrogen, haloalkyl, halogen, -CN, -SO3H, -C(O)R3, -C(O)OR3, -OR3, -N(H)C(O)R3, -N(H)CO2R3, and -N(H)C(O)C(H)(R3)CO2H wherein each R3 is independently alkyl, cycloalkyl, heteroalkyl, heterocycloalkyl, aryl, or heteroaryl. In certain embodiments, each R5 is independently selected from the group consisting of haloalkyl, halogen, -CN, -SO3H, -C(O)R3, -C(O)OR3, -OR3, -N(H)C(O)R3, -N(H)CO2R3, and -N(H)C(O)C(H)(R3)CO2H wherein each R3 is independently alkyl, cycloalkyl, heteroalkyl, heterocycloalkyl, aryl, or heteroaryl. In certain embodiments, each R5 is independently selected from the group consisting of -H, -CF3, -Br, -Cl, -F, -CN, - SO3H, -C(O)Me, -CO2Me, -Ome, -N(H)C(O)Me, -N(H)CO2Me, and N(H)C(O)C(H)(Me)CO2H. In certain embodiments, each R5 is independently selected from the group consisting of -CF3, - Br, -Cl, -F, -CN, -SO3H, -C(O)Me, -CO2Me, -OMe, -N(H)C(O)Me, -N(H)CO2Me, and N(H)C(O)C(H)(Me)CO2H. In certain embodiments, R5 is hydrogen. In certain embodiments, R5 is -Br. In certain embodiments, R5 is -Cl. In certain embodiments, R5 is -F. In certain embodiments, R5 is -CN. In certain embodiments, R5 is -SO3H. In certain embodiments, R5 is -C(O)Me. In certain embodiments, R5 is -OMe. [000263] In certain embodiments, X4 is -O-. In certain embodiments, X4 is -NR6- wherein R6 is hydrogen or an electron withdrawing group. In certain embodiments, R6 is hydrogen. In certain embodiments, R6 is an electron withdrawing group. The electron withdrawing group can be any electron withdrawing group deemed suitable to the person of skill in the art. In certain embodiments, R6 is independently selected from the group consisting of -C(O)R3, - 58
C(O)OR3, and -S(O)2R3, wherein each R3 is independently alkyl, cycloalkyl, heteroalkyl, heterocycloalkyl, aryl, or heteroaryl. In certain embodiments, R6 is independently selected from the group consisting of -C(O)R3, C(O)OR3, and -S(O)2R3 wherein each R3 is independently alkyl, cycloalkyl, heteroalkyl, heterocycloalkyl, aryl, or heteroaryl. In certain embodiments, R6 is independently selected from the group consisting of hydrogen, -CF3, -C(O)Me, -CO2Me, and -S(O)2CH3. In certain embodiments, R6 is independently selected from the group consisting of -C(O)Me, and -S(O)2R3. In certain embodiments, R6 is -C(O)Me. In certain embodiments, R6 is -S(O)2CH3. [000264] In certain embodiments, m is 1. In certain embodiments, m is 2. In certain embodiments, m is 3. In certain embodiments, m is 4. [000265] Non-limiting examples of include: X5 R5 O , , , , , , , and . [000266] In certain embodiments, is . 59
[000267] In certain embodiments, is . [000268] In certain embodiments, L1 comprises a peptide. In certain embodiments, L1 comprises a dipeptide. In certain embodiments, L1 comprises a tripeptide or a tetrapeptide. In certain embodiments, L1 comprises natural and non-natural amino acids. In certain embodiments, L1 comprises at least one natural amino acid selected from alanine, β-alanine, arginine, asparagine, aspartic acid, cysteine, cystine, glutamic acid, glutamine, glycine, phenylalanine, histidine, isoleucine, lysine, leucine, methionine, proline, serine, threonine, valine, tryptophan, and tyrosine. In certain embodiments, L1 comprises at least one non-natural amino acid selected from sulfoalanine, hydroxyproline (Hyp), beta-alanine, citrulline (Cit), ornithine (Orn), norleucine (Nle), 3-nitrotyrosine, nitroarginine, pyroglutamic acid (Pyr), naphtylalanine (Nal), 2,4-diaminobutyric acid (DAB), methionine sulfoxide, and methionine sulfone. In certain embodiments, L1 comprises at least one natural amino acid selected from alanine, β-alanine, arginine, asparagine, aspartic acid, cysteine, cystine, glutamic acid, glutamine, glycine, phenylalanine, histidine, isoleucine, lysine, leucine, methionine, proline, serine, threonine, valine, tryptophan, and tyrosine and further comprises at least one non- natural amino acid from sulfoalanine, hydroxyproline (Hyp), beta-alanine, citrulline (Cit), ornithine (Orn), norleucine (Nle), 3-nitrotyrosine, nitroarginine, pyroglutamic acid (Pyr), naphtylalanine (Nal), 2,4-diaminobutyric acid (DAB), methionine sulfoxide, and methionine sulfone. [000269] In certain embodiments, L1 comprises a dipeptide selected from the group consisting of -Phe-Lys-, -Val-Ala-, -Val-Lys-, -Ala-Lys-, -Val-Cit-, -Phe-Cit-, -Leu-Cit-, -Ile- Cit-, -Phe-Arg-, and -Trp-Cit-. In particular embodiments, L1 comprises -Val-Ala-. In particular embodiments, L1 comprises -Val-Cit-. [000270] In certain embodiments, L1 comprises a peptide-self immolative group selected from the group consisting of -Phe-Lys-PABC-, -Val-Ala-PABC-, -Val-Lys-PABC-, -Ala-Lys- PABC-, -Val-Cit-PABC-, -Phe-Cit-PABC-, -Leu-Cit-PABC-, -Ile-Cit-PABC-, -Phe-Arg- PABC-, -Trp-Cit-PABC-, and Val-Glu-PABC. In particular embodiments, L1 comprises -Val- Ala-PABC-. In particular embodiments, L1 comprises -Val-Cit-PABC-. The peptide and/or self immolative group can be in either orientation with respect to W2 or SG1 or (SG2). PABC refers 60
to para-aminobenzyloxycarbonyl (PABC), or the corresponding divalent group: . [000271] In certain embodiments, L1 is -Val-Cit-. In certain embodiments, L1 is -Val-Cit- PABC- wherein the -PABC- is covalently bound to W2 and -Val- is covalently bound to SG1 or SG2. [000272] In an alternative embodiment, L1 is absent. 1.3.2.4. Spacer Groups [000273] Spacer groups facilitate spacing of the conjugating group from the other groups of the compounds described herein. This spacing can lead to more efficient conjugation. The spacer group can also stabilize the conjugating group and lead to improved overall IFNα conjugate properties. In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein, SG1 is any divalent spacer group. [000274] In certain embodiments, SG1 is , , , , , , , , , , , , , or ; wherein a and c are an integer independently selected from 0, 1, 2, 3, 4, 5, and 6; and b is an integer selected from 1, 2, 3, 4, 5, and 6. 61
[000275] In certain embodiments, SG1 is , , , , , , , , , or ; wherein a is an integer selected from 0, 1, 2, 3, 4, 5, and 6; and b is an integer selected from 1, 2, 3, 4, 5, and 6. [000276] Non-limiting examples of SG1 include , , , , , and . [000277] In certain embodiments, is . [000278] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein, SG2 is any trivalent spacer group. [000279] In certain embodiments, SG2 is or ; wherein a is independently an integer selected from 0, 1, 2, 3, 4, 5, and 6. [000280] In certain embodiments, SG2 is . [000281] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W1 is -C(O)CH2CH2NH-, W2 is , and RG, L1, and 62
SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is , and RG, L1, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is , RG is or , and SG1 is or , and L1 is as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is , RG is or , L1 is a pH-sensitive linker, and SG1 is as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is , RG is or , L1 is , and SG1 is as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is , RG is or , L1 is , and SG1 is as defined herein. In certain embodiments, W1 63
is -C(O)CH2CH2NH-, W2 is , RG is or , L1 is , and SG1 is . In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is , RG is or , L1 is , and SG1 is . [000282] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W1 is -C(O)CH2CH2NH-, W2 is , and RG, L1, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is , and RG, L1, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is selected from , , and , and RG, L1, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is selected from 64
, , and , RG is or , L1 is a protease cleavable linker, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is selected from , , and , RG is or , L1 is a cathepsin B cleavable linker, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is selected from , , and , RG is or , L1 is comprises a dipeptide, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is selected from , , and , RG is or , L1 is comprises -Val-Cit-, and SG1 is as defined herein. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is selected from 65
, , and , RG is or , L1 comprises a peptide-self immolative group, and SG1 is . In certain embodiments, W1 is -C(O)(CH2CH2)NH-, W2 is selected from , , and , RG is or , L1 is -Val-Cit-PABC-, and SG1 is . [000283] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W1 is -C(O)CH2CH2CH2CH2NH-, W2 is absent, and RG, L1, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2CH2CH2NH-, W2 is absent, RG is or , L1 is a protease cleavable linker, and SG1 is as defined herein. In certain embodiments, W1 is -C(O)CH2CH2CH2CH2NH-, W2 is absent, RG is or , L1 is a cathepsin B cleavable linker, and SG1 is as defined herein. In certain embodiments, W1 is -C(O)CH2CH2CH2CH2NH-, W2 is absent, RG 66
is or , L1 is comprises a dipeptide, and SG1 is as defined herein. In certain embodiments, W1 is -C(O)CH2CH2CH2CH2NH-, W2 is absent, RG is or , L1 is comprises -Val-Cit-, and SG1 is as defined herein. In certain embodiments, W1 is -C(O)CH2CH2CH2CH2NH-, W2 is absent, RG is or , L1 comprises a peptide-self immolative group, and SG1 is . In certain embodiments, W1 is -C(O)CH2CH2CH2CH2NH-, W2 is absent, RG is or , L1 is -Val-Cit-PABC-, and SG1 is . [000284] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W1 is -C(O)(C1-6alkylene)NH-, W2 is , and RG, L1, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)(C1-6alkylene)NH- and W2 is . In certain embodiments, W1 is -C(O)CH2CH2NH- and W2 is . In certain embodiments, W1 is -C(O)CH2CH2NH- and W2 is . In certain 67
embodiments, W1 is -C(O)CH2CH2NH- and W2 is . In certain embodiments, W1 is -C(O)CH2CH2NH- and W2 is . In certain embodiments, W1 is -C(O)CH2CH2NH- and W2 is . In certain embodiments, W1 is -C(O)CH2CH2NH- and W2 is . In certain embodiments, W1 is -C(O)(CH2CH2)NH-, W2 is , RG is or , SG is , and L1 is absent. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is , RG is or , SG is . and L1 is absent. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is , RG is or , SG is , and L1 is 68
absent. In certain embodiments, W1 is -C(O)CH2CH2NH-, W2 is , RG is or , SG is , and L1 is absent. [000285] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L1), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W1 is -C(O)(C1-6alkylene)-, W2 is , and RG, L1, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2CH2CH2-, W2 is , and RG, L1, and SG1 are as defined herein. In certain embodiments, W1 O N 1-6 is -C(O)CH2CH2CH2CH2-, W2 is R4 , and RG, L1, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2CH2CH2-, W2 is , and RG, L1, and SG1 are as defined herein. In certain embodiments, W1 is -C(O)CH2CH2CH2CH2-, W2 is , RG is or , and SG1 is , and L1 is as defined herein. In certain embodiments, W1 is -C(O)CH2CH2CH2CH2-, W2 is , RG is or , L1 is absent, and SG1 is . In certain 69
embodiments, W1 is -C(O)CH2CH2CH2CH2-, W2 is , RG is or , L1 is absent, and SG1 is . [000286] In certain embodiments, provided herein is a conjugate comprising a linker according to Formula (L2), or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein W1 is -C(O)(C1-6alkylene)-, W2 is , and RG, L1, and SG2 are as defined herein. In certain embodiments, W1 is -C(O)(C1-6alkylene)-, W2 is , and RG, L1, and SG2 are as defined herein. In certain embodiments, W1 is -C(O)(C1-6alkylene)-, W2 is , and RG, L1, and SG2 are as defined herein. In certain embodiments, W1 is -C(O)(C1-6alkylene)-, W2 is , and RG is or , L1, and SG2 is as defined herein. In certain embodiments, W1 is -C(O)(C1-6alkylene)-, W2 is , and RG is or 70
, L1 is absent, and SG2 is as defined herein. In certain embodiments, W1 is -C(O)(C1-6alkylene)-, W2 is , and RG is or , L1 is absent, and SG2 is . In certain embodiments, W1 is -C(O)(C1-6alkylene)-, W2 is , and RG is or , L1 is absent, and SG2 is . 1.3.3. Conjugating Groups and Residues Thereof [000287] In an embodiment, provided herein is a conjugate or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; according to any of the following formulas: 71
, , , , 72
, , , 73
, , , and where COMP indicates a residue of the IFNα polypeptide, MM indicates a masking moiety, and L1 is as defined herein. 74
[000288] In any of the foregoing embodiments, MM is a polymer (POLY), for example a water-soluble polymer. In any of the foregoing embodiments, POLY is , wherein n1 is an integer from 1 to 10,000, inclusive, and represents attachment to the remainder of the compound or conjugate. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 5000. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 2500. In certain embodiments, including any of the foregoing, n1 is an integer between 1 to 1500. In certain embodiments, including any of the foregoing, n1 is an integer between 100 to 1000. In certain embodiments, n1 is an integer from 100 to 700. In certain embodiments, n1 is an integer from 300 to 700. In certain embodiments, n1 is an integer from 400 to 500. In certain embodiments, n1 is an integer from 600 to 700. In certain embodiments, including any of the foregoing, n1 is an integer between 100 to 500. [000289] The present disclosure encompasses each and every regioisomer of the conjugate structures depicted below: 75
76
wherein COMP is an IFNα polypeptide. [000290] In any of the foregoing embodiments, n1 is an integer between 300 and 800, inclusive. In any of the foregoing embodiments, n1 is an integer between 400 and 600, inclusive. In any of the foregoing embodiments, n1 is an integer between 400 and 500, inclusive. [000291] In any of the foregoing embodiments, n is an integer from 1 to 8. In any of the foregoing embodiments, n is 1. In any of the foregoing embodiments, n is 2. In any of the 77
foregoing embodiments, n is 3. In any of the foregoing embodiments, n is 4. In any of the foregoing embodiments, n is 6. In any of the foregoing embodiments, n is 8. [000292] The present disclosure encompasses the conjugate structures depicted below: 78
80
81
82
wherein n is an integer from 1 to 8 and COMP is an IFNα polypeptide. [000293] In any of the foregoing embodiments, the bracketed structure can be covalently bonded to one or more non-natural amino acids or modified amino acids of the IFNα polypeptide, wherein the one or more non-natural amino acids or modified amino acids are located at sites selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156 of SEQ ID NO: 33. In the foregoing embodiment, the one or more non-natural amino acid or modified amino acid is two non-natural amino acids or modified amino acids, respectively. In the foregoing embodiment, the one or more non-natural amino acid or modified amino acid is three non-natural amino acids or modified amino acids, respectively. In the foregoing embodiment, the one or more non-natural amino acid or modified amino acid is four non-natural amino acids or modified amino acids, respectively. In the foregoing embodiment, the one or more non-natural amino acid or modified amino acid is five or more non-natural amino acids or modified amino acids, respectively. [000294] In any of the foregoing embodiments, the bracketed structure can be covalently bonded to one or more non-natural amino acids or modified amino acids of the IFNα polypeptide, wherein the one or more non-natural amino acids or modified amino acids are located at sites selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33. In any of the foregoing embodiments, the bracketed structure can be covalently bonded to two non-natural amino acids or modified amino acids at sites selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33. In any of the foregoing embodiments, the bracketed structure can be covalently bonded to three non- natural amino acids or modified amino acids at sites selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33. In any of the foregoing embodiments, the bracketed structure can be covalently bonded to four non-natural amino acids or modified amino acids at sites selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33. In any of the foregoing embodiments, the bracketed structure can be covalently bonded to five or more non-natural amino acids or modified amino acids at sites selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33. [000295] In certain embodiments, the conjugate comprises a non-natural amino acid or modified amino acid located at amino acid position H7. In certain embodiments, the conjugate comprises a non-natural amino acid or modified amino acid located at amino acid position Q40. In certain embodiments, the conjugate comprises a non-natural amino acid or modified amino 83
acid located at amino acid position E41. In certain embodiments, the conjugate comprises a non-natural amino acid or modified amino acid located at amino acid position N45. In certain embodiments, the conjugate comprises a non-natural amino acid or modified amino acid located at amino acid position E51. In certain embodiments, the conjugate comprises a non- natural amino acid or modified amino acid located at amino acid position N156. [000296] In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at H7 and Q40. In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at H7 and Q40 and further comprising at least one additional non-natural amino acid or modified amino acid at a position selected from E51, N45, and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at H7, Q40 and E51. In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at H7, Q40 and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at H7, Q40, N45, and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at H7 and E51. In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at H7, E51 and N156. [000297] In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at Q40 and E51. In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at Q40 and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at E51 and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at Q40, E51, and N156. [000298] In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at Q40 and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids at Q40 and N156 and further comprising at least one non-natural amino acid or modified amino acid at a position selected from H7 and E51. [000299] In some embodiments, the conjugate comprises an IFNα polypeptide comprising at least one non-natural amino acid or modified amino acid in a position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprising at least one non- 84
natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136. [000300] In some embodiments, the conjugate comprises an IFNα polypeptide comprising at least two non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprising at least one non- natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136. [000301] In some embodiments, the conjugate comprises an IFNα polypeptide comprising at least three non-natural amino acids or modified amino acids in positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156 and further comprising at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136. [000302] In some embodiments, the conjugate comprises an IFNα polypeptide comprising non-natural amino acids or modified amino acids in positions Q40 and N156 and further comprising at least one non-natural amino acid or modified amino acid at a position selected from the group consisting of: D2, L3, Q5, T6, E42, Q46, Q48, K49, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, and S136. [000303] In any of the foregoing embodiments, the non-natural or modified amino acid is a non-natural amino acid. In any of the foregoing embodiments, the non-natural or modified amino acid is a modified amino acid. [000304] In certain embodiments, one or more linkers and/or masking moieties are conjugated to the one or more non-natural amino acids or modified amino acids. In certain embodiments, the non-natural amino acid residue or modified amino acid residue comprises a residue of a moiety selected from the group consisting of amino, carboxy, acetyl, hydrazino, hydrazido, semicarbazido, sulfanyl, azido and alkynyl. In certain embodiments, the modified amino acid residue is selected from the group consisting of: p-acetyl-L-phenylalanine, O- methyl-L-tyrosine, 3-methyl-phenylalanine, O-4-allyl-L-tyrosine, 4-propyl-L-tyrosine, fluorinated phenylalanine, isopropyl-L-phenylalanine, p-azido-L-phenylalanine, p-acetyl-L- phenylalanine, p-benzoyl-L-phenylalanine, p-iodo-phenylalanine, p-bromophenylalanine, p- amino-L-phenylalanine, isopropyl-L-phenylalanine, p-propargyloxy-phenylalanine, and p- azidomethyl-L-phenylalanine residues. In certain embodiments, the modified amino acid 85
residue is para-azido-L-phenylalanine. In certain embodiments, the modified amino acid is para-azidomethyl-L-phenylalanine (pAMF). In certain embodiments, the modified amino acid is para-azidomethyl-L-phenylalanine (pAMF) and is located at an amino acid position selected from amino acid positions: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156 of SEQ ID NO: 33. In certain embodiments, the modified amino acid is para- azidomethyl-L-phenylalanine (pAMF) and is located at an amino acid position selected from amino acid positions: H7, Q40, E41, N45, E51, and N156 of SEQ ID NO: 33. [000305] In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p-azidomethyl-L-phenylalanine residue in at least one amino acid position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156. [000306] In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p-azidomethyl-L-phenylalanine residue in at least one amino acid position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p-azidomethyl-L-phenylalanine residue in at least two amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p- azidomethyl-L-phenylalanine residue in at least three amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p-azidomethyl-L-phenylalanine residue in at least four amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p- azidomethyl-L-phenylalanine residue in at least five amino acid positions selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p-azidomethyl-L-phenylalanine residue in positions H7, Q40, E41, N45, E51, and N156. [000307] In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p-azidomethyl-L-phenylalanine residue at H7. In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p-azidomethyl-L-phenylalanine residue at Q40. In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p-azidomethyl-L- phenylalanine residue at E41. In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p-azidomethyl-L-phenylalanine residue at N45. In some 86
embodiments, the conjugate comprises an IFNα polypeptide comprising p-azidomethyl-L- phenylalanine residue at E51. In some embodiments, the conjugate comprises an IFNα polypeptide comprising a p-azidomethyl-L-phenylalanine residue at N156. [000308] In some embodiments, the conjugate comprises an IFNα polypeptide comprising p- azidomethyl-L-phenylalanine residues at H7 and Q40. In some embodiments, the conjugate comprises an IFNα polypeptide comprising p-azidomethyl-L-phenylalanine residues at H7 and Q40 and further comprising at least one additional p-azidomethyl-L-phenylalanine residue at a position selected from E51, N45, and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising p-azidomethyl-L-phenylalanine residues at H7, Q40 and E51. In some embodiments, the conjugate comprises an IFNα polypeptide comprising p- azidomethyl-L-phenylalanine residues at H7, Q40 and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising p-azidomethyl-L-phenylalanine residues at H7, Q40, N45, and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising p-azidomethyl-L-phenylalanine residues at H7 and E51. In some embodiments, the conjugate comprises an IFNα polypeptide comprising p-azidomethyl-L- phenylalanine residues at H7, E51 and N156. [000309] In some embodiments, the conjugate comprises an IFNα polypeptide comprising p- azidomethyl-L-phenylalanine residues at Q40 and E51. In some embodiments, the conjugate comprises an IFNα polypeptide comprising p-azidomethyl-L-phenylalanine residues at Q40 and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising p- azidomethyl-L-phenylalanine residues at E51 and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising p-azidomethyl-L-phenylalanine residues at Q40, E51, and N156. [000310] In some embodiments, the conjugate comprises an IFNα polypeptide comprising p- azidomethyl-L-phenylalanine residues at Q40 and N156. In some embodiments, the conjugate comprises an IFNα polypeptide comprising p-azidomethyl-L-phenylalanine residues at Q40 and N156 and further comprising at least one p-azidomethyl-L-phenylalanine residues at a position selected from H7 and E51. [000311] In certain embodiments, the masking moiety is a water soluble polymer (POLY) selected from the group consisting of is polyethylene glycol (PEG), poly(propylene glycol) (PPG), copolymers of ethylene glycol and propylene glycol, poly(oxyethylated polyol), poly(olefinic alcohol), poly(vinylpyrrolidone), poly(hydroxyalkylmethacrylamide), poly(hydroxyalkylmethacrylate), poly(saccharides), poly(a-hydroxy acid), poly(vinyl alcohol), polyphosphazene, polyoxazolines (POZ), poly(N-acryloylmorpholine), and combinations 87
thereof. In certain embodiments, the water soluble polymer is PEG. In certain embodiments, the PEG has an average molecular weight of between about 5KDa and about 50 KDa. In certain embodiments, the PEG is selected from the group consisting of a linear or branched PEG molecule having an average molecular weight of 10Kda, 20Kda, 30Kda, or 40Kda. In certain embodiments, the PEG has an average molecular weight of 30Kda. In certain embodiments, the PEG has an average molecular weight of 40Kda. In certain embodiments, the conjugate has an extended half-life compared to an identical polypeptide lacking the water-soluble polymer. [000312] In certain embodiments, provided herein are IFNα polypeptides comprising one or more non-natural amino acids or modified amino acids. These non-natural amino acids or modified amino acids can facilitate conjugation to a masking moiety or linker to form conjugates. In certain embodiments, the non-natural amino acid or modified amino acid is at a position selected from the group consisting of: D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156. In certain embodiments, the non-natural amino acid or modified amino acid is at a position selected from the group consisting of: H7, Q40, E41, N45, E51, and N156. In certain embodiments, a polynucleotide is provided, encoding one of these IFNα polypeptides. In certain embodiments, the polynucleotide encodes a TAG codon to facilitate incorporation of a non-natural amino acid or modified amino acid according to the expression techniques described herein. Any non-natural amino acid or modified amino acid can be incorporated at the TAG position. In certain embodiments, the non-natural amino acid or modified amino acid is one described herein. In certain embodiments, the non-natural or modified amino acid is a modified amino acid and the modified amino acid is p- azidomethylphenylalanine. [000313] In certain embodiments, the IFNα polypeptide has an amino acid sequence according to SEQ ID NO: 1, SEQ ID NO: 2, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22, or SEQ ID NO: 23, SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 29, SEQ ID NO: 30, SEQ ID NO: 31, SEQ ID NO: 35, SEQ ID NO: 36, and SEQ ID NO: 37. [000314] In certain embodiments, the IFNα polypeptide has an amino acid sequence selected from SEQ ID NO: 11, SEQ ID NO: 13, SEQ ID NO: 16, and SEQ ID NO: 18. In certain embodiments, the IFNα polypeptide conjugate comprises a PEG having an average molecular weight of 20Kda, 30Kda or 40Kda. 88
[000315] In certain embodiments, the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 11. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 11. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 11. [000316] In certain embodiments, the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 16. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 16. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 16. [000317] In certain embodiments, the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 18. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 18. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 18. [000318] In certain embodiments, the polypeptides have an amino acid sequence at least 90% identical to SEQ ID NO: 33. In certain embodiments, the polypeptides have an amino acid sequence at least 95% identical to SEQ ID NO: 33. In certain embodiments, the polypeptides have an amino acid sequence at least 98% identical to SEQ ID NO: 33. [000319] In certain embodiments, the TAG (*) position of the above amino acid sequences indicates a non-natural amino acid or modified amino acid. In certain embodiments, the non- natural amino acid or modified amino acid is one described herein. In certain embodiments, the non-natural or modified amino acid is a modified amino acid and the modified amino acid is p-azidomethylphenylalanine. 1.4. Vectors, Host Cells, and Recombinant Methods [000320] Also provided are isolated nucleic acids encoding IFNα polypeptide, vectors and host cells comprising the nucleic acids, and recombinant techniques for the production of the IFNα polypeptide and cytokines. [000321] For recombinant production of the IFNα polypeptides, the nucleic acid encoding it may be isolated and inserted into a replicable vector for further cloning (i.e., amplification of the DNA) or expression. In some aspects, the nucleic acid may be produced by homologous recombination, for example as described in U.S. Patent No.5,204,244. [000322] Many different vectors are known in the art. The vector components generally include, but are not limited to, one or more of the following: a signal sequence, an origin of replication, one or more marker genes, an enhancer element, a promoter, and a transcription termination sequence, for example as described in U.S. Patent No.5,534,615. 89
[000323] Illustrative examples of suitable host cells are provided below. These host cells are not meant to be limiting. [000324] Suitable host cells include any prokaryotic (e.g., bacterial), lower eukaryotic (e.g., yeast), or higher eukaryotic (e.g., mammalian) cells. Suitable prokaryotes include eubacteria, such as Gram-negative or Gram-positive organisms, for example, Enterobacteriaceae such as Escherichia (E. coli), Enterobacter, Erwinia, Klebsiella, Proteus, Salmonella (S. typhimurium), Serratia (S. marcescans), Shigella, Bacilli (B. subtilis and B. licheniformis), Pseudomonas (P. aeruginosa), and Streptomyces. One useful E. coli cloning host is E. coli 294, although other strains such as E. coli B, E. coli X1776, and E. coli W3110 are suitable. [000325] In addition to prokaryotes, eukaryotic microbes such as filamentous fungi or yeast are also suitable cloning or expression hosts for IFNα polypeptide-encoding vectors. Saccharomyces cerevisiae, or common baker’s yeast, is a commonly used lower eukaryotic host microorganism. However, a number of other genera, species, and strains are available and useful, such as Schizosaccharomyces pombe, Kluyveromyces (K. lactis, K. fragilis, K. bulgaricus K. wickeramii, K. waltii, K. drosophilarum, K. thermotolerans, and K. marxianus), Yarrowia, Pichia pastoris, Candida (C. albicans), Trichoderma reesia, Neurospora crassa, Schwanniomyces (S. occidentalis), and filamentous fungi such as, for example Penicillium, Tolypocladium, and Aspergillus (A. nidulans and A. niger). [000326] Useful mammalian host cells include COS-7 cells, HEK293 cells; baby hamster kidney (BHK) cells; Chinese hamster ovary (CHO); mouse cells; African green monkey kidney cells (VERO-76), and the like. [000327] The host cells used to produce the IFNα polypeptides may be cultured in a variety of media. Commercially available media such as, for example, Ham’s F10, Minimal Essential Medium (MEM), RPMI-1640, and Dulbecco’s Modified Eagle’s Medium (DMEM) are suitable for culturing the host cells. In addition, any of the media described in Ham et al., Meth. Enz., 1979, 58:44; Barnes et al., Anal. Biochem., 1980, 102:255; and U.S. Patent Nos. 4,767,704, 4,657,866, 4,927,762, 4,560,655, and 5,122,469, or WO 90/03430 and WO 87/00195 may be used. [000328] Any of these media may be supplemented as necessary with hormones and/or other growth factors (such as insulin, transferrin, or epidermal growth factor), salts (such as sodium chloride, calcium, magnesium, and phosphate), buffers (such as HEPES), nucleotides (such as adenosine and thymidine), antibiotics, trace elements (defined as inorganic compounds usually present at final concentrations in the micromolar range), and glucose or an equivalent energy 90
source. Any other necessary supplements may also be included at appropriate concentrations that would be known to those skilled in the art. [000329] The culture conditions, such as temperature, pH, and the like, are those previously used with the host cell selected for expression, and will be apparent to the ordinarily skilled artisan. [000330] In some embodiments, a IFNα polypeptide is produced by a method comprising the step of culturing a host cell described herein. In certain embodiments, the host cell comprises a nucleic acid, vector, or expression vector described herein for producing the IFNα polypeptide. In certain embodiments, the IFNα polypeptide variant comprises one or more non- natural amino acids or modified amino acids as described herein. In certain embodiments, the host cell further comprises a nucleic acid, vector, or expression vector encoding an aminoacyl tRNA synthetase (RS) specific for the non-natural amino acid or modified amino acid. In certain embodiments, the host cell further comprises a nucleic acid, vector, or expression vector encoding a tRNA specific for the non-natural amino acid or modified amino acid. In certain embodiments, any or each nucleic acid, vector, or expression vector is codon optimized for the host cell. In certain embodiments, the non-natural or modified amino acid is a modified amino acid and the modified amino acid is p-azidomethylphenylalanine. In certain embodiments, the host cell is E. coli. In certain embodiments, the host cells, for instance E. coli host cells, have an oxidative cytoplasm, for instance as described in PCT/US2019/060345. [000331] When using recombinant techniques, the IFNα polypeptides can be produced intracellularly, in the periplasmic space, or directly secreted into the medium. If the IFNα polypeptide is produced intracellularly, as a first step, the particulate debris, either host cells or lysed fragments, is removed, for example, by centrifugation or ultrafiltration. For example, Carter et al. (Bio/Technology, 1992, 10:163-167) describes a procedure for isolating polypeptides which are secreted to the periplasmic space of E. coli. Briefly, cell paste is thawed in the presence of sodium acetate (pH 3.5), EDTA, and phenylmethylsulfonylfluoride (PMSF) over about 30 min. Cell debris can be removed by centrifugation. [000332] In some embodiments, the IFNα polypeptide is produced in a cell-free system. In some aspects, the cell-free system is an in vitro transcription and translation system as described in Yin et al., mAbs, 2012, 4:217-225, incorporated by reference in its entirety. In some aspects, the cell-free system utilizes a cell-free extract from a eukaryotic cell or from a prokaryotic cell. In some aspects, the prokaryotic cell is E. coli. Cell-free expression of the IFNα polypeptide may be useful, for example, where the IFNα polypeptide accumulates in a cell as an insoluble aggregate, or where yields from periplasmic expression are low. 91
[000333] Where the IFNα polypeptide is secreted into the medium, supernatants from such expression systems are generally first concentrated using a commercially available protein concentration filter, for example, an Amicon® or Millipore® Pellcon® ultrafiltration unit. A protease inhibitor such as PMSF may be included in any of the foregoing steps to inhibit proteolysis and antibiotics may be included to prevent the growth of adventitious contaminants. [000334] The IFNα polypeptide composition prepared from the cells can be purified using, for example, hydroxyapatite chromatography, gel electrophoresis, dialysis, and affinity chromatography, with affinity chromatography being a particularly useful purification technique. [000335] The matrix to which the affinity ligand is attached is most often agarose, but other matrices are available. Mechanically stable matrices such as controlled pore glass or poly(styrenedivinyl)benzene allow for faster flow rates and shorter processing times than can be achieved with agarose. [000336] Other techniques for protein purification, such as fractionation on an ion-exchange column, ethanol precipitation, Reverse Phase HPLC, chromatography on silica, chromatography on heparin Sepharose®, chromatofocusing, SDS-PAGE, and ammonium sulfate precipitation are also available, and can be applied by one of skill in the art. [000337] Following any preliminary purification step(s), the mixture comprising the IFNα polypeptide of interest and contaminants may be subjected to low pH hydrophobic interaction chromatography using an elution buffer at a pH between about 2.5-4.5, generally performed at low salt concentrations (e.g., from about 0-0.25 M salt). [000338] In some embodiments, the IFNα polypeptide is conjugated, for instance as described below. 1.5. Conjugation [000339] The conjugates can be prepared by standard techniques. In certain embodiments, an IFNα is contacted with a masking moiety or linker precursor under conditions suitable for forming a bond from the IFNα polypeptide to the masking moiety to form an IFNα-masking moiety conjugate. In certain embodiments, an IFNα polypeptide is contacted with a linker precursor under conditions suitable for forming a bond from the IFNα polypeptide to the linker. The resulting IFNα polypeptide-linker is contacted with a masking moiety precursor under conditions suitable for forming a bond from the IFNα -linker to the masking moiety to form an IFNα-linker-masking moiety conjugate. In certain embodiments, a masking moiety precursor is contacted with a linker precursor under conditions suitable for forming a bond from the 92
masking moiety to the linker. The resulting masking moiety-linker is contacted with an IFNα under conditions suitable for forming a bond from the masking moiety-linker to the IFNα to form an IFNα-linker-masking moiety conjugate. Suitable linkers for preparing the IFNα conjugates are disclosed herein, and exemplary conditions for conjugation are described in the Examples below. [000340] In some embodiments, an IFNα conjugate is prepared by contacting an IFNα as disclosed herein with a linker precursor having a structure selected from: , , , 93
O O O O O H O O N O N N O H N 19 H O O N N n1 H O H HN H2N O and . [000341] In any of the foregoing embodiments, n1 is an integer between 300 and 800, inclusive. In any of the foregoing embodiments, n1 is an integer between 400 and 600, inclusive. In any of the foregoing embodiments, n1 is an integer between 400 and 500, inclusive. [000342] In some embodiments, an IFNα conjugate is prepared by contacting an IFNα as disclosed herein with a linker precursor having a structure selected from: 94
LP1, LP2, LP3, LP4, LP5, 95
LP6, LP7, and LP8. 1.6. Pharmaceutical Compositions and Methods of Administration [000343] The IFNα polypeptides or conjugates provided herein can be formulated into pharmaceutical compositions using methods available in the art and those disclosed herein. Any of the IFNα polypeptides or conjugates provided herein can be provided in the appropriate pharmaceutical composition and be administered by a suitable route of administration. [000344] The methods provided herein encompass administering pharmaceutical compositions comprising at least one IFNα polypeptides or conjugates provided herein and one or more compatible and pharmaceutically acceptable carriers. In this context, the term “pharmaceutically acceptable” means approved by a regulatory agency of the Federal or a state government or listed in the U.S. Pharmacopeia or other generally recognized pharmacopeia for 96
use in animals, and more particularly in humans. The term “carrier” includes a diluent, excipient, or vehicle with which the therapeutic is administered. Such pharmaceutical carriers can be sterile liquids, such as water and oils, including those of petroleum, animal, vegetable or synthetic origin, such as peanut oil, soybean oil, mineral oil, sesame oil and the like. Water can be used as a carrier when the pharmaceutical composition is administered intravenously. Saline solutions and aqueous dextrose and glycerol solutions can also be employed as liquid carriers, particularly for injectable solutions. Examples of suitable pharmaceutical carriers are described in Martin, E.W., Remington’s Pharmaceutical Sciences. [000345] In clinical practice, the pharmaceutical compositions or IFNα polypeptide or conjugate provided herein may be administered by any route known in the art. In certain embodiments, a pharmaceutical composition or IFNα polypeptide or conjugate provided herein is administered parenterally. [000346] The compositions for parenteral administration can be emulsions or sterile solutions. Parenteral compositions may include, for example, propylene glycol, polyethylene glycol, vegetable oils, and injectable organic esters (e.g., ethyl oleate). These compositions can also contain wetting, isotonizing, emulsifying, dispersing and stabilizing agents. Sterilization can be carried out in several ways, for example using a bacteriological filter, by radiation or by heating. Parenteral compositions can also be prepared in the form of sterile solid compositions which can be dissolved at the time of use in sterile water or any other injectable sterile medium. [000347] In certain embodiments, a composition provided herein is a pharmaceutical composition or a single unit dosage form. Pharmaceutical compositions and single unit dosage forms provided herein comprise a prophylactically or therapeutically effective amount of one or more prophylactic or therapeutic IFNα polypeptide or conjugate. [000348] Typical pharmaceutical compositions and dosage forms comprise one or more excipients. Suitable excipients are well-known to those skilled in the art of pharmacy, and non- limiting examples of suitable excipients include starch, glucose, lactose, sucrose, gelatin, malt, rice, flour, chalk, silica gel, sodium stearate, glycerol monostearate, talc, sodium chloride, dried skim milk, glycerol, propylene, glycol, water, ethanol and the like. Whether a particular excipient is suitable for incorporation into a pharmaceutical composition or dosage form depends on a variety of factors well known in the art including, but not limited to, the way in which the dosage form will be administered to a subject and the specific IFNα polypeptide or conjugate in the dosage form. The composition or single unit dosage form, if desired, can also contain minor amounts of wetting or emulsifying agents, or pH buffering agents. 97
[000349] Lactose free compositions provided herein can comprise excipients that are well known in the art and are listed, for example, in the U.S. Pharmocopeia (USP) SP (XXI)/NF (XVI). In general, lactose free compositions comprise an active ingredient, a binder/filler, and a lubricant in pharmaceutically compatible and pharmaceutically acceptable amounts. Exemplary lactose free dosage forms comprise an active ingredient, microcrystalline cellulose, pre gelatinized starch, and magnesium stearate. [000350] Components of the pharmaceutical composition can be supplied either separately or mixed together in unit dosage form, for example, as a dry lyophilized powder or water free concentrate. Where the composition is to be administered by infusion, it can be dispensed with an infusion bottle containing sterile pharmaceutical grade water or saline. Where the composition is administered by injection, an ample of sterile water for injection or saline can be provided so that the ingredients may be mixed prior to administration. [000351] In some embodiments, the pharmaceutical composition is supplied as a dry sterilized lyophilized powder that is capable of being reconstituted to the appropriate concentration for administration to a subject. In some embodiments, IFNα polypeptides or conjugates are supplied as a water free concentrate. [000352] In another embodiment, the pharmaceutical composition is supplied in liquid form. In some embodiments, the pharmaceutical composition is provided in liquid form and is substantially free of surfactants and/or inorganic salts. [000353] In some embodiments, the pharmaceutical composition is formulated as a salt form. Pharmaceutically acceptable salts include those formed with anions such as those derived from hydrochloric, phosphoric, acetic, oxalic, tartaric acids, etc., and those formed with cations such as those derived from sodium, potassium, ammonium, calcium, ferric hydroxides, isopropylamine, triethylamine, 2-ethylamino ethanol, histidine, procaine, etc. [000354] Further encompassed herein are anhydrous pharmaceutical compositions and dosage forms comprising an IFNα polypeptide or conjugate, since water can facilitate the degradation of some IFNα polypeptide or conjugate. [000355] Anhydrous pharmaceutical compositions and dosage forms provided herein can be prepared using anhydrous or low moisture containing ingredients and low moisture or low humidity conditions. Pharmaceutical compositions and dosage forms that comprise lactose and at least one active ingredient that comprises a primary or secondary amine can be anhydrous if substantial contact with moisture and/or humidity during manufacturing, packaging, and/or storage is expected. 98
[000356] An anhydrous pharmaceutical composition should be prepared and stored such that its anhydrous nature is maintained. Accordingly, anhydrous compositions can be packaged using materials known to prevent exposure to water such that they can be included in suitable formulary kits. Examples of suitable packaging include, but are not limited to, hermetically sealed foils, plastics, unit dose containers (e.g., vials), blister packs, and strip packs. [000357] Further provided are pharmaceutical compositions and dosage forms that comprise one or more excipients that reduce the rate by which an IFNα polypeptide or conjugate will decompose. Such excipients, which are referred to herein as “stabilizers,” include, but are not limited to, antioxidants such as ascorbic acid, pH buffers, or salt buffers. [000358] Parenteral Dosage Forms [000359] In certain embodiments, provided are parenteral dosage forms. Parenteral dosage forms can be administered to subjects by various routes including, but not limited to, subcutaneous, intravenous (including bolus injection), intramuscular, and intraarterial. Because their administration typically bypasses subjects’ natural defenses against contaminants, parenteral dosage forms are typically, sterile or capable of being sterilized prior to administration to a subject. Examples of parenteral dosage forms include, but are not limited to, solutions ready for injection, dry products ready to be dissolved or suspended in a pharmaceutically acceptable vehicle for injection, suspensions ready for injection, and emulsions. [000360] Suitable vehicles that can be used to provide parenteral dosage forms are well known to those skilled in the art. Examples include, but are not limited to: Water for Injection USP; aqueous vehicles such as, but not limited to, Sodium Chloride Injection, Ringer’s Injection, Dextrose Injection, Dextrose and Sodium Chloride Injection, and Lactated Ringer’s Injection; water miscible vehicles such as, but not limited to, ethyl alcohol, polyethylene glycol, and polypropylene glycol; and non-aqueous vehicles such as, but not limited to, corn oil, cottonseed oil, peanut oil, sesame oil, ethyl oleate, isopropyl myristate, and benzyl benzoate. [000361] Excipients that increase the solubility of one or more of the IFNα polypeptides or conjugates disclosed herein can also be incorporated into the parenteral dosage forms. 1.7. Dosage and Unit Dosage Forms [000362] In human therapeutics, the doctor will determine the posology which he considers most appropriate according to a preventive or curative treatment and according to the age, weight, stage of the infection and other factors specific to the subject to be treated. 99
[000363] The amount of the IFNα polypeptide or conjugate or composition which will be effective in the prevention or treatment of a disorder or one or more symptoms thereof will vary with the nature and severity of the disease or condition, and the route by which the IFNα polypeptide or conjugate is administered. The frequency and dosage will also vary according to factors specific for each subject depending on the specific therapy (e.g., therapeutic or prophylactic agents) administered, the severity of the disorder, disease, or condition, the route of administration, as well as age, body, weight, response, and the past medical history of the subject. Effective doses may be extrapolated from dose-response curves derived from in vitro or animal model test systems. [000364] The dose can be administered according to a suitable schedule, for example, once, two times, three times, or for times weekly. It may be necessary to use dosages of the IFNα polypeptide or conjugate outside the ranges disclosed herein in some cases, as will be apparent to those of ordinary skill in the art. Furthermore, it is noted that the clinician or treating physician will know how and when to interrupt, adjust, or terminate therapy in conjunction with subject response. [000365] Different therapeutically effective amounts may be applicable for different diseases and conditions, as will be readily known by those of ordinary skill in the art. Similarly, amounts sufficient to prevent, manage, treat or ameliorate such disorders, but insufficient to cause, or sufficient to reduce, adverse effects associated with the IFNα polypeptide or conjugate provided herein are also encompassed by the herein described dosage amounts and dose frequency schedules. Further, when a subject is administered multiple dosages of a composition provided herein, not all of the dosages need be the same. For example, the dosage administered to the subject may be increased to improve the prophylactic or therapeutic effect of the composition or it may be decreased to reduce one or more side effects that a particular subject is experiencing. [000366] In certain embodiments, treatment or prevention can be initiated with one or more loading doses of an IFNα polypeptide or conjugate or composition provided herein followed by one or more maintenance doses. [000367] In certain embodiments, a dose of an IFNα polypeptide or conjugate or composition provided herein can be administered to achieve a steady-state concentration of the IFNα polypeptide or conjugate in blood or serum of the subject. The steady-state concentration can be determined by measurement according to techniques available to those of skill or can be based on the physical characteristics of the subject such as height, weight and age. [000368] Therapeutic Applications 100
[000369] For therapeutic applications, IFNα polypeptides or conjugates disclosed herein are administered to a mammal, generally a human, in a pharmaceutically acceptable dosage form such as those known in the art and those discussed above. For example, the IFNα polypeptides or conjugates disclosed herein may be administered to a human intravenously as a bolus or by continuous infusion over a period of time, by intravenous, intramuscular, intraperitoneal, intra- cerebrospinal, subcutaneous, intra-articular, intrasynovial, intrathecal, or intratumoral routes. The IFNα polypeptides or conjugates can also be suitably administered by peritumoral, intralesional, or perilesional routes, to exert local as well as systemic therapeutic effects. [000370] A therapeutically effective amount of the IFNα polypeptide or conjugate or composition is an amount that is effective to reduce the severity, the duration and/or the symptoms of a particular disease or condition. The amount of the IFNα polypeptide or conjugate or composition that will be therapeutically effective in the prevention, management, treatment and/or amelioration of a particular disease can be determined by standard clinical techniques. The precise amount of the IFNα polypeptide or conjugate or composition to be administered will depend, in part, on the route of administration, the seriousness of the particular disease or condition, and should be decided according to the judgment of the practitioner and each subject’s circumstances. 1.8. Methods of Treatment [000371] For therapeutic applications, the IFNα polypeptides and conjugates provided herein can be administered to a mammal, generally a human, for the treatment of any disease, disorder, or condition that would benefit from the stimulation of amplification of the immune response. In certain embodiments, the disease or condition is abnormal cellular proliferation. In certain embodiments, the disease, disorder, or condition is cancer. [000372] Any suitable cancer may be treated with the IFNα polypeptides and conjugates provided herein. Illustrative suitable cancers include, for example, acute lymphoblastic leukemia (ALL), acute myeloid leukemia (AML), adrenocortical carcinoma, anal cancer, appendix cancer, astrocytoma, basal cell carcinoma, brain tumor, bile duct cancer, bladder cancer, bone cancer, breast cancer (including triple-negative breast cancer, or TNBC), bronchial tumor, carcinoma of unknown primary origin, cardiac tumor, cervical cancer, chordoma, colon cancer, colorectal cancer, craniopharyngioma, ductal carcinoma, embryonal tumor, endometrial cancer, ependymoma, esophageal cancer, esthesioneuroblastoma, fallopian tube carcinoma, fibrous histiocytoma, Ewing sarcoma, eye cancer, germ cell tumor, gallbladder cancer, gastric cancer, gastrointestinal carcinoid tumor, gastrointestinal stromal tumor, 101
gestational trophoblastic disease, glioma, head and neck cancer, hepatocellular cancer, histiocytosis, Hodgkin lymphoma, hypopharyngeal cancer, intraocular melanoma, islet cell tumor, Kaposi sarcoma, kidney cancer, Langerhans cell histiocytosis, laryngeal cancer, lip and oral cavity cancer, liver cancer, lobular carcinoma in situ, lung cancer, macroglobulinemia, malignant fibrous histiocytoma, melanoma, Merkel cell carcinoma, mesothelioma, metastatic squamous neck cancer with occult primary, midline tract carcinoma involving NUT gene, mouth cancer, multiple endocrine neoplasia syndrome, multiple myeloma, mycosis fungoides, myelodysplastic syndrome, myelodysplastic/myeloproliferative neoplasm, nasal cavity and par nasal sinus cancer, nasopharyngeal cancer, neuroblastoma, non-small cell lung cancer (NSCLC), oropharyngeal cancer, osteosarcoma, ovarian cancer, pancreatic cancer, papillomatosis, paraganglioma, parathyroid cancer, penile cancer, pharyngeal cancer, pheochromocytomas, pituitary tumor, pleuropulmonary blastoma, primary central nervous system lymphoma, primary peritoneal carcinoma, prostate cancer, rectal cancer, renal cell cancer, renal pelvis and ureter cancer, retinoblastoma, rhabdoid tumor, salivary gland cancer, Sezary syndrome, skin cancer, small cell lung cancer, small intestine cancer, soft tissue sarcoma, spinal cord tumor, stomach cancer, T-cell lymphoma, teratoid tumor, testicular cancer, throat cancer, thymoma and thymic carcinoma, thyroid cancer, urethral cancer, uterine cancer, vaginal cancer, vulvar cancer, and Wilms tumor. [000373] In some embodiments, the disease to be treated with the IFNα polypeptides and conjugates provided herein is melanoma, gastric cancer, colorectal cancer, renal cell carcinoma, cervical cancer, non-small cell lung carcinoma, ovarian cancer, uterine cancer, fallopian tube carcinoma, primary peritoneal carcinoma, uterine corpus carcinoma, endometrial carcinoma, prostate cancer, and breast cancer. In certain embodiments, the disease to be treated is breast cancer. In certain embodiments, the disease to be treated is melanoma. [000374] In certain embodiments, a IFNα polypeptide or IFNα conjugates described herein or a pharmaceutical composition thereof treats the disease or condition, for example, cancer, by activating anti-tumor immunity. In certain embodiments, a IFNα polypeptide or IFNα conjugate described herein or a pharmaceutical composition thereof treats the disease or condition, for example, cancer, by inducing or enhancing anti-tumor immune memory. [000375] In certain embodiments, the disease or condition is a viral infection, for example hepatitis B (HBV) or hepatitis C (HCV). In one embodiment, the viral infection is hepatitis C, including drug resistant and multidrug resistant forms of HCV or HBV and related disease states, conditions, or complications of an HCV or HBV infection, including cirrhosis and related hepatotoxicities, In one embodiment, the HBV or HCV is chronic. 102
1.9. Combination Therapy [000376] In certain embodiments, a INFα polypeptide or conjugate as described herein is administered with a second active agent, for example, for the treatment of cancer. In certain embodiments, the second active agent is an immune checkpoint inhibitor, including but not limited to, a PD-1 inhibitor, a PD-L1 inhibitor, a PD-L2 inhibitor, a CTLA-4 inhibitor, or a LAG-3 inhibitor. [000377] In certain embodiments, the immune checkpoint inhibitor is a PD-1 inhibitor. In certain embodiments, the immune checkpoint inhibitor is a PD-1 immune checkpoint inhibitor selected from, but not limited to, nivolumab (Opdivo), pembrolizumab (Keytruda), and cemiplimab (Libtayo). In certain embodiments, the PD-1 immune checkpoint inhibitor is pembrolizumab (Keytruda). [000378] In certain embodiments, the immune checkpoint inhibitor is a PD-L1 inhibitor. In certain embodiments, the immune checkpoint inhibitor is a PD-1 immune checkpoint inhibitor selected from, but not limited to, atezolizumab (Texentriq), durvalumab (Imfinzi), and Avelumab (Bavencio). [000379] In certain embodiments, the immune checkpoint inhibitor is a CTLA-4 immune checkpoint inhibitor. In certain embodiments, the immune checkpoint inhibitor is a PD-1 immune checkpoint inhibitor selected from, but not limited to, ipilimumab (Yervoy). [000380] In certain embodiments, the immune checkpoint inhibitor is a LAG-3 immune checkpoint inhibitor, for example, Relatlimab. [000381] In certain embodiments, a INFα polypeptide or conjugate as described herein is administered with an immune checkpoint inhibitor selected from a PD-1 inhibitor, a PD-L1 inhibitor, a PD-L2 inhibitor, a CTLA-4 inhibitor, or a LAG-3 inhibitor for the treatment of melanoma. In one embodiment, the immune checkpoint inhibitor is a PD-1 inhibitor, for example, pembrolizumab (Keytruda). [000382] In certain embodiments, a IFNα polypeptide or conjugate as described herein is administered in combination with a second active agent for the treatment of hepatitis B or hepatitis C, including, but not limited to ribavirin. In certain embodiments, a IFNα polypeptide or conjugate as described herein is administered in combination ribavirin for the treatment of hepatitis C. [000383] Additional second active agents that can be administered in combination with a IFNα polypeptide or conjugate as described herein for the treatment of hepatitis C include, but are not limited to, a protease inhibitor (such as telaprevir (Incivek), boceprevir (Victrelis), and 103
simeprevir (Olysio); a NS5A inhibitor (such as daclatasvir (Daklinza) and velpatasvir (Epclusa); a NS5B inhibitor (such as dasabuvir (Exviera) and sofosbuvir (Sovaldi)); or a combination drug (such as Harvoni (ledipasvir/sofosbuvir), Viekira Pak (ombitasvir/paritaprevir/ritonavir/dasabuvir), Viekirax (ombitasvir/paritaprevir/ritonavir), Mavyret (paritaprevir and glecaprevir), Technivie (ombitasvir/paritaprevir/ritonavir), Epclusa (sofosbuvir/velpatasvir) and Zepatier (elbasvir and grazoprevir)). [000384] Additional second active agents that can be administered in combination with a IFNα polypeptide or conjugate as described herein for the treatment of hepatitis B include, but are not limited to, a nucleoside reverse transcriptase inhibitor, for example, Epivir (Lamivudine), Hepsera (Adefovir dipivoxil), Baraclude (Entecavir), Tyzeka (Telbivudine), Viread (Tenofovir), Vemlidy (tenofovir alfenamide), and Levovir (Cledvudine). 1.10. Diagnostic Applications [000385] In some embodiments, the IFNα polypeptides or conjugates provided herein are used in diagnostic applications. For example, an IFNα polypeptide or conjugate disclosed herein that is specific for a given receptor may be useful in assays for the given receptor. In some aspects, the IFNα polypeptide or conjugate can be used to detect the expression of the given receptor in various cells and tissues. These assays may be useful, for example, diagnosing cancer, infection and autoimmune disease. [000386] In the methods, the formation of a complex between the IFNα polypeptide or conjugate and receptor can be detected by any method known to those of skill in the art. Examples include assays that use secondary reagents for detection, ELISA’s and immunoprecipitation and agglutination assays. A detailed description of these assays is, for example, given in Harlow and Lane, Antibodies: A Laboratory Manual (Cold Spring Harbor Laboratory, New York 1988555-612, WO 96/13590 to Maertens and Stuyver, Zrein et al. (1998) and WO 96/29605. [000387] For in situ diagnosis, the IFNα polypeptide or conjugate may be administered to a subject by methods known in the art such as, for example, intravenous, intranasal, intraperitoneal, intracerebral, intraarterial injection such that a specific binding between the IFNα polypeptide or conjugate and receptor may occur. The IFNα polypeptide or conjugate/receptor complex may conveniently be detected through a label attached to the IFNα polypeptide or conjugate or any other art-known method of detection. 104
[000388] In some diagnostic applications, the IFNα polypeptide or conjugate may be labeled with a detectable moiety. Suitable detectable moieties include, but are not limited to radioisotopes, fluorescent labels, and enzyme-substrate labels. 1.11. Kits [000389] In some embodiments, an IFNα polypeptide or conjugate as described herein can be provided in a kit, i.e., a packaged combination of reagents in predetermined amounts with instructions for performing a procedure. In some embodiments, the procedure is a diagnostic assay. In other embodiments, the procedure is a therapeutic procedure. EXAMPLES EXAMPLE 1 INTERFERON ALPHA PARA-AZIDOMETHYLPHENYLALANINE (PAMF) SCAN Expression of IFNα-His6 Variants [000390] IFNα variants were expressed in a cell-free protein synthesis reaction as described in Zawada et al. Biotechnol. Bioeng., 2011, 108:1570-1578. Briefly, cell-free extracts were added to a premix containing cell-free reaction components (Groff et al., mAbs, 2014, 6:671- 678) and 10ug/mL plasmid DNA template. Cell free reactions, at a final volume of 1 mL, were incubated at 22° C for 16 h on a shaker at 650 rpm in 48-well plates. [000391] The expressed variants were purified from clarified cell-free reactions via immobilized metal ion affinity chromatography (IMAC). Purified IFNα-His6 variants were quantified via high throughput capillary electrophoresis using the LabChip GXII® (Perkin Elmer) against an IFNα standard curve, according to the manufacturer's instructions. Thermal Stability of pAMF substituted variants by Differential Scanning Fluorimetry (DSF) [000392] Thermal stability of selected IFNα variants was determined by differential scanning fluorimetry (DSF) as previously described in He et al. J Pharm Sci, 2010, 99:1707-1720. Briefly, in a CFX384 (Bio-Rad Laboratories), variants were heated in the presence of Sypro Orange protein stain (Millipore). Fluorescence was monitored and Tm was determined by the minimum of the negative derivative. The thermal stability of selected single-site IFNα variants is provided in Table 4. 105
[000393] Table 4. Thermal stability of selected single-site IFNa variants
Kinetic Analysis of Selected IFNα Variants [000394] Kinetic binding experiments were performed at 25 °C in 1×HBS-EP+ buffer on a Biacore T200 instrument. Anti-human Fc antibody was immobilized onto a CM5 chip using amine coupling chemistry. Human IFNAR1 or human IFNAR2 C-terminally fused to human Fc (hIFNAR1-Fc, hIFNAR2-Fc, respectively) were captured on an anti-human Fc surface. IFNα variants were bound to IFNAR1 at concentrations from 80 nM to 10 uM and allowed to dissociate. The data was fit using Biacore T200 Evaluation Software using a steady state 106
binding model. IFNα variants were bound to IFNAR2 at concentrations from 0.8 nM to 100 nM. The data was fit with the Biacore T200 Evaluation software using a 1:1 Langmuir binding model. The binding affinity of selected single-site IFNα variants to hIFNAR1 and hIFNAR2 are provided in Table 5. [000395] Table 5. Binding affinity of selected single-site IFNα variants to hIFNAR1 and hIFNAR2
107
HEK-blue IFNα/β Reporter Assay for human IFNα [000396] HEK-Blue IFNα/β Reporter Cells (Invivogen, Cat# hkb-ifnab) were maintained in complete DMEM/F-12 Media (Corning) with 100IU Penicillin/100μg/mL Streptomycin (Corning), 2mM GlutaMax (Gibco), 10% h.i. FBS (Sigma), 100μg/mL Normocin (Invivogen), and HEK-Blue Selection antibiotics mix (Invivogen). On assay day, cells were harvested with Accutase, counted, and resuspend at 0.5 x 106 cells/mL in HEK-Blue Detection media (Invivogen). Cells (25 μL) were seeded per well in 384-well clear-bottom plate. Cells were treated with 25 μL of serial dilution of IFNα samples (1:6 serial dilution of 1000 nM starting concentration) and then incubated at 37 °C, 5% CO2 for 16 hours. The plates were then read on a SpectraMax M5 plate reader for absorbance 640 nM. Untreated cells in control wells were used to subtract background absorbance from treated wells. Data was fitted with non-linear regression analysis, using log (agonist) vs. response, variable slope, 4-parameter fit equation using GraphPad Prism. Data was expressed as % relative signal vs. dose of IFNα samples in nM. The results are provided in Table 6 and Table 7. [000397] IFNα samples in Table 6 were conjugated with LP5 or PEG (2x20 kD) or LP9 (DBCO-PEG4-amine), a residual linker remaining after the PEG mask is released through protease cleavage. The potency of LP5-conjugated IFNα variants was attenuated compared to unconjugated IFNα variants and an IFNα control without pAMF incorporation. By comparison, LP9-conjugated IFNα variants had in most cases similar potency as un-conjugated IFNα variants. The structure of LP5 and LP9 are provided below and the conjugation was performed as described herein. [000398] IFNα samples in Table 7 were conjugated with LP6 (20kD PEG) or LP11, a residual linker remaining after the PEG mask is released through protease cleavage. The potency of LP6-conjugated IFNα variants was attenuated compared to unconjugated IFNα variants and an IFNα control without pAMF incorporation. By comparison, LP11-conjugated IFNα variants had in most cases similar potency as un-conjugated IFNα variants. The structure of LP6 and LP11 are provided below and the conjugation was performed as described herein. 108
LP5 LP6, LP9, and LP11. [000399] Table 6. Activity of IFNα variants in HEK-blue IFNα/β Reporter Assay HEK blue EC50 (nM)
09
110
[000401] Table 7. Activity of IFNα variants in HEK-blue IFNα/β Reporter Assay
PEGylation and catB release of DBCO-valcit-pAB-mPEG from selected IFNα variants [000402] Single non-natural amino acid containing IFNα variants were PEGylated by reacting a 3:1 molar excess of DBCO-valcit-pAB-mPEG at 1 mg/mL IFNα in 10 mM citrate, 9% sucrose, pH 6.0, for 16 hours at room temp. To understand the dependence of PEG cleavage on the PEG-linker-length and site of conjugation, cathepsin B forced release experiments were performed directly on the PEGylation reaction. Recombinant human cathepsin B (catB, 953- CY-010, R&D Systems) was pre-activated with 5 mM DTT in 10 mM MES pH 5.0 at room temperature for 15 min. In vitro catB PEG release assays were performed with 20 uM conjugated IFNa and 240 nM activated catB in 50 mM sodium phosphate, pH 6.0 at 40 °C for 16 hours. Table 8 shows the %PEG release calculated by SDS-PAGE gel densitometry of IFNα variants conjugated to DBCO-valcit-pAB-PEGs with varying linker lengths. The structures of LP2, LP6, LP7, and LP8 are provided below. 111
112
[000403] Table 8. Cathepsin B PEG release efficiency of IFNα variants conjugated to DBCO-valcit-pAB-PEGs with varying linker lengths
EXAMPLE 2 CELL-FREE EXPRESSION OF INTERFERON ALPHA VARIANTS IN STIRRED TANK [000404] Cell free protein synthesis reactions were carried using the XpressCF+® system as described previously (Zawada, J. F. et al. Biotechnol. Bioeng.108, 1570–1578 (2011)). Briefly, cell free reactions were prepared by the addition of 37.5% v/v S30 extract, 3 ug/mL plasmid encoding IFN variants and a supermix containing amino-acids, NMPs and small molecules for energy generation (Cai, Q. et al. Biotechnol. Prog.31, 823–831 (2015)). Four macromolecular reagents were individually over-expressed in E. coli and added to the XpressCF+® reaction as reagent lysates at <1% v/v each: T7 RNA polymerase, E. coli peptide deformylase, and the orthogonal tRNA synthetase / tRNA pair from M. jannaschii which have been engineered for incorporation of the modified amino acid para-azidomethyl-L-phenylalanine (pAMF) at the TAG amber codon (Zimmerman, E. S. et al. Bioconjug. Chem.25, 351–361 (2014)). Large scale reactions were carried out in a DASbox stirred tank (Eppendorf) at 250 mL volume with pH, DO and temperature control. Reactions were run with a temperature of 25 °C, pH was controlled at 7.3 using 1 M citrate and 1 M KOH and DO was maintained at 20%. 113
EXAMPLE 3 PURIFICATION OF INTERFERON ALPHA VARIANTS Clarification and IMAC affinity capture [000405] The XpressCF+® expression of IFN variants with His SUMO tag were clarified by centrifugation at 10,000 rpm for 20 minutes (Beckman, JLA-10.500 rotor) and filtered through a 0.22-μm membrane filter. The clarified material is loaded onto a HisTrap Excel affinity column equilibrated with 15 mM Tris-acetate, 500 mM NaCl, 1mM DTT, pH 7.5. After 20 column volumes was applied to wash unbound impurities, the bound proteins were eluted with 20mM Tris-acetate, 300mM imidazole, 1mM DTT, pH 7.5. The eluted fractions were analyzed by 4-12% SDS-PAGE gel electrophoresis and protein concentrations were determined by measured absorbance at 280 nm. Removal of His SUMO tag and anion exchange purification [000406] Ulp1 protease was mixed with the purified protein and incubated at room temperature for 1 hour. The digested reaction was analyzed by 4-12% SDS-PAGE to verify full cleavage of the His SUMO tag prior to HiPrep Desalting column with Sephadex G-25 resin for rapid buffer exchange into 20 mM Tris-acetate, 150mM NaCl, 1mM DTT, pH 7.5. IMAC affinity polish and buffer exchange [000407] For the final purification step, the desalted IFN variants were applied to a HisTrap Exel affinity column equilibrated with 15 mM Tris-acetate, 150mM NaCl, 1mM DTT pH 7.5 as a flow through chromatography process. The target IFNα variants were eluted from the column without adsorption whereas the remaining contaminants were strongly bound. A 15 column wash with 15mM Tris-acetate, 150mM NaCl, 1mM DTT pH 7.5 was applied and the collected flow through and wash fractions were pooled. An Amicon Ultra-15, 3kD centrifugal filter was used to concentrate and buffer exchange the IFN variants into PBS buffer, 6% sucrose, 1mM DTT, pH 7.2. 114
EXAMPLE 4 SYNTHESIS OF MPEG-DBCO LINKERS Dibenzocyclooctyne-amine (LP10): [000408] Dibenzocyclooctyne-amine was obtained from Click Chemistry Tools (CAS 1255942-06-3; Cat #A103-25). DBCO-(PEG)4-NH2 (LP9): [000409] DBCO-(PEG)4-NH2 (LP9) was obtained from Click Chemistry Tools (CAS 1255942-08-5; Cat # A103P). 115
DBCO-Fmoc-mPEG (20 kDa) (LP1): [000410] DBCO-Fmoc-mPEG (20 kDa) (LP1) was synthesized as shown below in Scheme 1. Scheme 1: [000411] Step 1: In a 1000 mL flask equipped with a magnetic stir bar, mPEG-NH2 (20,000 kDa) (23.98 g, 1.2 mmol) in anhydrous toluene (250 mL) was added. The mixture was azeotropically dried under reduced pressure at 45 °C on a rotary evaporator, lyophilized overnight, and then dissolved in anhydrous DCM (200 mL). A solution of 9-(hydroxymethyl)- 9H-fluorene-2-carboxylic acid (compound 3) (0.72 g, 2.99 mmol) and HOBt (0.61 g, 4.5 mmol) was dissolved in anhydrous DMF (15 mL) and added to the mPEG-NH2 (20 kDa) solution. Thereafter, DCC (0.93 g, 4.5 mmol) was added. The reaction was stirred at room temperature for one day under N2 atmosphere. Reaction progress was monitored by analytical HPLC-ELSD (column: Jupiter C4: LC column 250 mm × 4.6 mm × 5µm (Vendor: Phenomenex, part # 00G167-E0), mobile Phase: acetonitrile and water with 0.1% TFA (90% water to 10% water in 50 min, flow rate, 1 mL/minute). Thereafter, HPLC showed completion of the reaction, solvents were removed under reduced pressure, and the crude PEG product was added to isopropanol (600 mL) with gentle heating (35 °C). To this was added 200 mL MTBE and then the mixture was cooled to 10 °C. Solids were filtered and washed with cold IPA (100 mL) and MTBA (50 mL). Crude PEGylated amide 4 as an off-white solid was dried under vacuum. The PEGylated amide 4 was dissolved in DCM (300 mL), added to a Capto® S resin (prewashed 116
with water; 1 L), and stirred overnight. After filtration, the solvent was entirely removed. MALDI-TOF, 1NMR, and HPLC-ELSD confirmed the desired Compound 4 in good purity. [000412] Step 2: To an oven-dried 250 mL flask equipped with a magnetic stir bar, Fmoc PEGylated amide compound 4 (11.4 g, 0.52 mmol) (azeotropically dried with 100 mL toluene removed at 50° C under vacuum prior to use) and anhydrous DCM (70 mL) were added. The clear solution was flushed with argon and then triphosgene (231.9 mg, 0.78 mmol) and pyridine (0.06 mL, 0.73 mmol) were added sequentially. The reaction mixture was stirred at room temperature for two hours under nitrogen. DCM and pyridine were removed under reduced pressure. The chloroformate intermediate (not shown) was dissolved in 50 mL of DCM, and DBCO amine (432 mg, 1.56 mmol) was added in one portion. The reaction was stirred at room temperature for two hours under an N2 atmosphere. Solvent was removed to dryness and the solids were dissolved in 15 mL of DCM and precipitated via IPA (500 mL). The precipitation was repeated twice. Solids were filtered and dried under vacuum. Compound LP1 was confirmed by 1H NMR (CDCl3), MALDI-TOF, SDS-PAGE, and analytical ELSD-HPLC. 117
mPEG (20kDa)-valcit-pAB-(PEG)4-DBCO (LP2): [000413] mPEG (20 kDa)-valcit-pAB-(PEG)4-DBCO (LP2) was synthesized as shown below in Scheme 2. Scheme 2: [000414] Step 1: To a 100 mL round bottom flask equipped with magnetic stir bar, was added [4-[[(2S)-2-[[(2S)-2-(9H-fluoren-9-ylmethoxycarbonylamino)-3-methyl-butanoyl]amino]-5- ureido-pentanoyl]amino]phenyl]methyl (4-nitrophenyl) carbonate (compound 1) (0.7 g, 1.3 mmol), DBCO-(PEG)4-amine compound 4 (909 mg, 2.98 mmol), and anhydrous DMF (15 mL). The clear solution was flushed with argon and then DIPEA (0.3 mL) was added. The reaction was stirred at room temperature for 4 hrs. under N2 atm. LC-MS showed completion of the reaction. Solvent was removed to dryness and the crude material was precipitated and washed with water and MTBE to afford the Fmoc protected intermediate (800 mg) (not shown). This intermediate (800 mg) was dissolved in DCM (10 mL), DMF (2 mL), and piperidine (0.5 mL). The reaction mixture was stirred at ambient temperature for 1 hour under N2 atm. LC-MS showed completion of the reaction. Volatiles were removed under reduced pressure. The crude free amine was purified by preparative reverse phase-high performance liquid chromatography 118
using an Ultro 120 (7 µm) C18Q, 150x20 mmID column. Solvent system used; Solvent A: water containing 10 mm NH4OAc; Solvent B: acetonitrile containing 10 mm NH4OAc; Gradient mode from 10% Solvent B to 90% solvent B, over 20 minutes, 10 mL/min), pure fractions were collected and lyophilized to give compound 5500 mg as a white solid. LC-MS (ESI) m/z 929.5 (M+H). [000415] A mixture of mPEG (20KDa)-SC (Layson Bio) (19.0 g, 1 eq) and NH2-Val-cit- pAB-(PEG)4-DBCO compound 5 (1.26 g, 1.3 eq) was dissolved in a mixture of freshly distilled DCM (80 mL) and DMF (10 mL). DIPEA (486 µL, 3 eq) was added and reaction mixture was stirred at room temperature for 18 h and the reaction was monitored by HPLC. iPrOH was added, the resulting precipitate was centrifuged, and washed 2 times with iPrOH and 2 times with MTBE. The product was dried in vacuum pump for 24 h to give 18.1 g (92%) of LP2 as a white solid. MALDI-Tof and 1HNMR & HPLC-ELSD data showed the desired product in good purity. mPEG (30K)-Valcitp-pAB-(PEG)4-DBCO (LP3): [000416] mPEG (30 kDa)-val-citp-pAB-(PEG)4-DBCO (LP3) was synthesized as described above using commercially available m-PEG-succinimidyl carbonate (30 kDa). MALDI-Tof and 1HNMR & HPLC-ELSD data showed the desired product in good purity. DBCO PEG linkers (LP4 and LP5) 119
[000417] DBCO-C6-NHS ester (in about 10% excess) and mPEG-amine was dissolved in anhydrous DCM and the reaction was stirred for 24 hours while monitoring the consumption of DBCO-C6-NHS ester. The compounds were purified by repeated crystallization from MTBE until no DBCO-C6-NHS ester was detected by HPLC. DBCO PEG compounds LP4 and LP5 were confirmed by 1H NMR (CDCl3), MALDI-TOF, and analytical ELSD-HPLC. EXAMPLE 5 INTERFERON ALPHA PEGYLATION [000418] A stock solution of a conjugate selected from LP1-LP11 (5 mM) was mixed with a final concentration of 1-50 mg/mL protein incorporated with pAMF in 1xPBS at DBCO to pAMF ratio of 2-50. The mixture was incubated at 22-35 oC for 4-24 hr. The PEG density was measured by 4-12% Bis-tris SDS-PAGE gel. EXAMPLE 6 PURIFICATION OF INTERFERON ALPHA PEG CONJUGATES [000419] The reaction consisting of conjugated IFNα and unreacted PEG was further processed by a cation exchange column packed with Capto SP resin (Cytiva). Dilution of the IFNa/PEG reaction prior to purification was performed with binding buffer (10 mM citric acid, pH 4.5) and bound to the Capto SP column with a 4 minute residence time during the load. A linear gradient with elution buffer (10 mM citric acid, 500 mM NaCl, pH 4.5) was performed over 8 CV and the target elution fractions were collected and buffer exchanged into 10 mM acetic acid or PBS, 9% sucrose, pH 4.5 by Amicon Ultra-15, 10kD. SDS-PAGE 4-12% [000420] Analysis of the unconjugated and conjugated IFNα variants by SDS-PAGE showed a single protein band at the correct molecular weight. 120
Analytical HPLC SEC [000421] Monomer percentage and presence of impurities were checked by HPLC-SEC, performed with Ultimate 3000 system and Sepax Zenix-C SEC-150 (7.8 x 300 mm) for the unconjugated IFNα variants, while the PEGylated IFNα were analyzed with Sepax SRT SEC- 300. The unconjugated and PEGylated IFNα proteins eluted as a single peak on the analytical size-exclusion chromatogram with a reported monomer content percentage of >90%. PEG density analysis using gel densitometry [000422] Gel densitometry analysis was used to estimate PEG density.1-4 ug of PEGylated IFNα was loaded on 4-12% Bis-tris SDS-PAGE (NuPAGE™ Invitrogen). The gel ran in 1x NuPAGE™ MES SDS Running Buffer (Invitrogen) with constant voltage at 400 volts for 35 minutes. The gel image was scanned using Bio-Rad Gel DOC EZ Imager and exported for densitometry analysis using ImageQuant TL 7.0 (GE Health). The PEGylated IFNα migrated slower than unpegylated IFNα. EXAMPLE 7 IN VITRO STUDIES OF INTERFERON ALPHA PAMF MULTI-SITE VARIANTS Thermal Stability [000423] IFNα variants with two or more pAMF site substitutions were expressed as described in Example 2. The thermal stability of select variants was determined as described in Example 1 and the results are provided in Table 9. [000424] Table 9. Thermal stability of selected IFNα variants with multi-pAMF sites
121
Kinetic Analysis of Selected IFNα Variants [000425] The binding affinity of selected multi-site IFNα variants wherein the variants are unconjugated or conjugated to PEG are provided in Table 10. The binding affinity was determined as described in Example 1. [000426] Table 10. Binding affinity of selected multi-site IFNα variants to hIFNAR2
HEK-blue IFNα/β Reporter Assay for human IFNα [000427] The in vitro potency of selected multi-site IFNα variants in the HEK-blue IFNα/β reporter assay was determined as described in Example 1. The potencies of the IFNα variants conjugated to different PEGs (LP1, LP2, LP3) were significantly attenuated compared to an unaltered IFNα variant, while IFNα variants conjugated to LP9 or LP10 only showed slightly reduction in potency. The results are provided in Table 11A and Table 11B. [000428] Table 11A: Activity of multi-site IFNα variants in the HEK-blue IFNα/β Reporter Assay Conjugate HEk-blue EC50
122
[000429] Selected multi-site IFNα variants were also conjugated to LP1, LP2 and LP4 and in vitro activity of the conjugates were evaluated in the HEK-blue IFNα/β reporter assay. The potencies of Conjugate 12, Conjugate 11 and Conjugate 31 were significantly attenuated compared to an unaltered IFNα variant. Conjugate 16 and Conjugate 18 showed similar activity as Conjugate 1, which is moderately reduced. Conjugate 2 showed only slightly reduction in activity compared to unconjugated IFNa. The results are provided in Table 11B and the dose response curves are shown in FIG.5A. [000430] Table 11B: Activity of multi-site IFNα variants in the HEK-blue IFNα/β Reporter Assay
B16-Blue IFNα/β Reporter Assay for mouse IFNα 124
[000431] B16-Blue IFNα/β Reporter Cells (Invivogen, Cat# hkb-ifnab) were maintained in complete DMEM/F-12 Media (Corning) with 100IU Penicillin/100μg/mL Streptomycin (Corning), 2mM GlutaMax (Gibco), 10% h.i. FBS (Sigma), 100 μg/mL Normocin (Invivogen), and HEK-Blue selection antibiotics mix (Invivogen). On assay day, cells were harvested with Accutase, counted and resuspend at 0.5 x 106 cells/mL 25μL of cells were seeded per well in 384-well clear-bottom plate. Cells were treated with 25 uL of serial dilution of mIFNα samples (1:6 serial dilution of 1000nM starting concentration). After incubation at 37°C, 5% CO2 for 16 hours, 5 μL of induced B16-Blue™ IFN-α/β cells supernatant was taken out and put into a new 384-well clear-bottom plate.180 μl of resuspended QUANTI-Blue solution per well were then added and the plates were incubated at 37 °C incubator for 1-5 h. The plates were then read on a SpectraMax M5 plate reader for absorbance 640 nM. Cell-only control wells were used to subtract background absorbance from treated wells. Data was fitted with non-linear regression analysis, using log (agonist) vs. response, variable slope, 4-parameter fit equation using GraphPad Prism. Data was expressed as % relative signal vs. dose of mIFNα samples in nM. The results for select multi-site IFNα variants are provided in Table 12. EXAMPLE 8 PK OF HLE-INTERFERON VARIANTS IN C57Bl/6 MICE [000432] In-vivo plasma exposure of different Interferon variants was tested in naïve C57Bl/6 mice in a single dose study. Conjugate 1, Conjugate 13, Conjugate 18 and Conjugate 33 were dosed IV at 3 mg/kg dose. Plasma samples were collected at different timepoints to assess levels of total hIFNa2b throughout the course of 7 days. FIG. 6 illustrates plasma concentrations of hIFNa2b from each test article. At equivalent dose of 3 mg/kg, Conjugate 33 had the fastest plasma clearance and was not detected 4 hours post dosing. In contrast Conjugate 1, Conjugate 13 and Conjugate 18 had a greater half-life and were detectable throughout the 7-day period. Conjugate 1 and Conjugate 13 had similar half-life extension, whereas Conjugate 18 had a faster clearance (FIG.6). EXAMPLE 9 IN VIVO ACTIVITY COMPARISON OF HLE-INTERFERON VARIANTS IN XENOGRAFT TUMOR MODEL [000433] FIG. 1A and FIG. 1B illustrate the effects of Conjugate 1, Conjugate 16 and Conjugate 18 on MDA-MB-231 tumor growth up until the end of the study at day 41 post 125
treatment. The effect of a 3 mg/kg dose is shown in FIG.1A and the effect of a 10 mg/kg dose is shown in FIG.1B. Analysis of tumor sizes was done on day 41, when the mean of vehicle- treated tumors reached the study endpoint (>1,500 mm3). At equivalent doses of 3 mg/kg (based on interferon equivalents), Conjugate 1 induced greater tumor growth suppression (47% TGI) compared to Conjugate 18 (34% TGI) and Conjugate 16 (36% TGI) (FIG. 1A). At equivalent doses of 10 mg/kg, Conjugate 1, Conjugate 18, and Conjugate 16 showed similar tumor growth suppression (51% TGI and 51% TGI and 45% TGI, respectively) (FIG.1B). [000434] FIGS.1C, 1D, and 1E illustrate the effects of Conjugate 1, Conjugate 13, Conjugate 11, and Conjugate 31 on MDA-MB-231 tumor growth up until the end of the study at day 44 post treatment. FIG. 1C and FIG. 1D show the effect of a 3 mg/kg and 15 mg/kg dose, respectively. FIG.1E is a graph showing the tumor size on day 44. Analysis of tumor sizes was done on day 44, when the mean of vehicle-treated tumors reached >1,200 mm3. Overall Conjugate 11, Conjugate 13, and Conjugate 31 exhibited trends of dose-dependent anti-tumor activity. At equivalent doses of 3 mg/kg, Conjugate 1 induced greater tumor growth suppression (50% TGI) compared to Conjugate 11 (10% TGI), Conjugate 13 (37% TGI) (FIG. 1C). At equivalent doses of 15 mg/kg, Conjugate 11 and Conjugate 13 induced similar activity (38% TGI and 46% TGI, respectively) (FIG.1D). Conjugate 31, which has a non-releasable PEG, did not demonstrate significant effect when dosed at 3 mg/kg and 15 mg/kg (0% and 12% TGI) (FIG.1E) compared to other IFNα-variants, indicating that cleavage of PEG is essential to confer anti-tumor activity. EXAMPLE 10 ANTI-TUMOR ACTIVITY COMPARISON OF HLE-INTERFERON VARIANTS IN HUMANIZED MOUSE XENOGRAFT MODEL [000435] NSG mice were engrafted intraperitoneally with 5x106 human PBMCs and implanted with 5x106 MDA-MB-231 tumor cells in the mammary fat pad. Mice were randomized and enrolled into the study 7 days post implant, with tumor sizes around 100 mm3 - 150 mm3. Tumor-bearing mice from both studies were administered three weekly doses (qwx3) of the test articles at doses ranging from 0.05 mg/kg to 3 mg/kg. All treatments were well tolerated with normal body weight gain throughout the course of the study. [000436] FIGS. 2A-2E summarize results illustrating dose-dependent effects of different IFNα variants on growth of MDA-MB-231 tumors up in a mouse model engrafted with human PBMCs. At equivalent doses of 3 mg/kg, all test articles demonstrated prolonged tumor stasis 126
with evidence of regrowth after day 21 for Conjugate 13, Conjugate 18, and Conjugate 2 and day 31 for Conjugate 1. Analysis of tumors on day 32 of study revealed that at the 3 mg/kg dose, Conjugate 1 induced mildly greater tumor growth suppression (104% TGI) compared to Conjugate 18 (91% TGI), Conjugate 13 (83% TGI) and Conjugate 2 (82% TGI) (FIG.2A). At equivalent doses of 1 mg/kg, activity of Conjugate 1 and Conjugate 18 was the greatest and nearly comparable (95% and 92% TGI, respectively) followed by Conjugate 13 (70% TGI) and Conjugate 2 (51% TGI) (FIG.2B). Treatment with Conjugate 1 demonstrated potent and dose-dependent tumor growth suppression at 0.1 mg/kg and 0.05 mg/kg doses (81% TGI at 0.1 mg/kg and 60% TGI at 0.05 mg/kg) (FIGS. 2C and 2D). Conjugate 18 demonstrated comparable anti-tumor activity at 0.1 mg/kg and 0.05 mg/kg dose (59% and 52% TGI, respectively) (FIG.2C and 2D). A similar trend was seen with Conjugate 13, inducing similar effect at 0.1 mg/kg and 0.05 mg/kg dose (32% and 40% TGI, respectively) (FIG.2C and 2D). Activity of Conjugate 2 was the least, with 23% TGI at 0.1 mg/kg dose (FIG. 2C and 2E). Overall, Conjugate 1 demonstrated the greatest TGI amongst all IFNα variants tested at equivalent doses (FIG.2E). EXAMPLE 11 TOLERABILITY OF HLE-INTERFERON VARIANTS IN GOLDEN SYRIAN HAMSTERS [000437] In vivo tolerability of HLE-Interferon variants was tested in naive Golden Syrian hamsters in a single dose study. Briefly Conjugate 18, Conjugate 13, Conjugate 2, and Conjugate 1 were dosed IV at doses ranging from 3 mg/kg to 45 mg/kg. Animal body weights were monitored regularly for 7 days throughout the course of the study. Serum samples were collected at different timepoints to assess plasma levels of each test article. Additionally on Day 2 and Day 7 of study, serum samples were analyzed for liver enzymes that are indicators of hepatocyte injury. [000438] FIG. 3A illustrates serum concentrations of each test article. At all tested doses, Conjugate 13, Conjugate 18, Conjugate 1 was detectable until Day 7 post treatment. Treatment with Conjugate 18 and Conjugate 13 followed linear dose-dependent pharmacokinetics (FIG. 3A). At equivalent dose of 15 mg/kg, Conjugate 13 and Conjugate 1 showed similar PK profile and had greater half-life extension compared to Conjugate 18 and Conjugate 2 (FIG. 3B). Overall Conjugate 2 had the fastest clearance, with total interferon levels undetectable in 2 out of 3 animals by day 3 of study (FIG.3B). 127
[000439] As shown in FIG.3C, hamsters treated with Conjugate 18 showed dose dependent body weight loss (-7% at 3 mg/kg on Day 7, -10.4% loss at 15 mg/kg on Day 7, -20.6% at 45mg/kg on Day 6). Overall, a single dose of 45 mg/kg of Conjugate 18 was not tolerated (FIG. 3C). Similarly a single dose of 15 mg/kg Conjugate 1 also induced 12.4% reduction in hamster body weight on Day 7 post treatment (FIG.3D). Conversely, a single dose of Conjugate 13 did not result in any hamster body weight loss up to a dose of 45 mg/kg (FIG.3C). An equivalent dose, corresponding to 15 mg/kg of the cytokine, of Conjugate 1 induced maximum body weight loss (12.4%), followed by Conjugate 18 (10.4%), Conjugate 2 (3.4%) and Conjugate 13 (1%) (FIG.3D). [000440] FIGS.3E-J illustrate effects of all test articles on induction of liver enzymes AST, ALT, and ALP compared to the vehicle-treated serum on 2 days and 7 days of the study. Analysis of serum revealed treatment with Conjugate 18 at all doses resulted in early induction of ALT (8.5-20 fold) on day 2 of the study that returned to baseline levels by day 7 (FIG.3E and 3H). Treatment with Conjugate 18 at all doses caused induction of AST by 4-7 fold on study day 2 that returned to baseline levels by day 7 for lower doses (FIG.3F and 3I). However, on study day 7, 45 mg/kg of Conjugate 18 treatment significantly induced serum levels of both AST & ALP (~15 fold and ~20 fold respectively) (FIG.3I and 3J). Treatment with Conjugate 13 resulted in initial induction of ALT in a dose-dependent manner (~3-8 fold) and returned to baseline at end of study on day 7 (FIG.3E and 3H). Conjugate 13 did not induce significant changes in AST levels on day 2. However, a 45 mg/kg dose of Conjugate 13 resulted in ~5 fold increase in AST levels on day 7 of study (FIG.3I). Treatment with 15 mg/kg of Conjugate 1 resulted in a time-dependent increase in levels of AST (4 fold on day 2, 9 fold on day 7) and ALP (1.5 fold on day 2, 4 fold on day 7) (FIG.3F, 3G, 3I, and 3J). Similar to Conjugate 13 and Conjugate 18, Conjugate 1 also caused initial induction of ALT on day 2 (~10.5 fold) that returned to baseline by day 7 (FIG.3E and 3H). Unlike the effect with Conjugate 1, Conjugate 13 and Conjugate 18, 15 mg/kg Conjugate 2 did not significantly alter liver enzyme levels at both day 2 and day 7 of the study (FIG.3E-3J). [000441] FIGS.3K-3L demonstrate the effect of all test articles on platelet and reticulocyte counts. Analysis of whole blood was done on Day 7 post treatment. Analysis revealed that treatment with Conjugate 18 resulted in greater than 50% reduction in reticulocyte count at the lowest dose 3 mg/kg. At doses greater than 3 mg/kg, Conjugate 18 showed dose-dependent decrease in reticulocyte count (92% reduction at 15 mg/kg and 99% reduction at 45 mg/kg dose) (FIG.3K). Treatment with Conjugate 1 resulted in ~97% decrease in reticulocyte count. Treatment with Conjugate 13 showed evidence of dose-dependent loss in reticulocyte count 128
(~30% reduction at 15 mg/kg and ~50% reduction at 45 mg/kg dose) (FIG.3K). In contrast, Conjugate 2 did not alter reticulocyte count on Day 7 (FIG.3K). FIG.3L illustrates the effect of all IFNa variants on platelet count on Day 7. Treatment with Conjugate 18 showed reduction in platelet counts at all doses (40-55% reduction). Treatment with Conjugate 13 and Conjugate 2 did not result in platelet loss. Conjugate 1 at 15 mg/kg induced ~60% reduction in platelet count (FIG.3L). EXAMPLE 12 IN VITRO ACTIVITY OF MOUSE SURROGATE [000442] Mouse IFNa molecules with pAMF incorporated at the same sites as in human IFNa were made in order to study the efficacy of the IFNa variants in mouse models with intact immune system. [000443] The in vitro activity of mouse IFNa molecules were evaluated using a B16-Blue IFNα/β Reporter assay. In brief, B16-Blue IFNα/β Reporter Cells (Invivogen, Cat# hkb-ifnab) were maintained in complete DMEM/F-12 Media (Corning) with 100IU Penicillin/100μg/mL Streptomycin (Corning), 2mM GlutaMax (Gibco), 10% FBS (Sigma), 100 μg/mL Normocin (Invivogen), and HEK-Blue selection antibiotics mix (Invivogen). On assay day, cells were harvested with Accutase, counted and resuspend at 0.5 x 106 cells/mL 25μL of cells were seeded per well in 384-well clear-bottom plate. Cells were treated with 25 uL of serial dilution of mIFNα samples (1:6 serial dilution of 1000nM starting concentration). After incubation at 37°C, 5% CO2 for 16 hours, 5 μL of induced B16-Blue™ IFN-α/β cells supernatant was taken out and put into a new 384-well clear-bottom plate. 180 μl of resuspended QUANTI-Blue solution per well were then added and the plates were incubated at 37 °C incubator for 1-5 h. The plates were then read on a SpectraMax M5 plate reader for absorbance 640 nM. Cell-only control wells were used to subtract background absorbance from treated wells. Data was fitted with non-linear regression analysis, using log (agonist) vs. response, variable slope, 4- parameter fit equation using GraphPad Prism. Data was expressed as % relative signal vs. dose of mIFNα samples in nM. The results for select multi-site IFNα variants are provided in Table 12. The dose response curves are shown in Fig 1B. 129
[000444] Table 12. Activity of multi-site mouse IFNα variants in the B16-Blue IFNα/β Reporter Assay
EXAMPLE 13 MOUSE SURROGATE EFFICACY AND PD IN SYNGENEIC MOUSE TUMOR MODEL MC38-HCEA [000445] The anti-tumor activity of different mouse HLE-IFNα variants was additionally examined in a model system with intact innate and adaptive immune compartments using syngeneic mouse tumor model. Briefly, C57/Bl6 mice were implanted subcutaneously with B16F10 cells and enrolled into the study with tumor size around 130 mm3. Tumor-bearing mice were administered 3 weekly (qwx3) doses of mouse surrogates Conjugate 36, Conjugate 37, Conjugate 34 and Conjugate 35 at doses ranging from 1 mg/kg to 10 mg/kg. All treatments were well tolerated throughout the course of the study. [000446] FIG.4A-4G illustrates the effects of different test articles on B16F10 tumor growth. Analysis of tumor sizes was done on study day 10, when the mean of vehicle-treated tumors reached the study endpoint. Overall treatment with Conjugate 36 resulted in the most potent anti-tumor activity compared to all other test articles (FIG. 4D). At equivalent doses of 3 mg/kg, Conjugate 36 showed significantly greater TGI (90%) followed by Conjugate 37 (73%), Conjugate 34 (65%) and Conjugate 35 (25%) (FIG.4A). At the 1 mg/kg dose, Conjugate 36 induced greater tumor growth suppression compared to Conjugate 34 (79% and 44% 130
respectively) (FIG. 4B). At equivalent doses of both 3 mg/kg and 10 mg/kg, Conjugate 37 showed significantly greater TGI compared to Conjugate 35: 73% and 25% TGI, respectively at 3 mg/kg; 81% and 35% TGI respectively, at 10 mg/kg, indicating that cleavage of PEG is essential to confer anti-tumor activity (FIG.4C). [000447] Anti-tumor activity of Conjugate 37 was also evaluated in MC38 cells expressing a human tumor associated antigen. Briefly, C57/Bl6 mice were implanted subcutaneously with 1x106 MC38-hCEA cells and enrolled into the study with tumor size around 150 mm3. Tumor bearing mice were administered 3 weekly (qwx3) doses of Conjugate 37 at doses ranging from 0.1 mg/kg to 3 mg/kg. All treatments were well tolerated throughout the course of the study. FIG.4H illustrates that treatment with Conjugate 37 elicits potent dose-dependent anti-tumor activity and results in high complete response rates in this model. Analysis of tumor sizes done on study day 17 revealed 48% TGI at 0.1 mg/kg, 96% TGI at 1 mg/kg. At the end of the study, 12.5% mice in the 1 mg/kg dose group and 100% mice in the 3 mg/kg dose group were tumor free. Tumor free mice from Conjugate 37 treatment demonstrated no recurrence of tumors when rechallenged with 5x106 MC38-hCEA cells (FIG.4I) suggesting formation of treatment- induced anti-tumor immune memory. [000448] The effect of Conjugate 37 on the tumor microenvironment was evaluated in the syngeneic mouse tumor model. Briefly, MC38-hCEA tumor bearing mice were administered a single dose of vehicle or 3 mg/kg of Conjugate 37 IV. Tumors were harvested 3 days post treatment and analyzed for infiltration of cytotoxic immune cells and levels of cytolytic enzymes in the TME. FIGS.4E-4G illustrates the effect of Conjugate 37 on immune activation in TME. Analysis revealed that Conjugate 37 treatment increased percentage of CD8-T cells and - Granzyme B levels - in tumor infiltrating CD8 T-cells and NK cells, compared to vehicle- treated tumors (FIGS.4E-4G). EXAMPLE 14 COMBINATION OF HLE-INTERFERON VARIANT WITH CHECKPOINT INHIBITOR IN B16F10 SYNGENEIC MOUSE TUMOR MODEL [000449] The anti-tumor activity of HLE-Interferon variant in combination with mouse anti- PD-1 antibody was examined in the B16F10 syngeneic mouse tumor model. Briefly, C57/Bl6 mice were implanted subcutaneously with 1x106 B16F10 cells and enrolled into the study with tumor size around 70 mm3. Tumor bearing mice were administered 3 weekly (qwx3) doses of Conjugate 37 at 3 mg/kg as a single-agent or in combination with 10 mg/kg anti-PD-1. Mouse 131
anti-PD-1 antibody, clone RMP1-14, was dosed every 4 days for 3 weeks. All treatments were well tolerated throughout the course of the study. FIG. 7A illustrates single-agent and combination anti-tumor activity of Conjugate 37 and anti-PD-1 on B16F10 tumor growth. Analysis of tumors on study day 10 when the vehicle group tumor volume reached ~1,000 mm3 revealed anti-PD-1 treatment alone did not induce anti-tumor activity (FIG.7A). Analysis of tumors on study day 24 revealed single agent treatment with Conjugate 37 revealed potent anti- tumor activity. Conjugate 37 combined with anti-PD-1 exhibited trends of greater anti-tumor activity compared to treatment with Conjugate 37 alone (FIG. 7A and FIG. 7B) suggesting reversal of anti-PD-1 resistance with Conjugate 37 in this model. FIG.7C demonstrates mice treated with combination of Conjugate 37 and anti-PD-1 exhibit longer time to reach tumor volume of 500 mm3 compared to mice treated with Conjugate 37 alone. EXAMPLE 15 PRODUCTION OF INTERFERON ALPHA CONTAINING 3 PAMF AMINO ACIDS IN E. COLI CELLS WITH HIGH DENSITY FERMENTATION [000450] Interferon alpha (IFNα) production was demonstrated in an E. coli strain engineered with an oxidizing cytoplasm (E. coli Snuggle). In order to achieve high levels of Amber suppression sufficient for the incorporation of 3 non-natural amino acids (NNAAs) into a single protein, the genes involved in NNAA incorporation were expressed on a plasmid (RS plasmid), while genes for expression of the protein of interest were expressed on a second higher copy plasmid (product plasmid). [000451] To generate the RS plasmid, a gene having the coding sequence (CDS) for an aminoacyl tRNA synthetase (RS) specific for para-azidomethylphenylalanine (pAMF) was cloned into a medium copy (p15a origin) plasmid with a carbenicillin (Carb) selection cassette behind a constitutive promoter followed by an inducible T7 promoter (T7p) and a strong ribosome binding site (RBS). Three copies of a tRNA specific for para- azidomethylphenylalanine (pAMF) were cloned behind the pAMF RS CDS, with 23 nucleotide non-coding DNA spacers before each tRNA sequence. [000452] To generate the product plasmid, the coding sequence for human interferon alpha (IFNα) with an N-terminal HisSUMO tag and TAG codons at the positions coding for amino acids 40, 51, and N156 (HisSUMO-IFNα Q40/E51/N156 TAG) was codon optimized for E. coli. The construct was cloned behind a T7p and strong RBS into a high copy (pUC origin) plasmid with a kanamycin (Kan) selection cassette. An additional copy of the pAMF tRNA 132
was cloned behind the HisSUMO-IFNα Q40/E51/N156 TAG coding sequence on the product plasmid. [000453] The E. coli strain for expression of IFNα was generated by transforming the E. coli Snuggle strain with both the RS plasmid and product plasmid. Transformations were plated on LB agar containing 50 μg/mL kanamycin and 100 μg/mL carbenicillin. Single colonies were picked and transferred into culture tubes with 3 mL of TB media containing 50 μg/mL kanamycin and 100 μg/mL carbenicillin for overnight growth at 37° C. The culture tube was used to inoculate a shake flask with I17-SF shake flask media containing 50 μg/mL of kanamycin and 100 μg/mL of carbenicillin at 8% (v/v) seeding density. The shake flask was harvested once the culture achieved an OD 595 nm greater than 3. Glycerol was added to the shake flask to a final concentration of 16-20% (v/v). The cell bank was collected and aliquoted into 2 mL vials, flash frozen in liquid nitrogen and stored at -80°C. [000454] To express the IFNα, the fermentation process began by taking a 2 mL vial of the cell bank and inoculating a shake flask with I17-SF shake flask media (as described in Hanson, J.; Groff, D.; Carlos, A.; Usman, H.; Fong, K.; Yu, A.; Armstrong, S.; Dwyer, A.; Masikat, M.R.; Yuan, D.; et al. An Integrated In Vivo/In Vitro Protein Production Platform for Site- Specific Antibody Drug Conjugates. Bioengineering 2023, 10, 304, which is incorporated herein by reference in its entirety) containing 50 μg/mL of kanamycin and 100 μg/mL of carbenicillin at about 8% (v/v) seeding density. Once an OD 595 nm of 3-4 was reached, the shake flask culture was used to inoculate a 1 L bioreactor at a seeding density of 6% (v/v) in batched media. The batched media consisted of 2.4% (v/v) 5x I17 Media in DI H2O, 50 μg/mL of kanamycin, 100 μg/mL of carbenicillin, and 0.1% (v/v) A204 antifoam. The bioreactor temperature, dissolved oxygen and pH setpoints at inoculation were 37° C, 30% and 7, respectively. [000455] Once the cells grew to an OD 595 nm between 3-5 during the batch phase, the fed batch phase began by feeding 5x I17 media at an exponential rate of 0.15 h-1. After 21 hours in the fed batch phase, the temperature decreased to 25° C, and the exponential feed rate decreased to 0.02 h-1. [000456] One hour later, the induction phase began by adding pAMF to a target concentration of 4 mM and L-Arabinose to a target concentration of 4 g/L based on the culture volume in the bioreactor prior to induction. The induction phase took 24 hours before the bioreactor was harvested. At the end of the fermentation, the culture was collected and centrifuged at 18,592 xG and 2-8° C for 15 min in a floor centrifuge. The supernatant was discarded, and the cell pellets were resuspended with DPBS at a concentration of 16.67% (w/w). The cell resuspension 133
was then passed twice through an Avestin Homogenizer (EmulsiFlex-C5) at 17,000 Psi to disrupt the cells and generate the crude lysate. The crude lysate was clarified by centrifuging at 18,000-20,000 xG and 2-8° C for 30 minutes in a floor centrifuge. The supernatant (clarified lysate) was collected and aliquoted, flash frozen in liquid nitrogen and stored at -80°C. [000457] Lysate supernatants were applied to Ni-NTA resin that had been pre-equilibrated with PBS. After application of the supernatant, the resin was washed with PBS containing 10 mM imidazole before the protein was eluted across several fractions with PBS containing 200 mM imidazole. The purest fractions were identified by analysis via SDS-PAGE then pooled and concentrated in 10 kDa MWCO Amicon centrifuge filters. Samples were quantified by adjusting the absorbance at 280 nm according to the calculated molar absorbance of the protein and considering the % purity calculated by gel densitometry analysis from an SDS-PAGE gel. [000458] The presence of full-length protein was verified using intact LC-MS analysis. Protein samples were digested with Ulp1 (1:20 w/w ratio) for 1 hour at 22oC, after which 10- 15 pmol of each protein sample was injected onto a reverse phase column via the autosampler of an Agilent 1200 series HPLC. A gradient method starting at 10% A (0.1% formic acid in H2O) and increasing to 95% B (0.1% formic acid in acetonitrile) over 10 minutes was used to separate proteins prior to introduction into the MS source. After separation, proteins were analyzed using an Agilent 6520 QTOF mass spectrometer operating in positive ion MS (Seg) mode in a mass range of 500-3200 m/z with a scan rate of 1 spectra per second. Data analysis was performed in Agilent MassHunter Qualitative analysis software. Proteins were identified by the existence of corresponding peaks in the total ion and UV (280, 254, and 214 nM) chromatograms. The averaged mass spectrum of the entire IFNα chromatogram peak was used for protein identification. It was deconvoluted using the MassHunter BioConfirm Protein Deconvolution function using the Maximum Entropy algorithm with a mass range of 10,000- 60,000 Daltons and a 1.00 Dalton mass step. Peak identities were confirmed by comparing the deconvoluted peak mass to the predicted protein mass that had been calculated using gpmaw3. Peaks were integrated, and the integrated peak areas were used to calculate the percent truncation at position N156 according to the following formula:
EXAMPLE 16 PRODUCTION OF INTERFERON ALPHA CONTAINING 3 PAMF AMINO ACIDS IN E. COLI CELLS WITH HIGH DENSITY FERMENTATION [000459] To generate the product plasmid, each product gene was synthesized and then cloned into pJ411. This vector has a kanamycin resistance marker and a pUC high copy origin of replication, and the expression cassette has a T7 promoter for high level transcription. Plasmid sequence was verified by sequencing. [000460] For the incorporation of nnAAs into proteins of interest, the selected codons in product genes where nnAAs would be incorporated were substituted with the amber codon “TAG.” The gene for 3XnnAA-IFNα was mutated to contain 3 TAG codons at the positions coding for amino acids 40, 51, and N156 as described in Example 15. The coding sequence for the pAMF RS was cloned into a medium copy pJ434 plasmid behind a constitutive Pc0 promoter. (Groff, D.; Armstrong, S.; Rivers, P.J.; Zhang, J.; Yang, J.; Green, E.; Rozzelle, J.; Liang, S.; Kittle, J.D.; Steiner, A.R.; et al. Engineering toward a Bacterial “Endoplasmic Reticulum” for the Rapid Expression of Immunoglobulin Proteins. MAbs 2014, 6, 671–678, incorporated herein by reference in its entirety). Three copies of the amber suppressor tRNA (AS tRNA) were included on the pJ434 plasmid 3’ to the pAMF RS coding sequence after a 20 base pair spacer sequence. [000461] All plasmids were transformed into E. coli SBDG419 (Hanson, J.; Groff, D.; Carlos, A.; Usman, H.; Fong, K.; Yu, A.; Armstrong, S.; Dwyer, A.; Masikat, M.R.; Yuan, D.; et al. An Integrated In Vivo/In Vitro Protein Production Platform for Site-Specific Antibody Drug Conjugates. Bioengineering 2023, 10, 304, incorporated herein by reference in its entirety) and plated on selective media. For wild type proteins, only the high copy product plasmid was transformed. For nnAA-containing proteins, each product plasmid was co-transformed with an RS plasmid. [000462] For initial screening experiments, protein expressions were performed in shake flasks. For each experiment, a single colony was picked from transformation plates into 2-3 mL of Terrific Broth (Teknova, Hollister, CA) containing appropriate antibiotic(s). After overnight incubation at 37°C in a shaking incubator at 250 RPM, cultures were inoculated into 135
50 mL of fresh Terrific Broth + antibiotics in a shake flask. Cultures were incubated at 37°C in a shaking incubator at 250 RPM until the optical density at 595 nm (OD595) reached 1.0-2.0. At this time, arabinose was added to induce expression of the protein(s) of interest. For cultures expressing nnAA proteins, pAMF was added at 2 mM. Cultures were transferred to 25°C for expression. After expression for 16-18 hours, cells were harvested by centrifugation for 5 minutes at ~7000g. Cells were resuspended in 10 mL per gram of wet cells in phosphate buffered saline (PBS) containing 0.1 mg/mL lysozyme and benzonase. After incubation on ice for 30 minutes, cells were lysed by sonication. Soluble lysates were isolated by centrifugation at >20,000g for 30 minutes. [000463] High-density fermentations for subsequent analysis and characterization were then performed according to the process described in Example 15. [000464] For purification of HisSUMO-IFNα and HisSUMO-3XnnAA-IFNα, lysates were centrifuged at 30,000g for 30 minutes, and the supernatants were applied to Cytiva Ni Sepharose excel resin that had been pre-equilibrated with PBS. After application of the supernatant, the resin was washed with PBS containing 10 mM imidazole before the protein was eluted across several fractions with PBS containing 400 mM imidazole. Protein was digested with Ulp1 (1:20 w/w ratio) for 1 hour. [000465] After complete Ulp1 digestion, proteins were polished with two additional column steps. Proteins were first buffer exchanged into 20 mM Tris, 300 mM sodium chloride, pH 7.5 with Cytiva Sephadex G-25 fine resin. Then they were applied back onto Cytiva Ni Sepharose excel resin and the flowthrough contained the target protein. The final pool was concentrated and buffer exchanged into PBS, 9% sucrose, pH 6 with Amicon centrifuge filters (10 kDa MWCO) for conjugation. [000466] For the PEG conjugation step, the conjugation reaction was diluted with water, adjusted to pH 5 using 1 M acetic acid, and applied to Cytiva Capto SP ImpRes resin. Mobile phase A consisted of 10 mM citric acid, pH 5 and mobile phase B consisted of 10 mM citric acid, 300 mM sodium chloride, pH 5. Protein was eluted using a linear gradient from 0% to 100% mobile phase B. Targeted fractions were collected and buffer exchanged into PBS, 9% sucrose, pH 6 with Amicon centrifuge filters (10 kDa MWCO) for further testing. [000467] For analytical SEC analysis, product quality was assessed by injection of 30-50 μg of protein onto an analytical SEC column (Zenix-C SEC-300, 3 μM, 7.8 × 300 mm, Sepax Technologies) with a mobile phase of 50 mM sodium phosphate, 140 mM NaCl, 5% isopropanol, pH 6.5. 136
[000468] For LC-MS analysis, samples (10-50 pmol) were injected into an Agilent 1200 series system with a Binary SL pump. Mobile phases A was 0.1% formic acid in water and mobile phase B was 0.1% formic acid in acetonitrile. Proteins were separated using an Agilent PLRP-S HPLC column (2.1 x 50 mm, 1000 Å, 5 micrometers) heated at 80°C in Agilent column oven with flow rate of 0.3 mL/min. The gradient started with 10% mobile phase B and was kept at 10% B for the first 2 min. and was then ramped up to 70% B in 8 min. Mobile phase B was then increased to 95% by 11 min. and was held for 3 min. before it was decreased to 10% by 14.5 min. Data was acquired on an Agilent 6520A Accurate Mass Q-TOF MS with mass detection range of 500 – 3200 m/z. The source gas temperature was at 325°C, drying gas flow at 8 L/min, nebulizer at 30 psig. Capillary voltage was set at 4000 V, fragmentor voltage at 250 V and skimmer 1 was set at 65 V. The instrument mode used was 2 GHz, Standard (3200 m/z). [000469] Spectra from all peaks on the total ion chromatogram were extracted and deconvoluted using Maximum Entropy algorithm from MassHunter Qualitative (B6.00) from Agilent. A mass range of 10,000-100,000 Da was searched. Adduct use in the deconvolution setting was proton. Peaks were filtered by setting a signal-to-noise ratio of > 30.0. Top 90% of the peak height was used to calculate average mass. [000470] For conjugation to DBCO amine, protein concentrations were brought to 1 mg/mL in DPBS. The DBCO-amine was added at a drug to pAMF ratio of 3:1, and 500 mM NaCl was added to the reaction to improve DBCO-amine solubility. The conjugation reaction was incubated overnight at 30°C prior to LC-MS analysis. [000471] For conjugation to DBCO-PEG, proteins were dialyzed into 1x DPBS + 9% Sucrose prior to conjugation. The PEG of interest was prepared in water as a 5mM stock solution. Targeting the final protein concentration at 1mg/ml, the protein was formulated in DPBS buffer, and 3 molar equivalents of PEG were added per mole of pAMF. Conjugation reactions were incubated at 25°C (3XnnAA-IFNα) overnight in a Thermomixer (Fisher scientific, Allentown Pennsylvania) with agitation at 450rpm.. [000472] After conjugate cleanup, PEGylated proteins were analyzed via SDS-PAGE. PEG- to-protein ratios were calculated by gel densitometry analysis using the Lane and Bands image analysis tools in the Image Lab software (version 5.2.1, Bio-Rad). [000473] For analysis of in vitro cell activation, IFNa/b HEK-Blue cells were obtained from Invivogen (catalog number hkb-ifnab) and were plated in 384-well plates at a density of 12,500 cells/well in HEK-Blue detection media (Invivogen, catalog number hb-det2). The cells were treated with various concentrations of test articles and incubated at 37°C with 5% CO2 137
overnight. Reporter activity was determined via absorbance readings at 640nm in a M5 Spectramax (Molecular Devices). Data was plotted as % cell activation vs. protein concentration using GraphPad Prism, with error bars indicating standard deviation of replicates. [000474] Interferon α (IFNα) is a cytokine protein that has successfully been used for the treatment of ulcerative melanoma and renal cell carcinoma and has shown promise as a treatment for leukemias and solid tumors. Conditional activation of cytokines such as IFNα through PEGylation can reduce toxicity and extend drug half-life while maintaining anti-tumor efficacy. This strategy requires the attachment of multiple, releasable PEGs that initially mask receptor binding in a prodrug form. This required expression of a variant of IFNα containing several nnAAs, unlike the other E. coli nnAA proteins produced in this paper with a single nnAA. Incorporation of multiple nnAAs into a single protein chain is known to reduce the amount of full length protein produced, but this effect can be overcome by increasing the cellular level of AS tRNA which helps overcome translational termination at TAG codons. [000475] A construct was designed that would result in the incorporation of pAMF at 3 solvent-exposed sites on IFNα. Because the variant contained several nnAA sites, the product plasmid was co-transformed into E. coli SBDG419 with an RS plasmid containing 3 copies of the pAMF tRNA in order to increase intracellular AS tRNA levels and amber suppression efficiency (FIGS.10A-10B). Initial test expressions revealed that the nnAA-containing IFNα variant (3XnnAA-IFNα) expressed at levels comparable to the wild-type protein (FIG.10C). However, analysis of the unlabeled protein revealed significant truncation (18%) at the C- terminal pAMF site (FIGS. 11A-11C). Therefore, an additional copy of the AS tRNA was cloned at the 3’ end of the IFNα coding sequence of the high copy product plasmid (FIG.10B). The 3XnnAA-IFNα protein that had been expressed in shake flasks using this construct was purified and analyzed the sample via LC-MS, which revealed that the addition of tRNA to the product plasmid improve amber suppression efficiency and significantly reduced the truncation observed at the C-terminal pAMF site (FIG.11C). As a final confirmation of the incorporation of three pAMF residues into the protein, a conjugation reaction with a small molecule DBCO- amine was performed. Analysis by intact LC-MS showed that >96% of each sample had been labeled with three DBCO-amine molecules (FIGS. 11A-11C), confirming the successful incorporation of three nnAAs. [000476] After optimization of its expression in shake flasks, a fermentation process for production of the 3XnnAA-IFNα protein at scale was tested and optimized. Initially strain and fermentation conditions were tested at 250 mL scale in a Dasbox bioreactor. Initial 138
experiments, however, led to inconsistent feeding rates due to pump flow rate constraints in the smaller scale fermenters. Therefore, the feed rate was changed to a target of 0.15 μ during the pre-induction phase of the fed-batch program. [000477] After cell harvest and lysis, 3XnnAA-IFNα lysates were analyzed for protein production via SDS-PAGE, which showed the presence of a band that ran at the same position as a 3XnnAA-IFNα standard (FIG.9A). Protein was purified from the soluble lysate from this initial fermentation, and the Ulp1-cleaved protein was analyzed via LC-MS for identity confirmation. The deconvoluted spectra predominantly consisted of a main peak, with the presence of 1% truncated product (FIGS.11A-11C). In fed batch fermentation, the 3XnnAA- IFNα was produced at a titer of approximately 540 mg/L (FIG.9D). [000478] Fed-batch fermentation of this strain was scaled to 500 mL to produce sufficient material for downstream processing and activity analysis. Growth rates, OD595 and product titers compared well with previous fermentations. The 3XnnAA-IFNα protein was expressed at titers of 600 mg/L as calculated by purification of the protein over IMAC resin (FIG.9D). After Ulp1 cleavage, the protein was then conjugated with a releasable DBCO-PEG molecule. Analysis of the conjugated sample by SDS-PAGE showed the reaction had gone to >90% completion. The conjugate was cleaned up using a cation exchange chromatography step which resulted in material with a final purity of 93.6% (FIG.9D) and a final PEG-to-protein ratio of 2.97 (FIG.9D). [000479] The finalized unconjugated 3XnnAA-IFNα and PEGylated material were then analyzed for in vitro activity. The 3XnnAA-IFNα protein retained similar in vitro cell activation activity to a wild type IFNα standard (FIG. 9C). The PEGylated 3XnnAA-IFNα showed no cell activation activity, demonstrating effective activity attenuation (expected activity of a prodrug) through site-specific PEGylation at multiple sites. The successful activity attenuation of 3XnnAA-IFNα through site specific PEGylation suggests that the E. coli SBDG419 production strain could feasibly be used to generate mutants of other therapeutic proteins containing multiple conjugatable pAMF handles. EXAMPLE 17 EFFECT OF HLE-INTERFERON VARIANTS ON IMMUNE MODULATION IN THE TUMOR MICROENVIRONMENT [000480] The effect of HLE-Interferon variants on immune modulation in the tumor microenvironment was evaluated in the syngeneic mouse tumor model. Briefly, MC38-hTrop2 139
tumor bearing mice were administered IV a single dose of vehicle or 1 mg/kg of Conjugate 37 or 1 mg/kg of Conjugate 34. Tumors and lymph nodes were harvested 3 days post treatment and analyzed for activation of innate and adaptive immune cells. FIGS.12A-F illustrate the effect of Conjugate 37 and Conjugate 34 on different cell types in the TME. Analysis revealed both HLE-Interferon variants increased Granzyme B levels in tumor infiltrating CD8 T-cells and NK cells compared to vehicle-treated tumors (FIGS.12A-12B). Additionally Conjugate 37 and Conjugate 34 treatment increased activation of multiple innate immune cells in TME including monocytes, dendritic cells and plasmacytoid dendritic cells (FIGS. 12C-12E). Lymph node analysis revealed that treatment with Conjugate 34 resulted in a similar increase in levels of GranzymeB in CD8 T-cells from both tumor-draining and non-draining lymph node. In contrast Conjugate 37 treatment resulted in a greater increase in GranzymeB levels in CD8 T-cells from tumor-draining lymph node compared to a non-draining lymph node (FIG 12F). EXAMPLE 18 EVALUATION OF FUNCTIONAL ROLE OF CD8 T-CELLS IN FORMATION OF HLE- INTERFERON VARIANT INDUCED ANTI-TUMOR IMMUNE MEMORY [000481] The functional role of CD8 T-cells in formation of HLE-Interferon variant induced anti-tumor immune memory was evaluated in tumor free (complete responder) mice obtained from Conjugate 37 treatment. Complete responder mice were treated with 300 ^g anti-CD8 antibody or Isotype antibody and rechallenged with 5x106 MC38-hCEA cells on D0. CD8 depletion was conducted prior to rechallenge and maintained throughout the course of the study. Conjugate 37-treated complete responder mice that received Isotype control antibody demonstrated no recurrence of tumors when rechallenged with MC38-hCEA cells. In contrast CD8 depletion ablated Conjugate 37-induced anti-tumor immune memory in complete responder mice as seen by formation of tumors when rechallenged with MC38-hCEA cells (FIG 13). EXAMPLE 19 PK-PD PROFILE IN NON-HUMAN PRIMATES [000482] To understand the potential toxicity and pharmacokinetic (PK)-pharmacodynamic (PD) relationship of HLE-Interferon variants described herein prior to clinical development, HLE-Interferon variants are tested in a non-human primate system, such as cynomolgus 140
monkey. HLE-Interferon variants are dosed to cynomolgus monkeys by slow intravenous infusion and subcutaneous injection. The monkeys are dosed at different dose levels (e.g., 6.75 mg/kg or 20.25 mg/kg) to understand the toxicity and PK-PD relationship. Blood is drawn at various timepoints to monitor drug exposure and proof of mechanism. Ultimately, the HLE- Interferon variants’ PK profile is aligned with observed toxicity and markers of mechanism to generate a better understanding of the drug safety profile. [000483] PK profile of the drugs is evaluated by measuring concentration of total Interferon and PEGylated Interferon. Presence of higher concentrations of non-PEGylated Interferon can contribute to interferon-mediated toxicity. Proof of mechanism and PD profile for HLE- Interferon variants is evaluated by measuring levels of different mechanism of action and inflammatory cytokines in the circulation. Blood clinical chemistry is evaluated to monitor for signs of liver or kidney toxicity. Blood hematology, such as various red and white blood cells, is also monitored to detect hematological toxicities. [000484] Half-life extension and masking efficiency of HLE-Interferon variants can be inferred by the PK-PD profile. Additionally, safety of the HLE-Interferon variants can be partially represented by the drug’s Highest Non-Severely Toxic Dose (HNSTD), or dose at which no or limited toxicities are observed. The higher the HNSTD, the higher probability of the drug safety. EXAMPLE 20 SEQUENCES Table 13 provides sequences referred to herein. [000485] In certain embodiments, SEQ ID NO.2-31 and 33-38 are preceded by a methionine amino acid. In Table 13, the sequence of the indicated HisSUMO fusions (SEQ. ID NO: 39) are not shown for clarity. As described herein, the cleavable HisSUMO fusion facilitates expression and purification. In certain embodiments, provided herein are IFNα polypeptides according to any of SEQ ID NOS. 2-38. In certain embodiments, provided herein are IFNα polypeptides according to any of SEQ ID NOS: 2-38 fused to HisSUMO SEQ ID NO: 39. [000486] Table 13. Sequence table Conjugate No. Name Amino acid sequence
[000487] All patents and patent publications referred to herein are hereby incorporated by reference. Certain modifications and improvements will occur to those skilled in the art upon a reading of the foregoing description. It should be understood that all such modifications and improvements have been deleted herein for the sake of conciseness and readability but are properly within the scope of the following claims. 146
Claims
CLAIMS What is claimed is: 1. An interferon alpha (IFNα) polypeptide comprising at least one non-natural amino acid at a position selected from the group consisting of D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156 relative to SEQ ID NO: 33, wherein the IFNα polypeptide specifically binds to IFN-α receptor (IFNAR).
2. The interferon alpha (IFNα) polypeptide of claim 1, comprising at least one non-natural amino acid at a position selected from the group consisting of H7, Q40, N45, E41, E51, and N156 relative to SEQ ID NO: 33, wherein the IFNα polypeptide specifically binds to IFN-α receptor (IFNAR).
3. The IFNα polypeptide of claim 1, wherein the IFNα polypeptide comprises an amino acid sequence having at least 90% sequence identity to SEQ ID NO: 33.
4. The IFNα polypeptide of claim 1, wherein the polypeptide has at least 95% sequence identity to SEQ ID NO: 33.
5. The IFNα polypeptide of claim 1, wherein the polypeptide has at least 98% sequence identity to SEQ ID NO: 33.
6. The IFNα polypeptide of any one of claims 1-5, comprising at least two, three, or four non-natural amino acids at positions selected from the group consisting of D2, L3, Q5, T6, H7, Q40, E41, E42, N45, Q46, Q48, K49, E51, A75, D77, E78, T79, L80, V103, T106, E107, L110, M111, K131, E132, K133, K134, Y135, S136, and N156.
7. The IFNα polypeptide of any one of claims 1-6, comprising at least two non-natural amino acids at positions selected from the group consisting of H7, Q40, N45, E41, E51, and N156.
8. The IFNα polypeptide of any one of claims 1-6, comprising at least three non-natural amino acids at positions selected from the group consisting of H7, Q40, N45, E41, E51, and N156.
9. The IFNα polypeptide of any one of claims 1-6, comprising at least four non-natural amino acids at positions selected from the group consisting of H7, Q40, N45, E41, E51, and N156.
10. The IFNα polypeptide of any one of claims 1-9, wherein the IFNα polypeptide comprises an amino acid sequence having a non-natural amino acid at position H7.
11. The IFNα polypeptide of any one of claims 1-10, wherein the IFNα polypeptide comprises an amino acid sequence having a non-natural amino acid at position E51. 147
12. The IFNα polypeptide of any one of claims 1-11, wherein the IFNα polypeptide comprises an amino acid sequence having a non-natural amino acid at position Q40.
13. The IFNα polypeptide of any one of claims 1-12, wherein the IFNα polypeptide comprises an amino acid sequence having a non-natural amino acid at position N156.
14. The IFNα polypeptide of any one of claims 1-9, wherein the IFNα polypeptide comprises an amino acid sequence having non-natural amino acids at positions Q40 and N156.
15. The IFNα polypeptide of claim 14, wherein the polypeptide further comprises at least one non-natural amino acid at a position selected from the group consisting of H7 and E51.
16. The IFNα polypeptide of claim 1, wherein the IFNα polypeptide has an amino acid sequence according to SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 5, SEQ ID NO: 6, SEQ ID NO: 7, or SEQ ID NO: 8.
17. The IFNα polypeptide of any one of claims 1-16, wherein the non-natural amino acid comprises a residue of a moiety selected from the group consisting of amino, carboxy, acetyl, hydrazino, hydrazido, semicarbazido, sulfanyl, azido and alkynyl.
18. The IFNα polypeptide of any one of claims 1-16, wherein the non-natural amino acid is selected from the group consisting of p-acetyl-L-phenylalanine, O-methyl-L- tyrosine, 3-methyl-phenylalanine, O-4-allyl-L-tyrosine, 4-propyl-L-tyrosine, fluorinated phenylalanine, isopropyl-L-phenylalanine, p-azido-L-phenylalanine, p- acyl-L-phenylalanine, p-benzoyl-L-phenylalanine, p-iodo-phenylalanine, p- bromophenylalanine, p-amino-L-phenylalanine, isopropyl-L-phenylalanine, p- propargyloxy-phenylalanine, and p-azidomethyl-L-phenylalanine residues.
19. The IFNα polypeptide of any one of claims 1-15, wherein the non-natural amino acid is a p-azidomethyl-L-phenylalanine residue.
20. The IFNα polypeptide of claim 20, wherein the non-natural amino acid at position Q40 is a p-azidomethyl-L-phenylalanine residue.
21. The IFNα polypeptide of claim 20, wherein the non-natural amino acid at position N156 is a p-azidomethyl-L-phenylalanine residue.
22. The IFNα polypeptide of claim 20, wherein the non-natural amino acid at positions Q40 and N156 are p-azidomethyl-L-phenylalanine residues.
23. An interferon alpha (IFNα) conjugate comprising an IFNα polypeptide of any one of the preceding claims, wherein the at least non-natural amino acid residue is linked to a masking moiety optionally via a linker. 148
24. An interferon alpha (IFNα) conjugate comprising: an IFNα polypeptide; and, at least one masking moiety; wherein the IFNα polypeptide is site-specifically linked to the at least one masking moiety with a protease cleavable linker and the masking moiety is a water-soluble polymer or carbohydrate.
25. An interferon alpha (IFNα) conjugate comprising: an IFNα polypeptide; and, at least one masking moiety; wherein the IFNα polypeptide is site-specifically linked to the at least one masking moiety with a cathepsin B cleavable linker.
26. The IFNα conjugate of claim 24 or 25, wherein the IFNα polypeptide is a IFNα polypeptide of any one of claims 1-23.
27. The IFNα conjugate of claim 25 or 26, wherein the masking moiety comprises a water- soluble polymer, peptide, or a carbohydrate.
28. The IFNα conjugate of claim 27, wherein the masking moiety comprises a water- soluble polymer.
29. The IFNα conjugate of claim 28, wherein the water-soluble polymer is a residue of polyethylene glycol (PEG), methoxypolyethylene glycol (mPEG), poly(propylene glycol) (PPG), copolymers of ethylene glycol and propylene glycol, poly(oxyethylated polyol), poly(olefinic alcohol), poly(vinylpyrrolidone), poly(hydroxyalkylmethacrylamide), poly(hydroxyalkylmethacrylate), poly(saccharides), poly(a-hydroxy acid), poly(vinyl alcohol), polyphosphazene, polyoxazolines (POZ), poly(N-acryloylmorpholine), and combinations thereof.
30. The IFNα conjugate of claim 29, wherein the water-soluble polymer is PEG.
31. The IFNα conjugate of claim 29, wherein the water-soluble polymer is mPEG.
32. The IFNα conjugate of claim 30 or 31, wherein the PEG or mPEG has an average molecular weight between about 5KDa and about 50 KDa.
33. The IFNα conjugate of claim 32, wherein the PEG or mPEG has an average molecular weight between about 20KDa and about 40 KDa.
34. The IFNα conjugate of any one of claims 30-33, wherein the PEG or mPEG is selected from the group consisting of a linear or branched PEG molecule or mPEG molecule having an average molecular weight of about 10KDa, about 20KDa, about 30KDa, or about 40KDa. 149
35. The IFNα conjugate of any one of claims 30-34, wherein the PEG or mPEG has an average molecular weight of about 20KDa.
36. The IFNα conjugate of claim 35, wherein the PEG or mPEG has an average molecular weight of about 40KDa.
37. The IFNα conjugate of claim 28, wherein the water-soluble polymer is of the formula wherein n1 is an integer between 1 and 1,000 and R1 is methyl or hydrogen.
38. The IFNα conjugate of claim 37, wherein n1 is an integer between 200 and 800.
39. The IFNα conjugate of claim 38, wherein n1 is an integer between 300 and 700.
40. The IFNα conjugate of any one of claims 23-25, wherein the IFNα conjugate has an amino acid sequence according to SEQ ID NO: 1, SEQ ID NO: 2, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22, or SEQ ID NO: 23, SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 29, SEQ ID NO: 30, SEQ ID NO: 31, SEQ ID NO: 35, SEQ ID NO: 36, or SEQ ID NO: 37.
41. The IFNα conjugate of claim 40, wherein the IFNα conjugate has an amino acid sequence according to SEQ ID NO: 11, SEQ ID NO: 13, SEQ ID NO: 16, or SEQ ID NO: 18.
42. The IFNα conjugate of claim 41, wherein the IFNα conjugate comprises mPEG having an average molecular weight of 20KDa or 40KDa.
43. The IFNα conjugate of any one of claims 23 and 26-42, wherein the at least one non- natural amino acid is conjugated to the water-soluble polymer via a linker.
44. The IFNα conjugate of claim 43, wherein the linker is a protease cleavable linker or a pH-sensitive linker.
45. The IFNα conjugate of claim 43, wherein the protease cleavable linker is a cathepsin B linker.
46. The IFNα conjugate of claim 43, wherein the linker is of the formula (L1): 150
wherein RG is a reactive group residue; W1 and W2 are independently absent or a divalent attaching group; L1 is absent, a protease cleavable linker, or a pH-sensitive linker; SG1 is a divalent spacer group; is a bond to the IFNα polypeptide; and is a bond to the masking moiety.
47. The IFNα conjugate of claim 43, wherein the linker is of the formula (L2): wherein RG is a reactive group residue; W1 and W2 are independently absent or a divalent attaching group; L1 is absent, a protease cleavable linker, or a pH-sensitive linker; SG2 is a trivalent spacer group; is a bond to the IFNα polypeptide; and is a bond to the masking moiety. 151
48. The IFNα conjugate of claim 46 or 47, wherein RG is , , , , , , , , , , or .
49. The IFNα conjugate of claim 48, wherein RG is or .
50. The IFNα conjugate of any one of claims 46-49, wherein W1 is -C(O)-C1- 6alkylene-, -C(O)(C1-6alkylene)NR4-, -C(O)(C1-6alkylene)O-, or -C(O)(C1- 6alkylene)S-; wherein R4 is hydrogen or C1-6alkyl, RG is connected to W1 at -C(O)-, and the C1-6alkylene is optionally substituted with one, two, or three substituents selected from halogen, alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy.
51. The IFNα conjugate of claim 50, wherein W1 is -C(O)-C1-6alkylene- or -C(O)(C1- 6alkylene)NH-.
52. The IFNα conjugate of claim 51, wherein W1 is -C(O)-(CH2CH2)4- or -C(O)- (CH2CH2)2NH-.
53. The IFNα conjugate of any one of claims 46-52, wherein W2 is ; 152
wherein X1 is absent, a divalent water-soluble polymer, -C1-6alkylene-, - NR4(C1-6alkylene)-, or -O(C1-6alkylene)-; X2 is absent or -C1-6alkylene-; X3 is absent, -NR4-, or -O-; R4 is independently hydrogen or C1-6alkyl; and wherein the C1-6alkylene of X1 or X2 is optionally substituted with one, two, or three substituents selected from a halogen, alkyl, haloalkyl, hydroxyl, amino, alkylamino, and alkoxy; and wherein the carbonyl is attached to W1.
54. The IFNα conjugate of claim 53, wherein the divalent water-soluble polymer is wherein R1 is hydrogen or methyl and n2 is an integer between 1 and 50, inclusive.
55. The IFNα conjugate of claim 54, wherein n2 is an integer between 1 and 10.
56. The IFNα conjugate of claim 54, wherein n2 is an integer between 10 and 20.
57. The IFNα conjugate of claim 55, wherein n2 is 4, 5, or 6.
58. The IFNα conjugate of claim 53, wherein X1 is absent.
59. The IFNα conjugate of claim 53, wherein X1, X2, and X3 are absent.
60. The IFNα conjugate of claim 53, wherein W2 is .
61. The IFNα conjugate of claim 53, wherein W2 is .
62. The IFNα conjugate of any one of claims 46-61, wherein L1 comprises a compound of the formula: ; wherein R5 is hydrogen, an electron donating group, or an electron withdrawing group; X4 is -O- or -NR6-; 153
X5 is a linker; R6 is hydrogen or an electron withdrawing group; and m is an integer selected from 1 to 4.
63. The IFNα conjugate of claim 62, wherein X5 includes an ester wherein the carbonyl carbon of the ester functional group is covalently bound to the fluorene.
64. The IFNα conjugate of claim 62 or 63, wherein R5 is hydrogen or halogen.
65. The IFNα conjugate of claim 64, wherein R5 is hydrogen.
66. The IFNα conjugate of claim 62, of the formula .
67. The IFNα conjugate of any one of claims 46-66, wherein L1 comprises a peptide.
68. The IFNα conjugate of any one of claims 46-66, wherein L1 comprises a dipeptide.
69. The IFNα conjugate of claim 68, wherein L1 comprises a dipeptide selected from the group consisting of -Phe-Lys-, -Val-Ala-, -Val-Lys-, -Ala-Lys-, -Val-Cit-, -Phe-Cit-, - Leu-Cit-, -Ile-Cit-, -Phe-Arg-, and -Trp-Cit-.
70. The IFNα conjugate of claim 69, wherein L1 comprises -Val-Cit-.
71. The IFNα conjugate of claim 68, wherein L1 comprises a peptide-self immolative group selected from the group consisting of -Phe-Lys-PABC-, -Val-Ala-PABC-, -Val-Lys- PABC-, -Ala-Lys-PABC-, -Val-Cit-PABC-, -Phe-Cit-PABC-, -Leu-Cit-PABC-, -Ile- Cit-PABC-, -Phe-Arg-PABC-, -Trp-Cit-PABC-, and Val-Glu-PABC.
72. The IFNα conjugate of claim 71, wherein L1 comprises-Val-Cit-PABC-.
73. The IFNα conjugate of any one of claims 46 and 48-72, wherein SG1 is , , , , , , , , , or ; wherein a is an integer selected from 0, 1, 2, 3, 4, 5, and 6; b is an integer selected from 1, 2, 3, 4, 5, and 6; and is a bond to the masking moiety. 154
74. The IFNα conjugate of claim 73, wherein SG1 is selected from , , , , , and ; wherein is a bond to the masking moiety.
75. The IFNα conjugate of any one of claims 47-72, wherein SG2 is or ; wherein a is independently an integer selected from 0, 1, 2, 3, 4, 5, and 6 and is a bond to the masking moiety.
76. The IFNα conjugate of claim 46, of the formula: , , 155
, , , , 156
, , , 157
, or ; wherein COMP is the IFNα polypeptide; and MM is a masking moiety; or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer, thereof.
77. The IFNα conjugate of claim 47, of the formula: wherein COMP is the IFNα polypeptide; and MM is a masking moiety; or a pharmaceutically acceptable salt, solvate, stereoisomer or regioisomer thereof.
78. The IFNα conjugate of any one of claims 46-77, wherein the masking moiety is a water- soluble polymer of the formula wherein n1 is an integer between 1 and 1,000 and R1 is methyl or hydrogen.
79. The IFNα conjugate of claim 78, wherein n1 is an integer between 200 and 800.
80. The IFNα conjugate of claim 78, wherein n1 is an integer between 300 and 700. 158
81. The IFNα conjugate of claim 78, of the formula: , , , 159
, , , 160
, , , 161
, 162
. 163
or a pharmaceutically acceptable salt, solvate, stereoisomer, or regioisomer thereof; wherein n is an integer from 1 to 8.
82. The IFNα conjugate of any one of claims 23-81, wherein the conjugate has an extended half-life compared to an identical conjugate lacking the masking moiety.
83. A nucleic acid encoding the IFNα polypeptide of any one of claims 1-22 or IFNα conjugate of any one of claims 23-82.
84. A pharmaceutical composition comprising an IFNα polypeptide of any one of claims 1-22 or IFNα conjugate of any one of claims 23-82.
85. A method for treating or preventing a disease, disorder, or condition in a subject in need thereof, the method comprising: administering to the subject, a therapeutically effective amount of an IFNα polypeptide of any one of claims 1-22, an IFNα conjugate of any one of claims 23-82, or the pharmaceutical composition of claim 84.
86. The method of claim 85, wherein the disease is cancer.
87. The method of claim 86, wherein the cancer is selected from a cancer of the oral cavity the digestive system the respiratory system, the bones, joints, skin, the breast, the genital system, the urinary system, the eye, the nervous system, the endocrine system and the hematopoietic system.
88. The method of claim 86 or 87, wherein the cancer is a cancer selected from the group consisting of: melanoma, lung cancer, primary mediastinal B-cell lymphoma (PMBCL), a mismatch repair deficient (dMMR) solid tumor, colon cancer, stomach cancer, esophageal cancer, cervical cancer, liver cancer, uterine cancer, cutaneous squamous cell carcinoma (cSCC), and triple-negative breast cancer.
89. The method of any one of claims 86-88, wherein the cancer is breast cancer or melanoma.
90. The method of claim any one of claims 86-88, wherein the cancer is colorectal cancer.
91. The method of claim any one of claims 86-88, wherein the cancer is liver cancer.
92. The method of claim any one of claims 86-88, wherein the cancer is pancreatic cancer.
93. The method of claim any one of claims 86-88, wherein the cancer is a solid tumor cancer.
94. The method of claim any one of claims 85-93, further comprising the administration of one of more additional therapeutic agents selected from an immune checkpoint inhibitor.
95. The method of claim 94, wherein the immune checkpoint inhibitor is a PD-1 inhibitor, a PD-L1 inhibitor, a PD-L2 inhibitor, a CTLA-4 inhibitor, or a LAG-3 inhibitor. 164
96. The method of claim 95, wherein the immune checkpoint inhibitor is a PD-1 inhibitor.
97. The method of any one of claims 85-96, wherein the IFNα polypeptide of any one of claims 1-22, the IFNα conjugate of any one of claims 23-82, or the pharmaceutical composition of claim 84 treats the disease or condition by activating anti-tumor immunity.
98. The method of any one of claims 85-96, wherein the IFNα polypeptide of any one of claims 1-22, the IFNα conjugate of any one of claims 23-82, or the pharmaceutical composition of claim 84 treats the disease or condition by inducing or enhancing anti- tumor immune memory.
99. The method of claim 85, wherein the condition is a viral infection.
100. The method of claim 99, wherein the viral infection is hepatitis C.
101. The method of claim 99 or 100, further comprising the administration of ribavirin.
102. Use of a therapeutically effective amount of an IFNα polypeptide of any one of claims 1-22, IFNα conjugate of any one of claims 23-82, or the pharmaceutical composition of claim 84 in the preparation of medicament for the treatment or prevention of a disease, disorder, or condition.
103. The use of claim 102, wherein the disease is cancer.
104. The use of claim 103, wherein the cancer is selected from a cancer of the oral cavity the digestive system the respiratory system, the bones, joints, skin, the breast, the genital system, the urinary system, the eye, the nervous system, the endocrine system and the hematopoietic system.
105. The use of claim 103 or 104, wherein the cancer is a cancer selected from the group consisting of: melanoma, lung cancer, primary mediastinal B-cell lymphoma (PMBCL), a mismatch repair deficient (dMMR) solid tumor, colon cancer, stomach cancer, esophageal cancer, cervical cancer, liver cancer, uterine cancer, cutaneous squamous cell carcinoma (cSCC), and triple-negative breast cancer.
106. The use of any one of claims 103-105, wherein the cancer is breast cancer or melanoma.
107. The use of claim any one of claims 103-105, wherein the cancer is colorectal cancer.
108. The use of claim any one of claims 103-105, wherein the cancer is liver cancer.
109. The use of claim any one of claims 103-105, wherein the cancer is pancreatic cancer. 165
110. The use of claim any one of claims 103-105, wherein the cancer is a solid tumor cancer.
111. The use of claim any one of claims 102-110, further comprising the administration of one of more additional therapeutic agents selected from an immune checkpoint inhibitor.
112. The use of claim 111, wherein the immune checkpoint inhibitor is a PD-1 inhibitor, a PD-L1 inhibitor, a PD-L2 inhibitor, a CTLA-4 inhibitor, or a LAG-3 inhibitor.
113. The use of claim 112, wherein the immune checkpoint inhibitor is a PD-1 inhibitor.
114. The use of any one of claims 102-113, wherein the IFNα polypeptide of any one of claims 1-22, the IFNα conjugate of any one of claims 23-82, or the pharmaceutical composition of claim 84 treats the disease or condition by activating anti-tumor immunity.
115. The use of any one of claims 102-113, wherein the IFNα polypeptide of any one of claims 1-22, the IFNα conjugate of any one of claims 23-82, or the pharmaceutical composition of claim 84 treats the disease or condition by inducing or enhancing anti- tumor immune memory.
116. The use of claim 115, wherein the condition is a viral infection.
117. The use of claim 116, wherein the viral infection is hepatitis C.
118. The use of claim 115 or 116, further comprising the administration of ribavirin.
119. A method of making an IFNα polypeptide of any one of claims 1-22, or an IFNα conjugate of any one of claims 23-82, the method comprising culturing host cells expressing a coding sequence for the IFNα polypeptide and generating the IFNα polypeptide.
120. The method of claim 119, wherein the E. coli cells have an oxidative cytoplasm. 166
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202263422795P | 2022-11-04 | 2022-11-04 | |
| US63/422,795 | 2022-11-04 | ||
| US202363493439P | 2023-03-31 | 2023-03-31 | |
| US63/493,439 | 2023-03-31 |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| WO2024098023A2 true WO2024098023A2 (en) | 2024-05-10 |
| WO2024098023A3 WO2024098023A3 (en) | 2024-07-25 |
Family
ID=89223897
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2023/078726 Ceased WO2024098023A2 (en) | 2022-11-04 | 2023-11-03 | Interferon alpha polypeptides and conjugates |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2024098023A2 (en) |
Citations (24)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4289872A (en) | 1979-04-06 | 1981-09-15 | Allied Corporation | Macromolecular highly branched homogeneous compound based on lysine units |
| US4560655A (en) | 1982-12-16 | 1985-12-24 | Immunex Corporation | Serum-free cell culture medium and process for making same |
| WO1987000195A1 (en) | 1985-06-28 | 1987-01-15 | Celltech Limited | Animal cell culture |
| US4657866A (en) | 1982-12-21 | 1987-04-14 | Sudhir Kumar | Serum-free, synthetic, completely chemically defined tissue culture media |
| US4767704A (en) | 1983-10-07 | 1988-08-30 | Columbia University In The City Of New York | Protein-free culture medium |
| WO1990003430A1 (en) | 1988-09-23 | 1990-04-05 | Cetus Corporation | Cell culture medium for enhanced cell growth, culture longevity and product expression |
| US4927762A (en) | 1986-04-01 | 1990-05-22 | Cell Enterprises, Inc. | Cell culture medium with antioxidant |
| US5122469A (en) | 1990-10-03 | 1992-06-16 | Genentech, Inc. | Method for culturing Chinese hamster ovary cells to improve production of recombinant proteins |
| US5204244A (en) | 1987-10-27 | 1993-04-20 | Oncogen | Production of chimeric antibodies by homologous recombination |
| US5229490A (en) | 1987-05-06 | 1993-07-20 | The Rockefeller University | Multiple antigen peptide system |
| WO1993021259A1 (en) | 1992-04-14 | 1993-10-28 | Cornell Research Foundation Inc. | Dendritic based macromolecules and method of production |
| WO1996013590A2 (en) | 1994-10-21 | 1996-05-09 | Innogenetics N.V. | New sequences of hepatitis c virus genotypes and their use as prophylactic, therapeutic and diagnostic agents |
| US5534615A (en) | 1994-04-25 | 1996-07-09 | Genentech, Inc. | Cardiac hypertrophy factor and uses therefor |
| WO1996021469A1 (en) | 1995-01-10 | 1996-07-18 | Shearwater Polymers, Inc. | Multi-armed, monofunctional, and hydrolytically stable derivatives of poly(ethylene glycol) and related polymers for modification of surfaces and molecules |
| WO1996029605A2 (en) | 1995-03-14 | 1996-09-26 | Corixa Corporation | Compounds and methods for the dectection of t. cruzi infection |
| US5643575A (en) | 1993-10-27 | 1997-07-01 | Enzon, Inc. | Non-antigenic branched polymer conjugates |
| US20030143596A1 (en) | 2001-11-07 | 2003-07-31 | Shearwater Corporation | Branched polymers and their conjugates |
| WO2008106186A2 (en) | 2007-02-28 | 2008-09-04 | Serina Therapeutics, Inc. | Activated polyoxazolines and compositions comprising the same |
| US8431558B2 (en) | 2004-11-01 | 2013-04-30 | The Regents Of The University Of California | Compositions and methods for modification of biomolecules |
| US20130189287A1 (en) | 2011-12-23 | 2013-07-25 | Paul Scherrer Institut | Enzymatic conjugation of polypeptides |
| US20130251783A1 (en) | 2011-09-14 | 2013-09-26 | Universitat Heidelberg | Liposomes containing permeation enhancers for oral drug delivery |
| US8703936B2 (en) | 2010-02-12 | 2014-04-22 | The Regents Of The University Of California | Compositions and methods for modification of biomolecules |
| US9145361B2 (en) | 2011-03-25 | 2015-09-29 | Life Technologies Corporation | SDP-containing heterobifunctional agents |
| US9222940B2 (en) | 2010-04-27 | 2015-12-29 | Synaffix B.V. | Fused cyclooctyne compounds and their use in metal-free click reactions |
Family Cites Families (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP2005517648A (en) * | 2001-12-07 | 2005-06-16 | インターミューン インコーポレイテッド | Compositions and methods for treating hepatitis virus infections |
| SG135176A1 (en) * | 2004-02-02 | 2007-09-28 | Ambrx Inc | Modified human four helical bundle polypeptides and their uses |
| CN101247821B (en) * | 2005-06-03 | 2013-01-23 | Ambrx公司 | Improved human interferon molecules and uses thereof |
| CN104693300B (en) * | 2013-12-05 | 2021-07-20 | 北京大学 | Modified pegylated recombinant human interferon α2b |
-
2023
- 2023-11-03 WO PCT/US2023/078726 patent/WO2024098023A2/en not_active Ceased
Patent Citations (26)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4289872A (en) | 1979-04-06 | 1981-09-15 | Allied Corporation | Macromolecular highly branched homogeneous compound based on lysine units |
| US4560655A (en) | 1982-12-16 | 1985-12-24 | Immunex Corporation | Serum-free cell culture medium and process for making same |
| US4657866A (en) | 1982-12-21 | 1987-04-14 | Sudhir Kumar | Serum-free, synthetic, completely chemically defined tissue culture media |
| US4767704A (en) | 1983-10-07 | 1988-08-30 | Columbia University In The City Of New York | Protein-free culture medium |
| WO1987000195A1 (en) | 1985-06-28 | 1987-01-15 | Celltech Limited | Animal cell culture |
| US4927762A (en) | 1986-04-01 | 1990-05-22 | Cell Enterprises, Inc. | Cell culture medium with antioxidant |
| US5229490A (en) | 1987-05-06 | 1993-07-20 | The Rockefeller University | Multiple antigen peptide system |
| US5204244A (en) | 1987-10-27 | 1993-04-20 | Oncogen | Production of chimeric antibodies by homologous recombination |
| WO1990003430A1 (en) | 1988-09-23 | 1990-04-05 | Cetus Corporation | Cell culture medium for enhanced cell growth, culture longevity and product expression |
| US5122469A (en) | 1990-10-03 | 1992-06-16 | Genentech, Inc. | Method for culturing Chinese hamster ovary cells to improve production of recombinant proteins |
| WO1993021259A1 (en) | 1992-04-14 | 1993-10-28 | Cornell Research Foundation Inc. | Dendritic based macromolecules and method of production |
| US5643575A (en) | 1993-10-27 | 1997-07-01 | Enzon, Inc. | Non-antigenic branched polymer conjugates |
| US5534615A (en) | 1994-04-25 | 1996-07-09 | Genentech, Inc. | Cardiac hypertrophy factor and uses therefor |
| WO1996013590A2 (en) | 1994-10-21 | 1996-05-09 | Innogenetics N.V. | New sequences of hepatitis c virus genotypes and their use as prophylactic, therapeutic and diagnostic agents |
| WO1996021469A1 (en) | 1995-01-10 | 1996-07-18 | Shearwater Polymers, Inc. | Multi-armed, monofunctional, and hydrolytically stable derivatives of poly(ethylene glycol) and related polymers for modification of surfaces and molecules |
| US5932462A (en) | 1995-01-10 | 1999-08-03 | Shearwater Polymers, Inc. | Multiarmed, monofunctional, polymer for coupling to molecules and surfaces |
| WO1996029605A2 (en) | 1995-03-14 | 1996-09-26 | Corixa Corporation | Compounds and methods for the dectection of t. cruzi infection |
| US20030143596A1 (en) | 2001-11-07 | 2003-07-31 | Shearwater Corporation | Branched polymers and their conjugates |
| US8431558B2 (en) | 2004-11-01 | 2013-04-30 | The Regents Of The University Of California | Compositions and methods for modification of biomolecules |
| WO2008106186A2 (en) | 2007-02-28 | 2008-09-04 | Serina Therapeutics, Inc. | Activated polyoxazolines and compositions comprising the same |
| US8703936B2 (en) | 2010-02-12 | 2014-04-22 | The Regents Of The University Of California | Compositions and methods for modification of biomolecules |
| US9222940B2 (en) | 2010-04-27 | 2015-12-29 | Synaffix B.V. | Fused cyclooctyne compounds and their use in metal-free click reactions |
| US9145361B2 (en) | 2011-03-25 | 2015-09-29 | Life Technologies Corporation | SDP-containing heterobifunctional agents |
| US20130251783A1 (en) | 2011-09-14 | 2013-09-26 | Universitat Heidelberg | Liposomes containing permeation enhancers for oral drug delivery |
| US20130189287A1 (en) | 2011-12-23 | 2013-07-25 | Paul Scherrer Institut | Enzymatic conjugation of polypeptides |
| US20140356385A1 (en) | 2011-12-23 | 2014-12-04 | Innate Pharma | Enzymatic conjugation of antibodies |
Non-Patent Citations (18)
| Title |
|---|
| "Current Protocols in Molecular Biology", 2022, WILEY PERIODICALS LLC |
| "UniProt", Database accession no. P48551 |
| ALTSCHUL ET AL.: "Nucleic Acids Res", vol. 25, 2007, pages: 3389 - 3402 |
| BARNES ET AL., ANAL. BIOCHEM., vol. 102, 1980, pages 255 |
| BUTT ET AL., PROTEIN EXPR PURIF., vol. 43, no. 1, 2009, pages 1 - 9 |
| CAI, Q ET AL., BIOTECHNOL. PROG., vol. 31, 2015, pages 823 - 831 |
| CARTER ET AL., BIOITECHNOLOGY, vol. 10, 1992, pages 163 - 167 |
| CREIGHTON: "Proteins: Structures and Molecular Properties", 1993, W. H. FREEMAN & CO. |
| GROFF ET AL., MABS, vol. 6, 2014, pages 671 - 678 |
| GROFF, D.ARMSTRONG, S.RIVERS, P.J.ZHANG, J.YANG, J.GREEN, E.ROZZELLE, J.LIANG, S.KITTLE, J.D.STEINER, A.R. ET AL.: "Engineering toward a Bacterial ''Endoplasmic Reticulum'' for the Rapid Expression of Immunoglobulin Proteins", MABS, vol. 6, 2014, pages 671 - 678, XP055241828, DOI: 10.4161/mabs.28172 |
| HAM ET AL., METH. ENZ., vol. 58, 1979, pages 138 - 161 |
| HANSON, J.GROFF, D.CARLOS, A.USMAN, H.FONG, K.YU, A.ARMSTRONG, S.DWYER, A.MASIKAT, M.R.YUAN, D. ET AL.: "An Integrated In Vivo/In Vitro Protein Production Platform for Site-Specific Antibody Drug Conjugates", BIOENGINEERING, vol. 10, 2023, pages 304 |
| HARLOWLANE: "Antibodies: A Laboratory Manual", 1988, COLD SPRING HARBOR LABORATORY, pages: 555 - 612 |
| HE ET AL., J PHARM SCI, vol. 99, 2010, pages 1707 - 1720 |
| SLEIJFER, S ET AL., PHARMACY WORLD AND SCIENCE, vol. 27, 2005, pages 423 - 431 |
| YIN ET AL., MABS, vol. 4, 2012, pages 217 - 225 |
| ZAWADA, J. F. ET AL., BIOTECHNOL. BIOENG., vol. 108, 2011, pages 1570 - 1578 |
| ZIMMERMAN. E. S. ET AL., BIOCONJUG. CHEM., vol. 25, 2014, pages 351 - 361 |
Also Published As
| Publication number | Publication date |
|---|---|
| WO2024098023A3 (en) | 2024-07-25 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US11845808B2 (en) | Peptide inhibitors of interleukin-23 receptor and their use to treat inflammatory diseases | |
| CN113660957B (en) | Compositions, methods and uses comprising antibody-TLR agonist conjugates | |
| AU2021258008A1 (en) | Conjugates of an IL-15 moiety and a polymer | |
| AU2021225133A1 (en) | Conjugates of an IL-2 moiety and a polymer | |
| KR101651703B1 (en) | FGF21 Mutants and Uses Thereof | |
| KR20220097921A (en) | Therapeutic derivatives of interleukin-22 | |
| WO2024044780A1 (en) | Interleukin-18 variants and uses thereof | |
| CN110536899A (en) | Insulin analog complexes with reduced affinity for insulin receptors and uses thereof | |
| CN116457023A (en) | antibody-TLR agonist conjugates, methods and uses thereof | |
| WO2024098023A2 (en) | Interferon alpha polypeptides and conjugates | |
| WO2023244517A1 (en) | Interleukin-2 prodrugs | |
| HK40096444A (en) | Antibody-tlr agonist conjugates, methods and uses thereof | |
| CA3144697C (en) | Conjugates of an il-2 moiety and a polymer | |
| HK40053166A (en) | Conjugates of an il-2 moiety and a polymer | |
| HK40062074A (en) | Compositions containing, methods and uses of antibody-tlr agonist conjugates | |
| WO2005085283A1 (en) | Modified interleukin-11 and medicinal composition containing the same | |
| HK1235666A1 (en) | Conjugates of il-15 and branched polyethylene glycol | |
| HK1235666B (en) | Conjugates of il-15 and branched polyethylene glycol | |
| HK1177465A1 (en) | Physiologically active polypeptide conjugate having prolonged in vivo half-life |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| 122 | Ep: pct application non-entry in european phase |
Ref document number: 23825288 Country of ref document: EP Kind code of ref document: A2 |