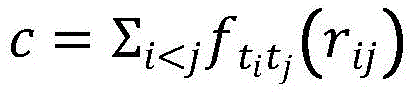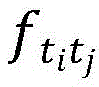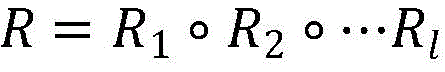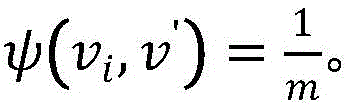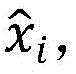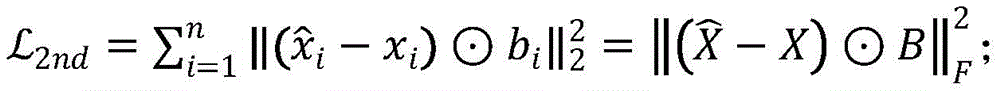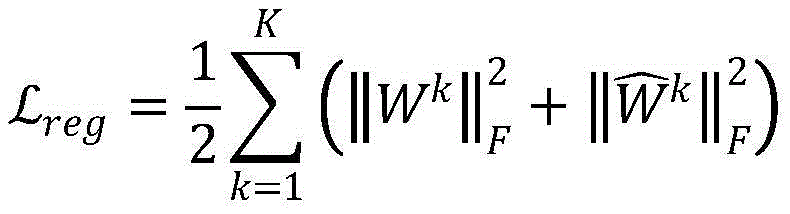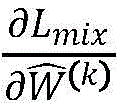CN112863634A - Traditional Chinese medicine prescription recommendation method and system based on new crown protein heterogeneous network clustering - Google Patents
Traditional Chinese medicine prescription recommendation method and system based on new crown protein heterogeneous network clustering Download PDFInfo
- Publication number
- CN112863634A CN112863634A CN202110038417.7A CN202110038417A CN112863634A CN 112863634 A CN112863634 A CN 112863634A CN 202110038417 A CN202110038417 A CN 202110038417A CN 112863634 A CN112863634 A CN 112863634A
- Authority
- CN
- China
- Prior art keywords
- chinese medicine
- traditional chinese
- protein
- compound
- clustering
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Granted
Links
Images
Classifications
-
- G—PHYSICS
- G16—INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR SPECIFIC APPLICATION FIELDS
- G16H—HEALTHCARE INFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR THE HANDLING OR PROCESSING OF MEDICAL OR HEALTHCARE DATA
- G16H20/00—ICT specially adapted for therapies or health-improving plans, e.g. for handling prescriptions, for steering therapy or for monitoring patient compliance
- G16H20/10—ICT specially adapted for therapies or health-improving plans, e.g. for handling prescriptions, for steering therapy or for monitoring patient compliance relating to drugs or medications, e.g. for ensuring correct administration to patients
-
- G—PHYSICS
- G06—COMPUTING OR CALCULATING; COUNTING
- G06F—ELECTRIC DIGITAL DATA PROCESSING
- G06F18/00—Pattern recognition
- G06F18/20—Analysing
- G06F18/22—Matching criteria, e.g. proximity measures
-
- G—PHYSICS
- G16—INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR SPECIFIC APPLICATION FIELDS
- G16B—BIOINFORMATICS, i.e. INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR GENETIC OR PROTEIN-RELATED DATA PROCESSING IN COMPUTATIONAL MOLECULAR BIOLOGY
- G16B15/00—ICT specially adapted for analysing two-dimensional or three-dimensional molecular structures, e.g. structural or functional relations or structure alignment
- G16B15/30—Drug targeting using structural data; Docking or binding prediction
-
- G—PHYSICS
- G16—INFORMATION AND COMMUNICATION TECHNOLOGY [ICT] SPECIALLY ADAPTED FOR SPECIFIC APPLICATION FIELDS
- G16C—COMPUTATIONAL CHEMISTRY; CHEMOINFORMATICS; COMPUTATIONAL MATERIALS SCIENCE
- G16C20/00—Chemoinformatics, i.e. ICT specially adapted for the handling of physicochemical or structural data of chemical particles, elements, compounds or mixtures
- G16C20/50—Molecular design, e.g. of drugs
Landscapes
- Engineering & Computer Science (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Chemical & Material Sciences (AREA)
- Health & Medical Sciences (AREA)
- Theoretical Computer Science (AREA)
- Life Sciences & Earth Sciences (AREA)
- Physics & Mathematics (AREA)
- Spectroscopy & Molecular Physics (AREA)
- General Health & Medical Sciences (AREA)
- Bioinformatics & Computational Biology (AREA)
- Medicinal Chemistry (AREA)
- Evolutionary Biology (AREA)
- Medical Informatics (AREA)
- Crystallography & Structural Chemistry (AREA)
- Data Mining & Analysis (AREA)
- Pharmacology & Pharmacy (AREA)
- Public Health (AREA)
- Artificial Intelligence (AREA)
- Epidemiology (AREA)
- Primary Health Care (AREA)
- General Physics & Mathematics (AREA)
- Computer Vision & Pattern Recognition (AREA)
- Evolutionary Computation (AREA)
- General Engineering & Computer Science (AREA)
- Computing Systems (AREA)
- Biophysics (AREA)
- Biotechnology (AREA)
- Investigating Or Analysing Biological Materials (AREA)
- Medical Treatment And Welfare Office Work (AREA)
Abstract
本发明公开一种基于新冠蛋白质异构网络聚类的中药处方推荐方法及系统,包括:构建候选中药与其化合物的边连接,和化合物与新冠蛋白质靶标的边连接,根据上述边连接构建中药‑化合物‑蛋白质异构网;确定中药‑化合物‑蛋白质异构网的元路径,计算元路径下异构节点间的相似度,以此得到中药‑蛋白质的转移概率矩阵,根据转移概率矩阵得到异构节点的嵌入向量;根据由嵌入向量计算的中药和蛋白质的欧式距离对蛋白质进行聚类,得到以结构蛋白质、非结构蛋白质及辅助蛋白质为中心的中药聚类组合;根据中药数据库对中药聚类组合中的候选中药进行筛选,得到推荐中药。通过中药‑化合物‑蛋白质异构网,基于新冠蛋白进行聚类来推荐治疗COVID‑19的潜在中药。
The invention discloses a method and system for recommending traditional Chinese medicine prescriptions based on novel coronavirus protein heterogeneous network clustering, including: constructing edge connections between candidate traditional Chinese medicines and their compounds, and edge connections between compounds and novel coronavirus protein targets, and constructing traditional Chinese medicine-compounds according to the edge connections ‑Protein Heterogeneous Network; determine the meta-path of the traditional Chinese medicine-compound-protein heterogeneous network, and calculate the similarity between the heterogeneous nodes under the meta-path, so as to obtain the transition probability matrix of the traditional Chinese medicine-protein, and obtain the heterogeneous nodes according to the transition probability matrix Embedding vector; according to the Euclidean distance between Chinese medicine and protein calculated by the embedding vector, the protein is clustered, and the traditional Chinese medicine clustering combination centered on structural protein, non-structural protein and auxiliary protein is obtained; according to the traditional Chinese medicine database, the traditional Chinese medicine clustering combination is The candidate Chinese medicines were screened and recommended Chinese medicines were obtained. Through the TCM-compound-protein heterogeneous network, clustering based on SARS-CoV-2 was used to recommend potential TCMs for the treatment of COVID-19.
Description
技术领域technical field
本发明涉及药物推荐技术领域,特别是涉及一种基于新冠蛋白质异构网络聚类的中药处方推荐方法及系统。The invention relates to the technical field of drug recommendation, in particular to a method and system for recommending traditional Chinese medicine prescriptions based on novel coronavirus protein heterogeneous network clustering.
背景技术Background technique
本部分的陈述仅仅是提供了与本发明相关的背景技术信息,不必然构成在先技术。The statements in this section merely provide background information related to the present invention and do not necessarily constitute prior art.
中药药物重定位是将已经批准应用于临床的中药单体或中药复方,通过进一步研究,扩大其临床适应症、发现其新的作用靶点或机制的研究。新冠病毒的基因组不到3万个碱基,能够转录29种蛋白质,其中有16种非结构蛋白、4 种结构蛋白和9种辅助蛋白,明确这29种蛋白质与化合物之间的相互作用,寻找作用靶点,对药物研发至关重要。由于新冠病毒通过多种蛋白质作用于人体,如果通过单靶点治疗则不会产生良好的治疗效果,因此研究者大多采用多靶点的治疗手段治疗新冠肺炎患者,中药恰好就具有多成分-多靶点的特点。利用网络药理学,在“药物-化合物-靶点”相互作用网络基础上分析药物的作用机制及其主要成分,通过对多靶点的研究,治疗效果提升,从而提高药物重定位的成功率。TCM drug repositioning is the study of expanding the clinical indications and discovering new targets or mechanisms of TCM monomers or TCM compounds that have been approved for clinical use. The genome of the new coronavirus is less than 30,000 bases, and it can transcribe 29 proteins, including 16 non-structural proteins, 4 structural proteins and 9 auxiliary proteins. Targets of action are crucial to drug development. Since the new coronavirus acts on the human body through a variety of proteins, if it is treated with a single target, it will not produce a good therapeutic effect. Therefore, researchers mostly use multi-target therapy to treat patients with new coronary pneumonia. Traditional Chinese medicine happens to have multiple components-multiple characteristics of the target. Using network pharmacology, the mechanism of drug action and its main components are analyzed on the basis of the "drug-compound-target" interaction network. Through the study of multiple targets, the therapeutic effect can be improved, thereby improving the success rate of drug repositioning.
为了挖掘可替代治疗COVID-19的潜在方剂,在文献Identifying potentialtreatments of COVID-19from Traditional Chinese Medicine(TCM)by using a data-driven approach中,Ren利用数据挖掘和关联网络对古代方剂中高频药材和方剂进行挖掘。通过分子对接的方法,探索高频CMs与配伍中药有效成分的虚拟筛选。此外,在文献Analysis on herbal medicines utilized fortreatment of COVID-19中,Luo通过使用复杂系统熵和无监督层次聚类,8种核心草药组合和10种新配方可能成为COVID-19的有用候选药物。这些寻找潜在中药方剂的方法都采用了较为传统的数据挖掘和聚类方法,以频率为依据发现潜在方剂,但是没有考虑草药、化合物以及蛋白质之间的交互作用。In order to mine potential prescriptions for alternative treatment of COVID-19, in the document Identifying potentialtreatments of COVID-19 from Traditional Chinese Medicine (TCM) by using a data-driven approach, Ren used data mining and association networks to identify high-frequency medicinal materials and prescriptions in ancient prescriptions to dig. Through the method of molecular docking, the virtual screening of high-frequency CMs and compatible traditional Chinese medicines was explored. Furthermore, in the literature Analysis on herbal medicines utilized for treatment of COVID-19, Luo, by using complex system entropy and unsupervised hierarchical clustering, 8 core herbal combinations and 10 new formulations may become useful drug candidates for COVID-19. These methods for finding potential TCM formulas all use more traditional data mining and clustering methods to discover potential formulas based on frequency, but do not consider the interactions among herbs, compounds, and proteins.
发明内容SUMMARY OF THE INVENTION
为了解决上述问题,本发明提出了一种基于新冠蛋白质异构网络聚类的中药处方推荐方法及系统,通过中药-化合物-蛋白质异构网,基于新冠蛋白进行聚类,得到分别以结构蛋白、非结构蛋白及辅助蛋白为中心的中药聚类组合,根据聚类结果来推荐治疗COVID-19的潜在中药。In order to solve the above problems, the present invention proposes a method and system for recommending traditional Chinese medicine prescriptions based on novel coronavirus protein heterogeneous network clustering. A clustering combination of traditional Chinese medicines centered on non-structural proteins and auxiliary proteins, and the potential traditional Chinese medicines for the treatment of COVID-19 are recommended according to the clustering results.
为了实现上述目的,本发明采用如下技术方案:In order to achieve the above object, the present invention adopts the following technical solutions:
第一方面,本发明提供一种基于新冠蛋白质异构网络聚类的中药处方推荐方法,包括:In a first aspect, the present invention provides a method for recommending traditional Chinese medicine prescriptions based on novel coronavirus protein heterogeneous network clustering, including:
构建候选中药与其化合物的边连接,以及化合物与新冠蛋白质靶标的边连接,根据上述边连接构建中药-化合物-蛋白质异构网;Construct edge connections between candidate traditional Chinese medicines and their compounds, as well as edge connections between compounds and new crown protein targets, and construct a traditional Chinese medicine-compound-protein heterogeneous network based on the above edge connections;
确定中药-化合物-蛋白质异构网的元路径,计算元路径下异构节点间的相似度,以此得到中药-蛋白质的转移概率矩阵,根据转移概率矩阵得到异构节点的嵌入向量;Determine the meta-path of the Chinese medicine-compound-protein heterogeneous network, calculate the similarity between the heterogeneous nodes under the meta-path, and obtain the transition probability matrix of the Chinese medicine-protein, and obtain the embedding vector of the heterogeneous nodes according to the transition probability matrix;
根据由嵌入向量计算的中药和蛋白质的欧式距离对蛋白质进行聚类,得到分别以结构蛋白质、非结构蛋白质及辅助蛋白质为中心的中药聚类组合;According to the Euclidean distance of the Chinese medicine and the protein calculated by the embedding vector, the protein is clustered, and the Chinese medicine clustering combination centered on the structural protein, the non-structural protein and the auxiliary protein is obtained respectively;
根据中药数据库对中药聚类组合中的候选中药进行筛选,得到推荐中药。According to the traditional Chinese medicine database, the candidate traditional Chinese medicines in the traditional Chinese medicine clustering combination are screened, and the recommended traditional Chinese medicines are obtained.
第二方面,本发明提一种基于新冠蛋白质异构网络聚类的中药处方推荐系统,包括:In the second aspect, the present invention provides a traditional Chinese medicine prescription recommendation system based on novel coronavirus protein heterogeneous network clustering, including:
网络构建模块,被配置为构建候选中药与其化合物的边连接,以及化合物与新冠蛋白质靶标的边连接,根据上述边连接构建中药-化合物-蛋白质异构网;The network building block is configured to construct the edge connection between the candidate traditional Chinese medicine and its compound, as well as the edge connection between the compound and the new crown protein target, and construct the traditional Chinese medicine-compound-protein heterogeneous network according to the above-mentioned edge connection;
相似度计算模块,被配置为确定中药-化合物-蛋白质异构网的元路径,计算元路径下异构节点间的相似度,以此得到中药-蛋白质的转移概率矩阵,根据转移概率矩阵得到异构节点的嵌入向量;The similarity calculation module is configured to determine the meta-path of the traditional Chinese medicine-compound-protein heterogeneous network, and calculate the similarity between the heterogeneous nodes under the meta-path, so as to obtain the transition probability matrix of the traditional Chinese medicine-protein, and obtain the heterogeneity according to the transition probability matrix. Embedding vector for constructing nodes;
聚类模块,被配置为根据由嵌入向量计算的中药和蛋白质的欧式距离对蛋白质进行聚类,得到分别以结构蛋白质、非结构蛋白质及辅助蛋白质为中心的中药聚类组合;The clustering module is configured to cluster the proteins according to the Euclidean distance of the traditional Chinese medicine and the protein calculated by the embedding vector, and obtain the traditional Chinese medicine clustering combination centered on the structural protein, the non-structural protein and the auxiliary protein respectively;
推荐模块,被配置为根据中药数据库对中药聚类组合中的候选中药进行筛选,得到推荐中药。The recommendation module is configured to screen the candidate traditional Chinese medicines in the traditional Chinese medicine clustering combination according to the traditional Chinese medicine database to obtain the recommended traditional Chinese medicines.
第三方面,本发明提供一种电子设备,包括存储器和处理器以及存储在存储器上并在处理器上运行的计算机指令,所述计算机指令被处理器运行时,完成第一方面所述的方法。In a third aspect, the present invention provides an electronic device, comprising a memory, a processor, and computer instructions stored in the memory and executed on the processor, and when the computer instructions are executed by the processor, the method described in the first aspect is completed .
第四方面,本发明提供一种计算机可读存储介质,用于存储计算机指令,所述计算机指令被处理器执行时,完成第一方面所述的方法。In a fourth aspect, the present invention provides a computer-readable storage medium for storing computer instructions, and when the computer instructions are executed by a processor, the method described in the first aspect is completed.
与现有技术相比,本发明的有益效果为:Compared with the prior art, the beneficial effects of the present invention are:
本发明通过建立中药-化合物-蛋白质异构网,基于新冠蛋白进行聚类来推荐治疗COVID-19的潜在中药。首先将候选中药及其包含的化合物建立边连接,利用分子对接建立化合物和新冠蛋白质靶标之间的边连接,以此得到中药-化合物-蛋白质异构网;其次利用结构化网络嵌入模型SDNE得到所有节点的嵌入向量;最终通过计算蛋白质与中药的欧氏距离,得到分别以结构蛋白、非结构蛋白及辅助蛋白为中心的中药聚类组合,这些中药相应的新冠蛋白可能存在潜在的疗效。另外,本发明还通过计算聚类组合中的中药和诊疗指南中中药的余弦相似度,得到指南中药中与推荐中药相似度最高且用于推荐治疗的潜在中药,这些结果具有替代诊疗指南中中药的可能性,并且为治疗COVID-19提供潜在的候选中药。The present invention recommends potential Chinese medicines for treating COVID-19 by establishing a Chinese medicine-compound-protein heterogeneous network and clustering based on the novel coronavirus protein. First, the candidate traditional Chinese medicines and the compounds contained in them are established to establish edge connections, and molecular docking is used to establish the edge connections between the compounds and the new crown protein target, so as to obtain the traditional Chinese medicine-compound-protein heterogeneous network; secondly, the structured network embedding model SDNE is used to obtain all the The embedding vector of the node; finally, by calculating the Euclidean distance between the protein and the traditional Chinese medicine, the clustering combination of the traditional Chinese medicine centered on the structural protein, the non-structural protein and the auxiliary protein is obtained, and the corresponding new crown protein of these traditional Chinese medicine may have potential curative effect. In addition, the present invention also calculates the cosine similarity between the traditional Chinese medicine in the cluster combination and the traditional Chinese medicine in the diagnosis and treatment guide, and obtains the potential traditional Chinese medicine with the highest similarity with the recommended traditional Chinese medicine in the guide traditional Chinese medicine and used for the recommended treatment, and these results can replace the traditional Chinese medicine in the diagnosis and treatment guide. possibility and provide a potential candidate traditional Chinese medicine for the treatment of COVID-19.
由于一些异构节点之间没有边直接连接,导致一阶相似度不足以保存网络结构,为解决该问题,本发明引入二阶相似度,通过二阶相似度保留全局信息,实现通过一阶相似度和二阶相似度同时保留全局结构和局部结构。Because there is no direct connection between some heterogeneous nodes, the first-order similarity is not enough to preserve the network structure. In order to solve this problem, the present invention introduces the second-order similarity, retains the global information through the second-order similarity, and realizes the first-order similarity through the first-order similarity. degree and second-order similarity preserve both global and local structure.
本发明建立的异构网包含多个异构节点和关系,深度挖掘中药、化合物、蛋白质三者的交互信息,产生更优的聚类结果。The heterogeneous network established by the invention includes a plurality of heterogeneous nodes and relationships, and the interactive information of traditional Chinese medicine, compounds and proteins is deeply excavated to generate better clustering results.
本发明附加方面的优点将在下面的描述中部分给出,部分将从下面的描述中变得明显,或通过本发明的实践了解到。Advantages of additional aspects of the invention will be set forth in part in the description which follows, and in part will become apparent from the description which follows, or may be learned by practice of the invention.
附图说明Description of drawings
构成本发明的一部分的说明书附图用来提供对本发明的进一步理解,本发明的示意性实施例及其说明用于解释本发明,并不构成对本发明的不当限定。The accompanying drawings forming a part of the present invention are used to provide further understanding of the present invention, and the exemplary embodiments of the present invention and their descriptions are used to explain the present invention, and do not constitute an improper limitation of the present invention.
图1为本发明实施例1提供的基于新冠蛋白质异构网络聚类的中药处方推荐方法流程图;1 is a flowchart of a method for recommending traditional Chinese medicine prescriptions based on novel coronavirus protein heterogeneous network clustering provided in Example 1 of the present invention;
图2(a)-2(f)为本发明实施例1提供的可替代治疗新冠的中药处方的推荐流示意图。Figures 2(a)-2(f) are schematic diagrams of the recommended flow of Chinese medicine prescriptions that can replace the new crown treatment provided in Example 1 of the present invention.
具体实施方式:Detailed ways:
下面结合附图与实施例对本发明做进一步说明。The present invention will be further described below with reference to the accompanying drawings and embodiments.
应该指出,以下详细说明都是示例性的,旨在对本发明提供进一步的说明。除非另有指明,本文使用的所有技术和科学术语具有与本发明所属技术领域的普通技术人员通常理解的相同含义。It should be noted that the following detailed description is exemplary and intended to provide further explanation of the invention. Unless otherwise defined, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs.
需要注意的是,这里所使用的术语仅是为了描述具体实施方式,而非意图限制根据本发明的示例性实施方式。如在这里所使用的,除非上下文另外明确指出,否则单数形式也意图包括复数形式,此外,还应当理解的是,术语“包括”和“具有”以及他们的任何变形,意图在于覆盖不排他的包含,例如,包含了一系列步骤或单元的过程、方法、系统、产品或设备不必限于清楚地列出的那些步骤或单元,而是可包括没有清楚地列出的或对于这些过程、方法、产品或设备固有的其它步骤或单元。It should be noted that the terminology used herein is for the purpose of describing specific embodiments only, and is not intended to limit the exemplary embodiments according to the present invention. As used herein, unless the context clearly dictates otherwise, the singular is intended to include the plural as well, furthermore, it is to be understood that the terms "including" and "having" and any conjugations thereof are intended to cover the non-exclusive A process, method, system, product or device comprising, for example, a series of steps or units is not necessarily limited to those steps or units expressly listed, but may include those steps or units not expressly listed or for such processes, methods, Other steps or units inherent to the product or equipment.
在不冲突的情况下,本发明中的实施例及实施例中的特征可以相互组合。Embodiments of the invention and features of the embodiments may be combined with each other without conflict.
实施例1Example 1
如图1和图2(a)-(f)所示,本实施例提供一种基于新冠蛋白质异构网络聚类的中药处方推荐方法,包括:As shown in Figure 1 and Figure 2(a)-(f), this embodiment provides a method for recommending traditional Chinese medicine prescriptions based on novel coronavirus protein heterogeneous network clustering, including:
S1:构建候选中药与其化合物的边连接,以及化合物与新冠蛋白质靶标的边连接,根据上述边连接构建中药-化合物-蛋白质异构网;S1: Construct the edge connection between the candidate traditional Chinese medicine and its compound, as well as the edge connection between the compound and the new crown protein target, and construct the traditional Chinese medicine-compound-protein heterogeneous network according to the above edge connection;
S2:确定中药-化合物-蛋白质异构网的元路径,计算元路径下异构节点间的相似度,以此得到中药-蛋白质的转移概率矩阵,根据转移概率矩阵得到异构节点的嵌入向量;S2: Determine the meta-path of the traditional Chinese medicine-compound-protein heterogeneous network, calculate the similarity between the heterogeneous nodes under the meta-path, so as to obtain the transition probability matrix of the traditional Chinese medicine-protein, and obtain the embedding vector of the heterogeneous nodes according to the transition probability matrix;
S3:根据由嵌入向量计算的中药和蛋白质的欧式距离对蛋白质进行聚类,得到分别以结构蛋白质、非结构蛋白质及辅助蛋白质为中心的中药聚类组合;S3: Cluster the proteins according to the Euclidean distance between the Chinese medicine and the protein calculated by the embedding vector, and obtain the Chinese medicine clustering combination centered on the structural protein, the non-structural protein and the auxiliary protein respectively;
S4:根据中药数据库对中药聚类组合中的候选中药进行筛选,得到推荐中药。S4: Screen the candidate traditional Chinese medicines in the traditional Chinese medicine clustering combination according to the traditional Chinese medicine database, and obtain the recommended traditional Chinese medicines.
在本实施例中,从中药系统药理学数据库及分析平台TCMS中获取482个候选中药、13448个化合物结构和相应的ADME信息;然后通过OpenBabel将其转换成PDBQT格式。In this example, 482 candidate traditional Chinese medicines, 13,448 compound structures and corresponding ADME information were obtained from the traditional Chinese medicine system pharmacology database and analysis platform TCMS; then they were converted into PDBQT format through OpenBabel.
从蛋白数据带(PROTEIN DATA BAND,RSCB)中下载现有的SARS-CoV-2 结构,包括S、nsp5、nsp7、nsp8、nsp9、nsp10、nsp12、nsp15、nsp16;针对结构不明确的SARS-CoV-2,本实施例从实验室下载预测蛋白结构,预测蛋白结构为E、N、M、orf3a、orf6、orf7a、orf8、orf10、nsp1、nsp2、nsp3、nsp4、nsp6、 nsp13、nsp14;总共得到24个蛋白结构,使用OpenBabel对下载的pdb文件进行脱水和氢化操作,并将其转换为PDBQT格式。Download the existing SARS-CoV-2 structure from PROTEIN DATA BAND (RSCB), including S, nsp5, nsp7, nsp8, nsp9, nsp10, nsp12, nsp15, nsp16; for SARS-CoV with unclear structure -2, in this example, the predicted protein structure is downloaded from the laboratory, and the predicted protein structure is E, N, M, orf3a, orf6, orf7a, orf8, orf10, nsp1, nsp2, nsp3, nsp4, nsp6, nsp13, nsp14; a total of For 24 protein structures, the downloaded pdb files were dehydrated and hydrogenated using OpenBabel and converted to PDBQT format.
中药靶标是指体内具有药效功能并能被中药作用的生物大分子,如某些蛋白质和核酸等生物大分子,目前中药的靶标主要是蛋白质。中药与靶蛋白的相互作用是很多中药发挥生物学功能的基础,由于目前没有针对新冠的特效药,因此本实施例通过研究新冠病毒编码的蛋白质寻找打到靶标上的中药。The target of traditional Chinese medicine refers to the biological macromolecules that have pharmacodynamic functions in the body and can be acted on by traditional Chinese medicine, such as some biological macromolecules such as proteins and nucleic acids. The interaction between traditional Chinese medicines and target proteins is the basis for many traditional Chinese medicines to exert their biological functions. Since there is currently no specific drug for the new crown virus, this example searches for the traditional Chinese medicine that hits the target by studying the protein encoded by the new coronavirus.
SARS-CoV-2属于冠状病毒科,为不分节段的单股正链RNA病毒,它编码 29种蛋白质,其中有4种结构蛋白(S、E、M和N蛋白),16个非结构蛋白 (nsp1-nsp16)以及9种辅助蛋白;其中,4种结构蛋白S、E、M和N形成冠状病毒外层,保护内部的RNA;S蛋白是病毒进入宿主细胞的重要决定因子,与ACE2具有较高的结合强度;N蛋白与病毒基因组RNA结合,将RNA包装成核糖核蛋白(RNP)复合体;E蛋白是包膜蛋白,将病毒内部的遗传物质包裹起来。16个非结构蛋白(nsp1-nsp16),用于形成病毒复制酶转录酶复合物,这部分蛋白一般是传统小分子抗病毒中药的靶点,非结构蛋白中NSP7、NSP8和 NSP12形成核糖核酸(RNA)依赖的RNA聚合酶复合物,在病毒的复制和转录周期中发挥重要作用;NSP5是3C蛋白水解酶,可以使大多数其他NSP蛋白质从蛋白链上解放出来,被认为是抗新冠肺炎病毒中药研究的重要靶点。SARS-CoV-2 belongs to the family Coronaviridae and is a non-segmented single-stranded positive-stranded RNA virus. It encodes 29 proteins, including 4 structural proteins (S, E, M and N proteins) and 16 non-structural proteins. Proteins (nsp1-nsp16) and 9 accessory proteins; among them, 4 structural proteins S, E, M and N form the outer layer of the coronavirus and protect the internal RNA; the S protein is an important determinant for the virus to enter the host cell, and ACE2 It has a high binding strength; the N protein binds to the viral genome RNA and packages the RNA into a ribonucleoprotein (RNP) complex; the E protein is an envelope protein that wraps the genetic material inside the virus. 16 non-structural proteins (nsp1-nsp16) are used to form viral replicase transcriptase complexes. These proteins are generally the targets of traditional small-molecule antiviral Chinese medicines. Among the non-structural proteins, NSP7, NSP8 and NSP12 form ribonucleic acid ( RNA)-dependent RNA polymerase complex, which plays an important role in the replication and transcription cycle of the virus; NSP5 is a 3C proteolytic enzyme, which can liberate most other NSP proteins from the protein chain, and is considered to be anti-COVID-19 virus An important target for Chinese medicine research.
对于异构图;图被定义为G=(V,E,w),其中V={v1,v2,…,vn}表示n个顶点;表示边;w表示边权重的集合,如果vi和vj中间没有边相连,wij=0,否则wij>0;每一个节点vi∈V都属于一个特定的节点类型每一个边 eij∈E都属于关系类型集合R:φ(e)∈R中的一个特定的关系类型,当节点类型的总数目|A|>1或者边的类型的总数目|R|>1时,得到的图是异构图。For heterogeneous graphs; the graph is defined as G=(V,E,w), where V={v 1 ,v 2 ,...,v n } denotes n vertices; Represents an edge; w represents the set of edge weights, if there is no edge connected between vi and v j, w ij = 0 , otherwise w ij >0; each node vi ∈V belongs to a specific node type Each edge e ij ∈ E belongs to the set of relation types R: φ(e)∈R for a particular relation type, when the total number of node types |A|>1 or the total number of edge types |R|> 1, the resulting graph is a heterogeneous graph.
那么,本实施例获取候选中药、中药所包含的化合物以及新冠蛋白质靶标后,根据候选中药与其化合物的边连接,以及化合物与新冠蛋白质靶标的边连接,构建中药-化合物-蛋白质异构网G=(V,E);具体地:Then, after obtaining the candidate traditional Chinese medicine, the compound contained in the traditional Chinese medicine, and the new crown protein target in this example, according to the edge connection between the candidate traditional Chinese medicine and its compound, and the edge connection between the compound and the new crown protein target, the traditional Chinese medicine-compound-protein heterogeneous network G= (V,E); specifically:
首先,将482个中药及其包含的13448化合物建立边连接,如果中药包含某些化合物,那么该中药和这些化合物之间有边连接;First, establish edge connections between 482 traditional Chinese medicines and the 13448 compounds they contain. If the traditional Chinese medicines contain certain compounds, there are edge connections between the traditional Chinese medicines and these compounds;
然后,建立化合物和24中新冠蛋白质之间的边连接;本实施例使用分子对接的方式对化合物-蛋白质的结合亲和力进行判断,使用打分函数c计算结合亲和力,即:Then, establish the edge connection between the compound and the new crown protein in 24; in this example, the molecular docking method is used to judge the binding affinity of the compound-protein, and the scoring function c is used to calculate the binding affinity, namely:
其中,是空间相互作用的加权和,rij是原子间的距离。in, is the weighted sum of spatial interactions, and r ij is the distance between atoms.
从热力学的观点来看,配体-受体相互作用是一个综合平衡的过程,生物分子的稳定构象是自由能最低的构象,本实施例将化合物及蛋白分别对接10次,并取10次平均值为对接的结果。当结合自由能,即结合亲和力小于-7kcal/mol 时,该化合物与蛋白视为可以有效结合,反之则无效,将结合自由能小于-7kcal/mol的化合物和蛋白质建立边连接。From the point of view of thermodynamics, the ligand-receptor interaction is a comprehensive equilibrium process. The stable conformation of biomolecules is the conformation with the lowest free energy. In this example, the compound and the protein are docked 10 times respectively, and the average of 10 times is taken. The value is the result of the docking. When the binding free energy, that is, the binding affinity is less than -7kcal/mol, the compound and the protein are considered to be able to bind effectively, otherwise, it is invalid, and the compound with the binding free energy less than -7kcal/mol is connected with the protein.
中药-化合物-蛋白质异构网G中,异构节点之间通过多种元路径连接,元路径是异构网络的核心,是建模一个特定关系时,带有边类型的节点类型序列。一条元路径ρ被定义为由边的类型R1,…Rl连接的节点类型A1,…Al+1的路径序列,如表示一个节点类型的综合关系,表示关系的合成器。元路径描述节点之间的语义关系,不同的元路径描述节点之间不同的语义关系,这个语义关系的开发是多种后续任务的基石。In the traditional Chinese medicine-compound-protein heterogeneous network G, heterogeneous nodes are connected by a variety of meta-paths. The meta-path is the core of the heterogeneous network and is a sequence of node types with edge types when modeling a specific relationship. A meta-path ρ is defined as a sequence of paths of node types A 1 ,...A l+1 connected by edge types R 1 ,...R l , such as Represents a node type comprehensive relationship, A compositor representing a relationship. Meta-paths describe the semantic relationship between nodes, and different meta-paths describe different semantic relationships between nodes. The development of this semantic relationship is the cornerstone of various subsequent tasks.
本实施例构建四条元路径,包括中药-化合物-中药、中药-化合物-蛋白质、蛋白质-化合物-中药、蛋白质-化合物-蛋白质;其中,中药->化合物->中药这条元路径中表示两种中药包含同一种化合物,中药->化合物->蛋白质表示中药通过某一种化合物打到某蛋白质上,蛋白质->化合物->中药表示蛋白质与某中药中的化合物结合,蛋白质->化合物->蛋白质表示一种化合物可以打到两种蛋白质上。In this example, four meta-paths are constructed, including Chinese medicine-compound-Chinese medicine, Chinese medicine-compound-protein, protein-compound-Chinese medicine, protein-compound-protein; wherein, the meta-path of Chinese medicine->compound->Chinese medicine represents two Chinese medicine contains the same compound, Chinese medicine->compound->protein means that Chinese medicine hits a protein through a certain compound, protein->compound->traditional medicine means that protein is combined with a compound in a Chinese medicine, protein->compound->protein Indicates that a compound can hit two proteins.
确定元路径后,采用随机游走方法计算节点vi和vj之间的邻近度,例如给定元路径p:H->C->H,一个中药节点h1连接三个化合物,其中仅有一个化合物连接着中药h2,并且该化合物还对接中药h3,那么中药h1到中药h2的转移概率为 After the meta-path is determined, the random walk method is used to calculate the proximity between nodes v i and v j . For example, given a meta-path p: H->C->H, a traditional Chinese medicine node h 1 connects three compounds, of which only There is a compound connected to traditional Chinese medicine h 2 , and this compound is also connected to traditional Chinese medicine h 3 , then the transition probability of traditional Chinese medicine h 1 to traditional Chinese medicine h 2 is
由于一些节点之间没有边直接连接,所以仅用一阶相似度不足以保存网络结构,一阶相似度代表两个节点之间的局部邻近度,如果两个节点之间有连接,则边的权重代表节点vi和vj之间的一阶相似度;否则,节点vi和vj的一阶相似度为0;而二阶相似度衡量没有边连接的两个节点之间的相似度,节点vi和vj的二阶相似度是邻近网络结构之间的相似性。Since there is no direct connection between some nodes, only the first-order similarity is not enough to preserve the network structure. The first-order similarity represents the local proximity between two nodes. If there is a connection between the two nodes, the edge The weight represents the first-order similarity between nodes v i and v j ; otherwise, the first-order similarity between nodes v i and v j is 0; while the second-order similarity measures the similarity between two nodes without edge connections , the second-order similarity of nodes v i and v j is the similarity between adjacent network structures.
因此本实施例考虑二阶相似度,通过二阶相似度来保留全局信息。其中,本实施例采用pu=(wu,1,…,wu,|V|)表示u与所有其他顶点的一阶相似度,则u 和v之间的二阶相似度由pu和pv决定;如果节点vi和vj共同的邻居越多,则这两个节点越相似;如果没有顶点与u和v都连接,则u和v之间的二阶相似度为0。Therefore, in this embodiment, the second-order similarity is considered, and the global information is preserved through the second-order similarity. Among them, in this embodiment, p u =(w u ,1,...,w u ,|V|) is used to represent the first-order similarity between u and all other vertices, then the second-order similarity between u and v is represented by p u and p v ; if the nodes v i and v j have more neighbors in common, the two nodes are more similar; if there is no vertex connected to both u and v, the second-order similarity between u and v is 0.
本实施例基于确定的元路径HCH、HCP、PCH、PCP,采用随机游走方法计算两节点之间的邻近度,构建中药-蛋白质的关系矩阵A,矩阵A的每一个元素是两节点之间的转移概率,行向量即为每个节点的嵌入向量,矩阵A提供了每个节点的邻域结构信息。其中,随机游走方法包括:给定图G=(V,E),选定初始节点,随机选择一个邻居节点,将初始节点移动到邻居节点上;然后把当前节点作为出发点,重复以上过程,被随机选出来的节点序列构成一个随机游走过程。Based on the determined meta-paths HCH, HCP, PCH, and PCP, this embodiment uses a random walk method to calculate the proximity between two nodes, and constructs a traditional Chinese medicine-protein relationship matrix A, where each element of the matrix A is between two nodes. The transition probability of , the row vector is the embedding vector of each node, and the matrix A provides the neighborhood structure information of each node. Among them, the random walk method includes: given a graph G=(V, E), select an initial node, randomly select a neighbor node, move the initial node to the neighbor node; then use the current node as the starting point, repeat the above process, The randomly selected node sequence constitutes a random walk process.
中药-蛋白质的转移概率定义如下:The TCM-protein transition probability is defined as follows:
其中,s(vi,vj|ρ)表示在元路径ρ下,节点vi和节点vj的相似度,l是元路径长度的阈值。Among them, s(v i , v j |ρ) represents the similarity between node v i and node v j under the meta-path ρ, and l is the threshold of the meta-path length.
对于基于相似度的随机游走事件,有下面的属性:For similarity-based random walk events, there are the following properties:
其中,是通过关系类型ψ(vi,v′)从vi到v′的转移概率,如果从vi出发有m条边属于关系类型ψ(vi,v′),那么 in, is the transition probability from v i to v' through the relation type ψ(vi , v' ), if there are m edges from v i belonging to the relation type ψ(vi , v'), then
本实施例定义基于中药元路径的相似度计算如下:This embodiment defines the similarity calculation based on the meta-path of traditional Chinese medicine as follows:
其中,ρ[i:j]是从节点vi到节点vj,基于元路径的相似度计算是一个动态设计方法。Among them, ρ[i:j] is from node v i to node v j , and the similarity calculation based on meta-path is a dynamic design method.
在本实施例中,使用随机游走计算两节点之间的相似度,进而得到中药-蛋白质的转移概率矩阵T,转移概率矩阵的第i行代表第i个节点的向量,根据中药-蛋白质的转移概率矩阵得到异构节点的嵌入向量,构建基于相似度的元路径中药图嵌入;具体地:给定网络G=(V,E),寻映射f,使得f:vi→yi,其中yi的维度是d,且yi∈Rd,d<<V。因此,映射f对每个节点向量进行降维,尽管节点类型不一样,最终得到的向量维度相同。这个过程是中药网络的嵌入,得到的嵌入向量仍然保留原来网络的结构及语义关系。In this embodiment, a random walk is used to calculate the similarity between two nodes, and then the transition probability matrix T of traditional Chinese medicine-protein is obtained, and the i-th row of the transition probability matrix is obtained. The vector representing the ith node, the embedding vector of heterogeneous nodes is obtained according to the transition probability matrix of traditional Chinese medicine-protein, and the similarity-based meta-path traditional Chinese medicine graph embedding is constructed; specifically: given the network G=(V, E), find Map f such that f: v i →y i , where the dimension of y i is d, and y i ∈ R d , d<<V. Therefore, the mapping f reduces the dimension of each node vector, although the node types are different, the final vector dimension is the same. This process is the embedding of the traditional Chinese medicine network, and the obtained embedding vector still retains the structure and semantic relationship of the original network.
由于只有少数的化合物能打到蛋白质靶点上,导致只有少数的中药节点和蛋白质节点之间有边,因此在中药-蛋白质的转移概率矩阵T中,大多数元素为 0,即转移概率矩阵是一个稀疏矩阵;此外,本实施例构建的异构网络还具有高度的非线性性,为了捕捉高度的非线性结构,同时解决稀疏问题,本实施例采用结构化深度网络嵌入(SDNE),SDNE不仅能捕捉非线性结构,还可以保留局部和全局的网络结构。SDNE是一个实现网络嵌入的半监督模型,探索一阶和二阶相似度,SDNE由监督部分和无监督部分组成。无监督的组成部分是一个深度自编码器,可以捕捉重构输入数据ti后的二阶相似度,且二阶相似度保留了全局结构;监督部分可以通过拉普拉斯特征映像保留一阶相似度。Since only a few compounds can hit the protein target, there are only a few edges between TCM nodes and protein nodes, so in the TCM-protein transition probability matrix T, most elements are 0, that is, the transition probability matrix is A sparse matrix; in addition, the heterogeneous network constructed in this embodiment also has a high degree of nonlinearity. In order to capture the highly nonlinear structure and solve the sparse problem at the same time, this embodiment adopts structured deep network embedding (SDNE), SDNE not only It can capture the nonlinear structure, and can also preserve the local and global network structure. SDNE is a semi-supervised model that implements network embeddings to explore first- and second-order similarities, SDNE consists of a supervised part and an unsupervised part. The unsupervised component is a deep autoencoder, which can capture the second-order similarity after reconstructing the input data t i , and the second-order similarity preserves the global structure; the supervised part can preserve the first-order through the Laplacian feature map similarity.
由此,本实施例采用SDNE构建中药-化合物-蛋白质异构网的目标函数,具体地:Thus, the present embodiment adopts SDNE to construct the objective function of traditional Chinese medicine-compound-protein heterogeneous network, specifically:
深度自编码器由编码器和解码器组成,编码器将输入压缩成潜在空间表征,解码器重构来自潜在空间表征的输入。给定输入xi,每一层的隐藏表示如下:A deep autoencoder consists of an encoder and a decoder, the encoder compresses the input into a latent space representation, and the decoder reconstructs the input from the latent space representation. Given an input x i , the hidden representation of each layer is as follows:
其中,σ是sigmoid函数;W(k)是第k层的权重矩阵;b(k)第k层的偏差。where σ is the sigmoid function; W (k) is the weight matrix of the kth layer; b (k) is the bias of the kth layer.
本实施例通过逆转编码器的计算过程得到输出假设第k层是节点vi的表示向量(仅考虑全局信息),那么从第k层开始解码,最终得到解码后的向量那么,自编码器的误差就是输入节点vi的邻接向量的重构误差。因此,自编码器的目标是尽量减小输入和输出的重构误差。损失函数如下:This embodiment obtains the output by reversing the calculation process of the encoder Assuming that the kth layer is the representation vector of the node v i (only global information is considered), then start decoding from the kth layer, and finally get the decoded vector Then, the error of the autoencoder is the reconstruction error of the adjacency vector of the input node v i . Therefore, the goal of an autoencoder is to minimize the reconstruction error of the input and output. The loss function is as follows:
由于本实施例构建的网络具有稀疏性,即转移概率矩阵T中零元素的数量远远多于非零元素的数量。如果直接使用T作为输入,则更容易重构T中的零元素;为了解决稀疏性带来的问题,对非零元的重构误差施加了比零元更大的惩罚;修正后的目标函数如下:Due to the sparseness of the network constructed in this embodiment, that is, the number of zero elements in the transition probability matrix T is far more than the number of non-zero elements. If T is used directly as input, it is easier to reconstruct the zero elements in T; in order to solve the problem caused by sparsity, a larger penalty is imposed on the reconstruction error of non-zero elements than zero elements; the modified objective function as follows:
其中,⊙是Hadamardproduct,表示对应元素相乘;邻接矩阵中的0对应b=1,非0元素的b>1;这样的目的是对有边连接的节点赋予更高权重。Among them, ⊙ is Hadamard product, which means that the corresponding elements are multiplied; 0 in the adjacency matrix corresponds to b=1, and b>1 for non-zero elements; the purpose of this is to give higher weights to nodes connected by edges.
以上获得二阶相似度的损失函数,该模型的无监督部分可以通过重建顶点之间的二阶邻近度来保持全局网络结构;除此之外,本实施例还利用一阶相似度捕捉局部结构。一阶相似度损失函数定义如下:The above loss function to obtain the second-order similarity, the unsupervised part of the model can maintain the global network structure by reconstructing the second-order proximity between vertices; in addition, this embodiment also uses the first-order similarity to capture the local structure . The first-order similarity loss function is defined as follows:
一阶相似度损失函数借鉴拉普拉斯映射(LaplacianEigenmap)的思想。LE 通过构建相似关系图重构局部特征结构。如果节点vi和vj有边连接,那么在embedding space中的距离也比较接近。和是自编码器对节点vi和vj的编码结果,LE可以保持一条边的两个节点在嵌入空间中的表示相对接近;在上式中s(vi,vj)是基于元路径利用随机游走得到的节点vi到节点vj的转移概率。The first-order similarity loss function draws on the idea of Laplacian Eigenmap. LE reconstructs the local feature structure by building a similarity graph. If nodes v i and v j are connected by edges, the distances in the embedding space are also relatively close. and is the encoding result of nodes v i and v j by the autoencoder, LE can keep the representation of two nodes of an edge relatively close in the embedding space; in the above formula s(vi , v j ) is based on the meta-path utilization The transition probability from node v i to node v j obtained by random walk.
为了同时保留全局结构和局部结构,本实施例同时优化L1和L2:In order to preserve the global structure and local structure at the same time, this embodiment optimizes L1 and L2 at the same time:
其中,添加Lreg防止过拟合:Among them, adding Lreg prevents overfitting:
使用随机梯度下降算法(ASGD)优化目标函数,目标是最小化也就是需要求偏导和 The objective function is optimized using stochastic gradient descent (ASGD) with the goal of minimizing That is to say, the need for partial guidance and
通过SDNE模型得到每个节点的嵌入向量后,计算向量之间的欧氏距离,根据欧式距离对中药节点和蛋白质节点进行聚类,每个蛋白取欧氏距离最近的前10个中药,分别对三种蛋白组合,即结构蛋白、非结构蛋白及辅助蛋白输出距离最小的中药去重,即得到以结构蛋白质、非结构蛋白质及辅助蛋白质为中心的中药聚类组合,去重后的中药对相应的新冠蛋白可能存在潜在的疗效,聚类结果如表1所示。After obtaining the embedding vector of each node through the SDNE model, calculate the Euclidean distance between the vectors, and cluster the traditional Chinese medicine nodes and protein nodes according to the Euclidean distance. The combination of three proteins, that is, the traditional Chinese medicine with the smallest output distance of structural protein, non-structural protein and auxiliary protein, is deduplicated, that is, the clustering combination of traditional Chinese medicine centered on structural protein, non-structural protein and auxiliary protein is obtained. The new coronavirus protein may have potential therapeutic effects, and the clustering results are shown in Table 1.
本实施例从Herbnet数据库中爬取所有中药的特征,包括药性、归经、功效、主治、中药配伍等,共涉及689个特征,将去重后的所有候选中药与Herbnet 数据库中的中药信息进行匹配,发现除贯叶金丝桃、黄柏之外的21种中药都可以在Herbnet中找到相应的中药信息,则这21种中药视为推荐中药。The present embodiment crawls the characteristics of all traditional Chinese medicines from the Herbnet database, including medicinal properties, meridians, efficacy, indications, compatibility of traditional Chinese medicines, etc., involving a total of 689 characteristics. Matching, it is found that 21 kinds of traditional Chinese medicines except Hypericum perforatum and Phellodendron phellodendri can find corresponding traditional Chinese medicine information in Herbnet, then these 21 kinds of traditional Chinese medicines are regarded as recommended traditional Chinese medicines.
21种推荐中药具有一定的可能性来代替诊疗指南中与其相似的中药,对这 21种中药和中国第八版诊疗指南方案中涉及到的76个中药建立97*689维的0-1 矩阵,如果中药有相应的特征,对应元素记为1,否则为0;然后计算21个推荐中药和诊疗指南中药的余弦相似度,得到指南中药中与推荐中药相似度前10 且用于推荐治疗的潜在中药。可替代治疗新冠的潜在中药如表2所示。The 21 recommended Chinese medicines have a certain possibility to replace the similar Chinese medicines in the diagnosis and treatment guidelines. A 97*689-dimensional 0-1 matrix was established for these 21 Chinese medicines and the 76 Chinese medicines involved in the Chinese eighth edition diagnosis and treatment guidelines. If the traditional Chinese medicine has corresponding characteristics, the corresponding element is recorded as 1, otherwise it is 0; then calculate the cosine similarity of the 21 recommended traditional Chinese medicines and the traditional Chinese medicine in the diagnosis and treatment guide, and obtain the top 10 similarity between the guide traditional Chinese medicine and the recommended traditional Chinese medicine and the potential for the recommended treatment. traditional Chinese medicine. Potential traditional Chinese medicines that can be used as an alternative for the treatment of COVID-19 are shown in Table 2.
表1:基于新冠蛋白的中药聚类结果Table 1: Clustering results of traditional Chinese medicine based on SARS-CoV-2
表2:可替代治疗新冠的潜在中药Table 2: Potential Chinese Medicines for Alternative Treatment of COVID-19
实施例2Example 2
本实施例提供一种基于新冠蛋白质异构网络聚类的中药处方推荐系统,包括:This embodiment provides a traditional Chinese medicine prescription recommendation system based on novel coronavirus protein heterogeneous network clustering, including:
网络构建模块,被配置为构建候选中药与其化合物的边连接,以及化合物与新冠蛋白质靶标的边连接,根据上述边连接构建中药-化合物-蛋白质异构网;The network building block is configured to construct the edge connection between the candidate traditional Chinese medicine and its compound, as well as the edge connection between the compound and the new crown protein target, and construct the traditional Chinese medicine-compound-protein heterogeneous network according to the above-mentioned edge connection;
相似度计算模块,被配置为确定中药-化合物-蛋白质异构网的元路径,计算元路径下异构节点间的相似度,以此得到中药-蛋白质的转移概率矩阵,根据转移概率矩阵得到异构节点的嵌入向量;The similarity calculation module is configured to determine the meta-path of the traditional Chinese medicine-compound-protein heterogeneous network, and calculate the similarity between the heterogeneous nodes under the meta-path, so as to obtain the transition probability matrix of the traditional Chinese medicine-protein, and obtain the heterogeneity according to the transition probability matrix. Embedding vector for constructing nodes;
聚类模块,被配置为根据由嵌入向量计算的中药和蛋白质的欧式距离对蛋白质进行聚类,得到分别以结构蛋白质、非结构蛋白质及辅助蛋白质为中心的中药聚类组合;The clustering module is configured to cluster the proteins according to the Euclidean distance of the traditional Chinese medicine and the protein calculated by the embedding vector, and obtain the traditional Chinese medicine clustering combination centered on the structural protein, the non-structural protein and the auxiliary protein respectively;
推荐模块,被配置为根据中药数据库对中药聚类组合中的候选中药进行筛选,得到推荐中药。The recommendation module is configured to screen the candidate traditional Chinese medicines in the traditional Chinese medicine clustering combination according to the traditional Chinese medicine database to obtain the recommended traditional Chinese medicines.
此处需要说明的是,上述模块对应于实施例1中所述的步骤,上述模块与对应的步骤所实现的示例和应用场景相同,但不限于上述实施例1所公开的内容。需要说明的是,上述模块作为系统的一部分可以在诸如一组计算机可执行指令的计算机系统中执行。It should be noted here that the foregoing modules correspond to the steps described in
在更多实施例中,还提供:In further embodiments, there is also provided:
一种电子设备,包括存储器和处理器以及存储在存储器上并在处理器上运行的计算机指令,所述计算机指令被处理器运行时,完成实施例1中所述的方法。为了简洁,在此不再赘述。An electronic device includes a memory, a processor, and computer instructions stored on the memory and executed on the processor, and when the computer instructions are executed by the processor, the method described in
应理解,本实施例中,处理器可以是中央处理单元CPU,处理器还可以是其他通用处理器、数字信号处理器DSP、专用集成电路ASIC,现成可编程门阵列FPGA或者其他可编程逻辑器件、分立门或者晶体管逻辑器件、分立硬件组件等。通用处理器可以是微处理器或者该处理器也可以是任何常规的处理器等。It should be understood that, in this embodiment, the processor may be a central processing unit CPU, and the processor may also be other general-purpose processors, digital signal processors DSP, application-specific integrated circuits ASIC, off-the-shelf programmable gate array FPGA or other programmable logic devices , discrete gate or transistor logic devices, discrete hardware components, etc. A general purpose processor may be a microprocessor or the processor may be any conventional processor or the like.
存储器可以包括只读存储器和随机存取存储器,并向处理器提供指令和数据、存储器的一部分还可以包括非易失性随机存储器。例如,存储器还可以存储设备类型的信息。The memory may include read-only memory and random access memory and provide instructions and data to the processor, and a portion of the memory may also include non-volatile random access memory. For example, the memory may also store device type information.
一种计算机可读存储介质,用于存储计算机指令,所述计算机指令被处理器执行时,完成实施例1中所述的方法。A computer-readable storage medium for storing computer instructions, when the computer instructions are executed by a processor, the method described in
实施例1中的方法可以直接体现为硬件处理器执行完成,或者用处理器中的硬件及软件模块组合执行完成。软件模块可以位于随机存储器、闪存、只读存储器、可编程只读存储器或者电可擦写可编程存储器、寄存器等本领域成熟的存储介质中。该存储介质位于存储器,处理器读取存储器中的信息,结合其硬件完成上述方法的步骤。为避免重复,这里不再详细描述。The method in
本领域普通技术人员可以意识到,结合本实施例描述的各示例的单元即算法步骤,能够以电子硬件或者计算机软件和电子硬件的结合来实现。这些功能究竟以硬件还是软件方式来执行,取决于技术方案的特定应用和设计约束条件。专业技术人员可以对每个特定的应用来使用不同方法来实现所描述的功能,但是这种实现不应认为超出本申请的范围。Those of ordinary skill in the art can realize that the unit, that is, the algorithm step of each example described in conjunction with this embodiment, can be implemented by electronic hardware or a combination of computer software and electronic hardware. Whether these functions are performed in hardware or software depends on the specific application and design constraints of the technical solution. Skilled artisans may implement the described functionality using different methods for each particular application, but such implementations should not be considered beyond the scope of this application.
以上仅为本发明的优选实施例而已,并不用于限制本发明,对于本领域的技术人员来说,本发明可以有各种更改和变化。凡在本发明的精神和原则之内,所作的任何修改、等同替换、改进等,均应包含在本发明的保护范围之内。The above are only preferred embodiments of the present invention, and are not intended to limit the present invention. For those skilled in the art, the present invention may have various modifications and changes. Any modification, equivalent replacement, improvement, etc. made within the spirit and principle of the present invention shall be included within the protection scope of the present invention.
上述虽然结合附图对本发明的具体实施方式进行了描述,但并非对本发明保护范围的限制,所属领域技术人员应该明白,在本发明的技术方案的基础上,本领域技术人员不需要付出创造性劳动即可做出的各种修改或变形仍在本发明的保护范围以内。Although the specific embodiments of the present invention have been described above in conjunction with the accompanying drawings, they do not limit the scope of protection of the present invention. Those skilled in the art should understand that on the basis of the technical solutions of the present invention, those skilled in the art do not need to pay creative work. Various modifications or deformations that can be made are still within the protection scope of the present invention.
Claims (10)
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202110038417.7A CN112863634B (en) | 2021-01-12 | 2021-01-12 | Traditional Chinese medicine prescription recommendation method and system based on new crown protein heterogeneous network clustering |
Applications Claiming Priority (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CN202110038417.7A CN112863634B (en) | 2021-01-12 | 2021-01-12 | Traditional Chinese medicine prescription recommendation method and system based on new crown protein heterogeneous network clustering |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| CN112863634A true CN112863634A (en) | 2021-05-28 |
| CN112863634B CN112863634B (en) | 2022-09-20 |
Family
ID=76002991
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| CN202110038417.7A Active CN112863634B (en) | 2021-01-12 | 2021-01-12 | Traditional Chinese medicine prescription recommendation method and system based on new crown protein heterogeneous network clustering |
Country Status (1)
| Country | Link |
|---|---|
| CN (1) | CN112863634B (en) |
Cited By (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN114121181A (en) * | 2021-11-12 | 2022-03-01 | 东南大学 | An attention mechanism-based heterogeneous graph neural network target prediction method for traditional Chinese medicine |
| CN114678063A (en) * | 2022-02-24 | 2022-06-28 | 华南理工大学 | A method for predicting drugs that interact with novel coronavirus proteins |
| CN120388610B (en) * | 2025-04-22 | 2026-02-10 | 广东工业大学 | Gene Design and Evolutionary Path Inference Method Based on Adaptive Constrained Deep Model |
Citations (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN106529205A (en) * | 2016-11-03 | 2017-03-22 | 中南大学 | Drug target relation prediction method based on drug substructure and molecule character description information |
| CN109887540A (en) * | 2019-01-15 | 2019-06-14 | 中南大学 | A Drug-Target Interaction Prediction Method Based on Heterogeneous Network Embedding |
| CN111462833A (en) * | 2019-01-20 | 2020-07-28 | 深圳智药信息科技有限公司 | Virtual drug screening method and device, computing equipment and storage medium |
| CN111554360A (en) * | 2020-04-27 | 2020-08-18 | 大连理工大学 | Drug relocation prediction method based on biomedical literature and domain knowledge data |
| CN111785320A (en) * | 2020-06-28 | 2020-10-16 | 西安电子科技大学 | Drug-target interaction prediction method based on multi-layer network representation learning |
| CN111916145A (en) * | 2020-07-24 | 2020-11-10 | 湖南大学 | Novel coronavirus target prediction and drug discovery method based on graph representation learning |
-
2021
- 2021-01-12 CN CN202110038417.7A patent/CN112863634B/en active Active
Patent Citations (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN106529205A (en) * | 2016-11-03 | 2017-03-22 | 中南大学 | Drug target relation prediction method based on drug substructure and molecule character description information |
| CN109887540A (en) * | 2019-01-15 | 2019-06-14 | 中南大学 | A Drug-Target Interaction Prediction Method Based on Heterogeneous Network Embedding |
| CN111462833A (en) * | 2019-01-20 | 2020-07-28 | 深圳智药信息科技有限公司 | Virtual drug screening method and device, computing equipment and storage medium |
| CN111554360A (en) * | 2020-04-27 | 2020-08-18 | 大连理工大学 | Drug relocation prediction method based on biomedical literature and domain knowledge data |
| CN111785320A (en) * | 2020-06-28 | 2020-10-16 | 西安电子科技大学 | Drug-target interaction prediction method based on multi-layer network representation learning |
| CN111916145A (en) * | 2020-07-24 | 2020-11-10 | 湖南大学 | Novel coronavirus target prediction and drug discovery method based on graph representation learning |
Non-Patent Citations (2)
| Title |
|---|
| FEIFEI GUO ET AL.: "Exploration of the mechanism of traditional Chinese medicine by AI approach using unsupervised machine learning for cellular functional similarity of compounds in heterogeneous networks, XiaoErFuPi granules as an example", 《ELSEVIER》 * |
| 徐婷 等: "一种基于异构网络算法的药物-蛋白关联性研究方法", 《智能计算机与应用》 * |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN114121181A (en) * | 2021-11-12 | 2022-03-01 | 东南大学 | An attention mechanism-based heterogeneous graph neural network target prediction method for traditional Chinese medicine |
| CN114121181B (en) * | 2021-11-12 | 2024-03-29 | 东南大学 | Heterogeneous graph neural network traditional Chinese medicine target prediction method based on attention mechanism |
| CN114678063A (en) * | 2022-02-24 | 2022-06-28 | 华南理工大学 | A method for predicting drugs that interact with novel coronavirus proteins |
| CN120388610B (en) * | 2025-04-22 | 2026-02-10 | 广东工业大学 | Gene Design and Evolutionary Path Inference Method Based on Adaptive Constrained Deep Model |
Also Published As
| Publication number | Publication date |
|---|---|
| CN112863634B (en) | 2022-09-20 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| Gu et al. | REDDA: Integrating multiple biological relations to heterogeneous graph neural network for drug-disease association prediction | |
| Meng et al. | Drug repositioning based on similarity constrained probabilistic matrix factorization: COVID-19 as a case study | |
| US20230037376A1 (en) | Generating enhanced graphical user interfaces for presentation of anti-infective design spaces for selecting drug candidates | |
| Berger et al. | Computational biology in the 21st century: Scaling with compressive algorithms | |
| Böckenhauer et al. | Algorithmic aspects of bioinformatics | |
| CN114582508B (en) | A method to predict potentially associated circular RNA-disease pairs based on GCN and ensemble learning | |
| CN112863634B (en) | Traditional Chinese medicine prescription recommendation method and system based on new crown protein heterogeneous network clustering | |
| CN114496084A (en) | Efficient prediction method for association relation between circRNA and miRNA | |
| Mäkinen et al. | Genome-scale algorithm design: bioinformatics in the era of high-throughput sequencing | |
| CN114566210A (en) | Medicine reuse prediction method, device, storage medium and computer equipment | |
| Jacox et al. | Resolution and reconciliation of non-binary gene trees with transfers, duplications and losses | |
| CN116646002A (en) | Multi-non-coding RNA and disease association prediction method, device, equipment and medium | |
| Yang et al. | Signaling repurposable drug combinations against COVID-19 by developing the heterogeneous deep herb-graph method | |
| Ray et al. | Deep variational graph autoencoders for novel host-directed therapy options against COVID-19 | |
| CN114141361A (en) | Traditional Chinese medicine prescription recommendation method based on symptom term mapping and deep learning | |
| CN116884473B (en) | Protein function prediction model generation method and device | |
| Yang et al. | An FM-index based high-throughput memory-efficient FPGA accelerator for paired-end short-read mapping | |
| Malusare et al. | Improving molecule generation and drug discovery with a knowledge-enhanced generative model | |
| Mohanty et al. | A review on planted (l, d) Motif Discovery algorithms for Medical Diagnose | |
| Onokpasa et al. | RNA secondary structures: from ab initio prediction to better compression, and back | |
| CN117059196B (en) | Small molecule drug design method, device, electronic equipment and storage medium | |
| Zhang et al. | DRML-Ensemble: drug repurposing method based on feature construction of multi-layer ensemble | |
| Jurisica et al. | Knowledge discovery in proteomics | |
| He et al. | DRTerHGAT: A drug repurposing method based on the ternary heterogeneous graph attention network | |
| CN114822681A (en) | Virus-drug association prediction method based on recommendation system |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| PB01 | Publication | ||
| PB01 | Publication | ||
| SE01 | Entry into force of request for substantive examination | ||
| SE01 | Entry into force of request for substantive examination | ||
| GR01 | Patent grant | ||
| GR01 | Patent grant |