Journal Description
SynBio
SynBio
is an international, peer-reviewed, open access journal on synthetic biology, biological parts, devices, and systems, published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- Rapid Publication: first decisions in 16 days; acceptance to publication in 5.8 days (median values for MDPI journals in the first half of 2024).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
- SynBio is a companion journal of IJMS.
Latest Articles
Can Methylococcus capsulatus Revolutionize Methane Capture and Utilization for Sustainable Energy Production?
SynBio 2024, 2(3), 311-328; https://doi.org/10.3390/synbio2030019 - 4 Sep 2024
Abstract
►
Show Figures
Methane is the second largest contributor to global warming after carbon dioxide. Once it is released into the atmosphere, methane lingers for over 10 years, during which it traps heat, contributes to the formation of ground-level ozone, and affects air quality adversely. Conversely,
[...] Read more.
Methane is the second largest contributor to global warming after carbon dioxide. Once it is released into the atmosphere, methane lingers for over 10 years, during which it traps heat, contributes to the formation of ground-level ozone, and affects air quality adversely. Conversely, methane has some benefits that could be harnessed to address its impact on the environment while utilizing it for good. Methane’s significant role in global warming and potential for energy production and other beneficial applications necessitate the adoption of innovative solutions to remediate the gas from the atmosphere and harness some of its benefits. This article explores Methylococcus capsulatus, a methanotrophic bacterium, and its potential for revolutionizing sustainable methane capture and utilization. With its unique metabolic abilities, M. capsulatus efficiently oxidizes methane, making it a promising candidate for biotechnological applications. We review current research in its current and potential applications in methane capture and utilization, emphasizing key characteristics, implementation challenges, benefits, and limitations in methane capture and conversion. We also highlight the importance of interdisciplinary collaborations and technological advancements in synthetic biology to maximize its energy production potential. Our article analyzes M. capsulatus’ role in addressing methane-related environmental concerns and advancing sustainable energy solutions.
Full article
Open AccessArticle
The Crystal Structure of Thermal Green Protein Q66E (TGP-E) and Yellow Thermostable Protein (YTP-E) E148D
by
Matthew R. Anderson, Caitlin M. Padgett, Victoria O. Ogbeifun and Natasha M. DeVore
SynBio 2024, 2(3), 298-310; https://doi.org/10.3390/synbio2030018 - 23 Aug 2024
Abstract
►▼
Show Figures
Thermal green protein Q66E (TGP-E) has previously shown increased thermal stability compared to thermal green protein (TGP), a thermal stable fluorescent protein produced through consensus and surface protein engineering. In this paper, we describe the protein crystal structure of TGP-E to 2.0 Å.
[...] Read more.
Thermal green protein Q66E (TGP-E) has previously shown increased thermal stability compared to thermal green protein (TGP), a thermal stable fluorescent protein produced through consensus and surface protein engineering. In this paper, we describe the protein crystal structure of TGP-E to 2.0 Å. This structure reveals alterations in the hydrogen bond network near the chromophore that may result in the observed increase in thermal stability. We compare the very stable TGP-E protein to the structure of a yellow mutant version of this protein YTP-E E148D. The structure of this mutant protein reveals the rationale for the observed low quantum yield and directions for future protein engineering efforts.
Full article

Graphical abstract
Graphical abstract
Full article ">Figure 1
<p>Absorbance of TGP-E (green dashed) and YTP-E E148D (yellow dashed) normalized to the highest absorbance after 280 nm. The emission spectra of TGP-E (green solid) and YTP-E E148D (yellow solid) with excitation normalized to the highest fluorescent value.</p> Full article ">Figure 2
<p>(<b>A</b>) pH stability of YTP-E E148D plotted as percent fluorescence. The sample with the highest fluorescence (pH 10) was set to 100% fluorescence. This data was performed as duplicate trials in triplicate. (<b>B</b>) The chemical stability of YTP-E E148D was assayed as % fluorescence in increasing concentrations of guanidine hydrochloride. One-hundred percent fluorescence was set to the fluorescence with 0 M guanidine hydrochloride present. (<b>C</b>) The thermostability was measured using a real-time PCR instrument. Three rounds of temperature increase to 100 °C followed by rapid temperature return to 25 °C were repeated. Temperature is plotted on the right axis with a grey line. Each cycle of temperature increases to 100 °C followed by a decrease to 25 °C is plotted in a different colour (cycle 1 is red, cycle 2 is orange, and cycle 3 is green). The left axis measures the percent fluorescence with the first cycle percent fluorescence at 25 °C prior to heating set to 100%. This assay was performed in quadruplicate.</p> Full article ">Figure 3
<p>(<b>A</b>) TGP-E global structure alignment of molecule A (pink) and molecule B (green). (<b>B</b>) Chromophore and S146 and L213 electron density shown as blue netting (2mFo-DFc contoured at 1σ) of TGP-E molecule B.</p> Full article ">Figure 4
<p>(<b>A</b>) Overlay of TGP-E molecule A (pink), molecule B (green), and TGP molecule A (purple: PDB: 4TZA) with the chromophore hydrogen bond network shown near the chromophore phenol. (<b>B</b>) Overlay of TGP-E molecule A (pink), molecule B (green), and TGP molecule A (purple: PDB: 4TZA) with the chromophore hydrogen bond network shown near the altered portion of the chromophore. For both A and B, hydrogen bonds with differing lengths are coloured orange (TGP-E) and yellow (TGP), respectively. Hydrogen bonds with the same length in all molecules are coloured black. Key residues interacting with the chromophore are shown as sticks and labelled.</p> Full article ">Figure 5
<p>(<b>A</b>) Chromophore of molecule A YTPE E148D with electron density (2mFo-DFc contoured at 1σ) (<b>B</b>) Chromophore of molecule B YTPE E148D with electron density (2mFo-DFc contoured at 1σ) (<b>C</b>) Chromophore of molecule C YTPE E148D with electron density (2mFo-DFc contoured at 1σ) (<b>D</b>) Chromophore of molecule D YTPE E148D with electron density (2mFo-DFc contoured at 1σ).</p> Full article ">Figure 6
<p>An overlay of YTP-E E148D molecules A (cyan), B (salmon), C (yellow), and D (pink). Hydrogen bond interactions and distances to nearby residues are labelled with the same colour as the corresponding molecule.</p> Full article ">Figure 7
<p>(<b>A</b>) An overlay of YTP-E E148D molecule A (cyan) with TGP-E (green). Hydrogen bond distances are coloured to match the molecule colour. The water molecules shown are from TGP-E. (<b>B</b>) An overlay of YTP-E E148D molecule A (cyan) with YTP (RCSB PDB 1YFP) (orange). Hydrogen bond distances and residue names are coloured to match the molecule colour.</p> Full article ">
Full article ">Figure 1
<p>Absorbance of TGP-E (green dashed) and YTP-E E148D (yellow dashed) normalized to the highest absorbance after 280 nm. The emission spectra of TGP-E (green solid) and YTP-E E148D (yellow solid) with excitation normalized to the highest fluorescent value.</p> Full article ">Figure 2
<p>(<b>A</b>) pH stability of YTP-E E148D plotted as percent fluorescence. The sample with the highest fluorescence (pH 10) was set to 100% fluorescence. This data was performed as duplicate trials in triplicate. (<b>B</b>) The chemical stability of YTP-E E148D was assayed as % fluorescence in increasing concentrations of guanidine hydrochloride. One-hundred percent fluorescence was set to the fluorescence with 0 M guanidine hydrochloride present. (<b>C</b>) The thermostability was measured using a real-time PCR instrument. Three rounds of temperature increase to 100 °C followed by rapid temperature return to 25 °C were repeated. Temperature is plotted on the right axis with a grey line. Each cycle of temperature increases to 100 °C followed by a decrease to 25 °C is plotted in a different colour (cycle 1 is red, cycle 2 is orange, and cycle 3 is green). The left axis measures the percent fluorescence with the first cycle percent fluorescence at 25 °C prior to heating set to 100%. This assay was performed in quadruplicate.</p> Full article ">Figure 3
<p>(<b>A</b>) TGP-E global structure alignment of molecule A (pink) and molecule B (green). (<b>B</b>) Chromophore and S146 and L213 electron density shown as blue netting (2mFo-DFc contoured at 1σ) of TGP-E molecule B.</p> Full article ">Figure 4
<p>(<b>A</b>) Overlay of TGP-E molecule A (pink), molecule B (green), and TGP molecule A (purple: PDB: 4TZA) with the chromophore hydrogen bond network shown near the chromophore phenol. (<b>B</b>) Overlay of TGP-E molecule A (pink), molecule B (green), and TGP molecule A (purple: PDB: 4TZA) with the chromophore hydrogen bond network shown near the altered portion of the chromophore. For both A and B, hydrogen bonds with differing lengths are coloured orange (TGP-E) and yellow (TGP), respectively. Hydrogen bonds with the same length in all molecules are coloured black. Key residues interacting with the chromophore are shown as sticks and labelled.</p> Full article ">Figure 5
<p>(<b>A</b>) Chromophore of molecule A YTPE E148D with electron density (2mFo-DFc contoured at 1σ) (<b>B</b>) Chromophore of molecule B YTPE E148D with electron density (2mFo-DFc contoured at 1σ) (<b>C</b>) Chromophore of molecule C YTPE E148D with electron density (2mFo-DFc contoured at 1σ) (<b>D</b>) Chromophore of molecule D YTPE E148D with electron density (2mFo-DFc contoured at 1σ).</p> Full article ">Figure 6
<p>An overlay of YTP-E E148D molecules A (cyan), B (salmon), C (yellow), and D (pink). Hydrogen bond interactions and distances to nearby residues are labelled with the same colour as the corresponding molecule.</p> Full article ">Figure 7
<p>(<b>A</b>) An overlay of YTP-E E148D molecule A (cyan) with TGP-E (green). Hydrogen bond distances are coloured to match the molecule colour. The water molecules shown are from TGP-E. (<b>B</b>) An overlay of YTP-E E148D molecule A (cyan) with YTP (RCSB PDB 1YFP) (orange). Hydrogen bond distances and residue names are coloured to match the molecule colour.</p> Full article ">
Open AccessArticle
The Natural Evolution of RNA Viruses Provides Important Clues about the Origin of SARS-CoV-2 Variants
by
Hiroshi Arakawa
SynBio 2024, 2(3), 285-297; https://doi.org/10.3390/synbio2030017 - 16 Aug 2024
Abstract
►▼
Show Figures
Despite the recent pandemic, the origin of its causative agent, SARS-CoV-2, remains controversial. This study identifies several prototype SARS-CoV-2 variants (proto-variants) that are descendants of the Wuhan variant. A thorough evaluation of the evolutionary histories of the genomes of these proto-variants reveals that
[...] Read more.
Despite the recent pandemic, the origin of its causative agent, SARS-CoV-2, remains controversial. This study identifies several prototype SARS-CoV-2 variants (proto-variants) that are descendants of the Wuhan variant. A thorough evaluation of the evolutionary histories of the genomes of these proto-variants reveals that most mutations in proto-variants were biased toward mutations that change the amino acid sequence. While these nonsynonymous substitutions (N mutations) were common in SARS-CoV-2 proto-variants, nucleotide changes that do not result in an amino acid change, termed synonymous substitutions (S mutations), dominate the mutations found in other RNA viruses. The N mutation bias in the SARS-CoV2 proto-variants was found in the spike gene as well as several other genes. The analysis of the ratio of N to S mutations in general RNA viruses revealed that the probability that an RNA virus spontaneously evolves a proto-variant is between 1.5 × 10−9 and 2.7 × 10−26 under natural conditions. These results suggest that SARS-CoV-2 variants did not emerge via a canonical route.
Full article

Graphical abstract
Graphical abstract
Full article ">Figure 1
<p>(<b>A</b>) Phylogenetic tree. The ancestral type of each SARS-CoV-2 variant was named its “proto-variant”. The evolutionary relationships among proto-variants are expanded below. The scale shows their evolutionary distance. (<b>B</b>) Percentage differences among proto-variants, RaTG13, and SARS-CoV-1. Differences among proto-variants are highlighted in orange.</p> Full article ">Figure 2
<p>N and S mutations. (<b>A</b>) N and S mutations of each gene among SARS-CoV-1, RaTG13, and proto-Wuhan. (<b>B</b>) Those from proto-Wuhan to the different proto-variants.</p> Full article ">Figure 3
<p>dN and dS (<b>A</b>) dN and dS in each gene of SARS-CoV-1, RaTG13, and proto-Wuhan. (<b>B</b>) dN and dS from proto-Wuhan to the respective proto-variants. dN is the N mutations per N sites, and dS is the S mutations per S sites. When there is no selective pressure and mutations are introduced randomly, the ratio of dN to dS is 1.</p> Full article ">Figure 4
<p>dN/dS values of proto-variants and various viruses. (<b>A</b>) dN/dS of mammals and various viruses. Each dot indicates the dN/dS of the entire coding region of each genome. (<b>B</b>) dN/dS of SARS-CoV-1 vs. RaTG13 vs. proto-Wuhan (left). dN/dS of proto-Wuhan vs. proto-variants (right). (<b>C</b>) dN/dS of each gene of RNA viruses and retroviruses. Each dot indicates the dN/dS of the coding region of the respective gene. (<b>D</b>) dN/dS of each gene in SARS-CoV-1 vs. RaTG13 vs. proto-Wuhan. Data on mammals are from Nikolaev et al. [<a href="#B19-synbio-02-00017" class="html-bibr">19</a>], data on non-coronaviruses are from Lin et al. [<a href="#B20-synbio-02-00017" class="html-bibr">20</a>], and data on coronaviruses are from Kistler et al. [<a href="#B21-synbio-02-00017" class="html-bibr">21</a>].</p> Full article ">Figure 5
<p>The probability of the natural emergence of proto-variants expected from the dN/dS ratios of general RNA viruses. (<b>A</b>) N and S mutation ratios of proto-variants expected from general RNA viruses. By transforming the dN/dS equation, the ratio of N mutations/S mutations can be expressed as a multiplication of dN/dS and N sites/S sites. (<b>B</b>) The probability of the natural emergence of proto-variants and their genes. The number of N and S mutations (in brackets) and their probabilities are shown below the genes. Supposing the N mutation ratio is 15.8% and the number of N and S mutations are “n” and “s”, the probability “P” can be calculated based on the Bernoulli trial via the binomial distribution formula: P = <sub>n+s</sub>C<sub>n</sub> 0.158<sup>n</sup> × 0.842<sup>s</sup>. Here, <sub>n+s</sub>C<sub>n</sub> means (n+s)!/(n! × s!). Only genes with probabilities less than 1% were colored in red. The heatmap’s color intensity indicates the probability; probabilities less than one in a million were considered almost impossible and highlighted in deep red.</p> Full article ">
Full article ">Figure 1
<p>(<b>A</b>) Phylogenetic tree. The ancestral type of each SARS-CoV-2 variant was named its “proto-variant”. The evolutionary relationships among proto-variants are expanded below. The scale shows their evolutionary distance. (<b>B</b>) Percentage differences among proto-variants, RaTG13, and SARS-CoV-1. Differences among proto-variants are highlighted in orange.</p> Full article ">Figure 2
<p>N and S mutations. (<b>A</b>) N and S mutations of each gene among SARS-CoV-1, RaTG13, and proto-Wuhan. (<b>B</b>) Those from proto-Wuhan to the different proto-variants.</p> Full article ">Figure 3
<p>dN and dS (<b>A</b>) dN and dS in each gene of SARS-CoV-1, RaTG13, and proto-Wuhan. (<b>B</b>) dN and dS from proto-Wuhan to the respective proto-variants. dN is the N mutations per N sites, and dS is the S mutations per S sites. When there is no selective pressure and mutations are introduced randomly, the ratio of dN to dS is 1.</p> Full article ">Figure 4
<p>dN/dS values of proto-variants and various viruses. (<b>A</b>) dN/dS of mammals and various viruses. Each dot indicates the dN/dS of the entire coding region of each genome. (<b>B</b>) dN/dS of SARS-CoV-1 vs. RaTG13 vs. proto-Wuhan (left). dN/dS of proto-Wuhan vs. proto-variants (right). (<b>C</b>) dN/dS of each gene of RNA viruses and retroviruses. Each dot indicates the dN/dS of the coding region of the respective gene. (<b>D</b>) dN/dS of each gene in SARS-CoV-1 vs. RaTG13 vs. proto-Wuhan. Data on mammals are from Nikolaev et al. [<a href="#B19-synbio-02-00017" class="html-bibr">19</a>], data on non-coronaviruses are from Lin et al. [<a href="#B20-synbio-02-00017" class="html-bibr">20</a>], and data on coronaviruses are from Kistler et al. [<a href="#B21-synbio-02-00017" class="html-bibr">21</a>].</p> Full article ">Figure 5
<p>The probability of the natural emergence of proto-variants expected from the dN/dS ratios of general RNA viruses. (<b>A</b>) N and S mutation ratios of proto-variants expected from general RNA viruses. By transforming the dN/dS equation, the ratio of N mutations/S mutations can be expressed as a multiplication of dN/dS and N sites/S sites. (<b>B</b>) The probability of the natural emergence of proto-variants and their genes. The number of N and S mutations (in brackets) and their probabilities are shown below the genes. Supposing the N mutation ratio is 15.8% and the number of N and S mutations are “n” and “s”, the probability “P” can be calculated based on the Bernoulli trial via the binomial distribution formula: P = <sub>n+s</sub>C<sub>n</sub> 0.158<sup>n</sup> × 0.842<sup>s</sup>. Here, <sub>n+s</sub>C<sub>n</sub> means (n+s)!/(n! × s!). Only genes with probabilities less than 1% were colored in red. The heatmap’s color intensity indicates the probability; probabilities less than one in a million were considered almost impossible and highlighted in deep red.</p> Full article ">
Open AccessReview
Tropical Fruit Virus Resistance in the Era of Next-Generation Plant Breeding
by
Marcella Silva Vieira, Rafael Lara Rezende Cabral, Luíza Favaratto, Laiane Silva Maciel, André da Silva Xavier, Francisco Murilo Zerbini and Patricia M. B. Fernandes
SynBio 2024, 2(3), 267-284; https://doi.org/10.3390/synbio2030016 - 8 Jul 2024
Cited by 1
Abstract
►▼
Show Figures
Plant viral diseases constitute a major contributor to agricultural production losses, significantly impacting the economies of exporting countries by more than USD 30 billion annually. Understanding and researching the biology and genomics of viruses is crucial for developing virus-resistant genetically edited or genetically
[...] Read more.
Plant viral diseases constitute a major contributor to agricultural production losses, significantly impacting the economies of exporting countries by more than USD 30 billion annually. Understanding and researching the biology and genomics of viruses is crucial for developing virus-resistant genetically edited or genetically modified plants. Genetic modifications can be targeted to specific regions within genes of target plants which are important or essential for the virus to establish a systemic infection, thus fostering resistance or enabling plants to effectively respond to invading agents while preserving their yield. This review provides an overview of viral incidence and diversity in tropical fruit crops and aims to examine the current state of the knowledge on recent research efforts aimed at reducing or eliminating the damage caused by viral diseases, with emphasis on genetically edited products that have reached the market in recent years.
Full article

Figure 1
Figure 1
<p>Evolution of the strategies for conferring resistance to viruses in plants. The scheme illustrates the two main landscapes in plant breeding to confer resistance to viral infections in plants (non–precision and precision breeding), considering the advances with the new breeding technologies (NBTs) and some particularities between strategies. (<b>A</b>) Classic breeding methods have, as their source of antiviral resistance, plant genotypes that contain canonical or similar resistance (R) genes, which through targeted crossings/backcrosses are introduced into productive, but susceptible, elite genotypes. (<b>B</b>) With the advent of genetic engineering, the prospects for obtaining virus–resistant plants have expanded, as even genomic sequences from the virus itself can be used to achieve the pathogen–derived resistance phenotype (PDR) or genes derived from heterologous organisms (HoDR), which may not be sexually compatible with the recipient plant. Both RNA interference (RNAi) using constructs with viral reverse complementary (RC) sequences to generate hairpin RNA and heterologous expression using coding sequences (CDS) from distinct sources are efficient and precise strategies that have been employed to obtain virus–resistant GM crops. (<b>C</b>) NBTs allow for an unprecedented level of precision in breeding, with the specific editing of regulatory and/or gene–coding sequences to obtain genome-edited (GEd) crops. Among the most widespread NBTs, CRISPR/Cas technology enables precise modification of the plant genome to introduce resistance against specific viral infections. Created with <a href="http://BioRender.com" target="_blank">BioRender.com</a>.</p> Full article ">
<p>Evolution of the strategies for conferring resistance to viruses in plants. The scheme illustrates the two main landscapes in plant breeding to confer resistance to viral infections in plants (non–precision and precision breeding), considering the advances with the new breeding technologies (NBTs) and some particularities between strategies. (<b>A</b>) Classic breeding methods have, as their source of antiviral resistance, plant genotypes that contain canonical or similar resistance (R) genes, which through targeted crossings/backcrosses are introduced into productive, but susceptible, elite genotypes. (<b>B</b>) With the advent of genetic engineering, the prospects for obtaining virus–resistant plants have expanded, as even genomic sequences from the virus itself can be used to achieve the pathogen–derived resistance phenotype (PDR) or genes derived from heterologous organisms (HoDR), which may not be sexually compatible with the recipient plant. Both RNA interference (RNAi) using constructs with viral reverse complementary (RC) sequences to generate hairpin RNA and heterologous expression using coding sequences (CDS) from distinct sources are efficient and precise strategies that have been employed to obtain virus–resistant GM crops. (<b>C</b>) NBTs allow for an unprecedented level of precision in breeding, with the specific editing of regulatory and/or gene–coding sequences to obtain genome-edited (GEd) crops. Among the most widespread NBTs, CRISPR/Cas technology enables precise modification of the plant genome to introduce resistance against specific viral infections. Created with <a href="http://BioRender.com" target="_blank">BioRender.com</a>.</p> Full article ">
Open AccessArticle
Efficient Stereoselective Biotransformation of Prochiral Carbonyls by Endophytic Fungi from Handroanthus impetiginosus
by
Valmore Henrique Pereira dos Santos, Monielly Vasconcellos Pereira de Souza, Maurício Moraes Victor, Valéria Belli Riatto and Eliane Oliveira Silva
SynBio 2024, 2(3), 254-266; https://doi.org/10.3390/synbio2030015 - 5 Jul 2024
Abstract
►▼
Show Figures
Endophytic microorganisms are promising sources for new biocatalysts as they must deal with their host plants’ chemicals by developing adaptative strategies, such as enzymatic pathways. As part of our efforts in selecting endophytic strains as biocatalysts, this study describes the screening of endophytic
[...] Read more.
Endophytic microorganisms are promising sources for new biocatalysts as they must deal with their host plants’ chemicals by developing adaptative strategies, such as enzymatic pathways. As part of our efforts in selecting endophytic strains as biocatalysts, this study describes the screening of endophytic fungi isolated from Handroanthus impetiginosus leaves for selective bioreduction of Acetophenone. The bioreductions were monitored by chiral gas chromatography and conducted to the selection of the endophyte Talaromyces sp. H4 as capable of reducing acetophenone to (S)-1-phenylethanol in excellent conversion and enantiomeric excess rates. The influence of seven parameters on the stereoselective bioreduction of acetophenone by Talaromyces sp. H4 was studied: reaction time, inoculum charge, shaking, pH, temperature, substrate concentration, and co-solvent. The optimal conditions were then used to reduce substituted acetophenones and Acetophenone scale-up, which furnished (S)-1-Phenylethanol in 73% yield and 96% ee. The results highlight the endophytic fungus Talaromyces sp. H4 as an excellent biocatalyst for stereoselective reduction of prochiral carbonyls.
Full article

Figure 1
Figure 1
<p>Chiral GC-FID chromatogram of bioreduction extract of acetophenone (<b>1</b>) by <span class="html-italic">Talaromyces</span> sp. H4. GC Rt <span class="html-italic">(R)-1-Phenyl ethanol</span> (<span class="html-italic">R</span>-<b>1a</b>): 18.319 min, and Rt <span class="html-italic">(S)-1-Phenyl ethanol</span> (<span class="html-italic">S</span>-<b>1a</b>): 18.648 min.</p> Full article ">Figure 2
<p>Comparative stereochemistry of alcohol dehydrogenases for producing Prelog and <span class="html-italic">anti</span>-Prelog chiral alcohols.</p> Full article ">
<p>Chiral GC-FID chromatogram of bioreduction extract of acetophenone (<b>1</b>) by <span class="html-italic">Talaromyces</span> sp. H4. GC Rt <span class="html-italic">(R)-1-Phenyl ethanol</span> (<span class="html-italic">R</span>-<b>1a</b>): 18.319 min, and Rt <span class="html-italic">(S)-1-Phenyl ethanol</span> (<span class="html-italic">S</span>-<b>1a</b>): 18.648 min.</p> Full article ">Figure 2
<p>Comparative stereochemistry of alcohol dehydrogenases for producing Prelog and <span class="html-italic">anti</span>-Prelog chiral alcohols.</p> Full article ">
Open AccessArticle
Metformin Lowers Plasma Triacylglycerol Levels in Mice with Impaired Carnitine Biosynthesis and Fatty Liver
by
Bodil Bjørndal, Tra-My Thi Le, Elin Strand, Lise Madsen and Rolf K. Berge
SynBio 2024, 2(3), 240-253; https://doi.org/10.3390/synbio2030014 - 3 Jul 2024
Abstract
The antidiabetic drug metformin has a wide range of metabolic effects and may also reduce the risk of obesity-related diseases. The aim of the current study was to investigate if metformin could counteract meldonium-induced fatty liver. Four groups of male C57BL/6J mice were
[...] Read more.
The antidiabetic drug metformin has a wide range of metabolic effects and may also reduce the risk of obesity-related diseases. The aim of the current study was to investigate if metformin could counteract meldonium-induced fatty liver. Four groups of male C57BL/6J mice were fed a low-fat control diet, or low-fat diets supplemented with metformin, meldonium, or metformin and meldonium for three weeks. Meldonium treatment led to 5.2-fold higher hepatic triacylglycerol (TAG) levels compared to control, and metformin lowered the meldonium-induced lipid accumulation insignificantly by 21%. Mice treated with metformin and meldonium demonstrated significantly lower weight gain, visceral adipose tissue weight and plasma levels of TAG compared to meldonium alone. The hepatic mRNA level of carnitine palmitoyl transferase 1 was increased 2-fold with combined meldonium and metformin treatment compared to meldonium treatment (p < 0.001). Increased hepatic expression of genes involved in fatty acid oxidation and lipid transport was observed in the combination group compared to control, and increased gene expression of the mitochondrial uncoupling protein UCP2 was observed compared to the meldonium group. In addition, the product of fatty acid oxidation, acetylcarnitine, increased in plasma in metformin-treated mice. Altogether, metformin treatment influenced hepatic lipid metabolism and lowered plasma TAG in meldonium-induced fatty liver in mice.
Full article
(This article belongs to the Special Issue Feature Paper Collection in Synthetic Biology)
►▼
Show Figures

Figure 1
Figure 1
<p>Hepatic lipid levels in low-fat-diet-fed mice (control) and mice treated with metformin, meldonium or meldonium and metformin for 21 days. (<b>A</b>) Total liver lipids, (<b>B</b>) triacylglycerol (TAG), (<b>C</b>) Cholesterol, (<b>D</b>) Phospholipids (PL). Mean values with standard deviation are shown (<span class="html-italic">n</span> = 8). Statistical difference between mean values was determined by one-way ANOVA and Tukey’s post hoc test, and dissimilar letters indicate statistically significant mean difference between groups (<span class="html-italic">p</span> < 0.05).</p> Full article ">Figure 2
<p>Hepatic lipid droplets in low-fat-diet-fed mice (control) and mice treated with metformin, meldonium or meldonium and metformin for 21 days. Representative images of liver cryo-sections stained with Oil Red O and hematoxylin and captured at 40× magnification using a light microscope (<span class="html-italic">n</span> = 3–4). Scale bar indicates 100 μm.</p> Full article ">Figure 3
<p>Plasma levels of L-carnitine precursors, L-carnitine and acylcarnitines in low-fat-diet-fed mice (control) and mice treated with metformin, meldonium or meldonium and metformin for 21 days. (<b>A</b>) Trimethyllysine, (<b>B</b>) g-butyrobetaine, (<b>C</b>) free L-carnitine, (<b>D</b>) palmitoylcarnitine and (<b>E</b>) acetylcarnitine. Mean values with standard deviation are shown (<span class="html-italic">n</span> = 8–10). Statistical difference between mean values was determined by one-way ANOVA and Tukey’s post hoc test, and dissimilar letters indicate statistically significant mean difference between groups (<span class="html-italic">p</span> < 0.05).</p> Full article ">Figure 4
<p>Plasma lipid levels in low-fat-diet-fed mice (control) and mice treated with metformin, meldonium or meldonium and metformin for 21 days. (<b>A</b>) Triacylglycerol (TAG), (<b>B</b>) total cholesterol, (<b>C</b>) phospholipid (PL) and (<b>D</b>) non-esterified fatty acids (NEFAs). Mean values with standard deviation are shown (<span class="html-italic">n</span> = 8–10). Statistical difference between mean values was determined by one-way ANOVA and Tukey’s post hoc test, and dissimilar letters indicate statistically significant mean difference between groups (<span class="html-italic">p</span> < 0.05).</p> Full article ">Figure 5
<p>Hepatic enzyme activities in control low-fat-diet-fed C57BL/6 mice, and mice treated with metformin, meldonium or meldonium and metformin for 21 days. (<b>A</b>) β-oxidation of palmitoyl-coenzyme A (CoA) analyzed in fresh liver homogenates. (<b>B</b>) ACOX1 activity, (<b>C</b>) fatty acid synthase (FAS) activity and (<b>D</b>) GPAT activity analyzed in frozen liver samples. Mean values with standard deviation are shown (<span class="html-italic">n</span> = 6). Statistical difference between mean values was determined by one-way ANOVA and Tukey’s post hoc test, and dissimilar letters indicate statistically significant mean difference between groups (<span class="html-italic">p</span> < 0.05).</p> Full article ">
<p>Hepatic lipid levels in low-fat-diet-fed mice (control) and mice treated with metformin, meldonium or meldonium and metformin for 21 days. (<b>A</b>) Total liver lipids, (<b>B</b>) triacylglycerol (TAG), (<b>C</b>) Cholesterol, (<b>D</b>) Phospholipids (PL). Mean values with standard deviation are shown (<span class="html-italic">n</span> = 8). Statistical difference between mean values was determined by one-way ANOVA and Tukey’s post hoc test, and dissimilar letters indicate statistically significant mean difference between groups (<span class="html-italic">p</span> < 0.05).</p> Full article ">Figure 2
<p>Hepatic lipid droplets in low-fat-diet-fed mice (control) and mice treated with metformin, meldonium or meldonium and metformin for 21 days. Representative images of liver cryo-sections stained with Oil Red O and hematoxylin and captured at 40× magnification using a light microscope (<span class="html-italic">n</span> = 3–4). Scale bar indicates 100 μm.</p> Full article ">Figure 3
<p>Plasma levels of L-carnitine precursors, L-carnitine and acylcarnitines in low-fat-diet-fed mice (control) and mice treated with metformin, meldonium or meldonium and metformin for 21 days. (<b>A</b>) Trimethyllysine, (<b>B</b>) g-butyrobetaine, (<b>C</b>) free L-carnitine, (<b>D</b>) palmitoylcarnitine and (<b>E</b>) acetylcarnitine. Mean values with standard deviation are shown (<span class="html-italic">n</span> = 8–10). Statistical difference between mean values was determined by one-way ANOVA and Tukey’s post hoc test, and dissimilar letters indicate statistically significant mean difference between groups (<span class="html-italic">p</span> < 0.05).</p> Full article ">Figure 4
<p>Plasma lipid levels in low-fat-diet-fed mice (control) and mice treated with metformin, meldonium or meldonium and metformin for 21 days. (<b>A</b>) Triacylglycerol (TAG), (<b>B</b>) total cholesterol, (<b>C</b>) phospholipid (PL) and (<b>D</b>) non-esterified fatty acids (NEFAs). Mean values with standard deviation are shown (<span class="html-italic">n</span> = 8–10). Statistical difference between mean values was determined by one-way ANOVA and Tukey’s post hoc test, and dissimilar letters indicate statistically significant mean difference between groups (<span class="html-italic">p</span> < 0.05).</p> Full article ">Figure 5
<p>Hepatic enzyme activities in control low-fat-diet-fed C57BL/6 mice, and mice treated with metformin, meldonium or meldonium and metformin for 21 days. (<b>A</b>) β-oxidation of palmitoyl-coenzyme A (CoA) analyzed in fresh liver homogenates. (<b>B</b>) ACOX1 activity, (<b>C</b>) fatty acid synthase (FAS) activity and (<b>D</b>) GPAT activity analyzed in frozen liver samples. Mean values with standard deviation are shown (<span class="html-italic">n</span> = 6). Statistical difference between mean values was determined by one-way ANOVA and Tukey’s post hoc test, and dissimilar letters indicate statistically significant mean difference between groups (<span class="html-italic">p</span> < 0.05).</p> Full article ">
Open AccessReview
Recombinant Protein Expression and Its Biotechnological Applications in Chlorella spp.
by
Chuchi Chen and Valerie C. A. Ward
SynBio 2024, 2(2), 223-239; https://doi.org/10.3390/synbio2020013 - 6 Jun 2024
Abstract
Recombinant protein expression is a fundamental aspect of both synthetic biology and biotechnology as well as a field unto itself. Microalgae, with their eukaryotic cellular machinery, high lipid content, cost-effective cultivation conditions, safety profile for human consumption, and environmentally friendly attributes, are a
[...] Read more.
Recombinant protein expression is a fundamental aspect of both synthetic biology and biotechnology as well as a field unto itself. Microalgae, with their eukaryotic cellular machinery, high lipid content, cost-effective cultivation conditions, safety profile for human consumption, and environmentally friendly attributes, are a promising system for protein expression or metabolic engineering for sustainable chemical production. Amongst the incredible diversity of microalgae species, Chlorella spp. are heavily studied due to their high growth efficiency, potential for low-cost cultivation, and well-characterized scale-up process for large-scale cultivation. This review aims to comprehensively examine the ongoing advancements in the bioengineering of Chlorella spp. for recombinant protein production and its biotechnological applications. This includes genetic elements such as promoters, terminators, reporters and markers, enhancers, and tags successfully used in Chlorella spp.
Full article
(This article belongs to the Special Issue Feature Paper Collection in Synthetic Biology)
►▼
Show Figures

Figure 1
Figure 1
<p>Cumulative tally of studies expressing recombinant proteins using <span class="html-italic">Chlorella</span> sp.</p> Full article ">Figure 2
<p>Comparison of transformation methods used for <span class="html-italic">Chlorella</span> spp. genome engineering. (<b>Left</b>) Agrobacterium-mediated transformation randomly integrates the T-DNA into the host nuclear genome. (<b>Center</b>) EP, PEG-mediated transformation, and MPB can be used for random integration of DNA into the nuclear genome, and in the case of MPB, into the chloroplast genome. (<b>Right</b>) Site-directed methods for both nuclear and chloroplast transformation have recently been developed. Nuclear transformation into the nitrate reductase locus allows for selection of auxotrophic transformants.</p> Full article ">Figure 3
<p>Organization of genetic elements for expression in <span class="html-italic">Chlorella</span> spp. The promoter (inducible or constitutive) is placed upstream of an optional 5′UTR and transit peptide. This is followed by the coding sequence, which may be interrupted by an intron to enhance expression. A 2A tag can be used for multigene expression using a single promoter. The terminator containing the polyA signal is needed for mRNA transport to the ER for translation.</p> Full article ">
<p>Cumulative tally of studies expressing recombinant proteins using <span class="html-italic">Chlorella</span> sp.</p> Full article ">Figure 2
<p>Comparison of transformation methods used for <span class="html-italic">Chlorella</span> spp. genome engineering. (<b>Left</b>) Agrobacterium-mediated transformation randomly integrates the T-DNA into the host nuclear genome. (<b>Center</b>) EP, PEG-mediated transformation, and MPB can be used for random integration of DNA into the nuclear genome, and in the case of MPB, into the chloroplast genome. (<b>Right</b>) Site-directed methods for both nuclear and chloroplast transformation have recently been developed. Nuclear transformation into the nitrate reductase locus allows for selection of auxotrophic transformants.</p> Full article ">Figure 3
<p>Organization of genetic elements for expression in <span class="html-italic">Chlorella</span> spp. The promoter (inducible or constitutive) is placed upstream of an optional 5′UTR and transit peptide. This is followed by the coding sequence, which may be interrupted by an intron to enhance expression. A 2A tag can be used for multigene expression using a single promoter. The terminator containing the polyA signal is needed for mRNA transport to the ER for translation.</p> Full article ">
Open AccessArticle
Structural Evolution of the Pharmaceutical Peptide Octreotide upon Controlled Relative Humidity and Temperature Variation
by
Maria Athanasiadou, Christina Papaefthymiou, Angelos Kontarinis, Maria Spiliopoulou, Dimitrios Koutoulas, Marios Konstantopoulos, Stamatina Kafetzi, Kleomenis Barlos, Kostas K. Barlos, Natalia Dadivanyan, Detlef Beckers, Thomas Degen, Andrew N. Fitch and Irene Margiolaki
SynBio 2024, 2(2), 205-222; https://doi.org/10.3390/synbio2020012 - 4 Jun 2024
Cited by 1
Abstract
►▼
Show Figures
Octreotide is the first synthetic peptide hormone, consisting of eight amino acids, that mimics the activity of somatostatin, a natural hormone in the body. During the past decades, advanced instrumentation and crystallographic software have established X-Ray Powder Diffraction (XRPD) as a valuable tool
[...] Read more.
Octreotide is the first synthetic peptide hormone, consisting of eight amino acids, that mimics the activity of somatostatin, a natural hormone in the body. During the past decades, advanced instrumentation and crystallographic software have established X-Ray Powder Diffraction (XRPD) as a valuable tool for extracting structural information from biological macromolecules. The latter was demonstrated by the successful structural determination of octreotide at a remarkably high d-spacing resolution (1.87 Å) (PDB code: 6vc1). This study focuses on the response of octreotide to different humidity levels and temperatures, with a particular focus on the stability of the polycrystalline sample. XRPD measurements were accomplished employing an Anton Paar MHC-trans humidity-temperature chamber installed within a laboratory X’Pert Pro diffractometer (Malvern Panalytical). The chamber is employed to control and maintain precise humidity and temperature levels of samples during XRPD data collection. Pawley analysis of the collected data sets revealed that the octreotide polycrystalline sample is remarkably stable, and no structural transitions were observed. The compound retains its orthorhombic symmetry (space group: P212121, a = 18.57744(4) Å, b = 30.17338(6) Å, c = 39.70590(9) Å, d ~ 2.35 Å). However, a characteristic structural evolution in terms of lattice parameters and volume of the unit cell is reported mainly upon controlled relative humidity variation. In addition, an improvement in the signal-to-noise ratio in the XRPD data under a cycle of dehydration/rehydration is reported. These results underline the importance of considering the impact of environmental factors, such as humidity and temperature, in the context of structure-based drug design, thereby contributing to the development of more effective and stable pharmaceutical products.
Full article

Figure 1
Figure 1
<p>Pawley fits of XRPD data of polycrystalline octreotide at ambient conditions (capillary mode) and selected rH levels (95%, 70%, 60%, 40%, and 30%). The data extend up to ~2.35 Å resolution. They were collected employing a laboratory X-ray powder diffractometer (X’Pert Pro, Malvern Panalytical) equipped with an Anton Paar MHC-trans humidity-temperature chamber [λ = 1.540598 Å, RT]. In each panel, the black and red lines represent the experimental data and the calculated profiles, respectively, while the blue line corresponds to the difference between the experimental and calculated profiles. The vertical bars indicate the Bragg reflections compatible with this space group (<span class="html-italic">P</span>2<sub>1</sub>2<sub>1</sub>2<sub>1</sub>, lattice parameters at ambient conditions: a = 18.608(2) Å, b = 30.254(3) Å, and c = 39.794(6) Å).</p> Full article ">Figure 2
<p>Upper panel: Pawley fit of the XRPD synchrotron data of octreotide. The data were collected on ID22 at ESRF and extend up to ~ 2.35 Å resolution [λ = 1.3007899(8) Å, RT]. The black, red, and lower green lines represent the experimental data, the calculated pattern, and the difference between the experimental and calculated profiles, respectively. The orange vertical bars correspond to Bragg reflections compatible with this space group (<span class="html-italic">P</span>2<sub>1</sub>2<sub>1</sub>2<sub>1</sub>, a = 18.57744(4) Å, b = 30.17338(6) Å, and c = 39.70590(9) Å). To highlight the enhanced d-spacing resolution, the profile was systematically multiplied by factors of 5 and 16, as indicated in the figure. Lower panel: Magnification of the 2θ range from 4.3° to 5.6°, emphasizing the enhanced angular resolution of the diffraction pattern. The background intensity has been subtracted for clarity.</p> Full article ">Figure 3
<p>Surface plots of laboratory XRPD data of the octreotide polycrystalline precipitate upon gradual dehydration/rehydration cycles from 95% to 60% rH (<b>left</b>), 95% to 40% rH (<b>middle</b>), and 95% to 30% (<b>right</b>). Alterations of the peak positions and intensities are evident upon gradual dehydration and rehydration cycles. Significant peak shifts become evident upon dehydration, particularly below 70% rH. Upon rehydration and above 75% rH, the sample effectively recovers to its initial state.</p> Full article ">Figure 4
<p>Evolution of normalized unit-cell parameters upon gradual dehydration and rehydration of the octreotide polycrystalline sample from 95% to 60% rH (<b>upper panel</b>), 40% rH (<b>middle panel</b>), and 30% rH (<b>lower panel</b>). Purple, red, green, and blue symbols correspond to the extracted parameters of the unit-cell volume V, the a axis, the b axis, and the c axis, respectively. The lines are guides to the eye.</p> Full article ">Figure 5
<p>Magnification of the laboratory XRPD data in the 4–8° 2<span class="html-italic">θ</span> range reveals significant peak shifts at 75% and 70% rH (RT), along with the subsequent recovery of the sample after rehydration.</p> Full article ">Figure 6
<p>Following a complete dehydration and rehydration cycle, a comparative view of XRPD data at 95% rH reveals a pronounced improvement of the XRPD data in terms of signal-to-noise ratio.</p> Full article ">Figure 7
<p>Surface plots of XRPD data of octreotide polycrystalline precipitate upon gradual heating/cooling cycles at specific rH levels. A noticeable shift in the diffraction peak positions and intensities is observed at 75% and 65% rH. The latter observation may be attributed more to the effect of humidity than temperature. In addition, the XRPD data collected upon rH variation at ambient temperature described above indicate that at rH lower than 75%, the sample exhibits a slight alteration in terms of unit-cell dimensions, yet crystallinity is maintained. The latter suggests that temperature does not significantly impact the structural integrity of the polycrystalline peptide sample.</p> Full article ">Figure 8
<p>Evolution of normalized unit-cell parameters upon gradual heating and cooling cycles of the octreotide polycrystalline sample from 294.15 K to 318.15 K at selected rH levels. Turquoise, pink, dark blue, light blue, brown, and green symbols correspond to the rH levels of 95%, 85%, 75%, 65%, 55%, and 45%, respectively. The lines are guides to the eye.</p> Full article ">Figure 9
<p>Optical microscopy images of the polycrystalline octreotide sample.</p> Full article ">Figure 10
<p>(<b>a</b>) Empty Kapton foil holder (<b>left</b>) and Kapton foil holder filled with the polycrystalline octreotide sample (<b>right</b>). (<b>b</b>) View of the interior of the humidity chamber containing the multiple-position sample holder. (<b>c</b>) The configuration of the X’Pert Pro diffractometer equipped with the MHC-trans humidity and temperature chamber for in situ XRPD data collection in transmission mode.</p> Full article ">
<p>Pawley fits of XRPD data of polycrystalline octreotide at ambient conditions (capillary mode) and selected rH levels (95%, 70%, 60%, 40%, and 30%). The data extend up to ~2.35 Å resolution. They were collected employing a laboratory X-ray powder diffractometer (X’Pert Pro, Malvern Panalytical) equipped with an Anton Paar MHC-trans humidity-temperature chamber [λ = 1.540598 Å, RT]. In each panel, the black and red lines represent the experimental data and the calculated profiles, respectively, while the blue line corresponds to the difference between the experimental and calculated profiles. The vertical bars indicate the Bragg reflections compatible with this space group (<span class="html-italic">P</span>2<sub>1</sub>2<sub>1</sub>2<sub>1</sub>, lattice parameters at ambient conditions: a = 18.608(2) Å, b = 30.254(3) Å, and c = 39.794(6) Å).</p> Full article ">Figure 2
<p>Upper panel: Pawley fit of the XRPD synchrotron data of octreotide. The data were collected on ID22 at ESRF and extend up to ~ 2.35 Å resolution [λ = 1.3007899(8) Å, RT]. The black, red, and lower green lines represent the experimental data, the calculated pattern, and the difference between the experimental and calculated profiles, respectively. The orange vertical bars correspond to Bragg reflections compatible with this space group (<span class="html-italic">P</span>2<sub>1</sub>2<sub>1</sub>2<sub>1</sub>, a = 18.57744(4) Å, b = 30.17338(6) Å, and c = 39.70590(9) Å). To highlight the enhanced d-spacing resolution, the profile was systematically multiplied by factors of 5 and 16, as indicated in the figure. Lower panel: Magnification of the 2θ range from 4.3° to 5.6°, emphasizing the enhanced angular resolution of the diffraction pattern. The background intensity has been subtracted for clarity.</p> Full article ">Figure 3
<p>Surface plots of laboratory XRPD data of the octreotide polycrystalline precipitate upon gradual dehydration/rehydration cycles from 95% to 60% rH (<b>left</b>), 95% to 40% rH (<b>middle</b>), and 95% to 30% (<b>right</b>). Alterations of the peak positions and intensities are evident upon gradual dehydration and rehydration cycles. Significant peak shifts become evident upon dehydration, particularly below 70% rH. Upon rehydration and above 75% rH, the sample effectively recovers to its initial state.</p> Full article ">Figure 4
<p>Evolution of normalized unit-cell parameters upon gradual dehydration and rehydration of the octreotide polycrystalline sample from 95% to 60% rH (<b>upper panel</b>), 40% rH (<b>middle panel</b>), and 30% rH (<b>lower panel</b>). Purple, red, green, and blue symbols correspond to the extracted parameters of the unit-cell volume V, the a axis, the b axis, and the c axis, respectively. The lines are guides to the eye.</p> Full article ">Figure 5
<p>Magnification of the laboratory XRPD data in the 4–8° 2<span class="html-italic">θ</span> range reveals significant peak shifts at 75% and 70% rH (RT), along with the subsequent recovery of the sample after rehydration.</p> Full article ">Figure 6
<p>Following a complete dehydration and rehydration cycle, a comparative view of XRPD data at 95% rH reveals a pronounced improvement of the XRPD data in terms of signal-to-noise ratio.</p> Full article ">Figure 7
<p>Surface plots of XRPD data of octreotide polycrystalline precipitate upon gradual heating/cooling cycles at specific rH levels. A noticeable shift in the diffraction peak positions and intensities is observed at 75% and 65% rH. The latter observation may be attributed more to the effect of humidity than temperature. In addition, the XRPD data collected upon rH variation at ambient temperature described above indicate that at rH lower than 75%, the sample exhibits a slight alteration in terms of unit-cell dimensions, yet crystallinity is maintained. The latter suggests that temperature does not significantly impact the structural integrity of the polycrystalline peptide sample.</p> Full article ">Figure 8
<p>Evolution of normalized unit-cell parameters upon gradual heating and cooling cycles of the octreotide polycrystalline sample from 294.15 K to 318.15 K at selected rH levels. Turquoise, pink, dark blue, light blue, brown, and green symbols correspond to the rH levels of 95%, 85%, 75%, 65%, 55%, and 45%, respectively. The lines are guides to the eye.</p> Full article ">Figure 9
<p>Optical microscopy images of the polycrystalline octreotide sample.</p> Full article ">Figure 10
<p>(<b>a</b>) Empty Kapton foil holder (<b>left</b>) and Kapton foil holder filled with the polycrystalline octreotide sample (<b>right</b>). (<b>b</b>) View of the interior of the humidity chamber containing the multiple-position sample holder. (<b>c</b>) The configuration of the X’Pert Pro diffractometer equipped with the MHC-trans humidity and temperature chamber for in situ XRPD data collection in transmission mode.</p> Full article ">
Open AccessArticle
Density and Composition of Cohabiting Bacteria in Chlorella vulgaris CCAP 211/21A Is Influenced by Changes in Nutrient Supply
by
Wasayf J. Almalki, Alison O. Nwokeoji and Seetharaman Vaidyanathan
SynBio 2024, 2(2), 190-204; https://doi.org/10.3390/synbio2020011 - 17 May 2024
Abstract
►▼
Show Figures
Microalgae have considerable potential as a renewable feedstock for biochemical and bioethanol production that can be employed in processes associated with carbon capture. Large-scale microalgae cultivations are often non-axenic and are often cohabited by bacteria. A better understanding of the influence of cohabiting
[...] Read more.
Microalgae have considerable potential as a renewable feedstock for biochemical and bioethanol production that can be employed in processes associated with carbon capture. Large-scale microalgae cultivations are often non-axenic and are often cohabited by bacteria. A better understanding of the influence of cohabiting bacteria on microalgae productivity is required to develop sustainable synthetic co-culture processes at scale. Nutrient limitation is a frequently employed strategy in algal cultivations to accumulate energy reserves, such as lipids and carbohydrates. Here, a non-axenic culture of an estuarine green microalga, Chlorella vulgaris CCAP 211/21A, was studied under nutrient replete and deplete conditions to assess how changes in nutrient supply influenced the cohabiting bacterial population and its association with intracellular carbohydrate accumulations in the alga. Nutrient limitation resulted in a maximum carbohydrate yield of 47%, which was 74% higher than that in nutrient replete conditions. However, the latter condition elicited a 2-fold higher carbohydrate productivity. Three cohabiting bacterial isolates were cultivable from the three culture conditions tested. These isolates were identified using the 16S rRNA gene sequence to belong to Halomonas sp. and Muricauda sp. The composition of the bacterial population varied significantly between the growth conditions and time points. In all cases and at all time points, the dominant species was Halomonas isolates. Nutrient depletion resulted in an apparent loss of Muricauda sp. This finding demonstrates that nutrient supply can be used to control cohabiting bacterial populations in algal cultures, which will enable the development of synthetic co-culture strategies for improving algae productivity.
Full article

Figure 1
Figure 1
<p>Growth characteristics of non-axenic <span class="html-italic">C. vulgaris</span> in <span class="html-italic">f/2</span> and modified <span class="html-italic">f/2</span> media cultured in batch mode for 7 days; (<b>A</b>) cell count, (<b>B</b>) optical density (OD<sub>595nm</sub>), (<b>C</b>) dry cell weight of algae, (<b>D</b>) nitrate consumption over the cultivation period, (<b>E</b>) phosphate consumption over the cultivation period, (<b>F</b>) median specific growth rate of algae over the cultivation period, and (<b>G</b>) maximum biomass productivity.</p> Full article ">Figure 2
<p>(<b>a</b>). An SEM image of non-axenic <span class="html-italic">C. vulgaris</span> culture, showing <span class="html-italic">Halomonas</span> (A,B) and <span class="html-italic">Muricauda</span> (C) cells in the co-culture, alongside <span class="html-italic">C. vulgaris</span> cells. (<b>b</b>). Three species of co-habiting bacteria isolated from <span class="html-italic">C. vulgaris</span> culture after 7 days of cultivation on <span class="html-italic">f/2</span> + R2A agar plate incubated at room temperature.</p> Full article ">Figure 3
<p>Phylogenetic tree of isolated cohabiting bacteria (<b>A</b>): <span class="html-italic">Halomonas</span> (A,B) and (<b>B</b>): <span class="html-italic">Muricauda</span> (C) from <span class="html-italic">C. vulgaris</span> culture and closely related species based on 16SrRNA sequencing, distance was estimated with Tamura-Nei model and branch support were assessed with 500 bootstraps using Mega 11.0 software.</p> Full article ">Figure 4
<p>Growth rate by colonies count of bacterial isolates that grow in different media for optimal growth (<b>A</b>) <span class="html-italic">Halomonas</span> sp. WSR2, (<b>B</b>) <span class="html-italic">Halomonas</span> sp. WS1, (<b>C</b>) <span class="html-italic">Muricuda</span> sp. WSR, and (<b>D</b>) µmax of the growth rate of all three species.</p> Full article ">Figure 5
<p>Distribution of three species of bacteria at three isolation points during microalgae cultivation in <span class="html-italic">f/2</span>, <span class="html-italic">f/4,</span> and <span class="html-italic">2f</span> media. Nutrient depletion increases carbohydrate yields but repletion increases productivity with an apparent loss of a bacterial isolate from the cohabiting population.</p> Full article ">Figure 6
<p>Time profile of carbohydrate yield (%DCW) of <span class="html-italic">C. vulgaris</span> in <span class="html-italic">f/2</span>, <span class="html-italic">f/4</span>, and <span class="html-italic">2f</span> media. The cohabiting bacterial distribution in the three media at a time point when the maximum yield was observed is plotted on top, as well as the maximum carbohydrate productivity at this time point, plotted alongside.</p> Full article ">Figure 7
<p>Agarose gel electrophoresis of microbial genomic DNA of <span class="html-italic">Halomonas</span> sp. WSR2 (A), <span class="html-italic">Halomonas</span> sp. WS1 (B), and <span class="html-italic">Muricuda</span> sp. WSR (C).</p> Full article ">
<p>Growth characteristics of non-axenic <span class="html-italic">C. vulgaris</span> in <span class="html-italic">f/2</span> and modified <span class="html-italic">f/2</span> media cultured in batch mode for 7 days; (<b>A</b>) cell count, (<b>B</b>) optical density (OD<sub>595nm</sub>), (<b>C</b>) dry cell weight of algae, (<b>D</b>) nitrate consumption over the cultivation period, (<b>E</b>) phosphate consumption over the cultivation period, (<b>F</b>) median specific growth rate of algae over the cultivation period, and (<b>G</b>) maximum biomass productivity.</p> Full article ">Figure 2
<p>(<b>a</b>). An SEM image of non-axenic <span class="html-italic">C. vulgaris</span> culture, showing <span class="html-italic">Halomonas</span> (A,B) and <span class="html-italic">Muricauda</span> (C) cells in the co-culture, alongside <span class="html-italic">C. vulgaris</span> cells. (<b>b</b>). Three species of co-habiting bacteria isolated from <span class="html-italic">C. vulgaris</span> culture after 7 days of cultivation on <span class="html-italic">f/2</span> + R2A agar plate incubated at room temperature.</p> Full article ">Figure 3
<p>Phylogenetic tree of isolated cohabiting bacteria (<b>A</b>): <span class="html-italic">Halomonas</span> (A,B) and (<b>B</b>): <span class="html-italic">Muricauda</span> (C) from <span class="html-italic">C. vulgaris</span> culture and closely related species based on 16SrRNA sequencing, distance was estimated with Tamura-Nei model and branch support were assessed with 500 bootstraps using Mega 11.0 software.</p> Full article ">Figure 4
<p>Growth rate by colonies count of bacterial isolates that grow in different media for optimal growth (<b>A</b>) <span class="html-italic">Halomonas</span> sp. WSR2, (<b>B</b>) <span class="html-italic">Halomonas</span> sp. WS1, (<b>C</b>) <span class="html-italic">Muricuda</span> sp. WSR, and (<b>D</b>) µmax of the growth rate of all three species.</p> Full article ">Figure 5
<p>Distribution of three species of bacteria at three isolation points during microalgae cultivation in <span class="html-italic">f/2</span>, <span class="html-italic">f/4,</span> and <span class="html-italic">2f</span> media. Nutrient depletion increases carbohydrate yields but repletion increases productivity with an apparent loss of a bacterial isolate from the cohabiting population.</p> Full article ">Figure 6
<p>Time profile of carbohydrate yield (%DCW) of <span class="html-italic">C. vulgaris</span> in <span class="html-italic">f/2</span>, <span class="html-italic">f/4</span>, and <span class="html-italic">2f</span> media. The cohabiting bacterial distribution in the three media at a time point when the maximum yield was observed is plotted on top, as well as the maximum carbohydrate productivity at this time point, plotted alongside.</p> Full article ">Figure 7
<p>Agarose gel electrophoresis of microbial genomic DNA of <span class="html-italic">Halomonas</span> sp. WSR2 (A), <span class="html-italic">Halomonas</span> sp. WS1 (B), and <span class="html-italic">Muricuda</span> sp. WSR (C).</p> Full article ">
Open AccessArticle
Construction of an Elastin-like Polypeptide Gene in a High Copy Number Plasmid Using a Modified Method of Recursive Directional Ligation
by
Derek W. Nelson, Alexander Connor, Yu Shen and Ryan J. Gilbert
SynBio 2024, 2(2), 174-189; https://doi.org/10.3390/synbio2020010 - 5 May 2024
Abstract
►▼
Show Figures
Elastin-like polypeptides (ELPs) are popular biomaterials due to their reversible, temperature-dependent phase separation and their tunability, which is achievable by evolving procedures in recombinant technology. In particular, recursive direction ligation by plasmid reconstruction (PRe-RDL) is the predominant cloning technique used to generate ELPs
[...] Read more.
Elastin-like polypeptides (ELPs) are popular biomaterials due to their reversible, temperature-dependent phase separation and their tunability, which is achievable by evolving procedures in recombinant technology. In particular, recursive direction ligation by plasmid reconstruction (PRe-RDL) is the predominant cloning technique used to generate ELPs of varying lengths. Pre-RDL provides precise control over the number of (VPGXG)n repeat units in an ELP due to the selection of type IIS restriction enzyme (REs) sites in the reconstructed pET expression plasmid, which is a low-to-medium copy number plasmid. While Pre-RDL can be used to seamlessly repeat essentially any gene sequence and overcome limitations of previous cloning practices, we modified the Pre-RDL technique, where a high copy number plasmid (pBluescript II SK(+)—using a new library of type IIS REs) was used instead of a pET plasmid. The modified technique successfully produced a diblock ELP gene of 240 pentapeptide repeats from 30 pentapeptide “monomers” composed of alanine, tyrosine, and leucine X residues. This study found that the large, GC-rich ELP gene compromised plasmid yields in pBluescript II SK(+) and favored higher plasmid yields in the pET19b expression plasmid. Additionally, the BL21 E. coli strain expression consistently provided a higher transformation efficiency and higher plasmid yield than the high cloning efficiency strain TOP10 E. coli. We hypothesize that the plasmid/high GC gene ratio may play a significant role in these observations, and not the total plasmid size or the total plasmid GC content. While expression of the final gene resulted in a diblock ELP with a phase separation temperature of 34.5 °C, future work will need to investigate RDL techniques in additional plasmids to understand the primary driving factors for improving yields of plasmids with large ELP-encoding genes.
Full article

Figure 1
Figure 1
<p>Schematic of the GTMs-RDL methodology used in this study. A pBluescript II SK(+) plasmid containing an ELP gene undergoes two parallel digest reactions to create two gene/plasmid fragments of interest. Due to the design of the original ELP gene insert, ligating these two fragments together results in doubling the ELP gene size without a scar sequence in the original pBluescript SK(+) plasmid. This can be repeated as needed or moved to an expression plasmid once the final gene size is achieved. Not shown is the presence of the XbaI restriction enzyme site before the BmsI site to facilitate gene insertion into the pBluescript II SK(+) plasmid. The ELP gene is shown in gray. The enzyme binding sites are highlighted and enzyme cut sites are outlined in brown for AdeI, red for BmsI, blue for NcoI, purple for BpiI, and green for BamHI. * indicates the stop codon sequence (tag).</p> Full article ">Figure 2
<p>Representative images of agarose gels demonstrating the bands of interest during GTMs-RDL (<b>A</b>) and the confirmation of the ligation of those bands of interest in pBluescript II SK(+) for the final synthesis of the ELP-L<sub>4</sub>Y<sub>4</sub> gene (<b>B</b>). For example, bands of interest from lanes 1 and 4 in (<b>A</b>) were ligated together to produce the ELP-L<sub>2</sub> gene shown in lane 6 of (<b>B</b>). NcoI and BamHI restriction enzymes were used in (<b>B</b>) to remove each ELP gene from its plasmid. The white arrow in (<b>B</b>) indicates the successful ligation of the ELP-L<sub>4</sub>Y<sub>4</sub> gene in lane 4 from pBluescript II SK(+) to pET19b in lane 8. Orange indicates ELP genes encoding tyrosine residues and cyan indicates ELP genes encoding leucine residues. * indicates a scenario where a PagI restriction enzyme is needed to remove overlapping bands with the band of interest.</p> Full article ">Figure 3
<p>ELP plasmid yield is dependent on gene size, plasmid selection, and bacterial strain. Increasing ELP gene size correlates with decreased plasmid yields (<b>A</b>). Gene sizes correlate with monomer, dimer, tetramer, and octamer ELP genes in this study, left to right on the x-axis (<b>A</b>). Additionally, octamer pET19b plasmids (gray) resulted in higher yields compared to octamer pBluescript II SK(+) plasmids (blue), and BL21 <span class="html-italic">E. coli</span> also improved plasmid yields compared to TOP10 <span class="html-italic">E. coli</span> strains (<b>B</b>). Lastly, the final ELP octamer gene (cyan and orange arrows labeled L, Y) is shown in pBluescript II SK(+) (<b>C</b>) and pET19b (<b>D</b>) to represent GC content and the ELP gene size relative to each plasmid as a whole. Brown–Forsythe and Welch ANOVA tests were performed for both (<b>A</b>,<b>B</b>). Statistical significance is indicated by * (<span class="html-italic">p</span> < 0.05), *** (<span class="html-italic">p</span> < 0.001), or **** (<span class="html-italic">p</span> < 0.0001).</p> Full article ">Figure 4
<p>ELP-L<sub>4</sub>Y<sub>4</sub> was expressed, purified, and characterized for turbidimetry and particle formation. After three rounds of ITC, each ELP batch was assessed for purity via an Agilent Bioanalyzer, where the red brackets indicate where impurities were detected. (<b>A</b>) Purity was assessed and reported as a percentage by an area under the curve analysis of the fluorescent signal, and the molecular weight was assessed by associating integration time with an Agilent protein ladder. (<b>B</b>) These data were used to simulate a PAGE gel where all values are displayed in kDa. (<b>C</b>) Purified ELP was also assessed for turbidimetry, where red indicates heating and blue indicates cooling at 1 °C/min. Red and blue shading indicates the standard deviation among three separate batches of ELP. (<b>D</b>) Phase-separated ELP-L<sub>4</sub>Y<sub>4</sub> was visualized using phase contrast at 20× magnification where the scale bar is 20 µm. The time-lapse shows both coalescing (black arrows) and non-coalescing (white arrows) particles.</p> Full article ">Figure 5
<p>After completing the last round of GTMs-RDL, modification of the gene termini can be performed in a single digest reaction to integrate a short therapeutic or bioactive moiety. Due to the design criteria for GTMs-RDL, modification at the N-terminus will always result in methionine and a specified amino acid (U) between the inserted fragment and the ELP gene. Modification of the C-terminus does not have any limitations on the encoded amino acid sequence. The ELP gene is shown in gray. The enzyme binding sites are highlighted and enzyme cut sites are outlined in brown for XbaI, red for BmsI, blue for NcoI, purple for BpiI, and green for BamHI. * indicates the stop codon sequence (tag).</p> Full article ">
<p>Schematic of the GTMs-RDL methodology used in this study. A pBluescript II SK(+) plasmid containing an ELP gene undergoes two parallel digest reactions to create two gene/plasmid fragments of interest. Due to the design of the original ELP gene insert, ligating these two fragments together results in doubling the ELP gene size without a scar sequence in the original pBluescript SK(+) plasmid. This can be repeated as needed or moved to an expression plasmid once the final gene size is achieved. Not shown is the presence of the XbaI restriction enzyme site before the BmsI site to facilitate gene insertion into the pBluescript II SK(+) plasmid. The ELP gene is shown in gray. The enzyme binding sites are highlighted and enzyme cut sites are outlined in brown for AdeI, red for BmsI, blue for NcoI, purple for BpiI, and green for BamHI. * indicates the stop codon sequence (tag).</p> Full article ">Figure 2
<p>Representative images of agarose gels demonstrating the bands of interest during GTMs-RDL (<b>A</b>) and the confirmation of the ligation of those bands of interest in pBluescript II SK(+) for the final synthesis of the ELP-L<sub>4</sub>Y<sub>4</sub> gene (<b>B</b>). For example, bands of interest from lanes 1 and 4 in (<b>A</b>) were ligated together to produce the ELP-L<sub>2</sub> gene shown in lane 6 of (<b>B</b>). NcoI and BamHI restriction enzymes were used in (<b>B</b>) to remove each ELP gene from its plasmid. The white arrow in (<b>B</b>) indicates the successful ligation of the ELP-L<sub>4</sub>Y<sub>4</sub> gene in lane 4 from pBluescript II SK(+) to pET19b in lane 8. Orange indicates ELP genes encoding tyrosine residues and cyan indicates ELP genes encoding leucine residues. * indicates a scenario where a PagI restriction enzyme is needed to remove overlapping bands with the band of interest.</p> Full article ">Figure 3
<p>ELP plasmid yield is dependent on gene size, plasmid selection, and bacterial strain. Increasing ELP gene size correlates with decreased plasmid yields (<b>A</b>). Gene sizes correlate with monomer, dimer, tetramer, and octamer ELP genes in this study, left to right on the x-axis (<b>A</b>). Additionally, octamer pET19b plasmids (gray) resulted in higher yields compared to octamer pBluescript II SK(+) plasmids (blue), and BL21 <span class="html-italic">E. coli</span> also improved plasmid yields compared to TOP10 <span class="html-italic">E. coli</span> strains (<b>B</b>). Lastly, the final ELP octamer gene (cyan and orange arrows labeled L, Y) is shown in pBluescript II SK(+) (<b>C</b>) and pET19b (<b>D</b>) to represent GC content and the ELP gene size relative to each plasmid as a whole. Brown–Forsythe and Welch ANOVA tests were performed for both (<b>A</b>,<b>B</b>). Statistical significance is indicated by * (<span class="html-italic">p</span> < 0.05), *** (<span class="html-italic">p</span> < 0.001), or **** (<span class="html-italic">p</span> < 0.0001).</p> Full article ">Figure 4
<p>ELP-L<sub>4</sub>Y<sub>4</sub> was expressed, purified, and characterized for turbidimetry and particle formation. After three rounds of ITC, each ELP batch was assessed for purity via an Agilent Bioanalyzer, where the red brackets indicate where impurities were detected. (<b>A</b>) Purity was assessed and reported as a percentage by an area under the curve analysis of the fluorescent signal, and the molecular weight was assessed by associating integration time with an Agilent protein ladder. (<b>B</b>) These data were used to simulate a PAGE gel where all values are displayed in kDa. (<b>C</b>) Purified ELP was also assessed for turbidimetry, where red indicates heating and blue indicates cooling at 1 °C/min. Red and blue shading indicates the standard deviation among three separate batches of ELP. (<b>D</b>) Phase-separated ELP-L<sub>4</sub>Y<sub>4</sub> was visualized using phase contrast at 20× magnification where the scale bar is 20 µm. The time-lapse shows both coalescing (black arrows) and non-coalescing (white arrows) particles.</p> Full article ">Figure 5
<p>After completing the last round of GTMs-RDL, modification of the gene termini can be performed in a single digest reaction to integrate a short therapeutic or bioactive moiety. Due to the design criteria for GTMs-RDL, modification at the N-terminus will always result in methionine and a specified amino acid (U) between the inserted fragment and the ELP gene. Modification of the C-terminus does not have any limitations on the encoded amino acid sequence. The ELP gene is shown in gray. The enzyme binding sites are highlighted and enzyme cut sites are outlined in brown for XbaI, red for BmsI, blue for NcoI, purple for BpiI, and green for BamHI. * indicates the stop codon sequence (tag).</p> Full article ">
Open AccessReview
Crafting Genetic Diversity: Unlocking the Potential of Protein Evolution
by
Vamsi Krishna Gali, Kang Lan Tee and Tuck Seng Wong
SynBio 2024, 2(2), 142-173; https://doi.org/10.3390/synbio2020009 - 7 Apr 2024
Abstract
►▼
Show Figures
Genetic diversity is the foundation of evolutionary resilience, adaptive potential, and the flourishing vitality of living organisms, serving as the cornerstone for robust ecosystems and the continuous evolution of life on Earth. The landscape of directed evolution, a powerful biotechnological tool inspired by
[...] Read more.
Genetic diversity is the foundation of evolutionary resilience, adaptive potential, and the flourishing vitality of living organisms, serving as the cornerstone for robust ecosystems and the continuous evolution of life on Earth. The landscape of directed evolution, a powerful biotechnological tool inspired by natural evolutionary processes, has undergone a transformative shift propelled by innovative strategies for generating genetic diversity. This shift is fuelled by several factors, encompassing the utilization of advanced toolkits like CRISPR-Cas and base editors, the enhanced comprehension of biological mechanisms, cost-effective custom oligo pool synthesis, and the seamless integration of artificial intelligence and automation. This comprehensive review looks into the myriad of methodologies employed for constructing gene libraries, both in vitro and in vivo, categorized into three major classes: random mutagenesis, focused mutagenesis, and DNA recombination. The objectives of this review are threefold: firstly, to present a panoramic overview of recent advances in genetic diversity creation; secondly, to inspire novel ideas for further innovation in genetic diversity generation; and thirdly, to provide a valuable resource for individuals entering the field of directed evolution.
Full article
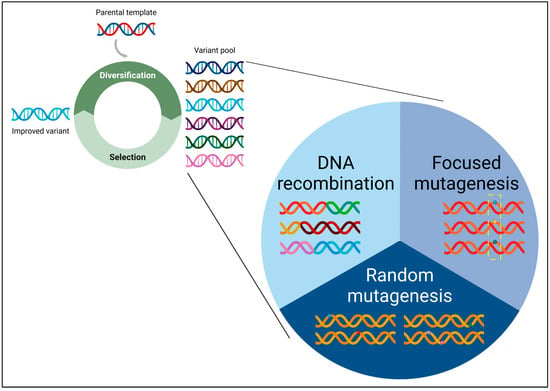
Figure 1
Figure 1
<p>(<b>Top left</b>) The directed evolution cycle. The parental gene of interest (GOI) undergoes mutagenesis to generate a diverse pool of genetic variants. This pool is then subjected to a selection process targeting the desired phenotype, enabling the identification of improved variant(s). This iterative cycle is repeated until the desired trait is successfully achieved. (<b>Bottom right</b>) Classification of genetic diversity creation methods. The diverse methods for generating a genetically varied gene pool can be systematically categorized into three main classes: random mutagenesis, focused mutagenesis, and DNA recombination. Random mutagenesis involves the introduction of random mutations throughout the starting parental gene sequence. Focused mutagenesis targets mutations to specific pre-selected region(s) or amino acid residue(s) within the starting parental gene sequence. DNA recombination generates chimeric sequences by combining segments from a set of either homologous or non-homologous parental sequences.</p> Full article ">Figure 2
<p>Illustration of the experimental procedure of CReasPy-cloning. Initially, a yeast is transformed with two plasmids—pCas9 and pgRNA—enabling the expression of the Cas9 nuclease and a guide RNA (gRNA). These plasmids carry the TRP1 and URA3 selection markers, respectively. Following this, the yeast undergoes simultaneous transformation with the target genome to be cloned and a linear DNA fragment containing yeast elements (CEN-HIS3, with or without ARS). The linear DNA fragment has recombination arms homologous to each side of the target locus. Upon entry into the cell, the Cas9/gRNA complex cleaves the target genome, and the yeast homologous recombination system repairs it using the provided linear DNA fragment as a template. Consequently, the bacterial genome incorporates the yeast elements precisely at the designated locus and is now carried by the yeast as an artificial chromosome.</p> Full article ">Figure 3
<p>An overview of the PTO-QuickStep method. Initially, megaprimers (coloured in blue) are generated in a PCR using a set of PTO oligonucleotides containing phosphorothioate linkages (indicated with letters ‘P’). Subsequently, iodoethanol treatment is applied to the megaprimers, breaking the phosphorothioate linkages and exposing 3′-overhangs. In the second step, these treated megaprimers anneal to the destination or recipient vector at the target locus, initiating the amplification of the entire plasmid. Moving to the third step, DpnI is employed to remove the methylated or hemimethylated destination or recipient vector without the gene insert (shown as dotted circle). After DpnI digestion, the newly synthesized plasmids undergo transformation into <span class="html-italic">E. coli</span>, where any nicks are repaired <span class="html-italic">in vivo</span>.</p> Full article ">Figure 4
<p>A schematic representation of the random mutagenesis method tailored for <span class="html-italic">Pichia pastoris</span>. Initially, the circular protein expression vector undergoes repeated amplification through strand displacement reactions facilitated by Phi29 DNA polymerase. Mutations are intentionally introduced by adding Mn<sup>2+</sup> to lower the fidelity of the polymerase, a process known as error-prone rolling circle amplification (epRCA). Following this, subsequent amplification, achieved through Phi29 DNA polymerase (or multiple displacement amplification, MDA), yields microgram quantities of mutated DNA. This mutated DNA is then utilized for transformation into <span class="html-italic">P. pastoris</span> to enable enzyme production.</p> Full article ">Figure 5
<p>Graphic summary of the MutaT7 mutagenesis system and its derivatives. The T7 RNA polymerase fusion (T7RNAP) selectively binds to the T7 promoter, initiating transcription and traversing the gene of interest. As the fusion carries a base editor (BE), mutations are randomly introduced into the gene, represented by blue vertical stripes. The fusion halts and disengages from the DNA upon encountering a dCas9 molecule bound to a specific sequence dictated by the CRISPR RNA (crRNA). The termination process is also facilitated by the transactivating CRISPR RNA (tracrRNA). In the absence of dCas9, the movement of the fusion protein can be halted by incorporating one or multiple T7 terminators.</p> Full article ">Figure 6
<p>Targeted random mutagenesis using chimeric proteins comprising a base editor (BE) and a guide protein (GP), following the general BE-GP protein architecture. BE is the mutagenic agent, introducing random mutations through its base-editing activity (<span class="html-italic">e.g.</span>, cytidine and adenosine deamination). GP, with DNA-binding capability, guides or leads the BE to its target locus within the gene of interest (GOI) to effect mutagenesis. Typical BE choices include cytidine deaminase (CDA) or error-prone DNA polymerase (Pol). Frequently used GP candidates are T7 RNA polymerase (RNAP) or catalytically dead Cas9 (dCas9)/Cas9 nickase (nCas9). BE is tethered to GP via gene fusion or protein/protein or protein/RNA interactions through the utilization of SRC homology domain 3 (SH3) and the MS2 bacteriophage coat protein. In some methods, an accessory protein such as uracil DNA glycosylase inhibitor (UGI) is required.</p> Full article ">Figure 7
<p>Schematic representation of the VEGAS platform for directed evolution, a technique for engineering DNA sequences in mammalian cells. This approach relies on the use of the Sindbis virus for efficient and mutagenic viral propagation in mammalian cell culture. To establish a robust directed evolution platform that harnesses the replicative and mutagenic potential of the Sindbis virus, artificial selective pressure must be applied. A crucial aspect involves the requirement for 240 copies of each structural protein (E1, E2, and capsid) in each Sindbis viral particle to form a functional unit capable of maturation and propagation. Without this envelope, the virus cannot mature or propagate. By strategically introducing limitations on the transcription of the structural genome, selective pressure can be applied to the transgenic Sindbis virus. In the VEGAS platform, the structural genome of the Sindbis virus is cloned into the mammalian expression vector pSSG, under the regulation of the tetracycline operator sequence. The structural genome elements of the Sindbis genome are then replaced with a transgene encoding a tetracycline transactivator. Propagation and selection can then be performed in mammalian cell culture, by infecting cells transfected with pSSG with the pTSin packaged virus.</p> Full article ">Figure 8
<p>Schematic representation of the TRIAD process for generating deletion libraries. In the first step, the TransDel insertion library is formed through <span class="html-italic">in vitro</span> transposition of the engineered transposon TransDel into the target sequence on circular plasmid DNA. In the second step, MlyI digestion is applied to eliminate TransDel along with 3 base pairs of the original target sequence, creating a single break per variant. The third step involves self-ligation, leading to the reconstitution of the target sequence minus 3 base pairs. This results in a library of single variants, each featuring a deletion of one triplet. Alternatively, DNA cassette Dels can be inserted between the breaks in the target sequence to produce insertion libraries. MlyI digestion removes the DNA cassette Del along with 3 or 6 additional base pairs of the original target sequence, depending on the used DNA cassette. Subsequent self-ligation reestablishes the target sequence, now with a deletion of 2 or 3 triplets. This versatile approach also allows for the creation of insertion libraries.</p> Full article ">Figure 9
<p>Multi-site-directed mutagenesis methods. (<b>A</b>) Nicking mutagenesis: This process begins with the wildtype (WT) plasmid dsDNA containing a 7-base-pair BbvCI recognition site, which is selectively nicked by Nt.BbvCI. The resulting nicked strand undergoes degradation by Exonuclease III (ExoIII), creating a single-stranded DNA (ssDNA) template. To eliminate insufficiently digested DNA, Exonuclease I (ExoI) is employed. Next, phosphorylated mutagenic primers are annealed to the ssDNA parental template. The mutagenic strand is then synthesized through the collaborative action of a polymerase and ligase. Following this synthesis, the WT template strand is nicked by Nb.BbvCI and subsequently digested by ExoIII. The introduction of a second primer initiates the synthesis of the complementary mutant strand, resulting in the generation of mutagenized dsDNA. (<b>B</b>) Combinatorial codon mutagenesis: This process initiates with two parallel PCR reactions. In one, mutagenic reverse primers are employed, while in the other, mutagenic forward primers are utilized. In the subsequent step, a third PCR is employed to combine the fragments generated from the preceding PCR reactions. (<b>C</b>) Plasmid recombineering: This method involves the direct <span class="html-italic">in vivo</span> incorporation of synthetic oligonucleotides carrying desired mutations into a gene of interest. These oligonucleotides are introduced into <span class="html-italic">E. coli</span> cells via electroporation and can recombine with resident plasmids, facilitated by the lambda phage protein Beta.</p> Full article ">Figure 10
<p>Schematic representation of the <span class="html-italic">In vitro</span> CRISPR/Cas9-mediated Mutagenic (ICM) system for site-directed mutagenesis: The target plasmid undergoes initial cleavage by the Cas9 protein and specific sgRNA complex at both sides of the mutational position, removing the wildtype sequence. Subsequent digestion with T5 exonuclease generates 15 nt sticky ends. Primers carrying the desired mutation are then annealed to create DNA fragments with 15 nt sticky ends that complement the digested plasmid. These fragments are inserted into the linearized vector through transformation into the host cell.</p> Full article ">
<p>(<b>Top left</b>) The directed evolution cycle. The parental gene of interest (GOI) undergoes mutagenesis to generate a diverse pool of genetic variants. This pool is then subjected to a selection process targeting the desired phenotype, enabling the identification of improved variant(s). This iterative cycle is repeated until the desired trait is successfully achieved. (<b>Bottom right</b>) Classification of genetic diversity creation methods. The diverse methods for generating a genetically varied gene pool can be systematically categorized into three main classes: random mutagenesis, focused mutagenesis, and DNA recombination. Random mutagenesis involves the introduction of random mutations throughout the starting parental gene sequence. Focused mutagenesis targets mutations to specific pre-selected region(s) or amino acid residue(s) within the starting parental gene sequence. DNA recombination generates chimeric sequences by combining segments from a set of either homologous or non-homologous parental sequences.</p> Full article ">Figure 2
<p>Illustration of the experimental procedure of CReasPy-cloning. Initially, a yeast is transformed with two plasmids—pCas9 and pgRNA—enabling the expression of the Cas9 nuclease and a guide RNA (gRNA). These plasmids carry the TRP1 and URA3 selection markers, respectively. Following this, the yeast undergoes simultaneous transformation with the target genome to be cloned and a linear DNA fragment containing yeast elements (CEN-HIS3, with or without ARS). The linear DNA fragment has recombination arms homologous to each side of the target locus. Upon entry into the cell, the Cas9/gRNA complex cleaves the target genome, and the yeast homologous recombination system repairs it using the provided linear DNA fragment as a template. Consequently, the bacterial genome incorporates the yeast elements precisely at the designated locus and is now carried by the yeast as an artificial chromosome.</p> Full article ">Figure 3
<p>An overview of the PTO-QuickStep method. Initially, megaprimers (coloured in blue) are generated in a PCR using a set of PTO oligonucleotides containing phosphorothioate linkages (indicated with letters ‘P’). Subsequently, iodoethanol treatment is applied to the megaprimers, breaking the phosphorothioate linkages and exposing 3′-overhangs. In the second step, these treated megaprimers anneal to the destination or recipient vector at the target locus, initiating the amplification of the entire plasmid. Moving to the third step, DpnI is employed to remove the methylated or hemimethylated destination or recipient vector without the gene insert (shown as dotted circle). After DpnI digestion, the newly synthesized plasmids undergo transformation into <span class="html-italic">E. coli</span>, where any nicks are repaired <span class="html-italic">in vivo</span>.</p> Full article ">Figure 4
<p>A schematic representation of the random mutagenesis method tailored for <span class="html-italic">Pichia pastoris</span>. Initially, the circular protein expression vector undergoes repeated amplification through strand displacement reactions facilitated by Phi29 DNA polymerase. Mutations are intentionally introduced by adding Mn<sup>2+</sup> to lower the fidelity of the polymerase, a process known as error-prone rolling circle amplification (epRCA). Following this, subsequent amplification, achieved through Phi29 DNA polymerase (or multiple displacement amplification, MDA), yields microgram quantities of mutated DNA. This mutated DNA is then utilized for transformation into <span class="html-italic">P. pastoris</span> to enable enzyme production.</p> Full article ">Figure 5
<p>Graphic summary of the MutaT7 mutagenesis system and its derivatives. The T7 RNA polymerase fusion (T7RNAP) selectively binds to the T7 promoter, initiating transcription and traversing the gene of interest. As the fusion carries a base editor (BE), mutations are randomly introduced into the gene, represented by blue vertical stripes. The fusion halts and disengages from the DNA upon encountering a dCas9 molecule bound to a specific sequence dictated by the CRISPR RNA (crRNA). The termination process is also facilitated by the transactivating CRISPR RNA (tracrRNA). In the absence of dCas9, the movement of the fusion protein can be halted by incorporating one or multiple T7 terminators.</p> Full article ">Figure 6
<p>Targeted random mutagenesis using chimeric proteins comprising a base editor (BE) and a guide protein (GP), following the general BE-GP protein architecture. BE is the mutagenic agent, introducing random mutations through its base-editing activity (<span class="html-italic">e.g.</span>, cytidine and adenosine deamination). GP, with DNA-binding capability, guides or leads the BE to its target locus within the gene of interest (GOI) to effect mutagenesis. Typical BE choices include cytidine deaminase (CDA) or error-prone DNA polymerase (Pol). Frequently used GP candidates are T7 RNA polymerase (RNAP) or catalytically dead Cas9 (dCas9)/Cas9 nickase (nCas9). BE is tethered to GP via gene fusion or protein/protein or protein/RNA interactions through the utilization of SRC homology domain 3 (SH3) and the MS2 bacteriophage coat protein. In some methods, an accessory protein such as uracil DNA glycosylase inhibitor (UGI) is required.</p> Full article ">Figure 7
<p>Schematic representation of the VEGAS platform for directed evolution, a technique for engineering DNA sequences in mammalian cells. This approach relies on the use of the Sindbis virus for efficient and mutagenic viral propagation in mammalian cell culture. To establish a robust directed evolution platform that harnesses the replicative and mutagenic potential of the Sindbis virus, artificial selective pressure must be applied. A crucial aspect involves the requirement for 240 copies of each structural protein (E1, E2, and capsid) in each Sindbis viral particle to form a functional unit capable of maturation and propagation. Without this envelope, the virus cannot mature or propagate. By strategically introducing limitations on the transcription of the structural genome, selective pressure can be applied to the transgenic Sindbis virus. In the VEGAS platform, the structural genome of the Sindbis virus is cloned into the mammalian expression vector pSSG, under the regulation of the tetracycline operator sequence. The structural genome elements of the Sindbis genome are then replaced with a transgene encoding a tetracycline transactivator. Propagation and selection can then be performed in mammalian cell culture, by infecting cells transfected with pSSG with the pTSin packaged virus.</p> Full article ">Figure 8
<p>Schematic representation of the TRIAD process for generating deletion libraries. In the first step, the TransDel insertion library is formed through <span class="html-italic">in vitro</span> transposition of the engineered transposon TransDel into the target sequence on circular plasmid DNA. In the second step, MlyI digestion is applied to eliminate TransDel along with 3 base pairs of the original target sequence, creating a single break per variant. The third step involves self-ligation, leading to the reconstitution of the target sequence minus 3 base pairs. This results in a library of single variants, each featuring a deletion of one triplet. Alternatively, DNA cassette Dels can be inserted between the breaks in the target sequence to produce insertion libraries. MlyI digestion removes the DNA cassette Del along with 3 or 6 additional base pairs of the original target sequence, depending on the used DNA cassette. Subsequent self-ligation reestablishes the target sequence, now with a deletion of 2 or 3 triplets. This versatile approach also allows for the creation of insertion libraries.</p> Full article ">Figure 9
<p>Multi-site-directed mutagenesis methods. (<b>A</b>) Nicking mutagenesis: This process begins with the wildtype (WT) plasmid dsDNA containing a 7-base-pair BbvCI recognition site, which is selectively nicked by Nt.BbvCI. The resulting nicked strand undergoes degradation by Exonuclease III (ExoIII), creating a single-stranded DNA (ssDNA) template. To eliminate insufficiently digested DNA, Exonuclease I (ExoI) is employed. Next, phosphorylated mutagenic primers are annealed to the ssDNA parental template. The mutagenic strand is then synthesized through the collaborative action of a polymerase and ligase. Following this synthesis, the WT template strand is nicked by Nb.BbvCI and subsequently digested by ExoIII. The introduction of a second primer initiates the synthesis of the complementary mutant strand, resulting in the generation of mutagenized dsDNA. (<b>B</b>) Combinatorial codon mutagenesis: This process initiates with two parallel PCR reactions. In one, mutagenic reverse primers are employed, while in the other, mutagenic forward primers are utilized. In the subsequent step, a third PCR is employed to combine the fragments generated from the preceding PCR reactions. (<b>C</b>) Plasmid recombineering: This method involves the direct <span class="html-italic">in vivo</span> incorporation of synthetic oligonucleotides carrying desired mutations into a gene of interest. These oligonucleotides are introduced into <span class="html-italic">E. coli</span> cells via electroporation and can recombine with resident plasmids, facilitated by the lambda phage protein Beta.</p> Full article ">Figure 10
<p>Schematic representation of the <span class="html-italic">In vitro</span> CRISPR/Cas9-mediated Mutagenic (ICM) system for site-directed mutagenesis: The target plasmid undergoes initial cleavage by the Cas9 protein and specific sgRNA complex at both sides of the mutational position, removing the wildtype sequence. Subsequent digestion with T5 exonuclease generates 15 nt sticky ends. Primers carrying the desired mutation are then annealed to create DNA fragments with 15 nt sticky ends that complement the digested plasmid. These fragments are inserted into the linearized vector through transformation into the host cell.</p> Full article ">
Open AccessArticle
Saccharomyces cerevisiae as a Host for Chondroitin Production
by
Márcia R. Couto, Joana L. Rodrigues, Oscar Dias and Lígia R. Rodrigues
SynBio 2024, 2(2), 125-141; https://doi.org/10.3390/synbio2020008 - 3 Apr 2024
Abstract
►▼
Show Figures
Chondroitin is a glycosaminoglycan that has gained widespread use in nutraceuticals and pharmaceuticals, mainly for treating osteoarthritis. Traditionally, it has been extracted from animal cartilage but recently, biotechnological processes have emerged as a commercial alternative to avoid the risk of viral or prion
[...] Read more.
Chondroitin is a glycosaminoglycan that has gained widespread use in nutraceuticals and pharmaceuticals, mainly for treating osteoarthritis. Traditionally, it has been extracted from animal cartilage but recently, biotechnological processes have emerged as a commercial alternative to avoid the risk of viral or prion contamination and offer a vegan-friendly source. Typically, these methods involve producing the chondroitin backbone using pathogenic bacteria and then modifying it enzymatically through the action of sulfotransferases. Despite the challenges of expressing active sulfotransferases in bacteria, the use of eukaryotic microorganisms is still limited to a few works using Pichia pastoris. To create a safer and efficient biotechnological platform, we constructed a biosynthetic pathway for chondroitin production in S. cerevisiae as a proof-of-concept. Up to 125 mg/L and 200 mg/L of intracellular and extracellular chondroitin were produced, respectively. Furthermore, as genome-scale models are valuable tools for identifying novel targets for metabolic engineering, a stoichiometric model of chondroitin-producing S. cerevisiae was developed and used in optimization algorithms. Our research yielded several novel targets, such as uridine diphosphate (UDP)-N-acetylglucosamine pyrophosphorylase (QRI1), glucosamine-6-phosphate acetyltransferase (GNA1), or N-acetylglucosamine-phosphate mutase (PCM1) overexpression, that might enhance chondroitin production and guide future experimental research to develop more efficient host organisms for the biotechnological production process.
Full article
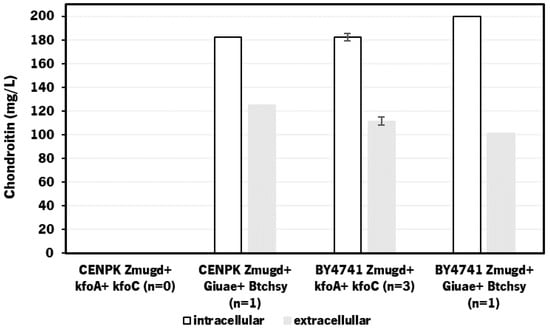
Figure 1
Figure 1
<p>Chondroitin production in engineered <span class="html-italic">Saccharomyces cerevisiae</span> CEN.PK2-1C (CENPK) and BY4741 strains. Btchsy: chondroitin synthase/polymerase (CHSY) from <span class="html-italic">Bos taurus</span>; Giuae: UDP-<span class="html-italic">N</span>-acetylglucosamine 4-epimerase (UAE) from <span class="html-italic">Giardia intestinalis</span>; kfoA and kfoC: UAE and CHSY, respectively, from <span class="html-italic">Escherichia coli</span> K4 (serotype O5:K4(L):H4); Zmugd: uridine diphosphate (UDP)-glucose 6-dehydrogenase (UGD) from <span class="html-italic">Zymomonas mobilis</span>.</p> Full article ">Figure 2
<p>Pathways involved in chondroitin production in engineered <span class="html-italic">Saccharomyces cerevisiae</span>, and competing pathways that redirect the metabolic flux from chondroitin precursors. Compound abbreviations: CoA—coenzyme A; NAD<sup>+</sup>—nicotinamide adenine dinucleotide, oxidized form; NADH—nicotinamide adenine dinucleotide, reduced form; PPi—diphosphate; GPI—glycosylphosphatidylinositol; UDP—uridine diphosphate; UTP—uridine triphosphate; Gene descriptions: <span class="html-italic">ALG11</span>—Alpha-1,2-mannosyltransferase; <span class="html-italic">ALG12</span>—Alpha-1,6-mannosyltransferase; <span class="html-italic">ALG9</span>—mannosyltransferase; <span class="html-italic">ALG5</span>—UDP-glucose:dolichyl-phosphate glucosyltransferase; <span class="html-italic">ATH1</span>—acid trehalase; <span class="html-italic">ATG26</span>—UDP-glucose:sterol glucosyltransferase; <span class="html-italic">CDA1</span>, <span class="html-italic">CDA2</span>—chitin deacetylase; <span class="html-italic">CHS1</span>, <span class="html-italic">CHS2</span>, <span class="html-italic">CHS3</span>—chitin synthases; <span class="html-italic">CHSY</span>—chondroitin synthase; <span class="html-italic">DPM1</span>—dolichol phosphate mannose synthase; <span class="html-italic">EMI2</span>—hexokinase; <span class="html-italic">ERI1</span>—endoplasmic reticulum-associated Ras Inhibitor; <span class="html-italic">FBA1</span>—fructose 1,6-bisphosphate aldolase; <span class="html-italic">FBP1</span>—fructose-1,6-bisphosphatase; <span class="html-italic">FKS1</span>, <span class="html-italic">FKS3</span>—1,3-β-D-glucan synthase; <span class="html-italic">GAL10</span>—UDP-glucose-4-epimerase; <span class="html-italic">GLG1</span>, <span class="html-italic">GLG2</span>—glycogenin glucosyltransferase; <span class="html-italic">GLK1</span>—glucokinase; <span class="html-italic">GNA1</span>—glucosamine-6-phosphate <span class="html-italic">N</span>-acetyltransferase; <span class="html-italic">GPD1, GPD2</span>—glycerol-3-phosphate dehydrogenases; <span class="html-italic">GPI1</span>, <span class="html-italic">GPI10</span>, <span class="html-italic">GPI14</span>, <span class="html-italic">GPI15</span>, <span class="html-italic">GPI18</span>, <span class="html-italic">GPI19</span>—GPI anchor proteins; <span class="html-italic">GPP1</span>, <span class="html-italic">GPP2</span>—glycerol-3-phosphate phosphatases; <span class="html-italic">GSC2</span>—1,3-β-glucan synthase; <span class="html-italic">GSY1</span>, <span class="html-italic">GSY2</span>—glycogen synthases; <span class="html-italic">GUT1</span>—glycerol kinase; <span class="html-italic">GUT2</span>—glycerol-3-phosphate dehydrogenase; <span class="html-italic">HXK1</span>, <span class="html-italic">HXK2</span>—hexokinases; <span class="html-italic">INO1</span>—inositol-3-phosphate synthase; <span class="html-italic">INM1</span>, <span class="html-italic">INM2</span>—inositol monophosphatases; <span class="html-italic">NTH1</span>, <span class="html-italic">NTH2</span>—neutral trehalases; <span class="html-italic">PBN1</span>—glycosylphosphatidylinositol-mannosyltransferase I; <span class="html-italic">PCM1</span>—<span class="html-italic">N</span>-acetylglucosamine-phosphate mutase; <span class="html-italic">PFK1, PFK2</span>—phosphofructokinase; <span class="html-italic">PGI1</span>—phosphoglucose isomerase; <span class="html-italic">PMT1</span>, <span class="html-italic">PMT2</span>, <span class="html-italic">PMT3</span>, <span class="html-italic">PMT4</span>, <span class="html-italic">PMT5</span>, <span class="html-italic">PMT6</span>, <span class="html-italic">PMT7</span>—protein <span class="html-italic">O</span>-mannosyltransferases; <span class="html-italic">PSA1</span>—guanosine diphosphate(GDP)-mannose pyrophosphorylase; <span class="html-italic">QRI1</span>—UDP-<span class="html-italic">N</span>-acetylglucosamine pyrophosphorylase; <span class="html-italic">SEC53</span>—phosphomannomutase; <span class="html-italic">SMP3</span>—alpha 1,2-mannosyltransferase; <span class="html-italic">SPT14</span>—UDP-glycosyltransferase; <span class="html-italic">SUC2</span>—invertase; <span class="html-italic">TPS1</span>—trehalose-6-phosphate synthase; <span class="html-italic">TPS2</span>—trehalose-phosphatase; <span class="html-italic">TPS3</span>—trehalose-6-phosphatase; <span class="html-italic">UAE</span>—UDP-<span class="html-italic">N</span>-acetylglucosamine 4’-epimerase; <span class="html-italic">UGD</span>—UDP-glucose 6-dehydrogenase; <span class="html-italic">ZWF1</span>—glucose-6-phosphate dehydrogenase.</p> Full article ">Figure 3
<p>Frequency and expression values of genes in the solutions from optimization of <span class="html-italic">Saccharomyces cerevisiae</span> model for chondroitin production using MEWpy tool, limiting number of modifications to 10. The mutant expression (in dots) represents the average expression value. Mutant expressions higher than 1 represent overexpression while values of expression lower than 1 represent underexpression. Deletion is represented using a light grey bar. Gene descriptions: <span class="html-italic">AAT1</span>—mitochondrial aspartate aminotransferase; <span class="html-italic">ABZ1</span>—para-aminobenzoate synthase; <span class="html-italic">ADE3</span>—cytoplasmic trifunctional enzyme; ALD2—cytoplasmic aldehyde dehydrogenase; <span class="html-italic">ALG1</span>—mannosyltransferase; <span class="html-italic">APE1</span>—vacuolar aminopeptidase; <span class="html-italic">ARG8</span>—acetylornithine aminotransferase; <span class="html-italic">ASN1</span>—asparagine synthetase; <span class="html-italic">ARO2</span>—bifunctional chorismate synthase and flavin reductase; BDH1—NAD-dependent (R,R)-butanediol dehydrogenase; <span class="html-italic">CAB4/5</span>—subunits of the CoA-synthesizing protein complex; <span class="html-italic">COX16</span>—mitochondrial inner membrane protein; <span class="html-italic">CRD1</span>—cardiolipin synthase; <span class="html-italic">CTA1</span>—catalase A; <span class="html-italic">CTS1</span>—endochitinase; <span class="html-italic">DAL7</span>—malate synthase; <span class="html-italic">DFR1</span>—dihydrofolate reductase; <span class="html-italic">DNF2</span>—aminophospholipid translocase; ECM4—S-glutathionyl-(chloro)hydroquinone reductase; <span class="html-italic">DPS1</span>- aspartyl-tRNA synthetase; <span class="html-italic">ELO2</span>—fatty acid elongase; <span class="html-italic">ENO1</span>—enolase I; <span class="html-italic">ERG10</span>—acetyl-CoA C-acetyltransferase; <span class="html-italic">FAA1</span>—long chain fatty acyl-CoA synthetase; <span class="html-italic">FDH1</span>—NAD(+)-dependent formate dehydrogenase; FOL1—multifunctional enzyme of the folic acid biosynthesis pathway; <span class="html-italic">GTT1</span>—glutathione S-transferase; <span class="html-italic">GWT1</span>—phosphatidylinositol glycan anchored wall transfer protein; <span class="html-italic">HMG1</span>—3-hydroxy-3-methylglutaryl-CoA reductase; <span class="html-italic">LYP1</span>—lysine permease; <span class="html-italic">MAL31</span>—maltose permease; <span class="html-italic">MAE1</span>—mitochondrial malic enzyme; <span class="html-italic">NAT1</span>—subunit of protein N-terminal acetyltransferase; <span class="html-italic">PCM1</span>—N-acetylglucosamine-phosphate mutase; <span class="html-italic">PDC6</span>—minor isoform of pyruvate decarboxylase; <span class="html-italic">PGM2</span>—phosphoglucomutase; <span class="html-italic">PHO89</span>—plasma membrane Na+/Pi cotransporter; PMP2—proteolipid associated with plasma membrane H(+)-ATPase; <span class="html-italic">PMT2</span>—Protein O-mannosyltransferase; <span class="html-italic">POF1</span>—nicotinamide mononucleotide-specific adenylyltransferase; <span class="html-italic">POT1</span>—3-ketoacyl-CoA thiolase; <span class="html-italic">PUS2</span>—mitochondrial tRNA:pseudouridine synthase; <span class="html-italic">QCR7</span>—subunit 7 of ubiquinol cytochrome-c reductase; <span class="html-italic">QRI1</span>—uridine diphosphate-<span class="html-italic">N</span>-acetylglucosamine pyrophosphorylase; <span class="html-italic">SCS7</span>—sphingolipid alpha-hydroxylase; <span class="html-italic">SER1</span>—3-phosphoserine aminotransferase; SLC1—1-acyl-sn-glycerol-3-phosphate acyltransferase; <span class="html-italic">SPE1</span>—ornithine decarboxylase; <span class="html-italic">SOR2</span>—sorbitol dehydrogenase; <span class="html-italic">TAZ1</span>—lyso-phosphatidylcholine acyltransferase; <span class="html-italic">YAH1</span>—ferredoxin of the mitochondrial matrix; <span class="html-italic">YJU3</span>—monoglyceride lipase.</p> Full article ">Figure 4
<p>Frequency and expression values of genes in the solutions from optimization of <span class="html-italic">Saccharomyces cerevisiae</span> model using MEWpy tool, limiting number of modifications to 3. The mutant expression (in dots) represents the average expression value. Mutant gene expression values higher than 1 represent overexpression, while values of expression lower than 1 represent underexpression. Deletions are represented in light grey bars. Gene descriptions: <span class="html-italic">ARO2</span>—bifunctional chorismate synthase and flavin reductase; <span class="html-italic">CDS1</span>—phosphatidate cytidylyltransferase; <span class="html-italic">CTM1</span>—cytochrome c lysine methyltransferase; <span class="html-italic">ERG11</span>—lanosterol 14-alpha-demethylase; <span class="html-italic">GUA1</span>—guanosine monophosphate synthase; <span class="html-italic">HAM1</span>—nucleoside triphosphate pyrophosphohydrolase; <span class="html-italic">HMG2</span>—3-hydroxy-3-methylglutaryl-CoA reductase; <span class="html-italic">POT1</span>—3-ketoacyl-CoA thiolase; <span class="html-italic">QRI1</span>—uridine diphosphate-<span class="html-italic">N</span>-acetylglucosamine pyrophosphorylase; <span class="html-italic">SAH1</span>—S-adenosyl-L-homocysteine hydrolase; <span class="html-italic">SDH2</span>—iron–sulfur protein subunit of succinate dehydrogenase; <span class="html-italic">THI21</span>—hydroxymethylpyrimidine (HMP) and HMP-phosphate kinase; <span class="html-italic">YDR341C</span>—arginyl-tRNA synthetase; <span class="html-italic">YNK1</span>—nucleoside diphosphate kinase.</p> Full article ">
<p>Chondroitin production in engineered <span class="html-italic">Saccharomyces cerevisiae</span> CEN.PK2-1C (CENPK) and BY4741 strains. Btchsy: chondroitin synthase/polymerase (CHSY) from <span class="html-italic">Bos taurus</span>; Giuae: UDP-<span class="html-italic">N</span>-acetylglucosamine 4-epimerase (UAE) from <span class="html-italic">Giardia intestinalis</span>; kfoA and kfoC: UAE and CHSY, respectively, from <span class="html-italic">Escherichia coli</span> K4 (serotype O5:K4(L):H4); Zmugd: uridine diphosphate (UDP)-glucose 6-dehydrogenase (UGD) from <span class="html-italic">Zymomonas mobilis</span>.</p> Full article ">Figure 2
<p>Pathways involved in chondroitin production in engineered <span class="html-italic">Saccharomyces cerevisiae</span>, and competing pathways that redirect the metabolic flux from chondroitin precursors. Compound abbreviations: CoA—coenzyme A; NAD<sup>+</sup>—nicotinamide adenine dinucleotide, oxidized form; NADH—nicotinamide adenine dinucleotide, reduced form; PPi—diphosphate; GPI—glycosylphosphatidylinositol; UDP—uridine diphosphate; UTP—uridine triphosphate; Gene descriptions: <span class="html-italic">ALG11</span>—Alpha-1,2-mannosyltransferase; <span class="html-italic">ALG12</span>—Alpha-1,6-mannosyltransferase; <span class="html-italic">ALG9</span>—mannosyltransferase; <span class="html-italic">ALG5</span>—UDP-glucose:dolichyl-phosphate glucosyltransferase; <span class="html-italic">ATH1</span>—acid trehalase; <span class="html-italic">ATG26</span>—UDP-glucose:sterol glucosyltransferase; <span class="html-italic">CDA1</span>, <span class="html-italic">CDA2</span>—chitin deacetylase; <span class="html-italic">CHS1</span>, <span class="html-italic">CHS2</span>, <span class="html-italic">CHS3</span>—chitin synthases; <span class="html-italic">CHSY</span>—chondroitin synthase; <span class="html-italic">DPM1</span>—dolichol phosphate mannose synthase; <span class="html-italic">EMI2</span>—hexokinase; <span class="html-italic">ERI1</span>—endoplasmic reticulum-associated Ras Inhibitor; <span class="html-italic">FBA1</span>—fructose 1,6-bisphosphate aldolase; <span class="html-italic">FBP1</span>—fructose-1,6-bisphosphatase; <span class="html-italic">FKS1</span>, <span class="html-italic">FKS3</span>—1,3-β-D-glucan synthase; <span class="html-italic">GAL10</span>—UDP-glucose-4-epimerase; <span class="html-italic">GLG1</span>, <span class="html-italic">GLG2</span>—glycogenin glucosyltransferase; <span class="html-italic">GLK1</span>—glucokinase; <span class="html-italic">GNA1</span>—glucosamine-6-phosphate <span class="html-italic">N</span>-acetyltransferase; <span class="html-italic">GPD1, GPD2</span>—glycerol-3-phosphate dehydrogenases; <span class="html-italic">GPI1</span>, <span class="html-italic">GPI10</span>, <span class="html-italic">GPI14</span>, <span class="html-italic">GPI15</span>, <span class="html-italic">GPI18</span>, <span class="html-italic">GPI19</span>—GPI anchor proteins; <span class="html-italic">GPP1</span>, <span class="html-italic">GPP2</span>—glycerol-3-phosphate phosphatases; <span class="html-italic">GSC2</span>—1,3-β-glucan synthase; <span class="html-italic">GSY1</span>, <span class="html-italic">GSY2</span>—glycogen synthases; <span class="html-italic">GUT1</span>—glycerol kinase; <span class="html-italic">GUT2</span>—glycerol-3-phosphate dehydrogenase; <span class="html-italic">HXK1</span>, <span class="html-italic">HXK2</span>—hexokinases; <span class="html-italic">INO1</span>—inositol-3-phosphate synthase; <span class="html-italic">INM1</span>, <span class="html-italic">INM2</span>—inositol monophosphatases; <span class="html-italic">NTH1</span>, <span class="html-italic">NTH2</span>—neutral trehalases; <span class="html-italic">PBN1</span>—glycosylphosphatidylinositol-mannosyltransferase I; <span class="html-italic">PCM1</span>—<span class="html-italic">N</span>-acetylglucosamine-phosphate mutase; <span class="html-italic">PFK1, PFK2</span>—phosphofructokinase; <span class="html-italic">PGI1</span>—phosphoglucose isomerase; <span class="html-italic">PMT1</span>, <span class="html-italic">PMT2</span>, <span class="html-italic">PMT3</span>, <span class="html-italic">PMT4</span>, <span class="html-italic">PMT5</span>, <span class="html-italic">PMT6</span>, <span class="html-italic">PMT7</span>—protein <span class="html-italic">O</span>-mannosyltransferases; <span class="html-italic">PSA1</span>—guanosine diphosphate(GDP)-mannose pyrophosphorylase; <span class="html-italic">QRI1</span>—UDP-<span class="html-italic">N</span>-acetylglucosamine pyrophosphorylase; <span class="html-italic">SEC53</span>—phosphomannomutase; <span class="html-italic">SMP3</span>—alpha 1,2-mannosyltransferase; <span class="html-italic">SPT14</span>—UDP-glycosyltransferase; <span class="html-italic">SUC2</span>—invertase; <span class="html-italic">TPS1</span>—trehalose-6-phosphate synthase; <span class="html-italic">TPS2</span>—trehalose-phosphatase; <span class="html-italic">TPS3</span>—trehalose-6-phosphatase; <span class="html-italic">UAE</span>—UDP-<span class="html-italic">N</span>-acetylglucosamine 4’-epimerase; <span class="html-italic">UGD</span>—UDP-glucose 6-dehydrogenase; <span class="html-italic">ZWF1</span>—glucose-6-phosphate dehydrogenase.</p> Full article ">Figure 3
<p>Frequency and expression values of genes in the solutions from optimization of <span class="html-italic">Saccharomyces cerevisiae</span> model for chondroitin production using MEWpy tool, limiting number of modifications to 10. The mutant expression (in dots) represents the average expression value. Mutant expressions higher than 1 represent overexpression while values of expression lower than 1 represent underexpression. Deletion is represented using a light grey bar. Gene descriptions: <span class="html-italic">AAT1</span>—mitochondrial aspartate aminotransferase; <span class="html-italic">ABZ1</span>—para-aminobenzoate synthase; <span class="html-italic">ADE3</span>—cytoplasmic trifunctional enzyme; ALD2—cytoplasmic aldehyde dehydrogenase; <span class="html-italic">ALG1</span>—mannosyltransferase; <span class="html-italic">APE1</span>—vacuolar aminopeptidase; <span class="html-italic">ARG8</span>—acetylornithine aminotransferase; <span class="html-italic">ASN1</span>—asparagine synthetase; <span class="html-italic">ARO2</span>—bifunctional chorismate synthase and flavin reductase; BDH1—NAD-dependent (R,R)-butanediol dehydrogenase; <span class="html-italic">CAB4/5</span>—subunits of the CoA-synthesizing protein complex; <span class="html-italic">COX16</span>—mitochondrial inner membrane protein; <span class="html-italic">CRD1</span>—cardiolipin synthase; <span class="html-italic">CTA1</span>—catalase A; <span class="html-italic">CTS1</span>—endochitinase; <span class="html-italic">DAL7</span>—malate synthase; <span class="html-italic">DFR1</span>—dihydrofolate reductase; <span class="html-italic">DNF2</span>—aminophospholipid translocase; ECM4—S-glutathionyl-(chloro)hydroquinone reductase; <span class="html-italic">DPS1</span>- aspartyl-tRNA synthetase; <span class="html-italic">ELO2</span>—fatty acid elongase; <span class="html-italic">ENO1</span>—enolase I; <span class="html-italic">ERG10</span>—acetyl-CoA C-acetyltransferase; <span class="html-italic">FAA1</span>—long chain fatty acyl-CoA synthetase; <span class="html-italic">FDH1</span>—NAD(+)-dependent formate dehydrogenase; FOL1—multifunctional enzyme of the folic acid biosynthesis pathway; <span class="html-italic">GTT1</span>—glutathione S-transferase; <span class="html-italic">GWT1</span>—phosphatidylinositol glycan anchored wall transfer protein; <span class="html-italic">HMG1</span>—3-hydroxy-3-methylglutaryl-CoA reductase; <span class="html-italic">LYP1</span>—lysine permease; <span class="html-italic">MAL31</span>—maltose permease; <span class="html-italic">MAE1</span>—mitochondrial malic enzyme; <span class="html-italic">NAT1</span>—subunit of protein N-terminal acetyltransferase; <span class="html-italic">PCM1</span>—N-acetylglucosamine-phosphate mutase; <span class="html-italic">PDC6</span>—minor isoform of pyruvate decarboxylase; <span class="html-italic">PGM2</span>—phosphoglucomutase; <span class="html-italic">PHO89</span>—plasma membrane Na+/Pi cotransporter; PMP2—proteolipid associated with plasma membrane H(+)-ATPase; <span class="html-italic">PMT2</span>—Protein O-mannosyltransferase; <span class="html-italic">POF1</span>—nicotinamide mononucleotide-specific adenylyltransferase; <span class="html-italic">POT1</span>—3-ketoacyl-CoA thiolase; <span class="html-italic">PUS2</span>—mitochondrial tRNA:pseudouridine synthase; <span class="html-italic">QCR7</span>—subunit 7 of ubiquinol cytochrome-c reductase; <span class="html-italic">QRI1</span>—uridine diphosphate-<span class="html-italic">N</span>-acetylglucosamine pyrophosphorylase; <span class="html-italic">SCS7</span>—sphingolipid alpha-hydroxylase; <span class="html-italic">SER1</span>—3-phosphoserine aminotransferase; SLC1—1-acyl-sn-glycerol-3-phosphate acyltransferase; <span class="html-italic">SPE1</span>—ornithine decarboxylase; <span class="html-italic">SOR2</span>—sorbitol dehydrogenase; <span class="html-italic">TAZ1</span>—lyso-phosphatidylcholine acyltransferase; <span class="html-italic">YAH1</span>—ferredoxin of the mitochondrial matrix; <span class="html-italic">YJU3</span>—monoglyceride lipase.</p> Full article ">Figure 4
<p>Frequency and expression values of genes in the solutions from optimization of <span class="html-italic">Saccharomyces cerevisiae</span> model using MEWpy tool, limiting number of modifications to 3. The mutant expression (in dots) represents the average expression value. Mutant gene expression values higher than 1 represent overexpression, while values of expression lower than 1 represent underexpression. Deletions are represented in light grey bars. Gene descriptions: <span class="html-italic">ARO2</span>—bifunctional chorismate synthase and flavin reductase; <span class="html-italic">CDS1</span>—phosphatidate cytidylyltransferase; <span class="html-italic">CTM1</span>—cytochrome c lysine methyltransferase; <span class="html-italic">ERG11</span>—lanosterol 14-alpha-demethylase; <span class="html-italic">GUA1</span>—guanosine monophosphate synthase; <span class="html-italic">HAM1</span>—nucleoside triphosphate pyrophosphohydrolase; <span class="html-italic">HMG2</span>—3-hydroxy-3-methylglutaryl-CoA reductase; <span class="html-italic">POT1</span>—3-ketoacyl-CoA thiolase; <span class="html-italic">QRI1</span>—uridine diphosphate-<span class="html-italic">N</span>-acetylglucosamine pyrophosphorylase; <span class="html-italic">SAH1</span>—S-adenosyl-L-homocysteine hydrolase; <span class="html-italic">SDH2</span>—iron–sulfur protein subunit of succinate dehydrogenase; <span class="html-italic">THI21</span>—hydroxymethylpyrimidine (HMP) and HMP-phosphate kinase; <span class="html-italic">YDR341C</span>—arginyl-tRNA synthetase; <span class="html-italic">YNK1</span>—nucleoside diphosphate kinase.</p> Full article ">
Open AccessArticle
Development of SynBio Tools for Pseudomonas chlororaphis: A Versatile Non-Pathogenic Bacterium Host
by
Miguel Angel Bello-González, Leidy Patricia Bedoya-Perez, Miguel Alberto Pantoja-Zepeda and Jose Utrilla
SynBio 2024, 2(2), 112-124; https://doi.org/10.3390/synbio2020007 - 27 Mar 2024
Abstract
►▼
Show Figures
Pseudomonas chlororaphis ATCC 9446 is a non-pathogenic bacterium associated with the rhizosphere. It is commonly used as a biocontrol agent against agricultural pests. This organism can grow on a variety of carbon sources, has a robust secondary metabolism, and produces secondary metabolites with
[...] Read more.
Pseudomonas chlororaphis ATCC 9446 is a non-pathogenic bacterium associated with the rhizosphere. It is commonly used as a biocontrol agent against agricultural pests. This organism can grow on a variety of carbon sources, has a robust secondary metabolism, and produces secondary metabolites with antimicrobial properties. This makes it an alternative host organism for synthetic biology applications. However, as a novel host there is a need for well-characterized molecular tools that allow fine control of gene expression and exploration of its metabolic potential. In this work we developed and characterized expression vectors for P. chlororaphis. We used two different promoters: the exogenously induced lac-IPTG promoter, and LuxR-C6-AHL, which we evaluated for its auto-inducible capacities, as well as using an external addition of C6-AHL. The expression response of these vectors to the inducer concentration was characterized by detecting a reporter fluorescent protein (YFP: yellow fluorescent protein). Furthermore, the violacein production operon was evaluated as a model heterologous pathway. We tested violacein production in shake flasks and a 3 L fermenter, showing that P. chlororaphis possesses a vigorous aromatic amino acid metabolism and was able to produce 1 g/L of violacein in a simple batch reactor experiment with minimal medium using only glucose as the carbon source. We compared the experimental results with the predictions of a modified genome scale model. The presented results show the potential of P. chlororaphis as a novel host organism for synthetic biology applications.
Full article
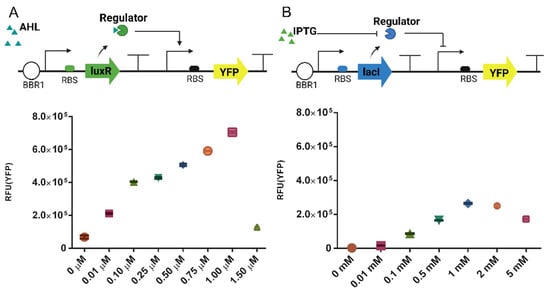
Figure 1
Figure 1
<p>Plasmid design and characterization of dose response to inducer concentration. (<b>A</b>) Displays the organization of the SAY-AHL vector. BBR1: origin of replication, luxR regulators, YFP reporter gene. The chart displays the expression response of YFP, measured in relative fluorescence units (RFU), to a gradient of concentration of exogenous C6-AHL. (<b>B</b>) Displays the organization of the SAY-IPTG vector. BBR1 origin of replication, lacI regulators. The chart displays the expression response of YFP, measured in RFUs, to a gradient of concentration of exogenous IPTG. Normalized data and optical density (OD) at sampling points are presented in <a href="#app1-synbio-02-00007" class="html-app">Figure S5</a>.</p> Full article ">Figure 2
<p>Auto-induced expression mediated by AHL synthase phzI. (<b>A</b>) Auto-induction circuit mediated by homologous AHL synthase phzI and Δ<span class="html-italic">phzI</span> mutant. (<b>B</b>) Fluorescence in <span class="html-italic">P. chlororaphis</span>, using exogenous addition of C6-AHL at 1 µM on wild-type <span class="html-italic">P. chlororaphis</span> and Δ<span class="html-italic">phzI</span> mutant. (<b>C</b>) Test of auto-induction capacity of SAY-AHL-YFP vector in wild-type (WT) and Δ<span class="html-italic">phzI</span> mutant strains. SAY-IPTG at 0 mM induction is shown as a reference. or better data visualization, two different scales are used in <span class="html-italic">Y</span>-axis at panels (<b>B</b>,<b>C</b>).</p> Full article ">Figure 3
<p>Violacein production in <span class="html-italic">P. chlororaphis</span>. (<b>A</b>) Violacein production operon in SAY-AHL-Vio and SAY-IPTG-Vio plasmids. (<b>B</b>) Violacein production M9 medium with sucrose 50 g/L or (<b>C</b>) glucose 50 g/L as the carbon source.</p> Full article ">Figure 4
<p>Violacein production in a 3 L fermenter in batch mode. (<b>A</b>) Total violacein production. (<b>B</b>) Normalized violacein production by measured protein.</p> Full article ">Figure 5
<p>Genome scale model simulations. Simulation of growth on glucose and sucrose at different substrate uptake rates. (<b>A</b>) Pathway reconstructed in the model iMA1267_VIO from tryptophan to violacein. (<b>B</b>) Effect of the carbon source consumption rate on the growth rate. The graph shows that with glucose (red) as the only carbon source, the growth rate is lower compared to sucrose (blue). We see how ammonium consumption in both carbon sources is very similar (light blue and yellow). (<b>C</b>) The violacein production envelope was calculated using the iMA1267_VIO model, modified with the incorporation of the violacein pathway. Thus, the green line and the shaded area represent the potential solution area for the model’s production with the production of violacein and biomass.</p> Full article ">
<p>Plasmid design and characterization of dose response to inducer concentration. (<b>A</b>) Displays the organization of the SAY-AHL vector. BBR1: origin of replication, luxR regulators, YFP reporter gene. The chart displays the expression response of YFP, measured in relative fluorescence units (RFU), to a gradient of concentration of exogenous C6-AHL. (<b>B</b>) Displays the organization of the SAY-IPTG vector. BBR1 origin of replication, lacI regulators. The chart displays the expression response of YFP, measured in RFUs, to a gradient of concentration of exogenous IPTG. Normalized data and optical density (OD) at sampling points are presented in <a href="#app1-synbio-02-00007" class="html-app">Figure S5</a>.</p> Full article ">Figure 2
<p>Auto-induced expression mediated by AHL synthase phzI. (<b>A</b>) Auto-induction circuit mediated by homologous AHL synthase phzI and Δ<span class="html-italic">phzI</span> mutant. (<b>B</b>) Fluorescence in <span class="html-italic">P. chlororaphis</span>, using exogenous addition of C6-AHL at 1 µM on wild-type <span class="html-italic">P. chlororaphis</span> and Δ<span class="html-italic">phzI</span> mutant. (<b>C</b>) Test of auto-induction capacity of SAY-AHL-YFP vector in wild-type (WT) and Δ<span class="html-italic">phzI</span> mutant strains. SAY-IPTG at 0 mM induction is shown as a reference. or better data visualization, two different scales are used in <span class="html-italic">Y</span>-axis at panels (<b>B</b>,<b>C</b>).</p> Full article ">Figure 3
<p>Violacein production in <span class="html-italic">P. chlororaphis</span>. (<b>A</b>) Violacein production operon in SAY-AHL-Vio and SAY-IPTG-Vio plasmids. (<b>B</b>) Violacein production M9 medium with sucrose 50 g/L or (<b>C</b>) glucose 50 g/L as the carbon source.</p> Full article ">Figure 4
<p>Violacein production in a 3 L fermenter in batch mode. (<b>A</b>) Total violacein production. (<b>B</b>) Normalized violacein production by measured protein.</p> Full article ">Figure 5
<p>Genome scale model simulations. Simulation of growth on glucose and sucrose at different substrate uptake rates. (<b>A</b>) Pathway reconstructed in the model iMA1267_VIO from tryptophan to violacein. (<b>B</b>) Effect of the carbon source consumption rate on the growth rate. The graph shows that with glucose (red) as the only carbon source, the growth rate is lower compared to sucrose (blue). We see how ammonium consumption in both carbon sources is very similar (light blue and yellow). (<b>C</b>) The violacein production envelope was calculated using the iMA1267_VIO model, modified with the incorporation of the violacein pathway. Thus, the green line and the shaded area represent the potential solution area for the model’s production with the production of violacein and biomass.</p> Full article ">
Open AccessReview
Expanding the Biosynthetic Toolbox: The Potential and Challenges of In Vitro Type II Polyketide Synthase Research
by
Max A. J. Rivers and Andrew N. Lowell
SynBio 2024, 2(1), 85-111; https://doi.org/10.3390/synbio2010006 - 7 Mar 2024
Abstract
►▼
Show Figures
Type II polyketide synthase (PKS) systems are a rich source of structurally diverse polycyclic aromatic compounds with clinically relevant antibiotic and chemotherapeutic properties. The enzymes responsible for synthesizing the polyketide core, known collectively as the minimal cassette, hold potential for applications in synthetic
[...] Read more.
Type II polyketide synthase (PKS) systems are a rich source of structurally diverse polycyclic aromatic compounds with clinically relevant antibiotic and chemotherapeutic properties. The enzymes responsible for synthesizing the polyketide core, known collectively as the minimal cassette, hold potential for applications in synthetic biology. The minimal cassette provides polyketides of different chain lengths, which interact with other enzymes that are responsible for the varied cyclization patterns. Additionally, the type II PKS enzyme clusters offer a wide repertoire of tailoring enzymes for oxidations, glycosylations, cyclizations, and rearrangements. This review begins with the variety of chemical space accessible with type II PKS systems including the recently discovered highly reducing variants that produce polyalkenes instead of the archetypical polyketide motif. The main discussion analyzes the previous approaches with an emphasis on further research that is needed to characterize the minimal cassette enzymes in vitro. Finally, the potential type II PKS systems hold the potential to offer new tools in biocatalysis and synthetic biology, particularly in the production of novel antibiotics and biofuels.
Full article
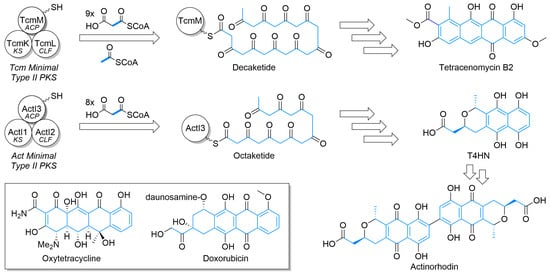
Figure 1
Figure 1
<p>The basic construction of the polyketides of tetracenomycin and actinorhodin by type II polyketide synthases and elaboration to aromatic polycycle. The minimal type II polyketide synthase (PKS) cassette, composed of an acyl carrier protein (ACP), ketosynthase (KS), and chain length factor (CLF), work in concert to produce various length polyketides that are subsequently elaborated by other enzymes to the natural product. Carbon atoms from the starter unit or malonyl extender units are highlighted in blue. <b>Inset</b>—Structures of oxytetracycline and doxorubicin.</p> Full article ">Figure 2
<p>Comparison of the core architectures and components of fatty acid and polyketide synthase proteins with example natural products from each class. All types rely on a core KS to conduct a Claisen reaction to extend the growing chain by an acetate unit. Different oxidation states can be achieved by the incorporation of additional domains, either within a megasynthase or as discrete proteins. A legend of abbreviations is shown on the right of the Figure. Domains marked with an asterisk are not always present.</p> Full article ">Figure 3
<p>Major classes of type II PKS products, benzoisochromanquinones, anthracyclines (see also <a href="#synbio-02-00006-f001" class="html-fig">Figure 1</a> for doxorubicin), tetracenomycins (also tetracenomycin B2 in <a href="#synbio-02-00006-f001" class="html-fig">Figure 1</a>), aureolic acids, tetracyclines (also oxytetracycline in <a href="#synbio-02-00006-f001" class="html-fig">Figure 1</a>), and angucyclines.</p> Full article ">Figure 4
<p>Examples of type II PKS-derived natural products including extended angular polyphenols, polycycles with different starter units, unusual cyclization patterns, and structures with other atom incorporation beyond C, H, and O.</p> Full article ">Figure 5
<p>Examples of type II PKS products with post-cyclization rearrangement or atom excision.</p> Full article ">Figure 6
<p>Claisen-like condensation mechanism of the minimal cassette in type II PKS to grow the polyketide chain. Once the KS is loaded with a starter unit (<b>A</b>) via one of several mechanisms, the ACP interacts with malonyl-CoA or transacylases from primary metabolism to obtain a malonyl for chain extension (<b>B</b>). Decarboxylation activates the malonyl as an enolate on the ACP, which reacts with the thioester on the KS to form a tetrahedral intermediate (<b>C</b>). Collapse of the tetrahedral intermediate results in a polyketide extended by one acetate unit attached to the ACP (<b>D</b>). Continued extension involves the transfer of the extended polyketide back to the KS (<b>E</b>) and the acquisition of another malonyl by the ACP to reenter the extension cycle (<b>E</b> to <b>B</b>). Once the ACP-bound polyketide chain is at the appropriate length, it can no longer be accepted back onto the KS (see later Figure) causing dissociation of the ACP from the KS-CLF dimer (<b>F</b>) and enabling delivery of the polyketide chain to cyclases for tailoring to various ring systems.</p> Full article ">Figure 7
<p>(<b>A</b>) The structures of coenzyme A (the phosphopantetheine is highlighted in blue) and the abbreviated form of the malonyl-CoA unit. (<b>B</b>) Conversion of <span class="html-italic">apo</span>-ACP to <span class="html-italic">holo</span>-ACP by the action of PPTase. The squiggly line terminating in a thiol in the cartoon to the right represents the flexible phosphopantetheine arm.</p> Full article ">Figure 8
<p>Cartoon depicting the gating residues [<a href="#B16-synbio-02-00006" class="html-bibr">16</a>] for octaketide (<b>top</b>) production in the actinorhodin system and (<b>bottom</b>) the corresponding residues for tridecaketide production in the griseorhodin system. Residues from the CLF are shown in green while residues from the KS are shown in blue. The larger threonine112 and phenylalanine109 (<b>top</b>) cause growth to stop at the octaketide because the octaketide cannot be accommodated in the binding tunnel and thus cannot be transferred back to the KS for a continued extension. These are replaced by smaller residues (<b>bottom</b>) for the tridecaketide, which terminate at methionine143 and tyrosine140.</p> Full article ">Figure 9
<p>Natural products that utilize HR type II PKS in their biosynthesis. The polyene(s) derived from the HR type II PKS are highlighted in blue. Simocyclinone D8 also includes a canonical type II PKS core (blue bonds). <span class="html-italic">Inset</span>: The structure of cispentacin, a non-canonical amino acid formed through the action of a type-II-PKS-like minimal cassette [<a href="#B191-synbio-02-00006" class="html-bibr">191</a>].</p> Full article ">
<p>The basic construction of the polyketides of tetracenomycin and actinorhodin by type II polyketide synthases and elaboration to aromatic polycycle. The minimal type II polyketide synthase (PKS) cassette, composed of an acyl carrier protein (ACP), ketosynthase (KS), and chain length factor (CLF), work in concert to produce various length polyketides that are subsequently elaborated by other enzymes to the natural product. Carbon atoms from the starter unit or malonyl extender units are highlighted in blue. <b>Inset</b>—Structures of oxytetracycline and doxorubicin.</p> Full article ">Figure 2
<p>Comparison of the core architectures and components of fatty acid and polyketide synthase proteins with example natural products from each class. All types rely on a core KS to conduct a Claisen reaction to extend the growing chain by an acetate unit. Different oxidation states can be achieved by the incorporation of additional domains, either within a megasynthase or as discrete proteins. A legend of abbreviations is shown on the right of the Figure. Domains marked with an asterisk are not always present.</p> Full article ">Figure 3
<p>Major classes of type II PKS products, benzoisochromanquinones, anthracyclines (see also <a href="#synbio-02-00006-f001" class="html-fig">Figure 1</a> for doxorubicin), tetracenomycins (also tetracenomycin B2 in <a href="#synbio-02-00006-f001" class="html-fig">Figure 1</a>), aureolic acids, tetracyclines (also oxytetracycline in <a href="#synbio-02-00006-f001" class="html-fig">Figure 1</a>), and angucyclines.</p> Full article ">Figure 4
<p>Examples of type II PKS-derived natural products including extended angular polyphenols, polycycles with different starter units, unusual cyclization patterns, and structures with other atom incorporation beyond C, H, and O.</p> Full article ">Figure 5
<p>Examples of type II PKS products with post-cyclization rearrangement or atom excision.</p> Full article ">Figure 6
<p>Claisen-like condensation mechanism of the minimal cassette in type II PKS to grow the polyketide chain. Once the KS is loaded with a starter unit (<b>A</b>) via one of several mechanisms, the ACP interacts with malonyl-CoA or transacylases from primary metabolism to obtain a malonyl for chain extension (<b>B</b>). Decarboxylation activates the malonyl as an enolate on the ACP, which reacts with the thioester on the KS to form a tetrahedral intermediate (<b>C</b>). Collapse of the tetrahedral intermediate results in a polyketide extended by one acetate unit attached to the ACP (<b>D</b>). Continued extension involves the transfer of the extended polyketide back to the KS (<b>E</b>) and the acquisition of another malonyl by the ACP to reenter the extension cycle (<b>E</b> to <b>B</b>). Once the ACP-bound polyketide chain is at the appropriate length, it can no longer be accepted back onto the KS (see later Figure) causing dissociation of the ACP from the KS-CLF dimer (<b>F</b>) and enabling delivery of the polyketide chain to cyclases for tailoring to various ring systems.</p> Full article ">Figure 7
<p>(<b>A</b>) The structures of coenzyme A (the phosphopantetheine is highlighted in blue) and the abbreviated form of the malonyl-CoA unit. (<b>B</b>) Conversion of <span class="html-italic">apo</span>-ACP to <span class="html-italic">holo</span>-ACP by the action of PPTase. The squiggly line terminating in a thiol in the cartoon to the right represents the flexible phosphopantetheine arm.</p> Full article ">Figure 8
<p>Cartoon depicting the gating residues [<a href="#B16-synbio-02-00006" class="html-bibr">16</a>] for octaketide (<b>top</b>) production in the actinorhodin system and (<b>bottom</b>) the corresponding residues for tridecaketide production in the griseorhodin system. Residues from the CLF are shown in green while residues from the KS are shown in blue. The larger threonine112 and phenylalanine109 (<b>top</b>) cause growth to stop at the octaketide because the octaketide cannot be accommodated in the binding tunnel and thus cannot be transferred back to the KS for a continued extension. These are replaced by smaller residues (<b>bottom</b>) for the tridecaketide, which terminate at methionine143 and tyrosine140.</p> Full article ">Figure 9
<p>Natural products that utilize HR type II PKS in their biosynthesis. The polyene(s) derived from the HR type II PKS are highlighted in blue. Simocyclinone D8 also includes a canonical type II PKS core (blue bonds). <span class="html-italic">Inset</span>: The structure of cispentacin, a non-canonical amino acid formed through the action of a type-II-PKS-like minimal cassette [<a href="#B191-synbio-02-00006" class="html-bibr">191</a>].</p> Full article ">
Open AccessArticle
Pangenome-Scale Mathematical Modelling of ANAMMOX Bacteria Metabolism
by
Roman G. Bielski and M. Ahsanul Islam
SynBio 2024, 2(1), 70-84; https://doi.org/10.3390/synbio2010005 - 8 Feb 2024
Abstract
►▼
Show Figures
Removal of fixed nitrogen compounds such as ammonium and nitrite from wastewater is of critical importance for balancing the nitrogen cycle and protecting aquatic environments from eutrophication. ANaerobic AMMonium OXidising (ANAMMOX) bacteria have recently been employed for fixed nitrogen removal purposes in wastewater
[...] Read more.
Removal of fixed nitrogen compounds such as ammonium and nitrite from wastewater is of critical importance for balancing the nitrogen cycle and protecting aquatic environments from eutrophication. ANaerobic AMMonium OXidising (ANAMMOX) bacteria have recently been employed for fixed nitrogen removal purposes in wastewater treatment processes. These specialised bacteria convert ammonium and nitrite into nitrogen gas anaerobically, thereby reducing the amount of energy required for aeration in conventional wastewater treatment processes. However, slow growth rates of ANAMMOX remain a major obstacle towards their widespread use in industrial wastewater treatment processes. Thus, a pangenome-scale, constraint-based metabolic model, iRB399, of ANAMMOX bacteria has been developed to design strategies for accelerating their growth. The main metabolic limitation was identified in the energy metabolism of these bacteria, concerning the production of ATP. The extremely low efficiency of the electron transport chain combined with very high growth-associated maintenance energy is likely to be responsible for the slow growth of ANAMMOX. However, different ANAMMOX species were found to conserve energy using a variety of different redox couples, and the modelling simulations revealed their comparative advantages under different growth conditions. iRB399 also identified dispensable catabolic reactions that have demonstrably beneficial effects on enhancing the growth rates of ANAMMOX bacteria. Thus, the pangenome-scale model will not only help identify and overcome metabolic limitations of ANNAMOX bacteria, but also provide a valuable resource for designing efficient ANNAMOX-based wastewater treatment processes.
Full article
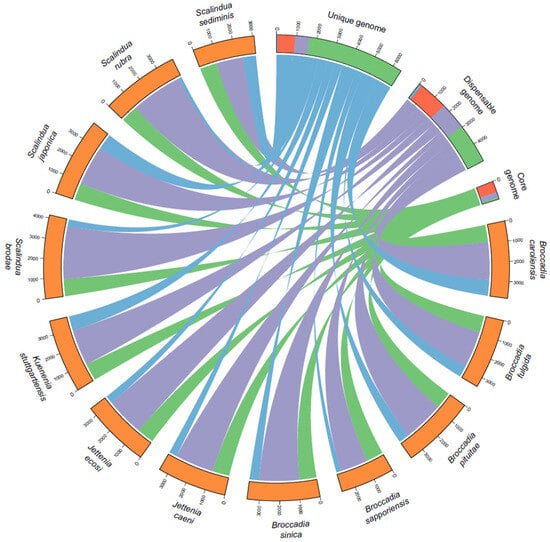
Figure 1
Figure 1
<p>The gene distribution in the ANAMMOX pangenome. The Figure was generated using CIRCOS [<a href="#B29-synbio-02-00005" class="html-bibr">29</a>]. The functional distribution of genes is shown in the core, dispensable and unique genomes as blue for transporters, red for metabolic genes, purple for non-metabolic genes, and green for hypothetical genes. The width of the ribbons represents the number of genes belonging to the unique, dispensable or core genome.</p> Full article ">Figure 2
<p>Prevalence of genes in the ANAMMOX pangenome related to energy metabolism. The number of genes found in each genome is shown by differing shade of green.</p> Full article ">Figure 3
<p>Distribution of genes enabling energy metabolism pathways in the ANAMMOX pangenome involving 7 different electron donors (along x-axis) coupled to 5 different electron sinks/acceptors (along y-axis). Colouration is used to show whether the redox couple has been observed promoting growth in the literature or in <span class="html-italic">i</span>RB399 or both.</p> Full article ">Figure 4
<p>In silico ANAMMOX growth simulation with <span class="html-italic">i</span>RB399. ANAMMOX growth rates (<b>left</b>) and ATP production fluxes (<b>right</b>) were simulated using <span class="html-italic">i</span>RB399 and the COBRA toolbox [<a href="#B35-synbio-02-00005" class="html-bibr">35</a>], and constrained by the availability of electron donors (5 mmol/gDCW.h of donor in each experiment) with an excess of nitric oxide (purple), nitrate (yellow) and nitrite (blue) as electron acceptors.</p> Full article ">Figure 5
<p>Ratio of electron donor to electron acceptor consumption in growth simulations of ANAMMOX using <span class="html-italic">i</span>RB399 and the COBRA toolbox [<a href="#B35-synbio-02-00005" class="html-bibr">35</a>]. Electron acceptors used are nitric oxide (purple), nitrate (yellow) and nitrite (blue). In each case, the uptake flux of the electron donor was constrained to 5 mmol/gDCW.h.</p> Full article ">Figure 6
<p>Simulation of metabolic fluxes in the central carbon metabolism under different substrate constraints. (<b>A</b>) shows the effect of glucose supplementation with nitrite as the electron sink, (<b>B</b>) shows the effect of ammonium and nitrite, and (<b>C</b>) shows the effect of ammonium and nitric oxide. Abbreviations: 1,3BPG, 1,3-Bisphosphoglyceric acid; 10-CHO-THF, 10-Formyltetrahydrofolate; 2GP, 2-Phosphoglycerate; 3GP, 3-Phosphoglycerate; 5,10-CHO=THF, 5,10-Methenyltetrahydrofolate; 5-10-CH2-THF, 5,10-Methylenetetrahydrofolate; 5-CH3-THF, 5-Formyltetrahydrofolate; CH3-CoFeSP, Methylcorrinoid protein; DHAP, Dihydroxyacetone phosphate; E4P, Erythrose 4-phosphate; F6P, Fructose 6-phosphate; FBP, Fructose bisphosphate; G6P, Glucose 6-phosphate; GAP, Glyceraldehyde 3-phosphate; PEP, Phosphoenolpyruvate; Pyr, Pyruvate; R5P, ribose-5-phosphate; Ru5P, Ribulose 5-phosphate; S7P, Sedoheptulose 7-phosphate; X5P, D-Xylulose 5-phosphate; α-KG, α-ketoglutarate.</p> Full article ">Figure 7
<p>Analysis of the thermodynamic efficiency of the ANAMMOX electron transport chain (ETC) expressed in % efficiency. Thermodynamic efficiency was measured by the number of protons translocated per electron transferred through the ETC. This is compared with the ammonium uptake flux to demonstrate the sensitivity of the system to both inputs. The sensitivity of ATP flux (<b>left</b>) and the growth rate (<b>right</b>) with ammonium uptake flux and ETC efficiency is compared to further demonstrate the similar pattern observed between them.</p> Full article ">
<p>The gene distribution in the ANAMMOX pangenome. The Figure was generated using CIRCOS [<a href="#B29-synbio-02-00005" class="html-bibr">29</a>]. The functional distribution of genes is shown in the core, dispensable and unique genomes as blue for transporters, red for metabolic genes, purple for non-metabolic genes, and green for hypothetical genes. The width of the ribbons represents the number of genes belonging to the unique, dispensable or core genome.</p> Full article ">Figure 2
<p>Prevalence of genes in the ANAMMOX pangenome related to energy metabolism. The number of genes found in each genome is shown by differing shade of green.</p> Full article ">Figure 3
<p>Distribution of genes enabling energy metabolism pathways in the ANAMMOX pangenome involving 7 different electron donors (along x-axis) coupled to 5 different electron sinks/acceptors (along y-axis). Colouration is used to show whether the redox couple has been observed promoting growth in the literature or in <span class="html-italic">i</span>RB399 or both.</p> Full article ">Figure 4
<p>In silico ANAMMOX growth simulation with <span class="html-italic">i</span>RB399. ANAMMOX growth rates (<b>left</b>) and ATP production fluxes (<b>right</b>) were simulated using <span class="html-italic">i</span>RB399 and the COBRA toolbox [<a href="#B35-synbio-02-00005" class="html-bibr">35</a>], and constrained by the availability of electron donors (5 mmol/gDCW.h of donor in each experiment) with an excess of nitric oxide (purple), nitrate (yellow) and nitrite (blue) as electron acceptors.</p> Full article ">Figure 5
<p>Ratio of electron donor to electron acceptor consumption in growth simulations of ANAMMOX using <span class="html-italic">i</span>RB399 and the COBRA toolbox [<a href="#B35-synbio-02-00005" class="html-bibr">35</a>]. Electron acceptors used are nitric oxide (purple), nitrate (yellow) and nitrite (blue). In each case, the uptake flux of the electron donor was constrained to 5 mmol/gDCW.h.</p> Full article ">Figure 6
<p>Simulation of metabolic fluxes in the central carbon metabolism under different substrate constraints. (<b>A</b>) shows the effect of glucose supplementation with nitrite as the electron sink, (<b>B</b>) shows the effect of ammonium and nitrite, and (<b>C</b>) shows the effect of ammonium and nitric oxide. Abbreviations: 1,3BPG, 1,3-Bisphosphoglyceric acid; 10-CHO-THF, 10-Formyltetrahydrofolate; 2GP, 2-Phosphoglycerate; 3GP, 3-Phosphoglycerate; 5,10-CHO=THF, 5,10-Methenyltetrahydrofolate; 5-10-CH2-THF, 5,10-Methylenetetrahydrofolate; 5-CH3-THF, 5-Formyltetrahydrofolate; CH3-CoFeSP, Methylcorrinoid protein; DHAP, Dihydroxyacetone phosphate; E4P, Erythrose 4-phosphate; F6P, Fructose 6-phosphate; FBP, Fructose bisphosphate; G6P, Glucose 6-phosphate; GAP, Glyceraldehyde 3-phosphate; PEP, Phosphoenolpyruvate; Pyr, Pyruvate; R5P, ribose-5-phosphate; Ru5P, Ribulose 5-phosphate; S7P, Sedoheptulose 7-phosphate; X5P, D-Xylulose 5-phosphate; α-KG, α-ketoglutarate.</p> Full article ">Figure 7
<p>Analysis of the thermodynamic efficiency of the ANAMMOX electron transport chain (ETC) expressed in % efficiency. Thermodynamic efficiency was measured by the number of protons translocated per electron transferred through the ETC. This is compared with the ammonium uptake flux to demonstrate the sensitivity of the system to both inputs. The sensitivity of ATP flux (<b>left</b>) and the growth rate (<b>right</b>) with ammonium uptake flux and ETC efficiency is compared to further demonstrate the similar pattern observed between them.</p> Full article ">
Open AccessArticle
Transcriptomic Investigation in CRISPR/Cas9-Mediated GRIK1-, GRIK2-, and GRIK4-Gene-Knockout Human Neuroblastoma Cells
by
Tsung-Ming Hu, Shih-Hsin Hsu, Hsin-Yao Tsai and Min-Chih Cheng
SynBio 2024, 2(1), 56-69; https://doi.org/10.3390/synbio2010004 - 5 Feb 2024
Cited by 1
Abstract
►▼
Show Figures
The glutamate ionotropic kainate receptors, encoded by the GRIK gene family, are composed of four subunits and function as ligand-activated ion channels. They play a critical role in regulating synaptic transmission and various synaptic receptors’ processes, as well as in the pathophysiology of
[...] Read more.
The glutamate ionotropic kainate receptors, encoded by the GRIK gene family, are composed of four subunits and function as ligand-activated ion channels. They play a critical role in regulating synaptic transmission and various synaptic receptors’ processes, as well as in the pathophysiology of schizophrenia. However, their functions and mechanisms of action need to be better understood and are worthy of exploration. To further understand the exact role of the kainate receptors in vitro, we generated kainate-receptor-knockout (KO) isogenic SH-SY5Y cell lines using the CRISPR/Cas9-mediated gene editing method. We conducted RNA sequencing (RNA-seq) to determine the differentially expressed genes (DEGs) in the isogenic edited cells and used rhodamine-phalloidin staining to quantitate filamentous actin (F-actin) in differentiated edited cells. The RNA-seq and the Gene Ontology enrichment analysis revealed that the genetic deletion of the GRIK1, GRIK2, and GRIK4 genes disturbed multiple genes involved in numerous signal pathways, including a converging pathway related to the synaptic membrane. An enrichment analysis of gene–disease associations indicated that DEGs in the edited cell lines were associated with several neuropsychiatric disorders, especially schizophrenia. In the morphology study, fluorescent images show that less F-actin was expressed in differentiated SH-SY5Y cells with GRIK1, GRIK2, or GRIK4 deficiency than wild-type cells. Our data indicate that kainate receptor deficiency might disturb synaptic-membrane-associated genes, and elucidating these genes should shed some light on the pathophysiology of schizophrenia. Furthermore, the transcriptomic profiles for kainate receptor deficiency of SH-SY5Y cells contribute to emerging evidence for the novel mechanisms underlying the effect of kainate receptors and the pathophysiology of schizophrenia. In addition, our data suggest that kainate-receptor-mediated F-actin remodeling may be a candidate mechanism underlying schizophrenia.
Full article
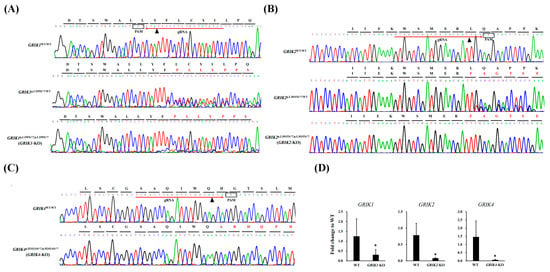
Figure 1
Figure 1
<p>Generation and characterization of <span class="html-italic">GRIK1</span>-KO, <span class="html-italic">GRIK2</span>-KO, and <span class="html-italic">GRIK4</span>-KO SH-SY5Y cell lines. (<b>A</b>) The schema of the gRNA target site of the <span class="html-italic">GRIK1</span> gene (red arrow). Sanger sequencing analysis of wild-type (<span class="html-italic">GRIK1</span><sup>WT/WT</sup>) and two edited cell lines (<span class="html-italic">GRIK1</span><sup>p.L25Pfs*?/WT</sup> and <span class="html-italic">GRIK1</span><sup>p.L25Pfs*?/p.L25Pfs*?</sup>). (<b>B</b>) The schema of the gRNA target site of the <span class="html-italic">GRIK2</span> gene (red arrow). Sanger sequencing analysis of wild-type (<span class="html-italic">GRIK2</span><sup>WT/WT</sup>) and two edited cell lines (<span class="html-italic">GRIK2</span><sup>p.L301Ffs*?/WT</sup> and <span class="html-italic">GRIK2</span><sup>p.L301Ffs*?/p.L301Ffs*</sup>). (<b>C</b>) The schema of the gRNA target site of the <span class="html-italic">GRIK4</span> gene (red arrow). Sanger sequencing analysis of wild-type (<span class="html-italic">GRIK4</span><sup>WT/WT</sup>) and one edited cell line (<span class="html-italic">GRIK4</span> <sup>p.H343Afs*?/p.H343Afs*?</sup>). (<b>D</b>) RT-qPCR assay showing the expression of <span class="html-italic">GRIK1</span>, <span class="html-italic">GRIK2</span>, and <span class="html-italic">GRIK4</span> genes in <span class="html-italic">GRIK1</span><sup>p.L25Pfs*?/p.L25Pfs*?</sup> (<span class="html-italic">GRIK1</span>-KO), <span class="html-italic">GRIK2</span><sup>p.L301Ffs*?/p.L301Ffs*</sup> (<span class="html-italic">GRIK2</span>-KO), and <span class="html-italic">GRIK4</span> <sup>p.H343Afs*?/p.H343Afs*?</sup> (<span class="html-italic">GRIK4</span>-KO) SH-SY5Y cell lines, respectively, compared to WT cells. The <span class="html-italic">GAPDH</span> gene was used as the endogenous gene for normalization. The data are expressed as fold change to WT ± SD (* <span class="html-italic">p</span> < 0.05, <span class="html-italic">n</span> = 6). Arrowhead indicates the predicted double-strand break site. PAM means the protospacer adjacent motif.</p> Full article ">Figure 2
<p>RNA-seq analysis. (<b>A</b>) Volcano plot analysis of DEGs between <span class="html-italic">GRIK1</span><sup>p.L25Pfs*?/p.L25Pfs*?</sup> (<span class="html-italic">GRIK1</span>-KO), <span class="html-italic">GRIK2</span><sup>p.L301Ffs*?/p.L301Ffs*</sup> (<span class="html-italic">GRIK2</span>-KO), and <span class="html-italic">GRIK4</span> <sup>p.H343Afs*?/p.H343Afs*?</sup> (<span class="html-italic">GRIK4</span>-KO) SH-SY5Y cell lines, respectively, compared to WT cells. (<b>B</b>) Hierarchical clustered heatmap showing the expression patterns of significant (fold change ≤−2 or ≥2; <span class="html-italic">p</span> < 0.05) DEGs in each edited cell line. (<b>C</b>) Top 30 GO terms enriched in <span class="html-italic">GRIK1</span>-KO SH-SY5Y cell lines. (<b>D</b>) Top 30 GO terms enriched in <span class="html-italic">GRIK2</span>-KO SH-SY5Y cell lines. (<b>E</b>) Top 30 GO terms enriched in <span class="html-italic">GRIK4</span>-KO SH-SY5Y cell lines. BP is the biological process; CC is the cell component; MF is the molecular function.</p> Full article ">Figure 3
<p>RT-qPCR assay showing the expression of six schizophrenia-associated genes (<span class="html-italic">ARC</span>, <span class="html-italic">GRIA2</span>, <span class="html-italic">GRIA4</span>, <span class="html-italic">GABRB3</span>, <span class="html-italic">GRM8</span>, and <span class="html-italic">KCNJ3</span>) in edited SH-SY5Y cells (<span class="html-italic">GRIK1</span>-KO, <span class="html-italic">GRIK2</span>-KO, and <span class="html-italic">GRIK4</span>-KO) and WT cells. The <span class="html-italic">GAPDH</span> gene was used as the endogenous gene for normalization. The data are expressed as fold change to WT ± SD (* <span class="html-italic">p</span> < 0.05, <span class="html-italic">n</span> = 6).</p> Full article ">Figure 4
<p>Representative phase-contrast and fluorescent images of differentiated SH-SY5Y cells harboring <span class="html-italic">GRIK1</span>, <span class="html-italic">GRIK2</span>, or <span class="html-italic">GRIK4</span> deficiency. (<b>A</b>) Phase-contrast. (<b>B</b>) Fluorescent images. Cells are labeled green with the neuronal marker MAP2, red with the cytoskeleton F-actin, and blue with DAPI.</p> Full article ">
<p>Generation and characterization of <span class="html-italic">GRIK1</span>-KO, <span class="html-italic">GRIK2</span>-KO, and <span class="html-italic">GRIK4</span>-KO SH-SY5Y cell lines. (<b>A</b>) The schema of the gRNA target site of the <span class="html-italic">GRIK1</span> gene (red arrow). Sanger sequencing analysis of wild-type (<span class="html-italic">GRIK1</span><sup>WT/WT</sup>) and two edited cell lines (<span class="html-italic">GRIK1</span><sup>p.L25Pfs*?/WT</sup> and <span class="html-italic">GRIK1</span><sup>p.L25Pfs*?/p.L25Pfs*?</sup>). (<b>B</b>) The schema of the gRNA target site of the <span class="html-italic">GRIK2</span> gene (red arrow). Sanger sequencing analysis of wild-type (<span class="html-italic">GRIK2</span><sup>WT/WT</sup>) and two edited cell lines (<span class="html-italic">GRIK2</span><sup>p.L301Ffs*?/WT</sup> and <span class="html-italic">GRIK2</span><sup>p.L301Ffs*?/p.L301Ffs*</sup>). (<b>C</b>) The schema of the gRNA target site of the <span class="html-italic">GRIK4</span> gene (red arrow). Sanger sequencing analysis of wild-type (<span class="html-italic">GRIK4</span><sup>WT/WT</sup>) and one edited cell line (<span class="html-italic">GRIK4</span> <sup>p.H343Afs*?/p.H343Afs*?</sup>). (<b>D</b>) RT-qPCR assay showing the expression of <span class="html-italic">GRIK1</span>, <span class="html-italic">GRIK2</span>, and <span class="html-italic">GRIK4</span> genes in <span class="html-italic">GRIK1</span><sup>p.L25Pfs*?/p.L25Pfs*?</sup> (<span class="html-italic">GRIK1</span>-KO), <span class="html-italic">GRIK2</span><sup>p.L301Ffs*?/p.L301Ffs*</sup> (<span class="html-italic">GRIK2</span>-KO), and <span class="html-italic">GRIK4</span> <sup>p.H343Afs*?/p.H343Afs*?</sup> (<span class="html-italic">GRIK4</span>-KO) SH-SY5Y cell lines, respectively, compared to WT cells. The <span class="html-italic">GAPDH</span> gene was used as the endogenous gene for normalization. The data are expressed as fold change to WT ± SD (* <span class="html-italic">p</span> < 0.05, <span class="html-italic">n</span> = 6). Arrowhead indicates the predicted double-strand break site. PAM means the protospacer adjacent motif.</p> Full article ">Figure 2
<p>RNA-seq analysis. (<b>A</b>) Volcano plot analysis of DEGs between <span class="html-italic">GRIK1</span><sup>p.L25Pfs*?/p.L25Pfs*?</sup> (<span class="html-italic">GRIK1</span>-KO), <span class="html-italic">GRIK2</span><sup>p.L301Ffs*?/p.L301Ffs*</sup> (<span class="html-italic">GRIK2</span>-KO), and <span class="html-italic">GRIK4</span> <sup>p.H343Afs*?/p.H343Afs*?</sup> (<span class="html-italic">GRIK4</span>-KO) SH-SY5Y cell lines, respectively, compared to WT cells. (<b>B</b>) Hierarchical clustered heatmap showing the expression patterns of significant (fold change ≤−2 or ≥2; <span class="html-italic">p</span> < 0.05) DEGs in each edited cell line. (<b>C</b>) Top 30 GO terms enriched in <span class="html-italic">GRIK1</span>-KO SH-SY5Y cell lines. (<b>D</b>) Top 30 GO terms enriched in <span class="html-italic">GRIK2</span>-KO SH-SY5Y cell lines. (<b>E</b>) Top 30 GO terms enriched in <span class="html-italic">GRIK4</span>-KO SH-SY5Y cell lines. BP is the biological process; CC is the cell component; MF is the molecular function.</p> Full article ">Figure 3
<p>RT-qPCR assay showing the expression of six schizophrenia-associated genes (<span class="html-italic">ARC</span>, <span class="html-italic">GRIA2</span>, <span class="html-italic">GRIA4</span>, <span class="html-italic">GABRB3</span>, <span class="html-italic">GRM8</span>, and <span class="html-italic">KCNJ3</span>) in edited SH-SY5Y cells (<span class="html-italic">GRIK1</span>-KO, <span class="html-italic">GRIK2</span>-KO, and <span class="html-italic">GRIK4</span>-KO) and WT cells. The <span class="html-italic">GAPDH</span> gene was used as the endogenous gene for normalization. The data are expressed as fold change to WT ± SD (* <span class="html-italic">p</span> < 0.05, <span class="html-italic">n</span> = 6).</p> Full article ">Figure 4
<p>Representative phase-contrast and fluorescent images of differentiated SH-SY5Y cells harboring <span class="html-italic">GRIK1</span>, <span class="html-italic">GRIK2</span>, or <span class="html-italic">GRIK4</span> deficiency. (<b>A</b>) Phase-contrast. (<b>B</b>) Fluorescent images. Cells are labeled green with the neuronal marker MAP2, red with the cytoskeleton F-actin, and blue with DAPI.</p> Full article ">
Open AccessArticle
Generation of New Glycoanalogues of Polyene Antibiotics by Synthetic Biology—Testing Current Technical Boundaries
by
Mark Hogan, Yuhao Song, Jimmy Muldoon and Patrick Caffrey
SynBio 2024, 2(1), 31-55; https://doi.org/10.3390/synbio2010003 - 4 Jan 2024
Cited by 1
Abstract
►▼
Show Figures
A number of antifungal drugs are based on polyene macrolides that cause severe side effects. Most of these compounds contain a single aminodeoxysugar, D-mycosamine. Toxicity can be reduced by increasing the extent of glycosylation. The aromatic heptaene 67-121C and two analogues of the
[...] Read more.
A number of antifungal drugs are based on polyene macrolides that cause severe side effects. Most of these compounds contain a single aminodeoxysugar, D-mycosamine. Toxicity can be reduced by increasing the extent of glycosylation. The aromatic heptaene 67-121C and two analogues of the degenerate heptaene nystatin have a second sugar attached to the C4′ hydroxyl of mycosamine. Another nystatin analogue has L-digitoxose as a second sugar attached to C35 on the macrolactone ring. The pentaene selvamicin has 4-O-methyl-L-digitoxose at C27, the equivalent position. To assist the production of new antifungals by synthetic biology, we explore further the utility of three classes of polyene glycosyltransferase: extending glycosyltransferases that form disaccharide-containing polyenes, glycosyltransferases that add the L-digitoxose sugars of nystatin A3 and selvamicin, and mycosaminyltransferases that add the primary aminodeoxysugar. In addition, we combine enzymatic hyperglycosylation with a known chemical method for adding sugars to the C3′ amino group of mycosamine. This was used to convert the disaccharide-containing 67-121C heptaene to forms containing branched trisaccharide or tetrasaccharide chains. These analogues are of interest for testing as anti-Leishmania drugs.
Full article
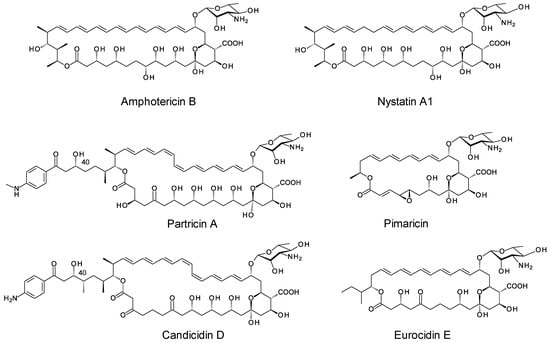
Figure 1
Figure 1
<p>Examples of glycosylated polyene macrolides.</p> Full article ">Figure 2
<p>Naturally ocurring disaccharide-containing polyenes. 67-121C is produced by <span class="html-italic">Couchioplanes caeruleus</span> DSM43634, NPP A1 is obtained from <span class="html-italic">Pseudonocardia autotrophica</span> KCTC9441, nystatin P1 is obtained from <span class="html-italic">Pseudonocardia</span> species P1.</p> Full article ">Figure 3
<p>Examples of polyenes containing two unlinked monosaccharides.</p> Full article ">Figure 4
<p>Overview of chemical glycosylation of a polyene with D-glucose. The rearrangement reaction gives a linear fructosyl chain that can cyclize to give α or β anomers of pyranosyl and furanosyl forms. Preobrazhenskaya and co-workers [<a href="#B37-synbio-02-00003" class="html-bibr">37</a>] found that the linear open chain fructosyl form exists in equilibrium with the four cyclized forms.</p> Full article ">Figure 5
<p>Opportunities for generating polyene glycoanalogues by synthetic biology. (<b>A</b>) Polyene macrolactones can be modified at three points by enzymatic glycosylation, chemical glycosylation of sugar amino groups is also practicable. (<b>B</b>) Hypothetical example of a glycoanalogue that might be obtained by combining these methods.</p> Full article ">Figure 6
<p>Alignment of N-terminal sequences of PegA and NypY showing methionines encoded by two plausible start codons.</p> Full article ">Figure 7
<p>HPLC analysis of candicidins. (<b>A</b>) Candicidin complex from <span class="html-italic">S. albidoflavus</span> containing the empty pIAGO vector. (<b>B</b>) Candicidins from <span class="html-italic">S. albidoflavus</span> containing pIAGO-pegA1. (<b>C</b>) Candicidins from <span class="html-italic">S. albidoflavus</span> containing pIAGO-pegA2. New heptaene peaks resulting from enzymatic mannosylation are marked with red stars.</p> Full article ">Figure 8
<p>Structures of mannosyl candidicidins.</p> Full article ">Figure 9
<p>HPLC of 67-121 polyenes extracted from <span class="html-italic">C. caeruleus</span>.</p> Full article ">Figure 10
<p>Trisaccharides and tetrasaccharides formed by chemical glycosylation of 67-121C.</p> Full article ">Figure 11
<p>Analysis of chemically glucosylated 67-121 polyenes by LC-MS. The chromatography step partially resolved the heptaene complex into two peaks containing trisaccharide- and disaccharide-modified polyenes, respectively. (<b>A</b>) The red trace shows total ion count (TIC) versus time. The more polar leading peak (shaded grey) has an RT of 3.394 min. (<b>B</b>) Mass spectrum of leading peak showing ions with a mass appropriate for a trisaccharide-modified 67-121 polyene, (M + H)<sup>+</sup> = 1451.6932, (M + Na)<sup>+</sup> = 1473.6748. (<b>C</b>) The red trace shows TIC versus time. The less polar trailing peak (shaded grey) has an RT of 3.478 min. (<b>D</b>) Mass spectrum of second peak showing ions with masses appropriate for disaccharide-containing 67-121 polyenes (M + H)<sup>+</sup> = 1289.6414, (M + Na)<sup>+</sup> = 1311.6230. In this analysis it is not possible to distinguish between unmodified 67-121C and glucose-modified 67-121A.</p> Full article ">Figure 12
<p>Chemical modification of 67−121 complex with lactose. Panel (<b>A</b>) shows analysis of control unmodified 67−121 polyenes. Panel (<b>B</b>) shows material modified with lactose. New heptaene peaks appearing after chemical modification are marked with stars.</p> Full article ">Figure 13
<p>Mass spectra of tetrasaccharide-containing (<b>A</b>) and trisaccharide-containing (<b>B</b>) aromatic heptaenes after modification of the 67−121C/67−121A complex with lactose.</p> Full article ">Figure 14
<p>Structures of pentaene predicted for <span class="html-italic">Sacc. gloriosae</span> and <span class="html-italic">S. nodosus</span> M57 pentaene.</p> Full article ">Figure 15
<p>EurN and EurDI replace AmphN and AmphDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>. The red lines indicate that the chromosomal <span class="html-italic">amphN</span> and <span class="html-italic">amphDI</span> genes have been deleted. Enzymes in blue text are encoded by a complementing pIAGO construct. This convention is used in Figures 18, 20, 22 and 24.</p> Full article ">Figure 16
<p>HPLC evidence for modification of amphotericin heptaene aglycones by EurN and EurDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>. Panel (<b>A</b>) Heptaenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO (empty vector control). Panel (<b>B</b>) Heptaenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO-EurDI-DII-NM. Panel (<b>C</b>) amphotericin B standard isolated from <span class="html-italic">S. nodosus</span>. Heptaenes were detected by monitoring at A<sub>405</sub>.</p> Full article ">Figure 17
<p>HPLC evidence for modification of amphotericin tetraene aglycone by EurN and EurDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>. Panel (<b>A</b>) Tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO (empty vector control). Panel (<b>B</b>) Tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO-EurDI-DII-NM. Red triangles highlight polar, possibly glycosylated, tetraenes with retention times different from that of amphotericin A. Panel (<b>C</b>) 8-deoxyamphoteronolide A isoforms isolated from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDIII</span>. Panel (<b>D</b>) Amphotericin A standard isolated from <span class="html-italic">S. nodosus</span>. Tetraenes were detected by monitoring at A<sub>320</sub>.</p> Full article ">Figure 18
<p>EurN P450 acts on 8-deoxy-16-descarboxyl-16-methylamphoteronolide A tetraene but EurDI does not efficiently mycosaminylate 8-deoxy-amphoteronolide A.</p> Full article ">Figure 19
<p>HPLC evidence for glycosylation of amphotericin heptaene aglycones by GloDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>. Panel (<b>A</b>) Heptaenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO (empty vector control). Panel (<b>B</b>) Heptaenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO-GloDI. Panel (<b>C</b>) Mycosaminylated 16-descarboxyl-16-methyl-amphotericin B standard isolated from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphNM</span>. Heptaenes were detected by monitoring at A<sub>405</sub>.</p> Full article ">Figure 20
<p>GloDI catalyses glycosylation of 8-deoxy-16-descarboxyl-16-methyl-amphoteronolide B in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>.</p> Full article ">Figure 21
<p>HPLC evidence for glycosylation amphotericin tetraene aglycones by GloDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>. Panel (<b>A</b>) Tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO (empty vector control). Panel (<b>B</b>) Tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO-GloDI. Panel (<b>C</b>) Mycosaminylated 8-deoxy-16-descarboxyl-16-methyl-amphotericin A standard isolated from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphNM</span>. Tetraenes were detected by monitoring at A<sub>320</sub>.</p> Full article ">Figure 22
<p>GloDI catalyses glycosylation of the tetraene 8-deoxy-16-descarboxyl-16-methyl-amphoteronolide A in the <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> mutant. AmphL does not C8 hydroxylate 8-deoxy-8-deoxy-16-descarboxyl-16-methyl-amphotericin A [<a href="#B39-synbio-02-00003" class="html-bibr">39</a>].</p> Full article ">Figure 23
<p>HPLC evidence for activity of GloDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-DII-NM</span>. Heptaenes and tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-DII-NM</span> pIAGO-GloDI are shown in Panels (<b>B</b>,<b>D</b>). Panels (<b>A</b>,<b>C</b>) show heptaenes and tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO control.</p> Full article ">Figure 24
<p>In the mycosamine-deficient <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-DII-NM</span>, GloDI transfers a neutral deoxyhexose to 8-deoxy-16-descarboxyl-16-methyl-amphoteronolide B heptaene but not to 8-deoxy-16-descarboxyl-16-methyl-amphoteronolide A tetraene.</p> Full article ">Figure 25
<p>Testing for antifungal activity in methanol extracts of <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> containing pIAGO, pIAGO-GloDI, pIAGO-EurDI-DII-N-M, and <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-DII-NM</span> pIAGO-GloDI.</p> Full article ">
<p>Examples of glycosylated polyene macrolides.</p> Full article ">Figure 2
<p>Naturally ocurring disaccharide-containing polyenes. 67-121C is produced by <span class="html-italic">Couchioplanes caeruleus</span> DSM43634, NPP A1 is obtained from <span class="html-italic">Pseudonocardia autotrophica</span> KCTC9441, nystatin P1 is obtained from <span class="html-italic">Pseudonocardia</span> species P1.</p> Full article ">Figure 3
<p>Examples of polyenes containing two unlinked monosaccharides.</p> Full article ">Figure 4
<p>Overview of chemical glycosylation of a polyene with D-glucose. The rearrangement reaction gives a linear fructosyl chain that can cyclize to give α or β anomers of pyranosyl and furanosyl forms. Preobrazhenskaya and co-workers [<a href="#B37-synbio-02-00003" class="html-bibr">37</a>] found that the linear open chain fructosyl form exists in equilibrium with the four cyclized forms.</p> Full article ">Figure 5
<p>Opportunities for generating polyene glycoanalogues by synthetic biology. (<b>A</b>) Polyene macrolactones can be modified at three points by enzymatic glycosylation, chemical glycosylation of sugar amino groups is also practicable. (<b>B</b>) Hypothetical example of a glycoanalogue that might be obtained by combining these methods.</p> Full article ">Figure 6
<p>Alignment of N-terminal sequences of PegA and NypY showing methionines encoded by two plausible start codons.</p> Full article ">Figure 7
<p>HPLC analysis of candicidins. (<b>A</b>) Candicidin complex from <span class="html-italic">S. albidoflavus</span> containing the empty pIAGO vector. (<b>B</b>) Candicidins from <span class="html-italic">S. albidoflavus</span> containing pIAGO-pegA1. (<b>C</b>) Candicidins from <span class="html-italic">S. albidoflavus</span> containing pIAGO-pegA2. New heptaene peaks resulting from enzymatic mannosylation are marked with red stars.</p> Full article ">Figure 8
<p>Structures of mannosyl candidicidins.</p> Full article ">Figure 9
<p>HPLC of 67-121 polyenes extracted from <span class="html-italic">C. caeruleus</span>.</p> Full article ">Figure 10
<p>Trisaccharides and tetrasaccharides formed by chemical glycosylation of 67-121C.</p> Full article ">Figure 11
<p>Analysis of chemically glucosylated 67-121 polyenes by LC-MS. The chromatography step partially resolved the heptaene complex into two peaks containing trisaccharide- and disaccharide-modified polyenes, respectively. (<b>A</b>) The red trace shows total ion count (TIC) versus time. The more polar leading peak (shaded grey) has an RT of 3.394 min. (<b>B</b>) Mass spectrum of leading peak showing ions with a mass appropriate for a trisaccharide-modified 67-121 polyene, (M + H)<sup>+</sup> = 1451.6932, (M + Na)<sup>+</sup> = 1473.6748. (<b>C</b>) The red trace shows TIC versus time. The less polar trailing peak (shaded grey) has an RT of 3.478 min. (<b>D</b>) Mass spectrum of second peak showing ions with masses appropriate for disaccharide-containing 67-121 polyenes (M + H)<sup>+</sup> = 1289.6414, (M + Na)<sup>+</sup> = 1311.6230. In this analysis it is not possible to distinguish between unmodified 67-121C and glucose-modified 67-121A.</p> Full article ">Figure 12
<p>Chemical modification of 67−121 complex with lactose. Panel (<b>A</b>) shows analysis of control unmodified 67−121 polyenes. Panel (<b>B</b>) shows material modified with lactose. New heptaene peaks appearing after chemical modification are marked with stars.</p> Full article ">Figure 13
<p>Mass spectra of tetrasaccharide-containing (<b>A</b>) and trisaccharide-containing (<b>B</b>) aromatic heptaenes after modification of the 67−121C/67−121A complex with lactose.</p> Full article ">Figure 14
<p>Structures of pentaene predicted for <span class="html-italic">Sacc. gloriosae</span> and <span class="html-italic">S. nodosus</span> M57 pentaene.</p> Full article ">Figure 15
<p>EurN and EurDI replace AmphN and AmphDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>. The red lines indicate that the chromosomal <span class="html-italic">amphN</span> and <span class="html-italic">amphDI</span> genes have been deleted. Enzymes in blue text are encoded by a complementing pIAGO construct. This convention is used in Figures 18, 20, 22 and 24.</p> Full article ">Figure 16
<p>HPLC evidence for modification of amphotericin heptaene aglycones by EurN and EurDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>. Panel (<b>A</b>) Heptaenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO (empty vector control). Panel (<b>B</b>) Heptaenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO-EurDI-DII-NM. Panel (<b>C</b>) amphotericin B standard isolated from <span class="html-italic">S. nodosus</span>. Heptaenes were detected by monitoring at A<sub>405</sub>.</p> Full article ">Figure 17
<p>HPLC evidence for modification of amphotericin tetraene aglycone by EurN and EurDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>. Panel (<b>A</b>) Tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO (empty vector control). Panel (<b>B</b>) Tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO-EurDI-DII-NM. Red triangles highlight polar, possibly glycosylated, tetraenes with retention times different from that of amphotericin A. Panel (<b>C</b>) 8-deoxyamphoteronolide A isoforms isolated from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDIII</span>. Panel (<b>D</b>) Amphotericin A standard isolated from <span class="html-italic">S. nodosus</span>. Tetraenes were detected by monitoring at A<sub>320</sub>.</p> Full article ">Figure 18
<p>EurN P450 acts on 8-deoxy-16-descarboxyl-16-methylamphoteronolide A tetraene but EurDI does not efficiently mycosaminylate 8-deoxy-amphoteronolide A.</p> Full article ">Figure 19
<p>HPLC evidence for glycosylation of amphotericin heptaene aglycones by GloDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>. Panel (<b>A</b>) Heptaenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO (empty vector control). Panel (<b>B</b>) Heptaenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO-GloDI. Panel (<b>C</b>) Mycosaminylated 16-descarboxyl-16-methyl-amphotericin B standard isolated from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphNM</span>. Heptaenes were detected by monitoring at A<sub>405</sub>.</p> Full article ">Figure 20
<p>GloDI catalyses glycosylation of 8-deoxy-16-descarboxyl-16-methyl-amphoteronolide B in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>.</p> Full article ">Figure 21
<p>HPLC evidence for glycosylation amphotericin tetraene aglycones by GloDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span>. Panel (<b>A</b>) Tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO (empty vector control). Panel (<b>B</b>) Tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO-GloDI. Panel (<b>C</b>) Mycosaminylated 8-deoxy-16-descarboxyl-16-methyl-amphotericin A standard isolated from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphNM</span>. Tetraenes were detected by monitoring at A<sub>320</sub>.</p> Full article ">Figure 22
<p>GloDI catalyses glycosylation of the tetraene 8-deoxy-16-descarboxyl-16-methyl-amphoteronolide A in the <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> mutant. AmphL does not C8 hydroxylate 8-deoxy-8-deoxy-16-descarboxyl-16-methyl-amphotericin A [<a href="#B39-synbio-02-00003" class="html-bibr">39</a>].</p> Full article ">Figure 23
<p>HPLC evidence for activity of GloDI in <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-DII-NM</span>. Heptaenes and tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-DII-NM</span> pIAGO-GloDI are shown in Panels (<b>B</b>,<b>D</b>). Panels (<b>A</b>,<b>C</b>) show heptaenes and tetraenes from <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> pIAGO control.</p> Full article ">Figure 24
<p>In the mycosamine-deficient <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-DII-NM</span>, GloDI transfers a neutral deoxyhexose to 8-deoxy-16-descarboxyl-16-methyl-amphoteronolide B heptaene but not to 8-deoxy-16-descarboxyl-16-methyl-amphoteronolide A tetraene.</p> Full article ">Figure 25
<p>Testing for antifungal activity in methanol extracts of <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-NM</span> containing pIAGO, pIAGO-GloDI, pIAGO-EurDI-DII-N-M, and <span class="html-italic">S. nodosus</span> Δ<span class="html-italic">amphDI-DII-NM</span> pIAGO-GloDI.</p> Full article ">
Open AccessArticle
Chemo-Enzymatic Synthesis of Bioactive Carbazole Derivatives
by
Saad Alrashdi, Federica Casolari, Kwaku Kyeremeh and Hai Deng
SynBio 2024, 2(1), 21-30; https://doi.org/10.3390/synbio2010002 - 4 Jan 2024
Abstract
►▼
Show Figures
Carbazoles are key scaffolds of either antimicrobial/antiviral alkaloid natural products or therapeutics. As such, access to structurally diverse indole-containing carbazoles has attracted considerable attention. In this report, a pilot study is described using biotransformation to provide carbazoles that contain various acyl substituents. The
[...] Read more.
Carbazoles are key scaffolds of either antimicrobial/antiviral alkaloid natural products or therapeutics. As such, access to structurally diverse indole-containing carbazoles has attracted considerable attention. In this report, a pilot study is described using biotransformation to provide carbazoles that contain various acyl substituents. The biotransformation system contains the thiamine-diphosphate (ThDP)-dependent enzyme NzsH, the FabH-like 3-ketoacyl-ACP synthase NzsJ, and the aromatase/cyclase NzsI, encoded in the biosynthetic gene cluster (nzs) of the bacterial carbazole alkaloid natural product named neocarazostatin A. The utilization of a range of acyl-SNACs (synthetic acyl-thioester analogues of the native substrate) together with indole-3-pyruvate and pyruvate in the designed biotransformation system allows production of carbazole derivatives. Our results demonstrate that this three-enzyme system displays a considerable substrate profile toward acyl donors for production of carbazoles with different acyl substituents. Finally, two more enzymes were included in the biotransformation system: the tryptophan synthase stand-alone β-subunit variant, PfTrpB, generated from directed evolution in the literature, and a commercially available L-amino acid oxidase (LAAO). The addition of these two enzymes allows the transformation to start with indole building blocks to provide carbazoles with modifications in the indole ring system.
Full article
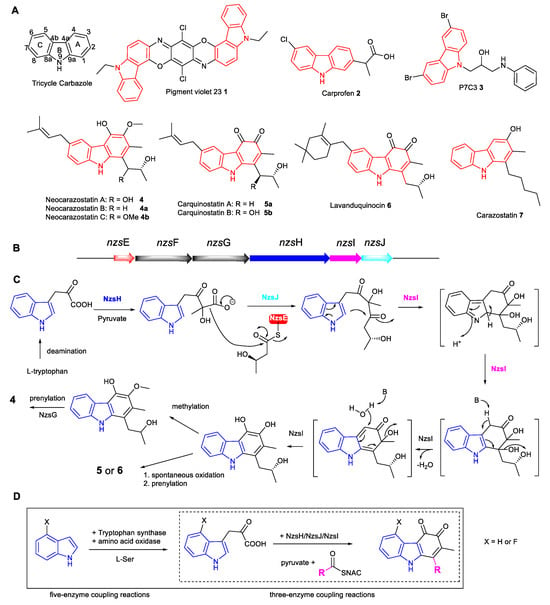
Figure 1
Figure 1
<p>(<b>A</b>) Representative structures of tricyclic carbazole, utility molecules containing carbazole motifs, and one group of bacterial metabolites containing simple carbazole scaffolds. (<b>B</b>) The conserved essential genes responsible for the production of this group of bacterial carbazoles. The conserved genes are color-coded. <span class="html-italic">nzs</span>E: red, <span class="html-italic">nzs</span>H: blue, <span class="html-italic">nzs</span>I: pink, and <span class="html-italic">nzs</span>J: light blue. (<b>C</b>) The proposed pathways of bacterial carbazoles starting from indole-3-pyruvate catalyzed by three enzyme homologues (NzsH, NzsJ and NzsI in the case of 4). The corresponding conserved enzymes are color-coded. NzsE: red, NzsH: blue, NzsI: pink, and NzsJ: light blue. (<b>D</b>) Chemo-enzymatic synthesis of structurally diverse carbazole molecules in this study.</p> Full article ">Figure 2
<p>HPLC analysis confirming production of iso-valeryl carbazole (CA) derivative. (<b>A</b>) A biotransformation scheme starting from indole-3-pyruvate and pyruvate together with isovaleryl-SNAC to generate isovaleryl-CAs. (<b>B</b>) HPLC traces of the biotransformation from indole-3-pyruvate and pyruvate together with isovaleryl-SNACs to isovaleryl-CA. (i). indole-3-pyruvate as a control. (ii). The production of acyloins generated by NzsH-catalyzed system. (iii). The production of isovaleryl-CA generated by the three-enzyme biotransformation system, including NzsH, J, and I.</p> Full article ">Figure 3
<p>(<b>A</b>) One-pot 5-enzymatic system to generate isovaleryl-carbazole derivatives. (<b>B</b>) Extracted ion chromatography of HR-LCMS analysis confirming production of isovaleryl-carbazole (red) and 4-fluoro-isovaleryl carbazole (green). (i) the enzymatic system in the absence of indole as control; (ii) the enzymatic system in the presence of 4H-indole; (iii) the enzymatic system in the presence of 4-fluoro-indole.</p> Full article ">
<p>(<b>A</b>) Representative structures of tricyclic carbazole, utility molecules containing carbazole motifs, and one group of bacterial metabolites containing simple carbazole scaffolds. (<b>B</b>) The conserved essential genes responsible for the production of this group of bacterial carbazoles. The conserved genes are color-coded. <span class="html-italic">nzs</span>E: red, <span class="html-italic">nzs</span>H: blue, <span class="html-italic">nzs</span>I: pink, and <span class="html-italic">nzs</span>J: light blue. (<b>C</b>) The proposed pathways of bacterial carbazoles starting from indole-3-pyruvate catalyzed by three enzyme homologues (NzsH, NzsJ and NzsI in the case of 4). The corresponding conserved enzymes are color-coded. NzsE: red, NzsH: blue, NzsI: pink, and NzsJ: light blue. (<b>D</b>) Chemo-enzymatic synthesis of structurally diverse carbazole molecules in this study.</p> Full article ">Figure 2
<p>HPLC analysis confirming production of iso-valeryl carbazole (CA) derivative. (<b>A</b>) A biotransformation scheme starting from indole-3-pyruvate and pyruvate together with isovaleryl-SNAC to generate isovaleryl-CAs. (<b>B</b>) HPLC traces of the biotransformation from indole-3-pyruvate and pyruvate together with isovaleryl-SNACs to isovaleryl-CA. (i). indole-3-pyruvate as a control. (ii). The production of acyloins generated by NzsH-catalyzed system. (iii). The production of isovaleryl-CA generated by the three-enzyme biotransformation system, including NzsH, J, and I.</p> Full article ">Figure 3
<p>(<b>A</b>) One-pot 5-enzymatic system to generate isovaleryl-carbazole derivatives. (<b>B</b>) Extracted ion chromatography of HR-LCMS analysis confirming production of isovaleryl-carbazole (red) and 4-fluoro-isovaleryl carbazole (green). (i) the enzymatic system in the absence of indole as control; (ii) the enzymatic system in the presence of 4H-indole; (iii) the enzymatic system in the presence of 4-fluoro-indole.</p> Full article ">
Open AccessReview
Synthetic Proteins in Dental Applications
by
Christian Andrea Lopez-Ayuso, Benjamin Aranda-Herrera, Dulce Guzman-Rocha, Patricia Alejandra Chavez-Granados and Rene Garcia-Contreras
SynBio 2024, 2(1), 1-20; https://doi.org/10.3390/synbio2010001 - 27 Dec 2023
Abstract
►▼
Show Figures
Biotechnology and artificial intelligence have sparked a revolution in dentistry, with a focus on restoring natural tissue functions. This transformation has given rise to bioactive materials, inspired by biomimetics, aimed at replicating the processes found in nature. As synthetic biology advances, there is
[...] Read more.
Biotechnology and artificial intelligence have sparked a revolution in dentistry, with a focus on restoring natural tissue functions. This transformation has given rise to bioactive materials, inspired by biomimetics, aimed at replicating the processes found in nature. As synthetic biology advances, there is a heightened focus on signaling systems crucial for bio-based diagnostics and therapeutics. Dentistry now harnesses synthetic proteins for tissue regeneration and dental material enhancement. A current research priority is bacterial biofilm inhibition, vital for dental health. Given the role of Streptococcus mutans in dental caries, the development of synthetic antimicrobial peptides targeting this bacterium is underway. The balance of dental enamel between demineralization and remineralization impacts caries formation. Factors such as the presence of hydroxyapatite and salivary peptides influence enamel health. Recent studies have spotlighted salivary protein-inspired peptides for enhanced remineralization. In the realm of bone regeneration, synthetic proteins like bone morphogenetic proteins (BMP) have been spotlighted, earning FDA approval. Research is currently delving into peptides such as cementum protein 1 peptide (CEMP-1-p1) and parathyroid hormone variants like PTH (1-34), underscoring their potential in advancing dental and bone health.
Full article
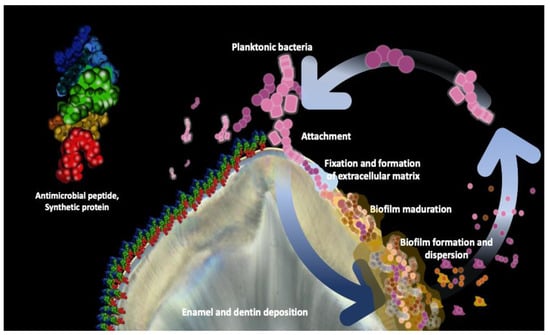
Figure 1
Figure 1
<p>Enamel biofilm formation and antimicrobial synthetic protein bioactivity. The right side shows the development by which planktonic bacteria adhere to the tooth surface and begin to form an extracellular matrix, subsequently forming communities co-aggregated with other bacteria giving rise to a more complex and dense structure called bacterial biofilm, which matures and disperses. On the left is a representative image of synthetic proteins. Their highly specific amphipathic α-helical structure interacts with hydroxyapatite through electrostatic forces, which inhibit bacterial adhesion and biofilm formation. This, in turn, promotes the exchange of mineralized ions, facilitating the remineralization process.</p> Full article ">Figure 2
<p>Understanding the significance of hydroxyapatite in ion exchange of calcium and phosphate for biomimetic remineralization. Its structure is mainly composed of repeating units of calcium phosphate; the presence of hydroxyl ions gives it unique biochemical properties, which makes it biocompatible and bioactive. The release of calcium and phosphate ions favors remineralization of tooth enamel, which is why many synthesized peptides are based on mimicking the function of this compound.</p> Full article ">Figure 3
<p><b>Dentin-pulp complex biomimetic remineralization by synthetic peptide.</b> The yellow diagrams presented in the graph represent the dentinal tissue. Intrafibrillar collagen mineralization plays a crucial role in improving the adhesion of current dental restorations. To achieve this, the use of synthetic proteins that are incorporated into collagen was investigated. These synthetic proteins promote a calcium response and, upon self-assembly, induce the formation of hydroxyapatite, which favors the remineralization of the affected tissue, increasing the protection of the dentin-pulp complex.</p> Full article ">Figure 4
<p><b>Synthetic proteins for bone regeneration.</b> This illustrative diagram showcases a collection of synthetic proteins and peptides that hold notable importance in the realm of bone regeneration within the field of dentistry—synthetic proteins for bone regeneration. Among these compounds, Bone Morphogenetic Proteins (BMPs), Abaloparatide, PTHrP (Parathyroid Hormone-Related Protein), PTH (Parathyroid Hormone), the peptide CEMP-1-p1 (Cementum-Related Peptide), rhBMP-7 (Recombinant Human Bone Morphogenetic Protein-7), rhBMP-2 (Recombinant Human Bone Morphogenetic Protein-2), and rhGDF-5 (Recombinant Human Growth/Differentiation Factor-5) are featured. Additionally, Platelet-Derived Growth Factor (PDGF) is included. Collectively, these substances represent pivotal elements in the promotion and enhancement of bone regeneration within the field of dentistry.</p> Full article ">
<p>Enamel biofilm formation and antimicrobial synthetic protein bioactivity. The right side shows the development by which planktonic bacteria adhere to the tooth surface and begin to form an extracellular matrix, subsequently forming communities co-aggregated with other bacteria giving rise to a more complex and dense structure called bacterial biofilm, which matures and disperses. On the left is a representative image of synthetic proteins. Their highly specific amphipathic α-helical structure interacts with hydroxyapatite through electrostatic forces, which inhibit bacterial adhesion and biofilm formation. This, in turn, promotes the exchange of mineralized ions, facilitating the remineralization process.</p> Full article ">Figure 2
<p>Understanding the significance of hydroxyapatite in ion exchange of calcium and phosphate for biomimetic remineralization. Its structure is mainly composed of repeating units of calcium phosphate; the presence of hydroxyl ions gives it unique biochemical properties, which makes it biocompatible and bioactive. The release of calcium and phosphate ions favors remineralization of tooth enamel, which is why many synthesized peptides are based on mimicking the function of this compound.</p> Full article ">Figure 3
<p><b>Dentin-pulp complex biomimetic remineralization by synthetic peptide.</b> The yellow diagrams presented in the graph represent the dentinal tissue. Intrafibrillar collagen mineralization plays a crucial role in improving the adhesion of current dental restorations. To achieve this, the use of synthetic proteins that are incorporated into collagen was investigated. These synthetic proteins promote a calcium response and, upon self-assembly, induce the formation of hydroxyapatite, which favors the remineralization of the affected tissue, increasing the protection of the dentin-pulp complex.</p> Full article ">Figure 4
<p><b>Synthetic proteins for bone regeneration.</b> This illustrative diagram showcases a collection of synthetic proteins and peptides that hold notable importance in the realm of bone regeneration within the field of dentistry—synthetic proteins for bone regeneration. Among these compounds, Bone Morphogenetic Proteins (BMPs), Abaloparatide, PTHrP (Parathyroid Hormone-Related Protein), PTH (Parathyroid Hormone), the peptide CEMP-1-p1 (Cementum-Related Peptide), rhBMP-7 (Recombinant Human Bone Morphogenetic Protein-7), rhBMP-2 (Recombinant Human Bone Morphogenetic Protein-2), and rhGDF-5 (Recombinant Human Growth/Differentiation Factor-5) are featured. Additionally, Platelet-Derived Growth Factor (PDGF) is included. Collectively, these substances represent pivotal elements in the promotion and enhancement of bone regeneration within the field of dentistry.</p> Full article ">
Open AccessArticle
3DNA: A Tool for Sculpting Brick-Based DNA Nanostructures
by
Shikhar Kumar Gupta, Foram Joshi, Amay Agrawal, Sourav Deb, Martin Sajfutdinow, Dixita Limbachiya, David M. Smith and Manish K. Gupta
SynBio 2023, 1(3), 226-238; https://doi.org/10.3390/synbio1030016 - 18 Dec 2023
Abstract
►▼
Show Figures
To assist in the speed and accuracy of designing brick-based DNA nanostructures, we introduce a lightweight software suite 3DNA that can be used to generate complex structures. Currently, implementation of this fabrication strategy involves working with generalized, typically commercial CAD software, ad-hoc sequence-generating
[...] Read more.
To assist in the speed and accuracy of designing brick-based DNA nanostructures, we introduce a lightweight software suite 3DNA that can be used to generate complex structures. Currently, implementation of this fabrication strategy involves working with generalized, typically commercial CAD software, ad-hoc sequence-generating scripts, and visualization software, which must often be integrated together with an experimental lab setup for handling the hundreds or thousands of constituent DNA sequences. 3DNA encapsulates the solutions to these challenges in one package by providing a customized, easy-to-use molecular canvas and back-end functionality to assist in both visualization and sequence design. The primary motivation behind this software is enabling broader use of the brick-based method for constructing rigid, 3D DNA-based nanostructures, first introduced in 2012. 3DNA is developed to provide a streamlined, real-time workflow for designing and implementing this type of 3D nanostructure by integrating different visualization and design modules. Due to its cross-platform nature, it can be used on the most popular desktop environments, i.e., Windows, Mac OS X, and various flavors of Linux. 3DNA utilizes toolbar-based navigation to create a user-friendly GUI and includes a customized feature to analyze the constituent DNA sequences. Finally, the oligonucleotide sequences themselves can either be created on the fly by a random sequence generator, or selected from a pre-existing set of sequences making up a larger molecular canvas.
Full article
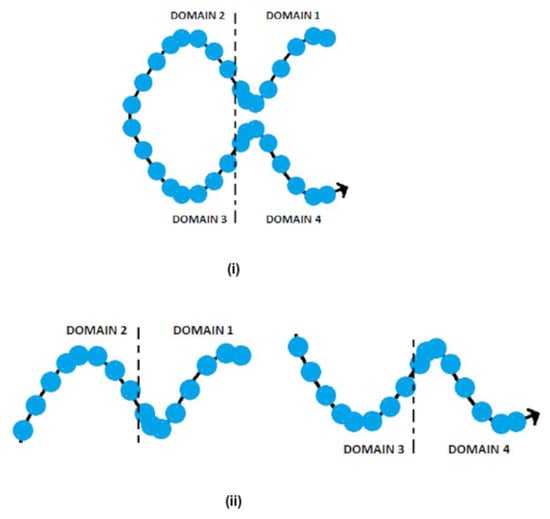
Figure 1
Figure 1
<p>The single-stranded DNA structure of (<b>i</b>) full bricks having four 8-nt domains, and (<b>ii</b>) half bricks having two 8-nt domains [<a href="#B50-synbio-01-00016" class="html-bibr">50</a>].</p> Full article ">Figure 2
<p>The full brick structure formation utilizing complementary DNA strands (indicated in identical colors) present in the respective domains [<a href="#B50-synbio-01-00016" class="html-bibr">50</a>].</p> Full article ">Figure 3
<p>The introduction of the 3DNA Software GUI.</p> Full article ">Figure 4
<p>The flowchart for the detailed process of different modules in 3DNA.</p> Full article ">Figure 5
<p>The canvas dimension input window.</p> Full article ">Figure 6
<p>The front-end view of a <math display="inline"><semantics> <mrow> <mn>6</mn> <mo>×</mo> <mn>6</mn> <mo>×</mo> <mn>6</mn> </mrow> </semantics></math> (48 bp) cuboid in the 3D modular canvas.</p> Full article ">Figure 7
<p>Advanced panel for added features.</p> Full article ">Figure 8
<p>The crystal structure formation of the <math display="inline"><semantics> <mrow> <mn>6</mn> <mo>×</mo> <mn>6</mn> <mo>×</mo> <mn>6</mn> </mrow> </semantics></math> (48 bp) cuboid (<a href="#synbio-01-00016-f006" class="html-fig">Figure 6</a>) along different directions. (<b>i</b>) elongated along <span class="html-italic">Z</span> axis, (<b>ii</b>) elongated along <span class="html-italic">X</span> axis, (<b>iii</b>) elongated along <span class="html-italic">Y</span> axis, and (<b>iv</b>) crystal formation in <math display="inline"><semantics> <mrow> <mi>X</mi> <mi>Y</mi> </mrow> </semantics></math> plane along <span class="html-italic">X</span> and <span class="html-italic">Y</span> axes.</p> Full article ">Figure 9
<p>The visualization canvas of the <math display="inline"><semantics> <mrow> <mn>6</mn> <mo>×</mo> <mn>6</mn> <mo>×</mo> <mn>6</mn> </mrow> </semantics></math> (48 bp) cuboid (<a href="#synbio-01-00016-f006" class="html-fig">Figure 6</a>) provides three different rendering modules. (<b>i</b>) Full canvas visualization, (<b>ii</b>) Plane visualization with color differentiation markers, and (<b>iii</b>) Elementary visualization.</p> Full article ">Figure 10
<p>The Bar Diagram analysis of the 6 H<math display="inline"><semantics> <mrow> <mspace width="3.33333pt"/> <mo>×</mo> <mspace width="3.33333pt"/> <mn>6</mn> </mrow> </semantics></math> H<math display="inline"><semantics> <mrow> <mspace width="3.33333pt"/> <mo>×</mo> <mspace width="3.33333pt"/> <mn>48</mn> </mrow> </semantics></math> BP cuboid in 432 domains [<a href="#B50-synbio-01-00016" class="html-bibr">50</a>].</p> Full article ">
<p>The single-stranded DNA structure of (<b>i</b>) full bricks having four 8-nt domains, and (<b>ii</b>) half bricks having two 8-nt domains [<a href="#B50-synbio-01-00016" class="html-bibr">50</a>].</p> Full article ">Figure 2
<p>The full brick structure formation utilizing complementary DNA strands (indicated in identical colors) present in the respective domains [<a href="#B50-synbio-01-00016" class="html-bibr">50</a>].</p> Full article ">Figure 3
<p>The introduction of the 3DNA Software GUI.</p> Full article ">Figure 4
<p>The flowchart for the detailed process of different modules in 3DNA.</p> Full article ">Figure 5
<p>The canvas dimension input window.</p> Full article ">Figure 6
<p>The front-end view of a <math display="inline"><semantics> <mrow> <mn>6</mn> <mo>×</mo> <mn>6</mn> <mo>×</mo> <mn>6</mn> </mrow> </semantics></math> (48 bp) cuboid in the 3D modular canvas.</p> Full article ">Figure 7
<p>Advanced panel for added features.</p> Full article ">Figure 8
<p>The crystal structure formation of the <math display="inline"><semantics> <mrow> <mn>6</mn> <mo>×</mo> <mn>6</mn> <mo>×</mo> <mn>6</mn> </mrow> </semantics></math> (48 bp) cuboid (<a href="#synbio-01-00016-f006" class="html-fig">Figure 6</a>) along different directions. (<b>i</b>) elongated along <span class="html-italic">Z</span> axis, (<b>ii</b>) elongated along <span class="html-italic">X</span> axis, (<b>iii</b>) elongated along <span class="html-italic">Y</span> axis, and (<b>iv</b>) crystal formation in <math display="inline"><semantics> <mrow> <mi>X</mi> <mi>Y</mi> </mrow> </semantics></math> plane along <span class="html-italic">X</span> and <span class="html-italic">Y</span> axes.</p> Full article ">Figure 9
<p>The visualization canvas of the <math display="inline"><semantics> <mrow> <mn>6</mn> <mo>×</mo> <mn>6</mn> <mo>×</mo> <mn>6</mn> </mrow> </semantics></math> (48 bp) cuboid (<a href="#synbio-01-00016-f006" class="html-fig">Figure 6</a>) provides three different rendering modules. (<b>i</b>) Full canvas visualization, (<b>ii</b>) Plane visualization with color differentiation markers, and (<b>iii</b>) Elementary visualization.</p> Full article ">Figure 10
<p>The Bar Diagram analysis of the 6 H<math display="inline"><semantics> <mrow> <mspace width="3.33333pt"/> <mo>×</mo> <mspace width="3.33333pt"/> <mn>6</mn> </mrow> </semantics></math> H<math display="inline"><semantics> <mrow> <mspace width="3.33333pt"/> <mo>×</mo> <mspace width="3.33333pt"/> <mn>48</mn> </mrow> </semantics></math> BP cuboid in 432 domains [<a href="#B50-synbio-01-00016" class="html-bibr">50</a>].</p> Full article ">
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics

Conferences
Special Issues
Special Issue in
SynBio
AI-Powered Synthetic Biology: Emerging Technologies and Biosecurity
Guest Editor: Masahito YamagataDeadline: 20 December 2024
Special Issue in
SynBio
Programmable Proteins in Synthetic Biology
Guest Editor: Masahito YamagataDeadline: 30 December 2024




