<p>Examples of the structure of retrotransposons. (<b>a</b>) Structure of LINE-1: Autonomous elements (such as LINE-1) encode the protein activities essential for retrotransposition, such as a reverse transcriptase and an endonuclease. The arrow at the 5′ end of retrotransposons indicates the transcription start site from their internal promoter. (<b>b</b>) Structure of Alu: Nonautonomous elements (such as Alu) do not encode proteins and their retrotransposition relies on proteins encoded by autonomous elements [<a href="#B15-epigenomes-08-00035" class="html-bibr">15</a>]. ORF: open reading frame; UTR: untranslated-region sequences; EN: endonuclease; Pol II: RNA polymerase II promoter; Pol III: RNA polymerase III promoter (bars labeled A and B); C: denotes cysteine-rich domain (encoded by ORF2); A-rich: adenosine-rich (AR) linker.</p> Full article ">
Epigenetic Mechanisms in Diabetes Research
A topical collection in Epigenomes (ISSN 2075-4655).
Viewed by 14118
Editor
Interests: diabetes; clinical and molecular epigenetics; chromatin; DNA and RNA methylation; histones; transcriptional regulation; computational epigenomics; metabolic disease
Special Issues, Collections and Topics in MDPI journals
Topical Collection Information
Dear Colleagues,
The rising global epidemic of obesity, cardiovascular disease and diabetes underscores the current challenge of characterizing the metabolically responsive epigenome. Recent studies investigating the molecular mechanisms linking fat and glucose metabolism with the epigenome highlight important regulatory determinants that underpin the expression of genes implicated in disease yet remain poorly understood. Epigenomes is now accepting submissions for a collection on "Epigenetic Mechanisms in Diabetes Research". This Topical Collection is edited by Professor Sam El-Osta from Monash University and will accept original research results and reviews, as well as special interest topics related to methods and resources of exceptional interest on topics related to metabolic regulation by epigenetic mechanisms in health and disease. You can find all the information related to the new Topical Collection at the following link: https://www.mdpi.com/journal/epigenomes/special_issues/diabetes.
We kindly invite you to submit your unpublished work to this Epigenomes Topical Collection, "Epigenetic Mechanisms in Diabetes Research".
I do hope you will join me in this Topical Collection.
Prof. Dr. Assam El-Osta
Collection Editor
Manuscript Submission Information
Manuscripts should be submitted online at www.mdpi.com by registering and logging in to this website. Once you are registered, click here to go to the submission form. Manuscripts can be submitted until the deadline. All submissions that pass pre-check are peer-reviewed. Accepted papers will be published continuously in the journal (as soon as accepted) and will be listed together on the collection website. Research articles, review articles as well as short communications are invited. For planned papers, a title and short abstract (about 100 words) can be sent to the Editorial Office for announcement on this website.
Submitted manuscripts should not have been published previously, nor be under consideration for publication elsewhere (except conference proceedings papers). All manuscripts are thoroughly refereed through a single-blind peer-review process. A guide for authors and other relevant information for submission of manuscripts is available on the Instructions for Authors page. Epigenomes is an international peer-reviewed open access quarterly journal published by MDPI.
Please visit the Instructions for Authors page before submitting a manuscript. The Article Processing Charge (APC) for publication in this open access journal is 1500 CHF (Swiss Francs). Submitted papers should be well formatted and use good English. Authors may use MDPI's English editing service prior to publication or during author revisions.
2024
Jump to: 2021
<p>Examples of the structure of retrotransposons. (<b>a</b>) Structure of LINE-1: Autonomous elements (such as LINE-1) encode the protein activities essential for retrotransposition, such as a reverse transcriptase and an endonuclease. The arrow at the 5′ end of retrotransposons indicates the transcription start site from their internal promoter. (<b>b</b>) Structure of Alu: Nonautonomous elements (such as Alu) do not encode proteins and their retrotransposition relies on proteins encoded by autonomous elements [<a href="#B15-epigenomes-08-00035" class="html-bibr">15</a>]. ORF: open reading frame; UTR: untranslated-region sequences; EN: endonuclease; Pol II: RNA polymerase II promoter; Pol III: RNA polymerase III promoter (bars labeled A and B); C: denotes cysteine-rich domain (encoded by ORF2); A-rich: adenosine-rich (AR) linker.</p> Full article ">
<p>Regulation of SASP by BET proteins and identification of Brd4-associated proteins during senescence in NIT-1 cells. (<b>A</b>) Luminex assays measuring the concentration of SASP factors Pai1 and Igfbp3 in the 24 h conditioned media collected from NIT-1 cells control (vehicle, DMSO), or senescent cells (0.25 µM etoposide treatment) at 11 days post-etoposide washout. Following drug washout, cells were treated once with vehicle or iBET-762 (5 µM) until harvest. (<b>B</b>) A repeated experiment as in (<b>A</b>) except for assaying Igfpb3 secretion by normalizing the amount in the conditioned media to the viable cell counts at the end of the assay at day 7 post-etoposide washout. (<b>C</b>) Immunoprecipitation (IP) of Brd4 followed by Western blots for Brd4 and p53 from 100 µg of nuclear extract protein of control or senescent NIT-1 cells at day 6 post-etoposide treatment. Input is 3, 5, and 10 µg of nuclear extract and rabbit IgG was the species matched control IP. (<b>D</b>) Mass spectrometry was performed on IgG control and Brd4 IPs from senescent NIT-1 cell nuclear extracts in two independent experiments on senescent NIT-1 cells at day 6 post-etoposide treatment. The number of proteins identified uniquely or in common in the datasets are shown. Brd4 IP experiment 1 and 2 identified a total of 24 common accessions (22 different proteins). (<b>E</b>) Table listing the 22 different proteins in the Brd4 IP1 and IP2. Brd4 is highlighted in red, subunits of the Ino80 complex are in blue, and the Mediator complex is in green. ns, not significant, * <span class="html-italic">p</span> < 0.05, ** <span class="html-italic">p</span> < 0.005, one-way ANOVA.</p> Full article ">Figure 2
<p>Characterization of Ino80 protein and <span class="html-italic">INO80</span> gene expression during T1D in NOD mice and humans, respectively. (<b>A</b>) Western blot analysis validating Ino80 antibody on a recombinant GST-tagged partial Ino80 protein (~90 kDa in size) with 200, 100, 50, 25 ng loaded per lane. (<b>B</b>) Western blot of Ino80 on 10 or 20 µg of nuclear extracts from human 293T cells as a control, or control and senescent NIT-1 cells at day 5 post-etoposide washout. Kap1 was a loading control. (<b>C</b>) Western blot of Ino80 on 10 µg of total RIPA protein extracts from 293T cells, C57BL6 3-month-old male testis and isolated islets from female NOD mice at indicated ages, including a 16-week-old diabetic NOD female. Beta-actin was a loading control. The 180 kDa and 130 kDa Ino80 bands from each sample were quantified relative to the beta-actin bands and plotted. (<b>D</b>) Immunohistochemistry and fluorescence staining of Ino80 along with insulin and DAPI as a nuclear counterstain on euglycemic NOD female mouse pancreas sections at 6 weeks, 14 weeks, and 17 weeks of age. Scale bars indicate 50 µm. (<b>E</b>) Analysis of publicly available single-cell RNA-seq data from PANC-DB (Human Pancreas Analysis Project) using CellxGene, comparing expression of <span class="html-italic">INO80</span> with <span class="html-italic">BRD4</span> across all beta cells in control donors (n = 27,160) or in autoantibody-positive (AA+) and T1D donors (n = 9238). Dots are colored according to <span class="html-italic">BRD4</span> expression levels. <span class="html-italic">Y</span>-axis reports binned expression level of <span class="html-italic">INO80</span> read-counts while the <span class="html-italic">X</span>-axis reports binned expression level of <span class="html-italic">BRD4</span> read-counts.</p> Full article ">Figure 3
<p>RNAi knockdown of <span class="html-italic">Cdkn1a</span> during senescence in NIT-1 cells. (<b>A</b>,<b>B</b>) Viability (gated R3) and transfection efficiency (gated R5) histograms and quantifications of NIT-1 cells transfected with 25 nM siGLO-FITC or mock transfected, 24 h post-transfection. (<b>C</b>) qRT-PCR analysis of <span class="html-italic">Cdkn1a</span> or <span class="html-italic">Ppia</span> as a control gene in control or etoposide induced senescent NIT-1 cells (72 h post-etoposide treatment) at 24 h post-transfection with control non-targeting siRNA (siCtrl), or <span class="html-italic">Cdkn1a</span> siRNA. N = 3 or 4 biological replicates per KD, error bars are S.D. ns = not significant, ** <span class="html-italic">p</span> < 0.005, one-way ANOVAs. (<b>D</b>) Western blot analysis of p21 on whole cell protein extracts prepared from NIT-1 cells transfected with the indicated siRNAs at 24 h or 72 h post-transfection. Beta-actin was as a loading control. (<b>E</b>) Western blot analysis of p21 levels after transfection with control or <span class="html-italic">Cdkn1a</span> siRNA at 72 h post-transfection in control/non-senescent NIT-1 cells in n = 3 biological replicates per transfection. Vinculin was a loading control. Plot shows quantification of p21 normalized to vinculin, error bars are S.D. * <span class="html-italic">p</span> < 0.05 two-tailed <span class="html-italic">t</span>-test. (<b>F</b>) Upper plot: viability assay using trypan blue staining and automated cell counting (Bio-Rad TC-20) of control or senescent NIT-1 cells transfected with the indicated siRNAs at 24 h post-transfection. Data are n = 10–12 biological replicates (upper plot), error bars are S.D. <span class="html-italic">Lower plot</span>: viability assay as in upper plot except 48 h following a second transfection which was conducted 2 days after the first transfection in control or senescent NIT-1 cells. Data are n = 3 or 4 biological replicates, error bars are S.D. ns = not significant, ** <span class="html-italic">p</span> < 0.005, *** <span class="html-italic">p</span> < 0.0005, two-tailed T-tests with multiple comparisons corrections (<b>C</b>,<b>E</b>) or two-way ANOVA in (<b>F</b>).</p> Full article ">Figure 4
<p>SASP gene activation is unaffected by KD of <span class="html-italic">Cdkn1a</span> in NIT-1 cells Control or senescent NIT-1 cells were transfected with indicated siRNAs and harvested 24 h post-transfection for qRT-PCR analysis of SASP genes <span class="html-italic">Igfbp3</span> and <span class="html-italic">Serpine1</span> (encoding Pai1). Data are n = 3 or 4 biological replicates, error bars are S.D. ns = not significant, ** <span class="html-italic">p</span> < 0.005, **** <span class="html-italic">p</span> < 0.0001, two-way ANOVA.</p> Full article ">
2021
Jump to: 2024
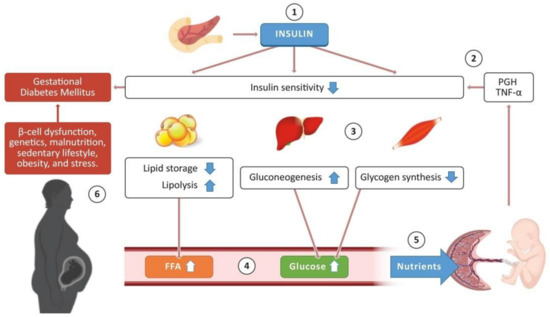
<p>Schematic representation of the glucose-insulin metabolism in pregnancy. The pancreas releases insulin, which promotes different processes in some tissues to regulate glucose concentration in the bloodstream (1). During a healthy pregnancy, the placenta produces the placental growth hormone (PGH) and proinflammatory cytokines such as tumor necrosis factor-alpha (TNF-α), promoting a decrease in insulin sensitivity in adipose tissue, liver, and skeletal muscle (2). Consequently, adipose tissue stops lipid storage and activates lipolysis; the liver enhances the production of endogenous glucose (gluconeogenesis), and glycogen synthesis decreases in skeletal muscle (3). These processes promote an increase in free fatty acids (FFA) and glucose concentration in the bloodstream (4), which are required as nutrients for the development of the placenta and the fetus (5). Nevertheless, some pregnant women present risk factors that trigger the development of gestational diabetes mellitus (6). Some images in this figure were taken from <a href="https://biorender.com/" target="_blank">https://biorender.com/</a> (last accessed on 25 March 2021).</p> Full article ">Figure 2
<p>GDM and obesity are chronic diseases often present in pregnant women. These may induce an aberrant epigenetic flow, triggering pregnancy complications and postnatal metabolic disorders for the baby, such as macrosomia, hypoglycemia, and hyperinsulinemia. Environmental factors (such as diet, activity, or stress) may increase the incidence of type 2 diabetes mellitus (T2DM), obesity, cardiovascular diseases (CVD), and metabolic syndrome; they may also promote the inheritance of aberrant epigenetic marks. Some images in this figure were taken from <a href="https://biorender.com/" target="_blank">https://biorender.com/</a> (last accessed on 25 March 2021).</p> Full article ">Figure 3
<p>Alterations of DNA methylation and miRNA expression in the placenta from women with GDM will alter the expression and function of genes involved in metabolic and cellular pathways. These alterations will have potential implications for the offspring such as hyperinsulinemia, hypoglycemia, macrosomia, and lower viability of pancreatic β-cells. These neonate disorders may, in turn, produce an increased susceptibility to developing obesity, diabetes, metabolic syndrome, and cardiovascular diseases later in life. Some images in this figure were taken from <a href="https://biorender.com/" target="_blank">https://biorender.com/</a> (last accessed on 25 March 2021).</p> Full article ">