WO2025147583A1 - Engineered pd-1 ligands - Google Patents
Engineered pd-1 ligands Download PDFInfo
- Publication number
- WO2025147583A1 WO2025147583A1 PCT/US2025/010194 US2025010194W WO2025147583A1 WO 2025147583 A1 WO2025147583 A1 WO 2025147583A1 US 2025010194 W US2025010194 W US 2025010194W WO 2025147583 A1 WO2025147583 A1 WO 2025147583A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- seq
- engineered
- protein
- icd
- sequence
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/705—Receptors; Cell surface antigens; Cell surface determinants
- C07K14/70503—Immunoglobulin superfamily
- C07K14/70532—B7 molecules, e.g. CD80, CD86
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P37/00—Drugs for immunological or allergic disorders
- A61P37/02—Immunomodulators
- A61P37/06—Immunosuppressants, e.g. drugs for graft rejection
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/01—Fusion polypeptide containing a localisation/targetting motif
- C07K2319/03—Fusion polypeptide containing a localisation/targetting motif containing a transmembrane segment
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/01—Fusion polypeptide containing a localisation/targetting motif
- C07K2319/035—Fusion polypeptide containing a localisation/targetting motif containing a signal for targeting to the external surface of a cell, e.g. to the outer membrane of Gram negative bacteria, GPI- anchored eukaryote proteins
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/31—Fusion polypeptide fusions, other than Fc, for prolonged plasma life, e.g. albumin
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/90—Fusion polypeptide containing a motif for post-translational modification
- C07K2319/91—Fusion polypeptide containing a motif for post-translational modification containing a motif for glycosylation
- C07K2319/912—Fusion polypeptide containing a motif for post-translational modification containing a motif for glycosylation containing a GPI (phosphatidyl-inositol glycane) anchor
Definitions
- PD-L1 Programmed death-ligand 1
- CD274 also known as CD274 or B7-H1
- PD-L1 has been shown to suppress adaptive immunity by binding to its natural ligand, programmed cell death protein 1 (PD-1).
- PD-1 is encoded by the CD279 gene and expressed on the surface of immune cells such as T cells.
- the binding of PD-L1 to PD-1 transmits an inhibitory signal to the immune cells. This inhibitory signal dampens effector and helper T cells’ proliferation and functional activity, such as cytokine secretion and cytotoxicity.
- PD-L1/PD-1 interaction also reduces apoptosis of regulatory T cells, further enhancing the immunosuppressive function of these cells.
- the dysregulation of the PD-L1/PD-1 immune checkpoint pathway may lead to autoimmunity in humans.
- low PD-1/PD-L1 expression levels can lead autoreactive T cells to target and induce apoptosis in hepatocytes, triggering a harmful autoimmune response against the liver.
- supplemental PD-L1 expression in a vulnerable tissue is a promising strategy to inhibit autoimmune responses against the tissue, PD-L1 has a short half-life. Exogenous expression of the wildtype protein may not be sufficient to fully suppress autoimmune activity in cells.
- the present disclosure provides an engineered programmed cell death 1 ligand (PD-L1) protein comprising an extracellular sequence of a PD-L1 and a transmembrane domain (TM) and/or an intracellular domain (ICD) that is heterologous to PD-L1, wherein the engineered PD-L1 protein has an increased half-life than the corresponding wildtype PD- L1 when expressed in a mammalian cell.
- PD-L1 programmed cell death 1 ligand
- the PD-L1 is human PD-L1, optionally wherein the extracellular sequence comprises SEQ ID NO:24 or an amino acid sequence at least 95% identical thereto.
- the extracellular sequence comprises amino acids 1- 219, 1-210, or 1-200 of SEQ ID NO:24, or a variant amino acid sequence at least 95% identical to amino acids 1-219, 1-210, or 1-200 of SEQ ID NO:24.
- the amino acid sequence at least 95% identical to SEQ ID NO:24 comprises one or more mutations relative to SEQ ID NO: 1 selected from N-to-Q mutations, I54E/Q, Y56H/F, E58F/M, R113T, M115L, S117G/A, G119K, G120V, A121W, R238A, an insertion ofW between wildtype residues A121 and D122, and a deletion between amino acids 230-238, optionally wherein the variant amino acid sequence comprises (i) an extracellular sequence of SEQ ID NO:54, 55, 56, 57, 67, 68, or 79, or (ii) SEQ ID NO:73, 74, 75, 76, 77, 78, or 80.
- the TM domain comprises a wildtype PD-L1 TM sequence, optionally SEQ ID NO:25.
- the TM domain comprises a TM sequence from integrin alpha-M (ITGAM), optionally SEQ ID NO:31, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO: 18, with or without the signal sequence shown in SEQ ID NO:23.
- the TM domain comprises a TM sequence from carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM1), optionally SEQ ID NO:32, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO: 19, with or without the signal sequence shown in SEQ ID NO:23.
- the TM domain comprises a TM sequence from integrin alpha-1 (ITGA1), optionally SEQ ID NO:33, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:20, with or without the signal sequence shown in SEQ ID NO:23.
- the ICD is a PD-L1 -derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:26, 34, 35, or 36.
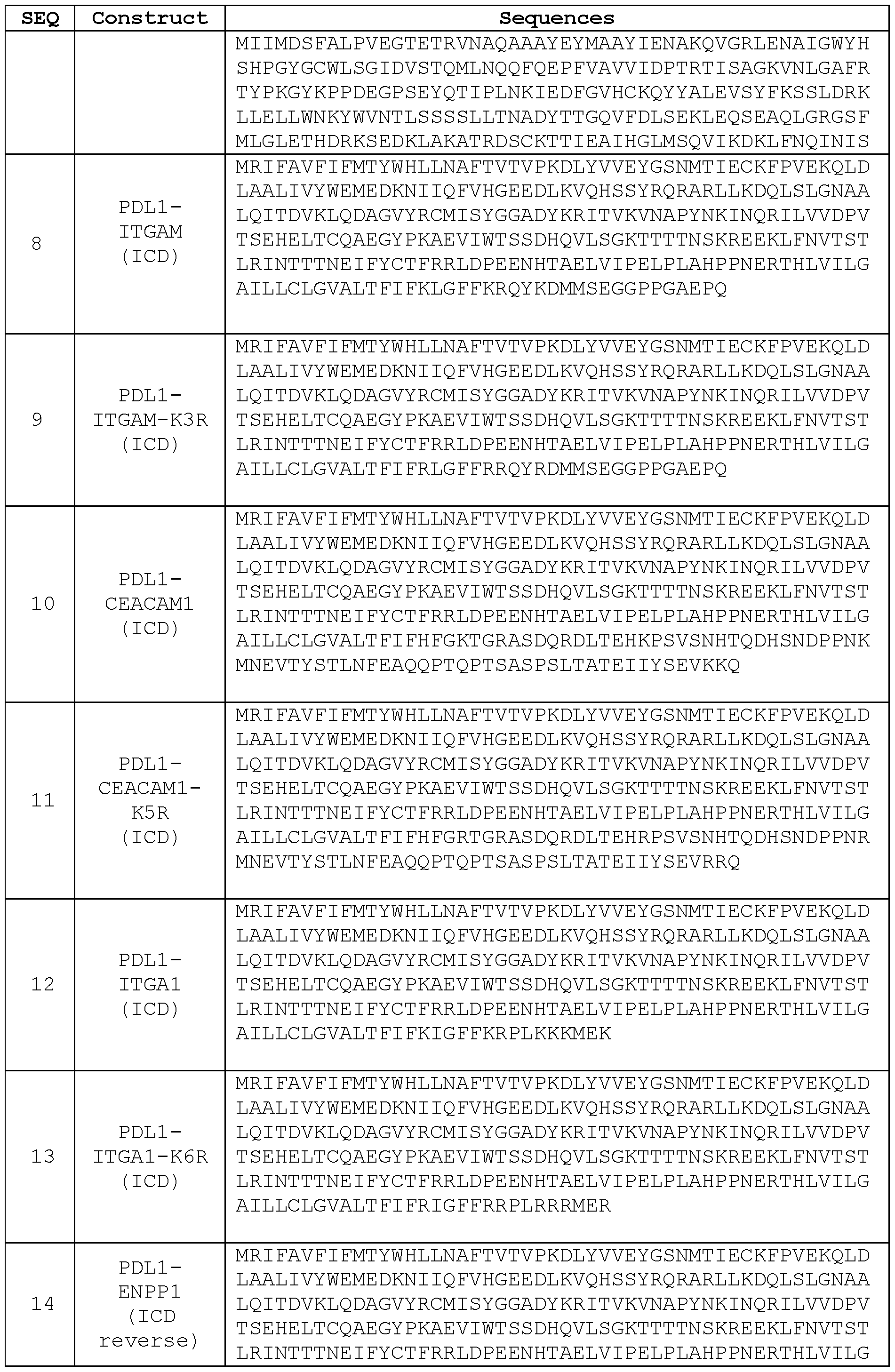
- the engineered PD-L1 protein comprises SEQ ID NO:2, 3, or 4, with or without the signal sequence shown in SEQ ID NO: 23.
- the ICD comprises a heterologous polypeptide sequence fused, optionally through a peptide linker, to the C-terminus of a PD-L1 -derived ICD sequence.
- the heterologous polypeptide is a CMTM6 sequence, optionally SEQ ID NO:47, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:6, with or without the signal sequence shown in SEQ ID NO:23.
- the heterologous polypeptide is a CSN5 sequence, optionally SEQ ID NO:48, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:7, with or without the signal sequence shown in SEQ ID NO:23.
- the ICD comprises an ITGAM-derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:37 or 38.
- the engineered PD-L1 protein comprises SEQ ID NO:8 or 9, with or without the signal sequence shown in SEQ ID NO:23.
- the ICD comprises a CEACAM1 -derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:39 or 40.
- the engineered PD-L1 protein comprises SEQ ID NO: 10, 11, 67, or 68, with or without the signal sequence shown in SEQ ID NO: 23.
- the ICD comprises an ITGA1 -derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:41 or 42.
- the engineered PD-L1 protein comprises SEQ ID NO: 12 or 13, with or without the signal sequence shown in SEQ ID NO: 23.
- the ICD comprises an ENPP1 -derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:43 or 44.
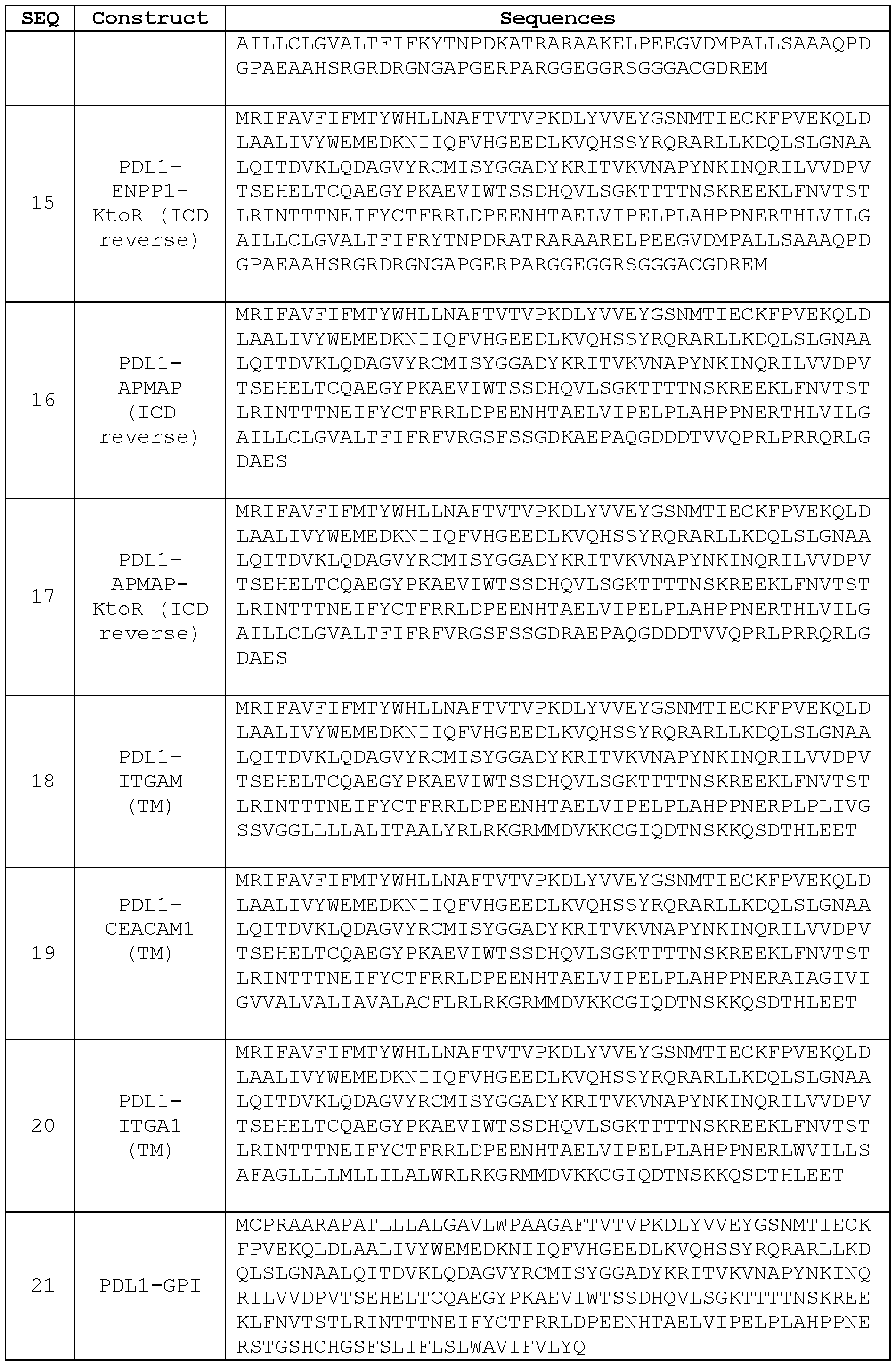
- the engineered PD-L1 protein comprises SEQ ID NO: 14 or 15, with or without the signal sequence shown in SEQ ID NO: 23.
- the ICD comprises an APMAP-derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:45 or 46.
- the engineered PD-L1 protein comprises SEQ ID NO: 16 or 17, with or without the signal sequence shown in SEQ ID NO: 23.
- the extracellular sequence of a PD-L1 is fused to a GPI anchor, optionally comprising SEQ ID NO:50, further optionally wherein the engineered PD- L1 protein comprises SEQ ID NO:21.
- the extracellular sequence of a PD-L1 is fused, with or without a peptide linker, to a GPI-anchored protein, optionally NT5E.
- the GPI-anchored protein comprises SEQ ID NO:51, optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:22.
- the engineered PD-L1 protein comprises SEQ ID NO:5, with or without the signal sequence shown in SEQ ID NO:23, or comprising SEQ ID NO:73, 74, 76, 77, or 80.
- the engineered PD-L1 protein herein further comprises a signal sequence, wherein the signal sequence comprises a PD-Ll signal sequence, optionally SEQ ID NO:23; or a NT5E signal sequence, optionally SEQ ID NO:29.
- the present disclosure provides a nucleic acid molecule comprising a coding sequence for the engineered PD-L1 protein herein.
- the nucleic acid molecule is DNA, mRNA, or circular RNA.
- a recombinant virus comprising the nucleic acid molecule herein.
- the present disclosure provides a lipid nanoparticle comprising an mRNA or circular RNA herein.
- the present disclosure provides a pharmaceutical composition
- a pharmaceutical composition comprising the nucleic acid molecule, recombinant virus, or lipid nanoparticle herein, and a pharmaceutically acceptable carrier.
- the present disclosure provides a method of treating an autoimmune condition of a tissue in a patient in need thereof, comprising administering the pharmaceutical composition herein to the patient such that the engineered PD-L1 protein is expressed in the tissue.
- a method of treating an autoimmune condition of a tissue in a patient in need thereof comprising administering the pharmaceutical composition herein to the patient such that the engineered PD-L1 protein is expressed in the tissue.
- use of the nucleic acid molecule, recombinant virus, or lipid nanoparticle herein for the manufacture of a medicament for treating an autoimmune condition in a patient in need thereof.
- the nucleic acid molecule, recombinant virus, lipid nanoparticle, or pharmaceutical composition herein for use in treating an autoimmune condition in a patient in need thereof.
- the tissue is the liver and the autoimmune condition is autoimmune hepatitis.
- the present disclosure provides a method of preventing rejection of an organ transplant in a patient in need thereof, comprising administering the pharmaceutical composition herein to the transplant such that the engineered PD-L1 protein is expressed in the transplant.
- a method of preventing rejection of an organ transplant in a patient in need thereof comprising administering the pharmaceutical composition herein to the transplant such that the engineered PD-L1 protein is expressed in the transplant.
- use of the nucleic acid molecule, recombinant virus, or lipid nanoparticle herein for the manufacture of a medicament for preventing rejection of an organ transplant in a patient in need thereof.
- the nucleic acid molecule, recombinant virus, lipid nanoparticle, or pharmaceutical composition herein for use in preventing rejection of an organ transplant in a patient in need thereof.
- FIG. 1 is a schematic showing the function of the PD-1 ligand, PD-L1, in the inhibition of aberrant immune responses and prevention of apoptosis of hepatocytes expressing PD-L1 at their cell surface.
- FIG. 2 is a series of flow cytometry plots demonstrating the effectiveness of detecting PD-L1 expression on CHO-K1 cells using an anti-PD-Ll antibody (top) or Alexa Fluor 647-conjugated human PD-1 extracellular domain (ECD) (bottom).
- FIGs. 3A-B are bar graphs showing the expression levels of various engineered PD-L1 fusion proteins expressed from DNA plasmids in CHO-K1 cells, as compared to wildtype (WT) PD-L1 (FIG. 3A), and the relative binding ability of various engineered PD- L1 fusion proteins to PD-1 compared to wildtype PD-L1 (FIG. 3B).
- FIG. 4 is a set of heat maps showing the change over a 96-hour period in the percentage of PD-L1 -positive, viable CHO-K1 cells transfected with plasmid DNA encoding a PD-L1 chimera (left) and the geometric mean expression of the PD-L1 chimeras in positive cells (right).
- FIG. 5 is a set of heat maps showing the change over a nine-day period in the percentage of PD-L1 -positive, viable OVCAR-3 cells transfected with plasmid DNA encoding a PD-L1 chimera (left) and the geometric mean expression of the PD-L1 chimeras in positive cells (right).
- FIG. 6 is a set of heat maps showing the change over a seven-day period in the percentage of PD-L1 positive, viable CHO-K1 cells transfected with circular RNA (circRNA) encoding a PD-L1 chimera (left), the geometric mean expression of the PD-L1 chimeras in positive cells (center), and the net mean fluorescence intensity (MFI) of the PD-L1 chimeras on Day 7 (right).
- circRNA circular RNA
- MFI mean fluorescence intensity
- FIG. 7 is a set of heat maps showing the change over a seven-day period in the percentage of PD-L1 positive, viable HEK293T cells transfected with circRNA encoding a PD-L1 chimera (left), the geometric mean expression of the PD-L1 chimeras in positive cells (center), and the net MFI of the PD-L1 chimera on Day 7 (right).
- FIG. 8 is a set of graphs showing expression of PD-L1 after six hours as determined by flow cytometry (left two graphs), the change over 48 hours in the percentage of PD-L1 -positive cells (center two graphs), and the MFI of PD-L1 + cells (right two graphs) in either hepatocytes (top row of graphs) or immune cells (bottom row of graphs).
- the cells were obtained from the liver of mice treated with mRNA encoding PD-L1.
- the mRNA was delivered using an ionizable lipid formulation, LRN1.
- FIG. 9 is a line graph showing the amounts of PD-L1 detected in the liver of mice by an ELISA assay.
- the cells were obtained from the liver of mice treated with mRNA encoding PD-L1.
- the mRNA was delivered using an ionizable lipid formulation, LRN1.
- FIG. 10 is a set of line graphs showing the change over 120 hours in the percentage of PD-L1 -positive cells (left) and the MFI of PD-L1 + cells (right).
- the cells were obtained from the liver of mice pre-dosed with anti-IFNaR-1 monoclonal antibody and administered circular RNA encoding PD-L1.
- the circular RNA was delivered with an ionizable lipid formulation, LRN1.
- FIG. 11 is a line graph showing the change over seven days in the percentage of PD-L1 -positive cells.
- the cells were obtained from the liver of mice pre-dosed with anti- IFNaR-1 monoclonal antibody and administered circRNA encoding PD-L1 variants.
- the circular RNA was delivered with an ionizable lipid formulation (LP-01).
- FIG. 12 is a line graph showing the MFI and derived EC50 values of CHO cells transfected with mRNAs encoding PD-L1 molecules bearing various point mutations, as measured by flow cytometry.
- FIG. 13 is a plot showing the PD-L1 cell surface expression (as indicated by MFI) and PD-L1 soluble levels in the cell culture supernatant of CHO cells transfected with mRNAs encoding PD-L1 molecules bearing various point mutations.
- the present disclosure provides engineered PD-L1 proteins that have improved longevity while retaining wildtype PD-Ll’s expression levels and affinity for PD-1.
- autoimmune diseases e.g., autoimmune hepatitis
- PD-1 and PD-L1 have been observed to express PD-1 and PD-L1 at low levels, which may contribute to T cell-mediated destruction of healthy tissues (see, e.g., Agina et al., Clin Exp Hepatol. (2019) 5(3):256-64; Jilkova et al., Cells. (2021) 10(10):2671).
- Expressing the present PD-L1 proteins in specific tissues can repress aberrant immune responses against those tissues, thereby preserving viability of healthy cells.
- the present PD-L1 proteins may be expressed in liver parenchymal and residential cells to alleviate autoimmune hepatitis or other immune-driven liver diseases (e.g., primary biliary cholangitis). Additionally, the present PD-L1 proteins may also be expressed, for example, in kidney cells to alleviate lupus nephritis; in the kidney, skin, or central nervous system to alleviate systemic lupus erythematosus (SLE); in the synovium to alleviate rheumatoid arthritis; in pancreatic cells to alleviate type 1 diabetes; in cells of the salivary or lacrimal glands to alleviate Sjogren’s syndrome; in the thyroid to alleviate Graves’ disease; in oligodendrocytes to alleviate multiple sclerosis; in the central or peripheral nervous system to alleviate myasthenia gravis or neuromyelitis optica; in the intestines to alleviate Crohn’s disease, ulcerative colitis or celiac disease;
- the present PD-L1 -derived proteins can be expressed at levels approximating wildtype PD-L1 on the cell surface in several cell types and effectively bind to PD-1, indicating their potential use as a therapeutic. Additionally, these engineered proteins exhibit longer half-lives than wildtype PD-L1, suggesting their potential enhanced therapeutic efficacy compared to supplementation with wildtype PD-L1.
- the engineered PD-L1 proteins herein are derived from PD-L1 but may contain only a partial, rather than the entire, sequence of wildtype PD-L1. These polypeptides are cell surface proteins when expressed in mammalian cells.
- “derived from” means that the sequence is the same as or similar to the original sequence; a derived sequence may be longer or shorter than, or have the same length as, the original sequence.
- PD-L1 refers to human PD-L1.
- a human PD-L1 polypeptide sequence may be found at the UniProt database (Identifier No.
- Q9NZQ7-1) may have the following sequence:
- LGVALTFIFR LRKGRMMDVK KCGIQDTNSK KQSDTHLEET SEQ ID NO : 1
- the extracellular region spans amino acids 1-238.
- the extracellular region includes a signal sequence (amino acids 1-18; underlined) and two distinct extracellular domains (ECDs).
- the first ECD is an IgV-like domain that spans amino acids 19-127 and the second ECD is an IgC-like domain that spans amino acids 133-225.
- the transmembrane domain (TM) spans amino acids 239-259 (boldface and underlined).
- the intracellular domain (ICD) spans amino acids 260-290.
- the signal sequence is removed in the mature polypeptide.
- the signal sequence, the extracellular region minus the signal sequence, the TM domain, and the ICD domain of the above sequence are assigned SEQ ID NOs:23-26, respectively.
- the IgV-like domain and IgC-like domain sequences are assigned SEQ ID NOs:27 and 28, respectively.
- PD-L1 is predicted to possess a matrix metalloproteinase cleavage site that spans amino acids 230-238 (boxed; SEQ ID NO:71).
- a PD-L1 amino acid position recited herein refers to the position in SEQ ID NO: 1 or a corresponding position in a variant of SEQ ID NO: 1 (e.g., a naturally occurring polymorphic variant or a genetically engineered variant).
- PD-L1 proteins are fusion proteins with components from different sources, those proteins also are called “PD-L1 chimeras” or “PD-L1 fusion proteins” herein.
- polypeptides extracellular, transmembrane, and intracellular regions of the engineered PD-L1 proteins herein are described below.
- the extracellular region of the present engineered PD-L1 polypeptides may comprise a signal sequence that triggers the translocation of the newly synthesized PD-L1 protein into the endoplasmic reticulum for trafficking and expression to the cell surface.
- the signal sequence may be cleaved off before the protein reaches the cell membrane.
- the signal sequence comprises PD-Ll’s wildtype signal sequence MRIFAVFIFMTYWHLLNA (SEQ ID NO:23) or a functional variant thereof.
- the signal sequence may be derived from another cell surface protein or an artificial sequence.
- the signal sequence may comprise a signal sequence from human 5 ’ -nucleotidase NT5E (a.k.a. CD73), MCPRAARAPATLLLALGAVLWPAAGA (SEQ ID NO:29), or a functional variant thereof.
- the extracellular region comprises mutations in the IgV- like domain and/or the IgC-like domain sequence relative to wildtype PD-L1.
- the mutations are made in the IgC-like domain sequence.
- the IgC-like domain comprises an N-to-Q substitution at one, two, or all three positions corresponding to residue 192, 200, and 219 of SEQ ID NO: 1.
- the extracellular region comprises PD-Ll’s extracellular region with N192Q, N200Q, and N219Q substitutions.
- the extracellular region comprises SEQ ID NO:30.
- the extracellular region of the present engineered PD-L1 polypeptide comprises a part (e.g., amino acid 1 to amino acid 210, 211, 212, 213, 214, 215, 216, 217, or 218) or the entirety (i.e., amino acids 1 to 219) of SEQ ID NO:24.
- the extracellular region comprises an additional phenylalanine (F) at the N- terminus.
- the ICD of the engineered PD-L1 protein may be derived from PD-L1 or another cell surface protein, or may be completely artificial (i.e., not derived from a naturally occurring protein). Where the ICD is derived from a naturally occurring protein, it may contain the wildtype ICD or a variant thereof. The variant may, for example, contain mutations (e.g., substitution) that reduce or eradicate the signal transduction function of the ICD.
- the variant may be the “PD-L1 ICD K5R” variant (SEQ ID NO:34), which is identical to the wildtype ICD sequence except that all five lysine residues in the wildtype sequence have been substituted by arginines.
- the variant may be the “PD-L1 ICD K263R” variant (SEQ ID NO:35), which is identical to the wildtype ICD sequence except that the first lysine residue in the wildtype sequence (corresponding to K263 in SEQ ID NO:1) has been substituted by arginine.
- amino acid residues other than lysines e.g., 2, 3, 4, or more additional amino acid residues are substituted in the same or similar manner.
- the variant may be the “PD-L1 ICD KtoR,DEtoR” variant (SEQ ID NO:36), which is identical to the wildtype ICD sequence except that all lysines (5), aspartic acids (3), and glutamic acid (2) residues in the wildtype sequence have been substituted by arginines; these mutations increase the net positive charge of the ICD and help to increase interaction with the membrane and resist degradation.
- the ICD of the engineered PD-L1 protein comprises an ICD from ectonucleotide pyrophosphatase/phosphodiesterase family member 1 (ENPP1) or a variant thereof.
- ENPP1 ectonucleotide pyrophosphatase/phosphodiesterase family member 1
- the ICD from ENPP1 may be in a reverse orientation in the engineered protein.
- the ICD of the engineered protein comprises SEQ ID NO:43 (reverse of a wildtype human ENPP1 ICD), or a variant thereof.
- the variant comprises one or more substitutions (e.g., K to R) relative to the wildtype ICD sequence.
- the variant may be the “ENPP1 ICD (reverse) KtoR” variant (SEQ ID NO:44), which is identical to SEQ ID NO:43 except that all three lysine residues in it have been substituted by arginines.
- the ICD of the engineered PD-L1 protein comprises an ICD from adipocyte plasma membrane-associated protein (APMAP) or a variant thereof.
- the ICD from APMAP may be in a reverse orientation in the engineered protein.
- the ICD of the engineered protein comprises SEQ ID NO:45 (reverse of a wildtype human APMAP ICD), or a variant thereof.
- the variant comprises one or more substitutions (e.g., K to R) relative to the wildtype ICD sequence.
- the variant may be the “APMAP ICD (reverse) KtoR” variant (SEQ ID NO:46), which is identical to SEQ ID NO:45 except that the lysine residue in it has been substituted by arginine.
- the ICD of the engineered PD-L1 protein may comprise an ICD sequence of another cell surface protein with a long protein half-life (i.e., longer than 15 hours).
- additional cell surface proteins are SLC2A2, NT5E, CD47, ABCC6, STX4, FAS, and NPTN.
- the ICD of the engineered PD-L1 protein may comprise the polypeptide sequence of a second cell surface proteins with a long protein half-life (i.e., longer than 15 hours) or of another protein selected based on useful functions (e.g., proteins know to interact with the PD-L1 protein, stabilize the PD-L1 protein, or both).
- this polypeptide sequence may be fused to the ICD described above, with or without a peptide linker.
- the ICD of the engineered PD-L1 protein may comprise a wildtype PD-L1 ICD fused through a peptide linker to CKLF-like MARVEL transmembrane domain-containing protein 6 (CMTM6) (e.g., SEQ ID NO:47).
- the ICD of the engineered PD-L1 protein may comprise a wildtype PD-L1 ICD fused through a peptide linker to COP9 signalosome complex subunit 5 (CSN5) (e.g., SEQ ID NO:48).
- the peptide linker may be a flexible linker such as a GS linker.
- a GS linker is rich in glycine and serine (more than 50% of the residues are glycine and/or serine).
- the peptide linker comprises GGGGSGGGSGGGS (SEQ ID NO:49).
- the engineered PD-L1 protein is anchored in the cell membrane through a glycophosphatidylinositol (GPI) anchor.
- GPI glycophosphatidylinositol
- the extracellular region of the protein is fused to a GPI anchor sequence, e.g., SEQ ID NO:50.
- the present polypeptide comprises, consists of, or consists essentially of a PD-L1 protein sequence with one or more (e.g., one, two, three, four, five, six, seven, eight, nine, ten or more) mutations, such as substitutions.
- the engineered PD-L1 sequence is termed “PDL1-K5R,” with five K-to-R mutations (underlined) in the ICD relative to wildtype PD-L1 :
- LGVALTFI FR LRRGRMMDVR RCGIQDTNSR RQSDTHLEET SEQ ID NO : 2
- the engineered PD-L1 sequence is termed “PDL1-
- K263R contains one K-to-R mutation (underlined) in the ICD relative to wildtype PD-
- the engineered PD-L1 sequence is termed “PDL1- KtoR,DEtoR” and contains ten amino acid substitutions relative to wildtype PD-L1, where all lysines, aspartic acids, and glutamic acids in the wildtype PD-L1 ICD have been substituted with arginines (underlined):
- the engineered PD-L1 sequence is termed “PDL1 (3NQ)” and contains three N-to-Q mutations (underlined) in the IgC-like domain relative to wildtype PD-
- the engineered PD-L1 sequence is termed “PDL1-CMTM6 fusion with linker,” in which a wildtype PD-L1 sequence is fused at its C-terminus to a CMTM6 polypeptide sequence (boldfaced; SEQ ID NO:47) through a peptide linker of SEQ ID NO:49 (underlined):
- CEACAM1 in which the ICD of a wildtype PD-L1 sequence is replaced with an ICD from CEACAM1 (italic; SEQ ID NO:39): MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC
- the engineered PD-L1 sequence is termed “PDL1-ENPP1
- the reversed APMAP ICD sequence is mutated, where the lysine residue is substituted by arginine (underlined), and the engineered PD-L1 sequence is called “PDL1 -APMAP -KtoR (ICD reverse)”:
- TM in which the TM domain of a wildtype PD-L1 is replaced with a TM domain from
- the engineered PD-L1 sequence is called “PDL1-CEACAM1
- TM in which the TM domain of a wildtype PD-L1 is replaced with a TM domain from CEACAM1 (boldface; SEQ ID NO:32):
- the engineered PD-L1 sequence is called “PDL1-ITGA1
- TM in which the TM domain of a wildtype PD-L1 is replaced with a TM domain from
- the engineered PD-L1 sequence comprises the PD-Ll’s extracellular portion that interacts with PD-1 fused to a GPI anchor.
- the engineered PD-L1 sequence is called “PDL1-GPI” and comprises the extracellular region of a wildtype PD-L1 in which the signal sequence is replaced with that of a human NT5E (underlined; SEQ ID NO:29) and the C-terminus of the extracellular region is fused to a GPI anchor with the sequence of SEQ ID NO:50 (italic):
- the engineered PD-L1 sequence comprises the PD-Ll’s extracellular portion that interacts with PD-1 fused to a GPI-anchorable protein such as NT5E.
- the engineered PD-L1 sequence is called “PDL1-NT5E fusion” and comprises the extracellular region of a wildtype PD-L1 in which the signal sequence is replaced with that of a human NT5E (underlined; SEQ ID NO:29) and the C- terminus of the extracellular region is fused to a mature sequence of NT5E (i.e., without the signal sequence; italic; SEQ ID NO:51) through a short peptide linker with the sequence of SEQ ID NO: 52 (boxed):
- Variants of the above-described sequences are also within the present disclosure. Variants may be deletions, insertions, and/or substitutions. In some embodiments, the substitutions are conservative substitutions: for example, an aliphatic amino acid (e.g., glycine, alanine, valine, leucine, or isoleucine) may be substituted by another aliphatic amino acid; a hydroxyl or sulfur/selenium-containing amino acid (e.g., serine, cysteine, selenocysteine, threonine, or methionine) may be substituted by another hydroxyl or sulfur/selenium-containing amino acid; an aromatic amino acid (e.g., phenylalanine, tyrosine, or tryptophan) may be substituted by another aromatic amino acid; a branched chain amino acid (e.g., isoleucine, leucine, or valine) may be substituted by another branched chain amino acid;
- a negatively charged amino acid e.g., aspartic acid or glutamic acid
- a positively charged amino acid e.g., lysine, arginine and histidine
- a nonpolar amino acid e.g., alanine, glycine, isoleucine, leucine, methionine, phenylalanine, proline, valine, or tryptophan
- a nonpolar amino acid e.g., alanine, glycine, isoleucine, leucine, methionine, phenylalanine, proline, valine, or tryptophan
- the engineered PD-L1 sequence comprises, consists of, or consists essentially of, one of SEQ ID NOs:2-22, 67, and 68 (with or without the signal sequence (i.e., SEQ ID NO:23)), or comprises one of SEQ ID NOs:73-78, and 80.
- the present disclosure provides pharmaceutical compositions comprising a nucleic acid for expressing an engineered PD-L1 protein herein and a pharmaceutically acceptable carrier (e.g., water and phosphate-buffered solution).
- a pharmaceutically acceptable carrier e.g., water and phosphate-buffered solution.
- the pharmaceutical composition comprises a viral vector comprising the nucleic acid.
- the pharmaceutical composition comprises an RNA (e.g., mRNA or circular RNA) encoding an engineered PD-L1, wherein the RNA is encapsulated in an LNP.
- the pharmaceutical composition comprises a nucleic acid for expressing a polypeptide sequence comprises the sequence of PDL1-K5R, PDL1- APMAP-KtoR (ICD Reverse), PDLl-KtoR,DEtoR, PDL1-GPI, and PDL1-CEACAM1-K5R (ICD), or PDL1-APMAP (ICD reverse) or an amino acid sequence at least 95% (e.g., at least 96, 97, 98, or 99%) identical thereto.
- CH0-K1 epithelial cells were transfected with plasmid DNA encoding the various chimeric proteins.
- CHO-K1 cells were seeded in 24-well plates at 40,000 cells per well and grown overnight at 37°C prior to transfection with 500 ng of plasmid DNA and 1.5 pL Lipofectamine2000TM (Invitrogen) per well. The cells were then incubated at 37°C in a 5% CO2 incubator.
- This Example describes a study evaluating the half-life of the PD-L1 chimeras in transfected cells.
- CHO-K1 cells were seeded in a 24-well plate and transfected as described in Example 1. The transfected cells were incubated at 37°C for 24 hours, 48 hours, 72 hours, and 96 hours in a 5% CO2 incubator. At each timepoint, cells were washed with PBS containing 2 mM EDTA and detached with Accutase®.
- Detached cells were then stained for viability and PD-L1 expression and analyzed by FACS as described in Example 1, except anti-PD-Ll antibody clone 29EA3 (Biolegend) was used to detect PD-L1 at 1 :400 dilution for this and all subsequent experiments.
- the ten chimeras were PD-L1-K5R (degradation stabilized control), PD-L1- KtoR,DEtoR, PDL1-ENPP1 (ICD reverse), PDLl-ENPPl-KtoR (ICD reverse), PDL1- CMTM6 fusion with linker, PDL1-CEACAM1 (ICD), PDL1-CEACAM1-K5R (ICD), PDL1-APMAP (ICD reverse), PDLl-APMAP-KtoR (ICD reverse), and PDL1-GPI.
- This experiment was completed twice with CHO-K1 cells and both transfection replicates yielded consistent results.
- OVCAR-3 ovarian adenocarcinoma cells were seeded in a 96-well plate at 30,000 cells per well and grown overnight at 37°C. The cells were then transfected with 200 ng of plasmid DNA encoding the PD-L1 fusion proteins and 0.6 pL Lipofectamine2000TM (1 :3 ratio) per well. The transfected cells were incubated at 37°C for 1 day, 5 days, or 9 days in a 5% CO2 incubator with cell culture medium replaced on Day 5 for cells that were cultured for 9 days.
- DNA vectors carry safety risks because vector DNA may be incorporated into the host genome.
- circRNA also termed herein “eRNATM” for “Endless RNATM”
- eRNATM circular RNA
- CHO-K1 cells were seeded in a 96-well plate with 35,000 cells per well and grown overnight at 37°C. The cells were then transfected with 0.2 pmol eRNA encoding the PD-L1 chimeras and 0.6 pL LipofectamineTM MessengerMAXTM Transfection Reagent (Thermo Fisher) (1 :3 ratio) per well.
- the cells were cultured for 1, 2, 3, and 7 days at 37°C in a 5% CO2 incubator. For cells that were cultured for 7 days, they were split 1 : 10 on Day 3 and the culture medium was replaced on Day 5. The cells were detached, stained, and analyzed by FACS as described above.
- MFI median fluorescent intensity
- HEK293T cells a human embryonic kidney cell line.
- HEK293T cells were seeded in a 96-well plate at 15,000 cells per well and grown overnight at 37°C.
- the cells were then transfected with 35 ng eRNA encoding the PD-L1 chimeras and 0.1 pL LipofectamineTM MessengerMAXTM Transfection Reagent (2: 1 ratio) per well.
- the cells were then incubated for 1, 3, and 7 days at 37°C in a 5% CO2 incubator. For cells that were cultured for 7 days, the culture medium was replaced on Day 5.
- Cells were detached by pipetting. Detached cells were stained and analyzed by FACS as described above.
- the cells were stained with anti-CD45-AF700 (BioLegend; 1 :400), anti-CD26-APC (BioLegend; 1 :200), and anti- PD-L1-PE (BioLegend; 1 :200).
- the stained cells were washed with FACS buffer and samples were analyzed by flow cytometry.
- CHO cells were transfected with Nl-Methylpseudouridine-modified mRNAs encoding various PD-L1 mutations shown in Table 2.
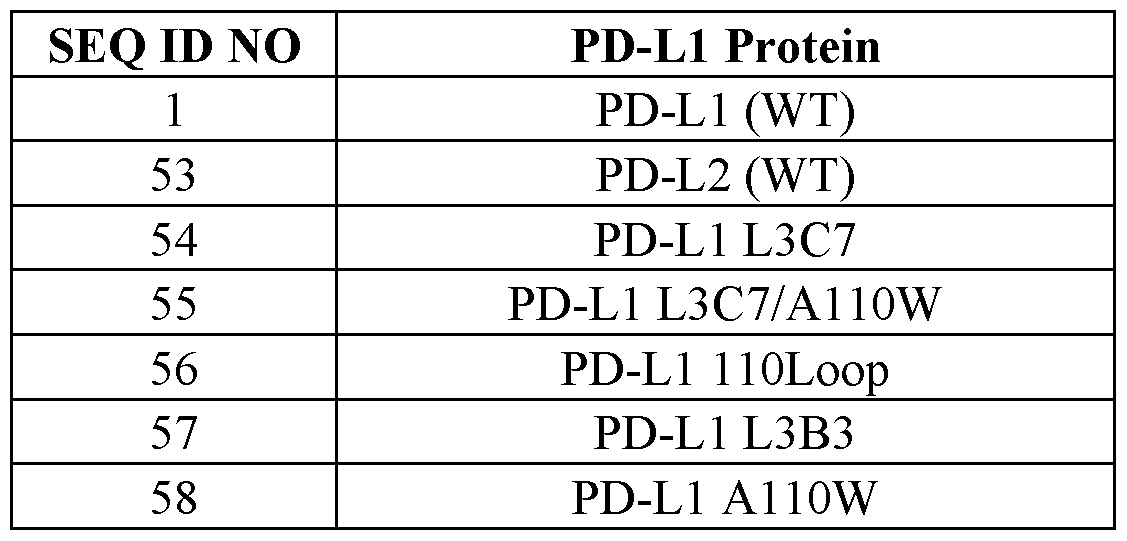
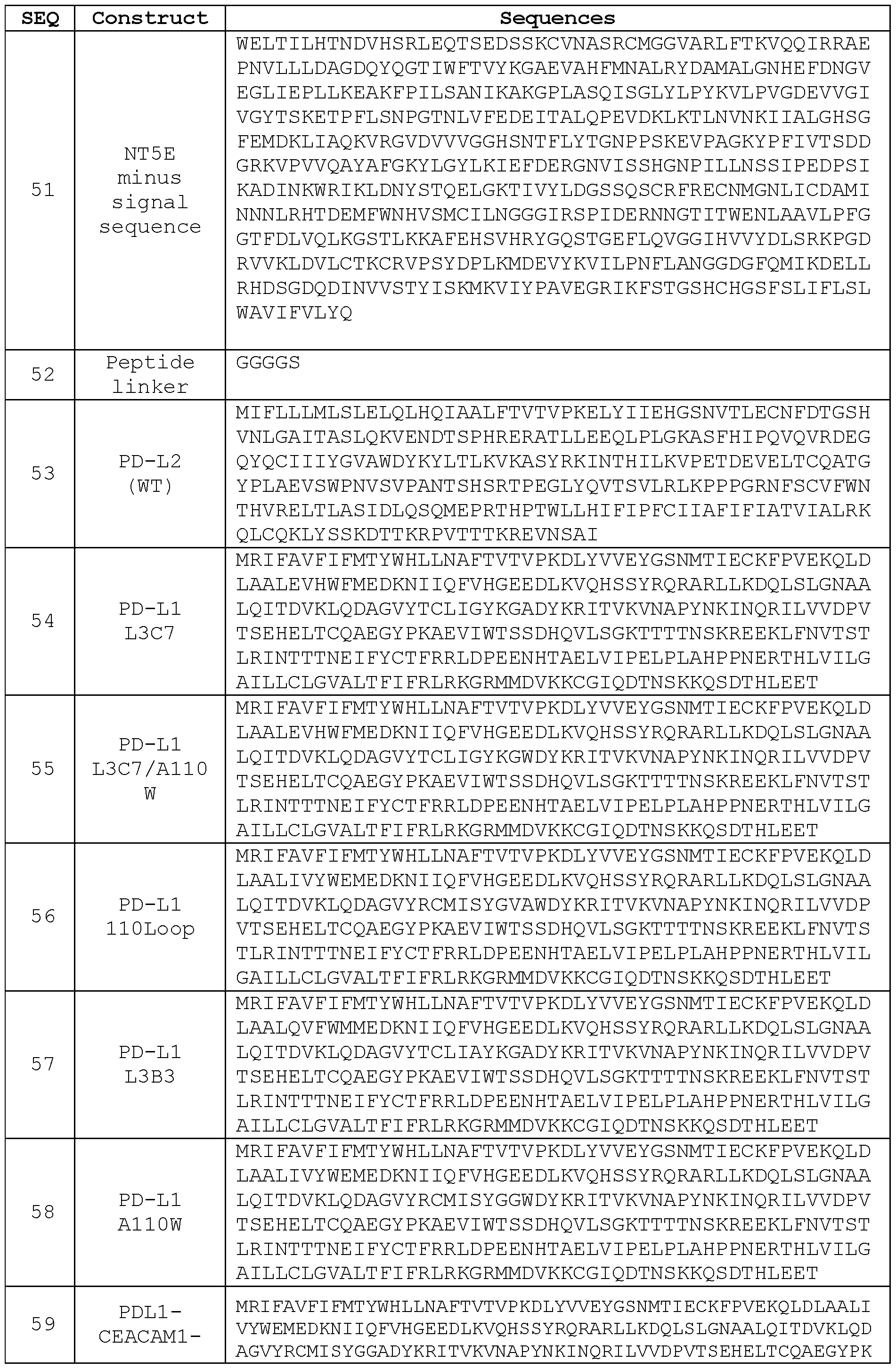
- the PD-L1 L3C7 variant (ECD shown in SEQ ID NO: 75) has the following mutations relative to wildtype PD-L1 : I54E, Y56H, E58F, R113T, Ml 15L, SI 17G, and G119K.
- the PD-L1 L3C7/A110W variant (ECD shown in SEQ ID NO:76) has one additional A121W mutation relative to PD-L1 L3C7.
- PD-L1 1 lOLoop (ECD shown in SEQ ID NO: 77) differs from wildtype PD-L1 in that the former has a G120V mutation, and an insertion of W between wildtype residues A121 and D122.
- the PD-L1 L3B3 variant (ECD shown in SEQ ID NO:78) has the following mutations relative to wildtype PD-L1 : I54Q, Y56F, E58M, R113T, Ml 15L, SI 17A, and G119K.
- the PD-L1 Al 10W variant has an A121W mutation relative to wildtype PD-L1. (The PD-L1 Al 10W variant was so named because A121 in PD-L1 corresponds to Al 10 in PD-L2.)
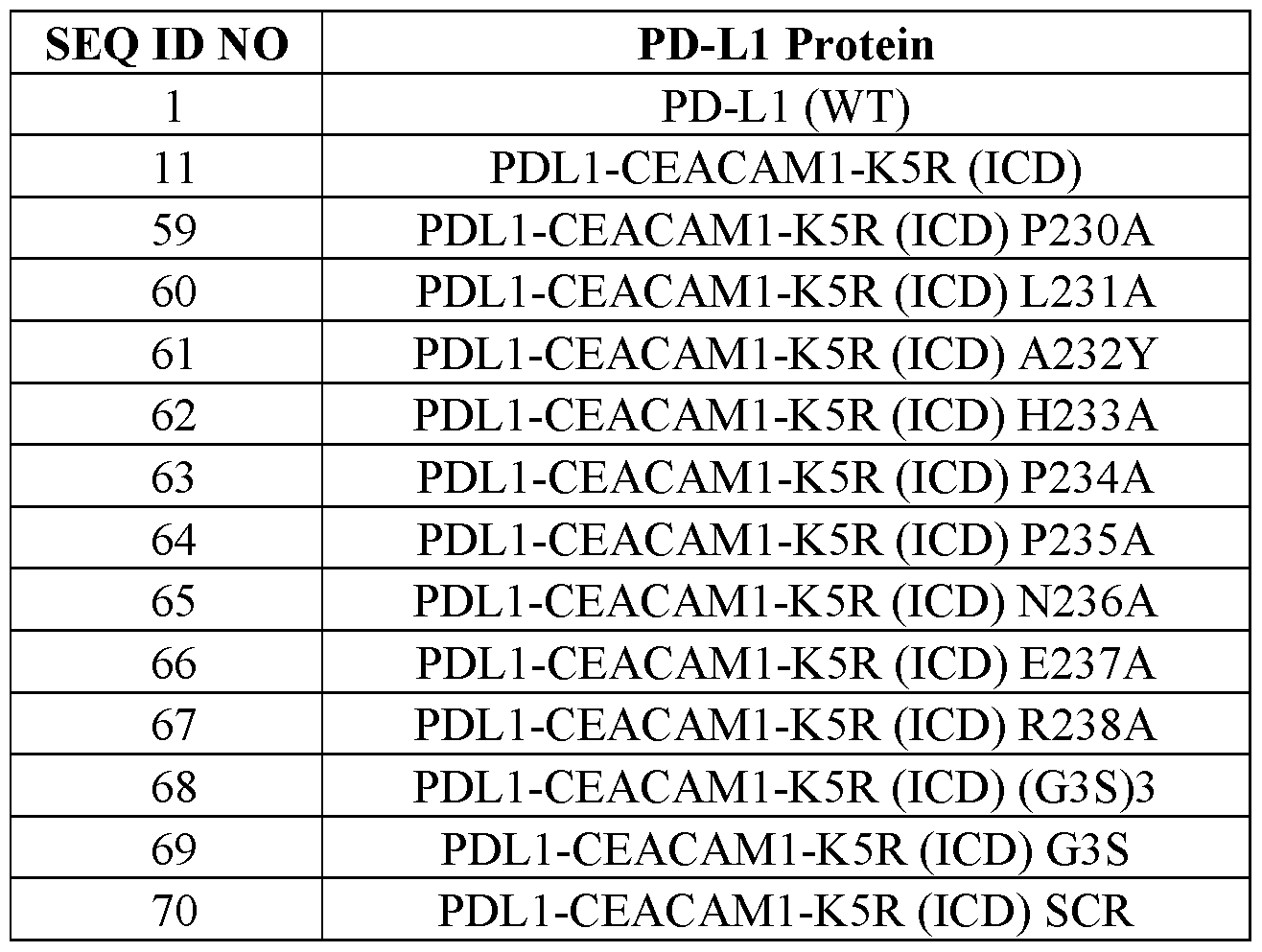
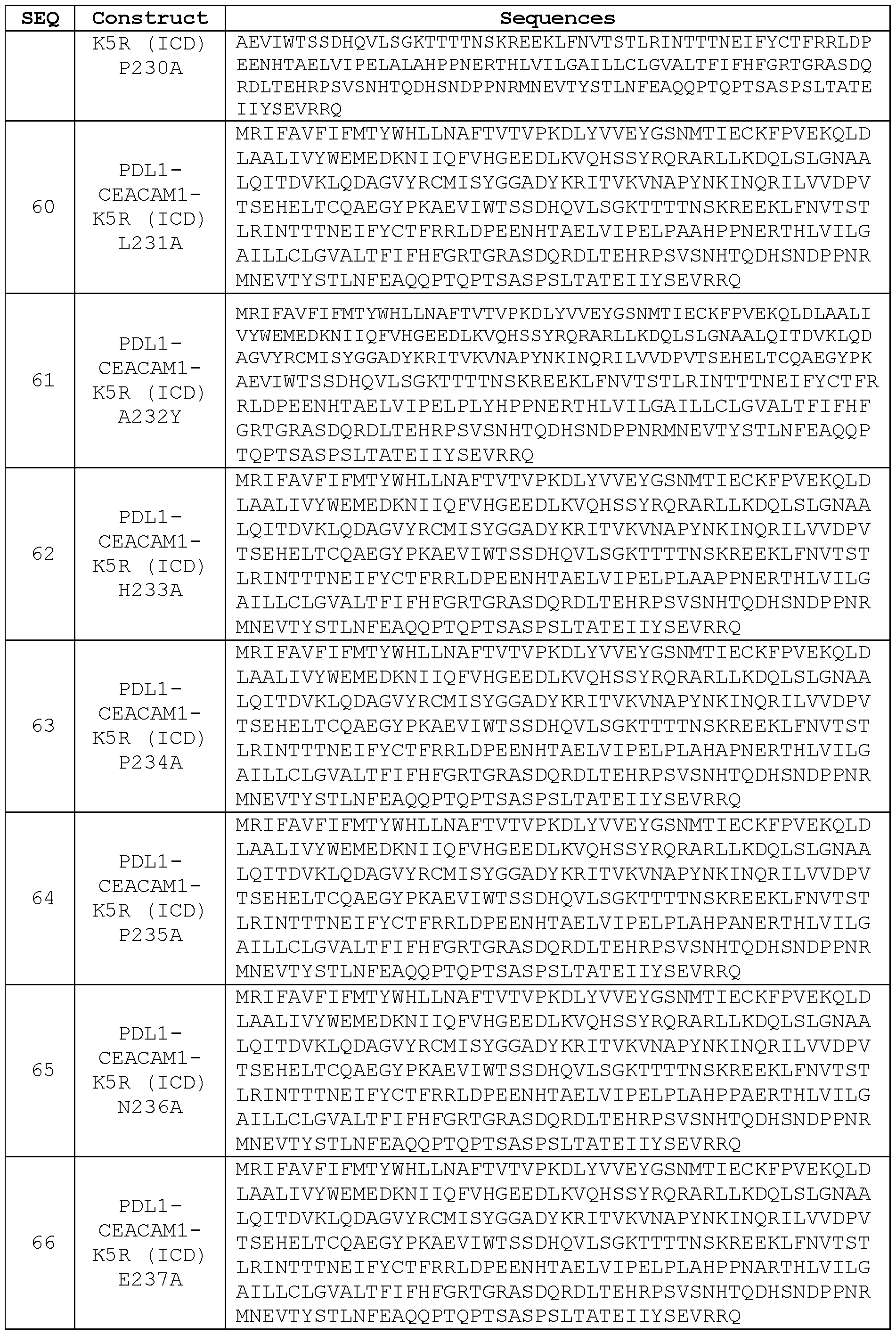
- the mutations were done in the background of CEACAM1-K5R (ICD) PD-L1.
- K5R G 3 S (SEQ ID NO:81) and (G 3 S) 3 (SEQ ID NO:72) represent the PD-L1 chimeras where the cleavage site has been substituted by one or three G 3 S (SEQ ID NO: 81) linkers, respectively, whereas the cleavage site for K5R SCR has been scrambled using the same amino acid composition.
Landscapes
- Health & Medical Sciences (AREA)
- Immunology (AREA)
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- Engineering & Computer Science (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Health & Medical Sciences (AREA)
- Medicinal Chemistry (AREA)
- General Chemical & Material Sciences (AREA)
- Toxicology (AREA)
- Pharmacology & Pharmacy (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Animal Behavior & Ethology (AREA)
- Transplantation (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
- Cell Biology (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Zoology (AREA)
- Gastroenterology & Hepatology (AREA)
- Biochemistry (AREA)
- Biophysics (AREA)
- Genetics & Genomics (AREA)
- Molecular Biology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
- Peptides Or Proteins (AREA)
Abstract
The present disclosure provides polypeptides derived from PD-1 ligands, nucleic acids encoding these polypeptides, and methods of using the polypeptides and nucleic acids for therapeutic purposes.
Description
ENGINEERED PD-1 LIGANDS
CROSS-REFERENCE TO RELATED APPLICATION
[0001] This application claims priority from U.S. Patent Application No. 63/617,869, filed January 5, 2024. The disclosure of the aforementioned priority application is incorporated by reference herein in its entirety.
SEQUENCE LISTING
[0002] The instant application contains a Sequence Listing that has been submitted electronically as an XML file and is hereby incorporated by reference in its entirety. Said electronic file, created on December 31, 2024, is named 125388_WO002_SL.xml and is 87,784 bytes in size.
BACKGROUND OF THE INVENTION
[0003] Programmed death-ligand 1 (PD-L1), also known as CD274 or B7-H1, is a cell surface protein that plays an important role in immune regulation. PD-L1 has been shown to suppress adaptive immunity by binding to its natural ligand, programmed cell death protein 1 (PD-1). PD-1 is encoded by the CD279 gene and expressed on the surface of immune cells such as T cells. The binding of PD-L1 to PD-1 transmits an inhibitory signal to the immune cells. This inhibitory signal dampens effector and helper T cells’ proliferation and functional activity, such as cytokine secretion and cytotoxicity. PD-L1/PD-1 interaction also reduces apoptosis of regulatory T cells, further enhancing the immunosuppressive function of these cells.
[0004] The dysregulation of the PD-L1/PD-1 immune checkpoint pathway may lead to autoimmunity in humans. For example, in C/U /-/-deficient rodent models or patients with autoimmune hepatitis, low PD-1/PD-L1 expression levels can lead autoreactive T cells to target and induce apoptosis in hepatocytes, triggering a harmful autoimmune response against the liver. Although supplemental PD-L1 expression in a vulnerable tissue is a promising strategy to inhibit autoimmune responses against the tissue, PD-L1 has a short half-life. Exogenous expression of the wildtype protein may not be sufficient to fully suppress autoimmune activity in cells. Thus, there remains a need for developing therapeutically useful engineered PD-L1 proteins that can be stably expressed in cells to mimic the levels of PD-1 -binding activity of wildtype PD-L1 while exhibiting an extended half-life in vivo.
SUMMARY OF THE INVENTION
[0005] The present disclosure provides an engineered programmed cell death 1 ligand (PD-L1) protein comprising an extracellular sequence of a PD-L1 and a transmembrane domain (TM) and/or an intracellular domain (ICD) that is heterologous to PD-L1, wherein the engineered PD-L1 protein has an increased half-life than the corresponding wildtype PD- L1 when expressed in a mammalian cell.
[0006] In some embodiments, the PD-L1 is human PD-L1, optionally wherein the extracellular sequence comprises SEQ ID NO:24 or an amino acid sequence at least 95% identical thereto. In some embodiments, the extracellular sequence comprises amino acids 1- 219, 1-210, or 1-200 of SEQ ID NO:24, or a variant amino acid sequence at least 95% identical to amino acids 1-219, 1-210, or 1-200 of SEQ ID NO:24. In further embodiments, the amino acid sequence at least 95% identical to SEQ ID NO:24 comprises one or more mutations relative to SEQ ID NO: 1 selected from N-to-Q mutations, I54E/Q, Y56H/F, E58F/M, R113T, M115L, S117G/A, G119K, G120V, A121W, R238A, an insertion ofW between wildtype residues A121 and D122, and a deletion between amino acids 230-238, optionally wherein the variant amino acid sequence comprises (i) an extracellular sequence of SEQ ID NO:54, 55, 56, 57, 67, 68, or 79, or (ii) SEQ ID NO:73, 74, 75, 76, 77, 78, or 80. [0007] In some embodiments, the TM domain comprises a wildtype PD-L1 TM sequence, optionally SEQ ID NO:25. In further embodiments, the TM domain comprises a TM sequence from integrin alpha-M (ITGAM), optionally SEQ ID NO:31, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO: 18, with or without the signal sequence shown in SEQ ID NO:23. In certain embodiments, the TM domain comprises a TM sequence from carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM1), optionally SEQ ID NO:32, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO: 19, with or without the signal sequence shown in SEQ ID NO:23. In certain embodiments, the TM domain comprises a TM sequence from integrin alpha-1 (ITGA1), optionally SEQ ID NO:33, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:20, with or without the signal sequence shown in SEQ ID NO:23.
[0008] In some embodiments, the ICD is a PD-L1 -derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:26, 34, 35, or 36. In some embodiments, the engineered PD-L1 protein comprises SEQ ID NO:2, 3, or 4, with or without the signal sequence shown in SEQ ID NO: 23.
[0009] In some embodiments, the ICD comprises a heterologous polypeptide sequence fused, optionally through a peptide linker, to the C-terminus of a PD-L1 -derived ICD sequence. In certain embodiments, the heterologous polypeptide is a CMTM6 sequence, optionally SEQ ID NO:47, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:6, with or without the signal sequence shown in SEQ ID NO:23. In certain embodiments, the heterologous polypeptide is a CSN5 sequence, optionally SEQ ID NO:48, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:7, with or without the signal sequence shown in SEQ ID NO:23.
[0010] In some embodiments, the ICD comprises an ITGAM-derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:37 or 38. In some embodiments, the engineered PD-L1 protein comprises SEQ ID NO:8 or 9, with or without the signal sequence shown in SEQ ID NO:23.
[0011] In some embodiments, the ICD comprises a CEACAM1 -derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:39 or 40. In some embodiments, the engineered PD-L1 protein comprises SEQ ID NO: 10, 11, 67, or 68, with or without the signal sequence shown in SEQ ID NO: 23.
[0012] In some embodiments, the ICD comprises an ITGA1 -derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:41 or 42. In some embodiments, the engineered PD-L1 protein comprises SEQ ID NO: 12 or 13, with or without the signal sequence shown in SEQ ID NO: 23.
[0013] In some embodiments, the ICD comprises an ENPP1 -derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:43 or 44. In some embodiments, the engineered PD-L1 protein comprises SEQ ID NO: 14 or 15, with or without the signal sequence shown in SEQ ID NO: 23.
[0014] In some embodiments, the ICD comprises an APMAP-derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:45 or 46. In some embodiments, the engineered PD-L1 protein comprises SEQ ID NO: 16 or 17, with or without the signal sequence shown in SEQ ID NO: 23.
[0015] In some embodiments, the extracellular sequence of a PD-L1 is fused to a GPI anchor, optionally comprising SEQ ID NO:50, further optionally wherein the engineered PD- L1 protein comprises SEQ ID NO:21.
[0016] In some embodiments, the extracellular sequence of a PD-L1 is fused, with or without a peptide linker, to a GPI-anchored protein, optionally NT5E. In further embodiments, the GPI-anchored protein comprises SEQ ID NO:51, optionally wherein the
engineered PD-L1 protein comprises SEQ ID NO:22. In some embodiments, the engineered PD-L1 protein comprises SEQ ID NO:5, with or without the signal sequence shown in SEQ ID NO:23, or comprising SEQ ID NO:73, 74, 76, 77, or 80.
[0017] In some embodiments, the engineered PD-L1 protein herein further comprises a signal sequence, wherein the signal sequence comprises a PD-Ll signal sequence, optionally SEQ ID NO:23; or a NT5E signal sequence, optionally SEQ ID NO:29.
[0018] In another aspect, the present disclosure provides a nucleic acid molecule comprising a coding sequence for the engineered PD-L1 protein herein. In some embodiments, the nucleic acid molecule is DNA, mRNA, or circular RNA. Also provided herein is a recombinant virus comprising the nucleic acid molecule herein.
[0019] In another aspect, the present disclosure provides a lipid nanoparticle comprising an mRNA or circular RNA herein.
[0020] In another aspect, the present disclosure provides a pharmaceutical composition comprising the nucleic acid molecule, recombinant virus, or lipid nanoparticle herein, and a pharmaceutically acceptable carrier.
[0021] In another aspect, the present disclosure provides a method of treating an autoimmune condition of a tissue in a patient in need thereof, comprising administering the pharmaceutical composition herein to the patient such that the engineered PD-L1 protein is expressed in the tissue. Also provided is use of the nucleic acid molecule, recombinant virus, or lipid nanoparticle herein for the manufacture of a medicament for treating an autoimmune condition in a patient in need thereof. Also provided is the nucleic acid molecule, recombinant virus, lipid nanoparticle, or pharmaceutical composition herein for use in treating an autoimmune condition in a patient in need thereof. In some embodiments, the tissue is the liver and the autoimmune condition is autoimmune hepatitis.
[0022] In another aspect, the present disclosure provides a method of preventing rejection of an organ transplant in a patient in need thereof, comprising administering the pharmaceutical composition herein to the transplant such that the engineered PD-L1 protein is expressed in the transplant. Also provided is use of the nucleic acid molecule, recombinant virus, or lipid nanoparticle herein for the manufacture of a medicament for preventing rejection of an organ transplant in a patient in need thereof. Also provided is the nucleic acid molecule, recombinant virus, lipid nanoparticle, or pharmaceutical composition herein for use in preventing rejection of an organ transplant in a patient in need thereof.
[0023] Other features, objectives, and advantages of the invention are apparent in the detailed description that follows. It should be understood, however, that the detailed
description, while indicating embodiments and aspects of the invention, is given by way of illustration only, not limitation. Various changes and modifications within the scope of the invention will become apparent to those skilled in the art from the detailed description.
BRIEF DESCRIPTION OF THE FIGURES
[0024] FIG. 1 is a schematic showing the function of the PD-1 ligand, PD-L1, in the inhibition of aberrant immune responses and prevention of apoptosis of hepatocytes expressing PD-L1 at their cell surface.
[0025] FIG. 2 is a series of flow cytometry plots demonstrating the effectiveness of detecting PD-L1 expression on CHO-K1 cells using an anti-PD-Ll antibody (top) or Alexa Fluor 647-conjugated human PD-1 extracellular domain (ECD) (bottom).
[0026] FIGs. 3A-B are bar graphs showing the expression levels of various engineered PD-L1 fusion proteins expressed from DNA plasmids in CHO-K1 cells, as compared to wildtype (WT) PD-L1 (FIG. 3A), and the relative binding ability of various engineered PD- L1 fusion proteins to PD-1 compared to wildtype PD-L1 (FIG. 3B).
[0027] FIG. 4 is a set of heat maps showing the change over a 96-hour period in the percentage of PD-L1 -positive, viable CHO-K1 cells transfected with plasmid DNA encoding a PD-L1 chimera (left) and the geometric mean expression of the PD-L1 chimeras in positive cells (right).
[0028] FIG. 5 is a set of heat maps showing the change over a nine-day period in the percentage of PD-L1 -positive, viable OVCAR-3 cells transfected with plasmid DNA encoding a PD-L1 chimera (left) and the geometric mean expression of the PD-L1 chimeras in positive cells (right).
[0029] FIG. 6 is a set of heat maps showing the change over a seven-day period in the percentage of PD-L1 positive, viable CHO-K1 cells transfected with circular RNA (circRNA) encoding a PD-L1 chimera (left), the geometric mean expression of the PD-L1 chimeras in positive cells (center), and the net mean fluorescence intensity (MFI) of the PD-L1 chimeras on Day 7 (right).
[0030] FIG. 7 is a set of heat maps showing the change over a seven-day period in the percentage of PD-L1 positive, viable HEK293T cells transfected with circRNA encoding a PD-L1 chimera (left), the geometric mean expression of the PD-L1 chimeras in positive cells (center), and the net MFI of the PD-L1 chimera on Day 7 (right).
[0031] FIG. 8 is a set of graphs showing expression of PD-L1 after six hours as determined by flow cytometry (left two graphs), the change over 48 hours in the percentage
of PD-L1 -positive cells (center two graphs), and the MFI of PD-L1+ cells (right two graphs) in either hepatocytes (top row of graphs) or immune cells (bottom row of graphs). The cells were obtained from the liver of mice treated with mRNA encoding PD-L1. The mRNA was delivered using an ionizable lipid formulation, LRN1.
[0032] FIG. 9 is a line graph showing the amounts of PD-L1 detected in the liver of mice by an ELISA assay. The cells were obtained from the liver of mice treated with mRNA encoding PD-L1. The mRNA was delivered using an ionizable lipid formulation, LRN1. [0033] FIG. 10 is a set of line graphs showing the change over 120 hours in the percentage of PD-L1 -positive cells (left) and the MFI of PD-L1+ cells (right). The cells were obtained from the liver of mice pre-dosed with anti-IFNaR-1 monoclonal antibody and administered circular RNA encoding PD-L1. The circular RNA was delivered with an ionizable lipid formulation, LRN1.
[0034] FIG. 11 is a line graph showing the change over seven days in the percentage of PD-L1 -positive cells. The cells were obtained from the liver of mice pre-dosed with anti- IFNaR-1 monoclonal antibody and administered circRNA encoding PD-L1 variants. The circular RNA was delivered with an ionizable lipid formulation (LP-01).
[0035] FIG. 12 is a line graph showing the MFI and derived EC50 values of CHO cells transfected with mRNAs encoding PD-L1 molecules bearing various point mutations, as measured by flow cytometry.
[0036] FIG. 13 is a plot showing the PD-L1 cell surface expression (as indicated by MFI) and PD-L1 soluble levels in the cell culture supernatant of CHO cells transfected with mRNAs encoding PD-L1 molecules bearing various point mutations.
DETAILED DESCRIPTION OF THE INVENTION
[0037] The present disclosure provides engineered PD-L1 proteins that have improved longevity while retaining wildtype PD-Ll’s expression levels and affinity for PD-1. By supplying the present engineered PD-L1 protein to cells or tissues that are susceptible for autoimmune reactions, it is now possible to locally suppress autoimmunity without systemic immunosuppression.
[0038] Patients with certain autoimmune diseases (e.g., autoimmune hepatitis) have been observed to express PD-1 and PD-L1 at low levels, which may contribute to T cell-mediated destruction of healthy tissues (see, e.g., Agina et al., Clin Exp Hepatol. (2019) 5(3):256-64; Jilkova et al., Cells. (2021) 10(10):2671). Expressing the present PD-L1 proteins in specific tissues can repress aberrant immune responses against those tissues, thereby preserving
viability of healthy cells. For example, the present PD-L1 proteins may be expressed in liver parenchymal and residential cells to alleviate autoimmune hepatitis or other immune-driven liver diseases (e.g., primary biliary cholangitis). Additionally, the present PD-L1 proteins may also be expressed, for example, in kidney cells to alleviate lupus nephritis; in the kidney, skin, or central nervous system to alleviate systemic lupus erythematosus (SLE); in the synovium to alleviate rheumatoid arthritis; in pancreatic cells to alleviate type 1 diabetes; in cells of the salivary or lacrimal glands to alleviate Sjogren’s syndrome; in the thyroid to alleviate Graves’ disease; in oligodendrocytes to alleviate multiple sclerosis; in the central or peripheral nervous system to alleviate myasthenia gravis or neuromyelitis optica; in the intestines to alleviate Crohn’s disease, ulcerative colitis or celiac disease; and in skin cells to alleviate psoriasis. The present PD-L1 proteins may also be expressed in organ transplants to prevent (including reducing) graft rejection in patients receiving the transplants.
[0039] Due to the design of their sequences, the present PD-L1 -derived proteins can be expressed at levels approximating wildtype PD-L1 on the cell surface in several cell types and effectively bind to PD-1, indicating their potential use as a therapeutic. Additionally, these engineered proteins exhibit longer half-lives than wildtype PD-L1, suggesting their potential enhanced therapeutic efficacy compared to supplementation with wildtype PD-L1.
I. Engineered PD-L1 Proteins
[0040] The engineered PD-L1 proteins herein are derived from PD-L1 but may contain only a partial, rather than the entire, sequence of wildtype PD-L1. These polypeptides are cell surface proteins when expressed in mammalian cells. As used herein, “derived from” means that the sequence is the same as or similar to the original sequence; a derived sequence may be longer or shorter than, or have the same length as, the original sequence.
[0041] Unless otherwise indicated, PD-L1 as used herein refers to human PD-L1. A human PD-L1 polypeptide sequence may be found at the UniProt database (Identifier No.
Q9NZQ7-1) and may have the following sequence:
1 MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL
51 AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ
101 ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE
151 HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN
201 TTTNEI FYCT FRRLDPEENH TAELVIPEL|P LAHPPNER|TH LVILGAILLC
251 LGVALTFIFR LRKGRMMDVK KCGIQDTNSK KQSDTHLEET ( SEQ ID NO : 1 )
[0042] In the above sequence, the various PD-L1 domains are delineated as follows. The extracellular region spans amino acids 1-238. The extracellular region includes a signal
sequence (amino acids 1-18; underlined) and two distinct extracellular domains (ECDs). The first ECD is an IgV-like domain that spans amino acids 19-127 and the second ECD is an IgC-like domain that spans amino acids 133-225. The transmembrane domain (TM) spans amino acids 239-259 (boldface and underlined). The intracellular domain (ICD) spans amino acids 260-290. The signal sequence is removed in the mature polypeptide. The signal sequence, the extracellular region minus the signal sequence, the TM domain, and the ICD domain of the above sequence are assigned SEQ ID NOs:23-26, respectively. The IgV-like domain and IgC-like domain sequences are assigned SEQ ID NOs:27 and 28, respectively. PD-L1 is predicted to possess a matrix metalloproteinase cleavage site that spans amino acids 230-238 (boxed; SEQ ID NO:71).
[0043] Unless otherwise noted, a PD-L1 amino acid position recited herein refers to the position in SEQ ID NO: 1 or a corresponding position in a variant of SEQ ID NO: 1 (e.g., a naturally occurring polymorphic variant or a genetically engineered variant).
[0044] To the extent that some of the present engineered PD-L1 proteins are fusion proteins with components from different sources, those proteins also are called “PD-L1 chimeras” or “PD-L1 fusion proteins” herein.
[0045] The polypeptides’ extracellular, transmembrane, and intracellular regions of the engineered PD-L1 proteins herein are described below.
A. Extracellular Region
[0046] The extracellular region of the present engineered PD-L1 polypeptides may comprise a signal sequence that triggers the translocation of the newly synthesized PD-L1 protein into the endoplasmic reticulum for trafficking and expression to the cell surface. The signal sequence may be cleaved off before the protein reaches the cell membrane. In some embodiments, the signal sequence comprises PD-Ll’s wildtype signal sequence MRIFAVFIFMTYWHLLNA (SEQ ID NO:23) or a functional variant thereof. In some embodiments, the signal sequence may be derived from another cell surface protein or an artificial sequence. For example, the signal sequence may comprise a signal sequence from human 5 ’ -nucleotidase NT5E (a.k.a. CD73), MCPRAARAPATLLLALGAVLWPAAGA (SEQ ID NO:29), or a functional variant thereof.
[0047] The extracellular region of the present engineered PD-L1 polypeptides comprises a PD-L1 extracellular region for their binding to the wildtype ligand PD-1. In some embodiments, the extracellular region comprises both the IgV-like domain (e.g., SEQ ID NO:27) and the IgC-like domain (e.g., SEQ ID NO:28) of PD-L1. The two domains may be linked by the wildtype peptide sequence (amino acids 128-132 of SEQ ID NO:1) or by
another peptide linker, for example a glycine/serine-rich sequence (e.g., a linker containing a G4S motif (SEQ ID NO: 52)). The stem region of the extracellular region may be the wildtype stem region of PD-L1 (amino acids 226-238 of SEQ ID NO: 1) or a functional analog thereof. In some embodiments, the extracellular region of the present polypeptides comprises SEQ ID NO:24.
[0048] In some embodiments, the extracellular region comprises mutations in the IgV- like domain and/or the IgC-like domain sequence relative to wildtype PD-L1. In further embodiments, the mutations are made in the IgC-like domain sequence. By way of example, the IgC-like domain comprises an N-to-Q substitution at one, two, or all three positions corresponding to residue 192, 200, and 219 of SEQ ID NO: 1. In some embodiments, the extracellular region comprises PD-Ll’s extracellular region with N192Q, N200Q, and N219Q substitutions. In further embodiments, the extracellular region comprises SEQ ID NO:30.
[0049] In some embodiments, the extracellular region of the present engineered PD-L1 polypeptide comprises a part (e.g., amino acid 1 to amino acid 210, 211, 212, 213, 214, 215, 216, 217, or 218) or the entirety (i.e., amino acids 1 to 219) of SEQ ID NO:24. In some embodiments, the extracellular region comprises an additional phenylalanine (F) at the N- terminus.
[0050] In some embodiments, the extracellular region is a variant of a wildtype extracellular region of PD-L1; for example, the variant is at least 95% (e.g., at least 96, 97, 98, or 99%) identical to the wildtype extracellular region. In some embodiments, the variant has improved biological activities such as increased affinity for PD-1 and/or increased resistance to protease cleavage.
[0051] In some embodiments, the engineered PD-L1 polypeptide herein has increased affinity for PD-1. The extracellular region of such a polypeptide may comprise one or more mutations selected from I54E/Q, Y56H/F, E58F/M, R113T, Ml 15L, SI 17G/A, G119K, G120V, A121W, and an insertion of W between wildtype residues A121 and D122, relative to SEQ ID NO: 1. In further embodiments, the extracellular region of the polypeptide an extracellular sequence of SEQ ID NO:54, 55, 56, 57, or 79. For example, the extracellular region may comprise amino acid 19 or 20 to amino acid 238 of SEQ ID NO:54, 55, 57, or 79, or amino acid 19 or 20 to amino acid 239 of SED ID NO:56. In some embodiments, the extracellular region comprises SEQ ID NO:75, 76, 77, 78, or 80.
[0052] In some embodiments, the engineered PD-L1 polypeptide herein has increased resistance to protease cleavage at its hinge region such that it has an increased ability to
remain membrane-bound. In some embodiments, the metalloprotease recognition site (e.g., amino acids 230-238 in SEQ ID NO: 1) is mutated to achieve this purpose. In some embodiments, the engineered polypeptide may comprise a mutation at R238 (e.g., R238A) relative to SEQ ID NO: 1; an exemplary polypeptide may comprise an extracellular region comprising SEQ ID NO:73. In other embodiments, the metalloprotease recognition site may be replaced by protease-resistant sequences; for example, amino acids 230-238 may be replaced by a G/S-rich peptide linker (i.e., 50% or more of the amino acids in the linker are glycine and/or serine). In further embodiments, amino acids 230-238 may be replaced by (G3S)3 (SEQ ID NO:72); an exemplary polypeptide may comprise an extracellular region comprising SEQ ID NO:74.
[0053] In some embodiments, the engineered PD-L1 polypeptide comprises both mutation(s) that increase PD-1 -binding affinity and mutation(s) that increase resistance to protease cleavage. For example, the engineered PD-L1 polypeptide comprises two or more mutations selected from I54E/Q, Y56H/F, E58F/M, R113T, Ml 15L, SI 17G/A, G119K, G120V, A121W, R238A, and an insertion of W between wildtype residues A121 and D122, relative to SEQ ID NO: 1. In some embodiments, the engineered PD-L1 polypeptide comprise mutations at all of the aforementioned positions with the indicated substitutions and insertion.
B. Transmembrane Region
[0054] The transmembrane region of the present polypeptides contains a hydrophobic region sufficient to anchor the PD-Ll-derived polypeptide within the cell membrane. This region may comprise an artificial sequence or may be derived from any transmembrane (TM) protein with a long protein half-life (i.e., longer than 15 hours). In some embodiments, the TM region comprises the TM region of PD-L1, e.g., SEQ ID NO:25, or a functional equivalent thereof. In some embodiments, the TM region comprises a TM region from other TM proteins. For example, the TM region herein may comprise a TM sequence from integrin-alpha M (ITGAM), e.g., SEQ ID NO:31; a TM sequence from carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM1), e.g., SEQ ID NO:32; or a TM sequence from integrin-alpha 1 (ITGA1), e.g., SEQ ID NO:33; or a functional variant thereof. [0055] In additional embodiments, the TM domain may be derived from other cell surface proteins known to express on the target tissue with long half-lives. Example of the additional cell surface proteins are solute carrier family 2 member 2 (SLC2A2), 5’- nucleotidase ecto (NT5E), CD47, ATP binding cassette subfamily C member (6ABCC6), syntaxin 4 (STX4), FAS, neuroplastin (NPTN), ectonucleotide
pyrophosphatase/phosphodiesterase 1 (ENPP1), and adipocyte plasma membrane associated protein (APMAP).
C. Intracellular Region
[0056] The ICD of the engineered PD-L1 protein may be derived from PD-L1 or another cell surface protein, or may be completely artificial (i.e., not derived from a naturally occurring protein). Where the ICD is derived from a naturally occurring protein, it may contain the wildtype ICD or a variant thereof. The variant may, for example, contain mutations (e.g., substitution) that reduce or eradicate the signal transduction function of the ICD. The variant may also be a wildtype ICD placed in a reverse orientation that maintains the original topology and sequence orientation of the partner protein; that is, the C-terminus of the wildtype ICD becomes the N-terminus of the ICD in the engineered protein and the N- terminus of the wildtype ICD becomes the C-terminus of the ICD in the engineered protein. [0057] In some embodiments, the ICD of the engineered PD-L1 protein comprises a wildtype PD-L1 ICD (e.g., SEQ ID NO:26) or a variant thereof. In some embodiments, the variant comprises one or more substitutions relative to the wildtype ICD sequence. For example, lysine may be substituted by arginine to prevent ubiquitination and stabilize the protein. In some embodiments, the variant may be the “PD-L1 ICD K5R” variant (SEQ ID NO:34), which is identical to the wildtype ICD sequence except that all five lysine residues in the wildtype sequence have been substituted by arginines. In some embodiments, the variant may be the “PD-L1 ICD K263R” variant (SEQ ID NO:35), which is identical to the wildtype ICD sequence except that the first lysine residue in the wildtype sequence (corresponding to K263 in SEQ ID NO:1) has been substituted by arginine. In some embodiments, amino acid residues other than lysines (e.g., 2, 3, 4, or more additional amino acid residues) are substituted in the same or similar manner. In some embodiments, the variant may be the “PD-L1 ICD KtoR,DEtoR” variant (SEQ ID NO:36), which is identical to the wildtype ICD sequence except that all lysines (5), aspartic acids (3), and glutamic acid (2) residues in the wildtype sequence have been substituted by arginines; these mutations increase the net positive charge of the ICD and help to increase interaction with the membrane and resist degradation.
[0058] The ICD of the engineered PD-L1 protein may comprise an ICD derived from another protein with a long protein half-life (i.e., longer than 15 hours). For example, the ICD comprises an ICD from ITGAM (e.g., SEQ ID NO:37) or a variant thereof. In some embodiments, the variant comprises one or more substitutions (e.g., K to R) relative to the wildtype ICD sequence. In some embodiments, the variant may be the “ITGAM ICD K3R”
variant (SEQ ID NO:38), which is identical to the wildtype ICD sequence except that all three lysine residues in the wildtype sequence have been substituted by arginines.
[0059] In some embodiments, the ICD of the engineered PD-L1 protein comprises an ICD from CEACAM1 (e.g., SEQ ID NO:39) or a variant thereof. In some embodiments, the variant comprises one or more substitutions (e.g., K to R) relative to the wildtype ICD sequence. In some embodiments, the variant may be the “CEACAM1 ICD K5R” variant (SEQ ID NO:40), which is identical to the wildtype ICD sequence except that all five lysine residues in the wildtype sequence have been substituted by arginines.
[0060] In some embodiments, the ICD of the engineered PD-L1 protein comprises an ICD from ITGA1 (e.g., SEQ ID NO:41) or a variant thereof. In some embodiments, the variant comprises one or more substitutions (e.g., K to R) relative to the wildtype ICD sequence. In some embodiments, the variant may be the “ITGA1 ICD K6R” variant (SEQ ID NO:42), which is identical to the wildtype ICD sequence except that all six lysine residues in the wildtype sequence have been substituted by arginines.
[0061] In some embodiments, the ICD of the engineered PD-L1 protein comprises an ICD from ectonucleotide pyrophosphatase/phosphodiesterase family member 1 (ENPP1) or a variant thereof. The ICD from ENPP1 may be in a reverse orientation in the engineered protein. In some embodiments, the ICD of the engineered protein comprises SEQ ID NO:43 (reverse of a wildtype human ENPP1 ICD), or a variant thereof. In some embodiments, the variant comprises one or more substitutions (e.g., K to R) relative to the wildtype ICD sequence. For example, the variant may be the “ENPP1 ICD (reverse) KtoR” variant (SEQ ID NO:44), which is identical to SEQ ID NO:43 except that all three lysine residues in it have been substituted by arginines.
[0062] In some embodiments, the ICD of the engineered PD-L1 protein comprises an ICD from adipocyte plasma membrane-associated protein (APMAP) or a variant thereof. The ICD from APMAP may be in a reverse orientation in the engineered protein. In some embodiments, the ICD of the engineered protein comprises SEQ ID NO:45 (reverse of a wildtype human APMAP ICD), or a variant thereof. In some embodiments, the variant comprises one or more substitutions (e.g., K to R) relative to the wildtype ICD sequence. For example, the variant may be the “APMAP ICD (reverse) KtoR” variant (SEQ ID NO:46), which is identical to SEQ ID NO:45 except that the lysine residue in it has been substituted by arginine.
[0063] In certain cases, the ICD of the engineered PD-L1 protein may comprise an ICD sequence of another cell surface protein with a long protein half-life (i.e., longer than 15
hours). Examples of such additional cell surface proteins are SLC2A2, NT5E, CD47, ABCC6, STX4, FAS, and NPTN.
[0064] In certain cases, the ICD of the engineered PD-L1 protein may comprise the polypeptide sequence of a second cell surface proteins with a long protein half-life (i.e., longer than 15 hours) or of another protein selected based on useful functions (e.g., proteins know to interact with the PD-L1 protein, stabilize the PD-L1 protein, or both). For example, this polypeptide sequence may be fused to the ICD described above, with or without a peptide linker. For example, the ICD of the engineered PD-L1 protein may comprise a wildtype PD-L1 ICD fused through a peptide linker to CKLF-like MARVEL transmembrane domain-containing protein 6 (CMTM6) (e.g., SEQ ID NO:47). In another example, the ICD of the engineered PD-L1 protein may comprise a wildtype PD-L1 ICD fused through a peptide linker to COP9 signalosome complex subunit 5 (CSN5) (e.g., SEQ ID NO:48). The peptide linker may be a flexible linker such as a GS linker. A GS linker is rich in glycine and serine (more than 50% of the residues are glycine and/or serine). In some embodiments, the peptide linker comprises GGGGSGGGSGGGS (SEQ ID NO:49).
D. Glycophosphatidylinositol Anchorage
[0065] In some embodiments, the engineered PD-L1 protein is anchored in the cell membrane through a glycophosphatidylinositol (GPI) anchor. For example, the extracellular region of the protein is fused to a GPI anchor sequence, e.g., SEQ ID NO:50.
[0066] In some embodiments, the extracellular region of the engineered PD-L1 protein is fused to a polypeptide that can be anchored by GPI. An example of such a protein is NT5E. A human NT5E sequence (without the signal sequence) is SEQ ID NO:51. The GPL anchorable polypeptide sequence may be used to the extracellular region through a peptide linker such as a GS linker, for example, GGGGS (SEQ ID NO:52).
E. Examples of Engineered PD-L1 Proteins
[0067] In some embodiments, the present polypeptide comprises, consists of, or consists essentially of a PD-L1 protein sequence with one or more (e.g., one, two, three, four, five, six, seven, eight, nine, ten or more) mutations, such as substitutions. In one embodiment, the engineered PD-L1 sequence is termed “PDL1-K5R,” with five K-to-R mutations (underlined) in the ICD relative to wildtype PD-L1 :
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC
LGVALTFI FR LRRGRMMDVR RCGIQDTNSR RQSDTHLEET
( SEQ ID NO : 2 )
[0068] In another embodiment, the engineered PD-L1 sequence is termed “PDL1-
K263R” and contains one K-to-R mutation (underlined) in the ICD relative to wildtype PD-
MR I FAVE I FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC LGVALTFI FR LRRGRMMDVK KCGIQDTNSK KQSDTHLEET ( SEQ ID NO : 3 )
[0069] In another embodiments, the engineered PD-L1 sequence is termed “PDL1- KtoR,DEtoR” and contains ten amino acid substitutions relative to wildtype PD-L1, where all lysines, aspartic acids, and glutamic acids in the wildtype PD-L1 ICD have been substituted with arginines (underlined):
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC LGVALTFI FR LRRGRMMRVR RCGIQRTNSR RQSRTHLRRT ( SEQ ID NO 4 )
[0070] In one embodiment, the engineered PD-L1 sequence is termed “PDL1 (3NQ)” and contains three N-to-Q mutations (underlined) in the IgC-like domain relative to wildtype PD-
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FQVTSTLRIQ TTTNEI FYCT FRRLDPEEQH TAELVIPELP LAHPPNERTH LVILGAILLC LGVALTFI FR LRKGRMMDVK KCGIQDTNSK KQSDTHLEET ( SEQ ID NO : 5 )
[0071] In one embodiment, the engineered PD-L1 sequence is termed “PDL1-CMTM6 fusion with linker,” in which a wildtype PD-L1 sequence is fused at its C-terminus to a CMTM6 polypeptide sequence (boldfaced; SEQ ID NO:47) through a peptide linker of SEQ ID NO:49 (underlined):
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC
LGVALTFI FR LRKGRMMDVK KCGIQDTNSK KQSDTHLEET GGGGSGGGSG
GGSMENGAVY SPTTEEDPGP ARGPRSGLAA YFFMGRLPLL RRVLKGLQLL LSLLAFICEE WSQCTLCGG LYFFEFVSCS AFLLSLLILI VYCTPFYERV DTTKVKSSDF YITLGTGCVF LLASIIFVST HDRTSAEIAA IVFGFIASFM FLLDFITMLY EKRQESQLRK PENTTRAEAL TEPLNA ( SEQ ID NO : 6 )
[0072] In one embodiment, the engineered PD-L1 sequence is termed “PDL1-CSN5 fusion with linker,” in which a wildtype PD-L1 sequence is fused at this C-terminus to a
CSN5 polypeptide sequence (boldfaced; SEQ ID N0:51) through a peptide linker of SEQ ID
NO:49 (underlined):
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC LGVALTFI FR LRKGRMMDVK KCGIQDTNSK KQSDTHLEET GGGGSGGGSG
GGSMAASGSG MAQKTWELAN NMQEAQSIDE IYKYDKKQQQ EILAAKPWTK DHHYFKYCKI SALALLKMVM HARSGGNLEV MGLMLGKVDG ETMIIMDSFA LPVEGTETRV NAQAAAYEYM AAYIENAKQV GRLENAIGWY HSHPGYGCWL SGIDVSTQML NQQFQEPFVA WIDPTRTIS AGKVNLGAFR TYPKGYKPPD EGPSEYQTIP LNKIEDFGVH CKQYYALEVS YFKSSLDRKL LELLWNKYWV NTLSSSSLLT NADYTTGQVF DLSEKLEQSE AQLGRGSFML GLETHDRKSE DKLAKATRDS CKTTIEAIHG LMSQVIKDKL FNQINIS ( SE( ! ID NO : 7 )
[0073] In one embodiment, the engineered PD-L1 sequence is termed “PDL1-ITGAM
(ICD),” in which the ICD of a wildtype PD-L1 sequence is replaced with an ICD from
ITGAM (italic; SEQ ID NO:37):
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC LGVALTFI FK LGFFKRQYKD MMSEGGPPGA EPQ ( SEQ ID NO : 8 )
In a related embodiment, the ITGAM ICD sequence is mutated, where all three lysine residues are substituted by arginine residues (underlined), and the engineered PD-L1 sequence is called “PDL1 -ITGAM K3R (ICD)”:
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN
TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC LGVALTFI FR LGFFRRQYRD MMSEGGPPGA EPQ ( SEQ i: | NO : 9 )
[0074] In one embodiment, the engineered PD-L1 sequence is termed “PDL1-
CEACAM1 (ICD),” in which the ICD of a wildtype PD-L1 sequence is replaced with an ICD from CEACAM1 (italic; SEQ ID NO:39):
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC
LGVALTFI FH FGKTGRASDQ RDLTEHKPSV SNHTQDHSND PPNRMNEVTY STLNFEAQQP TQPTSASPSL TATEIIYSEV KKQ ( SEQ I] | NO : 10 )
In a related embodiment, the CEACAM1 ICD sequence is mutated, where all five lysine residues are substituted by arginine residues (underlined), and the engineered PD-L1 sequence is called “PDL1-CEACAM1 K5R (ICD)”:
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC
LGVALTFI FH FGRTGRASDQ RDLTEHRPSV SNHTQDHSND PPNRMNEVTY STLNFEAQQP TQPTSASPSL TATETTYSEV RRQ ( SEQ I] | NO : 11 )
In another related embodiment, PDL1-CEACAM1 K5R (ICD) is further mutated in the region corresponding to amino acids 230-238 of SEQ ID NO: 1 to make it more resistant to protease cleavage. For example, the PD-L1 extracellular region comprises an R238A mutation (boxed), and the engineered PD-L1 sequence is called “PDL1-CEACAM1 K5R
(ICD) R238A”:
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVIPELP LAHPPNERTH LVILGAILLC
LGVALTFI FH FGRTGRASDQ RDLTEHRPSV SNHTQDHSND PPNRMNEVTY STLNFEAQQP TQPTSASPSL TATEIIYSEV RRQ ( SEQ I] | NO : 67 )
In another example, the region corresponding to amino acids 230-238 of SEQ ID NO: 1 in PDL1-CEACAM1 K5R (ICD) is replaced by (G3S)3 (SEQ ID NO:72; boxed), and the engineered PD-L1 sequence is called “PD-L1-CEACAM1 K5R (ICD) (G3S)3”:
MRIFAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMTIEC KFPVEKQLDL AALIVYWEME DKNIIQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVl|GGGS GGGSGGGS|TH LVILGAILLC LGVALTFI FH FGRTGRASDQ RDLTEHRPSV SNHTQDHSND PPNRMNEVTY STLNFEAQQP TQPTSASPSL TATEIIYSEV RRQ ( SEQ ID NO : 68 )
[0075] In one embodiment, the engineered PD-L1 sequence is termed “PDL1-ITGA1 (ICD),” in which the ICD of a wildtype PD-L1 sequence is replaced with an ICD from ITGA1 (italic; SEQ ID NO:41):
MRI FAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMT IEC KFPVEKQLDL
AALIVYWEME DKNI IQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ
ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE
HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVI PELP LAHPPNERTH LVILGAILLC LGVALT FI FK IGFFKRPLKK KMEK ( SEQ D NO : 12 )
In a related embodiment, the ITGA1 ICD sequence is mutated, where all six lysine residues are substituted by arginine residues (underlined), and the engineered PD-L1 sequence is called “PDL1-ITGA1 K6R (ICD)”:
MRI FAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMT IEC KFPVEKQLDL AALIVYWEME DKNI IQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN
TTTNEI FYCT FRRLDPEENH TAELVI PELP LAHPPNERTH LVILGAILLC LGVALT FI FR IGFFRRPLRR RMER ( SEQ D NO : 13 )
[0076] In one embodiment, the engineered PD-L1 sequence is termed “PDL1-ENPP1
(ICD reverse),” in which the ICD of a wildtype PD-L1 is replaced with an ICD from ENPP1 in a reverse orientation (italic; SEQ ID NO:43):
MRI FAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMT IEC KFPVEKQLDL AALIVYWEME DKNI IQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVI PELP LAHPPNERTH LVILGAILLC
LGVALT FI FK YTNPDKATRA RAAKELPEEG VDMPALLSAA AQPDGPAEAA HSRGRDRGNG APGERPARGG EGGRSGGGAC GDREM ( SEQ ID NO : 14 )
In a related embodiment, the reversed ENPP1 ICD sequence is mutated, where all three lysine residues are substituted by arginine residues (underlined), and the engineered PD-L1 sequence is called “PDLl-ENPPl-KtoR (ICD reverse)”:
MRI FAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMT IEC KFPVEKQLDL AALIVYWEME DKNI IQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVI PELP LAHPPNERTH LVILGAILLC
LGVALT FI FR YTNPDRATRA RAARELPEEG VDMPALLSAA AQPDGPAEAA HSRGRDRGNG APGERPARGG EGGRSGGGAC GDREM ( SEQ ID NO : 15 )
[0077] In one embodiment, the engineered PD-L1 sequence is termed “PDL1-APMAP
(ICD reverse),” in which the ICD of a wildtype PD-L1 is replaced with an ICD from APMAP in a reverse orientation (italic; SEQ ID NO:45):
MRI FAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMT IEC KFPVEKQLDL
AALIVYWEME DKNI IQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ
ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE
HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN
TTTNEI FYCT FRRLDPEENH TAELVI PELP LAHPPNERTH LVILGAILLC
LGVALT FI FR FVRGSFSSGD KAEPAQGDDD TWQPRLPRR QRLGDAES ( SEQ ID NO : 16 )
In a related embodiment, the reversed APMAP ICD sequence is mutated, where the lysine residue is substituted by arginine (underlined), and the engineered PD-L1 sequence is called “PDL1 -APMAP -KtoR (ICD reverse)”:
MRI FAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMT IEC KFPVEKQLDL AALIVYWEME DKNI IQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVI PELP LAHPPNERTH LVILGAILLC LGVALT FI FR FVRGSFSSGD RAEPAQGDDD TWQPRLPRR QRLGDAES
NO : 17 )
[0078] In one embodiment, the engineered PD-L1 sequence is called “PDL1-ITGAM
(TM),” in which the TM domain of a wildtype PD-L1 is replaced with a TM domain from
ITGAM (boldface; SEQ ID NO:31):
MRI FAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMT IEC KFPVEKQLDL AALIVYWEME DKNI IQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE
HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVI PELP LAHPPNERPL PLIVGSSVGG
LLLLALITAA LYRLRKGRMM DVKKCGIQDT NSKKQSDTHL EET
( SEQ ID NO 18 )
[0079] In one embodiment, the engineered PD-L1 sequence is called “PDL1-CEACAM1
(TM),” in which the TM domain of a wildtype PD-L1 is replaced with a TM domain from CEACAM1 (boldface; SEQ ID NO:32):
MRI FAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMT IEC KFPVEKQLDL
AALIVYWEME DKNI IQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ
ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVI PELP LAHPPNERAI AGIVIGWAL
VALIAVALAC FLRLRKGRMM DVKKCGIQDT NSKKQSDTHL EET
( SEQ ID NO 19 )
[0080] In one embodiment, the engineered PD-L1 sequence is called “PDL1-ITGA1
(TM),” in which the TM domain of a wildtype PD-L1 is replaced with a TM domain from
ITGA1 (boldface; SEQ ID NO:33):
MRI FAVFI FM TYWHLLNAFT VTVPKDLYVV EYGSNMT IEC KFPVEKQLDL AALIVYWEME DKNI IQFVHG EEDLKVQHSS YRQRARLLKD QLSLGNAALQ ITDVKLQDAG VYRCMISYGG ADYKRITVKV NAPYNKINQR ILVVDPVTSE HELTCQAEGY PKAEVIWTSS DHQVLSGKTT TTNSKREEKL FNVTSTLRIN TTTNEI FYCT FRRLDPEENH TAELVI PELP LAHPPNERLW VILLSAFAGL LLLMLLILAL WRLRKGRMMD VKKCGIQDTN SKKQSDTHLE ET
( SEQ ID NQ : 20 )
[0081] In some embodiments, the engineered PD-L1 sequence comprises the PD-Ll’s extracellular portion that interacts with PD-1 fused to a GPI anchor. In a further embodiment, the engineered PD-L1 sequence is called “PDL1-GPI” and comprises the extracellular region of a wildtype PD-L1 in which the signal sequence is replaced with that of a human NT5E (underlined; SEQ ID NO:29) and the C-terminus of the extracellular region is fused to a GPI anchor with the sequence of SEQ ID NO:50 (italic):
MCPRAARAPA TLLLALGAVL WPAAGAFTVT VPKDLYVVEY GSNMTIECKF PVEKQLDLAA LIVYWEMEDK NI IQFVHGEE DLKVQHSSYR QRARLLKDQL SLGNAALQIT DVKLQDAGVY RCMISYGGAD YKRITVKVNA PYNKINQRIL VVDPVTSEHE LTCQAEGYPK AEVIWTSSDH QVLSGKTTTT NSKREEKLFN VTSTLRINTT TNEIFYCTFR RLDPEENHTA ELVIPELPLA HPPNERS7GS HCHGSFSLIF LSLWAVIFVL YQ ( SEQ ID NO : 21 )
[0082] In some embodiments, the engineered PD-L1 sequence comprises the PD-Ll’s extracellular portion that interacts with PD-1 fused to a GPI-anchorable protein such as NT5E. In a further embodiment, the engineered PD-L1 sequence is called “PDL1-NT5E fusion” and comprises the extracellular region of a wildtype PD-L1 in which the signal sequence is replaced with that of a human NT5E (underlined; SEQ ID NO:29) and the C- terminus of the extracellular region is fused to a mature sequence of NT5E (i.e., without the signal sequence; italic; SEQ ID NO:51) through a short peptide linker with the sequence of SEQ ID NO: 52 (boxed):
MCPRAARAPA TLLLALGAVL WPAAGAFTVT VPKDLYVVEY GSNMTIECKF PVEKQLDLAA LIVYWEMEDK NI IQFVHGEE DLKVQHSSYR QRARLLKDQL SLGNAALQIT DVKLQDAGVY RCMISYGGAD YKRITVKVNA PYNKINQRIL VVDPVTSEHE LTCQAEGYPK AEVIWTSSDH QVLSGKTTTT NSKREEKLFN VTSTLRINTT TNEIFYCTFR RLDPEENHTA ELVIPELPLA HPPNER|GGGG| ^WELTILHTN DVHSRLEQTS EDSSKCVNAS RCMGGVARLF TKVQQIRRAE PNVLLLDAGD QYQGTIWFTV YKGAEVAHFM NALRYDAMAL GNHEFDNGVE GLIEPLLKEA KFPILSANIK AKGPLASQIS GLYLPYKVLP VGDEWGIVG YTSKETPFLS NPGTNLVFED EITALQPEVD KLKTLNVNKI IALGHSGFEM DKLIAQKVRG VDVWGGHSN TFLYTGNPPS KEVPAGKYPF IVTSDDGRKV PVVQAYAFGK YLGYLKIEFD ERGNVISSHG NPILLNSSIP EDPSIKADIN KWRIKLDNYS TQELGKTIVY LDGSSQSCRF RECNMGNLIC DAMINNNLRH TDEMFWNHVS MCILNGGGIR SPIDERNNGT ITWENLAAVL PFGGTFDLVQ LKGSTLKKAF EHSVHRYGQS TGEFLQVGGI HWYDLSRKP GDRVVKLDVL CTKCRVPSYD PLKMDEVYKV ILPNFLANGG DGFQMIKDEL LRHDSGDQDI NVVSTYISKM KVIYPAVEGR IKFSTGSHCH GSFSLIFLSL WAVIFVLYQ ( SEQ ID NO : 22 )
[0083] Variants of the above-described sequences are also within the present disclosure. Variants may be deletions, insertions, and/or substitutions. In some embodiments, the substitutions are conservative substitutions: for example, an aliphatic amino acid (e.g., glycine, alanine, valine, leucine, or isoleucine) may be substituted by another aliphatic amino
acid; a hydroxyl or sulfur/selenium-containing amino acid (e.g., serine, cysteine, selenocysteine, threonine, or methionine) may be substituted by another hydroxyl or sulfur/selenium-containing amino acid; an aromatic amino acid (e.g., phenylalanine, tyrosine, or tryptophan) may be substituted by another aromatic amino acid; a branched chain amino acid (e.g., isoleucine, leucine, or valine) may be substituted by another branched chain amino acid; a basic amino acid (e.g., histidine, lysine, or arginine) may be substituted by another basic amino acid; and an acidic amino acid or an amide thereof (e.g., aspartate, glutamate, asparagine, or glutamine) may be substituted by another acidic amino acid or its amide. In some embodiments, a negatively charged amino acid (e.g., aspartic acid or glutamic acid) may be substituted by another negatively charged amino acid, or a positively charged amino acid (e.g., lysine, arginine and histidine) may be substituted by another positively charged amino acid. In some embodiments, a nonpolar amino acid (e.g., alanine, glycine, isoleucine, leucine, methionine, phenylalanine, proline, valine, or tryptophan) may be substituted by another nonpolar amino acid.
[0084] In some embodiments, the engineered PD-L1 sequence comprises, consists of, or consists essentially of, one of SEQ ID NOs:2-22, 67, and 68 (with or without the signal sequence (i.e., SEQ ID NO:23)), or comprises one of SEQ ID NOs:73-78, and 80.
[0085] In some embodiments, the engineered PD-L1 sequence comprises, consists of, or consists essentially of, an amino acid sequence that is at least 95% (e.g., at least 96, 97, 98, or 99%) identical to one of SEQ ID NOs:2-22, 67, and 68 (with or without the signal sequence), or comprises an amino acid sequence that is at least 95% (e.g., at least 96, 97, 98, or 99%) identical to one of SEQ ID NOs:73-78 and 80. The percent identity of two amino acid sequences (or of two nucleic acid sequences) may be obtained by, e.g., BLAST® using default parameters (available at the U.S. National Library of Medicine’s National Center for Biotechnology Information website). In some embodiments, the length of a reference sequence aligned for comparison purposes is at least 30%, (e.g., at least 40, 50, 60, 70, 80, or 90%) of the reference sequence. In some embodiments, the engineered PD-L1 sequence comprises one or more (e.g., two, three, four, five, six, seven, eight, nine, ten, fifteen, or twenty or more) amino acid mutations (e.g., conservative substitutions) relative to one of SEQ ID NOs:2-22, 67, and 68 (with or without the signal sequence) or one of SEQ ID NOs:73-78 and 80.
[0086] In some embodiments, the engineered PD-L1 sequence comprises a functional equivalent or variant of any of the above-described polypeptide or peptide sequences. The term “functional equivalent” refers to a sequence that serves a biological function that is the
same as or similar to that of the reference sequence. The term “functional variant” refers to a sequence having sequence variations, such as deletions, insertions, and/or substitutions (e.g., conservative substitutions) relative to the reference sequence that do not affect the sequence’s desired biological function. The biological function may be a function related to, e.g., the production, stability, half-life, or affinity for PD-1. For example, the engineered PD-L1 sequence may comprise a functional equivalent or variant of the signal, extracellular, transmembrane, and/or intracellular sequence described above, including those found in SEQ ID NOs:2-22, 67, 68, 73-78, and 80.
[0087] In some embodiments, the engineered PD-L1 polypeptide comprises both mutation(s) that increase PD-1 -binding affinity and mutation(s) that increase resistance to protease cleavage, in addition to mutations shown in one or more of SEQ ID NOs:2-22. For example, the engineered PD-L1 polypeptide comprises two or more mutations selected from N-to-Q mutations, I54E/Q, Y56H/F, E58F/M, R113T, M115L, S117G/A, G119K, G120V, A121W, R238A, an insertion of W between wildtype residues A121 and D122, and replacement of the metalloproteinase cleavage site by a G/S-rich linker (e.g., SEQ ID NO: 72), relative to SEQ ID NO: 1.
II. Expression of Engineered PD-L1 Proteins
[0088] The present disclosure provides expression constructs suitable for expressing the present engineered PD-L1 proteins in cells that are targeted by aberrant immune activity. Expression of the engineered PD-L1 protein will protect the expressing cells from dysregulated T cell activity as the PD-L1 protein binds to PD-1 expressed on T cells and activates the immune checkpoint pathway in the T cells. The engineered PD-L1 protein may be introduced to a cell of interest through a nucleic acid encoding it as further described below.
A. DNA Encoding Engineered PD-L1 Proteins
[0089] In some embodiments, the nucleic acid is DNA (e.g., circular or linear DNA) in the form of, for example, plasmid DNA or a viral vector (e.g., an adenoviral vector, an adeno- associated viral (AAV) vector, a lentiviral vector, or a herpes simplex viral (HSV) vector). The DNA comprises an expression cassette for the PD-L1 protein and may comprise a constitute or inducible promoter for expressing the protein in a cell linked operably to the protein-coding sequence. The expression cassette may also comprise other one or more transcriptional regulatory elements, such as an enhancer, a polyadenylation signal, a transcription-stabilizing element such as woodchuck post-transcriptional element (WPRE), or
any combination thereof. In some embodiments, the promoter in the expression cassette is tissue or cell type specific. For example, the promoter may be liver-specific and may be derived from, for example, a gene for albumin, alpha- 1 -antitrypsin, or transthyretin. In another example, the promoter may be specific for neuronal cells and may be derived from, for example, a gene for NSE, synapsin, CAMKiia and MECPs. In some embodiments, the promoter may be a ubiquitous promoter, such as a CMV, CAG, or Ubc promoter. The PD-L1 expression cassette may exist episomally in the cell or integrated stably (e.g., at a site-specific manner) into the genome of the cell.
[0090] In some embodiments, the DNA may be incorporated into the host cell genome through, for example, gene editing technology such as CRISPR technology.
B. RNA Encoding Engineered PD-L1 Proteins
[0091] In some embodiments, the nucleic acid encoding the engineered PD-L1 for introduction into the cell of interest is RNA. The RNA may be unmodified chemically, or may be chemically modified to, e.g., increase stability, improve production yield, and/or reduce anti-RNA innate immunity in the patient.
[0092] In some embodiments, the RNA may include one or more modified nucleotides including, without limitation, 2’-O-alkyl (e.g., 2’-O-methyl, 2’-O-ethyl, or 2’-O-propryl) nucleotides), 2’-O-aminoalkyl (e.g., 2’-O-aminomethyl, 2’-O-aminoethyl, or 2’-O- aminopropryl) nucleotides, 2’ -deoxy nucleotides, 2’-O-methoxyethyl nucleotides, 2’ -deoxy - 2’ -fluoro nucleotides, pseudo-uridine such as Nl-methyl-pseudouridine and 5 ’methoxyuridine, and the like.
[0093] In some embodiments, the RNA may include one or more alternate internucleotide linkages such as a phosphorothioate linkage and a phosphotriester linkage.
[0094] In some embodiments, the RNA is mRNA. In further embodiments, the mRNA contains one or more nucleotide and/or internucleotide linkage modifications.
[0095] In some embodiments, the RNA is circular RNA. Circular RNA is a singlestranded RNA where the 5’ and 3’ ends present in an RNA molecule have been joined covalently. See, e.g., WO 2019/118919.
[0096] The RNA herein may be delivered to the cell of interest through a lipid nanoparticle (LNP). In some embodiments, the LNP comprises an ionizable lipid (e.g., a cationic lipid), a non-cationic lipid, a conjugated lipid that inhibits aggregation of particles, and a sterol. The amounts of these components can be varied independently to achieve desired properties. For example, the LNP comprises an ionizable lipid (e.g., a cationic lipid) in an amount from about 20% (mol) to about 90% (mol), from about 20% (mol) to about 70%
(mol), from about 30% (mol) to about 60% (mol), or from about 40% (mol) to about 50% (mol), or from about 50% (mol) to about 90% (mol), of the total lipids; a non-cationic lipid in an amount from about 5% (mol) to about 30% (mol) of the total lipids; a conjugated lipid in an amount from about 0.01% (mol) to about 10% (mol) of the total lipids; a PEG-lipid in an amount from about 0.5% (mol) to about 20% (mol); and a sterol in an amount from about 20% (mol) to about 50% (mol) of the total lipids. The ratio of total lipids to nucleic acid can be varied as desired. For example, the total lipids to nucleic acid (mass or weight) ratio can be from about 10: 1 to about 30: 1. In some embodiments, the LNP may be conjugated to a targeting moiety such as an antibody specific for an antigen expressed on the targeted tissue, or a sugar moiety such as N-acetylgalactosamine (GalNAc) for liver targeting.
[0097] In certain cases, the ionizable lipid may be LRN28 or LRN1. See, e.g., WO 2023/044006. LRN1 comprises Formula (i)
[0099] An RNA encoding the engineered PD-L1 protein may also be delivered by a viral vector such as a Sendai viral vector or an alphavirus vector (e.g., Venezuelan equine encephalitis viral vector or an Eastern equine encephalitis viral vector). See also Wong et al., Mol Ther. (2023) 31(11):3127-45.
III. Pharmaceutical Use
[0100] The engineered PD-L1 protein of the present disclosure may be introduced to a target cell in need of protection from autoimmunity. In some embodiments, a nucleic acid for
expressing the protein may be introduced to the cells or tissue of interest systemically, e.g., through intravenous injection. In some embodiments, the carrier for the nucleic acid (e.g., the viral vector or LNP) may preferentially target the cells or tissue of interest. For example, the viral vector may have tropism for the tissue of interest (e.g., AAV9 for the central nervous system and AAV2 for the ocular system). In another example, the LNP may be conjugated to a tissue-homing moiety (e.g., a GalNAc moiety for targeting the liver). The LNP can also be conjugated to other tissue-homing moieties, such as peptides, small molecules, antibodies or fragments thereof, non-antibody scaffolds, and endogenous ligands. For example, an antibody targeting a cell surface receptor on a specific tissue or cell type can be used to direct the LNP specifically to that tissue or cell type. In other embodiments, the nucleic acid or a vector carrying it (e.g., a viral vector) may be delivered locally (e.g., portal vein for liver delivery) to the tissue of interest.
[0101] Accordingly, the present disclosure provides pharmaceutical compositions comprising a nucleic acid for expressing an engineered PD-L1 protein herein and a pharmaceutically acceptable carrier (e.g., water and phosphate-buffered solution). In some embodiments, the pharmaceutical composition comprises a viral vector comprising the nucleic acid. In some embodiments, the pharmaceutical composition comprises an RNA (e.g., mRNA or circular RNA) encoding an engineered PD-L1, wherein the RNA is encapsulated in an LNP.
[0102] In some embodiments, the pharmaceutical composition can be used to treat tissues affected by autoimmunity. For example, the tissue of interest may be the liver (in, e.g., autoimmune hepatis or other immune-driven liver diseases), pancreas (in, e.g., type 1 diabetes mellitus), joints (in, e.g., rheumatoid arthritis), skin (in, e.g., psoriasis and SLE), salivary and/or lacrimal glands (in, e.g., Sjogren’s syndrome), kidney (in, e.g., SLE), central nervous system (in, e.g., multiple sclerosis and SLE), thyroid (in, e.g., Graves’ disease), intestines (in, e.g., Crohn’s disease, ulcerative colitis, or celiac disease), or other tissues affected by autoimmune diseases. In some embodiments, the present engineered PD-L1 may be used to treat autoimmune hepatitis, lupus, Type 1 diabetes, Rheumatoid arthritis, psoriasis, Sjogren’s syndrome, Graves’ disease, multiple sclerosis, myasthenia gravis, neuromyelitis optica, Crohn’s disease, ulcerative colitis, celiac disease, or other autoimmune diseases. In some embodiments, the pharmaceutical composition can be used to prevent tissue rejection by host immune cells in response to an organ transplantation.
[0103] The pharmaceutical composition, DNA, or RNA herein may be delivered to a subject (e.g., a human patient) in need thereof in a therapeutically effective amount. A
“therapeutically effective amount” refers to an amount for effectively treating a disease, for example, preventing the onset of the disease (i.e., prophylactic treatment), preventing or ameliorating one or more symptoms of the disease, and/or arresting or reversing disease progress. For example, a therapeutically effective amount of a pharmaceutical composition, or DNA or RNA herein is an amount sufficient to reduce incidence of aberrant immune responses in the tissue of interest. In some embodiments, the amount does not cause systemic immunosuppression.
[0104] In some embodiments, the pharmaceutical composition comprises a nucleic acid for expressing a polypeptide sequence comprises the sequence of PDL1-K5R, PDL1- APMAP-KtoR (ICD Reverse), PDLl-KtoR,DEtoR, PDL1-GPI, and PDL1-CEACAM1-K5R (ICD), or PDL1-APMAP (ICD reverse) or an amino acid sequence at least 95% (e.g., at least 96, 97, 98, or 99%) identical thereto.
[0105] Unless otherwise defined herein, scientific and technical terms used in connection with the present disclosure shall have the meanings that are commonly understood by those of ordinary skill in the art. Exemplary methods and materials are described below, although methods and materials similar or equivalent to those described herein can also be used in the practice or testing of the present disclosure. In case of conflict, the present specification, including definitions, will control. Further, unless otherwise required by context, singular terms shall include pluralities and plural terms shall include the singular. Throughout this specification and embodiments, the words “have” and “comprise,” or variations such as “has,” “having,” “comprises,” or “comprising,” will be understood to imply the inclusion of a stated integer or group of integers but not the exclusion of any other integer or group of integers. All publications and other references mentioned herein are incorporated by reference in their entirety, as if each individual reference were specifically and individually indicated to be incorporated by reference in its entirety. Although a number of documents are cited herein, this citation does not constitute an admission that any of these documents forms part of the common general knowledge in the art. As used herein, the term “approximately” or “about” as applied to one or more values of interest refers to a value that is similar to a stated reference value. In certain embodiments, the term refers to a range of values that fall within 10%, 9%, 8%, 7%, 6%, 5%, 4%, 3%, 2%, 1%, or less in either direction (greater than or less than) of the stated reference value unless otherwise stated or otherwise evident from the context.
[0106] According to the present disclosure, back-references in the dependent claims are meant as short-hand writing for a direct and unambiguous disclosure of each and every
combination of claims that is indicated by the back-reference. Any compound disclosed herein can be used in any of the treatment method here, wherein the individual to be treated is as defined anywhere herein. Further, headers herein are created for ease of organization and are not intended to limit the scope of the claimed invention in any manner. [0107] In order for the present disclosure to be better understood, the following examples are set forth. These examples are for illustration only and are not to be construed as limiting the scope of the present disclosure in any manner.
EXAMPLES [0108] The studies described in the following Examples aimed to achieve immune reprogramming through expressing an immunomodulatory molecule derived from PD-L1 in a target tissue to repress an aberrant immune response. Supplying the target cells with PD-L1- derived inhibitory signals was shown to locally suppress autoimmunity without systemic immunosuppression. Example 1: Expression of PD-L1 Chimeras from Plasmid DNA
[0109] To address the short half-life of PD-L1, we engineered and evaluated a series of PD-L1 chimeric proteins for their duration of action and therapeutic relevance. Twenty-one engineered PD-L1 proteins were generated. They are shown in Table 1 below.
[0110] To test the expression and ability of the engineered PD-L1 chimeras to bind to PD-1, CH0-K1 epithelial cells were transfected with plasmid DNA encoding the various chimeric proteins. To do so, CHO-K1 cells were seeded in 24-well plates at 40,000 cells per well and grown overnight at 37°C prior to transfection with 500 ng of plasmid DNA and 1.5 pL Lipofectamine2000™ (Invitrogen) per well. The cells were then incubated at 37°C in a 5% CO2 incubator. After 24 hours, the cells were washed in PBS with 2 mM EDTA and then treated with Accutase® Cell Detachment Solution (STEMCELL Technologies) at 37°C for 10 minutes. Cell detachment was monitored under the microscope. Cell culture medium containing fetal bovine serum (FBS) was then added to stop Accutase®.
[0111] The detached cells were transferred to a non-coated 96-well U bottom plate and spun down. The cells were then stained for 20 minutes at 4°C in the dark with an APC- conjugated mouse IgGi antibody for human PD-L1 (MIH2, BioLegend) or an Alexa Fluor® 647-conjugated recombinant human PD-1 (R&D Systems), which contains only the extracellular domain (ECD) of human PD-1. The stained cells were washed with PBS with 2% FBS to remove excess antibody and analyzed by FACS.
[0112] The data show that both detection systems were similarly sensitive in detecting PD-L1 expression in transfected cells (FIG. 2). Most of the engineered PD-L1 fusion proteins in Table 1 showed expression levels comparable to wildtype PD-L1 in transfected cells (FIG. 3A). All engineered PD-L1 proteins retained PD-1 binding affinity, suggesting that the extracellular domain of the protein folded correctly to bind PD-1 (FIG. 3B).
Example 2: Longevity of PD-L1 Chimeras Expressed from Plasmid DNA
[0113] This Example describes a study evaluating the half-life of the PD-L1 chimeras in transfected cells. CHO-K1 cells were seeded in a 24-well plate and transfected as described in Example 1. The transfected cells were incubated at 37°C for 24 hours, 48 hours, 72 hours, and 96 hours in a 5% CO2 incubator. At each timepoint, cells were washed with PBS containing 2 mM EDTA and detached with Accutase®. Detached cells were then stained for viability and PD-L1 expression and analyzed by FACS as described in Example 1, except
anti-PD-Ll antibody clone 29EA3 (Biolegend) was used to detect PD-L1 at 1 :400 dilution for this and all subsequent experiments.
[0114] Out of the 21 PD-L1 chimeras, ten showed a significantly improved expression window based on the change in percent positive cells (FIG. 4, left panel) and mean expression levels in positive cells (FIG. 4, right panel) between the 24-hour and 96-hour timepoints. The ten chimeras were PD-L1-K5R (degradation stabilized control), PD-L1- KtoR,DEtoR, PDL1-ENPP1 (ICD reverse), PDLl-ENPPl-KtoR (ICD reverse), PDL1- CMTM6 fusion with linker, PDL1-CEACAM1 (ICD), PDL1-CEACAM1-K5R (ICD), PDL1-APMAP (ICD reverse), PDLl-APMAP-KtoR (ICD reverse), and PDL1-GPI. This experiment was completed twice with CHO-K1 cells and both transfection replicates yielded consistent results.
[0115] To confirm that the effects of the PD-L1 fusion proteins were not specific to CHO-K1 cells, this study was repeated using OVCAR-3 ovarian adenocarcinoma cells. In this experiment, OVCAR-3 cells were seeded in a 96-well plate at 30,000 cells per well and grown overnight at 37°C. The cells were then transfected with 200 ng of plasmid DNA encoding the PD-L1 fusion proteins and 0.6 pL Lipofectamine2000™ (1 :3 ratio) per well. The transfected cells were incubated at 37°C for 1 day, 5 days, or 9 days in a 5% CO2 incubator with cell culture medium replaced on Day 5 for cells that were cultured for 9 days. Cells were detached, stained, and analyzed by FACS as described above for CHO-K1 cells. [0116] The data obtained from OVCAR-3 are consistent with those obtained from CHO- K1 cells. Again, the following PD-L1 chimeras demonstrated longer half-lives than wildtype PD-L1 : PD-L1-K5R (stabilized control), PDL1-ENPP1 (ICD reverse), PDLl-ENPPl-KtoR (ICD reverse), PDL1-CMTM6 fusion with linker, PDL1-CEACAM1 (ICD), POLICE AC AMI -K5R (ICD), PDL1-APMAP (ICD reverse), PDLl-APMAP-KtoR (ICD reverse), and PDL1-GPI (FIG. 5).
Example 3: Longevity of PD-L1 Chimeras Expressed from Circular RNA
[0117] For therapeutic use, DNA vectors carry safety risks because vector DNA may be incorporated into the host genome. Thus, we next evaluated expression of the PD-L1 chimeras from circular RNA (circRNA, also termed herein “eRNA™” for “Endless RNA™”), which is more stable than conventional linear mRNA. In this study, CHO-K1 cells were seeded in a 96-well plate with 35,000 cells per well and grown overnight at 37°C. The cells were then transfected with 0.2 pmol eRNA encoding the PD-L1 chimeras and 0.6 pL Lipofectamine™ MessengerMAX™ Transfection Reagent (Thermo Fisher) (1 :3 ratio) per
well. The cells were cultured for 1, 2, 3, and 7 days at 37°C in a 5% CO2 incubator. For cells that were cultured for 7 days, they were split 1 : 10 on Day 3 and the culture medium was replaced on Day 5. The cells were detached, stained, and analyzed by FACS as described above.
[0118] The net median fluorescent intensity (MFI) data show that many of the PD-L1 chimeras exhibited improved expression windows as compared to wildtype PD-L1 (FIG. 6). The net MFI value is calculated as the percent positive cells multiplied by the mean expression in positive cells on Day 7. In this experiment, PDL1-K5R, PDLl-APMAP-KtoR (ICD reverse), PDL1-APMAP (ICD reverse), PDLl-KtoR,DEtoR, and PDL1-GPI were among the best performing chimeras.
[0119] To evaluate the consistency of the PD-L1 chimeras’ longevity in different cell lines, the study was repeated in HEK293T cells, a human embryonic kidney cell line. HEK293T cells were seeded in a 96-well plate at 15,000 cells per well and grown overnight at 37°C. The cells were then transfected with 35 ng eRNA encoding the PD-L1 chimeras and 0.1 pL Lipofectamine™ MessengerMAX™ Transfection Reagent (2: 1 ratio) per well. The cells were then incubated for 1, 3, and 7 days at 37°C in a 5% CO2 incubator. For cells that were cultured for 7 days, the culture medium was replaced on Day 5. Cells were detached by pipetting. Detached cells were stained and analyzed by FACS as described above.
[0120] Expressed from circular RNA, many of the PD-L1 chimera fusion proteins exhibited improved expression windows in HEK293T cells based on their Net MFI (FIG. 7). In this experiment, PD-L1-K5R, PDL1-GPI, PDL1-CEACAM1 (TM), PDL1-CEACAM1- K5R (ICD), PDL1-ITGAM (TM), and PD-Ll-KtoR,DEtoR were among the best performers.
Example 4: Validation of in vivo Expression of Human PD-L1 from mRNA in Mice [0121] Given the successful expression of PD-L1 chimeras from plasmid DNA and eRNA, we next evaluated whether stable expression would similarly be possible from mRNA in vivo. On Day 0, C57BL/6 mice were injected intravenously with 1 mg/kg mRNA (NlMePsU-modified) encoding wildtype human PD-L1 and encapsulated into LNPs containing LRN1. PBS was administered as a negative control.
[0122] Liver samples were taken at 6 hours, 24 hours, and 48 hours post-administration. The left lateral lobe of each liver was harvested in Miltenyi MACS Tissue Storage Solution and kept on ice until further processing. The liver lobes were ex vivo perfused using the Miltenyi Liver Perfusion Kit according to the manufacturer’s instructions. Hepatocytes were collected by centrifugation at 30 g for 5 minutes. For flow cytometry analysis, about IxlO6
cells were stained per sample. A live-dead stain (Ghost Dye™ UV 450) was added to the cells for 10 minutes, followed by antibody staining for 20 minutes. The cells were stained with anti-CD45-AF700 (BioLegend; 1 :400), anti-CD26-APC (BioLegend; 1 :200), and anti- PD-L1-PE (BioLegend; 1 :200). The stained cells were washed with FACS buffer and samples were analyzed by flow cytometry.
[0123] Hepatocytes were defined as CD26+CD45 SSChlFSChl and immune cells as CD26+/ CD45+. The percent of PD-L1 positive cells at each time point, as well as the MFI of the PD-L1+ cells at each time point, were measured for both cell types. The data show that both murine hepatocytes and immune cells from treated mice expressed human PD-L1 (FIG. 8). Additionally, lysates of the murine livers were submitted to ELISA analysis using an anti-human PD-L1 ELISA kit (R&D Systems). The ELISA assay indicated that PD-L1 levels decrease over time (FIG. 9).
[0124] These results suggest the potential for PD-L1 supplementation via RNA as a means to clinically address autoimmune hepatitis or other immune-driven diseases.
Example 5: Expression of PD-L1 Chimeras in vivo
[0125] Based on multiple sets of in vitro data described above — plasmid DNA expression in CHO-K1 cells, plasmid DNA expression in OVCAR3 cells, eRNA expression in CHO-K1 cells, and eRNA expression in HEK293T cells — the following PD-L1 chimeras were selected for in vivo studies: PDLl-APMAP-KtoR (ICD Reverse), PDLl-KtoR,DEtoR, PDL1-GPI, and PDL1-CEACAM1-K5R (ICD).
[0126] In this study, C57BL/6 mice were pre-dosed with 5 mg/kg anti-IFNaR-1 monoclonal antibody (clone 5A3, BioXCell) on Day -1 and Day 0. On Day 0, the mice were injected intravenously with 0.75 mg/kg eRNA encoding one of the PD-L1 variants and encapsulated into LNPs containing an ionizable lipid formulation (LRN1). PBS was administered as a negative control. Liver samples were taken at 6 hours, 48 hours, and 120 hours post-administration and prepared for flow cytometry analysis as described in Example 4. Hepatocytes were defined as CD26+CD45'SSChlFSChl. The percent of PD-L1 positive cells at each time point, as well as the MFI of the PD-L1+ cells at each time point were measured. The data show that murine hepatocytes from treated mice expressed human PD- L1 (FIG. 10). Dotted lines represent values obtained in the prior experiment (FIG. 8) with 1 mg/kg mRNA encoding wildtype PD-L1. Expression of half-life engineered PD-L1 proteins PD-Ll-APMAP-KtoR (ICD reverse), PD-L1-CEACAM1-K5R (ICD), and PD-L1-
KtoR,DEtoR led to sustained expression up to 120 hours with less-steep decline of expression percentage and expression levels compared to wildtype PD-L1.
Example 6: Expression of PD-L1 Chimeras in Hepatocytes
[0127] We next sought to assess expression of CEACAM1-K5R (ICD) PD-L1 from circRNA using an LNP that demonstrates preferential tropism to the liver (LP-01) in vivo at two different doses. As controls, we used LP-01 encapsulated GLP-2 circRNA and wildtype PD-L1 5 ’-methoxyuridine (5MoU) mRNA.
[0128] For C57BL/6 mice dosed with circRNA, these mice were pre-dosed with 5 mg/kg anti-IFNaR-1 monoclonal antibody (clone 5A3, BioXCell) on Day 0. One hour after the antibody administration, the mice were injected intravenously with (a) 2.0 mg/kg circRNA encoding GLP-2 (as a negative control), (b) 1.0 mg/kg circRNA encoding the CEACAM1- K5R PD-L1 variant, (c) 3.0 mg/kg circRNA encoding the CEACAM1-K5R PD-L1 variant, (d) 1.0 mg/kg 5MoU mRNA encoding wildtype PD-L1, and (e) PBS buffer (no LNP control). Plasma and liver biopsies were taken on Days 1, 2, 4, and 7 for analysis.
[0129] The data show that circRNA-encoded expression of the CEACAM1-K5R PD-L1 variant at either dose persisted in murine hepatocytes compared to mRNA-encoded wildtype PD-L1, particularly at a circRNA dose of 3 mg/kg (FIG. 11). The circRNA delivered with LP-01 at the 3 mg/kg dose led to expression on more than 90% hepatocytes up to day 4. Similar expression kinetics was observed for the 5MoU mRNA encoding wildtype PD-L1 and the circRNA encoding the CEACAM1-K5R variant delivered at 1 mg/kg.
Example 7: PD-L1 Affinity Mutants
[0130] In an effort to improve potency and generate durable efficacy, we generated several PD-L1 molecules harboring point mutations that were known to bind PD-1 with higher affinity (Liang et al., Oncotarget (2017) 8(51): 88360-75). In addition, we expressed wildtype PD-L2 which exhibits a higher affinity for PD-1 (relative to wildtype PD-L1) (Youngnak et al., Biochem Biophys Res Comm. (2003) 307(3):672-77).
[0131] Briefly, CHO cells were transfected with Nl-Methylpseudouridine-modified mRNAs encoding various PD-L1 mutations shown in Table 2. The PD-L1 L3C7 variant (ECD shown in SEQ ID NO: 75) has the following mutations relative to wildtype PD-L1 : I54E, Y56H, E58F, R113T, Ml 15L, SI 17G, and G119K. The PD-L1 L3C7/A110W variant (ECD shown in SEQ ID NO:76) has one additional A121W mutation relative to PD-L1 L3C7. PD-L1 1 lOLoop (ECD shown in SEQ ID NO: 77) differs from wildtype PD-L1 in that
the former has a G120V mutation, and an insertion of W between wildtype residues A121 and D122. The PD-L1 L3B3 variant (ECD shown in SEQ ID NO:78) has the following mutations relative to wildtype PD-L1 : I54Q, Y56F, E58M, R113T, Ml 15L, SI 17A, and G119K. The PD-L1 Al 10W variant has an A121W mutation relative to wildtype PD-L1. (The PD-L1 Al 10W variant was so named because A121 in PD-L1 corresponds to Al 10 in PD-L2.)
[0132] Twenty-four hours post transfection, cells were detached using trypsin/PBS, incubated with AF647-labeled PD-1 and measured by flow cytometry. The data show that PD-L2 (WT), PD-L1 L3C7, and PD-L1 L3C7/A110W demonstrated improved binding affinity compared to wildtype PD-L1 in both ECso and maximal binding across two independent experiments (FIG. 12). PD-L1 Al 10W mutation was detrimental to PD-L1 affinity by itself, but outperformed wildtype PD-L1 when combined with the L3C7 mutations.
Example 8: Cleavage-Resistant PD-L1 Mutants
[0133] Proteolytic cleavage of PD-L1, which produces soluble PD-L1, may diminish the durability of T cell inhibition (Orme et al., Oncoimmunology (2020) 9(l):el744980). We next designed PD-L1 with point mutations (alanine scanning) within the predicted matrix metalloproteinase cleavage site on PD-L1, i.e., amino acids 230-238 (SEQ ID NO:71), to see whether these mutations would result in reduced production of soluble PD-L1. The mutations were done in the background of CEACAM1-K5R (ICD) PD-L1.
[0134] Briefly, CHO cells were transfected with mRNAs encoding various PD-L1 constructs shown in Table 3. Twenty-four hours post transfection, supernatants were collected to measure soluble PD-L1 and the cells were harvested to measure the cell surface expression of the PD-L1 variants. K5R G3S (SEQ ID NO:81) and (G3S)3 (SEQ ID NO:72) represent the PD-L1 chimeras where the cleavage site has been substituted by one or three
G3S (SEQ ID NO: 81) linkers, respectively, whereas the cleavage site for K5R SCR has been scrambled using the same amino acid composition.
Table 3. PD-L1 Protease Resistant Constructs
[0135] The data show that substitution of the cleavage site with 3x G3S linker (i.e.,
(G3S)3; SEQ ID NO: 72) (ECD shown in SEQ ID NO: 74) or with the single R238A point mutation (ECD shown in SEQ ID NO:73) led to lower levels of soluble PD-L1 and higher cell surface expression (FIG. 13). Based on this data, we believe that mutating the protease cleavage site of our chimeric PD-L1 molecules can be used to maximize the biologic activity ofPD-Ll.
SEQUENCES
[0137] The table below lists the sequences described in the present disclosure (SEQ: SEQ
Claims
1. An engineered programmed cell death 1 ligand (PD-L1) protein comprising an extracellular sequence of a PD-L1 and a transmembrane domain (TM) and/or an intracellular domain (ICD) that is heterologous to PD-L1, wherein the engineered PD-L1 protein has an increased half-life than the corresponding wildtype PD-L1 when expressed in a mammalian cell.
2. The engineered PD-L1 protein of claim 1, wherein the PD-L1 is human PD-L1, optionally wherein the extracellular sequence comprises amino acids 1-219, 1-210, or 1-200 of SEQ ID NO:24, or a variant amino acid sequence at least 95% identical to amino acids 1- 219, 1-210, or 1-200 of SEQ ID NO:24.
3. The engineered PD-L1 protein of claim 2, wherein the variant amino acid sequence comprises one or more mutations relative to SEQ ID NO: 1 selected from N-to-Q mutations, I54E/Q, Y56H/F, E58F/M, R113T, M115L, S117G/A, G119K, G120V, A121W, R238A, an insertion of W between wildtype residues A121 and D122, and a deletion between amino acids 230-238, optionally wherein the variant amino acid sequence comprises (i) an extracellular sequence of SEQ ID NO:54, 55, 56, 57, 67, 68, or 79, or (ii) SEQ ID NO:73, 74, 75, 76, 77, 78, or 80.
4. The engineered PD-L1 protein of any one of claims 1-3, wherein the TM domain comprises a wildtype PD-L1 TM sequence, optionally SEQ ID NO:25.
5. The engineered PD-L1 protein of any one of claims 1-3, wherein the TM domain comprises a TM sequence from integrin alpha-M (ITGAM), optionally SEQ ID NO:31, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO: 18, with or without the signal sequence shown in SEQ ID NO:23.
6. The engineered PD-L1 protein of any one of claims 1-3, wherein the TM domain comprises a TM sequence from carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM1), optionally SEQ ID NO:32, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO: 19, with or without the signal sequence shown in SEQ ID NO:23.
7. The engineered PD-L1 protein of any one of claims 1-3, wherein the TM domain comprises a TM sequence from integrin alpha-1 (ITGA1), optionally SEQ ID NO:33, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:20, with or without the signal sequence shown in SEQ ID NO:23.
8. The engineered PD-L1 protein of any one of claims 1-7, wherein the ICD is a PD-L1- derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:26, 34, 35, or 36.
9. The engineered PD-L1 protein of claim 8, comprising SEQ ID NO:2, 3, or 4, with or without the signal sequence shown in SEQ ID NO:23.
10. The engineered PD-L1 protein of any one of claims 1-9, wherein the ICD comprises a heterologous polypeptide sequence fused, optionally through a peptide linker, to the C- terminus of a PD-L1 -derived ICD sequence.
11. The engineered PD-L1 protein of claim 10, wherein the heterologous polypeptide is a CMTM6 sequence, optionally SEQ ID NO:47, further optionally wherein the engineered PD- L1 protein comprises SEQ ID NO:6, with or without the signal sequence shown in SEQ ID NO:23.
12. The engineered PD-L1 protein of claim 10, wherein the heterologous polypeptide is a CSN5 sequence, optionally SEQ ID NO:48, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:7, with or without the signal sequence shown in SEQ ID NO:23.
13. The engineered PD-L1 protein of any one of claims 1-7, wherein the ICD comprises an ITGAM-derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:37 or 38.
14. The engineered PD-L1 protein of claim 13, comprising SEQ ID NO:8 or 9, with or without the signal sequence shown in SEQ ID NO:23.
15. The engineered PD-L1 protein of any one of claims 1-7, wherein the ICD comprises an CEACAM1 -derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:39 or 40.
16. The engineered PD-L1 protein of claim 15, comprising SEQ ID NO: 10, 11, 67, or 68, with or without the signal sequence shown in SEQ ID NO:23.
17. The engineered PD-L1 protein of any one of claims 1-7, wherein the ICD comprises an ITGAl-derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:41 or 42.
18. The engineered PD-L1 protein of claim 17, comprising SEQ ID NO: 12 or 13, with or without the signal sequence shown in SEQ ID NO:23.
19. The engineered PD-L1 protein of any one of claims 1-7, wherein the ICD comprises an ENPPl-derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:43 or 44.
20. The engineered PD-L1 protein of claim 19, comprising SEQ ID NO: 14 or 15, with or without the signal sequence shown in SEQ ID NO:23.
21. The engineered PD-L1 protein of any one of claims 1-7, wherein the ICD comprises an APMAP-derived ICD sequence, optionally wherein the ICD comprises SEQ ID NO:45 or 46.
22. The engineered PD-L1 protein of claim 21, comprising SEQ ID NO: 16 or 17, with or without the signal sequence shown in SEQ ID NO:23.
23. The engineered PD-L1 protein of any one of claims 1-3, wherein the extracellular sequence of a PD-L1 is fused to a GPI anchor, optionally comprising SEQ ID NO:50, further optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:21.
24. The engineered PD-L1 protein of any one of claims 1-3, wherein the extracellular sequence of a PD-L1 is fused, with or without a peptide linker, to a GPI-anchored protein, optionally NT5E.
25. The engineered PD-L1 protein of claim 24, wherein the GPI-anchored protein comprises SEQ ID NO:51, optionally wherein the engineered PD-L1 protein comprises SEQ ID NO:22.
26. An engineered programmed cell death 1 ligand (PD-L1) protein comprising SEQ ID NO:5, with or without the signal sequence shown in SEQ ID NO:23, or comprising SEQ ID NO:73, 74, 76, 77, or 80.
27. The engineered PD-L1 protein of any one of claims 1-26, further comprising a signal sequence, wherein the signal sequence comprises a PD-L1 signal sequence, optionally SEQ ID NO:23; or a NT5E signal sequence, optionally SEQ ID NO:29.
28. A nucleic acid molecule comprising a coding sequence for the engineered PD-L1 protein of any one of the preceding claims.
29. The nucleic acid molecule of claim 28, wherein the nucleic acid molecule is DNA, mRNA, or circular RNA.
30. A recombinant virus comprising the nucleic acid molecule of claim 28 or 29.
31. A lipid nanoparticle comprising an mRNA or circular RNA of claim 29.
32. A pharmaceutical composition comprising the nucleic acid molecule, recombinant virus, or lipid nanoparticle of any one of claims 28-31, and a pharmaceutically acceptable carrier.
33. A method of treating an autoimmune condition of a tissue in a patient in need thereof, comprising administering the pharmaceutical composition of claim 32 to the patient such that the engineered PD-L1 protein is expressed in the tissue.
34. Use of the nucleic acid molecule, recombinant virus, or lipid nanoparticle of any one of claims 28-31 for the manufacture of a medicament for treating an autoimmune condition in a patient in need thereof in a method of claim 33.
35. The nucleic acid molecule, recombinant virus, lipid nanoparticle, or pharmaceutical composition of any one of claims 28-32 for use in treating an autoimmune condition in a patient in need thereof in a method of claim 33.
36. The method, use, or nucleic acid molecule, recombinant virus, lipid nanoparticle, or pharmaceutical composition for use of any one of claims 33-35, wherein the tissue is the liver and the autoimmune condition is autoimmune hepatitis.
37. A method of preventing rejection of an organ transplant in a patient in need thereof, comprising administering the pharmaceutical composition of claim 32 to the transplant such that the engineered PD-L1 protein is expressed in the transplant.
38. Use of the nucleic acid molecule, recombinant virus, lipid nanoparticle, or pharmaceutical composition of any one of claims 28-32 for the manufacture of a medicament for preventing rejection of an organ transplant in a patient in need thereof in a method of claim 37.
39. The nucleic acid molecule, recombinant virus, lipid nanoparticle, or pharmaceutical composition of any one of claims 28-32 for use in preventing rejection of an organ transplant in a patient in need thereof in a method of claim 37.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202463617869P | 2024-01-05 | 2024-01-05 | |
| US63/617,869 | 2024-01-05 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025147583A1 true WO2025147583A1 (en) | 2025-07-10 |
Family
ID=94393890
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2025/010194 Pending WO2025147583A1 (en) | 2024-01-05 | 2025-01-03 | Engineered pd-1 ligands |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025147583A1 (en) |
Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2019118919A1 (en) | 2017-12-15 | 2019-06-20 | Flagship Pioneering, Inc. | Compositions comprising circular polyribonucleotides and uses thereof |
| WO2020257710A1 (en) * | 2019-06-21 | 2020-12-24 | Entelexo Biotherapeutics Inc. | Platforms, compositions, and methods for therapeutics delivery |
| WO2021159016A1 (en) * | 2020-02-05 | 2021-08-12 | Diadem Biotherapeutics Inc. | Artificial synapses |
| WO2022245841A1 (en) * | 2021-05-17 | 2022-11-24 | Wisconsin Alumni Research Foundation | Synthetic protein for inducing immune tolerance |
| WO2023044006A1 (en) | 2021-09-17 | 2023-03-23 | Flagship Pioneering Innovations Vi, Llc | Compositions and methods for producing circular polyribonucleotides |
-
2025
- 2025-01-03 WO PCT/US2025/010194 patent/WO2025147583A1/en active Pending
Patent Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2019118919A1 (en) | 2017-12-15 | 2019-06-20 | Flagship Pioneering, Inc. | Compositions comprising circular polyribonucleotides and uses thereof |
| WO2020257710A1 (en) * | 2019-06-21 | 2020-12-24 | Entelexo Biotherapeutics Inc. | Platforms, compositions, and methods for therapeutics delivery |
| WO2021159016A1 (en) * | 2020-02-05 | 2021-08-12 | Diadem Biotherapeutics Inc. | Artificial synapses |
| WO2022245841A1 (en) * | 2021-05-17 | 2022-11-24 | Wisconsin Alumni Research Foundation | Synthetic protein for inducing immune tolerance |
| WO2023044006A1 (en) | 2021-09-17 | 2023-03-23 | Flagship Pioneering Innovations Vi, Llc | Compositions and methods for producing circular polyribonucleotides |
Non-Patent Citations (6)
| Title |
|---|
| AGINA ET AL., CLIN EXP HEPATOL., vol. 5, no. 3, 2019, pages 256 - 64 |
| JILKOVA ET AL., CELLS., vol. 10, no. 10, 2021, pages 2671 |
| LIANG ET AL., ONCOTARGET, vol. 8, no. 51, 2017, pages 88360 - 75 |
| ORME ET AL., ONCOIMMUNOLOGY, vol. 9, no. 1, 2020, pages e1744980 |
| WONG ET AL., MOL THER., vol. 31, no. 11, 2023, pages 3127 - 45 |
| YOUNGNAK ET AL., BIOCHEM BIOPHYS RES COMM., vol. 307, no. 3, 2003, pages 672 - 77 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| KR101993714B1 (en) | Compositions and methods of use in treating metabolic disorders | |
| KR102211177B1 (en) | Interleukin-15 fusion protein for tumor target therapy | |
| JP6704358B2 (en) | Compositions and methods of use for treating metabolic disorders | |
| CN107922489B (en) | Kappa Myeloma Antigen Chimeric Antigen Receptor and Uses Thereof | |
| EP4556567A2 (en) | Rationally-designed synthetic peptide shuttle agents for delivering polypeptide cargos from an extracellular space to the cytosol and/or nucleus of a target eukaryotic cell, uses thereof, methods and kits relating to same | |
| KR20250139414A (en) | Ultra-long acting insulin-fc fusion proteins and methods of use | |
| JP2016523268A (en) | Highly stable T cell receptor and its production and use | |
| KR20210090157A (en) | cell penetrating peptide | |
| ES2971719T3 (en) | Production of MHC II/CII complexes | |
| JP5689680B2 (en) | Targeted induction of protein aggregation in crossbeta structures | |
| CA3137809C (en) | Hla-dr/cii peptide complexes for treating arthritis | |
| WO2011102845A1 (en) | Rage fusion protein compositions and methods of use | |
| KR102348838B1 (en) | Active TRAIL trimer and tumor targeting peptide multi-displayed on ferritin nanocage and use in anti-cancer agent thereof | |
| JP4342314B2 (en) | Cleavage agents for specific delivery to the lesion site | |
| WO2025147583A1 (en) | Engineered pd-1 ligands | |
| CN113645990B (en) | Improved prostate apoptosis response-4 (PAR-4) polypeptides and methods of producing and using the same | |
| JP2026503660A (en) | Engineered repair regulatory T cells | |
| JP2007508806A (en) | Novel therapeutic fusion protein | |
| CN112236159B (en) | OCA-B peptide conjugates and therapeutic methods | |
| EP3496741B1 (en) | Novel therapy to achieve glycemic control | |
| KR20210005612A (en) | Gene therapy for oxidative stress | |
| HK40063235B (en) | Hla-dr/cii peptide complexes for treating arthritis | |
| HK40064159B (en) | Production of mhc ii/cii complexes | |
| NZ728101B2 (en) | Compositions and methods of use for treating metabolic disorders |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 25702367 Country of ref document: EP Kind code of ref document: A1 |