WO2025133290A1 - Protein for immune regulation - Google Patents
Protein for immune regulation Download PDFInfo
- Publication number
- WO2025133290A1 WO2025133290A1 PCT/EP2024/088159 EP2024088159W WO2025133290A1 WO 2025133290 A1 WO2025133290 A1 WO 2025133290A1 EP 2024088159 W EP2024088159 W EP 2024088159W WO 2025133290 A1 WO2025133290 A1 WO 2025133290A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- seq
- amino acid
- tafa4
- substitution
- mutated
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/52—Cytokines; Lymphokines; Interferons
- C07K14/521—Chemokines
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K38/00—Medicinal preparations containing peptides
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/30—Non-immunoglobulin-derived peptide or protein having an immunoglobulin constant or Fc region, or a fragment thereof, attached thereto
Definitions
- the present invention relates to TAFA4 polypeptide, fusion protein, pharmaceutical composition and methods for treating inflammatory disease.
- the skin serves as an important boundary between the internal milieu and the environment, preventing contact with potentially harmful antigens.
- an inflammatory response is induced to eliminate the antigen.
- This response leads to a dermal infiltrate that consists predominantly of T cells, polymophonuclear cells, and macrophages.
- this inflammatory response triggered by the pathogen, is under tight control and will be halted upon elimination of the pathogen.
- the inflammatory response occurs in absence of pathogen.
- UV radiation causes sunburn-like damage characterized by the destruction of the epidermis and inflammation of the underneath dermal papilla.
- Tissue-specific signals constantly shape resident macrophage functional identity and promote their maintenance throughout the lifespan, either by local self-renewal or by the recruitment of additional monocyte-derived cells.
- the (a) one or more immunoglobulin constant domains or fragments thereof are selected from the group comprising or consisting of IgA, IgD, IgE, IgG and IgM, preferably IgG.
- the invention further relates to a fusion protein comprising a plurality of the fusion polypeptides according to the invention.
- the invention further relates to a nucleic acid sequence encoding the fusion polypeptide according to the invention or the fusion protein according to the invention.
- the invention further relates to a vector comprising a nucleic acid sequence encoding the fusion polypeptide or the fusion protein according to the invention.
- the invention further relates to a cell or cell population comprising one or more fusion polypeptides according to the invention, or one or more fusion proteins according to the invention, or one or more nucleic acid sequences according to the invention, or one or more vectors according to the invention.
- the invention further relates to a pharmaceutical composition
- a pharmaceutical composition comprising one or more fusion polypeptides according to the invention, a fusion protein according to the invention, one or more nucleic acid sequences according to the invention, one or more vectors according to the invention, or one or more cells or cell population according to the invention; and one or more pharmaceutically acceptable excipients or carriers.
- the invention further relates to the fusion polypeptide according to the invention, the fusion protein according to the invention, the nucleic acid sequence according to the invention, the vector according to the invention, the cell or the cell population the invention or the pharmaceutical composition according to the invention, for use as a medicament.
- the invention further relates to the fusion polypeptide according to the invention, the fusion protein according to the invention, the nucleic acid sequence according to the invention, the vector according to the invention, the cell or the cell population according to the invention or the pharmaceutical composition according to the invention, for use for treating disease associated with the modulation of macrophage in a subject in need thereof
- the disease is an inflammation disease, more preferably a skin inflammation disease.
- the pharmaceutical composition is formulated for topical, intramuscular, subcutaneous, intradermal, intravenous or oral administration, preferably the topical administration is accomplished via a transdermal device or patch device.
- administering refers to providing a therapeutic agent (e.g., a compound of the invention) alone or as part of a pharmaceutically acceptable composition, to for example a patient in whom/which the condition, symptom, or disease is to be treated.
- a therapeutic agent e.g., a compound of the invention
- amino acid substitution is a substitution of one amino acid for another.
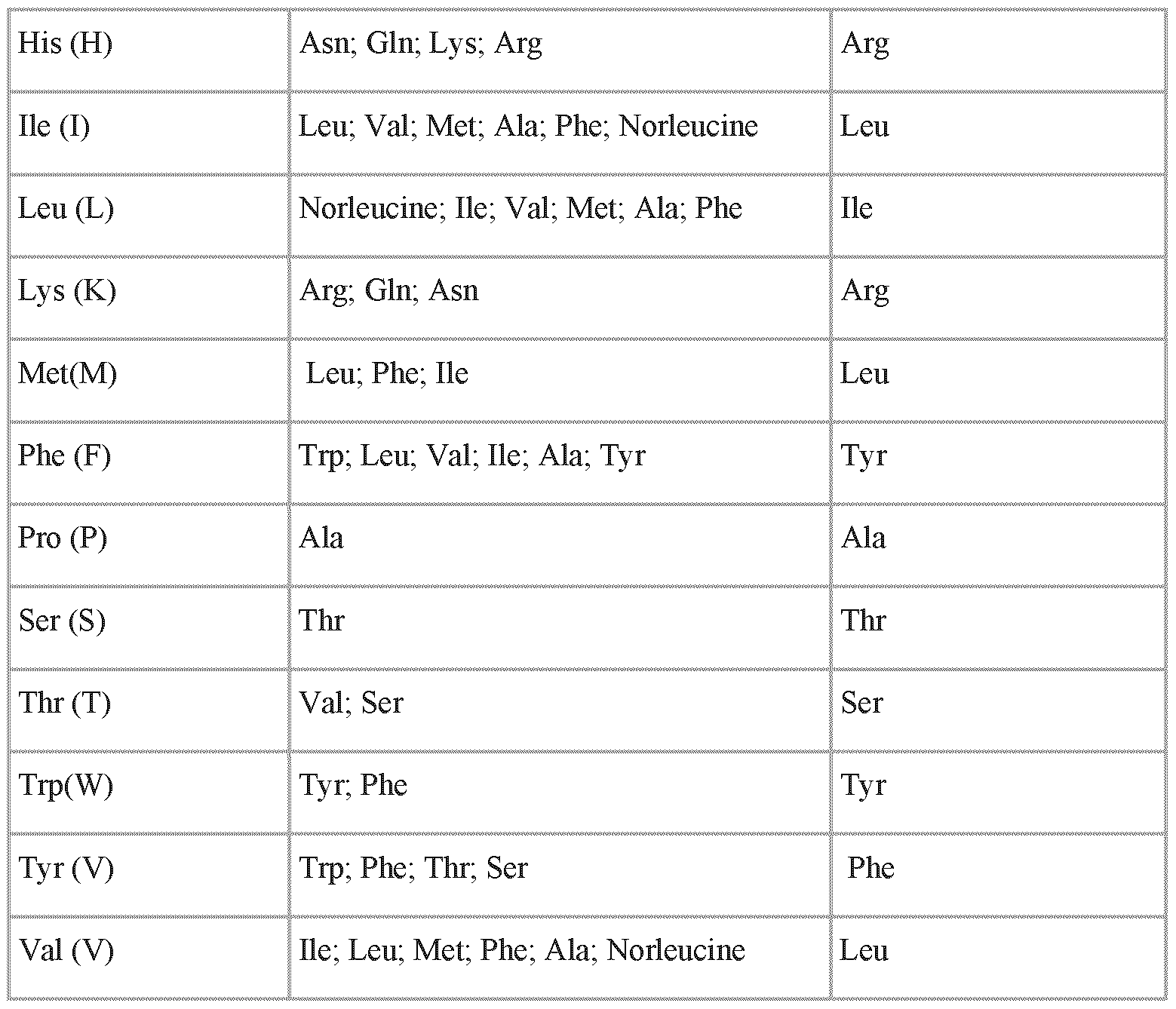
- a conservative amino acid substitution is a substitution of one amino acid for another with similar characteristics.
- Conservative amino acid substitutions include substitutions within the following groups: valine, alanine and glycine; leucine, valine, and isoleucine; aspartic acid and glutamic acid; asparagine and glutamine; serine, cysteine, and threonine; lysine and arginine; and phenylalanine and tyrosine.
- the nonpolar hydrophobic amino acids include alanine, leucine, isoleucine, valine, proline, phenylalanine, tryptophan and methionine.
- Antibody and “immunoglobulin”, as used herein, may be used interchangeably and refer to a protein having a combination of two heavy and two light chains whether or not it possesses any relevant specific immunoreactivity. “Antibodies” refers to such assemblies which have significant known specific immunoreactive activity to an antigen of interest. Antibodies and immunoglobulins comprise light and heavy chains, with or without an interchain covalent linkage between them. Basic immunoglobulin structures in vertebrate systems are relatively well understood. The generic term “immunoglobulin” comprises five distinct classes of antibody that can be distinguished biochemically. Although the following discussion will generally be directed to the IgG class of immunoglobulin molecules, all five classes of antibodies are within the scope of the present invention.
- the light and heavy chains are covalently bonded to each other, and the “tail” regions of the two heavy chains are bonded to each other by covalent disulfide linkages or non-covalent linkages when the immunoglobulins are generated either by hybridomas, B cells or genetically engineered host cells.
- the amino acid sequences run from an N- terminus at the forked ends of the Y configuration to the C -terminus at the bottom of each chain.
- heavy chains are classified as gamma (y), mu (p), alpha (a), delta (5) or epsilon (e) with some subclasses among them (e.g., yl- y4).
- immunoglobulin subclasses or “isotypes” e.g., IgGl, IgG2, IgG3, IgG4, IgAl, etc.
- isotypes e.g., IgGl, IgG2, IgG3, IgG4, IgAl, etc.
- Modified versions of each of these classes and isotypes are readily discernable to the skilled artisan in view of the instant disclosure and, accordingly, are within the scope of the present invention.
- the variable region of an antibody allows the antibody to selectively recognize and specifically bind epitopes on antigens.
- VL domain light chain variable domain
- VH domain heavy chain variable domain
- This quaternary antibody structure forms the antigen binding site presents at the end of each arm of the “Y”. More specifically, the antigen binding site is defined by three complementarity determining regions (CDRs) on each of the VH and VL chains.
- CDRs complementarity determining regions
- Combined treatment refers to a treatment that uses more than one medication.
- the combined therapy may be dual therapy or bi-therapy.
- the medications used in the combined treatment according to the invention are administered to the subject simultaneously, separately or sequentially.
- administration simultaneously refers to administration of 2 active ingredients by the same route and at the same time or at substantially the same time.
- administration separately refers to an administration of 2 active ingredients at the same time or at substantially the same time by different routes.
- administration sequentially refers to an administration of 2 active ingredients at different times, the administration route being identical or different.
- Comprise or a variant thereof (e.g., “comprises”, “comprising”) refers to according to common patent application drafting terminology. Hence, “comprise” preceded by an object and followed by a constituent means that the presence of a constituent in the object is required (typically as a component of a composition), but without excluding the presence of any further constituent(s) in the object. Moreover, any occurrence of “comprise” or a variant thereof herein also encompasses narrower expression “substantially consist of’, further narrower expression “consist of’ and any variants thereof (e.g., “consists of’, “consisting of’), unless otherwise stated.
- Control sequences refers collectively to promoter sequences, polyadenylation signals, transcription termination sequences, upstream regulatory domains, origins of replication, internal ribosome entry sites ("IRES"), enhancers, and the like, which collectively provide for the replication, transcription and translation of a coding sequence in a recipient cell. Not all of these control sequences need always be present so long as the selected coding sequence is capable of being replicated, transcribed and translated in an appropriate host cell.
- “Expression cassette” refers to a unit cassette capable of expressing a gene of interest, as the transgene, operably linked to a downstream side of a promoter, including a promoter and a gene of interest. Various factors that can help the efficient expression of the gene of interest may be included inside or outside such a gene expression cassette.
- the gene expression cassette may include, but is not limited to, a transcription termination signal, a ribosome binding site, and a translation termination signal in addition to a promoter operably linked to the gene of interest.
- “Homology” or “identity” refers to the subunit sequence identity between two polymeric molecules, e.g., between two nucleic acid molecules, such as, two DNA molecules or two RNA molecules, or between two polypeptide molecules. When a subunit position in both of the two molecules is occupied by the same monomeric subunit; e.g., if a position in each of two DNA molecules is occupied by adenine, then they are homologous or identical at that position.
- the homology between two sequences is a direct function of the number of matching or homologous positions; e.g., if half (e.g., five positions in a polymer ten subunits in length) of the positions in two sequences are homologous, the two sequences are 50% homologous; if 90% of the positions (e.g., 9 of 10), are matched or homologous, the two sequences are 90% homologous.
- homologous or “identical” when used in a relationship between the sequences of two or more polypeptides or of two or more nucleic acid molecules, refers to the degree of sequence relatedness between polypeptides or nucleic acid molecules, as determined by the number of matches between strings of two or more amino acid or nucleotide residues. “Identity” measures the percent of identical matches between the smaller of two or more sequences with gap alignments (if any) addressed by a particular mathematical model or computer program (i.e., "algorithms"). Identity of related polypeptides can be readily calculated by known methods. Such methods include, but are not limited to, those described in Computational Molecular Biology, Lesk, A.
- Preferred methods for determining identity are designed to give the largest match between the sequences tested. Methods of determining identity are described in publicly available computer programs. Preferred computer program methods for determining identity between two sequences include the GCG program package, including GAP (Devereux et al., Nucl. Acid. Res. ⁇ 2, 387 (1984); Genetics Computer Group, University of Wisconsin, Madison, Wis.), BLASTP, BLASTN, and FASTA (Altschul et al., J. Mol. Biol. 215, 403-410 (1990)). The BLASTX program is publicly available from the National Center for Biotechnology Information (NCBI) and other sources (BLAST Manual, Altschul et al. NCB/NLM/NIH Bethesda, Md.
- NCBI National Center for Biotechnology Information
- Immunoglobulin includes a protein having a combination of two heavy and two light chains whether or not it possesses any relevant specific immunoreactivity
- Inflammatory disease is used herein in the broadest sense and includes all diseases and pathological conditions having etiologies associated with a systemic or local abnormal and/or uncontrolled inflammatory response. For instance, over-expression of proinflammatory cytokines without proper controls leads to a variety of inflammatory diseases and disorders. This term includes both acute inflammatory diseases and chronic inflammatory diseases.
- Linker refers to a sequence of at least one amino acid that can link the TAFA4 polypeptide with the immunoglobulin chain or a fragment thereof.
- Linkers are well known to one of ordinary skill in the art and typically comprises 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20 or more amino acids. It can refer to a single covalent bond or a moiety comprising series of stable covalent bonds, the moiety often incorporating 1-40 plural valent atoms selected from the group consisting of C, N, O, S and P, that covalently attach a reactive group or bioactive group to the probe of the invention.
- the number of plural valent atoms in a linker may be, for example, 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 25, 30 or a larger number up to 40 or more.
- a linker may be linear or non-linear; some linkers have pendant side chains or pendant functional groups (or both). Examples of such pendant moieties are hydrophilicity modifiers, for example solubilizing groups like, e.g. sulfo (-SO3H or -SO3-), carboxy (-COOH or -COO-), hydroxy.
- the linker is composed of any combination of single, double, triple or aromatic carbon-carbon bonds, carbon-nitrogen bonds, nitrogen-nitrogen bonds, carbon-oxygen bonds and carbon-sulfur bonds.
- Linkers may by way of example consist of a combination of moieties selected from alkyl, -C(O)NH-, -C(O)O-, -NH-, -S- , -O-, -C(O) -, -S(O)n- where n is 0, 1 or 2; -O-, 5- or 6- membered monocyclic rings and optional pendant functional groups, for example sulfo, hydroxy and carboxy.
- hinge or “hinge region” or “hinge domain” refers, in antibodies, to the flexible portion of a heavy chain located between the CHI domain and the CH2 domain. It is approximately 25 amino acids long, and is divided into an “upper hinge,” a “middle hinge” or “core hinge,” and a “lower hinge.”
- a “hinge subdomain” refers to the upper hinge, middle (or core) hinge or the lower hinge.
- mammal refers to any animal classified as a mammal, including humans, other higher primates, domestic and farm animals, and zoo, sports, or pet animals, such as dogs, cats, cattle, horses, sheep, pigs, goats, rabbits, mouse, rat, etc. Preferably, the mammal is human.
- Nucleic acid or “Polynucleotide” refer to a polymeric form of nucleotides of any length, either ribonucleotides or deoxyribonucleotides. Thus, these terms include, but are not limited to, single-, double- or multi-stranded DNA or RNA, genomic DNA, cDNA, DNA-RNA hybrids, or a polymer comprising purine and pyrimidine bases, or other natural, chemically or biochemically modified, non-natural, or derivatized nucleotide bases.
- the backbone of the polynucleotide can comprise sugars and phosphate groups (as may typically be found in RNA or DNA), or modified or substituted sugar or phosphate groups.
- the backbone of the polynucleotide can comprise a polymer of synthetic subunits such as phosphoramidates and thus can be an oligodeoxynucleoside phosphoramidate (P-NH2) or a mixed phosphoramidate- phosphodiester oligomer.
- P-NH2 oligodeoxynucleoside phosphoramidate
- a double -stranded polynucleotide can be obtained from the single stranded polynucleotide product of chemical synthesis either by synthesizing the complementary strand and annealing the strands under appropriate conditions, or by synthesizing the complementary strand de novo using a DNA polymerase with an appropriate primer.
- operably linked refers to functional linkage between a regulatory sequence and a heterologous nucleic acid sequence resulting in expression of the latter.
- a first nucleic acid sequence is operably linked with a second nucleic acid sequence when the first nucleic acid sequence is placed in a functional relationship with the second nucleic acid sequence.
- a promoter is operably linked to a coding sequence if the promoter affects the transcription or expression of the coding sequence.
- operably linked DNA sequences are contiguous and, where necessary to join two protein coding regions, in the same reading frame.
- Patient refers to a subject who/which is awaiting the receipt of, or is receiving medical care or was/is/will be the object of a medical procedure, or is monitored for the development of the targeted disease or condition, such as, for example, an infectious disease.
- Protein refers to a compound comprised of amino acid residues covalently linked by peptide bonds.
- a protein or peptide must contain at least two amino acids, and no limitation is placed on the maximum number of amino acids that can comprise a protein's or peptide's sequence.
- Polypeptides include any peptide or protein comprising two or more amino acids joined to each other by peptide bonds.
- the term refers to both short chains, which also commonly are referred to in the art as peptides, oligopeptides and oligomers, for example, and to longer chains, which generally are referred to in the art as proteins, of which there are many types.
- Polypeptides include, for example, biologically active fragments, substantially homologous polypeptides, oligopeptides, homodimers, heterodimers, variants of polypeptides, modified polypeptides, derivatives, analogs, fusion proteins, among others.
- the polypeptides include natural peptides, recombinant peptides, synthetic peptides, or a combination thereof.
- Proteinogenic amino acids comprise Alanine (A), Arginine (R), Asparagine (N), Aspartic acid (D), Cysteine (C), Glutamine (Q), Glutamic acid (E), Glycine (G), Histidine (H), Isoleucine (I), Leucine (L), Lysine (K), Methionine (M), Phenylalanine (F), Proline (P), Serine (S), Threonine (T), Tryptophan (W), Tyrosine (Y), Valine (V).
- “Pharmaceutically acceptable” means that the ingredients of a composition are compatible with each other and not deleterious to the patient to which/whom it is administered.
- “Pharmaceutically acceptable carrier” refers to an excipient that does not produce an adverse, allergic or other untoward reaction when administered to an animal, preferably a human. It includes any and all solvents, dispersion media, coatings, antibacterial and antifungal agents, isotonic and absorption delaying agents and the like. For human administration, preparations should meet sterility, pyrogenicity, general safety and purity standards as required by regulatory offices, such as, e.g., FDA Office or EMA.
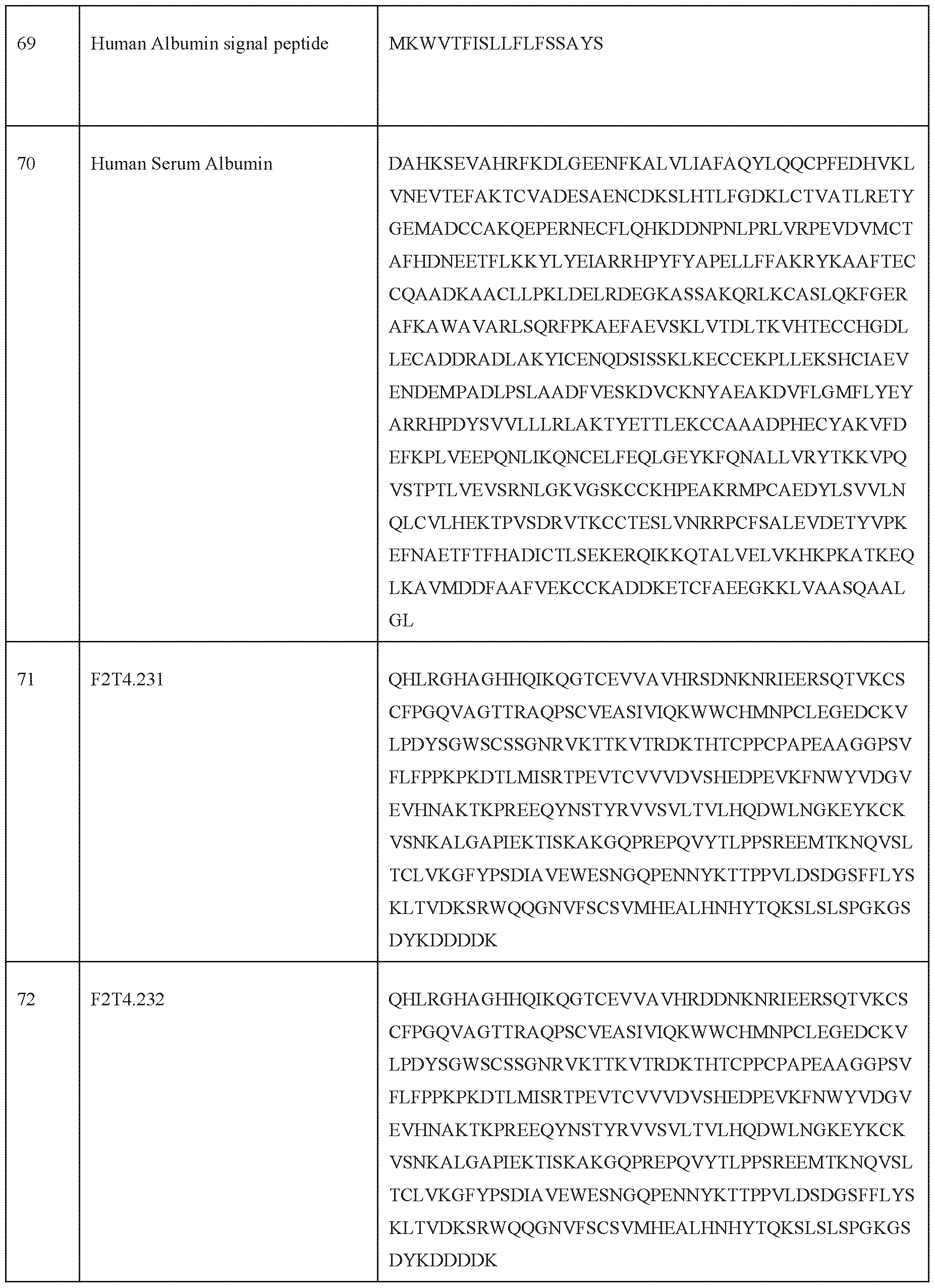
- Examples of pharmaceutically acceptable carriers include, but are not limited to, ion exchangers, alumina, aluminum stearate, lecithin, serum proteins, such as human serum albumin, buffer substances such as phosphates, glycine, sorbic acid, potassium sorbate, partial glyceride mixtures of saturated vegetable fatty acids, water, salts or electrolytes, such as protamine sulfate, di sodium hydrogen phosphate, potassium hydrogen phosphate, sodium chloride, zinc salts, colloidal silica, magnesium trisilicate, polyvinyl pyrrolidone, cellulose-based substances (for example sodium carboxymethylcellulose), polyethylene glycol, polyacrylates, waxes, polyethylene- polyoxypropylene- block polymers, polyethylene glycol and wool fat.
- ion exchangers alumina, aluminum stearate, lecithin
- serum proteins such as human serum albumin
- buffer substances such as phosphates, glycine, sorbic acid, potassium sorbate
- “Pharmaceutical composition” refers to a composition comprising an active agent in association with a pharmaceutically acceptable vehicle or excipient.
- a pharmaceutical composition is for therapeutic use, and relates to health. Especially, a pharmaceutical composition may be indicated for treating or preventing a disease.
- polynucleotide refers to a chain of nucleotides.
- nucleic acids are polymers of nucleotides.
- nucleic acids and polynucleotides as used herein are interchangeable.
- nucleic acids are polynucleotides, which can be hydrolyzed into the monomeric "nucleotides.”
- the monomeric nucleotides can be hydrolyzed into nucleosides.
- polynucleotides include, but are not limited to, all nucleic acid sequences which are obtained by any means available in the art, including, without limitation, recombinant means, i.e. , the cloning of nucleic acid sequences from a recombinant library or a cell genome, using ordinary cloning technology and PCR, and the like, and by synthetic means.
- Promoter sequence which is used herein in its ordinary sense to refer to a nucleotide region comprising a DNA regulatory sequence, wherein the regulatory sequence is derived from a gene which is capable of binding RNA polymerase and initiating transcription of a downstream (3 '-direction) coding sequence.
- Transcription promoters can include "inducible promoters” (where expression of a polynucleotide sequence operably linked to the promoter is induced by an analyte, cofactor, regulatory protein, etc.), “repressible promoters” (where expression of a polynucleotide sequence operably linked to the promoter is induced by an analyte, cofactor, regulatory protein, etc.), and “constitutive promoters”.
- the core of signal sequence or “signal sequence”, or “signal peptide” refers to a long stretch of hydrophobic amino acids (about 5-16 residues long) that has a tendency to form a single alpha-helix and is also referred to as the "h-region".
- signal peptides may begin with a short positively charged stretch of amino acids, which may help to enforce proper topology of the polypeptide during translocation by what is known as the positive-inside rule. Because of its close location to the N- terminus it is called the "n-region". At the end of the signal peptide there may be typically a stretch of amino acids that may be recognized and cleaved by signal peptidase and therefore named cleavage site.
- the signal peptidase may cleave either during or after completion of translocation to generate a free signal peptide and a mature protein.
- the free signal peptides can be then digested by specific proteases.
- signal sequences may act via the co-translational pathway, which is initiated when the signal peptide emerges from the ribosome and is recognized by the signal-recognition particle (SRP).
- SRP signal-recognition particle
- the SRP bound to the signal sequence then halts further translation (translational arrest) and directs the signal sequence-ribosome-mRNA complex to the SRP receptor, which is present on the surface of the ER.
- the signal sequence used to be is inserted into the translocon. Ribosomes are then physically docked onto the cytoplasmic face of the translocon and protein synthesis resumes.
- “Selected from” refers to according to common patent application drafting terminology, to introduce a list of elements among which one or more item(s) is (are) selected. Any occurrence of “selected from” in the specification may be replaced by “selected from the group comprising or consisting of’ and reciprocally without changing the meaning thereof.
- the term "Silenced Fc” refers to a genetically engineered Fc domain comprising mutations that abrogate binding of the Fc domain to Fc receptors (FcyR, FcR) while maintain the ability of the Fc domain to binds to neonatal Fc receptor (FcRn). Such silenced Fc exhibits extended half-life as described in Borrok M.J, et al. J Pharm Sci. 2017. [54] “Therapeutic agent”, “active pharmaceutical ingredient” and “active ingredient” refer to a compound for therapeutic use and relating to health. Especially, a therapeutic agent (e.g., a compound of the invention) may be indicated for treating a disease.
- a therapeutic agent e.g., a compound of the invention
- An active ingredient may also be indicated for improving the therapeutic activity of another therapeutic agent. It describes a molecule or a substance, preferably a biological molecule such as for example an oligonucleotide, a siRNA, a miRNA, a DNA fragment, an aptamer, an antibody and the like, or a chemical entity, whose administration to a subject slows down or stops the progression, aggravation, or deterioration of one or more symptoms of a disease, or condition; alleviates the symptoms of a disease or condition; cures a disease or condition.
- a biological molecule such as for example an oligonucleotide, a siRNA, a miRNA, a DNA fragment, an aptamer, an antibody and the like, or a chemical entity, whose administration to a subject slows down or stops the progression, aggravation, or deterioration of one or more symptoms of a disease, or condition; alleviates the symptoms of a disease or condition; cures a disease or condition.
- “Therapeutically effective amount” refers to level or amount of agent that is aimed at, without causing significant negative or adverse side effects to the target, (1) delaying or preventing the onset of the targeted pathologic condition or disorder; (2) slowing down or stopping the progression, aggravation, or deterioration of one or more symptoms of the targeted pathologic condition or disorder; (3) bringing about ameliorations of the symptoms of the targeted pathologic condition or disorder; (4) reducing the severity or incidence of the targeted pathologic condition or disorder; (5) curing the targeted pathologic condition or disorder.
- An effective amount may be administered prior to the onset of the targeted pathologic condition or disorder, for a prophylactic or preventive action. Alternatively, or additionally, the effective amount may be administered after initiation of the targeted pathologic condition or disorder, for a therapeutic action.
- Treating refers to reducing or alleviating at least one adverse effect or symptom of a disease, disorder or condition associated with a deficiency in or absence of an organ, tissue or cell function.
- Those in need of treatment include those already with the disorder as well as those prone to have the disorder or those in whom the disorder is to be prevented.
- a subject is successfully “treated” for the targeted pathologic condition or disorder if, after receiving a therapeutic amount of coated vector as described herein, the subject shows observable and/or measurable improvement in one or more of the following: reduction in the number of pathogenic cells; reduction in the percent of total cells that are pathogenic; relief to some extent of one or more of the symptoms associated with the targeted pathologic condition or disorder; reduced morbidity and mortality, and/or improvement in quality of life issues.
- the above parameters for assessing successful treatment and improvement in the disease are readily measurable by routine procedures familiar to a physician.
- variant refers to nucleic acid or amino acid sequences that typically differs from a nucleic acid or an amino acid sequence specifically disclosed herein in one or more substitutions, deletions, additions and/or insertions. Such variants may be naturally occurring or may be synthetically generated, for example, by modifying one or more of the nucleic acid or amino acid sequences of the invention and evaluating one or more biological activities of the encoded polypeptide as described herein and/or using any of a number of techniques well known in the art.
- nucleic acid or amino acid sequence comprising at least 85%, 90%, 95%, 96%, 97%, 98%, 99% or more sequence 10 identity with the reference nucleic acid or amino acid sequence.
- amino acid sequence of a polypeptide to create an equivalent, or even an improved, variant or portion of a polypeptide of the invention, one skilled in the art will typically change one or more of the codons of the encoding DNA sequence.
- certain amino acids may be substituted for other amino acids in a protein structure without appreciable loss of its ability to bind other polypeptides (e.g., antigens) or cells.
- variant polypeptides differ from a native sequence by substitution, deletion or addition of five amino acids or fewer. Variants may also (or alternatively) be modified by, for example, the deletion or addition of amino acids that have minimal influence on the immunogenicity, secondary structure and hydropathic nature of the polypeptide
- Vector refers to a DNA or RNA molecule that comprises a polynucleotide sequence that encodes a peptide, a polypeptide or a protein.
- a vector generally contains regulatory elements capable of directing expression of the encoding polynucleotide sequence, also called transgene, in the cells into which the nucleic acid molecule is introduced.
- transgene refers to a polynucleotide that is introduced into a cell and is capable of being transcribed into RNA and optionally, translated and/or expressed under appropriate conditions. In certain aspects, it confers a desired property to a cell into which it was introduced, or otherwise leads to a desired technical effect, here typically a therapeutic effect.
- a transgene may contain sequence coding for one or more proteins or one or more fragments of proteins.
- Wild-type refers to a nucleic acid or polypeptide refers to a nucleic acid or a polypeptide that occurs in, or is produced by, a biological organism as that biological organism exists in nature.
- the invention relates to a mutated TAFA4 polypeptide or a variant thereof, comprising at least one cysteine substitution with any other amino acid residue.
- the invention relates to a mutated TAFA4 polypeptide or a variant thereof, comprising a cysteine substitution with any other amino acid residue at one or more of the following amino acid positions: amino acid position C25 and amino acid position C26, by reference to SEQ ID NO: 3.
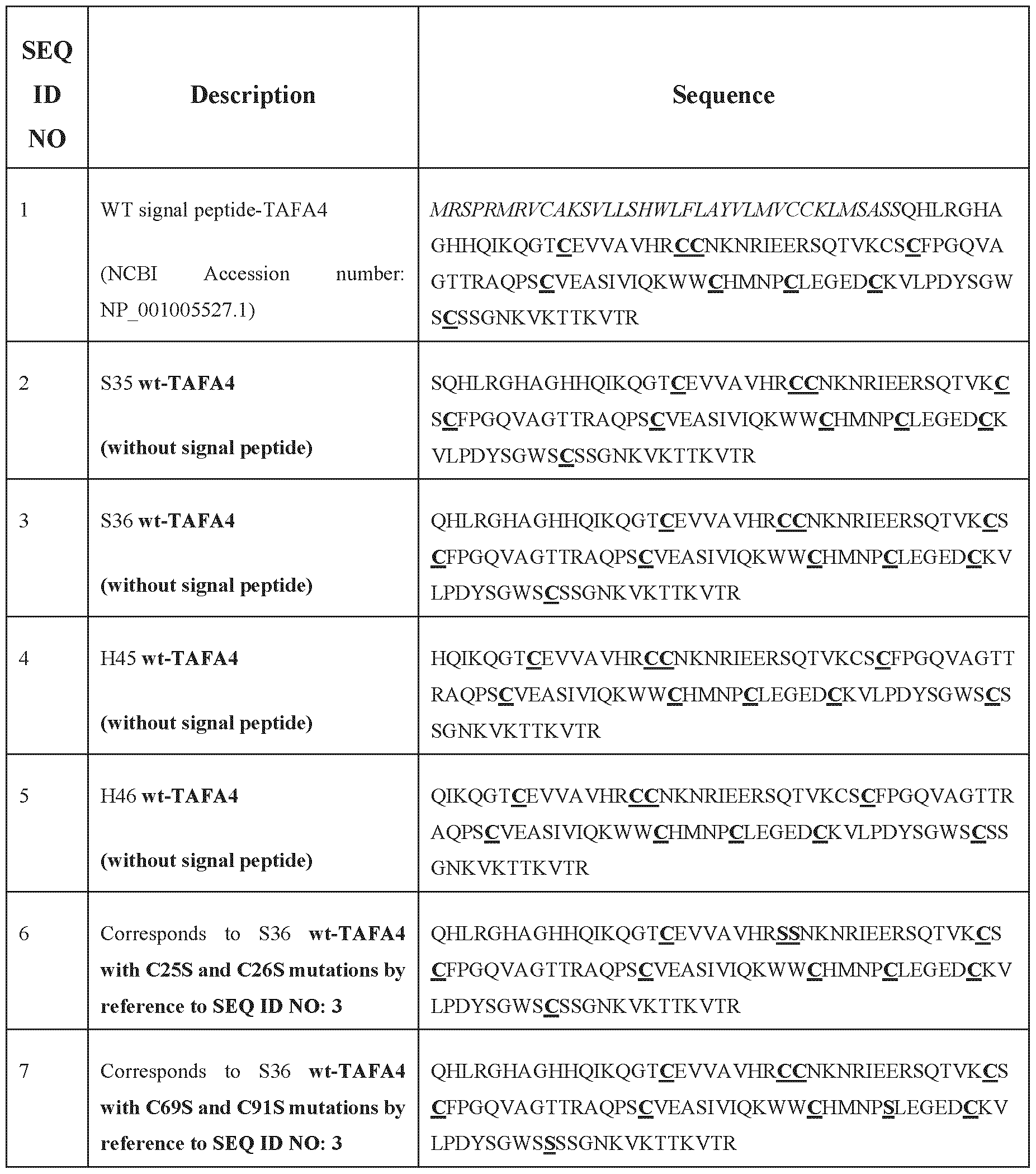
- TAFA4 or “TAFA4 polypeptide” designates a polypeptide belonging to the family of TAFA chemokine-like proteins, by reference to the amino acid sequence of SEQ ID NO: 3; which correspond to the human TAFA4 amino acid sequence without its predicted signal peptide sequence.
- the TAFA4 polypeptide can comprise a signal peptide allowing its secretion.
- the TAFA4 polypeptide without signal peptide corresponds to the secreted form of the polypeptide, such as TAFA4 comprising or consisting of SEQ ID NO: 3.
- the TAFA4 polypeptide is the wild-type form of human TAFA4 polypeptide or any variant thereof.
- the wild-type TAFA4 polypeptide is selected from the group comprising or consisting to the access number: NP_001005527.1 and XP 011531673.1.
- the TAFA4 polypeptide refers to proteins, polypeptides or peptides with the amino acid sequence of which is substantially identical (i.e., largely but not wholly identical) to the sequence of the protein, polypeptide, or peptide, e.g., at least about 80% identical or at least about 85% identical, e.g., preferably at least about 90% identical, e.g., at least 91% identical, 92% identical, more preferably at least about 93% identical, e.g., at least 94% identical, even more preferably at least about 95% identical, e.g., at least 96% identical, yet more preferably at least about 97% identical, e.g., at least 98% identical, and most preferably at least 99% identical to the sequence of the protein, polypeptide, or peptide, e.g., to the sequence of a corresponding TAFA4.
- the TAFA4 polypeptide may comprise a signal peptide which is cleaved, allowing to form an isoform.
- the TAFA4 polypeptide may be an isoform of said protein, polypeptide, or peptide.
- Isoforms are commonly used in the field and are well- known to the skilled artisan, who will know how to obtain all the isoforms of TAFA4, as described in Wang, et al., which published a cleavage of the signal sequence at position S36 by reference to SEQ ID NO: 1 (Cell Mol Immunol 2015, 12 (5), 615-624) (as in the sequence SEQ ID NO: 3).
- the TAFA4 polypeptide comprises a signal peptide which is cleaved at position selected from the group comprising or consisting of: S35, S36, H45 and H46 by reference to SEQ ID NO: 1.
- the TAFA4 polypeptide comprises a signal peptide which is cleaved at position S35 by reference to the SEQ ID NO: 1, and once cleaved, the TAFA4 polypeptide comprise or consists of the SEQ ID NO: 2.
- the TAFA4 polypeptide comprises a signal peptide which is cleaved at position S36 by reference to the SEQ ID NO: 1, and once cleaved, the TAFA4 polypeptide comprise or consists of the SEQ ID NO: 3.
- the TAFA4 polypeptide comprises a signal peptide which is cleaved at position H45 by reference to the SEQ ID NO: 1, and once cleaved, the TAFA4 polypeptide comprise or consists of the SEQ ID NO: 4.
- the TAFA4 polypeptide comprises a signal peptide which is cleaved at position H46 by reference to the SEQ ID NO: 1, and once cleaved, the TAFA4 polypeptide comprise or consists of the SEQ ID NO: 5.
- the TAFA4 polypeptide may be a variant thereof, i.e. may be conveniently denoted as "modified”, or as “mutated” or “mutant”, or as comprising one or more mutations, i.e., comprising one or more amino acid sequences changes compared to the amino acid sequence of TAFA4 that has not been so-mutated, such as, particularly, compared to the amino acid sequence of wild-type TAFA4.
- Said variant may comprise the same number of amino acids as any TAFA4 polypeptide defined above, more preferably as SEQ ID NO: 3, and thus the mutations and positions described herein are the same for the variant.
- said variant may comprise a different number of amino acids as SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or a variant thereof comprises one or more cysteine substitutions with any other amino acid residue.
- cysteine substitutions it is meant that the mutated TAFA4 polypeptide comprise at least one substituted cysteine residue, at least two substituted cysteines residue, at least three substituted cysteines residue, at least four substituted cysteines residue, at least five substituted cysteines residue, at least six substituted cysteines residue, at least seven substituted cysteines residue, at least eight substituted cysteines residue, at least nine substituted cysteines residue or at least ten substituted cysteines residue, by reference to SEQ ID NO:3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with any other amino acid residue, preferably a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue, which consists to residue 25 with any other amino acid residue, preferably a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 26 with any other amino acid residue, preferably a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue, which consists to residue 26 with any other amino acid residue, preferably a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with any other amino acid residue, preferably a serine residue (i.e., a C25S mutations), by reference to SEQ ID NO: 3; and a substitution of the cysteine residue 26 with any other amino acid residue, preferably a serine residue (i.e., a C26S mutations), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue, which consists to cysteine residue 25 with any other amino acid residue, preferably a serine residue (i.e., a C25S mutations), by reference to SEQ ID NO: 3; and a substitution of the cysteine residue, which consists to cysteine residue 26 with any other amino acid residue, preferably a serine residue (i.e., a C26S mutations), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises substitutions of the cysteine residues, said cysteine residues is selected from the group consisting of: residue 25, and residue 26, by reference to SEQ ID NO: 3.
- residue 25, and the residue 26, by reference to SEQ ID NO:3, are substituted by the same amino acid.
- residue 25, and the residue 26, by reference to SEQ ID NO:3, are substituted by two different amino acids.
- the amino acid substitution is a non-conservative amino acid substitution.
- the amino acid substituting the cysteine residue with a non-conservative substitution is selected from the group comprising or consisting of: aliphatic amino acids, aromatic amino acids, negatively charged amino acids, positively charged amino acids and special amino acid.
- the aliphatic amino acid is selected from the group consisting of: alanine, leucine, proline, valine, and isoleucine.
- the aromatic amino acid is selected from the group consisting of: tyrosine, tryptophane, and phenylalanine.
- the negatively charged amino acid is selected from the group consisting of: aspartic acid, and glutamic acid.
- the positively charged amino acid is selected from the group consisting of: arginine, histidine, and lysine.
- the mutated TAFA4 polypeptide or variant thereof comprises two substitutions of the cysteine residues, said cysteine residues are selected in the group consisting of: C25 and C26, by reference to SEQ ID NO:3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 74 with a serine residue (i.e., a C74S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 91 with a serine residue (i.e., a C91S mutation), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises one or more substitution of the cysteine residue 25, by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26, by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17, by reference to SEQ ID NO: 3, and/or a substitution of the cysteine residue 40, by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25, by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26, by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17, by reference to SEQ ID NO: 3, and/or a substitution of the cysteine residue 40, by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17 with a serine residue (i.e., a C17S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 40 with a serine residue (i.e., a C40S mutation), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 69 with a serine residue (i.e., a C69S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 80 with a serine residue (i.e., a C80S mutation), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17 with a serine residue (i.e., a C17S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 57 with a serine residue (i.e., a C57S mutation), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17 with a serine residue (i.e., a C17S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 57 with a serine residue (i.e., a C57S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 40 with a serine residue (i.e., a C40S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 42 with a serine residue (i.e., a C42S mutation), by reference to SEQ ID NO: 3.
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 69 with a serine residue (i.e., a C69S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 74 with a serine residue (i.e., a C74S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 80 with a serine residue (i.e., a C80S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 91 with a serine residue (i.e., a C91S mutation), by reference to SEQ ID NO:
- the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17 with a serine residue (i.e., a C17S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 57 with a serine residue (i.e., a C57S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 40 with a serine residue (i.e., a C40S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 42 with a serine residue (i.e., a C42S mutation), by reference to SEQ ID NO: 3,
- the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group comprising or consisting of SEQ ID NO: 6, SEQ ID NO: 7, SEQ
- SEQ ID NO: 58 SEQ ID NO: 59, SEQ ID NO: 60 and more preferably the SEQ ID NO: 6.
- the mutated TAFA4 polypeptide has the amino acid sequence selected from the group comprising or consisting of: SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, and more preferably the SEQ ID NO: 6.
- the (b) one or more mutated TAFA4 polypeptides or variants thereof comprising a cysteine substitution with an amino acid residue selected from the group consisting of serine (S), aspartic acid (D), tyrosine (Y), glutamic acid (E), arginine (R), glycine (G), and lysine (K), at amino acid position C25, by reference to SEQ ID NO: 3, and a cysteine substitution with an amino acid residue selected from the group consisting of serine (S), aspartic acid (D), tyrosine (Y), glutamic acid (E), arginine (R), glycine (G), and lysine (K), at amino acid position C26, by reference to SEQ ID NO: 3.
- the (b) one or more mutated TAFA4 polypeptides comprise:
- cysteine substitution at position C25 by reference to SEQ ID NO: 3 selected from the group consisting of C25S, C25D, C25Y, C25E, C25R, and C25G substitutions, preferably a C25S substitution, and
- the (b) one or more mutated TAFA4 polypeptides comprise: (i) a cysteine substitution at position C25 by reference to SEQ ID NO: 3 selected from the group consisting of C25S, C25D, C25Y, C25E, C25R, and C25G substitutions, preferably a C25S substitution, and
- cysteine substitution at position C26 by reference to SEQ ID NO: 3 selected from the group consisting of C26S, C26D, C26Y, C26E, C26R, and C26G, preferably a C26S substitution, wherein said (b) one or more mutated TAFA4 polypeptides do not comprise cysteine substitutions on positions other than C25, C26, Cl 7, and C40 by reference to SEQ ID NO: 3.
- the (b) one or more mutated TAFA4 polypeptides do not comprise cysteine substitutions on positions other than C25, C26, Cl 7, and C40 by reference to SEQ ID NO: 3.
- the (b) one or more mutated TAFA4 polypeptides comprise a K96R substitution, by reference to SEQ ID NO: 3.
- the (b) one or more mutated TAFA4 polypeptides comprise:
- cysteine substitution at position C25 by reference to SEQ ID NO: 3 selected from the group consisting of C25S, C25D, C25Y, C25E, C25R, and C25G substitutions, preferably a C25S substitution;
- the present invention thus relates to a fusion polypeptide, comprising:
- one or more mutated TAFA4 polypeptides comprise:
- cysteine substitution at position C25 by reference to SEQ ID NO: 3 selected from the group consisting of C25S, C25D, C25Y, C25E, C25R, and C25G substitutions, preferably a C25S substitution;
- the mutated TAFA4 polypeptide or a variant thereof further comprising a mutation with any other amino acid residue at one or more of the following amino acid positions: amino acid position K96 and amino acid position D85 by reference to SEQ ID NO: 3.
- these additional mutations preferably at the position K96 and D85, by reference to SEQ ID NO: 3, induce the enhancement in stability can be assessed by the increase in the protein's half-life, in particular by slowing down proteolysis.
- the one or more mutations at the amino acid position K96 and/or D85 by reference to SEQ ID NO: 3, correspond to a substitution.
- the mutation at the amino acid position K96 by reference to SEQ ID NO: 3 can change the susceptibility for protease cleavage.
- the mutation at the amino acid position K96 by reference to SEQ ID NO: 3 is a substitution into R (arginine) or into H (histidine), more preferably the mutated TAFA4 polypeptide or variant thereof comprises or consists of the sequence SEQ ID NO: 20 or SEQ ID NO: 58.
- the mutation at the amino acid position D85 by reference to SEQ ID NO: 3 can change the glycosylation side in order to improve yield and solubility, and avoid aggregation.
- the mutated TAFA4 polypeptide or a variant thereof, further comprising a mutation with any other amino acid residue comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group comprising or consisting of: SEQ ID NO: 20, SEQ ID NO: 21 and SEQ ID NO: 58.
- the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135.
- the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, and SEQ ID NO: 60.
- the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135.
- the mutated TAFA4 polypeptide does not have the amino acid sequence selected from the group consisting of SEQ ID NO: 83, SEQ ID NO: 84, SEQ ID NO: 85, SEQ ID NO: 86, SEQ ID NO: 107, SEQ ID NO: 108, SEQ ID NO: 109, SEQ ID NO: 110, SEQ ID NO: 111, SEQ ID NO: 112, and SEQ ID NO: 113.
- the mutated TAFA4 polypeptide comprises a signal peptide, such as the wild-type TAFA4 precursor polypeptide with the sequence of SEQ ID NO: 1, which corresponds to the human TAFA4 amino acid sequence with a signal peptide (NCBI Accession number: NP_001005527.1).
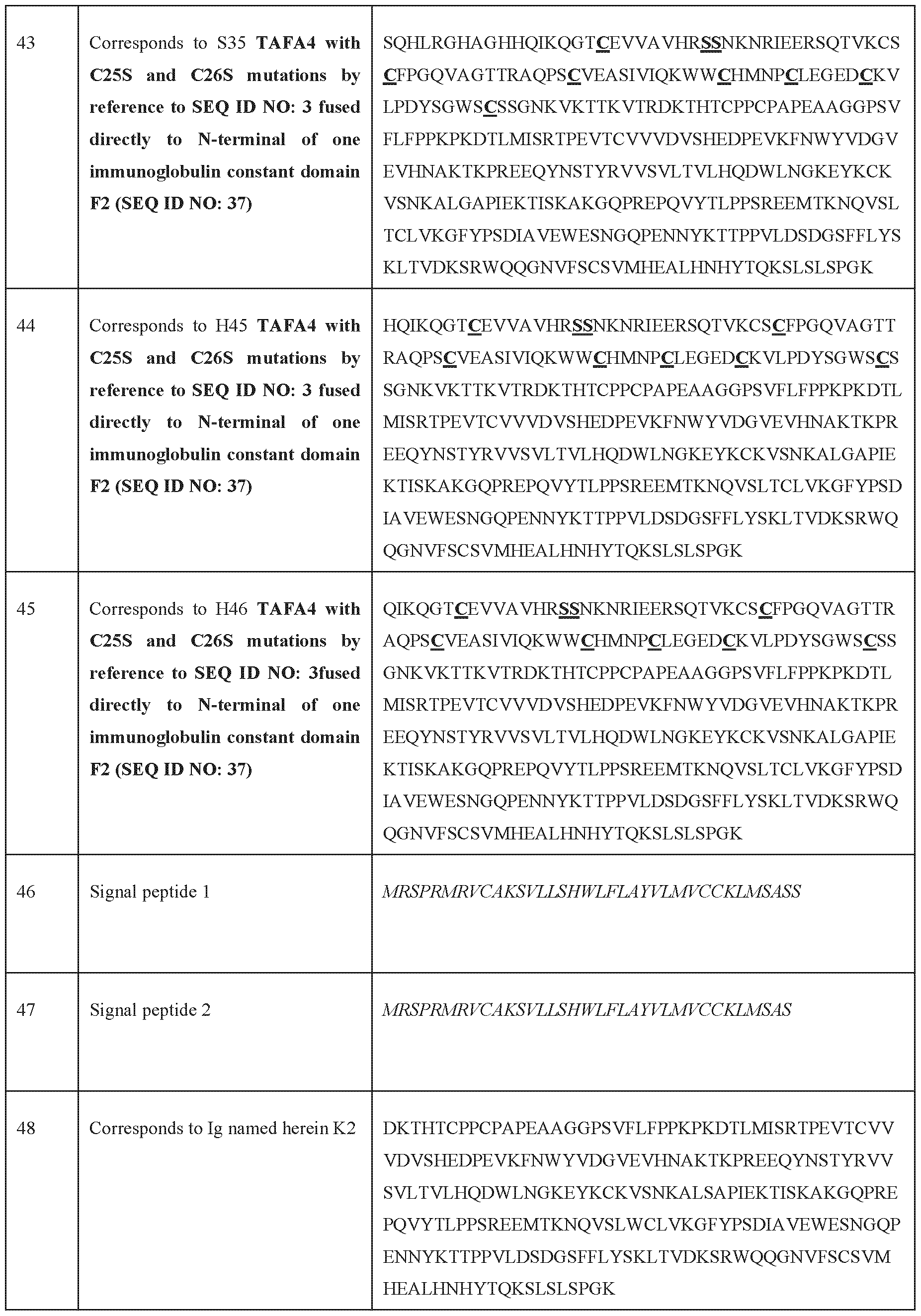
- the signal peptide in italic before and has the sequence of SEQ ID NO: 46, SEQ ID NO: 47, SEQ ID NO: 68 or SEQ ID NO: 69.
- SEQ ID NO 46 MRSPRMRVCAKSVLLSHWLFLAYVLMVCCKLMSASS
- SEQ ID NO 47 MRSPRMRVCAKSVLLSHWLFLAYVLMVCCKLMSAS [143] SEQ ID NO: 68:
- Signal peptides are commonly used in the field and are well-known to the skilled artisan, who will know how to choose a signal peptide.
- the TAFA4 polypeptide may contain additional amino acids, e. g., as a result of manipulations of the nucleic acid construct such as the addition of a restriction site, as long as these additional amino acids do not render the signal peptide or the TAFA4 polypeptide non-functional.
- the additional amino acids can be cleaved or can be retained by the mature polypeptide as long as retention does not result in a non-functional polypeptide.
- the signal peptide suitable in the context of the invention can be the signal peptide sequences of human TAFA4, the signal peptide sequences of mouse TAFA4, the signal peptide sequences of human Albumin, the signal peptide sequences of CPYm3, the signal peptide sequences of mouse IgG-H, the signal peptide sequences of Oncostatin-M, the signal peptide sequences of huIL-2, the signal peptide sequences of huIgG-kappa and the signal peptide sequences of muIgG-lambda.
- the invention further relates to a fusion polypeptide, comprising:
- the (a) one or more domains may be selected from the group comprising or consisting of an immunoglobulin constant domain or a fragment thereof, a serum albumin, and a linker.
- the (a) one or more domains may be selected from the group comprising or consisting of one domain, two domains, three domains, four domains, five domains, six domains.
- the (a) one or more domain are identical.
- the (b) one or more mutated TAFA4 polypeptides or variants thereof may be selected from the group comprising or consisting of two mutated TAFA4 polypeptides or variants thereof, three mutated TAFA4 polypeptides or variants thereof, four mutated TAFA4 polypeptides or variants thereof, five mutated TAFA4 polypeptides or variants thereof, six mutated TAFA4 polypeptides or variants thereof, preferably one mutated TAFA4 polypeptide or variant thereof.
- the second mutated TAFA4 polypeptide has the amino acid sequence comprising or consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135. and more preferably the SEQ ID NO: 6 depending on the first selected sequence.
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 114, and SEQ ID NO: 115.
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81.
- SEQ ID NO: 100 SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, and SEQ ID NO: 115.
- the fusion polypeptide comprises or consists of at least
- SEQ ID NO: 76 SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80,
- SEQ ID NO: 90 SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94,
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, and SEQ ID NO: 82.
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, , and SEQ ID NO: 115.
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, and SEQ ID NO: 106.
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80, preferably sequence selected from the group consisting of SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80.
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80.
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 78, and SEQ ID NO: 79.
- the fusion polypeptide has at least 90% sequence identity with the amino acid sequence of SEQ ID NO: 73.
- the fusion polypeptide has at least 90% sequence identity with the amino acid sequence of SEQ ID NO: 74.
- the fusion polypeptide has at least 90% sequence identity with the amino acid sequence of SEQ ID NO: 78.
- the fusion polypeptide has at least 90% sequence identity with the amino acid sequence of SEQ ID NO: 79.
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 114 and SEQ ID NO: 115. [180] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 40, SEQ ID NO: 42, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64, SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, and SEQ ID NO: 120.
- the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 40, SEQ ID NO: 42, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64, SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, and SEQ ID NO: 120.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64, SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64, SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 114, and SEQ ID NO: 115.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 114, and SEQ ID NO: 115.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, and SEQ ID NO: 82.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76,
- SEQ ID NO: 82 SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90,
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76,
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78,
- SEQ ID NO: 79 SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 87,
- SEQ ID NO: 98 SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, and SEQ ID NO: 106.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, and SEQ ID NO: 82.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, , and SEQ ID NO: 115.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, and SEQ ID NO: 106.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80, preferably sequence selected from the group consisting of SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76,
- SEQ ID NO: 77 SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 78, and SEQ ID NO: 79. In some embodiments, the fusion polypeptide has the amino acid sequence of SEQ ID NO: 73. In some embodiments, the fusion polypeptide has the amino acid sequence of SEQ ID NO: 74. In some embodiments, the fusion polypeptide has the amino acid sequence of SEQ ID NO: 78. In some embodiments, the fusion polypeptide has the amino acid sequence of SEQ ID NO: 79.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 114 and SEQ ID NO: 115.
- the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
- the (a) domain is selected from the group consisting of an immunoglobulin or parts thereof, i.e., immunoglobulin light chain, immunoglobulin heavy chain, immunoglobulin constant domain (or region), immunoglobulin crystallizable fragment (Fc), immunoglobulin hinge region, or any combination thereof.
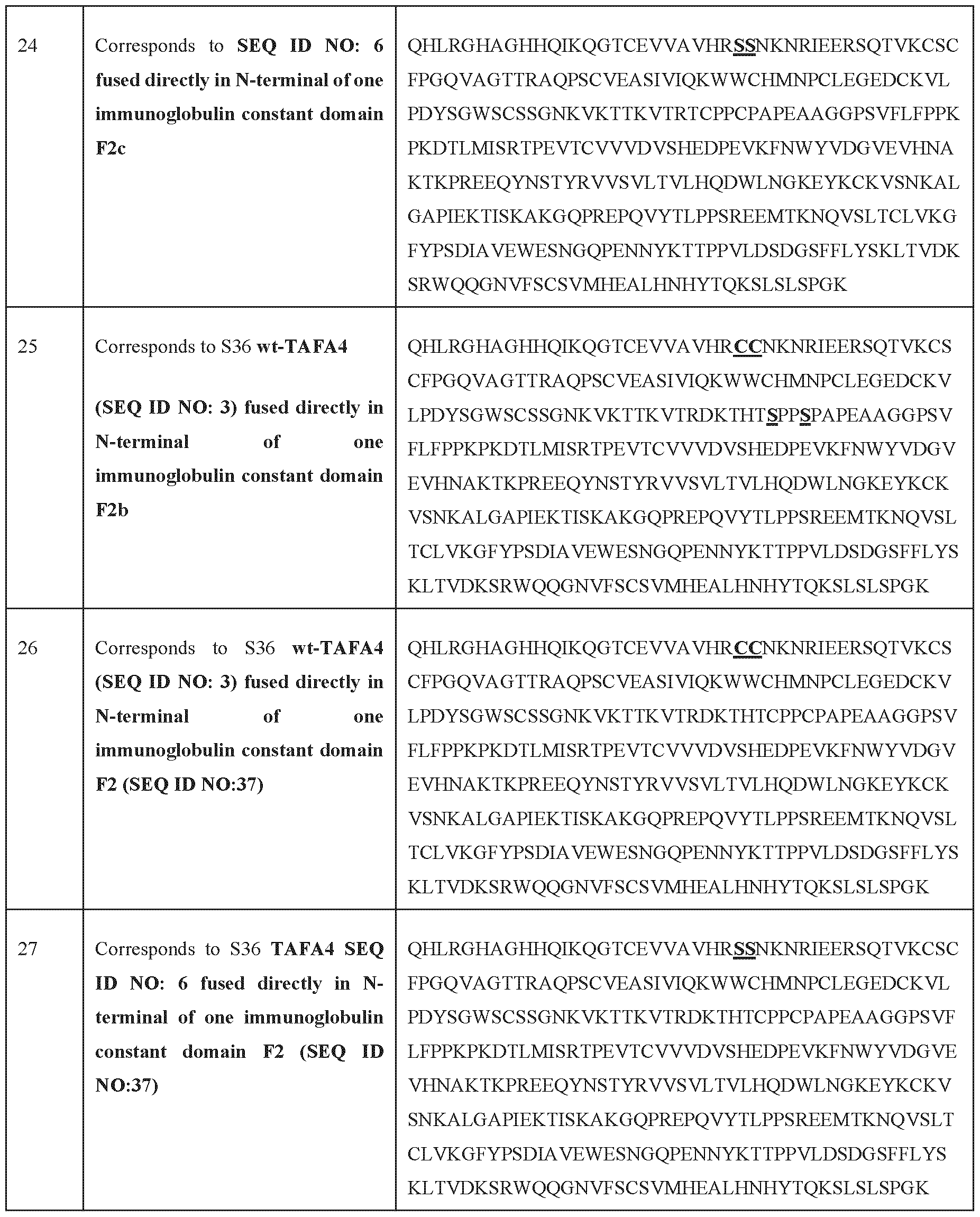
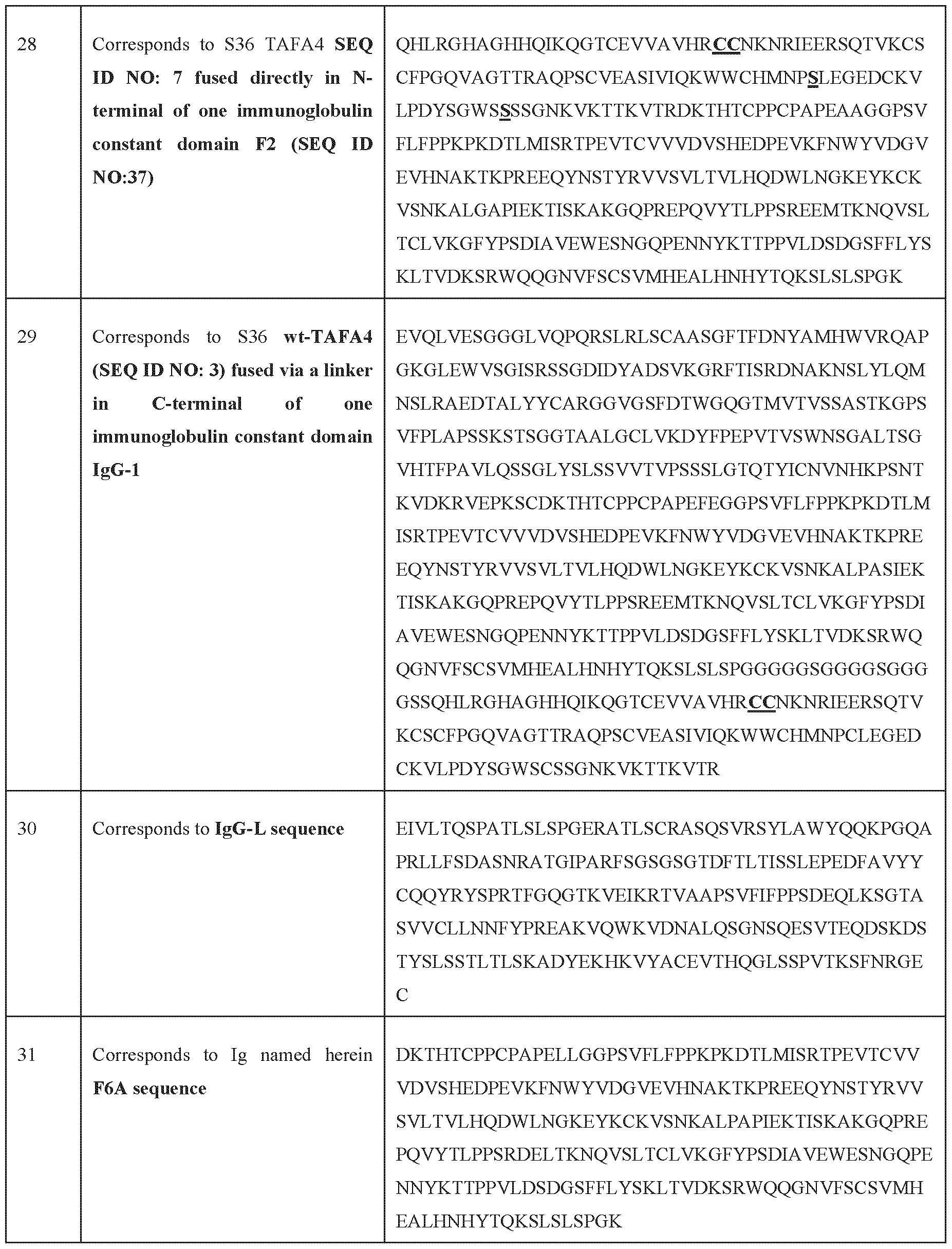
- immunoglobulins particularly constant domains of immunoglobulins (Fc regions, illustratively and not limiting as depicted in SEQ ID NO: 118; SEQ ID NO: 35; SEQ ID NO: 36; SEQ ID NO: 37; SEQ ID NO: 38, or SEQ ID NO: 39), into fusion proteins is a well-established technique in the field. This approach leverages, illustratively and non-limitatively, the stability, prolonged half-life, and effector functions conferred by the Fc region.
- the design and production of such fusion proteins are widely known and accessible to persons skilled in the art, with numerous methods available, including recombinant DNA technology, vector systems, and expression in mammalian, bacterial, or yeast systems.
- the use of Fc fusion proteins is extensively documented in scientific literature and supported by commercially available tools and reagents.
- the (a) domain is an immunoglobulin.
- the (a) domain is an immunoglobulin constant domain, or a fragment thereof.
- the fusion polypeptide comprises:
- the fusion polypeptide comprises:
- TAFA4 polypeptides or variants thereof selected from the group consisting of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 14, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135.
- the fusion polypeptide comprises or consists of from N terminal to C terminal: (a) an immunoglobulin constant domain or fragment thereof, (bl) a first mutated TAFA4 polypeptide or variant thereof and (b2) a second mutated TAFA4 polypeptide or variant thereof.
- (a) the immunoglobulin constant domain or fragment thereof, (bl) the first mutated TAFA4 polypeptide or variant thereof and (b2) the second mutated TAFA4 polypeptide or variant thereof may be directly or linked via a linker between each other.
- the (a) one or more immunoglobulin constant domains or fragments thereof are selected from the group comprising or consisting of the CH2 and CH3 domains of a Fc domain.
- the (a) one or more immunoglobulin constant domains or fragments thereof are selected from the group comprising or consisting of an IgA, IgD, IgE, IgG and IgM, more preferably the CH2 and CH3 domains of the human IgG Fc domain.
- the one or more mutated TAFA4 polypeptides or variants thereof and/or (c) the wild-type TAFA4 polypeptide or a variant thereof are fused directly at the N-terminal of (a) the one or more immunoglobulin constant domains or fragments thereof, more preferably to the heavy chain of the immunoglobulin chain.
- FcT4 SEQ ID NO: 27 FcT4 SEQ ID NO: 26 and rT4 SEQ ID NO: 3 reduce the production of the inflammatory cytokines TNFa and IL-6 by proinflammatory Ml macrophages (FIGURES 6-A and 6-B).
- FcT4 SEQ ID NO: 27 was more efficient than FcT4 SEQ ID NO: 26 and rT4 SEQ ID NO: 3 in reducing the production of IL-12 (p40 or p70) by these macrophages (FIGURES 6-C and 6-D).
Landscapes
- Chemical & Material Sciences (AREA)
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- General Health & Medical Sciences (AREA)
- Gastroenterology & Hepatology (AREA)
- Biochemistry (AREA)
- Biophysics (AREA)
- Zoology (AREA)
- Genetics & Genomics (AREA)
- Medicinal Chemistry (AREA)
- Molecular Biology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Toxicology (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
- Peptides Or Proteins (AREA)
Abstract
PROTEIN FOR IMMUNE REGULATION The present invention relates to a relates to TAFA4 polypeptide, fusion protein, pharmaceutical composition and methods for treating inflammatory disease.
Description
PROTEIN FOR IMMUNE REGULATION
FIELD OF INVENTION
[1] The present invention relates to TAFA4 polypeptide, fusion protein, pharmaceutical composition and methods for treating inflammatory disease.
BACKGROUND OF INVENTION
[2] The skin serves as an important boundary between the internal milieu and the environment, preventing contact with potentially harmful antigens. In the case of antigen/pathogen penetration, an inflammatory response is induced to eliminate the antigen. This response leads to a dermal infiltrate that consists predominantly of T cells, polymophonuclear cells, and macrophages. Normally, this inflammatory response, triggered by the pathogen, is under tight control and will be halted upon elimination of the pathogen. In certain cases, the inflammatory response occurs in absence of pathogen. For instance, UV radiation causes sunburn-like damage characterized by the destruction of the epidermis and inflammation of the underneath dermal papilla. These events lead to the rapid activation of mechanisms orchestrated by resident dermal macrophages and infiltrating monocytes to resolve inflammation and promote tissue repair. Excessive inflammation can however lead to chronic tissue damage followed by excessive collagen deposition resulting in unresolved fibrotic scars. Tissue-specific signals constantly shape resident macrophage functional identity and promote their maintenance throughout the lifespan, either by local self-renewal or by the recruitment of additional monocyte-derived cells.
[3] In the last years, it has been discovered that the protein TAFA4 might be a candidate for treatment of chronic pain, such as inflammatory disease. Especially, it has been discovered that TAFA4 polypeptide can be used for reducing skin inflammation in a subject in need thereof (W02020064907). However, there is a need to optimize the
activity and efficiency of such polypeptide so as to improve the therapeutic effect and/or the productivity.
[4] TAFA4 belongs to the TAFA family, which consists of five closely related genes responsible for producing small secreted proteins. These proteins comprise several cysteine residues at specific locations.
[5] TAFA4 is considered as a chemokine-like protein with potential implications in myeloid cell recruitment or activation in living organisms, particularly in the context of the skin inflammation.
[6] It is known that TAFA4 protein conformation plays a key role in the catalytic activity. In this context, the inventors have developed a mutated TAFA4 polypeptide including one or more substitutions of cysteine residue, leading to surprising enhanced activity and high potency of the TAFA4 polypeptide, especially when compared to the wild-type (WT) polypeptide.
[7] Moreover, the production of TAFA4 polypeptide is a complex endeavor due to its high cysteine content (10 cysteines in total). It is therefore quite challenging to develop new polypeptide being easily produced but maintaining/improving activity so as to improve the therapeutic efficacy.
[8] In this context, the inventors have successfully developed a TAFA4 polypeptide that enhances production efficiency, notably by reducing the occurrence of aggregates.
SUMMARY
[9] This invention thus relates to a fusion polypeptide, comprising:
(a) one or more domains, and
(b) one or more mutated TAFA4 polypeptides or variants thereof, comprising a cysteine substitution with any other amino acid residue at one or more of the following
amino acid positions: amino acid position C25 and amino acid position C26, by reference to SEQ ID NO: 3.
[10] In some embodiments, the cysteine substitution at position C25 of the (b) one or more mutated TAFA4 polypeptides is a substitution of the cysteine residue by a serine residue by reference to SEQ ID NO: 3, such as a C25S substitution.
[11] In some embodiments, the cysteine substitution at position C26 of the (b) one or more mutated TAFA4 polypeptides is a substitution of the cysteine residue by a serine residue by reference to SEQ ID NO: 3, such as a C26S substitution.
[12] In some embodiments, the one or more mutated TAFA4 polypeptide comprise a cysteine substitution with any other amino acid residue at the amino acid position C25 by reference to SEQ ID NO: 3, preferably a C25S substitution, and a cysteine substitution with any other amino acid residue amino acid position C26 by reference to SEQ ID NO: 3, preferably a C26S substitution.
[13] In some less preferred embodiments, the (b) one or more mutated TAFA4 polypeptides further comprise a substitution, such as a cysteine substitution, with any other amino acid residue at one or more of the following positions: amino acid position Cl 7, amino acid position C25, amino acid C26, amino acid position C40, amino acid position C42, amino acid position C57, amino acid position C69, amino acid position C74, amino acid position C80 and amino acid position C91 by reference to SEQ ID NO: 3.
[14] In some embodiments, the (b) one or more mutated TAFA4 polypeptides comprise at least 90% sequence identity to the amino acid sequences selected from the group comprising or consisting of: SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, more preferably the SEQ ID NO: 6.
[15] In some embodiments, the (b) one or more mutated TAFA4 polypeptides are fused directly or via a linker at the N-terminal or the C-terminal of the (a) one or more domains.
[16] In some embodiments, the (a) one or more domains correspond to one or more immunoglobulin constant domains or fragments thereof, preferably said one or more immunoglobulin constant domains or fragments thereof comprise the CH2 and CH3 domains of a Fc domain.
[17] In some embodiments, the (a) one or more immunoglobulin constant domains or fragments thereof are selected from the group comprising or consisting of IgA, IgD, IgE, IgG and IgM, preferably IgG.
[18] The invention further relates to a fusion protein comprising a plurality of the fusion polypeptides according to the invention.
[19] The invention further relates to a nucleic acid sequence encoding the fusion polypeptide according to the invention or the fusion protein according to the invention.
[20] The invention further relates to a vector comprising a nucleic acid sequence encoding the fusion polypeptide or the fusion protein according to the invention.
[21] The invention further relates to a cell or cell population comprising one or more fusion polypeptides according to the invention, or one or more fusion proteins according to the invention, or one or more nucleic acid sequences according to the invention, or one or more vectors according to the invention.
[22] The invention further relates to a pharmaceutical composition comprising one or more fusion polypeptides according to the invention, a fusion protein according to the invention, one or more nucleic acid sequences according to the invention, one or more vectors according to the invention, or one or more cells or cell population according to the invention; and one or more pharmaceutically acceptable excipients or carriers.
[23] The invention further relates to the fusion polypeptide according to the invention, the fusion protein according to the invention, the nucleic acid sequence according to the invention, the vector according to the invention, the cell or the cell population the invention or the pharmaceutical composition according to the invention, for use as a medicament.
[24] The invention further relates to the fusion polypeptide according to the invention, the fusion protein according to the invention, the nucleic acid sequence according to the invention, the vector according to the invention, the cell or the cell population according to the invention or the pharmaceutical composition according to the invention, for use for treating disease associated with the modulation of macrophage in a subject in need thereof
[25] In some embodiments, the disease is an inflammation disease, more preferably a skin inflammation disease.
[26] In some embodiments, the pharmaceutical composition is formulated for topical, intramuscular, subcutaneous, intradermal, intravenous or oral administration, preferably the topical administration is accomplished via a transdermal device or patch device.
DEFINITIONS
[27] In the present invention, the following terms have the following meanings:
[28] “About” refers to mean approximately, roughly, around, or in the region of. The term “about” preceding a figure means more or less 10 % of the value of the figure. When the term “about” is used in conjunction with a numerical range, it modifies that range by extending the boundaries above and below the numerical values set forth by 10%.
[29] “Administration", or a variant thereof e.g., “administering”), refers to providing a therapeutic agent (e.g., a compound of the invention) alone or as part of a
pharmaceutically acceptable composition, to for example a patient in whom/which the condition, symptom, or disease is to be treated.
[30] “Amino acid substitution” is a substitution of one amino acid for another. A conservative amino acid substitution is a substitution of one amino acid for another with similar characteristics. Conservative amino acid substitutions include substitutions within the following groups: valine, alanine and glycine; leucine, valine, and isoleucine; aspartic acid and glutamic acid; asparagine and glutamine; serine, cysteine, and threonine; lysine and arginine; and phenylalanine and tyrosine. The nonpolar hydrophobic amino acids include alanine, leucine, isoleucine, valine, proline, phenylalanine, tryptophan and methionine. The polar neutral amino acids include glycine, serine, threonine, cysteine, tyrosine, asparagine and glutamine. The positively charged (i.e., basic) amino acids include arginine, lysine and histidine. The negatively charged (i.e., acidic) amino acids include aspartic acid and glutamic acid. Any substitution of one member of the above- mentioned polar, basic, or acidic groups by another member of the same group can be deemed a conservative substitution. By contrast, a non-conservative substitution is a substitution of one amino acid for another with dissimilar characteristics.
[31] “Antibody” and “immunoglobulin”, as used herein, may be used interchangeably and refer to a protein having a combination of two heavy and two light chains whether or not it possesses any relevant specific immunoreactivity. “Antibodies” refers to such assemblies which have significant known specific immunoreactive activity to an antigen of interest. Antibodies and immunoglobulins comprise light and heavy chains, with or without an interchain covalent linkage between them. Basic immunoglobulin structures in vertebrate systems are relatively well understood. The generic term “immunoglobulin” comprises five distinct classes of antibody that can be distinguished biochemically. Although the following discussion will generally be directed to the IgG class of immunoglobulin molecules, all five classes of antibodies are within the scope of the present invention. With regard to IgG, immunoglobulins comprise two identical light polypeptide chains of molecular weight of about 23 kDa, and two identical heavy chains of molecular weight of about 53-70 kDa. The four chains are joined by disulfide bonds in a “Y” configuration wherein the light chains bracket the heavy
chains starting at the mouth of the “Y” and continuing through the variable region. The light chains of an antibody are classified as either kappa (K) or lambda (X). Each heavy chain class may be bonded with either a K or X, light chain. In general, the light and heavy chains are covalently bonded to each other, and the “tail” regions of the two heavy chains are bonded to each other by covalent disulfide linkages or non-covalent linkages when the immunoglobulins are generated either by hybridomas, B cells or genetically engineered host cells. In the heavy chain, the amino acid sequences run from an N- terminus at the forked ends of the Y configuration to the C -terminus at the bottom of each chain. Those skilled in the art will appreciate that heavy chains are classified as gamma (y), mu (p), alpha (a), delta (5) or epsilon (e) with some subclasses among them (e.g., yl- y4). It is the nature of this chain that determines the “class” of the antibody as IgG, IgM, IgA IgD or IgE, respectively. The immunoglobulin subclasses or “isotypes” e.g., IgGl, IgG2, IgG3, IgG4, IgAl, etc.) are well characterized and are known to confer functional specialization. Modified versions of each of these classes and isotypes are readily discernable to the skilled artisan in view of the instant disclosure and, accordingly, are within the scope of the present invention. As indicated above, the variable region of an antibody allows the antibody to selectively recognize and specifically bind epitopes on antigens. That is, the light chain variable domain (VL domain) and heavy chain variable domain (VH domain) of an antibody combine to form the variable region that defines a three-dimensional antigen binding site. This quaternary antibody structure forms the antigen binding site presents at the end of each arm of the “Y”. More specifically, the antigen binding site is defined by three complementarity determining regions (CDRs) on each of the VH and VL chains.
[32] “Combined treatment”, “combined preparation”, “combined therapy” or “therapy combination” refer to a treatment that uses more than one medication. The combined therapy may be dual therapy or bi-therapy. The medications used in the combined treatment according to the invention are administered to the subject simultaneously, separately or sequentially. As used herein, the term “administration simultaneously” refers to administration of 2 active ingredients by the same route and at the same time or at substantially the same time. The term “administration separately” refers to an administration of 2 active ingredients at the same time or at substantially the
same time by different routes. The term “administration sequentially” refers to an administration of 2 active ingredients at different times, the administration route being identical or different.
[33] “Comprise” or a variant thereof (e.g., “comprises”, “comprising”) refers to according to common patent application drafting terminology. Hence, “comprise” preceded by an object and followed by a constituent means that the presence of a constituent in the object is required (typically as a component of a composition), but without excluding the presence of any further constituent(s) in the object. Moreover, any occurrence of “comprise” or a variant thereof herein also encompasses narrower expression “substantially consist of’, further narrower expression “consist of’ and any variants thereof (e.g., “consists of’, “consisting of’), unless otherwise stated.
[34] “Control sequences” refers collectively to promoter sequences, polyadenylation signals, transcription termination sequences, upstream regulatory domains, origins of replication, internal ribosome entry sites ("IRES"), enhancers, and the like, which collectively provide for the replication, transcription and translation of a coding sequence in a recipient cell. Not all of these control sequences need always be present so long as the selected coding sequence is capable of being replicated, transcribed and translated in an appropriate host cell.
[35] “Expression cassette” refers to a unit cassette capable of expressing a gene of interest, as the transgene, operably linked to a downstream side of a promoter, including a promoter and a gene of interest. Various factors that can help the efficient expression of the gene of interest may be included inside or outside such a gene expression cassette. The gene expression cassette may include, but is not limited to, a transcription termination signal, a ribosome binding site, and a translation termination signal in addition to a promoter operably linked to the gene of interest.
[36] “Homology” or “identity” refers to the subunit sequence identity between two polymeric molecules, e.g., between two nucleic acid molecules, such as, two DNA molecules or two RNA molecules, or between two polypeptide molecules. When a subunit position in both of the two molecules is occupied by the same monomeric subunit;
e.g., if a position in each of two DNA molecules is occupied by adenine, then they are homologous or identical at that position. The homology between two sequences is a direct function of the number of matching or homologous positions; e.g., if half (e.g., five positions in a polymer ten subunits in length) of the positions in two sequences are homologous, the two sequences are 50% homologous; if 90% of the positions (e.g., 9 of 10), are matched or homologous, the two sequences are 90% homologous. Thus, the term "homologous" or "identical", when used in a relationship between the sequences of two or more polypeptides or of two or more nucleic acid molecules, refers to the degree of sequence relatedness between polypeptides or nucleic acid molecules, as determined by the number of matches between strings of two or more amino acid or nucleotide residues. “Identity” measures the percent of identical matches between the smaller of two or more sequences with gap alignments (if any) addressed by a particular mathematical model or computer program (i.e., "algorithms"). Identity of related polypeptides can be readily calculated by known methods. Such methods include, but are not limited to, those described in Computational Molecular Biology, Lesk, A. M., ed., Oxford University Press, New York, 1988; Biocomputing: Informatics and Genome Projects, Smith, D. W., ed., Academic Press, New York, 1993; Computer Analysis of Sequence Data, Part 1, Griffin, A. M., and Griffin, H. G., eds., Humana Press, New Jersey, 1994; Sequence Analysis in Molecular Biology, von Heinje, G., Academic Press, 1987; Sequence Analysis Primer, Gribskov, M. and Devereux, J., eds., M. Stockton Press, New York, 1991; and Carillo et al., SIAM J. Applied Math. 48, 1073 (1988). Preferred methods for determining identity are designed to give the largest match between the sequences tested. Methods of determining identity are described in publicly available computer programs. Preferred computer program methods for determining identity between two sequences include the GCG program package, including GAP (Devereux et al., Nucl. Acid. Res. \2, 387 (1984); Genetics Computer Group, University of Wisconsin, Madison, Wis.), BLASTP, BLASTN, and FASTA (Altschul et al., J. Mol. Biol. 215, 403-410 (1990)). The BLASTX program is publicly available from the National Center for Biotechnology Information (NCBI) and other sources (BLAST Manual, Altschul et al. NCB/NLM/NIH Bethesda, Md. 20894; Altschul etal., supra). The well-known Smith Waterman algorithm may also be used to determine identity.
[37] “Immunoglobulin” includes a protein having a combination of two heavy and two light chains whether or not it possesses any relevant specific immunoreactivity
[38] “Inflammatory disease” is used herein in the broadest sense and includes all diseases and pathological conditions having etiologies associated with a systemic or local abnormal and/or uncontrolled inflammatory response. For instance, over-expression of proinflammatory cytokines without proper controls leads to a variety of inflammatory diseases and disorders. This term includes both acute inflammatory diseases and chronic inflammatory diseases.
[39] ’’Linker’’ refers to a sequence of at least one amino acid that can link the TAFA4 polypeptide with the immunoglobulin chain or a fragment thereof. Linkers are well known to one of ordinary skill in the art and typically comprises 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20 or more amino acids. It can refer to a single covalent bond or a moiety comprising series of stable covalent bonds, the moiety often incorporating 1-40 plural valent atoms selected from the group consisting of C, N, O, S and P, that covalently attach a reactive group or bioactive group to the probe of the invention. The number of plural valent atoms in a linker may be, for example, 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 25, 30 or a larger number up to 40 or more. A linker may be linear or non-linear; some linkers have pendant side chains or pendant functional groups (or both). Examples of such pendant moieties are hydrophilicity modifiers, for example solubilizing groups like, e.g. sulfo (-SO3H or -SO3-), carboxy (-COOH or -COO-), hydroxy. In one embodiment, the linker is composed of any combination of single, double, triple or aromatic carbon-carbon bonds, carbon-nitrogen bonds, nitrogen-nitrogen bonds, carbon-oxygen bonds and carbon-sulfur bonds. Linkers may by way of example consist of a combination of moieties selected from alkyl, -C(O)NH-, -C(O)O-, -NH-, -S- , -O-, -C(O) -, -S(O)n- where n is 0, 1 or 2; -O-, 5- or 6- membered monocyclic rings and optional pendant functional groups, for example sulfo, hydroxy and carboxy.
[40] The term "hinge" or "hinge region" or "hinge domain" refers, in antibodies, to the flexible portion of a heavy chain located between the CHI domain and the CH2 domain. It is approximately 25 amino acids long, and is divided into an "upper hinge," a
"middle hinge" or "core hinge," and a "lower hinge." A "hinge subdomain" refers to the upper hinge, middle (or core) hinge or the lower hinge.
[41] "Mammal" refers to any animal classified as a mammal, including humans, other higher primates, domestic and farm animals, and zoo, sports, or pet animals, such as dogs, cats, cattle, horses, sheep, pigs, goats, rabbits, mouse, rat, etc. Preferably, the mammal is human.
[42] “Nucleic acid” or “Polynucleotide” refer to a polymeric form of nucleotides of any length, either ribonucleotides or deoxyribonucleotides. Thus, these terms include, but are not limited to, single-, double- or multi-stranded DNA or RNA, genomic DNA, cDNA, DNA-RNA hybrids, or a polymer comprising purine and pyrimidine bases, or other natural, chemically or biochemically modified, non-natural, or derivatized nucleotide bases. The backbone of the polynucleotide can comprise sugars and phosphate groups (as may typically be found in RNA or DNA), or modified or substituted sugar or phosphate groups. Alternatively, the backbone of the polynucleotide can comprise a polymer of synthetic subunits such as phosphoramidates and thus can be an oligodeoxynucleoside phosphoramidate (P-NH2) or a mixed phosphoramidate- phosphodiester oligomer. In addition, a double -stranded polynucleotide can be obtained from the single stranded polynucleotide product of chemical synthesis either by synthesizing the complementary strand and annealing the strands under appropriate conditions, or by synthesizing the complementary strand de novo using a DNA polymerase with an appropriate primer.
[43] "Operably linked" refers to functional linkage between a regulatory sequence and a heterologous nucleic acid sequence resulting in expression of the latter. For example, a first nucleic acid sequence is operably linked with a second nucleic acid sequence when the first nucleic acid sequence is placed in a functional relationship with the second nucleic acid sequence. For instance, a promoter is operably linked to a coding sequence if the promoter affects the transcription or expression of the coding sequence. Generally, operably linked DNA sequences are contiguous and, where necessary to join two protein coding regions, in the same reading frame.
[44] “Patient” refers to a subject who/which is awaiting the receipt of, or is receiving medical care or was/is/will be the object of a medical procedure, or is monitored for the development of the targeted disease or condition, such as, for example, an infectious disease.
[45] "Peptide," "Polypeptide," and "Protein" are used interchangeably, and refer to a compound comprised of amino acid residues covalently linked by peptide bonds. A protein or peptide must contain at least two amino acids, and no limitation is placed on the maximum number of amino acids that can comprise a protein's or peptide's sequence. Polypeptides include any peptide or protein comprising two or more amino acids joined to each other by peptide bonds. As used herein, the term refers to both short chains, which also commonly are referred to in the art as peptides, oligopeptides and oligomers, for example, and to longer chains, which generally are referred to in the art as proteins, of which there are many types. "Polypeptides" include, for example, biologically active fragments, substantially homologous polypeptides, oligopeptides, homodimers, heterodimers, variants of polypeptides, modified polypeptides, derivatives, analogs, fusion proteins, among others. The polypeptides include natural peptides, recombinant peptides, synthetic peptides, or a combination thereof. Proteinogenic amino acids comprise Alanine (A), Arginine (R), Asparagine (N), Aspartic acid (D), Cysteine (C), Glutamine (Q), Glutamic acid (E), Glycine (G), Histidine (H), Isoleucine (I), Leucine (L), Lysine (K), Methionine (M), Phenylalanine (F), Proline (P), Serine (S), Threonine (T), Tryptophan (W), Tyrosine (Y), Valine (V).
[46] “Pharmaceutically acceptable” means that the ingredients of a composition are compatible with each other and not deleterious to the patient to which/whom it is administered.
[47] “Pharmaceutically acceptable carrier” refers to an excipient that does not produce an adverse, allergic or other untoward reaction when administered to an animal, preferably a human. It includes any and all solvents, dispersion media, coatings, antibacterial and antifungal agents, isotonic and absorption delaying agents and the like. For human administration, preparations should meet sterility, pyrogenicity, general safety and purity standards as required by regulatory offices, such as, e.g., FDA Office or EMA.
Examples of pharmaceutically acceptable carriers include, but are not limited to, ion exchangers, alumina, aluminum stearate, lecithin, serum proteins, such as human serum albumin, buffer substances such as phosphates, glycine, sorbic acid, potassium sorbate, partial glyceride mixtures of saturated vegetable fatty acids, water, salts or electrolytes, such as protamine sulfate, di sodium hydrogen phosphate, potassium hydrogen phosphate, sodium chloride, zinc salts, colloidal silica, magnesium trisilicate, polyvinyl pyrrolidone, cellulose-based substances (for example sodium carboxymethylcellulose), polyethylene glycol, polyacrylates, waxes, polyethylene- polyoxypropylene- block polymers, polyethylene glycol and wool fat.
[48] “Pharmaceutical composition” refers to a composition comprising an active agent in association with a pharmaceutically acceptable vehicle or excipient. A pharmaceutical composition is for therapeutic use, and relates to health. Especially, a pharmaceutical composition may be indicated for treating or preventing a disease.
[49] "Polynucleotide" refers to a chain of nucleotides. Furthermore, nucleic acids are polymers of nucleotides. Thus, nucleic acids and polynucleotides as used herein are interchangeable. One skilled in the art has the general knowledge that nucleic acids are polynucleotides, which can be hydrolyzed into the monomeric "nucleotides." The monomeric nucleotides can be hydrolyzed into nucleosides. As used herein polynucleotides include, but are not limited to, all nucleic acid sequences which are obtained by any means available in the art, including, without limitation, recombinant means, i.e. , the cloning of nucleic acid sequences from a recombinant library or a cell genome, using ordinary cloning technology and PCR, and the like, and by synthetic means.
[50] "Promoter" sequence, which is used herein in its ordinary sense to refer to a nucleotide region comprising a DNA regulatory sequence, wherein the regulatory sequence is derived from a gene which is capable of binding RNA polymerase and initiating transcription of a downstream (3 '-direction) coding sequence. Transcription promoters can include "inducible promoters" (where expression of a polynucleotide sequence operably linked to the promoter is induced by an analyte, cofactor, regulatory protein, etc.), "repressible promoters" (where expression of a polynucleotide sequence
operably linked to the promoter is induced by an analyte, cofactor, regulatory protein, etc.), and "constitutive promoters”.
[51] The terms "The core of signal sequence" or “signal sequence”, or “signal peptide” refers to a long stretch of hydrophobic amino acids (about 5-16 residues long) that has a tendency to form a single alpha-helix and is also referred to as the "h-region". In addition, signal peptides may begin with a short positively charged stretch of amino acids, which may help to enforce proper topology of the polypeptide during translocation by what is known as the positive-inside rule. Because of its close location to the N- terminus it is called the "n-region". At the end of the signal peptide there may be typically a stretch of amino acids that may be recognized and cleaved by signal peptidase and therefore named cleavage site. In this case, the signal peptidase may cleave either during or after completion of translocation to generate a free signal peptide and a mature protein. The free signal peptides can be then digested by specific proteases. In Eukaryotes signal sequences may act via the co-translational pathway, which is initiated when the signal peptide emerges from the ribosome and is recognized by the signal-recognition particle (SRP). The SRP bound to the signal sequence then halts further translation (translational arrest) and directs the signal sequence-ribosome-mRNA complex to the SRP receptor, which is present on the surface of the ER. Once membrane-targeting is completed, the signal sequence used to be is inserted into the translocon. Ribosomes are then physically docked onto the cytoplasmic face of the translocon and protein synthesis resumes.
[52] “Selected from” refers to according to common patent application drafting terminology, to introduce a list of elements among which one or more item(s) is (are) selected. Any occurrence of “selected from” in the specification may be replaced by “selected from the group comprising or consisting of’ and reciprocally without changing the meaning thereof.
[53] The term "Silenced Fc” (Fc-silent) refers to a genetically engineered Fc domain comprising mutations that abrogate binding of the Fc domain to Fc receptors (FcyR, FcR) while maintain the ability of the Fc domain to binds to neonatal Fc receptor (FcRn). Such silenced Fc exhibits extended half-life as described in Borrok M.J, et al. J Pharm Sci. 2017.
[54] “Therapeutic agent”, “active pharmaceutical ingredient” and “active ingredient” refer to a compound for therapeutic use and relating to health. Especially, a therapeutic agent (e.g., a compound of the invention) may be indicated for treating a disease. An active ingredient may also be indicated for improving the therapeutic activity of another therapeutic agent. It describes a molecule or a substance, preferably a biological molecule such as for example an oligonucleotide, a siRNA, a miRNA, a DNA fragment, an aptamer, an antibody and the like, or a chemical entity, whose administration to a subject slows down or stops the progression, aggravation, or deterioration of one or more symptoms of a disease, or condition; alleviates the symptoms of a disease or condition; cures a disease or condition.
[55] “Therapeutically effective amount” refers to level or amount of agent that is aimed at, without causing significant negative or adverse side effects to the target, (1) delaying or preventing the onset of the targeted pathologic condition or disorder; (2) slowing down or stopping the progression, aggravation, or deterioration of one or more symptoms of the targeted pathologic condition or disorder; (3) bringing about ameliorations of the symptoms of the targeted pathologic condition or disorder; (4) reducing the severity or incidence of the targeted pathologic condition or disorder; (5) curing the targeted pathologic condition or disorder. An effective amount may be administered prior to the onset of the targeted pathologic condition or disorder, for a prophylactic or preventive action. Alternatively, or additionally, the effective amount may be administered after initiation of the targeted pathologic condition or disorder, for a therapeutic action.
[56] “Treatment” refers to both therapeutic treatment and prophylactic or preventative measures; wherein the object is to prevent or slow down (lessen) the targeted pathologic condition or disorder. Treating refers to reducing or alleviating at least one adverse effect or symptom of a disease, disorder or condition associated with a deficiency in or absence of an organ, tissue or cell function. Those in need of treatment include those already with the disorder as well as those prone to have the disorder or those in whom the disorder is to be prevented. A subject is successfully “treated” for the targeted pathologic condition or disorder if, after receiving a therapeutic amount of coated vector as described herein, the subject shows observable and/or measurable improvement in one or more of
the following: reduction in the number of pathogenic cells; reduction in the percent of total cells that are pathogenic; relief to some extent of one or more of the symptoms associated with the targeted pathologic condition or disorder; reduced morbidity and mortality, and/or improvement in quality of life issues. The above parameters for assessing successful treatment and improvement in the disease are readily measurable by routine procedures familiar to a physician.
[57] “Variant” refers to nucleic acid or amino acid sequences that typically differs from a nucleic acid or an amino acid sequence specifically disclosed herein in one or more substitutions, deletions, additions and/or insertions. Such variants may be naturally occurring or may be synthetically generated, for example, by modifying one or more of the nucleic acid or amino acid sequences of the invention and evaluating one or more biological activities of the encoded polypeptide as described herein and/or using any of a number of techniques well known in the art. When referring to percentage of identity, it is meant a nucleic acid or amino acid sequence comprising at least 85%, 90%, 95%, 96%, 97%, 98%, 99% or more sequence 10 identity with the reference nucleic acid or amino acid sequence. When it is desired to alter the amino acid sequence of a polypeptide to create an equivalent, or even an improved, variant or portion of a polypeptide of the invention, one skilled in the art will typically change one or more of the codons of the encoding DNA sequence. For example, certain amino acids may be substituted for other amino acids in a protein structure without appreciable loss of its ability to bind other polypeptides (e.g., antigens) or cells. Since it is the binding capacity and nature of a protein that defines that protein's biological functional activity, certain amino acid sequence substitutions can be made in a protein sequence, and, of course, its underlying DNA coding sequence, and nevertheless obtain a protein with similar properties. It is thus contemplated that various changes may be made in the peptide sequences of the present invention, or corresponding DNA sequences that encode said peptides without appreciable loss of their biological utility or activity. In many instances, a polypeptide variant will contain one or more conservative substitutions. A variant may also, or alternatively, contain nonconservative changes. In a preferred embodiment, variant polypeptides differ from a native sequence by substitution, deletion or addition of five amino acids or fewer. Variants may also (or alternatively) be modified by, for example,
the deletion or addition of amino acids that have minimal influence on the immunogenicity, secondary structure and hydropathic nature of the polypeptide
[58] “Vector” refers to a DNA or RNA molecule that comprises a polynucleotide sequence that encodes a peptide, a polypeptide or a protein. A vector generally contains regulatory elements capable of directing expression of the encoding polynucleotide sequence, also called transgene, in the cells into which the nucleic acid molecule is introduced. The term “transgene” refers to a polynucleotide that is introduced into a cell and is capable of being transcribed into RNA and optionally, translated and/or expressed under appropriate conditions. In certain aspects, it confers a desired property to a cell into which it was introduced, or otherwise leads to a desired technical effect, here typically a therapeutic effect. A transgene may contain sequence coding for one or more proteins or one or more fragments of proteins.
[59] "Wild-type" refers to a nucleic acid or polypeptide refers to a nucleic acid or a polypeptide that occurs in, or is produced by, a biological organism as that biological organism exists in nature.
[60] Unless specifically indicated or implied, the terms “a”, “an”, and “the” signify “at least one” or “one or more” as used herein.
DETAILED DESCRIPTION
[61] The invention relates to a mutated TAFA4 polypeptide or a variant thereof, comprising at least one cysteine substitution with any other amino acid residue.
[62] The invention relates to a mutated TAFA4 polypeptide or a variant thereof, comprising a cysteine substitution with any other amino acid residue at one or more of the following amino acid positions: amino acid position C25 and amino acid position C26, by reference to SEQ ID NO: 3.
[63] “TAFA4” or “TAFA4 polypeptide” designates a polypeptide belonging to the family of TAFA chemokine-like proteins, by reference to the amino acid sequence of
SEQ ID NO: 3; which correspond to the human TAFA4 amino acid sequence without its predicted signal peptide sequence. The TAFA4 polypeptide can comprise a signal peptide allowing its secretion. In other words, the TAFA4 polypeptide without signal peptide corresponds to the secreted form of the polypeptide, such as TAFA4 comprising or consisting of SEQ ID NO: 3.
[64] SEQ ID NO: 3
QHLRGHAGHH QIKQGTCEVV AVHRCCNKNR IEERSQTVKC SCFPGQVAGT TRAQPSCVEA SIVIQKWWCH MNPCLEGEDC KVLPDYSGWS CSSGNKVKTT KVTR.
[65] In some embodiments, the TAFA4 polypeptide is the wild-type form of human TAFA4 polypeptide or any variant thereof.
[66] In some embodiments, the wild-type TAFA4 polypeptide is selected from the group comprising or consisting to the access number: NP_001005527.1 and XP 011531673.1.
[67] The TAFA4 polypeptide refers to proteins, polypeptides or peptides with the amino acid sequence of which is substantially identical (i.e., largely but not wholly identical) to the sequence of the protein, polypeptide, or peptide, e.g., at least about 80% identical or at least about 85% identical, e.g., preferably at least about 90% identical, e.g., at least 91% identical, 92% identical, more preferably at least about 93% identical, e.g., at least 94% identical, even more preferably at least about 95% identical, e.g., at least 96% identical, yet more preferably at least about 97% identical, e.g., at least 98% identical, and most preferably at least 99% identical to the sequence of the protein, polypeptide, or peptide, e.g., to the sequence of a corresponding TAFA4.
[68] In some embodiments, the TAFA4 polypeptide may comprise a signal peptide which is cleaved, allowing to form an isoform.
[69] In some embodiments, the TAFA4 polypeptide may be an isoform of said protein, polypeptide, or peptide. Isoforms are commonly used in the field and are well- known to the skilled artisan, who will know how to obtain all the isoforms of TAFA4, as
described in Wang, et al., which published a cleavage of the signal sequence at position S36 by reference to SEQ ID NO: 1 (Cell Mol Immunol 2015, 12 (5), 615-624) (as in the sequence SEQ ID NO: 3).
[70] In some embodiment, the TAFA4 polypeptide comprises a signal peptide which is cleaved at position selected from the group comprising or consisting of: S35, S36, H45 and H46 by reference to SEQ ID NO: 1.
[71] In some embodiments, in the TAFA4 polypeptide comprises a signal peptide which is cleaved at position S35 by reference to the SEQ ID NO: 1, and once cleaved, the TAFA4 polypeptide comprise or consists of the SEQ ID NO: 2.
[72] In some embodiments, in the TAFA4 polypeptide comprises a signal peptide which is cleaved at position S36 by reference to the SEQ ID NO: 1, and once cleaved, the TAFA4 polypeptide comprise or consists of the SEQ ID NO: 3.
[73] In some embodiments, in the TAFA4 polypeptide comprises a signal peptide which is cleaved at position H45 by reference to the SEQ ID NO: 1, and once cleaved, the TAFA4 polypeptide comprise or consists of the SEQ ID NO: 4.
[74] In some embodiments, in the TAFA4 polypeptide comprises a signal peptide which is cleaved at position H46 by reference to the SEQ ID NO: 1, and once cleaved, the TAFA4 polypeptide comprise or consists of the SEQ ID NO: 5.
[75] In some embodiments, the TAFA4 polypeptide may be a variant thereof, i.e. may be conveniently denoted as "modified", or as "mutated" or "mutant", or as comprising one or more mutations, i.e., comprising one or more amino acid sequences changes compared to the amino acid sequence of TAFA4 that has not been so-mutated, such as, particularly, compared to the amino acid sequence of wild-type TAFA4. Said variant may comprise the same number of amino acids as any TAFA4 polypeptide defined above, more preferably as SEQ ID NO: 3, and thus the mutations and positions described herein are the same for the variant. Alternatively, said variant may comprise a different number of amino acids as SEQ ID NO: 3. In this case, the skilled artisan in the art will know how to place the mutations and positions described herein in the variant.
[76] In some embodiments, the mutated TAFA4 polypeptide or a variant thereof comprises one or more cysteine substitutions with any other amino acid residue.
[77] By “one or more cysteine substitutions”, it is meant that the mutated TAFA4 polypeptide comprise at least one substituted cysteine residue, at least two substituted cysteines residue, at least three substituted cysteines residue, at least four substituted cysteines residue, at least five substituted cysteines residue, at least six substituted cysteines residue, at least seven substituted cysteines residue, at least eight substituted cysteines residue, at least nine substituted cysteines residue or at least ten substituted cysteines residue, by reference to SEQ ID NO:3.
[78] In one embodiment, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with any other amino acid residue, preferably a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3.
[79] In some embodiment, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue, which consists to residue 25 with any other amino acid residue, preferably a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3.
[80] In another embodiment, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 26 with any other amino acid residue, preferably a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3.
[81] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue, which consists to residue 26 with any other amino acid residue, preferably a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3.
[82] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with any other amino acid residue, preferably a serine residue (i.e., a C25S mutations), by reference to SEQ ID NO: 3; and a substitution of the cysteine residue 26 with any other amino acid residue, preferably a serine residue (i.e., a C26S mutations), by reference to SEQ ID NO: 3.
[83] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue, which consists to cysteine residue 25 with any other amino acid residue, preferably a serine residue (i.e., a C25S mutations), by reference to SEQ ID NO: 3; and a substitution of the cysteine residue, which consists to cysteine residue 26 with any other amino acid residue, preferably a serine residue (i.e., a C26S mutations), by reference to SEQ ID NO: 3.
[84] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises substitutions of the cysteine residues, said cysteine residues is selected from the group consisting of: residue 25, and residue 26, by reference to SEQ ID NO: 3.
[85] In an embodiment, the residue 25, and the residue 26, by reference to SEQ ID NO:3, are substituted by the same amino acid.
[86] In another embodiment, the residue 25, and the residue 26, by reference to SEQ ID NO:3, are substituted by two different amino acids.
[87] In some embodiments, the amino acid substitution is a non-conservative amino acid substitution.
[88] In some embodiments, the amino acid substituting the cysteine residue with a non-conservative substitution is selected from the group comprising or consisting of: aliphatic amino acids, aromatic amino acids, negatively charged amino acids, positively charged amino acids and special amino acid.
[89] In some embodiments, the aliphatic amino acid is selected from the group consisting of: alanine, leucine, proline, valine, and isoleucine.
[90] In some embodiments, the aromatic amino acid is selected from the group consisting of: tyrosine, tryptophane, and phenylalanine.
[91] In some embodiments, the negatively charged amino acid is selected from the group consisting of: aspartic acid, and glutamic acid.
[92] In some embodiments, the positively charged amino acid is selected from the group consisting of: arginine, histidine, and lysine.
[93] In some embodiments, the special amino acid is selected from the group consisting of glycine, and proline.
[94] In some embodiments, the amino acid substitution is a conservative amino acid substitution.
[95] In some embodiments, the amino acid substituting the cysteine residue with a conservative substitution is selected from the group comprising or consisting of: polar amino acids and sulfur-containing amino acids, such as a methionine. In some embodiments, the amino acid substituting the cysteine residue with a conservative substitution is a methionine residue.
[96] In some embodiments, the polar amino acid substituting the cysteine residue is selected from the group comprising or consisting of: serine, threonine, asparagine and glutamine, more preferably serine.
[97] In some embodiments, the sulfur-containing amino acid substituting the cysteine residue is methionine.
[98] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25, said substitution of the cysteine residue 25 is selected from the group consisting of: C25S (i.e., S for serine), C25D (i.e., D for aspartic acid), C25Y (i.e., Y for tyrosine), C25E (i.e., E for glutamic acid), C25R (i.e., R for arginine), and C25G (i.e., G for glycine), by reference to SEQ ID NO: 3.
[99] In a preferred embodiment, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3.
[100] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 26, said substitution of the cysteine residue 26 is selected from the group consisting of: C26S (i.e., S for serine), C26D (i.e.,
D for aspartic acid), C26Y (i.e., Y for tyrosine), C26E (i.e., E for glutamic acid), C26R (i.e., R for arginine), C26G (i.e., G for glycine), and C26K (i.e., K for Lysine) by reference to SEQ ID NO: 3.
[101] In another preferred embodiment, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3.
[ 102] In certain less preferred embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, and further comprises a cysteine substitution with any other amino acid residue at one or more of the following amino acid positions: amino acid position Cl 7, amino acid position C26, amino acid position C40, amino acid position C42, amino acid position C57, amino acid position C69, amino acid position C74, amino acid position C80 and amino acid position C91 by reference to SEQ ID NO: 3.
[103] In certain less preferred embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, and further comprises a cysteine substitution with any other amino acid residue at one or more of the following amino acid positions: amino acid position C17, amino acid position C25, amino acid position C40, amino acid position C42, amino acid position C57, amino acid position C69, amino acid position C74, amino acid position C80 and amino acid position C91 by reference to SEQ ID NO: 3.
[104] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises two substitutions of the cysteine residues, said cysteine residues are selected in the group consisting of: C25 and C26, by reference to SEQ ID NO:3.
[105] In certain less preferred embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution
of the cysteine residue 40 with a serine residue (i.e., a C40S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 42 with a serine residue (i.e., a C42S mutation), by reference to SEQ ID NO: 3.
[ 106] In certain less preferred embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 74 with a serine residue (i.e., a C74S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 91 with a serine residue (i.e., a C91S mutation), by reference to SEQ ID NO: 3.
[107] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises one or more substitution of the cysteine residue 25, by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26, by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17, by reference to SEQ ID NO: 3, and/or a substitution of the cysteine residue 40, by reference to SEQ ID NO: 3. In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25, by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26, by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17, by reference to SEQ ID NO: 3, and/or a substitution of the cysteine residue 40, by reference to SEQ ID NO: 3. In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises exactly four substitutions, said four substitutions being of the cysteine residue 25, by reference to SEQ ID NO: 3, of the cysteine residue 26, by reference to SEQ ID NO: 3, of the cysteine residue 17, by reference to SEQ ID NO: 3, and of the cysteine residue 40, by reference to SEQ ID NO: 3. In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17 with a serine residue (i.e., a C17S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 40 with a serine residue (i.e., a C40S mutation), by reference to SEQ ID NO: 3.
[ 108] In certain less preferred embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 69 with a serine residue (i.e., a C69S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 80 with a serine residue (i.e., a C80S mutation), by reference to SEQ ID NO: 3.
[ 109] In certain less preferred embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17 with a serine residue (i.e., a C17S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 57 with a serine residue (i.e., a C57S mutation), by reference to SEQ ID NO: 3.
[110] In certain less preferred embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17 with a serine residue (i.e., a C17S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 57 with a serine residue (i.e., a C57S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 40 with a serine residue (i.e., a C40S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 42 with a serine residue (i.e., a C42S mutation), by reference to SEQ ID NO: 3.
[111] In certain less preferred embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 69 with a serine residue (i.e., a C69S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 74 with a serine residue (i.e., a C74S
mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 80 with a serine residue (i.e., a C80S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 91 with a serine residue (i.e., a C91S mutation), by reference to SEQ ID NO: 3.
[112] In certain less preferred embodiments, the mutated TAFA4 polypeptide or variant thereof comprises a substitution of the cysteine residue 25 with a serine residue (i.e., a C25S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 26 with a serine residue (i.e., a C26S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 17 with a serine residue (i.e., a C17S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 57 with a serine residue (i.e., a C57S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 40 with a serine residue (i.e., a C40S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 42 with a serine residue (i.e., a C42S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 69 with a serine residue (i.e., a C69S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 74 with a serine residue (i.e., a C74S mutation), by reference to SEQ ID NO: 3, a substitution of the cysteine residue 80 with a serine residue (i.e., a C80S mutation), by reference to SEQ ID NO: 3, and a substitution of the cysteine residue 91 with a serine residue (i.e., a C91S mutation), by reference to SEQ ID NO: 3.
[113] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group comprising or consisting of SEQ ID NO: 6, SEQ ID NO: 7, SEQ
ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID
NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID
NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID
NO: 58, SEQ ID NO: 59, SEQ ID NO: 60 and more preferably the SEQ ID NO: 6.
[114] In some embodiments, the mutated TAFA4 polypeptide has the amino acid sequence selected from the group comprising or consisting of: SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17,
SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, and more preferably the SEQ ID NO: 6.
[115] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof, comprising a cysteine substitution with an amino acid residue selected from the group consisting of serine (S), aspartic acid (D), tyrosine (Y), glutamic acid (E), arginine (R), glycine (G), and lysine (K), at one or more of the following amino acid positions: amino acid position C25 and amino acid position C26, by reference to SEQ ID NO: 3.
[116] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof, comprising a cysteine substitution with an amino acid residue selected from the group consisting of serine (S), aspartic acid (D), tyrosine (Y), glutamic acid (E), arginine (R), glycine (G), and lysine (K), at amino acid position C25, by reference to SEQ ID NO: 3, and a cysteine substitution with an amino acid residue selected from the group consisting of serine (S), aspartic acid (D), tyrosine (Y), glutamic acid (E), arginine (R), glycine (G), and lysine (K), at amino acid position C26, by reference to SEQ ID NO: 3.
[117] In some embodiments, the (b) one or more mutated TAFA4 polypeptides comprise:
(i) a cysteine substitution at position C25 by reference to SEQ ID NO: 3 selected from the group consisting of C25S, C25D, C25Y, C25E, C25R, and C25G substitutions, preferably a C25S substitution, and
(ii) a cysteine substitution at position C26 by reference to SEQ ID NO: 3 selected from the group consisting of C26S, C26D, C26Y, C26E, C26R, and C26G, preferably a C26S substitution.
[118] In some embodiments, the (b) one or more mutated TAFA4 polypeptides comprise:
(i) a cysteine substitution at position C25 by reference to SEQ ID NO: 3 selected from the group consisting of C25S, C25D, C25Y, C25E, C25R, and C25G substitutions, preferably a C25S substitution, and
(ii) a cysteine substitution at position C26 by reference to SEQ ID NO: 3 selected from the group consisting of C26S, C26D, C26Y, C26E, C26R, and C26G, preferably a C26S substitution, wherein said (b) one or more mutated TAFA4 polypeptides do not comprise cysteine substitutions on positions other than C25, C26, Cl 7, and C40 by reference to SEQ ID NO: 3.
[119] In some embodiments, the (b) one or more mutated TAFA4 polypeptides do not comprise cysteine substitutions on positions other than C25, C26, Cl 7, and C40 by reference to SEQ ID NO: 3.
[120] In some embodiments, the (b) one or more mutated TAFA4 polypeptides comprise a K96R substitution, by reference to SEQ ID NO: 3.
[121] In some embodiments, the (b) one or more mutated TAFA4 polypeptides comprise:
(i) a cysteine substitution at position C25 by reference to SEQ ID NO: 3 selected from the group consisting of C25S, C25D, C25Y, C25E, C25R, and C25G substitutions, preferably a C25S substitution;
(ii) a cysteine substitution at position C26 by reference to SEQ ID NO: 3 selected from the group consisting of C26S, C26D, C26Y, C26E, C26R, and C26G, preferably a C26S substitution; and
(iii) a K96R substitution, by reference to SEQ ID NO: 3, wherein said (b) one or more mutated TAFA4 polypeptides do not comprise cysteine substitutions on positions other than C25, C26, Cl 7, and C40 by reference to SEQ ID NO: 3.
[122] The present invention thus relates to a fusion polypeptide, comprising:
(a) one or more immunoglobulin constant domains or fragments thereof, and
(b) one or more mutated TAFA4 polypeptides comprise:
(i) a cysteine substitution at position C25 by reference to SEQ ID NO: 3 selected from the group consisting of C25S, C25D, C25Y, C25E, C25R, and C25G substitutions, preferably a C25S substitution;
(ii) a cysteine substitution at position C26 by reference to SEQ ID NO: 3 selected from the group consisting of C26S, C26D, C26Y, C26E, C26R, and C26G, preferably a C26S substitution; and
(iii) a K96R substitution, by reference to SEQ ID NO: 3, wherein said (b) one or more mutated TAFA4 polypeptides do not comprise substitutions on positions other than C25, C26 and K96 by reference to SEQ ID NO: 3.
[123] In some embodiments, the mutated TAFA4 polypeptide or a variant thereof, further comprising a mutation with any other amino acid residue at one or more of the following amino acid positions: amino acid position K96 and amino acid position D85 by reference to SEQ ID NO: 3.
[124] In some embodiments, these additional mutations, preferably at the position K96 and D85, by reference to SEQ ID NO: 3, induce the enhancement in stability can be assessed by the increase in the protein's half-life, in particular by slowing down proteolysis.
[125] In some embodiments, the one or more mutations at the amino acid position K96 and/or D85 by reference to SEQ ID NO: 3, correspond to a substitution.
[126] In some embodiments, the mutation at the amino acid position K96 by reference to SEQ ID NO: 3, can change the susceptibility for protease cleavage.
[127] In some embodiments, the mutation at the amino acid position K96 by reference to SEQ ID NO: 3, is a substitution into R (arginine) or into H (histidine), more preferably the mutated TAFA4 polypeptide or variant thereof comprises or consists of the sequence SEQ ID NO: 20 or SEQ ID NO: 58. In some embodiments, the mutation at the amino acid position D85 by reference to SEQ ID NO: 3, can change the glycosylation side in order to improve yield and solubility, and avoid aggregation.
[128] In some embodiments, the mutation at the amino acid position D85 by reference to SEQ ID NO: 3, is a substitution into N (asparagine). More preferably the mutated TAFA4 polypeptide or variant thereof comprises or consists of the sequence SEQ ID NO: 21.
[129] In some embodiments, the mutated TAFA4 polypeptide or a variant thereof, further comprising a mutation with any other amino acid residue comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group comprising or consisting of: SEQ ID NO: 20, SEQ ID NO: 21 and SEQ ID NO: 58.
[130] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 14, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135. In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135. In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group
consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, and SEQ ID NO: 60. In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135.
[131] In some embodiments, the mutated TAFA4 polypeptide or variant thereof does not comprises or consists of an amino acid sequence selected from the group consisting of SEQ ID NO: 7, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 83, SEQ ID NO: 84, SEQ ID NO: 85, SEQ ID NO: 86, SEQ ID NO: 107, SEQ ID NO: 108, SEQ ID NO: 109, SEQ ID NO: 110, SEQ ID NO: 111, SEQ ID NO: 112, and SEQ ID NO: 113.
[132] In some embodiments, the mutated TAFA4 polypeptide has the amino acid sequence selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO:
59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135. In some embodiments, the mutated TAFA4 polypeptide has the amino acid sequence selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, and SEQ ID NO:
60. In some embodiments, the mutated TAFA4 polypeptide has the amino acid sequence selected from the group consisting of SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135
[133] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8,
SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135, and preferably selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135. In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group comprising or consisting of SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60 and more preferably the SEQ ID NO: 6.
[134] In some embodiments, the mutated TAFA4 polypeptide or variant thereof comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID
NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135, and preferably selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO:
129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID
NO: 134, and SEQ ID NO: 135. In some embodiments, the mutated TAFA4 polypeptide has the amino acid sequence selected from the group comprising or consisting of SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135, and preferably selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO:
127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135.
[135] In some embodiments, the mutated TAFA4 polypeptide has the amino acid sequence selected from the group comprising or consisting of: SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, and more preferably the SEQ ID NO: 6.
[136] In some embodiments, the mutated TAFA4 polypeptide does not have the amino acid sequence selected from the group consisting of SEQ ID NO: 83, SEQ ID NO: 84, SEQ ID NO: 85, SEQ ID NO: 86, SEQ ID NO: 107, SEQ ID NO: 108, SEQ ID NO: 109, SEQ ID NO: 110, SEQ ID NO: 111, SEQ ID NO: 112, and SEQ ID NO: 113.
[137] Some of these further improvements are schematically depicted in FIGURE 5B.
[138] In some embodiments, the mutated TAFA4 polypeptide comprises a signal peptide, such as the wild-type TAFA4 precursor polypeptide with the sequence of SEQ ID NO: 1, which corresponds to the human TAFA4 amino acid sequence with a signal peptide (NCBI Accession number: NP_001005527.1).
[139] SEQ ID NO: 1
MRSPJ^RVCAKSVLLSHWLFLAYVIMVCCKLMSASSQHLRGHAGHHQIIAQGECEN VAVHRCCNKNRIEERSQTVKCSCFPGQVAGTTRAQPSCVEASIVIQKWWCHM NPCLEGEDCKVLPD YSGWSC S SGNK VKTTK VTR
[140] In some embodiments, the signal peptide, in italic before and has the sequence of SEQ ID NO: 46, SEQ ID NO: 47, SEQ ID NO: 68 or SEQ ID NO: 69.
[141] SEQ ID NO 46: MRSPRMRVCAKSVLLSHWLFLAYVLMVCCKLMSASS
[142] SEQ ID NO 47: MRSPRMRVCAKSVLLSHWLFLAYVLMVCCKLMSAS
[143] SEQ ID NO: 68:
MRVCAKWVLLSRWLVLTYVLMVCCKLMSASS
[144] SEQ ID NO: 69
MKWVTFISLLFLF S SAYS
[145] Signal peptides are commonly used in the field and are well-known to the skilled artisan, who will know how to choose a signal peptide.
[146] Those skilled artisans in the art will further understand that the TAFA4 polypeptide may contain additional amino acids, e. g., as a result of manipulations of the nucleic acid construct such as the addition of a restriction site, as long as these additional amino acids do not render the signal peptide or the TAFA4 polypeptide non-functional. The additional amino acids can be cleaved or can be retained by the mature polypeptide as long as retention does not result in a non-functional polypeptide.
[147] It is well known in the art that signal sequences are ubiquitous and interchangeable in function, as their role is solely to guide the mRNA-ribosome complex to the translocon complex, without presuming any interaction of the signal sequence to the protein to be synthesized.
[148] For example, the signal peptide suitable in the context of the invention can be the signal peptide sequences of human TAFA4, the signal peptide sequences of mouse TAFA4, the signal peptide sequences of human Albumin, the signal peptide sequences of CPYm3, the signal peptide sequences of mouse IgG-H, the signal peptide sequences of Oncostatin-M, the signal peptide sequences of huIL-2, the signal peptide sequences of huIgG-kappa and the signal peptide sequences of muIgG-lambda.
[149] The invention further relates to a fusion polypeptide, comprising:
(a) one or more domains, and
(b) one or more mutated TAFA4 polypeptides or variants thereof according to the invention, as described herein above.
[150] According to the invention, the (a) one or more domains may be selected from the group comprising or consisting of an immunoglobulin constant domain or a fragment thereof, a serum albumin, and a linker.
[151] In some embodiments, the (a) one or more domains may be selected from the group comprising or consisting of one domain, two domains, three domains, four domains, five domains, six domains.
[152] In some embodiments, the (a) one or more domain are identical.
[153] In some embodiments, the (a) one or more domains are different.
[154] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof may be selected from the group comprising or consisting of two mutated TAFA4 polypeptides or variants thereof, three mutated TAFA4 polypeptides or variants thereof, four mutated TAFA4 polypeptides or variants thereof, five mutated TAFA4 polypeptides or variants thereof, six mutated TAFA4 polypeptides or variants thereof, preferably one mutated TAFA4 polypeptide or variant thereof.
[155] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof fused to (a) one or more domains are identical.
[156] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof fused to the (a) one or more domains are different. In other words, the (b) one or more mutated TAFA4 polypeptides or variants thereof have different amino acid sequences. For example, the first mutated TAFA4 polypeptide has the amino acid sequence comprising or consisting of SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60 and the second mutated TAFA4 polypeptide has the amino acid sequence comprising or consisting of SEQ ID NO: 6, SEQ ID NO: 7, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 13, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16, SEQ ID NO: 17, SEQ ID NO: 18, SEQ ID
NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60 and more preferably the SEQ ID NO: 6 depending on the first selected sequence.
[157] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof fused to the (a) one or more domains are different. In other words, the (b) one or more mutated TAFA4 polypeptides or variants thereof have different amino acid sequences. For example, the first mutated TAFA4 polypeptide has the amino acid sequence comprising or consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135. and the second mutated TAFA4 polypeptide has the amino acid sequence comprising or consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135. and more preferably the SEQ ID NO: 6 depending on the first selected sequence.
[158] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof fused to the (a) one or more domains are different. In other words, the (b) one or more mutated TAFA4 polypeptides or variants thereof have different amino acid sequences. For example, the first mutated TAFA4 polypeptide has the amino acid sequence comprising or consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60 and the second mutated TAFA4 polypeptide has the amino acid sequence comprising or consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60 and more preferably the SEQ ID NO: 6 depending on the first selected sequence.
[159] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof are fused directly to the (a) one or more domains via a linker at the N- terminal or the C-terminal.
[160] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof are fused directly at the N-terminal or at the C-terminal of the (a) one or more domains.
[161] In some embodiments, the fusion polypeptide further comprises (c) one or more wildtype TAFA4 polypeptides or variants thereof
[162] In some embodiments, the (c) one or more wildtype TAFA4 polypeptided or variants thereof are fused directly to the (a) one or more domains via a linker at the N- terminal or the C-terminal.
[163] In some embodiments, the (c) one or more wildtype TAFA4 polypeptides or variants thereof are fused directly at the N-terminal or at the C-terminal of the (a) one or more domains.
[164] In some embodiments, the fusion polypeptide comprises or consists of at least
90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64,
SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72,
SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77,
SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82,
SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123. In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64, SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 114, and SEQ ID NO: 115. In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence
identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 114, and SEQ ID NO: 115.
[165] In some embodiments, the fusion polypeptide comprises or consists of at least
90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, and SEQ ID NO: 82. In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79,
SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 87, SEQ ID NO: 88,
SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93,
SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98,
SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
[166] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82.
[167] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72,
SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81.
[168] In some embodiments, the fusion polypeptide comprises or consists of at least
90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75,
SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80,
SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89,
SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94,
SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99,
SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
[169] In some embodiments, the fusion polypeptide comprises or consists of at least
90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75,
SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80,
SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89,
SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94,
SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99,
SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, and SEQ ID NO: 115.
[170] In some embodiments, the fusion polypeptide comprises or consists of at least
90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75,
SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80,
SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89,
SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94,
SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99,
SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, and SEQ ID NO: 106.
[171] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, and SEQ ID NO: 82.
[172] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
[173] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, , and SEQ ID NO: 115.
[174] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, and SEQ ID NO: 106.
[175] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of
SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80, preferably sequence selected from the group consisting of SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80.
[176] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80.
[177] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80
[178] In a preferred embodiment, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 78, and SEQ ID NO: 79. In some embodiments, the fusion polypeptide has at least 90% sequence identity with the amino acid sequence of SEQ ID NO: 73. In some embodiments, the fusion polypeptide has at least 90% sequence identity with the amino acid sequence of SEQ ID NO: 74. In some embodiments, the fusion polypeptide has at least 90% sequence identity with the amino acid sequence of SEQ ID NO: 78. In some embodiments, the fusion polypeptide has at least 90% sequence identity with the amino acid sequence of SEQ ID NO: 79.
[179] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 114 and SEQ ID NO: 115.
[180] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
[181] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 40, SEQ ID NO: 42, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64, SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, and SEQ ID NO: 120.
[182] In some embodiments, the fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 40, SEQ ID NO: 42, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64, SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, and SEQ ID NO: 120.
[183] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64, SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123. In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64, SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID
NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 114, and SEQ ID NO: 115. In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 114, and SEQ ID NO: 115.
[184] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, and SEQ ID NO: 82.
[185] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76,
SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81,
SEQ ID NO: 82, SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90,
SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95,
SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
[186] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76,
SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81,
SEQ ID NO: 82, SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90,
SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95,
SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100,
SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, and SEQ ID NO: 115.
[187] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76,
SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81,
SEQ ID NO: 82, SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90,
SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95,
SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, and SEQ ID NO: 106.
[188] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82.
[189] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81.
[190] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78,
SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 87,
SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92,
SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97,
SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO:
114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
[191] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78,
SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 87,
SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92,
SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97,
SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, and SEQ ID NO: 115.
[192] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78,
SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, SEQ ID NO: 82, SEQ ID NO: 87,
SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92,
SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97,
SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, and SEQ ID NO: 106.
[193] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO: 81, and SEQ ID NO: 82.
[194] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO:
104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
[195] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, SEQ ID NO: 106, SEQ ID NO: 114, , and SEQ ID NO: 115.
[196] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 87, SEQ ID NO: 88, SEQ ID NO: 89, SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 93, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96, SEQ ID NO: 97, SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 101, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104, SEQ ID NO: 105, and SEQ ID NO: 106.
[197] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80, preferably sequence selected from the group consisting of SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80.
[198] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80.
[199] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71,
SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76,
SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80
[200] In a preferred embodiment, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 78, and SEQ ID NO: 79. In some embodiments, the fusion polypeptide has the amino acid sequence of SEQ ID NO: 73. In some embodiments, the fusion polypeptide has the amino acid sequence of SEQ ID NO: 74. In some embodiments, the fusion polypeptide has the amino acid sequence of SEQ ID NO: 78. In some embodiments, the fusion polypeptide has the amino acid sequence of SEQ ID NO: 79.
[201] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 114 and SEQ ID NO: 115.
[202] In some embodiments, the fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 119, SEQ ID NO: 120, SEQ ID NO: 121, SEQ ID NO: 122, and SEQ ID NO: 123.
[203] In some embodiments, the (a) domain is selected from the group consisting of an immunoglobulin or parts thereof, i.e., immunoglobulin light chain, immunoglobulin heavy chain, immunoglobulin constant domain (or region), immunoglobulin crystallizable fragment (Fc), immunoglobulin hinge region, or any combination thereof.
[204] The incorporation of immunoglobulins, particularly constant domains of immunoglobulins (Fc regions, illustratively and not limiting as depicted in SEQ ID NO: 118; SEQ ID NO: 35; SEQ ID NO: 36; SEQ ID NO: 37; SEQ ID NO: 38, or SEQ ID NO: 39), into fusion proteins is a well-established technique in the field. This approach leverages, illustratively and non-limitatively, the stability, prolonged half-life, and effector functions conferred by the Fc region. The design and production of such fusion proteins are widely known and accessible to persons skilled in the art, with numerous methods available, including recombinant DNA technology, vector systems, and expression in mammalian, bacterial, or yeast systems. The use of Fc fusion proteins is extensively documented in scientific literature and supported by commercially available tools and reagents.
[205] In some embodiments, the (a) domain is an immunoglobulin.
[206] In some embodiments, the (a) domain is an immunoglobulin constant domain, or a fragment thereof.
[207] In some embodiments, the fusion polypeptide, comprises:
(a) one or more immunoglobulin constant domains or fragments thereof, and
(b) one or more mutated TAFA4 polypeptides or variants thereof, comprising a cysteine substitution with any other amino acid residue at one or more of the following amino acid positions: amino acid position C25 and amino acid position C26 by reference to SEQ ID NO: 3.
[208] In some embodiments, the fusion polypeptide, comprises:
(a) one or more immunoglobulin constant domains or fragments thereof, and
(b) one or more mutated TAFA4 polypeptides or variants thereof selected from the group consisting of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 14, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135.
[209] In some embodiments, the fusion polypeptide comprises or consists of from N terminal to C terminal: (a) an immunoglobulin constant domain or fragment thereof, (bl) a first mutated TAFA4 polypeptide or variant thereof and (b2) a second mutated TAFA4 polypeptide or variant thereof. In this embodiment, (a) the immunoglobulin constant domain or fragment thereof, (bl) the first mutated TAFA4 polypeptide or variant thereof and (b2) the second mutated TAFA4 polypeptide or variant thereof may be directly or linked via a linker between each other.
[210] In some embodiments, the (a) one or more immunoglobulin constant domains or fragments thereof are selected from the group comprising or consisting of the CH2 and CH3 domains of a Fc domain.
[211] In some embodiments, the (a) one or more immunoglobulin constant domains or fragments thereof are selected from the group comprising or consisting of an IgA, IgD, IgE, IgG and IgM, more preferably the CH2 and CH3 domains of the human IgG Fc domain.
[212] In some embodiments, the (a) one or more immunoglobulin constant domains or fragments thereof may be selected from the group comprising or consisting of IgG-1, IgG-2, IgG-3, or IgG-4 subtypes, IgA (including IgA-1 and IgA-2), IgE, IgD or IgM.
[213] In some embodiments, the Fc region comprises one or more amino acid modifications which may be introduced into the Fc region, thereby generating an Fc region variant. The Fc region variant may comprise a human Fc region sequence (e.g., a human IgGl, IgG2, IgG3 or IgG4 Fc region) comprising an amino acid modification (e.g., a substitution) at one or more amino acid positions.
[214] In certain aspects, antibody variants having one or more amino acid substitutions that may be introduced into an antibody of interest, e.g., in the variable and/or constant regions, and the products screened for a desired activity.
[215] In some embodiments, one or more amino acid modifications may be introduced into the Fc region of an antibody provided herein, thereby generating an Fc region variant. The Fc region variant may comprise a human Fc region sequence (e.g., a human IgGl, IgG2, IgG3 or IgG4 Fc region) comprising an amino acid modification (e.g., a substitution) at one or more amino acid positions.
[216] In some embodiments, an antibody herein has effector functions.
[217] In some embodiment, an antibody herein has enhanced effector functions.
[218] In some embodiments, an antibody variant comprises an Fc region with one or more amino acid substitutions which improve ADCC, e.g., substitutions at positions 298,
333, and/or 334 of the Fc region (EU numbering of residues), or amino acids 233-239 (see e.g. US6737056), or at position 235, 243, 300 and 396 (see e.g. US10711069).
[219] In some embodiments, an antibody provided herein is altered to increase or decrease the extent to which the antibody is glycosylated. Addition or deletion of glycosylation sites to an antibody may be conveniently accomplished by altering the amino acid sequence such that one or more glycosylation sites are created or removed. Where the antibody comprises an Fc region, the oligosaccharide attached thereto may be altered. Native antibodies produced by mammalian cells typically comprise a branched, biantennary oligosaccharide that is generally attached by an N-linkage to Asn297 of the CH2 domain of the Fc region. See, e.g., Wright et al. TIBTECH 15:26-32 (1997). The oligosaccharide may include various carbohydrates, e.g., mannose, N-acetyl glucosamine (GlcNAc), galactose, and sialic acid, as well as a fucose attached to a GlcNAc in the "stem" of the biantennary oligosaccharide structure.
[220] In some embodiments, modifications of the oligosaccharide in an antibody of the invention may be made in order to create antibody variants with certain improved properties.
[221] In some embodiments, antibody variants are provided having a non-fucosylated oligosaccharide, i.e. an oligosaccharide structure that lacks fucose attached (directly or indirectly) to an Fc region. Such non-fucosylated oligosaccharide (also referred to as "afucosylated" oligosaccharide) particularly is an N-linked oligosaccharide which lacks a fucose residue attached to the first GlcNAc in the stem of the biantennary oligosaccharide structure.
[222] In some embodiments, antibody variants are provided having an increased proportion of non-fucosylated oligosaccharides in the Fc region as compared to a native or parent antibody. For example, the proportion of non-fucosylated oligosaccharides may be at least about 20%, at least about 40%, at least about 60%, at least about 80%, or even about 100% (i.e. no fucosylated oligosaccharides are present). The percentage of non- fucosylated oligosaccharides is the (average) amount of oligosaccharides lacking fucose residues, relative to the sum of all oligosaccharides attached to Asn 297 (e. g. complex,
hybridand high mannose structures) as measured by MALDI-TOF mass spectrometry, as described in W02006082515, for example. Asn297 refers to the asparagine residue located at about position 297 in the Fc region (EU numbering of Fc region residues); however, Asn297 may also be located about ± 3 amino acids upstream or downstream of position 297, i.e., between positions 294 and 300, due to minor sequence variations in antibodies.
[223] In some embodiments, antibody variants are provided with bisected oligosaccharides, e.g., in which a biantennary oligosaccharide attached to the Fc region of the antibody is bisected by GlcNAc.
[224] Such antibodies having an increased proportion of non-fucosylated oligosaccharides in the Fc region may have improved FcyRIIIa receptor binding and/or improved effector function, in particular improved ADCC function. See, e.g., W02003035835; W02003055993. Examples of cell lines capable of producing antibodies with reduced fucosylation include:
In some embodiments, Lecl3 CHO cells deficient in protein fucosylation (see Ripka et al. Arch. Biochem. Biophys. 249:533-545 (1986); W02003035835; and W02004056312, especially at Example 11)
In some embodiments, knockout cell lines, such as alpha- 1,6-fucosyltransferase gene, FUT8, knockout CHO cells (see, e.g., Yamane-Ohnuki et al. Biotech. Bioeng. 87:614-622 (2004); Kanda, Y. et al., Biotechnol. Bioeng., 94(4):680-688 (2006); and W02003085107), cells with reduced or abolished activity of a GDP- fucose synthesis or transporter protein (see, e.g., W02003085119, W02003084570, W02003085118, W02003085102).
[225] In some embodiments, antibody variants are provided with bisected oligosaccharides, e.g., in which a biantennary oligosaccharide attached to the Fc region of the antibody is bisected by GlcNAc. Such antibody variants may have reduced fucosylation and/or improved ADCC function as described above. Examples of such antibody variants are described, e.g., in Umana et al., NatBiotechnol 17, 176-180 (1999);
Ferrara et al., Biotechn Bioeng 93, 851-861 (2006); WO9954342; W02004065540, W02003011878.
[226] In some embodiments, antibody variants with at least one galactose residue in the oligosaccharide attached to the Fc region are also provided. Such antibody variants may have improved CDC function. Such antibody variants are described, e.g., in WO199730087; WO199858964; and WO199922764.
[227] In some embodiments, an antibody herein lacks effector function.
[228] In some embodiments, the invention comprises an antibody variant that possesses some but not all effector functions, which make it a desirable candidate for applications in which the half-life of the antibody in vivo is important yet certain effector functions (such as complement-dependent cytotoxicity (CDC) and antibody-dependent cell mediated cytotoxicity (ADCC)) are unnecessary or deleterious.
[229] In some embodiments, an antibody variant comprises an Fc region with one or more amino acid substitutions which diminish FcyR binding, e.g., substitutions at positions 234 and 235 of the Fc region (EU numbering of residues).
[230] In some embodiments, the substitutions are L234A and L235A (LALA). In certain aspects, the antibody variant further comprises D265A and/or P329G in an Fc region derived from a human IgGl Fc region (see e.g. Xu, et al. Cellular immunology 2000, 200 (1), 16-26)
[231] In some embodiments, the substitutions are L234A, L235A and P329G (LALAPG) in an Fc region derived from a human IgGl Fc region (See, e.g., W02012130831).
[232] In some embodiments, the substitutions are L234A, L235A and P329S (LALAPS) in an Fc region derived from a human IgGl Fc region (See, e.g., WO1999051642).
[233] In some embodiments, the substitutions are L234A, L235A and G236R (STR) in an Fc region derived from a human IgGl Fc region (See, e.g., WO2021234402)
[234] In some embodiments, the substitutions are L234A, L235A and D265A (LALADA) in an Fc region derived from a human IgGl Fc region or F234, L235E and D265A in an Fc region derived from a human IgG4 Fc region (see, e.g. WO2021234402).
[235] In some embodiments, the antibodies may have a modification at position N297 to reduce or eliminate ADCC activity, such as N297G, N297A or N297Q (see e.g. WO2014153063).
[236] In some embodiments, Fc mutants include Fc mutants with substitutions at two or more of amino acid positions 265, 269, 270, 297 and 327, including the so-called "DANA" Fc mutant with substitution of residues 265 and 297 to alanine (US7332581). Certain antibody variants with improved or diminished binding to certain selected FcRs are described (See, e.g., US6737056; W02004056312, and Shields et al., J Biol. Chem. 9(2): 6591-6604(2001).
[237] In some embodiments, the antibody Fc lacks effector function.
In some embodiments, alterations are made in the Fc region that result in altered (i.e., either improved or diminished) Clq binding and/or Complement Dependent Cytotoxicity (CDC), e.g., as described in US6194551, WO9951642, and Idusogie et al. J Immunol. 164: 4178-4184 (2000). Examples of Fc region variants are described also in Duncan & Winter, Nature 322:738-40 (1988); W01988007089 and WO9429351.
[238] In some embodiments, alterations are made in the Fc region that result in altered (i.e., either improved or diminished) Clq binding and/or Complement Dependent Cytotoxicity (CDC), e.g., as described in US6194551, WO9951642, and Idusogie et al. J Immunol. 164: 4178-4184 (2000).
[239] In some embodiments, in vitro and/or in vivo cytotoxicity assays can be conducted to confirm the reduction/depletion of CDC and/or ADCC activities. For example, Fc receptor (FcR) binding assays can be conducted to ensure that the antibody lacks FcyR binding (hence likely lacking ADCC activity), but retains FcRn binding ability. The primary cells for mediating ADCC, NK cells, express FcyRIII only, whereas monocytes express FcyRI, FcyRII and FcyRIII. FcR expression on hematopoietic cells is
summarized in Table 3 on page 464 of Ravetch and Kinet, Annu. Rev. Immunol. 9:457- 492 (1991).
[240] In some embodiments, non-limiting examples of in vitro assays to assess ADCC activity of a molecule of interest is described in WO1988004936. Alternatively, nonradioactive assays methods may be employed (see, for example, ACTI™ non-radioactive cytotoxicity assay for flow cytometry (CellTechnology, Inc. Mountain View, CA; and CytoTox 96® non-radioactive cytotoxicity assay (Promega, Madison, WI). Useful effector cells for such assays include peripheral blood mononuclear cells (PBMC) and Natural Killer (NK) cells. Alternatively, or additionally, ADCC activity of the molecule of interest may be assessed in vivo, e.g., in an animal model such as that disclosed in Clynes et al. Proc. Nat. Acad. Sci. USA 95:652-656 (1998).
[241] In some embodiments, Clq binding assays may also be carried out to confirm that the antibody is unable to bind Clq and hence lacks CDC activity. See, e.g., Clq and C3c binding ELISA in W02006029879 and W02005100402. To assess complement activation, a CDC assay may be performed (see, for example, Gazzano- Santoro et al., J Immunol. Methods 202: 163 (1996); Cragg, M.S. et al., Blood 101 :1045-1052 (2003); and Cragg, M.S. and M.J. Glennie, Blood 103:2738-2743 (2004)).
[242] In some embodiments, the Fc of the invention contains several mutations located at the interface between the CH2 and CH3 domains, such as T250Q/M428L and M252Y/S254T/T256E + H433K/N434F By this the serum half-life of an Fc-containing molecule could be further extended.
[243] Fc region residues critical to the mouse Fc-mouse FcRn interaction have been identified by site-directed mutagenesis (see e.g. Dall'Acqua, W.F., et al. J. Immunol 169 (2002) 5171-5180). Residues 1253, H310, H433, N434, and H435 (EU index numbering) are involved in the interaction (Medesan, C., et al., Eur. J. Immunol. 26 (1996) 2533; Firan, M., et al., Int. Immunol. 13 (2001) 993; Kim, J.K., et al., Eur. J. Immunol. 24 (1994) 542).
[244] Residues 1253, H310, and H435 were found to be critical for the interaction of human Fc with murine FcRn (Kim, J.K., et al., Eur. J. Immunol. 29 (1999) 2819). Studies
of the human Fc-human FcRn complex have shown that residues 1253, S254, H435, and Y436 are crucial for the interaction (Firan, M., et al., Int. Immunol. 13 (2001) 993; Shields, R.L., et al., J. Biol. Chem. 276 (2001) 6591-6604). In Yeung, Y.A., et al. (J. Immunol. 182 (2009) 7667-7671) various mutants of residues 248 to 259 and 301 to 317 and 376 to 382 and 424 to 437 have been reported and examined.
[245] In some embodiments, FcRn binding and in vivo clearance/half-life determinations can be performed using methods known in the art (see, e.g., Petkova, S.B. et al.,Int'l. Immunol. 18(12): 1759-1769 (2006); WO2013120929).
[246] In some embodiments, the invention comprises an Fc region with one or more amino acid substitutions, which reduce FcRn binding, e.g., substitutions at positions 253, and/or 310, and/or 435 of the Fc-region (EU numbering of residues). In certain embodiments, the fusion protein variant comprises an Fc region with the amino acid substitutions at positions 253, 310 and 435.
[247] In some embodiments, the substitutions are 1253 A, H310A and H435A in an Fc region derived from a human IgGl Fc-region. In certain embodiments, an antibody variant comprises an Fc region with one or more amino acid substitutions, which reduce FcRn binding, e.g., substitutions at positions 310, and/or 433, and/or 436 of the Fc region (EU numbering of Fc residues).
[248] In some embodiments, the antibody variant comprises an Fc region with the amino acid substitutions at positions 310, 433 and 436.
[249] In some embodiments, the substitutions are H310A, H433 A and Y436A in an Fc region derived from a human IgGl Fc-region (see, e.g., WO2014177460).
[250] In some embodiments, an antibody variant comprises an Fc region with one or more amino acid substitutions which increase FcRn binding, e.g., substitutions at positions 252, and/or 254, and/or 35 256 of the Fc region (EU numbering of Fc residues).
[251] In some embodiments, the antibody variant comprises an Fc region with amino acid substitutions at positions 252, 254, and 256.
[252] In some embodiments, the substitutions are M252Y, S254T and T256E in an Fc region derived from a human IgGl Fc-region (See e.g. teachings of W02002060919).
[253] In some embodiments, the (a) one or more immunoglobulin constant domains or fragments thereof may be selected from the group comprising or consisting of: IgG-lL (i.e., SEQ ID NO: 30), F6A (i.e., SEQ ID NO: 31), F6B (i.e., SEQ ID NO: 32), F7A (i.e., SEQ ID NO: 33), F7B (i.e., SEQ ID NO: 34), Fl (i.e., SEQ ID NO: 35), F2 (i.e., SEQ ID NO: 36), F3 (i.e., SEQ ID NO: 37), F5 (i.e., SEQ ID NO: 38), and F4 (i.e., SEQ ID NO: 39), K2 (i.e., SEQ ID NO: 48), H2 (i.e., SEQ ID NO: 52).
[254] In some embodiments, the (a) one or more immunoglobulin constant domains or fragments thereof is a Fc domain. In this case, the fusion polypeptide is named TAFA4- Fc or Fc-TAFA4 or FcT4 or T4Fc fusion protein.
[255] In some embodiments, the (a) one or more Fc domains are selected from the group comprising or consisting of: human IgG-1 hinge and Fc, human IgG4 with the wildtype or stabilized hinge region, derivatives of human IgG-1 which contain mutations that severely reduce the binding to Fc-receptors and complement
[256] In some embodiments, the sequence of the human IgG-1 hinge and Fc is selected from the group comprising or consisting of: SEQ ID NO: 30-32.
[257] In some embodiments, the sequence of the human IgG-4 hinge and Fc is selected from the group comprising or consisting of: SEQ ID NO: 33-34.
[258] In some embodiments, the sequence of derivatives of human IgG-1 which contain mutations that severely reduce the binding to Fc-receptors and complement is selected from the group comprising or consisting of: SEQ ID NO: 35-39.
[259] In some embodiments, the sequence of the human IgG is selected from the group comprising or consisting of: SEQ ID NO: 48 and SEQ ID NO: 52.
[260] Derivatives of human IgG-1 are well known in the art. IgG and more precisely IgG-1 are commonly used in the field and are well-known to the skilled artisan, who will be able to routinely prepare said derivatives of human IgG-1.
[261] In some embodiments, the (a) one or more immunoglobulin constant domains or fragments thereof are a silenced immunoglobulin constant domain.
[262] In some embodiments, the silenced immunoglobulin constant domains or fragmenst thereof are an Fc-silenced.
[263] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof and/or (c) the wild-type TAFA4 polypeptide or variant thereof are fused via a linker at the C -terminal of the (b) one or more immunoglobulin constant domains or fragments thereof, more preferably to the heavy chain of the immunoglobulin chain.
[264] In some embodiments, the b) one or more mutated TAFA4 polypeptides or variants thereof and/or the (c) wild-type TAFA4 polypeptide or a variant thereof are fused via a linker at the N-terminal of the (a) one or more immunoglobulin constant domains or fragments thereof, more preferably to the heavy chain of the immunoglobulin chain.
[265] In some embodiments, the linker sequence is Gly/Ser linker.
[266] In some embodiments, the linker sequence Gly/Ser is selected in the group comprising or consisting of: (gly4ser)3, (gly4ser)4, (gly4ser), (gly3ser), gly3, and (gly3ser2)3.
[267] In some embodiments, (b) the one or more mutated TAFA4 polypeptides or variants thereof and/or (c) the wild-type TAFA4 polypeptide or variant thereof are fused directly at the C-terminal of (a) the one or more one or more immunoglobulin constant domains or fragments thereof, more preferably to the heavy chain of (a) the immunoglobulin chain.
[268] In some embodiments, (b) the one or more mutated TAFA4 polypeptides or variants thereof and/or (c) the wild-type TAFA4 polypeptide or a variant thereof are fused directly at the N-terminal of (a) the one or more immunoglobulin constant domains or fragments thereof, more preferably to the heavy chain of the immunoglobulin chain.
[269] In some embodiments, (a) the one or more immunoglobulin constant domains or fragments thereof are a silenced immunoglobulin constant domain (Fc-silent).
[270] In some embodiments, (a) the one or more immunoglobulin constant domains (Fc) or fragments thereof are a silenced Fc derived from a human IgG-1 antibody.
[271] In some embodiments, the fusion polypeptide comprises or consists of the sequence selected from the group comprising or consisting of SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 29, SEQ ID
NO: 40, SEQ ID NO: 41, SEQ ID NO: 42, SEQ ID NO: 43, SEQ ID NO: 44, SEQ ID
NO: 45, SEQ ID NO: 49, SEQ ID NO: 50, SEQ ID NO: 51, SEQ ID NO: 53, SEQ ID
NO: 54, SEQ ID NO: 55 and SEQ ID NO: 61.
[272] In some embodiments, (a) the one or more immunoglobulin constant domains (Fc) or fragments thereof are having one or more amino acid substitutions.
[273] In some embodiments, the one or more amino acid substitutions may be introduced into (a) the one or more immunoglobulin constant domains (Fc) or fragments, or the variable regions of the antibody.
[274] In some embodiments, the one or more substitutions are a conservative substitution, as shown in Table 1 under the heading of "exemplary substitutions”, or of
[276] In some embodiments, substitutions, insertions, or deletions may occur so long as such alterations do not substantially reduce the ability of the protein to bind its receptor. For example, conservative alterations (e.g., conservative substitutions as provided herein) that do not substantially reduce binding affinity may be made. Such alterations may, for example, be outside of receptor contacting residues.
[277] In some embodiments, a useful method for identification of residues or regions of a protein that may be targeted for mutagenesis is called "alanine scanning mutagenesis" as described by Cunningham and Wells (1989) Science, 244: 1081-1085. In this method, a residue or group of target residues (e.g., charged residues such as arg, asp, his, lys, and glu) are identified and replaced by a neutral or negatively charged amino acid (e.g., alanine or polyalanine) to determine whether the interaction of the antibody with antigen is affected.
[278] In some embodiments, further substitutions may be introduced at the amino acid locations demonstrating functional sensitivity to the initial substitutions. Alternatively, or additionally, a crystal structure of a protein-receptor complex may be used to identify
contact points between the protein and receptor. Such contact residues and neighboring residues may be targeted or eliminated as candidates for substitution. Variants may be screened to determine whether they contain the desired properties.
[279] In some embodiments, the (a) domain is a serum albumin.
[280] In some embodiments of the invention, the fusion polypeptide, comprises:
(a) one or more serum albumins or fragments thereof, and
(b) one or more mutated TAFA4 polypeptides or variants thereof, comprising a cysteine substitution with any other amino acid residue at one or more of the following amino acid positions: amino acid position C25 and amino acid position C26 by reference to SEQ ID NO: 3.
[281] In some embodiments, the serum albumin is human serum albumin, such as human serum albumin comprising or consisting of SEQ ID NO: 70, or fragment thereof or variant thereof.
[282] SEQ ID NO: 70
DAHKSEVAHRFKDLGEENFKALVLIAFAQYLQQCPFEDHVKLVNEVTEFAKTC VADESAENCDKSLHTLFGDKLCTVATLRETYGEMADCCAKQEPERNECFLQH
KDDNPNLPRLVRPEVDVMCTAFHDNEETFLKKYLYEIARRHPYFYAPELLFFAK RYKAAFTECCQAADKAACLLPKLDELRDEGKASSAKQRLKCASLQKFGERAF KAWAVARLSQRFPKAEFAEVSKLVTDLTKVHTECCHGDLLECADDRADLAKY ICENQDSISSKLKECCEKPLLEKSHCIAEVENDEMPADLPSLAADFVESKDVCKN YAEAKDVFLGMFLYEYARRHPDYSVVLLLRLAKTYETTLEKCCAAADPHECY AKVFDEFKPLVEEPQNLIKQNCELFEQLGEYKFQNALLVRYTKKVPQVSTPTLV EVSRNLGKVGSKCCKHPEAKRMPCAEDYLSVVLNQLCVLHEKTPVSDRVTKC CTESLVNRRPCFSALEVDETYVPKEFNAETFTFHADICTLSEKERQIKKQTALVE LVKHKPKATKEQLKAVMDDFAAFVEKCCKADDKETCFAEEGKKLVAASQAA LGL
[283] In some embodiments, the (a) domain is a linker.
[284] In some embodiments, the fusion polypeptide, comprises:
(a) one or more linkers, and
(b) one or more mutated TAFA4 polypeptides or variants thereof, comprising a cysteine substitution with any other amino acid residue at one or more of the following amino acid positions: amino acid position C25 and amino acid position C26 by reference to SEQ ID NO: 3, and
Optionally (c) one or more wild-type TAFA4 polypeptides or variants thereof
[285] In some embodiments of the present invention, the mutatedTAFA4 polypeptide fused to the Fc fragment may be linked directly to any amino acid in the hinge region or via a linker. For example, flexible linker sequences include those disclosed in Huston et al., 1988, PNAS 85:5879-5883;Wright & Deonarain, Mol. Immunol. 2007, 44(10:2860- 2869;Alfthan et al., Prot. Eng., 1995,8(7): 725-731; Luo et al., J. Biochem., 1995, 118(4):825-831;Tang et al., 1996, J. Biol. Chem. 271 (26): 15682-15686; and Turner et al., 1997, JIMM 205, 42-54.
[286] In some embodiments, the mutatedTAFA4 polypeptide is fused to the the amino acids sequences of the hinges of an IgGl, IgG2, IgG3 and IgG4 molecule. Hinges are well defined in the literature and therefore the person skilled in the art knows how to fuse said polypeptide with the Fc fragment of the hinge. In some embodiments, combinations of charged or hydrophobic residues may be incorporated into the hinge to confer multimerization properties, as seen for example in Richter et al., 2001, Prot. Eng. 14(10):775-783 for use of charged or ionic tails, e.g., acidic tails as linkers and Kostelnyet al., 1992, J. Immunol. 5(1): 1547-1553 for leucine zipper sequences. Other modified hinge regions may be entirely synthetic and may be designed to possess desired properties such as length, composition and flexibility. A number of modified hinge regions has already been described for example, in US5677425, WO1999015549, W02005003170 and W02005003169.
[287] Examples of suitable linker regions include, but are not limited to, flexible linker sequences and rigid linker sequences. Flexible linker sequences include those disclosed in Huston et al., 1988, PNAS 85:5879-5883;Wright & Deonarain, Mol. Immunol. 2007, 44(10:2860-2869;Alfthan et al., Prot. Eng., 1995,8(7): 725-731; Luo et al., J. Biochem.,
1995, 118(4): 825-83 l;Tang et al., 1996, J. Biol. Chem. 271 (26): 15682-15686; and Turner et al., 1997, JIMM 205, 42-54. Moreover, examples of suitable linker regions are described in the patent application W02009040562, in Table 1 and Table 2.
[288] Alternatively, combinations of charged or hydrophobic residues may be incorporated into the hinge to confer multimerization properties, see for example, Richter et al., 2001, Prot. Eng. 14(10):775-783 for use of charged or ionic tails, e.g., acidic tails as linkers and Kostelnyet al., 1992, J. Immunol. 5(1): 1547-1553 for leucine zipper sequences. Other modified hinge regions may be entirely synthetic and may be designed to possess desired properties such as length, composition and flexibility. A number of modified hinge regions haves already been described for example, in US5677425, WO 1999015549, W02005003170 and W02005003169.
[289] In some embodiments, the (b) one or more mutated TAFA4 polypeptides or variants thereof may be selected from the group comprising or consisting of two mutated TAFA4 polypeptides or variants thereof, three mutated TAFA4 polypeptides or variants thereof, four mutated TAFA4 polypeptides or variants thereof, five mutated TAFA4 polypeptides or variants thereof, six mutated TAFA4 polypeptides or variants thereof, preferably one mutated TAFA4 polypeptide or variant thereof.
[290] In some embodiments, the fusion polypeptide has the sequence consisting or comprising the SEQ ID NO: 22 or SEQ ID NO: 23.
[291] The invention also relates to a fusion protein comprising one or more fusion polypeptides according to the invention as described herein above, more preferably a plurality of the fusion polypeptides as described herein above.
[292] In some embodiments, the one or more fusion polypeptides may be selected from the group comprising or consisting of one fusion polypeptides, two fusion polypeptides, three fusion polypeptides, four fusion polypeptides, five fusion polypeptides, six fusions polypeptides, more preferably two fusion polypeptides forming a dimer.
[293] In some embodiments, the more than one fusion polypeptides are identical. In one embodiment, the dimer is a homodimer, meaning that the two fusion polypeptides are identical.
[294] In some embodiments, the more than one fusion polypeptides are different. In one embodiment, the dimer is a heterodimer, meaning that the two fusion polypeptides are different.
[295] In some embodiments, the two fusion polypeptides are different regarding the (a) domains, such as the (a) more than one immunoglobulin or fragment thereof are different (FIGURE 5C-2).
[296] In some embodiments, the two fusion polypeptides are different regarding the (b) more than one TAFA4 polypeptides or variants thereof which are different.
[297] In some embodiments, the two fusion polypeptides are different regarding the (b) number of TAFA4 polypeptides or variants thereof fused to the (a) domain of each fusion polypeptide defining a monomer (FIGURE 5C-2 and 5C-3).
[298] In some embodiments, the fusion protein comprises or consists to:
• one fusion polypeptide, and
• (a) one or more domains, such as one or more immunoglobulins or fragments thereof, said immunoglobulins or fragments thereof does not comprise one or more TAFA4 polypeptides or variants thereof.
[299] In some embodiments, the fusion protein comprises or consists to:
• one fusion polypeptide, said fusion polypeptide comprises more than one TAFA4 polypeptides or variants thereof, and
• (a) one or more domains, such as one or more immunoglobulins or fragments thereof, said immunoglobulins or fragments thereof do not comprise one or more TAFA4 polypeptides or variants thereof.
[300] The invention also relates to a nucleic acid sequence encoding a mutated TAFA4 polypeptide or a variant thereof according to the invention as described herein, a fusion polypeptide according to the invention as described herein or a fusion protein according to the invention as described herein.
[301] In some embodiments, the nucleic acid sequence is in form of DNA.
[302] In some embodiments, the nucleic acid sequence is in form of RNA.
[303] In some embodiments, the nucleic acid encodes for at least one expression cassette.
[304] In some embodiments, the expression cassette can be monocistronic, or polycistronic, such as bicistronic, tricistronic, tetraci str onic, pentacistronic, hexaci stronic, heptaci str onic, octaci stronic, nonaci stronic, decacistronic.
[305] In some embodiments, expression cassette comprises as a nucleic acid sequence of interest at least one the transgene, such as a mutated TAFA4 polypeptide or a variant thereof according to the invention as described herein, a fusion polypeptide according to the invention as described herein or a fusion protein according to the invention as described herein.
[306] In some embodiments, the transgene, may include an open reading frame encoding for a polypeptide or protein.
[307] In some embodiments, the expression cassette further comprises one or more signal peptides, allowing the secretion of the polypeptide.
[308] In some embodiments, the expression cassette further comprises one or more TAGs, allowing to tag the polypeptide.
[309] In some embodiments, the nucleic acid sequence of the present invention can include control sequences.
[310] In some embodiments, the expression cassette can comprise a promoter sequence. For example, the promoter may be a ubiquitous and/or constitutive promoter. Promoters are well known to the skilled artisan, who will know to select a suitable promoter depending on the nucleic acid sequence of interest to be expressed and on the cells in which expression is desired.
[311] Nucleic acid can be integrated into an expression cassette. Said cassette of expression are commonly used in the field and are well-known to the skilled artisan.
[312] This invention relates to a vector comprising a nucleic acid sequence encoding to a mutated TAFA4 polypeptide or a variant thereof according to the invention as described herein, a fusion polypeptide according to the invention as described herein or a fusion protein according to the invention as described herein.
[313] In some embodiments, the vector is in form of DNA.
[314] In some embodiments, the vector is in form of RNA.
[315] In some embodiments, the vector is selected from the group comprising or consisting of: a plasmid, a cosmid, an episome, a bacterial artificial chromosome, a viral vector.
[316] In some embodiments, the vector is plasmid-based or virus-based, and are configured to carry the essential sequences for incorporating foreign nucleic acid, for selection and for transfer of the nucleic acid into a host cell.
[317] In some embodiments, the vector of the present invention can include control sequences.
[318] In some embodiments, the nucleic acid sequence of the present invention is included in a suitable vector. Typically, the vector is a viral vector, and more particularly an adeno-associated virus (AAV), a retrovirus, bovine papilloma virus, an adenovirus vector, a lentiviral vector, a vaccinia virus, a polyoma virus, or an infective virus. In some embodiments, the vector is an AAV vector.
[319] In some embodiments, the Retroviruses may be chosen as gene delivery vectors due to their ability to integrate their genes into the host genome, transferring a large amount of foreign genetic material, infecting a broad spectrum of species and cell types and for being packaged in special cell- lines. In order to construct a retroviral vector, a nucleic acid encoding a gene of interest is inserted into the viral genome in the place of certain viral sequences to produce a virus that is replication-defective. In order to produce virions, a packaging cell line is constructed containing the gag, pol, and/or env genes but without the LTR and/or packaging components. When a recombinant plasmid containing a cDNA, together with the retroviral LTR and packaging sequences is introduced into this cell line (by calcium phosphate precipitation for example), the packaging sequence allows the RNA transcript of the recombinant plasmid to be packaged into viral particles, which are then secreted into the culture media. The media containing the recombinant retroviruses is then collected, optionally concentrated, and used for gene transfer. Retroviral vectors are able to infect a broad variety of cell types.
[320] In some embodiments, the vector is a lentiviral vector. Lentiviruses are complex retroviruses, which, in addition to the common retroviral genes gag, pol, and env, contain other genes with regulatory or structural function. The higher complexity enables the virus to modulate its life cycle, as in the course of latent infection. Some examples of lentivirus include the Human Immunodeficiency Viruses (HIV 1, HIV 2) and the Simian Immunodeficiency Virus (SIV). Lentiviral vectors have been generated by multiply attenuating the HIV virulence genes, for example, the genes env, vif, vpr, vpu and nef are deleted making the vector biologically safe. Lentiviral vectors are known in the art, see, e.g.. U.S. Pat. Nos. 6,013,516 and 5,994,136. herein by reference. Recombinant lentivirus capable of infecting a non-dividing cell wherein a suitable host cell is transfected with two or more vectors carrying the packaging functions, namely gag, pol and env, as well as rev and tat is described in U.S. Pat. No. 5,994,136. This describes a first vector that can provide a nucleic acid encoding a viral gag and a pol gene and another vector that can provide a nucleic acid encoding a viral env to produce a packaging cell. Introducing a vector providing a heterologous gene into that packaging cell yields a producer cell which releases infectious viral particles carrying the foreign gene of interest. The env preferably
is an amphotropic envelope protein which allows transduction of cells of human and other species.
[321] Vectors are commonly used in the field and are well-known to the skilled artisan, who will know how to design a vector.
[322] The invention also relates to a cell or cell population comprising one or more mutated TAFA4 polypeptides or variants thereof according to the invention as described herein, one or more fusion polypeptides according to the invention as described herein, a fusion protein according to the invention as described herein, one or more nucleic acid sequences according to the invention as described herein, or one or more vectors according to the invention as described herein.
[323] In some embodiments, the cell or cell population is prokaryote cell.
[324] In some embodiments, the cell or cell population is eucaryote cell.
[325] In some embodiments, the cell or cell population is selected from the group comprising or consisting of: CHO, HEK and HeLa.
[326] The fusion protein of the invention may be produced by any suitable means, as will be apparent to those of skill in the art. In order to produce sufficient amounts of polypeptides or functional equivalents thereof for use in accordance with the present invention, expression may conveniently be achieved by culturing under appropriate conditions recombinant host cells containing the polypeptide of the invention. In particular, the fusion protein is produced by recombinant means, by expression from an encoding nucleic acid sequence. Systems for cloning and expression of a polypeptide in a variety of different host cells are well known. When expressed in recombinant form, the fusion protein is in particular generated by expression from an encoding nucleic acid in a host cell. Any host cell may be used, depending upon the individual requirements of a particular system. Suitable host cells include bacteria mammalian cells, plant cells, yeast and baculovirus systems. Mammalian cell lines available in the art for expression of a heterologous polypeptide include Chinese hamster ovary cells (CHO cells). HeLa cells, baby hamster kidney cells and many others. Bacteria are also preferred hosts for the
production of recombinant protein, due to the ease with which bacteria may be manipulated and grown. A common, preferred bacterial host is E. coli.
[327] In some embodiments, the fusion protein of the invention is produced in Chinese hamster ovary cells (CHO cells).
[328] In some embodiments, the fusion protein of the invention is produced in Human embryonic kidney cells (HEK cells) and more particularly in HEK293 cells.
[329] The invention also relates to a pharmaceutical composition comprising: a. one or more mutated TAFA4 polypeptides or variants thereof according to the invention as described herein, one or more fusion polypeptides according to the invention as described herein, one or more fusion proteins according to the invention as described herein, one or more nucleic acid sequences according to the invention as described herein, one or more vectors according to the invention as described herein, or one or more cells or cell population according to the invention as described herein, and b. one or more pharmaceutically acceptable excipients or carriers.
[330] In some embodiments, the pharmaceutical composition further comprises sustained-release matrices, such as biodegradable polymers.
[331] The invention also relates to mutated TAFA4 polypeptide or variant thereof according to the invention as described herein, or the fusion polypeptide according to the invention as described herein, or the fusion protein according to the invention as described herein, or the pharmaceutical composition according to the invention as described herein, for use as a medicament.
[332] In some embodiments, the mutated TAFA4 polypeptide or a variant thereof, according to the invention as described herein, or the fusion polypeptide according to the invention as described herein, or the fusion protein according to the invention as described herein, for use according to the present invention is administered to the subject in need thereof in the form of a pharmaceutical composition. Typically, mutated TAFA4
polypeptide or a variant thereof, or the fusion polypeptide according to the invention as described herein, or the fusion protein according to the invention as described herein, for use according to the present invention may be combined with pharmaceutically acceptable excipients, and optionally sustained-release matrices, such as biodegradable polymers, to form pharmaceutical compositions.
[333] In some embodiments, the mutated TAFA4 or a variant thereof according to the invention, the fusion polypeptide according to the invention as described herein, the fusion protein according to the invention as described herein or the pharmaceutical composition according to the invention as described herein, are formulated for topical, intramuscular, intravenous, oral, sublingual, subcutaneous, transdermal, local or rectal administration.
[334] In some embodiments, suitable unit administration forms comprise oral-route forms such as tablets, gel capsules, powders, granules and oral suspensions or solutions, sublingual and buccal administration forms, aerosols, implants, subcutaneous, transdermal, topical, intraperitoneal, intramuscular, intravenous, subdermal, intradermal, transdermal, intrathecal and intranasal administration forms and rectal administration forms.
[335] In some embodiments, the pharmaceutical compositions are formulated for topical administration. The topical pharmaceutically acceptable carrier is any substantially nontoxic carrier conventionally usable for topical administration of pharmaceuticals in which the active ingredient of the invention will remain stable and bioavailable when applied directly to skin. For example, carriers such as those known in the art effective for penetrating the keratin layer of the skin into the stratum comeum may be useful in delivering the active ingredient of the invention to the area of interest. Such carriers include liposomes. The active ingredient of the invention can be dispersed or emulsified in a medium in a conventional manner to form a liquid preparation or mixed with a semi-solid (gel) or solid carrier to form a paste, powder, ointment, cream, lotion or the like. Suitable topical pharmaceutically acceptable carriers include water, buffered saline, petroleum jelly (vaseline), petrolatum, mineral oil, vegetable oil, animal oil, organic and inorganic waxes, such as microcrystalline, paraffin and ozocerite wax, natural
polymers, such as xanthanes, gelatin, cellulose, collagen, starch, or gum arabic, synthetic polymers, alcohols, polyols, and the like. The carrier can be a water miscible carrier composition. Such water miscible, topical pharmaceutically acceptable carrier composition can include those made with one or more appropriate ingredients outset of therapy. Because dermatologic conditions to be treated may be visible, the topical carrier can also be a topical cosmetically acceptable carrier. The topical cosmetically acceptable carrier will be any substantially non-toxic carrier conventionally usable for topical administration of cosmetics in which active ingredient of the invention will remain stable and bioavailable when applied directly to the skin surface. Suitable cosmetically acceptable carriers are known to those of skill in the art and include, but are not limited to, cosmetically acceptable liquids, creams, oils, lotions, ointments, gels, or solids, such as conventional cosmetic night creams, foundation creams, suntan lotions, sunscreens, hand lotions, make-up and make-up bases, masks and the like. Topical cosmetically acceptable carriers may be similar or identical in nature to the above described topical pharmaceutically acceptable carriers. The compositions can contain other ingredients conventional in cosmetics including perfumes, estrogen, vitamins A, C or E, alphahydroxy or alpha-keto acids such as pyruvic, lactic or glycolic acids, lanolin, vaseline, aloe vera, methyl or propyl paraben, pigments and the like.
[336] In some embodiments, it may be desirable to have a delivery system that controls the release of active ingredient of the invention to the skin and adheres to or maintains itself on the wound for an extended period of time to increase the contact time of the active ingredient of the invention on the skin. Sustained or delayed release of active ingredient of the invention provides a more efficient administration resulting in less frequent and/or decreased dosage of active ingredient of the invention and better patient compliance. Examples of suitable carriers for sustained or delayed release in a moist environment include gelatin, gum arabic, xanthane polymers. Pharmaceutical carriers capable of releasing the active ingredient of the invention when exposed to any oily, fatty, waxy, or moist environment on the area being treated, include thermoplastic or flexible thermoset resin or elastomer including thermoplastic resins such as polyvinyl halides, polyvinyl esters, polyvinylidene halides and halogenated polyolefins, elastomers such as brasiliensis, polydienes, and halogenated natural and synthetic rubbers, and flexible
thermoset resins such as polyurethanes, epoxy resins and the like. Controlled delivery systems are described, for example, in U.S. Pat. No. 5,427,778 which provides gel formulations and viscous solutions for delivery of the active ingredient of the invention to a skin site. Gels have the advantages of having a high water content to keep the skin moist, the ability to absorb skin exudate, easy application and easy removal by washing. Preferably, the sustained or delayed release carrier is a gel, liposome, microsponge or microsphere.
[337] In some embodiments, the pharmaceutical composition according to the invention as described herein may be administrated in combination with another active principle. In some embodiments, the topical administration for cutaneous treatment is accomplished via a transdermal device or patch device.
[338] In some embodiments, the mutated TAFA4 polypeptide or variant thereof according to the invention as described herein, or the fusion polypeptide according to the invention as described herein, or the fusion protein according to the invention as described herein, or the pharmaceutical composition according to the invention as described herein is administered in combination with classical treatment of skin inflammation, meaning any active agent used for the treatment of skin inflammation.
[339] In some embodiments, the classical treatment of skin inflammation refers to calcineurin inhibitor such as tacrolimus and pimecrolimus; dapsone; retinoids; methotrexate; and glucocorticoid.
[340] In some embodiments, the mutated TAFA4 polypeptide or variant thereof according to the invention as described herein, or the fusion polypeptide according to the invention as described herein, or the fusion protein according to the invention as described herein, or the pharmaceutical composition according to the invention as described herein is administered in combination with another active agent such as a glucocorticoid. Specific non-limiting examples of glucocorticoids include dexamethasone, triamcinolone acetonide (AZMACORT®), beclomethasone, dipropionate (VANCERIL®), flunisolide (AEROBID®), fluticasone propionate (FLOVENT®), prednisone, methylprednisolone and mometasonefuroate (ASMANEX®, TWISTHALER®).
[341] In some embodiments, the present invention relates to the mutated TAFA4 or a variant thereof according to the invention as described herein, the fusion polypeptide according to the invention as described herein, a fusion protein according to the invention as described herein or a pharmaceutical composition according to the invention as described herein, for use for treating a disease associated with the modulation of macrophage, such as the activation or deactivation of macrophage, in a subject in need thereof
[342] In some embodiments, the present invention relates to the mutated TAFA4 or a variant thereof according to the invention as described herein, the fusion polypeptide according to the invention as described herein, a fusion protein according to the invention as described herein or a pharmaceutical composition according to the invention as described herein, for use for treating an inflammation disease in a subject in need thereof
[343] In some embodiments, the mutated TAFA4 or variant thereof according to the invention, the fusion polypeptide according to the invention as described herein, the fusion protein according to the invention as described herein or a pharmaceutical composition according to the invention, for use according to the present invention is used for preventing skin fibrosis.
[344] In some embodiments, the present invention relates to a method of treating a disease associated with the modulation of macrophage, such as an activation or desactivation of macrophage, in a subject in need thereof
[345] In some embodiments, the present invention relates to a method of treating an inflammatory disease in a subject in need thereof comprising administering to the subject a therapeutically effective amount of the mutated TAFA4 polypeptide or the variant according to the invention thereof as described herein, the fusion polypeptide according to the invention as described herein or the fusion protein according to the invention as described herein.
[346] In some embodiments, the present invention relates to a method for preventing skin fibrosis.
[347] In some embodiments, the present invention relates to the use of the mutated TAFA4 polypeptide or the variant according to the invention thereof as described herein, the fusion polypeptide according to the invention as described herein or the fusion protein according to the invention as described herein for the manufacture of a medicament.
[348] In some embodiments, the medicament is for treating disease associated with the modulation of macrophage, such as an activation or desactivation of macrophage, in a subject in need thereof
[349] In some embodiments, the medicament is for treating an inflammatory disease.
BRIEF DESCRIPTION OF THE DRAWINGS
[350] Figures 1A-1D are a combination of graphs. Fig. 1-A shows the predicted structure of the signal sequence containing TAFA4, by using the AlphaFold-2 algorithm (see Jumper, J et al. Highly accurate protein structure prediction with AlphaFold. Nature (2021), and Varadi, M et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Research (2022)). The S36 and H46 sites by reference to SEQ ID NO: 1 are indicated with a black and white arrowhead respectively. Fig. IB shows these sites on the amino acid sequence. Fig. 1C shows the MS spectrum derived from expressing the sequence from SEQ ID NO: 2 in CHO cells. Fig. ID shows a gel of SDS-PAGE, showing the WT TAFA4 comprising the tag HIS6 in the non-reducing (-DTT) and reducing conditions (+DTT) using CCB or CBB plus the antibody aHIS to detect the protein in the gel.
[351] Figures 2A-B are a combination of graphs. Fig. 2-A shows the position of the cysteine and the prediction of the disulfide bond between them. Fig. 2-B shows the possible mutated position of the mutated TAFA4 polypeptide or variant thereof.
[352] Figures 3A-D are a combination of graphs. Fig. 3-A shows the sequence of the wildtype TAFA4 rT4 SEQ3 (SEQ ID NO:3) and the mutated TAFA4 rT4 SEQ6 (SEQ ID NO: 6). Fig. 3-B shows the SDS-PAGE. Fig. 3-C and 3-D shows the chromatogram.
[353] Figure 4A-C are a combination of graphs. Fig. 4-A shows chromatogram for FcT4 SEQ 26, Fig. 4-B shows chromatogram for FcT4 SEQ 27 and Fig. 4-C shows chromatogram for FcT4 SEQ 28.
[354] Figure 5A-B are a combination of graphs. Fig. 5A shows the sequence of some the FcT4. Fig5-A shows TAFA4 fused to an Fc (1), an asymmetrical Fc (e.g. Knobs-in- Holes) as a monomer (2) or a dimer (3), to the C -terminus of an antibody heavy chain (4) or light chain (5), or to the N-terminus of an antibody heavy chain (6) or light-chain (7).
[355] Figure 6A-F are a combination of graphs showing the quantification of cytokine production, TNF-a (Fig. 6-A), IL-6 (Fig. 6-B), IL-12p70 (Fig. 6-C), IL-12p40 (Fig. 6-D), IL-10 (Fig. 6-E), IL-IRA (Fig. 6-F) in presence of ICT, rT4 SEQ3, FcT4 SEQ26 or G=FcT4 SEQ 27.
[356] Figure 7A-E are a combination of graphs. Fig. 7A shows the survival rate in presence of PBS rT4 SEQ3 and FcT4 SEQ27. Fig. 7B shows pathological Score rate in presence of PBS rT4 SEQ3 and FcT4 SEQ27. Fig. 7C shows the quantification of cytokine production in presence of PBS rT4 SEQ3 and FcT4 SEQ27 of IL-10 (Fig. 7-C), IFN-y (Fig. 7-D) and ILip (Fig. 7-E).
[357] Figure 8 is a scheme depicting the variations analyzed having only one mutation either at position 25 or position 26, with comparison on of yield and format.
[358] Figure 9 is a combination of schemes depicting the variations analysed having two cysteine to serine mutations at positions other than 25 and 26, with comparison on of yield and format.
[359] Figure 10 is a combination of schemes depicting the variations analyzed having more than two cysteine to serine mutations at positions either other than 25 and 26 or including the 25 and 26 position, with comparison on of yield and format.
[360] Figure 11 is a combination of schemes depicting the variations analysed having more mutations on top of the two cysteine to serine mutations at positions 25 and 26, with comparison on of yield and format.
[361] Figure 12 is a combination of schemes depicting schematic sequence variation at position 25 and 26 in the TAFA4 molecule fused to an Fc, with comparison of yield and format by SDS-CE and aSEC.
[362] Figure 13 is a histogram showing IL-IRA production in Ml macrophages.
EXAMPLES
[363] The present invention is further illustrated by the following examples.
Example 1: Improved TAFA4-variants
Materials and Methods
[364] For transient gene expression, the QMCF Technology was used. Various expression constructs were designed and all coding sequences were cloned into pQMCF expression vector, coding regions were verified by sequencing. CHOEBNALT-85-1E9 cells were transiently transfected with the expression plasmid by chemical transfection according to QMCF Technology cell handling manual. Transfected cells were cultivated in CHO TF (Xell AG) growth medium for up to 10 days, after which cells were removed and supernatant clarified by filtration or centrifugation. Secreted proteins were purified by standard Protein A affinity capture and further polished by preparative size-exclusion chromatography. After production and purification on standard Protein A affinity capture and further polished by preparative size-exclusion chromatography, protein solutions were analysed by Size Exclusion Chromatography (aSEC). Purified proteins were analysed either by SDS-PAGE (NuPAGE Bis-Tris 4-12% gradient gels, Thermo Fisher Scientific) or Labchip (PerkinElmer) in the presence of Coomassie brilliant blue (CBB) staining and optionally a C-terminal anti-hexahistidine antibody (aHIS), following manufacturer’s instructions, either in non-reducing conditions (-DTT) or reducing conditions (+DTT). Buffer systems may vary between Tris-glycine (Laemli buffer) and Tris-tricine (Tricine buffer, results in better resolution of small polypeptides).
Results
[365] TAFA4 is produced in different isoforms by reference to the amino acid sequence of SEQ ID NO: 3. A sequence prediction algorithm such as SignalP version 6 (https://dtu.biolib.com/SignalP-6) predicts a signal sequence cleaved after S35 (producing TAFA4 SEQ ID NO: 2) or S36 (producing TAFA4 SEQ ID NO: 3) by reference to SEQ ID NO: 1). Wang, et al., however published a cleavage of the signal sequence position H46 by reference to SEQ ID NO: 1 (Cell Mol Immunol 2015, 12 (5), 615-624) (producing TAFA4 SEQ ID NO: 4).
[366] The wildtype (WT) TAFA4 gene coding sequence (SEQ ID NO: 1) was cloned in an expression vector with a promoter that is known to work in eukaryotic cells for transient expression (CMV promoter), such as HEK293 and CHO cells, with a C -terminal hexa-histidine fusion. After purification, the final product was analysed by MALDI- TOFF.
[367] Two clear signals were identified: one derived from a cleavage at either S35 (SEQ ID NO: 2) or S36 (SEQ ID NO: 3) by reference to SEQ ID NO: 1, and another compatible with cleavage at either H45 (SEQ ID NO: 4) or H46 (SEQ ID NO: 5) by reference to SEQ ID NO: 1 (FIGURE 1-A to 1-C). All these forms we considered as mature TAFA4 and retained activity.
[368] Upon expression of WT TAFA4-(HIS)6 in HEK293 or CHO cells, WT TAFA4 was secreted. A non-reducing sodium-dodecyl sulphate polyacrylamide lab gel (SDfS- PAGE gel) analysis revealed about 25% of the protein as a dimer (around 25 in the gel), and 75% as a monomer (around 13 kDa in the gel). A Western Blot developed with a C- terminal anti-hexahistidine antibody revealed significant cross-linking of the molecule with other proteins, resulting in a multitude of higher molecular weight bands revealed by the antibody. These bands disappear after reduction of the sample before running the SDS-PAGE, indicating these interactions were due to disulfide bond formation. This was even more clear using a tricine buffer system allowing a better resolution of smaller polypeptides. This is shown in FIGURE 2-A.
[369] We performed MS analysis of fragments obtained from non-reduced and reduced TAFA4 lysed with Trypsin and Chymotrypsin and identified possible disulphide bonds withing the TAFA4 molecule (FIGURE 2-A).
[370] FIGURE 2-A illustrates a screening campaign of mutants where various putative disulphide bonds were mutated by changing the cysteines to the homologous serine residue.
[371] Of particular interest was the sequence given in SEQ ID NO: 6, with a double cysteine removal at position 25 and 26 (by reference to SEQ ID NO: 3), as shown in FIGURE 3-B. A reducing SDS-PAGE shows a surprising increase in yield and homogeneity (FIGURE 3-B), while an analytical size exclusion chromatography (aSEC) on a high-pressure system shows a surprising increase in monomeric protein for TAFA SEQ ID NO: 6 as compared to the wild-type sequence (27% versus 6% respectively at 12 minutes), as shown in FIGURE 1-D.
Example 2: Improved TAFA4-Fc Fusion proteins
Results
[372] A fusion protein comprising the TAFA4 sequence fused to an Fc part is named FcT4. WT TAFA4 sequence, named rT4 SEQ3 (SEQ ID NO: 3), fused with an Fc-part where the effector functions are knocked-out (Fc-KO) is given in SEQ ID NO: 26. The TAFA4 double cysteine mutant (C25S and C26S by reference to sequence SEQ ID NO: 3), corresponding to SEQ ID NO: 6, fused to an Fc-KO is given in SEQ ID NO: 27. Another double cysteine mutant (C74S and C91S by reference to sequence SEQ ID NO: 3, corresponding to SEQ ID NO: 7, fused to the same Fc-KO is given in SEQ ID NO: 28. These fusions are schematically represented in FIGURE 4A-C.
[373] These 3 fusion proteins were expressed in CHO cells. From the resulting supernatant the Fc-fusions were isolated by protein-A affinity chromatography. The isolated TAFA4-Fc variants were analysed by SDS-PAGE, as a non-reduced (NR) or reduced (R) sample (M: markers size). These protein-A isolated fractions were also
analysed by analytical aSEC on HPLC (FIGURE 4). The results show a clear improvement in both yield and monomeric fraction of FcT4 SEQ ID NO: 27, with 85mAU, as compared to the other fusions FcT4 SEQ ID NO: 26 (wild type, with 42 mAU) and FcT4 SEQ ID NO: 28, another double cysteine mutant, with 14 mAU). The monomeric fraction on the chromatogram is indicated by a triangle. The expected band on a non-reducing SDS-PAGE (NR) is indicated by an arrow.
[374] The chromatogram of FcT4 SEQ ID NO: 26 shows a large proportion of aggregates (17%, RT (Retention Time) = 6.2 min), tetramers (6%, RT=7.7 min) and dimers (8%, RT=8.5 min). Also, the FcT4 SEQ ID NO: 26 monomeric peak is not homogeneous and is clearly a mixture of conformations (FIGURE 4-A). The chromatogram of FcT4 SEQ ID NO: 27 shows a small proportion of aggregates (7%, RT=6.2 min), and dimers (8%, RT=8.8 min). Surprisingly, the monomeric band is now highly homogeneous (FIGURE 4-B). Also, on a non-reducing SDS-PAGE the FcT4 SEQ ID NO: 27 fusion protein runs as a homogenous band, and with a higher yield as compared to the wild-type FcT4 SEQ ID NO: 26. The yield of FcT4 SEQ ID NO: 28 is much lower as compared to the others (FIGURE 4-C). The aSEC method shows that FcT4 SEQ ID NO: 26 comes out of purification as a much cleaner and more homogenous preparation than FcT4 SEQ ID NO: 26 and FcT4 SEQ ID NO: 28 with much less aggregates and no tetramers.
[375] The gel-inlets in FIGURE 4 compare the purities of FcT4 SEQ ID NO: 26 and of FcT4 SEQ ID NO: 27. This gel in non-reducing conditions demonstrates that FcT4 SEQ ID NO: 27 is a pure molecule with the expected molecular weight, while FcT4 SEQ ID NO: 26 is a mix of two molecules with different MWs (Molecular Weight), the low intensity band appearing at the expected MW while the high intensity band appearing at a lower than expected MW. In reducing conditions, the two fusion proteins appear as a single band of high intensity at the expected MW for a monomer of the fusion protein. These results indicate that FcT4 SEQ ID NO: 27 is the expected product while FcT4 SEQ ID NO: 26 is a mix of two conformations.
[376] In conclusion, we designed and isolated a novel fusion protein with improved yield, and a much-improved portion of the desired form-of-interest of a TAFA4-Fc fusion protein.
[377] Further, other parameters were tested, as described hereinbelow.
[378] While mutating both the cysteines at position 25 and 26 greatly improved yield, format (monomeric) and activity of the mutant molecule, we examined whether a single cysteine mutation could be enough. A variant was constructed with only a Cysteine to Serine replacement either at position 25 (C25S, SEQ ID NO: 8) or position 26 (C26S, SEQ ID NO: 9) (Figure 8-(i)). These variants were fused to an Fc part of an IgG (SEQ ID NO: 114 and SEQ ID NO: 115) and expressed with a good yield, all as the band corresponding to the active form on an SDS-PAGE analysis (Figure 8-(ii)), and remained largely monomeric when analyzed on an analytical size exclusion (aSEC) column (Figure 8-(iii))
[379] Mutating one or two cysteines at position 25 and /or 26 improves yield, format and activity, while any other two cysteines to serine mutations would not deliver such an effect (Figure 9).
[380] Using the stabilized form of TAFA4(C25S, C26S) and adding additional cysteine mutations to this form (SEQ ID NO: 12-19), or mutating 4 (SEQ ID NO: 12-16), 6 (SEQ ID NO: 17-18) , or 10 (SEQ ID NO: 19) cysteines all resulted in low or incomplete expression (Figure 10), with product formed mainly in multimers, or less stable products.
[381] Using the stabilized form of TAFA4(C25S, C26S) we varied either the N- terminus (SEQ ID NO: 44; SEQ ID NO: 45) , K96 that was found vulnerable for protease clipping (SEQ ID NO: 61, 63 - 65) , introduced a putative glycosylation site (SEQ ID NO: 62, 64), the serine residues at 25-26 (SEQ ID NO: 59, 64) or varied the C -terminal part of the TAFA4 molecule (SEQ ID NO: 66-67) (Figure ll-(i)).
[382] These variants were analyzed for yield, forming the active form (Figure ll-(ii)), and monomeric behavior on an aSEC (Figure ll-(iii)).
[383] One variant F2T4.215 (SEQ ID NO: 64) having two additional mutations gave a remarkable and surprising increase in yield, while the two predecessors F2T4.203 (SEQ ID NO: 81) and F2T4.212 (SEQ ID NO: 61) having only one of these additional mutations did not.
[384] Based on the surprising and remarkable improvement seen with F2T4.215, we further varied position 25 and 26. Representative but not limiting examples (SEQ ID NO: 71-82) are schematically depicted in Figure 12-(i). Expression yields of all variants was higher than the original C25S-C26S mutations, as visualized on an SDS-CE (Figure 12- (ii)), quantified as a protA captured titer and analysed for monomeric content by analytical size exclusion HPLC (Figure 12-iv).
[385] Most surprisingly, we isolated variants that were even improved as compared to F2T4.215. In particular F2T4.233 (SEQ ID NO: 73), F2T4.234 (SEQ ID NO: 74), F2T4.238 (SEQ ID NO: 78) and F2T4.239 (SEQ ID NO: 79) surprisingly improved yield up to 200% over F2T4.201, with a highly monomeric protein preparation.
Example 3: In vitro activity assay
Materials and Methods
[386] Monocytes were isolated from human blood and cultured for six days with M- CSF (50ng/mL) to induce their differentiation into macrophages. These macrophages were then treated with TFN-y (50ng/mL) + LPS (10 ng/mL) for two days to induce proinflammatory polarization (Ml). During the polarization process, various molecules were added to the culture, including FcT4 SEQ ID NO: 26 and FcT4 SEQ ID NO: 27 and their corresponding isotype control antibody, and rT4 (recombinant TAFA4 from R&D Systems, 5099-TA-050) with the SEQ ID NO: 3. All molecules were tested at the same molar concentration and serially diluted by 1/3. After 2 days, the supernatants were collected and analyzed by Cytometric Bead Array (Legendplex, Biolegend) to quantify cytokine production: quantification of TNF-a, IL-6, IL-12p70, IL-12p40, IL-10 and IL-
IRA production in culture supernatant of human monocyte-derived macrophages (Ml) activated.
Results
[387] FcT4 SEQ ID NO: 27, FcT4 SEQ ID NO: 26 and rT4 SEQ ID NO: 3 reduce the production of the inflammatory cytokines TNFa and IL-6 by proinflammatory Ml macrophages (FIGURES 6-A and 6-B). Surprisingly, FcT4 SEQ ID NO: 27 was more efficient than FcT4 SEQ ID NO: 26 and rT4 SEQ ID NO: 3 in reducing the production of IL-12 (p40 or p70) by these macrophages (FIGURES 6-C and 6-D). In addition, FcT4 SEQ ID NO: 27 was more potent than FcT4 SEQ ID NO: 26 or rT4 SEQ ID NO: 3 in inducing the production of the anti-inflammatory cytokines IL- 10 (FIGURE 6-E). While rT4 SEQ ID NO: 3 did not stimulate the production of IL-IRA by inflammatory macrophages, treatment with FcT4 SEQ ID NO: 27 and FcT4 SEQ ID NO: 26 was able to stimulate the production of this anti-inflammatory cytokine. Moreover, FcT4 SEQ ID NO: 27 was more active than FcT4 SEQ ID NO: 26 in stimulating the production of IL- 1RA (FIGURE 6-F).
[388] These data show that FcT4 SEQ ID NO: 27, the fusion protein bearing 2 mutated TAFA4 (C25S and C26S by reference to SEQ ID NO: 3), has a stronger antiinflammatory effect than the fusion protein bearing FcT4 SEQ ID NO: 26, and an even stronger effect than rT4 SEQ ID NO: 3, demonstrating the unexpected improvement provided by the fusion format and the 2 C to S mutations introduced in TAFA4.
[389] FcT4 SEQ ID NO: 27 has a strong anti-inflammatory effect on human macrophages.
[390] To measure cytokine production in activated macrophages, CD14+ monocytes were isolated from PBMCs and differentiated into MO macrophages over six days with M-CSF (50 ng/ml). On day 6, M0 macrophages were polarized to the Ml phenotype by stimulating them with TFNy (50 ng/ml) and LPS (10 ng/ml) for 48 hours. During polarization, Ml macrophages were treated with recombinant TAFA4 (rT4), F2T4.201, and ICT T4.1 at 384.3 nM. Untreated Ml macrophages and Ml macrophages treated with
Dexamethasone (3 pM) were included as controls. After 48 hours of treatment, supernatants were collected for cytokine quantification.
[391] Cytokines quantification revealed that F2T4.201 treatment significantly increased IL-IRA production in Ml macrophages compared to both untreated cells and those treated with its isotype control. Dexamethasone, serving as a positive control, reduced IL-IRA production, as expected. In contrast, treatment with recombinant TAFA4 had an effect on IL- IRA production similar to that of untreated cells, indicating minimal impact (Figure 13).
Example 4: In vivo activity assay
Materials and Methods
[392] Nine-week-old male C57BL/6 mice were i.p. injected with LD50 (lethal dose 50%) of LPS at day 0 (DO), in the presence of FcT4 SEQ ID NO: 27, rT4.000 molecules or vehicle (PBS). The survival rate (FIGURE 7-A) and pathological scores (FIGURE 7- B) were monitored for four days. Results from 2 independent experiments are shown with a total of 20 mice per group, (c) Production of IL-10, IFN-y (FIGURE 7-D) and IL-ip (FIGURE 7-E) in the serum of mice was monitored 6 hours post LPS injection. Each dot represents the data obtained for one mouse, n=10 mice per group from two independent experiments were analysed. Data are expressed as mean values ± S.E.M. Nine-week-old male C57BL/6 mice were studied in an in vivo model of LPS-induced endotoxic shock. LPS (from Escherichia coli strain 055 :B5; Sigma Aldrich) was diluted in PBS and injected intraperitoneally at a lethal dose of 50% (LD50, 10 pg per gram body weight). Mice were co-injected with LPS and FcT4 SEQ ID NO: 27, rT4 SEQ ID NO: 3 molecules or vehicle (PBS). rT4 SEQ ID NO: 3 corresponds to recombinant TAFA4 (R&D Systems, 5099-TA-050). FcT4 SEQ ID NO: 27 and rT4 SEQ ID NO: 3 molecules were injected at the dose of 2.1 picomoles per gram body weight. The injections were administered between 9 and 10 am and the mice were examined every 24 hours for body weight and survival rate until day 4. A pathological score was calculated as following: each day a score was attributed to each mice. A score of +2 was attributed for dead mice, +1 for mice
that keep losing weight, or +0 for mice that start recovering weight. At the end of the experiment, a cumulative pathological score (from 0 to 8) was calculated for each mouse.
[393] Different cohorts of mice were used for blood collection and clinical follow-up. For the analysis of serum cytokine production, blood samples were collected retro- orbitally six hours after LPS injection and processed to remove red blood cells. The serum samples were then stored at -80 °C until the cytokines were quantified. All samples were diluted by half before quantification. Cytokine concentrations were determined using the CBA kit (BD Biosciences) according to the manufacturer’s protocol.
Results
[394] We found that treatment with FcT4 SEQ ID NO: 27 resulted in better survival (FIGURE 7 -A) and lower pathologic score (FIGURE 7-B) compared to control mice after induction of endotoxic shock. A similar trend was also observed when the mice were treated with rT4 SEQ ID NO: 3, but the effect was smaller than in mice treated with FcT4 SEQ ID NO: 27. This better clinical outcome was associated with increased IL- 10 levels and lower IL-ip and IFN-g levels in the serum of rT4 SEQ ID NO: 3 and FcT4 SEQ ID NO: 27-treated mice compared to untreated controls (FIGURE 7-C, 7-D and 7-E). Importantly, FcT4 SEQ ID NO: 27 treatment modulated the production of inflammatory cytokines in vivo more efficiently than rT4 SEQ ID NO: 3 treatment, leading to a lower mortality rate.
[395] These data show that in vivo treatment with FcT4 SEQ ID NO: 27 modulates the systemic inflammatory response more efficiently than treatment with rT4 SEQ ID NO: 3 in a model of endotoxic shock. Moreover, this potent effect on cytokine production - upregulation of IL-10 and downregulation of IFN-g and IL-lb - is associated with a higher survival rate and a positive clinical response in mice treated with FcT4 SEQ ID NO: 27. These results confirm the ones observed in vitro on the inflammatory activity of activated macrophages. The fusion protein format and the 2 C to S mutations in TAFA4 of FcT4 SEQ ID NO: 27 lead to a strong increase in anti-inflammatory activity compared to WT TAFA4.
[396] Treatment with FcT4 SEQ ID NO: 27 protects mice from systemic inflammation and death in a model of septic shock.
Claims
1. A fusion polypeptide, comprising:
(a) one or more domains, and
(b) one or more mutated TAFA4 polypeptides or variants thereof, comprising a cysteine substitution with any other amino acid residue at one or more of the following amino acid positions: amino acid position C25 and amino acid position C26, by reference to SEQ ID NO: 3.
2. The fusion polypeptide according to claim 1, wherein the (b) one or more mutated TAFA4 polypeptides or variants thereof, comprise a cysteine substitution with an amino acid residue selected from the group consisting of serine (S), aspartic acid (D), tyrosine (Y), glutamic acid (E), arginine (R), glycine (G), and lysine (K), at one or more of the following amino acid positions: amino acid position C25 and amino acid position C26, by reference to SEQ ID NO: 3.
3. The fusion polypeptide according to claim 1 or 2, wherein the cysteine substitution at position C25 of the (b) one or more mutated TAFA4 polypeptides is a substitution of the cysteine residue by a serine residue by reference to SEQ ID NO: 3, such as a C25S substitution.
4. The fusion polypeptide according to any one of claims 1 to 3, wherein the cysteine substitution at position C26 of the (b) one or more mutated TAFA4 polypeptides is a substitution of the cysteine residue by a serine residue by reference to SEQ ID NO: 3, such as a C26S substitution.
5. The fusion polypeptide according to any one of claims 1 to 4, wherein the (b) one or more mutated TAFA4 polypeptides comprise
(i) a cysteine substitution with any other amino acid residue at the amino acid position C25 by reference to SEQ ID NO: 3, and
(ii) a cysteine substitution with any other amino acid residue amino acid position C26 by reference to SEQ ID NO: 3.
6. The fusion polypeptide according to any one of claims 1 to 5, wherein the (b) one or more mutated TAFA4 polypeptides comprise:
(i) a cysteine substitution at position C25 by reference to SEQ ID NO: 3 selected from the group consisting of C25S, C25D, C25Y, C25E, C25R, and C25G substitutions, preferably a C25S substitution, and
(ii) a cysteine substitution at position C26 by reference to SEQ ID NO: 3 selected from the group consisting of C26S, C26D, C26Y, C26E, C26R, and C26G, preferably a C26S substitution.
7. The fusion polypeptide according to any one of claims 1 to 6, wherein the (b) one or more mutated TAFA4 polypeptides do not comprise cysteine substitutions on positions other than C25, C26, C17, and C40, by reference to SEQ ID NO: 3.
8. The fusion polypeptide according to any one of claims 1 to 7, wherein the (b) one or more mutated TAFA4 polypeptides comprise a K96R substitution, by reference to SEQ ID NO: 3.
9. The fusion polypeptide according to any one of claims 1 to 8, wherein the (b) one or more mutated TAFA4 polypeptides comprise:
(i) a cysteine substitution at position C25 by reference to SEQ ID NO: 3 selected from the group consisting of C25S, C25D, C25Y, C25E, C25R, and C25G substitutions, preferably a C25S substitution;
(ii) a cysteine substitution at position C26 by reference to SEQ ID NO: 3 selected from the group consisting of C26S, C26D, C26Y, C26E, C26R, and C26G, preferably a C26S substitution; and
(iii) a K96R substitution, by reference to SEQ ID NO: 3, wherein said (b) one or more mutated TAFA4 polypeptides do not comprise cysteine substitutions on positions other than C25, C26, C17, and C40 by reference to SEQ ID NO: 3.
10. The fusion polypeptide according to any one of claims 1 to 9, wherein the (b) one or more mutated TAFA4 polypeptides thereof comprise at least 90% sequence
I l l identity to the amino acid sequences selected from the group comprising or consisting of: SEQ ID NO: 6, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 57, SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135.
11. The fusion polypeptide according to any one of claims 1 to 9, wherein the (b) one or more mutated TAFA4 polypeptides thereof comprise at least 90% sequence identity to the amino acid sequences selected from the group comprising or consisting of SEQ ID NO: 124, SEQ ID NO: 125, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128, SEQ ID NO: 129, SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 133, SEQ ID NO: 134, and SEQ ID NO: 135.
12. The fusion polypeptide according to any one of claims 1 to 11, wherein the (b) one or more mutated TAFA4 polypeptides are fused directly or via a linker at the N- terminal or the C-terminal of the (a) one or more domain.
13. The fusion polypeptide according to any one of claims 1 to 12, wherein (a) one or more domains correspond to one or more immunoglobulin constant domains or fragments thereof, preferably said one or more immunoglobulin constant domains or fragments thereof comprise the CH2 and CH3 domains of a Fc domain.
14. The fusion polypeptide according to claim 13, wherein the (a) one or more immunoglobulin constant domains or a fragment thereof are selected from the group comprising or consisting of: IgA, IgD, IgE, IgG and IgM, preferably IgG.
15. The fusion polypeptide according to any one of claims 1 to 14, wherein said fusion polypeptide comprises or consists of at least 90% sequence identity to the amino acid sequences selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 40, SEQ ID NO: 42, SEQ ID NO: 61, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64, SEQ ID NO: 65, SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80, SEQ ID NO:
81, SEQ ID NO: 82, SEQ ID NO: 106, SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 119, and SEQ ID NO: 120.
16. The fusion polypeptide according to any one of claims 1 to 14, wherein said fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 27, SEQ ID NO: 64, SEQ ID NO: 82, SEQ ID NO: 71, SEQ ID NO: 72, SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 77, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80.
17. The fusion polypeptide according to any one of claims 1 to 14, wherein said fusion polypeptide has an amino acid sequence selected from the group consisting of SEQ ID NO: 73, SEQ ID NO: 74, SEQ ID NO: 78, and SEQ ID NO: 79.
18. A fusion protein comprising a plurality of the fusion polypeptides according to any one of claims 1 to 17.
19. A nucleic acid sequence encoding the fusion polypeptide according to any one of claims 1 to 17 or the fusion protein according to claim 18.
20. A vector comprising a nucleic acid sequence encoding the fusion polypeptide or the fusion protein according to claim 19.
21. A cell or cell population comprising one or more fusion polypeptides according to any one of claims 1 to 17, or one or more fusion proteins according to claim 18, or one or more nucleic acid sequences according to claim 19, or one or more vectors according to claim 20.
22. A pharmaceutical composition comprising:
• one or more fusion polypeptides according to any one of claims 1 to 17, a fusion protein according to claim 18, one or more nucleic acid sequence according to claim 19, one or more vector according to claim 20, or one or more cell or cell population according to claim 21; and one or more pharmaceutically acceptable excipients or carriers.
23. The fusion polypeptide according to any one of claims 1 to 17, the fusion protein according to claim 18, the nucleic acid sequence according to claim 19, the vector according to claim 20, the cell or the cell population according to claim 21 or the pharmaceutical composition according to claim 22, for use as a medicament.
24. The fusion polypeptide according to any one of claims 1 to 17, the fusion protein according to claim 18, the nucleic acid sequence according to claim 19, the vector according to claim 20, the cell or the cell population according to claim 21 or the pharmaceutical composition according to claim 22, for use for treating disease associated with the modulation of macrophage in a subject in need thereof
25. The fusion polypeptide, the fusion protein, the nucleic acid sequence, the vector, the cell or the cell population or the pharmaceutical composition according to claim 24 for use, wherein the disease is an inflammation disease, more preferably a skin inflammation disease.
26. The pharmaceutical composition according to claim 22 or the pharmaceutical composition for use according to any one of claims 23 to 25, wherein the pharmaceutical composition is formulated for topical, intramuscular, subcutaneous, intradermal, intravenous or oral administration, preferably the topical administration is accomplished via a transdermal device or patch device.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP23307315.4 | 2023-12-21 | ||
| EP23307315 | 2023-12-21 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025133290A1 true WO2025133290A1 (en) | 2025-06-26 |
Family
ID=89620804
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/EP2024/088159 Pending WO2025133290A1 (en) | 2023-12-21 | 2024-12-20 | Protein for immune regulation |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025133290A1 (en) |
Citations (39)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1988004936A1 (en) | 1987-01-08 | 1988-07-14 | International Genetic Engineering, Inc. | Chimeric antibody with specificity to human b cell surface antigen |
| WO1988007089A1 (en) | 1987-03-18 | 1988-09-22 | Medical Research Council | Altered antibodies |
| WO1994029351A2 (en) | 1993-06-16 | 1994-12-22 | Celltech Limited | Antibodies |
| US5427778A (en) | 1987-09-18 | 1995-06-27 | Ethicon, Inc. | Gel formulations containing growth factors and acrylamide polymer |
| WO1997030087A1 (en) | 1996-02-16 | 1997-08-21 | Glaxo Group Limited | Preparation of glycosylated antibodies |
| US5677425A (en) | 1987-09-04 | 1997-10-14 | Celltech Therapeutics Limited | Recombinant antibody |
| WO1998058964A1 (en) | 1997-06-24 | 1998-12-30 | Genentech, Inc. | Methods and compositions for galactosylated glycoproteins |
| WO1999015549A2 (en) | 1997-09-19 | 1999-04-01 | Celltech Therapeutics Limited | Peptide sequences as hinge regions in proteins like immunoglobulin fragments and their use in medicine |
| WO1999022764A1 (en) | 1997-10-31 | 1999-05-14 | Genentech, Inc. | Methods and compositions comprising glycoprotein glycoforms |
| WO1999051642A1 (en) | 1998-04-02 | 1999-10-14 | Genentech, Inc. | Antibody variants and fragments thereof |
| WO1999054342A1 (en) | 1998-04-20 | 1999-10-28 | Pablo Umana | Glycosylation engineering of antibodies for improving antibody-dependent cellular cytotoxicity |
| US5994136A (en) | 1997-12-12 | 1999-11-30 | Cell Genesys, Inc. | Method and means for producing high titer, safe, recombinant lentivirus vectors |
| US6013516A (en) | 1995-10-06 | 2000-01-11 | The Salk Institute For Biological Studies | Vector and method of use for nucleic acid delivery to non-dividing cells |
| US6194551B1 (en) | 1998-04-02 | 2001-02-27 | Genentech, Inc. | Polypeptide variants |
| WO2002060919A2 (en) | 2000-12-12 | 2002-08-08 | Medimmune, Inc. | Molecules with extended half-lives, compositions and uses thereof |
| WO2003011878A2 (en) | 2001-08-03 | 2003-02-13 | Glycart Biotechnology Ag | Antibody glycosylation variants having increased antibody-dependent cellular cytotoxicity |
| WO2003035835A2 (en) | 2001-10-25 | 2003-05-01 | Genentech, Inc. | Glycoprotein compositions |
| WO2003055993A1 (en) | 2001-12-25 | 2003-07-10 | Kyowa Hakko Kogyo Co., Ltd. | Composition of antibody specifically binding to cd20 |
| WO2003085107A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | Cells with modified genome |
| WO2003084570A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | DRUG CONTAINING ANTIBODY COMPOSITION APPROPRIATE FOR PATIENT SUFFERING FROM FcϜRIIIa POLYMORPHISM |
| WO2003085118A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | Process for producing antibody composition |
| WO2003085119A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | METHOD OF ENHANCING ACTIVITY OF ANTIBODY COMPOSITION OF BINDING TO FcϜ RECEPTOR IIIa |
| WO2003085102A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | Cell with depression or deletion of the activity of protein participating in gdp-fucose transport |
| US6737056B1 (en) | 1999-01-15 | 2004-05-18 | Genentech, Inc. | Polypeptide variants with altered effector function |
| WO2004056312A2 (en) | 2002-12-16 | 2004-07-08 | Genentech, Inc. | Immunoglobulin variants and uses thereof |
| WO2004065540A2 (en) | 2003-01-22 | 2004-08-05 | Glycart Biotechnology Ag | Fusion constructs and use of same to produce antibodies with increased fc receptor binding affinity and effector function |
| WO2005003170A2 (en) | 2003-07-01 | 2005-01-13 | Celltech R & D Limited | Modified antibody fragments |
| WO2005003169A2 (en) | 2003-07-01 | 2005-01-13 | Celltech R & D Limited | Modified antibody fab fragments |
| WO2005100402A1 (en) | 2004-04-13 | 2005-10-27 | F.Hoffmann-La Roche Ag | Anti-p-selectin antibodies |
| WO2006029879A2 (en) | 2004-09-17 | 2006-03-23 | F.Hoffmann-La Roche Ag | Anti-ox40l antibodies |
| WO2006082515A2 (en) | 2005-02-07 | 2006-08-10 | Glycart Biotechnology Ag | Antigen binding molecules that bind egfr, vectors encoding same, and uses thereof |
| WO2009040562A1 (en) | 2007-09-26 | 2009-04-02 | Ucb Pharma S.A. | Dual specificity antibody fusions |
| WO2012130831A1 (en) | 2011-03-29 | 2012-10-04 | Roche Glycart Ag | Antibody fc variants |
| WO2013120929A1 (en) | 2012-02-15 | 2013-08-22 | F. Hoffmann-La Roche Ag | Fc-receptor based affinity chromatography |
| WO2014153063A1 (en) | 2013-03-14 | 2014-09-25 | Amgen Inc. | AGLYCOSYLATED Fc-CONTAINING POLYPEPTIDES |
| WO2014177460A1 (en) | 2013-04-29 | 2014-11-06 | F. Hoffmann-La Roche Ag | Human fcrn-binding modified antibodies and methods of use |
| WO2020064907A1 (en) | 2018-09-27 | 2020-04-02 | INSERM (Institut National de la Santé et de la Recherche Médicale) | Methods and pharmaceutical composition reducing skin inflammation |
| US10711069B2 (en) | 2006-12-08 | 2020-07-14 | Macrogenics, Inc. | Methods for the treatment of disease using immunoglobulins having Fc regions with altered affinities for FcγRactivating and FcγRinhibiting |
| WO2021234402A2 (en) | 2020-05-21 | 2021-11-25 | Mabsolve Limited | Modified immunoglobulin fc regions |
-
2024
- 2024-12-20 WO PCT/EP2024/088159 patent/WO2025133290A1/en active Pending
Patent Citations (40)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1988004936A1 (en) | 1987-01-08 | 1988-07-14 | International Genetic Engineering, Inc. | Chimeric antibody with specificity to human b cell surface antigen |
| WO1988007089A1 (en) | 1987-03-18 | 1988-09-22 | Medical Research Council | Altered antibodies |
| US5677425A (en) | 1987-09-04 | 1997-10-14 | Celltech Therapeutics Limited | Recombinant antibody |
| US5427778A (en) | 1987-09-18 | 1995-06-27 | Ethicon, Inc. | Gel formulations containing growth factors and acrylamide polymer |
| WO1994029351A2 (en) | 1993-06-16 | 1994-12-22 | Celltech Limited | Antibodies |
| US6013516A (en) | 1995-10-06 | 2000-01-11 | The Salk Institute For Biological Studies | Vector and method of use for nucleic acid delivery to non-dividing cells |
| WO1997030087A1 (en) | 1996-02-16 | 1997-08-21 | Glaxo Group Limited | Preparation of glycosylated antibodies |
| WO1998058964A1 (en) | 1997-06-24 | 1998-12-30 | Genentech, Inc. | Methods and compositions for galactosylated glycoproteins |
| WO1999015549A2 (en) | 1997-09-19 | 1999-04-01 | Celltech Therapeutics Limited | Peptide sequences as hinge regions in proteins like immunoglobulin fragments and their use in medicine |
| WO1999022764A1 (en) | 1997-10-31 | 1999-05-14 | Genentech, Inc. | Methods and compositions comprising glycoprotein glycoforms |
| US5994136A (en) | 1997-12-12 | 1999-11-30 | Cell Genesys, Inc. | Method and means for producing high titer, safe, recombinant lentivirus vectors |
| WO1999051642A1 (en) | 1998-04-02 | 1999-10-14 | Genentech, Inc. | Antibody variants and fragments thereof |
| US6194551B1 (en) | 1998-04-02 | 2001-02-27 | Genentech, Inc. | Polypeptide variants |
| WO1999054342A1 (en) | 1998-04-20 | 1999-10-28 | Pablo Umana | Glycosylation engineering of antibodies for improving antibody-dependent cellular cytotoxicity |
| US7332581B2 (en) | 1999-01-15 | 2008-02-19 | Genentech, Inc. | Polypeptide variants with altered effector function |
| US6737056B1 (en) | 1999-01-15 | 2004-05-18 | Genentech, Inc. | Polypeptide variants with altered effector function |
| WO2002060919A2 (en) | 2000-12-12 | 2002-08-08 | Medimmune, Inc. | Molecules with extended half-lives, compositions and uses thereof |
| WO2003011878A2 (en) | 2001-08-03 | 2003-02-13 | Glycart Biotechnology Ag | Antibody glycosylation variants having increased antibody-dependent cellular cytotoxicity |
| WO2003035835A2 (en) | 2001-10-25 | 2003-05-01 | Genentech, Inc. | Glycoprotein compositions |
| WO2003055993A1 (en) | 2001-12-25 | 2003-07-10 | Kyowa Hakko Kogyo Co., Ltd. | Composition of antibody specifically binding to cd20 |
| WO2003085118A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | Process for producing antibody composition |
| WO2003085107A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | Cells with modified genome |
| WO2003085102A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | Cell with depression or deletion of the activity of protein participating in gdp-fucose transport |
| WO2003084570A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | DRUG CONTAINING ANTIBODY COMPOSITION APPROPRIATE FOR PATIENT SUFFERING FROM FcϜRIIIa POLYMORPHISM |
| WO2003085119A1 (en) | 2002-04-09 | 2003-10-16 | Kyowa Hakko Kogyo Co., Ltd. | METHOD OF ENHANCING ACTIVITY OF ANTIBODY COMPOSITION OF BINDING TO FcϜ RECEPTOR IIIa |
| WO2004056312A2 (en) | 2002-12-16 | 2004-07-08 | Genentech, Inc. | Immunoglobulin variants and uses thereof |
| WO2004065540A2 (en) | 2003-01-22 | 2004-08-05 | Glycart Biotechnology Ag | Fusion constructs and use of same to produce antibodies with increased fc receptor binding affinity and effector function |
| WO2005003170A2 (en) | 2003-07-01 | 2005-01-13 | Celltech R & D Limited | Modified antibody fragments |
| WO2005003169A2 (en) | 2003-07-01 | 2005-01-13 | Celltech R & D Limited | Modified antibody fab fragments |
| WO2005100402A1 (en) | 2004-04-13 | 2005-10-27 | F.Hoffmann-La Roche Ag | Anti-p-selectin antibodies |
| WO2006029879A2 (en) | 2004-09-17 | 2006-03-23 | F.Hoffmann-La Roche Ag | Anti-ox40l antibodies |
| WO2006082515A2 (en) | 2005-02-07 | 2006-08-10 | Glycart Biotechnology Ag | Antigen binding molecules that bind egfr, vectors encoding same, and uses thereof |
| US10711069B2 (en) | 2006-12-08 | 2020-07-14 | Macrogenics, Inc. | Methods for the treatment of disease using immunoglobulins having Fc regions with altered affinities for FcγRactivating and FcγRinhibiting |
| WO2009040562A1 (en) | 2007-09-26 | 2009-04-02 | Ucb Pharma S.A. | Dual specificity antibody fusions |
| WO2012130831A1 (en) | 2011-03-29 | 2012-10-04 | Roche Glycart Ag | Antibody fc variants |
| WO2013120929A1 (en) | 2012-02-15 | 2013-08-22 | F. Hoffmann-La Roche Ag | Fc-receptor based affinity chromatography |
| WO2014153063A1 (en) | 2013-03-14 | 2014-09-25 | Amgen Inc. | AGLYCOSYLATED Fc-CONTAINING POLYPEPTIDES |
| WO2014177460A1 (en) | 2013-04-29 | 2014-11-06 | F. Hoffmann-La Roche Ag | Human fcrn-binding modified antibodies and methods of use |
| WO2020064907A1 (en) | 2018-09-27 | 2020-04-02 | INSERM (Institut National de la Santé et de la Recherche Médicale) | Methods and pharmaceutical composition reducing skin inflammation |
| WO2021234402A2 (en) | 2020-05-21 | 2021-11-25 | Mabsolve Limited | Modified immunoglobulin fc regions |
Non-Patent Citations (43)
| Title |
|---|
| "Biocomputing: Informatics and Genome Projects", 1993, ACADEMIC PRESS |
| ALFTHAN ET AL., PROT. ENG., vol. 8, no. 7, 1995, pages 725 - 731 |
| ALTSCHUL ET AL., J. MOI. BIOL., vol. 215, 1990, pages 403 - 410 |
| BORROK M.J ET AL., J PHARM SCI., 2017 |
| CARILLO ET AL., SIAM J. APPLIED MATH., vol. 48, 1988, pages 1073 |
| CELL MOL IMMUNOL, vol. 12, no. 5, 2015, pages 615 - 624 |
| CLYNES ET AL., PROC. NAT. ACAD. SCI. USA, vol. 95, 1998, pages 652 - 656 |
| CRAGG, M.S. ET AL., BLOOD, vol. 101, 2003, pages 1045 - 1052 |
| CRAGG, M.S.M.J. GLENNIE, BLOOD, vol. 103, 2004, pages 2738 - 2743 |
| CUNNINGHAMWELLS, SCIENCE, vol. 244, 1989, pages 1081 - 1085 |
| DALL'ACQUA, W.F. ET AL., J. IMMUNOL, vol. 169, 2002, pages 5171 - 5180 |
| DEVEREUX ET AL., NUCL. ACID. RES., 1984, pages 387 |
| DUNCANWINTER, NATURE, vol. 322, 1988, pages 738 - 40 |
| FERRARA ET AL., BIOTECHN BIOENG, vol. 93, 2006, pages 851 - 861 |
| FIRAN, M. ET AL., INT. IMMUNOL., vol. 13, 2001, pages 993 |
| GAZZANO-SANTORO ET AL., J IMMUNOL. METHODS, vol. 202, 1996, pages 163 |
| HUSTON ET AL., PNAS, vol. 85, 1988, pages 5879 - 5883 |
| IDUSOGIE ET AL., J IMMUNOL., vol. 164, 2000, pages 4178 - 4184 |
| JUMPER, JET: "Highly accurate protein structure prediction with AlphaFold", NATURE, 2021 |
| KANDA, Y. ET AL., BIOTECHNOL. BIOENG., vol. 94, no. 4, pages 680 - 688 |
| KIM, J.K. ET AL., EUR. J. IMMUNOL., vol. 29, 1999, pages 2819 |
| KIM, J.K. ET AL., J. IMMUNOL., vol. 24, 1994, pages 542 |
| KOSTELNYET, J. IMMUNOL., vol. 5, no. 1, 1992, pages 1547 - 1553 |
| LUO ET AL., J. BIOCHEM., vol. 118, no. 4, 1995, pages 825 - 831 |
| MEDESAN, C. ET AL., EUR. J. IMMUNOL., vol. 26, 1996, pages 2533 |
| PETKOVA, S.B. ET AL., INT'1. IMMUNOL., vol. 18, no. 12, 2006, pages 1759 - 1769 |
| RAVETCHKINET, ANNU. REV. IMMUNOL., vol. 9, 1991, pages 457 - 492 |
| RICHTER ET AL., PROT. ENG., vol. 14, no. 10, 2001, pages 775 - 783 |
| RIPKA ET AL., ARCH. BIOCHEM. BIOPHYS., vol. 249, 1986, pages 533 - 545 |
| SHIELDS ET AL., J BIOL. CHEM., vol. 9, no. 2, 2001, pages 6591 - 6604 |
| SHIELDS, R.L. ET AL., J. BIOL. CHEM., vol. 276, 2001, pages 6591 - 6604 |
| TANG ET AL., J. BIOL. CHEM., vol. 271, no. 26, 1996, pages 15682 - 15686 |
| TURNER ET AL., JIMM, vol. 205, 1997, pages 42 - 54 |
| UMANA ET AL., NAT BIOTECHNOL, vol. 17, 1999, pages 176 - 180 |
| VARADI, M ET AL.: "AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models", NUCLEIC ACIDS RESEARCH, 2022 |
| VON HEINJE, G.: "Sequence Analysis in Molecular Biology", 1987, ACADEMIC PRESS |
| WENYAN WANG ET AL: "FAM19A4 is a novel cytokine ligand of formyl peptide receptor 1 (FPR1) and is able to promote the migration and phagocytosis of macrophages", CELLULAR & MOLECULAR IMMUNOLOGY, vol. 12, no. 5, 1 September 2015 (2015-09-01), London, pages 615 - 624, XP055705364, ISSN: 1672-7681, DOI: 10.1038/cmi.2014.61 * |
| WRIGHT ET AL., TIBTECH, vol. 15, 1997, pages 26 - 32 |
| WRIGHTDEONARAIN, MOL. IMMUNOL., vol. 44, 2007 |
| XU ET AL., CELLULAR IMMUNOLOGY, vol. 200, no. 1, 2000, pages 16 - 26 |
| Y TOM TANG: "TAFA: a novel secreted family with conserved cysteine residues and restricted expression in the brain", GENOMICS, vol. 83, no. 4, 1 April 2004 (2004-04-01), US, pages 727 - 734, XP093164339, ISSN: 0888-7543, DOI: 10.1016/j.ygeno.2003.10.006 * |
| YAMANE-OHNUKI ET AL., BIOTECH. BIOENG., vol. 87, 2004, pages 614 - 622 |
| YEUNG, Y.A. ET AL., J. IMMUNOL., vol. 182, 2009, pages 7667 - 7671 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| AU2022201204B2 (en) | Stabilization of Fc-containing polypeptides | |
| JP2023159173A (en) | Modified IL-2 FC fusion protein | |
| EP2161287B1 (en) | OPTIMIZED TACI-Fc FUSION PROTEINS | |
| KR102276157B1 (en) | Hyperglycosylated human blood coagulation factor VIII fusion protein and its preparation and use | |
| EP2504360B1 (en) | Monomeric antibody fc | |
| US8871907B2 (en) | Glycosylated immunoglobulin and immunoadhesin comprising the same | |
| KR102865175B1 (en) | Ultra-long-acting insulin-FC fusion protein and methods of use | |
| JP7488303B2 (en) | Extracellular domain of the alpha subunit of the IgE Fc receptor, pharmaceutical compositions containing same, and methods for producing same | |
| KR20180026734A (en) | Stabilized soluble fusion-pre-RSV F polypeptide | |
| WO2018144784A1 (en) | MONOMERIC HUMAN IgG1 Fc AND BISPECIFIC ANTIBODIES | |
| CN108727486A (en) | Long-acting nerve growth factor, preparation method and combinations thereof | |
| WO2025133290A1 (en) | Protein for immune regulation | |
| JPH08333394A (en) | Rat obesity gene, its gene product and its production | |
| JP7796415B2 (en) | Fusion proteins containing erythropoietin polypeptides | |
| TW202528351A (en) | Fc fragment polypeptide for producing therapeutically active fusion polypeptides and conjugates | |
| RU2812283C2 (en) | Designed il-2-fc fusion proteins | |
| KR20240049337A (en) | GDF15 fusion protein and its uses | |
| US20220332847A1 (en) | RECOMBINANT IgG Fc MULTIMERS FOR THE TREATMENT OF IMMUNE COMPLEX-MEDIATED KIDNEY DISORDERS | |
| TW202542190A (en) | Fusion proteins comprising acid alpha-glucosidase enzymes and methods thereof | |
| HK40061872B (en) | Insulin-fc fusion protein and application thereof | |
| CN119968390A (en) | Pig antibody mutant | |
| KR20220008788A (en) | Fusion protein comprising glp-1 and glp-2 and use thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24837031 Country of ref document: EP Kind code of ref document: A1 |