WO2025111030A2 - Use of novel genes for the control of nematode pests - Google Patents
Use of novel genes for the control of nematode pests Download PDFInfo
- Publication number
- WO2025111030A2 WO2025111030A2 PCT/US2024/036850 US2024036850W WO2025111030A2 WO 2025111030 A2 WO2025111030 A2 WO 2025111030A2 US 2024036850 W US2024036850 W US 2024036850W WO 2025111030 A2 WO2025111030 A2 WO 2025111030A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- plant
- nucleic acid
- protein
- cell
- nematode
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8241—Phenotypically and genetically modified plants via recombinant DNA technology
- C12N15/8261—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield
- C12N15/8271—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance
- C12N15/8279—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance for biotic stress resistance, pathogen resistance, disease resistance
- C12N15/8285—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance for biotic stress resistance, pathogen resistance, disease resistance for nematode resistance
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/195—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria
- C07K14/32—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria from Bacillus (G)
- C07K14/325—Bacillus thuringiensis crystal peptides, i.e. delta-endotoxins
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
-
- Y—GENERAL TAGGING OF NEW TECHNOLOGICAL DEVELOPMENTS; GENERAL TAGGING OF CROSS-SECTIONAL TECHNOLOGIES SPANNING OVER SEVERAL SECTIONS OF THE IPC; TECHNICAL SUBJECTS COVERED BY FORMER USPC CROSS-REFERENCE ART COLLECTIONS [XRACs] AND DIGESTS
- Y02—TECHNOLOGIES OR APPLICATIONS FOR MITIGATION OR ADAPTATION AGAINST CLIMATE CHANGE
- Y02A—TECHNOLOGIES FOR ADAPTATION TO CLIMATE CHANGE
- Y02A40/00—Adaptation technologies in agriculture, forestry, livestock or agroalimentary production
- Y02A40/10—Adaptation technologies in agriculture, forestry, livestock or agroalimentary production in agriculture
- Y02A40/146—Genetically Modified [GMO] plants, e.g. transgenic plants
Definitions
- This invention relates to the field of molecular biology. Provided are methods for the control of nematode pests using novel pest genes.
- Plant pests are a major factor in the loss of the world's important agricultural crops. Some estimates claim that 40% of global crop production is lost due to infestations of invertebrate pests including nematodes. In addition to losses in field crops, nematode pests are also a burden to vegetable and fruit growers, to producers of ornamental flowers, and to home gardeners. Plant-infesting nematodes, a majority of which are root feeders, are found in association with most plants. Some are endoparasitic, living and feeding within the tissue of the roots, tubers, buds, seeds, etc. Others are ectoparasitic, feeding externally through plant walls. A single endoparasitic nematode can kill a plant or reduce its productivity. Endoparasitic root feeders include such economically important pests as the root-knot nematodes
- Nematodes damage caused by nematodes can drastically decrease a plant's uptake of nutrients and water. Nematodes have the greatest impact on crop productivity when they attack the roots of seedlings immediately after seed germination. Nematode feeding also creates open wounds that provide entry to a wide variety of plant-pathogenic fungi and bacteria. These microbial infections can be more economically damaging than the direct effects of nematode feeding.
- Cyst nematodes are responsible for direct loss in soybean yield and indirect loss due to cost of pesticides and non-optimal use of land for crop rotation.
- Soybean cyst nematode Heterodera glycines
- Economically significant densities of cyst nematodes usually cause stunting of crop plants. The stunted plants have smaller root systems, show symptoms of mineral deficiencies in their leaves, and wilt easily.
- Traditional practices for managing nematode infestations include maintaining proper fertility and soil pH levels in nematode- infested land; controlling plant diseases that aid nematode invasion, as well as controlling insect and weed pests; using sanitation practices such as plowing, planting, and cultivating of nematode-infested fields only after working non-infested fields; cleaning equipment thoroughly after working in infested fields; not using seed from plants grown on infested land for planting non-infested fields unless the seed has been properly cleaned; rotating infested fields and alternating host crops with non-host crops, such as, corn, oat and alfalfa and planting resistant or tolerant plant varieties.
- Nematodes are difficult pests to control without the use of chemical pesticides or fumigants (e.g., nematicides) and these approaches come with many challenges such as nematode resistance, regulatory restrictions, etc.
- transgenic plants and/or plant parts expressing the polypeptides of the one or more embodiments were found to be capable of inhibiting the ability of nematode pests to survive, grow and reproduce, or of limiting nematode-related damage or loss to crop plants.
- transgenic nematode-resistant plants which express any one or more of said proteins and to methods of using the transgenic plants alone or in combination with other nematode control strategies to confer maximal nematode control efficiency with minimal environmental impact.
- Plants and plant parts expressing the proteins described herein are highly tolerant or resistant to nematode infestation.
- a method of controlling a nematode pest comprising contacting the nematode pest with a protein comprising any one of SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof is provided for.
- the protein can be a Cry21, Cry21 Aa, Cry21Aa2 or Cry21 Aal protein, for example as demonstrated by, SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, or a nematode-active fragment thereof.
- Cry21, Cry21 Aa2 or Cry21 Aal protein homologue having at least about 40%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95% identity with SEQ ID NOs: 1, 7, 9, 13, 15, 21 , 23, 33, 35, 39, 41 , 51 , 53, and any functional fragments thereof.
- a method of controlling a nematode pest comprising contacting the nematode pest with a transgenic plant or plant part comprising a heterologous nucleic acid molecule that directs expression of a Cry21 protein of the invention in the transgenic plant or plant part, wherein the transgenic plant or plant part controls the nematode pest compared to a plant or plant part of the same type that does not express the Cry21 protein is provided for.
- the nematode pest is selected from the group consisting of Criconemella, Ditylenchus, Globodera, Helicotylenchus, Heterodera, Longidorus, Meloidogyne, Paratrichodorus, Pratylenchus, Radolpholus, Rotelynchus, Rotylenchulus, Tylenchulus and Xiphinema.
- Such nematode pests selected from these genera can be cyst- forming nematodes.
- the cyst-forming nematodes are in the genus Heterodera.
- the nematode pest is Heterodera glycines (soybean cyst nematode).
- the nematode pest is Pratylenchus brachyurus but not SCN and/or both.
- the transgenic plant or plant part is selected from the group consisting of alfalfa, apple, apricot, Arabidopsis, artichoke, asparagus, avocado, banana, barley, beans, beet, blackberry, blueberry, Brassica, broccoli, Brussels sprouts, cabbage, canola, carrot, cassava, cauliflower, a cereal, celery, cherry, citrus, Clementine, coffee, com, cotton, cucumber, eggplant, endive, eucalyptus, figs, grape, grapefruit, groundnuts, ground cherry, kiwifruit, lettuce, leek, lemon, lime, pine, maize, mango, melon, millet, mushroom, nut oat, okra, onion, orange, an ornamental plant or flower or tree, papaya, parsley, pea, peach, peanut, peat, pepper, persimmon, pineapple, plantain, plum, pomegranate, potato, pumpkin, radicchio, radish,
- the plant part is a root.
- the root is a soybean root.
- the Ciy21 protein is a Cry21 Aa, Cry21Aal, Cry21 Aa2, r Cry21Ba protein.
- the Cry21 protein comprises an amino acid sequence that is the translation product of a nucleotide sequence whose complement hybridizes to SEQ ID NOs: 2, 8, 10, 14, 16, 22, 24, 34, 36, 40, 42, 52, or 54 under high-stringency conditions.
- a Cry21 protein of the invention comprises SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof or a nematode-active homologue thereof having at least 60%, 65%, 70%, 75%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99 sequence identity with SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof.
- the Cry21 protein comprises SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof or a nematode-active homologue having at least 60%, 65%, 70%, 75% 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% sequence identity with SEQ ID NOs: 1, 7, 9, 13, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof.
- a transgenic plant of the invention further comprises or expresses at least one additional pesticidal agent, for example without limitation, a patatin, a Bacillus thuringiensis insecticidal protein, a Bacillus thuringiensis nematicidal protein, a Xenorhabdus insecticidal protein, a Photorhabdus insecticidal protein, a Bacillus laterosporous insecticidal protein, a Bacillus sphearicus insecticidal protein, a vegetative insecticidal protein, VLP3 and/or an RNAi molecule that targets a nematode pest.
- additional pesticidal agent for example without limitation, a patatin, a Bacillus thuringiensis insecticidal protein, a Bacillus thuringiensis nematicidal protein, a Xenorhabdus insecticidal protein, a Photorhabdus insecticidal protein, a Bacillus laterosporous insecticid
- a Bacillus thuringiensis nematicidal protein is selected from the group consisting of a Cry5, a Cry6, a Cryl 3, a Cryl 4, a, Cry21 , and a Cry55.
- a method of conferring nematode resistance to a plant and/or a plant part comprising inserting in the plant and/or a plant part a heterologous nucleic acid molecule encoding a Cry21 protein, wherein the plant and/or plant part expresses the Cry21 protein at a nematode-inhibiting level so as to confer nematode resistance to the plant and/or plant part compared to the same type of plant and/or plant part not expressing the Cry21 protein is provided for.
- Such insertion may occur via transformation, gene editing or through breeding.
- a Cry21 protein comprises SEQ ID NO: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof or a Pratylenchus brachyurus nematode- active homologue thereof having at least 60%, 65%, 70%, 75%, 80%, 81 %, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99 sequence identity with SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof.
- the Cry21 protein comprises SEQ ID NOs: 1, 7, 9, 13, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof or a Pratylenchus brachyurus nematode-active homologue having at least 60%, 65%, 70%, 75% 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99% sequence identity with SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof.
- a method of reducing nematode infectivity to a plant and/or plant part comprising contacting the nematode with a Cry21 protein, wherein nematode infectivity is reduced compared to infectivity of a plant and/or plant part by a nematode not contacted with a Cry21 protein is provided for.
- transgenic soybean plant or plant part thereof comprising a heterologous nucleic acid molecule encoding a Cry 21 protein, wherein the transgenic soybean plant or plant part is resistant to nematode infestation is provided for.
- a method of producing a soybean plant protected against nematode infestation comprising transforming a soybean plant cell with a nucleic acid molecule encoding a Cry21 protein; and regenerating a transformed soybean plant from the soybean plant cell, wherein the transformed plant is protected against nematode infestation is provided for.
- a method of producing a soybean plant protected against nematode infestation comprising crossing a first parent soybean plant with a second parent soybean plant, wherein the first or second parent soybean plant comprises a heterologous nucleic acid molecule encoding a Cry21 protein of the invention, thereby producing a plurality of progeny plants; and selecting from the plurality of progeny plants, a transgenic plant that is protected against nematode infestation is provided for.
- Another aspect provides for a method of reducing nematode cyst development on roots of a plant infectable by a nematode is provided, comprising introducing into cells of the root a nucleic acid molecule capable of directing the expression of a Cry21 protein, thereby reducing nematode cyst development on roots of the plant.
- a method of reducing nematode cyst development on roots of a plant infectable by a nematode comprising introducing into plant cells a nucleic acid molecule capable of directing the expression of a Cry 21 protein to the roots, thereby reducing nematode cyst development on roots of the plant
- Another aspect provided for a method of controlling or preventing nematode growth comprising providing a nematode pest with plant material comprising a heterologous DNA capable of directing expression of a Cry21 protein, wherein said plant inhibits a nematode biological activity.
- Another aspect provides for a method of providing a grower with a means of controlling nematode pests is provided, the method comprising supplying seed to a grower, wherein the seed comprises a heterologous nucleic acid molecule that encodes a Cry21 protein and wherein the seed is capable of producing a plant that is resistant to nematode infestation.
- Another aspect provides for a method of suppressing growth of a plant-pathogenic nematode population in a location capable of supporting growth of the nematode population comprising growing in the location a population of transgenic soybean plants comprising a heterologous nucleic acid molecule capable of directing expression of a Cry21 protein, wherein growth of the plant-pathogenic nematode population is suppressed.
- Another aspect provides for a method of controlling any one of soy cyst nematode, lesion nematode, root knot nematode or reniform nematode (“target pest”) comprising providing a transgenic soybean plant comprising an expression cassette having any one of SEQ ID Nos: 2, 8, 10, 14, 16, 22, 24, 34, 36, 40, 42, 52, or 54 operably linked to a promoter capable of driving expression of an encoded Cry21 protein to levels sufficient to inhibit nematodes, wherein the proliferation of target pest feeding on the soybean plant is reduced compared to target pest feeding on a non-transgenic soybean plant not comprising the expression cassette.
- Another aspect provides for a method of improving plant yield in nematode infested fields, comprising expressing in the plant a Cry21 protein, wherein plant yield is improved compared to yield of a plant of the same type not expressing a Cry 21 protein.
- Another aspect provides for a method of increasing the vigor or yield in a transgenic soybean plant exposed to a population of nematodes comprising: introgressing a transgenic soybean event into a soybean plant resulting in a transgenic soybean plant, wherein the transgenic soybean event comprises a heterologous DNA sequence encoding a Cry 21 protein that confers upon the transgenic soybean event resistance to nematodes; and growing the transgenic soybean plant or progeny thereof at a location where nematode infestation is yield limiting to a soybean plant not comprising the heterologous nucleic acid molecule encoding the Cry21 protein, whereby the transgenic soybean plant has increased vigor or yield compared to the control plant.
- Another aspect provides for a method of improving yield of a soybean field, comprising: introducing into a soybean plant a nucleic acid molecule capable of directing expression of a Cry 21 protein, thus producing a transgenic plant; and cultivating a plurality of transgenic seeds from the transgenic plant in a field producing or resulting in a soybean field comprising a plurality of transgenic soybean plants having enhanced resistance to nematode infestation, thereby improving yield of the soybean field.
- Another aspect provides for a recombinant expression cassette comprising a heterologous promoter sequence operatively linked to a nucleic acid molecule encoding a Ciy21 protein. Further, the present invention provides a recombinant vector comprising such an expression cassette. Still further, another aspect provides for a transgenic host cell comprising such an expression cassette. A transgenic host cell according to this aspect may be a plant cell. Even further, another aspect provides for a transgenic plant or plant part comprising such a plant cell.
- a nematicidal composition comprising an effective nematode-controlling amount of a Cry21 protein and an acceptable agricultural carrier.
- the agricultural carrier is a transgenic plant.
- the transgenic plant is a transgenic soybean plant and the Cry21 protein is a Cry21 protein having at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, or 99% sequence identity with SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof.
- the Cry21 protein comprises either SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53 and any functional fragments thereof.
- a further aspect provides for a method of producing a nematode-resistant transgenic plant, comprising introducing a nucleic acid molecule encoding a Cry 21 protein into a plant cell thereby making a transgenic plant cell; regenerating a transgenic plant from the transgenic plant cell, wherein the Cry21 protein is expressible in the transgenic plant in an effective amount to control nematodes.
- the plant is a soybean plant.
- the Cry21 protein is a protein having at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 97%, or 99% sequence identity with SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof .
- the Cry21 protein comprises SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof.
- the nematode is a target pest.
- the nematode is Heterodera glycines, Pratylenchus brachyurus, Rotylenchulus reniformis, or Meloidogyne sp.
- Another aspect comprises a method of controlling a nematode pest, comprising contacting the nematode pest with a Cry 21 protein comprising an amino acid sequence having 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 98%, 99%, 100% sequence identity to any one of SEQ ID NOs: 1, 3, 7, 9, 11, 13, 15, 17, 21, 23, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53 or a nematicidally-effective variant or fragment thereof.
- Another aspect includes a method of claim 40, wherein the nematode pest is selected from any one of the species consisting of;
- Criconemella Ditylenchus, Globodera, Helicotylenchus, Heterodera, Longidorus, Meloidogyne, Paratrichodorus, Pratylenchus, Radolpholus, Rotelynchus, Rotylenchulus, Tylenchulus and Xiphinema.
- Another aspect includes an expression cassette comprising a nucleic acid encoding a Cry21 protein comprising a amino acid sequence having 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 98%, 99%, 100% sequence identity to any one of SEQ ID NOs: , 3, 7, 9, 11, 13, 15, 17, 21, 23, 25, 31, 33, 35, 37, 39, 41, 43, 45, 47, 49, 51, 53 or a nematicidally-effective variant or fragment thereof and wherein the nucleic acid is operably linked to said nucleic acid in the expression cassette and/or is further in a plant expression vector.
- a further embodiment includes a nucleic acid comprises a nucleic acid sequence having sequence having 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 98%, 99%, 100% sequence identity to any one of SEQ ID NOs: 2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26, 28, 30, 32, 34, 36, 38, 40, 42, 44, 46, 48, 50 52, or 54.
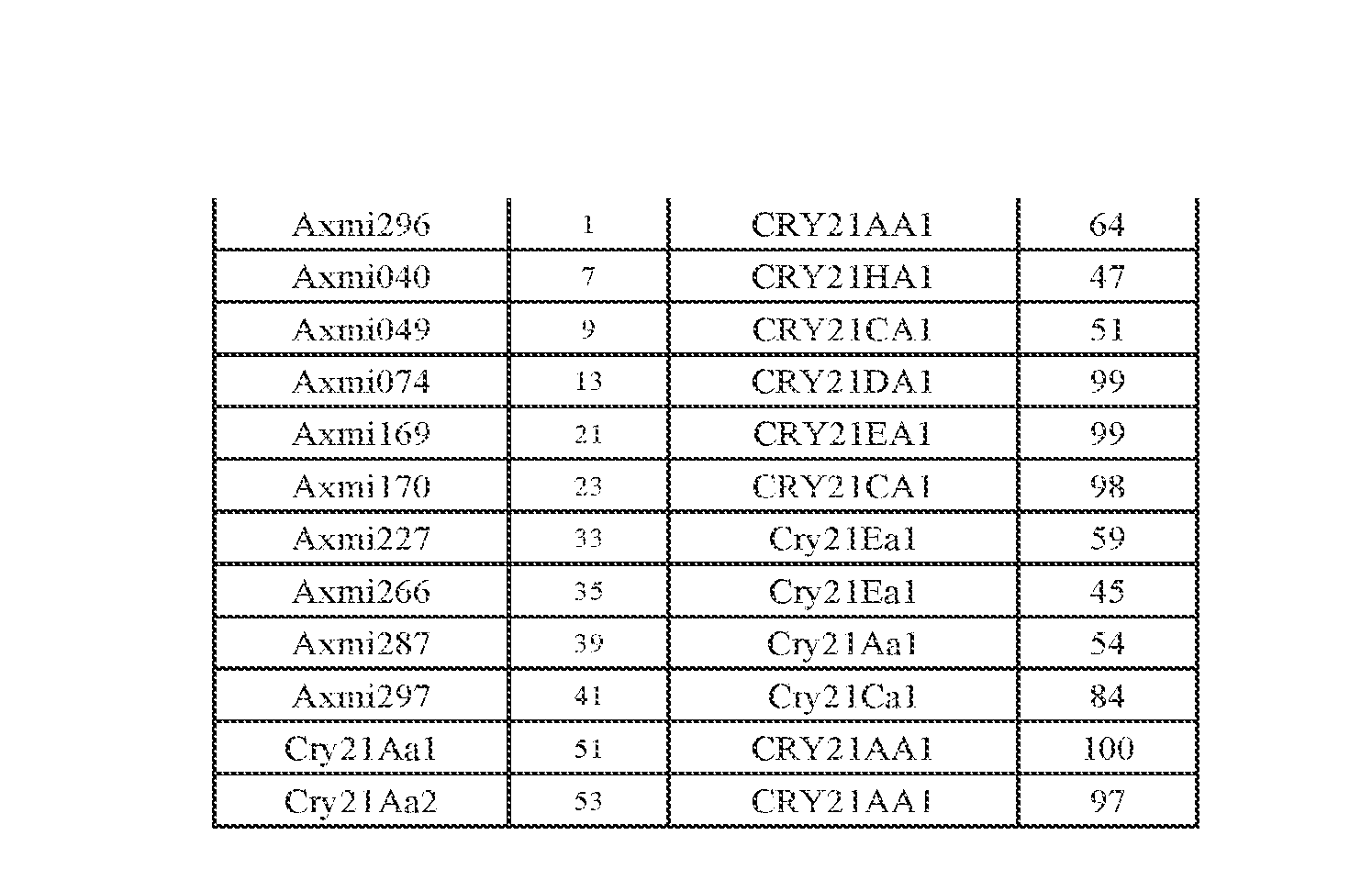
- SEQ ID NO: 1 discloses the amino acid sequence of a Cry21 homolog, Axmi296.
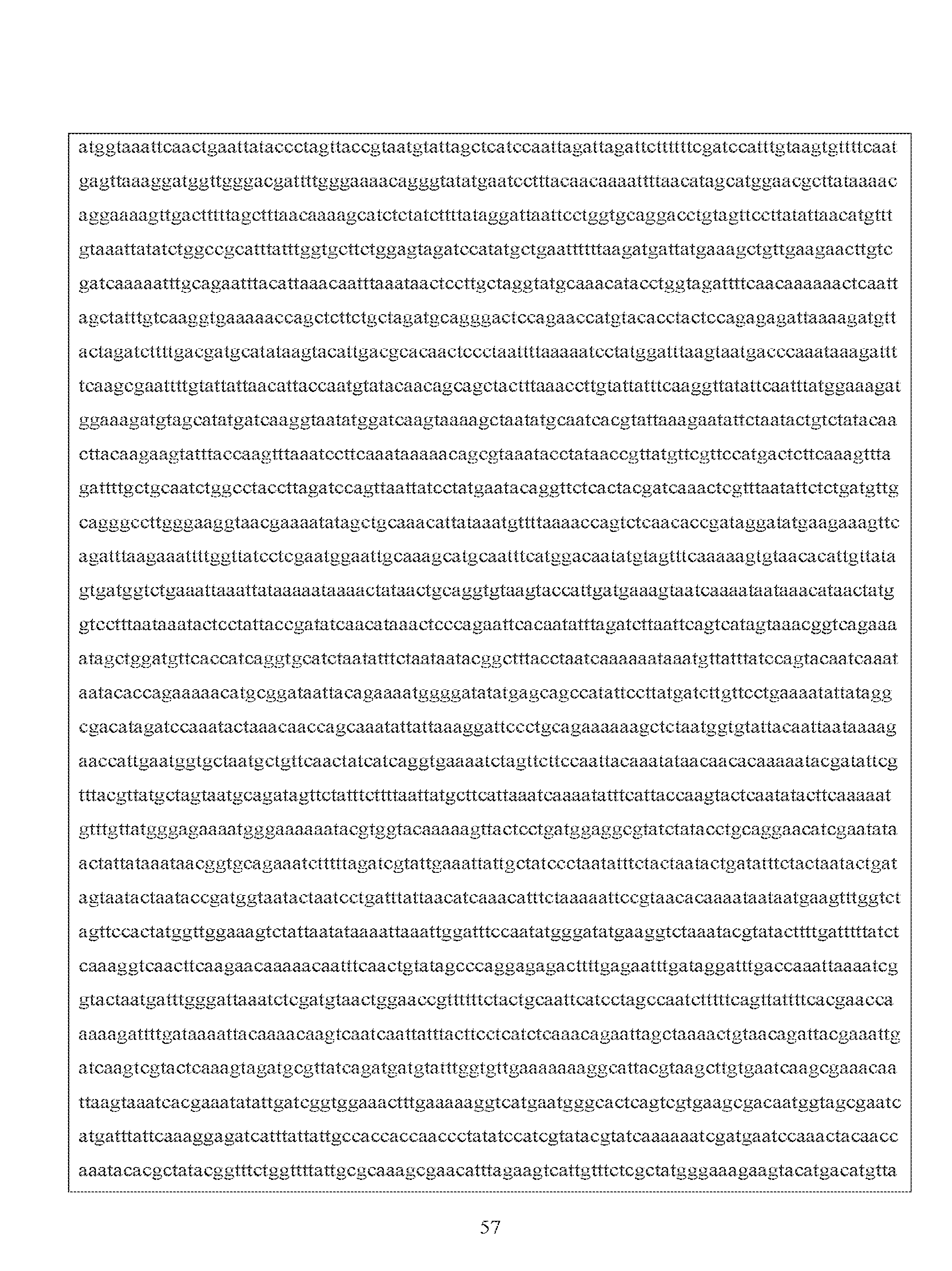
- SEQ ID NO:2 discloses the nucleotide sequence of a Cry21homolog, Axmi296.
- SEQ ID NO:3 discloses the amino acid sequence of AxmiOlS.
- SEQ ID NO:4 discloses the nucleotide sequence of AxmiOl 8.
- SEQ ID NO: 5 discloses the amino acid sequence of Axmi021.
- SEQ ID NO:6 discloses the nucleotide sequence of Axmi021.
- SEQ ID NO:7 discloses the amino acid sequence of Axmi040.
- SEQ ID NO: 8 discloses the nucleotide sequence of Axmi040.
- SEQ ID NO:9 discloses the amino acid sequence of Axmi049.
- SEQ ID NO: 10 discloses the nucleotide sequence of Axmi049.
- SEQ ID NO: 11 discloses the amino acid sequence of Axmi071.
- SEQ ID NO: 12 discloses the nucleotide sequence of Axmi071.
- SEQ ID NO: 13 discloses the amino acid sequence of Axmi074.
- SEQ ID NO: 14 discloses the nucleotide sequence of Axmi074.
- SEQ ID NO: 15 discloses the amino acid sequence of AxmiOSS.
- SEQ ID NO: 16 discloses the nucleotide sequence of Axmi088.
- SEQ ID NO:17 discloses the amino acid sequence of Axmi 104.
- SEQ ID NO: 18 discloses the nucleotide sequence of Axmi 104.
- SEQ ID NO: 19 discloses the amino acid sequence of Axmi 155.
- SEQ ID NO:20 discloses the nucleotide sequence of Axmil 55.
- SEQ ID NO:21 discloses the amino acid sequence of Axmi 169.
- SEQ ID NO:22 discloses the nucleotide sequence of Axmil 69.
- SEQ ID NO:23 discloses the amino acid sequence of Axmi 170.
- SEQ ID NO:24 discloses the nucleotide sequence of Axmi 170.
- SEQ ID NO:25 discloses the amino acid sequence of Axmi215.
- SEQ ID NO:26 discloses the nucleotide sequence of Axmi215.
- SEQ ID NO: 27 discloses the amino acid sequence of Axmi218.
- SEQ ID NO:28 discloses the nucleotide sequence of Axmi218.
- SEQ ID NO:29 discloses the amino acid sequence of Axmi219.
- SEQ ID NO:30 discloses the nucleotide sequence of Axmi219.
- SEQ ID NO: 31 discloses the amino acid sequence of Axmi220.
- SEQ ID NO:32 discloses the nucleotide sequence of Axmi220.
- SEQ ID NO: 33 discloses the amino acid sequence of Axmi227.
- SEQ ID NO:34 discloses the nucleotide sequence of Axmi227.
- SEQ ID NO:35 discloses the amino acid sequence of Axmi266.
- SEQ ID NO:36 discloses the nucleotide sequence of Axmi266.
- SEQ ID NO:37 discloses the amino acid sequence of Axmi280.
- SEQ ID NO:38 discloses the nucleotide sequence of Axmi280.
- SEQ ID NO:39 discloses the amino acid sequence of Axmi287.
- SEQ ID NO:40 discloses the nucleotide sequence of Axmi287.
- SEQ ID NO:41 discloses the amino acid sequence of Axmi297.
- SEQ ID NO:42 discloses the nucleotide sequence of Axmi297.
- SEQ ID NO:43 discloses the amino acid sequence of Axmi299.
- SEQ ID NO:44 discloses the nucleotide sequence of Axmi299.
- SEQ ID NO:45 discloses the amino acid sequence of Axmi340.
- SEQ ID NO:46 discloses the nucleotide sequence of Axmi340.
- SEQ ID NO:47 discloses the amino acid sequence of Axmi350.
- SEQ ID NO:48 discloses the nucleotide sequence of Axmi350.
- SEQ ID NO:49 discloses the amino acid sequence of Axmi380.
- SEQ ID NO:50 discloses the nucleotide sequence of Axmi380.
- SEQ ID NO: 51 discloses the amino acid sequence of native Cry21 Aal.
- SEQ ID NO:52 discloses the nucleotide sequence of native Cry21Aal.
- SEQ ID NO:53 discloses the amino acid sequence of native Cry21Aa2.
- SEQ ID NO: 54 discloses the protein sequence of native Cry21Aa2.
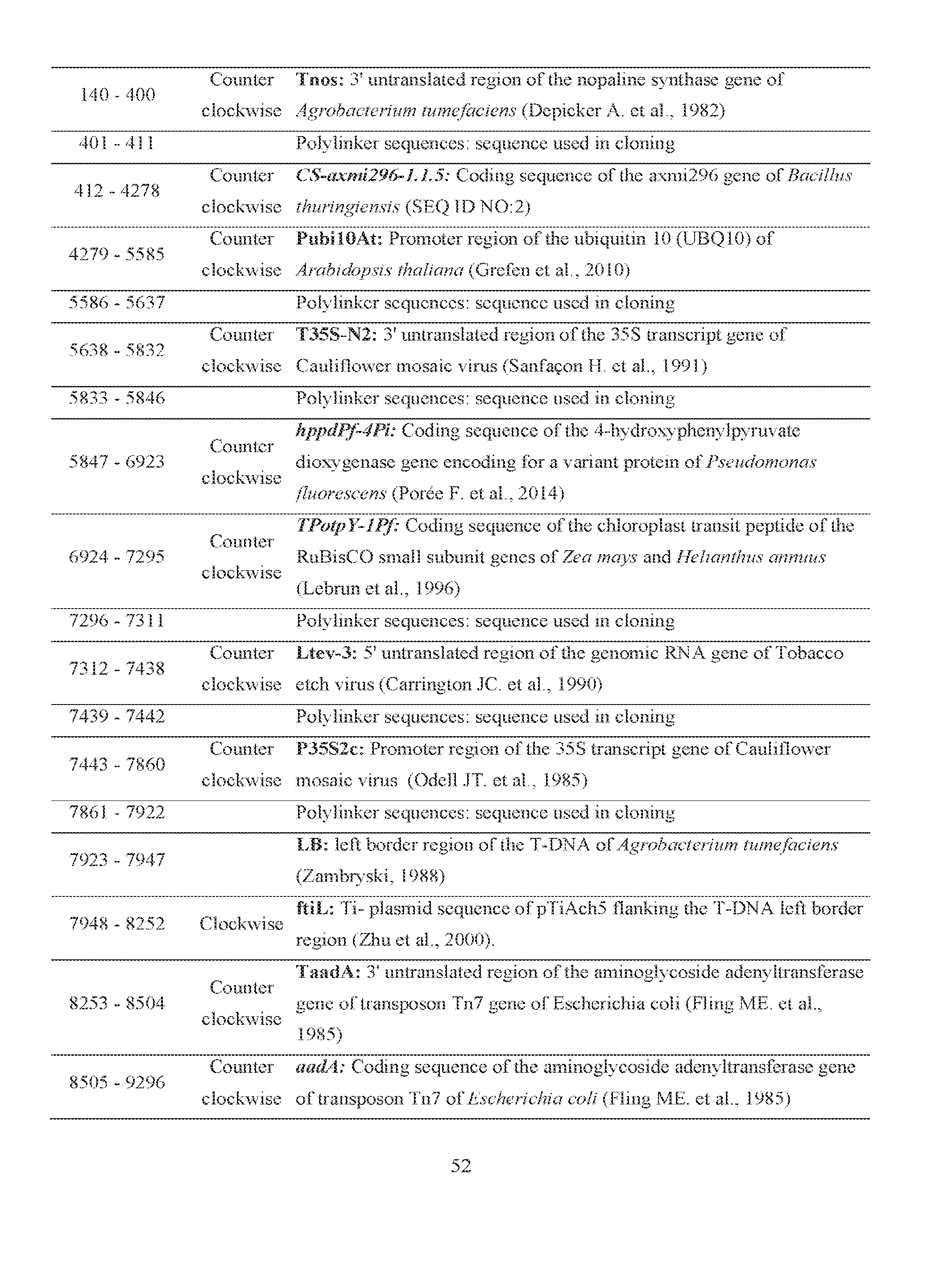
- FIG. 1 depicts the plant transformation vector for the expression of Axmi296 in planta.
- FIG. 2 depicts a NEEDLE Sequence alignment of Axmi296 and Cry21 Aa2.
- FIGs. 3a and 3b depict the effectiveness of Axmi296 against Pratylenchus brachyurus in 60-day greenhouse assays.
- FIGs. 4a and 4b depict results showing Axmi296 provides increased yield benefit in Soybean Cyst Nematode (SCN) field trials over SCN non-resistant controls.
- SCN Soybean Cyst Nematode
- FIG. 4c depicts results from SCN field trials comparing nematode reduction between the novel Cry21 homolog (Axmi296) with wild-type Thome.
- FIG. 4d depicts results from SCN field trials comparing yield between the novel Cry21 homolog (Axmi296) with wild-type Thome.
- FIG. 5 depicts a photo of field showing Axmi296 plants (left) growing next to non- resistant controls soybean plants (right) in an SCN-infested field.
- Cry21 proteins of the invention have a toxic effect on nematodes by disrupting or deleting feeding, inhibiting the ability of nematode pests to survive, grow and reproduce which may or may not cause death of the nematode, or of limiting nematode-related damage or loss to crop plants.
- Associated with/operatively linked refer to two nucleic acid sequences that are related physically or functionally.
- a promoter or regulatory DNA sequence is said to be “associated with” a DNA sequence that codes for an RNA or a protein if the two sequences are operatively linked or situated such that the regulator DNA sequence will affect the expression level of the coding or structural DNA sequence.
- the term “contacting” refers to a process by which a Cry21 protein of the invention or transgenic plant or plant part expressing the Cry21 protein of the embodiments or aspects are delivered or administered to target nematode pests or nematode pest populations. Contacting describes physical proximity of Cry21 proteins or transgenic plants or plant parts expressing a Cry 21 protein and the target nematode so that they interact.
- the transgenic plants or plant parts may be contacted with a target nematode or nematode population by planting transgenic seed, transgenic seedlings, cuttings, plant runners, tubers, and the like in a location capable of supporting growth of a nematode pest or nematode pest population.
- a “chimeric gene” is a recombinant nucleic acid sequence in which a promoter or regulatory nucleic acid sequence is operatively linked to, or associated with, a nucleic acid sequence that codes for an mRNA or which is expressed as a protein, such that the regulator nucleic acid sequence is able to regulate transcription or expression of the associated nucleic acid sequence.
- the regulator nucleic acid sequence of the chimeric gene is not normally operatively linked to the associated nucleic acid sequence as found in nature.
- a “coding sequence” is a nucleic acid sequence that is transcribed into RNA such as mRNA, rRNA, tRNA, snRNA, sense RNA or antisense RNA. The RNA is then translated in an organism to produce a protein.
- control means to inhibit, through a toxic effect, the ability of nematode pests to survive, grow, feed, and/or reproduce, or to limit nematode-related damage or loss in crop plants.
- control may or may not mean killing the nematodes.
- nucleic acid coding sequences or amino acid sequences of different Cry21 genes or proteins are aligned with each other, the nucleic or amino acids that “correspond to” certain enumerated positions are those that align with these positions but that are not necessarily in these exact numerical positions relative to the particular Cry21's respective nucleic acid coding sequence or amino acid sequence.
- a toxin means that the toxin comes in contact with a nematode or nematode population, resulting in a toxic effect and control of the nematode or nematode population.
- the toxin can be delivered in many recognized ways, e.g., orally by ingestion by the nematode or by contact with the nematode via transgenic plant expression, formulated protein compositions), sprayable protein composition(s), a bait matrix, or any other art-recognized toxin delivery system.
- economic threshold is defined as the pest nematode population that produces incremental damage equal to the cost of controlling or preventing that damage. It is the level of nematode population where the benefit of nematode control is equal to its cost.
- economic threshold may be defined as the nematode pest damage level where the value of incremental reduction in crop yield is equal to the cost of preventing its occurrence. In other words, economic threshold attempts to determine the point at which it becomes economically feasible to control a nematode pest population.
- a “nematode-controlling effective amount” or interchangeably a “pesticidal amount” as used herein refers to the concentration of a Cry 21 protein or functional fragment thereof capable of inhibiting, through a toxic effect, the ability of nematodes to survive, grow, feed and/or reproduce, or of reducing or preventing nematode-related damage or loss in crop plants. “Nematode-controlling effective amount” may or may not mean killing the nematodes.
- “Expression cassette” as used herein means a nucleic acid sequence capable of directing expression of a particular nucleotide sequence in an appropriate host cell, comprising a promoter operably linked to the nucleotide sequence of interest which is operably linked to termination signals. It also typically comprises sequences required for proper translation of the nucleotide sequence.
- the expression cassette comprising the nucleotide sequence of interest may be chimeric, meaning that at least one of its components is heterologous with respect to at least one of its other components.
- the expression cassette may also be one that is naturally occurring but has been obtained in a recombinant form useful for heterologous expression.
- the expression cassette is heterologous with respect to the host, i.e., the particular nucleic acid sequence of the expression cassette does not occur naturally in the host cell and must have been introduced into the host cell or an ancestor of the host cell by a transformation event
- the expression of the nucleotide sequence in the expression cassette may be under the control of a constitutive promoter or of an inducible promoter that initiates transcription only when the host cell is exposed to some particular external stimulus.
- the promoter can also be specific to a particular tissue, or organ, or stage of development.
- a “gene” is a defined region that is located within a genome and that, besides the aforementioned coding nucleic acid sequence, comprises other, primarily regulatory, nucleic acid sequences responsible for the control of the expression, that is to say the transcription and translation, of the coding portion.
- a gene may also comprise other 5' and 3' untranslated sequences and termination sequences. Further elements that may be present are, for example, introns.
- Nematicidal is defined as a toxic biological activity capable of controlling nematodes, preferably by killing them.
- a nucleic acid sequence is “isocoding with” a reference nucleic acid sequence when the nucleic acid sequence encodes a polypeptide having the same amino acid sequence as the polypeptide encoded by the reference nucleic acid sequence.
- a native coding sequence from Bacillus spp. that encodes a Cry21 protein is isocoding with a coding sequence codon optimized for expression in a plant that encodes the same Cry21 protein.
- An “isolated” nucleic acid molecule or an isolated protein or toxin is a nucleic acid molecule or protein or toxin that, by the hand of man, exists apart from its native environment and is therefore not a product of nature.
- An isolated nucleic acid molecule or protein or toxin may exist in a purified form or may exist in a non-native environment such as, for example, a recombinant host cell or a transgenic plant.
- the term “native” refers to a coding sequence or gene that is naturally present in the genome of a cell or plant.
- Naturally occurring is used herein to describe an object that can be found in nature as distinct from being artificially produced by man.
- a protein or nucleotide sequence present in an organism which can be isolated from a source in nature and which has not been intentionally modified by man in the laboratory, is naturally occurring.
- a “plant” is any plant at any stage of development, particularly a seed plant.
- a “plant cell” is a structural and physiological unit of a plant, comprising a protoplast and a cell wall.
- the plant cell may be in the form of an isolated single cell or a cultured cell, or as a part of higher organized unit such as, for example, plant tissue, a plant organ, or a whole plant.
- Plant material refers to leaves, stems, roots, flowers or flower parts, fruits, pollen, egg cells, zygotes, seeds, cuttings, cell or tissue cultures, or any other part or product of a plant.
- a “plant organ” is a distinct and visibly structured and differentiated part of a plant such as a root, stem, leaf, flower bud, or embryo.
- a “plant part” may be any part of a plant and include a plant cell, plant material, plant organ or plant tissue.
- Plant tissue as used herein means a group of plant cells organized into a structural and functional unit Any tissue of a plant in planta or in culture is included. This term includes, but is not limited to, whole plants, plant organs, plant seeds, tissue culture and any groups of plant cells organized into structural and/or functional units. The use of this term in conjunction with, or in the absence of, any specific type of plant tissue as listed above or otherwise embraced by this definition is not intended to be exclusive of any other type of plant tissue.
- a “promoter” is an untranslated DNA sequence upstream of the coding region that contains the binding site for RNA polymerase II and initiates transcription of the DNA.
- the promoter region may also include other elements that act as regulators of gene expression.
- Regulatory elements refer to sequences involved in controlling the expression of a nucleotide sequence. Regulatory elements comprise a promoter operably linked to the nucleotide sequence of interest and termination signals. They also typically encompass sequences required for proper translation of the nucleotide sequence.
- resistant means a transgenic soybean variety that prevents a majority of nematodes from surviving and/or reproducing upon their attempted infestation.
- substantially identical in the context of two nucleic acid or protein sequences, refers to two or more sequences or subsequences that have at least 60%, at least 80%, at least 90%, at least 95%, and at least 99% nucleotide or amino acid residue identity, when compared and aligned for maximum correspondence, as measured using one of the following sequence comparison algorithms or by visual inspection.
- the substantial identity exists over a region of the sequences that is at least about 50 residues in length, over a region of at least about 100 residues, or the sequences are substantially identical over at least about 150 residues.
- the sequences are substantially identical over the entire length of the coding regions.
- substantially identical nucleic acid or protein sequences perform substantially the same function.
- a pairwise sequence alignment is generated between those two sequences. Pairwise alignments in this first step can be generated by various tools known to a person skilled in the art, like e.g. programs “Blast” (Altschul etal. J. Mol. Biol. 215:403-410), “Blast2” (“gapped Blast”) (Altschul et al, Nucleic Acids Res. 25:3389-3402.), programs from The European Molecular Biology Open Software Suite (EMBOSS, Trends in Genetics 16 (6), 276 (2000)) like “Water”, “Matcher” or “Needle”, or by visual inspection.
- Blast Altschul etal. J. Mol. Biol. 215:403-410
- Blast2 gapped Blast
- EMBOSS European Molecular Biology Open Software Suite
- a percent-identity value can be determined from the alignment produced. Percent-identity between the two sequences can be calculated from the complete alignment produced, or from a region out of the alignment, e.g. the region of the alignment showing the sequence of this invention over its complete length, or the region showing the other sequence over its complete length, or from a region showing only parts of the sequences.
- the alignment region from which a percent-identity value is calculated preferably has a length of at least 100 positions, more preferably of at least 150 positions, and most preferably of more than 200 positions.
- percent-identity For determination of percent-identity, first the sum over all positions is calculated, in which both sequences are showing identical residues in the alignment region, and this sum is then divided by the length of the alignment region, whereby positions in which a sequence has an introduced gap are either component of said length (length of alignment region), or are subtracted from said length (length of alignment region - total number of gaps in alignment region). The obtained value is then multiplied with 100 to result in percent-identity (% identity).
- Needleman and Wunsch J. Mol. Biol. (1979) 48, p. 443-453
- Percent-identity is then determined from the complete alignment produced and is calculated as follows: percent-identity — (sum of positions showing identical residues in alignment x 100) / ( length of alignment - total number of gaps in alignment ). This value can also be obtained directly from EMBOSS program “Needle” as program labeled “longest identity” when the parameter option “-nobrief* is applied. [00134] For nucleotide sequences encoding for a protein the pairwise alignment shall be made over the complete length of the coding region of the sequence of one or more embodiments from start to stop codon excluding introns. Introns present in the other sequence may also be removed for the pairwise alignment to allow comparison with the sequence of this invention.
- hybridizing specifically to refers to the binding, duplexing, or hybridizing of a molecule only to a particular nucleotide sequence under stringent conditions when that sequence is present in a complex mixture (e.g., total cellular) DNA or RNA.
- Bod(s) substantially refers to complementary hybridization between a probe nucleic acid and a target nucleic acid and embraces minor mismatches that can be accommodated by reducing the stringency of the hybridization media to achieve the desired detection of the target nucleic acid sequence.
- “Stringent hybridization conditions” and “stringent hybridization wash conditions” in the context of nucleic acid hybridization experiments such as Southern and Northern hybridizations are sequence dependent and are different under different environmental parameters. Longer sequences hybridize specifically at higher temperatures. An extensive guide to the hybridization of nucleic acids is found in Tijssen (1993J Laboratory Techniques in Biochemistry and Molecular Biology-Hybridization "with Nucleic Acid Probes part I chapter 2 “Overview of principles of hybridization and the strategy of nucleic acid probe assays” Elsevier, New York.
- highly stringent hybridization and wash conditions are selected to be about 5° C. lower than the thermal melting point (Tm) for the specific sequence at a defined ionic strength and pH.
- Tm thermal melting point
- a probe will hybridize to its target subsequence, but to no other sequences.
- a nucleic acid molecule is said to be the “complement” of another nucleic acid molecule if they exhibit complete complementarity.
- molecules are said to exhibit “complete complementarity” when every nucleotide of one of the molecules is complementary to a nucleotide of the other.
- Two molecules are said to be “minimally complementary” if they can hybridize to one another with sufficient stability to permit them to remain annealed to one another under at least conventional “low-stringency” conditions.
- the molecules are said to be “complementary” if they can hybridize to one another with sufficient stability to permit them to remain annealed to one another under conventional “high-stringency” conditions.
- the Tm is the temperature (under defined ionic strength and pH) at which 50% of the target sequence hybridizes to a perfectly matched probe.
- Very stringent conditions are selected to be equal to the Tm for a particular probe.
- An example of stringent hybridization conditions for hybridization of complementary nucleic acids which have more than 100 complementary residues on a filter in a Southern or northern blot is 50% formamide with 1 mg of heparin at 42° C., with the hybridization being carried out overnight.
- An example of highly stringent wash conditions is 0.15M NaCl at 72° C. for about 15 minutes.
- An example of stringent wash conditions is a 0.2xSSC wash at 65° C. for 15 minutes (see, Sambrook, infra, for a description of SSC buffer).
- a high stringency wash is preceded by a low stringency wash to remove background probe signal.
- An example medium stringency wash for a duplex of, e.g., more than 100 nucleotides, is 1 X SSC at 45° C. for 15 minutes.
- An example low stringency wash for a duplex of, e.g., more than 100 nucleotides, is 4-6*SSC at 40° C. for 15 minutes.
- stringent conditions typically involve salt concentrations of less than about 1.0 M Na ion, typically about 0.01 to 1.0 M Na ion concentration (or other salts) at pH 7.0 to 8.3, and the temperature is typically at least about 30° C.
- Stringent conditions can also be achieved with the addition of destabilizing agents such as formamide.
- destabilizing agents such as formamide.
- a signal to noise ratio of 2x (or higher) than that observed for an unrelated probe in the particular hybridization assay indicates detection of a specific hybridization.
- Nucleic acids that do not hybridize to each other under stringent conditions are still substantially identical if the proteins that they encode are substantially identical. This occurs, e.g., when a copy of a nucleic acid is created using the maximum codon degeneracy permitted by the genetic code.
- a reference nucleotide sequence preferably hybridizes to the reference nucleotide sequence in 7% sodium dodecyl sulfate (SDS), 0.5 M NaPO4, 1 mMEDTA at 50° C. with washing in 2xSSC, 0.1% SDS at 50° C., more desirably in 7% sodium dodecyl sulfate (SDS), 0.5 M NaPO4, 1 mM EDTA at 50° C.
- a further indication that two nucleic acid sequences or proteins are substantially identical is that the protein encoded by the first nucleic acid is immunologically cross reactive with, or specifically binds to, the protein encoded by the second nucleic acid.
- a protein is typically substantially identical to a second protein, for example, where the two proteins differ only by conservative substitutions.
- Synthetic refers to a nucleotide sequence comprising structural characters that are not present in the natural sequence. For example, a Cry21 coding sequence, not naturally found in Bacillus, that resembles more closely the G+C content and the normal codon distribution of dicot and/or monocot genes is said to be synthetic.
- Transformation is a process for introducing heterologous nucleic acid into a host cell or organism.
- transformation means the stable integration of a DNA molecule into the genome of an organism of interest.
- Transformed/transgenic/recombinant refer to a host organism such as a bacterium or a plant into which a heterologous nucleic acid molecule has been introduced.
- the nucleic acid molecule can be stably integrated into the genome of the host or the nucleic acid molecule can also be present as an extrachromosomal molecule. Such an extrachromosomal molecule can be auto-replicating.
- Transformed cells, tissues, or plants are understood to encompass not only the end product of a transformation process, but also transgenic progeny thereof.
- non- transformed refers to a wild-type organism, e.g., a bacterium or plant, which does not contain the heterologous nucleic acid molecule.
- a “Cry21 protein” in the context of the invention means a Bacillus Cry insecticidal protein (that is a member of the Cry21 class including for example without limitation, Cry21Aal, Ciy21Aa2, and their homologues.
- a nematode-active “homologue” as used herein means that the indicated protein or polypeptide is active against nematodes and bears a defined relationship to other members of the Cry21 class of proteins. This defined relationship may include but is not limited to, 1) proteins which are at least 60%, or at least 70%, or at least 80%, or at least 90% identical at the sequence level to another member of the Cry21 class of proteins while also retaining nematicidal activity.
- nucleotides are indicated by their bases by the following standard abbreviations: adenine (A), cytosine (C), thymine (T), and guanine (G)
- Amino acids are likewise indicated by the following standard abbreviations: alanine (Ala; A), arginine (Arg; R), asparagine (Asn; N), aspartic acid (Asp; D), cysteine (Cys; C), glutamine (Gin; Q), glutamic acid (Glu; E), glycine (Gly; G), histidine (His; H), isoleucine (lie; 1), leucine (Leu; L), lysine (Lys; K), methionine (Met; M), phenylalanine (Phe; F), proline (Pro; P), serine (Ser; S), threonine (Thr; T), tryptophan (Trp; W), tyrosine (T
- the materials and methods of the subject invention are useful for killing or controlling nematodes; retarding growth or reproduction of nematodes; reducing nematode populations; and/or reducing or retarding damage to plants caused by infestation of nematode pests.
- One embodiment provides methods of controlling nematode pests of crop plants such as soybean by using transgenic crop plants expressing a Cry 21 protein.
- compositions that can be used to control nematode pests, for example, without limitation, target pest, Meloidogyne spp. (for example, Meloidogyne incoginita and Meloidogyne javanica, Meloidogyne hapla, Meloidogyne arenari), Heterodera spp. (for example, Heterodera glycines, Heterodera carotae, Heterodera schachtii, Heterodora avenae and Heterodora trifolii), Globodera spp.
- target pest for example, without limitation, target pest, Meloidogyne spp.
- Heterodera spp. for example, Heterodera glycines, Heterodera carotae
- Ditylenchus spp. Ditylenchus dipsaci, Dolichodorus spp., Hemicriconemoides spp., Hemicycliophora spp., Hirschmaniella spp., Hypsoperine spp., Macroposthonia spp., Melinius spp., Punctodera spp., Quinisulcius spp., Scutellonema spp., Xiphinema spp , and Tylenchorhynchus spp.
- One embodiment provides for a method of controlling a nematode pest, comprising contacting the nematode pest with a Cry21 protein comprising SEQ ID Nos: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53, and any functional fragments thereof.
- the nematode is selected from the group consisting of Criconemella, Ditylenchus, Globodera, Helicotylenchus, Heterodera, Longidorus, Meloidogyne, Paratrichodorus, Pratylenchus, Radolpholus, Rotelynchus, Rotylenchulus, Tylenchulus and Xiphinema.
- the nematode is a cyst forming nematode.
- the nematode is in the genus Pratylenchus.
- the nematode is Pratylenchus brachyurus.
- the contacting step is carried out with a plant or plant part transformed with at least one nucleic acid molecule encoding the Cry21 protein.
- the plant or plant part is a soybean plant or plant part.
- the soybean plant part is a soybean root
- the transgenic plant or plant part is selected from the group consisting of alfalfa, apple, apricot, Arabidopsis, artichoke, asparagus, avocado, banana, barley, beans, beet, blackberry, blueberry, Brassica, broccoli, Brussels sprouts, cabbage, canola, carrot, cassava, cauliflower, a cereal, celery, cherry, citrus, Clementine, coffee, corn, cotton, cucumber, eggplant, endive, eucalyptus, figs, grape, grapefruit, groundnuts, ground cherry, kiwifruit, lettuce, leek, lemon, lime, pine, maize, mango, melon, millet, mushroom, nut oat, okra, onion, orange, an ornamental plant or flower or tree, papaya, parsley, pea, peach, peanut, peat, pepper, persimmon, pineapple, plantain, plum, pomegranate, potato, pumpkin, radicchio, radish, rap
- transgenic seed of a transgenic plant of one or more embodiments wherein the transgenic seed comprises a heterologous nucleic acid molecule encoding a Cry21 protein of one or more embodiments.
- Another embodiment also encompasses recombinant vectors and expression cassettes comprising the Cry21 nucleic acid sequences of the invention.
- the nucleic acid sequences are comprised in expression cassettes comprising regulatory elements for expression of the Cry21 nucleotide sequences in a transgenic host cell capable of expressing the nucleotide sequences.
- regulatory elements usually comprise promoter and termination signals and preferably also comprise elements allowing efficient translation of polypeptides encoded by the nucleic acid sequences of the present invention.
- Vectors comprising the nucleic acid sequences are usually capable of replication in particular host cells, such as extrachromosomal molecules, and are therefore used to amplify the nucleic acid sequences of this invention in the host cells.
- host cells for such vectors are microorganisms, such as bacteria, in particular E. coll.
- host cells for such recombinant vectors are endophytes or epiphytes.
- a host cell for such vectors is a eukaryotic cell, such as a plant cell. Such plant cells may be soybean cells or maize cells.
- such vectors are viral vectors and are used for replication of the nucleotide sequences in particular host cells, e.g. insect cells or plant cells.
- Recombinant vectors are also used for transformation of the nucleotide sequences of this invention into transgenic host cells, whereby the nucleotide sequences are stably integrated into the DNA of such transgenic host cells.
- transgenic host cells are prokaryotic cells.
- transgenic host cells are eukaryotic cells, such as yeast cells, insect cells, or plant cells.
- the transgenic host cells are plant cells, such as soybean cells or maize cells.
- the Cry21 nucleotide sequences of the invention can be modified by incorporation of random mutations in a technique known as in vitro recombination or DNA shuffling to increase nematode activity for example.
- This technique is described in Stemmer etal., Nature 370:389-391 (1994) and U.S. Pat No. 5,605,793, which are incorporated herein by reference. Millions of mutant copies of a nucleotide sequence are produced based on an original nucleotide sequence of this invention and variants with improved properties, such as increased nematicidal activity, enhanced stability, or different specificity or range of target nematode pests are recovered.
- the method encompasses forming a mutagenized double-stranded polynucleotide from a template double-stranded polynucleotide comprising a nucleotide sequence of this invention, wherein the template double-stranded polynucleotide has been cleaved into double-stranded-random fragments of a desired size, and comprises the steps of adding to the resultant population of double-stranded random fragments one or more single or double-stranded oligonucleotides, wherein said oligonucleotides comprise an area of identity and an area of heterology to the double-stranded template polynucleotide; denaturing the resultant mixture of double-stranded random fragments and oligonucleotides into single-stranded fragments; incubating the resultant population of single-stranded fragments with a polymerase under conditions which result in the annealing of said single-stranded fragments at said areas of identity to form pairs of
- the concentration of a single species of double-stranded random fragment in the population of double-stranded random fragments is less than 1% by weight of the total DNA.
- the template double-stranded polynucleotide comprises at least about 100 species of polynucleotides.
- the size of the double-stranded random fragments is from about 5 by to 5 kb.
- the fourth step of the method comprises repeating the second and the third steps for at least 10 cycles.
- the Cry21 nucleotide sequences of the invention can be modified through N- or preferably C-terminal deletions to encode functional fragments.
- the term functional fragments relate to consecutive amino acids for Cry21 proteins that are at least 10, 20, 30, 40, 50, or 60 consecutive amino acids of SEQ ID NOs: 1, 7, 9, 13, 21, 23, 33, 35, 39, 41, 51, or 53.
- Preferred deletions would remove the C-terminal crystallization domain downstream of a conserved amino acid motif called “DRIE ” or “DRIE” which is also highlighted in FIG. 2 and which has previously been described as “Block 5” in Schnepf et al 1998.
- Functional fragments with C-terminal deletions of the oystallization domain are expected to result in toxic core proteins, which are independent from proteolytic activation.
- At least one of the Cry21 nucleotide sequences of the invention is inserted into an appropriate expression cassette, comprising a promoter and termination signals. Expression of the nucleotide sequence is constitutive, or an inducible promoter responding to various types of stimuli to initiate transcription is used.
- the cell in which the toxin is expressed is a microorganism, such as a virus, a bacteria, or a fungus.
- a virus such as a baculovirus
- the insecticidal toxin thus produced is used as an insecticidal agent.
- baculoviruses engineered to include the nucleotide sequence are used to infect insects in vivo and kill them either by expression of the insecticidal toxin or by a combination of viral infection and expression of the insecticidal toxin.
- Bacterial cells are also hosts for the expression of the nucleotide sequences of the invention.
- non-pathogenic symbiotic bacteria which are able to live and replicate within plant tissues, so-called endophytes, or non-pathogenic symbiotic bacteria, which are capable of colonizing the phyllosphere or the rhizosphere, so-called epiphytes, are used.
- Such bacteria include bacteria of the genera Agrobacterium, Alcaligenes, Azospirillum, Azotobacter, Bacillus, Clavibacter, Enterobacter, Erwinia, Flavobacter, Klebsiella, Pseudomonas, Rhizobium, Serratia, Streptomyces and Xanthomonas.
- Symbiotic fungi such as Trichoderma and Gliocladium are also possible hosts for expression of the inventive nucleotide sequences for the same purpose.
- the expression vectors pKK223-3 and pKK223-2 can be used to express heterologous genes in E. coli, either in transcriptional or translational fusion, behind the tac or trc promoter.
- the simplest procedure is to insert the operon into a vector such as pKK223-3 in transcriptional fusion, allowing the cognate ribosome binding site of the heterologous genes to be used.
- Alternate systems for overexpression rely for example, on yeast vectors and include the use of Pichia, Saccharomyces and Kluyveromyces (Sreekrishna, In: Industrial Microorganisms: Basic and Applied Molecular Genetics, Baltz, Hegeman, and Skatrud eds., American Society for Microbiology, Washington (1993); Dequin & Bane, Biotechnology L2: 173-177 (1994); van den Berg et al., Biotechnology 8: 135-139 (1990)).
- At least one Cry21 protein of the invention is expressed in a higher organism, e.g., a plant.
- transgenic plants expressing effective amounts of the toxins protect themselves from nematode pests.
- the nematode starts feeding on such a transgenic plant, it also ingests the expressed Ciy21 toxin. This may deter the nematode from further feeding in the plant tissue, may harm or kill the nematode or may reduce the nematodes ability to reproduce.
- a nucleotide sequence of the present invention is inserted into an expression cassette, which is then stably integrated in the genome of the plant.
- Plants transformed in accordance with the invention may be monocots or dicots and include, but are not limited to, maize, wheat, barley, rye, sweet potato, bean, pea, chicory, lettuce, cabbage, cauliflower, broccoli, turnip, radish, spinach, asparagus, onion, garlic, pepper, celery, squash, pumpkin, hemp, zucchini, apple, pear, quince, melon, plum, cherry, peach, nectarine, apricot, strawberry, grape, raspberry, blackberry, pineapple, avocado, papaya, mango, banana, soybean, tomato, sorghum, sugarcane, sugar beet, sunflower, rapeseed, clover, tobacco, carrot, cotton, alfalfa, rice, potato, eggplant, cucumber, Arabidopsis, and woody plants such as coniferous and deciduous trees.
- nucleotide sequence Once a desired nucleotide sequence has been transformed into a particular plant species, it may be propagated in that species or moved into other varieties of the same species, particularly including commercial varieties, using traditional breeding techniques.
- a nucleotide sequence of the invention is expressed in transgenic plants, thus causing the biosynthesis of the corresponding toxin in the transgenic plants. In this way, transgenic plants with enhanced resistance to nematodes are generated.
- the nucleotide sequences of the invention may require modification and optimization. Although in many cases genes from microbial organisms can be expressed in plants at high levels without modification, low expression in transgenic plants may result from microbial nucleotide sequences having codons that are not preferred in plants. It is known in the art that all organisms have specific preferences for codon usage, and the codons of the nucleotide sequences described in this invention can be changed to conform with plant preferences, while maintaining the amino acids encoded thereby.
- coding sequences that have at least about 35% GC content, preferably more than about 45%, more preferably more than about 50%, and most preferably more than about 60%.
- preferred gene sequences may be adequately expressed in both monocotyledonous and dicotyledonous plant species, sequences can be modified to account for the specific codon preferences and GC content preferences of monocotyledons or dicotyledons as these preferences have been shown to differ (Murray et al. Nucl. Acids Res. 17:477-498 (1989)).
- the nucleotide sequences are screened for the existence of illegitimate splice sites that may cause message truncation. All changes required to be made within the nucleotide sequences such as those described above are made using well known techniques of site directed mutagenesis, PCR, and synthetic gene construction using the methods known in the art.
- the nucleotide sequences of the invention can be optimized for expression in any plant It is recognized that all or any part of the gene sequence may be optimized or synthetic. That is, synthetic or partially optimized sequences may also be used. [00163] For efficient initiation of translation, sequences adjacent to the initiating methionine encoding start codon may require modification. For example, they can be modified by the inclusion of sequences known to be effective in plants. Joshi has suggested an appropriate consensus for plants (NAR 15:6643-6653 (1987)) and Clonetech suggests a further consensus translation initiator (1993/1994 catalog, page 210). These consensuses are suitable for use with the nucleotide sequences of this invention.
- sequences are incorporated into constructions comprising the nucleotide sequences, up to and including the ATG (while leaving the second amino acid unmodified), or alternatively up to and including the GTC subsequent to the ATG (with the possibility of modifying the second amino acid of the transgene).
- the Cry21 toxin genes of the embodiments can be operably fused to a variety of promoters for expression in plants including constitutive, inducible, temporally regulated, developmentally regulated, chemically regulated, tissue-preferred and tissue-specific promoters to prepare recombinant DNA molecules, i.e., chimeric genes.
- the choice of promoter will vary depending on the temporal and spatial requirements for expression.
- expression of the nucleotide sequences encoding Cry21 proteins of the embodiments in leaves, in stalks or stems, in ears, in inflorescences e.g.
- a pesticidal sequence of the invention may be provided in an expression cassette for expression in a host cell of interest, e.g. a plant cell or a microbe.
- plant expression cassette is intended a DNA construct that is capable of resulting in the expression of a protein from an open reading frame in a plant cell. Typically, these contain a promoter and a coding sequence. Often, such constructs will also contain a 3' untranslated region. Such constructs may contain a signal sequence or leader sequence to facilitate co-translational or post-translational transport of the peptide to certain intracellular structures such as the chloroplast (or other plastid), endoplasmic reticulum, or Golgi apparatus.
- signal sequence is intended a sequence that is known or suspected to result in co- translational or post-translational peptide transport across the cell membrane. In eukaryotes, this typically involves secretion into the Golgi apparatus, with some resulting glycosylation.
- Insecticidal toxins of bacteria are often synthesized as protoxins, which are protolytically activated in the gut of the target pest (Chang (1987) Methods Enzymol. 153 : 507-516).
- the signal sequence is located in the native sequence, or may be derived from a sequence of the invention.
- leader sequence is intended any sequence that when translated, results in an amino acid sequence sufficient to trigger co-translational transport of the peptide chain to a subcellular organelle.
- this includes leader sequences targeting transport and/or glycosylation by passage into the endoplasmic reticulum, passage to vacuoles, plastids including chloroplasts, mitochondria, and the like.
- a polypeptide comprising an amino acid sequence of the present invention that is operably linked to a heterologous leader or signal sequence.
- plant transformation vector is intended a DNA molecule that is necessary for efficient transformation of a plant cell. Such a molecule may consist of one or more plant expression cassettes, and may be organized into more than one “vector” DNA molecule.
- binary vectors are plant transformation vectors that utilize two non-contiguous DNA vectors to encode all requisite cis- and trans-acting functions for transformation of plant cells (Kellens and Mullineaux (2000) Trends in Plant Science 5:446-451).
- Vector refers to a nucleic acid construct designed for transfer between different host cells.
- “Expression vector” refers to a vector that has the ability to incorporate, integrate and express heterologous DNA sequences or fragments in a foreign cell.
- the cassette will include 5' and/or 3' regulatory sequences operably linked to a sequence of the invention.
- operably linked is intended a functional linkage between a promoter and a second sequence, wherein the promoter sequence initiates and mediates transcription of the DNA sequence corresponding to the second sequence.
- operably linked means that the nucleic acid sequences being linked are contiguous and, where necessary to join two protein coding regions, contiguous and in the same reading frame.
- the nucleotide sequence is operably linked to a heterologous promoter capable of directing expression of said nucleotide sequence in a host cell, such as a microbial host cell or a plant host cell.
- the cassette may additionally contain at least one additional gene to be cotransformed into the organism. Alternatively, the additional gene(s) can be provided on multiple expression cassettes.
- the nucleotide sequence of the invention is operably linked to a heterologous promoter, e.g., a plant promoter.
- Such an expression cassette is provided with a plurality of restriction sites for insertion of the pesticidal sequence to be under the transcriptional regulation of the regulatory regions.
- the expression cassette will include in the 5'-3' direction of transcription, a transcriptional and translational initiation region (i.e., a promoter), a DNA sequence of the invention, and a translational and transcriptional termination region (i.e., termination region) functional in plants.
- the promoter may be native or analogous, or foreign or heterologous, to the plant host and/or to the DNA sequence of the invention. Additionally, the promoter may be the natural sequence or alternatively a synthetic sequence.
- the promoter is “native” or “homologous” to the plant host, it is intended that the promoter is found in the native plant into which the promoter is introduced. Where the promoter is foreign or heterologous to the DNA sequence of the invention, it is intended that the promoter is not the native or naturally occurring promoter for the operably linked DNA sequence of the invention.
- the promoter may be inducible or constitutive. It may be naturally occurring, may be composed of portions of various naturally occurring promoters, or may be partially or totally synthetic. Guidance for the design of promoters is provided by studies of promoter structure, such as that of Harley and Reynolds (1987) Nucleic Acids Res. 15:2343-2361. Also, the location of the promoter relative to the transcription start may be optimized. See, e.g., Roberts et al. (1979) Proc. Natl. Acad. Sci. USA, 76:760-764. Many suitable promoters for use in plants are well known in the art.
- suitable constitutive promoters for use in plants include: the promoters from plant viruses, such as the peanut chlorotic streak caulimovirus (PC1SV) promoter (U.S. Pat. No. 5,850,019); the 35S promoter from cauliflower mosaic virus (CaMV) (Odell et al. (1985) Nature 313:810-812); the 35S promoter described in Kay et al. (1987) Science 236: 1299-1302; promoters of Chlorella virus methyltransferase genes (U.S. Pat. No. 5,563,328) and the full- length transcript promoter from figwort mosaic virus (FMV) (U.S. Pat. No.
- PC1SV peanut chlorotic streak caulimovirus
- CaMV cauliflower mosaic virus
- FMV full- length transcript promoter from figwort mosaic virus
- Patent 5,510,474) maize H3 histone (Lepetit et al. (1992) Mol. Gen. Genet. 231:276-285 and Atanassova et al. (1992) Plant J. 2(3):291-300); Brassica napus ALS3 (PCT application WO97/41228); a plant ribulose-biscarboxylase/oxygenase (RuBisCO) small subunit gene; the circovirus (AU 689311) or the Cassava vein mosaic virus (CsVMV, US 7,053,205); promoters from soybean (Pbdc6 or Pbdc7, described in WO/2014/150449 or ubiquitin 3 promoter described in US Patent No. 7393948 and US Patent No. 8395021); and promoters of various Agrobacterium genes (see U.S. Pat. Nos. 4,771,002; 5,102,796; 5,182,200; and 5,428,147).
- Suitable inducible promoters for use in plants include: the promoter from the ACE1 system which responds to copper (Mett et al. (1993) PNAS 90:4567-4571); the promoter of the maize In2 gene which responds to benzenesulfonamide herbicide safeners (Hershey et al. (1991) Mol. Gen. Genetics 227:229-237 and Gatz et al. (1994) Mol. Gen. Genetics 243:32-38); and the promoter of the Tet repressor from TnlO (Gatz et al. (1991) Mol. Gen. Genet. 227:229-237).
- Another inducible promoter for use in plants is one that responds to an inducing agent to which plants do not normally respond.
- An exemplary inducible promoter of this type is the inducible promoter from a steroid hormone gene, the transcriptional activity of which is induced by a glucocorticosteroid hormone (Schena et al. (1991) Proc. Natl. Acad. Sci. USA 88:10421) or the recent application of a chimeric transcription activator, XVE, for use in an estrogen receptor- based inducible plant expression system activated by estradiol (Zuo et al. (2000) Plant J., 24:265-273).
- inducible promoters for use in plants are described in EP 332104, PCT WO 93/21334 and PCT WO 97/06269 which are herein incorporated by reference in their entirety. Promoters composed of portions of other promoters and partially or totally synthetic promoters can also be used. See, e.g., Ni et al. (1995) Plant J. 7:661-676 and PCT WO 95/14098 describing such promoters for use in plants.
- a promoter sequence specific for particular regions or tissues of plants can be used to express the nematicidal proteins of the invention, such as promoters specific for seeds (Datla, R et al., 1997, Biotechnology Ann. Rev. 3, 269-296), especially the napin promoter (EP 255 378 Al), the phaseolin promoter, the glutenin promoter, the helianthinin promoter (WO92/17580), the albumin promoter (WO98/45460), the oleosin promoter (WO98/45461), the SAT1 promoter or the SAT3 promoter (PCT/US98/06978).
- promoters specific for seeds such as promoters specific for seeds (Datla, R et al., 1997, Biotechnology Ann. Rev. 3, 269-296), especially the napin promoter (EP 255 378 Al), the phaseolin promoter, the glutenin promoter, the helianthinin promoter (WO92/17580), the albumin promoter (WO98/45460), the
- an inducible promoter advantageously chosen from the phenylalanine ammonia lyase (PAL), HMG-CoA reductase (HMG), chitinase, glucanase, proteinase inhibitor (PI), PR1 family gene, nopaline synthase (nos) and vspB promoters (US 5 670349, Table 3), the HMG2 promoter (US 5 670349), the apple beta-galactosidase (ABG1) promoter and the apple aminocyclopropane carboxylate synthase (ACC synthase) promoter (WO98/45445).
- Multiple promoters can be used in the constructs of the invention, including in succession.
- the promoter may include, or be modified to include, one or more enhancer elements.
- the promoter may include a plurality of enhancer elements. Promoters containing enhancer elements provide for higher levels of transcription as compared to promoters that do not include them. Suitable enhancer elements for use in plants include the PC1SV enhancer element (U.S. Pat. No. 5,850,019), the CaMV 35S enhancer element (U.S. Pat Nos. 5,106,739 and 5,164,316) and the FMV enhancer element (Maiti et al. (1997) Transgenic Res.
- TMV tobacco mosaic virus
- TSV tobacco etch virus
- Kirol. 64: 1590-1597 for example, or introns such as the adhl intron of maize or intron 1 of rice actin. See also PCT WO96/23898, WO2012/021794, WO2012/021797, WO2011/084370, and WO2011/028914.
- constructs can contain 5' and 3' untranslated regions.
- Such constructs may contain a signal sequence or leader sequence to facilitate co-translational or post-translational transport of the peptide of interest to certain intracellular structures such as the chloroplast (or other plastid), endoplasmic reticulum, or Golgi apparatus, or to be secreted.
- the construct can be engineered to contain a signal peptide to facilitate transfer of the peptide to the endoplasmic reticulum.
- 3' untranslated region is intended a polynucleotide located downstream of a coding sequence.
- Polyadenylation signal sequences and other sequences encoding regulatory signals capable of affecting the addition of polyadenylic acid tracts to the 3' end of the mRNA precursor are 3' untranslated regions.
- 5' untranslated region is intended a polynucleotide located upstream of a coding sequence.
- Enhancers are polynucleotides that act to increase the expression of a promoter region. Enhancers are well known in the art and include, but are not limited to, the SV40 enhancer region and the 35S enhancer element.
- the termination region may be native with the transcriptional initiation region, may be native with the operably linked DNA sequence of interest, may be native with the plant host, or may be derived from another source (i.e., foreign or heterologous to the promoter, the DNA sequence of interest, the plant host, or any combination thereof).
- Convenient termination regions are available from the Ti-plasmid of A. tumefaciens, such as the octopine synthase and nopaline synthase termination regions. See also Guerineau et al. (1991) Mol. Gen. Genet. 262: 141-144; Proudfoot (1991) Cell 64:671-674; Sanfacon et al. (1991) Genes Dev.
- the gene(s) may be optimized for increased expression in the transformed host cell (synthetic DNA sequence). That is, the genes can be synthesized using host cell-preferred codons for improved expression or may be synthesized using codons at a host-preferred codon usage frequency.
- Synthetic DNA sequences can be useful to simply remove unwanted restriction endonuclease sites, to facilitate DNA cloning strategies, to alter or remove any potential codon bias, to alter or improve GC content, to remove or alter alternate reading frames, and/or to alter or remove intron/exon splice recognition sites, polyadenylation sites, Shine-Delgamo sequences, unwanted promoter elements and the like that may be present in a native DNA sequence.
- the GC content of the gene will be increased. See, for example, Campbell and Gowri (1990) Plant Physiol. 92: 1-11 for a discussion of host-preferred codon usage.
- DNA sequences may be utilized to introduce other improvements to a DNA sequence, such as introduction of an intron sequence, creation of a DNA sequence that in expressed as a protein fusion to organelle targeting sequences, such as chloroplast transit peptides, apoplast/vacuolar targeting peptides, or peptide sequences that result in retention of the resulting peptide in the endoplasmic reticulum.
- organelle targeting sequences such as chloroplast transit peptides, apoplast/vacuolar targeting peptides, or peptide sequences that result in retention of the resulting peptide in the endoplasmic reticulum.
- the nematicidal protein is targeted to the chloroplast for expression.
- the expression cassette will additionally contain a nucleic acid encoding a transit peptide to direct the nematicidal protein to the chloroplasts.
- transit peptides are known in the art. See, for example, Von Heijne et al. (1991) Plant Mol. Biol. Rep. 9: 104-126; Clark et al. (1989) J. Biol. Chem. 264:17544-17550; Della-Cioppa et al. (1987) Plant Physiol. 84:965-968; Romer et al. (1993) Biochem. Biophys. Res. Commun. 196:1414-1421; and Shah et al. (1986) Science 233:478-481.
- Methods of the embodiments involve introducing a nucleotide construct into a plant
- introducing is intended to present to the plant the nucleotide construct in such a manner that the construct gains access to the interior of a cell of the plant.
- the methods of the invention do not require that a particular method for introducing a nucleotide construct to a plant is used, only that the nucleotide construct gains access to the interior of at least one cell of the plant.
- Methods for introducing nucleotide constructs into plants are known in the art including, but not limited to, stable transformation methods, transient transformation methods, gene editing and virus- mediated methods.
- Transgenic plants or “transformed plants” or “stably transformed” plants or cells or tissues refers to plants that have incorporated or integrated exogenous nucleic acid sequences or DNA fragments into the plant cell. These nucleic acid sequences include those that are exogenous, or not present in the untransformed plant cell, as well as those that may be endogenous, or present in the untransformed plant cell.
- “Heterologous” generally refers to the nucleic acid sequences that are not endogenous to the cell or part of the native genome in which they are present, and have been added to the cell by infection, transfection, microinjection, electroporation, microprojection, or the like.
- the transgenic plants of the invention express one or more of the novel toxin sequences disclosed herein.
- the protein or nucleotide sequence of the invention is advantageously combined in plants with other genes which encode proteins or RNAs that confer useful agronomic properties to such plants.
- genes which encode proteins or RNAs that confer useful agronomic properties on the transformed plants mention can be made of the DNA sequences encoding proteins which confer tolerance to one or more herbicides, and others which confer tolerance to certain insects, those which confer tolerance to certain diseases, DNAs that encodes RNAs that provide nematode or insect control, and the like.
- the transgenic plant further comprises one or more additional genes for insect resistance (e.g., Cryl, such as members of the CrylA, CrylB, CrylC, CrylD, CrylE, and Cry IF families; Cry2, such as members of the Cry2A family; Cry9, such as members of the Cry9A, Cry9B, Cry9C, Cry9D, Cry9E, and Cry9F families; etc.).
- Cryl such as members of the CrylA, CrylB, CrylC, CrylD, CrylE, and Cry IF families
- Cry2A family such as members of the Cry2A family
- Cry9 such as members of the Cry9A, Cry9B, Cry9C, Cry9D, Cry9E, and Cry9F families
- the transgenic plant may comprise any gene imparting an agronomic trait of interest.
- sequence encoding these enzymes is advantageously preceded by a sequence encoding a transit peptide, in particular the “optimized transit peptide” described in US Patent 5,510,471 or 5,633,448.
- Exemplary herbicide tolerance traits that can be combined with the nucleic acid sequence of the invention further include at least one ALS (acetolactate synthase) inhibitor (W02007/024782); a mutated Arabidopsis ALS/AHAS gene (U.S. Patent 6,855,533); genes encoding 2,4-D-monooxygenases conferring tolerance to 2,4-D (2,4-dichlorophenoxyacetic acid) by metabolization (U.S. Patent 6,153,401); and, genes encoding Dicamba monooxygenases conferring tolerance to dicamba (3,6-dichloro-2-methoxybenzoic acid) by metabolization (US 2008/0119361 and US 2008/0120739).
- ALS acetolactate synthase
- a mutated Arabidopsis ALS/AHAS gene U.S. Patent 6,855,533
- genes encoding 2,4-D-monooxygenases conferring tolerance to 2,4-D (2,
- the nucleic acid of the invention is stacked with one or more herbicide tolerant genes, including one or more HPPD inhibitor herbicide tolerant genes, and/or one or more genes tolerant to glyphosate and/or glufosinate.
- herbicide tolerant genes including one or more HPPD inhibitor herbicide tolerant genes, and/or one or more genes tolerant to glyphosate and/or glufosinate.
- Bt proteins proteins widely described in the literature and well known to those skilled in the art. Mention will also be made of proteins extracted from bacteria such as Photorhabdus (WO97/17432 & WO98/08932).
- the VIP3Aal9 protein of NCBI accession ABG20428 the VIP3Aa20 protein of NCBI accession ABG20429 (SEQ ID No. 2 in WO 2007/142840), the VIP3A proteins produced in the COT202 or COT203 cotton events (W02005/054479 and W02005/054480, respectively), the Cry proteins as described in WO2001/47952, the VIP3Aa protein or a toxic fragment thereof as described in Estruch et al. (1996), Proc Natl Acad Sci U SA.
- any variants or mutants of any one of these proteins differing in some (1-10, preferably 1-5) amino acids from any of the above sequences, particularly the sequence of their toxic fragment, or which are fused to a transit peptide, such as a plastid transit peptide, or another protein or peptide, is included herein.
- the nucleic acid of the invention can be combined in plants with one or more genes conferring a desirable trait, such as herbicide tolerance, insect tolerance, drought tolerance, nematode control, water use efficiency, nitrogen use efficiency, improved nutritional value, disease resistance, improved photosynthesis, improved fiber quality, stress tolerance, improved reproduction, and the like.
- a desirable trait such as herbicide tolerance, insect tolerance, drought tolerance, nematode control, water use efficiency, nitrogen use efficiency, improved nutritional value, disease resistance, improved photosynthesis, improved fiber quality, stress tolerance, improved reproduction, and the like.
- transgenic events which may be combined with the genes of the current invention in plants of the same species (e.g., by crossing or by re-transforming a plant containing another transgenic event with a chimeric gene of the invention), include Event BPS- C VI 27-9 (soybean, herbicide tolerance, deposited as NCIMB No. 41603, described in
- Event DAS21606-3 / 1606 (soybean, herbicide tolerance, deposited as PTA- 11028, described in WO2012/033794), Event DAS-44406-6 / pDAB8264.44.06.1 (soybean, herbicide tolerance, deposited as PTA-11336, described in WO2012/075426), Event DAS- 14536-7 /pDAB8291.45.36.2 (soybean, herbicide tolerance, deposited as PTA-11335, described in WO2012/075429), Event DAS68416 (soybean, herbicide tolerance, deposited as ATCC PTA- 10442, described in WO2011/066384 or WO2011/066360); Event DP-305423-1 (soybean, quality trait, not deposited, described in US-A 2008-312082 or W02008/054747); Event DP- 356043-5 (soybean, herbicide tolerance, deposited as ATCC PTA
- Event MON89788 (soybean, herbicide tolerance, deposited as ATCC PTA- 6708, described in US-A 2006-282915 or W02006/130436); Event SYHT0H2 / SYN-000H2-5 (soybean, herbicide tolerance, deposited as PTA-11226, described in WO2012/082548), event EE-GM3 / FG72 (soybean, herbicide tolerance, ATCC Accession No.
- PTA-11041) optionally stacked with event EE-GM1/LL27 or event EE-GM2/LL55 (WO2011/063413A2); Event DAS-68416-4 (soybean, herbicide tolerance, ATCC Accession N° PTA- 10442, WO2011/066360A1); Event DAS-68416-4 (soybean, herbicide tolerance, ATCC Accession No. PTA-10442, WO2011/066384A1); Event DAS-21606-3 (soybean, herbicide tolerance, ATCC Accession No. PTA-1 1028, WO2012/033794A2); Event MON-87712-4 (soybean, quality trait, ATCC Accession No.
- a method for producing a soybean plant or seed comprising a nucleotide sequence encoding SEQ ID NO 2, 8, 10, 14, 16, 22, 24, 34, 36, 40, 42, 52, 54, or any functional fragments thereof combined with another SCN resistance locus/gene, such as by combining a soybean plant or seed comprising a nucleotide sequence encoding SEQ ID NOs: 2, 8, 10, 14, 16, 22, 24, 34, 36, 40, 42, 52, 54, or any functional fragments thereof with another SCN resistance locus/gene occurring in the same soybean plant/seed, and planting seed comprising a nucleotide sequence encoding SEQ ID NOs: 1, 7, 9, 13, 15, 21, 23, 33, 35, 39, 41, 51, 53 and any functional fragments thereof and said other SCN resistance locus/gene.
- the plants, cells or seeds of the invention contain one or more other SCN resistance loci/genes that occur in soybean, to get a combination of different SCN resistance sources in the soybean plants, cells or seeds of the invention.
- soybean SCN resistance loci or genes are known and one or more of those can be combined with a plant comprising SEQ ID NO: 2, 8, 10, 14, 16, 22, 24, 34, 36, 40, 42, 52, 54, or any functional fragments thereof in the same plant, cell or seed, such as any one of the SCN resistance genes/loci from the resistance sources PI 88788, PI 548402 (Peking), PI 437654 (Hartwig or CystX®), or any combination thereof, or one or more of the native SCN resistance loci/genes rhgl, rhgl-b, rhg2, rhg3, Rhg4, Rhg5, qSCNl 1, cqSCN-003, cqSCN-005, cqSCN-006, cqS
- the plants or seeds of the invention are combined with one or more SCN resistance loci in soybean obtained from any one of SCN resistance sources PI 548316, PI 567305, PI 437654, PI 90763, PI 404198B, PI 88788, PI 468916 , PI 567516C, PI 209332, PI 438489B, PI 89772, Peking, Pl 548402, Pl 404198A, PI 561389B, Pl 629013, PI 507471, PI 633736, PI 507354, PI 404166, PI 437655, PI 467312, PI 567328, PI 22897, or PI 494182.
- Transformation of plant cells can be accomplished by one of several techniques known in the art.
- the pesticidal gene of the invention may be modified to obtain or enhance expression in plant cells.
- a construct that expresses such a protein would contain a promoter to drive transcription of the gene, as well as a 3' untranslated region to allow transcription termination and polyadenylation.
- the organization of such constructs is well known in the art
- the gene can be engineered to contain a signal peptide to facilitate transfer of the peptide to the endoplasmic reticulum.
- a plant expression cassette will be inserted into a plant transformation vector.
- This plant transformation vector may be comprised of one or more DNA vectors needed for achieving plant transformation.
- DNA vectors needed for achieving plant transformation.
- Binary vectors as well as vectors with helper plasmids are most often used for Agrobacterium-mediated transformation, where the size and complexity of DNA segments needed to achieve efficient transformation is quite large, and it is advantageous to separate functions onto separate DNA molecules.
- Binary vectors typically contain a plasmid vector that contains the cis-acting sequences required for T-DNA transfer (such as left border and right border), a selectable marker that is engineered to be capable of expression in a plant cell, and a “gene of interest” (a gene engineered to be capable of expression in a plant cell for which generation of transgenic plants is desired). Also present on this plasmid vector are sequences required for bacterial replication. The cis-acting sequences are arranged in a fashion to allow efficient transfer into plant cells and expression therein. For example, the selectable marker gene and the pesticidal gene are located between the left and right borders.
- a second plasmid vector contains the trans-acting factors that mediate T-DNA transfer from Agrobacterium to plant cells.
- This plasmid often contains the virulence functions (Vir genes) that allow infection of plant cells by Agrobacterium, and transfer of DNA by cleavage at border sequences and vir-mediated DNA transfer, as is understood in the art (Hellens and Mullineaux (2000) Trends in Plant Science 5:446-451).
- Several types of Agrobacterium strains e.g. LBA4404, GV3101, EHA101, EHA105, etc.
- the second plasmid vector is not necessary for transforming the plants by other methods such as microprojection, microinjection, electroporation, polyethylene glycol, etc.
- plant transformation methods involve transferring heterologous DNA into target plant cells (e.g. immature or mature embryos, suspension cultures, undifferentiated callus, protoplasts, etc.), followed by applying a maximum threshold level of appropriate selection (depending on the selectable marker gene) to recover the transformed plant cells from a group of untransformed cell mass.
- target plant cells e.g. immature or mature embryos, suspension cultures, undifferentiated callus, protoplasts, etc.
- a maximum threshold level of appropriate selection depending on the selectable marker gene
- Explants are typically transferred to a fresh supply of the same medium and cultured routinely.
- the transformed cells are differentiated into shoots after placing on regeneration medium supplemented with a maximum threshold level of selecting agent
- the shoots are then transferred to a selective rooting medium for recovering rooted shoot or plantlet.
- the transgenic plantlet then grows into a mature plant and produces fertile seeds (e.g.
- Transformation protocols as well as protocols for introducing nucleotide sequences into plants may vary depending on the type of plant or plant cell, i.e., monocot or dicot, targeted for transformation.
- Generation of transgenic plants may be performed by one of several methods, including, but not limited to, microinjection, electroporation, direct gene transfer, introduction of heterologous DNA by Agrobacterium into plant cells ⁇ Agrobacterium-mediated transformation), bombardment of plant cells with heterologous foreign DNA adhered to particles, ballistic particle acceleration, aerosol beam transformation (U.S. Published Application No. 20010026941; U.S. Patent No. 4,945,050; International Publication No. WO 91/00915; U.S. Published Application No. 2002015066), Lecl transformation, and various other non-particle direct-mediated methods to transfer DNA.
- heterologous foreign DNA Following integration of heterologous foreign DNA into plant cells, one then applies a maximum threshold level of appropriate selection in the medium to kill the untransformed cells and separate and proliferate the putatively transformed cells that survive from this selection treatment by transferring regularly to a fresh medium. By continuous passage and challenge with appropriate selection, one identifies and proliferates the cells that are transformed with the plasmid vector. Molecular and biochemical methods can then be used to confirm the presence of the integrated heterologous gene of interest into the genome of the transgenic plant.
- the cells that have been transformed may be grown into plants in accordance with conventional ways. See, for example, McCormick et al. (1986) Plant Cell Reports 5:81-84. These plants may then be grown, and either pollinated with the same transformed strain or different strains, and the resulting hybrid having constitutive expression of the desired phenotypic characteristic identified. Two or more generations may be grown to ensure that expression of the desired phenotypic characteristic is stably maintained and inherited and then seeds harvested to ensure expression of the desired phenotypic characteristic has been achieved. In this manner, the present invention provides transformed seed (also referred to as “transgenic seed”) having a nucleotide construct of the invention, for example, an expression cassette of the invention, stably incorporated into their genome.
- heterologous foreign DNA Following introduction of heterologous foreign DNA into plant cells, the transformation or integration of heterologous gene in the plant genome is confirmed by various methods such as analysis of nucleic acids, proteins and metabolites associated with the integrated gene.
- PCR analysis is a rapid method to screen transformed cells, tissue or shoots for the presence of incorporated gene at the earlier stage before transplanting into the soil (Sambrook and Russell (2001) Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY). PCR is carried out using oligonucleotide primers specific to the gene of interest or Agrobacterium vector background, etc.
- Plant transformation may be confirmed by Southern blot analysis of genomic DNA (Sambrook and Russell, 2001, supra). In general, total DNA is extracted from the transformant, digested with appropriate restriction enzymes, fractionated in an agarose gel and transferred to a nitrocellulose or nylon membrane. The membrane or “blof ’ is then probed with, for example, radiolabeled 32P target DNA fragment to confirm the integration of introduced gene into the plant genome according to standard techniques (Sambrook and Russell, 2001, supra).
- RNA is isolated from specific tissues of transformant, fractionated in a formaldehyde agarose gel, and blotted onto a nylon filter according to standard procedures that are routinely used in the art (Sambrook and Russell, 2001 , supra). Expression of RNA encoded by the pesticidal gene is then tested by hybridizing the filter to a radioactive probe derived from a pesticidal gene, by methods known in the art (Sambrook and Russell, 2001, supra).
- Methods described above by way of example may be utilized to generate transgenic plants, but the manner in which the transgenic plant cells are generated is not critical to this invention. Methods known or described in the art such as Agrpbacteroium-mediated transformation, biolistic transformation, and non-particle-mediated methods may be used at the discretion of the experimenter.
- Plants expressing a nematicidal protein may be isolated by common methods described in the art, for example by transformation of callus, selection of transformed callus, and regeneration of fertile plants from such transgenic callus. Ln such process, one may use any gene as a selectable marker so long as its expression in plant cells confers ability to identify or select for transformed cells.
- a number of markers have been developed for use with plant cells, such as resistance to chloramphenicol, the aminoglycoside G418, hygromycin, or the like.
- Other genes that encode a product involved in chloroplast metabolism may also be used as selectable markers.
- genes that provide resistance to plant herbicides such as glyphosate, bromoxynil, or imidazolinone may find particular use.
- Such genes have been reported (Stalker et al. (1985) J. Biol. Chem. 263:6310-6314 (bromoxynil resistance nitrilase gene); and Sathasivan et al. (1990) Nucl. Acids Res. 18:2188 (AHAS imidazolinone resistance gene).
- the genes disclosed herein are useful as markers to assess transformation of bacterial or plant cells.
- a transgene in a plant is detected by testing for pesticidal activity against a nematode pest.
- Fertile plants expressing a nematicidal protein may be tested for pesticidal activity against a nematode pest, and the plants showing optimal activity selected for further breeding Standard greenhouse methods have been developed to evaluate new soybean varieties for nematode resistance see, Niblack, T. L. et al. A standard greenhouse method for assessing soybean cyst nematode resistance in soybean: SCE08 (Standardized Cyst Evaluation 2008). Plant Health Prog. 10, 33 (2009) herein incorporated by reference.
- the present invention may be used for transformation of any plant species, including, but not limited to, monocots and dicots.
- plants of interest include, but are not limited to, maize, sorghum, wheat, sunflower, tomato, crucifers, peppers, potato, cotton, rice, soybean, sugarbeet, sugarcane, tobacco, barley, and oilseed rape, Brassica sp., alfalfa, rye, millet, safflower, peanuts, sweet potato, cassava, coffee, coconut, pineapple, citrus trees, cocoa, tea, banana, avocado, fig, guava, mango, olive, papaya, cashew, macadamia, almond, oats, vegetables, ornamentals, and conifers.
- Microorganisms can be genetically altered to contain a nucleotide sequence encoding SEQ ID NO: 2, 8, 10, 14, 16, 22, 24, 34, 36, 40, 42, 52, 54 or nematicidally-active variants or fragments thereof, and protein may be used for protecting agricultural crops and products from pests.
- whole, i.e., unlysed, cells of a toxin (pesticide)-producing organism are treated with reagents that prolong the activity of the toxin produced in the cell when the cell is applied to the environment of target pest(s).
- the pesticide is produced by introducing a pesticidal gene into a cellular host. Expression of the pesticidal gene results, directly or indirectly, in the intracellular production and maintenance of the pesticide. In one aspect of this invention, these cells are then treated under conditions that prolong the activity of the toxin produced in the cell when the cell is applied to the environment of the target pest(s). The resulting product retains the toxicity of the toxin.
- These naturally encapsulated pesticides may then be formulated in accordance with conventional techniques for application to the environment hosting a target pest, e.g., soil, water, and foliage of plants. See, for example EP A 0192319, and the references cited therein. Alternatively, one may formulate the cells expressing a gene of this invention such as to allow application of the resulting material as a pesticide.
- the active ingredients of the present invention are normally applied in the form of compositions and can be applied to the crop area or plant to be treated, simultaneously or in succession, with other compounds.
- These compounds can be fertilizers, weed killers, cryoprotectants, surfactants, detergents, pesticidal soaps, dormant oils, polymers, and/or time- release or biodegradable carrier formulations that permit long-term dosing of a target area following a single application of the formulation.
- Suitable carriers and adjuvants can be solid or liquid and correspond to the substances ordinarily employed in formulation technology, e.g. natural or regenerated mineral substances, solvents, dispersants, wetting agents, tackifiers, binders or fertilizers.
- the formulations may be prepared into edible “baits” or fashioned into pest “traps” to permit feeding or ingestion by a target pest of the pesticidal formulation.
- Methods of applying an active ingredient of the present invention or an agrochemical composition of the present invention that contains at least one of the nematicidal proteins disclosed herein as SEQ ID NO: SEQ ID NOs: 2, 8, 10, 14, 16, 22, 24, 34, 36, 40, 42, 52, 54, or nematicidal ly effective variants or fragments thereof, include leaf application, seed coating and soil application. The number of applications and the rate of application depend on the intensity of infestation by the corresponding pest.
- the composition may be formulated as a powder, dust, pellet, granule, spray, emulsion, colloid, solution, or such like, and may be prepared by such conventional means as desiccation, lyophilization, homogenation, extraction, filtration, centrifugation, sedimentation, or concentration of a culture of cells comprising the polypeptide.
- the polypeptide may be present in a concentration of from about 1% to about 99% by weight.
- Nematode pests may be killed or reduced in numbers in a given area by the methods of the invention, or may be prophylactically applied to an environmental area to prevent infestation by a susceptible pest.
- the pesticide compositions described may be made by formulating either the bacterial cell, the crystal and/or the spore suspension, or the isolated protein component with the desired agriculturally-acceptable carrier.
- the compositions may be formulated prior to administration in an appropriate means such as lyophilized, freeze-dried, desiccated, or in an aqueous carrier, medium or suitable diluent, such as saline or other buffer.
- the formulated compositions may be in the form of a dust or granular material, or a suspension in oil (vegetable or mineral), or water or oil/water emulsions, or as a wettable powder, or in combination with any other carrier material suitable for agricultural application.
- Suitable agricultural carriers can be solid or liquid and are well known in the art.
- agriculturally-acceptable carrier covers all adjuvants, inert components, dispersants, surfactants, tackifiers, binders, etc. that are ordinarily used in pesticide formulation technology; these are well known to those skilled in pesticide formulation.
- the formulations may be mixed with one or more solid or liquid adjuvants and prepared by various means, e.g., by homogeneously mixing, blending and/or grinding the pesticidal composition with suitable adjuvants using conventional formulation techniques. Suitable formulations and application methods are described in U.S. Patent No. 6,468,523, herein incorporated by reference.
- Methods for increasing plant yield comprise providing a plant or plant cell expressing a polynucleotide encoding the nematicidal polypeptide sequence disclosed herein and growing the plant or a seed thereof in a field infested with (or susceptible to infestation by) a nematode pest against which said polypeptide has nematicidal activity.
- the Cry21 polypeptide described herein has nematicidal activity against a Pratylenchus spp., and said field is infested with said Pratylenchus spp.
- the Pratylenchus spp In various embodiments, the Pratylenchus spp.
- the “yield” of the plant refers to the quality and/or quantity of biomass produced by the plant
- biomass is intended any measured plant product.
- An increase in biomass production is any improvement in the yield of the measured plant product.
- Increasing plant yield has several commercial applications. For example, increasing plant leaf biomass may increase the yield of leafy vegetables for human or animal consumption. Additionally, increasing leaf biomass can be used to increase production of plant-derived pharmaceutical or industrial products.
- An increase in yield can comprise any statistically significant increase including, but not limited to, at least a 1% increase, at least a 3% increase, at least a 5% increase, at least a 10% increase, at least a 20% increase, at least a 30%, at least a 50%, at least a 70%, at least a 100% or a greater increase in yield compared to a plant not expressing the pesticidal protein described herein.
- plant yield is increased as a result of improved nematode resistance of a plant expressing the nematicidal protein disclosed herein. Expression of the nematicidal protein results in a reduced ability of a pest to infest or feed.
- expression of the nematicidal protein results in improved root development (e.g., improved root or root hair growth), improved yield, faster emergence, improved plant stress management including increased stress tolerance and/or improved recovery from stress, increased mechanical strength, improved drought resistance, reduced fungal disease infection, and improved plant health compared to a plant not expressing the nematicidal protein of the invention.
- the plants can also be treated with one or more chemical compositions, including one or more herbicide, insecticides, or fungicides.
- exemplary chemical compositions include: Fruits/Vegetables Herbicides: Atrazine, Bromacil, Diuron, Glyphosate, Linuron, Metribuzin, Simazine, Trifluralin, Fluazifop, Glufosinate, Halosulfiiron Gowan, Paraquat, Propyzamide, Sethoxydim, Butafenacil, Halosulfuron, Indaziflam.
- Fruits/Vegetables Insecticides Aldicarb , Bacillus thuriengiensis, Carbaryl, Carbofuran, Chlorpyrifos, Cypermethrin, Deltamethrin, Abamectm, Cyfluthrin/beta-cyfluthrin, Esfenvalerate, Lambda-cyhalothrin, Acequinocyl, Bifenazate, Methoxyfenozide, Novaluron, Chromafenozide, Thiacloprid, Dinotefuran, Fluacrypyrim, Spirodiclofen, Gamma-cyhalothrin, Spiromesifen, Spinosad, Rynaxypyr, Cyazypyr, Triflumuron, Spirotetramat, Imidacloprid, Flubendiamide, Thiodicarb, Metaflumizone, Sulfoxaflor, Cyflumetofen
- Cereals Herbicides 2.4-D, Amidosulfuron, Bromoxynil, Carfentrazone-E, Chlorotoluron, Chlorsulfuron, Clodinafop-P, Clopyralid, Dicamba, Diclofop-M, Diflufenican, Fenoxaprop, Florasulam, Flucarbazone-NA, Flufenacet, Flupyrosulfuron-M, Fluroxypyr, Flurtamone, Glyphosate, lodosulfuron, Ioxynil, Isoproturon, MCPA, Mesosulfuron, Metsulfuron, Pendimethalin, Pinoxaden, Propoxycarbazone, Prosulfocarb, Pyroxsulam, Sulfosulfuron, Thifensulfuron, Tralkoxydim, Triasulfuron, Tribenuron, Trifluralin, Tritosulfuron.
- Cereals Fungicides Azoxystrobin, Bixafen, Boscalid, Carbendazim, Chlorothalonil, Cyflufenamid, Cyproconazole, Cyprodinil, Dimoxystrobin, Epoxiconazole, Fenpropidin, Fenpropimorph, Fluopyram, Fluoxastrobin, Fluquinconazole, Fluxapyroxad, Isopyrazam, Kresoxim-methyl, Metconazole, Metrafenone, Penthiopyrad, Picoxystrobin, Prochloraz, Propiconazole, Proquinazid, Prothioconazole, Pyraclostrobin, Quinoxyfen, Spiroxamine, Tebuconazole, Thiophanate-methyl , Trifloxystrobin.
- Cereals Insecticides Dimethoate, Lambda-cyhalthrin, Deltamethrin, alpha- Cypermethrin, B-cyfluthrin, Bifenthrin, Imidacloprid, Clothianidin, Thiamethoxam, Thiacloprid, Acetamiprid, Dinetofuran, Clorphyriphos, Pirimicarb, Methiocarb, Sulfoxaflor; Maize Herbicides: Atrazine, Alachlor, Bromoxynil, Acetochlor, Dicamba, Clopyralid, (S- )Dimethenamid, Glufosinate, Glyphosate, Isoxaflutole, (S-)Metolachlor, Mesotrione, Nicosulfuron, Primisulfuron, Rimsulfuron, Sulcotrione, Foramsulfuron, Topramezone, Tembotrione, Saflufen
- Maize Insecticides Carbofuran, Chlorpyrifos, Bifenthrin, Fipronil, Imidacloprid, Lambda-Cyhalothrin, Tefluthrin, Terbufos, Thiamethoxam, Clothianidin, Spiromesifen, Flubendiamide, Triflumuron, Rynaxypyr, Deltamethrin, Thiodicarb, B-Cyfluthrin, Cypermethrin, Bifenthrin, Lufenuron, Tebupirimphos, Ethiprole, Cyazypyr, Thiacloprid, Acetamiprid, Dinetofuran, Avermectin; Maize Fungicides: Azoxystrobin, Bixafen, Boscalid, Cyproconazole, Dimoxystrobin, Epoxiconazole, Fenitropan, Fluopyram, Fluoxastrobin, Fluxapyroxad,
- Rice Herbicides Butachlor, Propanil, Azimsulfuron, Bensulfuron, Cyhalofop, Daimuron, Fentrazamide, Imazosulfuron, Mefenacet, Oxaziclomefone, Pyrazosulfuron, Pyributicarb, Quinclorac, Thiobencarb, Indanofan, Flufenacet, Fentrazamide, Halosulfuron, Oxaziclomefone, Benzobicyclon, Pyriftalid, Penoxsulam, Bispyribac, Oxadiargyl, Ethoxysulfuron, Pretilachlor, Mesotrione, Tefuryltrione, Oxadiazone, Fenoxaprop, Pyrimisulfan.
- Rice Insecticides Diazinon, Fenobucarb, Benfuracarb, Buprofezin, Dinotefuran, Fipronil, Imidacloprid, Isoprocarb, Thiacloprid, Chromafenozide, Clothianidin, Ethiprole, Flubendiamide, Rynaxypyr, Deltamethrin, Acetamiprid, Thiamethoxam, Cyazypyr, Spinosad, Spinotoram, Emamectin-Benzoate, Cypermethrin, Chlorpyriphos, Etofenprox, Carbofuran, Benfuracarb, Sulfoxaflor.
- Rice Fungicides Azoxystrobin, Carbendazim, Carpropamid, Diclocymet, Difenoconazole, Edifenphos, Ferimzone, Gentamycin, Hexaconazole, Hymexazol, Iprobenfos (IBP), Isoprothiolane, Isotianil, Kasugamycin, Mancozeb, Metominostrobin, Orysastrobin, Pencycuron, Probenazole, Propiconazole, Propineb, Pyroquilon, Tebuconazole, Thiophanate- methyl, Tiadinil, Tricyclazole, Trifloxystrobin, Validamycin.
- Cotton Herbicides Diuron, Fluometuron, MSMA, Oxyfluorfen, Promettyn, Trifluralin, Carfentrazone, Clethodim, Fluazifop-butyl, Glyphosate, Norflurazon, Pendimethalin, Pyrithiobac-sodium, Trifl oxy sulfuron, Tepraloxydim, Glufosinate, Flumioxazin, Thidiazuron; Cotton Insecticides: Acephate, Aldicarb, Chlorpyrifos, Cypermethrin, Deltamethrin, Abamectin, Acetamiprid, Emamectin Benzoate, Imidacloprid, Indoxacarb, Lambda-Cyhalothrin, Spinosad, Thiodicarb, Gamma-Cyhalothrin, Spiromesifen, Pyridalyl, Flonicamid Flu
- Cotton Fungicides Azoxystrobin, Bixafen, Boscalid, Carbendazim, Chlorothalonil, Copper, Cyproconazole, Difenoconazole, Dimoxystrobin, Epoxiconazole, Fenamidone, Fluazinam, Fluopyram, Fluoxastrobin, Fluxapyroxad, Iprodione, Isopyrazam, Isotianil, Mancozeb, Maneb, Metomi nostrobin, Penthiopyrad, Picoxystrobin, Propineb, Prothioconazole, Pyraclostrobin, Quintozene, Tebuconazole, Tetraconazole, Thiophanate-methyl, Trifloxystrobin; Soybean Herbicides: Alachlor, Bentazone, Trifluralin, Chlorimuron-Ethyl, Cloransulam-Methyl, Fenoxaprop, Fomesafen, Fluazifenofen,
- Soybean Insecticides Lambda-cyhalothrin, Methomyl, Imidacloprid, Clothianidin, Thiamethoxam, Thiacloprid, Acetamiprid, Dinetofuran, Flubendiamide, Rynaxypyr, Cyazypyr, Spinosad, Spinotoram, Emamectin-Benzoate, Fipronil, Ethiprole, Deltamethrin, B-Cyfluthrin, gamma and lambda Cyhalothrin, 4-[[(6-Chlorpyridin-3-yl)methyl](2,2-difluorethyl)amino]furan- 2(5H)-on, Spirotetramat, Spinodiclofen, Triflumuron, Flonicamid, Thiodicarb, beta-Cyfluthrin.
- Soybean Fungicides Azoxystrobin, Bixafen, Boscalid, Carbendazim, Chlorothalonil, Copper, Cyproconazole, Difenoconazole, Dimoxystrobin, Epoxiconazole, Fluazinam, Fluopyram, Fluoxastrobin, Flutriafol, Fluxapyroxad, Isopyrazam, Iprodione, Isotianil, Mancozeb, Maneb, Metconazole, Metominostrobin, Myclobutanil, Penthiopyrad, Picoxystrobin, Propiconazole, Propineb, Prothioconazole, Pyraclostrobin, Tebuconazole, Tetraconazole, Thiophanate-methyl, Trifloxystrobin.
- Sugarbeet Herbicides Chloridazon, Desmedipham, Ethofumesate, Phenmedipham, Triallate, Clopyralid, Fluazifop, Lenacil, Metamitron, Quinmerac, Cycloxydim, Triflusulfuron, Tepraloxydim, Quizalofop; Sugarbeet Insecticides: Imidacloprid, Clothianidin, Thiamethoxam, Thiacloprid, Acetamiprid, Dinetofuran, Deltamethrin, B-Cyfluthrin, gamma/lambda Cyhalothrin, 4-[[(6-Chlorpyridin-3-yl)methyl](2,2-difluorethyl)amino]furan-2(5H)-on, Tefluthrin, Rynaxypyr, Cyaxypyr, Fipronil, Carbofuran.
- Canola Herbicides Clopyralid, Diclofop, Fluazifop, Glufosinate, Glyphosate, Metazachlor, Trifluralin Ethametsulfuron, Quinmerac, Quizalofop, Clethodim, Tepraloxydim; Canola Fungicides: Azoxystrobin, Bixafen, Boscalid, Carbendazim, Cyproconazole, Difenoconazole, Dimoxystrobin, Epoxiconazole, Fluazinam, Fluopyram, Fluoxastrobin, Flusilazole, Fluxapyroxad, Iprodione, Isopyrazam, Mepiquat-chloride, Metconazole, Metominostrobin, Paclobutrazole, Penthiopyrad., Picoxystrobin, Prochloraz, Prothioconazole, Pyraclostrobin, Tebuconazole
- Canola Insecticides Carbofuran, Thiacloprid, Deltamethrin, Imidacloprid, Clothianidin, Thiamethoxam, Acetamiprid, Dinetofiiran, B-Cyfluthrin, gamma and lambda Cyhalothrin, tau- Fluvaleriate, Ethiprole, Spinosad, Spinotoram, Flubendiamide, Rynaxypyr, Cyazypyr, 4-[[(6- Chlorpyridin-3-yl)methyl](2,2-difluorethyl)amino]furan-2(5H)-on.
- Soybean events expressing a novel Cry21homolog (SEQ ID NO:2 (Axmi296) and SEQ ID NO: 54 (Cry21Aa2), respectively) were developed through Agrobacterium-mediated transformation of Thome soybean plants using a construct containing a gene encoding a 4- hydroxyphenylpyruvate dioxygenase protein (HPPD) inhibitor tolerant herbicide gene (described in WO2014043435) and Cry21 coding sequences. Wild-type Thome soybean served as the non- nematode resistant control. Cry21, when expressed in soybean plants, reduces the number of Pratylenchus brachyurus that reproduce in the roots compared with wild-type plants.
- SEQ ID NO:2 Axmi296
- SEQ ID NO: 54 SEQ ID NO: 54
- This particular Cry21 (SEQ ID NO:2) gene shares approximately 65% sequence homology with Cry21 Aa. Alignments of the Cry21s indicated in this application against known Cry genes including Ciy21s indicate a potential new or different mode of action that could be combined with other nematode resistance genes to control nematodes.
- FIG. 1 depicts the plant transformation vector for the expression of the Axmi296 in planta.
- Table 1 shows the description of the genetics elements of the construct.
- Table 2 shows the references for the genetic elements of the construct.
- FIG. 2 depicts a NEEDLE Sequence alignment of Axmi296 and Cry21 Aa2.
- the “DRIE” motif separating the N-terminal domain from the C-terminal crystal domain, is indicated by Identical amino acids are shaded black, a consensus sequence (Cons) is given below the aligned sequences.
- Cons consensus sequence
- Soybean transformation is achieved using methods well-known in the art, such as the one described using the Agrobacterium tumefaciens mediated transformation soybean half-seed explants using essentially the method described by Paz et al. (2006), Plant Cell Rep. 25:206. Transformants are identified using tembotrione as selection marker. The appearance of green shoots was observed, and documented as an indicator of tolerance to the herbicide isoxaflutole or tembotrione.
- the tolerant transgenic shoots will show normal greening comparable to wild-type soybean shoots not treated with isoxaflutole or tembotrione, whereas wild-type soybean shoots treated with the same amount of isoxaflutole or tembotrione will be entirely bleached. This indicates that the presence of the HPPD protein enables the tolerance to HPPD inhibitor herbicides, like isoxaflutole or tembotrione.
- Tolerant green shoots are transferred to rooting media or grafted. Rooted plantlets are transferred to the greenhouse after an acclimation period. Plants containing the transgene are selected for in transformation media.
- FIGs. 3a and 3b show the effectiveness of Axmi296 against Pratylenchus brachyurus.
- Greenhouse data shows an increase Pratylenchus brachyurus mortality as compared to Thome WT (wild-type, a non-resistant control) and a slight advantage to soybean Ciyl4 (a known resistant control).
- FIGs. 4a and 4b show that Axmi296 provides increased yield benefit in SCN field trials (3 independent sites) over SCN non-resistant controls.
- FIG 4c shows field trial results of the novel Ciy21 homolog (Axmi296) reduces SCN cysts on roots 20%, compared with wild-type Thome.
- FIG. 4d shows field trial results of the novel Cry21 homolog (Axmi296) produces an average yield increase of about 2% compared with wild-type Thome in an SCN-infested field.
- FIG. 5 depicts Axmi296 plants (left) next to non-resistant controls soybean plants in and SCN-infested field (right).
- Table 3 depicts the novel Cry21 homolog and further Cry21 homologous sequences. Protein sequence identities to closest Cry holotypes, are as determined by pairwise alignments according Crickmore etai, 2021.
- Table 4 depicts the sequence listing of the novel Ciy21 homolog sequences disclosed herein.
Landscapes
- Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Engineering & Computer Science (AREA)
- Molecular Biology (AREA)
- Biophysics (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Zoology (AREA)
- Biochemistry (AREA)
- Wood Science & Technology (AREA)
- General Health & Medical Sciences (AREA)
- General Engineering & Computer Science (AREA)
- Biotechnology (AREA)
- Biomedical Technology (AREA)
- Crystallography & Structural Chemistry (AREA)
- Physics & Mathematics (AREA)
- Cell Biology (AREA)
- Plant Pathology (AREA)
- Gastroenterology & Hepatology (AREA)
- Microbiology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Medicinal Chemistry (AREA)
- Agricultural Chemicals And Associated Chemicals (AREA)
- Peptides Or Proteins (AREA)
- Breeding Of Plants And Reproduction By Means Of Culturing (AREA)
Abstract
Description
Claims
Priority Applications (3)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| AU2024386241A AU2024386241A1 (en) | 2023-07-07 | 2024-07-05 | Use of cry genes for the control of nematode pests |
| IL325704A IL325704A (en) | 2023-07-07 | 2025-12-31 | Use of cry genes for the control of nematode pests |
| CONC2026/0000073A CO2026000073A2 (en) | 2023-08-21 | 2026-01-06 | Use of novel genes for nematode pest control |
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363512422P | 2023-07-07 | 2023-07-07 | |
| US63/512,422 | 2023-07-07 | ||
| US202363520679P | 2023-08-21 | 2023-08-21 | |
| US63/520,679 | 2023-08-21 |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| WO2025111030A2 true WO2025111030A2 (en) | 2025-05-30 |
| WO2025111030A3 WO2025111030A3 (en) | 2025-10-09 |
Family
ID=95696864
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/036850 Pending WO2025111030A2 (en) | 2023-07-07 | 2024-07-05 | Use of novel genes for the control of nematode pests |
Country Status (4)
| Country | Link |
|---|---|
| AU (1) | AU2024386241A1 (en) |
| IL (1) | IL325704A (en) |
| UY (1) | UY40814A (en) |
| WO (1) | WO2025111030A2 (en) |
Citations (115)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US1506602A (en) | 1922-05-17 | 1924-08-26 | Nichols Henry | Vehicle wheel |
| US4535060A (en) | 1983-01-05 | 1985-08-13 | Calgene, Inc. | Inhibition resistant 5-enolpyruvyl-3-phosphoshikimate synthetase, production and use |
| WO1987007644A1 (en) | 1986-06-04 | 1987-12-17 | Diatech Limited | TRANSLATION OF mRNA |
| US4771002A (en) | 1984-02-24 | 1988-09-13 | Lubrizol Genetics, Inc. | Transcription in plants and bacteria |
| EP0332104A2 (en) | 1988-03-08 | 1989-09-13 | Ciba-Geigy Ag | Chemically regulatable DNA sequences and genes and uses thereof |
| US4940835A (en) | 1985-10-29 | 1990-07-10 | Monsanto Company | Glyphosate-resistant plants |
| US4945050A (en) | 1984-11-13 | 1990-07-31 | Cornell Research Foundation, Inc. | Method for transporting substances into living cells and tissues and apparatus therefor |
| US4971908A (en) | 1987-05-26 | 1990-11-20 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvyl-3-phosphoshikimate synthase |
| WO1991000915A1 (en) | 1989-07-11 | 1991-01-24 | Biotechnology Research & Development Corporation | Aerosol beam microinjector |
| WO1991002071A2 (en) | 1989-08-09 | 1991-02-21 | Dekalb Plant Genetics | Methods and compositions for the production of stably transformed, fertile monocot plants and cells thereof |
| US5039523A (en) | 1988-10-27 | 1991-08-13 | Mycogen Corporation | Novel Bacillus thuringiensis isolate denoted B.t. PS81F, active against lepidopteran pests, and a gene encoding a lepidopteran-active toxin |
| EP0451878A1 (en) | 1985-01-18 | 1991-10-16 | Plant Genetic Systems, N.V. | Modifying plants by genetic engineering to combat or control insects |
| US5094945A (en) | 1983-01-05 | 1992-03-10 | Calgene, Inc. | Inhibition resistant 5-enolpyruvyl-3-phosphoshikimate synthase, production and use |
| US5102796A (en) | 1983-04-15 | 1992-04-07 | Lubrizol Genetics, Inc. | Plant structural gene expression |
| EP0480762A2 (en) | 1990-10-12 | 1992-04-15 | Mycogen Corporation | Novel bacillus thuringiensis isolates active against dipteran pests |
| US5106739A (en) | 1989-04-18 | 1992-04-21 | Calgene, Inc. | CaMv 355 enhanced mannopine synthase promoter and method for using same |
| US5145783A (en) | 1987-05-26 | 1992-09-08 | Monsanto Company | Glyphosate-tolerant 5-endolpyruvyl-3-phosphoshikimate synthase |
| WO1992017580A1 (en) | 1991-04-08 | 1992-10-15 | Rhone-Poulenc Agrochimie | Chimeric plant genes based on upstream regulatory elements of helianthinin |
| US5164316A (en) | 1987-01-13 | 1992-11-17 | The University Of British Columbia | DNA construct for enhancing the efficiency of transcription |
| US5182200A (en) | 1985-04-22 | 1993-01-26 | Lubrizol Genetics, Inc. | T-dna promoters |
| US5188642A (en) | 1985-08-07 | 1993-02-23 | Monsanto Company | Glyphosate-resistant plants |
| WO1993021334A1 (en) | 1992-04-13 | 1993-10-28 | Zeneca Limited | Dna constructs and plants incorporating them |
| EP0255378B1 (en) | 1986-07-31 | 1994-04-13 | Calgene, Inc. | Seed specific transcriptional regulation |
| US5310667A (en) | 1989-07-17 | 1994-05-10 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvyl-3-phosphoshikimate synthases |
| US5312910A (en) | 1987-05-26 | 1994-05-17 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvyl-3-phosphoshikimate synthase |
| US5378619A (en) | 1989-10-31 | 1995-01-03 | Monsanto Company | Promoter for transgenic plants |
| US5380831A (en) | 1986-04-04 | 1995-01-10 | Mycogen Plant Science, Inc. | Synthetic insecticidal crystal protein gene |
| WO1995006128A2 (en) | 1993-08-25 | 1995-03-02 | Dekalb Genetics Corporation | Fertile, transgenic maize plants and methods for their production |
| WO1995014098A1 (en) | 1993-11-19 | 1995-05-26 | Biotechnology Research And Development Corporation | Chimeric regulatory regions and gene cassettes for expression of genes in plants |
| US5428147A (en) | 1983-04-15 | 1995-06-27 | Mycogen Plant Science, Inc. | Octopine T-DNA promoters |
| US5436391A (en) | 1991-11-29 | 1995-07-25 | Mitsubishi Corporation | Synthetic insecticidal gene, plants of the genus oryza transformed with the gene, and production thereof |
| US5463175A (en) | 1990-06-25 | 1995-10-31 | Monsanto Company | Glyphosate tolerant plants |
| US5510474A (en) | 1988-05-17 | 1996-04-23 | Mycogen Plant Science, Inc. | Plant ubiquitin promoter system |
| US5510471A (en) | 1991-03-05 | 1996-04-23 | Rhone-Poulenc Agrochimie | Chimeric gene for the transformation of plants |
| WO1996023898A1 (en) | 1995-01-31 | 1996-08-08 | Novo Nordisk A/S | A method of detecting biologically active substances |
| US5563328A (en) | 1992-08-19 | 1996-10-08 | Board Of Regents, University Of Nebraska-Lincoln | Promoters from chlorella virus genes providing for expression of genes in prokaryotic and eukaryotic hosts |
| WO1996038567A2 (en) | 1995-06-02 | 1996-12-05 | Rhone-Poulenc Agrochimie | Dna sequence of a gene of hydroxy-phenyl pyruvate dioxygenase and production of plants containing a gene of hydroxy-phenyl pyruvate dioxygenase and which are tolerant to certain herbicides |
| WO1997006269A1 (en) | 1995-08-03 | 1997-02-20 | Zeneca Limited | Inducible herbicide resistance |
| US5605793A (en) | 1994-02-17 | 1997-02-25 | Affymax Technologies N.V. | Methods for in vitro recombination |
| US5627061A (en) | 1990-08-31 | 1997-05-06 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvylshikimate-3-phosphate synthases |
| WO1997017432A1 (en) | 1995-11-06 | 1997-05-15 | Wisconsin Alumni Research Foundation | Insecticidal protein toxins from photorhabdus |
| US5641876A (en) | 1990-01-05 | 1997-06-24 | Cornell Research Foundation, Inc. | Rice actin gene and promoter |
| US5670349A (en) | 1993-08-02 | 1997-09-23 | Virginia Tech Intellectual Properties, Inc. | HMG2 promoter expression system and post-harvest production of gene products in plants and plant cell cultures |
| WO1997041228A2 (en) | 1996-05-01 | 1997-11-06 | Pioneer Hi-Bred International, Inc. | Use of the green fluorescent protein as a screenable marker for plant transformation |
| WO1998008932A1 (en) | 1996-08-29 | 1998-03-05 | Dow Agrosciences Llc | Insecticidal protein toxins from $i(photorhabdus) |
| AU689311B2 (en) | 1994-08-30 | 1998-03-26 | Commonwealth Scientific And Industrial Research Organisation | Plant transcription regulators from circovirus |
| WO1998045461A1 (en) | 1997-04-09 | 1998-10-15 | Rhone-Poulenc Agro | An oleosin 5' regulatory region for the modification of plant seed lipid composition |
| WO1998045445A1 (en) | 1997-04-09 | 1998-10-15 | The Minister Of Agriculture Fisheries And Food In Her Britannic Majesty's Government Of The United Kingdom Of Great Britain And Nothern Ireland | Inducible plant promoters |
| WO1998045460A1 (en) | 1997-04-09 | 1998-10-15 | Rhone-Poulenc Agro | A sunflower albumin 5' regulatory region for the modification of plant seed lipid composition |
| WO1998050427A1 (en) | 1997-05-05 | 1998-11-12 | Dow Agrosciences Llc | INSECTICIDAL PROTEIN TOXINS FROM $i(XENORHABDUS) |
| US5850019A (en) | 1996-08-06 | 1998-12-15 | University Of Kentucky Research Foundation | Promoter (FLt) for the full-length transcript of peanut chlorotic streak caulimovirus (PCLSV) and expression of chimeric genes in plants |
| US5880275A (en) | 1989-02-24 | 1999-03-09 | Monsanto Company | Synthetic plant genes from BT kurstaki and method for preparation |
| US6153401A (en) | 1986-08-29 | 2000-11-28 | Hoechst Schering Agrevo Gnbh | Microorganisms and plasmids for 2,4-dichlorophenoxyacetic acid (2,4-D) monooxygenase formation and process for the production of these plasmids and strains |
| US6218188B1 (en) | 1997-11-12 | 2001-04-17 | Mycogen Corporation | Plant-optimized genes encoding pesticidal toxins |
| WO2001047952A2 (en) | 1999-12-28 | 2001-07-05 | Bayer Cropscience N.V. | Insecticidal proteins from bacillus thuringiensis |
| US6281016B1 (en) | 1996-11-20 | 2001-08-28 | Monsanto Company | Broad-spectrum insect resistant transgenic plants |
| US6291156B1 (en) | 1997-04-03 | 2001-09-18 | Syngenta Participations Ag | Plant pest control |
| US20010026941A1 (en) | 1999-11-29 | 2001-10-04 | Held Bruce Marvin | Methods and compositions for the introduction of molecules into cells |
| US6326169B1 (en) | 1996-11-20 | 2001-12-04 | Monsanto Company | Polynucleotide compositions encoding Cry1Ac/Cry1F chimeric O-endotoxins |
| WO2002057664A2 (en) | 2001-01-09 | 2002-07-25 | Bayer Bioscience N.V. | Bacillus thuringiensis insecticidal proteins |
| US6468523B1 (en) | 1998-11-02 | 2002-10-22 | Monsanto Technology Llc | Polypeptide compositions toxic to diabrotic insects, and methods of use |
| US6566587B1 (en) | 1995-07-19 | 2003-05-20 | Bayer Cropscience S.A. | Mutated 5-enolpyruvylshikimate-3-phosphate synthase, gene coding for said protein and transformed plants containing said gene |
| WO2004055191A1 (en) | 2002-12-17 | 2004-07-01 | Biogemma | Expression of hydroxyphenylpyruvate dioxygenase in plastids of plants for herbicide tolerance |
| WO2004074443A2 (en) | 2003-02-18 | 2004-09-02 | Monsanto Technology Llc | Glyphosate resistant class i 5-enolpyruvylshikimate-3-phosphate synthase (epsps) |
| US6791014B2 (en) | 2000-08-11 | 2004-09-14 | Aventis Cropscience, S.A. | Use of HPPD inhibitors as selection agents in plant transformation |
| US6855533B2 (en) | 1995-04-20 | 2005-02-15 | Basf Corporation | Structure-based designed herbicide resistant products |
| WO2005054479A1 (en) | 2003-12-01 | 2005-06-16 | Syngenta Participations Ag | Insect resistant cotton plants and methods of detecting the same |
| WO2005054480A2 (en) | 2003-12-01 | 2005-06-16 | Syngenta Participations Ag | Insect resistant cotton plants and methods of detecting the same |
| US7053205B1 (en) | 1996-06-20 | 2006-05-30 | The Scripps Research Institute | Cassava vein mosaic virus promoter nucleic acid sequences and expression vectors |
| WO2006108674A2 (en) | 2005-04-08 | 2006-10-19 | Bayer Bioscience N.V. | Elite event a2704-12 and methods and kits for identifying such event in biological samples |
| WO2006108675A2 (en) | 2005-04-11 | 2006-10-19 | Bayer Bioscience N.V. | Elite event a5547-127 and methods and kits for identifying such event in biological samples |
| WO2006130436A2 (en) | 2005-05-27 | 2006-12-07 | Monsanto Technology Llc | Soybean event mon89788 and methods for detection thereof |
| WO2007140256A1 (en) | 2006-05-26 | 2007-12-06 | Monsanto Technology, Llc | Corn plant and seed corresponding to transgenic event mon89034 and methods for detection and use thereof |
| WO2007142840A2 (en) | 2006-06-03 | 2007-12-13 | Syngenta Participations Ag | Corn event mir162 |
| WO2008002872A2 (en) | 2006-06-28 | 2008-01-03 | Pioneer Hi-Bred International, Inc. | Soybean event 3560.4.3.5 and compositions and methods for the identification and/or detection thereof |
| WO2008054747A2 (en) | 2006-10-31 | 2008-05-08 | E. I. Du Pont De Nemours And Company | Soybean event dp-305423-1 and compositions and methods for the identification and/or detection thereof |
| US20080120739A1 (en) | 2006-06-06 | 2008-05-22 | Monsanto Technology Llc | Method for selection of transformed cells |
| US20080119361A1 (en) | 2006-06-06 | 2008-05-22 | Feng Paul C C | Methods for weed control |
| US7393948B1 (en) | 2007-04-03 | 2008-07-01 | Ms Technologies, Inc. | Ubiquitin regulatory nucleic acids, vectors, and methods of using same |
| US7405074B2 (en) | 2004-04-29 | 2008-07-29 | Pioneer Hi-Bred International, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| WO2008100353A2 (en) | 2006-11-29 | 2008-08-21 | Athenix Corporation | Improved grg23 epsp synthases: compositions and methods of use |
| US7527955B2 (en) | 2000-10-30 | 2009-05-05 | Pioneer Hi-Bred International, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US20090130071A1 (en) | 2007-11-15 | 2009-05-21 | Ai-Guo Gao | Soybean Plant And Seed Corresponding To Transgenic Event MON87701 And Methods For Detection Thereof |
| US20090137409A1 (en) | 2007-10-09 | 2009-05-28 | Athenix Corporation | Computational methods for synthetic gene design |
| WO2009102873A1 (en) | 2008-02-15 | 2009-08-20 | Monsanto Technology Llc | Soybean plant and seed corresponding to transgenic event mon87769 and methods for detection thereof |
| WO2009152359A2 (en) | 2008-06-11 | 2009-12-17 | Dow Agrosciences Llc | Constructs for expressing herbicide tolerance genes, related plants, and related trait combinations |
| WO2010024976A1 (en) | 2008-08-29 | 2010-03-04 | Monsanto Technology Llc | Soybean plant and seed corresponding to transgenic event mon87754 and methods for detection thereof |
| US20100080887A1 (en) | 2008-09-29 | 2010-04-01 | Monsanto Technology Llc | Soybean Transgenic Event MON87705 and Methods for Detection Thereof |
| US20100166723A1 (en) | 2008-12-15 | 2010-07-01 | Athenix Corporation | Genes encoding nematode toxins |
| WO2010080829A1 (en) | 2009-01-07 | 2010-07-15 | Basf Agrochemical Products B.V. | Soybean event 127 and methods related thereto |
| WO2011028914A1 (en) | 2009-09-04 | 2011-03-10 | Syngenta Participations Ag | Stacking of translational enhancer elements to increase polypeptide expression in plants |
| WO2011034704A1 (en) | 2009-09-17 | 2011-03-24 | Monsanto Technology Llc | Soybean transgenic event mon 87708 and methods of use thereof |
| US7923602B2 (en) | 2006-06-14 | 2011-04-12 | Athenix Corp. | AXMI-031, AXMI-039, AXMI-040 and AXMI-049, a family of novel delta endotoxin genes and methods for their use |
| WO2011063413A2 (en) | 2009-11-23 | 2011-05-26 | Bayer Bioscience N.V. | Herbicide tolerant soybean plants and methods for identifying same |
| WO2011066360A1 (en) | 2009-11-24 | 2011-06-03 | Dow Agrosciences Llc | Detection of aad-12 soybean event 416 |
| WO2011066384A1 (en) | 2009-11-24 | 2011-06-03 | Dow Agrosciences Llc | Aad-12 event 416, related transgenic soybean lines, and event-specific identification thereof |
| WO2011068567A1 (en) | 2009-07-10 | 2011-06-09 | Syngenta Participations Ag | Novel hydroxyphenylpyruvate dioxygenase polypeptides and methods of use |
| WO2011076345A1 (en) | 2009-12-22 | 2011-06-30 | Bayer Bioscience N.V. | Herbicide tolerant plants |
| WO2011085221A2 (en) | 2010-01-07 | 2011-07-14 | Basf Agro B.V., Arnhem (Nl), Zweigniederlassung Wadenswil | Herbicide-tolerant plants |
| WO2011084370A1 (en) | 2010-01-05 | 2011-07-14 | Syngenta Participations Ag | Constitutive synthetic plant promoters and methods of use |
| WO2011094199A1 (en) | 2010-01-26 | 2011-08-04 | Pioneer Hi-Bred International, Inc. | Polynucleotide and polypeptide sequences associated with herbicide tolerance |
| WO2011145015A1 (en) | 2010-05-04 | 2011-11-24 | Basf Se | Plants having increased tolerance to herbicides |
| WO2012021794A1 (en) | 2010-08-13 | 2012-02-16 | Pioneer Hi-Bred International, Inc. | Chimeric promoters and methods of use |
| WO2012033794A2 (en) | 2010-09-08 | 2012-03-15 | Dow Agrosciences Llc | Aad-12 event 1606 and related transgenic soybean lines |
| WO2012051199A2 (en) | 2010-10-12 | 2012-04-19 | Monsanto Technology Llc | Soybean plant and seed corresponding to transgenic event mon87712 and methods for detection thereof |
| WO2012056401A1 (en) | 2010-10-25 | 2012-05-03 | Rosetta Green Ltd. | ISOLATED POLYNUCLEOTIDES EXPRESSING OR MODULATING MICRORNAs OR TARGETS OF SAME, TRANSGENIC PLANTS COMPRISING SAME AND USES THEREOF IN IMPROVING NITROGEN USE EFFICIENCY, ABIOTIC STRESS TOLERANCE, BIOMASS, VIGOR OR YIELD OF A PLANT |
| WO2012075429A1 (en) | 2010-12-03 | 2012-06-07 | Dow Agrosciences Llc | Stacked herbicide tolerance event 8291.45.36.2, related transgenic soybean lines, and detection thereof |
| WO2012075426A1 (en) | 2010-12-03 | 2012-06-07 | Dow Agrosciences Llc | Stacked herbicide tolerance event 8264.44.06.1, related transgenic soybean lines, and detection thereof |
| WO2012082548A2 (en) | 2010-12-15 | 2012-06-21 | Syngenta Participations Ag | Soybean event syht0h2 and compositions and methods for detection thereof |
| WO2013010094A1 (en) | 2011-07-13 | 2013-01-17 | Dow Agrosciences Llc | Stacked herbicide tolerance event 8264.42.32.1, related transgenic soybean lines, and detection thereof |
| US8395021B2 (en) | 2007-05-08 | 2013-03-12 | The Ohio State University Research Foundation | Highly active soybean promoter from the SUBI-3 polyubiquitin gene and uses thereof |
| WO2014043435A1 (en) | 2012-09-14 | 2014-03-20 | Bayer Cropscience Lp | Hppd variants and methods of use |
| WO2014150449A2 (en) | 2013-03-15 | 2014-09-25 | Bayer Cropscience Lp | Constitutive soybean promoters |
| US9806978B2 (en) | 2009-10-26 | 2017-10-31 | Amazon Technologies, Inc. | Monitoring of replicated data instances |
| WO2020001404A1 (en) | 2018-06-26 | 2020-01-02 | 比亚迪股份有限公司 | Internally-guided turnout and rail transportation system having same |
Family Cites Families (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20120110706A1 (en) * | 2008-08-25 | 2012-05-03 | Dow Agrosciences Llc | Modified bacillus thuringiensis cry21 proteins for nematode control |
| US11535653B2 (en) * | 2016-10-21 | 2022-12-27 | Vestaron Corporation | Cleavable peptides and insecticidal and nematicidal proteins comprising same |
| WO2018119336A1 (en) * | 2016-12-22 | 2018-06-28 | Athenix Corp. | Use of cry14 for the control of nematode pests |
-
2024
- 2024-07-05 WO PCT/US2024/036850 patent/WO2025111030A2/en active Pending
- 2024-07-05 UY UY0001040814A patent/UY40814A/en unknown
- 2024-07-05 AU AU2024386241A patent/AU2024386241A1/en active Pending
-
2025
- 2025-12-31 IL IL325704A patent/IL325704A/en unknown
Patent Citations (141)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US1506602A (en) | 1922-05-17 | 1924-08-26 | Nichols Henry | Vehicle wheel |
| US4535060A (en) | 1983-01-05 | 1985-08-13 | Calgene, Inc. | Inhibition resistant 5-enolpyruvyl-3-phosphoshikimate synthetase, production and use |
| US4769061A (en) | 1983-01-05 | 1988-09-06 | Calgene Inc. | Inhibition resistant 5-enolpyruvyl-3-phosphoshikimate synthase, production and use |
| US5094945A (en) | 1983-01-05 | 1992-03-10 | Calgene, Inc. | Inhibition resistant 5-enolpyruvyl-3-phosphoshikimate synthase, production and use |
| US5428147A (en) | 1983-04-15 | 1995-06-27 | Mycogen Plant Science, Inc. | Octopine T-DNA promoters |
| US5102796A (en) | 1983-04-15 | 1992-04-07 | Lubrizol Genetics, Inc. | Plant structural gene expression |
| US4771002A (en) | 1984-02-24 | 1988-09-13 | Lubrizol Genetics, Inc. | Transcription in plants and bacteria |
| US4945050A (en) | 1984-11-13 | 1990-07-31 | Cornell Research Foundation, Inc. | Method for transporting substances into living cells and tissues and apparatus therefor |
| EP0451878A1 (en) | 1985-01-18 | 1991-10-16 | Plant Genetic Systems, N.V. | Modifying plants by genetic engineering to combat or control insects |
| US5182200A (en) | 1985-04-22 | 1993-01-26 | Lubrizol Genetics, Inc. | T-dna promoters |
| US5188642A (en) | 1985-08-07 | 1993-02-23 | Monsanto Company | Glyphosate-resistant plants |
| US4940835A (en) | 1985-10-29 | 1990-07-10 | Monsanto Company | Glyphosate-resistant plants |
| US5380831A (en) | 1986-04-04 | 1995-01-10 | Mycogen Plant Science, Inc. | Synthetic insecticidal crystal protein gene |
| WO1987007644A1 (en) | 1986-06-04 | 1987-12-17 | Diatech Limited | TRANSLATION OF mRNA |
| EP0255378B1 (en) | 1986-07-31 | 1994-04-13 | Calgene, Inc. | Seed specific transcriptional regulation |
| US6153401A (en) | 1986-08-29 | 2000-11-28 | Hoechst Schering Agrevo Gnbh | Microorganisms and plasmids for 2,4-dichlorophenoxyacetic acid (2,4-D) monooxygenase formation and process for the production of these plasmids and strains |
| US5164316A (en) | 1987-01-13 | 1992-11-17 | The University Of British Columbia | DNA construct for enhancing the efficiency of transcription |
| US4971908A (en) | 1987-05-26 | 1990-11-20 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvyl-3-phosphoshikimate synthase |
| US5145783A (en) | 1987-05-26 | 1992-09-08 | Monsanto Company | Glyphosate-tolerant 5-endolpyruvyl-3-phosphoshikimate synthase |
| US5312910A (en) | 1987-05-26 | 1994-05-17 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvyl-3-phosphoshikimate synthase |
| EP0332104A2 (en) | 1988-03-08 | 1989-09-13 | Ciba-Geigy Ag | Chemically regulatable DNA sequences and genes and uses thereof |
| US5510474A (en) | 1988-05-17 | 1996-04-23 | Mycogen Plant Science, Inc. | Plant ubiquitin promoter system |
| US5039523A (en) | 1988-10-27 | 1991-08-13 | Mycogen Corporation | Novel Bacillus thuringiensis isolate denoted B.t. PS81F, active against lepidopteran pests, and a gene encoding a lepidopteran-active toxin |
| US5880275A (en) | 1989-02-24 | 1999-03-09 | Monsanto Company | Synthetic plant genes from BT kurstaki and method for preparation |
| US5106739A (en) | 1989-04-18 | 1992-04-21 | Calgene, Inc. | CaMv 355 enhanced mannopine synthase promoter and method for using same |
| WO1991000915A1 (en) | 1989-07-11 | 1991-01-24 | Biotechnology Research & Development Corporation | Aerosol beam microinjector |
| US5310667A (en) | 1989-07-17 | 1994-05-10 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvyl-3-phosphoshikimate synthases |
| WO1991002071A2 (en) | 1989-08-09 | 1991-02-21 | Dekalb Plant Genetics | Methods and compositions for the production of stably transformed, fertile monocot plants and cells thereof |
| US5378619A (en) | 1989-10-31 | 1995-01-03 | Monsanto Company | Promoter for transgenic plants |
| US5641876A (en) | 1990-01-05 | 1997-06-24 | Cornell Research Foundation, Inc. | Rice actin gene and promoter |
| US5463175A (en) | 1990-06-25 | 1995-10-31 | Monsanto Company | Glyphosate tolerant plants |
| US5627061A (en) | 1990-08-31 | 1997-05-06 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvylshikimate-3-phosphate synthases |
| US5633435A (en) | 1990-08-31 | 1997-05-27 | Monsanto Company | Glyphosate-tolerant 5-enolpyruvylshikimate-3-phosphate synthases |
| EP0480762A2 (en) | 1990-10-12 | 1992-04-15 | Mycogen Corporation | Novel bacillus thuringiensis isolates active against dipteran pests |
| US5510471A (en) | 1991-03-05 | 1996-04-23 | Rhone-Poulenc Agrochimie | Chimeric gene for the transformation of plants |
| US5633448A (en) | 1991-03-05 | 1997-05-27 | Rhone-Poulenc Agrochimie | Chimeric gene for the transformation of plants |
| WO1992017580A1 (en) | 1991-04-08 | 1992-10-15 | Rhone-Poulenc Agrochimie | Chimeric plant genes based on upstream regulatory elements of helianthinin |
| US5436391A (en) | 1991-11-29 | 1995-07-25 | Mitsubishi Corporation | Synthetic insecticidal gene, plants of the genus oryza transformed with the gene, and production thereof |
| WO1993021334A1 (en) | 1992-04-13 | 1993-10-28 | Zeneca Limited | Dna constructs and plants incorporating them |
| US5563328A (en) | 1992-08-19 | 1996-10-08 | Board Of Regents, University Of Nebraska-Lincoln | Promoters from chlorella virus genes providing for expression of genes in prokaryotic and eukaryotic hosts |
| US5670349A (en) | 1993-08-02 | 1997-09-23 | Virginia Tech Intellectual Properties, Inc. | HMG2 promoter expression system and post-harvest production of gene products in plants and plant cell cultures |
| WO1995006128A2 (en) | 1993-08-25 | 1995-03-02 | Dekalb Genetics Corporation | Fertile, transgenic maize plants and methods for their production |
| WO1995014098A1 (en) | 1993-11-19 | 1995-05-26 | Biotechnology Research And Development Corporation | Chimeric regulatory regions and gene cassettes for expression of genes in plants |
| US5605793A (en) | 1994-02-17 | 1997-02-25 | Affymax Technologies N.V. | Methods for in vitro recombination |
| AU689311B2 (en) | 1994-08-30 | 1998-03-26 | Commonwealth Scientific And Industrial Research Organisation | Plant transcription regulators from circovirus |
| WO1996023898A1 (en) | 1995-01-31 | 1996-08-08 | Novo Nordisk A/S | A method of detecting biologically active substances |
| US6855533B2 (en) | 1995-04-20 | 2005-02-15 | Basf Corporation | Structure-based designed herbicide resistant products |
| WO1996038567A2 (en) | 1995-06-02 | 1996-12-05 | Rhone-Poulenc Agrochimie | Dna sequence of a gene of hydroxy-phenyl pyruvate dioxygenase and production of plants containing a gene of hydroxy-phenyl pyruvate dioxygenase and which are tolerant to certain herbicides |
| US6566587B1 (en) | 1995-07-19 | 2003-05-20 | Bayer Cropscience S.A. | Mutated 5-enolpyruvylshikimate-3-phosphate synthase, gene coding for said protein and transformed plants containing said gene |
| WO1997006269A1 (en) | 1995-08-03 | 1997-02-20 | Zeneca Limited | Inducible herbicide resistance |
| WO1997017432A1 (en) | 1995-11-06 | 1997-05-15 | Wisconsin Alumni Research Foundation | Insecticidal protein toxins from photorhabdus |
| WO1997041228A2 (en) | 1996-05-01 | 1997-11-06 | Pioneer Hi-Bred International, Inc. | Use of the green fluorescent protein as a screenable marker for plant transformation |
| US7053205B1 (en) | 1996-06-20 | 2006-05-30 | The Scripps Research Institute | Cassava vein mosaic virus promoter nucleic acid sequences and expression vectors |
| US5850019A (en) | 1996-08-06 | 1998-12-15 | University Of Kentucky Research Foundation | Promoter (FLt) for the full-length transcript of peanut chlorotic streak caulimovirus (PCLSV) and expression of chimeric genes in plants |
| WO1998008932A1 (en) | 1996-08-29 | 1998-03-05 | Dow Agrosciences Llc | Insecticidal protein toxins from $i(photorhabdus) |
| US6281016B1 (en) | 1996-11-20 | 2001-08-28 | Monsanto Company | Broad-spectrum insect resistant transgenic plants |
| US6326169B1 (en) | 1996-11-20 | 2001-12-04 | Monsanto Company | Polynucleotide compositions encoding Cry1Ac/Cry1F chimeric O-endotoxins |
| US6291156B1 (en) | 1997-04-03 | 2001-09-18 | Syngenta Participations Ag | Plant pest control |
| WO1998045445A1 (en) | 1997-04-09 | 1998-10-15 | The Minister Of Agriculture Fisheries And Food In Her Britannic Majesty's Government Of The United Kingdom Of Great Britain And Nothern Ireland | Inducible plant promoters |
| WO1998045460A1 (en) | 1997-04-09 | 1998-10-15 | Rhone-Poulenc Agro | A sunflower albumin 5' regulatory region for the modification of plant seed lipid composition |
| WO1998045461A1 (en) | 1997-04-09 | 1998-10-15 | Rhone-Poulenc Agro | An oleosin 5' regulatory region for the modification of plant seed lipid composition |
| WO1998050427A1 (en) | 1997-05-05 | 1998-11-12 | Dow Agrosciences Llc | INSECTICIDAL PROTEIN TOXINS FROM $i(XENORHABDUS) |
| US6218188B1 (en) | 1997-11-12 | 2001-04-17 | Mycogen Corporation | Plant-optimized genes encoding pesticidal toxins |
| US6468523B1 (en) | 1998-11-02 | 2002-10-22 | Monsanto Technology Llc | Polypeptide compositions toxic to diabrotic insects, and methods of use |
| US20010026941A1 (en) | 1999-11-29 | 2001-10-04 | Held Bruce Marvin | Methods and compositions for the introduction of molecules into cells |
| WO2001047952A2 (en) | 1999-12-28 | 2001-07-05 | Bayer Cropscience N.V. | Insecticidal proteins from bacillus thuringiensis |
| US6791014B2 (en) | 2000-08-11 | 2004-09-14 | Aventis Cropscience, S.A. | Use of HPPD inhibitors as selection agents in plant transformation |
| US8044261B2 (en) | 2000-10-30 | 2011-10-25 | E. I. Du Pont De Nemours | Glyphosate-N-acetyltransferase (GAT) genes |
| US8008547B2 (en) | 2000-10-30 | 2011-08-30 | E.I. Dupont De Nemours | Glyphosate-N-acetyltransferase (GAT) genes |
| US7531339B2 (en) | 2000-10-30 | 2009-05-12 | Verdia, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US8088972B2 (en) | 2000-10-30 | 2012-01-03 | Verdia, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US7527955B2 (en) | 2000-10-30 | 2009-05-05 | Pioneer Hi-Bred International, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US7666643B2 (en) | 2000-10-30 | 2010-02-23 | Pioneer Hi-Bred International, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US8021857B2 (en) | 2000-10-30 | 2011-09-20 | Pioneer Hi-Bred International, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US7709702B2 (en) | 2000-10-30 | 2010-05-04 | Pioneer Hi-Bred Int'l., Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US7998703B2 (en) | 2000-10-30 | 2011-08-16 | Verdia, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US7999152B2 (en) | 2000-10-30 | 2011-08-16 | Pioneer Hi-Bred International, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US7714188B2 (en) | 2000-10-30 | 2010-05-11 | Pioneer Hi-Bred Int'l., Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| WO2002057664A2 (en) | 2001-01-09 | 2002-07-25 | Bayer Bioscience N.V. | Bacillus thuringiensis insecticidal proteins |
| WO2004055191A1 (en) | 2002-12-17 | 2004-07-01 | Biogemma | Expression of hydroxyphenylpyruvate dioxygenase in plastids of plants for herbicide tolerance |
| WO2004074443A2 (en) | 2003-02-18 | 2004-09-02 | Monsanto Technology Llc | Glyphosate resistant class i 5-enolpyruvylshikimate-3-phosphate synthase (epsps) |
| WO2005054479A1 (en) | 2003-12-01 | 2005-06-16 | Syngenta Participations Ag | Insect resistant cotton plants and methods of detecting the same |
| WO2005054480A2 (en) | 2003-12-01 | 2005-06-16 | Syngenta Participations Ag | Insect resistant cotton plants and methods of detecting the same |
| US7666644B2 (en) | 2004-04-29 | 2010-02-23 | Verdia, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US7405074B2 (en) | 2004-04-29 | 2008-07-29 | Pioneer Hi-Bred International, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US7863503B2 (en) | 2004-04-29 | 2011-01-04 | Pioneer Hi-Bred International, Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| US8222489B2 (en) | 2004-04-29 | 2012-07-17 | Verdia Inc. | Glyphosate-N-acetyltransferase (GAT) genes |
| WO2006108674A2 (en) | 2005-04-08 | 2006-10-19 | Bayer Bioscience N.V. | Elite event a2704-12 and methods and kits for identifying such event in biological samples |
| US20080320616A1 (en) | 2005-04-08 | 2008-12-25 | Bayer Bioscience N.V. | Elite Event A2407-12 and Methods and Kits for Identifying Such Event in Biological Samples |
| WO2006108675A2 (en) | 2005-04-11 | 2006-10-19 | Bayer Bioscience N.V. | Elite event a5547-127 and methods and kits for identifying such event in biological samples |
| US20080196127A1 (en) | 2005-04-11 | 2008-08-14 | Bayer Bioscience N.V. | Elite Event A5547-127 and Methods and Kits For Identifying Such Event in Biological Samples |
| WO2006130436A2 (en) | 2005-05-27 | 2006-12-07 | Monsanto Technology Llc | Soybean event mon89788 and methods for detection thereof |
| US20060282915A1 (en) | 2005-05-27 | 2006-12-14 | Monsanto Technology Llc | Soybean event MON89788 and methods for detection thereof |
| WO2007140256A1 (en) | 2006-05-26 | 2007-12-06 | Monsanto Technology, Llc | Corn plant and seed corresponding to transgenic event mon89034 and methods for detection and use thereof |
| WO2007142840A2 (en) | 2006-06-03 | 2007-12-13 | Syngenta Participations Ag | Corn event mir162 |
| US20080119361A1 (en) | 2006-06-06 | 2008-05-22 | Feng Paul C C | Methods for weed control |
| US20080120739A1 (en) | 2006-06-06 | 2008-05-22 | Monsanto Technology Llc | Method for selection of transformed cells |
| US7923602B2 (en) | 2006-06-14 | 2011-04-12 | Athenix Corp. | AXMI-031, AXMI-039, AXMI-040 and AXMI-049, a family of novel delta endotoxin genes and methods for their use |
| WO2008002872A2 (en) | 2006-06-28 | 2008-01-03 | Pioneer Hi-Bred International, Inc. | Soybean event 3560.4.3.5 and compositions and methods for the identification and/or detection thereof |
| US20100184079A1 (en) | 2006-06-28 | 2010-07-22 | Pioneer Hi-Bred International, Inc. | Soybean event 3560.4.3.5 and compositions and methods for the identification and detection thereof |
| WO2008054747A2 (en) | 2006-10-31 | 2008-05-08 | E. I. Du Pont De Nemours And Company | Soybean event dp-305423-1 and compositions and methods for the identification and/or detection thereof |
| US20080312082A1 (en) | 2006-10-31 | 2008-12-18 | Kinney Anthony J | Soybean event dp-305423-1 and compositions and methods for the identification and/or detection thereof |
| WO2008100353A2 (en) | 2006-11-29 | 2008-08-21 | Athenix Corporation | Improved grg23 epsp synthases: compositions and methods of use |
| US7393948B1 (en) | 2007-04-03 | 2008-07-01 | Ms Technologies, Inc. | Ubiquitin regulatory nucleic acids, vectors, and methods of using same |
| US8395021B2 (en) | 2007-05-08 | 2013-03-12 | The Ohio State University Research Foundation | Highly active soybean promoter from the SUBI-3 polyubiquitin gene and uses thereof |
| US20090137409A1 (en) | 2007-10-09 | 2009-05-28 | Athenix Corporation | Computational methods for synthetic gene design |
| WO2009064652A1 (en) | 2007-11-15 | 2009-05-22 | Monsanto Technology Llc | Soybean plant and seed corresponding to transgenic event mon87701 and methods for detection thereof |
| US20090130071A1 (en) | 2007-11-15 | 2009-05-21 | Ai-Guo Gao | Soybean Plant And Seed Corresponding To Transgenic Event MON87701 And Methods For Detection Thereof |
| WO2009102873A1 (en) | 2008-02-15 | 2009-08-20 | Monsanto Technology Llc | Soybean plant and seed corresponding to transgenic event mon87769 and methods for detection thereof |
| US20110067141A1 (en) | 2008-02-15 | 2011-03-17 | Byron Froman | Soybean plant and seed corresponding to transgenic event mon87769 and methods for detection thereof |
| WO2009152359A2 (en) | 2008-06-11 | 2009-12-17 | Dow Agrosciences Llc | Constructs for expressing herbicide tolerance genes, related plants, and related trait combinations |
| WO2010024976A1 (en) | 2008-08-29 | 2010-03-04 | Monsanto Technology Llc | Soybean plant and seed corresponding to transgenic event mon87754 and methods for detection thereof |
| WO2010037016A1 (en) | 2008-09-29 | 2010-04-01 | Monsanto Technology Llc | Soybean transgenic event mon87705 and methods for detection thereof |
| US20100080887A1 (en) | 2008-09-29 | 2010-04-01 | Monsanto Technology Llc | Soybean Transgenic Event MON87705 and Methods for Detection Thereof |
| US20100166723A1 (en) | 2008-12-15 | 2010-07-01 | Athenix Corporation | Genes encoding nematode toxins |
| WO2010080829A1 (en) | 2009-01-07 | 2010-07-15 | Basf Agrochemical Products B.V. | Soybean event 127 and methods related thereto |
| WO2011068567A1 (en) | 2009-07-10 | 2011-06-09 | Syngenta Participations Ag | Novel hydroxyphenylpyruvate dioxygenase polypeptides and methods of use |
| WO2011028914A1 (en) | 2009-09-04 | 2011-03-10 | Syngenta Participations Ag | Stacking of translational enhancer elements to increase polypeptide expression in plants |
| WO2011034704A1 (en) | 2009-09-17 | 2011-03-24 | Monsanto Technology Llc | Soybean transgenic event mon 87708 and methods of use thereof |
| US9806978B2 (en) | 2009-10-26 | 2017-10-31 | Amazon Technologies, Inc. | Monitoring of replicated data instances |
| WO2011063413A2 (en) | 2009-11-23 | 2011-05-26 | Bayer Bioscience N.V. | Herbicide tolerant soybean plants and methods for identifying same |
| WO2011066360A1 (en) | 2009-11-24 | 2011-06-03 | Dow Agrosciences Llc | Detection of aad-12 soybean event 416 |
| WO2011066384A1 (en) | 2009-11-24 | 2011-06-03 | Dow Agrosciences Llc | Aad-12 event 416, related transgenic soybean lines, and event-specific identification thereof |
| WO2011076345A1 (en) | 2009-12-22 | 2011-06-30 | Bayer Bioscience N.V. | Herbicide tolerant plants |
| WO2011084370A1 (en) | 2010-01-05 | 2011-07-14 | Syngenta Participations Ag | Constitutive synthetic plant promoters and methods of use |
| WO2011085221A2 (en) | 2010-01-07 | 2011-07-14 | Basf Agro B.V., Arnhem (Nl), Zweigniederlassung Wadenswil | Herbicide-tolerant plants |
| WO2011094199A1 (en) | 2010-01-26 | 2011-08-04 | Pioneer Hi-Bred International, Inc. | Polynucleotide and polypeptide sequences associated with herbicide tolerance |
| WO2011094205A1 (en) | 2010-01-26 | 2011-08-04 | Pioneer Hi-Bred International, Inc. | Hppd-inhibitor herbicide tolerance |
| WO2011145015A1 (en) | 2010-05-04 | 2011-11-24 | Basf Se | Plants having increased tolerance to herbicides |
| WO2012021794A1 (en) | 2010-08-13 | 2012-02-16 | Pioneer Hi-Bred International, Inc. | Chimeric promoters and methods of use |
| WO2012021797A1 (en) | 2010-08-13 | 2012-02-16 | Pioneer Hi-Bred International, Inc. | Methods and compositions for targeting sequences of interest to the chloroplast |
| WO2012033794A2 (en) | 2010-09-08 | 2012-03-15 | Dow Agrosciences Llc | Aad-12 event 1606 and related transgenic soybean lines |
| WO2012051199A2 (en) | 2010-10-12 | 2012-04-19 | Monsanto Technology Llc | Soybean plant and seed corresponding to transgenic event mon87712 and methods for detection thereof |
| WO2012056401A1 (en) | 2010-10-25 | 2012-05-03 | Rosetta Green Ltd. | ISOLATED POLYNUCLEOTIDES EXPRESSING OR MODULATING MICRORNAs OR TARGETS OF SAME, TRANSGENIC PLANTS COMPRISING SAME AND USES THEREOF IN IMPROVING NITROGEN USE EFFICIENCY, ABIOTIC STRESS TOLERANCE, BIOMASS, VIGOR OR YIELD OF A PLANT |
| WO2012075429A1 (en) | 2010-12-03 | 2012-06-07 | Dow Agrosciences Llc | Stacked herbicide tolerance event 8291.45.36.2, related transgenic soybean lines, and detection thereof |
| WO2012075426A1 (en) | 2010-12-03 | 2012-06-07 | Dow Agrosciences Llc | Stacked herbicide tolerance event 8264.44.06.1, related transgenic soybean lines, and detection thereof |
| WO2012082548A2 (en) | 2010-12-15 | 2012-06-21 | Syngenta Participations Ag | Soybean event syht0h2 and compositions and methods for detection thereof |
| WO2013010094A1 (en) | 2011-07-13 | 2013-01-17 | Dow Agrosciences Llc | Stacked herbicide tolerance event 8264.42.32.1, related transgenic soybean lines, and detection thereof |
| WO2014043435A1 (en) | 2012-09-14 | 2014-03-20 | Bayer Cropscience Lp | Hppd variants and methods of use |
| WO2014150449A2 (en) | 2013-03-15 | 2014-09-25 | Bayer Cropscience Lp | Constitutive soybean promoters |
| WO2020001404A1 (en) | 2018-06-26 | 2020-01-02 | 比亚迪股份有限公司 | Internally-guided turnout and rail transportation system having same |
Non-Patent Citations (76)
| Title |
|---|
| "CURRENT Protocols in Protein Science", 1995, JOHN WILEY & SONS, INC., article "PCR 2: A Practical Approach" |
| "EMBOSS", TRENDS IN GENETICS, vol. 16, no. 6, 2000, pages 276 |
| "European Molecular Biology Open Software Suite", TRENDS IN GENETICS, vol. VII, no. 6, 2000, pages 276 |
| "Molecular Biology and Biotechnology", 1995, VCH_ PUBLISHERS, INC., article "a Comprehensive Desk Reference" |
| "NCBI", Database accession no. ABG20429 |
| "The Encyclopedia of Molecular Biology", 1999, WILEY-INTERSCIENCE |
| ALTSCHUL ET AL., J. MOL. BIOL., vol. 215, pages 403 - 410 |
| ALTSCHUL ET AL., NUCLEIC ACIDS RES., vol. 15, 1987, pages 2343 - 2361 |
| ATANASSOVA ET AL., PLANT J., vol. 2, no. 3, 1992, pages 291 - 300 |
| AYRESPARK, CRITICAL REVIEWS IN PLANT SCIENCE, vol. 13, 1994, pages 219 - 239 |
| BOMMINENIJAUHAR, MAYDICA, vol. 42, 1997, pages 107 - 120 |
| CAMPBELLGOVVRI, PLANT PHYSIOL., vol. 92, 1990, pages 1 - 11 |
| CARRINGTONFREED, J., vol. 64, 1990, pages 1590 - 1597 |
| CHANG, METHODS ENZYMOL., vol. 153, 1987, pages 507 - 516 |
| CHRISTENSEN ET AL., PLANT MOL. BIOL., vol. 12, 1989, pages 619 - 632 |
| CHRISTENSEN ET AL., PLANT MOL. BIOL., vol. 18, 1992, pages 675 - 689 |
| CLARK, J. BIOL. CHEM., vol. 264, 1989, pages 17544 - 17550 |
| CONCIBIDO ET AL., CROP SCIENCE, vol. 44, no. 3, 2004, pages 1121 - 1131 |
| CRICKMORE ET AL., JOURNAL OF INVERTEBRATE PATHOLOGY, vol. 1, 2021, pages 86 |
| DATLA, R. ET AL., BIOTECHNOLOGY ANN. REV., vol. 3, 1997, pages 269 - 296 |
| DELLA-CIOPPA ET AL., PLANT PHYSIOL., vol. 84, 1987, pages 965 - 968 |
| ESTRUCH ET AL., PROC NAT! ACAD SCI, vol. 93, no. 11, 1996, pages 5389 - 94 |
| FFRENCH-CONSTANTBOWEN, CELL MOL LIFE SCI., vol. 57, no. 5, 2000, pages 828 - 33 |
| GATZ ET AL., MOL. GEN. GENETICS, vol. 243, 1994, pages 173 - 177 |
| GREFEN ET AL., PLANT J, vol. 64, 2010, pages 355 - 365 |
| GUERINEAU ET AL., MOL. GEN. GENET., vol. 262, 1991, pages 141 - 144 |
| HAYMES ET AL.: "Nucleic Acid Hybridization", 1985, IRL PRESS, article "A. Practical Approach" |
| HELLENSMULLINEAUX, TRENDS IN PLANT SCIENCE, vol. 5, 2000, pages 446 - 451 |
| HERSHEY ET AL., MOL. GEN. GENETICS, vol. 227, 1991, pages 229 - 237 |
| HIEI ET AL., THE PLANT JOURNAL, vol. 6, 1994, pages 271 - 282 |
| ISHIDA ET AL., NATURE BIOTECHNOLOGY, vol. 14, 1996, pages 745 - 750 |
| J PROTEOME RES., vol. 8, no. 2, February 2009 (2009-02-01), pages 787 - 797 |
| JOSHI ET AL., NUCLEIC ACID RES., vol. 15, 1987, pages 9627 - 9639 |
| KAHN, T.W.DUCK, N.B.MCCARVILLE, M.T. ET AL.: "A Bacillus thuringiensis Cry protein controls soybean cyst nematode in transgenic soybean plants.", NOT COMMUN., vol. 12, 2021, pages 3380, Retrieved from the Internet <URL:https://doi.org/10.1038/s41467-021-23743-3> |
| KAY ET AL., SCIENCE, vol. 236, 1987, pages 1299 - 1302 |
| KAZI ET AL., THEOR. APPL. GEN., vol. 120, no. 3, 2010, pages 633 - 644 |
| KIM ET AL., THEOR. APPL. GENET., vol. 129, no. 12, 2016, pages 2295 - 2311 |
| KIMDIERS, CROP SCIENCE, vol. 53, 2013, pages 775 - 785 |
| LAST ET AL., THEOR. APPL. GENET., vol. 81, 1991, pages 581 - 588 |
| LEPETIT ET AL., MOL. GEN. GENET., vol. 231, 1992, pages 276 - 285 |
| M. D. HAYWARDN, O BOSEMARKI. ROMAGOSA: "Plant Breeding", vol. 1, 1993, CHAPMAN & HALL, article "Plant Breeding: Principles and Prospects" |
| MAITI ET AL., TRANSGENIC RES., vol. 6, 1997, pages 143 - 156 |
| MCBRIDE ET AL., PROC. NATL. ACAD. SCI., vol. 91, 1994, pages 7301 - 7305 |
| MCCORMICK ET AL., PLANT CELL REPORTS, vol. 5, 1986, pages 81 - 84 |
| MCELROY ET AL., PLANT CELL, vol. 2, 1990, pages 1261 - 1272 |
| METT ET AL., PNAS, vol. 90, 1993, pages 45067 - 4571 |
| MUNROE ET AL., GENE, vol. 91, 1990, pages 151 - 158 |
| MURRAY ET AL., NUCL. ACIDS RES., vol. 17, 1989, pages 477 - 498 |
| MURRAY ET AL., NUCLEIC ACIDS RES., vol. 17, 1989, pages 7891 - 7903 |
| NAR, vol. 15, 1987, pages 6643 - 6653 |
| NEEDLEMANWUNSCH, J. MOL. BIOL, vol. 48, 1979, pages 443 - 453 |
| NI ET AL., PLANT J., vol. 7, 1995, pages 661 - 676 |
| NIBLACK, T. L. ET AL.: "A standard greenhouse method for assessing soybean cyst nematode resistance in soybean: SCE08 (Standardized Cyst Evaluation", PLANT HEALTH PROG., vol. 10, 2008, pages 33 |
| ODELL ET AL., NATURE, vol. 313, 1985, pages 810 - 812 |
| PAZ ET AL., PLANT CELL REP, vol. 25, 2006, pages 206 |
| PROUDFOOT, CELL, vol. 64, 1991, pages 671 - 674 |
| ROBERTS ET AL., PROC. NATL. ACAD. SCI., vol. 76, 1979, pages 7760 - 764 |
| ROMER ET AL., BIOCHEM. BIOPHYS. RES. COMMUN., vol. 196, 1993, pages 1414 - 1421 |
| SAMBROOKRUSSELL: "Molecular Cloning: A Laboratory Manual", 2001, COLD SPRING HARBOR LABORATORY PRESS |
| SANFACON ET AL., GENES DEV., vol. 5, 1991, pages 141 - 149 |
| SATHASIVAN ET AL., NUCL. ACIDS RES., vol. 18, 1990, pages 2188 |
| SCHENA ET AL., PROC. NATL. ACAD. SCI., vol. 88, no. 1, 1991 |
| SCHNEPF ET AL., BLOCK 5, 1998 |
| SHAH ET AL., SCIENCE, vol. 233, 1986, pages 478 - 481 |
| STALKER ET AL., J. BIOL. CHEM., vol. 263, 1985, pages 6310 - 6314 |
| STEMMER ET AL., NATURE, vol. 370, 1994, pages 389 - 391 |
| SVAB ET AL., PROC. NATL. ACAD. SCI., vol. 87, 1990, pages 8526 - 8530 |
| SVAB, PROC. NATL., vol. 90, 1993, pages 913 - 917 |
| SVABMALIGA, EMBO J., vol. 12, 1993, pages 601 - 606 |
| TIJSSEN: "Laboratory Techniques in Biochemistry and Molecular Biology-Hybridization with Nucleic Acid Probes", 1993, ELSEVIER, article "Overview of principles of hybridization and the strategy of nucleic acid probe assays" |
| VAN DEN BERG ET AL., BIOTECHNOLOGY, vol. 8, 1990, pages 135 - 139 |
| VELTEN ET AL., FAROJ., vol. 3, 1984, pages 2723 - 2730 |
| VON HEIJNE ET AL., PLANT MOL. BIOL. REP., vol. 9, 1991, pages 104 - 126 |
| WATERFIELD ET AL., APPL ENVIRON MICROBIOL., vol. 67, no. 11, 2001, pages 5017 - 24 |
| WEBB ET AL., THEOR. APPL. GENET., vol. 91, 1995, pages 574 - 581 |
| ZUO ET AL., PLANT J., vol. 24, 2000, pages 265 - 273 |
Also Published As
| Publication number | Publication date |
|---|---|
| WO2025111030A3 (en) | 2025-10-09 |
| UY40814A (en) | 2024-12-31 |
| AU2024386241A1 (en) | 2026-01-08 |
| IL325704A (en) | 2026-02-01 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US10875897B2 (en) | AXMI205 variant proteins and methods for their use | |
| CN102421905B (en) | Insecticidal proteins and use their method | |
| CN108064233B (en) | Insecticidal protein and method of use | |
| US12054729B2 (en) | Use of CRY14 for the control of nematode pests | |
| CN114075267A (en) | Insecticidal proteins and methods of use thereof | |
| JP2020058357A (en) | Toxin gene and use method thereof | |
| CN104302662B (en) | Bacillus thuringiensis toxin gene AXMI335 and methods of use thereof | |
| CA2901160C (en) | Use of axmi184 for the control of rootworm insects | |
| CN114787360B (en) | Novel insect resistance genes and methods of use | |
| WO2025111030A2 (en) | Use of novel genes for the control of nematode pests | |
| WO2025090606A1 (en) | Use of novel genes for the control of nematode pests | |
| US11286498B2 (en) | Use of BP005 for the control of plant pathogens | |
| CN110267975A (en) | AXMI669 and AXMI991 toxin genes and methods of use thereof | |
| WO2015077525A1 (en) | Use of axmi-011 for the control of hemipteran insects | |
| WO2026010930A1 (en) | Use of axmi277 for the control of rotylenchulus reniformis nematode pests | |
| BR112019012897B1 (en) | METHODS FOR CONTROLLING PEST POPULATION, PROTECTING PLANTS AGAINST PESTS AND INCREASING PLANT PRODUCTION | |
| EP4461128A2 (en) | Novel insect resistant genes and methods of use | |
| CN120380151A (en) | Insect toxin genes and methods of use thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| WWE | Wipo information: entry into national phase |
Ref document number: AU2024386241 Country of ref document: AU |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 325704 Country of ref document: IL |
|
| ENP | Entry into the national phase |
Ref document number: 2024386241 Country of ref document: AU Date of ref document: 20240705 Kind code of ref document: A |
|
| REG | Reference to national code |
Ref country code: BR Ref legal event code: B01A Ref document number: 112026000256 Country of ref document: BR |