WO2025106766A1 - Fusion-modified enzyme constructs - Google Patents
Fusion-modified enzyme constructs Download PDFInfo
- Publication number
- WO2025106766A1 WO2025106766A1 PCT/US2024/056055 US2024056055W WO2025106766A1 WO 2025106766 A1 WO2025106766 A1 WO 2025106766A1 US 2024056055 W US2024056055 W US 2024056055W WO 2025106766 A1 WO2025106766 A1 WO 2025106766A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- seq
- amino acid
- streptavidin
- fusion construct
- acid sequence
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/10—Transferases (2.)
- C12N9/12—Transferases (2.) transferring phosphorus containing groups, e.g. kinases (2.7)
- C12N9/1241—Nucleotidyltransferases (2.7.7)
- C12N9/1276—RNA-directed DNA polymerase (2.7.7.49), i.e. reverse transcriptase or telomerase
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/10—Transferases (2.)
- C12N9/12—Transferases (2.) transferring phosphorus containing groups, e.g. kinases (2.7)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P19/00—Preparation of compounds containing saccharide radicals
- C12P19/26—Preparation of nitrogen-containing carbohydrates
- C12P19/28—N-glycosides
- C12P19/30—Nucleotides
- C12P19/34—Polynucleotides, e.g. nucleic acids, oligoribonucleotides
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y207/00—Transferases transferring phosphorus-containing groups (2.7)
- C12Y207/07—Nucleotidyltransferases (2.7.7)
- C12Y207/07049—RNA-directed DNA polymerase (2.7.7.49), i.e. telomerase or reverse-transcriptase
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/20—Fusion polypeptide containing a tag with affinity for a non-protein ligand
- C07K2319/22—Fusion polypeptide containing a tag with affinity for a non-protein ligand containing a Strep-tag
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/10—Processes for the isolation, preparation or purification of DNA or RNA
- C12N15/1096—Processes for the isolation, preparation or purification of DNA or RNA cDNA Synthesis; Subtracted cDNA library construction, e.g. RT, RT-PCR
Definitions
- Streptavidin is one of the most important protein conjugation reagents, as it facilitates bridging of molecules and anchoring to biotinylated surfaces or nanoparticles.
- its tetrameric structure makes it impractical for creating single-enzyme fusion constructs.
- streptavidin fusion proteins form tetramers, the four proteins of interest fused to streptavidin become compacted within a confined space, which leads to steric hindrance, and reduction in the activity of attached proteins.
- chemically-conjugated streptavidin proteins are commonly employed for affinity tagging of proteins with streptavidin.
- streptavidin fusion proteins have the potential to offer several advantages over conjugates, including greater homogeneity, ease of scale-up, and lower production costs.
- the present disclosure features a bifunctional fusion construct comprising an affinity domain operably linked to an enzyme domain, for example, a polymerase domain.
- the affinity domain comprises avidin or streptavidin.
- the present disclosure relates to a fusion construct comprising avidin or streptavidin operably linked to a functional protein or enzyme domain, wherein the avidin or streptavidin can form a tetramer with additional avidin or streptavidin molecules or additional avidin or streptavidin fusion enzymes.
- the fusion construct comprises streptavidin comprising an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO: 1.
- the functional protein or enzyme domain comprises a polymerase or reverse transcriptase.
- the functional protein or enzyme domain comprises Marathon Reverse Transcriptase (MRT), or a variant thereof.
- MRT Marathon Reverse Transcriptase
- the functional protein or enzyme domain comprises an amino acid sequence at least 90% identical to an amino acid sequence selected from the group consisting SEQ ID NOs: 3 and 8-12 or a functional fragment thereof.
- the fusion construct comprises an amino acid sequence as set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the disclosure relates to a nucleic acid molecule encoding a fusion construct comprising avidin or streptavidin operably linked to a functional protein or enzyme domain, wherein the streptavidin can form a tetramer with additional streptavidin molecules or additional streptavidin fusion enzymes.
- the nucleic acid molecule encodes a streptavidin fusion construct, wherein the streptavidin comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO:1
- the functional protein or enzyme domain comprises a polymerase, reverse transcriptase, helicase, ligase, receptor protein, kinase, phosphatase, exonuclease and non-specific endonuclease, restriction endonuclease, DNA/RNA methyl transferase, recombinase, terminal transferase, nucleic acid binding protein, protease, ribosomal protein, aminoacyl-tRNA synthetase, glycosyltransferase, fatty acid synthase, or an enzyme for food, biofuel or pharmaceutical industries.
- the functional protein or enzyme domain comprises MRT, or a variant thereof.
- the functional protein or enzyme domain comprises an amino acid sequence at least 90% identical to an amino acid sequence selected from the group consisting SEQ ID NOs: 3 and 8-12 or a functional fragment thereof.
- the streptavidin fusion enzyme comprises an amino acid sequence as set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the nucleic acid molecule comprises the nucleotide sequence set forth in SEQ ID NO: 17.
- the disclosure relates to a composition comprising at least one fusion construct, and further wherein the functional protein or enzyme domain maintains at least one wild type activity or function of the protein or enzyme when the avidin or streptavidin is complexed as a tetramer.
- the composition comprises a tetramer comprising at least one avidin or streptavidin fusion construct.
- the composition comprises a tetramer comprising at least two avidin or streptavidin fusion constructs.
- the composition comprises a tetramer comprising at least three avidin or streptavidin fusion constructs.
- the tetramer is linked or bound to a solid support.
- the solid support is coated with or attached to at least one biotin molecule.
- the solid support is comprised in at least one device.
- the at least one device comprises a device used for complementary DNA (cDNA) synthesis.
- cDNA complementary DNA
- the at least one device is used for nucleic acid synthesis or sequencing.
- the present disclosure relates to a method for producing a nucleic acid comprising contacting a first nucleic acid molecule with the fusion construct or a composition of the present disclosure under conditions sufficient for the synthesis of a second nucleic acid molecule complementary to all or a portion of the first nucleic acid molecule.
- the present disclosure relates to a method for producing an RNA comprising contacting a first RNA molecule with the fusion construct or a composition of the present disclosure under conditions sufficient for the synthesis of a second RNA molecule complementary to all or a portion of the first RNA molecule.
- the present disclosure relates to a method for producing cDNA, comprising contacting at least one mRNA molecule with the fusion construct or a composition of the present disclosure, under conditions sufficient for the synthesis of at least one nucleic acid molecule complementary to all or a portion of the at least one mRNA molecule.
- the disclosure relates to a method for determining the identity of each of a series of consecutive nucleotide residues of at least one target nucleic acid, comprising contacting the at least one target nucleic acid with the fusion construct or a composition of the present disclosure, under conditions sufficient for the synthesis of at least one nucleic acid molecule complementary to all or a portion of the at least one target nucleic acid, and for the determination of each nucleotide in the at least one complementary nucleic acid.
- Figure 1 comprising Figure 1 A and Figure IB, depicts an illustration of producing streptavidin-MarathonRT fusion protein tetramer containing reduced number of MarathonRT domains.
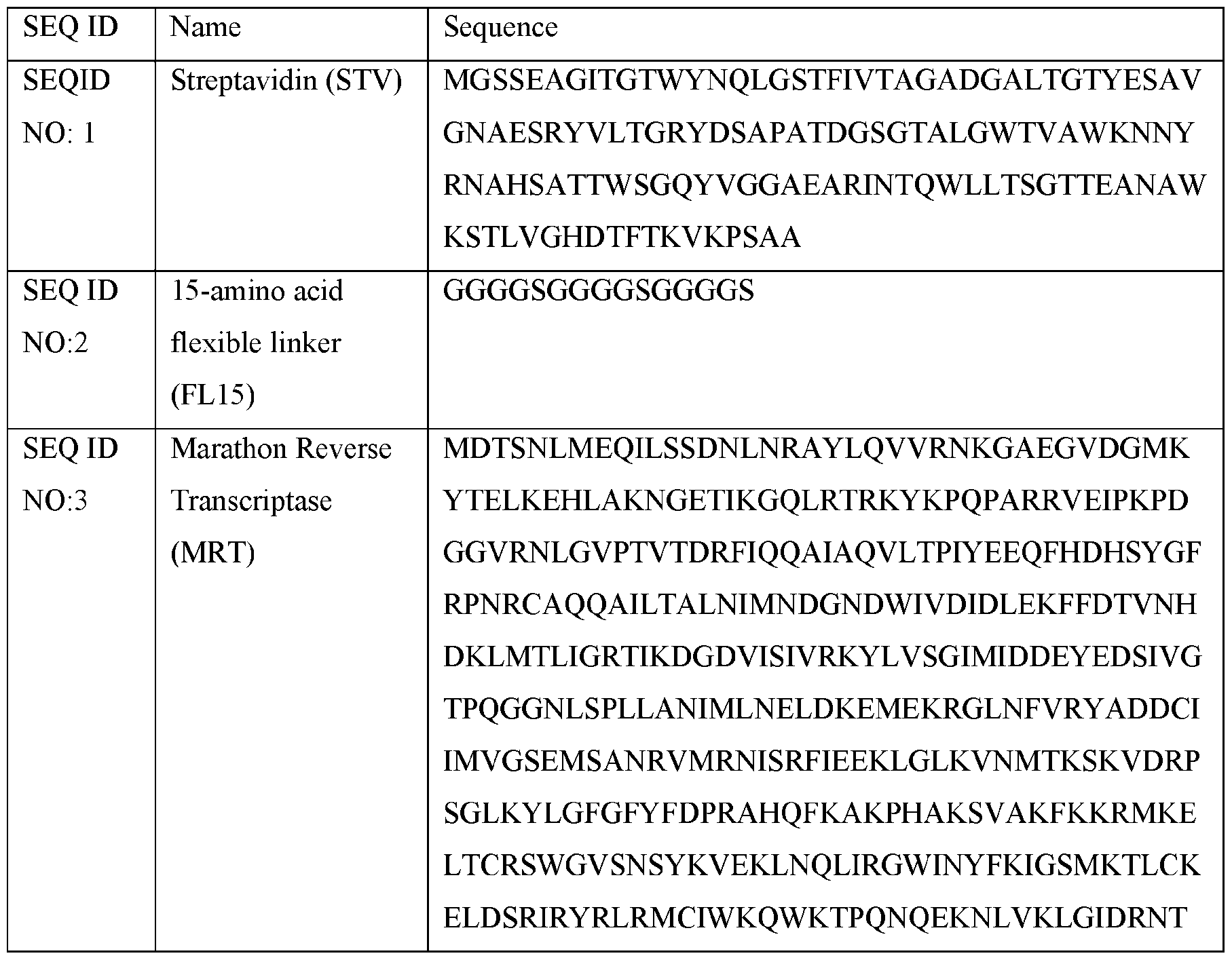
- a representative fusion protein is shown comprising streptavidin, STV (SEQ ID NO: 1) and Marathon Reverse Transcriptase, MRT, (SEQ ID NO:3 linked by a 15-amino acid flexible linker, FL15, (SEQ ID NO:2) to produce STV-FL15-MRT (SEQ ID NO:4) which was assembled into a tetramer comprising one molecule of STV-FL15-MRT and three molecules of STV, thus containing a reduced number of MRT domains.
- FIG. 1A depicts a representative illustration of the co-expression of STV protein (SEQ ID NO: 1) and STV-FL15-MRT fusion protein (SEQ ID NO:4).
- STV-FL15-MRT fusion protein (SEQ ID NO:4) comprises STV (SEQ ID NO:1) connected to MRT (SEQ ID N0:3) via a flexible linker of 15-amino acids (SEQ ID NO:2).
- Figure IB depicts a representative illustration of STV-FL15-MRT fusion protein (SEQ ID NO:4) tetramer containing reduced number of MRT domains.
- the expression level of STV (SEQ ID NO:1) and STV-FL15-MRT fusion protein (SEQ ID NO:4) can be individually adjusted.
- Figure 2 depicts the amino acid sequence of streptavidin-MarathonRT fusion protein containing a 15-amino acid linker (SEQ ID NO: 4). Shown is a representative amino acid sequence of STV connected to MRT via FL15 (SEQ ID NO:4). Underlined sequence: core sequence of STV 14-138 (STV) with enhanced solubility and full biotin binding activity (Sano et al., 1995, JBC, 270: 28204-28209) (SEQ ID NO: 1). Italicized sequence: FL15 (SEQ ID NO:2). Non-underlined, non-italicized sequence: wild-type MRT (SEQ ID NO:3).
- Figure 3 depicts long cDNA synthesis using streptavidin tetramer containing two (estimated) MarathonRT domains. Shown is representative data demonstrating cDNA synthesis from two RNA templates (left: HOTAIR; right: RepA D3) using wild type MRT (SEQ ID NO:3) or STV-FL15-MRT (SEQ ID NO:4) tetramer containing two (estimated) MRT domains.
- Figure 4 depicts tetrameric streptavidin-MarathonRT fusion proteins with a single or multiple MarathonRT molecules with a flexible 15-amino acid linker. Shown are representative illustrations of linear peptide and 3D STV- FL15-MRT fusion protein (SEQ ID NO:4). Left: linear peptide; middle tetrameric STV-FL15-MRT; right: a streptavidin tetramer with a STV-FL15-MRT monomer.
- Figure 5 depicts primer extension assays using STV-MRT fusion protein (with a 15-amino acid linker) containing four or one (estimated) MarathonRT domains. Shown are representative primer extension assay data demonstrating that the long, flexible linker, FL 15, (SEQ ID NO:2) between STV (SEQ ID NO: 1) and MRT (SEQ ID NO:3) greatly improves the enzyme activity of the fusion protein (SEQ ID NO:4); reducing the number of MRT domains further improves the enzyme activity.
- MRT (lane 1) (SEQ ID NO:3); STV-FL15-MRT fusion protein (SEQ ID NO:4) containing four (estimated) MRT domains (lane 2); STV-FL15-MRT fusion protein (SEQ ID NO:4) containing one (estimated) MRT domains (lane 3).
- Figure 6 depicts the amino acid sequence of streptavidin-MarathonRT fusion protein without a linker or with a 3 -amino acid linker. Shown are representative amino acid sequences of STV-MRT fusion protein without a linker (SEQ ID NO:6), top, or with a 3-amino acid linker, FL3, (SEQ ID NO:5), bottom STV-FL3-MRT (SEQ ID NO:7). Underlined sequences: core sequence of STV 14-138 with enhanced solubility and full biotin binding activity (Sano et al., 1995, JBC, 270: 28204-28209) (SEQ ID NO:1). Italicized sequence: 3- amino acid flexible linker, FL3, between (SEQ ID NO:5). Non-underlined, non-italicized sequence: wild-type MRT (SEQ ID NO:3).
- Figure 7 depicts tetrameric streptavidin-MarathonRT fusion protein without or with a 3-amino acid linker, Shown are representative illustrations (linear peptide and 3D protein tetramer) of tetrameric STV-MRT fusion protein without (SEQ ID NO:6) or with (SEQ ID NO:7) a 3-amino acid linker (SEQ ID NO:5).
- Figure 8 depicts representative primer extension assay data demonstrating that the tetrameric STV-MRT fusion protein without a linker (SEQ ID NO:6) (lane 2) or STV-MRT with a short linker (lane 3) (SEQ ID NO:7) is nearly inactive compared to WT-MRT (lane 1) (SEQ ID NO:3).
- the present disclosure is based, in part, on the development of a bifunctional fusion construct comprising an affinity domain linked to an enzyme domain, for example, a polymerase domain.
- the present disclosure features a fusion molecule comprising an enzyme fused to a monomer of avidin or streptavidin, which when expressed and combined with additional monomeric avidin or streptavidin in the correct proportions, yields an active avidin or streptavidin tetramer, in which the fused enzyme maintains at least one enzymatic activity, such as processivity, high speed cDNA synthesis, RNA unwinding, or dNTP hydrolysis.
- the present disclosure provides fusion constructs comprising an affinity domain fused to least one enzymatic domain.
- the affinity domain is capable of binding a target, e g., a substrate.
- the affinity domain comprises a protein or a protein fragment, e.g., avidin or streptavidin.
- the affinity domain comprises avidin or streptavidin tetramers comprising at least one avidin or streptavidin fusion construct comprising an avidin or streptavidin monomer fused to least one enzymatic domain.
- the monomeric avidin or streptavidin domain of the fusion construct retains the ability to bind to additional avidin or streptavidin monomers to form a tetramer.
- the streptavidin fusion construct comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO: 1.
- the avidin or streptavidin fusion construct comprises an enzymatic or catalytic domain, wherein the enzymatic or catalytic domain retains at least one wild type enzymatic or catalytic activity when the fused avidin or streptavidin domain is bound to additional avidin or streptavidin monomers to form a tetramer.
- Exemplary enzymes that can be fused to the monomeric avidin or streptavidin in the fusion construct include, but are not limited to, transcriptases, polymerases, or helicase motor enzymes, receptor proteins, ligases, kinases, phosphatases, exonucleases and non-specific endonucleases, restriction endonucleases, DNA/RNA methyl transferases, recombinases, terminal transferases, nucleic acid binding proteins, proteases, ribosomal proteins, aminoacyl-tRNA synthetases, glycosyltransferase, fatty acid synthases and enzymes for food, biofuel and pharmaceutical industries, such as palatase, cellulase and lipase.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r. maturase also known as Marathon Reverse Transcriptase (MRT), or a variant thereof, wherein the E.r. maturase or variant thereof comprises an amino acid sequence at least 90% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NOTO, SEQ ID NOT 1, SEQ ID NO: 12 or SEQ ID NOT 3, or a functional fragment thereof.
- E.r. maturase also known as Marathon Reverse Transcriptase (MRT)
- MRT Marathon Reverse Transcriptase
- the monomeric avidin or streptavidin is linked to the N- terminus of the enzymatic domain.
- the monomeric streptavidin is linked to the N-terminus of the enzymatic domain via a linker sequence.
- the linker is a flexible linker.
- the linker comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO:2.
- the monomeric avidin or streptavidin is linked to the C- terminus of the enzymatic domain. In one embodiment, the monomeric streptavidin is linked to the C-terminus of the enzymatic domain via a linker sequence. In some embodiments, the linker is a flexible linker.
- the disclosure provides compositions comprising at least one avidin or streptavidin fusion construct.
- the disclosure provides a avidin or streptavidin tetramer comprising at least one avidin or streptavidin fusion construct.
- the avidin or streptavidin tetramer comprises at least two avidin or streptavidin fusion constructs.
- the avidin or streptavidin tetramer comprises at least three avidin or streptavidin fusion constructs.
- the disclosure includes methods of use of the avidin or streptavidin fusion constructs or avidin or streptavidin tetramers comprising the avidin or streptavidin fusion construct of the disclosure.
- the avidin or streptavidin fusion constructs or avidin or streptavidin tetramers of the disclosure are attached to a solid support.
- the solid support is coated with biotin.
- the disclosure includes methods of using the avidin or streptavidin fusion constructs or avidin or streptavidin tetramers or a solid support or device comprising the avidin or streptavidin fusion constructs or avidin or streptavidin tetramers in molecular applications.
- the molecular applications nucleic acid sequencing applications, reverse transcription applications, nucleic acid synthesis applications, or in any molecular application that requires the catalytic activity of the enzymatic or catalytic domain of a avidin or streptavidin fusion construct of the present disclosure.
- autologous refers to a biological material derived from the same individual into whom the material will later be re-introduced.
- allogeneic refers to a biological material derived from a genetically different individual of the same species as the individual into whom the material will be introduced.
- cells and “population of cells” are used interchangeably and generally refer to a plurality of cells, i.e., more than one cell.
- the population may be a pure population comprising one cell type. Alternatively, the population may comprise more than one cell type. In the present disclosure, there is no limit on the number of cell types that a cell population may comprise.
- Encoding refers to the inherent property of specific sequences of nucleotides in a polynucleotide, such as a gene, a DNA, or an RNA, to serve as templates for synthesis of other polymers and macromolecules in biological processes having either a defined sequence of nucleotides (i.e., rRNA, tRNA and mRNA) or a defined sequence of amino acids and the biological properties resulting therefrom.
- a gene encodes a protein if transcription and translation of mRNA corresponding to that gene produces the protein in a cell or other biological system.
- Both the coding strand the nucleotide sequence of which is identical to the mRNA sequence and is usually provided in sequence listings, and the non-coding strand, used as the template for transcription of a gene or cDNA, can be referred to as encoding the protein or other product of that gene or cDNA.
- “Expression vector” refers to a vector comprising a recombinant polynucleotide comprising expression control sequences operatively linked to a nucleotide sequence to be expressed.
- An expression vector comprises sufficient cis-acting elements for expression; other elements for expression can be supplied by the host cell or in an in vitro expression system.
- Expression vectors include all those known in the art, such as cosmids, plasmids (e.g., naked or contained in liposomes) and viruses (e g., lentiviruses, retroviruses, adenoviruses, and adeno- associated viruses) that incorporate the recombinant polynucleotide.
- “Homologous” refers to the sequence similarity or sequence identity between two polypeptides or between two nucleic acid molecules. When a position in both of the two compared sequences is occupied by the same base or amino acid monomer subunit, e.g., if a position in each of two DNA molecules is occupied by adenine, then the molecules are homologous at that position.
- the percent of homology between two sequences is a function of the number of matching or homologous positions shared by the two sequences divided by the number of positions compared X 100. For example, if 6 of 10 of the positions in two sequences are matched or homologous then the two sequences are 60% homologous.
- the DNA sequences ATTGCC and TATGGC share 50% homology. Generally, a comparison is made when two sequences are aligned to give maximum homology.
- isolated means altered or removed from the natural state.
- a nucleic acid or a peptide naturally present in a living organism is not “isolated,” but the same nucleic acid or peptide partially or completely separated from the coexisting materials of its natural state is “isolated.”
- An isolated nucleic acid or protein can exist in substantially purified form, or can exist in a non-native environment such as, for example, a host cell.
- A refers to adenosine
- C refers to cytosine
- G refers to guanosine
- T refers to thymidine
- U refers to uridine.
- nucleotide sequence encoding an amino acid sequence includes all nucleotide sequences that are degenerate versions of each other and that encode the same amino acid sequence.
- the phrase nucleotide sequence that encodes a protein or an RNA may also include introns to the extent that the nucleotide sequence encoding the protein may in some versions contain an intron(s).
- nucleotide as used herein is defined as a chain of nucleotides.
- nucleic acids are polymers of nucleotides.
- nucleic acids and polynucleotides as used herein are interchangeable.
- nucleic acids are polynucleotides, which can be hydrolyzed into the monomeric “nucleotides.” The monomeric nucleotides can be hydrolyzed into nucleosides.
- polynucleotides include, but are not limited to, all nucleic acid sequences which are obtained by any means available in the art, including, without limitation, recombinant means, i.e., the cloning of nucleic acid sequences from a recombinant library or a cell genome, using ordinary cloning technology and PCR, and the like, and by synthetic means.
- recombinant means i.e., the cloning of nucleic acid sequences from a recombinant library or a cell genome, using ordinary cloning technology and PCR, and the like, and by synthetic means.
- peptide As used herein, the terms “peptide,” “polypeptide,” and “protein” are used interchangeably, and refer to a compound comprised of amino acid residues covalently linked by peptide bonds.
- a protein or peptide must contain at least two amino acids, and no limitation is placed on the maximum number of amino acids that can comprise the sequence of a protein or peptide.
- Polypeptides include any peptide or protein comprising two or more amino acids joined to each other by peptide bonds.
- the term refers to both short chains, which also commonly are referred to in the art as peptides, oligopeptides and oligomers, for example, and to longer chains, which generally are referred to in the art as proteins, of which there are many types.
- Polypeptides include, for example, biologically active fragments, substantially homologous polypeptides, oligopeptides, homodimers, heterodimers, variants of polypeptides, modified polypeptides, derivatives, analogs, fusion proteins, among others.
- the polypeptides include natural peptides, recombinant peptides, synthetic peptides, or a combination thereof.
- promoter as used herein is defined as a DNA sequence recognized by the synthetic machinery of the cell, or introduced synthetic machinery, required to initiate the specific transcription of a polynucleotide sequence.
- promoter/regulatory sequence means a nucleic acid sequence which is required for expression of a gene product operably linked to the promoter/regulatory sequence.
- this sequence may be the core promoter sequence and in other instances, this sequence may also include an enhancer sequence and other regulatory elements which are required for expression of the gene product.
- the promoter/regulatory sequence may, for example, be one which expresses the gene product in a conditional manner.
- a “constitutive” promoter is a nucleotide sequence which, when operably linked with a polynucleotide which encodes or specifies a gene product, causes the gene product to be produced in a cell under most or all physiological conditions of the cell.
- An “inducible” promoter is a nucleotide sequence which, when operably linked with a polynucleotide which encodes or specifies a gene product, causes the gene product to be produced in a cell substantially only when an inducer which corresponds to the promoter is present in the cell.
- a “vector” is a composition of matter which comprises an isolated nucleic acid and which can be used to deliver the isolated nucleic acid to the interior of a cell.
- vectors are known in the art including, but not limited to, linear polynucleotides, polynucleotides associated with ionic or amphiphilic compounds, plasmids, and viruses.
- the term “vector” includes an autonomously replicating plasmid or a virus.
- the term should also be construed to include non-plasmid and non-viral compounds which facilitate transfer of nucleic acid into cells, such as, for example, polylysine compounds, liposomes, and the like.
- viral vectors include, but are not limited to, adenoviral vectors, adeno-associated virus vectors, retroviral vectors, and the like.
- Wild type activity refers to the activity (e.g., catalytic, structural, etc.) displayed by a biological molecule as it occurs in nature.
- wild type activity refers to any documented, or yet undocumented, biological activity of a molecule as determined by techniques or assays commonly used in the field to determine the function of a molecule (e.g., catalytic, structural, etc.).
- Ranges throughout this disclosure, various aspects of the disclosure can be presented in a range format. It should be understood that the description in range format is merely for convenience and brevity and should not be construed as an inflexible limitation on the scope of the disclosure. Accordingly, the description of a range should be considered to have specifically disclosed all the possible subranges as well as individual numerical values within that range. For example, description of a range such as from 1 to 6 should be considered to have specifically disclosed subranges such as from 1 to 3, from 1 to 4, from 1 to 5, from 2 to 4, from 2 to 6, from 3 to 6 etc., as well as individual numbers within that range, for example, 1, 2, 2.7, 3, 4, 5, 5.3, and 6. This applies regardless of the breadth of the range.
- the present disclosure provides streptavidin fusion constructs comprising a streptavidin monomer fused to least one enzymatic domain as well as streptavidin tetramers comprising at least one streptavidin fusion construct comprising a streptavidin monomer fused to least one enzymatic domain.
- the streptavidin fusion construct comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO: 1.
- the monomeric streptavidin domain of the streptavidin fusion construct retains the ability to bind to additional streptavidin monomers to form a streptavidin tetramer.
- the streptavidin fusion construct comprises an enzymatic or catalytic domain, wherein the enzymatic or catalytic domain retains at least one wild type enzymatic or catalytic activity when the fused streptavidin domain is bound to additional streptavidin monomers to form a streptavidin tetramer.
- Exemplary enzymes that can be fused to the monomeric streptavidin in the streptavidin fusion construct include, but are not limited to, transcriptases, polymerases, helicase motor enzymes, receptor proteins, ligases, kinases, phosphatases, exonucleases and non-specific endonucleases, restriction endonucleases, DNA/RNA methyl transferases, recombinases, terminal transferases, nucleic acid binding proteins, proteases, ribosomal proteins, aminoacyl-tRNA synthetases, glycosyltransferase, fatty acid synthases and enzymes for food, biofuel and pharmaceutical industries, such as palatase, cellulase and lipase.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterinm rectale (E.r.) maturase also known as Marathon Reverse Transcriptase (MRT), or a variant thereof, wherein the E.r. maturase or variant thereof comprises an amino acid sequence at least 90% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO: 9, SEQ ID NOTO, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- E.r. Eubacterinm rectale
- MRT Marathon Reverse Transcriptase
- the monomeric streptavidin is linked to the N-terminus of the enzymatic domain. In one embodiment, the monomeric streptavidin is linked to the N-terminus of the enzymatic domain via a linker sequence. In one embodiment, the linker comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO:2.
- the disclosure provides compositions comprising at least one streptavidin fusion construct. In some embodiments, the disclosure provides a streptavidin tetramer comprising at least one streptavidin fusion construct. In one embodiment, the streptavidin tetramer comprises at least two streptavidin fusion constructs. In one embodiment, the streptavidin tetramer comprises at least three streptavidin fusion constructs.
- the disclosure includes methods of use of the streptavidin fusion constructs or streptavidin tetramers comprising the streptavidin fusion construct of the disclosure.
- the streptavidin fusion constructs or streptavidin tetramers of the disclosure are attached to a solid support.
- the solid support is coated with biotin.
- the disclosure includes methods of using the streptavidin fusion constructs or streptavidin tetramers or a solid support or device comprising the streptavidin fusion constructs or streptavidin tetramers in molecular applications.
- the molecular applications nucleic acid sequencing applications, reverse transcription applications, nucleic acid synthesis applications, or in any molecular application that requires the catalytic activity of the enzymatic or catalytic domain of a streptavidin fusion construct of the present disclosure.
- compositions comprising at least one avidin or streptavidin domain fusion construct comprising a avidin or streptavidin monomer linked to an enzymatic domain in a manner such that the monomer domain retains the ability to form a tetramer with additional avidin or streptavidin monomers while the enzymatic domain retains at least one enzymatic function.
- compositions comprising at least one avidin or streptavidin fusion construct, wherein the fusion construct is bound to or in a complex with at least one additional avidin or streptavidin fusion construct or avidin or streptavidin monomer to form a tetramer comprising at least one avidin or streptavidin fusion construct.
- the avidin or streptavidin domain fusion construct comprises an enzymatic or catalytic domain, wherein the enzymatic or catalytic domain retains at least one wild type enzymatic or catalytic activity when the fused streptavidin domain is bound to additional streptavidin monomers to form a streptavidin tetramer.
- Exemplary proteins or enzymes that can be fused to the monomeric streptavidin in the avidin or streptavidin domain fusion construct include, but are not limited to, transcriptases, polymerases, helicase motor enzymes, receptor proteins, ligases, kinases, phosphatases, exonucleases and non-specific endonucleases, restriction endonucleases, DNA/RNA methyl transferases, recombinases, terminal transferases, nucleic acid binding proteins, proteases, ribosomal proteins, aminoacyl- tRNA synthetases, glycosyltransferase, fatty acid synthases and enzymes for food, biofuel and pharmaceutical industries.
- Exemplary enzymes that can be incorporated into the avidin or streptavidin domain fusion construct of the disclosure include, but are not limited to, reverse transcriptases, translocating helicase enzymes, DNA polymerases, and RNA , ligases, kinases, phosphatases, exonucleases and non-specific endonucleases, restriction endonucleases, DNA/RNA methyl transferases, recombinases, terminal transferases, nucleic acid binding proteins, proteases, ribosomal proteins, aminoacyl-tRNA synthetases, glycosyltransferase, fatty acid synthases and enzymes for food, biofuel and pharmaceutical industries, such as palatase, cellulase and lipase.
- the enzymatic domain comprises a polymerase, e.g., a DNA or RNA polymerase.
- the polymerase is a DNA polymerase, e.g., a reverse transcriptase.
- the polymerase is a reverse transcriptase.
- the reverse transcriptase is derived from a virus.
- the reverse transcriptase may be derived from a retrovirus.
- the reverse transcriptase is an Avian Myeloblastosis Virus (AMV) reverse transcriptase.
- AMV Avian Myeloblastosis Virus
- the reverse transcriptase is a Human Immunodeficiency Virus reverse transcriptase.
- the reverse transcriptase is a Rous Sarcoma Virus reverse transcriptase. In some embodiments, the reverse transcriptase is a Moloney Murine Leukemia Virus (MMLV) reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is Maxima H Minus reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is SuperScript II reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is SuperScript III reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is SuperScript IV reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is PrimeScript reverse transcriptase.
- MMLV Moloney Murine Leukemia Virus
- the MMLV reverse transcriptase is GoScript reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is ProtoScript II reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is SMARTScribe reverse transcriptase. In one embodiment, the MMLV reverse transcriptase comprises SEQ ID NO: 22 or SEQ ID NO: 23, or a fragment or variant thereof. In one embodiment, the AMV reverse transcriptase comprises SEQ ID NO: 26 or SEQ ID NO: 27, or a fragment or variant thereof.
- the reverse transcriptase is derived from a mobile genetic element.
- the reverse transcriptase may be derived from a telomerase, e.g., a mammalian telomerase.
- the reverse transcriptase is derived from a human telomerase, e.g., TERT.
- the reverse transcriptase comprises SEQ ID NO: 28 or SEQ ID NO: 29, or a fragment or variant thereof.
- the reverse transcriptase may be derived from a non-long terminal repeat (non-LTR) retrotransposon or a group II intron.
- non-LTR non-long terminal repeat
- the reverse transcriptase is a non-LTR retrotransposon reverse transcriptase.
- the non-LTR retrotransposon reverse transcriptase is a Bombyx mori R2 RNA element reverse transcriptase.
- the Bombyx mori R2 RNA element reverse transcriptase comprises SEQ ID NO: 24 or SEQ ID NO: 25, or a fragment or variant thereof.
- the non-LTR retrotransposon reverse transcriptase is a human LI element reverse transcriptase.
- the reverse transcriptase is a group II intron reverse transcriptase, e.g., a maturase reverse transcriptase.
- the group II intron reverse transcriptase is a maturase. In one embodiment, the group II intron reverse transcriptase is a maturase encoded by Eubacterium rectale. In one embodiment, the group II intron reverse transcriptase is a maturase encoded by Roseburia intestinalis. In one embodiment, the Roseburia intestinalis reverse transcriptase comprises SEQ ID NO: 20 or SEQ ID NO: 21, or a fragment or variant thereof. In one embodiment, the group II intron reverse transcriptase is MarathonRT reverse transcriptase. In one embodiment, the group II intron reverse transcriptase is UltraMarathonRT. In one embodiment, the group II intron reverse transcriptase is Induro reverse transcriptase. In one embodiment, the group II intron reverse transcriptase is a TGIRT reverse transcriptase.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, also known as Marathon Reverse Transcriptase (MRT), or a fragment or variant thereof.
- E.r. maturase comprises an amino acid sequence identical to an amino acid sequence selected from the group consisting of SEQ ID NOs: 3, and 8-12.
- the fragment or variant of the E.r. maturase comprises a fragment or variant of an amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO: 9, SEQ ID NOTO, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
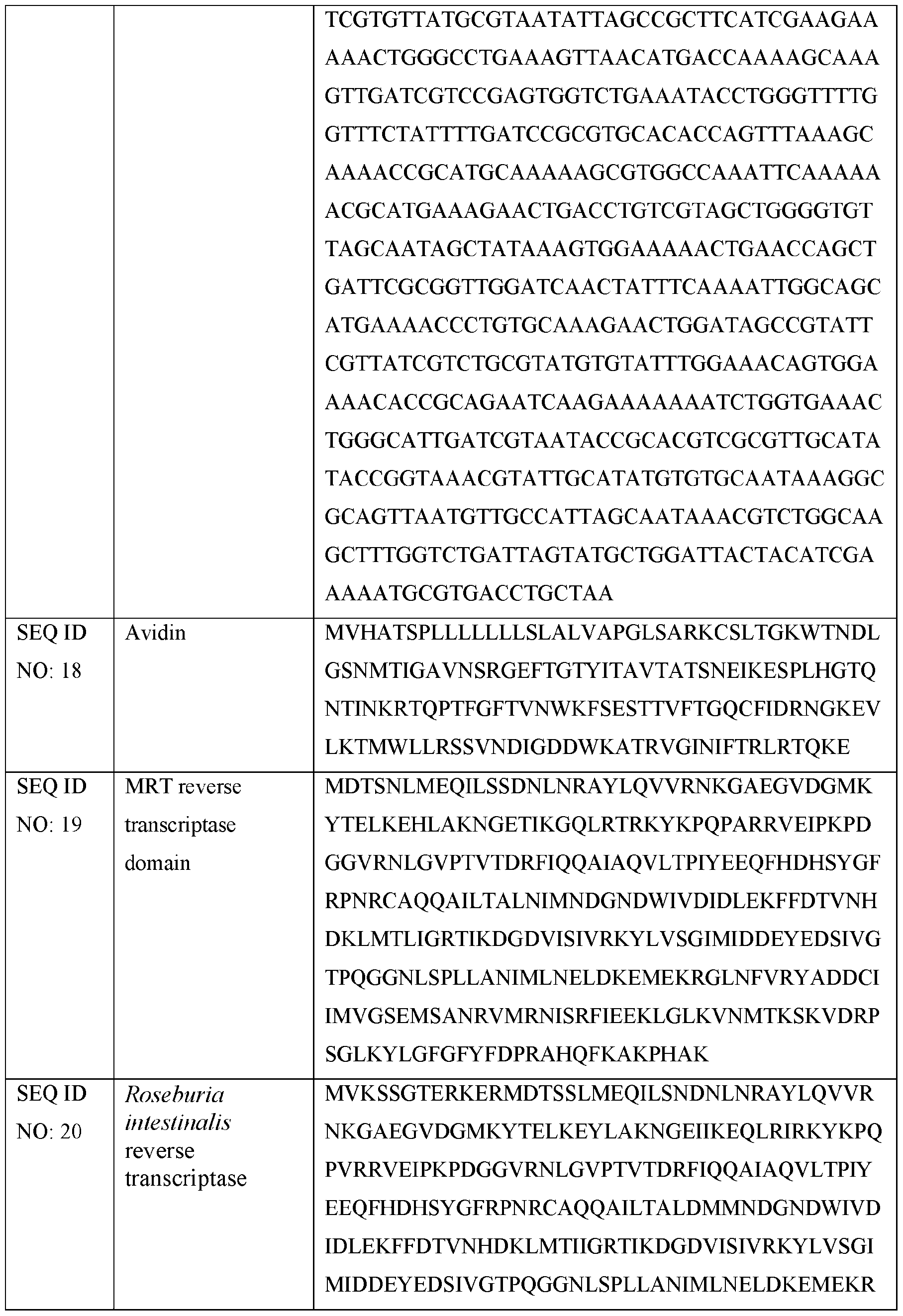
- the fragment or variant of the E.r. maturase comprises a fragment or variant of an amino acid sequence of SEQ ID NO: 19. In some embodiments, the fragment or variant of the E.r. maturase retains at least one wild type activity of the parental E.r. maturase. In some embodiments, the fragment or variant of the E.r. maturase has an enhanced activity as compared to the parental E.r. maturase. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale E.r.)' maturase, or a variant thereof, wherein the E.r.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectal e (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 70% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- E.r. Eubacterium rectal e
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.)' maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 75% identical to an amino acid sequence of SEQ ID N0:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 75% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 80% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 80% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 85% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 90% identical to an amino acid sequence of SEQ ID NO : 3 , SEQ ID NO : 8, SEQ ID NO : 9, SEQ ID NO : 10, SEQ ID NO : 11 , SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- E.r. Eubacterium rectale
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 90% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- Eubacterium rectale E.r.
- the E.r. maturase or variant thereof comprises an amino acid sequence at least 90% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r. maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 95% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 95% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 96% identical to an amino acid sequence of SEQ ID NO : 3 , SEQ ID NO : 8, SEQ ID NO : 9, SEQ ID NO : 10, SEQ ID NO : 11 , SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 96% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 97% identical to an amino acid sequence of SEQ ID NO : 3 , SEQ ID NO : 8, SEQ ID NO : 9, SEQ ID NO : 10, SEQ ID NO : 11 , SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 97% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.)' maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 98% identical to an amino acid sequence of SEQ ID NO : 3 , SEQ ID NO : 8, SEQ ID NO : 9, SEQ ID NO : 10, SEQ ID NO : 11 , SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 98% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 99% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 99% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 99.5% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 99.5% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
- the enzymatic domain comprises a functional fragment comprising at least 400 amino acid residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQIDNO:10, SEQIDNO:11, SEQIDNO:12or SEQIDNO:13.
- the enzymatic domain comprises a functional fragment comprising at least 350 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQIDNO:10, SEQIDNO:11, SEQIDNO:12or SEQIDNO:13.
- the enzymatic domain comprises a functional fragment comprising at least 300 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
- the enzymatic domain comprises a functional fragment comprising at least 250 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ IDNO:10, SEQIDNO:11, SEQIDNO:12or SEQIDNO:13.
- the enzymatic domain comprises a functional fragment comprising at least 200 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO:10, SEQ IDNO:11, SEQIDNO:12or SEQ ID NO:13.
- the enzymatic domain comprises a functional fragment comprising at least 150 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ IDNO:10, SEQ IDNO:11, SEQIDNO:12or SEQ ID NO:13.
- the enzymatic domain comprises a functional fragment comprising at least 100 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ IDNO:10, SEQIDNO:11, SEQIDNO:12or SEQIDNO:13.
- the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 400 amino acids residues ofSEQIDNO:3, SEQIDNO:8, SEQIDNO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
- the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 350 amino acids residues ofSEQIDNO:3, SEQIDNO:8, SEQIDNO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13. [0103] In one embodiment the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 300 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
- the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 250 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
- the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 200 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
- the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 150 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
- the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 100 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
- the reverse transcriptase has a sequence with at least 60% to 99.9% identity to a sequence listed in Example 3, e.g., at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 99.9% or greater identity to a sequence listed in Example 3.
- the enzymatic domain comprises a reverse transcriptase.
- the reverse transcriptase is derived from Eubacterium rectale (E.r.) maturase (also known as Marathon Reverse Transcriptase (MRT)).
- E.r. maturase is modified relative to unmodified E.r. maturase.
- the variant comprises one or more point mutations, insertion mutations, or deletion mutations, relative to wildtype E.r. maturase.
- the variant comprises a fusion protein comprising E.r. maturase, E.r. maturase mutant, or E.r. maturase domain.
- the full-length E.r. maturase comprises a “secondary” RNA binding site and DNA binding domain that can influence stability, specificity, and efficiency of reverse transcription of an RNA template.
- the reverse transcriptase comprises an E.r. maturase variant where one or more secondary RNA binding sites on the surface of the protein are mutated to reduce nonspecific binding of the reverse transcription protein to the RNA template, thereby promoting binding at the polymerase cleft and facilitating enzyme turnover.
- a variant of E.r. maturase comprises at least one point mutation selected from the group R58X, K59X, K61X, K163X, K216X, R217X, K338X, K342X, and R353X wherein X denotes any amino acid.
- a variant of E.r. maturase comprises at least one point mutation selected from the group R58A, K59A, K61 A, K163A, K216A, R217A, K338A, K342A, and R353A.
- the reverse transcriptase comprises an E.r. maturase variant (referred to herein as E.r. maturase mutl; and denoted as SEQ ID NO:8) comprising the point mutations of: R58A, K59A, K61A, and K163A, relative to wildtype E.r. maturase.
- E.r. maturase mutl referred to herein as E.r. maturase mutl
- SEQ ID NO:8 comprising the point mutations of: R58A, K59A, K61A, and K163A, relative to wildtype E.r. maturase.
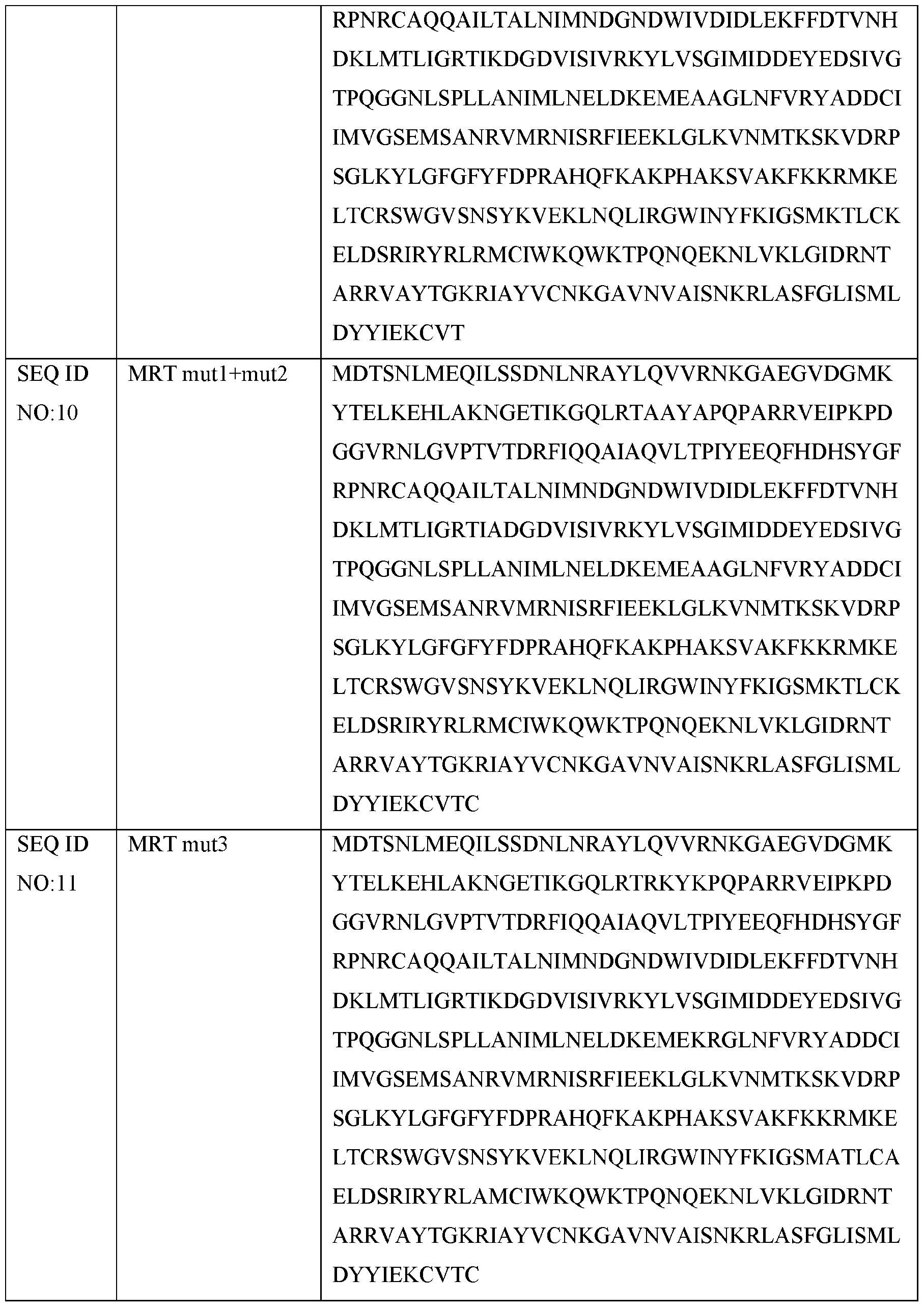
- the reverse transcriptase comprises an E.r. maturase variant (referred to herein as E.r. maturase mutl+mut2; and denoted as SEQ ID NOTO) comprising the point mutations of: R58A, K59A, K61A, K163A, K216A, and R217A, relative to wildtype E.r. maturase.
- E.r. maturase mutl+mut2 referred to herein as E.r. maturase mutl+mut2; and denoted as SEQ ID NOTO
- the reverse transcriptase comprises an E.r. maturase variant (referred to herein as E.r. maturase mut3; and denoted as SEQ ID NO: 11) comprising the point mutations of: K338A, K342A, and R353A relative to wildtype E.r. maturase.
- E.r. maturase mut3 referred to herein as E.r. maturase mut3; and denoted as SEQ ID NO: 11
- SEQ ID NO: 11 comprising the point mutations of: K338A, K342A, and R353A relative to wildtype E.r. maturase.
- the reverse transcriptase comprises an E.r. maturase variant comprising one or more mutations in the C-terminal DNA binding domain of E.r. maturase.
- a variant of E.r. maturase comprises at least one point mutation selected from the group K388X, R389X, K396X, K406X, R407X, and K423X, wherein X denotes any amino acid.
- a variant of E.r. maturase comprises at least one point mutation selected from the group K388A, R389A, K396A, K406A, R407A, and K423A.
- maturase comprises at least one point mutation selected from the group K388S, R389S, K396S, K406S, R407S, and K423S.
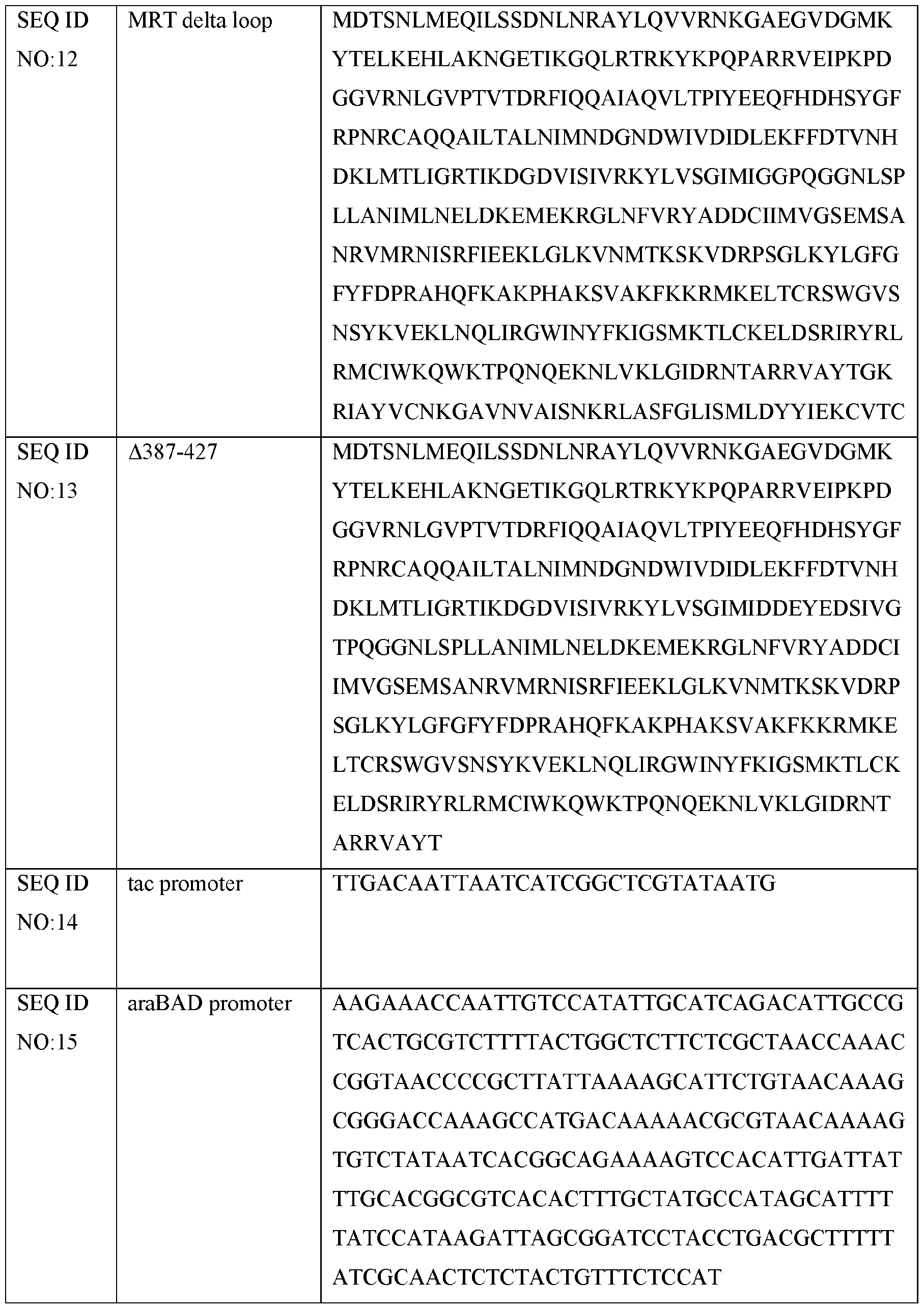
- the C-terminal sequence residues 387-427 are deleted relative to wildtype E.r. maturase.
- the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, comprising one or more mutations in the a-loop (AA180-196) of E.r maturase.
- the E.r. maturase variant comprises one or more mutations in the N-terminal region of the a-loop.
- the mutation is at least one selected from the group: M180X, I181X, D182X, D183X, E184X, Y185X, E186X, D187X, S188X, I189X, V190X, wherein X denotes any amino acid.
- the at least one point mutation (X) is selected from the group: alanine, polar amino acid (e.g., Gin), electrostatic amino acid (e.g. Glu), and a combination thereof.
- the a-loop is engineered to be more flexible by substituting positions in the N- terminal region with one or more glycines.
- the a-loop is engineered to be more stiff by substituting positions in the N-terminal region with one or more alanines.
- the mutation is a deletion of at least one residue of the a-loop.
- the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant in which residues 182-192 are substituted with two glycine residues (Aloop; SEQ ID NO: 12).
- E.r. maturase can perform reverse transcription at lower temperatures relative to other reverse transcriptases, and the engineering of a more thermostable E.r. maturase would enable amplification of RNA templates in a single reaction (i.e., without using DNA ⁇ DNA amplification reactions).
- thermophilic protein structure and function suggests that they tend to have larger numbers of side-chain hydrogen bonds and salt-bridges within rigid sections of the tertiary structure. Therefore, in one embodiment, the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r.
- the variant comprises at least one point mutation selected from the group LI IE (which can form a salt bridge with R56), L21E (which can form a salt bridge with K41), and S13E (which can form a salt bridge with K52).
- the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, engineered to comprise a proofreading (e g., 3'- 5' exonuclease) domain to enhance fidelity.
- the proofreading domain comprises an exonuclease domain.
- the proofreading domain is appended to the C-terminus of the E.r. maturase variant.
- the proofreading domain is appended to the C-terminus of the E.r. maturase variant through a linker molecule or sequence (see, for example, Ellefson, JW et al., 2016, Science, 352(6293): 1590-3).
- the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, wherein at least one fragment or domain of E.r. maturase is replaced with a fragment or domain from a maturase reverse transcriptase from a species other than Eubacterium rectale.
- the RT domain (finger and palm) of E.r. maturase reverse transcriptase is replaced with the RT domain from a thermophilic maturase reverse transcriptase to enhance thermostability.
- the a-loop of E.r. maturase is replaced by a longer a-loop from another maturase reverse transcriptase to enhance processivity.
- one or more amino acids are substituted with hydrophobic amino acids or charged amino acids in order to improve thermostability.
- the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, wherein one or more residues are substituted with one or more residues derived from a maturase enzyme from an organism other than Eubacterium rectale.
- the E.r. maturase variant can comprise one or more point mutations based on conserved residues in thermophilic maturases.
- the variant comprises at least one mutation selected from the group: A29X, V82X, E104X, I129X, I137X, T161X, I168X, I170X, VI 7 IX, and M337X, where X denotes any amino acid.
- the mutation is at least one selected from the group: A29X, V82X, E104X, I129X, I137X, T161X, I168X, I170X, V171X, and M337X, where X denotes any amino acid.
- the variant comprises at least one mutation selected from the group: A29S, V82I, E104P, I129Y, I137V, T161R, I168L, I170L, V171I, and M337T.
- the variant comprises a triple point mutation of A29S/V82I/E104P. In certain instances, these mutations improve upon the thermostability of the enzyme.
- the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, comprising one or more mutations in the thumb domain relative to wildtype E.r. maturase.
- the variant comprises at least one point mutation selected from the group consisting of K338X, K342X, and R353X, wherein X denotes any amino acid.
- the variant comprises at least one point mutation selected from the group consisting of K338A, K342A, and R353A.
- one or more mutations are incorporated on the surface of the thumb domain, optimizing its ability to clasp the template.
- the variant comprises at least one point mutation selected from the group consisting of S315X, E319X, and Q323X, wherein X denotes any amino acid.
- the variant comprises at least one point mutation selected from the group consisting of S315K, E319K, and Q323K.
- the enzymatic domain comprises a polypeptide comprising a reverse transcriptase.
- the reverse transcriptase is derived from E.r. maturase.
- the polypeptide comprises E.r. maturase, or a variant thereof.
- Exemplary amino acid sequences of the E.r. maturase-derived reverse transcriptases of the present disclosure include, but are not limited to, SEQ ID NO:3 (E.r. maturase), SEQ ID NO:8 (Er. maturase mutl), SEQ ID NO:9 (E.r. maturase mut2), SEQ ID NO: 10 (E.r.
- the present disclosure is not limited to these sequences. Rather the present disclosure encompasses any reverse transcriptase derived from E.r. maturase or a variant thereof.
- the polypeptide comprises a fragment of E.r. maturase or variant thereof that mimics the ability of E.r. maturase to perform reverse transcription.
- the polypeptide comprises a derivative of the E.r. maturase or variant thereof.
- the polypeptide comprises an amino acid sequence selected from a fragment or derivative of SEQ ID NO:3, a fragment or derivative of SEQ ID NO:8, a fragment or derivative of SEQ ID NO:9, a fragment or derivative of SEQ ID NO: 10, a fragment or derivative of SEQ ID NO: 11, and a fragment or derivative of SEQ ID NO: 12.
- the avidin or streptavidin domain fusion construct of the disclosure comprises one or more mutations in the catalytic active-site to reduce the fidelity of the enzyme, which will enhance its value for RNA structure mapping since structure-specific lesions that are used to probe RNA structure are flagged by misincorporation events. Similarly, mutations that increase the error rate of the enzyme can be used with certain RNA and transcriptome mapping experiments. Therefore, in some embodiments, the polypeptide comprises at least one mutation selected from the group: A225X, R114X, Y224X, I179X, M180X, I181X, E143X, K65X, L201X, wherein X denotes any amino acid.
- mutations at A225 (such as A225V, A225S, A225M or A225V), mutations at R114 (such as R114K, R114A), mutations at Y224 (such as Y224F), mutations at 1179 (such as I179F), mutations at Ml 80 (such as Ml 80V), mutations at 1181 (such as I181W), mutations at E143 (such as E143A or E143K), mutations at K65 (such as K65A), mutations at L201 (such as L201A or L201T), may be used, alone or in combination.
- A225 such as A225V, A225S, A225M or A225V
- mutations at R114 such as R114K, R114A
- mutations at Y224 such as Y224F
- mutations at 1179 such as I179F
- mutations at Ml 80 such as Ml 80V
- mutations at 1181 such as I181W
- mutations at E143 such as E143A or E143K
- the avidin or streptavidin domain fusion construct of the disclosure comprises one or more mutations in the primer/template binding interface to increase the binding affinity to primer/template, which will enhance its capability of capturing low abundance RNAs in single-cell or spatial transcriptomic RNA sequencing. Therefore, in some embodiments, the polypeptide comprises at least one mutation selected from the group: D14X, Q22X, N26X, E30X, D74X, Q91X, Q92X, Q96X, N116X, N197X, E304X, E319X, N322X, N330X, E422X, wherein X denotes any amino acid.
- mutations at D14 (such as D14K or D14R), mutations at Q22 (such as Q22K or Q22R), mutations at N26 (such as N26K or N26R), mutations at E30 (such as E30K or E30R), mutations at D74 (such as D74K or D74R), mutations at Q91 (such as Q91K or Q91R), mutations at Q92 (such as Q92K or Q92R), mutations at Q96 (such as Q96K or Q96R), mutations at N116 (such as N116K or N116R), mutations at N197 (such as N197K or N197R), mutations at E304 (such as E304K or E304R), mutations at E319 (such as E319K or E319R), mutations at N322 (such as N322K or N322R), mutations at N33O (such as N330K or N33OR), mutations at E422 (such as E422K or E422R), may be
- the enzymatic domain of the present disclosure comprises a polypeptide comprising Roseburia intestinalis (R.i.) maturase, or a variant or fragment thereof.
- the Ri. maturase comprises one or more mutations corresponding to one or more mutations described herein.
- Reverse transcriptases of the present disclosure may produce more product (e.g., full length product) at particular temperatures compared to other reverse transcriptases.
- comparisons of full length product synthesis are made at different temperatures (e.g., one temperature being lower, such as between 4° C and 42° C, and one temperature being higher, such as between 42° C and 78° C) while keeping all other reaction conditions similar or the same.
- the amount of full length product produced may be determined using techniques well known in the art, for example, by conducting a reverse transcription reaction at a first temperature (e.g., 37° C, 38° C, 39° C, 40° C, etc.) and determining the amount of full length transcript produced, conducting a second reverse transcription reaction at a temperature higher than the first temperature (e.g., 45° C, 50° C, 52.5° C, 55° C, etc.) and determining the amount of full length product produced, and comparing the amounts produced at the two temperatures.
- a convenient form of comparison is to determine the percentage of the amount of full length product at the first temperature that is produced at the second (i.e., elevated) temperature.
- reaction conditions used for the two reactions may be the same for both reactions. Suitable reaction conditions may be determined by those skilled in the art using routine techniques and examples of such conditions are provided herein.
- the reverse transcriptases of the disclosure may produce at least about 5%, at least 10%, at least 15%, at least 25%, at least 50%, at least 75%, at least 100%, or at least 200% more product or full length product compared to the corresponding control reverse transcriptase under the same reaction conditions and temperature.
- the reverse transcriptases of the disclosure may produce from about 10% to about 200%, from about 25% to about 200%, from about 50% to about 200%, from about 75% to about 200%, or from about 100% to about 200% more product or full length product compared to a control reverse transcriptase under the same reaction conditions and incubation temperature.
- the reverse transcriptases of the disclosure may produce at least 2 times, at least 3 times, at least 4 times, at least 5 times, at least 6 times, at least 7 times, at least 8 times, at least 9 times, at least 10 times, at least 25 times, at least 50 times, at least 75 times, at least 100 times, at least 150 times, at least 200 times, at least 300 times, at least 400 times, at least 500 times, at least 1000 times, at least 5,000 times, or at least 10,000 times more product or full length product compared to a control reverse transcriptase under the same reaction conditions and temperature.
- Reverse transcriptases of the present disclosure may have an increased thermostability at elevated temperatures as compared to corresponding control reverse transcriptases. They may show increased thermostability in the presence or absence an RNA template. In some instances, reverse transcriptases of the disclosure may show an increased thermostability in both the presence and absence of an RNA template. Those skilled in the art will appreciate that reverse transcriptase enzymes are typically more thermostable in the presence of an RNA template. The increase in thermostability may be measured by comparing suitable parameters of the modified or mutated reverse transcriptase of the disclosure to those of a corresponding un-modified or un-mutated reverse transcriptase.
- Suitable parameters to compare include, but are not limited to, the amount of product and/or full length product synthesized by the reverse transcriptases of the disclosure at an elevated temperature compared to the amount or product and/or full length product synthesized by a control reverse transcriptase at the same temperature, and/or the half-life of reverse transcriptase activity at an elevated temperature of a reverse transcriptase of the disclosure at an elevated temperature compared to that of a control reverse transcriptase.
- a reverse transcriptase of the disclosure may have an increase in thermostability at a particular temperature of at least about 1.5 fold (e.g., from about 1.5 fold to about 100 fold, from about 1.5 fold to about 50 fold, from about 1.5 fold to about 25 fold, from about 1.5 fold to about 10 fold) compared, for example, to the control reverse transcriptase.
- a reverse transcriptase of the disclosure may have an increase in thermostability at a particular temperature of at least about 10 fold (e.g., from about 10 fold to about 100 fold, from about 10 fold to about 50 fold, from about 10 fold to about 25 fold, or from about 10 fold to about 15 fold) compared, for example, to the control reverse transcriptase.
- a reverse transcriptase of the disclosure may have an increase in thermostability at a particular temperature of at least about 25 fold (e.g., from about 25 fold to about 100 fold, from about 25 fold to about 75 fold, from about 25 fold to about 50 fold, or from about 25 fold to about 35 fold) compared to the control reverse transcriptase.
- polypeptide of the present disclosure may be made using chemical methods.
- polypeptides can be synthesized by solid phase techniques (Roberge J Y et al (1995) Science 269: 202-204), cleaved from the resin, and purified by preparative high performance liquid chromatography. Automated synthesis may be achieved, for example, using the ABI 431 A Peptide Synthesizer (Perkin Elmer) in accordance with the instructions provided by the manufacturer.
- the polypeptide may be made by recombinant means or by cleavage from a longer polypeptide.
- the polypeptide may be confirmed by amino acid analysis or sequencing.
- the disclosure should also be construed to include any form of a polypeptide having substantial homology to a reverse transcriptase disclosed herein.
- a polypeptide which is “substantially homologous” is about 50% homologous, about 70% homologous, about 80% homologous, about 90% homologous, about 95% homologous, about 96% homologous, about 97% homologous, about 98% homologous, about 99% homologous, or about 99.5% homologous to an amino acid sequence of a reverse transcriptase disclosed herein.
- the enzymatic domain comprises a reverse transcriptase comprising an amino acid sequence that is about 50%, about 70%, about 80%, about 90%, about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 99.5% homologous to E.r. maturase or E.r. maturase variant described herein.
- the enzymatic domain comprises a reverse transcriptase comprising an amino acid sequence that is about 50%, about 70%, about 80%, about 90%, about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 99.5% identical to the amino acid sequence set forth in SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NOTO, SEQ ID NO l l, SEQ ID NOT2, or SEQ ID NOT3.
- the enzymatic domain comprises a reverse transcriptase comprising an amino acid sequence that is about 50%, about 70%, about 80%, about 90%, about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 99.5% identical to the amino acid sequence set forth in SEQ ID NOT, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NOTO, SEQ ID NOT 1, SEQ ID NOT2, or SEQ ID NOT3, wherein the reverse transcriptase comprises one more of the mutations described herein.
- the present disclosure provides a composition comprising an agent that improves RT activity of E.r. maturase or variants thereof.
- the composition comprises an agent that reduces non-specific binding of primers to the positively charged surface of E.r. maturase or variants thereof.
- the agent that reduces non-specific binding of primers to the positively charged surface of E.r. maturase or variants thereof comprises a peptide or protein, including, but not limited to, heparin.
- the enzymatic domain of the present disclosure can comprise any enzymatic domain applicable for molecular technology applications and industrial/pharmaceutical production well known in the field, and is not limited to the enzymatic domains described herein.
- the enzymatic domain of the present disclosure comprises any reverse transcriptase (RT) or RNA-dependent DNA polymerase (e.g., AMV/MAV RT, HIV RT, MuLV RT, Tth, MonsterScriptTM RT, KI enow fragment of C.
- RT reverse transcriptase
- RNA-dependent DNA polymerase e.g., AMV/MAV RT, HIV RT, MuLV RT, Tth, MonsterScriptTM RT, KI enow fragment of C.
- any DNA polymerase or DNA-dependent DNA polymerase e.g., Bst polymerase, Taq polymerase, Tth polymerase, Pfu polymerase, Pow polymerase, Vent polymerase, Pab polymerase, T4 DNA polymerase, Pol I and Klenow fragment, T7 DNA polymerase, terminal deoxynucleotidyl transferase, DNA Pol I, DNA Pol II, etc.
- any RNA polymerase or DNA-dependent RNA polymerase e.g., SP6 RNA polymerase, T7 RNA polymerase, etc.
- any nuclease e.g., restriction endonucleases, DNasel, exonuclease III, Bal31, exonuclease VII, RNase A, ribonuclease H, Phy I, CL3, Cereus, Phy M, RNase Tl, RNase T2, RNase
- RNA ligase e.g., T4 RNA ligase 1, T4 RNA ligase 2, truncated T4 RNA ligase 2, etc.
- any methylase e.g., dam methylase, dem methylase, etc.
- any other enzyme e.g., alkaline phosphatase, T4 polynucleotide kinase, tobacco acid pyrophosphatase, polyadenlyate polymerase, topoisomerase I, topoisomerase II, guanylyl transferase, any helicase, any primase, and recombinase, any protease, any nucleic acid binding protein, any aminoacyl-tRNA synthetase, any glycosyltransferase, any fatty acid synthase and any enzyme for food, biofuel and pharmaceutical industries, such as pal
- the enzymatic domain comprises a polypeptide.
- the polypeptide comprises a polypeptide variant wherein (i) one or more of the amino acid residues are substituted with a conserved or non-conserved amino acid residue, and such substituted amino acid residue may or may not be one encoded by the naturally occurring genetic code, (ii) there are one or more modified amino acid residues, e.g., residues that are modified by the attachment of substituent groups, (iii) the polypeptide is an alternative splice variant of the polypeptide of the present disclosure, (iv) fragments of the polypeptides and/or (v) the polypeptide is fused with another polypeptide, such as a leader or secretory sequence or a sequence which is employed for purification (for example, His-tag) or for detection (for example, Sv5 epitope tag).
- another polypeptide such as a leader or secretory sequence or a sequence which is employed for purification (for example, His-tag) or for detection (
- the fragments include polypeptides generated via proteolytic cleavage (including multi-site proteolysis) of an original sequence. Variants may be post- translationally, or chemically modified. Such variants are deemed to be within the scope of those skilled in the art from the teaching herein.
- the present disclosure includes amino acid sequences that are at least 60%, 65%, 70%, 72%, 74%, 76%, 78%, 80%, 90%, or 95% similar or identical to the original amino acid sequence.
- the degree of identity between two peptides is determined using computer algorithms and methods that are widely known to the persons skilled in the art.
- the identity between two amino acid sequences may be determined by using the BLASTP algorithm (BLAST Manual, Altschul, S., et al., NCBI NLM NIH Bethesda, Md. 20894, Altschul, S., et al., J. Mol. Biol. 215: 403-410 (1990)).
- the polypeptide of the disclosure can be post- translationally modified.
- post-translational modifications that fall within the scope of the present disclosure include signal peptide cleavage, glycosylation, acetylation, isoprenylation, proteolysis, myristoylation, protein folding and proteolytic processing, etc.
- Some modifications or processing events require introduction of additional biological machinery.
- processing events such as signal peptide cleavage and core glycosylation, are examined by adding canine microsomal membranes or Xenopus egg extracts (U.S. Pat. No. 6,103,489) to a standard translation reaction.
- the polypeptide of the disclosure may include unnatural amino acids formed by post-translational modification or by introducing unnatural amino acids during translation.
- a variety of approaches are available for introducing unnatural amino acids during protein translation.
- a polypeptide or protein of the disclosure may be conjugated with other molecules, such as proteins, to prepare fusion proteins. This may be accomplished, for example, by the synthesis of N-terminal or C-terminal fusion proteins provided that the resulting fusion protein retains the functionality of a reverse transcriptase.
- Cyclic derivatives of the polypeptides of the disclosure are also part of the present disclosure. Cyclization may allow the polypeptide to assume a more favorable conformation for association with other molecules. Cyclization may be achieved using techniques known in the art. For example, disulfide bonds may be formed between two appropriately spaced components having free sulfhydryl groups, or an amide bond may be formed between an amino group of one component and a carboxyl group of another component. Cyclization may also be achieved using an azobenzene-containing amino acid as described by Ulysse, L., et al., J. Am. Chem. Soc. 1995, 117, 8466-8467.
- the components that form the bonds may be side chains of amino acids, nonamino acid components or a combination of the two.
- cyclic peptides may comprise a beta-turn in the right position. Beta-turns may be introduced into the peptides of the disclosure by adding the amino acids Pro-Gly at the right position.
- a more flexible polypeptide may be prepared by introducing cysteines at the right and left position of the polypeptide and forming a disulfide bridge between the two cysteines.
- the two cysteines are arranged so as not to deform the beta-sheet and turn.
- the polypeptide is more flexible as a result of the length of the disulfide linkage and the smaller number of hydrogen bonds in the beta-sheet portion.
- the relative flexibility of a cyclic polypeptide can be determined by molecular dynamics simulations.
- the polypeptide comprises a reverse transcriptase fused to, or integrated into, a target protein, and/or a targeting domain capable of directing the chimeric protein to a desired location.
- the chimeric proteins may also comprise additional amino acid sequences or domains.
- the chimeric proteins are recombinant in the sense that the various components are from different sources, and as such are not found together in nature (i.e., are heterologous).
- the polypeptide of the disclosure may be synthesized by conventional techniques.
- the polypeptides or chimeric proteins may be synthesized by chemical synthesis using solid phase peptide synthesis. These methods employ either solid or solution phase synthesis methods (see for example, J. M. Stewart, and J. D. Young, Solid Phase Peptide Synthesis, 2nd Ed., Pierce Chemical Co., Rockford Ill. (1984) and G. Barany and R. B. Merrifield, The Peptides: Analysis Synthesis, Biology editors E. Gross and J. Meienhofer Vol. 2 Academic Press, New York, 1980, pp.
- a polypeptide of the disclosure may be synthesized using 9- fluorenyl methoxycarbonyl (Fmoc) solid phase chemistry with direct incorporation of phosphothreonine as the N-fluorenylmethoxy-carbonyl-O-benzyl-L-phosphothreonine derivative.
- Fmoc 9- fluorenyl methoxycarbonyl
- N-terminal or C-terminal fusion proteins comprising a polypeptide or chimeric protein of the disclosure conjugated with other molecules may be prepared by fusing, through recombinant techniques, the N-terminal or C-terminal of the polypeptide or chimeric protein, and the sequence of a selected protein or selectable marker with a desired biological function.
- the resultant fusion proteins comprise a reverse transcriptase fused to the selected protein or marker protein as described herein.
- proteins which may be used to prepare fusion proteins include immunoglobulins, glutathione-S-transf erase (GST), hemagglutinin (HA), and truncated myc.
- the polypeptide of the disclosure may be developed using a biological expression system.
- the use of these systems allows the production of large libraries of random peptide sequences and the screening of these libraries for peptide sequences that bind to particular proteins.
- Libraries may be produced by cloning synthetic DNA that encodes random peptide sequences into appropriate expression vectors (see Christian et al 1992, J. Mol. Biol. 227:711; Devlin et al, 1990 Science 249:404; Cwirla et al 1990, Proc. Natl. Acad, Sci. USA, 87:6378). Libraries may also be constructed by concurrent synthesis of overlapping peptides (see U.S. Pat. No. 4,708,871).
- the polypeptide of the disclosure may be converted into pharmaceutical salts by reacting with inorganic acids such as hydrochloric acid, sulfuric acid, hydrobromic acid, phosphoric acid, etc., or organic acids such as formic acid, acetic acid, propionic acid, glycolic acid, lactic acid, pyruvic acid, oxalic acid, succinic acid, malic acid, tartaric acid, citric acid, benzoic acid, salicylic acid, benezenesulfonic acid, and toluenesulfonic acids.
- inorganic acids such as hydrochloric acid, sulfuric acid, hydrobromic acid, phosphoric acid, etc.
- organic acids such as formic acid, acetic acid, propionic acid, glycolic acid, lactic acid, pyruvic acid, oxalic acid, succinic acid, malic acid, tartaric acid, citric acid, benzoic acid, salicylic acid, benezenesulfonic acid, and toluenesulfonic
- the present disclosure provides a fusion construct comprising an affinity domain, e g., a domain capable of binding to a specific target or substrate.
- the affinity domain comprises a protein or a fragment of a protein.
- the affinity domain comprises glutathione-S-transferase.
- the affinity domain comprises chitin binding protein.
- the affinity domain comprises maltose binding protein.
- the affinity domain comprises colicin E7 (CL7-tag).
- the affinity domain comprises a SNAP-tag.
- the affinity domain comprises a CLIP-tag.
- the affinity comprises a His tag, e.g., a tag comprising at least two histidine amino acids, e g., at least 2, 3, 4, 5, 6, 7, 8, or more histidine amino acids.
- the affinity domain comprises avidin.
- the affinity tag comprises streptavidin.
- the affinity domain comprises a monomer of avidin or streptavidin.
- the streptavidin domain comprises an amino acid sequence at least 70% identical to the amino acid sequence set forth in SEQ ID NO:1.
- the streptavidin domain comprises an amino acid sequence at least 75% identical to the amino acid sequence set forth in SEQ ID NO:1.
- the streptavidin domain comprises an amino acid sequence at least 80% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 85% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 90% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 95% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 96% identical to the amino acid sequence set forth in SEQ ID NO: 1.
- the streptavidin domain comprises an amino acid sequence at least 97% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 98% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 99% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 99.5% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 99.9% identical to the amino acid sequence set forth in SEQ ID NO:1.
- the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 40 amino acids residues of the amino acid sequence set forth in SEQ ID NO:1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 60 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 80 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 100 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 1.
- the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 40 amino acids residues of the amino acid sequence set forth in SEQ ID NO:1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 60 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 80 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 1.
- the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 100 amino acids residues of the amino acid sequence set forth in SEQ ID NO:1. In one embodiment, the streptavidin domain comprises the amino acid sequence of SEQ ID NO: 1.
- the avidin domain comprises an amino acid sequence at least 70% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 75% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 80% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 85% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 90% identical to the amino acid sequence set forth in SEQ ID NO: 18.
- the avidin domain comprises an amino acid sequence at least 95% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 96% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 97% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 98% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 99% identical to the amino acid sequence set forth in SEQ ID NO: 18.
- the avidin domain comprises an amino acid sequence at least 99.5% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 99.9% identical to the amino acid sequence set forth in SEQ ID NO: 18. [0157] In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 40 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 60 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18.
- the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 80 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 100 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 40 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 60 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18.
- the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 80 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 100 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises the amino acid sequence of SEQ ID NO: 18.
- the variants or fragments of the avidin or streptavidin domain of the present disclosure can comprise variations in the sequence that do not prevent or disrupt folding of the avidin or streptavidin monomer or the ability of the avidin or streptavidin monomer to form a tetramer in the presence of additional avidin or streptavidin monomers.
- the affinity domain is covalently attached to the enzymatic domain by a linker, e.g., a peptide linker.
- the affinity domain is linked to an enzymatic domain comprising a polymerase, e.g., a reverse transcriptase.
- the affinity domain is linked to a reverse transcriptase.
- the affinity domain linked to a polymerase is avidin.
- the affinity domain linked to a polymerase is streptavidin.
- the affinity domain linked to a reverse transcriptase is avidin.
- the affinity domain linked to a reverse transcriptase is streptavidin.
- the affinity domain comprising a monomer of avidin or streptavidin is linked to an enzymatic domain via a peptide linker sequence.
- the present disclosure provides a streptavidin fusion construct comprising a streptavidin domain.
- the streptavidin domain is linked to an enzymatic domain via a linker sequence.
- the present disclosure provides an avidin fusion construct comprising an avidin domain.
- the avidin domain is linked to an enzymatic domain via a linker sequence.
- the fusion construct comprises an avidin or streptavidin domain linked to an enzymatic domain via a linker sequence, e.g., an amino acid linker.
- the linker sequence is a flexible linker.
- the linker sequence comprises between 1-5, 5-10, 10-15, 15-20, 20-25, or 25-30 amino acids.
- the linker sequence comprises between 1-5 amino acids.
- the linker sequence comprises between 5-10 amino acids.
- the linker sequence comprises between 10-15 amino acids.
- the linker sequence comprises between 15-20 amino acids.
- the linker sequence comprises between 20-25 amino acids.
- the linker sequence comprises between 25-30 amino acids.
- the linker sequence comprises at least 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18. 19 or 20 amino acids, or more than 20 amino acids. In some embodiments, the linker sequence comprises at least 1 amino acid. In some embodiments, the linker sequence comprises at least 2 amino acids. In some embodiments, the linker sequence comprises at least 3 amino acids. In some embodiments, the linker sequence comprises at least 4 amino acids. In some embodiments, the linker sequence comprises at least 5 amino acids. In some embodiments, the linker sequence comprises at least 6 amino acids. In some embodiments, the linker sequence comprises at least 7 amino acids. In some embodiments, the linker sequence comprises at least 8 amino acids. In some embodiments, the linker sequence comprises at least 9 amino acids.
- the linker sequence comprises at least 10 amino acids. In some embodiments, the linker sequence comprises at least 11 amino acids. In some embodiments, the linker sequence comprises at least 12 amino acids. In some embodiments, the linker sequence comprises at least 13 amino acids. In some embodiments, the linker sequence comprises at least 14 amino acids. In some embodiments, the linker sequence comprises at least 15 amino acids. In some embodiments, the linker sequence comprises at least 16 amino acids. In some embodiments, the linker sequence comprises at least 17 amino acids. In some embodiments, the linker sequence comprises at least 18 amino acids. In some embodiments, the linker sequence comprises at least 19 amino acids. In some embodiments, the linker sequence comprises at least 20 amino acids.
- the linker sequence comprises a type of amino acid, e.g., a polar amino acid, a nonpolar amino acid, a hydrophobic amino acid, or a charged amino acid.
- the linker sequence comprises a polar amino acid, e.g., serine (S), threonine (T), cysteine (C), tyrosine (Y), asparagine (N), glutamine (Q), or glycine (G).
- the linker sequence comprises an S amino acid.
- the linker sequence comprises a T amino acid.
- the linker sequence comprises a C amino acid.
- the linker sequence comprises a Y amino acid.
- the linker sequence comprises an N amino acid. In some embodiments, the linker sequence comprises a Q amino acid. In some embodiments, the linker sequence comprises a G amino acid. In some embodiments, the linker sequence comprises at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 polar amino acids. In some embodiments, the linker sequence comprises at least 1 S amino acid. In some embodiments, the linker sequence comprises at least 2 S amino acids. In some embodiments, the linker sequence comprises at least 3 S amino acids. In some embodiments, the linker sequence comprises at least 4 S amino acids. In some embodiments, the linker sequence comprises at least 5 S amino acids. In some embodiments, the linker sequence comprises at least 6 S amino acids.
- the linker sequence comprises at least 7 S amino acids. In some embodiments, the linker sequence comprises at least 8 S amino acids. In some embodiments, the linker sequence comprises at least 9 S amino acids. In some embodiments, the linker sequence comprises at least 10 S amino acids. In some embodiments, the linker sequence comprises at least 1 G amino acid. In some embodiments, the linker sequence comprises at least 2 G amino acids. In some embodiments, the linker sequence comprises at least 3 G amino acids. In some embodiments, the linker sequence comprises at least 4 G amino acids. In some embodiments, the linker sequence comprises at least 5 G amino acids. Tn some embodiments, the linker sequence comprises at least 6 G amino acids. In some embodiments, the linker sequence comprises at least 7 G amino acids. In some embodiments, the linker sequence comprises at least 8 G amino acids. In some embodiments, the linker sequence comprises at least 9 G amino acids. In some embodiments, the linker sequence comprises at least 10 G amino acids.
- the linker sequence comprises a hydrophobic amino acid, e.g., alanine (A), valine (V), leucine (L), isoleucine (I), methionine (M), phenylalanine (F), tryptophan (W), or proline (P).
- the linker sequence comprises an A amino acid.
- the linker sequence comprises a V amino acid.
- the linker sequence comprises an L amino acid.
- the linker sequence comprises an I amino acid.
- the linker sequence comprises an M amino acid.
- the linker sequence comprises an F amino acid.
- the linker sequence comprises a W amino acid.
- the linker sequence comprises a P amino acid. In some embodiments, the linker sequence comprises at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 hydrophobic amino acids. In some embodiments, the linker sequence comprises at least 1 A amino acid. In some embodiments, the linker sequence comprises at least 2 A amino acids. In some embodiments, the linker sequence comprises at least 3 A amino acids. In some embodiments, the linker sequence comprises at least 4 A amino acids. In some embodiments, the linker sequence comprises at least 5 A amino acids. In some embodiments, the linker sequence comprises at least 6 A amino acids. In some embodiments, the linker sequence comprises at least 7 A amino acids. In some embodiments, the linker sequence comprises at least 8 A amino acids. In some embodiments, the linker sequence comprises at least 9 A amino acids. In some embodiments, the linker sequence comprises at least 10 A amino acids.
- the linker sequence comprises a charged amino acid, e.g., histidine (H), lysine (K), arginine (R), aspartate (D), or glutamate (E).
- the linker sequence comprises an H amino acid.
- the linker sequence comprises a K amino acid.
- the linker sequence comprises an R amino acid.
- the linker sequence comprises a D amino acid.
- the linker sequence comprises an E amino acid.
- the linker sequence comprises at least 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 charged amino acids.
- the linker sequence can comprise any sequence capable of linking the avidin or streptavidin domain to the enzymatic domain, and is not limited to the linker sequences described herein.
- the linker comprises an amino acid sequence at least 80% identical to the sequence set forth in SEQ ID NO: 2, e.g., at least 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 99.9% identical to SEQ ID NO: 2.
- the linker comprises an amino acid sequence at least 80% identical to the sequence set forth in SEQ ID NO:2.
- the linker comprises an amino acid sequence at least 85% identical to the sequence set forth in SEQ ID NO:2.
- the linker comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 95% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 96% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 97% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 98% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 99% identical to the sequence set forth in SEQ ID NO:2.
- the linker comprises an amino acid sequence at least 99.5% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 99.9% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises SEQ ID NO:2.
- the linker comprises an amino acid sequence at least 80% identical to the sequence set forth in SEQ ID NO: 5, e.g., at least 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 99.9% identical to SEQ ID NO: 5.
- the linker comprises an amino acid sequence at least 80% identical to the sequence set forth in SEQ ID NO:5.
- the linker comprises an amino acid sequence at least 85% identical to the sequence set forth in SEQ ID NO:5.
- the linker comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 95% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 96% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 97% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 98% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 99% identical to the sequence set forth in SEQ ID NO:5.
- the linker comprises an amino acid sequence at least 99.5% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 99.9% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises SEQ ID NO:5.
- the present disclosure provides a composition comprising at least one streptavidin fusion construct comprising a streptavidin domain operably linked to an enzymatic domain.
- the streptavidin domain is linked to the N- or C- terminus of the enzymatic domain via a linker sequence.
- the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to the amino acid sequence set forth in SEQ ID NO:1, and is linked at is C-terminus to the N-terminus of an amino acid sequence that is at least about 70% identical to the amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NOTO, SEQ ID NOT l, SEQ ID NO:12 or SEQ ID NO: 13 or a functional fragment thereof, via an amino acid sequence that is at least about 70% identical to the amino acid sequence set forth in SEQ ID NO:2.
- the streptavidin fusion construct comprises an amino acid sequence as set forth in SEQ ID NOT, SEQ ID NOT, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13 operably linked to a streptavidin monomeric domain comprising a sequence as set forth in SEQ ID NO: 1.
- the C terminus of SEQ ID NO: 1 is linked to the N-terminus of SEQ ID NOT, SEQ ID NOT, SEQ ID NO:9, SEQ ID NOTO, SEQ ID NOT 1, SEQ ID NO: 12 or SEQ ID NOT 3 via a linker sequence.
- the linker sequence comprises SEQ ID NOT.
- the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO:3.
- the linker comprises SEQ ID NO:2.
- the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO:8.
- the linker comprises SEQ ID NO:2.
- the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO:9.
- the linker comprises SEQ ID NO:2.
- the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO: 10.
- the linker comprises SEQ ID NO:2.
- the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO: 11.
- the linker comprises SEQ ID NO:2.
- the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO: 12.
- the linker comprises SEQ ID NO:2.
- the streptavidin fusion construct comprises an amino acid sequence that is at least 70% identical to the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the streptavidin fusion construct comprises an amino acid sequence that is at least 75% identical to the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the streptavidin fusion construct comprises an amino acid sequence that is at least 80% identical to the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the streptavidin fusion construct comprises an amino acid sequence that is at least 85% identical to the amino acid sequence set forth in SEQ ID NON, SEQ ID NO:6 or SEQ ID NO:7. [0180] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least 90% identical to the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the streptavidin fusion construct comprises an amino acid sequence that is at least 95% identical to the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 250 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 300 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 350 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 400 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 450 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 500 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7. [0188] In one embodiment, the streptavidin fusion construct comprises the amino acid sequence as set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
- the fusion constructs of the disclosure are linked to a peptide tag.
- the peptide tag of the present disclosure comprises any suitable peptide tag.
- the peptide tag can be removed or cleaved through protease treatment.
- the peptide tag is beneficial to verify that a protein multimer comprises the appropriate number of monomers.
- peptide tags of this nature are well known in the field and will consequently be able to select an appropriate peptide tag for use with the current disclosure. A skilled artisan will additional be able to select the appropriate protease for subsequent cleavage of the peptide tag.
- the peptide tag comprises a His6-Small Ubiquitin-like Modifier (SUMO) tag.
- SUMO His6-Small Ubiquitin-like Modifier
- the peptide tag is cleaved through protease treatment of the at least one protein multimer.
- the protease comprises SUMO-protease.
- the present disclosure provides a composition comprising at least one protein multimer, wherein the at least one protein multimer comprises at least one streptavidin fusion construct of the disclosure.
- the protein multimer comprises a streptavidin tetramer comprising at least one streptavidin fusion construct of the disclosure.
- the protein multimer comprises a streptavidin tetramer comprising at least two streptavidin fusion constructs of the disclosure.
- the protein multimer comprises a streptavidin tetramer comprising at least three streptavidin fusion constructs of the disclosure.
- the protein multimer comprises one streptavidin fusion construct of the disclosure in a tetrameric complex with three additional streptavidin monomers. In some embodiments, the protein multimer comprises two streptavidin fusion constructs of the disclosure in a tetrameric complex with two additional streptavidin monomers. In some embodiments, the protein multimer comprises three streptavidin fusion constructs of the disclosure in a tetrameric complex with one additional streptavidin monomer.
- one or more of the streptavidin domains of a streptavidin fusion construct or a streptavidin monomer incorporated into a tetramer of the disclosure comprise an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO: 1.
- the present disclosure provides a composition comprising an isolated nucleic acid encoding a streptavidin fusion construct of the disclosure.
- the composition comprises a nucleic acid encoding streptavidin fusion construct of the disclosure comprising a streptavidin domain fused to an enzymatic domain.
- the nucleic acid is DNA, RNA, mRNA, or cDNA.
- enzymatic domains that can be encoded by the nucleic acid molecules of the disclosure are described elsewhere herein and include, but are not limited to, reverse transcriptases, translocating helicase enzymes, DNA polymerases, and RNA polymerases.
- the enzymatic domain comprises a reverse transcriptase derived from E.r. maturase.
- the enzymatic domain comprises a reverse transcriptase, wherein the reverse transcriptase comprises E.r. maturase or variant thereof.
- the isolated nucleic acid encodes a reverse transcriptase comprising E.r. maturase, or a variant thereof, wherein the amino acid sequence of E.r. maturase, or a variant thereof, is at least about 90% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the isolated nucleic acid encodes a reverse transcriptase comprising E.r. maturase, or a variant thereof, wherein the amino acid sequence of E.r. maturase, or a variant thereof, is identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the isolated nucleic acid comprises a nucleotide sequence encoding an amino acid at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
- the isolated nucleic acid encodes an E.r. maturase variant comprising at least one point mutation selected from the group R58X, K59X, K61X, K163X, K216X, R217X, K338X, K342X, and R353X relative to wildtype E.r. maturase, wherein X denotes any amino acid.
- the nucleic acid encodes an E.r. maturase variant comprising at least one point mutation selected from the group R58A, K59A, K61 A, K163A, K216A, R217A, K338A, K342A, and R353A relative to wildtype E.r. maturase.
- the isolated nucleic acid encodes an E.r. maturase variant (referred to herein as E.r. maturase mutl; and denoted as SEQ ID NO: 8) comprising the point mutations of: R58A, K59A, K61A, and K163A, relative to wildtype E.r. maturase.
- E.r. maturase mutl referred to herein as E.r. maturase mutl
- SEQ ID NO: 8 comprising the point mutations of: R58A, K59A, K61A, and K163A, relative to wildtype E.r. maturase.
- the isolated nucleic acid encodes an E.r. maturase variant (referred to herein as E.r. maturase mut2; and denoted as SEQ ID NO:9) comprising the point mutations of: K216A and K217A, relative to wildtype E.r. maturase.
- E.r. maturase mut2 referred to herein as E.r. maturase mut2; and denoted as SEQ ID NO:9 comprising the point mutations of: K216A and K217A, relative to wildtype E.r. maturase.
- the isolated nucleic acid encodes an E.r. maturase variant (referred to herein as E.r. maturase mutl+mut2; and denoted as SEQ ID NO: 10) comprising the point mutations of: R58A, K59A, K61A, K163A, K216A, and R217A, relative to wildtype E.r. maturase.
- E.r. maturase mutl+mut2 referred to herein as E.r. maturase mutl+mut2; and denoted as SEQ ID NO: 10.
- the n isolated ucleic acid encodes an E.r. maturase variant (referred to herein as E.r. maturase mut3; and denoted as SEQ ID NO: 11) comprising the point mutations of: K338A, K342A, and R353A relative to wildtype E.r. maturase.
- E.r. maturase mut3 referred to herein as E.r. maturase mut3; and denoted as SEQ ID NO: 11
- the composition increases the expression of a biologically functional fragment of E.r. maturase.
- the composition comprises an isolated nucleic acid sequence encoding a biologically functional fragment of E.r. maturase.
- a biologically functional fragment is a portion or portions of a full-length sequence that retain at least one biological function of the full-length sequence.
- a biologically functional fragment of E.r. maturase comprises a peptide that retains at least one function of full length E.r. maturase.
- the isolated nucleic acid encodes a streptavidin fusion construct comprising an enzymatic domain comprising a peptide having substantial homology to a reverse transcriptase disclosed herein.
- the isolated nucleic acid sequence encodes a reverse transcriptase having at least 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% sequence identity to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID N0:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13 or a functional fragment thereof.
- the isolated nucleic acid sequence encoding a reverse transcriptase can be obtained using any of the many recombinant methods known in the art, such as, for example by screening libraries from cells expressing the gene, by deriving the gene from a vector known to include the same, or by isolating directly from cells and tissues containing the same, using standard techniques.
- the gene of interest can be produced synthetically, rather than cloned.
- the isolated nucleic acid may comprise any type of nucleic acid, including, but not limited to DNA and RNA.
- the composition comprises an isolated DNA molecule, including for example, an isolated cDNA molecule, encoding a reverse transcriptase.
- the composition comprises an isolated RNA molecule encoding a streptavidin fusion construct of the disclosure.
- the present disclosure provides an isolated nucleic acid encoding a streptavidin fusion construct comprising a nucleotide sequence encoding an amino acid at least about 90% identical to the amino acid sequence as set forth in SEQ ID NO: 1.
- the isolated nucleic acid comprises a nucleotide sequence encoding an amino acid at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% identical to the amino acid sequence as set forth in SEQ ID NO: 1 .
- the isolated nucleic acid comprises a nucleotide sequence encoding SEQ ID NO:1.
- the isolated nucleic acid encodes a streptavidin fusion construct comprising a nucleotide sequence encoding an amino acid at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% identical to the amino acid sequence as set forth in SEQ ID NO:1 linked to a nucleotide sequence encoding a reverse transcriptase having at least 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% sequence identity to an amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13 or a functional fragment thereof
- the isolated nucleic acid encodes a linker.
- the isolated nucleic acid comprises a nucleotide sequence encoding an amino acid at least about 90% identical to the amino acid sequence as set forth in SEQ ID NO:2.
- the isolated nucleic acid comprises a nucleotide sequence encoding SEQ ID NO:2.
- the isolated nucleic acid comprises a nucleotide sequence encoding an amino acid at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% identical to the amino acid sequence as set forth in SEQ ID NON, SEQ ID NO:6 or SEQ ID NO:7.
- the isolated nucleic acid comprises a nucleotide sequence encoding SEQ ID NON, SEQ ID NO:6 or SEQ ID NON.
- the linker sequence can comprise any sequence capable of linking the streptavidin domain to the enzymatic domain, wherein the enzymatic domain maintains at least one wild type catalytic activity in the oligomer, and is not limited to the linker sequences described herein.
- the nucleic acid molecules of the present disclosure can be modified to improve stability in serum or in growth medium for cell cultures. Modifications can be added to enhance stability, functionality, and/or specificity. For example, in order to enhance the stability, the 3’- residues may be stabilized against degradation, e.g., they may be selected such that they consist of purine nucleotides, particularly adenosine or guanosine nucleotides. Alternatively, substitution of pyrimidine nucleotides by modified analogues, e.g., substitution of uridine by 2’- deoxythymidine is tolerated and does not affect function of the molecule.
- the nucleic acid molecule may comprise at least one modified nucleotide analogue.
- the ends may be stabilized by incorporating modified nucleotide analogues.
- Non-limiting examples of nucleotide analogues include sugar- and/or backbone- modified ribonucleotides (i.e., include modifications to the phosphate-sugar backbone).
- the phosphodiester linkages of natural RNA may be modified to include at least one of a nitrogen or sulfur heteroatom.
- the phosphoester group connecting to adjacent ribonucleotides is replaced by a modified group, e.g., of phosphothioate group.
- the 2’ OH-group is replaced by a group selected from H, OR, R, halo, SH, SR, NH2, NHR, NR2 or ON, wherein R is Ci-Ce alkyl, alkenyl or alkynyl and halo is F, Cl, Br or I.
- nucleobase-modified ribonucleotides i.e., ribonucleotides, containing at least one non-naturally occurring nucleobase instead of a naturally occurring nucleobase.
- Bases may be modified to block the activity of adenosine deaminase.
- modified nucleobases include, but are not limited to, uridine and/or cytidine modified at the 5-position, e.g., 5-(2-amino)propyl uridine, 5-bromo uridine; adenosine and/or guanosines modified at the 8 position, e.g., 8-bromo guanosine; deaza nucleotides, e.g., 7-deaza-adenosine; O- and N-alkylated nucleotides, e.g., N6-methyl adenosine are suitable.
- the above modifications may be combined.
- the nucleic acid molecule comprises at least one of the following chemical modifications: 2’-H, 2’-O-methyl, or 2’-OH modification of one or more nucleotides.
- a nucleic acid molecule of the disclosure can have enhanced resistance to nucleases.
- a nucleic acid molecule can include, for example, 2’ -modified ribose units and/or phosphorothioate linkages.
- the 2’ hydroxyl group (OH) can be modified or replaced with a number of different “oxy” or “deoxy” substituents.
- the nucleic acid molecules of the disclosure can include 2’-O-methyl, 2’-fluorine, 2’-O-methoxyethyl, 2’-O-aminopropyl, 2’- amino, and/or phosphorothioate linkages.
- LNA locked nucleic acids
- ENA ethylene nucleic acids
- 2’-4’-ethylene-bridged nucleic acids e.g., 2’-4’-ethylene-bridged nucleic acids
- certain nucleobase modifications such as 2-amino-A, 2-thio (e.g., 2-thio-U), G-clamp modifications, can also increase binding affinity to a target.
- the nucleic acid molecule includes a 2’ -modified nucleotide, e.g., a 2’-deoxy, 2 ’-deoxy-2’ -fluoro, 2’-O-methyl, 2’-O-methoxyethyl (2’-O-MOE), 2’-O- aminopropyl (2’-O-AP), 2’-O-dimethylaminoethyl (2’-O-DMAOE), 2’-O-dimethylaminopropyl (2’-O-DMAP), 2’-O-dimethylaminoethyloxyethyl (2’-O-DMAEOE), or 2’-O-N- methylacetamido (2’-0-NMA).
- the nucleic acid molecule includes at least one 2’-O-methyl-modified nucleotide, and in some embodiments, all of the nucleotides of the nucleic acid molecule include a 2’
- the nucleic acid molecule of the disclosure may have one or more of the following properties:
- Nucleic acid agents discussed herein include otherwise unmodified RNA and DNA as well as RNA and DNA that have been modified, e.g., to improve efficacy, and polymers of nucleoside surrogates.
- Unmodified RNA refers to a molecule in which the components of the nucleic acid, namely sugars, bases, and phosphate moieties, are the same or essentially the same as that which occur in nature.
- the art has referred to rare or unusual, but naturally occurring, RNAs as modified RNAs, see, e.g., Limbach et al. (Nucleic Acids Res., 1994, 22:2183-2196).
- modified RNA refers to a molecule in which one or more of the components of the nucleic acid, namely sugars, bases, and phosphate moieties, are different from those which occur in nature. While they are referred to as “modified RNAs” they will of course, because of the modification, include molecules that are not, strictly speaking, RNAs.
- Nucleoside surrogates are molecules in which the ribophosphate backbone is replaced with a non-ribophosphate construct that allows the bases to be presented in the correct spatial relationship such that hybridization is substantially similar to what is seen with a ribophosphate backbone, e.g., non-charged mimics of the ribophosphate backbone.
- Modifications of the nucleic acid of the disclosure may be present at one or more of, a phosphate group, a sugar group, backbone, N-terminus, C-terminus, or nucleobase.
- the present disclosure also includes a vector in which the isolated nucleic acid of the present disclosure is inserted.
- the art is replete with suitable vectors that are useful in the present disclosure.
- the expression of natural or synthetic nucleic acids encoding a reverse transcriptase described herein is typically achieved by operably linking a nucleic acid encoding a streptavidin fusion construct of the disclosure to a promoter, and incorporating the construct into an expression vector.
- the vectors to be used are suitable for replication and, optionally, integration in host cells. Typical vectors contain transcription and translation terminators, initiation sequences, and promoters useful for regulation of the expression of the desired nucleic acid sequence.
- the isolated nucleic acid of the disclosure can be cloned into many types of vectors.
- the nucleic acid can be cloned into a vector including, but not limited to a plasmid, a phagemid, a phage derivative, an animal virus, and a cosmid.
- Vectors of particular interest include expression vectors, replication vectors, probe generation vectors, and sequencing vectors.
- the vector may be provided to a cell in the form of a viral vector.
- Viral vector technology is well known in the art and is described, for example, in Sambrook et al. (2012, Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Laboratory, New York), and in other virology and molecular biology manuals.
- Viruses, which are useful as vectors include, but are not limited to, retroviruses, adenoviruses, adeno- associated viruses, herpes viruses, and lentiviruses.
- a suitable vector contains an origin of replication functional in at least one organism, a promoter sequence, convenient restriction endonuclease sites, and one or more selectable markers, (e.g., WO 01/96584; WO 01/29058; and U.S. Pat. No. 6,326,193).
- retroviruses provide a convenient platform for gene delivery systems.
- a selected gene can be inserted into a vector and packaged in retroviral particles using techniques known in the art.
- the recombinant virus can then be isolated and delivered to cells.
- retroviral systems are known in the art.
- adenovirus vectors are used.
- a number of adenovirus vectors are known in the art.
- lentivirus vectors are used.
- vectors derived from retroviruses such as the lentivirus are suitable tools to achieve long-term gene transfer since they allow long-term, stable integration of a transgene and its propagation in daughter cells.
- Lentiviral vectors have the added advantage over vectors derived from onco-retroviruses such as murine leukemia viruses in that they can transduce non-proliferating cells, such as hepatocytes. They also have the added advantage of low immunogenicity.
- the composition includes a vector derived from an adeno-associated virus (AAV).
- Adeno-associated viral (AAV) vectors have become powerful gene delivery tools for the treatment of various disorders.
- AAV vectors possess a number of features that render them ideally suited for use, including a lack of pathogenicity, minimal immunogenicity, and the ability to transduce post-mitotic cells in a stable and efficient manner. Expression of a particular gene contained within an AAV vector can be specifically targeted to one or more types of cells by choosing the appropriate combination of AAV serotype, promoter, and delivery method.
- the vector also includes conventional control elements which are operably linked to the transgene in a manner which permits its transcription, translation and/or expression in a cell transfected with the plasmid vector or infected with the virus produced by the disclosure.
- operably linked sequences include both expression control sequences that are contiguous with the gene of interest and expression control sequences that act in trans or at a distance to control the gene of interest.
- Expression control sequences include appropriate transcription initiation, termination, promoter and enhancer sequences; efficient RNA processing signals such as splicing and polyadenylation (polyA) signals; sequences that stabilize cytoplasmic mRNA; sequences that enhance translation efficiency (i.e., Kozak consensus sequence); sequences that enhance protein stability; and when desired, sequences that enhance secretion of the encoded product.
- polyA polyadenylation
- a great number of expression control sequences, including promoters which are native, constitutive, inducible and/or tissuespecific, are known in the art and may be utilized.
- Additional promoter elements e.g., enhancers, regulate the frequency of transcriptional initiation.
- these are located in the region 30-110 bp upstream of the start site, although a number of promoters have recently been shown to contain functional elements downstream of the start site as well.
- the spacing between promoter elements frequently is flexible, so that promoter function is preserved when elements are inverted or moved relative to one another.
- tk thymidine kinase
- the spacing between promoter elements can be increased to 50 bp apart before activity begins to decline.
- individual elements can function either cooperatively or independently to activate transcription.
- a suitable promoter is the immediate early cytomegalovirus (CMV) promoter sequence.
- CMV immediate early cytomegalovirus
- This promoter sequence is a strong constitutive promoter sequence capable of driving high levels of expression of any polynucleotide sequence operatively linked thereto.
- Another example of a suitable promoter is Elongation Growth Factor -la (EF-la).
- constitutive promoter sequences may also be used, including, but not limited to the simian virus 40 (SV40) early promoter, mouse mammary tumor virus (MMTV), human immunodeficiency virus (HIV) long terminal repeat (LTR) promoter, MoMuLV promoter, an avian leukemia virus promoter, an Epstein-Barr virus immediate early promoter, a Rous sarcoma virus promoter, as well as human gene promoters such as, but not limited to, the actin promoter, the myosin promoter, the hemoglobin promoter, and the creatine kinase promoter. Further, the disclosure should not be limited to the use of constitutive promoters.
- Enhancer sequences found on a vector also regulate expression of the gene contained therein.
- enhancers are bound with protein factors to enhance the transcription of a gene.
- An enhancer may be located upstream or downstream of the gene it regulates. Enhancers may also be tissue-specific to enhance transcription in a specific cell or tissue type.
- the vector of the present disclosure comprises one or more enhancers to boost transcription of the gene present within the vector.
- the expression vector to be introduced into a cell can also comprise either a selectable marker gene or a reporter gene or both to facilitate identification and selection of expressing cells from the population of cells sought to be transfected or infected through viral vectors.
- the selectable marker may be carried on a separate piece of DNA and used in a cotransfection procedure. Both selectable markers and reporter genes may be flanked with appropriate regulatory sequences to enable expression in the host cells.
- Useful selectable markers include, for example, antibiotic-resistance genes, such as neo and the like.
- Reporter genes are used for identifying potentially transfected cells and for evaluating the functionality of regulatory sequences.
- a reporter gene is a gene that is not present in or expressed by the recipient organism or tissue and that encodes a polypeptide whose expression is manifested by some easily detectable property, e.g., enzymatic activity. Expression of the reporter gene is assayed at a suitable time after the DNA has been introduced into the recipient cells.
- Suitable reporter genes may include genes encoding luciferase, betagalactosidase, chloramphenicol acetyl transferase, secreted alkaline phosphatase, or the green fluorescent protein gene (e.g., Ui-Tei et al., 2000 FEBS Letters 479: 79-82).
- Suitable expression systems are well known and may be prepared using known techniques or obtained commercially.
- the construct with the minimal 5' flanking region showing the highest level of expression of reporter gene is identified as the promoter.
- Such promoter regions may be linked to a reporter gene and used to evaluate agents for the ability to modulate promoter- driven transcription.
- the vector can be readily introduced into a host cell, e.g., mammalian, bacterial, yeast, or insect cell by any method in the art.
- the expression vector can be transferred into a host cell by physical, chemical, or biological means.
- Physical methods for introducing a polynucleotide into a host cell include calcium phosphate precipitation, lipofection, particle bombardment, microinjection, electroporation, and the like. Methods for producing cells comprising vectors and/or exogenous nucleic acids are well-known in the art. See, for example, Sambrook et al. (2012, Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Laboratory, New York). A preferred method for the introduction of a polynucleotide into a host cell is calcium phosphate transfection.
- Biological methods for introducing a polynucleotide of interest into a host cell include the use of DNA and RNA vectors.
- Viral vectors, and especially retroviral vectors have become the most widely used method for inserting genes into mammalian, e.g., human cells.
- Other viral vectors can be derived from lentivirus, poxviruses, herpes simplex virus I, adenoviruses and adeno-associated viruses, and the like. See, for example, U.S. Pat. Nos. 5,350,674 and 5,585,362.
- Chemical means for introducing a polynucleotide into a host cell include colloidal dispersion systems, such as macromolecule complexes, nanocapsules, microspheres, beads, and lipid-based systems including oil-in-water emulsions, micelles, mixed micelles, and liposomes.
- colloidal dispersion systems such as macromolecule complexes, nanocapsules, microspheres, beads, and lipid-based systems including oil-in-water emulsions, micelles, mixed micelles, and liposomes.
- An exemplary colloidal system for use as a delivery vehicle in vitro and in vivo is a liposome (e g., an artificial membrane vesicle).
- an exemplary delivery vehicle is a liposome.
- lipid formulations is contemplated for the introduction of the nucleic acids into a host cell (in vitro, ex vivo or in vivo).
- the nucleic acid may be associated with a lipid.
- the nucleic acid associated with a lipid may be encapsulated in the aqueous interior of a liposome, interspersed within the lipid bilayer of a liposome, attached to a liposome via a linking molecule that is associated with both the liposome and the oligonucleotide, entrapped in a liposome, complexed with a liposome, dispersed in a solution comprising a lipid, mixed with a lipid, combined with a lipid, contained as a suspension in a lipid, contained or complexed with a micelle, or otherwise associated with a lipid.
- Lipid, lipid/DNA or lipid/expression vector associated compositions are not limited to any particular structure in solution.
- Lipids are fatty substances which may be naturally occurring or synthetic lipids.
- lipids include the fatty droplets that naturally occur in the cytoplasm as well as the class of compounds which contain long-chain aliphatic hydrocarbons and their derivatives, such as fatty acids, alcohols, amines, amino alcohols, and aldehydes.
- Lipids suitable for use can be obtained from commercial sources.
- DMPC dimyristyl phosphatidylcholine
- DCP dicetyl phosphate
- Choi cholesterol
- DMPG dimyristyl phosphatidylglycerol
- Stock solutions of lipids in chloroform or chloroform/methanol can be stored at about -20°C.
- Liposome is a generic term encompassing a variety of single and multilamellar lipid vehicles formed by the generation of enclosed lipid bilayers or aggregates. Liposomes can be characterized as having vesicular structures with a phospholipid bilayer membrane and an inner aqueous medium. Multilamellar liposomes have multiple lipid layers separated by aqueous medium. They form spontaneously when phospholipids are suspended in an excess of aqueous solution.
- compositions that have different structures in solution than the normal vesicular structure are also encompassed.
- the lipids may assume a micellar structure or merely exist as nonuniform aggregates of lipid molecules.
- lipofectamine-nucleic acid complexes are also contemplated.
- assays include, for example, “molecular biological” assays well known to those of skill in the art, such as Southern and Northern blotting, RT-PCR and PCR; “biochemical” assays, such as detecting the presence or absence of a particular peptide, e.g., by immunological means (ELISAs and Western blots) or by assays described herein to identify agents falling within the scope of the disclosure.
- molecular biological assays well known to those of skill in the art, such as Southern and Northern blotting, RT-PCR and PCR
- biochemical assays, such as detecting the presence or absence of a particular peptide, e.g., by immunological means (ELISAs and Western blots) or by assays described herein to identify agents falling within the scope of the disclosure.
- the present disclosure provides a delivery vehicle comprising a reverse transcriptase, or a nucleic acid molecule encoding a reverse transcriptase.
- exemplary delivery vehicles include, but are not limited to, microspheres, microparticles, nanoparticles, polymerosomes, liposomes, and micelles.
- the delivery vehicle is loaded with a reverse transcriptase, or a nucleic acid molecule encoding a reverse transcriptase.
- the delivery vehicle provides for controlled release, delayed release, or continual release of its loaded cargo.
- the delivery vehicle comprises a targeting moiety that targets the delivery vehicle to a particular location.
- the present disclosure provides a full-length cDNA derived from a full-length RNA, produced by a reverse transcriptase described herein.
- the RNA has significant secondary or tertiary structure, and/or is long (greater than or equal to 5,000 bases in length). For example, it is described herein that E.r. maturase and E.r. maturase-derived peptides described herein are highly processive reverse transcriptases.
- the RNA reverse transcribed into DNA is at least about 100, at least about 200, at least about 300, at least about 400, at least about 500, at least about 600, at least about 700, at least about 800, at least about 900, at least about 1000, at least about 2000, at least about 3000, at least about 4000, at least about 5000, at least about 6000, at least about 7000, at least about 8000, at least about 9000, or at least about 10000 bases in length.
- the DNA so reverse transcribed is at least about 100, at least about 200, at least about 300, at least about 400, at least about 500, at least about 600, at least about 700, at least about 800, at least about 900, at least about 1000, at least about 2000, at least about 3000, at least about 4000, at least about 5000, at least about 6000, at least about 7000, at least about 8000, at least about 9000, or at least about 10000 bases in length.
- the at least one protein tetramer, of the present disclosure is linked to a solid support, wherein the linkage to the solid support occurs via the interaction of the avidin or streptavidin tetramer with at least one component attached to the solid support, wherein the at least one component attached to the solid support has high binding affinity for the streptavidin tetramer.
- the at least one component attached to the solid support comprises biotin.
- the solid support is comprised in a device or procedure utilized in molecular applications.
- the device or procedure utilized in molecular applications comprises a device or procedure utilized in nucleic acid sequencing applications.
- the device or procedure utilized in molecular applications comprises a device or procedure utilized in any molecular application that requires the reverse transcriptase activity of the compositions of the present disclosure. In some embodiments, the device or procedure utilized in molecular applications comprises a device or procedure utilized in any molecular application that requires the catalytic activity of the compositions of the present disclosure.
- any device or procedure that employs reverse transcription as a method or step can utilize the compositions of the present disclosure.
- the compositions of the present disclosure are used to perform reverse transcription as part of an assay.
- the assay may be at least one selected from the group RT-PCR, qRT-PCR, capillary electrophoresis (CE) for RNA-structure mapping (such as SHAPE-seq or SHAPE-map, DMS-seq), in-cell sequencing, next-generation RNA sequencing (RNA-seq), nanopore sequencing, cDNA library synthesis, cDNA synthesis, and a combination thereof.
- compositions provide for reverse transcription at physiologic temperatures, or at lower temperatures relative to that required when using non-/:./' maturase-derived reverse transcriptases.
- the lower temperature of the reverse transcription reaction provides a decreased rate of degradation of the RNA molecule during the reaction, relative to the rate of degradation of an RNA molecule in a reverse transcription reaction that uses a non-E.r maturase-derived reverse transcriptase.
- compositions of the present disclosure provide for performing reverse transcription of a long and/or complex RNA molecule.
- the reverse transcriptase exhibits increased stability.
- the reverse transcriptase exhibits reduced turnover.
- the reverse transcriptases described herein have reduced turnover, thereby allowing the synthesis of longer reads and full-length DNA products.
- compositions provide for performing a single reaction amplification of RNA, made possible by the true thermocycling ability of the reverse transcriptases described herein.
- thermocycling ability of the reverse transcriptases described herein allows for the amplification of RNA without the need for DNA replication.
- compositions provided herein are utilized in a quantitative RT-PCR (qRT-PCR) device or procedure.
- qRT-PCR quantitative RT-PCR
- the formation of PCR products is monitored in each cycle of the PCR.
- the amplification is usually measured in thermocyclers which have additional devices for measuring fluorescence signals during the amplification reaction. See, for example, U.S. Pat. No. 6,174,670, and U.S. Pat. No. 8,137,616.
- the qRT-PCR procedure is carried out using a thermostable improved E.r. maturase enzyme, without a DNA->DNA polymerase.
- the compositions provided herein are utilized in a capillary electrophoresis (CE) for RNA-structure mapping device or procedure.
- CE capillary electrophoresis
- the application of capillary electrophoresis to RNA structure probing is an important step in increasing the throughput of RNA structure data.
- Gel electrophoresis typically resolves about a hundred bases of RNA at a time, and hence probing an RNA of several kilobases long might require running tens to hundreds of gels.
- Capillary electrophoresis allows the resolution of 300-650 bases from a structure probing experiment and multiple lanes can be run at the same time to increase the throughput of RNA structure probing.
- the readout of the probing experiment is typically through the reverse transcription of a 5' fluorescently labeled DNA primer that anneals specifically to the RNA of interest. If the RNA is several kilobases long, multiple primers are designed to anneal along the length of the transcript. Modification or cleavage of the RNA template results in premature stops in the primer extension reaction, leading to different lengths of the cDNA product which are resolved by capillary electrophoresis.
- Software tools such as CAFA and Shapefinder can automate the data acquisition from capillary electrophoresis and further improve speed and accuracy (see, for example, Wan, Y. et al., 2011, Nat Rev Genet., 12(9): 1-26).
- compositions provided herein are utilized in a nextgeneration RNA sequencing (RNA-seq) device or procedure.
- RNA-Seq nextgeneration RNA sequencing
- RNA-Seq High-throughput RNA sequencing
- a standard RNA-Seq library is generated from ligating sequencing adapters to double-stranded DNA.
- Another, more widely used method comprises incorporating dUTP in addition to dNTPs in the second strand DNA synthesis.
- the second strand DNA can be specifically digested by an Uracil-N-glycosylase (UNG) enzyme so that only the library strand containing the first strand cDNA will be sequenced and information on the direction of the transcripts can therefore be obtained (see M. Sultan et al., Biochemical and Biophysical Research Communications 422 (2012) 643-646; also see PCT Patent Application Number PCT/EP2016/069997).
- UNG Uracil-N-glycosylase
- the disclosure is also directed to methods for making one or more nucleic acid molecules and/or labeled nucleic acid molecules, comprising mixing one or more nucleic acid templates (e.g., one or more RNA templates or messenger RNA templates) with one or more compositions of the disclosure having reverse transcriptase activity and incubating the mixture under conditions sufficient to synthesize one or more first nucleic acid molecules complementary to all or a portion of the one or more nucleic acid templates, wherein at least one of the synthesized molecules are optionally labeled and/or comprise one or more labeled nucleotides and/or wherein the synthesized molecules may optionally be modified to contain one or more labels.
- nucleic acid templates e.g., one or more RNA templates or messenger RNA templates
- compositions of the disclosure having reverse transcriptase activity
- the one or more first nucleic acid molecules are single-stranded cDNA molecules.
- Nucleic acid templates suitable for reverse transcription according to this aspect of the disclosure include any nucleic acid molecule or population of nucleic acid molecules (e.g., RNA, mRNA), particularly those derived from a cell or tissue.
- a population of mRNA molecules are used to make a labeled cDNA library, in accordance with the disclosure.
- Exemplary sources of nucleic acid templates include viruses, virally infected cells, bacterial cells, fungal cells, plant cells and animal cells.
- the disclosure also concerns methods for making one or more double-stranded nucleic acid molecules (which may optionally be labeled).
- Such methods comprise (a) mixing one or more nucleic acid templates (e.g., RNA or mRNA, or a population of mRNA templates) with one or more polypeptides of the disclosure having reverse transcriptase activity; (b) incubating the mixture under conditions sufficient to make one or more first nucleic acid molecules complementary to all or a portion of the one or more templates; and (c) incubating the one or more first nucleic acid molecules under conditions sufficient to make one or more second nucleic acid molecules complementary to all or a portion of the one or more first nucleic acid molecules, thereby forming one or more double-stranded nucleic acid molecules comprising the first and second nucleic acid molecules.
- nucleic acid templates e.g., RNA or mRNA, or a population of mRNA templates
- polypeptides of the disclosure having reverse transcriptase activity
- the first and/or second nucleic acid molecules may be labeled (e.g., may comprise one or more of the same or different labeled nucleotides and/or may be modified to contain one or more of the same or different labels).
- labeled nucleotides may be used at one or both synthesis steps.
- Such methods may include the use of one or more DNA polymerases as part of the process of making the one or more double-stranded nucleic acid molecules.
- the disclosure also concerns compositions useful for making such double-stranded nucleic acid molecules.
- Such compositions comprise one or more reverse transcriptases of the disclosure and optionally one or more DNA polymerases, a suitable buffer and/or one or more nucleotides (e.g., including labeled nucleotides).
- the disclosure is also directed to nucleic acid molecules and/or labeled nucleic acid molecules (particularly single- or double-stranded cDNA molecules) produced according to the above-described methods and to kits comprising these nucleic acid molecules. Such molecules or kits may be used to detect nucleic acid molecules (for example by hybridization) or for diagnostic purposes.
- the streptavidin fusion construct can be coexpressed with the streptavidin monomer within the same host cell from their respective expression systems, allowing the streptavidin monomer to interact with the streptavidin moiety of the fusion protein, resulting in the formation of a tetramer, in which the presence of streptavidin monomer reduces the number of streptavidin fusion constructs in the tetramer.
- a streptavidin tetramer containing three, two or one streptavidin fusion construct can be created.
- the protein multimer of the present disclosure comprises one streptavidin fusion construct of the disclosure in a tetrameric complex with three additional streptavidin monomers.
- a bacterial expression system is utilized to generate the tetrameric complex.
- the bacterial expression system is a co-expression system.
- the bacterial co-expression system comprises at least one bacterial cell comprising at least one nucleic acid molecule encoding the streptavidin fusion construct of the present disclosure and the streptavidin monomer.
- the at least one nucleic acid molecule is a plasmid.
- the streptavidin fusion construct and the streptavidin monomer are operably connected to different promoter sequences, wherein the different promoter sequences lead to different levels of expression of the streptavidin fusion construct and the streptavidin monomer, wherein the different levels of expression lead to the formation of the tetramer comprising one streptavidin fusion molecule and three streptavidin monomers.
- the streptavidin fusion construct is operably connected to the tac promoter, wherein the tac promoter comprises the sequence set forth in SEQ ID NO: 14 and the streptavidin monomer is operably connected to the araBAD promoter, wherein the araBAD promoter comprises the sequence set forth in SEQ ID NO: 15.
- the bacterial co-expression system comprises at least one bacterial cell comprising distinct nucleic acid molecules encoding the avidin or streptavidin fusion construct and the avidin or streptavidin monomer.
- the distinct nucleic acid molecules are plasmids.
- the avidin or streptavidin fusion construct and the avidin or streptavidin monomer encoded on distinct plasmids are each operably connected to the same promoter sequence.
- the promoter sequence comprises the T7 promoter, wherein the T7 promoter comprises the sequence set forth in SEQ ID NO: 16.
- the distinct plasmids encoding the avidin or streptavidin fusion construct and the avidin or streptavidin monomer are present in the at least one bacterial cell in different ratios, wherein the different ratios lead to different levels of expression of the fusion construct and the monomer, wherein the different levels of expression lead to the formation of the tetramer comprising one fusion molecule and three monomers.
- the ratio comprises one plasmid encoding the fusion construct and two plasmids encoding the monomer (1 :2). In some embodiments, the ratio comprises one plasmid encoding the fusion construct and three plasmids encoding the monomer (1 :3). In some embodiments, the ratio comprises one plasmid encoding the fusion construct and four plasmids encoding the monomer (1 :4). In some embodiments, the ratio comprises one plasmid encoding the fusion construct and five plasmids encoding the monomer (1 :5). In some embodiments, the ratio comprises one plasmid encoding the fusion construct and 10 or more plasmids encoding the monomer ( ⁇ 1: 10).
- the plasmids used are low copy number plasmids, such as pl5A ori, wherein there are approximately 10 copies of the plasmids per cell.
- the plasmids used are high copy number plasmids, such as pUC18, wherein there are approximately 300-600 copies of the plasmids per cell.
- the distinct plasmids encoding the fusion construct and the monomer are present in the at least one bacterial cell in equal ratios.
- the fusion construct and the monomer are operably connected to different promoter sequences, wherein the different promoter sequences lead to different levels of expression of the fusion construct and the monomer, wherein the different levels of expression lead to the formation of the tetramer comprising one fusion molecule and three monomers.
- the fusion construct is operably connected to the tac promoter (SEQ ID NO: 14) and the monomer is operably connected the araBAD promoter (SEQ ID NO: 15). Kits
- kits for use in the production of the compositions of the present disclosure are also directed to kits for use in the production of the compositions of the present disclosure.
- the present disclosure provides a kit to produce the compositions comprising the at least one protein multimer.
- the kit comprises at least one fusion construct of the present disclosure and/or a nucleic acid molecule encoding the same.
- the kit comprises at least one protein multimer, wherein the at least one protein multimer comprises at least one avidin or streptavidin fusion construct of the disclosure.
- the protein multimer comprises an avidin or streptavidin tetramer comprising at least one avidin or streptavidin fusion construct of the disclosure.
- the protein multimer comprises an avidin or streptavidin tetramer comprising at least two avidin or streptavidin fusion constructs of the disclosure.
- the protein multimer comprises an avidin or streptavidin tetramer comprising at least three avidin or streptavidin fusion constructs of the disclosure.
- the kit comprises an expression system that comprises one or more a polynucleotide encoding a fusion construct and a monomer of the disclosure. In one embodiment, the kit comprises an expression system that comprises a first polynucleotide encoding a fusion construct of the disclosure and a second polynucleotide encoding an avidin or streptavidin monomer.
- the kit comprises an expression system that comprises a polynucleotide comprising or encoding a nucleic acid molecule that reduces non-specific binding.
- the kit comprises an expression system that comprises a polynucleotide encoding a protein or carbohydrate (e g., heparin) that reduces non-specific binding.
- the kit includes instructional material that describes the use of the kit to produce the compositions of the present disclosure, wherein the instructional material creates an increased functional relationship between the kit components and the individual using the kit.
- the kit is utilized by one person or entity.
- the kit is utilized by more than one person or entity.
- the kit is used without any additional compositions or methods.
- the kit is used with at least one additional composition or method.
- kits for use in the methods of the disclosure can be used for making nucleic acid molecules and/or labeled nucleic acid molecules (single- or double-stranded).
- Kits of the disclosure may comprise a carrier, such as a box or carton, having in close confinement therein one or more containers, such as vials, tubes, bottles and the like.
- a first container may contain one or more of the fusion constructs of the disclosure or one or more of the compositions of the disclosure.
- Kits of the disclosure may also comprise, in the same or different containers, at least one component selected from one or more DNA polymerases (e.g., thermostable DNA polymerases), a suitable buffer for nucleic acid synthesis and one or more nucleotides.
- kits of the disclosure may also comprise, in the same or different containers, an agent that reduces nonspecific binding of primers to the surface of E.r. maturase or variant thereof.
- kits of the disclosure may also comprise, in the same or different containers, an optimized reaction buffer as described elsewhere herein, or components used to produce the optimized reaction buffer. Alternatively, the components of the kit may be divided into separate containers.
- kits for use in methods of the disclosure can be used for making, sequencing or amplifying nucleic acid molecules (single- or doublestranded), e.g., at the particular temperatures described herein.
- Kits of the disclosure may comprise a carrier, such as a box or carton, having in close confinement therein one or more (e.g., one, two, three, four, five, ten, twelve, fifteen, etc.) containers, such as vials, tubes, bottles and the like.
- a first container contains one or more of the reverse transcriptase enzymes of the present disclosure.
- Kits of the disclosure may also comprise, in the same or different containers, one or more DNA polymerases (e.g., thermostable DNA polymerases), one or more (e.g., one, two, three, four, five, ten, twelve, fifteen, etc.) suitable buffers for nucleic acid synthesis, one or more nucleotides and one or more (e.g., one, two, three, four, five, ten, twelve, fifteen, etc.) oligonucleotide primers.
- kits of the disclosure may also comprise, in the same or different containers, an agent that reduces nonspecific binding of primers to the surface of E.r. maturase or variant thereof, as described elsewhere herein.
- kits of the disclosure may also comprise, in the same or different containers, an optimized reaction buffer as described elsewhere herein, or components used to produce the optimized reaction buffer.
- the components of the kit may be divided into separate containers (e.g., one container for each enzyme and/or component).
- Kits of the disclosure also may comprise instructions or protocols for carrying out the methods of the disclosure.
- the present disclosure provides a kit to use the compositions of the present disclosure, in a reverse transcription reaction.
- the kit comprises a composition of the present disclosure.
- the kit includes instructional material that describes the use of the kit to use the compositions of the present disclosure in a reverse transcription reaction, wherein the instructional material creates an increased functional relationship between the kit components and the individual using the kit.
- the kit is utilized by one person or entity.
- the kit is utilized by more than one person or entity.
- the kit is used without any additional compositions or methods.
- the kit is used with at least one additional composition or method.
- a fusion construct comprising an affinity domain operably linked to a polymerase domain.
- polymerase domain comprises a DNA polymerase, an RNA polymerase, or a reverse transcriptase.
- polymerase domain comprises a reverse transcriptase, e.g., a retroviral reverse transcriptase or a group II intron reverse transcriptase.
- polymerase domain comprises a group II intron reverse transcriptase, e.g., a MarathonRT or UltraMarathonRT reverse transcriptase, or a fragment or variant thereof.
- polymerase domain comprises MarathonRT or a fragment or variant thereof.
- polymerase domain comprises SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22, SEQ ID NO: 23, SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, or SEQ ID NO: 29, or a fragment or variant thereof.
- polymerase domain comprises SEQ ID NO: 3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, or SEQ ID NO: 12 or a fragment or variant thereof.
- a fusion construct comprising avidin or streptavidin operably linked to a functional protein or enzyme domain, wherein the avidin or streptavidin can form a tetramer with additional avidin or streptavidin molecules or additional avidin or streptavidin fusion enzymes.
- the functional protein or enzyme domain comprises a polymerase, reverse transcriptase, helicase, receptor protein, ligase, kinase, phosphatase, exonuclease and non-specific endonuclease, restriction endonuclease, DNA/RNA methyl transferase, recombinase, terminal transferase, nucleic acid binding protein, protease, ribosomal protein, aminoacyl-tRNA synthetase, glycosyltransferase, fatty acid synthase, or an enzyme for food, biofuel or pharmaceutical industries.
- the functional protein or enzyme domain comprises Marathon Reverse Transcriptase (MRT), or a variant thereof.
- a nucleic acid molecule encoding a fusion construct comprising avidin or streptavidin operably linked to a functional protein or enzyme domain, wherein the streptavidin can form a tetramer with additional streptavidin molecules or additional streptavidin fusion enzymes.
- nucleic acid molecule of any of embodiments 15-16 wherein the functional protein or enzyme domain comprises a polymerase, reverse transcriptase, helicase, receptor protein, ligase, kinase, phosphatase, exonuclease and non-specific endonuclease, restriction endonuclease, DNA/RNA methyl transferase, recombinase, terminal transferase, nucleic acid binding protein, protease, ribosomal protein, aminoacyl-tRNA synthetase, glycosyltransferase, fatty acid synthase, or an enzyme for food, biofuel or pharmaceutical industries.
- the functional protein or enzyme domain comprises a polymerase, reverse transcriptase, helicase, receptor protein, ligase, kinase, phosphatase, exonuclease and non-specific endonuclease, restriction endonuclease, DNA/RNA methyl transferase
- nucleic acid molecule of any of embodiments 15-17, wherein the functional enzyme domain comprises MRT, or a variant thereof.
- functional enzyme domain comprises an amino acid sequence at least 90% identical to an amino acid sequence selected from the group consisting SEQ ID NOs: 3 and 8-12 or a functional fragment thereof.
- nucleic acid molecule of any of embodiments 15-20 comprising the nucleotide sequence set forth in SEQ ID NO: 17.
- composition comprising at least one fusion construct of embodiment 1, and further wherein the functional enzyme domain maintains at least one wild type activity of the enzyme when the avidin or streptavidin is complexed as a tetramer.
- composition of embodiment 22, comprising a tetramer comprising at least one avidin or streptavidin fusion construct of embodiment 1.
- composition of embodiments 22-24 comprising a tetramer comprising at least three avidin or streptavidin fusion constructs of embodiment 1.
- composition of embodiments 22-26, wherein the solid support is coated with or attached to at least one biotin molecule.
- [0298] 28. The composition of embodiments 22-27, wherein the solid support is comprised in at least one device.
- composition of embodiments 22-28, wherein the at least one device comprises a device used for complementary DNA (cDNA) synthesis.
- cDNA complementary DNA
- a method for producing cDNA comprising contacting at least one mRNA molecule with the fusion construct of any one of embodiments 9-14, or a composition of any one of embodiments 22-30, under conditions sufficient for the synthesis of at least one nucleic acid molecule complementary to all or a portion of the at least one mRNA molecule.
- a method for determining the identity of each of a series of consecutive nucleotide residues of at least one target nucleic acid comprising contacting the at least one target nucleic acid with the fusion construct of any one of embodiments 9-14, or a composition of any one of embodiments 22-30, under conditions sufficient for the synthesis of at least one nucleic acid molecule complementary to all or a portion of the at least one target nucleic acid, and for the determination of each nucleotide in the at least one complementary nucleic acid.
- Example 1 Production of catalytically functional, affinity tagged-reverse-transcriptase fusion proteins
- Streptavidin is one of the most important protein conjugation reagents, as it facilitates bridging of molecules and anchoring to biotinylated surfaces or nanoparticles.
- its tetrameric structure makes it impractical for creating single-enzyme fusion constructs.
- chemically-conjugated streptavidin proteins are more commonly employed for affinity tagging of proteins with streptavidin.
- MRT (or any other RT) was functionalized at its N-terminus with another protein (such as an affinity tag like streptavidin) while still maintaining its full enzymatic activities in terms of processive, high speed cDNA synthesis, RNA unwinding and dNTP hydrolysis ( Figure 3- Figure 5).
- Another protein such as an affinity tag like streptavidin
- Example 2 Streptavidin-MarathonRT fusion protein is used in devices for direct RNA and DNA sequencing
- streptavidin-MarathonRT fusion protein is used in devices for direct RNA and DNA sequencing, resulting in new types of commercial products.
- affinity-tagged RT proteins are generated more cheaply because they can be created as overexpressed single-chain proteins instead of chemically-conjugated proteins.
- RTs mounted on a solid support in an industrial setting are re-used and recycled, which has great economic benefit.
Landscapes
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Health & Medical Sciences (AREA)
- Organic Chemistry (AREA)
- Wood Science & Technology (AREA)
- Engineering & Computer Science (AREA)
- Zoology (AREA)
- Genetics & Genomics (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Molecular Biology (AREA)
- General Health & Medical Sciences (AREA)
- Biochemistry (AREA)
- General Engineering & Computer Science (AREA)
- Microbiology (AREA)
- Biotechnology (AREA)
- Medicinal Chemistry (AREA)
- Biomedical Technology (AREA)
- General Chemical & Material Sciences (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Enzymes And Modification Thereof (AREA)
- Peptides Or Proteins (AREA)
Abstract
Provided are compositions comprising avidin or streptavidin fusion constructs comprising a monomer of avidin or streptavidin linked to an enzymatic domain, wherein the enzymatic domain maintains wild type catalytic activity when the avidin or streptavidin forms a tetrameric structure, as well as method of use thereof.
Description
FUSION-MODIFIED ENZYME CONSTRUCTS
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application claims priority to U.S. Provisional Patent Application No. 63/599,119, filed November 15, 2023, which is hereby incorporated herein by reference in its entirety.
STATEMENT REGARDING FEDERALLY SPONSORED RESEARCH OR DEVELOPMENT
[0002] This disclosure was made with government support under HG011087 awarded by the National Institutes of Health. The government has certain rights in the disclosure.
REFERENCE TO A "SEQUENCE LISTING" SUBMITTED AS AN XML FILE
[0003] The Sequence Listing written in the XML file: “047162-5356- OOWO_Sequencelisting.xml”; created on November 14, 2024, and 37,721 bytes in size, is hereby incorporated by reference.
BACKGROUND OF THE INVENTION
[0004] Streptavidin is one of the most important protein conjugation reagents, as it facilitates bridging of molecules and anchoring to biotinylated surfaces or nanoparticles. However, its tetrameric structure makes it impractical for creating single-enzyme fusion constructs. When streptavidin fusion proteins form tetramers, the four proteins of interest fused to streptavidin become compacted within a confined space, which leads to steric hindrance, and reduction in the activity of attached proteins. As a result, post-expression, chemically-conjugated streptavidin proteins are commonly employed for affinity tagging of proteins with streptavidin. However, streptavidin fusion proteins have the potential to offer several advantages over conjugates, including greater homogeneity, ease of scale-up, and lower production costs.
[0005] Thus, there is a need in the art for improved compositions and methods for fusing enzymes to streptavidin while maintaining the wild type catalytic activity of the enzymes. This disclosure satisfies this unmet need.
SUMMARY OF THE INVENTION
[0006] The present disclosure features a bifunctional fusion construct comprising an affinity domain operably linked to an enzyme domain, for example, a polymerase domain. In some embodiments, the affinity domain comprises avidin or streptavidin.
[0007] In one embodiment, the present disclosure relates to a fusion construct comprising avidin or streptavidin operably linked to a functional protein or enzyme domain, wherein the avidin or streptavidin can form a tetramer with additional avidin or streptavidin molecules or additional avidin or streptavidin fusion enzymes.
[0008] In one embodiment, the fusion construct comprises streptavidin comprising an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO: 1.
[0009] In one embodiment, the functional protein or enzyme domain comprises a polymerase or reverse transcriptase.
[0010] In one embodiment, the functional protein or enzyme domain comprises Marathon Reverse Transcriptase (MRT), or a variant thereof.
[0011] In one embodiment, the functional protein or enzyme domain comprises an amino acid sequence at least 90% identical to an amino acid sequence selected from the group consisting SEQ ID NOs: 3 and 8-12 or a functional fragment thereof.
[0012] In one embodiment, the fusion construct comprises an amino acid sequence as set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0013] In one embodiment, the disclosure relates to a nucleic acid molecule encoding a fusion construct comprising avidin or streptavidin operably linked to a functional protein or enzyme domain, wherein the streptavidin can form a tetramer with additional streptavidin molecules or additional streptavidin fusion enzymes.
[0014] In one embodiment, the nucleic acid molecule encodes a streptavidin fusion construct, wherein the streptavidin comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO:1
[0015] In one embodiment, the functional protein or enzyme domain comprises a polymerase, reverse transcriptase, helicase, ligase, receptor protein, kinase, phosphatase, exonuclease and non-specific endonuclease, restriction endonuclease, DNA/RNA methyl transferase, recombinase, terminal transferase, nucleic acid binding protein, protease, ribosomal
protein, aminoacyl-tRNA synthetase, glycosyltransferase, fatty acid synthase, or an enzyme for food, biofuel or pharmaceutical industries.
[0016] In one embodiment, the functional protein or enzyme domain comprises MRT, or a variant thereof.
[0017] In one embodiment, the functional protein or enzyme domain comprises an amino acid sequence at least 90% identical to an amino acid sequence selected from the group consisting SEQ ID NOs: 3 and 8-12 or a functional fragment thereof.
[0018] In one embodiment, the streptavidin fusion enzyme comprises an amino acid sequence as set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0019] In one embodiment, the nucleic acid molecule comprises the nucleotide sequence set forth in SEQ ID NO: 17.
[0020] In one embodiment, the disclosure relates to a composition comprising at least one fusion construct, and further wherein the functional protein or enzyme domain maintains at least one wild type activity or function of the protein or enzyme when the avidin or streptavidin is complexed as a tetramer.
[0021] In one embodiment, the composition comprises a tetramer comprising at least one avidin or streptavidin fusion construct.
[0022] In one embodiment, the composition comprises a tetramer comprising at least two avidin or streptavidin fusion constructs.
[0023] In one embodiment, the composition comprises a tetramer comprising at least three avidin or streptavidin fusion constructs.
[0024] In one embodiment, the tetramer is linked or bound to a solid support.
[0025] In one embodiment, the solid support is coated with or attached to at least one biotin molecule.
[0026] In one embodiment, the solid support is comprised in at least one device.
[0027] In one embodiment, the at least one device comprises a device used for complementary DNA (cDNA) synthesis.
[0028] In one embodiment, the at least one device is used for nucleic acid synthesis or sequencing.
[0029] In one embodiment, the present disclosure relates to a method for producing a nucleic acid comprising contacting a first nucleic acid molecule with the fusion construct or a
composition of the present disclosure under conditions sufficient for the synthesis of a second nucleic acid molecule complementary to all or a portion of the first nucleic acid molecule.
[0030] In one embodiment, the present disclosure relates to a method for producing an RNA comprising contacting a first RNA molecule with the fusion construct or a composition of the present disclosure under conditions sufficient for the synthesis of a second RNA molecule complementary to all or a portion of the first RNA molecule.
[0031] In one embodiment, the present disclosure relates to a method for producing cDNA, comprising contacting at least one mRNA molecule with the fusion construct or a composition of the present disclosure, under conditions sufficient for the synthesis of at least one nucleic acid molecule complementary to all or a portion of the at least one mRNA molecule.
[0032] In one embodiment, the disclosure relates to a method for determining the identity of each of a series of consecutive nucleotide residues of at least one target nucleic acid, comprising contacting the at least one target nucleic acid with the fusion construct or a composition of the present disclosure, under conditions sufficient for the synthesis of at least one nucleic acid molecule complementary to all or a portion of the at least one target nucleic acid, and for the determination of each nucleotide in the at least one complementary nucleic acid.
BRIEF DESCRIPTION OF THE DRAWINGS
[0033] The following detailed description of embodiments of the disclosure will be better understood when read in conjunction with the appended drawings. It should be understood that the disclosure is not limited to the precise arrangements and instrumentalities of the embodiments shown in the drawings.
[0034] Figure 1 comprising Figure 1 A and Figure IB, depicts an illustration of producing streptavidin-MarathonRT fusion protein tetramer containing reduced number of MarathonRT domains. A representative fusion protein is shown comprising streptavidin, STV (SEQ ID NO: 1) and Marathon Reverse Transcriptase, MRT, (SEQ ID NO:3 linked by a 15-amino acid flexible linker, FL15, (SEQ ID NO:2) to produce STV-FL15-MRT (SEQ ID NO:4) which was assembled into a tetramer comprising one molecule of STV-FL15-MRT and three molecules of STV, thus containing a reduced number of MRT domains. Figure 1A depicts a representative illustration of the co-expression of STV protein (SEQ ID NO: 1) and STV-FL15-MRT fusion protein (SEQ ID NO:4). STV-FL15-MRT fusion protein (SEQ ID NO:4) comprises STV (SEQ ID NO:1)
connected to MRT (SEQ ID N0:3) via a flexible linker of 15-amino acids (SEQ ID NO:2). Figure IB depicts a representative illustration of STV-FL15-MRT fusion protein (SEQ ID NO:4) tetramer containing reduced number of MRT domains. The expression level of STV (SEQ ID NO:1) and STV-FL15-MRT fusion protein (SEQ ID NO:4) can be individually adjusted.
[0035] Figure 2 depicts the amino acid sequence of streptavidin-MarathonRT fusion protein containing a 15-amino acid linker (SEQ ID NO: 4). Shown is a representative amino acid sequence of STV connected to MRT via FL15 (SEQ ID NO:4). Underlined sequence: core sequence of STV 14-138 (STV) with enhanced solubility and full biotin binding activity (Sano et al., 1995, JBC, 270: 28204-28209) (SEQ ID NO: 1). Italicized sequence: FL15 (SEQ ID NO:2). Non-underlined, non-italicized sequence: wild-type MRT (SEQ ID NO:3).
[0036] Figure 3 depicts long cDNA synthesis using streptavidin tetramer containing two (estimated) MarathonRT domains. Shown is representative data demonstrating cDNA synthesis from two RNA templates (left: HOTAIR; right: RepA D3) using wild type MRT (SEQ ID NO:3) or STV-FL15-MRT (SEQ ID NO:4) tetramer containing two (estimated) MRT domains.
[0037] Figure 4 depicts tetrameric streptavidin-MarathonRT fusion proteins with a single or multiple MarathonRT molecules with a flexible 15-amino acid linker. Shown are representative illustrations of linear peptide and 3D STV- FL15-MRT fusion protein (SEQ ID NO:4). Left: linear peptide; middle tetrameric STV-FL15-MRT; right: a streptavidin tetramer with a STV-FL15-MRT monomer.
[0038] Figure 5 depicts primer extension assays using STV-MRT fusion protein (with a 15-amino acid linker) containing four or one (estimated) MarathonRT domains. Shown are representative primer extension assay data demonstrating that the long, flexible linker, FL 15, (SEQ ID NO:2) between STV (SEQ ID NO: 1) and MRT (SEQ ID NO:3) greatly improves the enzyme activity of the fusion protein (SEQ ID NO:4); reducing the number of MRT domains further improves the enzyme activity. MRT (lane 1) (SEQ ID NO:3); STV-FL15-MRT fusion protein (SEQ ID NO:4) containing four (estimated) MRT domains (lane 2); STV-FL15-MRT fusion protein (SEQ ID NO:4) containing one (estimated) MRT domains (lane 3).
[0039] Figure 6 depicts the amino acid sequence of streptavidin-MarathonRT fusion protein without a linker or with a 3 -amino acid linker. Shown are representative amino acid sequences of STV-MRT fusion protein without a linker (SEQ ID NO:6), top, or with a 3-amino acid linker, FL3, (SEQ ID NO:5), bottom STV-FL3-MRT (SEQ ID NO:7). Underlined
sequences: core sequence of STV 14-138 with enhanced solubility and full biotin binding activity (Sano et al., 1995, JBC, 270: 28204-28209) (SEQ ID NO:1). Italicized sequence: 3- amino acid flexible linker, FL3, between (SEQ ID NO:5). Non-underlined, non-italicized sequence: wild-type MRT (SEQ ID NO:3).
[0040] Figure 7 depicts tetrameric streptavidin-MarathonRT fusion protein without or with a 3-amino acid linker, Shown are representative illustrations (linear peptide and 3D protein tetramer) of tetrameric STV-MRT fusion protein without (SEQ ID NO:6) or with (SEQ ID NO:7) a 3-amino acid linker (SEQ ID NO:5).
[0041] Figure 8 depicts representative primer extension assay data demonstrating that the tetrameric STV-MRT fusion protein without a linker (SEQ ID NO:6) (lane 2) or STV-MRT with a short linker (lane 3) (SEQ ID NO:7) is nearly inactive compared to WT-MRT (lane 1) (SEQ ID NO:3).
DETAILED DESCRIPTION
[0042] The present disclosure is based, in part, on the development of a bifunctional fusion construct comprising an affinity domain linked to an enzyme domain, for example, a polymerase domain. In one aspect, the present disclosure features a fusion molecule comprising an enzyme fused to a monomer of avidin or streptavidin, which when expressed and combined with additional monomeric avidin or streptavidin in the correct proportions, yields an active avidin or streptavidin tetramer, in which the fused enzyme maintains at least one enzymatic activity, such as processivity, high speed cDNA synthesis, RNA unwinding, or dNTP hydrolysis.
[0043] Accordingly, in some embodiments, the present disclosure provides fusion constructs comprising an affinity domain fused to least one enzymatic domain. In some embodiments, the affinity domain is capable of binding a target, e g., a substrate. In some embodiments, the affinity domain comprises a protein or a protein fragment, e.g., avidin or streptavidin. In some embodiments, the affinity domain comprises avidin or streptavidin tetramers comprising at least one avidin or streptavidin fusion construct comprising an avidin or streptavidin monomer fused to least one enzymatic domain. In some embodiments, the monomeric avidin or streptavidin domain of the fusion construct retains the ability to bind to additional avidin or streptavidin monomers to form a tetramer. In one embodiment, the
streptavidin fusion construct comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO: 1.
[0044] In some embodiments, the avidin or streptavidin fusion construct comprises an enzymatic or catalytic domain, wherein the enzymatic or catalytic domain retains at least one wild type enzymatic or catalytic activity when the fused avidin or streptavidin domain is bound to additional avidin or streptavidin monomers to form a tetramer. Exemplary enzymes that can be fused to the monomeric avidin or streptavidin in the fusion construct include, but are not limited to, transcriptases, polymerases, or helicase motor enzymes, receptor proteins, ligases, kinases, phosphatases, exonucleases and non-specific endonucleases, restriction endonucleases, DNA/RNA methyl transferases, recombinases, terminal transferases, nucleic acid binding proteins, proteases, ribosomal proteins, aminoacyl-tRNA synthetases, glycosyltransferase, fatty acid synthases and enzymes for food, biofuel and pharmaceutical industries, such as palatase, cellulase and lipase. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r. maturase also known as Marathon Reverse Transcriptase (MRT), or a variant thereof, wherein the E.r. maturase or variant thereof comprises an amino acid sequence at least 90% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NOTO, SEQ ID NOT 1, SEQ ID NO: 12 or SEQ ID NOT 3, or a functional fragment thereof.
[0045] In one embodiment, the monomeric avidin or streptavidin is linked to the N- terminus of the enzymatic domain. In one embodiment, the monomeric streptavidin is linked to the N-terminus of the enzymatic domain via a linker sequence. In some embodiments, the linker is a flexible linker. In one embodiment, the linker comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO:2.
[0046] In one embodiment, the monomeric avidin or streptavidin is linked to the C- terminus of the enzymatic domain. In one embodiment, the monomeric streptavidin is linked to the C-terminus of the enzymatic domain via a linker sequence. In some embodiments, the linker is a flexible linker.
[0047] In some embodiments, the disclosure provides compositions comprising at least one avidin or streptavidin fusion construct. In some embodiments, the disclosure provides a avidin or streptavidin tetramer comprising at least one avidin or streptavidin fusion construct. In one embodiment, the avidin or streptavidin tetramer comprises at least two avidin or streptavidin
fusion constructs. In one embodiment, the avidin or streptavidin tetramer comprises at least three avidin or streptavidin fusion constructs.
[0048] In some embodiments, the disclosure includes methods of use of the avidin or streptavidin fusion constructs or avidin or streptavidin tetramers comprising the avidin or streptavidin fusion construct of the disclosure. In some embodiments, the avidin or streptavidin fusion constructs or avidin or streptavidin tetramers of the disclosure are attached to a solid support. In some embodiments, the solid support is coated with biotin.
[0049] In some embodiments, the disclosure includes methods of using the avidin or streptavidin fusion constructs or avidin or streptavidin tetramers or a solid support or device comprising the avidin or streptavidin fusion constructs or avidin or streptavidin tetramers in molecular applications. In some embodiments, the molecular applications nucleic acid sequencing applications, reverse transcription applications, nucleic acid synthesis applications, or in any molecular application that requires the catalytic activity of the enzymatic or catalytic domain of a avidin or streptavidin fusion construct of the present disclosure.
Definitions
[0050] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this disclosure belongs. It is also to be understood that the terminology used herein is for the purpose of describing particular embodiments only, and is not intended to be limiting.
[0051] As used herein, each of the following terms has the meaning associated with it in this section.
[0052] The articles “a” and “an” are used herein to refer to one or to more than one (i.e., to at least one) of the grammatical object of the article. By way of example, “an element” means one element or more than one element.
[0053] “About” as used herein when referring to a measurable value such as an amount, a temporal duration, and the like, is meant to encompass variations of ±20%, ±10%, ±5%, ±1%, or ±0.1% from the specified value, as such variations are appropriate to perform the disclosed methods.
[0054] As used herein, “autologous” refers to a biological material derived from the same individual into whom the material will later be re-introduced.
[0055] As used herein, “allogeneic” refers to a biological material derived from a genetically different individual of the same species as the individual into whom the material will be introduced.
[0056] The terms “cells” and “population of cells” are used interchangeably and generally refer to a plurality of cells, i.e., more than one cell. The population may be a pure population comprising one cell type. Alternatively, the population may comprise more than one cell type. In the present disclosure, there is no limit on the number of cell types that a cell population may comprise.
[0057] “Encoding” refers to the inherent property of specific sequences of nucleotides in a polynucleotide, such as a gene, a DNA, or an RNA, to serve as templates for synthesis of other polymers and macromolecules in biological processes having either a defined sequence of nucleotides (i.e., rRNA, tRNA and mRNA) or a defined sequence of amino acids and the biological properties resulting therefrom. Thus, a gene encodes a protein if transcription and translation of mRNA corresponding to that gene produces the protein in a cell or other biological system. Both the coding strand, the nucleotide sequence of which is identical to the mRNA sequence and is usually provided in sequence listings, and the non-coding strand, used as the template for transcription of a gene or cDNA, can be referred to as encoding the protein or other product of that gene or cDNA.
[0058] “Expression vector” refers to a vector comprising a recombinant polynucleotide comprising expression control sequences operatively linked to a nucleotide sequence to be expressed. An expression vector comprises sufficient cis-acting elements for expression; other elements for expression can be supplied by the host cell or in an in vitro expression system. Expression vectors include all those known in the art, such as cosmids, plasmids (e.g., naked or contained in liposomes) and viruses (e g., lentiviruses, retroviruses, adenoviruses, and adeno- associated viruses) that incorporate the recombinant polynucleotide.
[0059] “Homologous” refers to the sequence similarity or sequence identity between two polypeptides or between two nucleic acid molecules. When a position in both of the two compared sequences is occupied by the same base or amino acid monomer subunit, e.g., if a position in each of two DNA molecules is occupied by adenine, then the molecules are homologous at that position. The percent of homology between two sequences is a function of the number of matching or homologous positions shared by the two sequences divided by the
number of positions compared X 100. For example, if 6 of 10 of the positions in two sequences are matched or homologous then the two sequences are 60% homologous. By way of example, the DNA sequences ATTGCC and TATGGC share 50% homology. Generally, a comparison is made when two sequences are aligned to give maximum homology.
[0060] “Isolated” means altered or removed from the natural state. For example, a nucleic acid or a peptide naturally present in a living organism is not “isolated,” but the same nucleic acid or peptide partially or completely separated from the coexisting materials of its natural state is “isolated.” An isolated nucleic acid or protein can exist in substantially purified form, or can exist in a non-native environment such as, for example, a host cell.
[0061] In the context of the present disclosure, the following abbreviations for the commonly occurring nucleic acid bases are used. “A” refers to adenosine, “C” refers to cytosine, “G” refers to guanosine, “T” refers to thymidine, and “U” refers to uridine.
[0062] Unless otherwise specified, a “nucleotide sequence encoding an amino acid sequence” includes all nucleotide sequences that are degenerate versions of each other and that encode the same amino acid sequence. The phrase nucleotide sequence that encodes a protein or an RNA may also include introns to the extent that the nucleotide sequence encoding the protein may in some versions contain an intron(s).
[0063] The term “polynucleotide” as used herein is defined as a chain of nucleotides. Furthermore, nucleic acids are polymers of nucleotides. Thus, nucleic acids and polynucleotides as used herein are interchangeable. One skilled in the art has the general knowledge that nucleic acids are polynucleotides, which can be hydrolyzed into the monomeric “nucleotides.” The monomeric nucleotides can be hydrolyzed into nucleosides. As used herein polynucleotides include, but are not limited to, all nucleic acid sequences which are obtained by any means available in the art, including, without limitation, recombinant means, i.e., the cloning of nucleic acid sequences from a recombinant library or a cell genome, using ordinary cloning technology and PCR, and the like, and by synthetic means.
[0064] As used herein, the terms “peptide,” “polypeptide,” and “protein” are used interchangeably, and refer to a compound comprised of amino acid residues covalently linked by peptide bonds. A protein or peptide must contain at least two amino acids, and no limitation is placed on the maximum number of amino acids that can comprise the sequence of a protein or peptide. Polypeptides include any peptide or protein comprising two or more amino acids joined
to each other by peptide bonds. As used herein, the term refers to both short chains, which also commonly are referred to in the art as peptides, oligopeptides and oligomers, for example, and to longer chains, which generally are referred to in the art as proteins, of which there are many types. “Polypeptides” include, for example, biologically active fragments, substantially homologous polypeptides, oligopeptides, homodimers, heterodimers, variants of polypeptides, modified polypeptides, derivatives, analogs, fusion proteins, among others. The polypeptides include natural peptides, recombinant peptides, synthetic peptides, or a combination thereof.
[0065] The term “promoter” as used herein is defined as a DNA sequence recognized by the synthetic machinery of the cell, or introduced synthetic machinery, required to initiate the specific transcription of a polynucleotide sequence.
[0066] As used herein, the term “promoter/regulatory sequence” means a nucleic acid sequence which is required for expression of a gene product operably linked to the promoter/regulatory sequence. In some instances, this sequence may be the core promoter sequence and in other instances, this sequence may also include an enhancer sequence and other regulatory elements which are required for expression of the gene product. The promoter/regulatory sequence may, for example, be one which expresses the gene product in a conditional manner.
[0067] A “constitutive” promoter is a nucleotide sequence which, when operably linked with a polynucleotide which encodes or specifies a gene product, causes the gene product to be produced in a cell under most or all physiological conditions of the cell.
[0068] An “inducible” promoter is a nucleotide sequence which, when operably linked with a polynucleotide which encodes or specifies a gene product, causes the gene product to be produced in a cell substantially only when an inducer which corresponds to the promoter is present in the cell.
[0069] A “vector” is a composition of matter which comprises an isolated nucleic acid and which can be used to deliver the isolated nucleic acid to the interior of a cell. Numerous vectors are known in the art including, but not limited to, linear polynucleotides, polynucleotides associated with ionic or amphiphilic compounds, plasmids, and viruses. Thus, the term “vector” includes an autonomously replicating plasmid or a virus. The term should also be construed to include non-plasmid and non-viral compounds which facilitate transfer of nucleic acid into cells, such as, for example, polylysine compounds, liposomes, and the like. Examples of viral vectors
include, but are not limited to, adenoviral vectors, adeno-associated virus vectors, retroviral vectors, and the like.
[0070] “Wild type activity” as used herein refers to the activity (e.g., catalytic, structural, etc.) displayed by a biological molecule as it occurs in nature. As used herein, “wild type activity” refers to any documented, or yet undocumented, biological activity of a molecule as determined by techniques or assays commonly used in the field to determine the function of a molecule (e.g., catalytic, structural, etc.).
[0071] Ranges: throughout this disclosure, various aspects of the disclosure can be presented in a range format. It should be understood that the description in range format is merely for convenience and brevity and should not be construed as an inflexible limitation on the scope of the disclosure. Accordingly, the description of a range should be considered to have specifically disclosed all the possible subranges as well as individual numerical values within that range. For example, description of a range such as from 1 to 6 should be considered to have specifically disclosed subranges such as from 1 to 3, from 1 to 4, from 1 to 5, from 2 to 4, from 2 to 6, from 3 to 6 etc., as well as individual numbers within that range, for example, 1, 2, 2.7, 3, 4, 5, 5.3, and 6. This applies regardless of the breadth of the range.
Description
[0072] In some embodiments, the present disclosure provides streptavidin fusion constructs comprising a streptavidin monomer fused to least one enzymatic domain as well as streptavidin tetramers comprising at least one streptavidin fusion construct comprising a streptavidin monomer fused to least one enzymatic domain. In one embodiment, the streptavidin fusion construct comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO: 1. In some embodiments, the monomeric streptavidin domain of the streptavidin fusion construct retains the ability to bind to additional streptavidin monomers to form a streptavidin tetramer.
[0073] In some embodiments, the streptavidin fusion construct comprises an enzymatic or catalytic domain, wherein the enzymatic or catalytic domain retains at least one wild type enzymatic or catalytic activity when the fused streptavidin domain is bound to additional streptavidin monomers to form a streptavidin tetramer. Exemplary enzymes that can be fused to the monomeric streptavidin in the streptavidin fusion construct include, but are not limited to,
transcriptases, polymerases, helicase motor enzymes, receptor proteins, ligases, kinases, phosphatases, exonucleases and non-specific endonucleases, restriction endonucleases, DNA/RNA methyl transferases, recombinases, terminal transferases, nucleic acid binding proteins, proteases, ribosomal proteins, aminoacyl-tRNA synthetases, glycosyltransferase, fatty acid synthases and enzymes for food, biofuel and pharmaceutical industries, such as palatase, cellulase and lipase. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterinm rectale (E.r.) maturase also known as Marathon Reverse Transcriptase (MRT), or a variant thereof, wherein the E.r. maturase or variant thereof comprises an amino acid sequence at least 90% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO: 9, SEQ ID NOTO, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
[0074] In one embodiment, the monomeric streptavidin is linked to the N-terminus of the enzymatic domain. In one embodiment, the monomeric streptavidin is linked to the N-terminus of the enzymatic domain via a linker sequence. In one embodiment, the linker comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO:2.
[0075] In some embodiments, the disclosure provides compositions comprising at least one streptavidin fusion construct. In some embodiments, the disclosure provides a streptavidin tetramer comprising at least one streptavidin fusion construct. In one embodiment, the streptavidin tetramer comprises at least two streptavidin fusion constructs. In one embodiment, the streptavidin tetramer comprises at least three streptavidin fusion constructs.
[0076] In some embodiments, the disclosure includes methods of use of the streptavidin fusion constructs or streptavidin tetramers comprising the streptavidin fusion construct of the disclosure. In some embodiments, the streptavidin fusion constructs or streptavidin tetramers of the disclosure are attached to a solid support. In some embodiments, the solid support is coated with biotin.
[0077] In some embodiments, the disclosure includes methods of using the streptavidin fusion constructs or streptavidin tetramers or a solid support or device comprising the streptavidin fusion constructs or streptavidin tetramers in molecular applications. In some embodiments, the molecular applications nucleic acid sequencing applications, reverse transcription applications, nucleic acid synthesis applications, or in any molecular application
that requires the catalytic activity of the enzymatic or catalytic domain of a streptavidin fusion construct of the present disclosure.
Compositions
[0078] The present disclosure provides compositions comprising at least one avidin or streptavidin domain fusion construct comprising a avidin or streptavidin monomer linked to an enzymatic domain in a manner such that the monomer domain retains the ability to form a tetramer with additional avidin or streptavidin monomers while the enzymatic domain retains at least one enzymatic function.
[0079] The present disclosure also provides compositions comprising at least one avidin or streptavidin fusion construct, wherein the fusion construct is bound to or in a complex with at least one additional avidin or streptavidin fusion construct or avidin or streptavidin monomer to form a tetramer comprising at least one avidin or streptavidin fusion construct.
Fusion Construct
Enzymatic domain
[0080] In some embodiments, the avidin or streptavidin domain fusion construct comprises an enzymatic or catalytic domain, wherein the enzymatic or catalytic domain retains at least one wild type enzymatic or catalytic activity when the fused streptavidin domain is bound to additional streptavidin monomers to form a streptavidin tetramer. Exemplary proteins or enzymes that can be fused to the monomeric streptavidin in the avidin or streptavidin domain fusion construct include, but are not limited to, transcriptases, polymerases, helicase motor enzymes, receptor proteins, ligases, kinases, phosphatases, exonucleases and non-specific endonucleases, restriction endonucleases, DNA/RNA methyl transferases, recombinases, terminal transferases, nucleic acid binding proteins, proteases, ribosomal proteins, aminoacyl- tRNA synthetases, glycosyltransferase, fatty acid synthases and enzymes for food, biofuel and pharmaceutical industries. Exemplary enzymes that can be incorporated into the avidin or streptavidin domain fusion construct of the disclosure include, but are not limited to, reverse transcriptases, translocating helicase enzymes, DNA polymerases, and RNA , ligases, kinases, phosphatases, exonucleases and non-specific endonucleases, restriction endonucleases, DNA/RNA methyl transferases, recombinases, terminal transferases, nucleic acid binding
proteins, proteases, ribosomal proteins, aminoacyl-tRNA synthetases, glycosyltransferase, fatty acid synthases and enzymes for food, biofuel and pharmaceutical industries, such as palatase, cellulase and lipase.
[0081] In some embodiments, the enzymatic domain comprises a polymerase, e.g., a DNA or RNA polymerase. In one embodiment, the polymerase is a DNA polymerase, e.g., a reverse transcriptase. In one embodiment, the polymerase is a reverse transcriptase. In some embodiments, the reverse transcriptase is derived from a virus. For example, the reverse transcriptase may be derived from a retrovirus. In some embodiments, the reverse transcriptase is an Avian Myeloblastosis Virus (AMV) reverse transcriptase. In some embodiments, the reverse transcriptase is a Human Immunodeficiency Virus reverse transcriptase. In some embodiments, the reverse transcriptase is a Rous Sarcoma Virus reverse transcriptase. In some embodiments, the reverse transcriptase is a Moloney Murine Leukemia Virus (MMLV) reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is Maxima H Minus reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is SuperScript II reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is SuperScript III reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is SuperScript IV reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is PrimeScript reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is GoScript reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is ProtoScript II reverse transcriptase. In some embodiments, the MMLV reverse transcriptase is SMARTScribe reverse transcriptase. In one embodiment, the MMLV reverse transcriptase comprises SEQ ID NO: 22 or SEQ ID NO: 23, or a fragment or variant thereof. In one embodiment, the AMV reverse transcriptase comprises SEQ ID NO: 26 or SEQ ID NO: 27, or a fragment or variant thereof.
[0082] In some embodiments, the reverse transcriptase is derived from a mobile genetic element. For example, the reverse transcriptase may be derived from a telomerase, e.g., a mammalian telomerase. In one embodiment, the reverse transcriptase is derived from a human telomerase, e.g., TERT. In one embodiment, the reverse transcriptase comprises SEQ ID NO: 28 or SEQ ID NO: 29, or a fragment or variant thereof. As another example, the reverse transcriptase may be derived from a non-long terminal repeat (non-LTR) retrotransposon or a group II intron. In one embodiment, the reverse transcriptase is a non-LTR retrotransposon reverse transcriptase. In one embodiment, the non-LTR retrotransposon reverse transcriptase is a
Bombyx mori R2 RNA element reverse transcriptase. In one embodiment, the Bombyx mori R2 RNA element reverse transcriptase comprises SEQ ID NO: 24 or SEQ ID NO: 25, or a fragment or variant thereof. In one embodiment, the non-LTR retrotransposon reverse transcriptase is a human LI element reverse transcriptase. In one embodiment, the reverse transcriptase is a group II intron reverse transcriptase, e.g., a maturase reverse transcriptase. In one embodiment, the group II intron reverse transcriptase is a maturase. In one embodiment, the group II intron reverse transcriptase is a maturase encoded by Eubacterium rectale. In one embodiment, the group II intron reverse transcriptase is a maturase encoded by Roseburia intestinalis. In one embodiment, the Roseburia intestinalis reverse transcriptase comprises SEQ ID NO: 20 or SEQ ID NO: 21, or a fragment or variant thereof. In one embodiment, the group II intron reverse transcriptase is MarathonRT reverse transcriptase. In one embodiment, the group II intron reverse transcriptase is UltraMarathonRT. In one embodiment, the group II intron reverse transcriptase is Induro reverse transcriptase. In one embodiment, the group II intron reverse transcriptase is a TGIRT reverse transcriptase.
[0083] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, also known as Marathon Reverse Transcriptase (MRT), or a fragment or variant thereof. In some embodiments, the E.r. maturase comprises an amino acid sequence identical to an amino acid sequence selected from the group consisting of SEQ ID NOs: 3, and 8-12. In some embodiments, the fragment or variant of the E.r. maturase comprises a fragment or variant of an amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO: 9, SEQ ID NOTO, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13. In some embodiments, the fragment or variant of the E.r. maturase comprises a fragment or variant of an amino acid sequence of SEQ ID NO: 19. In some embodiments, the fragment or variant of the E.r. maturase retains at least one wild type activity of the parental E.r. maturase. In some embodiments, the fragment or variant of the E.r. maturase has an enhanced activity as compared to the parental E.r. maturase. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale E.r.)' maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 70% identical to an amino acid sequence of SEQ ID NOT, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NOTO, SEQ ID NO: 11, SEQ ID NOT 2 or SEQ ID NOT 3, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectal e (E.r.)
maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 70% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0084] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.)' maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 75% identical to an amino acid sequence of SEQ ID N0:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 75% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0085] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 80% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 80% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0086] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 85% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 85% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0087] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 90% identical to an amino acid sequence of SEQ ID NO : 3 , SEQ ID NO : 8, SEQ ID NO : 9, SEQ ID NO : 10, SEQ ID NO : 11 , SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 90% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0088] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r. maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 95% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 95% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0089] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 96% identical to an amino acid sequence of SEQ ID NO : 3 , SEQ ID NO : 8, SEQ ID NO : 9, SEQ ID NO : 10, SEQ ID NO : 11 , SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 96% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0090] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 97% identical to an amino acid sequence of SEQ ID NO : 3 , SEQ ID NO : 8, SEQ ID NO : 9, SEQ ID NO : 10, SEQ ID NO : 11 , SEQ
ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 97% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0091] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.)' maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 98% identical to an amino acid sequence of SEQ ID NO : 3 , SEQ ID NO : 8, SEQ ID NO : 9, SEQ ID NO : 10, SEQ ID NO : 11 , SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 98% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0092] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 99% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 99% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0093] In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino acid sequence at least 99.5% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof. In one embodiment, the enzymatic domain comprises a reverse transcriptase comprising Eubacterium rectale (E.r.) maturase, or a variant thereof, wherein the E.r. maturase or variant thereof, comprises an amino
acid sequence at least 99.5% identical to an amino acid sequence of SEQ ID NO: 19 or a functional fragment thereof.
[0094] In one embodiment the enzymatic domain comprises a functional fragment comprising at least 400 amino acid residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQIDNO:10, SEQIDNO:11, SEQIDNO:12or SEQIDNO:13.
[0095] In one embodiment the enzymatic domain comprises a functional fragment comprising at least 350 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQIDNO:10, SEQIDNO:11, SEQIDNO:12or SEQIDNO:13.
[0096] In one embodiment the enzymatic domain comprises a functional fragment comprising at least 300 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
[0097] In one embodiment the enzymatic domain comprises a functional fragment comprising at least 250 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ IDNO:10, SEQIDNO:11, SEQIDNO:12or SEQIDNO:13.
[0098] In one embodiment the enzymatic domain comprises a functional fragment comprising at least 200 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO:10, SEQ IDNO:11, SEQIDNO:12or SEQ ID NO:13.
[0099] In one embodiment the enzymatic domain comprises a functional fragment comprising at least 150 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ IDNO:10, SEQ IDNO:11, SEQIDNO:12or SEQ ID NO:13.
[0100] In one embodiment the enzymatic domain comprises a functional fragment comprising at least 100 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ IDNO:10, SEQIDNO:11, SEQIDNO:12or SEQIDNO:13.
[0101] In one embodiment the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 400 amino acids residues ofSEQIDNO:3, SEQIDNO:8, SEQIDNO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
[0102] In one embodiment the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 350 amino acids residues ofSEQIDNO:3, SEQIDNO:8, SEQIDNO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
[0103] In one embodiment the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 300 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
[0104] In one embodiment the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 250 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
[0105] In one embodiment the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 200 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
[0106] In one embodiment the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 150 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
[0107] In one embodiment the enzymatic domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 100 amino acids residues of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13.
[0108] In some embodiments, the reverse transcriptase has a sequence with at least 60% to 99.9% identity to a sequence listed in Example 3, e.g., at least 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 99.9% or greater identity to a sequence listed in Example 3.
Marathon Reverse Transcriptase
[0109] In one embodiment, the enzymatic domain comprises a reverse transcriptase. In one embodiment, the reverse transcriptase is derived from Eubacterium rectale (E.r.) maturase (also known as Marathon Reverse Transcriptase (MRT)). In one embodiment, E.r. maturase is modified relative to unmodified E.r. maturase. For example, in certain embodiments, the variant comprises one or more point mutations, insertion mutations, or deletion mutations, relative to
wildtype E.r. maturase. In certain embodiments, the variant comprises a fusion protein comprising E.r. maturase, E.r. maturase mutant, or E.r. maturase domain.
[0110] The full-length E.r. maturase comprises a “secondary” RNA binding site and DNA binding domain that can influence stability, specificity, and efficiency of reverse transcription of an RNA template. In one embodiment, the reverse transcriptase comprises an E.r. maturase variant where one or more secondary RNA binding sites on the surface of the protein are mutated to reduce nonspecific binding of the reverse transcription protein to the RNA template, thereby promoting binding at the polymerase cleft and facilitating enzyme turnover. In one such embodiment, a variant of E.r. maturase comprises at least one point mutation selected from the group R58X, K59X, K61X, K163X, K216X, R217X, K338X, K342X, and R353X wherein X denotes any amino acid. In another such embodiment, a variant of E.r. maturase comprises at least one point mutation selected from the group R58A, K59A, K61 A, K163A, K216A, R217A, K338A, K342A, and R353A.
[0111] In one embodiment, the reverse transcriptase comprises an E.r. maturase variant (referred to herein as E.r. maturase mutl; and denoted as SEQ ID NO:8) comprising the point mutations of: R58A, K59A, K61A, and K163A, relative to wildtype E.r. maturase.
[0112] In one embodiment, the reverse transcriptase comprises an E.r. maturase variant (referred to herein as E.r. maturase mut2; and denoted as SEQ ID NO:9) comprising the point mutations of: K216A and K217A, relative to wildtype E.r. maturase.
[0113] In one embodiment, the reverse transcriptase comprises an E.r. maturase variant (referred to herein as E.r. maturase mutl+mut2; and denoted as SEQ ID NOTO) comprising the point mutations of: R58A, K59A, K61A, K163A, K216A, and R217A, relative to wildtype E.r. maturase.
[0114] In one embodiment, the reverse transcriptase comprises an E.r. maturase variant (referred to herein as E.r. maturase mut3; and denoted as SEQ ID NO: 11) comprising the point mutations of: K338A, K342A, and R353A relative to wildtype E.r. maturase.
[0115] In one embodiment, the reverse transcriptase comprises an E.r. maturase variant comprising one or more mutations in the C-terminal DNA binding domain of E.r. maturase. In one such embodiment, a variant of E.r. maturase comprises at least one point mutation selected from the group K388X, R389X, K396X, K406X, R407X, and K423X, wherein X denotes any amino acid. In another such embodiment, a variant of E.r. maturase comprises at least one point
mutation selected from the group K388A, R389A, K396A, K406A, R407A, and K423A. In another such embodiment, a variant of E.r. maturase comprises at least one point mutation selected from the group K388S, R389S, K396S, K406S, R407S, and K423S. In another such embodiment, the C-terminal sequence residues 387-427 are deleted relative to wildtype E.r. maturase.
[0116] In one embodiment, the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, comprising one or more mutations in the a-loop (AA180-196) of E.r maturase. In one embodiment, the E.r. maturase variant comprises one or more mutations in the N-terminal region of the a-loop. In one embodiment, the mutation is at least one selected from the group: M180X, I181X, D182X, D183X, E184X, Y185X, E186X, D187X, S188X, I189X, V190X, wherein X denotes any amino acid. In one such embodiment, the at least one point mutation (X) is selected from the group: alanine, polar amino acid (e.g., Gin), electrostatic amino acid (e.g. Glu), and a combination thereof. In another such embodiment, the a-loop is engineered to be more flexible by substituting positions in the N- terminal region with one or more glycines. In another such embodiment, the a-loop is engineered to be more stiff by substituting positions in the N-terminal region with one or more alanines. In one embodiment, the mutation is a deletion of at least one residue of the a-loop. In one embodiment, the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant in which residues 182-192 are substituted with two glycine residues (Aloop; SEQ ID NO: 12).
[0117] E.r. maturase can perform reverse transcription at lower temperatures relative to other reverse transcriptases, and the engineering of a more thermostable E.r. maturase would enable amplification of RNA templates in a single reaction (i.e., without using DNA^DNA amplification reactions). Analysis of thermophilic protein structure and function suggests that they tend to have larger numbers of side-chain hydrogen bonds and salt-bridges within rigid sections of the tertiary structure. Therefore, in one embodiment, the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, engineered to have Lys-Glu pairs at positions that are proximal in 3-D space, according to the structure of the enzyme (Zhao C et al., 2016, Nature structural & molecular biology, 23(6):558-65). In one such embodiment, the variant comprises at least one point mutation selected from the group
LI IE (which can form a salt bridge with R56), L21E (which can form a salt bridge with K41), and S13E (which can form a salt bridge with K52).
[0118] In one embodiment, the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, engineered to comprise a proofreading (e g., 3'- 5' exonuclease) domain to enhance fidelity. In one such embodiment, the proofreading domain comprises an exonuclease domain. In another such embodiment, the proofreading domain is appended to the C-terminus of the E.r. maturase variant. In another such embodiment, the proofreading domain is appended to the C-terminus of the E.r. maturase variant through a linker molecule or sequence (see, for example, Ellefson, JW et al., 2016, Science, 352(6293): 1590-3).
[0119] Maturase reverse transcriptases are generally conserved among species, but some may have additional, beneficial properties compared to others. Therefore, in one embodiment, the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, wherein at least one fragment or domain of E.r. maturase is replaced with a fragment or domain from a maturase reverse transcriptase from a species other than Eubacterium rectale. For example, in one embodiment, the RT domain (finger and palm) of E.r. maturase reverse transcriptase is replaced with the RT domain from a thermophilic maturase reverse transcriptase to enhance thermostability. In another embodiment, the a-loop of E.r. maturase is replaced by a longer a-loop from another maturase reverse transcriptase to enhance processivity. In one embodiment, one or more amino acids are substituted with hydrophobic amino acids or charged amino acids in order to improve thermostability.
[0120] In one embodiment, the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, wherein one or more residues are substituted with one or more residues derived from a maturase enzyme from an organism other than Eubacterium rectale. For example, in some embodiments, the E.r. maturase variant can comprise one or more point mutations based on conserved residues in thermophilic maturases. In one embodiment, the variant comprises at least one mutation selected from the group: A29X, V82X, E104X, I129X, I137X, T161X, I168X, I170X, VI 7 IX, and M337X, where X denotes any amino acid. In one embodiment, the mutation is at least one selected from the group: A29X, V82X, E104X, I129X, I137X, T161X, I168X, I170X, V171X, and M337X, where X denotes any amino acid. In one embodiment, the variant comprises at least one mutation selected from the
group: A29S, V82I, E104P, I129Y, I137V, T161R, I168L, I170L, V171I, and M337T. In one embodiment, the variant comprises a triple point mutation of A29S/V82I/E104P. In certain instances, these mutations improve upon the thermostability of the enzyme.
[0121] In one embodiment, the avidin or streptavidin domain fusion construct of the present disclosure comprises an E.r. maturase variant, comprising one or more mutations in the thumb domain relative to wildtype E.r. maturase.
[0122] In one embodiment, the variant comprises at least one point mutation selected from the group consisting of K338X, K342X, and R353X, wherein X denotes any amino acid. In another such embodiment, the variant comprises at least one point mutation selected from the group consisting of K338A, K342A, and R353A.
[0123] In one such embodiment, one or more mutations are incorporated on the surface of the thumb domain, optimizing its ability to clasp the template. In one such embodiment, the variant comprises at least one point mutation selected from the group consisting of S315X, E319X, and Q323X, wherein X denotes any amino acid. In another such embodiment, the variant comprises at least one point mutation selected from the group consisting of S315K, E319K, and Q323K.
[0124] In one embodiment, the enzymatic domain comprises a polypeptide comprising a reverse transcriptase. In one embodiment, the reverse transcriptase is derived from E.r. maturase. For example, in one embodiment, the polypeptide comprises E.r. maturase, or a variant thereof. Exemplary amino acid sequences of the E.r. maturase-derived reverse transcriptases of the present disclosure include, but are not limited to, SEQ ID NO:3 (E.r. maturase), SEQ ID NO:8 (Er. maturase mutl), SEQ ID NO:9 (E.r. maturase mut2), SEQ ID NO: 10 (E.r. maturase mutl+mut2), SEQ ID NO: 11 (E.r. maturase mut3) and SEQ ID NO: 12 (E.r. maturase Aloop). However, the present disclosure is not limited to these sequences. Rather the present disclosure encompasses any reverse transcriptase derived from E.r. maturase or a variant thereof.
[0125] In one embodiment, the polypeptide comprises a fragment of E.r. maturase or variant thereof that mimics the ability of E.r. maturase to perform reverse transcription. In one embodiment, the polypeptide comprises a derivative of the E.r. maturase or variant thereof. In certain embodiments, the polypeptide comprises an amino acid sequence selected from a fragment or derivative of SEQ ID NO:3, a fragment or derivative of SEQ ID NO:8, a fragment or
derivative of SEQ ID NO:9, a fragment or derivative of SEQ ID NO: 10, a fragment or derivative of SEQ ID NO: 11, and a fragment or derivative of SEQ ID NO: 12.
[0126] In one embodiment, the avidin or streptavidin domain fusion construct of the disclosure comprises one or more mutations in the catalytic active-site to reduce the fidelity of the enzyme, which will enhance its value for RNA structure mapping since structure-specific lesions that are used to probe RNA structure are flagged by misincorporation events. Similarly, mutations that increase the error rate of the enzyme can be used with certain RNA and transcriptome mapping experiments. Therefore, in some embodiments, the polypeptide comprises at least one mutation selected from the group: A225X, R114X, Y224X, I179X, M180X, I181X, E143X, K65X, L201X, wherein X denotes any amino acid. Specifically, mutations at A225 (such as A225V, A225S, A225M or A225V), mutations at R114 (such as R114K, R114A), mutations at Y224 (such as Y224F), mutations at 1179 (such as I179F), mutations at Ml 80 (such as Ml 80V), mutations at 1181 (such as I181W), mutations at E143 (such as E143A or E143K), mutations at K65 (such as K65A), mutations at L201 (such as L201A or L201T), may be used, alone or in combination.
[0127] In one embodiment, the avidin or streptavidin domain fusion construct of the disclosure comprises one or more mutations in the primer/template binding interface to increase the binding affinity to primer/template, which will enhance its capability of capturing low abundance RNAs in single-cell or spatial transcriptomic RNA sequencing. Therefore, in some embodiments, the polypeptide comprises at least one mutation selected from the group: D14X, Q22X, N26X, E30X, D74X, Q91X, Q92X, Q96X, N116X, N197X, E304X, E319X, N322X, N330X, E422X, wherein X denotes any amino acid. Specifically, mutations at D14 (such as D14K or D14R), mutations at Q22 (such as Q22K or Q22R), mutations at N26 (such as N26K or N26R), mutations at E30 (such as E30K or E30R), mutations at D74 (such as D74K or D74R), mutations at Q91 (such as Q91K or Q91R), mutations at Q92 (such as Q92K or Q92R), mutations at Q96 (such as Q96K or Q96R), mutations at N116 (such as N116K or N116R), mutations at N197 (such as N197K or N197R), mutations at E304 (such as E304K or E304R), mutations at E319 (such as E319K or E319R), mutations at N322 (such as N322K or N322R), mutations at N33O (such as N330K or N33OR), mutations at E422 (such as E422K or E422R), may be used, alone or in combination.
[0128] In one embodiment, the enzymatic domain of the present disclosure comprises a polypeptide comprising Roseburia intestinalis (R.i.) maturase, or a variant or fragment thereof. In one such embodiment, the Ri. maturase comprises one or more mutations corresponding to one or more mutations described herein.
[0129] Reverse transcriptases of the present disclosure may produce more product (e.g., full length product) at particular temperatures compared to other reverse transcriptases. In one aspect, comparisons of full length product synthesis are made at different temperatures (e.g., one temperature being lower, such as between 4° C and 42° C, and one temperature being higher, such as between 42° C and 78° C) while keeping all other reaction conditions similar or the same. The amount of full length product produced may be determined using techniques well known in the art, for example, by conducting a reverse transcription reaction at a first temperature (e.g., 37° C, 38° C, 39° C, 40° C, etc.) and determining the amount of full length transcript produced, conducting a second reverse transcription reaction at a temperature higher than the first temperature (e.g., 45° C, 50° C, 52.5° C, 55° C, etc.) and determining the amount of full length product produced, and comparing the amounts produced at the two temperatures. A convenient form of comparison is to determine the percentage of the amount of full length product at the first temperature that is produced at the second (i.e., elevated) temperature. The reaction conditions used for the two reactions (e.g., salt concentration, buffer concentration, pH, divalent metal ion concentration, nucleoside triphosphate concentration, template concentration, reverse transcriptase concentration, primer concentration, length of time the reaction is conducted, etc.) may be the same for both reactions. Suitable reaction conditions may be determined by those skilled in the art using routine techniques and examples of such conditions are provided herein.
[0130] The reverse transcriptases of the disclosure may produce at least about 5%, at least 10%, at least 15%, at least 25%, at least 50%, at least 75%, at least 100%, or at least 200% more product or full length product compared to the corresponding control reverse transcriptase under the same reaction conditions and temperature. The reverse transcriptases of the disclosure may produce from about 10% to about 200%, from about 25% to about 200%, from about 50% to about 200%, from about 75% to about 200%, or from about 100% to about 200% more product or full length product compared to a control reverse transcriptase under the same reaction conditions and incubation temperature. The reverse transcriptases of the disclosure may
produce at least 2 times, at least 3 times, at least 4 times, at least 5 times, at least 6 times, at least 7 times, at least 8 times, at least 9 times, at least 10 times, at least 25 times, at least 50 times, at least 75 times, at least 100 times, at least 150 times, at least 200 times, at least 300 times, at least 400 times, at least 500 times, at least 1000 times, at least 5,000 times, or at least 10,000 times more product or full length product compared to a control reverse transcriptase under the same reaction conditions and temperature.
[0131] Reverse transcriptases of the present disclosure may have an increased thermostability at elevated temperatures as compared to corresponding control reverse transcriptases. They may show increased thermostability in the presence or absence an RNA template. In some instances, reverse transcriptases of the disclosure may show an increased thermostability in both the presence and absence of an RNA template. Those skilled in the art will appreciate that reverse transcriptase enzymes are typically more thermostable in the presence of an RNA template. The increase in thermostability may be measured by comparing suitable parameters of the modified or mutated reverse transcriptase of the disclosure to those of a corresponding un-modified or un-mutated reverse transcriptase. Suitable parameters to compare include, but are not limited to, the amount of product and/or full length product synthesized by the reverse transcriptases of the disclosure at an elevated temperature compared to the amount or product and/or full length product synthesized by a control reverse transcriptase at the same temperature, and/or the half-life of reverse transcriptase activity at an elevated temperature of a reverse transcriptase of the disclosure at an elevated temperature compared to that of a control reverse transcriptase.
[0132] A reverse transcriptase of the disclosure may have an increase in thermostability at a particular temperature of at least about 1.5 fold (e.g., from about 1.5 fold to about 100 fold, from about 1.5 fold to about 50 fold, from about 1.5 fold to about 25 fold, from about 1.5 fold to about 10 fold) compared, for example, to the control reverse transcriptase. A reverse transcriptase of the disclosure may have an increase in thermostability at a particular temperature of at least about 10 fold (e.g., from about 10 fold to about 100 fold, from about 10 fold to about 50 fold, from about 10 fold to about 25 fold, or from about 10 fold to about 15 fold) compared, for example, to the control reverse transcriptase. A reverse transcriptase of the disclosure may have an increase in thermostability at a particular temperature of at least about 25 fold (e.g., from
about 25 fold to about 100 fold, from about 25 fold to about 75 fold, from about 25 fold to about 50 fold, or from about 25 fold to about 35 fold) compared to the control reverse transcriptase.
[0133] The polypeptide of the present disclosure may be made using chemical methods. For example, polypeptides can be synthesized by solid phase techniques (Roberge J Y et al (1995) Science 269: 202-204), cleaved from the resin, and purified by preparative high performance liquid chromatography. Automated synthesis may be achieved, for example, using the ABI 431 A Peptide Synthesizer (Perkin Elmer) in accordance with the instructions provided by the manufacturer. The polypeptide may be made by recombinant means or by cleavage from a longer polypeptide. The polypeptide may be confirmed by amino acid analysis or sequencing.
[0134] The disclosure should also be construed to include any form of a polypeptide having substantial homology to a reverse transcriptase disclosed herein. For example, a polypeptide which is “substantially homologous” is about 50% homologous, about 70% homologous, about 80% homologous, about 90% homologous, about 95% homologous, about 96% homologous, about 97% homologous, about 98% homologous, about 99% homologous, or about 99.5% homologous to an amino acid sequence of a reverse transcriptase disclosed herein.
[0135] In some embodiments, the enzymatic domain comprises a reverse transcriptase comprising an amino acid sequence that is about 50%, about 70%, about 80%, about 90%, about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 99.5% homologous to E.r. maturase or E.r. maturase variant described herein.
[0136] In some embodiments, the enzymatic domain comprises a reverse transcriptase comprising an amino acid sequence that is about 50%, about 70%, about 80%, about 90%, about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 99.5% identical to the amino acid sequence set forth in SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NOTO, SEQ ID NO l l, SEQ ID NOT2, or SEQ ID NOT3.
[0137] In some embodiments, the enzymatic domain comprises a reverse transcriptase comprising an amino acid sequence that is about 50%, about 70%, about 80%, about 90%, about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 99.5% identical to the amino acid sequence set forth in SEQ ID NOT, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NOTO, SEQ ID NOT 1, SEQ ID NOT2, or SEQ ID NOT3, wherein the reverse transcriptase comprises one more of the mutations described herein.
[0138] In one aspect, the present disclosure provides a composition comprising an agent that improves RT activity of E.r. maturase or variants thereof. For example, in some embodiments, the composition comprises an agent that reduces non-specific binding of primers to the positively charged surface of E.r. maturase or variants thereof. In some embodiments, the agent that reduces non-specific binding of primers to the positively charged surface of E.r. maturase or variants thereof comprises a peptide or protein, including, but not limited to, heparin.
Other enzymatic domains
[0139] A skilled artisan will appreciate that the enzymatic domain of the present disclosure can comprise any enzymatic domain applicable for molecular technology applications and industrial/pharmaceutical production well known in the field, and is not limited to the enzymatic domains described herein.
[0140] Accordingly, in some embodiments, the enzymatic domain of the present disclosure comprises any reverse transcriptase (RT) or RNA-dependent DNA polymerase (e.g., AMV/MAV RT, HIV RT, MuLV RT, Tth, MonsterScript™ RT, KI enow fragment of C. therm, viral RT, non-viral RT, etc.), any DNA polymerase or DNA-dependent DNA polymerase (e.g., Bst polymerase, Taq polymerase, Tth polymerase, Pfu polymerase, Pow polymerase, Vent polymerase, Pab polymerase, T4 DNA polymerase, Pol I and Klenow fragment, T7 DNA polymerase, terminal deoxynucleotidyl transferase, DNA Pol I, DNA Pol II, etc.), any RNA polymerase or DNA-dependent RNA polymerase (e.g., SP6 RNA polymerase, T7 RNA polymerase, etc.), any nuclease (e.g., restriction endonucleases, DNasel, exonuclease III, Bal31, exonuclease VII, RNase A, ribonuclease H, Phy I, CL3, Cereus, Phy M, RNase Tl, RNase T2, RNase U2, nuclease SI, Mung Bean nuclease, etc.), any DNA ligase (e.g., T4-DNA ligase, E. Coli DNA ligase, Thermostable DNA ligases, etc ), any RNA ligase (e.g., T4 RNA ligase 1, T4 RNA ligase 2, truncated T4 RNA ligase 2, etc.), any methylase (e.g., dam methylase, dem methylase, etc.), or any other enzyme (e.g., alkaline phosphatase, T4 polynucleotide kinase, tobacco acid pyrophosphatase, polyadenlyate polymerase, topoisomerase I, topoisomerase II, guanylyl transferase, any helicase, any primase, and recombinase, any protease, any nucleic acid binding protein, any aminoacyl-tRNA synthetase, any glycosyltransferase, any fatty acid synthase and any enzyme for food, biofuel and pharmaceutical industries, such as palatase, cellulase and lipase).
Modifications to enzymatic domain polypeptides
[0141] In one embodiment, the enzymatic domain comprises a polypeptide. In one embodiment, the polypeptide comprises a polypeptide variant wherein (i) one or more of the amino acid residues are substituted with a conserved or non-conserved amino acid residue, and such substituted amino acid residue may or may not be one encoded by the naturally occurring genetic code, (ii) there are one or more modified amino acid residues, e.g., residues that are modified by the attachment of substituent groups, (iii) the polypeptide is an alternative splice variant of the polypeptide of the present disclosure, (iv) fragments of the polypeptides and/or (v) the polypeptide is fused with another polypeptide, such as a leader or secretory sequence or a sequence which is employed for purification (for example, His-tag) or for detection (for example, Sv5 epitope tag). The fragments include polypeptides generated via proteolytic cleavage (including multi-site proteolysis) of an original sequence. Variants may be post- translationally, or chemically modified. Such variants are deemed to be within the scope of those skilled in the art from the teaching herein.
[0142] As known in the art the “similarity” between two polypeptides is determined by comparing the amino acid sequence and its conserved amino acid substitutes of one polypeptide to a sequence of a second polypeptide. Variants are defined to include polypeptide sequences different from the original sequence, for example, different from the original sequence in less than 40% of residues per segment of interest, different from the original sequence in less than 25% of residues per segment of interest, different by less than 10% of residues per segment of interest, or different from the original protein sequence in just a few residues per segment of interest and at the same time sufficiently homologous to the original sequence to preserve the functionality of the original sequence and/or the ability to perform reverse transcription. The present disclosure includes amino acid sequences that are at least 60%, 65%, 70%, 72%, 74%, 76%, 78%, 80%, 90%, or 95% similar or identical to the original amino acid sequence. The degree of identity between two peptides is determined using computer algorithms and methods that are widely known to the persons skilled in the art. The identity between two amino acid sequences may be determined by using the BLASTP algorithm (BLAST Manual, Altschul, S., et al., NCBI NLM NIH Bethesda, Md. 20894, Altschul, S., et al., J. Mol. Biol. 215: 403-410 (1990)).
[0143] In some embodiments, the polypeptide of the disclosure can be post- translationally modified. For example, post-translational modifications that fall within the scope of the present disclosure include signal peptide cleavage, glycosylation, acetylation, isoprenylation, proteolysis, myristoylation, protein folding and proteolytic processing, etc. Some modifications or processing events require introduction of additional biological machinery. For example, processing events, such as signal peptide cleavage and core glycosylation, are examined by adding canine microsomal membranes or Xenopus egg extracts (U.S. Pat. No. 6,103,489) to a standard translation reaction.
[0144] In some embodiments, the polypeptide of the disclosure may include unnatural amino acids formed by post-translational modification or by introducing unnatural amino acids during translation. A variety of approaches are available for introducing unnatural amino acids during protein translation.
[0145] A polypeptide or protein of the disclosure may be conjugated with other molecules, such as proteins, to prepare fusion proteins. This may be accomplished, for example, by the synthesis of N-terminal or C-terminal fusion proteins provided that the resulting fusion protein retains the functionality of a reverse transcriptase.
[0146] A peptide or protein of the disclosure may be phosphorylated using conventional methods such as the method described in Reedijk et al. (The EMBO Journal 11(4): 1365, 1992).
[0147] Cyclic derivatives of the polypeptides of the disclosure are also part of the present disclosure. Cyclization may allow the polypeptide to assume a more favorable conformation for association with other molecules. Cyclization may be achieved using techniques known in the art. For example, disulfide bonds may be formed between two appropriately spaced components having free sulfhydryl groups, or an amide bond may be formed between an amino group of one component and a carboxyl group of another component. Cyclization may also be achieved using an azobenzene-containing amino acid as described by Ulysse, L., et al., J. Am. Chem. Soc. 1995, 117, 8466-8467. The components that form the bonds may be side chains of amino acids, nonamino acid components or a combination of the two. In an embodiment of the disclosure, cyclic peptides may comprise a beta-turn in the right position. Beta-turns may be introduced into the peptides of the disclosure by adding the amino acids Pro-Gly at the right position.
[0148] It may be desirable to produce a cyclic polypeptide which is more flexible than the cyclic polypeptides having peptide bond linkages as described above. A more flexible
polypeptide may be prepared by introducing cysteines at the right and left position of the polypeptide and forming a disulfide bridge between the two cysteines. The two cysteines are arranged so as not to deform the beta-sheet and turn. The polypeptide is more flexible as a result of the length of the disulfide linkage and the smaller number of hydrogen bonds in the beta-sheet portion. The relative flexibility of a cyclic polypeptide can be determined by molecular dynamics simulations.
[0149] In some embodiments, the polypeptide comprises a reverse transcriptase fused to, or integrated into, a target protein, and/or a targeting domain capable of directing the chimeric protein to a desired location. The chimeric proteins may also comprise additional amino acid sequences or domains. The chimeric proteins are recombinant in the sense that the various components are from different sources, and as such are not found together in nature (i.e., are heterologous).
[0150] In some embodiments, the polypeptide of the disclosure may be synthesized by conventional techniques. For example, the polypeptides or chimeric proteins may be synthesized by chemical synthesis using solid phase peptide synthesis. These methods employ either solid or solution phase synthesis methods (see for example, J. M. Stewart, and J. D. Young, Solid Phase Peptide Synthesis, 2nd Ed., Pierce Chemical Co., Rockford Ill. (1984) and G. Barany and R. B. Merrifield, The Peptides: Analysis Synthesis, Biology editors E. Gross and J. Meienhofer Vol. 2 Academic Press, New York, 1980, pp. 3-254 for solid phase synthesis techniques; and M Bodansky, Principles of Peptide Synthesis, Springer- Verlag, Berlin 1984, and E. Gross and J. Meienhofer, Eds., The Peptides: Analysis, Synthesis, Biology, suprs, Vol 1, for classical solution synthesis). By way of example, a polypeptide of the disclosure may be synthesized using 9- fluorenyl methoxycarbonyl (Fmoc) solid phase chemistry with direct incorporation of phosphothreonine as the N-fluorenylmethoxy-carbonyl-O-benzyl-L-phosphothreonine derivative.
[0151] N-terminal or C-terminal fusion proteins comprising a polypeptide or chimeric protein of the disclosure conjugated with other molecules may be prepared by fusing, through recombinant techniques, the N-terminal or C-terminal of the polypeptide or chimeric protein, and the sequence of a selected protein or selectable marker with a desired biological function. The resultant fusion proteins comprise a reverse transcriptase fused to the selected protein or marker protein as described herein. Examples of proteins which may be used to prepare fusion proteins
include immunoglobulins, glutathione-S-transf erase (GST), hemagglutinin (HA), and truncated myc.
[0152] In some embodiments, the polypeptide of the disclosure may be developed using a biological expression system. The use of these systems allows the production of large libraries of random peptide sequences and the screening of these libraries for peptide sequences that bind to particular proteins. Libraries may be produced by cloning synthetic DNA that encodes random peptide sequences into appropriate expression vectors (see Christian et al 1992, J. Mol. Biol. 227:711; Devlin et al, 1990 Science 249:404; Cwirla et al 1990, Proc. Natl. Acad, Sci. USA, 87:6378). Libraries may also be constructed by concurrent synthesis of overlapping peptides (see U.S. Pat. No. 4,708,871).
[0153] In some embodiments, the polypeptide of the disclosure may be converted into pharmaceutical salts by reacting with inorganic acids such as hydrochloric acid, sulfuric acid, hydrobromic acid, phosphoric acid, etc., or organic acids such as formic acid, acetic acid, propionic acid, glycolic acid, lactic acid, pyruvic acid, oxalic acid, succinic acid, malic acid, tartaric acid, citric acid, benzoic acid, salicylic acid, benezenesulfonic acid, and toluenesulfonic acids.
Affinity Domain
[0154] The present disclosure provides a fusion construct comprising an affinity domain, e g., a domain capable of binding to a specific target or substrate. In some embodiments, the affinity domain comprises a protein or a fragment of a protein. In one embodiment, the affinity domain comprises glutathione-S-transferase. In one embodiment, the affinity domain comprises chitin binding protein. In one embodiment, the affinity domain comprises maltose binding protein. In one embodiment, the affinity domain comprises colicin E7 (CL7-tag). In one embodiment, the affinity domain comprises a SNAP-tag. In one embodiment, the affinity domain comprises a CLIP-tag. In one embodiment, the affinity comprises a His tag, e.g., a tag comprising at least two histidine amino acids, e g., at least 2, 3, 4, 5, 6, 7, 8, or more histidine amino acids. In one embodiment, the affinity domain comprises avidin. In one embodiment, the affinity tag comprises streptavidin. In one embodiment, the affinity domain comprises a monomer of avidin or streptavidin.
[01 5] In one embodiment, the streptavidin domain comprises an amino acid sequence at least 70% identical to the amino acid sequence set forth in SEQ ID NO:1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 75% identical to the amino acid sequence set forth in SEQ ID NO:1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 80% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 85% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 90% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 95% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 96% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 97% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 98% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 99% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 99.5% identical to the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence at least 99.9% identical to the amino acid sequence set forth in SEQ ID NO:1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 40 amino acids residues of the amino acid sequence set forth in SEQ ID NO:1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 60 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 80 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 100 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence
comprising at least 40 amino acids residues of the amino acid sequence set forth in SEQ ID NO:1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 60 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 80 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 1. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 100 amino acids residues of the amino acid sequence set forth in SEQ ID NO:1. In one embodiment, the streptavidin domain comprises the amino acid sequence of SEQ ID NO: 1.
[0156] In one embodiment, the avidin domain comprises an amino acid sequence at least 70% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 75% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 80% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 85% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 90% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 95% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 96% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 97% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 98% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 99% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 99.5% identical to the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the avidin domain comprises an amino acid sequence at least 99.9% identical to the amino acid sequence set forth in SEQ ID NO: 18.
[0157] In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 40 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 60 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 80 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is identical to an amino acid sequence comprising at least 100 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 40 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 60 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 80 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 100 amino acids residues of the amino acid sequence set forth in SEQ ID NO: 18. In one embodiment, the streptavidin domain comprises the amino acid sequence of SEQ ID NO: 18.
[0158] A skilled artisan will appreciate that in some embodiments, the variants or fragments of the avidin or streptavidin domain of the present disclosure can comprise variations in the sequence that do not prevent or disrupt folding of the avidin or streptavidin monomer or the ability of the avidin or streptavidin monomer to form a tetramer in the presence of additional avidin or streptavidin monomers.
[0159] In some embodiments, the affinity domain is covalently attached to the enzymatic domain by a linker, e.g., a peptide linker. In some embodiments, the affinity domain is linked to an enzymatic domain comprising a polymerase, e.g., a reverse transcriptase. In some embodiments, the affinity domain is linked to a reverse transcriptase. In some embodiments, the affinity domain linked to a polymerase is avidin. In some embodiments, the affinity domain
linked to a polymerase is streptavidin. In some embodiments, the affinity domain linked to a reverse transcriptase is avidin. In some embodiments, the affinity domain linked to a reverse transcriptase is streptavidin. In some embodiments, the affinity domain comprising a monomer of avidin or streptavidin is linked to an enzymatic domain via a peptide linker sequence. In some embodiments, the present disclosure provides a streptavidin fusion construct comprising a streptavidin domain. In some embodiments, the streptavidin domain is linked to an enzymatic domain via a linker sequence. In some embodiments, the present disclosure provides an avidin fusion construct comprising an avidin domain. In some embodiments, the avidin domain is linked to an enzymatic domain via a linker sequence.
Linker
[0160] In some embodiments, the fusion construct comprises an avidin or streptavidin domain linked to an enzymatic domain via a linker sequence, e.g., an amino acid linker. In some embodiments, the linker sequence is a flexible linker. In some embodiments, the linker sequence comprises between 1-5, 5-10, 10-15, 15-20, 20-25, or 25-30 amino acids. In some embodiments, the linker sequence comprises between 1-5 amino acids. In some embodiments, the linker sequence comprises between 5-10 amino acids. In some embodiments, the linker sequence comprises between 10-15 amino acids. In some embodiments, the linker sequence comprises between 15-20 amino acids. In some embodiments, the linker sequence comprises between 20-25 amino acids. In some embodiments, the linker sequence comprises between 25-30 amino acids. In some embodiments, the linker sequence comprises at least 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18. 19 or 20 amino acids, or more than 20 amino acids. In some embodiments, the linker sequence comprises at least 1 amino acid. In some embodiments, the linker sequence comprises at least 2 amino acids. In some embodiments, the linker sequence comprises at least 3 amino acids. In some embodiments, the linker sequence comprises at least 4 amino acids. In some embodiments, the linker sequence comprises at least 5 amino acids. In some embodiments, the linker sequence comprises at least 6 amino acids. In some embodiments, the linker sequence comprises at least 7 amino acids. In some embodiments, the linker sequence comprises at least 8 amino acids. In some embodiments, the linker sequence comprises at least 9 amino acids. In some embodiments, the linker sequence comprises at least 10 amino acids. In some embodiments, the linker sequence comprises at least 11 amino acids. In some embodiments, the
linker sequence comprises at least 12 amino acids. In some embodiments, the linker sequence comprises at least 13 amino acids. In some embodiments, the linker sequence comprises at least 14 amino acids. In some embodiments, the linker sequence comprises at least 15 amino acids. In some embodiments, the linker sequence comprises at least 16 amino acids. In some embodiments, the linker sequence comprises at least 17 amino acids. In some embodiments, the linker sequence comprises at least 18 amino acids. In some embodiments, the linker sequence comprises at least 19 amino acids. In some embodiments, the linker sequence comprises at least 20 amino acids.
[0161] In some embodiments, the linker sequence comprises a type of amino acid, e.g., a polar amino acid, a nonpolar amino acid, a hydrophobic amino acid, or a charged amino acid. In some embodiments, the linker sequence comprises a polar amino acid, e.g., serine (S), threonine (T), cysteine (C), tyrosine (Y), asparagine (N), glutamine (Q), or glycine (G). In some embodiments, the linker sequence comprises an S amino acid. In some embodiments, the linker sequence comprises a T amino acid. In some embodiments, the linker sequence comprises a C amino acid. In some embodiments, the linker sequence comprises a Y amino acid. In some embodiments, the linker sequence comprises an N amino acid. In some embodiments, the linker sequence comprises a Q amino acid. In some embodiments, the linker sequence comprises a G amino acid. In some embodiments, the linker sequence comprises at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 polar amino acids. In some embodiments, the linker sequence comprises at least 1 S amino acid. In some embodiments, the linker sequence comprises at least 2 S amino acids. In some embodiments, the linker sequence comprises at least 3 S amino acids. In some embodiments, the linker sequence comprises at least 4 S amino acids. In some embodiments, the linker sequence comprises at least 5 S amino acids. In some embodiments, the linker sequence comprises at least 6 S amino acids. In some embodiments, the linker sequence comprises at least 7 S amino acids. In some embodiments, the linker sequence comprises at least 8 S amino acids. In some embodiments, the linker sequence comprises at least 9 S amino acids. In some embodiments, the linker sequence comprises at least 10 S amino acids. In some embodiments, the linker sequence comprises at least 1 G amino acid. In some embodiments, the linker sequence comprises at least 2 G amino acids. In some embodiments, the linker sequence comprises at least 3 G amino acids. In some embodiments, the linker sequence comprises at least 4 G amino acids. In some embodiments, the linker sequence comprises at least
5 G amino acids. Tn some embodiments, the linker sequence comprises at least 6 G amino acids. In some embodiments, the linker sequence comprises at least 7 G amino acids. In some embodiments, the linker sequence comprises at least 8 G amino acids. In some embodiments, the linker sequence comprises at least 9 G amino acids. In some embodiments, the linker sequence comprises at least 10 G amino acids.
[0162] In some embodiments, the linker sequence comprises a hydrophobic amino acid, e.g., alanine (A), valine (V), leucine (L), isoleucine (I), methionine (M), phenylalanine (F), tryptophan (W), or proline (P). In some embodiments, the linker sequence comprises an A amino acid. In some embodiments, the linker sequence comprises a V amino acid. In some embodiments, the linker sequence comprises an L amino acid. In some embodiments, the linker sequence comprises an I amino acid. In some embodiments, the linker sequence comprises an M amino acid. In some embodiments, the linker sequence comprises an F amino acid. In some embodiments, the linker sequence comprises a W amino acid. In some embodiments, the linker sequence comprises a P amino acid. In some embodiments, the linker sequence comprises at least 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 hydrophobic amino acids. In some embodiments, the linker sequence comprises at least 1 A amino acid. In some embodiments, the linker sequence comprises at least 2 A amino acids. In some embodiments, the linker sequence comprises at least 3 A amino acids. In some embodiments, the linker sequence comprises at least 4 A amino acids. In some embodiments, the linker sequence comprises at least 5 A amino acids. In some embodiments, the linker sequence comprises at least 6 A amino acids. In some embodiments, the linker sequence comprises at least 7 A amino acids. In some embodiments, the linker sequence comprises at least 8 A amino acids. In some embodiments, the linker sequence comprises at least 9 A amino acids. In some embodiments, the linker sequence comprises at least 10 A amino acids.
[0163] In some embodiments, the linker sequence comprises a charged amino acid, e.g., histidine (H), lysine (K), arginine (R), aspartate (D), or glutamate (E). In some embodiments, the linker sequence comprises an H amino acid. In some embodiments, the linker sequence comprises a K amino acid. In some embodiments, the linker sequence comprises an R amino acid. In some embodiments, the linker sequence comprises a D amino acid. In some embodiments, the linker sequence comprises an E amino acid. In some embodiments, the linker
sequence comprises at least 1 , 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 charged amino acids.
[0164] A skilled artisan will appreciate that in some embodiments, the linker sequence can comprise any sequence capable of linking the avidin or streptavidin domain to the enzymatic domain, and is not limited to the linker sequences described herein.
[0165] In one embodiment, the linker comprises an amino acid sequence at least 80% identical to the sequence set forth in SEQ ID NO: 2, e.g., at least 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 99.9% identical to SEQ ID NO: 2. In one embodiment, the linker comprises an amino acid sequence at least 80% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 85% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 95% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 96% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 97% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 98% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 99% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 99.5% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises an amino acid sequence at least 99.9% identical to the sequence set forth in SEQ ID NO:2. In one embodiment, the linker comprises SEQ ID NO:2.
[0166] In one embodiment, the linker comprises an amino acid sequence at least 80% identical to the sequence set forth in SEQ ID NO: 5, e.g., at least 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.5%, or 99.9% identical to SEQ ID NO: 5. In one embodiment, the linker comprises an amino acid sequence at least 80% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 85% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an
amino acid sequence at least 95% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 96% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 97% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 98% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 99% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 99.5% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises an amino acid sequence at least 99.9% identical to the sequence set forth in SEQ ID NO:5. In one embodiment, the linker comprises SEQ ID NO:5.
Streptavidin Fusion Construct
[0167] The present disclosure provides a composition comprising at least one streptavidin fusion construct comprising a streptavidin domain operably linked to an enzymatic domain. In one embodiment, the streptavidin domain is linked to the N- or C- terminus of the enzymatic domain via a linker sequence.
[0168] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to the amino acid sequence set forth in SEQ ID NO:1, and is linked at is C-terminus to the N-terminus of an amino acid sequence that is at least about 70% identical to the amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NOTO, SEQ ID NOT l, SEQ ID NO:12 or SEQ ID NO: 13 or a functional fragment thereof, via an amino acid sequence that is at least about 70% identical to the amino acid sequence set forth in SEQ ID NO:2.
[0169] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence as set forth in SEQ ID NOT, SEQ ID NOT, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13 operably linked to a streptavidin monomeric domain comprising a sequence as set forth in SEQ ID NO: 1. In one embodiment, the C terminus of SEQ ID NO: 1 is linked to the N-terminus of SEQ ID NOT, SEQ ID NOT, SEQ ID NO:9, SEQ ID NOTO, SEQ ID NOT 1, SEQ ID NO: 12 or SEQ ID NOT 3 via a linker sequence. In some embodiments, the linker sequence comprises SEQ ID NOT.
[0170] Therefore, in one embodiment, the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO:3. In one embodiment, the linker comprises SEQ ID NO:2.
[0171] Therefore, in one embodiment, the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO:8. In one embodiment, the linker comprises SEQ ID NO:2.
[0172] Therefore, in one embodiment, the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO:9. In one embodiment, the linker comprises SEQ ID NO:2.
[0173] Therefore, in one embodiment, the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO: 10. In one embodiment, the linker comprises SEQ ID NO:2.
[0174] Therefore, in one embodiment, the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO: 11. In one embodiment, the linker comprises SEQ ID NO:2.
[0175] Therefore, in one embodiment, the streptavidin fusion construct comprises SEQ ID NO: 1 linked to the N-terminus of SEQ ID NO: 12. In one embodiment, the linker comprises SEQ ID NO:2.
[0176] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least 70% identical to the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0177] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least 75% identical to the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0178] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least 80% identical to the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0179] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least 85% identical to the amino acid sequence set forth in SEQ ID NON, SEQ ID NO:6 or SEQ ID NO:7.
[0180] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least 90% identical to the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0181] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least 95% identical to the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0182] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 250 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0183] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 300 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0184] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 350 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0185] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 400 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0186] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 450 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0187] In one embodiment, the streptavidin fusion construct comprises an amino acid sequence that is at least about 70% identical to an amino acid sequence comprising at least 500 amino acids residues of the amino acid sequence set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0188] In one embodiment, the streptavidin fusion construct comprises the amino acid sequence as set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
Peptide tag
[0189] In some embodiments, the fusion constructs of the disclosure, are linked to a peptide tag. In some embodiments, the peptide tag of the present disclosure comprises any suitable peptide tag. In some embodiments, the peptide tag can be removed or cleaved through protease treatment. In some embodiments, the peptide tag is beneficial to verify that a protein multimer comprises the appropriate number of monomers. A skilled artisan will appreciate that peptide tags of this nature are well known in the field and will consequently be able to select an appropriate peptide tag for use with the current disclosure. A skilled artisan will additional be able to select the appropriate protease for subsequent cleavage of the peptide tag.
[0190] In some embodiments the peptide tag comprises a His6-Small Ubiquitin-like Modifier (SUMO) tag. In one embodiment, following oligomerization of the at least one protein multimer, the peptide tag is cleaved through protease treatment of the at least one protein multimer. In one embodiment, the protease comprises SUMO-protease.
Streptavidin Tetramer
[0191] In some embodiments, the present disclosure provides a composition comprising at least one protein multimer, wherein the at least one protein multimer comprises at least one streptavidin fusion construct of the disclosure. In some embodiments, the protein multimer comprises a streptavidin tetramer comprising at least one streptavidin fusion construct of the disclosure. In some embodiments, the protein multimer comprises a streptavidin tetramer comprising at least two streptavidin fusion constructs of the disclosure. In some embodiments, the protein multimer comprises a streptavidin tetramer comprising at least three streptavidin fusion constructs of the disclosure.
[0192] In some embodiments, the protein multimer comprises one streptavidin fusion construct of the disclosure in a tetrameric complex with three additional streptavidin monomers. In some embodiments, the protein multimer comprises two streptavidin fusion constructs of the disclosure in a tetrameric complex with two additional streptavidin monomers. In some
embodiments, the protein multimer comprises three streptavidin fusion constructs of the disclosure in a tetrameric complex with one additional streptavidin monomer.
[0193] In one embodiment, one or more of the streptavidin domains of a streptavidin fusion construct or a streptavidin monomer incorporated into a tetramer of the disclosure comprise an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO: 1.
Nucleic acids
[0194] In one embodiment, the present disclosure provides a composition comprising an isolated nucleic acid encoding a streptavidin fusion construct of the disclosure. For example, in certain embodiments, the composition comprises a nucleic acid encoding streptavidin fusion construct of the disclosure comprising a streptavidin domain fused to an enzymatic domain. In certain embodiments, the nucleic acid is DNA, RNA, mRNA, or cDNA.
[0195] Exemplary enzymatic domains that can be encoded by the nucleic acid molecules of the disclosure are described elsewhere herein and include, but are not limited to, reverse transcriptases, translocating helicase enzymes, DNA polymerases, and RNA polymerases. In some embodiments, the enzymatic domain comprises a reverse transcriptase derived from E.r. maturase. In one embodiment, the enzymatic domain comprises a reverse transcriptase, wherein the reverse transcriptase comprises E.r. maturase or variant thereof.
[0196] In one embodiment, the isolated nucleic acid encodes a reverse transcriptase comprising E.r. maturase, or a variant thereof, wherein the amino acid sequence of E.r. maturase, or a variant thereof, is at least about 90% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
[0197] In one embodiment, the isolated nucleic acid encodes a reverse transcriptase comprising E.r. maturase, or a variant thereof, wherein the amino acid sequence of E.r. maturase, or a variant thereof, is identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
[0198] In one embodiment, the isolated nucleic acid comprises a nucleotide sequence encoding an amino acid at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%,
80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% identical to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13, or a functional fragment thereof.
[0199] In some embodiments, the isolated nucleic acid encodes an E.r. maturase variant comprising at least one point mutation selected from the group R58X, K59X, K61X, K163X, K216X, R217X, K338X, K342X, and R353X relative to wildtype E.r. maturase, wherein X denotes any amino acid. In some embodiments, the nucleic acid encodes an E.r. maturase variant comprising at least one point mutation selected from the group R58A, K59A, K61 A, K163A, K216A, R217A, K338A, K342A, and R353A relative to wildtype E.r. maturase.
[0200] In one embodiment, the isolated nucleic acid encodes an E.r. maturase variant (referred to herein as E.r. maturase mutl; and denoted as SEQ ID NO: 8) comprising the point mutations of: R58A, K59A, K61A, and K163A, relative to wildtype E.r. maturase.
[0201] In one embodiment, the isolated nucleic acid encodes an E.r. maturase variant (referred to herein as E.r. maturase mut2; and denoted as SEQ ID NO:9) comprising the point mutations of: K216A and K217A, relative to wildtype E.r. maturase.
[0202] In one embodiment, the isolated nucleic acid encodes an E.r. maturase variant (referred to herein as E.r. maturase mutl+mut2; and denoted as SEQ ID NO: 10) comprising the point mutations of: R58A, K59A, K61A, K163A, K216A, and R217A, relative to wildtype E.r. maturase.
[0203] In one embodiment, the n isolated ucleic acid encodes an E.r. maturase variant (referred to herein as E.r. maturase mut3; and denoted as SEQ ID NO: 11) comprising the point mutations of: K338A, K342A, and R353A relative to wildtype E.r. maturase.
[0204] In one embodiment, the isolated nucleic acid encodes an E.r. maturase variant comprising one or more mutations in the a-loop, C-terminal DNA binding domain, and/or thumb domain. In one embodiment, the isolated nucleic acid encodes an E.r. maturase variant engineered to have Lys-Glu pairs at positions that are proximal in 3-D space. In one embodiment, the isolated nucleic acid encodes an E.r. maturase variant, wherein one or more fragments or domains of E.r. maturase is replaced by one or more fragments or domains from a maturase reverse transcriptase from a species other than Eubacterium rectale.
[0205] In certain embodiments, the composition increases the expression of a biologically functional fragment of E.r. maturase. For example, in one embodiment, the composition comprises an isolated nucleic acid sequence encoding a biologically functional fragment of E.r. maturase. As would be understood in the art, a biologically functional fragment is a portion or portions of a full-length sequence that retain at least one biological function of the full-length sequence. Thus, a biologically functional fragment of E.r. maturase comprises a peptide that retains at least one function of full length E.r. maturase.
[0206] In some embodiments, the isolated nucleic acid encodes a streptavidin fusion construct comprising an enzymatic domain comprising a peptide having substantial homology to a reverse transcriptase disclosed herein. In certain embodiments, the isolated nucleic acid sequence encodes a reverse transcriptase having at least 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% sequence identity to an amino acid sequence of SEQ ID NO:3, SEQ ID NO: 8, SEQ ID N0:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13 or a functional fragment thereof.
[0207] The isolated nucleic acid sequence encoding a reverse transcriptase can be obtained using any of the many recombinant methods known in the art, such as, for example by screening libraries from cells expressing the gene, by deriving the gene from a vector known to include the same, or by isolating directly from cells and tissues containing the same, using standard techniques. Alternatively, the gene of interest can be produced synthetically, rather than cloned.
[0208] The isolated nucleic acid may comprise any type of nucleic acid, including, but not limited to DNA and RNA. For example, in one embodiment, the composition comprises an isolated DNA molecule, including for example, an isolated cDNA molecule, encoding a reverse transcriptase. In one embodiment, the composition comprises an isolated RNA molecule encoding a streptavidin fusion construct of the disclosure.
[0209] In one embodiment, the present disclosure provides an isolated nucleic acid encoding a streptavidin fusion construct comprising a nucleotide sequence encoding an amino acid at least about 90% identical to the amino acid sequence as set forth in SEQ ID NO: 1. In one embodiment, the isolated nucleic acid comprises a nucleotide sequence encoding an amino acid at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%
identical to the amino acid sequence as set forth in SEQ ID NO: 1 . In one embodiment, the isolated nucleic acid comprises a nucleotide sequence encoding SEQ ID NO:1.
[0210] In some embodiments, the isolated nucleic acid encodes a streptavidin fusion construct comprising a nucleotide sequence encoding an amino acid at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% identical to the amino acid sequence as set forth in SEQ ID NO:1 linked to a nucleotide sequence encoding a reverse transcriptase having at least 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% sequence identity to an amino acid sequence of SEQ ID NO:3, SEQ ID NO:8, SEQ ID NO:9, SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12 or SEQ ID NO: 13 or a functional fragment thereof.
[0211] In one embodiment, the isolated nucleic acid encodes a linker. In one embodiment, the isolated nucleic acid comprises a nucleotide sequence encoding an amino acid at least about 90% identical to the amino acid sequence as set forth in SEQ ID NO:2. In one embodiment, the isolated nucleic acid comprises a nucleotide sequence encoding SEQ ID NO:2.
[0212] In one embodiment, the isolated nucleic acid comprises a nucleotide sequence encoding an amino acid at least about 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% identical to the amino acid sequence as set forth in SEQ ID NON, SEQ ID NO:6 or SEQ ID NO:7. In one embodiment, the isolated nucleic acid comprises a nucleotide sequence encoding SEQ ID NON, SEQ ID NO:6 or SEQ ID NON.
[0213] A skilled artisan will appreciate that in some embodiments, the linker sequence can comprise any sequence capable of linking the streptavidin domain to the enzymatic domain, wherein the enzymatic domain maintains at least one wild type catalytic activity in the oligomer, and is not limited to the linker sequences described herein.
Modifications to nucleic acids
[0214] The nucleic acid molecules of the present disclosure can be modified to improve stability in serum or in growth medium for cell cultures. Modifications can be added to enhance stability, functionality, and/or specificity. For example, in order to enhance the stability, the 3’- residues may be stabilized against degradation, e.g., they may be selected such that they consist of purine nucleotides, particularly adenosine or guanosine nucleotides. Alternatively, substitution
of pyrimidine nucleotides by modified analogues, e.g., substitution of uridine by 2’- deoxythymidine is tolerated and does not affect function of the molecule.
[0215] In one embodiment of the present disclosure the nucleic acid molecule may comprise at least one modified nucleotide analogue. For example, the ends may be stabilized by incorporating modified nucleotide analogues.
[0216] Non-limiting examples of nucleotide analogues include sugar- and/or backbone- modified ribonucleotides (i.e., include modifications to the phosphate-sugar backbone). For example, the phosphodiester linkages of natural RNA may be modified to include at least one of a nitrogen or sulfur heteroatom. In exemplary backbone-modified ribonucleotides the phosphoester group connecting to adjacent ribonucleotides is replaced by a modified group, e.g., of phosphothioate group. In preferred sugar-modified ribonucleotides, the 2’ OH-group is replaced by a group selected from H, OR, R, halo, SH, SR, NH2, NHR, NR2 or ON, wherein R is Ci-Ce alkyl, alkenyl or alkynyl and halo is F, Cl, Br or I.
[0217] Other examples of modifications are nucleobase-modified ribonucleotides, i.e., ribonucleotides, containing at least one non-naturally occurring nucleobase instead of a naturally occurring nucleobase. Bases may be modified to block the activity of adenosine deaminase. Exemplary modified nucleobases include, but are not limited to, uridine and/or cytidine modified at the 5-position, e.g., 5-(2-amino)propyl uridine, 5-bromo uridine; adenosine and/or guanosines modified at the 8 position, e.g., 8-bromo guanosine; deaza nucleotides, e.g., 7-deaza-adenosine; O- and N-alkylated nucleotides, e.g., N6-methyl adenosine are suitable. The above modifications may be combined.
[0218] In some instances, the nucleic acid molecule comprises at least one of the following chemical modifications: 2’-H, 2’-O-methyl, or 2’-OH modification of one or more nucleotides. In certain embodiments, a nucleic acid molecule of the disclosure can have enhanced resistance to nucleases. For increased nuclease resistance, a nucleic acid molecule, can include, for example, 2’ -modified ribose units and/or phosphorothioate linkages. For example, the 2’ hydroxyl group (OH) can be modified or replaced with a number of different “oxy” or “deoxy” substituents. For increased nuclease resistance the nucleic acid molecules of the disclosure can include 2’-O-methyl, 2’-fluorine, 2’-O-methoxyethyl, 2’-O-aminopropyl, 2’- amino, and/or phosphorothioate linkages. Inclusion of locked nucleic acids (LNA), ethylene nucleic acids (ENA), e.g., 2’-4’-ethylene-bridged nucleic acids, and certain nucleobase
modifications such as 2-amino-A, 2-thio (e.g., 2-thio-U), G-clamp modifications, can also increase binding affinity to a target.
[0219] In one embodiment, the nucleic acid molecule includes a 2’ -modified nucleotide, e.g., a 2’-deoxy, 2 ’-deoxy-2’ -fluoro, 2’-O-methyl, 2’-O-methoxyethyl (2’-O-MOE), 2’-O- aminopropyl (2’-O-AP), 2’-O-dimethylaminoethyl (2’-O-DMAOE), 2’-O-dimethylaminopropyl (2’-O-DMAP), 2’-O-dimethylaminoethyloxyethyl (2’-O-DMAEOE), or 2’-O-N- methylacetamido (2’-0-NMA). In one embodiment, the nucleic acid molecule includes at least one 2’-O-methyl-modified nucleotide, and in some embodiments, all of the nucleotides of the nucleic acid molecule include a 2’-O-methyl modification.
[0220] In certain embodiments, the nucleic acid molecule of the disclosure may have one or more of the following properties:
[0221] Nucleic acid agents discussed herein include otherwise unmodified RNA and DNA as well as RNA and DNA that have been modified, e.g., to improve efficacy, and polymers of nucleoside surrogates. Unmodified RNA refers to a molecule in which the components of the nucleic acid, namely sugars, bases, and phosphate moieties, are the same or essentially the same as that which occur in nature. The art has referred to rare or unusual, but naturally occurring, RNAs as modified RNAs, see, e.g., Limbach et al. (Nucleic Acids Res., 1994, 22:2183-2196). Such rare or unusual RNAs, often termed modified RNAs, are typically the result of a post- transcriptional modification and are within the term unmodified RNA as used herein. Modified RNA, as used herein, refers to a molecule in which one or more of the components of the nucleic acid, namely sugars, bases, and phosphate moieties, are different from those which occur in nature. While they are referred to as “modified RNAs” they will of course, because of the modification, include molecules that are not, strictly speaking, RNAs. Nucleoside surrogates are molecules in which the ribophosphate backbone is replaced with a non-ribophosphate construct that allows the bases to be presented in the correct spatial relationship such that hybridization is substantially similar to what is seen with a ribophosphate backbone, e.g., non-charged mimics of the ribophosphate backbone.
[0222] Modifications of the nucleic acid of the disclosure may be present at one or more of, a phosphate group, a sugar group, backbone, N-terminus, C-terminus, or nucleobase.
Expression systems
[0223] The present disclosure also includes a vector in which the isolated nucleic acid of the present disclosure is inserted. The art is replete with suitable vectors that are useful in the present disclosure.
[0224] In brief summary, the expression of natural or synthetic nucleic acids encoding a reverse transcriptase described herein is typically achieved by operably linking a nucleic acid encoding a streptavidin fusion construct of the disclosure to a promoter, and incorporating the construct into an expression vector. The vectors to be used are suitable for replication and, optionally, integration in host cells. Typical vectors contain transcription and translation terminators, initiation sequences, and promoters useful for regulation of the expression of the desired nucleic acid sequence.
[0225] The isolated nucleic acid of the disclosure can be cloned into many types of vectors. For example, the nucleic acid can be cloned into a vector including, but not limited to a plasmid, a phagemid, a phage derivative, an animal virus, and a cosmid. Vectors of particular interest include expression vectors, replication vectors, probe generation vectors, and sequencing vectors.
[0226] Further, the vector may be provided to a cell in the form of a viral vector. Viral vector technology is well known in the art and is described, for example, in Sambrook et al. (2012, Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Laboratory, New York), and in other virology and molecular biology manuals. Viruses, which are useful as vectors include, but are not limited to, retroviruses, adenoviruses, adeno- associated viruses, herpes viruses, and lentiviruses. In general, a suitable vector contains an origin of replication functional in at least one organism, a promoter sequence, convenient restriction endonuclease sites, and one or more selectable markers, (e.g., WO 01/96584; WO 01/29058; and U.S. Pat. No. 6,326,193).
[0227] A number of viral based systems have been developed for gene transfer into cells. For example, retroviruses provide a convenient platform for gene delivery systems. A selected gene can be inserted into a vector and packaged in retroviral particles using techniques known in the art. The recombinant virus can then be isolated and delivered to cells. A number of retroviral systems are known in the art. In some embodiments, adenovirus vectors are used. A number of adenovirus vectors are known in the art. In one embodiment, lentivirus vectors are used.
[0228] For example, vectors derived from retroviruses such as the lentivirus are suitable tools to achieve long-term gene transfer since they allow long-term, stable integration of a
transgene and its propagation in daughter cells. Lentiviral vectors have the added advantage over vectors derived from onco-retroviruses such as murine leukemia viruses in that they can transduce non-proliferating cells, such as hepatocytes. They also have the added advantage of low immunogenicity. In one embodiment, the composition includes a vector derived from an adeno-associated virus (AAV). Adeno-associated viral (AAV) vectors have become powerful gene delivery tools for the treatment of various disorders. AAV vectors possess a number of features that render them ideally suited for use, including a lack of pathogenicity, minimal immunogenicity, and the ability to transduce post-mitotic cells in a stable and efficient manner. Expression of a particular gene contained within an AAV vector can be specifically targeted to one or more types of cells by choosing the appropriate combination of AAV serotype, promoter, and delivery method.
[0229] In certain embodiments, the vector also includes conventional control elements which are operably linked to the transgene in a manner which permits its transcription, translation and/or expression in a cell transfected with the plasmid vector or infected with the virus produced by the disclosure. As used herein, “operably linked” sequences include both expression control sequences that are contiguous with the gene of interest and expression control sequences that act in trans or at a distance to control the gene of interest. Expression control sequences include appropriate transcription initiation, termination, promoter and enhancer sequences; efficient RNA processing signals such as splicing and polyadenylation (polyA) signals; sequences that stabilize cytoplasmic mRNA; sequences that enhance translation efficiency (i.e., Kozak consensus sequence); sequences that enhance protein stability; and when desired, sequences that enhance secretion of the encoded product. A great number of expression control sequences, including promoters which are native, constitutive, inducible and/or tissuespecific, are known in the art and may be utilized.
[0230] Additional promoter elements, e.g., enhancers, regulate the frequency of transcriptional initiation. Typically, these are located in the region 30-110 bp upstream of the start site, although a number of promoters have recently been shown to contain functional elements downstream of the start site as well. The spacing between promoter elements frequently is flexible, so that promoter function is preserved when elements are inverted or moved relative to one another. In the thymidine kinase (tk) promoter, the spacing between promoter elements can be increased to 50 bp apart before activity begins to decline. Depending on the promoter, it
appears that individual elements can function either cooperatively or independently to activate transcription.
[0231] One example of a suitable promoter is the immediate early cytomegalovirus (CMV) promoter sequence. This promoter sequence is a strong constitutive promoter sequence capable of driving high levels of expression of any polynucleotide sequence operatively linked thereto. Another example of a suitable promoter is Elongation Growth Factor -la (EF-la). However, other constitutive promoter sequences may also be used, including, but not limited to the simian virus 40 (SV40) early promoter, mouse mammary tumor virus (MMTV), human immunodeficiency virus (HIV) long terminal repeat (LTR) promoter, MoMuLV promoter, an avian leukemia virus promoter, an Epstein-Barr virus immediate early promoter, a Rous sarcoma virus promoter, as well as human gene promoters such as, but not limited to, the actin promoter, the myosin promoter, the hemoglobin promoter, and the creatine kinase promoter. Further, the disclosure should not be limited to the use of constitutive promoters. Inducible promoters are also contemplated as part of the disclosure. The use of an inducible promoter provides a molecular switch capable of turning on expression of the polynucleotide sequence which it is operatively linked when such expression is desired, or turning off the expression when expression is not desired. Examples of inducible promoters include, but are not limited to a metallothionine promoter, a glucocorticoid promoter, a progesterone promoter, and a tetracycline promoter.
[0232] Enhancer sequences found on a vector also regulate expression of the gene contained therein. Typically, enhancers are bound with protein factors to enhance the transcription of a gene. An enhancer may be located upstream or downstream of the gene it regulates. Enhancers may also be tissue-specific to enhance transcription in a specific cell or tissue type. In one embodiment, the vector of the present disclosure comprises one or more enhancers to boost transcription of the gene present within the vector.
[0233] In order to assess the expression of E.r. maturase or a E.r. maturase-derived peptide, the expression vector to be introduced into a cell can also comprise either a selectable marker gene or a reporter gene or both to facilitate identification and selection of expressing cells from the population of cells sought to be transfected or infected through viral vectors. In other aspects, the selectable marker may be carried on a separate piece of DNA and used in a cotransfection procedure. Both selectable markers and reporter genes may be flanked with
appropriate regulatory sequences to enable expression in the host cells. Useful selectable markers include, for example, antibiotic-resistance genes, such as neo and the like.
[0234] Reporter genes are used for identifying potentially transfected cells and for evaluating the functionality of regulatory sequences. In general, a reporter gene is a gene that is not present in or expressed by the recipient organism or tissue and that encodes a polypeptide whose expression is manifested by some easily detectable property, e.g., enzymatic activity. Expression of the reporter gene is assayed at a suitable time after the DNA has been introduced into the recipient cells. Suitable reporter genes may include genes encoding luciferase, betagalactosidase, chloramphenicol acetyl transferase, secreted alkaline phosphatase, or the green fluorescent protein gene (e.g., Ui-Tei et al., 2000 FEBS Letters 479: 79-82). Suitable expression systems are well known and may be prepared using known techniques or obtained commercially. In general, the construct with the minimal 5' flanking region showing the highest level of expression of reporter gene is identified as the promoter. Such promoter regions may be linked to a reporter gene and used to evaluate agents for the ability to modulate promoter- driven transcription.
[0235] Methods of introducing and expressing genes into a cell are known in the art. In the context of an expression vector, the vector can be readily introduced into a host cell, e.g., mammalian, bacterial, yeast, or insect cell by any method in the art. For example, the expression vector can be transferred into a host cell by physical, chemical, or biological means.
[0236] Physical methods for introducing a polynucleotide into a host cell include calcium phosphate precipitation, lipofection, particle bombardment, microinjection, electroporation, and the like. Methods for producing cells comprising vectors and/or exogenous nucleic acids are well-known in the art. See, for example, Sambrook et al. (2012, Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Laboratory, New York). A preferred method for the introduction of a polynucleotide into a host cell is calcium phosphate transfection.
[0237] Biological methods for introducing a polynucleotide of interest into a host cell include the use of DNA and RNA vectors. Viral vectors, and especially retroviral vectors, have become the most widely used method for inserting genes into mammalian, e.g., human cells. Other viral vectors can be derived from lentivirus, poxviruses, herpes simplex virus I, adenoviruses and adeno-associated viruses, and the like. See, for example, U.S. Pat. Nos. 5,350,674 and 5,585,362.
[0238] Chemical means for introducing a polynucleotide into a host cell include colloidal dispersion systems, such as macromolecule complexes, nanocapsules, microspheres, beads, and lipid-based systems including oil-in-water emulsions, micelles, mixed micelles, and liposomes. An exemplary colloidal system for use as a delivery vehicle in vitro and in vivo is a liposome (e g., an artificial membrane vesicle).
[0239] In the case where a non-viral delivery system is utilized, an exemplary delivery vehicle is a liposome. The use of lipid formulations is contemplated for the introduction of the nucleic acids into a host cell (in vitro, ex vivo or in vivo). In another aspect, the nucleic acid may be associated with a lipid. The nucleic acid associated with a lipid may be encapsulated in the aqueous interior of a liposome, interspersed within the lipid bilayer of a liposome, attached to a liposome via a linking molecule that is associated with both the liposome and the oligonucleotide, entrapped in a liposome, complexed with a liposome, dispersed in a solution comprising a lipid, mixed with a lipid, combined with a lipid, contained as a suspension in a lipid, contained or complexed with a micelle, or otherwise associated with a lipid. Lipid, lipid/DNA or lipid/expression vector associated compositions are not limited to any particular structure in solution. For example, they may be present in a bilayer structure, as micelles, or with a “collapsed” structure. They may also simply be interspersed in a solution, possibly forming aggregates that are not uniform in size or shape. Lipids are fatty substances which may be naturally occurring or synthetic lipids. For example, lipids include the fatty droplets that naturally occur in the cytoplasm as well as the class of compounds which contain long-chain aliphatic hydrocarbons and their derivatives, such as fatty acids, alcohols, amines, amino alcohols, and aldehydes.
[0240] Lipids suitable for use can be obtained from commercial sources. For example, dimyristyl phosphatidylcholine (“DMPC”) can be obtained from Sigma, St. Louis, MO; dicetyl phosphate (“DCP”) can be obtained from K & K Laboratories (Plainview, NY); cholesterol (“Choi”) can be obtained from Calbiochem-Behring; dimyristyl phosphatidylglycerol (“DMPG”) and other lipids may be obtained from Avanti Polar Lipids, Inc. (Birmingham, AL). Stock solutions of lipids in chloroform or chloroform/methanol can be stored at about -20°C. Chloroform is used as the only solvent since it is more readily evaporated than methanol. “Liposome” is a generic term encompassing a variety of single and multilamellar lipid vehicles formed by the generation of enclosed lipid bilayers or aggregates. Liposomes can be
characterized as having vesicular structures with a phospholipid bilayer membrane and an inner aqueous medium. Multilamellar liposomes have multiple lipid layers separated by aqueous medium. They form spontaneously when phospholipids are suspended in an excess of aqueous solution. The lipid components undergo self-rearrangement before the formation of closed structures and entrap water and dissolved solutes between the lipid bilayers (Ghosh et al., 1991 Glycobiology 5: 505-10). However, compositions that have different structures in solution than the normal vesicular structure are also encompassed. For example, the lipids may assume a micellar structure or merely exist as nonuniform aggregates of lipid molecules. Also contemplated are lipofectamine-nucleic acid complexes.
[0241] Regardless of the method used to introduce exogenous nucleic acids into a host cell, in order to confirm the presence of the recombinant DNA sequence in the host cell, a variety of assays may be performed. Such assays include, for example, “molecular biological” assays well known to those of skill in the art, such as Southern and Northern blotting, RT-PCR and PCR; “biochemical” assays, such as detecting the presence or absence of a particular peptide, e.g., by immunological means (ELISAs and Western blots) or by assays described herein to identify agents falling within the scope of the disclosure.
[0242] In one embodiment, the present disclosure provides a delivery vehicle comprising a reverse transcriptase, or a nucleic acid molecule encoding a reverse transcriptase. Exemplary delivery vehicles include, but are not limited to, microspheres, microparticles, nanoparticles, polymerosomes, liposomes, and micelles. For example, in certain embodiments, the delivery vehicle is loaded with a reverse transcriptase, or a nucleic acid molecule encoding a reverse transcriptase. In certain embodiments, the delivery vehicle provides for controlled release, delayed release, or continual release of its loaded cargo. In certain embodiments, the delivery vehicle comprises a targeting moiety that targets the delivery vehicle to a particular location.
[0243] In one embodiment, the present disclosure provides a full-length cDNA derived from a full-length RNA, produced by a reverse transcriptase described herein. In one embodiment, the RNA has significant secondary or tertiary structure, and/or is long (greater than or equal to 5,000 bases in length). For example, it is described herein that E.r. maturase and E.r. maturase-derived peptides described herein are highly processive reverse transcriptases. In one embodiment, the RNA reverse transcribed into DNA is at least about 100, at least about 200, at least about 300, at least about 400, at least about 500, at least about 600, at least about 700, at
least about 800, at least about 900, at least about 1000, at least about 2000, at least about 3000, at least about 4000, at least about 5000, at least about 6000, at least about 7000, at least about 8000, at least about 9000, or at least about 10000 bases in length. In one embodiment, the DNA so reverse transcribed is at least about 100, at least about 200, at least about 300, at least about 400, at least about 500, at least about 600, at least about 700, at least about 800, at least about 900, at least about 1000, at least about 2000, at least about 3000, at least about 4000, at least about 5000, at least about 6000, at least about 7000, at least about 8000, at least about 9000, or at least about 10000 bases in length.
Molecular Applications
[0244] In some embodiments, the at least one protein tetramer, of the present disclosure, is linked to a solid support, wherein the linkage to the solid support occurs via the interaction of the avidin or streptavidin tetramer with at least one component attached to the solid support, wherein the at least one component attached to the solid support has high binding affinity for the streptavidin tetramer. In some embodiments, the at least one component attached to the solid support comprises biotin. In some embodiments, the solid support is comprised in a device or procedure utilized in molecular applications. In some embodiments, the device or procedure utilized in molecular applications comprises a device or procedure utilized in nucleic acid sequencing applications. In some embodiments, the device or procedure utilized in molecular applications comprises a device or procedure utilized in any molecular application that requires the reverse transcriptase activity of the compositions of the present disclosure. In some embodiments, the device or procedure utilized in molecular applications comprises a device or procedure utilized in any molecular application that requires the catalytic activity of the compositions of the present disclosure.
[0245] In some embodiments, any device or procedure that employs reverse transcription as a method or step can utilize the compositions of the present disclosure. In various embodiments, the compositions of the present disclosure are used to perform reverse transcription as part of an assay. In various embodiments, the assay may be at least one selected from the group RT-PCR, qRT-PCR, capillary electrophoresis (CE) for RNA-structure mapping (such as SHAPE-seq or SHAPE-map, DMS-seq), in-cell sequencing, next-generation RNA
sequencing (RNA-seq), nanopore sequencing, cDNA library synthesis, cDNA synthesis, and a combination thereof.
[0246] In certain aspects, the compositions provide for reverse transcription at physiologic temperatures, or at lower temperatures relative to that required when using non-/:./' maturase-derived reverse transcriptases. In certain instances, the lower temperature of the reverse transcription reaction provides a decreased rate of degradation of the RNA molecule during the reaction, relative to the rate of degradation of an RNA molecule in a reverse transcription reaction that uses a non-E.r maturase-derived reverse transcriptase.
[0247] In one embodiment, the compositions of the present disclosure provide for performing reverse transcription of a long and/or complex RNA molecule. In certain embodiments, the reverse transcriptase exhibits increased stability. In certain embodiments, the reverse transcriptase exhibits reduced turnover. In certain embodiments, the reverse transcriptases described herein have reduced turnover, thereby allowing the synthesis of longer reads and full-length DNA products.
[0248] In one embodiment, the compositions provide for performing a single reaction amplification of RNA, made possible by the true thermocycling ability of the reverse transcriptases described herein. For, example, the thermocycling ability of the reverse transcriptases described herein allows for the amplification of RNA without the need for DNA replication.
[0249] In one embodiment, the compositions provided herein are utilized in a quantitative RT-PCR (qRT-PCR) device or procedure. In qRT-PCR, the formation of PCR products is monitored in each cycle of the PCR. The amplification is usually measured in thermocyclers which have additional devices for measuring fluorescence signals during the amplification reaction. See, for example, U.S. Pat. No. 6,174,670, and U.S. Pat. No. 8,137,616. In one embodiment, the qRT-PCR procedure is carried out using a thermostable improved E.r. maturase enzyme, without a DNA->DNA polymerase.
[0250] In one embodiment, the compositions provided herein are utilized in a capillary electrophoresis (CE) for RNA-structure mapping device or procedure. The application of capillary electrophoresis to RNA structure probing is an important step in increasing the throughput of RNA structure data. Gel electrophoresis typically resolves about a hundred bases of RNA at a time, and hence probing an RNA of several kilobases long might require running
tens to hundreds of gels. Capillary electrophoresis allows the resolution of 300-650 bases from a structure probing experiment and multiple lanes can be run at the same time to increase the throughput of RNA structure probing. The readout of the probing experiment is typically through the reverse transcription of a 5' fluorescently labeled DNA primer that anneals specifically to the RNA of interest. If the RNA is several kilobases long, multiple primers are designed to anneal along the length of the transcript. Modification or cleavage of the RNA template results in premature stops in the primer extension reaction, leading to different lengths of the cDNA product which are resolved by capillary electrophoresis. Software tools such as CAFA and Shapefinder can automate the data acquisition from capillary electrophoresis and further improve speed and accuracy (see, for example, Wan, Y. et al., 2011, Nat Rev Genet., 12(9): 1-26).
[0251] In one embodiment, the compositions provided herein are utilized in a nextgeneration RNA sequencing (RNA-seq) device or procedure. High-throughput RNA sequencing (RNA-Seq) technology, enabled by recent developments in next generation sequencing, has become a powerful tool in analyzing gene expression profiles, detecting transcript variants, and understanding the function of non-coding regulatory RNAs. A standard RNA-Seq library is generated from ligating sequencing adapters to double-stranded DNA. There are two main classes of methods to prepare strand-specific RNA-Seq libraries. The first method comprises ligating different adapters to the 3' and 5' ends of the RNA molecules (see e.g. Ion Total RNA- Seq Kit v2 from Life Technologies). Another, more widely used method comprises incorporating dUTP in addition to dNTPs in the second strand DNA synthesis. Following adapter ligation, the second strand DNA can be specifically digested by an Uracil-N-glycosylase (UNG) enzyme so that only the library strand containing the first strand cDNA will be sequenced and information on the direction of the transcripts can therefore be obtained (see M. Sultan et al., Biochemical and Biophysical Research Communications 422 (2012) 643-646; also see PCT Patent Application Number PCT/EP2016/069997).
[0252] The disclosure is also directed to methods for making one or more nucleic acid molecules and/or labeled nucleic acid molecules, comprising mixing one or more nucleic acid templates (e.g., one or more RNA templates or messenger RNA templates) with one or more compositions of the disclosure having reverse transcriptase activity and incubating the mixture under conditions sufficient to synthesize one or more first nucleic acid molecules complementary to all or a portion of the one or more nucleic acid templates, wherein at least one of the
synthesized molecules are optionally labeled and/or comprise one or more labeled nucleotides and/or wherein the synthesized molecules may optionally be modified to contain one or more labels. In one embodiment, the one or more first nucleic acid molecules are single-stranded cDNA molecules. Nucleic acid templates suitable for reverse transcription according to this aspect of the disclosure include any nucleic acid molecule or population of nucleic acid molecules (e.g., RNA, mRNA), particularly those derived from a cell or tissue. In one aspect, a population of mRNA molecules (a number of different mRNA molecules, typically obtained from cells or tissue) are used to make a labeled cDNA library, in accordance with the disclosure. Exemplary sources of nucleic acid templates include viruses, virally infected cells, bacterial cells, fungal cells, plant cells and animal cells.
[0253] The disclosure also concerns methods for making one or more double-stranded nucleic acid molecules (which may optionally be labeled). Such methods comprise (a) mixing one or more nucleic acid templates (e.g., RNA or mRNA, or a population of mRNA templates) with one or more polypeptides of the disclosure having reverse transcriptase activity; (b) incubating the mixture under conditions sufficient to make one or more first nucleic acid molecules complementary to all or a portion of the one or more templates; and (c) incubating the one or more first nucleic acid molecules under conditions sufficient to make one or more second nucleic acid molecules complementary to all or a portion of the one or more first nucleic acid molecules, thereby forming one or more double-stranded nucleic acid molecules comprising the first and second nucleic acid molecules. In accordance with the disclosure, the first and/or second nucleic acid molecules may be labeled (e.g., may comprise one or more of the same or different labeled nucleotides and/or may be modified to contain one or more of the same or different labels). Thus, labeled nucleotides may be used at one or both synthesis steps. Such methods may include the use of one or more DNA polymerases as part of the process of making the one or more double-stranded nucleic acid molecules. The disclosure also concerns compositions useful for making such double-stranded nucleic acid molecules. Such compositions comprise one or more reverse transcriptases of the disclosure and optionally one or more DNA polymerases, a suitable buffer and/or one or more nucleotides (e.g., including labeled nucleotides).
[0254] The disclosure is also directed to nucleic acid molecules and/or labeled nucleic acid molecules (particularly single- or double-stranded cDNA molecules) produced according to the above-described methods and to kits comprising these nucleic acid molecules. Such
molecules or kits may be used to detect nucleic acid molecules (for example by hybridization) or for diagnostic purposes.
Methods for generation of protein tetramer
[0255] To produce the streptavidin tetramer that contains three or less streptavidin fusion constructs of the disclosure, the streptavidin fusion construct can be coexpressed with the streptavidin monomer within the same host cell from their respective expression systems, allowing the streptavidin monomer to interact with the streptavidin moiety of the fusion protein, resulting in the formation of a tetramer, in which the presence of streptavidin monomer reduces the number of streptavidin fusion constructs in the tetramer. By adjusting the relative expression levels of the streptavidin monomer to the streptavidin fusion construct, a streptavidin tetramer containing three, two or one streptavidin fusion construct can be created.
[0256] In some embodiments, the protein multimer of the present disclosure comprises one streptavidin fusion construct of the disclosure in a tetrameric complex with three additional streptavidin monomers.
[0257] In some embodiments, a bacterial expression system is utilized to generate the tetrameric complex. In some embodiments, the bacterial expression system is a co-expression system.
[0258] In some embodiments, the bacterial co-expression system comprises at least one bacterial cell comprising at least one nucleic acid molecule encoding the streptavidin fusion construct of the present disclosure and the streptavidin monomer. In some embodiments, the at least one nucleic acid molecule is a plasmid. In some embodiments, the streptavidin fusion construct and the streptavidin monomer are operably connected to different promoter sequences, wherein the different promoter sequences lead to different levels of expression of the streptavidin fusion construct and the streptavidin monomer, wherein the different levels of expression lead to the formation of the tetramer comprising one streptavidin fusion molecule and three streptavidin monomers. In some embodiments, the streptavidin fusion construct is operably connected to the tac promoter, wherein the tac promoter comprises the sequence set forth in SEQ ID NO: 14 and the streptavidin monomer is operably connected to the araBAD promoter, wherein the araBAD promoter comprises the sequence set forth in SEQ ID NO: 15.
[0259] In some embodiments, the bacterial co-expression system comprises at least one bacterial cell comprising distinct nucleic acid molecules encoding the avidin or streptavidin fusion construct and the avidin or streptavidin monomer. In some embodiments, the distinct nucleic acid molecules are plasmids. In some embodiments, the avidin or streptavidin fusion construct and the avidin or streptavidin monomer encoded on distinct plasmids are each operably connected to the same promoter sequence. In some embodiments, the promoter sequence comprises the T7 promoter, wherein the T7 promoter comprises the sequence set forth in SEQ ID NO: 16. In some embodiments, the distinct plasmids encoding the avidin or streptavidin fusion construct and the avidin or streptavidin monomer are present in the at least one bacterial cell in different ratios, wherein the different ratios lead to different levels of expression of the fusion construct and the monomer, wherein the different levels of expression lead to the formation of the tetramer comprising one fusion molecule and three monomers. In some embodiments, the ratio comprises one plasmid encoding the fusion construct and two plasmids encoding the monomer (1 :2). In some embodiments, the ratio comprises one plasmid encoding the fusion construct and three plasmids encoding the monomer (1 :3). In some embodiments, the ratio comprises one plasmid encoding the fusion construct and four plasmids encoding the monomer (1 :4). In some embodiments, the ratio comprises one plasmid encoding the fusion construct and five plasmids encoding the monomer (1 :5). In some embodiments, the ratio comprises one plasmid encoding the fusion construct and 10 or more plasmids encoding the monomer (< 1: 10).
[0260] In some embodiments, the plasmids used are low copy number plasmids, such as pl5A ori, wherein there are approximately 10 copies of the plasmids per cell. In some embodiments, the plasmids used are high copy number plasmids, such as pUC18, wherein there are approximately 300-600 copies of the plasmids per cell.
[0261] In some embodiments, the distinct plasmids encoding the fusion construct and the monomer are present in the at least one bacterial cell in equal ratios. In some embodiments, the fusion construct and the monomer are operably connected to different promoter sequences, wherein the different promoter sequences lead to different levels of expression of the fusion construct and the monomer, wherein the different levels of expression lead to the formation of the tetramer comprising one fusion molecule and three monomers. In some embodiments, the fusion construct is operably connected to the tac promoter (SEQ ID NO: 14) and the monomer is operably connected the araBAD promoter (SEQ ID NO: 15).
Kits
[0262] The disclosure is also directed to kits for use in the production of the compositions of the present disclosure. In various embodiments, the present disclosure provides a kit to produce the compositions comprising the at least one protein multimer.
[0263] In one embodiment, the kit comprises at least one fusion construct of the present disclosure and/or a nucleic acid molecule encoding the same.
[0264] In one embodiment, the kit comprises at least one protein multimer, wherein the at least one protein multimer comprises at least one avidin or streptavidin fusion construct of the disclosure. In some embodiments, the protein multimer comprises an avidin or streptavidin tetramer comprising at least one avidin or streptavidin fusion construct of the disclosure. In some embodiments, the protein multimer comprises an avidin or streptavidin tetramer comprising at least two avidin or streptavidin fusion constructs of the disclosure. In some embodiments, the protein multimer comprises an avidin or streptavidin tetramer comprising at least three avidin or streptavidin fusion constructs of the disclosure.
[0265] In one embodiment, the kit comprises an expression system that comprises one or more a polynucleotide encoding a fusion construct and a monomer of the disclosure. In one embodiment, the kit comprises an expression system that comprises a first polynucleotide encoding a fusion construct of the disclosure and a second polynucleotide encoding an avidin or streptavidin monomer.
[0266] In one embodiment, the kit comprises an expression system that comprises a polynucleotide comprising or encoding a nucleic acid molecule that reduces non-specific binding. In one embodiment, the kit comprises an expression system that comprises a polynucleotide encoding a protein or carbohydrate (e g., heparin) that reduces non-specific binding.
[0267] In one embodiment, the kit includes instructional material that describes the use of the kit to produce the compositions of the present disclosure, wherein the instructional material creates an increased functional relationship between the kit components and the individual using the kit. In one embodiment, the kit is utilized by one person or entity. In another embodiment, the kit is utilized by more than one person or entity. In one embodiment, the kit is
used without any additional compositions or methods. In another embodiment, the kit is used with at least one additional composition or method.
[0268] The disclosure is also directed to kits for use in the methods of the disclosure. Such kits can be used for making nucleic acid molecules and/or labeled nucleic acid molecules (single- or double-stranded). Kits of the disclosure may comprise a carrier, such as a box or carton, having in close confinement therein one or more containers, such as vials, tubes, bottles and the like. In kits of the disclosure, a first container may contain one or more of the fusion constructs of the disclosure or one or more of the compositions of the disclosure. Kits of the disclosure may also comprise, in the same or different containers, at least one component selected from one or more DNA polymerases (e.g., thermostable DNA polymerases), a suitable buffer for nucleic acid synthesis and one or more nucleotides. In one embodiment, kits of the disclosure may also comprise, in the same or different containers, an agent that reduces nonspecific binding of primers to the surface of E.r. maturase or variant thereof. In one embodiment, kits of the disclosure may also comprise, in the same or different containers, an optimized reaction buffer as described elsewhere herein, or components used to produce the optimized reaction buffer. Alternatively, the components of the kit may be divided into separate containers.
[0269] The disclosure is also directed to kits for use in methods of the disclosure. Such kits can be used for making, sequencing or amplifying nucleic acid molecules (single- or doublestranded), e.g., at the particular temperatures described herein. Kits of the disclosure may comprise a carrier, such as a box or carton, having in close confinement therein one or more (e.g., one, two, three, four, five, ten, twelve, fifteen, etc.) containers, such as vials, tubes, bottles and the like. In kits of the disclosure, a first container contains one or more of the reverse transcriptase enzymes of the present disclosure. Kits of the disclosure may also comprise, in the same or different containers, one or more DNA polymerases (e.g., thermostable DNA polymerases), one or more (e.g., one, two, three, four, five, ten, twelve, fifteen, etc.) suitable buffers for nucleic acid synthesis, one or more nucleotides and one or more (e.g., one, two, three, four, five, ten, twelve, fifteen, etc.) oligonucleotide primers. In one embodiment, kits of the disclosure may also comprise, in the same or different containers, an agent that reduces nonspecific binding of primers to the surface of E.r. maturase or variant thereof, as described elsewhere herein. In one embodiment, kits of the disclosure may also comprise, in the same or different containers, an optimized reaction buffer as described elsewhere herein, or components
used to produce the optimized reaction buffer. Alternatively, the components of the kit may be divided into separate containers (e.g., one container for each enzyme and/or component). Kits of the disclosure also may comprise instructions or protocols for carrying out the methods of the disclosure.
[0270] In various embodiments, the present disclosure provides a kit to use the compositions of the present disclosure, in a reverse transcription reaction. In one embodiment, the kit comprises a composition of the present disclosure. In one embodiment, the kit includes instructional material that describes the use of the kit to use the compositions of the present disclosure in a reverse transcription reaction, wherein the instructional material creates an increased functional relationship between the kit components and the individual using the kit. In one embodiment, the kit is utilized by one person or entity. In another embodiment, the kit is utilized by more than one person or entity. In one embodiment, the kit is used without any additional compositions or methods. In another embodiment, the kit is used with at least one additional composition or method.
ENUMERATED EMBODIMENTS
[0271] 1. A fusion construct comprising an affinity domain operably linked to a polymerase domain.
[0272] 2. The fusion construct of embodiment 1, wherein the polymerase domain comprises a DNA polymerase, an RNA polymerase, or a reverse transcriptase.
[0273] 3. The fusion construct of any of the preceding embodiments, wherein the polymerase domain comprises a reverse transcriptase, e.g., a retroviral reverse transcriptase or a group II intron reverse transcriptase.
[0274] 4. The fusion construct of any of the preceding embodiments, wherein the polymerase domain comprises a group II intron reverse transcriptase, e.g., a MarathonRT or UltraMarathonRT reverse transcriptase, or a fragment or variant thereof.
[0275] 5. The fusion construct of any of the preceding embodiments, wherein the polymerase domain comprises MarathonRT or a fragment or variant thereof.
[0276] 6. The fusion construct of any of the preceding embodiments, wherein the polymerase domain comprises UltraMarathonRT or a fragment or variant thereof.
[0277] 7. The fusion construct of any of the preceding embodiments, wherein the polymerase domain comprises SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22, SEQ ID NO: 23, SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, or SEQ ID NO: 29, or a fragment or variant thereof.
[0278] 8. The fusion construct of any of the preceding embodiments, wherein the polymerase domain comprises SEQ ID NO: 3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, or SEQ ID NO: 12 or a fragment or variant thereof.
[0279] 9. A fusion construct comprising avidin or streptavidin operably linked to a functional protein or enzyme domain, wherein the avidin or streptavidin can form a tetramer with additional avidin or streptavidin molecules or additional avidin or streptavidin fusion enzymes.
[0280] 10. The fusion construct of any of the preceding embodiments, wherein the construct comprises streptavidin comprising an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO: 1.
[0281] 11. The fusion construct of any of the preceding embodiments, wherein the functional protein or enzyme domain comprises a polymerase, reverse transcriptase, helicase, receptor protein, ligase, kinase, phosphatase, exonuclease and non-specific endonuclease, restriction endonuclease, DNA/RNA methyl transferase, recombinase, terminal transferase, nucleic acid binding protein, protease, ribosomal protein, aminoacyl-tRNA synthetase, glycosyltransferase, fatty acid synthase, or an enzyme for food, biofuel or pharmaceutical industries.
[0282] 12. The fusion construct of any of the preceding embodiments, wherein the functional protein or enzyme domain comprises Marathon Reverse Transcriptase (MRT), or a variant thereof.
[0283] 13. The fusion construct of any of the preceding embodiments, wherein the functional protein or enzyme domain comprises an amino acid sequence at least 90% identical to an amino acid sequence selected from the group consisting SEQ ID NOs: 3 and 8-12 or a functional fragment thereof.
[0284] 14. The fusion construct of any of the preceding embodiments, comprising an amino acid sequence as set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO: 7.
[0285] 15. A nucleic acid molecule encoding a fusion construct comprising avidin or streptavidin operably linked to a functional protein or enzyme domain, wherein the streptavidin can form a tetramer with additional streptavidin molecules or additional streptavidin fusion enzymes.
[0286] 16. The nucleic acid molecule of embodiment 15, encoding a streptavidin fusion construct, wherein the streptavidin comprises an amino acid sequence at least 90% identical to the sequence set forth in SEQ ID NO: 1.
[0287] 17. The nucleic acid molecule of any of embodiments 15-16, wherein the functional protein or enzyme domain comprises a polymerase, reverse transcriptase, helicase, receptor protein, ligase, kinase, phosphatase, exonuclease and non-specific endonuclease, restriction endonuclease, DNA/RNA methyl transferase, recombinase, terminal transferase, nucleic acid binding protein, protease, ribosomal protein, aminoacyl-tRNA synthetase, glycosyltransferase, fatty acid synthase, or an enzyme for food, biofuel or pharmaceutical industries.
[0288] 18. The nucleic acid molecule of any of embodiments 15-17, wherein the functional enzyme domain comprises MRT, or a variant thereof.
[0289] 19. The nucleic acid molecule of any of embodiments 15-18, wherein the functional enzyme domain comprises an amino acid sequence at least 90% identical to an amino acid sequence selected from the group consisting SEQ ID NOs: 3 and 8-12 or a functional fragment thereof.
[0290] 20. The nucleic acid molecule of any of embodiments 15-19, wherein the streptavidin fusion enzyme comprises an amino acid sequence as set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
[0291] 21. The nucleic acid molecule of any of embodiments 15-20, comprising the nucleotide sequence set forth in SEQ ID NO: 17.
[0292] 22. A composition comprising at least one fusion construct of embodiment 1, and further wherein the functional enzyme domain maintains at least one wild type activity of the enzyme when the avidin or streptavidin is complexed as a tetramer.
[0293] 23. The composition of embodiment 22, comprising a tetramer comprising at least one avidin or streptavidin fusion construct of embodiment 1.
[0294] 24. The composition of any of embodiments 22-23, comprising a tetramer comprising at least two avidin or streptavidin fusion constructs of embodiment 1.
[0295] 25. The composition of embodiments 22-24, comprising a tetramer comprising at least three avidin or streptavidin fusion constructs of embodiment 1.
[0296] 26. The composition of embodiments 22-25, wherein the tetramer is linked or bound to a solid support.
[0297] 27. The composition of embodiments 22-26, wherein the solid support is coated with or attached to at least one biotin molecule.
[0298] 28. The composition of embodiments 22-27, wherein the solid support is comprised in at least one device.
[0299] 29. The composition of embodiments 22-28, wherein the at least one device comprises a device used for complementary DNA (cDNA) synthesis.
[0300] 30. The composition of embodiments 22-29, wherein the at least one device is used for nucleic acid synthesis or sequencing.
[0301] 31. A method for producing cDNA, comprising contacting at least one mRNA molecule with the fusion construct of any one of embodiments 9-14, or a composition of any one of embodiments 22-30, under conditions sufficient for the synthesis of at least one nucleic acid molecule complementary to all or a portion of the at least one mRNA molecule.
[0302] 32. A method for determining the identity of each of a series of consecutive nucleotide residues of at least one target nucleic acid, comprising contacting the at least one target nucleic acid with the fusion construct of any one of embodiments 9-14, or a composition of any one of embodiments 22-30, under conditions sufficient for the synthesis of at least one nucleic acid molecule complementary to all or a portion of the at least one target nucleic acid, and for the determination of each nucleotide in the at least one complementary nucleic acid.
EXPERIMENTAL EXAMPLES
[0303] The disclosure is further described in detail by reference to the following experimental examples. These examples are provided for purposes of illustration only, and are not intended to be limiting unless otherwise specified. Thus, the disclosure should in no way be construed as being limited to the following examples, but rather, should be construed to encompass any and all variations which become evident as a result of the teaching provided herein.
[0304] Without further description, it is believed that one of ordinary skill in the art can, using the preceding description and the following illustrative examples, make and utilize the present disclosure and practice the claimed methods. The following working examples therefore are not to be construed as limiting in any way the remainder of the disclosure.
Example 1 : Production of catalytically functional, affinity tagged-reverse-transcriptase fusion proteins
[0305] Streptavidin is one of the most important protein conjugation reagents, as it facilitates bridging of molecules and anchoring to biotinylated surfaces or nanoparticles. However, its tetrameric structure makes it impractical for creating single-enzyme fusion constructs. As a result, post-expression, chemically-conjugated streptavidin proteins are more commonly employed for affinity tagging of proteins with streptavidin.
[0306] In this study, a tetrameric streptavidin-enzyme fusion that maintains strong biotin binding without losing activity of the enzyme (Marathon Reverse Transcriptase, MRT) due to streptavidin tetramerization was designed.
[0307] In this study, a single position on a reverse transcriptase (RT), in this case, Marathon Reverse Transcriptase, MRT, was functionalized without losing enzymatic and processivity properties.
[0308] First, a fusion construct, STV-FL15-MRT, (SEQ ID NO:4) that links streptavidin, STV, (SEQ ID NO: 1) and the N-terminus of MRT (SEQ ID NO:3), connected by a flexible GGGGSGGGGSGGGGS linker, FL15, (SEQ ID NO:2) was created (Figure 1 and Figure 2). The expression level of STV and STV-FL15-MRT fusion protein can be individually adjusted. Figure 1 is a representative illustration of streptavidin tetramer containing one MarathonRT domain.
[0309] Native streptavidin and streptavidin-MarathonRT fusion protein were coexpressed in E. coli in proportions that enabled three native streptavidin molecules (SEQ ID NO:1) to interact with one streptavidin moiety of the fusion protein (SEQ ID NO:4), resulting in the formation of a streptavidin tetramer that is only linked to one MRT (Figure 1 and Figure 2). This reduces the number of MRT moi eties incorporated, and thereby preserves its catalytic function. This was accomplished by adjusting the relative expression levels of the native streptavidin to the STV-FL15-MRT fusion protein, resulting in streptavidin tetramers containing
three, two or one MRT domain, which help minimize or eliminate steric hindrance between MRT domains. These were then tested and the optimal ratio was determined.
[0310] MRT (or any other RT) was functionalized at its N-terminus with another protein (such as an affinity tag like streptavidin) while still maintaining its full enzymatic activities in terms of processive, high speed cDNA synthesis, RNA unwinding and dNTP hydrolysis (Figure 3-Figure 5). The long, flexible linker (SEQ ID NO:2) between STV (SEQ ID NO: 1) and MRT (SEQ ID NO:3) greatly improved the enzyme activity of the fusion protein; reducing the number of MRT domains further improved the enzyme activity (Figure 5).
[0311] The problem of streptavidin tetramerization was overcome by co-expressing full- length fusion protein with monomeric streptavidin in the correct proportions to yield an active tetramer (Figure 1 -Figure 5).
[0312] Tetrameric STV-MRT fusion protein without a linker (SEQ ID NO:6) or with a short linker (SEQ ID NO:5) is nearly inactive (Figure 6-Figure 8).
[0313] This demonstrated controlled expression levels of streptavidin and streptavidin- MarathonRT fusion protein, respectively, resulting in different types of active fusion constructs that are useful for different purposes, as the number of MarathonRT proteins in the tetramer can be adjusted from 3 to 1 as needed.
[0314] By connecting an RT enzyme (such as MarathonRT) via a flexible linker with an affinity tag (such as streptavidin), two protein moieties folded independently during expression without damaging protein stability. In addition to providing a tagged RT, the process for creating the streptavidin fusion construct enabled scale-up and high yield of multimeric protein expression. This same approach/ strategy is applicable to any other protein of interest that is linked to streptavidin.
Example 2: Streptavidin-MarathonRT fusion protein is used in devices for direct RNA and DNA sequencing
[0315] By creating a functional affinity fusion construct of MarathonRT (or any other polymerase or helicase motor enzyme), utility is greatly expanded, enabling the incorporation of the enzyme into devices for DNA and RNA sequencing, RT-PCR on a solid support, RT-PCR that is accompanied by visualization, such as in spatial transcriptomics applications, long-read
sequencing library preparation methods that maintain long RNA integrity, and improved singlestranded DNA production by reusing the immobilized RT enzyme.
[0316] The streptavidin-MarathonRT fusion protein is used in devices for direct RNA and DNA sequencing, resulting in new types of commercial products. With the chemical matter and methods described above, affinity-tagged RT proteins are generated more cheaply because they can be created as overexpressed single-chain proteins instead of chemically-conjugated proteins.
[0317] The fusions described above enable active RTs to be put on a solid-support, enabling new types of devices such as flow-RT-PCR machines, or methods that require removal of an RT protein on a bead from a reaction, which make many processes faster because the protein does not need to be deactivated. RTs mounted on a solid support in an industrial setting are re-used and recycled, which has great economic benefit.
Example 3: Sequences of the present disclosure
[0318] The disclosures of each and every patent, patent application, and publication cited herein are hereby incorporated herein by reference in their entirety. While this disclosure has been disclosed with reference to specific embodiments, it is apparent that other embodiments and variations of this disclosure may be devised by others skilled in the art without departing from the true spirit and scope of the disclosure. The appended claims are intended to be construed to include all such embodiments and equivalent variations.
Claims
1. A fusion construct comprising an affinity domain operably linked to a polymerase domain, wherein the polymerase domain comprises a DNA polymerase, an RNA polymerase, or a reverse transcriptase, e.g., a group II intron reverse transcriptase, or a fragment or variant thereof.
2. The fusion construct of claim 1, wherein the polymerase domain comprises a group II intron reverse transcriptase, e.g., a MarathonRT or UltraMarathonRT reverse transcriptase, or a fragment or variant thereof.
3. The fusion construct of any of claims 1-2, wherein the polymerase domain comprises MarathonRT or a fragment or variant thereof.
4. The fusion construct of any of claims 1-3, wherein the polymerase domain comprises UltraMarathonRT or a fragment or variant thereof.
5. The fusion construct of claim 1, wherein the polymerase domain comprises SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22, SEQ ID NO: 23, SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, or SEQ ID NO: 29, or a fragment or variant thereof.
6. The fusion construct of any of claims 1-4, wherein the polymerase domain comprises SEQ ID NO: 3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO:
11, or SEQ ID NO: 12 or a fragment or variant thereof.
7. The fusion construct of any of claims 1-6, wherein the polymerase domain comprises an amino acid sequence at least 80% identical, e.g., at least 81%, 82%, 83%, 84%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.9% identical, to SEQ ID NO: 3, SEQ
ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11 , or SEQ ID NO: 12 or a fragment thereof.
8. The fusion construct of any of claims 1-7, wherein the polymerase domain comprises an amino acid sequence at least 90% identical, e.g., at least 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 99.9% identical, to SEQ ID NO: 3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, or SEQ ID NO: 12 or a fragment thereof.
9. The fusion construct of any of claims 1-8, wherein the polymerase domain comprises at least 1 amino acid substitution, e.g., at least 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 amino acid substitutions, relative to SEQ ID NO: 3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, or SEQ ID NO: 12 or a fragment thereof.
10. The fusion construct of any of the preceding claims, wherein the affinity domain and the polymerase domain are separated by a linker (e.g., an amino acid linker).
11. The fusion construct of any of the preceding claims, wherein the linker has a sequence of SEQ ID NO: 2 or SEQ ID NO: 5.
12. The fusion construct of any of the preceding claims, wherein the linker comprises an amino acid sequence at least 80% identical, e.g., at least 81%, 82%, 83%, 84%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.9% identical, to SEQ ID NO: 2.
13. The fusion construct of any of the preceding claims, wherein the linker comprises an amino acid sequence at least 80% identical, e.g., at least 81%, 82%, 83%, 84%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.9% identical, to SEQ ID NO: 5.
14. A fusion construct comprising avidin or streptavidin operably linked to an enzymatic domain, wherein the avidin or streptavidin can form a tetramer with additional avidin or streptavidin molecules or additional avidin or streptavidin fusion enzymes.
15. The fusion construct of claim 14, wherein the construct comprises streptavidin comprising an amino acid sequence at least 90% identical, e.g., 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.9% identical, to the sequence set forth in SEQ ID NO: 1.
16. The fusion construct of any of claims 14-15, wherein the construct comprises streptavidin comprising at least 1, 2, 3, 4, 5, 6, 7, 8, or more amino acid substitutions relative to the sequence set forth in SEQ ID NO: 1.
17. The fusion construct of any of claims 14-16, wherein the construct comprises avidin comprising an amino acid sequence at least 90% identical, e.g., 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, 99.9% identical, to the sequence set forth in SEQ ID NO: 18.
18. The fusion construct of any of claims 14-17, wherein the construct comprises avidin comprising at least 1, 2, 3, 4, 5, 6, 7, 8, or more amino acid substitutions relative to the sequence set forth in SEQ ID NO: 18.
19. The fusion construct of any of claims 14-18, wherein the functional enzymatic domain comprises a polymerase, reverse transcriptase, helicase, receptor protein, ligase, kinase, phosphatase, exonuclease and non-specific endonuclease, restriction endonuclease, DNA/RNA methyl transferase, recombinase, terminal transferase, nucleic acid binding protein, protease, ribosomal protein, aminoacyl-tRNA synthetase, glycosyltransferase, fatty acid synthase, or an enzyme for food, biofuel or pharmaceutical industries.
20. The fusion construct of any of claims 14-19, wherein the enzymatic domain comprises a polymerase, e.g., a DNA polymerase, RNA polymerase, or a reverse transcriptase.
21 . The fusion construct of any of claims 14-20, wherein the enzymatic domain comprises a reverse transcriptase, e.g., a group II intron reverse transcriptase, or a fragment or variant thereof.
22. The fusion construct of any of claims 14-21, wherein the enzymatic domain comprises a group II intron reverse transcriptase, e.g., a MarathonRT or UltraMarathonRT reverse transcriptase, or a fragment or variant thereof.
23. The fusion construct of any of claims 14-22, wherein the enzymatic domain comprises MarathonRT or a fragment or variant thereof.
24. The fusion construct of any of claims 14-23, wherein the enzymatic domain comprises UltraMarathonRT or a fragment or variant thereof.
25. The fusion construct of claim 14, wherein the enzymatic domain comprises SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22, SEQ ID NO: 23, SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, or SEQ ID NO: 29, or a fragment or variant thereof.
26. The fusion construct of any of claims 14-24 wherein the enzymatic domain comprises SEQ ID NO: 3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, or SEQ ID NO: 12 or a fragment or variant thereof.
27. The fusion construct of any of claims 14-26, wherein the enzymatic domain comprises an amino acid sequence at least 80% identical, e.g., at least 81%, 82%, 83%, 84%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.9% identical, to SEQ ID NO: 3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, or SEQ ID NO: 12 or a fragment thereof.
28. The fusion construct of any of claims 14-27, wherein the enzymatic domain comprises an amino acid sequence at least 90% identical, e.g., at least 91%, 92%, 93%,
94%, 95%, 96%, 97%, 98%, 99%, or 99.9% identical, to SEQ ID NO: 3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, or SEQ ID NO: 12 or a fragment thereof.
29. The fusion construct of any of claims 14-28, wherein the enzymatic domain comprises at least 1 amino acid substitution, e.g., at least 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 amino acid substitutions, relative to SEQ ID NO: 3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, or SEQ ID NO: 12 or a fragment thereof.
30. The fusion construct of any of the preceding claims, comprising an amino acid sequence as set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
31. A nucleic acid molecule encoding a fusion construct comprising avidin or streptavidin operably linked to an enzymatic domain, wherein the streptavidin can form a tetramer with additional streptavidin molecules or additional streptavidin fusion enzymes.
32. The nucleic acid molecule of claim 31, encoding a streptavidin fusion construct, wherein the streptavidin comprises an amino acid sequence at least 90% identical, e.g., at least 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 99.9% identical, to the sequence set forth in SEQ ID NO: 1.
33. The nucleic acid molecule of any of claims 31-32, wherein the enzymatic domain comprises a polymerase, reverse transcriptase, helicase, receptor protein, ligase, kinase, phosphatase, exonuclease and non-specific endonuclease, restriction endonuclease, DNA/RNA methyl transferase, recombinase, terminal transferase, nucleic acid binding protein, protease, ribosomal protein, aminoacyl-tRNA synthetase, glycosyltransferase, fatty acid synthase, or an enzyme for food, biofuel or pharmaceutical industries.
34. The nucleic acid molecule of any of claims 31-33, wherein the enzymatic domain comprises a polymerase, e.g., a DNA polymerase, RNA polymerase, or a reverse transcriptase.
35. The nucleic acid molecule of any of claims 31-34, wherein the enzymatic domain comprises a reverse transcriptase, e.g., a group II intron reverse transcriptase, or a fragment or variant thereof.
36. The nucleic acid molecule of any of claims 31-35, wherein the enzymatic domain comprises a group II intron reverse transcriptase, e.g., a MarathonRT or UltraMarathonRT reverse transcriptase, or a fragment or variant thereof.
37. The nucleic acid molecule of any of claims 31-36, wherein the enzymatic domain comprises MarathonRT or a fragment or variant thereof.
38. The nucleic acid molecule of any of claims 31-37, wherein the enzymatic domain comprises UltraMarathonRT or a fragment or variant thereof.
39. The nucleic acid molecule of claim 31, wherein the enzymatic domain comprises SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 21, SEQ ID NO: 22, SEQ ID NO: 23, SEQ ID NO: 24, SEQ ID NO: 25, SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, or SEQ ID NO: 29, or a fragment or variant thereof.
40. The nucleic acid molecule of any of claims 31-38 wherein the enzymatic domain comprises SEQ ID NO: 3, SEQ ID NO: 8, SEQ ID NO: 9, SEQ ID NO: 10, SEQ ID NO: 11, or SEQ ID NO: 12 or a fragment or variant thereof.
41. The nucleic acid molecule of any of claims 31-40, wherein the fusion construct comprises an amino acid sequence as set forth in SEQ ID NO:4, SEQ ID NO:6 or SEQ ID NO:7.
42. The nucleic acid molecule of any of claims 31-41, comprising the nucleotide sequence set forth in SEQ ID NO: 17.
43. A composition comprising at least one fusion construct of claim 1, wherein the enzymatic domain maintains at least one wild type activity of the enzyme when the avidin or streptavidin is complexed as a tetramer.
44. The composition of claim 43, comprising a tetramer comprising at least one avidin or streptavidin fusion construct of claim 1.
45. The composition of any of claims 43-44, comprising a tetramer comprising at least two avidin or streptavidin fusion constructs of claim 1.
46. The composition of any of claims 43-45, comprising a tetramer comprising at least three avidin or streptavidin fusion constructs of claim 1.
47. The composition of any of claims 43-46, wherein the tetramer is linked or bound to a solid support.
48. The composition of any of claims 43-47, wherein the solid support is coated with or attached to at least one biotin molecule.
49. The composition of any of claims 43-48, wherein the solid support is comprised in at least one device.
50. The composition of any of claims 43-49, wherein the at least one device comprises a device used for complementary DNA (cDNA) synthesis.
51. The composition of any of claims 43-50, wherein the at least one device is used for nucleic acid synthesis or sequencing.
52. A method for producing cDNA, comprising contacting at least one mRNA molecule with the fusion construct of any one of claims 1-30, or a composition of any
one of claims 43-51 , under conditions sufficient for the synthesis of at least one nucleic acid molecule complementary to all or a portion of the at least one mRNA molecule.
53. A method for determining the identity of each of a series of consecutive nucleotide residues of at least one target nucleic acid, comprising contacting the at least one target nucleic acid with the fusion construct of any one of claims 1-30, or a composition of any one of claims 43-51, under conditions sufficient for the synthesis of at least one nucleic acid molecule complementary to all or a portion of the at least one target nucleic acid, and for the determination of each nucleotide in the at least one complementary nucleic acid.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363599119P | 2023-11-15 | 2023-11-15 | |
| US63/599,119 | 2023-11-15 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025106766A1 true WO2025106766A1 (en) | 2025-05-22 |
Family
ID=95743535
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/056055 Pending WO2025106766A1 (en) | 2023-11-15 | 2024-11-15 | Fusion-modified enzyme constructs |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025106766A1 (en) |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20120214200A1 (en) * | 2010-08-30 | 2012-08-23 | Adelbert Grossmann | Prokaryotic expression construct |
| US20190352626A1 (en) * | 2017-01-30 | 2019-11-21 | KWS SAAT SE & Co. KGaA | Repair template linkage to endonucleases for genome engineering |
| US20210292769A1 (en) * | 2020-03-19 | 2021-09-23 | Evolve Biotech, Inc. | Methods and compositions for directed genome editing |
-
2024
- 2024-11-15 WO PCT/US2024/056055 patent/WO2025106766A1/en active Pending
Patent Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20120214200A1 (en) * | 2010-08-30 | 2012-08-23 | Adelbert Grossmann | Prokaryotic expression construct |
| US20190352626A1 (en) * | 2017-01-30 | 2019-11-21 | KWS SAAT SE & Co. KGaA | Repair template linkage to endonucleases for genome engineering |
| US20210292769A1 (en) * | 2020-03-19 | 2021-09-23 | Evolve Biotech, Inc. | Methods and compositions for directed genome editing |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US12031126B2 (en) | Methods and compositions for simultaneous editing of both strands of a target double-stranded nucleotide sequence | |
| US12509680B2 (en) | Methods and compositions for prime editing nucleotide sequences | |
| CN113373130B (en) | Cas12 protein, gene editing system containing Cas12 protein and application | |
| US20190352639A1 (en) | Genome editing reagents and their use | |
| US20230357733A1 (en) | Reverse Transcriptase and Methods of Use | |
| US20200140835A1 (en) | Engineered CRISPR-Cas9 Nucleases | |
| JP2022122910A (en) | An engineered Cas9 system for eukaryotic genome modification | |
| AU2018321105B2 (en) | Improved transposase polypeptide and uses thereof | |
| EP4133069A2 (en) | Compositions and methods for improved site-specific modification | |
| CA3225808A1 (en) | Context-specific adenine base editors and uses thereof | |
| WO2024251229A9 (en) | Cas enzyme and system and use thereof | |
| EP4534660A1 (en) | Engineered cas12f protein with expanded targetable range and uses thereof | |
| WO2025106766A1 (en) | Fusion-modified enzyme constructs | |
| US20220145332A1 (en) | Cell penetrating transposase | |
| WO2024059719A2 (en) | Compositions for preventing repetitive addition of switching oligonucleotides and nonspecific primer extension during cdna synthesis and methods of use thereof | |
| EA049378B1 (en) | METHODS AND COMPOSITIONS FOR EDITING NUCLEOTIDE SEQUENCES | |
| JP2025508794A (en) | Fusion proteins | |
| CN113403360A (en) | Hydrophobic interface-based in-vitro cell-free protein synthesis method, D2P kit and related application | |
| HK40104618A (en) | Compositions comprising a variant polypeptide and uses thereof | |
| BR122025004697B1 (en) | Systems for Modifying a Eukaryotic Genome, Nucleic Acid Plurality, Vector, and Fusion Protein |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24892287 Country of ref document: EP Kind code of ref document: A1 |