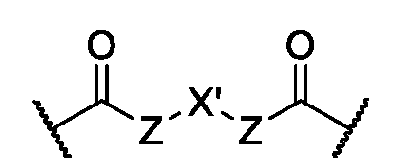WO2025101535A1 - Biomolecule cleanup methods, kits, and systems - Google Patents
Biomolecule cleanup methods, kits, and systems Download PDFInfo
- Publication number
- WO2025101535A1 WO2025101535A1 PCT/US2024/054605 US2024054605W WO2025101535A1 WO 2025101535 A1 WO2025101535 A1 WO 2025101535A1 US 2024054605 W US2024054605 W US 2024054605W WO 2025101535 A1 WO2025101535 A1 WO 2025101535A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- alkyl
- macromolecule
- optionally substituted
- proteins
- biomolecules
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K1/00—General methods for the preparation of peptides, i.e. processes for the organic chemical preparation of peptides or proteins of any length
- C07K1/14—Extraction; Separation; Purification
- C07K1/16—Extraction; Separation; Purification by chromatography
- C07K1/20—Partition-, reverse-phase or hydrophobic interaction chromatography
-
- B—PERFORMING OPERATIONS; TRANSPORTING
- B01—PHYSICAL OR CHEMICAL PROCESSES OR APPARATUS IN GENERAL
- B01D—SEPARATION
- B01D15/00—Separating processes involving the treatment of liquids with solid sorbents; Apparatus therefor
Definitions
- the methods provided herein provide for removal of mass-spectrometry incompatible components from biological samples.
- the methods provided herein provide for removal of mass-spectrometry incompatible components from biological samples.
- methods of isolating one or more biomolecules from a biological sample comprising contacting a population of biomolecules with one or more macromolecule structure(s), binding the biomolecules (e.g., proteins) to the macromolecule structure(s), and eluting the isolated biomolecules from the macromolecule structure(s).
- the macromolecule structures provided herein provide fast pulldown times, facile re-suspension, and are filter plate compatible, which may allow for automation of the methods provided herein.
- systems and kits performing the methods provided herein.
- a method of isolating one or more biomolecules from a biological sample comprising: a. contacting a population of biomolecules comprising one or more proteins in the biological sample with a macromolecule structure in the presence of an organic solvent to bind the one or more proteins to the macromolecule structure; and b.
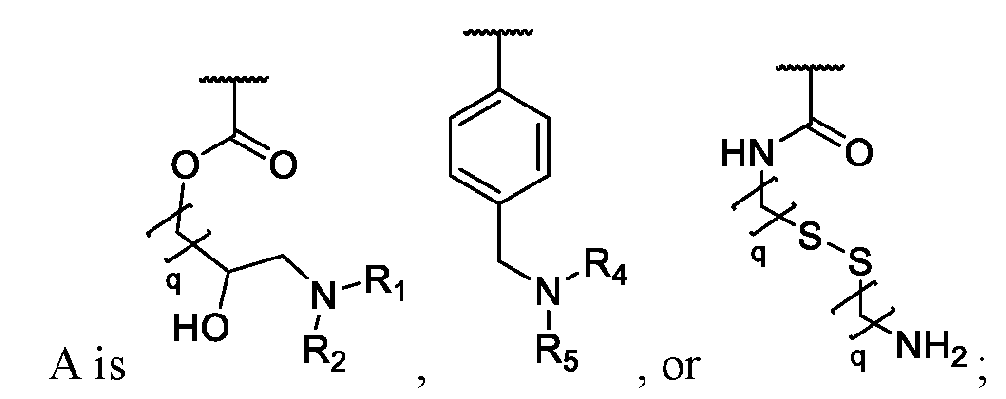
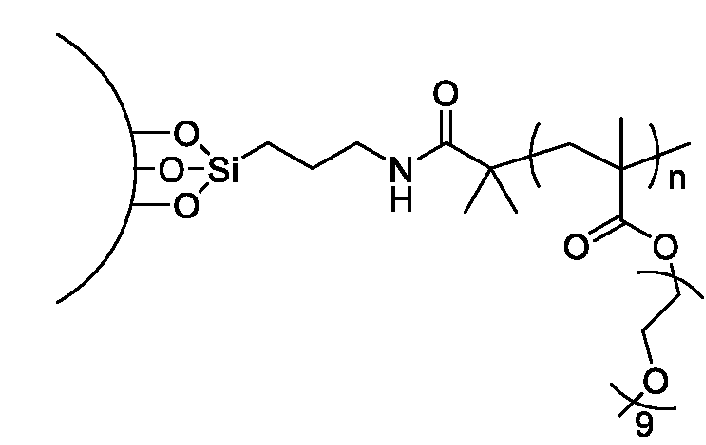
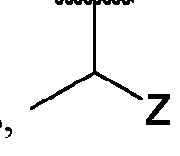
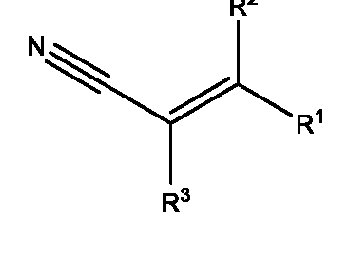
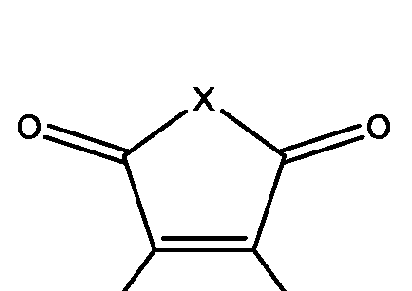
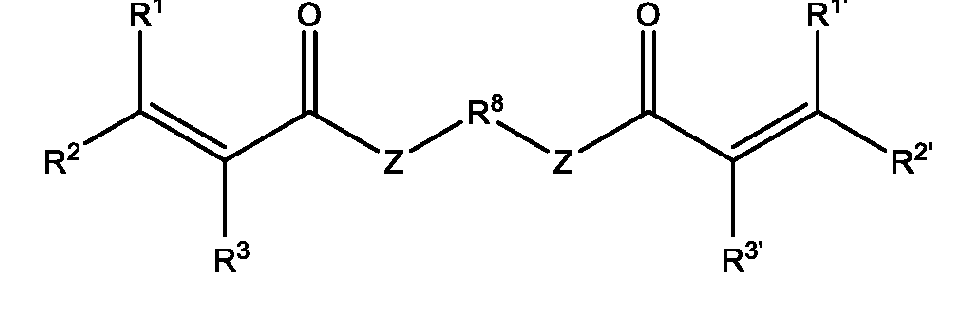
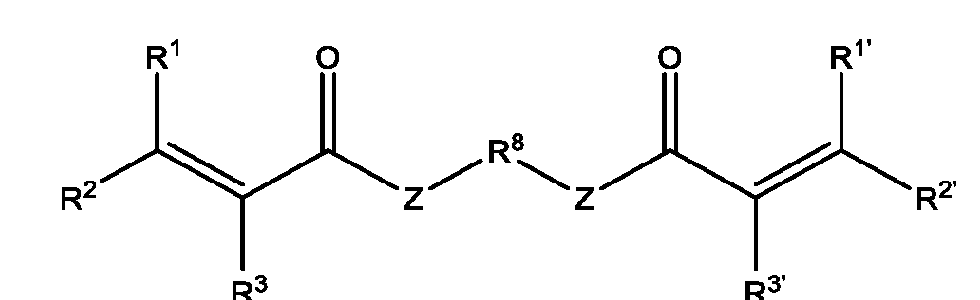
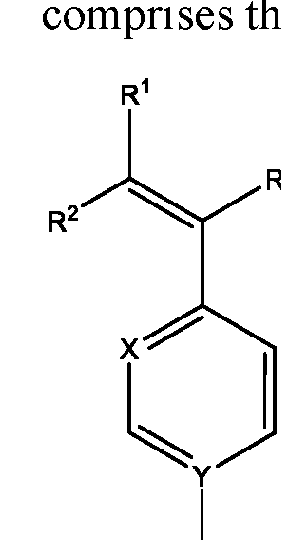
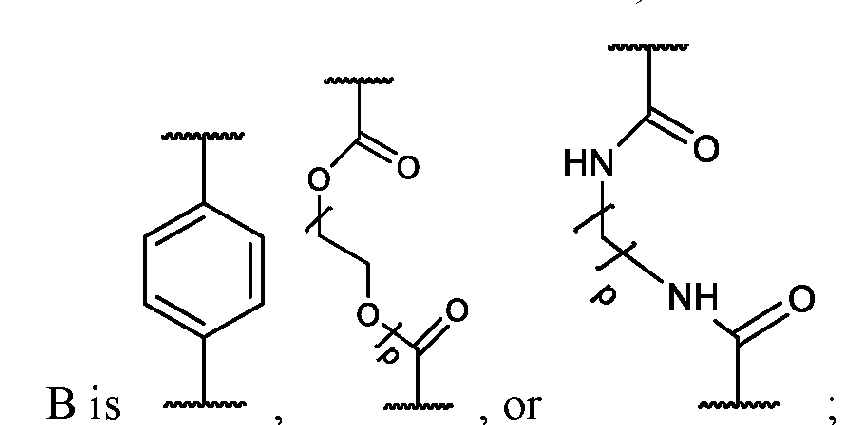
- the macromolecule structures comprise a surface and a macromolecule comprising recurring units of a first component and a second component, wherein the first component comprises a structure of Component (A) and the second component comprises a structure of Component (B’): wherein each of Yi, Y2, and Y3 is independently selected from hydrogen or Ci-Ce alkyl; each of Xi, X2, and X3 is independently selected from hydrogen or Ci-Ce alkyl;
- Ri is hydrogen, optionally substituted succinate, Ci-Ce alkyl substituted with sulfone, or phthalate;
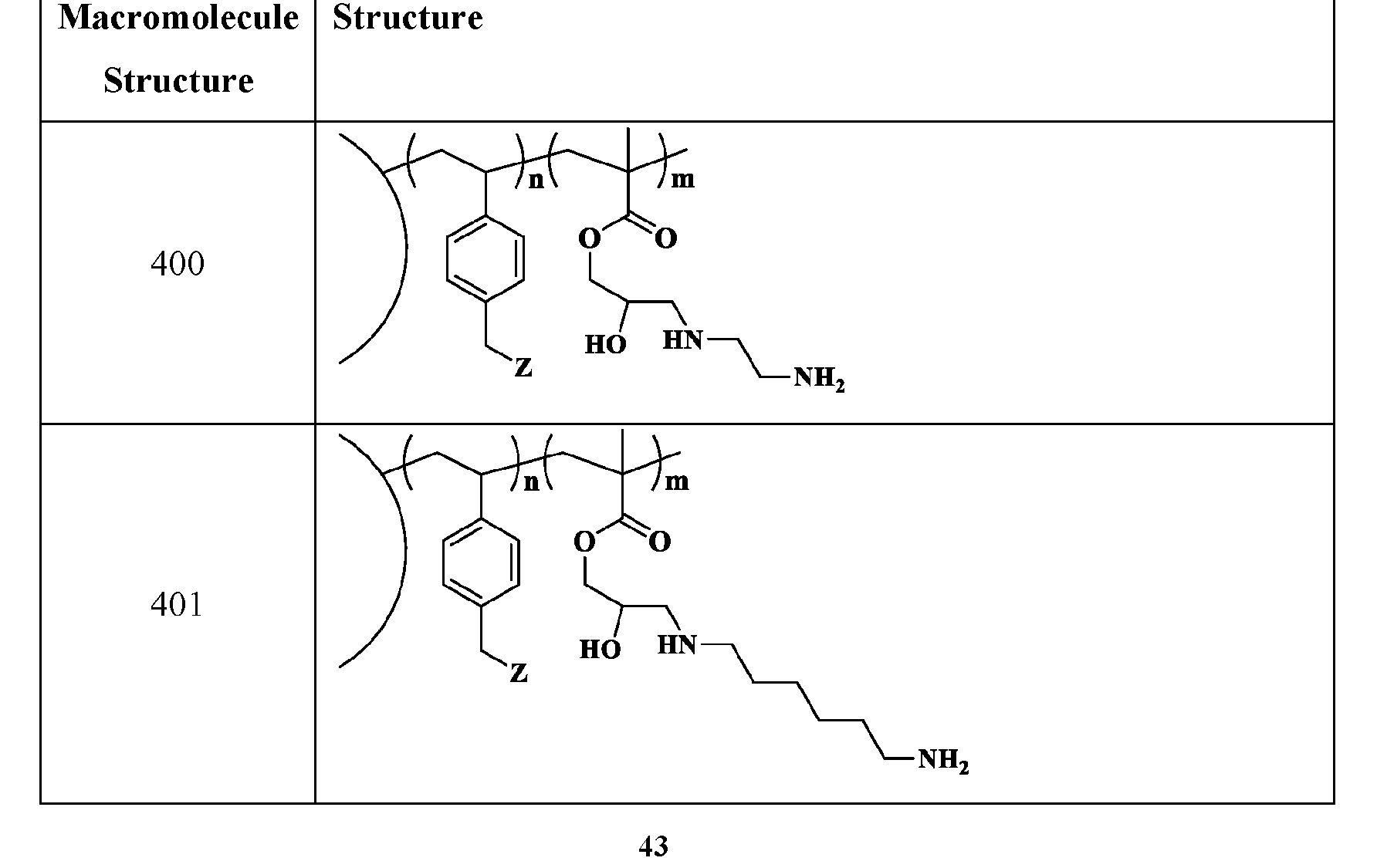
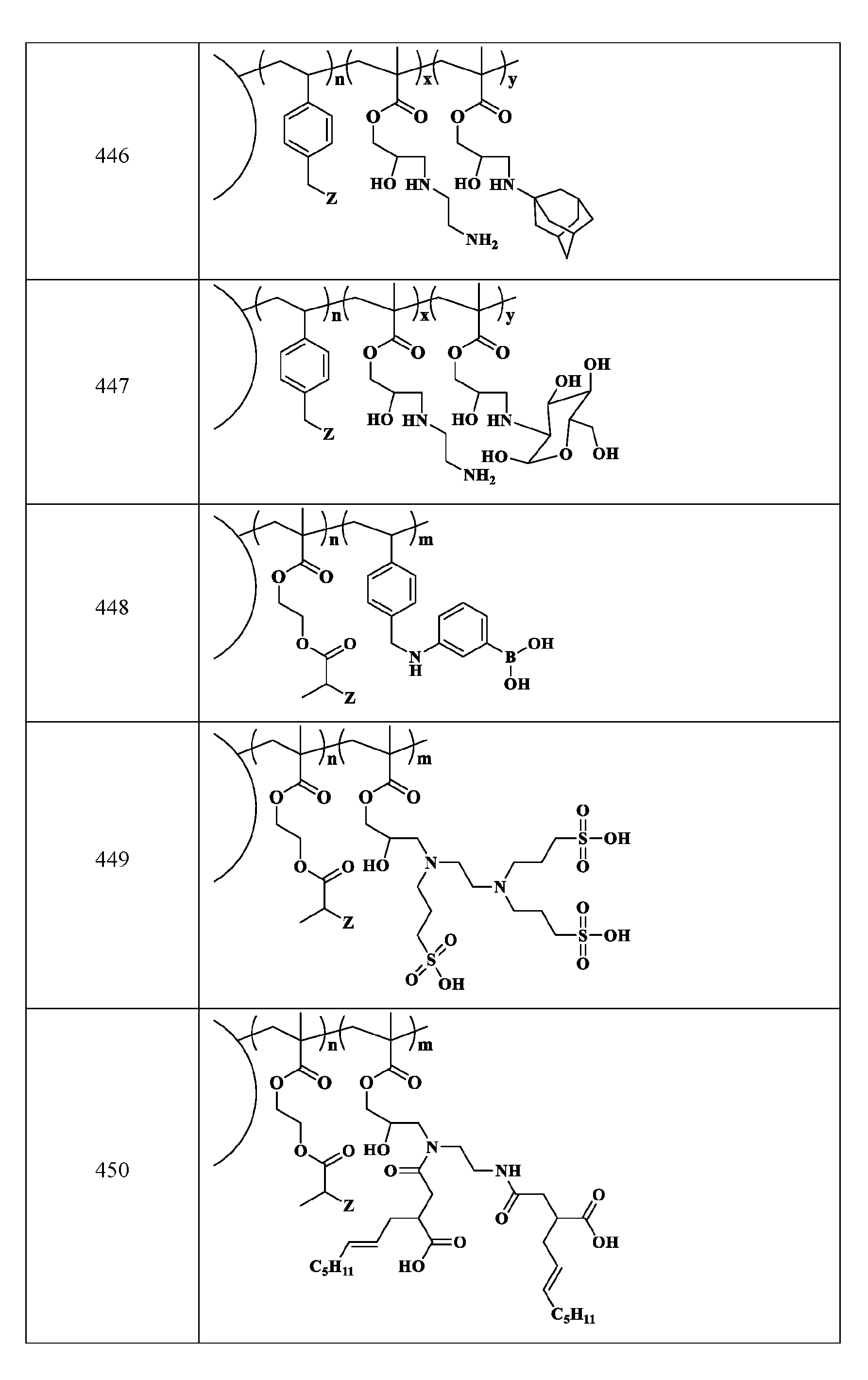
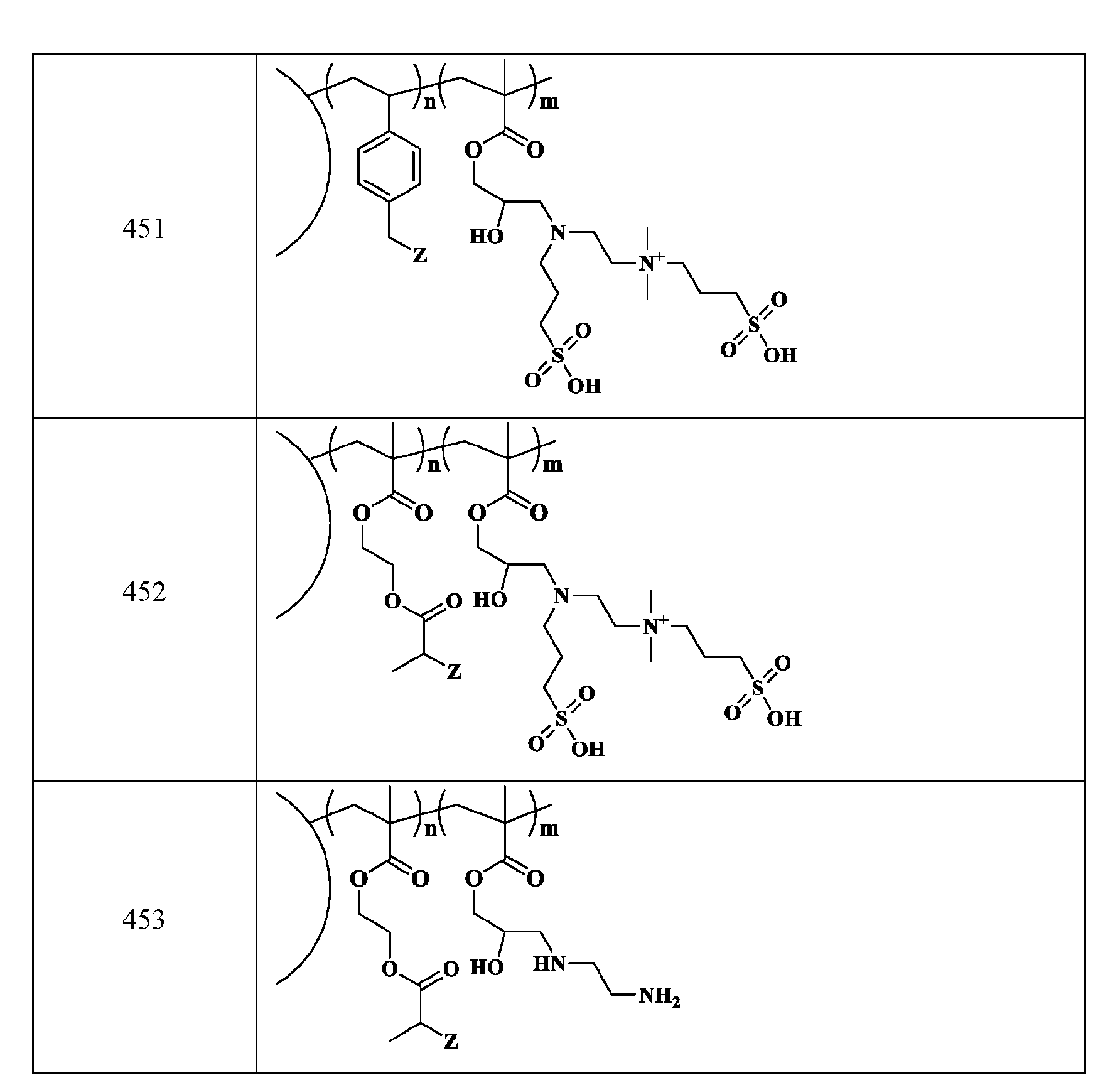
- R2 is C1-C12 alkyl substituted with amine, Ci-Ce alkyl substituted with hydroxy, optionally substituted aryl, optionally substituted heteroaryl, optionally substituted C3-C6 heterocycloalkyl, two or more fused 3-6 member rings; optionally substituted C3-C6 dicycloalkyl methane, Ci-Ce alkyl substituted with guanidine, Ci-Ce alkoxy, optionally substituted Ci-Ce alkyl substituted with disulfide, Ci-Ce alkyl substituted with thiol, optionally substituted succinate, optionally substituted Ci-Ce alkylamine, Ci-Ce alkyl acetamide, Ci-Ce aminophthalate, a boronic acid, C1-C11 optionally substituted cycloalkyl, or a monosaccharide; or
- Ri and R2 are taken together with the nitrogen to which they are attached to form an optionally substituted heterocycle;
- Ri and R2 are taken together with the nitrogen to which they are attached to form an azide group
- R4 is hydrogen or Ci-Ce alkyl substituted with thiol; Rs is succinate, optionally substituted aryl, or optionally substituted Ci-Ce alkyl substituted with disulfide; q is an integer between 1 and 6; and p is an integer between 1 and 20.
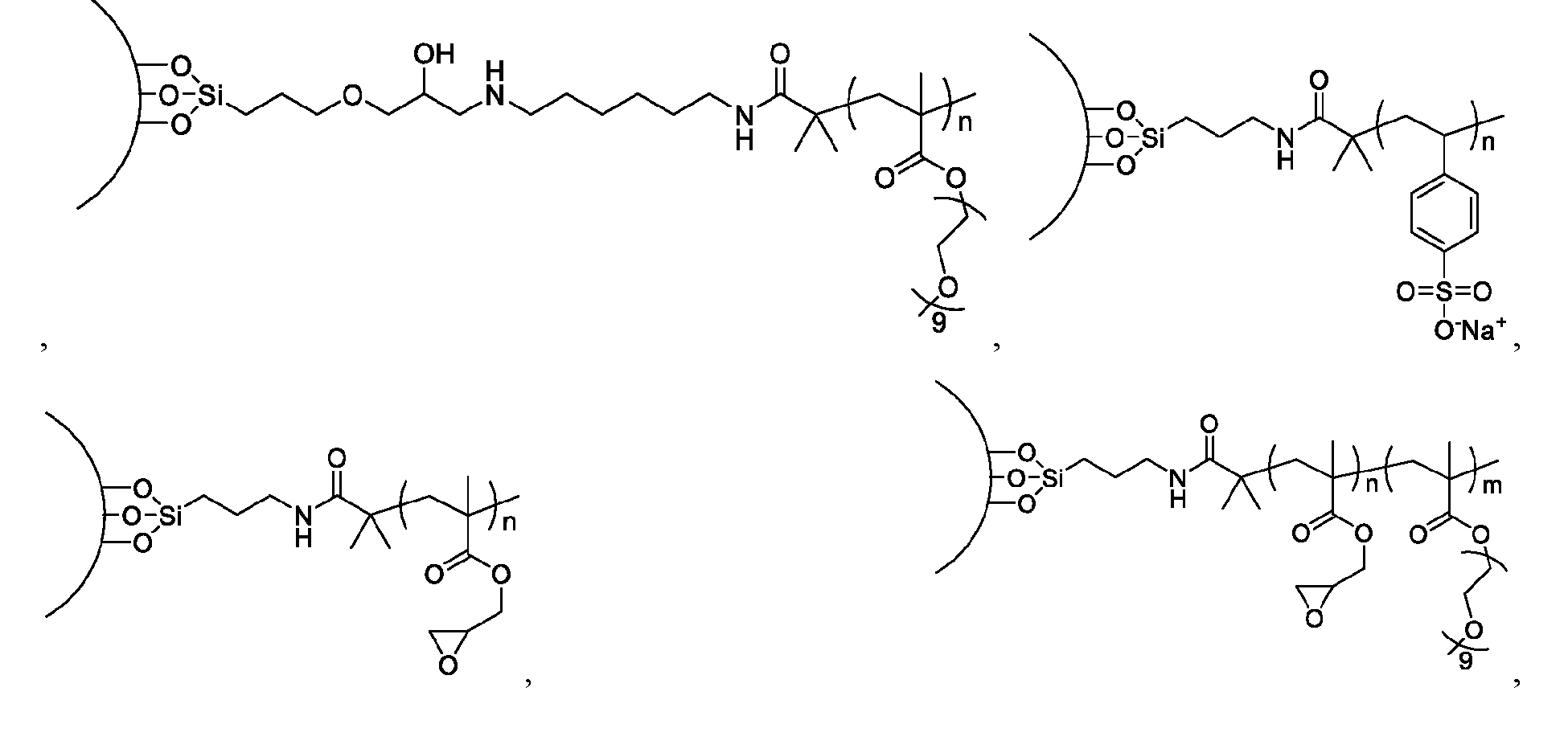
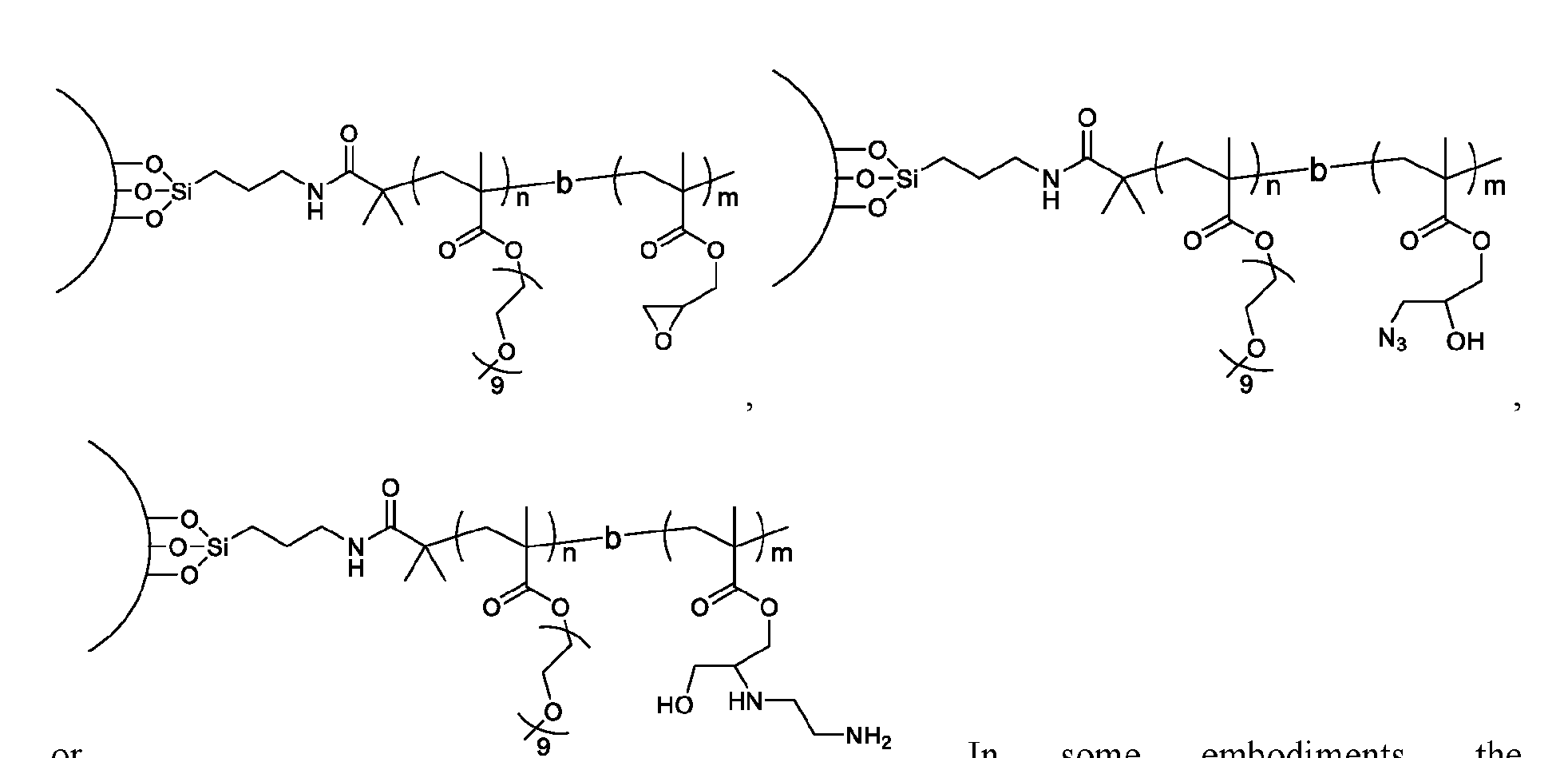
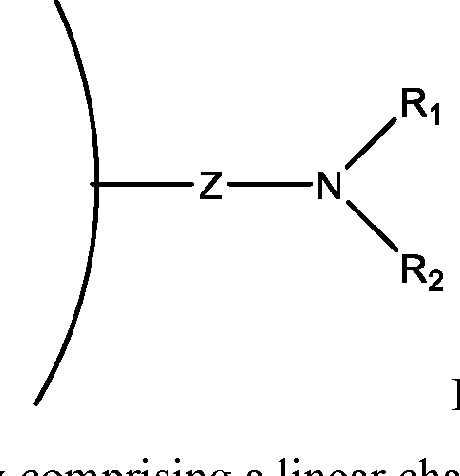
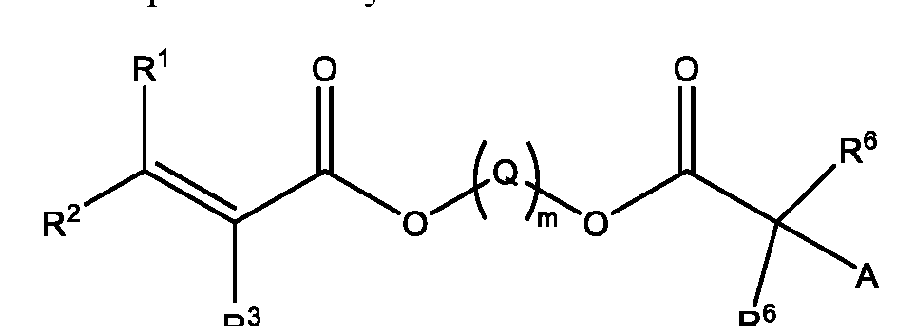
- a method of isolating one or more biomolecules from a biological sample comprising: a. contacting a population of biomolecules comprising one or more proteins in the biological sample with a macromolecule structure in the presence of an organic solvent to bind the one or more proteins to the macromolecule structure; and b. eluting the one or more proteins from the macromolecule structure, thereby providing the one or more isolated biomolecules, wherein, the macromolecule structure comprises:
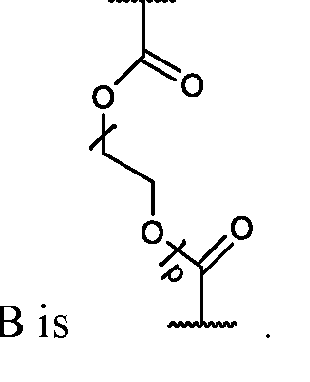
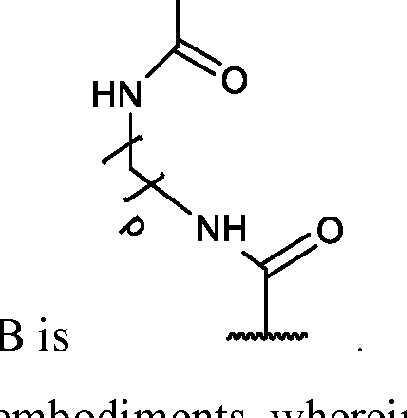
- each of X and Y is independently -C-, -O-, or -N-; each Z is independently -O- or -NH;
- Q is -CH2- or ethylene glycol
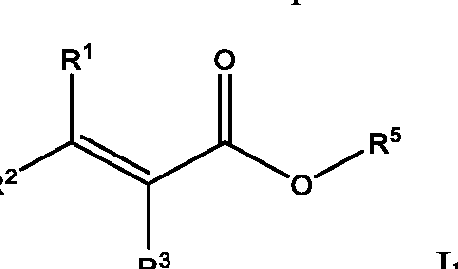
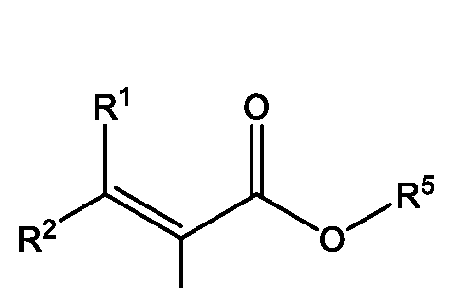
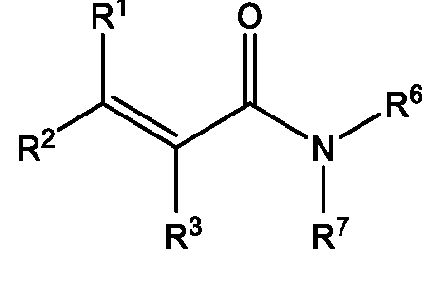
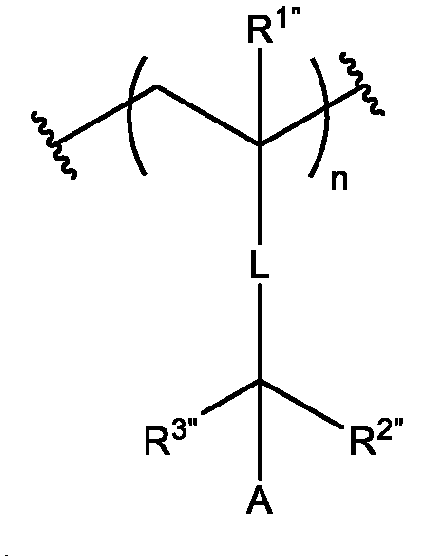
- A is a polymeric side chain comprising a recurring unit derived from a monomer represented by the structure: m is an integer selected from 1-20;
- R 1 , R 2 , R 1 , R 2 , and R 3 is independently selected from hydrogen or -Ci-Ce alkyl;
- R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl;
- R 4 is absent, hydrogen, sulfonate, carboxylate, C1-C4 alkylene, or Ci-Ce alkyl optionally substituted with halogen;
- R 5 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, - C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne;
- R 6 is hydrogen or Ci-Ce alkyl
- R 7 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, - C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne;
- R 8 is Ci-Ce alkyl, divalent metal, or symmetric or asymmetric disulfide
- R 9 is hydrogen or oxo; and n 1 is an integer selected from 1-100.
- R 5 is Ci-Cni ethylene glycol.
- nl is 9.
- the macromolecule chain comprises the recurring unit derived from a monomer represented by the structure: .
- R 3 is Ci-Ce alkyl.
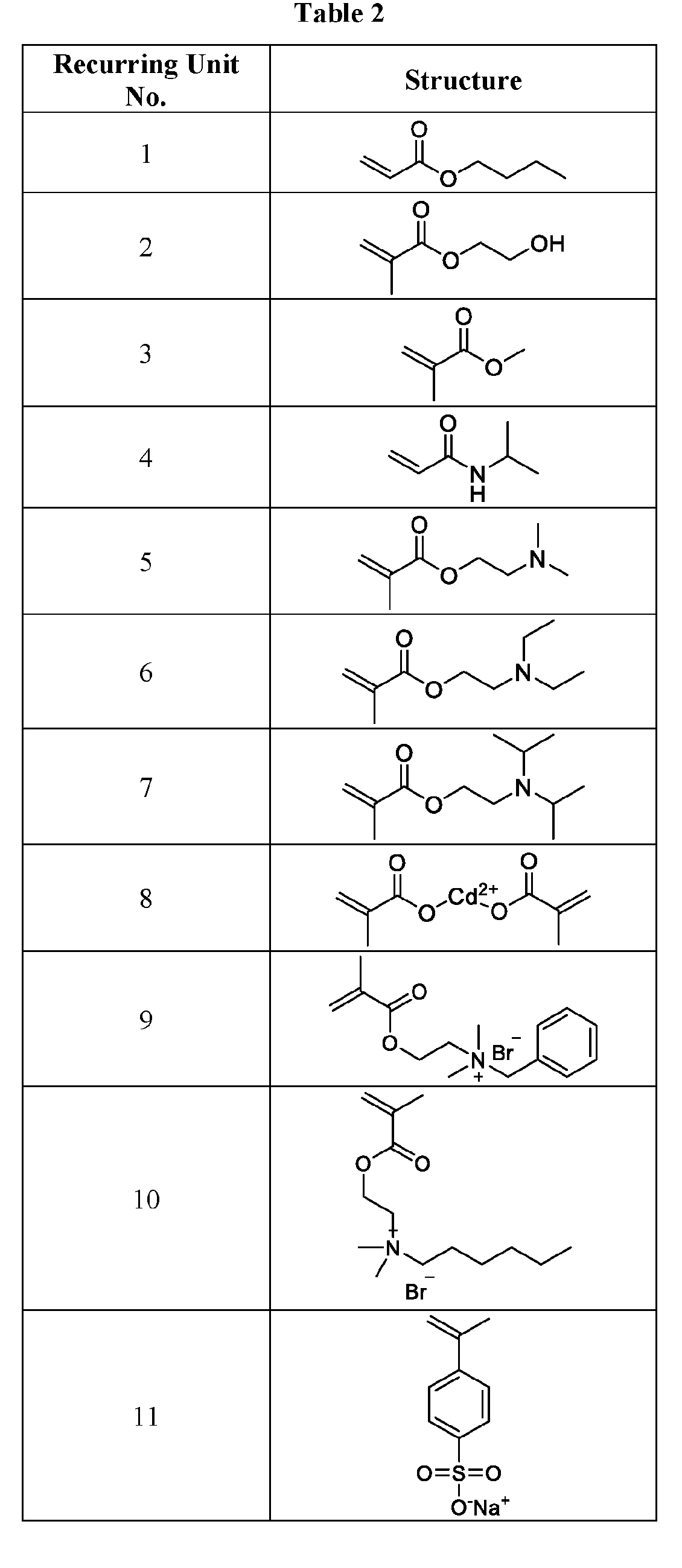
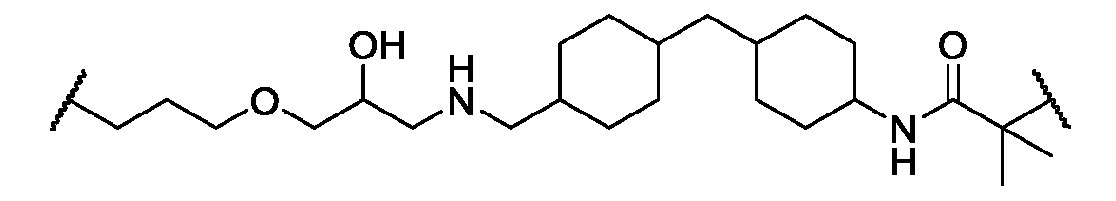
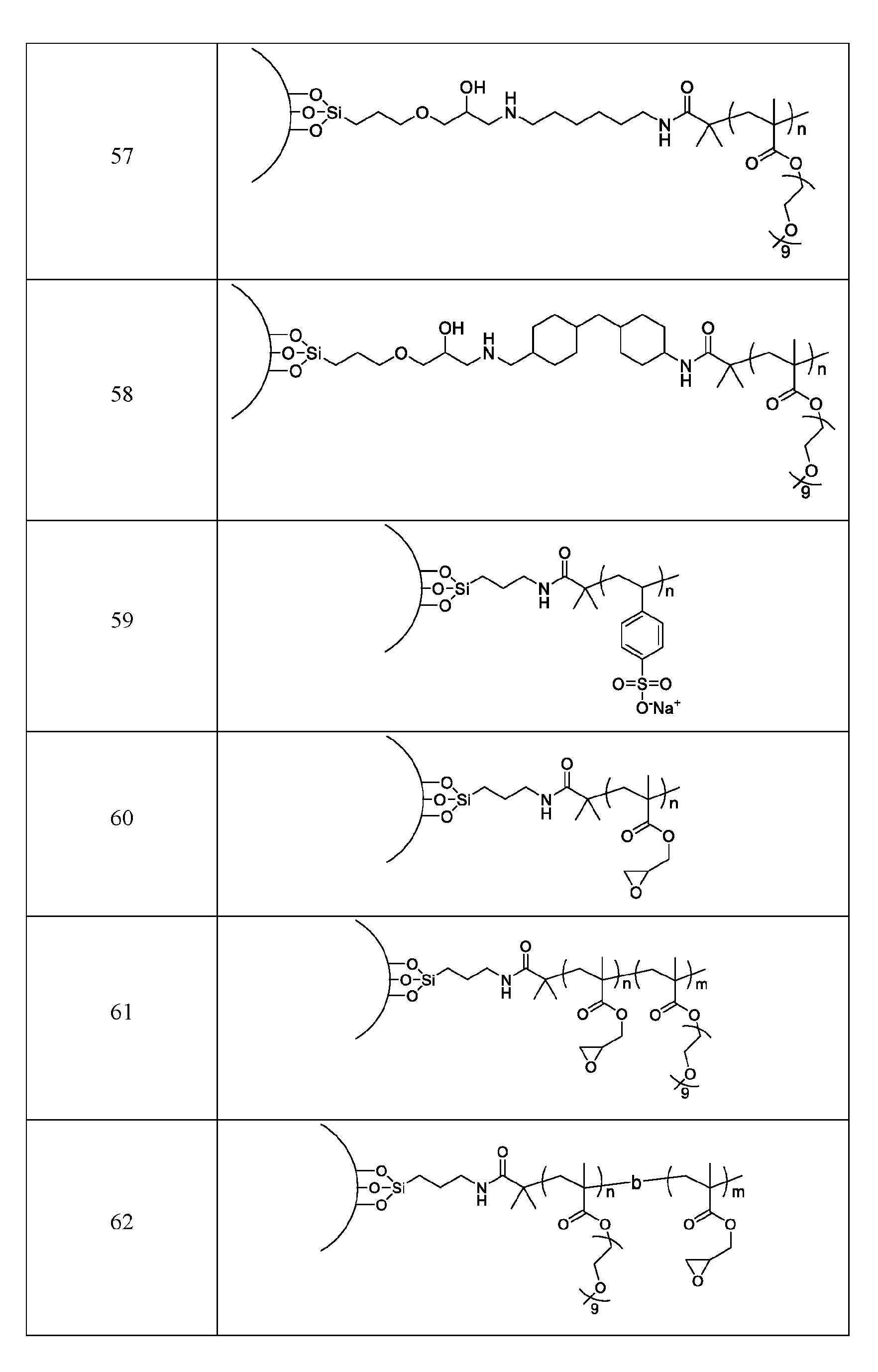
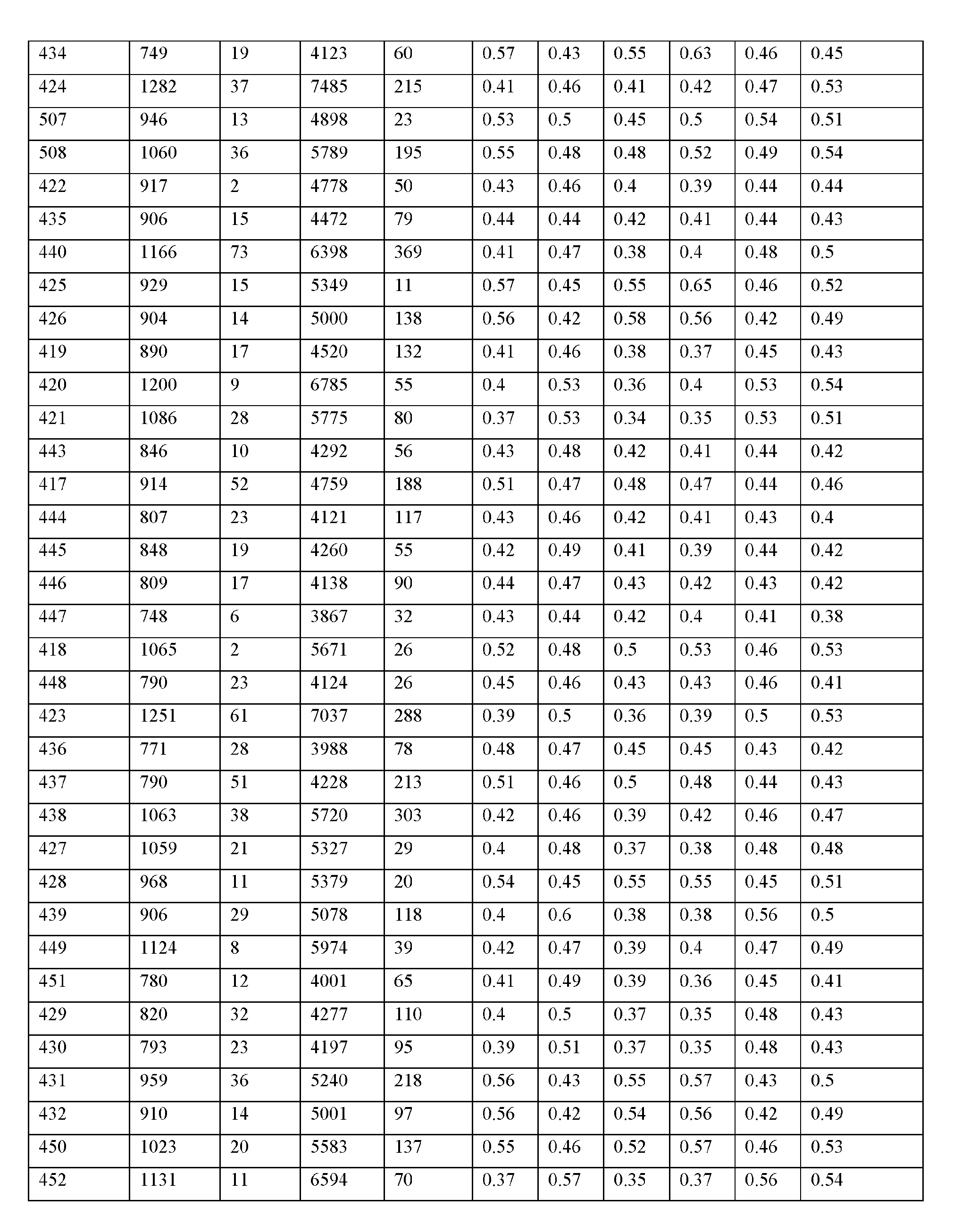
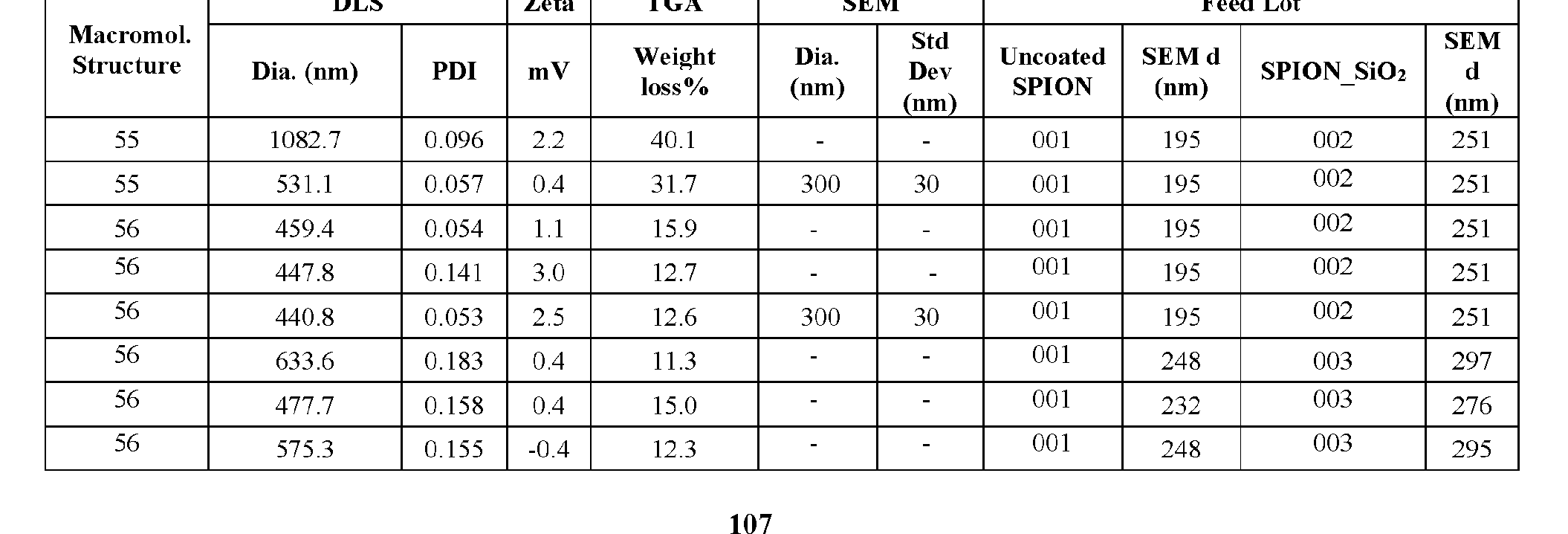
- the macromolecule structure is selected from Table 4 or Table 1 (e.g., as provided herein).
- the macromolecule structure comprises:
- the macromolecule structure comprises: the macromolecule structure comprises .
- the surface is a microparticle or a nanoparticle.
- the surface is a nanoparticle.
- the method further comprises separating the macromolecule structure from the one or more proteins after (a). In some embodiments, separating is magnetic separation, centrifugation, or a combination thereof.
- the method before (b) the method further comprises digesting one or more proteins bound to the macromolecule structure to form one or more peptides.
- (b) comprises eluting the one or more proteins (e.g., or peptides) from the macromolecule structure with an aqueous solution.
- the aqueous solution comprises an organic solvent. In some embodiments, the organic solvent comprises acetonitrile.
- the microparticle or nanoparticle comprises a zeta potential of -10 to 10 mV. In some embodiments the microparticle or nanoparticle comprises an average diameter of
- a method of isolating one or more biomolecules from a biological sample comprising: a. contacting the biomolecules comprising one or more proteins with a first macromolecule structure to provide a protein corona; b. optionally, digesting the biomolecules of the protein corona to form one or more peptides; c. separating the one or more peptides of the protein corona from the first macromolecule structure; d. contacting the one or more peptides with a second macromolecule structure and an organic solvent to bind at least a portion of the one or more peptides to the second macromolecule structure; and e. eluting the one or more peptides from the second macromolecule structure, thereby providing the one or more isolated biomolecules.
- a system for performing a method provided herein comprising: a. one or more macromolecule structure; b. a suspension solution; c. a biological sample comprising one or more proteins; and d. an automated system comprising a network of units with differentiated functions configured to perform the method.
- kits for performing a method provided herein comprising: a. one or more macromolecule structures; b. an organic solvent configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure; c. a washing agent configured to wash proteins bound to the macromolecule structure; and d. an elution agent configured to elute proteins from the macromolecule structure.
- FIG. 1 shows examples of the general design space of the epoxidated nanoparticle platform following addition of a functionalizing amine.
- FIG. 2 shows examples of the conversion schemes available after epoxidation of nanoparticles including reaction with glycidyls and azides to obtain diverse functionalities.
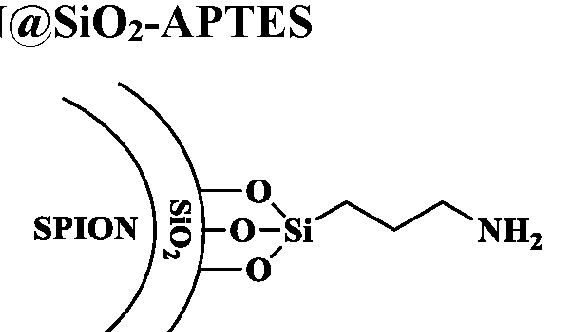
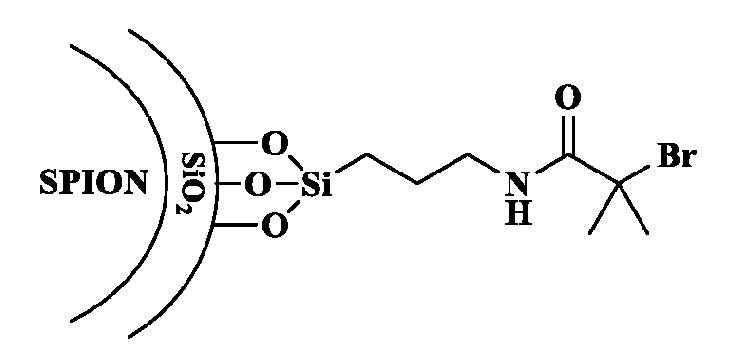
- FIG. 3 shows an example of a synthetic scheme for preparation of a macromolecule structure herein by contacting a monomer with an initiator surface.
- FIG. 4 shows an example of a synthetic scheme for preparation of a macromolecule structure herein by contacting a vinyl-functionalized surface with a cross-linking monomer and a second monomer.
- FIG. 5 shows examples of PEG structures that can be used in the macromolecule structures.
- FIG. 6a shows a schematic of an embodiment of a method of isolating biomolecules (e.g., proteins or peptides)
- biomolecules e.g., proteins or peptides
- FIG. 6b shows a schematic of an embodiment of a method of isolating biomolecules (e.g., proteins or peptides).
- biomolecules e.g., proteins or peptides.
- FIG. 7 illustrates a schematic of an embodiment of a method of isolating biomolecules for analysis.
- the peptide preparation may include a purification workflow using beads and an organic solvent.
- FIG. 8 illustrates examples of peptide yield (in micrograms) for various macromolecule structures.
- FIG. 9a illustrates examples of a quantity of unique peptides identified using liquid chromatography mass spectrometry for various macromolecule structures in comparison to commercially available surfaces (e.g., MPE or Cytiva beads).
- FIG. 9b illustrates a quantity of unique protein groups identified using liquid chromatography mass spectrometry for various macromolecule structures in comparison to commercially available surfaces (e.g., MPE or Cytiva beads).
- FIG. 10 illustrates an example of a difference in peptide identification rate (peptides/minute) between a macromolecule structure and MPE over elution time when performing liquid chromatography mass spectrometry.
- FIG. Ila illustrates an example of peptide yield (in micrograms) in a solution with beads compared to a solution with 5x the concentration of beads.
- FIG. 11b illustrates an example of the number of unique peptides identified using liquid chromatography mass spectrometry in a solution with beads compared to a solution with 5x the concentration of beads.
- FIG. 12a illustrates an example of beads in a nanoparticle (NP) solution being pulled down through the use of a magnet.
- FIG. 12b illustrates an example of the difference in rate of bead pulldown for two beads of different diameters.
- Amino refers to the -NH2 radical.
- Cyano refers to the -CN radical.
- Niro refers to the -NO2 radical.
- Haldroxyl refers to the -OH radical.
- Alkyl refers to an optionally substituted straight-chain, or optionally substituted branched-chain saturated hydrocarbon mono-radical, and preferably having from one to fifteen carbon atoms (z.e., C1-C15 alkyl).
- an alkyl comprises one to thirteen carbon atoms (z.e., C1-C13 alkyl).
- an alkyl comprises one to eight carbon atoms (z.e., Ci-Cs alkyl).
- an alkyl comprises one to five carbon atoms (z.e., C1-C5 alkyl).
- an alkyl comprises one to four carbon atoms (z.e., C1-C4 alkyl). In other embodiments, an alkyl comprises one to three carbon atoms (z.e., C1-C3 alkyl). In other embodiments, an alkyl comprises one to two carbon atoms (z.e., C1-C2 alkyl). Whenever it appears herein, a numerical range such as “C1-C3 alkyl” means that the alkyl group consists of 1 carbon atom, 2 carbon atoms, or 3 carbon atoms. In other embodiments, an alkyl comprises one carbon atom (z.e., Ci alkyl).
- an alkyl comprises five to fifteen carbon atoms (/.e., C5-C15 alkyl). In other embodiments, an alkyl comprises five to eight carbon atoms (/.e., C5- Cs alkyl). In other embodiments, an alkyl comprises two to five carbon atoms (/.e., C2-C5 alkyl). In other embodiments, an alkyl comprises three to five carbon atoms (/.e., C3-C5 alkyl).
- the alkyl group is selected from methyl, ethyl, 1 -propyl (//-propyl), 1 -methylethyl (/.w-propyl), 1 -butyl (//-butyl), 1 -methylpropyl (.scc-butyl), 2-m ethylpropyl (/.w-butyl),
- examples include, but are not limited to, methyl, ethyl, n-propyl, isopropyl, 2 -methyl- 1 -propyl, 2-methyl-2-propyl, 2- m ethyl- 1 -butyl, 3 -methyl- 1 -butyl, 2-m ethyl-3 -butyl, 2,2-dimethyl-l -propyl, 2-methyl-l -pentyl, 3 -methyl- 1 -pentyl, 4-methyl-l -pentyl, 2-methyl-2-pentyl, 3-methyl-2-pentyl, 4-methyl-2-pentyl,
- alkyl is attached to the rest of the molecule by a single bond.
- an alkyl group is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, sulfone, mercapto, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like.
- the alkyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or -OMe. In some embodiments, the alkyl is optionally substituted with halogen such as F. In some embodiments, the alkyl is unsubstituted.
- Ci-C x (or Ci-x) includes C1-C2, C1-C3... Ci-C x .
- a group designated as “C1-C4” indicates that there are one to four carbon atoms in the moiety, i.e. groups containing 1 carbon atom, 2 carbon atoms, 3 carbon atoms or 4 carbon atoms.
- C1-C4 alkyl indicates that there are one to four carbon atoms in the alkyl group, i.e., the alkyl group is selected from among methyl, ethyl, propyl, zso-propyl, //-butyl, iso- butyl, ec-butyl, and t-butyl.
- C0-C2 alkylene includes a direct bond, - CH2-, and -CH2CH2- linkages.
- Alkoxy refers to a radical bonded through an oxygen atom of the formula -O-alkyl, where alkyl is an alkyl chain as defined above. Unless stated otherwise specifically in the specification, an alkoxy group can be optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, an alkoxy is optionally substituted with oxo, halogen, -CN, - CF3, -OH, -OMe, -NH2, or -NO2.
- an alkoxy is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or -OMe. In some embodiments, the alkoxy is optionally substituted with halogen. In some embodiments, the alkoxy is unsubstituted.
- Alkenyl refers to an optionally substituted straight or branched hydrocarbon chain radical group containing at least one carbon-carbon double bond, and preferably having from two to twelve carbon atoms (i.e., C2-C12 alkenyl). In certain embodiments, an alkenyl comprises two to eight carbon atoms (z.e., C2-C8 alkenyl).
- an alkenyl comprises four to eight carbon atoms (z.e., C4-C6 alkenyl). In other embodiments, an alkenyl comprises six to eight carbon atoms (z.e., Ce-Cs alkenyl). In certain embodiments, an alkenyl comprises at least one double bond at the end of a carbon chain. In other embodiments, an alkenyl comprises at least one double bond in the middle of a carbon chain. The group can be in either the cis or trans configuration about the double bond(s), and should be understood to include both isomers.
- a numerical range such as “C2-C6 alkenyl” means that the alkenyl group can consist of 2 carbon atoms, 3 carbon atoms, 4 carbon atoms, 5 carbon atoms, or 6 carbon atoms.
- an alkenyl group is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like.
- an alkenyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, -OMe, -NH2, or -NO2.
- an alkenyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or -OMe.
- the alkenyl is optionally substituted with halogen.
- the alkenyl is attached to the rest of the molecule by a single bond, for example, ethenyl (z.e., vinyl), prop-l-enyl (z.e., allyl), but-l-enyl, pent-l-enyl, penta- 1,4-dienyl, and the like.
- an alkenyl group is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like.
- an alkenyl is optionally substituted with oxo, halogen, -CN, - CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, an alkenyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or -OMe. In some embodiments, the alkenyl is optionally substituted with halogen. In some embodiments, the alkenyl is unsubstituted.
- Alkynyl refers to an optionally substituted straight or branched hydrocarbon chain radical group containing at least one carbon-carbon triple bond, and preferably having from two to twelve carbon atoms (z.e., C2-C12 alkynyl).
- an alkynyl comprises two to eight carbon atoms (z.e., C2-C8 alkynyl).
- an alkynyl comprises two to six carbon atoms (z.e., C2-C6 alkynyl).
- an alkynyl comprises two to four carbon atoms (z.e., C2-C4 alkynyl).
- C2-C6 alkynyl means that the alkynyl group can consist of 2 carbon atoms, 3 carbon atoms, 4 carbon atoms, 5 carbon atoms, or 6 carbon atoms.
- the alkynyl is attached to the rest of the molecule by a single bond, for example, ethynyl, propynyl, butynyl, pentynyl, hexynyl, 2-propynyl, 2-butynyl, 1,3-butadiynyl, and the like.
- an alkynyl group is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like.
- an alkynyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, - OMe, -NH2, or -NO2.
- an alkynyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or -OMe.
- the alkynyl is optionally substituted with halogen.
- the alkynyl is unsubstituted.
- Alkylene or "alkylene chain” refers to an optionally substituted straight or branched divalent hydrocarbon chain linking the rest of the molecule to a radical group containing no unsaturation, and preferably having from one to twelve carbon atoms, for example, methylene, ethylene, propylene, zz-butylene, and the like.
- the alkylene chain is attached to the rest of the molecule through a single bond and to the radical group through a single bond.
- the points of attachment of the alkylene chain to the rest of the molecule and to the radical group can be through any two carbons within the chain.
- an alkylene comprises one to ten carbon atoms (i.e., Ci-Cs alkylene).
- an alkylene comprises one to eight carbon atoms (z.e., Ci-Cs alkylene). In other embodiments, an alkylene comprises one to five carbon atoms (z.e., C1-C5 alkylene). In other embodiments, an alkylene comprises one to four carbon atoms (z.e., C1-C4 alkylene). In other embodiments, an alkylene comprises one to three carbon atoms (z.e., C1-C3 alkylene). In other embodiments, an alkylene comprises one to two carbon atoms (z.e., C1-C2 alkylene). In other embodiments, an alkylene comprises one carbon atom (z.e., Ci alkylene).
- an alkylene comprises five to eight carbon atoms (z.e., Cs-Cs alkylene). In other embodiments, an alkylene comprises two to five carbon atoms (z.e., C2-C5 alkylene). In other embodiments, an alkylene comprises three to five carbon atoms (z.e., C3-C5 alkylene).
- an alkylene group can be optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like.
- an alkylene is optionally substituted with oxo, halogen, -CN, -CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, an alkylene is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or - OMe. In some embodiments, the alkylene is optionally substituted with halogen. In some embodiments, the alkylene is -CH2-, -CH2CH2-, or -CH2CH2CH2-. In some embodiments, the alkylene is -CH2-. In some embodiments, the alkylene is -CH2CH2-. In some embodiments, the alkylene is -CH2CH2CH2-. In some embodiments, the alkylene is unsubstituted.
- Aryl refers to a radical derived from a hydrocarbon ring system comprising at least one aromatic ring.
- an aryl comprises hydrogens and 5 to 30 carbon atoms.
- the aryl radical can be a monocyclic, bicyclic, tricyclic, or tetracyclic ring system, which can include fused (when fused with a cycloalkyl or heterocycloalkyl ring, the aryl is bonded through an aromatic ring atom) or bridged ring systems.
- the aryl is a 6- to 10- membered aryl.
- the aryl is a 6-membered aryl.
- Aryl radicals include, but are not limited to, aryl radicals derived from the hydrocarbon ring systems of anthrylene, naphthylene, phenanthrylene, anthracene, azulene, benzene, chrysene, fluoranthene, fluorene, indane, indene, naphthalene, phenalene, phenanthrene, pleiadene, pyrene, and triphenylene.
- the aryl is phenyl.
- an aryl can be optionally substituted, for example, with halogen, amino, alkylamino, aminoalkyl, nitrile, nitro, hydroxyl, alkyl, alkenyl, alkynyl, haloalkyl, heteroalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, -S(O)2NH-Ci-Cealkyl, and the like.
- an aryl is optionally substituted with halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, -NO2, - S(O) 2 NH 2 , -S(O) 2 NHCH3, -S(O) 2 NHCH 2 CH3, -S(O)2NHCH(CH 3 )2, -S(O)2N(CH 3 )2, or - S(O)2NHC(CH3)3.
- an aryl is optionally substituted with halogen, methyl, ethyl, -CN, -CF3, -OH, or -OMe.
- the aryl is optionally substituted with halogen.
- the aryl is substituted with alkyl, alkenyl, alkynyl, haloalkyl, or heteroalkyl, wherein each alkyl, alkenyl, alkynyl, haloalkyl, heteroalkyl is independently unsubstituted, or substituted with halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, or -NO2.
- the aryl is unsubstituted.
- alkyl refers to a radical of the formula -R c -aryl where R c is an alkylene chain as defined above, for example, methylene, ethylene, and the like.
- alkenyl refers to a radical of the formula -R d -aryl where R d is an alkenylene chain as defined above.
- Aralkynyl refers to a radical of the formula -R e -aryl, where R e is an alkynylene chain as defined above.
- Carbocycle refers to a saturated, unsaturated or aromatic rings in which each atom of the ring is carbon.
- Carbocycle can include 3- to 10-membered monocyclic rings and 6- to 12- membered bicyclic rings (such as spiro, fused, or bridged rings).
- Each ring of a bicyclic carbocycle can be selected from saturated, unsaturated, and aromatic rings.
- An aromatic ring e.g., phenyl, can be fused to a saturated or unsaturated ring, e.g., cyclohexane, cyclopentane, or cyclohexene.
- any combination of saturated, unsaturated and aromatic bicyclic rings, as valence permits, are included in the definition of carbocyclic.
- an aromatic ring e.g., phenyl
- a saturated or unsaturated ring e.g., cyclohexane, cyclopentane, or cyclohexene.
- a bicyclic carbocycle includes any combination of saturated, unsaturated and aromatic bicyclic rings, as valence permits.
- a bicyclic carbocycle includes any combination of ring sizes such as 4-5 fused ring systems, 5-5 fused ring systems, 5-6 fused ring systems, 6-6 fused ring systems, 5-7 fused ring systems, 6-5 fused ring systems, 6-7 fused ring systems, 5-8 fused ring systems, and 6-8 fused ring systems.
- Exemplary carbocycles include cyclopentyl, cyclohexyl, cyclohexenyl, adamantyl, phenyl, indanyl, and naphthyl.
- the term “unsaturated carbocycle” refers to carbocycles with at least one degree of unsaturation and excluding aromatic carbocycles.
- unsaturated carbocycles include cyclohexadiene, cyclohexene, and cyclopentene.
- saturated cyclaroalkyl refers to a saturated carbocycle.
- Exemplary carbocycles include cyclopropyl, cyclopentyl, cyclohexyl, cyclohexenyl, adamantyl, phenyl, indanyl, norborane, and naphthyl.
- Carbocycles can be optionally substituted by one or more substituents such as those substituents described herein.
- Cycloalkyl refers to a stable, partially or fully saturated, monocyclic or polycyclic carbocyclic ring, which can include fused (when fused with an aryl or a heteroaryl ring, the cycloalkyl is bonded through a non-aromatic ring atom), bridged, or spiro ring systems.
- Representative cycloalkyls include, but are not limited to, cycloalkyls having from three to fifteen carbon atoms (C3-C15 cycloalkyl), from three to ten carbon atoms (C3-C10 cycloalkyl), from three to eight carbon atoms (C3-C8 cycloalkyl), from three to six carbon atoms (C3-C6 cycloalkyl), from three to five carbon atoms (C3-C5 cycloalkyl), or three to four carbon atoms (C3-C4 cycloalkyl).
- the cycloalkyl is a 3- to 6-membered cycloalkyl.
- the cycloalkyl is a 5- to 6-membered cycloalkyl.
- Monocyclic cycloalkyls include, for example, cyclopropyl, cyclobutyl, cyclopentyl, cyclohexyl, cycloheptyl, and cyclooctyl.
- Polycyclic cycloalkyls or carbocycles include, for example, adamantyl, norbornyl, decalinyl, bicyclo[3.3.0]octane, bicyclo[4.3.0]nonane, cis-decalin, trans-decalin, bicyclo[2.1.1]hexane, bicyclo[2.2.1]heptane, bicyclo[2.2.2]octane, bicyclo[3.2.2]nonane, and bicyclo[3.3.2]decane, and 7,7-dimethyl-bicyclo[2.2.1]heptanyl.
- Partially saturated cycloalkyls include, for example, cyclopentenyl, cyclohexenyl, cycloheptenyl, and cyclooctenyl. Unless stated otherwise specifically in the specification, a cycloalkyl is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, alkyl, alkenyl, alkynyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like.
- a cycloalkyl is optionally substituted with oxo, halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, a cycloalkyl is optionally substituted with oxo, halogen, methyl, ethyl, -CN, - CF3, -OH, or -OMe. In some embodiments, the cycloalkyl is optionally substituted with halogen. In some embodiments, the cycloalkyl is unsubstituted.
- Cycloalkylalkyl refers to a radical of the formula -R c -cycloalkyl where R c is an alkylene chain as described above.
- Cycloalkylalkoxy refers to a radical bonded through an oxygen atom of the formula -O-R c -cycloalkyl where R c is an alkylene chain as described above.
- halo or halogen refers to halogen substituents such as bromo, chloro, fluoro and iodo substituents.
- haloalkyl or “haloalkane” refers to an alkyl radical, as defined above, that is substituted by one or more halogen radicals, for example, trifluoromethyl, di chloromethyl, bromomethyl, 2,2,2-trifluoroethyl, l-fluoromethyl-2-fluoroethyl, and the like.
- the alkyl part of the fluoroalkyl radical is optionally further substituted.
- halogen substituted alkanes include halomethane (e.g., chloromethane, bromomethane, fluoromethane, iodomethane), di-and trihalomethane (e.g., tri chloromethane, tribromomethane, trifluoromethane, triiodomethane), 1-haloethane, 2- haloethane, 1,2-dihaloethane, 1-halopropane, 2-halopropane, 3-halopropane, 1,2-dihalopropane, 1,3-dihalopropane, 2,3-dihalopropane, 1,2,3-trihalopropane, and any other suitable combinations of alkanes (or substituted alkanes) and halogens (e.g., Cl, Br, F, I, etc.).
- each halogen can be independently selected
- Fluoroalkyl refers to an alkyl radical, as defined above, that is substituted by one or more fluoro radicals, for example, trifluoromethyl, difluoromethyl, fluoromethyl, 2,2,2-trifluoroethyl, l-fhioromethyl-2-fluoroethyl, and the like.
- Disulfide refers to two sulfur atoms bonded to each other, where each sulfur comprises an optionally substituted alkyl chain.
- a disulfide may be R-S-S- R’.
- R and R’ may be identical.
- R and R’ are different.
- Each R and R’ may be independently selected from C1-C12 alkyl.
- R or R’ may be substituted with an amine, sulfone, or carboxylic acid.
- “Ci-Cx alkyl substituted with disulfide” as used herein may refer to a disulfide of the structure R-S-S-R’, where R and R’ may be identical or different.
- Each R and R’ may be independently selected from Ci-C y alkyl, such that the length of R and R’ is the length of the Ci-Cx alkyl.
- the term “heteroalkyl” refers to an alkyl group in which one or more skeletal atoms of the alkyl are selected from an atom other than carbon, e.g., oxygen, nitrogen (e.g., -NH-, -N(alkyl)- ), sulfur, or combinations thereof.
- a heteroalkyl is attached to the rest of the molecule at a carbon atom of the heteroalkyl.
- Heterocycloalkyl refers to a stable 3- to 24-membered partially or fully saturated ring radical comprising 2 to 23 carbon atoms and at least one ring heteroatoms. In some embodiments, a heterocycloalkyl contains from one to 8 heteroatoms selected from the group consisting of nitrogen, oxygen, phosphorous, and sulfur.
- heterocycloalkyls include, but are not limited to, heterocycloalkyls having from two to fifteen carbon atoms (C2-C15 heterocycloalkyl), from two to ten carbon atoms (C2-C10 heterocycloalkyl), from two to eight carbon atoms (C2-C8 heterocycloalkyl), from two to six carbon atoms (C2-C6 heterocycloalkyl), from two to five carbon atoms (C2-C5 heterocycloalkyl), or two to four carbon atoms (C2-C4 heterocycloalkyl).
- the heterocycloalkyl is a 3- to 6-membered heterocycloalkyl.
- the heterocycloalkyl is a 5- to 6-membered heterocycloalkyl.
- heterocycloalkyl radicals include, but are not limited to, aziridinyl, azetidinyl, dioxolanyl, thienyl[l,3]dithianyl, decahydroisoquinolyl, imidazolinyl, imidazolidinyl, isothiazolidinyl, isoxazolidinyl, morpholinyl, octahydroindolyl, octahydroisoindolyl, 2-oxopiperazinyl, 2-oxopiperidinyl, 2-oxopyrrolidinyl, oxazolidinyl, piperidinyl, piperazinyl, 4-piperidonyl, pyrrolidinyl, pyrazolidinyl, quinuclidinyl, thiazolidinyl,
- heterocycloalkyl also includes all ring forms of the carbohydrates, including but not limited to, the monosaccharides, the disaccharides, and the oligosaccharides. It is understood that when referring to the number of carbon atoms in a heterocycloalkyl, the number of carbon atoms in the heterocycloalkyl is not the same as the total number of atoms (including the heteroatoms) that make up the heterocycloalkyl (i.e. skeletal atoms of the heterocycloalkyl ring).
- a heterocycloalkyl is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, alkyl, alkenyl, alkynyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like.
- a heterocycloalkyl is optionally substituted with oxo, halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, or -NO2.
- a heterocycloalkyl is optionally substituted with oxo, halogen, methyl, ethyl, -CN, -CF3, -OH, or -OMe. In some embodiments, the heterocycloalkyl is optionally substituted with halogen. In some embodiments, the heterocycloalkyl is ub substituted.
- Heterocycle refers to a saturated, unsaturated or aromatic ring comprising one or more ring heteroatoms.
- exemplary heteroatoms include N, O, Si, P, B, and S atoms.
- Heterocycles include e.g., 3- to 10-membered monocyclic rings and 6- to 12-membered bicyclic rings (such as spiro, fused, or bridged rings).
- the heterocyclyl radical is a monocyclic, bicyclic, tricyclic or tetracyclic ring system, which optionally includes fused, bridged, or spirocyclic ring systems.
- heteroatoms in the heterocyclyl radical are optionally oxidized.
- One or more nitrogen atoms, if present, are optionally quaternized.
- the heterocyclyl radical can be partially or fully saturated.
- the heterocyclyl is attached to the rest of the molecule through any atom of the ring(s).
- heterocyclyl radicals include, but are not limited to, dioxolanyl, thienyl[l,3]dithianyl, decahydroisoquinolyl, imidazolinyl, imidazolidinyl, isothiazolidinyl, isoxazolidinyl, morpholinyl, octahydroindolyl, octahydroisoindolyl, 2-oxopiperazinyl, 2-oxopiperidinyl, 2-oxopyrrolidinyl, oxazolidinyl, piperidinyl, piperazinyl, 4-piperidonyl, pyrrolidinyl, pyrazolidinyl, quinuclidinyl, thiazolidinyl, tetrahydrofuryl, trithianyl, tetrahydropyranyl, thiomorpholinyl, thiamorpholinyl, 1-oxo-thio
- heterocyclyl is meant to include heterocyclyl radicals as defined above that are optionally substituted by one or more substituents.
- a heterocyclyl can be optionally substituted by one or more substituents selected from alkyl, alkenyl, alkynyl, halo, fluoroalkyl, oxo, thioxo, cyano, nitro, optionally substituted aryl, optionally substituted aralkyl, optionally substituted aralkenyl, optionally substituted aralkynyl, optionally substituted carbocyclyl, optionally substituted carbocyclylalkyl, optionally substituted heterocyclyl, optionally substituted heterocyclylalkyl, optionally substituted heteroaryl, optionally substituted heteroarylalkyl, -R b -OR a , -R b -OC(O)-R a , -R b -OR a , -R b -OC(O)
- Heteroaryl or “aromatic heterocycle” refers to a ring system radical comprising carbon atom(s) and one or more ring heteroatoms (e.g., selected from the group consisting of nitrogen, oxygen, phosphorous, silicon, and sulfur), and at least one aromatic ring.
- a heteroaryl is a 5- to 14-membered ring system radical comprising one to thirteen carbon atoms, one to six heteroatoms selected from the group consisting of nitrogen, oxygen, phosphorous, and sulfur.
- the heteroaryl radical can be a monocyclic, bicyclic, tricyclic, or tetracyclic ring system, which can include fused (when fused with a cycloalkyl or heterocycloalkyl ring, the heteroaryl is bonded through an aromatic ring atom) or bridged ring systems; and the nitrogen, carbon, or sulfur atoms in the heteroaryl radical can be optionally oxidized; the nitrogen atom can be optionally quaternized.
- the heteroaryl is a 5- to 10-membered heteroaryl.
- the heteroaryl is a 5- to 6-membered heteroaryl.
- Examples include, but are not limited to, azepinyl, acridinyl, benzimidazolyl, benzothiazolyl, benzindolyl, benzodi oxolyl, benzofuranyl, benzooxazolyl, benzothiazolyl, benzothiadiazolyl, benzo[b][l,4]dioxepinyl, 1,4-benzodioxanyl, benzonaphthofuranyl, benzoxazolyl, benzodi oxolyl, benzodioxinyl, benzopyranyl, benzopyranonyl, benzofuranyl, benzofuranonyl, benzothienyl (benzothiophenyl), benzotri azolyl, benzo[4,6]imidazo[l,2-a]pyridinyl, carbazolyl, cinnolinyl, dibenzofuranyl, dibenzothiophen
- a heteroaryl is optionally substituted, for example, with halogen, amino, nitrile, nitro, hydroxyl, alkyl, alkenyl, alkynyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like.
- a heteroaryl is optionally substituted with halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, or -NO2.
- a heteroaryl is optionally substituted with halogen, methyl, ethyl, -CN, -CF3, -OH, or -OMe. In some embodiments, the heteroaryl is optionally substituted with halogen. In some embodiments, the heteroaryl is unsubstituted.
- substituted refers to moieties having substituents replacing a hydrogen on one or more carbons or substitutable heteroatoms, e.g., NH, of the structure. It will be understood that “substitution” or “substituted with” includes the implicit proviso that such substitution is in accordance with permitted valence of the substituted atom and the substituent, and that the substitution results in a stable compound, i.e., a compound which does not spontaneously undergo transformation such as by rearrangement, cyclization, elimination, etc.
- substituted refers to moieties having substituents replacing two hydrogen atoms on the same carbon atom, such as substituting the two hydrogen atoms on a single carbon with an oxo, imino or thioxo group.
- substituted is contemplated to include all permissible substituents of organic compounds.
- the permissible substituents include acyclic and cyclic, branched and unbranched, carbocyclic and heterocyclic, aromatic and non-aromatic substituents of organic compounds.
- the permissible substituents can be one or more and the same or different for appropriate organic compounds.
- the heteroatoms such as nitrogen can have hydrogen substituents and/or any permissible substituents of organic compounds described herein which satisfy the valences of the heteroatoms.
- an optionally substituted group can be un- substituted (e.g., -CH2CH3), fully substituted (e.g., -CF2CF3), monosubstituted (e.g., -CH2CH2F) or substituted at a level anywhere in-between fully substituted and mono-substituted (e.g, -CH2CHF2, -CH2CF3, -CF2CH3, -CFHCHF2, etc ).
- biomolecule refers to biological components that may be involved in corona formation, including, but not limited to, for example, proteins, polypeptides, polysaccharides, a sugar, a lipid, a lipoprotein, a metabolite, an oligonucleotide, metabolome or combination thereof. It is contemplated that the biomolecule coronas of distinct particles may contain some of the same biomolecules, may contain distinct biomolecules with regard to the other sensor elements, and/or may differ in level or quantity, type or conformation of the biomolecule that binds to each sensor element. In one embodiment, the biomolecule is selected from the group of proteins, nucleic acids, lipids, and metabolomes.
- Ranges provided herein are understood to be shorthand for all of the values within the range.
- a range of 1 to 50 is understood to include any number, combination of numbers, or sub-range from the group consisting of 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, or 50, as well as all intervening decimal values between the aforementioned integers such as, for example, 1.1, 1.2, 1.3, 1.4, 1.5, 1.6, 1.7, 1.8, and 1.9.
- nested sub-ranges that extend from either end point of the range are specifically contemplated.
- a nested sub-range of an exemplary range of 1 to 50 can comprise 1 to 10, 1 to 20, 1 to 30, and 1 to 40 in one direction, or 50 to 40, 50 to 30, 50 to 20, and 50 to 10 in the other direction.
- the compounds and structures provided herein may be stereoisomeric.
- a compound or structure of the disclosure may form a stereoisomer.
- the stereoisomer may be a diastereomer (e.g., a cis/trans isomer, E/Z isomer, conformer, or rotamer).
- the stereoisomer may be an enantiomer (R,S enantiomers or +/- enantiomers).
- the compound or structure of the disclosure may be enantiopure (e.g., 100% pure).
- the compound or structure may form a racemic mixture of enantiomers (e.g., 50% pure).
- a compound or structure of the disclosure may stabilize as a stereoisomer, where the compound or structure of the disclosure comprises at least about 10%, about 15%, about 20%, about 25%, about 30%, about 35%, about 40%, about 45%, about 50%, about 55%, about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 98.5%, about 99%, about 99.1%, about 99.2%, about 99.3%, about 99.4%, about 99.5%, about 99.6%, about 99.7%, about 99.8%, about 99.9%, or more of a mixture of the compound or structure and the corresponding stereoisomer.
- the methods provided herein comprise utilizing macromolecule structures which may (e.g., selectively) bind to the one or more biomolecules.
- the binding of the one or more biomolecules to the macromolecule structures may be reversible, allowing for pull-down of the macromolecule structures (e.g., via magnetic pull-down or via centrifugation), followed by elution of the biomolecules to provide the isolated (e.g., or purified) biomolecules.
- the methods provided herein allow for clean-up of complex biological solutions, such as removal of surfactants, detergents, chaotropes, salts, and other impurities or contaminants without the need for other methods such as solid phase chromatography (e.g., reversed-phase/ion exchange) or suspension trapping, which may, among other potential disadvantages, be solvent intensive or can result in reduced yield of the desired biomolecules.
- the methods provided herein allow for the selective removal of mass spectrometry (MS) incompatible components from a biological sample.
- MS mass spectrometry
- the methods provided herein allow for isolation of biomolecules (e.g., proteins or peptides) with, for example, no solid residues or contaminants, high biomolecule (e.g., protein or peptide) recovery, no bias in biomolecule properties, and the ability to isolate large quantities of similar and unique biomolecules.
- the methods provided herein also, in some embodiments, provide macromolecule structures with fast pull-down times, easy re-suspension, and are filter plate compatible, which in some instances allow for automation of the methods herein.
- the methods disclosed herein may be faster than existing methods for removing impurities or contaminants.
- the methods disclosed herein are advantageous because they do not require a positive pressure source.
- the methods disclosed herein provide samples that are compatible with liquid chromatography mass spectrometry analysis.
- FIG. 6a shows a schematic of an embodiment of a method of isolating biomolecules (e.g., proteins or peptides) while FIG. 6b shows a schematic of an alternate embodiment of a method of isolating biomolecules (e.g., proteins or peptides).
- the methods herein comprise contacting a population of biomolecules (e.g., proteins or peptides) in a biological sample with a macromolecule in the presence of an organic solvent.
- the methods comprise eluting proteins from the macromolecule structure to provide isolated biomolecules (e.g., proteins or peptides).
- the methods provided herein comprise contacting a biological sample (e.g., comprising proteins or peptides) with a first macromolecule structure to provide a biomolecule (e.g., protein) corona.
- the methods provided herein comprise digesting the proteins or peptides of the biomolecule (e.g., protein) corona to form one or more peptides.
- the methods provided herein comprise separating peptides or proteins of the biomolecule (e.g., protein) corona from the first macromolecule structure.
- the methods herein comprise contacting peptides or proteins with a second macromolecule structure and an organic solvent to bind the peptides or proteins to the second macromolecule structure.
- the methods provided herein comprise eluting peptides or proteins from the second macromolecule structure to provide isolated biomolecules (e.g., peptides or proteins).
- the methods provided herein comprise contacting one or more biomolecules from a biological sample with a macromolecule structure.
- the contacting comprises binding of the biomolecule with a macromolecule structure.
- the methods comprise washing, as to remove any species not contacted to (e.g., or bound to) the macromolecule structures. This may result in, when reconstituted, a solution comprising a macromolecule structure comprising one or more biomolecules.
- the methods comprise eluting the biomolecule from the macromolecule structure (e.g., thereby providing the isolated biomolecule).
- isolating one or more biomolecules from a biological sample comprises purifying one or more biomolecules.
- preparing a biological sample for analysis includes removing one or more mass spectrometry incompatible components from the biological sample.
- isolating or purifying one or more biomolecules from a biological sample comprises removing at least 90% (e.g., at least 95%, at least 97.5%, at least 98%, at least 99%, at least 99.5%) of any other species from the biological sample.
- isolating or purifying one or more biomolecules from a biological sample comprises removing about 90% to about 99.9% (e.g., 90% to about 99%, 95% to about 99%, 95% to about 98%) of any other species from the biological sample.
- isolating or purifying one or more biomolecules from a biological sample comprises processing the biomolecules to a level suitable for further analysis or use, such as by mass spectrometry.
- the other species is a chemical reagent used for preparing a biological sample for mass spectrometry.
- the other species is a surfactant.
- the other species is a lysis reagent.
- the other species is a reducing agent.
- the other species is an alkylating agent.
- the other species is a buffering agent.
- the other species is a chaotrope.
- the other species is an organic solvent.
- isolating or purifying one or more biomolecules from a biological sample comprises removing at least 90% (e.g., at least 95%, at least 97.5%, at least 98%, at least 99%, at least 99.5%) of a surfactant.
- the surfactant is a synthetic compound.
- the surfactant is ionic.
- the surfactant is zwitterionic.
- Non-limiting examples of surfactants that may be removed include sodium lauryl sulfate, Triton X-100, TWEEN, NP-40, CHAPS, Octyl glucoside, deoxy cholate, RAPIGEST, and decyl maltoside.
- the methods disclosed herein may remove at least 90% of sodium lauryl sulfate.
- isolating or purifying one or more biomolecules from a biological sample comprises removing at least 90% (e.g., at least 95%, at least 97.5%, at least 98%, at least 99%, at least 99.5%) of a buffering agent.
- the buffering agent is a Good’s buffering agent.
- buffering agents include Tris, phosphate buffer, Tris, HEPES, MES, MOPS, TES, CAPS, Bicine, and Bis-Tris.
- at least 90% of CAPS may be removed using the methods disclosed herein.
- at least 90% of HEPES may be removed using the methods disclosed herein.
- isolating or purifying one or more biomolecules from a biological sample comprises removing at least 90% (e.g., at least 95%, at least 97.5%, at least 98%, at least 99%, at least 99.5%) of a chaotrope.
- chaotropes that may removed include urea, guanidine hydrochloride, sodium thiocyanate, and perchlorate salts.
- at least 90% of urea may be removed using the methods disclosed herein.
- an amount of surfactants in a composition comprising the population of biomolecules is greater than the amount of surfactants in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides).
- the amount of surfactants in a composition comprising the population of biomolecules is greater than the amount of surfactants in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 10 wt% (e.g., 20 wt%, 30 wt%, 50 wt%, 80 wt%, 90 wt%, 100 wt%).
- the amount of surfactants in a composition comprising the population of biomolecules is greater than the amount of surfactants in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 70 wt%.
- the amount of surfactants in a composition comprising the population of biomolecules is greater than the amount of surfactants in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at most 100 wt% (e.g., 99 wt%, 95 wt%, 90 wt%, 80 wt%, 70 wt%, 60 wt%).
- the amount of surfactants in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of surfactants in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by about 10 wt% to about 100 wt%, 10 wt% to about 90 wt%, 20 wt% to about 100 wt%, 30 wt% to about 100 wt%, 50 wt% to about 100 wt%, 50 wt% to about 80 wt%, or about 80 wt% to about 100 wt%.
- Non-limiting examples of surfactants that may be greater include sodium lauryl sulfate, Triton X-100, TWEEN, NP-40, CHAPS, Octyl glucoside, and decyl maltoside.
- the surfactant is sodium lauryl sulfate.
- the surfactant is CHAPS.
- the surfactant is a synthetic molecule.
- the methods provided herein comprise removing at least 90 wt% (e.g., at least 95 wt%, at least 97.5 wt%, at least 98 wt%, at least 99 wt%, at least 99.5 wt%) of an acid or base from the population of biomolecules.
- the methods provided herein comprise removing at least at least 90 wt% (e.g., at least 9 wt5%, at least 97.5 wt%, at least 98 wt%, at least 99 wt%, at least 99.5 wt%) of a surfactant from the population of biomolecules.
- the methods provided herein comprise removing at least 90 wt% (e.g., at least 95 wt%, at least 97.5 wt%, at least 98 wt%, at least 99 wt%, at least 99.5 wt%) of a lysis reagent from the population of biomolecules.
- an amount of a buffering agent in a composition comprising the population of biomolecules is greater than the amount of buffering agent in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides).
- the amount of buffering agent in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of buffering agent in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 10 wt% (e.g., 20 wt%, 30 wt%, 50 wt%, 80 wt%, 90 wt%, 100 wt%).
- the amount of buffering agent in a composition comprising the population of biomolecules is greater than the amount of buffering agent in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 70 wt%.
- the amount of buffering agent in a composition comprising the population of biomolecules is greater than the amount of buffering agent in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at most 100 wt% (e.g., 99 wt%, 95 wt%, 90 wt%, 80 wt%, 70 wt%, 60 wt%).
- the amount of buffering agent in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of buffering agent in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by about 10 wt% to about 100 wt%, 10 wt% to about 90 wt%, 20 wt% to about 100 wt%, 30 wt% to about 100 wt%, 50 wt% to about 100 wt%, 50 wt% to about 80 wt%, or about 80 wt% to about 100 wt%.
- Non-limiting examples of buffering agents that may be greater include Tris, phosphate buffer, Tris, HEPES, MES, MOPS, TES, CAPS, Bicine, and Bis-Tris.
- the buffering agent is CAPS.
- the buffering agent is HEPES.
- an amount of chaotropes in a composition comprising the population of biomolecules is greater than the amount of chaotropes in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides).
- the amount of chaotropes in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of chaotropes in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 10 wt% (e.g., 20 wt%, 30 wt%, 50 wt%, 80 wt%, 90 wt%, 100 wt%).
- the amount of chaotropes in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of chaotropes in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 70 wt%.
- the amount of chaotropes in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of chaotropes in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at most 100 wt% (e.g., 99 wt%, 95 wt%, 90 wt%, 80 wt%, 70 wt%, 60 wt%).
- the amount of chaotropes in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of chaotropes in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by about 10 wt% to about 100 wt%, 10 wt% to about 90 wt%, 20 wt% to about 100 wt%, 30 wt% to about 100 wt%, 50 wt% to about 100 wt%, 50 wt% to about 80 wt%, or about 80 wt% to about 100 wt%.
- Non-limiting examples of chaotropes that may be greater include urea, guanidine hydrochloride, sodium thiocyanate, and perchlorate salts.
- the chaotrope is urea.
- the methods provided herein comprise contacting a population of biomolecules in a biological sample with a macromolecule structure.
- contacting the population of biomolecules in a biological sample with a macromolecule structure occurs in the presence of an organic solvent, such as to bind one or more biomolecules (e.g., proteins or peptides) to the macromolecule structure.
- the organic solvent provides the necessary chemical environment for binding of the one or more biomolecules (e.g., proteins peptides) to the macromolecule structure.
- the one or more biomolecules may not bind to the macromolecule structure.
- contacting as provided herein comprises incubating.
- the methods herein comprise incubating the one or more biomolecules with a macromolecule structure.
- incubating provides for contacting (e.g., binding) of the biomolecule with the macromolecule structure.
- incubation occurs for any suitable period of time according to one of skill in the art. In some embodiments, incubation occurs for at least 10 seconds (e.g., at least 20 seconds, at least 30 seconds, at least 1 minute, at least 2 minutes, at least 4 minutes, at least 5 minutes, at least 8 minutes, at least 10 minutes).
- incubation occurs for at most 1 hour (e.g., at most 45 minutes, at most 30 minutes, at most 20 minutes, at most 10 minutes, at most 8 minutes, at most 6 minutes, at most 5 minutes, at most 4 minutes, at most 2 minutes). In some embodiments, incubation occurs for about 10 seconds to about 1 hours, about 10 seconds to about 30 minutes, about 1 minute to about 20 minutes, about 1 minute to about 10 minutes, or about 1 minute to about 5 minutes. In some embodiments, incubation occurs for about 8 minutes. In some embodiments, incubation occurs at the same time as separation, such as separation as described elsewhere herein. The temperature of incubation is not particularly limited, and may be performed, for example, at about room temperature.
- biomolecules may comprise proteins, polypeptides, polysaccharides, sugars, lipids, lipoproteins, metabolites, oligonucleotides, metabolomes, or any combination thereof.
- the biomolecules comprise proteins.
- the biomolecules comprise peptides (e.g., polypeptides).
- proteins provided herein comprise peptides. Unless otherwise stated in the present application, any reference to “protein” or “proteins” encompasses peptides obtained by fragmenting a protein, such as by trypsin digestion.
- the methods provided herein comprise preparing the macromolecule structures (provided herein).
- preparing the macromolecule structures comprises washing the macromolecule structures with water, buffer, or a combination thereof.
- preparing the macromolecule structures comprises suspending the macromolecule structures at a concentration of at least 5 mg/mL (e.g., at least 10 mg/mL, 20 mg/mL, 30 mg/mL, 40 mg/mL, 50 mg/mL, 60 mg/mL, 70 mg/mL, or 80 mg/mL).
- the methods provided herein further comprise eluting the one or more biomolecules (e.g., proteins or peptides) from the macromolecule structure.
- the methods provided herein comprise eluting the one or more proteins.
- eluting the one or more biomolecules (e.g., proteins) from the macromolecule structure provides one or more isolated biomolecules (e.g., proteins or peptides).
- the methods provided herein comprise eluting the one or more proteins from the macromolecule structure to provide one or more isolated biomolecules (e.g., proteins or peptides).
- eluting as provided herein comprises elution with an aqueous solution.
- the aqueous solution is configured to elute proteins from a macromolecule structure.
- the aqueous solution comprises an organic solvent.
- the aqueous solution comprises no more than 50 wt% (e.g., 40 wt%, 30 wt%, 20 wt%, 10 wt%, 5 wt%) of an organic solvent, such as an organic solvent as described elsewhere herein.
- the aqueous solution comprises no organic solvent or only trace amounts of organic solvent.
- the aqueous solution comprises at least 1 wt% (e.g., at least 5 wt%, at least 10 wt%, at least 20 wt%, at least 30 wt%) of an organic solvent, such as an organic solvent as described elsewhere herein.
- wt% refers to weight per weight (e.g., w/w%).
- wt% refers to weight per volume (e.g., w/v%).
- the aqueous solution comprises an organic solvent.
- the aqueous solution comprises no more than 50 v/v% (e.g., 40 v/v%, 30 v/v%, 20 v/v%, 10 v/v%, 5 v/v%) of an organic solvent, such as an organic solvent as described elsewhere herein. In some embodiments, the aqueous solution comprises no organic solvent or only trace amounts of organic solvent. In some embodiments, the aqueous solution comprises at least 1 v/v% (e.g., at least 5 v/v%, at least 10 v/v%, at least 20 v/v%, at least 30 v/v%) of an organic solvent, such as an organic solvent as described elsewhere herein. In some embodiments, the aqueous solution is pure water.
- the aqueous solution comprises a solution of acetonitrile and water. In certain embodiments, the aqueous solution comprises a solution of 10% or less acetonitrile by volume in water. In certain embodiments, the aqueous solution comprises a solution of 5% or less acetonitrile by volume in water. In certain embodiments, the aqueous solution comprises a solution of about 3% acetonitrile by volume in water.
- the elution may, in some embodiments, include using two or more eluants successfully.
- the eluants have different properties such that different fractions of biomolecules (e.g. proteins) are eluted.
- a series of eluants comprising different mixtures of water and acetonitrile may be successively applied to the macromolecule structure to remove fractions of biomolecules (e.g., proteins).
- these fractions may include more hydrophilic biomolecules.
- the skilled person guided by the teaching of the present application, can select other suitable eluants to fractionate the biomolecules.
- eluting as described herein comprise the successive application (e.g., to the macromolecule structures) of two or more eluents.
- the two or more eluents are different.
- the two or more eluents are the same.
- the combination of two or more successively applied different eluents allows for elution of the one or more peptides from the macromolecule structure(s).
- the two or more eluents comprise eluents of varying (e.g., increasing) polarity, such as by increasing the amount of water in the eluent, or such as by increasing the ratio of water to organic solvent.
- the two or more eluants may be separately analyzed. For example, the two or more separately may be separately analyzed by LC-MS/MS, which in some cases may increase the number of biomolecules identified/quantified in the sample.
- the methods provided herein further comprise separating the macromolecule structure from the one or more biomolecules (e.g., proteins or peptides) (e.g., such as after (a)).
- separating the macromolecule structure form the one or more biomolecules occurs after contacting a population of biomolecules in a biological sample with a macromolecule structure (e.g., after binding of the one or more biomolecules (e.g., proteins or peptides) with the macromolecule structure).
- the method further comprises separating the macromolecule structure from the population of biomolecules in a biological sample that are not contacted (e.g., bound) to the macromolecule structure (e.g., such as after (a)). In some instances, this separating provides the macromolecule structure comprising the one or more biomolecules (e.g., proteins or peptides) in the absence of other species from the biological sample.
- biomolecules e.g., proteins or peptides
- separating may comprise magnetic separation, centrifugation, or a combination thereof. In some embodiments, separating comprises magnetic separation. In some embodiments, separating comprises centrifugation. In some embodiments, separating comprises natural separation (e.g., such as by gravity). Separating may be completed for any duration according to one of skill in the art in order to separate the macromolecule structures from the supernatant. In some embodiments, centrifugation occurs at any speed according to one of skill in the art, such as necessary to separate ethe macromolecule structures from the supernatant.
- centrifugation occurs at a speed of at least 10 RPM (e.g., at least 20 RPM, at least 50 RPM, at least 100 RPM, at least 200 RPM, at least 400 RPM, at least 600 RPM, at least 1000 RPM). In some embodiments, centrifugation occurs at a speed of at most 6000 RPM (e.g., at most 5000 RPM, at most 3000 RPM, at most 1000 RPM, at most 600 RPM, at most 500 RPM, at most 300 RPM, at most 100 RPM).
- centrifugation occurs at a speed of about 10 RPM to about 6000 RPM, 10 RPM to about 5000 RPM, 10 RPM to about 1000 RPM, 100 RPM to about 1000 RPM, 200 RPM to about 600 RPM, or 300 RPM to about 600 RPM. In some embodiments, centrifugation occurs at about 600 RPM. In some embodiments, centrifugation occurs at a speed of 1000 RPM.
- separating herein may refer to pulling down of the macromolecule structures. In some instances, higher rates of pull-down are favorable as they may lead to enhanced rates of isolation of biomolecules (e.g., proteins or peptides).
- FIG. 12a illustrates an example of nanoparticles (NPs) in solution being pulled down through the use of a magnet.
- the rate of pulldown can be assessed through analytical techniques (e.g., UV-vis spectrometry), where a solution becomes more optically transparent as more nanoparticle beads are pulled down.
- FIG. 12b illustrates the difference in rate of bead pulldown for two beads of different diameters, macromolecule structure 1 comprising an average core diameter of about 140 nanometers and macromolecule structure 2 comprising an average core diameter of about 190 nanometers.
- a particle comprising a larger diameter has a higher pulldown rate in comparison to an otherwise similar particle comprising a smaller diameter.
- pulldown refers to the collection of nanoparticle beads, or other components, at the bottom of a vessel upon exposure to a magnet.
- a biological sample may comprise a plurality of proteins.
- a biological sample comprises plasma, serum, urine, cerebrospinal fluid, synovial fluid, tears, saliva, whole blood, milk, nipple aspirate, ductal lavage, vaginal fluid, nasal fluid, ear fluid, gastric fluid, pancreatic fluid, trabecular fluid, lung lavage, sweat, crevicular fluid, semen, prostatic fluid, sputum, fecal matter, bronchial lavage, fluid from swabbings, bronchial aspirants, fluidized solids, fine needle aspiration samples, tissue homogenates, lymphatic fluid, cell culture samples, or any combination thereof.
- the biological sample is plasma, serum, CSF, urine, tear, cell lysate, tissue lysate, cell homogenate, nipple aspirate, fecal matter, synovial fluid, saliva, or blood.
- a biological sample comprises plasma, serum, or blood.
- a biological sample comprises plasma.
- a biological sample comprises blood.
- a biological sample comprises biofluid.
- a biological sample comprises a cell-free sample.
- a biological sample comprises serum.
- the methods provided herein may further comprise digesting the one or more biomolecules (e.g., proteins or peptides) bound to the macromolecule structure.
- digesting the one or more biomolecules e.g., proteins
- Digesting may refer to breaking the biomolecule, such as a protein, into smaller components, such as peptides.
- digesting comprises breaking proteins from the biological solution into peptides.
- digesting comprise enzymolysis.
- digestion comprises digestion with trypsin, lysin, serine protease, or any combination thereof.
- digestion comprises digestion with trypsin.
- digestion comprises digestion with lysin.
- digestion comprises digestion with serine protease.
- the method may comprise denaturing of the one or more biomolecules (e.g., proteins).
- denaturing may result from contact with the organic solvent, such as an organic solvent as provided elsewhere herein.
- a denaturing agent is contacted with the biomolecules (e.g., proteins), such as chaotropic agents, surfactants, heavy metals, high salt concentrations, and the like.
- the one or more biomolecules (e.g., proteins) are denatured by applying urea.
- an organic solvent may comprise one or more of an alcohol, acetonitrile, dichloromethane, dimethylformamide (DMF), dimethyl sulfoxide (DMSO), ethylacetate, hexamethylphosphoramide (HPMA), tetrahydrofuran (THF), or any combination thereof.
- an alcohol comprises ethanol, methanol, n-propanol, isopropanol, n-butanol, sec-butanol, tert-butanol, pentanol, hexanol, cyclohexanol, or any combination thereof.
- the organic solvent comprise acetonitrile.
- the organic solvent comprises an aprotic organic solvent, such as acetonitrile and DMSO.
- the organic solvent is configured to cause protein binding to a macromolecule structure when contacted with proteins and the macromolecule structure. Without being bound to any particular theory, the binding to the macromolecule structure may be driven by hydrophobic/hydrophilic interactions, wherein the one or more biomolecules (e.g., proteins) favor the more hydrophilic surface of the macromolecule structure relative to the organic solvent.
- the organic solvent has an estimated partition coefficient using XLOGP3 of at least -1.5, at least, -1, or at least -0.5.
- the organic solvent has an estimated partition coefficient using XLOGP3 of no more than 1.5, no more than 1, no more than 0.5, or no more than 0. In some embodiments, the organic solvent has an estimated partition coefficient using XL0GP3 of about -1.5 to about 1.5. In some embodiments, the organic solvent has an estimated partition coefficient using XL0GP3 of about - 1 to about 1. In some embodiments, the organic solvent has an estimated partition coefficient using XL0GP3 of about -0.5 to about 0.5. For organic solvent mixtures, the weighted molar average of the estimated partition coefficient may be used.
- the methods are capable of isolating at least 10 (e.g., unique) biomolecules (e.g., proteins or peptides).
- the methods herein are capable of isolating at least 50 (e.g., at least 100, at least 250, at least 1,000, at least 2,500, at least 5,000, at least 7,500, at least 10,000, at least 12,500, at least 15,000, at least 17,500, at least 20,000) (e.g., unique) biomolecules (e.g., proteins or peptides).
- the methods herein are capable of isolating at most 200,000 (e.g., at most 150,000, at most 100,000, at most 50,000, at most 30,000, at most 25,000, at most 20,000, at most 15,000, at most 10,000, at most 5,000, at most 2,500, at most 1,000, at most 500, at most 100) (e.g., unique) biomolecules (e.g., proteins or peptides).
- biomolecules e.g., proteins or peptides.
- the methods herein are capable of isolating about 10 to about 50,000, about 100 to about 20,000, about 100 to about 15,000, about 100 to about 10,000, about 1,000 to about 20,000, about 1,000 to about 10,000, or about 1,000 to about 5,000 (e.g., unique) biomolecules (e.g., proteins or peptides). In some embodiments, the methods herein are capable of isolating at least 1,000 (e.g., unique) biomolecules (e.g., proteins or peptides). In some embodiments, the methods herein are capable of isolating no more than 25,000 (e.g., unique) biomolecules (e.g., proteins or peptides). The number of biomolecule that may be isolated by a macromolecule structure herein may be modified by the skilled artisan by changing the surface of the macromolecule structure or the number of unique macromolecule structures.
- isolating of biomolecules occurs in the absence of any solution comprising salts in an amount effective to precipitate the population of biomolecules (e.g., proteins).
- Contacting of the biomolecules provided herein may be achieved solely by introduction of the biomolecules and macromolecule structures in an organic solvent, such as an organic solvent described elsewhere herein.
- the methods provided herein further comprise identifying the one or more biomolecules (e.g., proteins or peptides), such as after isolation of the one or more biomolecules (e.g., proteins or peptides).
- the methods provided herein further comprise quantifying the one or more biomolecules (e.g., proteins or peptides), such as after isolation of the one or more biomolecules (e.g., proteins or peptides). The quantification may be, for example, absolute or relative amounts.
- the methods provided herein further comprise analyzing the one or more biomolecules (e.g., proteins or peptides) (e.g., such as after (b)).
- identification or analyzing comprises mass spectrometry (MS), liquid chromatography-mass spectrometry (LC-MS), protein sequencing (e.g., single-molecule protein sequencing), light scattering (e.g., dynamic light scattering (DLS), static light scattering (SLS), or circular dichroism (CD), ELISA, immunoaffinity techniques (e.g., proximity extension assays), Edman Degradation, or any combination thereof.
- identification or analyzing comprises mass spectrometry, liquid chromatography-mass spectrometry (LC-MS), protein sequencing, or any combination thereof.
- identifying or analyzing comprises mass spectrometry. In some embodiments, identifying or analyzing comprises liquid chromatography-mass spectrometry (LC-MS). In some embodiments, the LC-MS/MS includes data independent analy si s/acqui sition (DIA). In some embodiments, the LC-MS/MS includes data dependent analy si s/acqui sition (DDA). In some embodiments, identifying or analyzing comprises protein sequencing.
- LC-MS liquid chromatography-mass spectrometry
- DIA analy si s/acqui sition
- DDA data dependent analy si s/acqui sition
- identifying or analyzing comprises protein sequencing.
- FIG. 7 illustrates an example of a schematic of isolating biomolecules for analysis.
- a biological sample containing proteins is incubated with one or more different macromolecule structures to form a biomolecule (e.g., protein) corona.
- the biomolecule corona may be washed through pulling down macromolecule structures (e.g., protein corona) using a magnet, removing the supernatant, and resuspending the macromolecule structures (e.g., protein corona).
- Washed biomolecule (e.g., protein) corona may be subjected to digestion to form one or more peptides and the peptides may be purified prior to analysis via mass spectrometry.
- Purifying the macromolecule structures may comprise (a) adding the beads in a vessel, (b) incubating in a solution comprising from about 90% to about 100% organic solvent (e.g., acetonitrile), (c) applying a magnetic field to the vessel such that the macromolecule structures and peptides collect at the bottom of the vessel, (d) removing the supernatant, (e) resuspending the macromolecule structures and peptides in a solution comprising from about 90% to about 100% organic solvent (e.g., acetonitrile), (f) applying a magnetic field to the vessel such that the macromolecule structures and peptides collect at the bottom of the vessel, (g) removing the supernatant, (h) resuspending and incubating the contents of the vessel in a solution comprising water and optionally less than 10% organic solvent (e.g., about 3% acetonitrile solution in water), (i) applying a magnetic field to the vessel such that only the macromolecule structures collect at the bottom of the vessel
- the organic solvent (e.g., acetonitrile) comprises a purity of about 90% to about 100%. In some embodiments, the organic solvent (e.g., acetonitrile) comprises a purity of about 95% to about 100% (e.g., 96%, 97%, 98%, 99%).
- the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona at a temperature of about 4 degrees Celsius (°C) to about 90°C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 20°C to about 90°C.
- one or more components of the composition may be incubated at a temperature of about 20°C to about 50°C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 4°C to about 40°C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 25°C to about 40°C.
- the incubation time may be, for example, at least 1 minute, at least 5 minutes, at least 10 minute, at least 15 minutes, at least 20 minutes, at least 30 minutes, at least 45 minutes, or at least 1 hour. As a non-limiting example, the incubation may be at about 4°C to about 40°C for about 20 minutes to about 2 hours. As another example, incubation may be at about 25°C to about 40°C for about 20 minutes to about 90 minutes.
- the biological sample can be incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture.
- the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising less than about 40% by volume or weight of organ solvent.
- the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising less than about 25% by volume or weight of organ solvent.
- the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising less than about 10% by volume or weight of organ solvent. In some embodiments, the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising less than about 5% by volume or weight of organ solvent. In some embodiments, the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising less than about 1% by volume or weight of organ solvent. In some embodiments, the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising no more than trace amounts of organic solvent.
- a biomolecule e.g., protein
- the surfaces disclosed herein can be used to identify at least at least 100 (unique) (unique) protein groups, at least 200 (unique) protein groups, at least 300 (unique) protein groups, at least 400 (unique) protein groups, at least 500 (unique) protein groups, at least 600 (unique) protein groups, at least 700 (unique) protein groups, at least 800 (unique) protein groups, at least 900 (unique) protein groups, at least 1000 (unique) protein groups, at least 1100 (unique) protein groups, at least 1200 (unique) protein groups, at least 1300 (unique) protein groups, at least 1400 (unique) protein groups, at least 1500 (unique) protein groups, at least 1600 (unique) protein groups, at least 1700 (unique) protein groups, at least 1800 (unique) protein groups, at least 1900 (unique) protein groups, at least 2000 (unique) protein groups, at least 2100 (unique) protein groups, at least 2200 (unique) protein groups, at least 2300 (unique) protein groups
- (unique) protein groups from 6000 to 7000 (unique) protein groups, from 7000 to 8000 (unique) protein groups, from 8000 to 9000 (unique) protein groups, from 9000 to 10000 (unique) protein groups, from 10000 to 11000 (unique) protein groups, from 11000 to 12000 (unique) protein groups, from 12000 to 13000 (unique) protein groups, from 13000 to 14000 (unique) protein groups, from 14000 to 15000 (unique) protein groups, from 15000 to 16000 (unique) protein groups, from 16000 to 17000 (unique) protein groups, from 17000 to 18000 (unique) protein groups, from 18000 to 19000 (unique) protein groups, from 19000 to 20000 (unique) protein groups, from 20000 to 25000 (unique) protein groups, from 25000 to 30000 (unique) protein groups, from 10000 to 20000 (unique) protein groups, from 10000 to 50000 (unique) protein groups, from 20000 to 100000 (unique) protein groups, from
- the methods provided herein further comprise (e.g., such as before (a)), contacting the biological sample comprising a population of biomolecules with an additional macromolecule structure to provide a protein corona.
- biomolecule corona formation are disclosed in US2018/0172694, US2021/0285957, US2021/0285958, and W02022/020272, each of which is hereby incorporated by reference in their entirety.
- the biological sample may be incubated with silica-coated superparamagnetic iron oxide nanoparticles (SPIONs) to form a biomolecule corona.
- SPIONs superparamagnetic iron oxide nanoparticles
- the methods provided herein further comprise (e.g., such as before (a)), treating or digesting one or more proteins of the population of biomolecules to form one or more peptides or desorbed proteins.
- proteins from the protein corona are treated or digested.
- the digesting comprises enzymatic digestion.
- the digesting comprises trypsin digestion.
- the digestion may comprise incubating the protein corona with LysC and trypsin.
- the macromolecule structure and the additional macromolecule structure are the same.
- the macromolecule structure may be polymer-coated SPIONs, which are contacted with a biological sample to form a protein corona.
- the proteins from protein corona may be digested using trypsin, and then an organic solvent is added to bind the peptides to the same polymer-coated SPIONs.
- the bound peptides may be eluted for further analysis.
- the SPIONs used for protein corona formation may be replaced with new SPIONs have the same properties.
- the macromolecule structure and the additional macromolecule structure are different.
- a protein corona may be formed on a particle having a polyacrylamide coating, and then peptides obtained from the protein corona are bound to particles with a polyethylene glycol coating when contacted with a suitable organic solvent.
- the additional macromolecule structure may be removed before the biomolecules are contacted with the macromolecule structure and an organic solvent.
- both the additional macromolecule structure and the one or biomolecules are contacted with the macromolecule structure and organic solvent.
- the mass of the one or more isolated biomolecules is at least 100 nanograms (ng). In some embodiments, a mass of the one or more isolated biomolecules (e.g., proteins or peptides) is at least 500 ng (e.g., at least 1 microgram (pg), 10 pg, 50 pg, 100 pg, 200 pg). In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is at most 10 mg (e.g., 5 mg, 1 mg, 750 pg, 500 pg, 250 pg, 200 pg, 100 pg).
- the mass of the one or more isolated biomolecules is about 100 ng to about 10 mg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is about 100 ng to about 500 pg, about 100 ng to about 250 pg, about 100 ng to about 50 pg, about 10 pg to about 100 pg, or about 10 pg to about 50 pg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is about 0.1 pg to about 50 pg.
- the mass of the one or more isolated biomolecules is about 1 pg to about 10 pg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is about 1 pg to about 5 pg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is about 1 pg to about 2 pg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is sufficient for further analysis or identification, such as analyzing or identifying as described elsewhere herein.
- the methods provided herein comprise contacting the biomolecules (e.g., comprising one or more proteins or peptides) with a first macromolecule structure.
- contacting biomolecules with a first macromolecule structure provides a protein corona.
- the protein corona formation is reversible.
- contacting comprises incubating, such as incubating as provided elsewhere herein.
- the methods provided herein comprise separating the one or more peptides and proteins (e.g., biomolecules) of the protein corona from the first macromolecule structure.
- separating comprises separating as provided elsewhere herein, such as by centrifugation or applying a magnetic field.
- the methods provided herein comprise contacting the one or more peptides and proteins (e.g., biomolecules) with a second macromolecule structure and an organic solvent.
- contacting the one or more peptides and proteins (e.g., biomolecules) with a second macromolecule and an organic solvent provides for binding of at least a portion of the one or more peptide and proteins (e.g., biomolecules) to the second macromolecule structure.
- contacting comprises incubating, such as incubating as described elsewhere herein.
- the first macromolecule structure and the second macromolecule structure are the same. In other instances, the first macromolecule structure and the second macromolecule structure are different. In some embodiments, the second macromolecule structure is added to a composition comprising the first macromolecule structure. In other embodiments, the second macromolecule structure is added to a composition in absence of the first macromolecule structure.
- the first macromolecule structure or the second macromolecule structure comprises a poly(ethylene) glycol (PEG) group.
- PEG poly(ethylene) glycol
- the method further comprises eluting the one or more biomolecules (e.g., proteins or peptides) from the second macromolecule structure with an aqueous solution.
- eluting is elution as described elsewhere herein.
- macromolecule structures as described herein may be used in variable cleanup methods described herein (e.g., modification of solvent, incubation temperature, separation conditions, etc.).
- a protein may comprise one or more peptides.
- the macromolecule structures provided herein comprise different polymers that may affect the physicochemical properties of the surface such as size, surface charge, hydrophobicity, hydrophilicity, surface functionality, surface topography, surface curvature, porosity, shape, and any combination thereof.
- the changes in physicochemical properties may affect the binding properties of the macromolecule structures to other compounds, such as biomolecules (e.g., peptides or proteins), leading to increased effectiveness or decreased effectiveness in binding.
- the macromolecule structures may comprise a surface and a macromolecule comprising recurring units of a first component and optionally a second component, wherein the first component comprises a structure of Component (A), and wherein the second component is a cross-linking component that may comprises a structure of Component (B’):
- each of Yi, Y2, and Y3 is independently selected from hydrogen or Ci-Ce alkyl.
- Yi is hydrogen or Ci-Ce alkyl.
- Yi is hydrogen.
- Yi is Ci-Ce alkyl.
- Y2 is hydrogen or Ci-Ce alkyl.
- Y2 is hydrogen.
- Y2 is Ci-Ce alkyl.
- Y3 is hydrogen or Ci-Ce alkyl.
- Y3 is hydrogen.
- Y3 is Ci-Ce alkyl.
- Yi is C1-C3 alkyl and each of Y2 and Y3 are hydrogen.
- Yi is Ci alkyl (e.g., -CH3) and each of Y2 and Y3 are hydrogen.
- each of Xi, X2, and X3 is independently selected from hydrogen or Ci-Ce alkyl.
- Xi is hydrogen or Ci-Ce alkyl.
- Xi is hydrogen.
- Xi is Ci-Ce alkyl.
- X2 is hydrogen or Ci-Ce alkyl.
- X2 is hydrogen.
- X2 is Ci-Ce alkyl.
- X3 is hydrogen or Ci-Ce alkyl.
- X3 is hydrogen.
- X3 is Ci-Ce alkyl.
- Xi is C1-C3 alkyl and each of X2 and X3 are hydrogen. In some embodiments, Xi is Ci alkyl (e.g., -CH3) and each of X2 and X3 are hydrogen.
- B is In some embodiments.
- Ri is hydrogen, optionally substituted succinate, Ci-Ce alkyl substituted with sulfone, or phthalate. In some embodiments, Ri is hydrogen. In some embodiments, Ri is nitrogen. In some embodiments, Ri is optionally substituted succinate. In some embodiments, Ri is Ci-Ce alkyl. In some embodiments, Ri is Ci-Ce alkyl substituited with sulfone. In some embodiments, Ri is phthalate.
- R2 is C1-C12 alkyl substituted with amine, Ci-Ce alkyl substituted with hydroxy, optionally substituted aryl, optionally substituted heteroaryl, optionally substituted C3-C6 heterocycloalkyl, two or more fused 3-6 member rings; optionally substituted C3-C6 dicycloalkyl methane, Ci-Ce alkyl substituted with guanidine, Ci-Ce alkoxy, optionally substituted Ci-Ce alkyl substituted with disulfide, Ci-Ce alkyl substituted with thiol, optionally substituted succinate, optionally substituted Ci-Ce alkylamine, Ci-Ce alkyl acetamide, Ci-Ce aminophthalate, a boronic acid, C1-C11 optionally substituted cycloalkyl, or a monosaccharide.
- R2 is C1-C12 alkyl substituted with amine. In some embodiments, R2 is Ci-Ce alkyl substituted with hydroxy. In some embodiments, R2 is optionally substituted aryl. In some embodiments, R2 is optionally substituted heteroaryl. In some embodiments, R2 is optionally substituted C3-C6 heterocycloalkyl. In some embodiments, R2 is two or more fused 3- 6 membered rings. In some embodiments, R2 is optionally substituted Ci-Ce alkyl substituted with disulfide. In some embodiments, R2 is Ci-Ce alkyl substitute with thiol. In some embodiments, R2 is optionally substituted succinate.
- Ri and R2 are taken together with the nitrogen to which they are attached to form an optionally substituted heterocycle.
- Ri and R2 are taken together with the nitrogen to which they are attached to form an azide group.
- R 2 is Ci-Ce ether, C1-12 alkyl substituted with amine, optionally substituted Ci-Ce alkylamine, -(CH2)I-6 imidazole -, (CH2)I-6 pyridine, optionally substituted aryl, -(CH 2 )1-6N(CH3)2(CH2CH2CH 2 SOOOH), -(CH 2 )I-6N(CH2CH2CH 2 SOOOH)2, or C2-C6 alkyl substituted with hydroxy.
- R4 is hydrogen or Ci-Ce alkyl substituted with thiol. In some embodiments, R4 is hydrogen. In some embodiments, R4 is Ci-Ce alkyl substituted with thiol.
- Rs is succinate, optionally substituted aryl, or optionally substituted Ci-Ce alkyl substituted with disulfide. In some embodiments, Rs is succinate. In some embodiments, Rs is optionally substituted aryl. In some embodiments, Rs is optionally substituted Ci-Ce alkyl substituted with disulfide.
- q is an integer between 1 and 6. In some embodiments, q is an integer between 1 and 3. In some embodiments, q is 1. In some embodiments, q is 2. In some embodiments, q is 3. In some embodiments, q is 4. In some embodiments, q is 5. In some embodiments, q is 6.
- p is an integer between 1 and 20. In some embodiments, p is 1. In some embodiments, p is 2. In some embodiments, p is 3. In some embodiments, p is 4. In some embodiments, p is 5. In some embodiments, p is 6. In some embodiments, p is 7. In some embodiments, p is 8. In some embodiments, p is 9. In some embodiments, p is 10. In some embodiments, p is 11. In some embodiments, p is 12. In some embodiments, p is 13. In some embodiments, p is 14. In some embodiments, p is 15. In some embodiments, p is 16. In some embodiments, p is 17. In some embodiments, p is 18. In some embodiments, p is 19. In some embodiments, p is 20.
- the macromolecule structures as described herein and their preparations are macromolecule structures as described in PCT Patent Application No. PCT/US2023/075863, which is incorporated herein by reference in its entirety.
- the macromolecule structure may be modified surfaces, such as particles, as described in U.S. Publication No. 2021/0285957, which is incorporated herein by reference in its entirety.
- the macromolecule structures may be any of the particle types provided in Table 1 of U.S. Publication No. 2021/0285957.
- n, m, x, and y are integers representing the number of recurring units.
- m is an integer greater than 2.
- m is an integer between 2 and 2,000.
- m is an integer between 50 and 1,000.
- m is an integer between 100 and 200.
- n is an integer greater than 2.
- n is an integer between 2 and 200.
- n is an integer between 2 and 100.
- n is an integer between 2 and 20.
- the recurring units denoted by n and m are randomly copolymerized.
- x is an integer greater than 2.
- x is an integer between 2 and 50,000.
- y is an integer greater than 2.
- y is an integer between 2 and 50,000.
- n may refer interchangeably to
- the macromolecule structures provided herein can be prepared according to methods as described in Examples 6-8.
- FIG. 1 shows examples of the general design space of the epoxidated macromolecule structure platform following addition of a functionalizing amine
- FIG. 2 shows examples of the conversion schemes available after epoxidation of macromolecule structures including reaction with glycidyls and azides to obtain diverse functionalities.
- the macromolecule structures provided herein may comprise a surface, a tethering moiety coupled to the surface, and a macromolecule chain.
- the macromolecule structures provided herein comprise a surface and a macromolecule chain.
- the macromolecule structure provided herein comprises a surface.
- the macromolecule structure provided herein comprises a tethering moiety (e.g., coupled to the surface).
- the macromolecule structure provided herein comprises a macromolecule chain.
- a first end of the macromolecule chain is covalently attached to the tethering moiety.
- a second end of the macromolecule chain is not coupled to the surface.
- the macromolecule chain comprises one or more (e.g., distinct) recurring units derived from a monomer. [00141] In some embodiments, the macromolecule chain comprises two or more distinct recurring units derived from different monomer. In some embodiments, the macromolecule chain comprises at least two (e.g., at least three, at least four, at least five) distinct recurring units derived from different monomers. In some embodiments, the macromolecule chain comprises at most ten (e.g., at most nine, at most eight, at most six, at most four, at most three) distinct recurring units derived from different monomers. In some embodiments, the macromolecule chain comprises two distinct recurring units derived from different monomers.
- the macromolecule chain comprises three distinct recurring units derived from different monomers. In some embodiments, the macromolecule chain comprises four distinct recurring units derived from different monomers. In some embodiments, the macromolecule chain comprises five distinct recurring units derived from different monomers.
- the macromolecule chain is comprised of a single monomeric recurring unit.
- the macromolecule structures provided herein comprise crosslinked polymers (e.g., crosslinked macromolecule chains).
- the relative amount of crosslinking monomer relative to total monomer in the polymers by weight or number can be at least 0.1%, at least 0.5%, at least 1%, or at least 2%.
- the macromolecule structures provided herein comprise substantially uncross-linked polymers (e.g., uncrosslinked macromolecule chains).
- the relative amount of crosslinking monomer relative to total monomer in the polymers by weight or number can be less than 0.1%, less than 0.05%, less than 0.01%, or about 0%.
- the macromolecule structures provided herein comprise polymer brushes (e.g., PEG brushes).
- the polymer brushes may comprise sidechains with at least 5, at least 10, at least 15, or at least 20 recurring units derived from one or more monomers.
- a macromolecule structure comprising a moiety of Formula (IV): Formula (IV) wherein Z is a linking moiety comprising a linear chain with 2 to 20 atoms selected from the group consisting of carbon, oxygen, and nitrogen, and optional substituents on the carbon and any nitrogen atoms in the linear chain; Ri is hydrogen or succinate; and
- R2 is optionally substituted Ci-Ce alkyl substituted with disulfide or Ci-Ce alkyl substituted with thiol.
- Z is a linear chain with 2 to 20 atoms. In some embodiments, Z is a linear chain with 2 to 12 atoms. In some embodiments, Z is a linear chain with 2 to 6 atoms. In some embodiments, Z is a linear chain with 2 atoms. In some embodiments, Z is a linear chain with 3 atoms. In some embodiments, Z is a linear chain with 4 atoms. In some embodiments, Z is a linear chain with 5 atoms. In some embodiments, Z is a linear chain with 6 atoms. In some embodiments Z comprises carbon only. In some embodiments, Z is a C2-C6 alkyl chain. In some embodiments, Z is C3 alkyl. In some embodiments Z comprises oxygen, nitrogen, carbon, or a combination thereof. In some embodiments, Z comprises substituents on the linear chain.
- Z comprises a C2-C8 heteroalkyl, optionally substituted (e.g., with hydroxy). In some embodiments, Z comprises a C2-C8 alkoxy, optionally substituted (e.g., with hydroxy).
- Ri is hydrogen, optionally substituted succinate, optionally substituted glutarate, optionally substituted adipate, optionally substituted pimelate, optionally substituted suberate, optionally substituted azelate, or optionally substituted sebacate.
- Ri is optionally substituted glutarate.
- Ri is optionally substituted adipate.
- Ri is optionally substitute pimelate.
- Ri is optionally substituted suberate.
- Ri is optionally substituted azelate.
- Ri is optionally substitute sebecate.
- Ri is hydrogen.
- Ri is optionally substituted Ci-Ce alkyl.
- Ri is substituted with one or more oxo and -COOH.
- Ri is succinate.
- Ri is optionally substituted succinate.
- R2 is optionally substituted -Ci-Ce alkyl substituted with disulfide.
- R2 is Ci-Ce thiol.
- R2 is substituted -Ci-Ce alkyl substituted with disulfide.
- R2 is substituted di-Ci-Ce alkyl disulfide.
- R2 is -CH2CH2-S-S-CH2CH2NH2.
- R2 is C2 alkyl substituted with thiol (e.g., -(CH2)2SH).
- Z is attached to a surface.
- Z is covalently attached is a surface.
- a structure of Formula (IV) comprises
- a structure of Formula (IV) comprises (Compound 507). In some embodiments, a structure of
- Formula (IV) comprises (Compound 509).
- FIG. 5 illustrates examples of PEG structures that a macromolecule structure described herein may comprise.
- the PEG structure comprises from about 2 to about 20 repeating ethylene glycol units.
- a macromolecule structure comprising a PEG component with increasing number of ethylene glycol units may comprise a more positive Zeta (mV) value.
- a PEG polymer may be a thermo-responsive polymer.
- a thermo-responsive polymer is sensitive to thermal energy (e.g., heat or increase in temperature).
- exposure of a thermo-responsive polymer to heat may cause one or more physical properties of the thermo-responsive polymer to change.
- solubility of a thermos-responsive polymer may change upon exposure to heat.
- the thermo-responsive polymer has a lower critical solution temperature.
- the solubility may decrease as the temperature is raised above the lower critical solution temperature.
- the temperature may be modified to improve protein or peptide capture on a macromolecule surface.
- the temperature may be modified to improve isolation of a macromolecule surface and any bound biomolecules (e.g., peptides or proteins).
- the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
- R 4 is hydrogen, sulfonate, carboxylate, C1-C4 alkylene, amine (e.g., quaternary ammonium cation), or Ci-Ce alkyl optionally substituted with halogen.
- R 4 is hydrogen.
- R 4 is absent.
- R 4 is sulfonate.
- R 4 is amine.
- R 4 is a quaternary ammonium cation.
- R 4 is carboxylate.
- R 4 is C1-C4 alkylene.
- R 4 is Ci-Ce alkyl optionally substituted with halogen (e.g., haloalkyl).
- X is -C- or -N-. In some embodiments, X is -C-. In some embodiments, X is -N-. In some embodiments, Y is -C- or -N-. In some embodiments, Y is -C-. In some embodiments, Y is -N-.
- the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
- the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
- a recurring unit is derived from an acrylate monomer. In some embodiments, a recurring unit is derived from a methacrylate monomer.
- R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl.
- R 2 is hydrogen or Ci-Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl.
- R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is hydrogen. In some embodiments, R 3 is 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is 3-, 5-, or 6-membered heterocycle optionally substituted with one or more methyl. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl.
- R 5 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6- membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further substituted with amine, hydroxyl, aryl, or sulfonate, Ci- Cs alkoxy optionally substituted with one or more oxo or halogen, -C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted (e.g., with 2 or more fused 6 membered rings further optionally substituted), optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-
- R 5 is hydrogen. In some embodiments, R 5 is Ci-Ce alkyl. In some embodiments, R 5 is C1-C4 alkylyne. In some embodiments, R 5 is Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci- Cni ethylene glycol.
- R 5 is Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol.
- R 5 is Ci-Cs alkylamine.
- R 5 is Ci-Cs alkylamine substituted with amine.
- R 5 is Ci-Cs alkylamine substituted with hydroxyl and amine.
- R 5 is Ci-Cs alkylamine substituted with sulfonate.
- R 5 is Ci-Cs alkylamine substituted with aryl. In some embodiments, R 5 is Ci-Cs alkoxy. In some embodiments, R 5 is Ci-Cs alkoxy substituted with one or more oxo or halogen. In some embodiments, R 5 is Ci-Cs alkoxy substituted with one or more oxo. In some embodiments, R 5 is Ci-Cs alkoxy substituted with one or more halogen. In some embodiments, the alkylamine comprises a quaternary ammonium cation.
- R 5 is C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted (e.g., with 2 or more fused 6 membered rings further optionally substituted), optionally substituted benzyl, trimethoxysilane, or phosphorocholine.
- R 5 is C1-C3 alkyl substituted with pyrene.
- R 5 is C1-C3 alkyl substituted with 2 or more 5-6 membered rings further optionally substituted, such as with 2 or more fused 6 membered rings further optionally substituted.
- R 5 is C1-C3 alkyl substituted with optionally substituted benzyl. In some embodiments, R 5 is C1-C3 alkyl substituted with trimethoxysilane. In some embodiments, R 5 is C1-C3 alkyl substituted with phosphorocholine.
- R 5 is hydrogen, Ci-Ce alkyl, or C1-C3 alkyl substituted with pyrene or 2 or more fused 5-6 membered rings optionally further substituted. In some embodiments, R 5 is hydrogen, Ci-Ce alkyl, or C1-C3 alkyl substituted with pyrene or 2 or more fused 5-6 membered rings optionally further substituted and the macromolecule chain comprises two or more distinct recurring units.
- the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
- a recurring unit is derived from an acrylamide monomer.
- R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl.
- R 2 is hydrogen or Ci-Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl.
- R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is hydrogen. In some embodiments, R 3 is 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is 3-, 5-, or 6-membered heterocycle optionally substituted with one or more methyl. In some embodiments, R 3 is 3,4-dimethyl- l//- pyrrole-2, 5-dione. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl. [00164] In some embodiments, R 6 is hydrogen or Ci-Ce alkyl. In some embodiments, R 6 is hydrogen. In some embodiments, R 6 is Ci-Ce alkyl.
- R 7 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6- membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, -C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted (e.g., with 2 or more fused 6 membered rings further optionally substituted), optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne.
- R 7 is hydrogen. In some embodiments, R 7 is Ci-Ce alkyl. In some embodiments, R 7 is C1-C4 alkylyne. In some embodiments, R 7 is Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci- Cni ethylene glycol.
- R 7 is Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol.
- R 7 is Ci-Cs alkylamine.
- R 7 is Ci-Cs alkylamine substituted with amine.
- R 7 is Ci-Cs alkylamine substituted with sulfonate.
- R 7 is Ci-Cs alkoxy.
- R 7 is Ci-Cs alkoxy substituted with one or more oxo or halogen. In some embodiments, R 7 is Ci-Cs alkoxy substituted with one or more oxo. In some embodiments, R 7 is Ci-Cs alkoxy substituted with one or more halogen. In some embodiments, R 7 is Ci-Ce alkyl optionally substituted with hydroxyl, substituted benzene, or hydrogen.
- R 7 is C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted (e.g., with 2 or more fused 6 membered rings further optionally substituted), optionally substituted benzyl, trimethoxysilane, or phosphorocholine.
- R 7 is C1-C3 alkyl substituted with pyrene.
- R 7 is C1-C3 alkyl substituted with 2 or more 5-6 membered rings further optionally substituted, such as with 2 or more fused 6 membered rings further optionally substituted.
- R 7 is C1-C3 alkyl substituted with optionally substituted benzyl. In some embodiments, R 7 is C1-C3 alkyl substituted with trimethoxysilane. In some embodiments, R 7 is C1-C3 alkyl substituted with phosphorocholine.
- the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
- a recurring unit derived from a monomer can be a diene, such as a cis diene.
- the diene may act as a cross-linking monomer.
- R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl. In some embodiments, R 2 is hydrogen or Ci- Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl. In some embodiments, R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is hydrogen. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl. In some embodiments, R 1 is hydrogen or Ci-Ce alkyl.
- R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl. In some embodiments, R 2 is hydrogen or Ci-Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl. In some embodiments, R 3 is hydrogen, Ci- Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is hydrogen. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl.
- the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
- a recurring unit is derived from an acrylonitrile monomer.
- R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl. In some embodiments, R 2 is hydrogen or Ci- Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl. In some embodiments, R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is hydrogen. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl. In some embodiments, R 1 is hydrogen, R 2 is hydrogen, and R 3 is hydrogen.
- the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
- R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl. In some embodiments, R 2 is hydrogen or Ci- Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl.
- the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
- R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl.
- R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is hydrogen. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl.
- R 6 is hydrogen or Ci-Ce alkyl. In some embodiments, R 6 is hydrogen. In some embodiments, R 6 is Ci-Ce alkyl. In some embodiments, R 6 is methyl.
- Q is -CH2- or ethylene glycol. In some embodiments, Q is -CH2- . In some embodiments, Q is ethylene glycol.
- m is 1-20. In some embodiments, m is 1-10. In some embodiments, m is 1-5. In some embodiments, m is 5-10. In some embodiments, m is 10-20. In some embodiments, m is 2. In some embodiments, m is 1. In some embodiments, m is 3, 4, 5, 6, 7, 8, 9, or 10.
- A is a polymeric side chain comprising a recurring unit derived from a monomer represented by the structure:
- R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl. In some embodiments, R 2 is hydrogen or Ci- Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl. In some embodiments, R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is hydrogen. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl.
- R 1 is hydrogen
- R 2 is hydrogen
- R 3 is hydrogen
- R 5 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, -C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne.
- R 5 is polyethylene glycol. In some embodiments, R 5 is polyethylene glycol with a chain length of 9. In some embodiments, R 1 is hydrogen, R 2 is hydrogen, R 3 is methyl, Q is -CH2-, R 6 is methyl, and R 5 is polyethylene glycol.
- the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
- a recurring unit is derived from a dimethacrylate monomer.
- the dimethacrylate monomer is a cross-linking monomer.
- R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl. In some embodiments, R 2 is hydrogen or Ci- Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl. In some embodiments, R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is hydrogen. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl. In some embodiments, R 1 is hydrogen or Ci-Ce alkyl.
- R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl. In some embodiments, R 2 is hydrogen or Ci-Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl. In some embodiments, R 3 is hydrogen, Ci- Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is hydrogen. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl. In some embodiments, Z is -O- or -NH. In some embodiments, Z is -O-. In some embodiments, Z is -NH.
- R 8 is Ci-Ce alkyl, divalent metal, or symmetric or asymmetric disulfide. In some embodiments, R 8 is Ci-Ce alkyl. In some embodiments, R 8 is a divalent metal. In some embodiments, R 8 is cadmium(II). In some embodiments, R 8 is symmetric or asymmetric disulfide. In some embodiments, R 8 is a symmetric disulfide. In some embodiments, R 8 is symmetrical disulfide (e.g., CH2CH2S-SCH2CH2) or a divalent metal.
- n 1 is an integer selected from 1-100. In some embodiments, n 1 is an integer selected from 5-100. In some embodiments, n 1 is an integer selected from 5-50. In some embodiments, n 1 is an integer selected from 50-100. In some embodiments, n 1 is an integer selected from 10-40. In some embodiments, n 1 is an integer selected from 1-20. In some embodiments, n 1 is an integer selected from 1-10. In some embodiments, n 1 is an integer selected from 1-5. In some embodiments, n 1 is an integer selected from 5-10. In some embodiments, n 1 is an integer selected from 10-20. In some embodiments, n 1 is 9. In some embodiments, n 1 is an integer selected from 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10.
- any of the macromolecule chains provided herein may be terminated by a terminating group.
- the terminating group may be a halogen, epoxide, or olefin (e.g., alkylene).
- the terminating group is a halogen.
- the terminating group is a bromine.
- the terminating group is an iodine.
- the terminating group is a chloride.
- the terminating group is an epoxide.
- the terminating group is an olefin.
- the macromolecule chains of the macromolecule structures provided herein comprise a recurring unit derived from a polyalkylene glycol (PAG) methacrylate, polyalkylene glycol (PAG) acrylate, polyalkylene glycol (PAG) methacrylamide, polyalkylene glycol (PAG) acrylamide, polyalkylene glycol (PAG) vinyl ether, and combinations thereof, where the number of alkylene oxide units is from 1 to 100 (e.g., 5 to 100, 5 to 90, 8 to 50, 8 to 30, and 10 to 25), and wherein the polyalkylene glycol (e.g., polyethylene glycol) is optionally terminated with Ci-Cs alkoxy group, such as methoxy, ethoxy, propoxy, butoxy, and the like.
- PAG polyalkylene glycol
- PAG polyalkylene glycol
- PAG polyalkylene glycol
- PAG polyalkylene glycol
- PAG polyalkylene glycol
- PAG polyalkylene
- the macromolecule chains of the macromolecule structures provided herein comprise a recurring unit derived from a polyethylene glycol (PEG) methacrylate, polyethylene glycol (PEG) acrylate, polyethylene glycol (PEG) methacrylamide, polyethylene glycol (PEG) acrylamide, polyethylene glycol (PEG) vinyl ether, and combinations thereof, where the number of alkylene oxide units is from 1 to 100 (e.g., 5 to 100, 5 to 90, 8 to 50, 8 to 30, and 10 to 25), and wherein the polyalkylene glycol (e.g., polyethylene glycol) is optionally terminated with Ci-Cs alkoxy group, such as methoxy, ethoxy, propoxy, butoxy, and the like.
- the macromolecule chains of the macromolecule structures provided herein comprise a recurring unit derived from a monomer represented by a structure of Table 2. In some embodiments, macromolecule chains comprise two or more distinct recurring units derived from a monomer represented by a structure of Table 2.
- the recurring units provided herein are randomly distributed throughout the macromolecule chain. In some embodiments, the recurring units provided herein have a controlled distribution throughout the macromolecule chain. In some embodiments, the macromolecule chain is a homopolymer. In some embodiments, the macromolecule chain is a block copolymer. In some embodiments, the macromolecule chain is a random copolymer. The skilled artisan, guided by the disclosure herein, would understand how to provide polymers with randomly or controlled distributions of recurring units.
- the multiple recurring units may be present in equal or differing ratios.
- the ratio of distinct recurring units may be controlled according to methods known to one skilled in the art, including modifying the stoichiometry of monomers added.
- the macromolecule structures comprise (I) a surface and (II) a macromolecule chain coupled to the surface.
- the macromolecule chain comprises a recurring unit of Formula (I):
- R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl. In some embodiments, R 1 is methyl. In some embodiments, R 2 is hydrogen or Ci-Ce alkyl. In some embodiments, R 2 is Ci-Ce alkyl. In some embodiments, R 2 is methyl. In some embodiments, R 3 is hydrogen or Ci-Ce alkyl. In some embodiments, R 3 is hydrogen. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl. In some embodiments, R 1 , R 2 , and R 3 are Ci-Ce alkyl (e.g., methyl).
- n is an integer from 1 to 10,000. In some embodiments, n is an integer from 1 to 200. In some embodiments n is an integer of no more than 25,000 (e.g., no more than 20,000, no more than 15,000, no more than 10,000, no more than 5,000, no more than 2,500, no more than 1,000, no more than 500, no more than 100, no more than 50). In some embodiments, n is an integer from 1 to 5,000. In some embodiments, n is an integer from 1 to 2,500. In some embodiments, n is an integer from 1 to 1,000. In some embodiments, n is an integer of no more than 200. In some embodiments, n is an integer from 1 to 100. In some embodiments, n is an integer from 1 to 50.
- A is a polymeric side chain comprising any of the recurring units provided elsewhere herein.
- A is a poly(alkylene oxide) methacrylate or poly(alkylene oxide) acrylate.
- A is poly(ethylene glycol) methacrylate or poly(ethylene glycol) acrylate.
- A is a poly(alkylene oxide) methacrylamide or poly(alkylene oxide) acrylamide.
- A is poly(ethylene glycol) methacrylamide or poly(ethylene glycol) acrylamide.
- the macromolecule structure further comprises a cross-linking moiety.
- the cross-linking moiety comprises a structure represented by:
- the cross-linking moiety is a derived from a dimethacrylate monomer, such as a dimethacrylate monomer as described elsewhere herein.
- R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl.
- R 2 is hydrogen or Ci-Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl.
- R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl.
- R 3 is hydrogen. In some embodiments, R 3 is Ci-Ce alkyl. In some embodiments, R 3 is methyl. In some embodiments, R 1 is hydrogen or Ci-Ce alkyl. In some embodiments, R 1 is hydrogen. In some embodiments, R 1 is Ci-Ce alkyl. In some embodiments, R 2 is hydrogen or Ci-Ce alkyl. In some embodiments, R 2 is hydrogen. In some embodiments, R 2 is Ci-Ce alkyl. In some embodiments, R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R 3 is hydrogen.
- the tethering moiety comprises the structure:
- the macromolecule chains comprise from 1 to 1,000 recurring units. In some embodiments, the macromolecule chains comprise at least 1, at least 10, at least 50, at least 100, at least 250, at least 500, at least 750, or at least 1,000 recurring units. In some embodiments, the macromolecule chain comprises at most 2,500 recurring units. In some embodiments , the macromolecule chains comprise at most 1,000 recurring units. In some embodiments, the macromolecule chains comprise at most 750 recurring units. In some embodiments, the macromolecule chains comprise at most 500 recurring units. In some embodiments, the macromolecule chains comprise at most 250 recurring units. In some embodiments, the macromolecule chains comprise at most 100 recurring units.
- the macromolecule chains comprise about 1 to about 100 recurring units. In some embodiments, the macromolecule chains comprise about 1 to about 250 recurring units. In some embodiments, the macromolecule chains comprise about 1 to about 500 recurring units. In some embodiments, the macromolecule chains comprise about 1 to about 1,000 recurring units. In some embodiments, the macromolecule chains comprise about 100 to about 1,000 recurring units. In some embodiments, the macromolecule chains comprise about 1 to about 10 recurring units.
- the macromolecule chains comprise a molecular weight of about 0.1 kDa to about 500 kDa. In some embodiments, the macromolecule chains comprise a molecular weight of at least 0.1 kDa, at least 1 kDa, at least 5 kDa, at least 10 kDa, at least 20 kDa, at least 25 kDa, at least 50 kDa, at least 100 kDa, at least 250 kDa, or at least 500 kDa.
- the macromolecule chains comprise a molecular weight of no more than 1000 kDa, no more than 500 kDa, no more than 750 kDa, no more than 500 kDa, no more than 250 kDa, no more than 100 kDa, no more than 75 kDa, no more than 50 kDa, no more than 40 kDa, no more than 30 kDa, no more than 25 kDa, no more than 20 kDa, no more than 15 kDa, or no more than 10 kDa.
- the macromolecule chain comprises a molecular weight of about 0.1 to about 500 kDa, about 0.1 to about 250 kDa, about 0.1 kDa to about 100 kDa, about 0.1 kDa to about 70 kDa, 0.5 kDa to about 10 kDa, 0.5 kDa to about 15 kDa, or about 1 kDa toa bout 25 kDa.
- the macromolecule chain comprises a molecular weight of about 0.1 kDa to about 100 kDa.
- the macromolecule chain comprises a molecular weight of about 0.1 kDa to about 50 kDa.
- the macromolecular chain comprises a block co-polymer.
- the block co-polymer comprises a first block derived from a first monomer and a second block derived from a second monomer, wherein the first block is adjacent to the tethering moiety, and wherein the first monomer is more hydrophobic than the second monomer.
- the block co-polymer comprises a first block derived from a first monomer and a second block derived from a second monomer, wherein the first block is adjacent to the tethering moiety, and wherein the first monomer is less hydrophobic than the second monomer.
- the hydrophobicity of the first monomer and the second monomer can be determined by the estimated partition coefficient using XLOGP3.
- an absolute difference between the estimated partition coefficients of the first monomer and the second monomer is at least 0.3, at least 0.5, at least 0.8, at least 1, at least 1.5, or at least 2.
- an absolute difference between the estimated partition coefficients of the first monomer and the second monomer is no more than 3, no more than 2.5, no more than 2, no more than 1, or no more than 0.8.
- any of the macromolecule structures provided herein, such as used in the methods provided herein, comprise a surface.
- a surface is a particle.
- a particle is a nanoparticle or a microparticle.
- the particle is a nanoparticle.
- the particle is a microparticle.
- the particles provided herein have a diameter of at least 10 nm, at least 100 nm, at least 200 nm, at least 300 nm, at least 400 nm, at least 500 nm, at least 600 nm, at least 700 nm, at least 800 nm, or at least 900 nm.
- the particles provided herein have a diameter of no more than 5000 nm, no more than 4000 nm, no more than 3000 nm, no more than 2000 nm, no more than 1000 nm, no more than 750 nm, or no more than 500 nm.
- the particles provided herein have a diameter of from 10 nm to 50 nm, from 50 nm to 100 nm, from 100 nm to 150 nm, from 150 nm to 200 nm, from 200 nm to 250 nm, from 250 nm to 300 nm, from 300 nm to 350 nm, from 350 nm to 400 nm, from 400 nm to 450 nm, from 450 nm to 500 nm, from 500 nm to 550 nm, from 550 nm to 600 nm, from 600 nm to 650 nm, from 650 nm to 700 nm, from 700 nm to 750 nm, from 750 nm to 800 nm, from 800 nm to 850 nm, from 850 nm to 900 nm, from 100 nm to 300 nm, from 150 nm to 350 nm, from 200 nm to 400 n
- the particles have a diameter of about 100 to about 400 nm. In some embodiments, the particles have a size of about 100 to about 300 nm.
- the particle size (e.g., diameter) can be measured by dynamic light scattering (DLS) as an indirect measure of size.
- the DLS measurement can be an ‘intensity -weighted’ average, which means the size distribution that the mean is calculated from can be weighted by the sixth power of radius. This can be referred to herein as ‘z-average’ or ‘intensity-mean’.
- Particle size can also be measured by electron microscopy (e.g., SEM, TEM).
- the particles provided herein may comprise a diameter of about 100 nm to about 500 nm. In some embodiments, the particles comprise a diameter of about 100 nm to about 300 nm. In some embodiments, the particles comprise a diameter of about 100 nm to about 200 nm. In some embodiments, the particles comprise a diameter of about 150 nm to about 250 nm.
- the particle size effects pull-down rate. In some instances, the particle size does not affect the ability of the macromolecule structure to isolate biomolecules, such as the ability to isolate unique biomolecules.
- particles can have a homogenous size distribution or a heterogeneous size distribution.
- Poly dispersity index (PDI) which can be measured by techniques such as dynamic light scattering, is a measure of the size distribution.
- a low PDI indicates a more homogeneous size distribution and a higher PDI indicates a more heterogeneous size distribution.
- particles disclosed herein can have a PDI of less than 0.5, less than 0.4, less than 0.3, less than 0.2, less than 0.15, or less than 0.1.
- the particles disclosed herein have a PDI of less than 0.1.
- the particles may have a PDI of at least 0.5, or at least 0.75.
- Particles disclosed herein can have a range of different surface charges. Particles can be negatively charged, positively charged, or neutral in charge. In some embodiments, particles have a surface charge of -150 mV to -100 mV, -100 mV to -90 mV, -90 mV to -80 mV, -80 mV to -70 mV, -70 mV to -60 mV, -60 mV to -50 mV, -50 mV to -40 mV, -40 mV to -30 mV, -30 mV to - 20 mV, -20 mV to -10 mV, -10 mV to 0 mV, -10 mV to 10 mV, 0 mV to 10 mV, 10 mV to 20 mV, 20 mV to 30 mV, 30 mV to 40 mV, 40 mV to 50 mV, 50 mV, 50
- particles disclosed herein have a surface charge of -60 mV to 60 mV.
- the surface charge may be evaluated by zeta potential analysis at about neutral pH using an appropriate buffer (e.g., phosphate-buffered saline).
- particles may be spherical, colloidal, square shaped, rods, wires, cones, pyramids, or oblong.
- the surface comprises any suitable material according to one skilled in the art.
- the particle is magnetic, such as any magnetic material suitable according to one skilled in the art.
- the particle comprises a metal material.
- the metal material comprises any one of or any combination of gold, silver, copper, nickel, cobalt, palladium, platinum, iridium, osmium, rhodium, ruthenium, rhenium, vanadium, chromium, manganese, niobium, molybdenum, tungsten, tantalum, iron and cadmium.
- the particle comprises iron oxide.
- the particle is a superparamagnetic iron oxide particle. In some embodiments, the particle comprises superparamagnetic iron oxide nanoparticles. In some embodiments, the particle has a core-shell structure. In some embodiments, the particle has an iron oxide core. In some embodiments, the particle comprises magnetite. In some embodiments, the particle comprises a silica shell. In some embodiments, the particle comprises an iron oxide core with a silica shell. In some instances, the silica shell can be functionalized with the tethering moieties or macromolecule chains provided elsewhere herein. In some embodiments, the particle comprises iron oxide crystals. In some embodiments, the particle comprises polystyrene. In some embodiments, the particle comprises iron oxide crystals embedded in a polystyrene core.
- provided herein are systems for performing any of the methods provided herein.
- systems for isolating one or more biomolecules from a biological solution In some embodiments, provided herein are systems for isolating one or more proteins (e.g., peptides) from a biological solution.
- proteins e.g., peptides
- the systems provided herein comprise (a) one or more macromolecule structures (e.g., such as those described elsewhere herein), (b) a suspension solution, (c) a biological sample comprising one or more proteins, and (d) an automated system comprising a network of units with differentiated functions configured to perform any one of the methods provided herein.
- the systems comprise one or more macromolecule structures.
- the systems comprise a biological sample comprising one or more proteins.
- the systems comprise an automated system comprising a network of units with differentiated functions configured to perform any one of the methods provided herein (e.g., such as isolating one or more biomolecules (e.g., proteins or peptides) from a biological solution).
- a network of units with differentiated functions configured to perform any one of the methods provided herein (e.g., such as isolating one or more biomolecules (e.g., proteins or peptides) from a biological solution).
- the macromolecule structure(s) as described elsewhere herein, the suspension solution, and the biological sample comprising a concentration of protein may be incubated at a temperature of about 10 degrees Celsius (°C) to about 100 °C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 20°C to about 90°C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 20°C to about 50°C.
- the suspension solution may comprise Tris, EDTA, and CHAPS buffer.
- the suspension solution may be Tris, EDTA in 150 millimolar (mM) KC1 and 0.05% CHAPS buffer. In another example, the suspension solution may be lOmM Tris HC1 pH 7.4, 1 mM EDTA.
- the (e.g., automated) system provided herein comprises a network of units which may comprise differentiated functions in isolating one or more biomolecules from a biological sample.
- the network of units comprises (a) a first unit comprising a multichannel fluid transfer instrument for transferring fluids between units within the system, (b) a second unit comprising a support for storing a plurality of biological samples, and (c) a third unit comprising a support for an array plate possessing partitions that comprise the one or more macromolecule structure for binding of the one or more proteins within the biological sample with the one or more macromolecule structure.
- the network of units comprises a unit comprises a multichannel fluid transfer instrument for transferring fluids between units within the system.
- the network of units comprises a unit comprising a support for storing a plurality of biological samples.
- the network of units comprises a unit comprising a support for an array plate possessing partitions that comprise the one or more macromolecule structure for binding of the one or more proteins within the biological sample with the one or more macromolecule structure.
- the network of units may comprise a (e.g., fourth) unit comprising supports for storing a plurality of reagents.
- the network of units may comprise a (e.g., fifth) unit comprising supports for storing a reagent to be disposed of. In some embodiments, the network of units may comprise a (e.g., sixth) unit comprising supports for storing consumables used by the multichannel fluid transfer instrument.
- the system is programmed to perform a series of steps comprising contacting the biological sample with a specified partition of the array, optionally incubating the biological sample with the macromolecule structures contained within the partition of the array plate, removing components from a partition except the macromolecule structures and the one or more biomolecules (e.g., proteins or peptides) interacting with the macromolecule structure, and eluting the one or more biomolecules (e.g., proteins or peptides) from the macromolecule structures.
- the system is further programmed to prepare the eluted (e.g., isolated) biomolecules (e.g., proteins or peptides) for analysis, such as by mass spectrometry or any other analysis as described elsewhere herein.
- the first unit comprises a degree of mobility that enables access to all other units within the system. In some embodiments, the first unit comprises a capacity to perform pipetting functions.
- the support of the second and/or third unit comprises support for a single plate, a 6 well plate, a 12 well plate, a 96 well plate, or a rack of microtubes.
- the second and/or unit comprises a thermal unit capable of modulating the temperature of said support and a sample.
- the second and/or third unit comprises a rotational unit capable of physically agitating and/or mixing a sample.
- the system comprises a step wherein the array plate is transferred to a (e.g., seventh) unit that comprises a magnetized support and a thermal unit capable of modulating the temperature of said support and a sample and incubated for an additional amount of time.
- a (e.g., seventh) unit that comprises a magnetized support and a thermal unit capable of modulating the temperature of said support and a sample and incubated for an additional amount of time.
- the fourth unit comprises a set of reagents for: generating the array plate, washing the species of the biological sample that are not bound to the macromolecule structure(s), and/or preparing a sample for mass spectrometry.
- contacting the biological sample with a specified partition of the array comprises pipetting a specified volume of the biological sample into the specific partition of the array.
- contacting the biological sample with a specified partition of the array comprises pipetting a volume of at least 10 microliters, at least 20 microliters at least 50 microliters, at least 100 microliters, at least 250 microliters, at least 500 microliters, or at least 1000 microliters of the biological sample into the specific partition of the array. In some embodiments, contacting the biological sample with a specified partition of the array comprises pipetting a volume of no more than 1000 microliters, no more than 500 microliters, nor more than 250 microliters, no more than 150 microliters, no more than 100 microliters, no more than 75 microliters, no more than 50 microliters, or no more than 30 microliters.
- incubating the biological sample e.g., comprising biomolecules (e.g., proteins or peptides)
- the macromolecule structure(s) contained within the partition of the array plate comprises an incubation time of at least about 10 seconds, at least about 15 seconds, at least about 20 seconds, at least about 25 seconds, at least about 30 seconds, at least about 40 seconds, at least about 50 seconds, at least about 60 seconds, at least about 90 seconds, at least about 2 minutes, at least about 3 minutes, at least about 4 minutes, at least about 5 minutes, at least about 6 minutes, at least about 7 minutes, at least about 8 minutes, at least about 9 minutes, at least about 10 minutes, at least about 15 minutes, at least about 20 minutes, at least about 25 minutes, at least about 30 minutes, at least about 45 minutes, at least about 50 minutes, at least about 60 minutes, at least about 90 minutes, at least about 2 hours, at least about 3 hours, at least about 4 hours, at least about 5 hours, at least about 6 hours, at least about 7 hours
- incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the array plate comprises an incubation time of no more than 24 hours, no more than 12 hours, no more than 6 hours, no more than 3 hours, no more than 2 hours, no more than 90 minutes, no more than 75 minutes, no more than 60 minutes, no more than 45 minutes, or no more than 30 minutes.
- incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the array plate comprises an incubation time of 30 minutes and 3 hours.
- incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the substrate comprises an incubation temperature between about 4° C to about 40° C.
- incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the substrate may comprise an incubation temperature between about 4° C to about 37° C.
- Incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the substrate may comprise an incubation temperature between about 20° C to about 50° C.
- Incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the substrate may comprise an incubation temperature between about 4° C to about 100° C.
- the system is programmed to complete a series of wash steps.
- the wash steps are completed with an organic solvent or an aqueous solvent, such as an organic solvent or an aqueous solvent as described elsewhere herein.
- the wash steps are used to remove all components from a partition except the macromolecule structure(s) and the biomolecules (e.g., proteins or peptides) interacting with the macromolecule structure(s).
- the second unit can facilitate a transfer of the sample for mass spectrometry to a mass spectrometry unit.
- the system provide herein is an automated system.
- kits for performing any one of the methods provided herein are kits for performing any one of the methods provided herein.
- kits for isolating one or more biomolecules e.g., proteins or peptides
- kits for isolating one or more proteins from a biological solution In some embodiments, provided herein are kits for isolating one or more peptides from a biological solution.
- kits provided herein comprise (a) one or more macromolecule structure(s), (b) an organic solvent configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure, (c) a washing agent configured to wash proteins bound to the macromolecule structure, and (d) an elution agent configured to elute proteins from the macromolecule structure.
- kits provided herein comprise (a) a first macromolecule structure and optionally a second macromolecule structure, (b) an organic solvent configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure(s), (c) one or more washing agents configured to wash proteins bound to the first macromolecule structure and optionally the second macromolecule structure, (d) an elution agent configured to elute proteins from the first macromolecule structure, (e) a reducing agent, and (f) a protein digesting agent.
- kits provided herein comprise one or more macromolecule structure(s), such as any of the macromolecule structures provided elsewhere herein.
- kits provided herein comprise an organic solvent, such as an organic solvent as provided elsewhere herein.
- an organic solvent is configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure.
- kits provided herein comprise a washing agent, such as an organic solvent (or wash solvent) as provided elsewhere herein.
- the washing agent is configured to wash proteins bound to the macromolecule structure.
- kits provided herein comprise an elution agent, such as an aqueous solvent as provided elsewhere herein.
- an elution agent is configured to elute proteins from the macromolecule structure.
- kits provided herein comprise a protein digesting agent.
- a protein digesting agent produces one or more peptides.
- a protein digesting agent comprises trypsin, lysin, serine protease.
- a protein digesting agent comprises trypsin.
- kits provided herein comprise a reducing agent.
- the reducing agent comprises TCEP, dithiothreitol, beta-mercaptoethanol, glutathione, cysteine, or any combination thereof.
- Embodiment 1 A method of isolating one or more biomolecules from a biological sample, the method comprising: a. contacting a population of biomolecules comprising one or more proteins in the biological sample with a macromolecule structure in the presence of an organic solvent to bind the one or more proteins to the macromolecule structure; and b.
- the macromolecule structures comprise a surface and a macromolecule comprising recurring units of a first component and a second component, wherein the first component comprises a structure of Component (A) and the second component comprises a structure of Component (B’): wherein each of Yi, Y2, and Y3 is independently selected from hydrogen or Ci-Ce alkyl; each of Xi, X2, and X3 is independently selected from hydrogen or Ci-Ce alkyl;
- Ri is hydrogen, optionally substituted succinate, Ci-Ce alkyl substituted with sulfone, or phthalate;
- R2 is C1-C12 alkyl substituted with amine, Ci-Ce alkyl substituted with hydroxy, optionally substituted aryl, optionally substituted heteroaryl, optionally substituted C3-C6 heterocycloalkyl, two or more fused 3-6 member rings; optionally substituted C3-C6 dicycloalkyl methane, Ci-Ce alkyl substituted with guanidine, Ci-Ce alkoxy, optionally substituted Ci-Ce alkyl substituted with disulfide, Ci-Ce alkyl substituted with thiol, optionally substituted succinate, optionally substituted Ci-Ce alkylamine, Ci-Ce alkyl acetamide, Ci-Ce aminophthalate, a boronic acid, Ci-Cn optionally substituted cycloalkyl, or a monosaccharide; or
- Ri and R.2 are taken together with the nitrogen to which they are attached to form an optionally substituted heterocycle;
- Ri and R.2 are taken together with the nitrogen to which they are attached to form an azide group
- R4 is hydrogen or Ci-Ce alkyl substituted with thiol
- Rs is succinate, optionally substituted aryl, or optionally substituted Ci-Ce alkyl substituted with disulfide; q is an integer between 1 and 6; and p is an integer between 1 and 20.
- Embodiment 2 The method of embodiment 1, wherein
- Embodiment 3 The method of embodiment 1, wherein
- Embodiment 4 The method of embodiment 1, wherein
- Embodiment 5 The method of any one of the previous embodiments, wherein R 1 is hydrogen.
- Embodiment 6 The method of any one of embodiments 1-5, wherein Ri is -
- Embodiment 7 The method of any one of embodiments 1-5, wherein R 2 is Ci-Ce alkoxy, Ci-12 alkyl substituted with amine, optionally substituted Ci-Ce alkylamine, -(CH2)I-6 imidazole -, (CH2)I-6 pyridine, optionally substituted aryl, -(CH2)I- 6N(CH3)2(CH2CH 2 CH 2 SOOOH), -(CH 2 )I-6N(CH2CH2CH 2 SOOOH)2, or C 2 -C 6 alkyl substituted with hydroxy.
- Embodiment 8 A method of isolating one or more biomolecules from a biological sample, the method comprising: a. contacting a population of biomolecules comprising one or more proteins in the biological sample with a macromolecule structure in the presence of an organic solvent to bind the one or more proteins to the macromolecule structure; and b. eluting the one or more proteins from the macromolecule structure, thereby providing the one or more isolated biomolecules, wherein, the macromolecule structure comprises:
- each of X and Y is independently -C-, -O-, or -N-; each Z is independently -O- or -NH;
- Q is -CH2- or ethylene glycol
- A is a polymeric side chain comprising a recurring unit derived from a monomer represented by the structure: m is an integer selected from 1-20;
- R 1 , R 2 , R 1 , R 2 , and R 3 is independently selected from hydrogen or -Ci-Ce alkyl;
- R 3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl;
- R 4 is absent, hydrogen, sulfonate, carboxylate, C1-C4 alkylene, or Ci-Ce alkyl optionally substituted with halogen;
- R 6 is hydrogen or Ci-Ce alkyl
- R 7 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, - C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne;
- R 8 is Ci-Ce alkyl, divalent metal, or symmetric or asymmetric disulfide
- R 9 is hydrogen or oxo; and n 1 is an integer selected from 1-100.
- Embodiment 9 The method of embodiment 8, wherein the macromolecule chain comprises the recurring unit derived from a monomer represented by the structure:
- Embodiment 10 The method of embodiment 8 or 9, wherein R 1 and R 1 is hydrogen.
- Embodiment 11 The method of any one of embodiments 8-10, wherein R 2 and R 2 is hydrogen.
- Embodiment 12 The method of any one of embodiments 8-11, wherein R 3 and R 3 hydrogen or Ci-Ce alkyl.
- Embodiment 13 The method of any one of embodiments 8-11, wherein R 3 and R 3 are hydrogen.
- Embodiment 14 The method of any one of embodiments 8-13, wherein Z is O.
- Embodiment 15 The method of any one of embodiments 8-13, wherein Z is NH.
- Embodiment 16 The method of any one of embodiments 8-15, wherein R 8 is Ci-Ce alkyl.
- Embodiment 17 The method of any one of embodiments 8-16, wherein R 8 is Ci alkyl.
- Embodiment 18 The method of embodiment 8, wherein the macromolecule chain comprises the recurring unit derived from a monomer represented by the structure:
- Embodiment 19 The method of embodiment 18, wherein R 1 is hydrogen.
- Embodiment 20 The method of embodiment 18 or 19, wherein R 2 is hydrogen.
- Embodiment 21 The method of any one of embodiments 18-20, wherein R 3 is Ci-Ce alkyl.
- Embodiment 23 The method of any one of embodiments 18-22, wherein Q is CEE.
- Embodiment 25 The method of any one of embodiments 18-24, wherein R 6 is Ci-Ce alkyl.
- Embodiment 26 The method of any one of embodiments 18-25, wherein R 6 is Ci alkyl.
- Embodiment 27 The method of any one of embodiments 18-26, wherein R 5 is Ci-Cni ethylene glycol.
- Embodiment 28 The method of embodiment 27, wherein nl is from 1 to 25.
- Embodiment 29 The method of embodiment 28, wherein nl is 9.
- Embodiment 30 The method of embodiment 8, wherein the macromolecule chain comprises the recurring unit derived from a monomer represented by the structure:
- Embodiment 31 The method of embodiment 30, wherein R 1 is hydrogen.
- Embodiment 32 The method of embodiment 30 or 31, wherein R 2 is hydrogen.
- Embodiment 33 The method of any one of embodiments 30-32, wherein R 3 is Ci-Ce alkyl.
- Embodiment 34 The method of any one of embodiments 30-33, wherein R 3 is Ci alkyl.
- Embodiment 35 The method of any one of embodiments 30-34, wherein R 5 is Ci-Cni ethylene glycol.
- Embodiment 36 The method of embodiment 35, wherein nl is from 1 to 10.
- Embodiment 37 The method of any one of embodiments 30-34, wherein R 5 is a 3-, 5-, or
- 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo.
- Embodiment 38 The method of any one of embodiments 30-34, wherein R 5 is Ci-Cs alkyl substituted with one or more hydroxyl, amine, or azide.
- Embodiment 39 The method of embodiment 38, wherein R 5 is C3 alkyl substituted with a hydroxyl and an azide.
- Embodiment 40 The method of embodiment 38, wherein R 5 is C3 alkyl substituted with a hydroxyl and an amine.
- Embodiment 41 The method of embodiment 8, wherein the macromolecule chain comprises the recurring unit derived from a monomer represented by the structure:
- Embodiment 42 The method of embodiment 41, wherein R 1 is hydrogen.
- Embodiment 43 The method of embodiment 41 or 42, wherein R 2 is hydrogen.
- Embodiment 44 The method of any one of embodiments 41-43, wherein R 3 is Ci-Ce alkyl.
- Embodiment 45 The method of any one of embodiments 41-44, wherein R 3 is Ci alkyl.
- Embodiment 46 The method of any one of embodiments 41-45, wherein each of X and Y is -C-.
- Embodiment 47 The method of any one of embodiments 41-46, wherein R 4 is sulfonate.
- Embodiment 48 The method of any one of embodiments 8-47, wherein the macromolecule chain comprises one distinct recurring unit.
- Embodiment 49 The method of any one of embodiments 8-47, wherein the macromolecule chain comprises two distinct recurring units.
- Embodiment 50 The method of any one of the previous embodiments, wherein the macromolecule structure is selected from Table 4.
- Embodiment 51 The method of any one of the previous embodiments, wherein the macromolecule structure is selected from Table 1.
- Embodiment 52 The method of any one of the embodiments 8-50, wherein the macromolecule structure comprises:
- Embodiment 53 The method of any one of the previous embodiments, wherein the one or more proteins comprises peptides.
- Embodiment 54 The method of any one of the previous embodiments, wherein the surface is a particle.
- Embodiment 55 The method of any one of the previous embodiments, wherein the surface is a microparticle or a nanoparticle.
- Embodiment 56 The method of any one of the previous embodiments, wherein the surface is a nanoparticle.
- Embodiment 57 The method of any one of the previous embodiments, wherein the surface comprises iron oxide.
- Embodiment 58 The method of any one of the previous embodiments, wherein the surface comprises magnetite.
- Embodiment 59 The method of any one of the previous embodiments, wherein the surface comprises superparamagnetic iron oxide nanoparticle(s).
- Embodiment 60 The method of any one of the previous embodiments, wherein the surface comprises an iron oxide core.
- Embodiment 61 The method of any one of the previous embodiments, wherein the particle comprises an iron oxide core with a silica shell coating.
- Embodiment 62 The method of any one of the previous embodiments, wherein the macromolecule structure has a size of at least 100 nm.
- Embodiment 63 The method of any one of the previous embodiments, wherein the macromolecule structure has a size of about 100 nm to about 400 nm.
- Embodiment 64 The method of any one of the previous embodiments, wherein the method further comprises separating the macromolecule structure from the one or more proteins after (a).
- Embodiment 65 The method of embodiment 64, wherein separating is magnetic separation, centrifugation, or a combination thereof.
- Embodiment 66 The method of any one of the previous embodiments, wherein before (b) the method further comprises digesting one or more proteins bound to the macromolecule structure to form one or more peptides.
- Embodiment 67 The method of any one of the previous embodiments, wherein eluting the one or more proteins comprises eluting the one or more peptides.
- Embodiment 68 The method of any one of the previous embodiments, wherein the one or more isolated biomolecules comprises one or more isolated proteins.
- Embodiment 69 The method of any one of the previous embodiments, wherein the one or more isolated biomolecules comprises one or more isolated peptides.
- Embodiment 70 The method of any one of the previous embodiments, wherein after (a), the method further comprises washing the macromolecule structure comprising the one or more proteins with an organic solvent wash.
- Embodiment 71 The method of embodiment 70, wherein the organic solvent wash comprises at least 50 wt% (e.g., 80 wt%, 90 wt%, 95 wt%) of an organic solvent.
- Embodiment 72 The method of any one of the previous embodiments, wherein the organic solvent comprises an alcohol, acetonitrile, dichloromethane, dimethylformamide (DMF), dimethylsulfoxide (DMSO), ethylacetate, hexamethylphosphoramide (HPMA), or tetrahydrofuran.
- the organic solvent comprises an alcohol, acetonitrile, dichloromethane, dimethylformamide (DMF), dimethylsulfoxide (DMSO), ethylacetate, hexamethylphosphoramide (HPMA), or tetrahydrofuran.
- Embodiment 73 The method of any one of the previous embodiments, wherein the organic solvent comprises acetonitrile.
- Embodiment 74 The method of embodiment 72, wherein the alcohol is ethanol, n- propanol, isopropanol, n-butanol, sec-butanol, tert-butanol, or pentanol.
- Embodiment 75 The method of any one of the previous embodiments, wherein (b) comprises eluting the one or more proteins (e.g., or peptides) from the macromolecule structure with an aqueous solution.
- proteins e.g., or peptides
- Embodiment 76 The method of embodiment 75, wherein the aqueous solution comprises an organic solvent.
- Embodiment 77 The method of embodiment 75 or 76, wherein the aqueous solution comprises no more than 50 wt% (e.g., 40 wt%, 30 wt%, 20 wt%, 10 wt%, 5 wt%) of an organic solvent.
- Embodiment 78 The method of any one of the previous embodiments, wherein the biological sample is plasma, serum, CSF, urine, tear, cell lysate, tissue lysate, cell homogenate, tissue homogenate, nipple aspirate, fecal matter, synovial fluid, saliva or blood.
- Embodiment 79 The method of any one of the previous embodiments, where the method is capable of isolating from 100 to 20,000 (e.g., unique) peptides.
- Embodiment 80 The method of any one of embodiments 1-79, wherein the method is capable of isolating at least 1,000 (e.g., unique) peptides.
- Embodiment 81 The method of any one of embodiments 1-79, wherein the method is capable of isolating no more than 25,000 (e.g., unique) peptides.
- Embodiment 82 The method of any one of the previous embodiments, wherein isolating occurs in absence of a precipitation solution comprising salts in an amount effective to precipitate the population of biomolecules.
- Embodiment 83 The method of any one of the previous embodiments, wherein the method further comprises identifying the one or more peptides.
- Embodiment 84 The method of any one of the previous embodiments, wherein at least 90% of an amount of a surfactant is removed from the population of biomolecules.
- Embodiment 85 The method of any one of the previous embodiments, wherein at least 90% of an amount of a lysis reagent is removed from the population of biomolecules.
- Embodiment 86 The method of any one of the previous embodiments, wherein the method further comprises after (b), analyzing the one or more proteins.
- Embodiment 87 The method of embodiment 86, wherein the analyzing comprises mass spectrometry (MS), liquid chromatography-mass spectrometry (LC-MS), protein sequencing, or a combination thereof.
- MS mass spectrometry
- LC-MS liquid chromatography-mass spectrometry
- protein sequencing or a combination thereof.
- Embodiment 88 The method of embodiment 86 or 87, wherein analyzing is capable of identifying from 1 to 20,000 protein groups (e.g., from 1,000 to 10,000 protein groups, from 1,000 to 5,000 protein groups, from 1,200 to 2,200 protein groups).
- 1 to 20,000 protein groups e.g., from 1,000 to 10,000 protein groups, from 1,000 to 5,000 protein groups, from 1,200 to 2,200 protein groups.
- Embodiment 89 The method of any one of the previous embodiments, wherein before (a) the method further comprises contacting the biological sample comprising a population of proteins with an additional macromolecule structure to provide a protein corona.
- Embodiment 90 The method of any one of the previous embodiments, further comprising before (a) treating or digesting one or more proteins of the population of proteins to form one or more peptides or desorbed proteins.
- Embodiment 91 The method of embodiment 89 or 90, wherein the macromolecule structure and the additional macromolecule structure are the same.
- Embodiment 92 The method of embodiment 89 or 90, wherein the macromolecule structure and the additional macromolecule structure are different.
- Embodiment 93 The method of any one of the previous embodiments, wherein a mass of the one or more isolated proteins is at least 100 nanograms (ng) (e.g., 500 ng, 1 microgram (pg), 10 pg, 50 pg, 100 pg, 200 pg).
- ng nanograms
- pg 1 microgram
- 10 pg 50 pg
- 100 pg 200 pg
- Embodiment 94 A method of isolating one or more biomolecules from a biological sample, the method comprising: a. contacting the biomolecules comprising one or more proteins with a first macromolecule structure to provide a protein corona; b. optionally, digesting the biomolecules of the protein corona to form one or more peptides; c. separating the one or more peptides of the protein corona from the first macromolecule structure; d. contacting the one or more peptides with a second macromolecule structure and an organic solvent to bind at least a portion of the one or more peptides to the second macromolecule structure; and e. eluting the one or more peptides from the second macromolecule structure, thereby providing the one or more isolated biomolecules.
- Embodiment 95 The method of embodiment 94, wherein the first surface and the second macromolecule structure are different.
- Embodiment 96 The method of embodiment 94, wherein the first surface and the second macromolecule structure are the same.
- Embodiment 97 The method of any one of embodiments 94-96, wherein in (d) the second macromolecule structure is added to a composition comprising the first surface.
- Embodiment 98 The method of any one of embodiments 94-97, wherein the first macromolecule structure or the second macromolecule structure comprises a PEG group.
- Embodiment 99 The method of any one of embodiments 94-98, wherein the surface is a particle.
- Embodiment 100 The method of any one of embodiments 94-99, wherein the surface is a microparticle or a nanoparticle.
- Embodiment 101 The method of any one of embodiments 94-100, wherein the surface comprises iron oxide.
- Embodiment 102 The method of any one of embodiments 94-101, wherein the surface comprises magnetite.
- Embodiment 104 The method of any one of embodiments 94-103, wherein the method further comprises removing the one or more isolated proteins after (e).
- Embodiment 105 The method of any one of embodiments 94-104, wherein separating comprises magnetic separation, centrifugation, or a combination thereof.
- Embodiment 106 The method of any one of embodiments 94-105, wherein the organic solvent comprises an alcohol, acetonitrile, dichloromethane, dimethylformamide (DMF), dimethylsulfoxide (DMSO), ethylacetate, hexamethylphosphoramide (HPMA), or tetrahydrofuran.
- the organic solvent comprises an alcohol, acetonitrile, dichloromethane, dimethylformamide (DMF), dimethylsulfoxide (DMSO), ethylacetate, hexamethylphosphoramide (HPMA), or tetrahydrofuran.
- Embodiment 107 The method of any one of embodiments 94-106, wherein (d) comprises eluting the one or more proteins or the one or more peptides from the second surface with an aqueous solution.
- Embodiment 108 The method of any one of embodiments 94-107, wherein the biological sample is plasma, serum, CSF, urine, tear, cell lysate, tissue lysate, cell homogenate, tissue homogenate, nipple aspirate, fecal matter, synovial fluid, saliva or blood.
- the biological sample is plasma, serum, CSF, urine, tear, cell lysate, tissue lysate, cell homogenate, tissue homogenate, nipple aspirate, fecal matter, synovial fluid, saliva or blood.
- Embodiment 109 The method of any one of embodiments 94-108, where the method is capable of isolating at least 100 (e.g., unique) protein groups or peptides.
- Embodiment 110 The method of any one of embodiments 94-109, wherein isolation occurs in absence of a precipitation solution comprising salts in an amount effective to precipitate the one or more proteins or one or more peptides.
- Embodiment 111 The method of any one of embodiments 94-110, wherein the method further comprises identifying the one or more proteins or one or more peptides.
- Embodiment 112 The method of any one of embodiments 94-111, wherein a mass of the one or more isolated proteins or one or more isolated peptides is at least 100 nanograms (ng) (e.g., 500 ng, 1 microgram (pg), 10 pg, 50 pg, 100 pg, 200 pg).
- ng nanograms
- pg 1 microgram
- Embodiment 113 The method of any one of embodiments 1-112, wherein eluting comprises the successive application of two or more eluents.
- Embodiment 114 The method of embodiment 113, wherein the two or more eluents comprise eluents of varying polarity.
- Embodiment 115 A system for performing the method of any one of embodiments 1-114, the system comprising: a. one or more macromolecule structure; b. a suspension solution; c. a biological sample comprising one or more proteins; and d. an automated system comprising a network of units with differentiated functions configured to perform the method of any one of embodiments 1-114.
- Embodiment 116 The system of embodiment 115, wherein the network of units comprises: a. a first unit comprising a multichannel fluid transfer instrument for transferring fluids between units within the system; b. a second unit comprising a support for storing a plurality of biological samples; c. a third unit comprising a support for an array plate possessing partitions that comprise the one or more macromolecule structure for binding of the one or more proteins within the biological sample with the one or more macromolecule structures.
- Embodiment 117 The system of embodiment 115 or 116, wherein the network of units further comprises a fourth unit comprising supports for storing a plurality of reagents.
- Embodiment 118 The system of any one of embodiments 115-117, wherein the network of units further comprises a fifth unit comprising supports for storing a reagent to be disposed of.
- Embodiment 119 The system of any one of embodiments 115-118, wherein the network of units further comprises supports for storing consumables used by a multichannel fluid transfer instrument.
- Embodiment 120 A kit for performing the method of any one of embodiments 1 to 114 comprising: a. one or more macromolecule structures; b. an organic solvent configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure; c. a washing agent configured to wash proteins bound to the macromolecule structure; and d. an elution agent configured to elute proteins from the macromolecule structure.
- Initiator particle SPION@SiO 2 -APTES-Br (1.000 g), CuBr 2 (0.010 g), N,N,N’ ,N” ,N” - pentamethyldiethylenetriamine (PMDETA) (0.240 g), monomer Oligo(ethylene glycol) Methyl Ether Methacrylate (OEGMA, MW500) (5.000 g) and 25 mL of dimethylformamide (DMF) were added in a 3-neck flask. The mixture was sonicated for 15 min and purged with N2 for another 15 min.
- PMDETA N,N,N’ ,N” ,N” - pentamethyldiethylenetriamine
- OFDETA monomer Oligo(ethylene glycol) Methyl Ether Methacrylate
- DMF dimethylformamide
- Macromolecule structures 55-57 were synthesized according to modified versions of the syntheses described in Examples 1-3 and the scheme depicted in FIG. 3.
- the particle size was characterized by DLS, surface charge by zeta potential, and macromolecule chain% by TGA (weight loss%).
- the particle sizes were compared to the particle sizes before functionalization with the macromolecule chains. The resulting data is found in Table 5.
- Macromolecule structures 55, 56, and 59-61 were analyzed by thermogravimetric analysis for the % organics. The results of which are in Table 6. The % organics in the particles ranged from 9.14% to 15.88%, increased from the % organics found on the initiator surface before further functionalization.
- Table 7 is performance data for nanoparticles containing the surfaces disclosed in Table 1. Briefly, the nanoparticles were incubated with plasma samples at 37 °C for 1 hour to form protein coronas and then washed three times. The proteins were then lysed, alkylated, and digested before analysis using liquid chromatography tandem mass spectrometry.
- NP-1 through NP-5 are nanoparticles found in commercially available ProteographTM VI.2 kit.
- the Jaccard Index (JI) is a measure of similarity between in the protein groups (PG) identified between different nanoparticles.
- Example 6 Synthesis of P(GMA-co-D VB)-coated particle
- azobi si sobutyronitrile (AIBN) (450 mg) is dissolved in acetonitrile (4 mL).
- AIBN azobi si sobutyronitrile
- the round bottom flask is heated to 80 degrees Celsius, and once the flask reaches temperature, the AIBN solution is injected into the flask.
- the reaction suspension reacts at 80 degrees Celsius for 60 minutes.
- the reaction is monitored every 30 minutes until the particles have formed a size of about 350 nm and PDI of ⁇ 0.2.
- the reaction is quenched by adding benzoquinone (0.108 g) in acetonitrile (5 mL). Once the reaction is complete, the flask is removed from the heat and the particles are purified using THF.
- P(GMA-co-DVB)-coated particle (see Example 6) (1.10g) and DMF (220mL) are added. The mixture is mixed or sonicated at about 200 rpm for 20 minutes. The round bottom flask is cooled to 10 degrees Celsius while stirring under inert atmosphere, after which the flask is heated to 80 degrees Celsius. When the flask has reached 80 degrees Celsius, the ethylene diamine (6.86g) is injected into the flask using a syringe. The mixture is reacted for 16 hours. Once the reaction is complete, the flask is removed from the heat and the particles are purified using DMF
- Example 8 Synthesis of Macromolecule Structure 15 (P(GMA-co-DVB)-EDA-PA Particle [00265]
- P(GMA-co-DVB)-EDA-PA particles i.Og
- DMF N,N-dimethylformamide
- TEA triethylamine
- the mixture is mixed or sonicated at about 200 rpm for 20 minutes.
- phthalic anhydride (L50g) in DMF (lOmL) are combined.
- the round bottom flask is cooled to 10 degrees Celsius while stirring under inert atmosphere, after which the flask is heated to 80 degrees Celsius.
- a biological sample containing proteins and/or peptides is incubated in a well plate with a macromolecule structure (e.g., macromolecular structure 50 or 51 depicted in Table 1) to form a protein corona.
- a macromolecule structure e.g., macromolecular structure 50 or 51 depicted in Table 1
- the protein corona is then washed and resuspended in the well plate.
- the protein corona is then subjected to treatment to digest the proteins (or peptides) of the protein corona into peptides as described elsewhere herein.
- the peptides are then subjected to purification prior to isolating for analysis through mass spectrometry.
- Peptide purification is achieved through adding the digested proteins and peptides in a vessel (e.g., centrifuge tube) with a different macromolecule structure (e.g., macromolecule structures 425, 441, 55, and 56) and 100% acetonitrile.
- a vessel e.g., centrifuge tube
- macromolecule structure e.g., macromolecule structures 425, 441, 55, and 56
- the biomolecule and macromolecule structures are incubated for 8 minutes to form a biomolecule corona (comprising peptides) and then centrifuged at 600 rotations per minute.
- the biomolecule corona is pulled down in the vessel using a magnet, and the resulting supernatant is removed and the contents of the vessel are washed with 100% acetonitrile before another round of magnetic pulldown and removing the supernatant.
- the contents of the vessel e.g., biomolecule corona
- a 3% solution of acetonitrile in water are combined with a 3% solution of acetonitrile in water and incubated for 10 minutes at 28 degrees Celsius, thereby releasing the peptides from the macromolecule structures.
- the macromolecule structures are magnetically pulled down in the vessel and the purified, isolated peptides remain in the supernatant.
- the isolated peptides are collected and used downstream for analysis (e.g., mass spectrometry or LC-MS).
- Macromolecule structures 55 and 56 were synthesized according to schemes illustrated in FIG. 3 and FIG. 4, respectively. The size characterization of these particles is shown in Table
- Table 9a shows the results weight loss (%) for Macromolecule Structures 55-57 after 1 day at 40° Celsius in various solvent conditions as a proxy for stability of the particles.
- Table 9b shows the results weight loss (%) for Macromolecule Structures 55-57 after 8 days at 40° Celsius in various solvent conditions as a proxy for stability of the particles.
- each of the tested particles were found to be stable after 1 day and after 8 days (Table 9c).
- FIG. 8 illustrates peptide yield (in micrograms) for various macromolecule structures, where macromolecular structures of the disclosure yielding a mass of peptides between about 1 microgram to about 2 micrograms include macromolecule structures as described herein, such as described in Table 1 and Table 4.
- the macromolecular structures of the disclosure have improved peptide yield compared to commercially available beads, MPE and Cytva beads.
- MPE as a reference bead, was also found to yield between about 1 microgram to about 2 micrograms of peptide.
- FIG. 9a illustrates an example of a quantity of unique peptides identified for various macromolecule structures in comparison to reference surfaces (e.g., Monitored Multi-Flow, Positive Pressure, Evaporative Extraction module (MPE) used in the commercially available PROTEOGRAPH method or commercially available Cytiva beads).
- FIG. 9b illustrates an example of a quantity of unique protein groups identified for various macromolecule structures in comparison to reference surfaces (e.g., MPE or commercially available Cytiva beads).
- MPE Monitored Multi-Flow, Positive Pressure, Evaporative Extraction module
- FIG. 9b illustrates an example of a quantity of unique protein groups identified for various macromolecule structures in comparison to reference surfaces (e.g., MPE or commercially available Cytiva beads).
- commercially available Cytiva beads identify a lower amount of unique peptide groups and a lower amount of unique protein groups in comparison to the macromolecule structures of the disclosure.
- FIG. 10 illustrates an example of a difference in peptide identification rate (peptides/minute) between a sample obtained using the methods disclosed herein and a macromolecule structure of the disclosure (e.g., Macromolecule Structure 55) and a sample obtained using MPE.
- the peptides are analyzed by liquid chromatography mass spectrometry and the number of peptides identified over the retention time is shown.
- the rate of identification between a macromolecule of the disclosure and MPE is comparable over time.
- FIG. Ila illustrates peptide yield (in micrograms) in a solution with beads compared to a solution with 5x the concentration of beads. In some instances, a higher concentration of beads results in higher peptide yield across various macromolecule structures.
- FIG. 11b illustrates the number of unique peptides identified in a solution with beads compared to a solution with 5x the concentration of beads. In some instances, a higher concentration of beads results in a higher count of peptides across various macromolecule structures.
- beads are distributed into one or more containers, such as tubes. Any number of tubes may be used to contain beads (e.g., 2-10 tubes, each containing 4-20 beads per tube). The mass of beads per tube in some instances comprises 0.2-5 mg per bead. Each tube is filled with a solvent such as water to a volume of 100-1000 microliters before transfer to a plate, and one or more washings. Beads may then be resuspended to a concentration, such as 10-100 mg/mL and at a volume of 1-100 microliters.
- a solvent such as water
- Beads may then be resuspended to a concentration, such as 10-100 mg/mL and at a volume of 1-100 microliters.
- Binding Digested peptide samples are added to the wells of the plate (10-2000 microliters). A binding reagent is then added to each well, in a volume of 0.2-3000 mL, the wells incubated for 5-20 minutes, and beads resuspended in binding reagent comprising an organic solvent. The binding reagent is then removed.
- elution buffer is added (50-500 microliters) comprising 1-10% organic solvent in water, and then beads are incubated 5-20 minutes at 20-35 degrees C. After centrifugation (500-2000 rpm) and a 1-5 minute pull down, eluted peptides are transferred to a new plate.
Landscapes
- Chemical & Material Sciences (AREA)
- Analytical Chemistry (AREA)
- Organic Chemistry (AREA)
- Biophysics (AREA)
- Life Sciences & Earth Sciences (AREA)
- Biochemistry (AREA)
- Health & Medical Sciences (AREA)
- General Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Medicinal Chemistry (AREA)
- Molecular Biology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Peptides Or Proteins (AREA)
Abstract
Provided herein are methods of isolating biomolecules, such as proteins or peptides, from biological solutions comprising contacting the biological solution with one or more macromolecule structures in the presence of an organic solvent and eluting one or more biomolecules from the macromolecule structures. Also provided herein are systems and kits for carrying out said methods.
Description
BIOMOLECULE CLEANUP METHODS, KITS, AND SYSTEMS
CROSS-REFERENCE
[0001] This application claims the benefit of U.S. Provisional Application No. 63/596,934, filed November 7, 2023, which is incorporated herein by reference in its entirety.
BACKGROUND
[0002] Analysis of biomolecules in complex biological solutions by sensitive methods, such as mass spectrometry, often requires that the incompatible components are removed from the complex biological mixtures. Typically, the isolation of analytes from complex biological solutions relies on the laborious fine tuning of analytical parameters. There is a need for widely applicable methods to remove incompatible components of biological solutions in preparation for further analysis that don’t rely on typical methodologies such as reversed-phase or ion exchange chromatography or suspension trapping.
SUMMARY
[0003] Provided herein are methods for clean-up of complex biological solutions, such as removal of surfactants, detergents, chaotropes, salts, and other impurities or contaminants, without the need for methods such as solid phase chromatography or suspension trapping. In some instances, the methods provided herein provide for removal of mass-spectrometry incompatible components from biological samples. Provided herein are methods of isolating one or more biomolecules from a biological sample comprising contacting a population of biomolecules with one or more macromolecule structure(s), binding the biomolecules (e.g., proteins) to the macromolecule structure(s), and eluting the isolated biomolecules from the macromolecule structure(s). In some instances, the macromolecule structures provided herein provide fast pulldown times, facile re-suspension, and are filter plate compatible, which may allow for automation of the methods provided herein. Also provided herein are systems and kits performing the methods provided herein.
[0004] Provided herein, in some embodiments, is a method of isolating one or more biomolecules from a biological sample, the method comprising: a. contacting a population of biomolecules comprising one or more proteins in the biological sample with a macromolecule structure in the presence of an organic solvent to bind the one or more proteins to the macromolecule structure; and b. eluting the one or more proteins from the macromolecule structure, thereby providing the one or more isolated biomolecules,
wherein, the macromolecule structures comprise a surface and a macromolecule comprising recurring units of a first component and a second component, wherein the first component comprises a structure of Component (A) and the second component comprises a structure of Component (B’):
wherein each of Yi, Y2, and Y3 is independently selected from hydrogen or Ci-Ce alkyl; each of Xi, X2, and X3 is independently selected from hydrogen or Ci-Ce alkyl;
Ri is hydrogen, optionally substituted succinate, Ci-Ce alkyl substituted with sulfone, or phthalate;
R2 is C1-C12 alkyl substituted with amine, Ci-Ce alkyl substituted with hydroxy, optionally substituted aryl, optionally substituted heteroaryl, optionally substituted C3-C6 heterocycloalkyl, two or more fused 3-6 member rings; optionally substituted C3-C6 dicycloalkyl methane, Ci-Ce alkyl substituted with guanidine, Ci-Ce alkoxy, optionally substituted Ci-Ce alkyl substituted with disulfide, Ci-Ce alkyl substituted with thiol, optionally substituted succinate, optionally substituted Ci-Ce alkylamine, Ci-Ce alkyl acetamide, Ci-Ce aminophthalate, a boronic acid, C1-C11 optionally substituted cycloalkyl, or a monosaccharide; or
Ri and R2 are taken together with the nitrogen to which they are attached to form an optionally substituted heterocycle; or
Ri and R2 are taken together with the nitrogen to which they are attached to form an azide group;
R4 is hydrogen or Ci-Ce alkyl substituted with thiol;
Rs is succinate, optionally substituted aryl, or optionally substituted Ci-Ce alkyl substituted with disulfide;
q is an integer between 1 and 6; and p is an integer between 1 and 20.
Provided herein in some embodiments, is a method of isolating one or more biomolecules from a biological sample, the method comprising: a. contacting a population of biomolecules comprising one or more proteins in the biological sample with a macromolecule structure in the presence of an organic solvent to bind the one or more proteins to the macromolecule structure; and b. eluting the one or more proteins from the macromolecule structure, thereby providing the one or more isolated biomolecules, wherein, the macromolecule structure comprises:
(I) a surface;
(II) a tethering moiety coupled to the surface; and
(III) a macromolecule chain, wherein a first end of the macromolecule chain is covalently attached to the tethering moiety, and wherein the macromolecule chain comprises one or more distinct recurring units derived from a monomer represented by a structure selected from the group consisting of:
Q is -CH2- or ethylene glycol;
A is a polymeric side chain comprising a recurring unit derived from a monomer represented by the structure:
m is an integer selected from 1-20;
—is a single or double bond; each of R1, R2, R1 , R2 , and R3 is independently selected from hydrogen or -Ci-Ce alkyl;
R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl;
R4 is absent, hydrogen, sulfonate, carboxylate, C1-C4 alkylene, or Ci-Ce alkyl optionally substituted with halogen;
R5 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, - C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne;
R6 is hydrogen or Ci-Ce alkyl,
R7 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered
heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, - C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne;
R8 is Ci-Ce alkyl, divalent metal, or symmetric or asymmetric disulfide;
R9 is hydrogen or oxo; and n1 is an integer selected from 1-100. In some embodiments, R5 is Ci-Cni ethylene glycol. In some embodiments, nl is 9. In some embodiments, the macromolecule chain comprises the recurring unit derived from a monomer represented by the structure:
. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, the macromolecule structure is selected from Table 4 or Table 1 (e.g., as provided herein). In some embodiments, the macromolecule structure comprises:
macromolecule structure comprises: the macromolecule structure comprises
. In some embodiments, the surface is a microparticle or a nanoparticle. In some embodiments, the surface is a nanoparticle. In some embodiments, the method further comprises separating the macromolecule structure from the one or more proteins after (a). In some embodiments, separating is magnetic separation, centrifugation, or a combination thereof. In some embodiments, before (b) the method further comprises digesting one or more proteins bound to the macromolecule structure to form one or more peptides. In some embodiments, (b) comprises eluting the one or more proteins (e.g., or peptides) from the macromolecule structure with an aqueous solution. In some embodiments, the aqueous solution comprises an organic solvent. In some embodiments, the organic solvent comprises acetonitrile. In some embodiments, the microparticle or nanoparticle comprises a zeta potential of -10 to 10 mV. In some embodiments the microparticle or nanoparticle comprises an average diameter of
400 to 500 nm.
[0005] Provided herein, in some embodiments, is a method of isolating one or more biomolecules from a biological sample, the method comprising: a. contacting the biomolecules comprising one or more proteins with a first macromolecule structure to provide a protein corona; b. optionally, digesting the biomolecules of the protein corona to form one or more peptides;
c. separating the one or more peptides of the protein corona from the first macromolecule structure; d. contacting the one or more peptides with a second macromolecule structure and an organic solvent to bind at least a portion of the one or more peptides to the second macromolecule structure; and e. eluting the one or more peptides from the second macromolecule structure, thereby providing the one or more isolated biomolecules.
[0006] Provided herein, in some embodiments, is a system for performing a method provided herein, the system comprising: a. one or more macromolecule structure; b. a suspension solution; c. a biological sample comprising one or more proteins; and d. an automated system comprising a network of units with differentiated functions configured to perform the method.
[0007] Provided herein, in some embodiments, is a kit for performing a method provided herein, comprising: a. one or more macromolecule structures; b. an organic solvent configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure; c. a washing agent configured to wash proteins bound to the macromolecule structure; and d. an elution agent configured to elute proteins from the macromolecule structure.
BRIEF DESCRIPTION OF THE DRAWINGS
[0008] The novel features of the disclosure are set forth with particularity in the appended claims. A better understanding of the features and advantages of the present disclosure will be obtained by reference to the following detailed description that sets forth illustrative embodiments, in which the principles of the disclosure are utilized, and the accompanying drawings (also “Figure” and “FIG.” herein), of which:
[0009] FIG. 1 shows examples of the general design space of the epoxidated nanoparticle platform following addition of a functionalizing amine.
[0010] FIG. 2 shows examples of the conversion schemes available after epoxidation of nanoparticles including reaction with glycidyls and azides to obtain diverse functionalities.
[0011] FIG. 3 shows an example of a synthetic scheme for preparation of a macromolecule structure herein by contacting a monomer with an initiator surface.
[0012] FIG. 4 shows an example of a synthetic scheme for preparation of a macromolecule structure herein by contacting a vinyl-functionalized surface with a cross-linking monomer and a second monomer.
[0013] FIG. 5 shows examples of PEG structures that can be used in the macromolecule structures.
[0014] FIG. 6a shows a schematic of an embodiment of a method of isolating biomolecules (e.g., proteins or peptides)
[0015] FIG. 6b shows a schematic of an embodiment of a method of isolating biomolecules (e.g., proteins or peptides).
[0016] FIG. 7 illustrates a schematic of an embodiment of a method of isolating biomolecules for analysis. The peptide preparation may include a purification workflow using beads and an organic solvent.
[0017] FIG. 8 illustrates examples of peptide yield (in micrograms) for various macromolecule structures.
[0018] FIG. 9a illustrates examples of a quantity of unique peptides identified using liquid chromatography mass spectrometry for various macromolecule structures in comparison to commercially available surfaces (e.g., MPE or Cytiva beads).
[0019] FIG. 9b illustrates a quantity of unique protein groups identified using liquid chromatography mass spectrometry for various macromolecule structures in comparison to commercially available surfaces (e.g., MPE or Cytiva beads).
[0020] FIG. 10 illustrates an example of a difference in peptide identification rate (peptides/minute) between a macromolecule structure and MPE over elution time when performing liquid chromatography mass spectrometry.
[0021] FIG. Ila illustrates an example of peptide yield (in micrograms) in a solution with beads compared to a solution with 5x the concentration of beads.
[0022] FIG. 11b illustrates an example of the number of unique peptides identified using liquid chromatography mass spectrometry in a solution with beads compared to a solution with 5x the concentration of beads.
[0023] FIG. 12a illustrates an example of beads in a nanoparticle (NP) solution being pulled down through the use of a magnet.
[0024] FIG. 12b illustrates an example of the difference in rate of bead pulldown for two beads of different diameters.
DETAILED DESCRIPTION
Certain Definitions
[0025] As used herein and in the appended claims, the singular forms "a," "and," and "the" include plural referents unless the context clearly dictates otherwise. Thus, for example, reference to "an agent" includes a plurality of such agents, and reference to "the cell" includes reference to one or more cells (or to a plurality of cells) and equivalents thereof known to those skilled in the art, and so forth. When ranges are used herein for physical properties, such as molecular weight, or chemical properties, such as chemical formulae, all combinations and subcombinations of ranges and specific embodiments therein are intended to be included. The term "about" when referring to a number or a numerical range means that the number or numerical range referred to is an approximation within experimental variability (or within statistical experimental error), and thus the number or numerical range may vary between 1% and 15% of the stated number or numerical range. The term "comprising" (and related terms such as "comprise" or "comprises" or "having" or "including") is not intended to exclude that in other certain embodiments, for example, an embodiment of any composition of matter, composition, method, or process, or the like, described herein, may "consist of' or "consist essentially of' the described features.
[0026] “ Amino” refers to the -NH2 radical.
[0027] “Cyano” refers to the -CN radical.
[0028] “Nitro” refers to the -NO2 radical.
[0029] “ Oxo” refers to the =0 radical.
[0030] “Hydroxyl” refers to the -OH radical.
[0031] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as is commonly understood by one of skill in the art to which this disclosure belongs. All patents and publications referred to herein are incorporated by reference.
[0032] "Alkyl" refers to an optionally substituted straight-chain, or optionally substituted branched-chain saturated hydrocarbon mono-radical, and preferably having from one to fifteen carbon atoms (z.e., C1-C15 alkyl). In certain embodiments, an alkyl comprises one to thirteen carbon atoms (z.e., C1-C13 alkyl). In certain embodiments, an alkyl comprises one to eight carbon atoms (z.e., Ci-Cs alkyl). In other embodiments, an alkyl comprises one to five carbon atoms (z.e., C1-C5 alkyl). In other embodiments, an alkyl comprises one to four carbon atoms (z.e., C1-C4 alkyl). In other embodiments, an alkyl comprises one to three carbon atoms (z.e., C1-C3 alkyl). In other embodiments, an alkyl comprises one to two carbon atoms (z.e., C1-C2 alkyl). Whenever it appears herein, a numerical range such as “C1-C3 alkyl” means that the alkyl group consists of 1 carbon atom, 2 carbon atoms, or 3 carbon atoms. In other embodiments, an alkyl comprises one carbon atom (z.e., Ci alkyl). In other embodiments, an alkyl comprises five to fifteen carbon atoms
(/.e., C5-C15 alkyl). In other embodiments, an alkyl comprises five to eight carbon atoms (/.e., C5- Cs alkyl). In other embodiments, an alkyl comprises two to five carbon atoms (/.e., C2-C5 alkyl). In other embodiments, an alkyl comprises three to five carbon atoms (/.e., C3-C5 alkyl). In certain embodiments, the alkyl group is selected from methyl, ethyl, 1 -propyl (//-propyl), 1 -methylethyl (/.w-propyl), 1 -butyl (//-butyl), 1 -methylpropyl (.scc-butyl), 2-m ethylpropyl (/.w-butyl),
1.1 -dimethylethyl (tert-butyl), 1 -pentyl (//-pentyl). In other embodiments, examples include, but are not limited to, methyl, ethyl, n-propyl, isopropyl, 2 -methyl- 1 -propyl, 2-methyl-2-propyl, 2- m ethyl- 1 -butyl, 3 -methyl- 1 -butyl, 2-m ethyl-3 -butyl, 2,2-dimethyl-l -propyl, 2-methyl-l -pentyl, 3 -methyl- 1 -pentyl, 4-methyl-l -pentyl, 2-methyl-2-pentyl, 3-methyl-2-pentyl, 4-methyl-2-pentyl,
2.2-dimethyl-l -butyl, 3, 3 -dimethyl- 1 -butyl, 2-ethyl-l -butyl, n-butyl, isobutyl, sec-butyl, t-butyl, n-pentyl, isopentyl, neopentyl, tert-amyl, and hexyl, and longer alkyl groups, such as heptyl, octyl, and the like. The alkyl is attached to the rest of the molecule by a single bond. Unless stated otherwise specifically in the specification, an alkyl group is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, sulfone, mercapto, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, the alkyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, -OMe, -NH2, -NO2, or -C=CH. In some embodiments, the alkyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or -OMe. In some embodiments, the alkyl is optionally substituted with halogen such as F. In some embodiments, the alkyl is unsubstituted.
[0033] As used herein, Ci-Cx(or Ci-x) includes C1-C2, C1-C3... Ci-Cx. By way of example only, a group designated as “C1-C4” indicates that there are one to four carbon atoms in the moiety, i.e. groups containing 1 carbon atom, 2 carbon atoms, 3 carbon atoms or 4 carbon atoms. Thus, by way of example only, “C1-C4 alkyl” indicates that there are one to four carbon atoms in the alkyl group, i.e., the alkyl group is selected from among methyl, ethyl, propyl, zso-propyl, //-butyl, iso- butyl, ec-butyl, and t-butyl. Also, by way of example, C0-C2 alkylene includes a direct bond, - CH2-, and -CH2CH2- linkages.
[0034] "Alkoxy" refers to a radical bonded through an oxygen atom of the formula -O-alkyl, where alkyl is an alkyl chain as defined above. Unless stated otherwise specifically in the specification, an alkoxy group can be optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, an alkoxy is optionally substituted with oxo, halogen, -CN, - CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, an alkoxy is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or -OMe. In some embodiments, the alkoxy is optionally substituted with halogen. In some embodiments, the alkoxy is unsubstituted.
[0035] "Alkenyl" refers to an optionally substituted straight or branched hydrocarbon chain radical group containing at least one carbon-carbon double bond, and preferably having from two to twelve carbon atoms (i.e., C2-C12 alkenyl). In certain embodiments, an alkenyl comprises two to eight carbon atoms (z.e., C2-C8 alkenyl). In certain embodiments, an alkenyl comprises four to eight carbon atoms (z.e., C4-C6 alkenyl). In other embodiments, an alkenyl comprises six to eight carbon atoms (z.e., Ce-Cs alkenyl). In certain embodiments, an alkenyl comprises at least one double bond at the end of a carbon chain. In other embodiments, an alkenyl comprises at least one double bond in the middle of a carbon chain. The group can be in either the cis or trans configuration about the double bond(s), and should be understood to include both isomers. Examples include, but are not limited to, ethenyl (-CH=CH2), 1 -propenyl (-CH2CH=CH2), isopropenyl [-C(CH3)=CH2], butenyl, 1,3-butadienyl, and the like. Whenever it appears herein, a numerical range such as “C2-C6 alkenyl” means that the alkenyl group can consist of 2 carbon atoms, 3 carbon atoms, 4 carbon atoms, 5 carbon atoms, or 6 carbon atoms. Unless stated otherwise specifically in the specification, an alkenyl group is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, an alkenyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, an alkenyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or -OMe. In some embodiments, the alkenyl is optionally substituted with halogen. The alkenyl is attached to the rest of the molecule by a single bond, for example, ethenyl (z.e., vinyl), prop-l-enyl (z.e., allyl), but-l-enyl, pent-l-enyl, penta- 1,4-dienyl, and the like. Unless stated otherwise specifically in the specification, an alkenyl group is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, an alkenyl is optionally substituted with oxo, halogen, -CN, - CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, an alkenyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or -OMe. In some embodiments, the alkenyl is optionally substituted with halogen. In some embodiments, the alkenyl is unsubstituted.
[0036] "Alkynyl" refers to an optionally substituted straight or branched hydrocarbon chain radical group containing at least one carbon-carbon triple bond, and preferably having from two to twelve carbon atoms (z.e., C2-C12 alkynyl). In certain embodiments, an alkynyl comprises two to eight carbon atoms (z.e., C2-C8 alkynyl). In other embodiments, an alkynyl comprises two to six carbon atoms (z.e., C2-C6 alkynyl). In other embodiments, an alkynyl comprises two to four carbon atoms (z.e., C2-C4 alkynyl). Whenever it appears herein, a numerical range such as “C2-C6 alkynyl” means that the alkynyl group can consist of 2 carbon atoms, 3 carbon atoms, 4 carbon atoms, 5 carbon atoms, or 6 carbon atoms. The alkynyl is attached to the rest of the molecule by
a single bond, for example, ethynyl, propynyl, butynyl, pentynyl, hexynyl, 2-propynyl, 2-butynyl, 1,3-butadiynyl, and the like. Unless stated otherwise specifically in the specification, an alkynyl group is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, an alkynyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, - OMe, -NH2, or -NO2. In some embodiments, an alkynyl is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or -OMe. In some embodiments, the alkynyl is optionally substituted with halogen. In some embodiments, the alkynyl is unsubstituted.
[0037] "Alkylene" or "alkylene chain" refers to an optionally substituted straight or branched divalent hydrocarbon chain linking the rest of the molecule to a radical group containing no unsaturation, and preferably having from one to twelve carbon atoms, for example, methylene, ethylene, propylene, zz-butylene, and the like. The alkylene chain is attached to the rest of the molecule through a single bond and to the radical group through a single bond. The points of attachment of the alkylene chain to the rest of the molecule and to the radical group can be through any two carbons within the chain. In certain embodiments, an alkylene comprises one to ten carbon atoms (i.e., Ci-Cs alkylene). In certain embodiments, an alkylene comprises one to eight carbon atoms (z.e., Ci-Cs alkylene). In other embodiments, an alkylene comprises one to five carbon atoms (z.e., C1-C5 alkylene). In other embodiments, an alkylene comprises one to four carbon atoms (z.e., C1-C4 alkylene). In other embodiments, an alkylene comprises one to three carbon atoms (z.e., C1-C3 alkylene). In other embodiments, an alkylene comprises one to two carbon atoms (z.e., C1-C2 alkylene). In other embodiments, an alkylene comprises one carbon atom (z.e., Ci alkylene). In other embodiments, an alkylene comprises five to eight carbon atoms (z.e., Cs-Cs alkylene). In other embodiments, an alkylene comprises two to five carbon atoms (z.e., C2-C5 alkylene). In other embodiments, an alkylene comprises three to five carbon atoms (z.e., C3-C5 alkylene). Unless stated otherwise specifically in the specification, an alkylene group can be optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, an alkylene is optionally substituted with oxo, halogen, -CN, -CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, an alkylene is optionally substituted with oxo, halogen, -CN, -CF3, -OH, or - OMe. In some embodiments, the alkylene is optionally substituted with halogen. In some embodiments, the alkylene is -CH2-, -CH2CH2-, or -CH2CH2CH2-. In some embodiments, the alkylene is -CH2-. In some embodiments, the alkylene is -CH2CH2-. In some embodiments, the alkylene is -CH2CH2CH2-. In some embodiments, the alkylene is unsubstituted.
[0038] "Aryl" refers to a radical derived from a hydrocarbon ring system comprising at least one aromatic ring. In some embodiments, an aryl comprises hydrogens and 5 to 30 carbon atoms.
The aryl radical can be a monocyclic, bicyclic, tricyclic, or tetracyclic ring system, which can include fused (when fused with a cycloalkyl or heterocycloalkyl ring, the aryl is bonded through an aromatic ring atom) or bridged ring systems. In some embodiments, the aryl is a 6- to 10- membered aryl. In some embodiments, the aryl is a 6-membered aryl. Aryl radicals include, but are not limited to, aryl radicals derived from the hydrocarbon ring systems of anthrylene, naphthylene, phenanthrylene, anthracene, azulene, benzene, chrysene, fluoranthene, fluorene, indane, indene, naphthalene, phenalene, phenanthrene, pleiadene, pyrene, and triphenylene. In some embodiments, the aryl is phenyl. Unless stated otherwise specifically in the specification, an aryl can be optionally substituted, for example, with halogen, amino, alkylamino, aminoalkyl, nitrile, nitro, hydroxyl, alkyl, alkenyl, alkynyl, haloalkyl, heteroalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, -S(O)2NH-Ci-Cealkyl, and the like. In some embodiments, an aryl is optionally substituted with halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, -NO2, - S(O)2NH2, -S(O)2NHCH3, -S(O)2NHCH2CH3, -S(O)2NHCH(CH3)2, -S(O)2N(CH3)2, or - S(O)2NHC(CH3)3. In some embodiments, an aryl is optionally substituted with halogen, methyl, ethyl, -CN, -CF3, -OH, or -OMe. In some embodiments, the aryl is optionally substituted with halogen. In some embodiments, the aryl is substituted with alkyl, alkenyl, alkynyl, haloalkyl, or heteroalkyl, wherein each alkyl, alkenyl, alkynyl, haloalkyl, heteroalkyl is independently unsubstituted, or substituted with halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, the aryl is unsubstituted.
[0039] "Aralkyl" refers to a radical of the formula -Rc-aryl where Rc is an alkylene chain as defined above, for example, methylene, ethylene, and the like.
[0040] "Aralkenyl" refers to a radical of the formula -Rd-aryl where Rd is an alkenylene chain as defined above. " Aralkynyl" refers to a radical of the formula -Re-aryl, where Re is an alkynylene chain as defined above.
[0041] “Carbocycle” refers to a saturated, unsaturated or aromatic rings in which each atom of the ring is carbon. Carbocycle can include 3- to 10-membered monocyclic rings and 6- to 12- membered bicyclic rings (such as spiro, fused, or bridged rings). Each ring of a bicyclic carbocycle can be selected from saturated, unsaturated, and aromatic rings. An aromatic ring, e.g., phenyl, can be fused to a saturated or unsaturated ring, e.g., cyclohexane, cyclopentane, or cyclohexene. Any combination of saturated, unsaturated and aromatic bicyclic rings, as valence permits, are included in the definition of carbocyclic. In an exemplary embodiment, an aromatic ring, e.g., phenyl, can be fused to a saturated or unsaturated ring, e.g., cyclohexane, cyclopentane, or cyclohexene. A bicyclic carbocycle includes any combination of saturated, unsaturated and aromatic bicyclic rings, as valence permits. A bicyclic carbocycle includes any combination of ring sizes such as 4-5 fused ring systems, 5-5 fused ring systems, 5-6 fused ring systems, 6-6 fused
ring systems, 5-7 fused ring systems, 6-5 fused ring systems, 6-7 fused ring systems, 5-8 fused ring systems, and 6-8 fused ring systems. Exemplary carbocycles include cyclopentyl, cyclohexyl, cyclohexenyl, adamantyl, phenyl, indanyl, and naphthyl. The term “unsaturated carbocycle” refers to carbocycles with at least one degree of unsaturation and excluding aromatic carbocycles. Examples of unsaturated carbocycles include cyclohexadiene, cyclohexene, and cyclopentene. The term “saturated cyclaroalkyl” as used herein refers to a saturated carbocycle. Exemplary carbocycles include cyclopropyl, cyclopentyl, cyclohexyl, cyclohexenyl, adamantyl, phenyl, indanyl, norborane, and naphthyl. Carbocycles can be optionally substituted by one or more substituents such as those substituents described herein.
[0042] "Cycloalkyl" refers to a stable, partially or fully saturated, monocyclic or polycyclic carbocyclic ring, which can include fused (when fused with an aryl or a heteroaryl ring, the cycloalkyl is bonded through a non-aromatic ring atom), bridged, or spiro ring systems. Representative cycloalkyls include, but are not limited to, cycloalkyls having from three to fifteen carbon atoms (C3-C15 cycloalkyl), from three to ten carbon atoms (C3-C10 cycloalkyl), from three to eight carbon atoms (C3-C8 cycloalkyl), from three to six carbon atoms (C3-C6 cycloalkyl), from three to five carbon atoms (C3-C5 cycloalkyl), or three to four carbon atoms (C3-C4 cycloalkyl). In some embodiments, the cycloalkyl is a 3- to 6-membered cycloalkyl. In some embodiments, the cycloalkyl is a 5- to 6-membered cycloalkyl. Monocyclic cycloalkyls include, for example, cyclopropyl, cyclobutyl, cyclopentyl, cyclohexyl, cycloheptyl, and cyclooctyl. Polycyclic cycloalkyls or carbocycles include, for example, adamantyl, norbornyl, decalinyl, bicyclo[3.3.0]octane, bicyclo[4.3.0]nonane, cis-decalin, trans-decalin, bicyclo[2.1.1]hexane, bicyclo[2.2.1]heptane, bicyclo[2.2.2]octane, bicyclo[3.2.2]nonane, and bicyclo[3.3.2]decane, and 7,7-dimethyl-bicyclo[2.2.1]heptanyl. Partially saturated cycloalkyls include, for example, cyclopentenyl, cyclohexenyl, cycloheptenyl, and cyclooctenyl. Unless stated otherwise specifically in the specification, a cycloalkyl is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, alkyl, alkenyl, alkynyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, a cycloalkyl is optionally substituted with oxo, halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, a cycloalkyl is optionally substituted with oxo, halogen, methyl, ethyl, -CN, - CF3, -OH, or -OMe. In some embodiments, the cycloalkyl is optionally substituted with halogen. In some embodiments, the cycloalkyl is unsubstituted.
[0043] "Cycloalkylalkyl" refers to a radical of the formula -Rc-cycloalkyl where Rc is an alkylene chain as described above.
[0044] "Cycloalkylalkoxy" refers to a radical bonded through an oxygen atom of the formula -O-Rc-cycloalkyl where Rc is an alkylene chain as described above.
[0045] " Halo" or "halogen" refers to halogen substituents such as bromo, chloro, fluoro and iodo substituents.
[0046] As used herein, the term "haloalkyl" or “haloalkane” refers to an alkyl radical, as defined above, that is substituted by one or more halogen radicals, for example, trifluoromethyl, di chloromethyl, bromomethyl, 2,2,2-trifluoroethyl, l-fluoromethyl-2-fluoroethyl, and the like. In some embodiments, the alkyl part of the fluoroalkyl radical is optionally further substituted. Examples of halogen substituted alkanes (“haloalkanes”) include halomethane (e.g., chloromethane, bromomethane, fluoromethane, iodomethane), di-and trihalomethane (e.g., tri chloromethane, tribromomethane, trifluoromethane, triiodomethane), 1-haloethane, 2- haloethane, 1,2-dihaloethane, 1-halopropane, 2-halopropane, 3-halopropane, 1,2-dihalopropane, 1,3-dihalopropane, 2,3-dihalopropane, 1,2,3-trihalopropane, and any other suitable combinations of alkanes (or substituted alkanes) and halogens (e.g., Cl, Br, F, I, etc.). When an alkyl group is substituted with more than one halogen radicals, each halogen can be independently selected e.g., 1 -chloro,2-fluoroethane.
[0047] "Fluoroalkyl" refers to an alkyl radical, as defined above, that is substituted by one or more fluoro radicals, for example, trifluoromethyl, difluoromethyl, fluoromethyl, 2,2,2-trifluoroethyl, l-fhioromethyl-2-fluoroethyl, and the like.
[0048] “Hydroxyalkyl” refers to an alkyl radical, as defined above, that is substituted by one or more hydroxyls. In some embodiments, the alkyl is substituted with one hydroxyl. In some embodiments, the alkyl is substituted with one, two, or three hydroxyls. Hydroxyalkyl include, for example, hydroxymethyl, hydroxy ethyl, hydroxypropyl, hydroxybutyl, or hydroxypentyl. In some embodiments, the hydroxyalkyl is hydroxymethyl.
[0049] “Aminoalkyl” refers to an alkyl radical, as defined above, that is substituted by one or more amines. In some embodiments, the alkyl is substituted with one amine. In some embodiments, the alkyl is substituted with one, two, or three amines. Aminoalkyl include, for example, aminomethyl, aminoethyl, aminopropyl, aminobutyl, or aminopentyl. In some embodiments, the aminoalkyl is aminomethyl.
[0050] “Disulfide” refers to two sulfur atoms bonded to each other, where each sulfur comprises an optionally substituted alkyl chain. In some embodiments a disulfide may be R-S-S- R’. In some embodiments, R and R’ may be identical. In some embodiments, R and R’ are different. Each R and R’ may be independently selected from C1-C12 alkyl. In certain embodiments, R or R’ may be substituted with an amine, sulfone, or carboxylic acid. “Ci-Cx alkyl substituted with disulfide” as used herein may refer to a disulfide of the structure R-S-S-R’, where R and R’ may be identical or different. Each R and R’ may be independently selected from Ci-Cy alkyl, such that the length of R and R’ is the length of the Ci-Cx alkyl.
[0051] The term “heteroalkyl” refers to an alkyl group in which one or more skeletal atoms of the alkyl are selected from an atom other than carbon, e.g., oxygen, nitrogen (e.g., -NH-, -N(alkyl)- ), sulfur, or combinations thereof. A heteroalkyl is attached to the rest of the molecule at a carbon atom of the heteroalkyl. In one aspect, a heteroalkyl is a Ci-Ce heteroalkyl wherein the heteroalkyl is comprised of 1 to 6 carbon atoms and one or more atoms other than carbon, e.g., oxygen, nitrogen (e.g. -NH-, -N(alkyl)-), sulfur, or combinations thereof wherein the heteroalkyl is attached to the rest of the molecule at a carbon atom of the heteroalkyl. Examples of such heteroalkyl are, for example, -CH2OCH3, -CH2CH2OCH3, -CH2CH2OCH2CH2OCH3, or - CH(CH3)OCH3. Unless stated otherwise specifically in the specification, a heteroalkyl is optionally substituted for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, alkyl, alkenyl, alkynyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, a heteroalkyl is optionally substituted with oxo, halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, a heteroalkyl is optionally substituted with oxo, halogen, methyl, ethyl, -CN, -CF3, -OH, or -OMe. In some embodiments, the heteroalkyl is optionally substituted with halogen. In some embodiments, the heteroalkyl is unsubstituted.
[0052] “Heterocycloalkyl” refers to a stable 3- to 24-membered partially or fully saturated ring radical comprising 2 to 23 carbon atoms and at least one ring heteroatoms. In some embodiments, a heterocycloalkyl contains from one to 8 heteroatoms selected from the group consisting of nitrogen, oxygen, phosphorous, and sulfur. Unless stated otherwise specifically in the specification, the heterocycloalkyl radical can be a monocyclic, bicyclic, tricyclic, or tetracyclic ring system, which can include fused (when fused with an aryl or a heteroaryl ring, the heterocycloalkyl is bonded through a non-aromatic ring atom) or bridged ring systems; and the nitrogen, carbon, or sulfur atoms in the heterocycloalkyl radical can be optionally oxidized; the nitrogen atom can be optionally quatemized.
[0053] Representative heterocycloalkyls include, but are not limited to, heterocycloalkyls having from two to fifteen carbon atoms (C2-C15 heterocycloalkyl), from two to ten carbon atoms (C2-C10 heterocycloalkyl), from two to eight carbon atoms (C2-C8 heterocycloalkyl), from two to six carbon atoms (C2-C6 heterocycloalkyl), from two to five carbon atoms (C2-C5 heterocycloalkyl), or two to four carbon atoms (C2-C4 heterocycloalkyl). In some embodiments, the heterocycloalkyl is a 3- to 6-membered heterocycloalkyl. In some embodiments, the heterocycloalkyl is a 5- to 6-membered heterocycloalkyl. Examples of such heterocycloalkyl radicals include, but are not limited to, aziridinyl, azetidinyl, dioxolanyl, thienyl[l,3]dithianyl, decahydroisoquinolyl, imidazolinyl, imidazolidinyl, isothiazolidinyl, isoxazolidinyl, morpholinyl, octahydroindolyl, octahydroisoindolyl, 2-oxopiperazinyl, 2-oxopiperidinyl, 2-oxopyrrolidinyl, oxazolidinyl, piperidinyl, piperazinyl, 4-piperidonyl, pyrrolidinyl,
pyrazolidinyl, quinuclidinyl, thiazolidinyl, tetrahydrofuryl, trithianyl, tetrahydropyranyl, thiomorpholinyl, thiamorpholinyl, 1-oxo-thiomorpholinyl, 1,1-dioxo-thiomorpholinyl, 1,3- dihydroisobenzofuran- 1 -yl, 3 -oxo- 1 ,3 -dihydroisobenzofuran- 1 -yl, methyl-2-oxo- 1 ,3 -dioxol-4-yl, and 2-oxo- 1,3 -di oxol -4-yl. The term heterocycloalkyl also includes all ring forms of the carbohydrates, including but not limited to, the monosaccharides, the disaccharides, and the oligosaccharides. It is understood that when referring to the number of carbon atoms in a heterocycloalkyl, the number of carbon atoms in the heterocycloalkyl is not the same as the total number of atoms (including the heteroatoms) that make up the heterocycloalkyl (i.e. skeletal atoms of the heterocycloalkyl ring). Unless stated otherwise specifically in the specification, a heterocycloalkyl is optionally substituted, for example, with oxo, halogen, amino, nitrile, nitro, hydroxyl, alkyl, alkenyl, alkynyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, a heterocycloalkyl is optionally substituted with oxo, halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, a heterocycloalkyl is optionally substituted with oxo, halogen, methyl, ethyl, -CN, -CF3, -OH, or -OMe. In some embodiments, the heterocycloalkyl is optionally substituted with halogen. In some embodiments, the heterocycloalkyl is ub substituted.
[0054] “Heterocycle” or “heterocyclyl” refers to a saturated, unsaturated or aromatic ring comprising one or more ring heteroatoms. Exemplary heteroatoms include N, O, Si, P, B, and S atoms. Heterocycles include e.g., 3- to 10-membered monocyclic rings and 6- to 12-membered bicyclic rings (such as spiro, fused, or bridged rings). Unless stated otherwise specifically in the specification, the heterocyclyl radical is a monocyclic, bicyclic, tricyclic or tetracyclic ring system, which optionally includes fused, bridged, or spirocyclic ring systems. The heteroatoms in the heterocyclyl radical are optionally oxidized. One or more nitrogen atoms, if present, are optionally quaternized. The heterocyclyl radical can be partially or fully saturated. The heterocyclyl is attached to the rest of the molecule through any atom of the ring(s). Examples of such heterocyclyl radicals include, but are not limited to, dioxolanyl, thienyl[l,3]dithianyl, decahydroisoquinolyl, imidazolinyl, imidazolidinyl, isothiazolidinyl, isoxazolidinyl, morpholinyl, octahydroindolyl, octahydroisoindolyl, 2-oxopiperazinyl, 2-oxopiperidinyl, 2-oxopyrrolidinyl, oxazolidinyl, piperidinyl, piperazinyl, 4-piperidonyl, pyrrolidinyl, pyrazolidinyl, quinuclidinyl, thiazolidinyl, tetrahydrofuryl, trithianyl, tetrahydropyranyl, thiomorpholinyl, thiamorpholinyl, 1-oxo-thiomorpholinyl, and 1,1-dioxo-thiomorpholinyl. Unless stated otherwise specifically in the specification, the term "heterocyclyl" is meant to include heterocyclyl radicals as defined above that are optionally substituted by one or more substituents. For example, a heterocyclyl can be optionally substituted by one or more substituents selected from alkyl, alkenyl, alkynyl, halo, fluoroalkyl, oxo, thioxo, cyano, nitro, optionally
substituted aryl, optionally substituted aralkyl, optionally substituted aralkenyl, optionally substituted aralkynyl, optionally substituted carbocyclyl, optionally substituted carbocyclylalkyl, optionally substituted heterocyclyl, optionally substituted heterocyclylalkyl, optionally substituted heteroaryl, optionally substituted heteroarylalkyl, -Rb-ORa, -Rb-OC(O)-Ra, -Rb- OC(O)-ORa, -Rb-OC(O)-N(Ra)2, -Rb-N(Ra)2, -Rb-C(O)Ra, -Rb-C(O)ORa, -Rb-C(O)N(Ra)2, -Rb- CN, -Rb-O-Re-C(O)N(Ra)2, -Rb-N(Ra)C(O)ORa, -Rb-N(Ra)C(O)Ra, -Rb-N(Ra)S(O)tRa (where t is 1 or 2), -Rb-S(O)tRa (where t is 1 or 2), -Rb-S(O)tORa (where t is 1 or 2) and -Rb-S(O)tN(Ra)2 (where t is 1 or 2), where each Ra is independently hydrogen, alkyl (optionally substituted with halogen, hydroxy, methoxy, or trifluoromethyl), fluoroalkyl, cycloalkyl (optionally substituted with halogen, hydroxy, methoxy, or trifluoromethyl), cycloalkylalkyl (optionally substituted with halogen, hydroxy, methoxy, or trifluoromethyl), aryl (optionally substituted with halogen, hydroxy, methoxy, or trifluoromethyl), aralkyl (optionally substituted with halogen, hydroxy, methoxy, or trifluoromethyl), heterocyclyl (optionally substituted with halogen, hydroxy, methoxy, or trifluoromethyl), heterocyclylalkyl (optionally substituted with halogen, hydroxy, methoxy, or trifluoromethyl), heteroaryl (optionally substituted with halogen, hydroxy, methoxy, or trifluoromethyl), or heteroarylalkyl (optionally substituted with halogen, hydroxy, methoxy, or trifluoromethyl), each Rb is independently a direct bond or a straight or branched alkylene or alkenylene chain, and Re is a straight or branched alkylene or alkenylene chain, and where each of the above substituents is unsubstituted unless otherwise indicated.
[0055] “Heteroaryl” or “aromatic heterocycle” refers to a ring system radical comprising carbon atom(s) and one or more ring heteroatoms (e.g., selected from the group consisting of nitrogen, oxygen, phosphorous, silicon, and sulfur), and at least one aromatic ring. In some embodiments, a heteroaryl is a 5- to 14-membered ring system radical comprising one to thirteen carbon atoms, one to six heteroatoms selected from the group consisting of nitrogen, oxygen, phosphorous, and sulfur. The heteroaryl radical can be a monocyclic, bicyclic, tricyclic, or tetracyclic ring system, which can include fused (when fused with a cycloalkyl or heterocycloalkyl ring, the heteroaryl is bonded through an aromatic ring atom) or bridged ring systems; and the nitrogen, carbon, or sulfur atoms in the heteroaryl radical can be optionally oxidized; the nitrogen atom can be optionally quaternized. In some embodiments, the heteroaryl is a 5- to 10-membered heteroaryl. In some embodiments, the heteroaryl is a 5- to 6-membered heteroaryl. Examples include, but are not limited to, azepinyl, acridinyl, benzimidazolyl, benzothiazolyl, benzindolyl, benzodi oxolyl, benzofuranyl, benzooxazolyl, benzothiazolyl, benzothiadiazolyl, benzo[b][l,4]dioxepinyl, 1,4-benzodioxanyl, benzonaphthofuranyl, benzoxazolyl, benzodi oxolyl, benzodioxinyl, benzopyranyl, benzopyranonyl, benzofuranyl, benzofuranonyl, benzothienyl (benzothiophenyl), benzotri azolyl, benzo[4,6]imidazo[l,2-a]pyridinyl, carbazolyl, cinnolinyl,
dibenzofuranyl, dibenzothiophenyl, furanyl, furanonyl, isothiazolyl, imidazolyl, indazolyl, indolyl, indazolyl, isoindolyl, indolinyl, isoindolinyl, isoquinolyl, indolizinyl, isoxazolyl, naphthyridinyl, oxadiazolyl, 2-oxoazepinyl, oxazolyl, oxiranyl, 1-oxidopyridinyl, 1-oxidopyrimidinyl, 1-oxidopyrazinyl, 1-oxidopyridazinyl, 1 -phenyl -IH-pyrrolyl, phenazinyl, phenothiazinyl, phenoxazinyl, phthalazinyl, pteridinyl, purinyl, pyrrolyl, pyrazolyl, pyridinyl, pyrazinyl, pyrimidinyl, pyridazinyl, quinazolinyl, quinoxalinyl, quinolinyl, quinuclidinyl, isoquinolinyl, tetrahydroquinolinyl, thiazolyl, thiadiazolyl, triazolyl, tetrazolyl, triazinyl, and thiophenyl (i.e., thienyl). Unless stated otherwise specifically in the specification, a heteroaryl is optionally substituted, for example, with halogen, amino, nitrile, nitro, hydroxyl, alkyl, alkenyl, alkynyl, haloalkyl, alkoxy, aryl, cycloalkyl, heterocycloalkyl, heteroaryl, and the like. In some embodiments, a heteroaryl is optionally substituted with halogen, methyl, ethyl, -CN, -CF3, -OH, -OMe, -NH2, or -NO2. In some embodiments, a heteroaryl is optionally substituted with halogen, methyl, ethyl, -CN, -CF3, -OH, or -OMe. In some embodiments, the heteroaryl is optionally substituted with halogen. In some embodiments, the heteroaryl is unsubstituted.
[0056] The term “substituted” refers to moieties having substituents replacing a hydrogen on one or more carbons or substitutable heteroatoms, e.g., NH, of the structure. It will be understood that “substitution” or “substituted with” includes the implicit proviso that such substitution is in accordance with permitted valence of the substituted atom and the substituent, and that the substitution results in a stable compound, i.e., a compound which does not spontaneously undergo transformation such as by rearrangement, cyclization, elimination, etc. In certain embodiments, substituted refers to moieties having substituents replacing two hydrogen atoms on the same carbon atom, such as substituting the two hydrogen atoms on a single carbon with an oxo, imino or thioxo group. As used herein, the term “substituted” is contemplated to include all permissible substituents of organic compounds. In a broad aspect, the permissible substituents include acyclic and cyclic, branched and unbranched, carbocyclic and heterocyclic, aromatic and non-aromatic substituents of organic compounds. The permissible substituents can be one or more and the same or different for appropriate organic compounds. For purposes of this disclosure, the heteroatoms such as nitrogen can have hydrogen substituents and/or any permissible substituents of organic compounds described herein which satisfy the valences of the heteroatoms.
[0057] In some embodiments, substituents can include any substituents described herein, for example: halogen, hydroxy, oxo (=0), thioxo (=S), cyano (-CN), nitro (-NO2), imino (=N-H), oximo (=N-0H), hydrazine (=N-
NH2), -Rb-0Ra, -Rb-OC(O)-Ra, -Rb-OC(O)-ORa, -Rb-OC(O)-N(Ra)2, -Rb-N(Ra)2, -Rb-C(O)Ra, -R b-C(O)ORa, -Rb-C(O)N(Ra)2, -Rb-O-Rc-C(O)N(Ra)2, -Rb-N(Ra)C(O)ORa, -Rb-N(Ra)C(O)Ra, -Rb- N(Ra)S(O)tRa (where t is 1 or 2), -Rb-S(O)tRa (where t is 1 or 2), -Rb-S(O)tORa (where t is 1 or
2), and -Rb-S(O)tN(Ra)2 (where t is 1 or 2); and alkyl, alkenyl, alkynyl, aryl, aralkyl, aralkenyl, aralkynyl, cycloalkyl, cycloalkylalkyl, and heterocycle, any of which can be optionally substituted by alkyl, alkenyl, alkynyl, halogen, haloalkyl, haloalkenyl, haloalkynyl, oxo (=0), thioxo (=S), cyano (-CN), nitro (-NO2), imino (=N-H), oximo (=N-0H), hydrazine (=N-NH2), SF5, -Rb-0Ra, -Rb-OC(O)-Ra, -Rb-OC(O)-ORa, -Rb-0C(0)-N(Ra)2, -Rb-N(Ra)2, -Rb-C(O)Ra, -Rb- C(O)ORa, -Rb-C(0)N(Ra)2, -Rb-0-Rc-C(0)N(Ra)2, -Rb-N(Ra)C(0)0Ra, -Rb-N(Ra)C(0)Ra, -Rb-N (Ra)S(O)tRa (where t is 1 or 2), -Rb-S(O)tRa (where t is 1 or 2), -Rb-S(O)tORa (where t is 1 or 2) and -Rb-S(O)tN(Ra)2 (where t is 1 or 2); wherein each Ra is independently selected from hydrogen, alkyl, cycloalkyl, cycloalkylalkyl, aryl, aralkyl, and heterocycle, wherein each Ra, valence permitting, can be optionally substituted with alkyl, alkenyl, alkynyl, halogen, haloalkyl, haloalkenyl, haloalkynyl, oxo (=0), thioxo (=S), cyano (-CN), nitro (-NO2), imino (=N-H), oximo (=N-0H), hydrazine (=N-
NH2), -Rb-0Ra, -Rb-0C(0)-Ra, -Rb-0C(0)-0Ra, -Rb-0C(0)-N(Ra)2, -Rb-N(Ra)2, -Rb-C(0)Ra, -R b-C(0)0Ra, -Rb-C(0)N(Ra)2, -Rb-0-Rc-C(0)N(Ra)2, -Rb-N(Ra)C(0)0Ra, -Rb-N(Ra)C(0)Ra, -Rb- N(Ra)S(O)tRa (where t is 1 or 2), -Rb-S(O)tRa (where t is 1 or 2), -Rb-S(O)tORa (where t is 1 or 2) and -Rb-S(O)tN(Ra)2 (where t is 1 or 2); and wherein each Rb is independently selected from a direct bond or a straight or branched alkylene, alkenylene, or alkynylene chain, and each Rc is a straight or branched alkylene, alkenylene or alkynylene chain.
[0058] The term “optional” or “optionally” means that the subsequently described event or circumstance may or may not occur, and that the description includes instances where said event or circumstance occurs and instances in which it does not. For example, “optionally substituted alkyl” means either “alkyl” or “substituted alkyl” as defined above. Further, an optionally substituted group can be un- substituted (e.g., -CH2CH3), fully substituted (e.g., -CF2CF3), monosubstituted (e.g., -CH2CH2F) or substituted at a level anywhere in-between fully substituted and mono-substituted (e.g, -CH2CHF2, -CH2CF3, -CF2CH3, -CFHCHF2, etc ).
[0059] The term “biomolecule” refers to biological components that may be involved in corona formation, including, but not limited to, for example, proteins, polypeptides, polysaccharides, a sugar, a lipid, a lipoprotein, a metabolite, an oligonucleotide, metabolome or combination thereof. It is contemplated that the biomolecule coronas of distinct particles may contain some of the same biomolecules, may contain distinct biomolecules with regard to the other sensor elements, and/or may differ in level or quantity, type or conformation of the biomolecule that binds to each sensor element. In one embodiment, the biomolecule is selected from the group of proteins, nucleic acids, lipids, and metabolomes.
[0060] Ranges provided herein are understood to be shorthand for all of the values within the range. For example, a range of 1 to 50 is understood to include any number, combination of
numbers, or sub-range from the group consisting of 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, or 50, as well as all intervening decimal values between the aforementioned integers such as, for example, 1.1, 1.2, 1.3, 1.4, 1.5, 1.6, 1.7, 1.8, and 1.9. With respect to sub-ranges, “nested sub-ranges” that extend from either end point of the range are specifically contemplated. For example, a nested sub-range of an exemplary range of 1 to 50 can comprise 1 to 10, 1 to 20, 1 to 30, and 1 to 40 in one direction, or 50 to 40, 50 to 30, 50 to 20, and 50 to 10 in the other direction.
[0061] The compounds and structures provided herein may be stereoisomeric. In some cases, a compound or structure of the disclosure may form a stereoisomer. In some cases, the stereoisomer may be a diastereomer (e.g., a cis/trans isomer, E/Z isomer, conformer, or rotamer). In some cases, the stereoisomer may be an enantiomer (R,S enantiomers or +/- enantiomers). In some cases, the compound or structure of the disclosure may be enantiopure (e.g., 100% pure). In some cases, the compound or structure may form a racemic mixture of enantiomers (e.g., 50% pure). In some cases, a compound or structure of the disclosure may stabilize as a stereoisomer, where the compound or structure of the disclosure comprises at least about 10%, about 15%, about 20%, about 25%, about 30%, about 35%, about 40%, about 45%, about 50%, about 55%, about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 98.5%, about 99%, about 99.1%, about 99.2%, about 99.3%, about 99.4%, about 99.5%, about 99.6%, about 99.7%, about 99.8%, about 99.9%, or more of a mixture of the compound or structure and the corresponding stereoisomer.
Methods of Isolating Biomolecules
[0062] Provided herein are methods of isolating one or more biomolecules, such as a protein, a peptide, or a polypeptide, from a (complex) biological sample. In some instances, the methods provided herein comprise utilizing macromolecule structures which may (e.g., selectively) bind to the one or more biomolecules. The binding of the one or more biomolecules to the macromolecule structures may be reversible, allowing for pull-down of the macromolecule structures (e.g., via magnetic pull-down or via centrifugation), followed by elution of the biomolecules to provide the isolated (e.g., or purified) biomolecules. The methods provided herein allow for clean-up of complex biological solutions, such as removal of surfactants, detergents, chaotropes, salts, and other impurities or contaminants without the need for other methods such as solid phase chromatography (e.g., reversed-phase/ion exchange) or suspension trapping, which may, among other potential disadvantages, be solvent intensive or can result in reduced yield of the desired biomolecules. In some embodiments, the methods provided herein allow for the selective removal of mass spectrometry (MS) incompatible components from a biological sample.
[0063] The methods provided herein allow for isolation of biomolecules (e.g., proteins or peptides) with, for example, no solid residues or contaminants, high biomolecule (e.g., protein or peptide) recovery, no bias in biomolecule properties, and the ability to isolate large quantities of similar and unique biomolecules. The methods provided herein also, in some embodiments, provide macromolecule structures with fast pull-down times, easy re-suspension, and are filter plate compatible, which in some instances allow for automation of the methods herein. In some embodiments, the methods disclosed herein may be faster than existing methods for removing impurities or contaminants. In some embodiments, the methods disclosed herein are advantageous because they do not require a positive pressure source. In some embodiments, the methods disclosed herein provide samples that are compatible with liquid chromatography mass spectrometry analysis.
[0064] As an example, FIG. 6a shows a schematic of an embodiment of a method of isolating biomolecules (e.g., proteins or peptides) while FIG. 6b shows a schematic of an alternate embodiment of a method of isolating biomolecules (e.g., proteins or peptides). In some embodiments, the methods herein comprise contacting a population of biomolecules (e.g., proteins or peptides) in a biological sample with a macromolecule in the presence of an organic solvent. In some embodiments, the methods comprise eluting proteins from the macromolecule structure to provide isolated biomolecules (e.g., proteins or peptides). In some embodiments, the methods provided herein comprise contacting a biological sample (e.g., comprising proteins or peptides) with a first macromolecule structure to provide a biomolecule (e.g., protein) corona. In some embodiments, the methods provided herein comprise digesting the proteins or peptides of the biomolecule (e.g., protein) corona to form one or more peptides. In some embodiments, the methods provided herein comprise separating peptides or proteins of the biomolecule (e.g., protein) corona from the first macromolecule structure. In some embodiments, the methods herein comprise contacting peptides or proteins with a second macromolecule structure and an organic solvent to bind the peptides or proteins to the second macromolecule structure. In some embodiments, the methods provided herein comprise eluting peptides or proteins from the second macromolecule structure to provide isolated biomolecules (e.g., peptides or proteins).
[0065] In some instances, the methods provided herein comprise contacting one or more biomolecules from a biological sample with a macromolecule structure. In some embodiments, the contacting comprises binding of the biomolecule with a macromolecule structure. In some embodiments, the methods comprise washing, as to remove any species not contacted to (e.g., or bound to) the macromolecule structures. This may result in, when reconstituted, a solution comprising a macromolecule structure comprising one or more biomolecules. In some
embodiments, the methods comprise eluting the biomolecule from the macromolecule structure (e.g., thereby providing the isolated biomolecule).
[0066] Provided herein are methods of isolating one or more biomolecules from a biological sample. In some embodiments, isolating one or more biomolecules comprises purifying one or more biomolecules.
[0067] Provided herein, in some embodiments, are methods of preparing a biological sample for analysis, such as analysis as provided elsewhere herein (e.g., mass spectrometry). In some embodiments, preparing a biological sample for mass spectrometry includes removing one or more mass spectrometry incompatible components from the biological sample.
[0068] In some embodiments, isolating or purifying one or more biomolecules from a biological sample, comprises removing at least 90% (e.g., at least 95%, at least 97.5%, at least 98%, at least 99%, at least 99.5%) of any other species from the biological sample. In some embodiments, isolating or purifying one or more biomolecules from a biological sample, comprises removing about 90% to about 99.9% (e.g., 90% to about 99%, 95% to about 99%, 95% to about 98%) of any other species from the biological sample. In some embodiments, isolating or purifying one or more biomolecules from a biological sample comprises processing the biomolecules to a level suitable for further analysis or use, such as by mass spectrometry. In some embodiments, the other species is a chemical reagent used for preparing a biological sample for mass spectrometry. In some embodiments, the other species is a surfactant. In some embodiments, the other species is a lysis reagent. In some embodiments, the other species is a reducing agent. In some embodiments, the other species is an alkylating agent. In some embodiments, the other species is a buffering agent. In some embodiments, the other species is a chaotrope. In some embodiments, the other species is an organic solvent.
[0069] In some embodiments, isolating or purifying one or more biomolecules from a biological sample, comprises removing at least 90% (e.g., at least 95%, at least 97.5%, at least 98%, at least 99%, at least 99.5%) of a surfactant. In some embodiments, the surfactant is a synthetic compound. In some embodiments, the surfactant is ionic. In some embodiments, the surfactant is zwitterionic. Non-limiting examples of surfactants that may be removed include sodium lauryl sulfate, Triton X-100, TWEEN, NP-40, CHAPS, Octyl glucoside, deoxy cholate, RAPIGEST, and decyl maltoside. As an example, the methods disclosed herein may remove at least 90% of sodium lauryl sulfate.
[0070] In some embodiments, isolating or purifying one or more biomolecules from a biological sample comprises removing at least 90% (e.g., at least 95%, at least 97.5%, at least 98%, at least 99%, at least 99.5%) of a buffering agent. In some embodiments, the buffering agent is a Good’s buffering agent. Non-limiting examples of buffering agents that may be removed
include Tris, phosphate buffer, Tris, HEPES, MES, MOPS, TES, CAPS, Bicine, and Bis-Tris. For example, at least 90% of CAPS may be removed using the methods disclosed herein. As another example, at least 90% of HEPES may be removed using the methods disclosed herein.
[0071] In some embodiments, isolating or purifying one or more biomolecules from a biological sample, comprises removing at least 90% (e.g., at least 95%, at least 97.5%, at least 98%, at least 99%, at least 99.5%) of a chaotrope. Non-limiting examples of chaotropes that may removed include urea, guanidine hydrochloride, sodium thiocyanate, and perchlorate salts. As an example, at least 90% of urea may be removed using the methods disclosed herein.
[0072] In some embodiments, an amount of surfactants in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of surfactants in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides). In some embodiments, the amount of surfactants in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of surfactants in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 10 wt% (e.g., 20 wt%, 30 wt%, 50 wt%, 80 wt%, 90 wt%, 100 wt%). In some embodiments, the amount of surfactants in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of surfactants in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 70 wt%. In some embodiments, the amount of surfactants in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of surfactants in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at most 100 wt% (e.g., 99 wt%, 95 wt%, 90 wt%, 80 wt%, 70 wt%, 60 wt%). In some embodiments, the amount of surfactants in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of surfactants in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by about 10 wt% to about 100 wt%, 10 wt% to about 90 wt%, 20 wt% to about 100 wt%, 30 wt% to about 100 wt%, 50 wt% to about 100 wt%, 50 wt% to about 80 wt%, or about 80 wt% to about 100 wt%. Non-limiting examples of surfactants that may be greater include sodium lauryl sulfate, Triton X-100, TWEEN, NP-40, CHAPS, Octyl glucoside, and decyl maltoside. In some embodiments, the surfactant is sodium lauryl sulfate. In some embodiments, the surfactant is CHAPS. In some embodiments, the surfactant is a synthetic molecule.
[0073] In some embodiments, the methods provided herein comprise removing at least 90 wt% (e.g., at least 95 wt%, at least 97.5 wt%, at least 98 wt%, at least 99 wt%, at least 99.5 wt%) of an acid or base from the population of biomolecules.
[0074] In some embodiments, the methods provided herein comprise removing at least at least 90 wt% (e.g., at least 9 wt5%, at least 97.5 wt%, at least 98 wt%, at least 99 wt%, at least 99.5 wt%) of a surfactant from the population of biomolecules.
[0075] In some embodiments, the methods provided herein comprise removing at least 90 wt% (e.g., at least 95 wt%, at least 97.5 wt%, at least 98 wt%, at least 99 wt%, at least 99.5 wt%) of a lysis reagent from the population of biomolecules.
[0076] In some embodiments, an amount of a buffering agent in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of buffering agent in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides). In some embodiments, the amount of buffering agent in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of buffering agent in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 10 wt% (e.g., 20 wt%, 30 wt%, 50 wt%, 80 wt%, 90 wt%, 100 wt%). In some embodiments, the amount of buffering agent in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of buffering agent in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 70 wt%. In some embodiments, the amount of buffering agent in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of buffering agent in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at most 100 wt% (e.g., 99 wt%, 95 wt%, 90 wt%, 80 wt%, 70 wt%, 60 wt%). In some embodiments, the amount of buffering agent in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of buffering agent in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by about 10 wt% to about 100 wt%, 10 wt% to about 90 wt%, 20 wt% to about 100 wt%, 30 wt% to about 100 wt%, 50 wt% to about 100 wt%, 50 wt% to about 80 wt%, or about 80 wt% to about 100 wt%. Non-limiting examples of buffering agents that may be greater include Tris, phosphate buffer, Tris, HEPES, MES, MOPS, TES, CAPS, Bicine, and Bis-Tris. In some embodiments, the buffering agent is CAPS. In some embodiments, the buffering agent is HEPES.
[0077] In some embodiments, an amount of chaotropes in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of chaotropes in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides). In some embodiments, the amount of chaotropes in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of chaotropes in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 10 wt% (e.g., 20 wt%, 30 wt%, 50 wt%, 80 wt%, 90 wt%, 100 wt%). In some embodiments, the amount
of chaotropes in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of chaotropes in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at least 70 wt%. In some embodiments, the amount of chaotropes in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of chaotropes in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by at most 100 wt% (e.g., 99 wt%, 95 wt%, 90 wt%, 80 wt%, 70 wt%, 60 wt%). In some embodiments, the amount of chaotropes in a composition comprising the population of biomolecules (e.g., proteins or peptides) is greater than the amount of chaotropes in a composition comprising the one or more isolated biomolecules (e.g., proteins or peptides) by about 10 wt% to about 100 wt%, 10 wt% to about 90 wt%, 20 wt% to about 100 wt%, 30 wt% to about 100 wt%, 50 wt% to about 100 wt%, 50 wt% to about 80 wt%, or about 80 wt% to about 100 wt%. Non-limiting examples of chaotropes that may be greater include urea, guanidine hydrochloride, sodium thiocyanate, and perchlorate salts. In some embodiments, the chaotrope is urea.
[0078] In some embodiments, the methods provided herein comprise contacting a population of biomolecules in a biological sample with a macromolecule structure. In some embodiments, contacting the population of biomolecules in a biological sample with a macromolecule structure occurs in the presence of an organic solvent, such as to bind one or more biomolecules (e.g., proteins or peptides) to the macromolecule structure. In some instances, the organic solvent provides the necessary chemical environment for binding of the one or more biomolecules (e.g., proteins peptides) to the macromolecule structure. In some instances, in the absence of the organic solvent, the one or more biomolecules (e.g., proteins or peptides) may not bind to the macromolecule structure.
[0079] In some embodiments, contacting as provided herein comprises incubating. In some embodiments, the methods herein comprise incubating the one or more biomolecules with a macromolecule structure. In some embodiments, incubating provides for contacting (e.g., binding) of the biomolecule with the macromolecule structure. In some embodiments, incubation occurs for any suitable period of time according to one of skill in the art. In some embodiments, incubation occurs for at least 10 seconds (e.g., at least 20 seconds, at least 30 seconds, at least 1 minute, at least 2 minutes, at least 4 minutes, at least 5 minutes, at least 8 minutes, at least 10 minutes). In some embodiments, incubation occurs for at most 1 hour (e.g., at most 45 minutes, at most 30 minutes, at most 20 minutes, at most 10 minutes, at most 8 minutes, at most 6 minutes, at most 5 minutes, at most 4 minutes, at most 2 minutes). In some embodiments, incubation occurs for about 10 seconds to about 1 hours, about 10 seconds to about 30 minutes, about 1 minute to about 20 minutes, about 1 minute to about 10 minutes, or about 1 minute to about 5 minutes. In
some embodiments, incubation occurs for about 8 minutes. In some embodiments, incubation occurs at the same time as separation, such as separation as described elsewhere herein. The temperature of incubation is not particularly limited, and may be performed, for example, at about room temperature.
[0080] In any of the methods provided herein, biomolecules may comprise proteins, polypeptides, polysaccharides, sugars, lipids, lipoproteins, metabolites, oligonucleotides, metabolomes, or any combination thereof. In some embodiments, the biomolecules comprise proteins. In some embodiments, the biomolecules comprise peptides (e.g., polypeptides). In some embodiments, proteins provided herein comprise peptides. Unless otherwise stated in the present application, any reference to “protein” or “proteins” encompasses peptides obtained by fragmenting a protein, such as by trypsin digestion.
[0081] In some embodiments, the methods provided herein comprise preparing the macromolecule structures (provided herein). In some embodiments, preparing the macromolecule structures comprises washing the macromolecule structures with water, buffer, or a combination thereof. In some embodiments, preparing the macromolecule structures comprises suspending the macromolecule structures at a concentration of at least 5 mg/mL (e.g., at least 10 mg/mL, 20 mg/mL, 30 mg/mL, 40 mg/mL, 50 mg/mL, 60 mg/mL, 70 mg/mL, or 80 mg/mL). In some embodiments, preparing the macromolecule structures comprises suspending the macromolecule structures at a concentration of at most 100 mg/mL (e.g., 90 mg/mL, 80 mg/mL, 70 mg/mL, 60 mg/mL, 50 mg/mL, 40 mg/mL, 30 mg/mL, 20 mg/mL, or 10 mg/mL). In some embodiments, preparing the macromolecule structures comprises suspending the macromolecule structures at a concentration of from about 5 mg/mL to about 100 mg/mL, 5 mg/mL to about 80 mg/mL, 20 mg/mL, to about 50 mg/mL, or about 30 mg/mL to about 50 mg/mL).
[0082] In some embodiments, the methods provided herein further comprise eluting the one or more biomolecules (e.g., proteins or peptides) from the macromolecule structure. In some embodiments, the methods provided herein comprise eluting the one or more proteins. In some embodiments, eluting the one or more biomolecules (e.g., proteins) from the macromolecule structure provides one or more isolated biomolecules (e.g., proteins or peptides). In some embodiments, the methods provided herein comprise eluting the one or more proteins from the macromolecule structure to provide one or more isolated biomolecules (e.g., proteins or peptides). [0083] In some embodiments, eluting as provided herein comprises elution with an aqueous solution. In some embodiments, the aqueous solution is configured to elute proteins from a macromolecule structure. In some embodiments, the aqueous solution comprises an organic solvent. In some embodiments, the aqueous solution comprises no more than 50 wt% (e.g., 40 wt%, 30 wt%, 20 wt%, 10 wt%, 5 wt%) of an organic solvent, such as an organic solvent as
described elsewhere herein. In some embodiments, the aqueous solution comprises no organic solvent or only trace amounts of organic solvent. In some embodiments, the aqueous solution comprises at least 1 wt% (e.g., at least 5 wt%, at least 10 wt%, at least 20 wt%, at least 30 wt%) of an organic solvent, such as an organic solvent as described elsewhere herein. In some embodiments, wt% refers to weight per weight (e.g., w/w%). In some embodiments, wt% refers to weight per volume (e.g., w/v%). In some embodiments, the aqueous solution comprises an organic solvent. In some embodiments, the aqueous solution comprises no more than 50 v/v% (e.g., 40 v/v%, 30 v/v%, 20 v/v%, 10 v/v%, 5 v/v%) of an organic solvent, such as an organic solvent as described elsewhere herein. In some embodiments, the aqueous solution comprises no organic solvent or only trace amounts of organic solvent. In some embodiments, the aqueous solution comprises at least 1 v/v% (e.g., at least 5 v/v%, at least 10 v/v%, at least 20 v/v%, at least 30 v/v%) of an organic solvent, such as an organic solvent as described elsewhere herein. In some embodiments, the aqueous solution is pure water. In some instances, elution is not possible in the presence of an (e.g., 100%) organic solvent, such that the biomolecule (e.g., protein or peptide) will not elute from the macromolecule structure. In some embodiments, the aqueous solution comprises a solution of acetonitrile and water. In certain embodiments, the aqueous solution comprises a solution of 10% or less acetonitrile by volume in water. In certain embodiments, the aqueous solution comprises a solution of 5% or less acetonitrile by volume in water. In certain embodiments, the aqueous solution comprises a solution of about 3% acetonitrile by volume in water.
[0084] In some embodiments, eluting the one or more proteins comprises eluting the one or more peptides. In some embodiments, the one or more isolated biomolecules comprises one or more isolated proteins. In some embodiments, the one or more isolated biomolecules comprises one or more isolated peptides.
[0085] The elution may, in some embodiments, include using two or more eluants successfully. In some embodiments, the eluants have different properties such that different fractions of biomolecules (e.g. proteins) are eluted. As a non-limiting example, a series of eluants comprising different mixtures of water and acetonitrile may be successively applied to the macromolecule structure to remove fractions of biomolecules (e.g., proteins). When the relative amount of water increases with each eluant, these fractions may include more hydrophilic biomolecules. The skilled person, guided by the teaching of the present application, can select other suitable eluants to fractionate the biomolecules.
[0086] In some embodiments, eluting as described herein comprise the successive application (e.g., to the macromolecule structures) of two or more eluents. In some embodiments, the two or more eluents are different. In some embodiments, the two or more eluents are the same. In some
embodiments, the combination of two or more successively applied different eluents allows for elution of the one or more peptides from the macromolecule structure(s). In some embodiments, the two or more eluents comprise eluents of varying (e.g., increasing) polarity, such as by increasing the amount of water in the eluent, or such as by increasing the ratio of water to organic solvent. In some embodiments, the two or more eluants may be separately analyzed. For example, the two or more separately may be separately analyzed by LC-MS/MS, which in some cases may increase the number of biomolecules identified/quantified in the sample.
[0087] In some embodiments, the methods provided herein further comprise separating the macromolecule structure from the one or more biomolecules (e.g., proteins or peptides) (e.g., such as after (a)). In some embodiments, separating the macromolecule structure form the one or more biomolecules (e.g., proteins or peptides) occurs after contacting a population of biomolecules in a biological sample with a macromolecule structure (e.g., after binding of the one or more biomolecules (e.g., proteins or peptides) with the macromolecule structure).
[0088] In some embodiments, the method further comprises separating the macromolecule structure from the population of biomolecules in a biological sample that are not contacted (e.g., bound) to the macromolecule structure (e.g., such as after (a)). In some instances, this separating provides the macromolecule structure comprising the one or more biomolecules (e.g., proteins or peptides) in the absence of other species from the biological sample.
[0089] In any of the methods provided herein, separating may comprise magnetic separation, centrifugation, or a combination thereof. In some embodiments, separating comprises magnetic separation. In some embodiments, separating comprises centrifugation. In some embodiments, separating comprises natural separation (e.g., such as by gravity). Separating may be completed for any duration according to one of skill in the art in order to separate the macromolecule structures from the supernatant. In some embodiments, centrifugation occurs at any speed according to one of skill in the art, such as necessary to separate ethe macromolecule structures from the supernatant. In some embodiments, centrifugation occurs at a speed of at least 10 RPM (e.g., at least 20 RPM, at least 50 RPM, at least 100 RPM, at least 200 RPM, at least 400 RPM, at least 600 RPM, at least 1000 RPM). In some embodiments, centrifugation occurs at a speed of at most 6000 RPM (e.g., at most 5000 RPM, at most 3000 RPM, at most 1000 RPM, at most 600 RPM, at most 500 RPM, at most 300 RPM, at most 100 RPM). In some embodiments, centrifugation occurs at a speed of about 10 RPM to about 6000 RPM, 10 RPM to about 5000 RPM, 10 RPM to about 1000 RPM, 100 RPM to about 1000 RPM, 200 RPM to about 600 RPM, or 300 RPM to about 600 RPM. In some embodiments, centrifugation occurs at about 600 RPM. In some embodiments, centrifugation occurs at a speed of 1000 RPM.
[0090] In some instances, separating herein may refer to pulling down of the macromolecule structures. In some instances, higher rates of pull-down are favorable as they may lead to enhanced rates of isolation of biomolecules (e.g., proteins or peptides). Pull-down rates may be affected by properties such as particle size and polymer coating (e.g., surface charge). FIG. 12a illustrates an example of nanoparticles (NPs) in solution being pulled down through the use of a magnet. The rate of pulldown can be assessed through analytical techniques (e.g., UV-vis spectrometry), where a solution becomes more optically transparent as more nanoparticle beads are pulled down. FIG. 12b illustrates the difference in rate of bead pulldown for two beads of different diameters, macromolecule structure 1 comprising an average core diameter of about 140 nanometers and macromolecule structure 2 comprising an average core diameter of about 190 nanometers. In some embodiments, a particle comprising a larger diameter has a higher pulldown rate in comparison to an otherwise similar particle comprising a smaller diameter. In some embodiments, pulldown refers to the collection of nanoparticle beads, or other components, at the bottom of a vessel upon exposure to a magnet.
[0091] In some embodiments, in any of the embodiments provided herein, a biological sample may comprise a plurality of proteins. In some embodiments, a biological sample comprises plasma, serum, urine, cerebrospinal fluid, synovial fluid, tears, saliva, whole blood, milk, nipple aspirate, ductal lavage, vaginal fluid, nasal fluid, ear fluid, gastric fluid, pancreatic fluid, trabecular fluid, lung lavage, sweat, crevicular fluid, semen, prostatic fluid, sputum, fecal matter, bronchial lavage, fluid from swabbings, bronchial aspirants, fluidized solids, fine needle aspiration samples, tissue homogenates, lymphatic fluid, cell culture samples, or any combination thereof. In some embodiments, the biological sample is plasma, serum, CSF, urine, tear, cell lysate, tissue lysate, cell homogenate, nipple aspirate, fecal matter, synovial fluid, saliva, or blood. In some embodiments, a biological sample comprises plasma, serum, or blood. In some embodiments, a biological sample comprises plasma. In some embodiments, a biological sample comprises blood. In some embodiments, a biological sample comprises biofluid. In some embodiments, a biological sample comprises a cell-free sample. In some embodiments, a biological sample comprises serum. [0092] In some embodiments, the methods provided herein may further comprise digesting the one or more biomolecules (e.g., proteins or peptides) bound to the macromolecule structure. In some embodiments, digesting the one or more biomolecules (e.g., proteins) provides one or more peptides.
[0093] Digesting may refer to breaking the biomolecule, such as a protein, into smaller components, such as peptides. In some instances, digesting comprises breaking proteins from the biological solution into peptides. In some embodiments, digesting comprise enzymolysis. In some embodiments, digestion comprises digestion with trypsin, lysin, serine protease, or any
combination thereof. In some embodiments, digestion comprises digestion with trypsin. In some embodiments, digestion comprises digestion with lysin. In some embodiments, digestion comprises digestion with serine protease.
[0094] In some embodiments, the method may comprise denaturing of the one or more biomolecules (e.g., proteins). In some embodiments, denaturing may result from contact with the organic solvent, such as an organic solvent as provided elsewhere herein. In some embodiments, a denaturing agent is contacted with the biomolecules (e.g., proteins), such as chaotropic agents, surfactants, heavy metals, high salt concentrations, and the like. For example, the one or more biomolecules (e.g., proteins) are denatured by applying urea.
[0095] In some embodiments, the methods provided herein further comprise washing the macromolecule structure comprising the one or more biomolecules (e.g., proteins or peptides) with a wash solvent. In some embodiments, the wash solvent is an organic wash solvent. In some embodiments, the organic wash solvent comprises at least 50 wt% (e.g., 80 wt%, 90 wt%, 95 wt%) of an organic solvent, such as an organic solvent described elsewhere herein. In some embodiments, the organic wash solvent comprises at most 100% wt% (e.g., 99 wt%, 98 wt%, 95 wt%, 90 wt%, 80 wt%) of an organic solvent, such as an organic solvent described elsewhere herein. In some embodiments, the organic wash solvent comprises 100% of an organic solvent, such as an organic solvent as described elsewhere herein. In certain embodiments, an organic wash solvent comprise 100% acetonitrile.
[0096] In any of the methods provided herein, an organic solvent may comprise one or more of an alcohol, acetonitrile, dichloromethane, dimethylformamide (DMF), dimethyl sulfoxide (DMSO), ethylacetate, hexamethylphosphoramide (HPMA), tetrahydrofuran (THF), or any combination thereof. In some embodiments, an alcohol comprises ethanol, methanol, n-propanol, isopropanol, n-butanol, sec-butanol, tert-butanol, pentanol, hexanol, cyclohexanol, or any combination thereof. In some embodiments, the organic solvent comprise acetonitrile. In some embodiments, the organic solvent comprises an aprotic organic solvent, such as acetonitrile and DMSO. In some embodiments, the organic solvent is configured to cause protein binding to a macromolecule structure when contacted with proteins and the macromolecule structure. Without being bound to any particular theory, the binding to the macromolecule structure may be driven by hydrophobic/hydrophilic interactions, wherein the one or more biomolecules (e.g., proteins) favor the more hydrophilic surface of the macromolecule structure relative to the organic solvent. In some embodiments, the organic solvent has an estimated partition coefficient using XLOGP3 of at least -1.5, at least, -1, or at least -0.5. In some embodiments, the organic solvent has an estimated partition coefficient using XLOGP3 of no more than 1.5, no more than 1, no more than 0.5, or no more than 0. In some embodiments, the organic solvent has an estimated partition
coefficient using XL0GP3 of about -1.5 to about 1.5. In some embodiments, the organic solvent has an estimated partition coefficient using XL0GP3 of about - 1 to about 1. In some embodiments, the organic solvent has an estimated partition coefficient using XL0GP3 of about -0.5 to about 0.5. For organic solvent mixtures, the weighted molar average of the estimated partition coefficient may be used. In any of the embodiments provided herein, the methods are capable of isolating at least 10 (e.g., unique) biomolecules (e.g., proteins or peptides). In some embodiments, the methods herein are capable of isolating at least 50 (e.g., at least 100, at least 250, at least 1,000, at least 2,500, at least 5,000, at least 7,500, at least 10,000, at least 12,500, at least 15,000, at least 17,500, at least 20,000) (e.g., unique) biomolecules (e.g., proteins or peptides). In some embodiments, the methods herein are capable of isolating at most 200,000 (e.g., at most 150,000, at most 100,000, at most 50,000, at most 30,000, at most 25,000, at most 20,000, at most 15,000, at most 10,000, at most 5,000, at most 2,500, at most 1,000, at most 500, at most 100) (e.g., unique) biomolecules (e.g., proteins or peptides). In some embodiments, the methods herein are capable of isolating about 10 to about 50,000, about 100 to about 20,000, about 100 to about 15,000, about 100 to about 10,000, about 1,000 to about 20,000, about 1,000 to about 10,000, or about 1,000 to about 5,000 (e.g., unique) biomolecules (e.g., proteins or peptides). In some embodiments, the methods herein are capable of isolating at least 1,000 (e.g., unique) biomolecules (e.g., proteins or peptides). In some embodiments, the methods herein are capable of isolating no more than 25,000 (e.g., unique) biomolecules (e.g., proteins or peptides). The number of biomolecule that may be isolated by a macromolecule structure herein may be modified by the skilled artisan by changing the surface of the macromolecule structure or the number of unique macromolecule structures.
[0097] In certain embodiments, isolating of biomolecules occurs in the absence of any solution comprising salts in an amount effective to precipitate the population of biomolecules (e.g., proteins). Contacting of the biomolecules provided herein may be achieved solely by introduction of the biomolecules and macromolecule structures in an organic solvent, such as an organic solvent described elsewhere herein.
[0098] In some embodiments, the methods provided herein further comprise identifying the one or more biomolecules (e.g., proteins or peptides), such as after isolation of the one or more biomolecules (e.g., proteins or peptides). In some embodiments, the methods provided herein further comprise quantifying the one or more biomolecules (e.g., proteins or peptides), such as after isolation of the one or more biomolecules (e.g., proteins or peptides). The quantification may be, for example, absolute or relative amounts. In some embodiments, the methods provided herein further comprise analyzing the one or more biomolecules (e.g., proteins or peptides) (e.g., such as after (b)). As discussed herein, the methods provide for removal of components of biological
solutions that are incompatible for many analytical techniques, such as mass spectrometry. In some embodiments, identification or analyzing comprises mass spectrometry (MS), liquid chromatography-mass spectrometry (LC-MS), protein sequencing (e.g., single-molecule protein sequencing), light scattering (e.g., dynamic light scattering (DLS), static light scattering (SLS), or circular dichroism (CD), ELISA, immunoaffinity techniques (e.g., proximity extension assays), Edman Degradation, or any combination thereof. In some embodiments, identification or analyzing comprises mass spectrometry, liquid chromatography-mass spectrometry (LC-MS), protein sequencing, or any combination thereof. In some embodiments, identifying or analyzing comprises mass spectrometry. In some embodiments, identifying or analyzing comprises liquid chromatography-mass spectrometry (LC-MS). In some embodiments, the LC-MS/MS includes data independent analy si s/acqui sition (DIA). In some embodiments, the LC-MS/MS includes data dependent analy si s/acqui sition (DDA). In some embodiments, identifying or analyzing comprises protein sequencing.
[0099] FIG. 7 illustrates an example of a schematic of isolating biomolecules for analysis. For example, a biological sample containing proteins is incubated with one or more different macromolecule structures to form a biomolecule (e.g., protein) corona. The biomolecule corona may be washed through pulling down macromolecule structures (e.g., protein corona) using a magnet, removing the supernatant, and resuspending the macromolecule structures (e.g., protein corona). Washed biomolecule (e.g., protein) corona may be subjected to digestion to form one or more peptides and the peptides may be purified prior to analysis via mass spectrometry. Purifying the macromolecule structures may comprise (a) adding the beads in a vessel, (b) incubating in a solution comprising from about 90% to about 100% organic solvent (e.g., acetonitrile), (c) applying a magnetic field to the vessel such that the macromolecule structures and peptides collect at the bottom of the vessel, (d) removing the supernatant, (e) resuspending the macromolecule structures and peptides in a solution comprising from about 90% to about 100% organic solvent (e.g., acetonitrile), (f) applying a magnetic field to the vessel such that the macromolecule structures and peptides collect at the bottom of the vessel, (g) removing the supernatant, (h) resuspending and incubating the contents of the vessel in a solution comprising water and optionally less than 10% organic solvent (e.g., about 3% acetonitrile solution in water), (i) applying a magnetic field to the vessel such that only the macromolecule structures collect at the bottom of the vessel, and (j) collecting the supernatant comprising the purified peptides or proteins. In some embodiments, the organic solvent (e.g., acetonitrile) comprises a purity of about 90% to about 100%. In some embodiments, the organic solvent (e.g., acetonitrile) comprises a purity of about 95% to about 100% (e.g., 96%, 97%, 98%, 99%).
[00100] In some embodiments, the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona at a temperature of about 4 degrees Celsius (°C) to about 90°C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 20°C to about 90°C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 20°C to about 50°C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 4°C to about 40°C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 25°C to about 40°C. The incubation time may be, for example, at least 1 minute, at least 5 minutes, at least 10 minute, at least 15 minutes, at least 20 minutes, at least 30 minutes, at least 45 minutes, or at least 1 hour. As a non-limiting example, the incubation may be at about 4°C to about 40°C for about 20 minutes to about 2 hours. As another example, incubation may be at about 25°C to about 40°C for about 20 minutes to about 90 minutes.
[00101] The biological sample can be incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture. In some embodiments, the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising less than about 40% by volume or weight of organ solvent. In some embodiments, the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising less than about 25% by volume or weight of organ solvent. In some embodiments, the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising less than about 10% by volume or weight of organ solvent. In some embodiments, the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising less than about 5% by volume or weight of organ solvent. In some embodiments, the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising less than about 1% by volume or weight of organ solvent. In some embodiments, the biological sample is incubated with the one or more different macromolecule structures to form a biomolecule (e.g., protein) corona in an aqueous mixture comprising no more than trace amounts of organic solvent.
[00102] The surfaces disclosed herein can be used to identify at least at least 100 (unique) (unique) protein groups, at least 200 (unique) protein groups, at least 300 (unique) protein groups, at least 400 (unique) protein groups, at least 500 (unique) protein groups, at least 600 (unique) protein groups, at least 700 (unique) protein groups, at least 800 (unique) protein groups, at least
900 (unique) protein groups, at least 1000 (unique) protein groups, at least 1100 (unique) protein groups, at least 1200 (unique) protein groups, at least 1300 (unique) protein groups, at least 1400 (unique) protein groups, at least 1500 (unique) protein groups, at least 1600 (unique) protein groups, at least 1700 (unique) protein groups, at least 1800 (unique) protein groups, at least 1900 (unique) protein groups, at least 2000 (unique) protein groups, at least 2100 (unique) protein groups, at least 2200 (unique) protein groups, at least 2300 (unique) protein groups, at least 2400 (unique) protein groups, at least 2500 (unique) protein groups, at least 2600 (unique) protein groups, at least 2700 (unique) protein groups, at least 2800 (unique) protein groups, at least 2900 (unique) protein groups, at least 3000 (unique) protein groups, at least 3100 (unique) protein groups, at least 3200 (unique) protein groups, at least 3300 (unique) protein groups, at least 3400 (unique) protein groups, at least 3500 (unique) protein groups, at least 3600 (unique) protein groups, at least 3700 (unique) protein groups, at least 3800 (unique) protein groups, at least 3900 (unique) protein groups, at least 4000 (unique) protein groups, at least 4100 (unique) protein groups, at least 4200 (unique) protein groups, at least 4300 (unique) protein groups, at least 4400 (unique) protein groups, at least 4500 (unique) protein groups, at least 4600 (unique) protein groups, at least 4700 (unique) protein groups, at least 4800 (unique) protein groups, at least 4900 (unique) protein groups, at least 5000 (unique) protein groups, at least 10000 (unique) protein groups, at least 20000 (unique) protein groups, at least 100000 (unique) protein groups, from 100 to 5000 (unique) protein groups, from 200 to 4700 (unique) protein groups, from 300 to 4400 (unique) protein groups, from 400 to 4100 (unique) protein groups, from 500 to 3800 (unique) protein groups, from 600 to 3500 (unique) protein groups, from 700 to 3200 (unique) protein groups, from 800 to 2900 (unique) protein groups, from 900 to 2600 (unique) protein groups, from 1000 to 2300 (unique) protein groups, from 1000 to 3000 (unique) protein groups, from 3000 to 4000 (unique) protein groups, from 4000 to 5000 (unique) protein groups, from 5000 to 6000
(unique) protein groups, from 6000 to 7000 (unique) protein groups, from 7000 to 8000 (unique) protein groups, from 8000 to 9000 (unique) protein groups, from 9000 to 10000 (unique) protein groups, from 10000 to 11000 (unique) protein groups, from 11000 to 12000 (unique) protein groups, from 12000 to 13000 (unique) protein groups, from 13000 to 14000 (unique) protein groups, from 14000 to 15000 (unique) protein groups, from 15000 to 16000 (unique) protein groups, from 16000 to 17000 (unique) protein groups, from 17000 to 18000 (unique) protein groups, from 18000 to 19000 (unique) protein groups, from 19000 to 20000 (unique) protein groups, from 20000 to 25000 (unique) protein groups, from 25000 to 30000 (unique) protein groups, from 10000 to 20000 (unique) protein groups, from 10000 to 50000 (unique) protein groups, from 20000 to 100000 (unique) protein groups, from 2000 to 20000 (unique) protein
groups, from 1800 to 20000 (unique) protein groups, or from 10000 to 100000 (unique) protein groups. In some embodiments, the identified protein groups are quantified.
[00103] In some embodiments, the methods provided herein further comprise (e.g., such as before (a)), contacting the biological sample comprising a population of biomolecules with an additional macromolecule structure to provide a protein corona. Methods of biomolecule corona formation are disclosed in US2018/0172694, US2021/0285957, US2021/0285958, and W02022/020272, each of which is hereby incorporated by reference in their entirety. For example, the biological sample may be incubated with silica-coated superparamagnetic iron oxide nanoparticles (SPIONs) to form a biomolecule corona.
[00104] In some embodiments, the methods provided herein further comprise (e.g., such as before (a)), treating or digesting one or more proteins of the population of biomolecules to form one or more peptides or desorbed proteins. In some embodiments, proteins from the protein corona are treated or digested. In some embodiments, the digesting comprises enzymatic digestion. In some embodiments, the digesting comprises trypsin digestion. As a non-limiting example, the digestion may comprise incubating the protein corona with LysC and trypsin.
[00105] In some embodiments, the macromolecule structure and the additional macromolecule structure are the same. For example, the macromolecule structure may be polymer-coated SPIONs, which are contacted with a biological sample to form a protein corona. The proteins from protein corona may be digested using trypsin, and then an organic solvent is added to bind the peptides to the same polymer-coated SPIONs. The bound peptides may be eluted for further analysis. Alternatively, the SPIONs used for protein corona formation may be replaced with new SPIONs have the same properties. In some embodiments, the macromolecule structure and the additional macromolecule structure are different. For example, a protein corona may be formed on a particle having a polyacrylamide coating, and then peptides obtained from the protein corona are bound to particles with a polyethylene glycol coating when contacted with a suitable organic solvent. In some embodiments, the additional macromolecule structure may be removed before the biomolecules are contacted with the macromolecule structure and an organic solvent. In some embodiments, both the additional macromolecule structure and the one or biomolecules (e.g., proteins) are contacted with the macromolecule structure and organic solvent.
[00106] In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is at least 100 nanograms (ng). In some embodiments, a mass of the one or more isolated biomolecules (e.g., proteins or peptides) is at least 500 ng (e.g., at least 1 microgram (pg), 10 pg, 50 pg, 100 pg, 200 pg). In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is at most 10 mg (e.g., 5 mg, 1 mg, 750 pg, 500 pg, 250 pg, 200 pg, 100 pg). In some embodiments, the mass of the one or more isolated biomolecules
(e.g., proteins or peptides) is about 100 ng to about 10 mg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is about 100 ng to about 500 pg, about 100 ng to about 250 pg, about 100 ng to about 50 pg, about 10 pg to about 100 pg, or about 10 pg to about 50 pg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is about 0.1 pg to about 50 pg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is about 1 pg to about 10 pg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is about 1 pg to about 5 pg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is about 1 pg to about 2 pg. In some embodiments, the mass of the one or more isolated biomolecules (e.g., proteins or peptides) is sufficient for further analysis or identification, such as analyzing or identifying as described elsewhere herein.
[00107] In some instances, the concentration of macromolecule structures added to a biological solution is modified. In some instances, the concentration of macromolecule structure added to a biological solution has an effect on peptide yield. In some embodiments, increasing the concentration of macromolecule structure added to a biological solution increases the peptide yield. In some embodiments, provided herein are methods of isolating one or more biomolecules (e.g., proteins or peptides) from a biological sample comprising contacting the biological sample with multiple macromolecule structures. In some embodiments, the methods herein comprise contacting the biological sample with a first macromolecule structure followed by a second macromolecule structure.
[00108] In some embodiments, the methods provided herein comprise contacting the biomolecules (e.g., comprising one or more proteins or peptides) with a first macromolecule structure. In some embodiments, contacting biomolecules with a first macromolecule structure provides a protein corona. In some embodiments, the protein corona formation is reversible. In some embodiments, contacting comprises incubating, such as incubating as provided elsewhere herein.
[00109] In some embodiments, the methods provided herein comprise separating the one or more peptides and proteins (e.g., biomolecules) of the protein corona from the first macromolecule structure. In some embodiments, separating comprises separating as provided elsewhere herein, such as by centrifugation or applying a magnetic field.
[00110] In some embodiments, the methods provided herein comprise contacting the one or more peptides and proteins (e.g., biomolecules) with a second macromolecule structure and an organic solvent. In some instances, contacting the one or more peptides and proteins (e.g., biomolecules) with a second macromolecule and an organic solvent provides for binding of at least a portion of the one or more peptide and proteins (e.g., biomolecules) to the second
macromolecule structure. In some embodiments, contacting comprises incubating, such as incubating as described elsewhere herein.
[00111] In some embodiments, the methods provided herein further comprise eluting the one or more peptides and proteins (e.g., biomolecules) from the second macromolecule structure, thereby providing the one or more isolated biomolecules. In some embodiments, eluting comprises eluting as described elsewhere herein.
[00112] In some instances, the first macromolecule structure and the second macromolecule structure are the same. In other instances, the first macromolecule structure and the second macromolecule structure are different. In some embodiments, the second macromolecule structure is added to a composition comprising the first macromolecule structure. In other embodiments, the second macromolecule structure is added to a composition in absence of the first macromolecule structure.
[00113] In some embodiments, the first macromolecule structure or the second macromolecule structure comprises a poly(ethylene) glycol (PEG) group.
[00114] In some embodiments, the method further comprises eluting the one or more biomolecules (e.g., proteins or peptides) from the second macromolecule structure with an aqueous solution. In some embodiments, eluting is elution as described elsewhere herein.
[00115] In some embodiments, macromolecule structures as described herein may be used in variable cleanup methods described herein (e.g., modification of solvent, incubation temperature, separation conditions, etc.).
[00116] In any of the embodiments provided herein, a protein may comprise one or more peptides.
Macromolecule Structures
[00117] In some embodiments, the macromolecule structures provided herein comprise different polymers that may affect the physicochemical properties of the surface such as size, surface charge, hydrophobicity, hydrophilicity, surface functionality, surface topography, surface curvature, porosity, shape, and any combination thereof. The changes in physicochemical properties may affect the binding properties of the macromolecule structures to other compounds, such as biomolecules (e.g., peptides or proteins), leading to increased effectiveness or decreased effectiveness in binding.
[00118] In any of the methods provided herein, the macromolecule structures may comprise a surface and a macromolecule comprising recurring units of a first component and optionally a second component, wherein the first component comprises a structure of Component (A), and wherein the second component is a cross-linking component that may comprises a structure of Component (B’):
[00119] In some embodiments, each of Yi, Y2, and Y3 is independently selected from hydrogen or Ci-Ce alkyl. In some embodiments, Yi is hydrogen or Ci-Ce alkyl. In some embodiments, Yi is hydrogen. In some embodiments, Yi is Ci-Ce alkyl. In some embodiments, Y2 is hydrogen or Ci-Ce alkyl. In some embodiments, Y2 is hydrogen. In some embodiments, Y2 is Ci-Ce alkyl. In some embodiments, Y3 is hydrogen or Ci-Ce alkyl. In some embodiments, Y3 is hydrogen. In some embodiments, Y3 is Ci-Ce alkyl. In some embodiments, Yi is C1-C3 alkyl and each of Y2 and Y3 are hydrogen. In some embodiments, Yi is Ci alkyl (e.g., -CH3) and each of Y2 and Y3 are hydrogen.
[00120] In some embodiments, each of Xi, X2, and X3 is independently selected from hydrogen or Ci-Ce alkyl. In some embodiments, Xi is hydrogen or Ci-Ce alkyl. In some embodiments, Xi is hydrogen. In some embodiments, Xi is Ci-Ce alkyl. In some embodiments, X2 is hydrogen or Ci-Ce alkyl. In some embodiments, X2 is hydrogen. In some embodiments, X2 is Ci-Ce alkyl. In some embodiments, X3 is hydrogen or Ci-Ce alkyl. In some embodiments, X3 is hydrogen. In some embodiments, X3 is Ci-Ce alkyl. In some embodiments, Xi is C1-C3 alkyl and each of X2 and X3 are hydrogen. In some embodiments, Xi is Ci alkyl (e.g., -CH3) and each of X2 and X3 are hydrogen.
[00125] In some embodiments, Ri is hydrogen, optionally substituted succinate, Ci-Ce alkyl substituted with sulfone, or phthalate. In some embodiments, Ri is hydrogen. In some embodiments, Ri is nitrogen. In some embodiments, Ri is optionally substituted succinate. In some embodiments, Ri is Ci-Ce alkyl. In some embodiments, Ri is Ci-Ce alkyl substituited with sulfone. In some embodiments, Ri is phthalate.
[00126] In some embodiments, R2 is C1-C12 alkyl substituted with amine, Ci-Ce alkyl substituted with hydroxy, optionally substituted aryl, optionally substituted heteroaryl, optionally substituted C3-C6 heterocycloalkyl, two or more fused 3-6 member rings; optionally substituted C3-C6 dicycloalkyl methane, Ci-Ce alkyl substituted with guanidine, Ci-Ce alkoxy, optionally substituted Ci-Ce alkyl substituted with disulfide, Ci-Ce alkyl substituted with thiol, optionally substituted succinate, optionally substituted Ci-Ce alkylamine, Ci-Ce alkyl acetamide, Ci-Ce aminophthalate, a boronic acid, C1-C11 optionally substituted cycloalkyl, or a monosaccharide.
[00127] In some embodiments, R2 is C1-C12 alkyl substituted with amine. In some embodiments, R2 is Ci-Ce alkyl substituted with hydroxy. In some embodiments, R2 is optionally substituted aryl. In some embodiments, R2 is optionally substituted heteroaryl. In some embodiments, R2 is optionally substituted C3-C6 heterocycloalkyl. In some embodiments, R2 is two or more fused 3- 6 membered rings. In some embodiments, R2 is optionally substituted Ci-Ce alkyl substituted with disulfide. In some embodiments, R2 is Ci-Ce alkyl substitute with thiol. In some embodiments, R2 is optionally substituted succinate. In some embodiments, R2 is optionally substituted Ci-Ce alkylamine. In some embodiments, R2 is Ci-Ce alkyl acetamide. In some embodiments, R2 is Ci-
Ce aminophthalate. In some embodiments, R2 is a boronic acid. In some embodiments, R2 is Ci- C11 optionally substituted cycloalkyl. In some embodiments, R2 is a monosaccharide.
[00128] In some embodiments, Ri and R2 are taken together with the nitrogen to which they are attached to form an optionally substituted heterocycle.
[00129] In some embodiments, Ri and R2 are taken together with the nitrogen to which they are attached to form an azide group.
[00130] In some embodiments, R2 is Ci-Ce ether, C1-12 alkyl substituted with amine, optionally substituted Ci-Ce alkylamine, -(CH2)I-6 imidazole -, (CH2)I-6 pyridine, optionally substituted aryl, -(CH2)1-6N(CH3)2(CH2CH2CH2SOOOH), -(CH2)I-6N(CH2CH2CH2SOOOH)2, or C2-C6 alkyl substituted with hydroxy.
[00131] In some embodiments, R4 is hydrogen or Ci-Ce alkyl substituted with thiol. In some embodiments, R4 is hydrogen. In some embodiments, R4 is Ci-Ce alkyl substituted with thiol.
[00132] In some embodiments, Rs is succinate, optionally substituted aryl, or optionally substituted Ci-Ce alkyl substituted with disulfide. In some embodiments, Rs is succinate. In some embodiments, Rs is optionally substituted aryl. In some embodiments, Rs is optionally substituted Ci-Ce alkyl substituted with disulfide.
[00134] In some embodiments, q is an integer between 1 and 6. In some embodiments, q is an integer between 1 and 3. In some embodiments, q is 1. In some embodiments, q is 2. In some embodiments, q is 3. In some embodiments, q is 4. In some embodiments, q is 5. In some embodiments, q is 6.
In some embodiments p is an integer between 1 and 20. In some embodiments, p is 1. In some embodiments, p is 2. In some embodiments, p is 3. In some embodiments, p is 4. In some embodiments, p is 5. In some embodiments, p is 6. In some embodiments, p is 7. In some embodiments, p is 8. In some embodiments, p is 9. In some embodiments, p is 10. In some embodiments, p is 11. In some embodiments, p is 12. In some embodiments, p is 13. In some
embodiments, p is 14. In some embodiments, p is 15. In some embodiments, p is 16. In some embodiments, p is 17. In some embodiments, p is 18. In some embodiments, p is 19. In some embodiments, p is 20.
[00135] In some embodiments, the macromolecule structures as described herein and their preparations are macromolecule structures as described in PCT Patent Application No. PCT/US2023/075863, which is incorporated herein by reference in its entirety. In some embodiments, the macromolecule structure may be modified surfaces, such as particles, as described in U.S. Publication No. 2021/0285957, which is incorporated herein by reference in its entirety. For example, the macromolecule structures may be any of the particle types provided in Table 1 of U.S. Publication No. 2021/0285957.
[00136] In some embodiments, the macromolecule structures as described herein are described in Table 1. In some embodiments, n, m, x, and y, are integers representing the number of recurring units. In some embodiments, m is an integer greater than 2. In some embodiments, m is an integer between 2 and 2,000. In some embodiments, m is an integer between 50 and 1,000. In some embodiments, m is an integer between 100 and 200. In some embodiments, n is an integer greater than 2. In some embodiments, n is an integer between 2 and 200. In some embodiments, n is an integer between 2 and 100. In some embodiments, n is an integer between 2 and 20. In some embodiments, the recurring units denoted by n and m are randomly copolymerized. In some embodiments x is an integer greater than 2. In some embodiments, x is an integer between 2 and 50,000. In some embodiments y is an integer greater than 2. In some embodiments, y is an integer between 2 and 50,000.
[00137] In some embodiments,
as represented in the compounds herein (e.g., such as in Table 1) represent an attachment point for a unit of Component (A) or Component (B). In some embodiments,
as represented in the compounds herein (e.g., such as in Table 1) represent an attachment point for a unit of Component (A) or Component (B). In some embodiments,
represent attachment points for crosslinking with a monomer
nstances,
as used herein may refer
some instances,
n may refer interchangeably to
[00139] In some embodiments, the macromolecule structures provided herein can be prepared according to methods as described in Examples 6-8. FIG. 1 shows examples of the general design space of the epoxidated macromolecule structure platform following addition of a functionalizing amine, while FIG. 2 shows examples of the conversion schemes available after epoxidation of macromolecule structures including reaction with glycidyls and azides to obtain diverse functionalities.
[00140] In any of the methods, systems, and kits provided herein, the macromolecule structures provided herein may comprise a surface, a tethering moiety coupled to the surface, and a macromolecule chain. In some embodiments, the macromolecule structures provided herein comprise a surface and a macromolecule chain. In some embodiments, the macromolecule structure provided herein comprises a surface. In some embodiments, the macromolecule structure provided herein comprises a tethering moiety (e.g., coupled to the surface). In some embodiments, the macromolecule structure provided herein comprises a macromolecule chain. In some embodiments, a first end of the macromolecule chain is covalently attached to the tethering moiety. In some embodiments, a second end of the macromolecule chain is not coupled to the surface. In some embodiments, the macromolecule chain comprises one or more (e.g., distinct) recurring units derived from a monomer.
[00141] In some embodiments, the macromolecule chain comprises two or more distinct recurring units derived from different monomer. In some embodiments, the macromolecule chain comprises at least two (e.g., at least three, at least four, at least five) distinct recurring units derived from different monomers. In some embodiments, the macromolecule chain comprises at most ten (e.g., at most nine, at most eight, at most six, at most four, at most three) distinct recurring units derived from different monomers. In some embodiments, the macromolecule chain comprises two distinct recurring units derived from different monomers. In some embodiments, the macromolecule chain comprises three distinct recurring units derived from different monomers. In some embodiments, the macromolecule chain comprises four distinct recurring units derived from different monomers. In some embodiments, the macromolecule chain comprises five distinct recurring units derived from different monomers.
[00142] In some embodiments, the macromolecule chain is comprised of a single monomeric recurring unit.
[00143] In some embodiments, the macromolecule structures provided herein comprise crosslinked polymers (e.g., crosslinked macromolecule chains). For example, the relative amount of crosslinking monomer relative to total monomer in the polymers by weight or number can be at least 0.1%, at least 0.5%, at least 1%, or at least 2%. In some embodiments, the macromolecule structures provided herein comprise substantially uncross-linked polymers (e.g., uncrosslinked macromolecule chains). For example, the relative amount of crosslinking monomer relative to total monomer in the polymers by weight or number can be less than 0.1%, less than 0.05%, less than 0.01%, or about 0%. In some embodiments, the macromolecule structures provided herein comprise polymer brushes (e.g., PEG brushes). For example, the polymer brushes may comprise sidechains with at least 5, at least 10, at least 15, or at least 20 recurring units derived from one or more monomers.
[0001] In one aspect, described herein is a macromolecule structure comprising a moiety of Formula (IV):
Formula (IV) wherein Z is a linking moiety comprising a linear chain with 2 to 20 atoms selected from the group consisting of carbon, oxygen, and nitrogen, and optional substituents on the carbon and any nitrogen atoms in the linear chain;
Ri is hydrogen or succinate; and
R2 is optionally substituted Ci-Ce alkyl substituted with disulfide or Ci-Ce alkyl substituted with thiol.
[0002] In some embodiments, Z is a linear chain with 2 to 20 atoms. In some embodiments, Z is a linear chain with 2 to 12 atoms. In some embodiments, Z is a linear chain with 2 to 6 atoms. In some embodiments, Z is a linear chain with 2 atoms. In some embodiments, Z is a linear chain with 3 atoms. In some embodiments, Z is a linear chain with 4 atoms. In some embodiments, Z is a linear chain with 5 atoms. In some embodiments, Z is a linear chain with 6 atoms. In some embodiments Z comprises carbon only. In some embodiments, Z is a C2-C6 alkyl chain. In some embodiments, Z is C3 alkyl. In some embodiments Z comprises oxygen, nitrogen, carbon, or a combination thereof. In some embodiments, Z comprises substituents on the linear chain.
[0003] In some embodiments, Z comprises a C2-C8 heteroalkyl, optionally substituted (e.g., with hydroxy). In some embodiments, Z comprises a C2-C8 alkoxy, optionally substituted (e.g., with hydroxy).
[0004] In some embodiments, Ri is hydrogen, optionally substituted succinate, optionally substituted glutarate, optionally substituted adipate, optionally substituted pimelate, optionally substituted suberate, optionally substituted azelate, or optionally substituted sebacate. In some embodiments, Ri is optionally substituted glutarate. In some embodiments, Ri is optionally substituted adipate. In some embodiments, Ri is optionally substitute pimelate. In some embodiments, Ri is optionally substituted suberate. In some embodiments, Ri is optionally substituted azelate. In some embodiments, Ri is optionally substitute sebecate. In some embodiments, Ri is hydrogen. In some embodiments, Ri is optionally substituted Ci-Ce alkyl. In some embodiments, Ri is substituted with one or more oxo and -COOH. In some embodiments, Ri is succinate. In some embodiments, Ri is optionally substituted succinate. In some embodiments, R2 is optionally substituted -Ci-Ce alkyl substituted with disulfide. In some embodiments, R2 is Ci-Ce thiol. In some embodiments, R2 is substituted -Ci-Ce alkyl substituted with disulfide. In some embodiments, R2 is substituted di-Ci-Ce alkyl disulfide. In some embodiments, R2 is -CH2CH2-S-S-CH2CH2NH2. In some embodiments, R2 is C2 alkyl substituted with thiol (e.g., -(CH2)2SH). In some embodiments, Z is attached to a surface. In some embodiments, Z is covalently attached is a surface. [00144] In some embodiments, a structure of Formula (IV) comprises
(Compound 508). In some embodiments, a structure of Formula (IV) comprises
(Compound 507). In some embodiments, a structure of
[00145] FIG. 5 illustrates examples of PEG structures that a macromolecule structure described herein may comprise. In some embodiments, the PEG structure comprises from about 2 to about 20 repeating ethylene glycol units. In some embodiments, a macromolecule structure comprising a PEG component with increasing number of ethylene glycol units may comprise a more positive Zeta (mV) value.
[00146] In some embodiments, a PEG polymer may be a thermo-responsive polymer. In some embodiments, a thermo-responsive polymer is sensitive to thermal energy (e.g., heat or increase in temperature). In some embodiments, exposure of a thermo-responsive polymer to heat may cause one or more physical properties of the thermo-responsive polymer to change. For example, solubility of a thermos-responsive polymer may change upon exposure to heat. In some embodiments, the thermo-responsive polymer has a lower critical solution temperature. For example, the solubility may decrease as the temperature is raised above the lower critical solution temperature. In some embodiments, the temperature may be modified to improve protein or peptide capture on a macromolecule surface. In some embodiments, the temperature may be modified to improve isolation of a macromolecule surface and any bound biomolecules (e.g., peptides or proteins).
[00147] In some embodiments, the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci-Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally
substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, R4 is hydrogen, sulfonate, carboxylate, C1-C4 alkylene, amine (e.g., quaternary ammonium cation), or Ci-Ce alkyl optionally substituted with halogen. In some embodiments, R4 is hydrogen. In some embodiments, R4 is absent. In some embodiments, R4 is sulfonate. In some embodiments, R4 is amine. In some embodiments, R4 is a quaternary ammonium cation. In some embodiments, R4 is carboxylate. In some embodiments, R4 is C1-C4 alkylene. In some embodiments, R4 is Ci-Ce alkyl optionally substituted with halogen (e.g., haloalkyl). In some embodiments, X is -C- or -N-. In some embodiments, X is -C-. In some embodiments, X is -N-. In some embodiments, Y is -C- or -N-. In some embodiments, Y is -C-. In some embodiments, Y is -N-.
[00148] In some embodiments, the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
[00149] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci- Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, R9 is hydrogen or oxo. In some embodiments, R9 is hydrogen. In some embodiments, R9 is oxo. In some embodiments, X is -C- or -N-. In some embodiments, X is -C-. In some embodiments, X is -N-. In some embodiments, Y is -C-, -N-, or -O-. In some embodiments, Y is -C-. In some embodiments, Y is -O-. In some embodiments, Y is -N-. In some embodiments, = is a single bond or a double bond. In some embodiments, = is a single bond. In some embodiments, = is a double bond. In some embodiments, one of = is a single bond and one of = is a double bond. In some embodiments, both = are single bonds. In some embodiments, both = are double bonds.
[00150] In some embodiments, the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
[00151] In some embodiments, a recurring unit is derived from an acrylate monomer. In some embodiments, a recurring unit is derived from a methacrylate monomer.
[00152] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl.
[00153] In some embodiments, R2 is hydrogen or Ci-Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl.
[00154] In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is 3-, 5-, or 6-membered heterocycle optionally substituted with one or more methyl. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl.
[00155] In some embodiments, R5 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6- membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further substituted with amine, hydroxyl, aryl, or sulfonate, Ci- Cs alkoxy optionally substituted with one or more oxo or halogen, -C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted (e.g., with 2 or more fused 6 membered rings further optionally substituted), optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne. In some embodiments, R5 is C1-C12 alkylamine.
[00156] In some embodiments, R5 is hydrogen. In some embodiments, R5 is Ci-Ce alkyl. In some embodiments, R5 is C1-C4 alkylyne. In some embodiments, R5 is Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci- Cni ethylene glycol. In some embodiments, R5 is Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol. In some embodiments, R5 is Ci-Cs alkylamine. In some embodiments, R5 is Ci-Cs alkylamine substituted with amine. In some embodiments, R5 is Ci-Cs alkylamine substituted with hydroxyl and amine. In some embodiments, R5 is Ci-Cs alkylamine substituted with sulfonate. In some
embodiments, R5 is Ci-Cs alkylamine substituted with aryl. In some embodiments, R5 is Ci-Cs alkoxy. In some embodiments, R5 is Ci-Cs alkoxy substituted with one or more oxo or halogen. In some embodiments, R5 is Ci-Cs alkoxy substituted with one or more oxo. In some embodiments, R5 is Ci-Cs alkoxy substituted with one or more halogen. In some embodiments, the alkylamine comprises a quaternary ammonium cation.
[00157] In some embodiments, R5 is C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted (e.g., with 2 or more fused 6 membered rings further optionally substituted), optionally substituted benzyl, trimethoxysilane, or phosphorocholine. In some embodiments, R5 is C1-C3 alkyl substituted with pyrene. In some embodiments, R5 is C1-C3 alkyl substituted with 2 or more 5-6 membered rings further optionally substituted, such as with 2 or more fused 6 membered rings further optionally substituted. In some embodiments, R5 is C1-C3 alkyl substituted with optionally substituted benzyl. In some embodiments, R5 is C1-C3 alkyl substituted with trimethoxysilane. In some embodiments, R5 is C1-C3 alkyl substituted with phosphorocholine.
[00158] In some embodiments, R5 is hydrogen, Ci-Ce alkyl, or C1-C3 alkyl substituted with pyrene or 2 or more fused 5-6 membered rings optionally further substituted. In some embodiments, R5 is hydrogen, Ci-Ce alkyl, or C1-C3 alkyl substituted with pyrene or 2 or more fused 5-6 membered rings optionally further substituted and the macromolecule chain comprises two or more distinct recurring units.
[00159] In some embodiments, the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
[00160] In some embodiments, a recurring unit is derived from an acrylamide monomer.
[00161] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl.
[00162] In some embodiments, R2 is hydrogen or Ci-Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl.
[00163] In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is 3-, 5-, or 6-membered heterocycle
optionally substituted with one or more methyl. In some embodiments, R3 is 3,4-dimethyl- l//- pyrrole-2, 5-dione. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. [00164] In some embodiments, R6 is hydrogen or Ci-Ce alkyl. In some embodiments, R6 is hydrogen. In some embodiments, R6 is Ci-Ce alkyl.
[00165] In some embodiments, R7 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6- membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, -C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted (e.g., with 2 or more fused 6 membered rings further optionally substituted), optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne.
[00166] In some embodiments, R7 is hydrogen. In some embodiments, R7 is Ci-Ce alkyl. In some embodiments, R7 is C1-C4 alkylyne. In some embodiments, R7 is Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci- Cni ethylene glycol. In some embodiments, R7 is Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol. In some embodiments, R7 is Ci-Cs alkylamine. In some embodiments, R7 is Ci-Cs alkylamine substituted with amine. In some embodiments, R7 is Ci-Cs alkylamine substituted with sulfonate. In some embodiments, R7 is Ci-Cs alkoxy. In some embodiments, R7 is Ci-Cs alkoxy substituted with one or more oxo or halogen. In some embodiments, R7 is Ci-Cs alkoxy substituted with one or more oxo. In some embodiments, R7 is Ci-Cs alkoxy substituted with one or more halogen. In some embodiments, R7 is Ci-Ce alkyl optionally substituted with hydroxyl, substituted benzene, or hydrogen.
[00167] In some embodiments, R7 is C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted (e.g., with 2 or more fused 6 membered rings further optionally substituted), optionally substituted benzyl, trimethoxysilane, or phosphorocholine. In some embodiments, R7 is C1-C3 alkyl substituted with pyrene. In some embodiments, R7 is C1-C3 alkyl substituted with 2 or more 5-6 membered rings further optionally substituted, such as with 2 or more fused 6 membered rings further optionally substituted. In some embodiments, R7 is C1-C3 alkyl substituted with optionally substituted benzyl. In some embodiments, R7 is C1-C3 alkyl substituted with trimethoxysilane. In some embodiments, R7 is C1-C3 alkyl substituted with phosphorocholine.
[00168] In some embodiments, the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
[00169] In some embodiments, a recurring unit derived from a monomer can be a diene, such as a cis diene. In some embodiments, the diene may act as a cross-linking monomer.
[00170] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci- Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci-Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci- Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl.
[00171] In some embodiments, the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
[00172] In some embodiments, a recurring unit is derived from an acrylonitrile monomer.
[00173] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci- Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, R1 is hydrogen, R2 is hydrogen, and R3 is hydrogen.
[00174] In some embodiments, the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
[00175] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci- Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl.
[00176] In some embodiments, the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
[00177] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl.
[00178] In some embodiments, R2 is hydrogen or Ci-Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl.
[00179] In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl.
[00180] In some embodiments, R6 is hydrogen or Ci-Ce alkyl. In some embodiments, R6 is hydrogen. In some embodiments, R6 is Ci-Ce alkyl. In some embodiments, R6 is methyl.
[00181] In some embodiments, Q is -CH2- or ethylene glycol. In some embodiments, Q is -CH2- . In some embodiments, Q is ethylene glycol.
[00182] In some embodiments, m is 1-20. In some embodiments, m is 1-10. In some embodiments, m is 1-5. In some embodiments, m is 5-10. In some embodiments, m is 10-20. In some embodiments, m is 2. In some embodiments, m is 1. In some embodiments, m is 3, 4, 5, 6, 7, 8, 9, or 10.
[00183] In some embodiments, A is a polymeric side chain comprising a recurring unit derived from a monomer represented by the structure:
[00184] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci-
Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, R1 is hydrogen, R2 is hydrogen, and R3 is hydrogen. In some embodiments, R5 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, -C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne. In certain embodiments, R5 is polyethylene glycol. In some embodiments, R5 is polyethylene glycol with a chain length of 9. In some embodiments, R1 is hydrogen, R2 is hydrogen, R3 is methyl, Q is -CH2-, R6 is methyl, and R5 is polyethylene glycol.
[00185] In some embodiments, the macromolecule chains provided herein comprise a recurring unit derived from a monomer represented by the structure:
[00186] In some embodiments, a recurring unit is derived from a dimethacrylate monomer. In some embodiments, the dimethacrylate monomer is a cross-linking monomer.
[00187] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci- Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci-Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci- Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, Z is -O- or -NH. In some embodiments, Z is -O-. In some embodiments, Z is -NH. In some embodiments, R8 is Ci-Ce alkyl, divalent metal, or
symmetric or asymmetric disulfide. In some embodiments, R8 is Ci-Ce alkyl. In some embodiments, R8 is a divalent metal. In some embodiments, R8 is cadmium(II). In some embodiments, R8 is symmetric or asymmetric disulfide. In some embodiments, R8 is a symmetric disulfide. In some embodiments, R8 is symmetrical disulfide (e.g., CH2CH2S-SCH2CH2) or a divalent metal.
[00188] In any of the macromolecule structures provided herein, n1 is an integer selected from 1-100. In some embodiments, n1 is an integer selected from 5-100. In some embodiments, n1 is an integer selected from 5-50. In some embodiments, n1 is an integer selected from 50-100. In some embodiments, n1 is an integer selected from 10-40. In some embodiments, n1 is an integer selected from 1-20. In some embodiments, n1 is an integer selected from 1-10. In some embodiments, n1 is an integer selected from 1-5. In some embodiments, n1 is an integer selected from 5-10. In some embodiments, n1 is an integer selected from 10-20. In some embodiments, n1 is 9. In some embodiments, n1 is an integer selected from 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10.
[00189] In some embodiments, any of the macromolecule chains provided herein may be terminated by a terminating group. The terminating group may be a halogen, epoxide, or olefin (e.g., alkylene). In some embodiments, the terminating group is a halogen. In some embodiments, the terminating group is a bromine. In some embodiments, the terminating group is an iodine. In some embodiments, the terminating group is a chloride. In some embodiments, the terminating group is an epoxide. In some embodiments, the terminating group is an olefin.
[00190] In some embodiments, the macromolecule chains of the macromolecule structures provided herein comprise a recurring unit derived from a polyalkylene glycol (PAG) methacrylate, polyalkylene glycol (PAG) acrylate, polyalkylene glycol (PAG) methacrylamide, polyalkylene glycol (PAG) acrylamide, polyalkylene glycol (PAG) vinyl ether, and combinations thereof, where the number of alkylene oxide units is from 1 to 100 (e.g., 5 to 100, 5 to 90, 8 to 50, 8 to 30, and 10 to 25), and wherein the polyalkylene glycol (e.g., polyethylene glycol) is optionally terminated with Ci-Cs alkoxy group, such as methoxy, ethoxy, propoxy, butoxy, and the like. In some embodiments, the macromolecule chains of the macromolecule structures provided herein comprise a recurring unit derived from a polyethylene glycol (PEG) methacrylate, polyethylene glycol (PEG) acrylate, polyethylene glycol (PEG) methacrylamide, polyethylene glycol (PEG) acrylamide, polyethylene glycol (PEG) vinyl ether, and combinations thereof, where the number of alkylene oxide units is from 1 to 100 (e.g., 5 to 100, 5 to 90, 8 to 50, 8 to 30, and 10 to 25), and wherein the polyalkylene glycol (e.g., polyethylene glycol) is optionally terminated with Ci-Cs alkoxy group, such as methoxy, ethoxy, propoxy, butoxy, and the like.
[00191] In some embodiments, the macromolecule chains of the macromolecule structures provided herein comprise a recurring unit derived from a monomer represented by a structure of
Table 2. In some embodiments, macromolecule chains comprise two or more distinct recurring units derived from a monomer represented by a structure of Table 2.
[00192] In some embodiments, the macromolecule chains of the macromolecule structures provided herein comprise a recurring unit derived from a monomer represented by a structure of Table 3. In some embodiments, macromolecule chains comprises a recurring unit (e.g., one or more) derived from a monomer represented by a structure of Table 3.
[00193] In some embodiments, the recurring units provided herein are randomly distributed throughout the macromolecule chain. In some embodiments, the recurring units provided herein have a controlled distribution throughout the macromolecule chain. In some embodiments, the macromolecule chain is a homopolymer. In some embodiments, the macromolecule chain is a block copolymer. In some embodiments, the macromolecule chain is a random copolymer. The skilled artisan, guided by the disclosure herein, would understand how to provide polymers with randomly or controlled distributions of recurring units.
[00194] When multiple distinct recurring units are present in a macromolecule chain, the multiple recurring units may be present in equal or differing ratios. The ratio of distinct recurring units may be controlled according to methods known to one skilled in the art, including modifying the stoichiometry of monomers added.
[00195] In some embodiments, the macromolecule structures comprise (I) a surface and (II) a macromolecule chain coupled to the surface. In some embodiments, the macromolecule chain comprises a recurring unit of Formula (I):
[00196] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R1 is methyl. In some embodiments, R2 is hydrogen or Ci-Ce alkyl. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R2 is methyl. In some embodiments, R3 is hydrogen or Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, R1 , R2 , and R3 are Ci-Ce alkyl (e.g., methyl).
[00197] In some embodiments, n is an integer from 1 to 10,000. In some embodiments, n is an integer from 1 to 200. In some embodiments n is an integer of no more than 25,000 (e.g., no more than 20,000, no more than 15,000, no more than 10,000, no more than 5,000, no more than 2,500,
no more than 1,000, no more than 500, no more than 100, no more than 50). In some embodiments, n is an integer from 1 to 5,000. In some embodiments, n is an integer from 1 to 2,500. In some embodiments, n is an integer from 1 to 1,000. In some embodiments, n is an integer of no more than 200. In some embodiments, n is an integer from 1 to 100. In some embodiments, n is an integer from 1 to 50.
[00198] In some embodiments, A is a polymeric side chain comprising any of the recurring units provided elsewhere herein. In some embodiments, A is a poly(alkylene oxide) methacrylate or poly(alkylene oxide) acrylate. In some embodiments, A is poly(ethylene glycol) methacrylate or poly(ethylene glycol) acrylate. In some embodiments, A is a poly(alkylene oxide) methacrylamide or poly(alkylene oxide) acrylamide. In some embodiments, A is poly(ethylene glycol) methacrylamide or poly(ethylene glycol) acrylamide.
[00199] In some embodiments, L is a linker moiety. In some embodiments, the linker moiety is represented by the structure:
[00200] In some embodiments, each Z is independently -O- or -N-. In some embodiments, Z is -O-. In some embodiments, Z is -N-. In some embodiments, X’ is Ci-Ce alkyl. In some embodiments, X’ is methyl. In some embodiments, X’ is ethyl. In some embodiments, X’ is propyl.
[00201] In some embodiments, the macromolecule structure further comprises a cross-linking moiety. In some embodiments, the cross-linking moiety comprises a structure represented by:
[00202] In some embodiments, the cross-linking moiety is a derived from a dimethacrylate monomer, such as a dimethacrylate monomer as described elsewhere herein. In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci-Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci-Ce alkyl. In some embodiments, R2 is hydrogen. In some
embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, Z is -O- or -NH. In some embodiments, Z is -O-. In some embodiments, Z is -NH. In some embodiments, R8 is Ci-Ce alkyl, divalent metal, or symmetric or asymmetric disulfide. In some embodiments, R8 is Ci-Ce alkyl. In some embodiments, R8 is a divalent metal. In some embodiments, R8 is cadmium(II). In some embodiments, R8 is symmetric or asymmetric disulfide. In some embodiments, R8 is a symmetric disulfide. In some embodiments, the cross-linking moiety is ethyleneglycol dimethacrylate (EGDMA). In some embodiments, the cross-linking moiety is ethyleneglycol dimethacrylamide. In some embodiments, the cross-linking moiety is ethyleneglycol diacrylate. In some embodiments, the cross-linking moiety is ethyleneglycol diacrylamide.
[00204] In some embodiments, R1 is hydrogen or Ci-Ce alkyl. In some embodiments, R1 is hydrogen. In some embodiments, R1 is Ci-Ce alkyl. In some embodiments, R2 is hydrogen or Ci- Ce alkyl. In some embodiments, R2 is hydrogen. In some embodiments, R2 is Ci-Ce alkyl. In some embodiments, R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl. In some embodiments, R3 is hydrogen. In some embodiments, R3 is Ci-Ce alkyl. In some embodiments, R3 is methyl. In some embodiments, R4 is C1-C4 alkylene. In some embodiments, the cross-linking moiety is divinylbenzene.
[00205] In some embodiments, the macromolecule structures provided herein comprise a tethering moiety which couples the surface to the macromolecule chain. In some embodiments, the tethering moiety is C1-C20 heteroalkyl optionally substituted with one or more Ci-Ce alkyl, oxo, halo, or hydroxyl.
[00208] In some embodiments, the tethering moiety is C1-C12 alkoxy optionally substituted with one or more C1-C20 heteroalkyl, each of the C1-C12 alkoxy and C1-C20 heteroalkyl optionally substituted with one or more Ci-Ce alkyl, oxo, halo, or hydroxyl. In some embodiments, the tethering moiety is represented by the structure:
[00209] In some embodiments, Y is C1-C20 heteroalkyl. In some embodiments, Y is C1-C20 heteroalkyl optionally substituted with one or more Ci-Ce alkyl, oxo, halo, or hydroxyl. In some embodiments, p is an integer from 1 to 12. In some embodiments, p is an integer from 1 to 6. In some embodiments, p is an integer from 1 to 3. In some embodiments, p is 3. In some embodiments, the tethering moiety is represented by the structure:
[00211] In any of the macromolecule structures provided herein, R5 or R7 may comprise polyethylene glycol. In some embodiments, polyethylene glycol may have a chain length of about 1 to about 50. In some embodiments, the polyethylene glycol may have a chain length of no more than 100. In some embodiments, the polyethylene glycol may have a chain length of at least 5. In some embodiments, the polyethylene glycol may have a chain length of about 5 to about 100 or about 5 to about 50. In some embodiments, the polyethylene glycol may have a chain length of about 9.
[00212] In some embodiments, the polyethylene glycol provided herein is an oligoethylene glycol. In some embodiments, the polyethylene glycol is a diethylene glycol. In some embodiments, the polyethylene glycol is a triethylene glycol. In some embodiments, the polyethylene glycol is a tetraethylene glycol.
[00213] In some embodiments, the macromolecule structures provided herein comprise at least 5% w/w of a recurring unit provided herein. In some embodiments, the macromolecule structure comprises at least 10% w/w, at least 15% w/w, at least 20% w/w, at least 25% w/w, at least 30% w/w, at least 35% w/w, at least 40% w/w, at least 45% w/w, at least 50% w/w, at least 60% w/w, at least 75% w/w, at least 90% w/w, at least 95% w/w, at least 99% w/w, or about 100% w/w of
a recurring unit provided herein. In some embodiments, the macromolecule structure comprises at most 95% w/w, at most 90% w/w, at most 85% w/w, at most 75% w/w, at most 70% w/w, at most 65% w/w, at most 60% w/w, at most 55% w/w, at most 50% w/w, at most 45% w/w, at most 40% w/w, at most 35% w/w, or at most 30% w/w of a recurring unit provided herein. In some embodiments, the macromolecule structure comprises about 5% to about 95% w/w of a recurring unit provided herein. In some embodiments, the macromolecule structure comprises about 5% to about 75% w/w of a recurring unit provided herein. In some embodiments, the macromolecule structure comprises about 10% to about 50% w/w of a recurring unit provided herein. In certain embodiments, the macromolecule structure comprises at least 10% w/w of the recurring unit. In certain embodiments, the macromolecule structure comprises at most 50% w/w of the recurring unit. The weight percentage of the recurring unit in the macromolecule structure may be determined by thermogravimetric analysis (TGA).
[00214] In some embodiments, the macromolecule structures as described herein and their preparations are macromolecule structures as described in U.S. Patent Application No. 63/517,515, which is incorporated herein by reference in its entirety.
[00215] In some embodiments, the macromolecule structures provided herein may have a structure represented in Table 4.
[00216] In some embodiments, in Table 4, in any of the structures, “b” denotes a blockcopolymer structure. In some embodiments, the block copolymer structures depicted in Table 3 may alternatively be a random copolymer.
[00217] In some embodiments, the preparation of macromolecule structures provided herein is described in Examples 1-3. In some instances, a synthetic scheme of macromolecule structures provided herein is provided in FIG. 3. In some instances, a synthetic scheme as described by the methods provided herein, such as in Examples 1-3, is provided in FIG. 4. In some instances, a vinyl-functionalized surface is subject to a polymerization reaction in the present of a
diacrylamide cross-linking agent and hydroxyalkyl methacrylate monomer. In some instances, a polymer initiator is coupled to the hydroxyalkyl sidechain of the cross-linked polymer. In some instances, a second polymerization is performed to incorporate ethyleneglycol methacrylate monomers.
[00218] In any of the macromolecule structures provided herein, the macromolecule chains comprise from 1 to 1,000 recurring units. In some embodiments, the macromolecule chains comprise at least 1, at least 10, at least 50, at least 100, at least 250, at least 500, at least 750, or at least 1,000 recurring units. In some embodiments, the macromolecule chain comprises at most 2,500 recurring units. In some embodiments ,the macromolecule chains comprise at most 1,000 recurring units. In some embodiments, the macromolecule chains comprise at most 750 recurring units. In some embodiments, the macromolecule chains comprise at most 500 recurring units. In some embodiments, the macromolecule chains comprise at most 250 recurring units. In some embodiments, the macromolecule chains comprise at most 100 recurring units. In some embodiments, the macromolecule chains comprise about 1 to about 100 recurring units. In some embodiments, the macromolecule chains comprise about 1 to about 250 recurring units. In some embodiments, the macromolecule chains comprise about 1 to about 500 recurring units. In some embodiments, the macromolecule chains comprise about 1 to about 1,000 recurring units. In some embodiments, the macromolecule chains comprise about 100 to about 1,000 recurring units. In some embodiments, the macromolecule chains comprise about 1 to about 10 recurring units.
[00219] In any one of the macromolecule structures provided herein, the macromolecule chains comprise a molecular weight of about 0.1 kDa to about 500 kDa. In some embodiments, the macromolecule chains comprise a molecular weight of at least 0.1 kDa, at least 1 kDa, at least 5 kDa, at least 10 kDa, at least 20 kDa, at least 25 kDa, at least 50 kDa, at least 100 kDa, at least 250 kDa, or at least 500 kDa. In some embodiments, the macromolecule chains comprise a molecular weight of no more than 1000 kDa, no more than 500 kDa, no more than 750 kDa, no more than 500 kDa, no more than 250 kDa, no more than 100 kDa, no more than 75 kDa, no more than 50 kDa, no more than 40 kDa, no more than 30 kDa, no more than 25 kDa, no more than 20 kDa, no more than 15 kDa, or no more than 10 kDa. In some embodiments, the macromolecule chain comprises a molecular weight of about 0.1 to about 500 kDa, about 0.1 to about 250 kDa, about 0.1 kDa to about 100 kDa, about 0.1 kDa to about 70 kDa, 0.5 kDa to about 10 kDa, 0.5 kDa to about 15 kDa, or about 1 kDa toa bout 25 kDa. In some embodiments, the macromolecule chain comprises a molecular weight of about 0.1 kDa to about 100 kDa. In some embodiments, the macromolecule chain comprises a molecular weight of about 0.1 kDa to about 50 kDa.
[00220] In some embodiments, the macromolecular chain comprises a block co-polymer. In some embodiments, the block co-polymer comprises a first block derived from a first monomer
and a second block derived from a second monomer, wherein the first block is adjacent to the tethering moiety, and wherein the first monomer is more hydrophobic than the second monomer. In some embodiments, the block co-polymer comprises a first block derived from a first monomer and a second block derived from a second monomer, wherein the first block is adjacent to the tethering moiety, and wherein the first monomer is less hydrophobic than the second monomer. In some embodiments, the hydrophobicity of the first monomer and the second monomer can be determined by the estimated partition coefficient using XLOGP3. In some embodiments, an absolute difference between the estimated partition coefficients of the first monomer and the second monomer is at least 0.3, at least 0.5, at least 0.8, at least 1, at least 1.5, or at least 2. In some embodiments, an absolute difference between the estimated partition coefficients of the first monomer and the second monomer is no more than 3, no more than 2.5, no more than 2, no more than 1, or no more than 0.8.
[00221] In some embodiments, any of the macromolecule structures provided herein, such as used in the methods provided herein, comprise a surface. In some embodiments, a surface is a particle. In some embodiments, a particle is a nanoparticle or a microparticle. In some embodiments, the particle is a nanoparticle. In some embodiments, the particle is a microparticle. [00222] In some instances, the particles provided herein have a diameter of at least 10 nm, at least 100 nm, at least 200 nm, at least 300 nm, at least 400 nm, at least 500 nm, at least 600 nm, at least 700 nm, at least 800 nm, or at least 900 nm. In some embodiments, the particles provided herein have a diameter of no more than 5000 nm, no more than 4000 nm, no more than 3000 nm, no more than 2000 nm, no more than 1000 nm, no more than 750 nm, or no more than 500 nm. In some embodiments, the particles provided herein have a diameter of from 10 nm to 50 nm, from 50 nm to 100 nm, from 100 nm to 150 nm, from 150 nm to 200 nm, from 200 nm to 250 nm, from 250 nm to 300 nm, from 300 nm to 350 nm, from 350 nm to 400 nm, from 400 nm to 450 nm, from 450 nm to 500 nm, from 500 nm to 550 nm, from 550 nm to 600 nm, from 600 nm to 650 nm, from 650 nm to 700 nm, from 700 nm to 750 nm, from 750 nm to 800 nm, from 800 nm to 850 nm, from 850 nm to 900 nm, from 100 nm to 300 nm, from 150 nm to 350 nm, from 200 nm to 400 nm, from 250 nm to 450 nm, from 300 nm to 500 nm, from 350 nm to 550 nm, from 400 nm to 500 nm, from 400 nm to 600 nm, from 450 nm to 650 nm, from 500 nm to 700 nm, from 550 nm to 750 nm, from 600 nm to 800 nm, from 650 nm to 850 nm, from 700 nm to 900 nm, or from 10 nm to 900 nm. In some embodiments, the particles have a diameter of about 100 to about 400 nm. In some embodiments, the particles have a size of about 100 to about 300 nm. The particle size (e.g., diameter) can be measured by dynamic light scattering (DLS) as an indirect measure of size. The DLS measurement can be an ‘intensity -weighted’ average, which means the size distribution that the mean is calculated from can be weighted by the sixth power of radius. This
can be referred to herein as ‘z-average’ or ‘intensity-mean’. Particle size can also be measured by electron microscopy (e.g., SEM, TEM).
[00223] In certain examples, the particles provided herein may comprise a diameter of about 100 nm to about 500 nm. In some embodiments, the particles comprise a diameter of about 100 nm to about 300 nm. In some embodiments, the particles comprise a diameter of about 100 nm to about 200 nm. In some embodiments, the particles comprise a diameter of about 150 nm to about 250 nm.
[00224] In some embodiments, the particle size effects pull-down rate. In some instances, the particle size does not affect the ability of the macromolecule structure to isolate biomolecules, such as the ability to isolate unique biomolecules.
[00225] Additionally, particles can have a homogenous size distribution or a heterogeneous size distribution. Poly dispersity index (PDI), which can be measured by techniques such as dynamic light scattering, is a measure of the size distribution. A low PDI indicates a more homogeneous size distribution and a higher PDI indicates a more heterogeneous size distribution. For example, particles disclosed herein can have a PDI of less than 0.5, less than 0.4, less than 0.3, less than 0.2, less than 0.15, or less than 0.1. In particular embodiments, the particles disclosed herein have a PDI of less than 0.1. In some embodiments, the particles may have a PDI of at least 0.5, or at least 0.75.
[00226] Particles disclosed herein can have a range of different surface charges. Particles can be negatively charged, positively charged, or neutral in charge. In some embodiments, particles have a surface charge of -150 mV to -100 mV, -100 mV to -90 mV, -90 mV to -80 mV, -80 mV to -70 mV, -70 mV to -60 mV, -60 mV to -50 mV, -50 mV to -40 mV, -40 mV to -30 mV, -30 mV to - 20 mV, -20 mV to -10 mV, -10 mV to 0 mV, -10 mV to 10 mV, 0 mV to 10 mV, 10 mV to 20 mV, 20 mV to 30 mV, 30 mV to 40 mV, 40 mV to 50 mV, 50 mV to 60 mV, 60 mV to 70 mV, 70 mV to 80 mV, 80 mV to 90 mV, 90 mV to 100 mV, 100 mV to 110 mV, 110 mV to 120 mV, 120 mV to 130 mV, 130 mV to 140 mV, 140 mV to 150 mV, -150 mv to -100 mV, -100 mv to 0 mV, 0 mv to 100 mV, 100 mv to 150 mV. In particular examples, particles disclosed herein have a surface charge of -60 mV to 60 mV. The surface charge may be evaluated by zeta potential analysis at about neutral pH using an appropriate buffer (e.g., phosphate-buffered saline).
[00227] Various particle morphologies are consistent with the particle types of the present disclosure. For example, particles may be spherical, colloidal, square shaped, rods, wires, cones, pyramids, or oblong.
[00228] In some embodiments, the surface (e.g., particle) comprises any suitable material according to one skilled in the art. In some embodiments, the particle is magnetic, such as any magnetic material suitable according to one skilled in the art. In some embodiments, the particle
comprises a metal material. In some embodiments, the metal material comprises any one of or any combination of gold, silver, copper, nickel, cobalt, palladium, platinum, iridium, osmium, rhodium, ruthenium, rhenium, vanadium, chromium, manganese, niobium, molybdenum, tungsten, tantalum, iron and cadmium. In some embodiments, the particle comprises iron oxide. In some embodiments, the particle is a superparamagnetic iron oxide particle. In some embodiments, the particle comprises superparamagnetic iron oxide nanoparticles. In some embodiments, the particle has a core-shell structure. In some embodiments, the particle has an iron oxide core. In some embodiments, the particle comprises magnetite. In some embodiments, the particle comprises a silica shell. In some embodiments, the particle comprises an iron oxide core with a silica shell. In some instances, the silica shell can be functionalized with the tethering moieties or macromolecule chains provided elsewhere herein. In some embodiments, the particle comprises iron oxide crystals. In some embodiments, the particle comprises polystyrene. In some embodiments, the particle comprises iron oxide crystals embedded in a polystyrene core.
Systems
[00229] Provided herein are systems for performing any of the methods provided herein. In some embodiments, provided herein are systems for isolating one or more biomolecules from a biological solution. In some embodiments, provided herein are systems for isolating one or more proteins (e.g., peptides) from a biological solution.
[00230] In some embodiments, the systems provided herein comprise (a) one or more macromolecule structures (e.g., such as those described elsewhere herein), (b) a suspension solution, (c) a biological sample comprising one or more proteins, and (d) an automated system comprising a network of units with differentiated functions configured to perform any one of the methods provided herein. In some embodiments, the systems comprise one or more macromolecule structures. In some embodiments, the systems comprise a biological sample comprising one or more proteins. In some embodiments, the systems comprise an automated system comprising a network of units with differentiated functions configured to perform any one of the methods provided herein (e.g., such as isolating one or more biomolecules (e.g., proteins or peptides) from a biological solution).
[00231] In some embodiments, the macromolecule structure(s) as described elsewhere herein, the suspension solution, and the biological sample comprising a concentration of protein may be incubated at a temperature of about 10 degrees Celsius (°C) to about 100 °C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 20°C to about 90°C. In some embodiments, one or more components of the composition may be incubated at a temperature of about 20°C to about 50°C.
[00232] In some embodiments, the suspension solution may comprise Tris, EDTA, and CHAPS buffer. For example, the suspension solution may be Tris, EDTA in 150 millimolar (mM) KC1 and 0.05% CHAPS buffer. In another example, the suspension solution may be lOmM Tris HC1 pH 7.4, 1 mM EDTA.
[00233] In some embodiments, the (e.g., automated) system provided herein comprises a network of units which may comprise differentiated functions in isolating one or more biomolecules from a biological sample. In some embodiments, the network of units comprises (a) a first unit comprising a multichannel fluid transfer instrument for transferring fluids between units within the system, (b) a second unit comprising a support for storing a plurality of biological samples, and (c) a third unit comprising a support for an array plate possessing partitions that comprise the one or more macromolecule structure for binding of the one or more proteins within the biological sample with the one or more macromolecule structure. In some embodiments, the network of units comprises a unit comprises a multichannel fluid transfer instrument for transferring fluids between units within the system. In some embodiments, the network of units comprises a unit comprising a support for storing a plurality of biological samples. In some embodiments, the network of units comprises a unit comprising a support for an array plate possessing partitions that comprise the one or more macromolecule structure for binding of the one or more proteins within the biological sample with the one or more macromolecule structure. In some embodiments, the network of units may comprise a (e.g., fourth) unit comprising supports for storing a plurality of reagents. In some embodiments, the network of units may comprise a (e.g., fifth) unit comprising supports for storing a reagent to be disposed of. In some embodiments, the network of units may comprise a (e.g., sixth) unit comprising supports for storing consumables used by the multichannel fluid transfer instrument. In some embodiments, the system is programmed to perform a series of steps comprising contacting the biological sample with a specified partition of the array, optionally incubating the biological sample with the macromolecule structures contained within the partition of the array plate, removing components from a partition except the macromolecule structures and the one or more biomolecules (e.g., proteins or peptides) interacting with the macromolecule structure, and eluting the one or more biomolecules (e.g., proteins or peptides) from the macromolecule structures. In some embodiments, the system is further programmed to prepare the eluted (e.g., isolated) biomolecules (e.g., proteins or peptides) for analysis, such as by mass spectrometry or any other analysis as described elsewhere herein.
[00234] In some embodiments, the first unit comprises a degree of mobility that enables access to all other units within the system. In some embodiments, the first unit comprises a capacity to perform pipetting functions.
[00235] In some embodiments, the support of the second and/or third unit comprises support for a single plate, a 6 well plate, a 12 well plate, a 96 well plate, or a rack of microtubes. In some embodiments, the second and/or unit comprises a thermal unit capable of modulating the temperature of said support and a sample. In some embodiments, the second and/or third unit comprises a rotational unit capable of physically agitating and/or mixing a sample.
[00236] In some embodiments, the system comprises a step wherein the array plate is transferred to a (e.g., seventh) unit that comprises a magnetized support and a thermal unit capable of modulating the temperature of said support and a sample and incubated for an additional amount of time.
[00237] In some embodiments, the fourth unit comprises a set of reagents for: generating the array plate, washing the species of the biological sample that are not bound to the macromolecule structure(s), and/or preparing a sample for mass spectrometry. In some embodiments, contacting the biological sample with a specified partition of the array comprises pipetting a specified volume of the biological sample into the specific partition of the array.
[00238] In some embodiments, contacting the biological sample with a specified partition of the array comprises pipetting a volume of at least 10 microliters, at least 20 microliters at least 50 microliters, at least 100 microliters, at least 250 microliters, at least 500 microliters, or at least 1000 microliters of the biological sample into the specific partition of the array. In some embodiments, contacting the biological sample with a specified partition of the array comprises pipetting a volume of no more than 1000 microliters, no more than 500 microliters, nor more than 250 microliters, no more than 150 microliters, no more than 100 microliters, no more than 75 microliters, no more than 50 microliters, or no more than 30 microliters.
[00239] In some embodiments, incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the array plate comprises an incubation time of at least about 10 seconds, at least about 15 seconds, at least about 20 seconds, at least about 25 seconds, at least about 30 seconds, at least about 40 seconds, at least about 50 seconds, at least about 60 seconds, at least about 90 seconds, at least about 2 minutes, at least about 3 minutes, at least about 4 minutes, at least about 5 minutes, at least about 6 minutes, at least about 7 minutes, at least about 8 minutes, at least about 9 minutes, at least about 10 minutes, at least about 15 minutes, at least about 20 minutes, at least about 25 minutes, at least about 30 minutes, at least about 45 minutes, at least about 50 minutes, at least about 60 minutes, at least about 90 minutes, at least about 2 hours, at least about 3 hours, at least about 4 hours, at least about 5 hours, at least about 6 hours, at least about 7 hours, at least about 8 hours, at least about 9 hours, at least about 10 hours, at least about 12 hours, at least about 14 hours, at least about 15 hours, at least about 16 hours, at least about 17 hours, at least about 18 hours, at
least about 19 hours, at least about 20 hours, or at least about 24 hours. In some embodiments, incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the array plate comprises an incubation time of no more than 24 hours, no more than 12 hours, no more than 6 hours, no more than 3 hours, no more than 2 hours, no more than 90 minutes, no more than 75 minutes, no more than 60 minutes, no more than 45 minutes, or no more than 30 minutes. In some embodiments, incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the array plate comprises an incubation time of 30 minutes and 3 hours.
[00240] In some embodiments, incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the substrate comprises an incubation temperature between about 4° C to about 40° C. Incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the substrate may comprise an incubation temperature between about 4° C to about 37° C. Incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the substrate may comprise an incubation temperature between about 20° C to about 50° C. Incubating the biological sample (e.g., comprising biomolecules (e.g., proteins or peptides)) with the macromolecule structure(s) contained within the partition of the substrate may comprise an incubation temperature between about 4° C to about 100° C.
[00241] In some embodiments, the system is programmed to complete a series of wash steps. In some embodiments, the wash steps are completed with an organic solvent or an aqueous solvent, such as an organic solvent or an aqueous solvent as described elsewhere herein. In some embodiments, the wash steps are used to remove all components from a partition except the macromolecule structure(s) and the biomolecules (e.g., proteins or peptides) interacting with the macromolecule structure(s).
[00242] In some embodiments, the second unit can facilitate a transfer of the sample for mass spectrometry to a mass spectrometry unit.
[00243] In some embodiments, the system provide herein is an automated system.
Kits
[00244] Provided herein, in some embodiments, are kits for performing any one of the methods provided herein.
[00245] In some embodiments, provided herein are kits for isolating one or more biomolecules (e.g., proteins or peptides) from a biological solution. In some embodiments, provided herein are
kits for isolating one or more proteins from a biological solution. In some embodiments, provided herein are kits for isolating one or more peptides from a biological solution.
[00246] In some embodiments, the kits provided herein comprise (a) one or more macromolecule structure(s), (b) an organic solvent configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure, (c) a washing agent configured to wash proteins bound to the macromolecule structure, and (d) an elution agent configured to elute proteins from the macromolecule structure.
[00247] In some embodiments, the kits provided herein comprise (a) a first macromolecule structure and optionally a second macromolecule structure, (b) an organic solvent configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure(s), (c) one or more washing agents configured to wash proteins bound to the first macromolecule structure and optionally the second macromolecule structure, (d) an elution agent configured to elute proteins from the first macromolecule structure, (e) a reducing agent, and (f) a protein digesting agent.
[00248] In some embodiments, the kits provided herein comprise one or more macromolecule structure(s), such as any of the macromolecule structures provided elsewhere herein.
[00249] In some embodiments, the kits provided herein comprise an organic solvent, such as an organic solvent as provided elsewhere herein. In some embodiments, an organic solvent is configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure.
[00250] In some embodiments, the kits provided herein comprise a washing agent, such as an organic solvent (or wash solvent) as provided elsewhere herein. In some embodiments, the washing agent is configured to wash proteins bound to the macromolecule structure.
[00251] In some embodiments, the kits provided herein comprise an elution agent, such as an aqueous solvent as provided elsewhere herein. In some embodiments, an elution agent is configured to elute proteins from the macromolecule structure.
[00252] In some embodiments, the kits provided herein comprise a protein digesting agent. In some embodiments, a protein digesting agent produces one or more peptides. In some embodiments, a protein digesting agent comprises trypsin, lysin, serine protease. In some embodiments, a protein digesting agent comprises trypsin.
[00253] In some embodiments, the kits provided herein comprise a reducing agent. In some embodiments, the reducing agent comprises TCEP, dithiothreitol, beta-mercaptoethanol, glutathione, cysteine, or any combination thereof.
EMBODIMENTS
[00254] The following are exemplary embodiments of the disclosure herein:
Embodiment 1. A method of isolating one or more biomolecules from a biological sample, the method comprising: a. contacting a population of biomolecules comprising one or more proteins in the biological sample with a macromolecule structure in the presence of an organic solvent to bind the one or more proteins to the macromolecule structure; and b. eluting the one or more proteins from the macromolecule structure, thereby providing the one or more isolated biomolecules, wherein, the macromolecule structures comprise a surface and a macromolecule comprising recurring units of a first component and a second component, wherein the first component comprises a structure of Component (A) and the second component comprises a structure of Component (B’):
wherein each of Yi, Y2, and Y3 is independently selected from hydrogen or Ci-Ce alkyl; each of Xi, X2, and X3 is independently selected from hydrogen or Ci-Ce alkyl;
Ri is hydrogen, optionally substituted succinate, Ci-Ce alkyl substituted with sulfone, or phthalate;
R2 is C1-C12 alkyl substituted with amine, Ci-Ce alkyl substituted with hydroxy, optionally substituted aryl, optionally substituted heteroaryl, optionally substituted C3-C6 heterocycloalkyl, two or more fused 3-6 member rings; optionally substituted C3-C6 dicycloalkyl methane, Ci-Ce alkyl substituted with guanidine, Ci-Ce alkoxy, optionally substituted Ci-Ce alkyl substituted with disulfide, Ci-Ce alkyl substituted with thiol, optionally substituted succinate,
optionally substituted Ci-Ce alkylamine, Ci-Ce alkyl acetamide, Ci-Ce aminophthalate, a boronic acid, Ci-Cn optionally substituted cycloalkyl, or a monosaccharide; or
Ri and R.2 are taken together with the nitrogen to which they are attached to form an optionally substituted heterocycle; or
Ri and R.2 are taken together with the nitrogen to which they are attached to form an azide group;
R4 is hydrogen or Ci-Ce alkyl substituted with thiol;
Rs is succinate, optionally substituted aryl, or optionally substituted Ci-Ce alkyl substituted with disulfide;
q is an integer between 1 and 6; and p is an integer between 1 and 20.
Embodiment 5. The method of any one of the previous embodiments, wherein R1 is hydrogen.
Embodiment 6. The method of any one of embodiments 1-5, wherein Ri is -
(CH2)3SOOOH.
Embodiment 7. The method of any one of embodiments 1-5, wherein R2 is Ci-Ce alkoxy, Ci-12 alkyl substituted with amine, optionally substituted Ci-Ce alkylamine, -(CH2)I-6 imidazole -, (CH2)I-6 pyridine, optionally substituted aryl, -(CH2)I- 6N(CH3)2(CH2CH2CH2SOOOH), -(CH2)I-6N(CH2CH2CH2SOOOH)2, or C2-C6 alkyl substituted with hydroxy.
Embodiment 8. A method of isolating one or more biomolecules from a biological sample, the method comprising: a. contacting a population of biomolecules comprising one or more proteins in the biological sample with a macromolecule structure in the presence of an organic solvent to bind the one or more proteins to the macromolecule structure; and b. eluting the one or more proteins from the macromolecule structure, thereby providing the one or more isolated biomolecules, wherein, the macromolecule structure comprises:
(I) a surface;
(II) a tethering moiety coupled to the surface; and
(III) a macromolecule chain, wherein a first end of the macromolecule chain is covalently attached to the tethering moiety, and wherein the macromolecule chain comprises one or more distinct recurring units derived from a monomer represented by a structure selected from the group consisting of:
Q is -CH2- or ethylene glycol;
A is a polymeric side chain comprising a recurring unit derived from a monomer represented by the structure:
m is an integer selected from 1-20;
—is a single or double bond; each of R1, R2, R1 , R2 , and R3 is independently selected from hydrogen or -Ci-Ce alkyl;
R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl;
R4 is absent, hydrogen, sulfonate, carboxylate, C1-C4 alkylene, or Ci-Ce alkyl optionally substituted with halogen;
R5 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, - C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne;
R6 is hydrogen or Ci-Ce alkyl,
R7 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered
heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, - C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne;
R8 is Ci-Ce alkyl, divalent metal, or symmetric or asymmetric disulfide;
R9 is hydrogen or oxo; and n1 is an integer selected from 1-100.
Embodiment 9. The method of embodiment 8, wherein the macromolecule chain comprises the recurring unit derived from a monomer represented by the structure:
Embodiment 10. The method of embodiment 8 or 9, wherein R1 and R1 is hydrogen.
Embodiment 11. The method of any one of embodiments 8-10, wherein R2 and R2 is hydrogen.
Embodiment 12. The method of any one of embodiments 8-11, wherein R3 and R3 hydrogen or Ci-Ce alkyl.
Embodiment 13. The method of any one of embodiments 8-11, wherein R3 and R3 are hydrogen.
Embodiment 14. The method of any one of embodiments 8-13, wherein Z is O.
Embodiment 15. The method of any one of embodiments 8-13, wherein Z is NH.
Embodiment 16. The method of any one of embodiments 8-15, wherein R8 is Ci-Ce alkyl.
Embodiment 17. The method of any one of embodiments 8-16, wherein R8 is Ci alkyl.
Embodiment 18. The method of embodiment 8, wherein the macromolecule chain comprises the recurring unit derived from a monomer represented by the structure:
Embodiment 19. The method of embodiment 18, wherein R1 is hydrogen.
Embodiment 20. The method of embodiment 18 or 19, wherein R2 is hydrogen.
Embodiment 21. The method of any one of embodiments 18-20, wherein R3 is Ci-Ce alkyl.
Embodiment 22. The method of any one of embodiments 18-21, wherein R3 is Ci alkyl.
Embodiment 23. The method of any one of embodiments 18-22, wherein Q is CEE.
Embodiment 24. The method of any one of embodiments 18-23, wherein m is 2.
Embodiment 25. The method of any one of embodiments 18-24, wherein R6 is Ci-Ce alkyl.
Embodiment 26. The method of any one of embodiments 18-25, wherein R6 is Ci alkyl.
Embodiment 27. The method of any one of embodiments 18-26, wherein R5 is Ci-Cni ethylene glycol.
Embodiment 28. The method of embodiment 27, wherein nl is from 1 to 25.
Embodiment 29. The method of embodiment 28, wherein nl is 9.
Embodiment 30. The method of embodiment 8, wherein the macromolecule chain comprises the recurring unit derived from a monomer represented by the structure:
Embodiment 31. The method of embodiment 30, wherein R1 is hydrogen.
Embodiment 32. The method of embodiment 30 or 31, wherein R2 is hydrogen.
Embodiment 33. The method of any one of embodiments 30-32, wherein R3 is Ci-Ce alkyl.
Embodiment 34. The method of any one of embodiments 30-33, wherein R3 is Ci alkyl.
Embodiment 35. The method of any one of embodiments 30-34, wherein R5 is Ci-Cni ethylene glycol.
Embodiment 36. The method of embodiment 35, wherein nl is from 1 to 10.
Embodiment 37. The method of any one of embodiments 30-34, wherein R5 is a 3-, 5-, or
6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo.
Embodiment 38. The method of any one of embodiments 30-34, wherein R5 is Ci-Cs alkyl substituted with one or more hydroxyl, amine, or azide.
Embodiment 39. The method of embodiment 38, wherein R5 is C3 alkyl substituted with a hydroxyl and an azide.
Embodiment 40. The method of embodiment 38, wherein R5 is C3 alkyl substituted with a hydroxyl and an amine.
Embodiment 41. The method of embodiment 8, wherein the macromolecule chain comprises the recurring unit derived from a monomer represented by the structure:
Embodiment 42. The method of embodiment 41, wherein R1 is hydrogen.
Embodiment 43. The method of embodiment 41 or 42, wherein R2 is hydrogen.
Embodiment 44. The method of any one of embodiments 41-43, wherein R3 is Ci-Ce alkyl.
Embodiment 45. The method of any one of embodiments 41-44, wherein R3 is Ci alkyl.
Embodiment 46. The method of any one of embodiments 41-45, wherein each of X and Y is -C-.
Embodiment 47. The method of any one of embodiments 41-46, wherein R4 is sulfonate.
Embodiment 48. The method of any one of embodiments 8-47, wherein the macromolecule chain comprises one distinct recurring unit.
Embodiment 49. The method of any one of embodiments 8-47, wherein the macromolecule chain comprises two distinct recurring units.
Embodiment 50. The method of any one of the previous embodiments, wherein the macromolecule structure is selected from Table 4.
Embodiment 51. The method of any one of the previous embodiments, wherein the macromolecule structure is selected from Table 1.
Embodiment 52. The method of any one of the embodiments 8-50, wherein the macromolecule structure comprises:
Embodiment 53. The method of any one of the previous embodiments, wherein the one or more proteins comprises peptides.
Embodiment 54. The method of any one of the previous embodiments, wherein the surface is a particle.
Embodiment 55. The method of any one of the previous embodiments, wherein the surface is a microparticle or a nanoparticle.
Embodiment 56. The method of any one of the previous embodiments, wherein the surface is a nanoparticle.
Embodiment 57. The method of any one of the previous embodiments, wherein the surface comprises iron oxide.
Embodiment 58. The method of any one of the previous embodiments, wherein the surface comprises magnetite.
Embodiment 59. The method of any one of the previous embodiments, wherein the surface comprises superparamagnetic iron oxide nanoparticle(s).
Embodiment 60. The method of any one of the previous embodiments, wherein the surface comprises an iron oxide core.
Embodiment 61. The method of any one of the previous embodiments, wherein the particle comprises an iron oxide core with a silica shell coating.
Embodiment 62. The method of any one of the previous embodiments, wherein the macromolecule structure has a size of at least 100 nm.
Embodiment 63. The method of any one of the previous embodiments, wherein the macromolecule structure has a size of about 100 nm to about 400 nm.
Embodiment 64. The method of any one of the previous embodiments, wherein the method further comprises separating the macromolecule structure from the one or more proteins after (a).
Embodiment 65. The method of embodiment 64, wherein separating is magnetic separation, centrifugation, or a combination thereof.
Embodiment 66. The method of any one of the previous embodiments, wherein before (b) the method further comprises digesting one or more proteins bound to the macromolecule structure to form one or more peptides.
Embodiment 67. The method of any one of the previous embodiments, wherein eluting the one or more proteins comprises eluting the one or more peptides.
Embodiment 68. The method of any one of the previous embodiments, wherein the one or more isolated biomolecules comprises one or more isolated proteins.
Embodiment 69. The method of any one of the previous embodiments, wherein the one or more isolated biomolecules comprises one or more isolated peptides.
Embodiment 70. The method of any one of the previous embodiments, wherein after (a), the method further comprises washing the macromolecule structure comprising the one or more proteins with an organic solvent wash.
Embodiment 71. The method of embodiment 70, wherein the organic solvent wash comprises at least 50 wt% (e.g., 80 wt%, 90 wt%, 95 wt%) of an organic solvent.
Embodiment 72. The method of any one of the previous embodiments, wherein the organic solvent comprises an alcohol, acetonitrile, dichloromethane, dimethylformamide (DMF), dimethylsulfoxide (DMSO), ethylacetate, hexamethylphosphoramide (HPMA), or tetrahydrofuran.
Embodiment 73. The method of any one of the previous embodiments, wherein the organic solvent comprises acetonitrile.
Embodiment 74. The method of embodiment 72, wherein the alcohol is ethanol, n- propanol, isopropanol, n-butanol, sec-butanol, tert-butanol, or pentanol.
Embodiment 75. The method of any one of the previous embodiments, wherein (b) comprises eluting the one or more proteins (e.g., or peptides) from the macromolecule structure with an aqueous solution.
Embodiment 76. The method of embodiment 75, wherein the aqueous solution comprises an organic solvent.
Embodiment 77. The method of embodiment 75 or 76, wherein the aqueous solution comprises no more than 50 wt% (e.g., 40 wt%, 30 wt%, 20 wt%, 10 wt%, 5 wt%) of an organic solvent.
Embodiment 78. The method of any one of the previous embodiments, wherein the biological sample is plasma, serum, CSF, urine, tear, cell lysate, tissue lysate, cell homogenate, tissue homogenate, nipple aspirate, fecal matter, synovial fluid, saliva or blood.
Embodiment 79. The method of any one of the previous embodiments, where the method is capable of isolating from 100 to 20,000 (e.g., unique) peptides.
Embodiment 80. The method of any one of embodiments 1-79, wherein the method is capable of isolating at least 1,000 (e.g., unique) peptides.
Embodiment 81. The method of any one of embodiments 1-79, wherein the method is capable of isolating no more than 25,000 (e.g., unique) peptides.
Embodiment 82. The method of any one of the previous embodiments, wherein isolating occurs in absence of a precipitation solution comprising salts in an amount effective to precipitate the population of biomolecules.
Embodiment 83. The method of any one of the previous embodiments, wherein the method further comprises identifying the one or more peptides.
Embodiment 84. The method of any one of the previous embodiments, wherein at least 90% of an amount of a surfactant is removed from the population of biomolecules.
Embodiment 85. The method of any one of the previous embodiments, wherein at least 90% of an amount of a lysis reagent is removed from the population of biomolecules.
Embodiment 86. The method of any one of the previous embodiments, wherein the method further comprises after (b), analyzing the one or more proteins.
Embodiment 87. The method of embodiment 86, wherein the analyzing comprises mass spectrometry (MS), liquid chromatography-mass spectrometry (LC-MS), protein sequencing, or a combination thereof.
Embodiment 88. The method of embodiment 86 or 87, wherein analyzing is capable of identifying from 1 to 20,000 protein groups (e.g., from 1,000 to 10,000 protein groups, from 1,000 to 5,000 protein groups, from 1,200 to 2,200 protein groups).
Embodiment 89. The method of any one of the previous embodiments, wherein before (a) the method further comprises contacting the biological sample comprising a population of proteins with an additional macromolecule structure to provide a protein corona.
Embodiment 90. The method of any one of the previous embodiments, further comprising before (a) treating or digesting one or more proteins of the population of proteins to form one or more peptides or desorbed proteins.
Embodiment 91. The method of embodiment 89 or 90, wherein the macromolecule structure and the additional macromolecule structure are the same.
Embodiment 92. The method of embodiment 89 or 90, wherein the macromolecule structure and the additional macromolecule structure are different.
Embodiment 93. The method of any one of the previous embodiments, wherein a mass of the one or more isolated proteins is at least 100 nanograms (ng) (e.g., 500 ng, 1 microgram (pg), 10 pg, 50 pg, 100 pg, 200 pg).
Embodiment 94. A method of isolating one or more biomolecules from a biological sample, the method comprising: a. contacting the biomolecules comprising one or more proteins with a first macromolecule structure to provide a protein corona; b. optionally, digesting the biomolecules of the protein corona to form one or more peptides; c. separating the one or more peptides of the protein corona from the first macromolecule structure; d. contacting the one or more peptides with a second macromolecule structure and an organic solvent to bind at least a portion of the one or more peptides to the second macromolecule structure; and e. eluting the one or more peptides from the second macromolecule structure, thereby providing the one or more isolated biomolecules.
Embodiment 95. The method of embodiment 94, wherein the first surface and the second macromolecule structure are different.
Embodiment 96. The method of embodiment 94, wherein the first surface and the second macromolecule structure are the same.
Embodiment 97. The method of any one of embodiments 94-96, wherein in (d) the second macromolecule structure is added to a composition comprising the first surface.
Embodiment 98. The method of any one of embodiments 94-97, wherein the first macromolecule structure or the second macromolecule structure comprises a PEG group.
Embodiment 99. The method of any one of embodiments 94-98, wherein the surface is a particle.
Embodiment 100. The method of any one of embodiments 94-99, wherein the surface is a microparticle or a nanoparticle.
Embodiment 101. The method of any one of embodiments 94-100, wherein the surface comprises iron oxide.
Embodiment 102. The method of any one of embodiments 94-101, wherein the surface comprises magnetite.
Embodiment 103. The method of any one of embodiments 94-102, wherein the surface comprises superparamagnetic iron oxide nanoparticle(s).
Embodiment 104. The method of any one of embodiments 94-103, wherein the method further comprises removing the one or more isolated proteins after (e).
Embodiment 105. The method of any one of embodiments 94-104, wherein separating comprises magnetic separation, centrifugation, or a combination thereof.
Embodiment 106. The method of any one of embodiments 94-105, wherein the organic solvent comprises an alcohol, acetonitrile, dichloromethane, dimethylformamide (DMF), dimethylsulfoxide (DMSO), ethylacetate, hexamethylphosphoramide (HPMA), or tetrahydrofuran.
Embodiment 107. The method of any one of embodiments 94-106, wherein (d) comprises eluting the one or more proteins or the one or more peptides from the second surface with an aqueous solution.
Embodiment 108. The method of any one of embodiments 94-107, wherein the biological sample is plasma, serum, CSF, urine, tear, cell lysate, tissue lysate, cell homogenate, tissue homogenate, nipple aspirate, fecal matter, synovial fluid, saliva or blood.
Embodiment 109. The method of any one of embodiments 94-108, where the method is capable of isolating at least 100 (e.g., unique) protein groups or peptides.
Embodiment 110. The method of any one of embodiments 94-109, wherein isolation occurs in absence of a precipitation solution comprising salts in an amount effective to precipitate the one or more proteins or one or more peptides.
Embodiment 111. The method of any one of embodiments 94-110, wherein the method further comprises identifying the one or more proteins or one or more peptides.
Embodiment 112. The method of any one of embodiments 94-111, wherein a mass of the one or more isolated proteins or one or more isolated peptides is at least 100 nanograms (ng) (e.g., 500 ng, 1 microgram (pg), 10 pg, 50 pg, 100 pg, 200 pg).
Embodiment 113. The method of any one of embodiments 1-112, wherein eluting comprises the successive application of two or more eluents.
Embodiment 114. The method of embodiment 113, wherein the two or more eluents comprise eluents of varying polarity.
Embodiment 115. A system for performing the method of any one of embodiments 1-114, the system comprising: a. one or more macromolecule structure; b. a suspension solution; c. a biological sample comprising one or more proteins; and
d. an automated system comprising a network of units with differentiated functions configured to perform the method of any one of embodiments 1-114.
Embodiment 116. The system of embodiment 115, wherein the network of units comprises: a. a first unit comprising a multichannel fluid transfer instrument for transferring fluids between units within the system; b. a second unit comprising a support for storing a plurality of biological samples; c. a third unit comprising a support for an array plate possessing partitions that comprise the one or more macromolecule structure for binding of the one or more proteins within the biological sample with the one or more macromolecule structures.
Embodiment 117. The system of embodiment 115 or 116, wherein the network of units further comprises a fourth unit comprising supports for storing a plurality of reagents.
Embodiment 118. The system of any one of embodiments 115-117, wherein the network of units further comprises a fifth unit comprising supports for storing a reagent to be disposed of.
Embodiment 119. The system of any one of embodiments 115-118, wherein the network of units further comprises supports for storing consumables used by a multichannel fluid transfer instrument.
Embodiment 120. A kit for performing the method of any one of embodiments 1 to 114 comprising: a. one or more macromolecule structures; b. an organic solvent configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure; c. a washing agent configured to wash proteins bound to the macromolecule structure; and d. an elution agent configured to elute proteins from the macromolecule structure.
EXAMPLES
[00255] The following examples are provided to further illustrate some embodiments of the present disclosure but are not intended to limit the scope of the disclosure; it will be understood by their exemplary nature that other procedures, methodologies, or techniques known to those skilled in the art may alternatively be used.
Example 1: Synthesis of SPION@SiOi- APTES
[00256] Silica coated super paramagnetic iron oxide nanoparticles (SPION@SiO2, 002 or 003) were suspended in DMF and sonicated for 15 minutes. The nanoparticle solution was purged with N2 gas for 20 minutes and the particle solution was heated to 120°C for 4 hours. After cooling, the SPION@SiO2 were washed with DMF 3 times, then isolated. The isolated particles were dispersed in N,N-dimethylacetamide (DMAc) (2 L). To the solution, 4.50 g of (3- aminopropyltri ethoxy silane) (APTES) was added and the resulting solution was reacted at 120°C for 4 hours to obtain the SPION@SiO2- APTES product.
[00257] SPION@SiO2-APTES in DMF was washed two times with tetrahydrofuran (THF) then resuspended in THF (801 mL) with sonication for 15 minutes. In an ice bath (0°C), under N2, triethylamine (10.40 g) was added. After addition of the tri ethyl amine, 2-bromoisobutyryl bromide (2.509 g) was added dropwise at 0°C. The reaction was stirred overnight at room temperature (16 hours). The resulting material was washed 1 time with THF, 1 time with ethanol, 1 time with water, and then 1 time with THF. The final product was dried with N2.
[00258] Initiator particle SPION@SiO2-APTES-Br (1.000 g), CuBr2 (0.010 g), N,N,N’ ,N” ,N” - pentamethyldiethylenetriamine (PMDETA) (0.240 g), monomer Oligo(ethylene glycol) Methyl Ether Methacrylate (OEGMA, MW500) (5.000 g) and 25 mL of dimethylformamide (DMF) were added in a 3-neck flask. The mixture was sonicated for 15 min and purged with N2 for another 15 min. Separately, 0.500 g of L-ascorbic acid was dissolved in 10 mL of DMF and purged with N2
for at least 15 minutes. The mixture containing the monomer and nanoparticle suspension was heated to 35°C under N2 and the L-ascorbic acid solution was added via syringe pump at 0.05 mL/min. The reaction mixture was maintained at 35°C for 16 hours to obtain Compound 56.
Example 4: Characterization of Macromolecule Structures
[00259] Macromolecule structures 55-57 were synthesized according to modified versions of the syntheses described in Examples 1-3 and the scheme depicted in FIG. 3. The particle size was characterized by DLS, surface charge by zeta potential, and macromolecule chain% by TGA (weight loss%). The particle sizes were compared to the particle sizes before functionalization with the macromolecule chains. The resulting data is found in Table 5.
[00260] Macromolecule structures 55, 56, and 59-61 were analyzed by thermogravimetric analysis for the % organics. The results of which are in Table 6. The % organics in the particles ranged from 9.14% to 15.88%, increased from the % organics found on the initiator surface before further functionalization.
[00261] Provided herein Table 7 is performance data for nanoparticles containing the surfaces disclosed in Table 1. Briefly, the nanoparticles were incubated with plasma samples at 37 °C for 1 hour to form protein coronas and then washed three times. The proteins were then lysed, alkylated, and digested before analysis using liquid chromatography tandem mass spectrometry.
[00262] NP-1 through NP-5 are nanoparticles found in commercially available Proteograph™ VI.2 kit. The Jaccard Index (JI) is a measure of similarity between in the protein groups (PG) identified between different nanoparticles.
Example 6: Synthesis of P(GMA-co-D VB)-coated particle
[00263] In a 4-neck round bottom flask, vinyl-containing silica coated nanoparticles (2 g) are added and dispersed in acetonitrile (400 mL). The mixture of the round bottom flask is mixed or sonicated at about 200 rpm for 20 minutes. In a separate vessel, purified glycidyl methacrylate (GMA) (3.000 g) divinyl benzene (DVB) (3.000 g), and acetonitrile (5mL) are combined. The round bottom flask is cooled to 10 degrees Celsius while stirring. Using a plastic syringe, the mixture of GMA and DVB is added to the round bottom flask under an inert atmosphere of nitrogen. In a separate vial, azobi si sobutyronitrile (AIBN) (450 mg) is dissolved in acetonitrile (4 mL). The round bottom flask is heated to 80 degrees Celsius, and once the flask reaches temperature, the AIBN solution is injected into the flask. The reaction suspension reacts at 80 degrees Celsius for 60 minutes. The reaction is monitored every 30 minutes until the particles have formed a size of about 350 nm and PDI of < 0.2. The reaction is quenched by adding benzoquinone (0.108 g) in acetonitrile (5 mL). Once the reaction is complete, the flask is removed from the heat and the particles are purified using THF.
Example 7: Synthesis of Macromolecule Structure 1 (P(GMA-co-DVB)-EDA Particle
[00264] In a 4-neck round bottom flask, P(GMA-co-DVB)-coated particle (see Example 6) (1.10g) and DMF (220mL) are added. The mixture is mixed or sonicated at about 200 rpm for 20 minutes. The round bottom flask is cooled to 10 degrees Celsius while stirring under inert atmosphere, after which the flask is heated to 80 degrees Celsius. When the flask has reached 80 degrees Celsius, the ethylene diamine (6.86g) is injected into the flask using a syringe. The mixture is reacted for 16 hours. Once the reaction is complete, the flask is removed from the heat and the particles are purified using DMF
Example 8: Synthesis of Macromolecule Structure 15 (P(GMA-co-DVB)-EDA-PA Particle [00265] In a 4-neck round bottom flask, P(GMA-co-DVB)-EDA-PA particles (i.Og) (see Example 7), N,N-dimethylformamide (DMF) (190mL), and triethylamine (TEA) (1.44g) are added. The mixture is mixed or sonicated at about 200 rpm for 20 minutes. In a separate vessel, phthalic anhydride (L50g) in DMF (lOmL) are combined. The round bottom flask is cooled to 10 degrees Celsius while stirring under inert atmosphere, after which the flask is heated to 80 degrees Celsius. When the flask has reached 60 degrees Celsius, the phthalic anhydride solution is injected into the flask using a syringe. The mixture is reacted for 16 hours. Once the reaction is complete, the flask is removed from the heat and the particles are purified using ethanol and DMF.
Example 9: Clean Up Method
[00266] As exemplified in FIG. 7, a biological sample containing proteins and/or peptides is incubated in a well plate with a macromolecule structure (e.g., macromolecular structure 50 or 51 depicted in Table 1) to form a protein corona. Using a magnet, the protein corona is pulled down in the well plate while the supernatant is removed. The protein corona is then washed and resuspended in the well plate. The protein corona is then subjected to treatment to digest the proteins (or peptides) of the protein corona into peptides as described elsewhere herein. The peptides are then subjected to purification prior to isolating for analysis through mass spectrometry.
[00267] Peptide purification is achieved through adding the digested proteins and peptides in a vessel (e.g., centrifuge tube) with a different macromolecule structure (e.g., macromolecule structures 425, 441, 55, and 56) and 100% acetonitrile. The biomolecule and macromolecule structures are incubated for 8 minutes to form a biomolecule corona (comprising peptides) and then centrifuged at 600 rotations per minute. The biomolecule corona is pulled down in the vessel using a magnet, and the resulting supernatant is removed and the contents of the vessel are washed with 100% acetonitrile before another round of magnetic pulldown and removing the supernatant. The contents of the vessel (e.g., biomolecule corona) are combined with a 3% solution of acetonitrile in water and incubated for 10 minutes at 28 degrees Celsius, thereby releasing the peptides from the macromolecule structures. The macromolecule structures are magnetically pulled down in the vessel and the purified, isolated peptides remain in the supernatant. The isolated peptides are collected and used downstream for analysis (e.g., mass spectrometry or LC-MS).
Example 10: Particle Characterization for Use in Peptide Clean Up
[00268] Macromolecule structures 55 and 56 were synthesized according to schemes illustrated in FIG. 3 and FIG. 4, respectively. The size characterization of these particles is shown in Table
8.
[00269] Table 9a shows the results weight loss (%) for Macromolecule Structures 55-57 after 1 day at 40° Celsius in various solvent conditions as a proxy for stability of the particles. Similarly, Table 9b shows the results weight loss (%) for Macromolecule Structures 55-57 after 8 days at 40° Celsius in various solvent conditions as a proxy for stability of the particles. In comparison to the initial size and weight loss (%) of the particles, each of the tested particles were found to be stable after 1 day and after 8 days (Table 9c).
Example 11: Performance of macromolecule structures vs. commercially available beads [00270] FIG. 8 illustrates peptide yield (in micrograms) for various macromolecule structures, where macromolecular structures of the disclosure yielding a mass of peptides between about 1 microgram to about 2 micrograms include macromolecule structures as described herein, such as described in Table 1 and Table 4. The macromolecular structures of the disclosure have improved peptide yield compared to commercially available beads, MPE and Cytva beads. For example, MPE, as a reference bead, was also found to yield between about 1 microgram to about 2 micrograms of peptide.
[00271] FIG. 9a illustrates an example of a quantity of unique peptides identified for various macromolecule structures in comparison to reference surfaces (e.g., Monitored Multi-Flow, Positive Pressure, Evaporative Extraction module (MPE) used in the commercially available PROTEOGRAPH method or commercially available Cytiva beads). FIG. 9b illustrates an example of a quantity of unique protein groups identified for various macromolecule structures in comparison to reference surfaces (e.g., MPE or commercially available Cytiva beads). A correlation exists in performance of the beads in both quantity of unique peptides identified and quantity unique protein groups identified. For example, commercially available Cytiva beads identify a lower amount of unique peptide groups and a lower amount of unique protein groups in comparison to the macromolecule structures of the disclosure.
[00272] FIG. 10 illustrates an example of a difference in peptide identification rate (peptides/minute) between a sample obtained using the methods disclosed herein and a macromolecule structure of the disclosure (e.g., Macromolecule Structure 55) and a sample obtained using MPE. The peptides are analyzed by liquid chromatography mass spectrometry and the number of peptides identified over the retention time is shown. In some instances, the rate of identification between a macromolecule of the disclosure and MPE is comparable over time.
[00273] FIG. Ila illustrates peptide yield (in micrograms) in a solution with beads compared to a solution with 5x the concentration of beads. In some instances, a higher concentration of beads results in higher peptide yield across various macromolecule structures. FIG. 11b illustrates the number of unique peptides identified in a solution with beads compared to a solution with 5x the
concentration of beads. In some instances, a higher concentration of beads results in a higher count of peptides across various macromolecule structures.
Example 12: Peptide clean-up workflow
[00274] The general procedures of Examples 10 and 11 are followed with modification.
[00275] Particle Prep. First, beads are distributed into one or more containers, such as tubes. Any number of tubes may be used to contain beads (e.g., 2-10 tubes, each containing 4-20 beads per tube). The mass of beads per tube in some instances comprises 0.2-5 mg per bead. Each tube is filled with a solvent such as water to a volume of 100-1000 microliters before transfer to a plate, and one or more washings. Beads may then be resuspended to a concentration, such as 10-100 mg/mL and at a volume of 1-100 microliters.
[00276] Binding. Digested peptide samples are added to the wells of the plate (10-2000 microliters). A binding reagent is then added to each well, in a volume of 0.2-3000 mL, the wells incubated for 5-20 minutes, and beads resuspended in binding reagent comprising an organic solvent. The binding reagent is then removed.
[00277] Elution. An elution buffer is added (50-500 microliters) comprising 1-10% organic solvent in water, and then beads are incubated 5-20 minutes at 20-35 degrees C. After centrifugation (500-2000 rpm) and a 1-5 minute pull down, eluted peptides are transferred to a new plate.
[00278] While preferred embodiments of the present disclosure have been shown and described herein, it will be obvious to those skilled in the art that such embodiments are provided by way of example only. Numerous variations, changes, and substitutions will now occur to those skilled in the art without departing from the disclosure. It should be understood that various alternatives to the embodiments of the present disclosure may be employed in practicing the present disclosure. It is intended that the following claims define the scope of the present disclosure and that methods and structures within the scope of these claims and their equivalents be covered thereby. The section headings used herein are for organizational purposes only and are not to be construed as limiting the subject matter described.
Claims
1. A method of isolating one or more biomolecules from a biological sample, the method comprising:
(a) contacting a population of biomolecules comprising one or more proteins in the biological sample with a macromolecule structure in the presence of an organic solvent to bind the one or more proteins to the macromolecule structure; and
(b) eluting the one or more proteins from the macromolecule structure, thereby providing the one or more isolated biomolecules, wherein, the macromolecule structures comprise a surface and a macromolecule comprising recurring units of a first component and a second component, wherein the first component comprises a structure of Component (A) and the second component comprises a structure of Component (B’):
wherein each of Yi, Y2, and Y3 is independently selected from hydrogen or Ci-Ce alkyl; each of Xi, X2, and X3 is independently selected from hydrogen or Ci-Ce alkyl;
Ri is hydrogen, optionally substituted succinate, Ci-Ce alkyl substituted with sulfone, or phthalate;
R2 is C1-C12 alkyl substituted with amine, Ci-Ce alkyl substituted with hydroxy, optionally substituted aryl, optionally substituted heteroaryl, optionally substituted C3-C6 heterocycloalkyl, two or more fused 3-6 member rings; optionally substituted C3-C6 dicycloalkyl methane, Ci-Ce alkyl substituted with guanidine, Ci-Ce alkoxy, optionally substituted Ci-Ce alkyl substituted with disulfide, Ci-Ce alkyl substituted with thiol, optionally substituted succinate,
optionally substituted Ci-Ce alkylamine, Ci-Ce alkyl acetamide, Ci-Ce aminophthalate, a boronic acid, C1-C11 optionally substituted cycloalkyl, or a monosaccharide; or
Ri and R2 are taken together with the nitrogen to which they are attached to form an optionally substituted heterocycle; or
Ri and R2 are taken together with the nitrogen to which they are attached to form an azide group;
R4 is hydrogen or Ci-Ce alkyl substituted with thiol;
2. A method of isolating one or more biomolecules from a biological sample, the method comprising:
(a) contacting a population of biomolecules comprising one or more proteins in the biological sample with a macromolecule structure in the presence of an organic solvent to bind the one or more proteins to the macromolecule structure; and
(b) eluting the one or more proteins from the macromolecule structure, thereby providing the one or more isolated biomolecules, wherein, the macromolecule structure comprises:
(I) a surface;
(II) a tethering moiety coupled to the surface; and
(III) a macromolecule chain, wherein a first end of the macromolecule chain is covalently attached to the tethering moiety, and wherein the macromolecule chain comprises one or more distinct recurring units derived from a monomer represented by a structure selected from the group consisting of:
Q is -CH2- or ethylene glycol;
A is a polymeric side chain comprising a recurring unit derived from a monomer represented by the structure:
m is an integer selected from 1-20;
—is a single or double bond; each of R1, R2, R1 , R2 , and R3 is independently selected from hydrogen or -Ci-Ce alkyl;
R3 is hydrogen, Ci-Ce alkyl, or 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl;
R4 is absent, hydrogen, sulfonate, carboxylate, C1-C4 alkylene, or Ci-Ce alkyl optionally substituted with halogen;
R5 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, - C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne;
R6 is hydrogen or Ci-Ce alkyl,
R7 is hydrogen, Ci-Ce alkyl, Ci-Cs alkyl substituted with one or more hydroxyl, amine, azide, sulfonate, carbamate ester, asymmetrical disulfide, 3-, 5-, or 6-membered heterocycle optionally substituted with one or more Ci-Ce alkyl or oxo, or Ci-Cni ethylene glycol, Ci-Cs alkylamine further optionally substituted with amine or sulfonate, Ci-Cs alkoxy optionally substituted with one or more oxo or halogen, - C1-C3 alkyl optionally substituted with one or more pyrene, 2 or more fused 5-6 membered rings further optionally substituted, optionally substituted benzyl, trimethoxysilane, or phosphorocholine, or C1-C4 alkylyne;
R8 is Ci-Ce alkyl, divalent metal, or symmetric or asymmetric disulfide;
R9 is hydrogen or oxo; and n1 is an integer selected from 1-100.
3. The method of claim 2, wherein R5 is Ci-Cni ethylene glycol.
4. The method of claim 2 or 3, wherein nl is 9.
6. The method of any one of claims 2-5, wherein R3 is Ci-Ce alkyl.
7. The method of any one of claims 1-6, wherein the macromolecule structure is selected from Table 4 or Table 1.
10. The method of any one of claims 1-9, wherein the surface is a microparticle or a nanoparticle.
11. The method of any one of claims 1-10, wherein the surface is a nanoparticle.
12. The method of claim 10, wherein the microparticle or nanoparticle comprises a zeta potential of -10 to 10 mV.
13. The method of claim 10, wherein the microparticle or nanoparticle comprises an average diameter of 400 to 500 nm.
14. The method of any one of claims 1-13, wherein the method further comprises separating the macromolecule structure from the one or more proteins after (a).
15. The method of claim 12, wherein separating is magnetic separation, centrifugation, or a combination thereof.
16. The method of any one of the claims 1-13, wherein before (b) the method further comprises digesting one or more proteins bound to the macromolecule structure to form one or more peptides.
17. The method of any one of claims 1-15, wherein (b) comprises eluting the one or more proteins (e.g., or peptides) from the macromolecule structure with an aqueous solution.
18. The method of claim 15, wherein the aqueous solution comprises an organic solvent.
19. The method of any one of claims 1-16, wherein the organic solvent comprises acetonitrile.
20. A method of isolating one or more biomolecules from a biological sample, the method comprising:
(a) contacting the biomolecules comprising one or more proteins with a first macromolecule structure to provide a protein corona;
(b) optionally, digesting the biomolecules of the protein corona to form one or more peptides;
(c) separating the one or more peptides of the protein corona from the first macromolecule structure;
(d) contacting the one or more peptides with a second macromolecule structure and an organic solvent to bind at least a portion of the one or more peptides to the second macromolecule structure; and
(e) eluting the one or more peptides from the second macromolecule structure, thereby providing the one or more isolated biomolecules.
21. A system for performing the method of any one of claims 1-20, the system comprising:
(a) one or more macromolecule structure;
(b) a suspension solution;
(c) a biological sample comprising one or more proteins; and
(d) an automated system comprising a network of units with differentiated functions configured to perform the method of any one of claims 1-20.
22. A kit for performing the method of any one of claims 1-20 comprising:
(a) one or more macromolecule structures;
(b) an organic solvent configured to cause protein binding to the macromolecule structure when contacted with proteins and the macromolecule structure;
(c) a washing agent configured to wash proteins bound to the macromolecule structure; and
(d) an elution agent configured to elute proteins from the macromolecule structure.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363596934P | 2023-11-07 | 2023-11-07 | |
| US63/596,934 | 2023-11-07 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025101535A1 true WO2025101535A1 (en) | 2025-05-15 |
Family
ID=95696589
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/054605 Pending WO2025101535A1 (en) | 2023-11-07 | 2024-11-05 | Biomolecule cleanup methods, kits, and systems |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025101535A1 (en) |
Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20200041487A1 (en) * | 2003-05-22 | 2020-02-06 | Seer, Inc. | Systems and methods for discovery and analysis of markers |
| US20200124616A1 (en) * | 2014-09-10 | 2020-04-23 | Genentech, Inc. | Identification of immunogenic mutant peptides using genomic, transcriptomic and proteomic information |
| US20200284810A1 (en) * | 2017-09-05 | 2020-09-10 | Immunogen, Inc. | Methods for detection of folate receptor 1 in a patient sample |
| US20220082570A1 (en) * | 2020-09-10 | 2022-03-17 | Laboratory Corporation Of America Holdings | Methods and Systems for Detecting Prostaglandins by LC-MS/MS |
| US20220334123A1 (en) * | 2019-11-02 | 2022-10-20 | Seer, Inc. | Systems for protein corona analysis |
-
2024
- 2024-11-05 WO PCT/US2024/054605 patent/WO2025101535A1/en active Pending
Patent Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20200041487A1 (en) * | 2003-05-22 | 2020-02-06 | Seer, Inc. | Systems and methods for discovery and analysis of markers |
| US20200124616A1 (en) * | 2014-09-10 | 2020-04-23 | Genentech, Inc. | Identification of immunogenic mutant peptides using genomic, transcriptomic and proteomic information |
| US20200284810A1 (en) * | 2017-09-05 | 2020-09-10 | Immunogen, Inc. | Methods for detection of folate receptor 1 in a patient sample |
| US20220334123A1 (en) * | 2019-11-02 | 2022-10-20 | Seer, Inc. | Systems for protein corona analysis |
| US20220082570A1 (en) * | 2020-09-10 | 2022-03-17 | Laboratory Corporation Of America Holdings | Methods and Systems for Detecting Prostaglandins by LC-MS/MS |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| EP1649284B1 (en) | Process for preparing coated magnetic particles | |
| EP1750127B1 (en) | Method of affinity separation. | |
| JP3147389B2 (en) | Anion exchanger | |
| US20110257028A1 (en) | Method for selectively binding a substrate to sorbents by way of at least bivalent bonds | |
| SG174055A1 (en) | Sugar chain-capturing substance and use thereof | |
| Qin et al. | Rapid and selective extraction of multiple sulfonamides from aqueous samples based on Fe 3 O 4–chitosan molecularly imprinted polymers | |
| JP2014521078A (en) | Chromatographic composite materials | |
| JP6240965B2 (en) | Oriented loading system and method for orienting particles loaded in a well | |
| JP7765421B2 (en) | polymer particles | |
| JP2014029313A (en) | Chromatographic separation material | |
| Omer et al. | Preparation of QP4VP-b-LCP liquid crystal block copolymer and its application as a biosensor | |
| Wang et al. | Preparation of two zwitterionic polymer functionalized stationary phases and comparative evaluation under mixed-mode of reversed phase/hydrophilic interaction/ion exchange chromatography | |
| WO2025101535A1 (en) | Biomolecule cleanup methods, kits, and systems | |
| CN120513126A (en) | Bipedal silane-bound adsorbents for solid phase extraction and use thereof for oligonucleotide extraction | |
| AU2023356949A1 (en) | Functionalized macromolecules and systems, methods of synthesis, and uses thereof | |
| EP1735378A1 (en) | Photocrosslinked hydrogel blend surface coatings | |
| WO2025030029A1 (en) | Brush polymer macromolecule structures and methods for detecting biomolecules | |
| WO2004092731A1 (en) | Element having bioactive substance fixed thereto | |
| WO2025212750A1 (en) | Biomolecule preparation methods, kits, and systems | |
| JP6223347B2 (en) | Protein chromatography matrix with hydrophilic copolymer coating | |
| JP4929635B2 (en) | Maleimide group-containing porous crosslinked polystyrene particles and method for producing the same | |
| Zarei | Improved capillary electrophoresis through the use of smart materials | |
| CA2635896A1 (en) | Synthesis and use of cross-linked hydrophilic hollow spheres for encapsulating hydrophilic cargo | |
| JP2021515002A (en) | Bioconjugate of antibodies and functionalized magnetic nanoparticles | |
| CN101107525A (en) | Process for the preparation of coated polymer particles containing superparamagnetic crystals |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24889461 Country of ref document: EP Kind code of ref document: A1 |