WO2025083247A1 - Novel enzymes - Google Patents
Novel enzymes Download PDFInfo
- Publication number
- WO2025083247A1 WO2025083247A1 PCT/EP2024/079566 EP2024079566W WO2025083247A1 WO 2025083247 A1 WO2025083247 A1 WO 2025083247A1 EP 2024079566 W EP2024079566 W EP 2024079566W WO 2025083247 A1 WO2025083247 A1 WO 2025083247A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- amino acid
- dda
- substituted
- position corresponding
- acid position
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6869—Methods for sequencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y306/00—Hydrolases acting on acid anhydrides (3.6)
- C12Y306/04—Hydrolases acting on acid anhydrides (3.6) acting on acid anhydrides; involved in cellular and subcellular movement (3.6.4)
- C12Y306/04012—DNA helicase (3.6.4.12)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N11/00—Carrier-bound or immobilised enzymes; Carrier-bound or immobilised microbial cells; Preparation thereof
- C12N11/02—Enzymes or microbial cells immobilised on or in an organic carrier
Definitions
- the present invention relates to modified Dda helicases which can be used to control the movement of analytes such as polynucleotides.
- the modified Dda helicases are used in analyte detection and characterisation.
- Two of the essential components of analyte, especially polymer, characterization using nanopore sensing are (1) the control of polymer movement through the pore and (2) the discrimination of the composing building blocks as the polymer is moved through the pore.
- the narrowest part of the pore typically corresponds to the most discriminating part of the nanopore with respect to the change in measurement signal as a function of the analyte moving with respect to the nanopore.
- WO2015/055981, WO2015/166276, WO2016/055777, and PCT/EP2023/059821 incorporated by reference herein in their entirety, describe polynucleotide binding proteins, specifically Dda helicases, which can be used to control the movement of analytes with respect to a transmembrane protein pore such as the CsgG pores described herein.
- the inventors have surprisingly identified specific Dda mutants (known herein as modified Dda helicases or modified helicases) which have an improved ability to control the movement of an analyte through a pore.
- modified Dda helicases or modified helicases When sequencing a polynucleotide using a pore, the system jointly estimates the number and identity of bases/nucleotides passing through the pore. Better control over variability in the speed of movement can reduce one of the sources of statistical noise and simplify the estimation task. Runs of consecutive short dwells of a polynucleotide in the pore may trigger a failure to call the underlying nucleotides/bases resulting in a deletion error. Unusually long dwells may lead to insertion errors.
- Ensuring that each nucleotide/base spends a sufficient time interval in the pore is helpful for resolving statistical uncertainty in the nucleotide/base identity from noisy signal levels. Further information can be extracted from dependence of dwell times on nucleotide/base identities, for example via interactions with the motor enzyme. Reducing the overall variability in dwell times can help to extract more precise information through this channel. During regions in which signal levels provide limited information about movement (e.g., long homopolymer regions) multi-nucleotide/base dwell times can be used to infer the number of bases traversing the pore. Reducing variability in dwell times can make these inferences more precise.
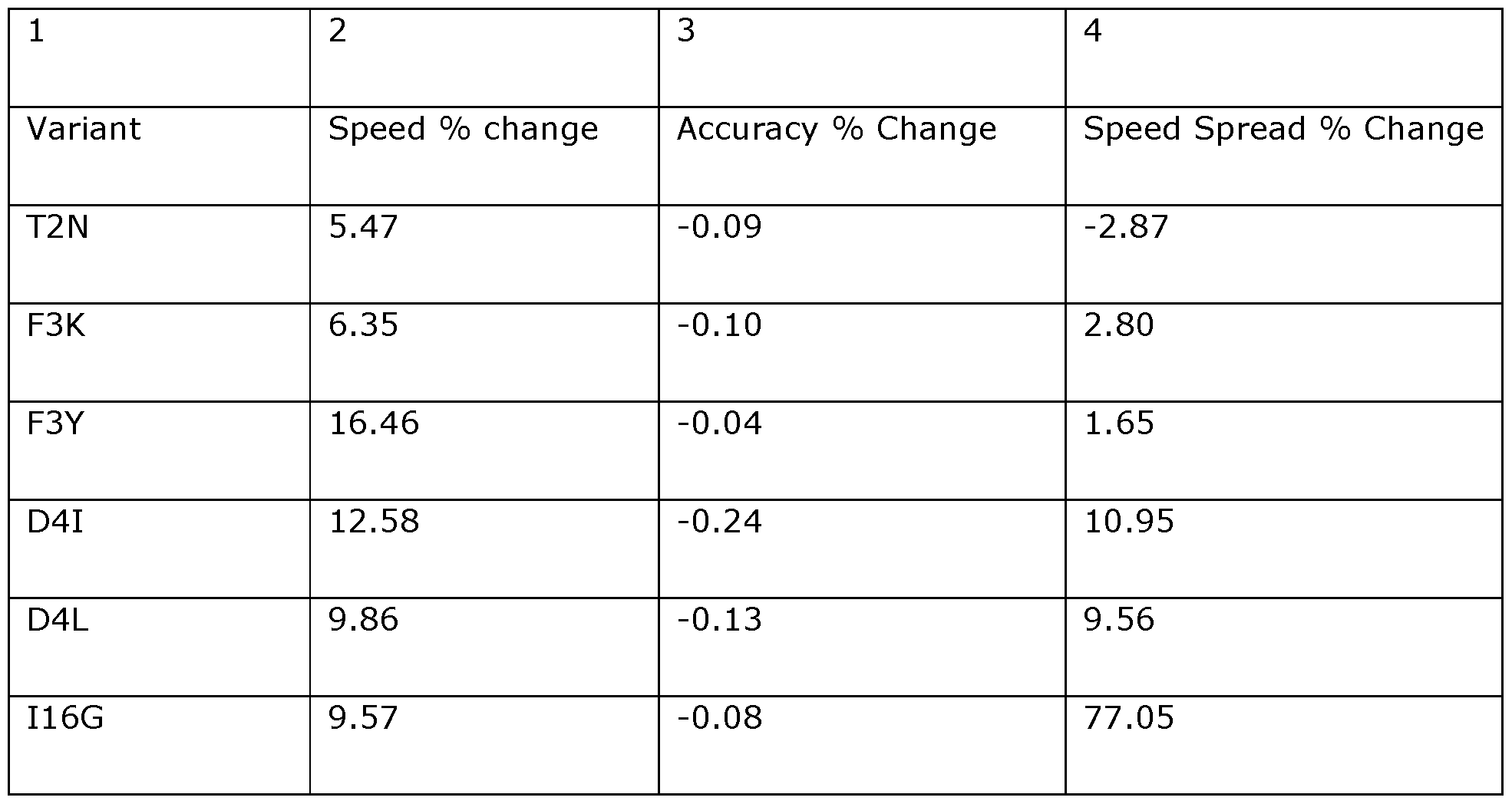
- the modified helicases of the invention display increased speed when used in methods of controlling the movement of an analyte through a transmembrane pore and in methods of characterising an analyte using a transmembrane pore.
- the speed at which an analyte passes through/relative to the pore may be increased by at least about 5%, at least about 10%, at least about 15%, at least about 20%, at least about 25%, at least about 30%, at least about 35%, at least about 40%, at least about 45%, at least about 50%, at least about 55%, at least about 60%, at least about 65%, at least about 70%, at least about 75%, at least about 80%, at least about 85%, at least about 90%, at least about 100%, at least about 150%, at least about 200%, at least about 300%, at least about 400%, at least about 500% or greater relative to the speed at which the analyte moves with respect to a pore when using a Dda helicase which does not comprise a mutation of the invention.
- the inventors have surprisingly found that these alterations in speed, such as increases in speed, have minimal or no effect on accuracy readings. This is particularly advantageous in a method of characterising an analyte wherein an analyte is contacted with the pore and a helicase of the invention such that the polynucleotide binding protein controls the movement of the target analyte through/relative to the pore.
- the modified helicases of the invention display improved accuracy when used in methods of controlling the movement of an analyte through a transmembrane pore and in methods of characterising an analyte using a transmembrane pore.
- accuracy is interpreted to mean raw read simplex accuracy; that is a single pass of a single molecule through a transmembrane pore.
- Accuracy is a useful measure to track platform improvements of sequencing devices. Accuracy can also refer to consensus accuracy or to the accuracy in detecting something specific such as a mutation in a polynucleotide analyte for example.
- accuracy is interpreted to mean the percentage of bases above a certain confidence level, where the confidence level has been pre-calibrated.
- the modified helicases of the invention display improved accuracy with minimal to no changes in speed.
- accuracy is improved to give less than 10% error, less than 5% error, less than 4 % error, less than 3% error, less than 2% error, less than 1% error, less than 0.1% error.
- the modified helicases identified by the inventors typically comprise a combination of mutations, namely one or more modifications in the part of the modified helicase which interacts with a transmembrane pore.
- Accuracy may also by influenced by the speed which the polymer translocates the pore under enzyme control and the speed may be altered by altering the concentration of ATP provided to the enzyme.
- the inventors have surprisingly realised that the enzyme can exhibit changes in speed during successive polymer translocations within the same sequencing run under the same conditions which can give rise to a decrease in accuracy.
- Accuracy may be influenced by a number of factors such as the nanopore shape and composition, the enzyme as well as the interaction between the enzyme and nanopore. It is also influenced by the speed at which the polymer translocates the pore under enzyme control and the translocation speed may be increased or lowered by altering the concentration of ATP provided to the enzyme.
- the inventors have surprisingly realised that changes in speed occur during successive polymer translocations within the same sequencing run under the same sequencing conditions, which can give rise to a decrease in sequencing accuracy.
- the variation in sequencing speed for a number of polymers may be measured to obtain a normalised speed distribution and the inventors have surprisingly realised that some modified enzymes can give rise to a lower normalised speed distribution and therefore an increased sequencing accuracy.
- the speed spread ratio of a modified helicase is the normalised speed distribution of the modified helicase divided by the normalised speed distribution of the control, unmodified helicase.
- the modified helicases of the invention display a decreased speed spread ratio when used in methods of controlling the movement of an analyte through a transmembrane pore and in methods of characterising an analyte using a transmembrane pore.
- the speed spread ratio may be decreased by at least about 21%, at least about 21.8%, at least about 22%, at least about 25%, at least about 30%, at least about 35%, at least about 40%, at least about 45%, at least about 50%, at least about at least about 55%, at least about 60%, at least about 65%, at least about 70%, at least about 75%, at least about 80%, at least about 85%, at least about 90% relative to the spread speed ratio of a Dda helicase which does not comprise a mutation of the invention.
- a decreased speed spread ratio indicates that the modified helicase of the invention has a narrower distribution of speeds than the control, unmodified helicase.
- the invention provides a modified DNA dependent ATPase (Dda) helicase, wherein the helicase (a) is one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: F3, D4, 116, K24, K25, H26, H27, V28, P33, T42, F44, 146, A48, A60, P62, A65, K67, K68, 169, K72, K76, S79, 184, P104, K108, V117, Y120, R122, L124, 1132, P134, W135, T137, N144, K145, E154, A157, Y158, P161, K166, Q170, T174, V176, N180, A181, V190, Y197, V200, V201, R207, T213, A214, L215, M219, V220, 1225, K227, L229, V237, M238, K247, L24
- V96K V96K
- V96T V96P
- V96D V96G
- V96S V96I
- V96N V96A
- V96H V96H
- V96M V96M
- V103K V103K
- V103N V103L
- V103I V103T
- V103R V103Y
- V103S V103D
- V103A V103E
- V103H V103M
- V103P V103P
- V103Q V103F
- G153H or G153P, - N155K, N155P, N155S, N155E, N155D, N155G, N155T, N155A, N155V, N155L, N155I, N155H, N155M, N155Q, or N155R,
- T210A T210M, T210P, T210Q, T210I, T210L, T210V, or T210F,
- N292E N292A, N292S, N292D, N292K, N292H, N292R, N292Q, or N292T,
- T359V T359M
- T359A T359I
- T359Y T359F
- T359E T359K
- T359N T359D
- T359P T359Q
- T359R T359S
- T359W T359W
- T359G T359G
- the invention also provides:
- a construct comprising a helicase of the invention and an additional polynucleotide binding moiety, wherein the helicase is attached to the polynucleotide binding moiety and the construct has the ability to control the movement of an analyte;
- polynucleotide which comprises a sequence which encodes a helicase of the invention or a construct of the invention
- a method of making a helicase of the invention or a construct of the invention which comprises expressing a polynucleotide of the invention, transfecting a cell with a vector of the invention or culturing a host cell of the invention;
- a method of controlling the movement of an analyte comprising contacting the analyte with a helicase of the invention or a construct of the invention and thereby controlling the movement of the analyte; a method of characterising a target analyte, comprising:
- a method of forming a sensor for characterising a target analyte comprising forming a complex between (a) a pore and (b) a helicase of the invention or a construct of the invention and thereby forming a sensor for characterising the target analyte;
- a sensor for characterising a target analyte comprising a complex between (a) a pore and (b) a helicase of the invention or a construct of the invention;
- an apparatus for characterising target analytes in a sample comprising (a) a plurality of pores and (b) a plurality of helicases of the invention or a plurality of constructs of the invention;
- a method of producing a construct of the invention comprising attaching a helicase of the invention to an additional polynucleotide binding moiety and thereby producing the construct; a series of two or more helicases attached to a polynucleotide, wherein at least one of the two or more helicases is a helicase of the invention; and a method of improving the movement of a target analyte with respect to a transmembrane pore when the movement is controlled by a DNA dependent ATPase (Dda) helicase, wherein the DNA dependent ATPase (Dda) helicase is modified according to the invention which improves the movement of the target analyte with respect to the transmembrane pore.
- Dda DNA dependent ATPase
- the I symbol means "or".
- Q87R/K means Q87R or Q87K.
- the I symbol means "and” such that Y51/N55 is Y51 and N55.
- “About” as used herein when referring to a measurable value such as an amount, a temporal duration, and the like, is meant to encompass variations of ⁇ 20 % or ⁇ 10 %, more preferably ⁇ 5 %, even more preferably ⁇ 1 %, and still more preferably ⁇ 0.1 % from the specified value, as such variations are appropriate to perform the disclosed methods.
- a “homologue” or “homologues” of a protein encompass peptides, oligopeptides, polypeptides, proteins and enzymes having amino acid substitutions, deletions and/or insertions relative to the unmodified or wild-type protein in question and having similar biological and functional activity as the unmodified protein from which they are derived.
- amino acid identity refers to the extent that sequences are identical on an amino acid-by-amino acid basis over a window of comparison.
- a "percentage of sequence identity” is calculated by comparing two optimally aligned sequences over the window of comparison, determining the number of positions at which the identical amino acid residue (e.g., Ala, Pro, Ser, Thr, Gly, Vai, Leu, He, Phe, Tyr, Trp, Lys, Arg, His, Asp, Glu, Asn, Gin, Cys and Met) occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison (i.e., the window size), and multiplying the result by 100 to yield the percentage of sequence identity.
- the identical amino acid residue e.g., Ala, Pro, Ser, Thr, Gly, Vai, Leu, He, Phe, Tyr, Trp, Lys, Arg, His, Asp, Glu, Asn, Gin, Cys and Met
- the present invention provides a modified Dda helicase.
- the one or more specific modifications are discussed in more detail below.
- Modifications according to the invention include one or more substitutions as discussed below.
- the invention provides a modified DNA dependent ATPase (Dda) helicase in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: F3, D4, 116, K24, K25, H26, H27, V28, P33, T42, F44, 146, A48, A60, P62, A65, K67, K68, 169, K72, K76, S79, 184, P104, K108, V117, Y120, R122, L124, 1132, P134, W135, T137, N144, K145, E154, A157, Y158, P161, K166, Q170, T174, V176,
- Dda DNA dependent ATPase
- corresponding positions or positions corresponding to specific amino acid positions may be determined by standard techniques in the art.
- the PILEUP and BLAST algorithms mentioned above can be used to align the sequence of a Dda helicase with Dda 1993 (SEQ ID NO: 118) and hence to identify corresponding residues.
- the modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising a substitution or modification at one or more of the 113 positions set out above (i.e., one or more of positions F3 to F437).
- the modified Dda helicase of the invention may comprise a modification or substitution at any number and combination of the 113 positions corresponding to the amino acid positions set out above.
- the modified helicase of the invention may comprise a modification or substitution at 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22,
- the modified Dda helicase is preferably one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: A60, P62, K68, K72, K76, N144, R207, V237, Y304, V309, G316, L319, G331, D333, L349, P373, S385, P391, M401, R405, A406, and H414.
- the modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising a modification or substitution at one or more of these 22 positions.
- the modified Dda helicase of the invention may comprise a modification or substitution at any number and combination of the positions corresponding to the 22 amino acid positions set out above.
- the modified Dda helicase of the invention may comprise a modification or substitution at 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, or 22 of the positions corresponding to the amino acid positions set out above (i.e., from A60 to H414).
- the modified Dda helicase is preferably one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: A60, P62, K68, K72, K76, N144, R207, V237, Y304, V309, G316, L319, G331, D333, L349, P373, S385, P391, M401, A406, and H414.
- the modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising a modification or substitution at one or more of these 21 positions.
- the modified Dda helicase of the invention may comprise a modification or substitution at any number and combination of the positions corresponding to the 21 amino acid positions set out above.
- the modified Dda helicase of the invention may comprise a modification or substitution at 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, or 21 of the positions corresponding to the amino acid positions set out above (i.e., from A60 to H414).
- the modified Dda helicase is preferably one in which the position corresponding to amino acid position R405 in Dda 1993 is modified or substituted.
- the modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising a modification or substitution at R405.
- the position corresponding to amino acid position F3 in Dda 1993 is preferably substituted with K (e.g., F3K) or Y (e.g., F3Y).
- the position corresponding to amino acid position D4 in Dda 1993 may be substituted with I, L, K, P, S, E, Y, T, F, M, V, A, G, H, N, or Q.
- the position corresponding to amino acid position D4 in Dda 1993 is preferably substituted with I (e.g., D4I) or L (e.g., D4L).
- the position corresponding to amino acid position 116 in Dda 1993 may be substituted with G, A, E, L, K, D, N, S, F, M, T, V, or Y.
- the position corresponding to amino acid position 116 in Dda 1993 is preferably substituted with G (e.g., I16G).
- the position corresponding to amino acid position K24 in Dda 1993 may be substituted with G, P, A, D, N, S, E, H, M, Q, R, or T.
- the position corresponding to amino acid position K24 in Dda 1993 is preferably substituted with G (e.g., K24G) or P (e.g., K24P).
- the position corresponding to amino acid position K25 in Dda 1993 may be substituted with A, N, G, D, E, I, L, M, P, Q, R, S, T, V, or H.
- the position corresponding to amino acid position K25 in Dda 1993 is preferably substituted with A (e.g., K25A).
- K e.g., H26K
- L e.g., H26L
- the position corresponding to amino acid position H27 in Dda 1993 may be substituted with F, I, V, L, Y, A, T, S, M, W, D, E, K, N, Q, or R.
- the position corresponding to amino acid position H27 in Dda 1993 is preferably substituted with F (e.g., H27F).
- the position corresponding to amino acid position V28 in Dda 1993 may be substituted with I, F, A, L, M, T, or Y.
- the position corresponding to amino acid position V28 in Dda 1993 is preferably substituted with I (e.g., V28I).
- the position corresponding to amino acid position P33 in Dda 1993 may be substituted with A, Y, F, G, E, I, K, L, M, Q, R, S, T, V, D, H, or W.
- the position corresponding to amino acid position P33 in Dda 1993 is preferably substituted with A (e.g., P33A) or Y (e.g., P33Y).
- the position corresponding to amino acid position T42 in Dda 1993 may be substituted with I, L, A, F, M, V, Y, D, E, K, N, P, Q, R, or S.
- the position corresponding to amino acid position T42 in Dda 1993 is preferably substituted with I (e.g., T42I).
- the position corresponding to amino acid position F44 in Dda 1993 may be substituted with K, A, R, Y, G, H, T, S, Q, L, N, E, M, V, D, P, I, or W.
- the position corresponding to amino acid position F44 in Dda 1993 is preferably substituted with K (e.g., F44K).
- the position corresponding to amino acid position 146 in Dda 1993 may be substituted with V, A, F, L, M, T, or Y.
- the position corresponding to amino acid position 146 in Dda 1993 is preferably substituted with V (e.g., I46V).
- the position corresponding to amino acid position A48 in Dda 1993 may be substituted with K, H, Y, R, E, N, Q, S, T, D, M, P, G, I, L, V, F, or W.
- the position corresponding to amino acid position A46 in Dda 1993 is preferably substituted with H (e.g., A48H), K (e.g., A48K) or Y (e.g., A48Y).
- the position corresponding to amino acid position A60 in Dda 1993 may be substituted with T, D, E, I, K, L, M, N, P, Q, R, S, V, or G.
- the position corresponding to amino acid position A60 in Dda 1993 is preferably substituted with T (e.g., A60T).
- the position corresponding to amino acid position P62 in Dda 1993 may be substituted with S, A, D, E, G, H, K, M, N, Q, R, or T.
- the position corresponding to amino acid position P62 in Dda 1993 is preferably substituted with S (e.g., P62S).
- the position corresponding to amino acid position A65 in Dda 1993 may be substituted with V, Q, K, G, R, Y, L, I, F, T, M, H, E, P, S, D, or N.
- the position corresponding to amino acid position A65 in Dda 1993 is preferably substituted with Q (e.g., A65Q) or V (e.g., A65V).
- the position corresponding to amino acid position K67 in Dda 1993 may be substituted with A, V, T, S, R, M, E, G, I, L, P, Q, D, H, N, F, or Y.
- the position corresponding to amino acid position K67 in Dda 1993 is preferably substituted with A (e.g., K67A) or V (e.g., K67V).
- the position corresponding to amino acid position K68 in Dda 1993 may be substituted with A, N, E, G, I, L, M, P, Q, R, S, T, V, D, or H.
- the position corresponding to amino acid position K68in Dda 1993 is preferably substituted with A (e.g., K68A).
- the position corresponding to amino acid position 169 in Dda 1993 may be substituted with E, V, R, Q, N, A, K, T, S, D, L, H, M, P, F, or Y.
- the position corresponding to amino acid position 169 in Dda 1993 is preferably substituted with E (e.g., I69E) or L (e.g., I69V).
- the position corresponding to amino acid position K72 in Dda 1993 may be substituted with A, E, G, I, L, M, P, Q, R, S, T, V, D, H, or N.
- the position corresponding to amino acid position K72 in Dda 1993 is preferably substituted with A (e.g., K72A).
- the position corresponding to amino acid position K76 in Dda 1993 may be substituted with M, Y, F, I, V, R, A, L, Q, S, T, W, D, E, H, N, or P.
- the position corresponding to amino acid position K76 in Dda 1993 is preferably substituted with M (e.g., K76M) or Y (e.g., K76Y).
- the position corresponding to amino acid position S79 in Dda 1993 may be substituted with T, K, R, V, E, A, D, I, L, M, N, P, Q, G, or H.
- the position corresponding to amino acid position S79 in Dda 1993 is preferably substituted with T (e.g., S79T).
- the position corresponding to amino acid position 184 in Dda 1993 may be substituted with L, V, F, A, M, T, or Y.
- the position corresponding to amino acid position 184 in Dda 1993 is preferably substituted with L (or e.g., I84L).
- the position corresponding to amino acid position P104 in Dda 1993 may be substituted with Y, K, E, F, H, I, L, M, Q, V, W, A, D, S, or T.
- the position corresponding to amino acid position P104 in Dda 1993 is preferably substituted with Y (e.g., P104Y).
- the position corresponding to amino acid position K108 in Dda 1993 may be substituted with Y, E, D, H, N, S, G, F, I, L, M, Q, V, W, A, P, R, or T.
- the position corresponding to amino acid position K108 in Dda 1993 is preferably substituted with D (e.g., K108D), E (e.g., K108E) or Y (e.g., K108Y).
- the position corresponding to amino acid position V117 in Dda 1993 may be substituted with A, S, E, G, I, K, L, M, P, Q, R, T, F, or Y.
- the position corresponding to amino acid position V117 in Dda 1993 is preferably substituted with A (e.g., V117A).
- the position corresponding to amino acid position Y120 in Dda 1993 may be substituted with V, L, I, M, F, A, T, S, H, Q, or W.
- the position corresponding to amino acid position Y120 in Dda 1993 is preferably substituted with V (e.g., Y120V).
- the position corresponding to amino acid position R122 in Dda 1993 may be substituted with E, K, N, D, L, T, S, A, V, I, P, Q, H, or M.
- the position corresponding to amino acid position R122 in Dda 1993 is preferably substituted with E (e.g., R122E).
- the position corresponding to amino acid position L124 in Dda 1993 may be substituted with T, A, D, E, I, K, M, N, P, Q, R, S, V, F, or Y.
- the position corresponding to amino acid position L124 in Dda 1993 is preferably substituted with T (e.g., L124T).
- the position corresponding to amino acid position 1132 in Dda 1993 may be substituted with T, V, L, A, D, E, K, M, N, P, Q, R, S, F, or Y.
- the position corresponding to amino acid position 1132 in Dda 1993 is preferably substituted with T (e.g., I132T).
- the position corresponding to amino acid position P134 in Dda 1993 may be substituted with D, E, K, S, L, A, R, I, N, V, T, G, H, or Q.
- the position corresponding to amino acid position P134 in Dda 1993 is preferably substituted with D (e.g., P134D).
- the position corresponding to amino acid position W135 in Dda 1993 may be substituted with D, H, T, N, E, G, S, K, Y, Q, R, F, A, P, or M.
- the position corresponding to amino acid position W135 in Dda 1993 is preferably substituted with D (e.g., W135D) or H (e.g., W135H).
- the position corresponding to amino acid position T137 in Dda 1993 may be substituted with K, R, I, V, A, D, E, H, M, N, P, Q, S, or L.
- the position corresponding to amino acid position T137 in Dda 1993 is preferably substituted with K (e.g., T137K)).
- the position corresponding to amino acid position N144 in Dda 1993 may be substituted with S, P, K, T, R, E, A, D, I, L, Q, Y, H, V, M, or G.
- the position corresponding to amino acid position N144 in Dda 1993 is preferably substituted with S (e.g., N144S).
- the position corresponding to amino acid position K145 in Dda 1993 may be substituted with D, A, H, R, G, N, E, P, Q, S, T, or M.
- the position corresponding to amino acid position K145 in Dda 1993 is preferably substituted with D (e.g., K145D).
- the position corresponding to amino acid position E154 in Dda 1993 may be substituted with N, S, V, I, T, D, G, H, K, Q, R, A, or P.
- the position corresponding to amino acid position E154 in Dda 1993 is preferably substituted with N (e.g., K154N).
- the position corresponding to amino acid position A157 in Dda 1993 may be substituted with E, D, S, K, P, N, H, Q, R, T, G, I, L, M, or V.
- the position corresponding to amino acid position A157 in Dda 1993 is preferably substituted with E (e.g., A157E).
- the position corresponding to amino acid position Y158 in Dda 1993 may be substituted with Q, S, E, L, D, A, I, K, F, N, G, R, V, T, H, P, M, or W.
- the position corresponding to amino acid position Y158 in Dda 1993 is preferably substituted with E (e.g., Y158E), Q (e.g., Y158Q) or S (e.g., Y158S).
- the position corresponding to amino acid position P161 in Dda 1993 may be substituted with E, D, K, S, N, R, Q, T, A, or H.
- the position corresponding to amino acid position P161 in Dda 1993 is preferably substituted with E (e.g., P161E).
- the position corresponding to amino acid position K166 in Dda 1993 may be substituted with N, P, D, E, G, H, Q, R, S, T, A, or M.
- the position corresponding to amino acid position K166 in Dda 1993 is preferably substituted with N (e.g., K166N).
- the position corresponding to amino acid position Q170 in Dda 1993 may be substituted with K, I, V, T, S, E, R, N, H, Y, F, A, D, M, or P.
- the position corresponding to amino acid position Q170 in Dda 1993 is preferably substituted with K (e.g., Q170K).
- the position corresponding to amino acid position T174 in Dda 1993 may be substituted with S, A, D, E, G, H, K, M, N, P, Q, R, I, L, or V.
- the position corresponding to amino acid position T174 in Dda 1993 is preferably substituted with S (e.g., T174S).
- the position corresponding to amino acid position V176 in Dda 1993 may be substituted with I, G, P, A, T, R, S, Q, K, L, E, F, M, or Y.
- the position corresponding to amino acid position V176 in Dda 1993 is preferably substituted with I (e.g., V176I).
- the position corresponding to amino acid position N180 in Dda 1993 may be substituted with D, E, K, A, G, H, P, Q, S, T, or R.
- the position corresponding to amino acid position N180 in Dda 1993 is preferably substituted with D (e.g., N180D).
- the position corresponding to amino acid position A181 in Dda 1993 may be substituted with G, N, L, S, R, T, D, K, Q, H, E, I, M, P, or V.
- the position corresponding to amino acid position A181 in Dda 1993 is preferably substituted with G (e.g., A180G) or N (e.g., A180N).
- the position corresponding to amino acid position V190 in Dda 1993 may be substituted with I, A, F, L, M, T, or Y.
- the position corresponding to amino acid position V190 in Dda 1993 is preferably substituted with I (e.g., V190I).
- the position corresponding to amino acid position Y197 in Dda 1993 may be substituted with K, P, R, S, E, G, D, T, L, N, I, V, F, A, H, M, Q, or W.
- the position corresponding to amino acid position Y197 in Dda 1993 is preferably substituted with K (e.g., Y197K).
- the position corresponding to amino acid position V200 in Dda 1993 may be substituted with I, T, F, K, D, L, N, E, S, Y, P, A, or M.
- the position corresponding to amino acid position V200 in Dda 1993 is preferably substituted with I (e.g., V200I).
- the position corresponding to amino acid position V201 in Dda 1993 may be substituted with F, Y, I, K, L, H, M, W, A, or T.
- the position corresponding to amino acid position V201 in Dda 1993 is preferably substituted with F (e.g., V201F).
- the position corresponding to amino acid position R.207 in Dda 1993 may be substituted with I, H, K, V, F, Y, L, T, E, A, M, N, Q, S, or D.
- the position corresponding to amino acid position R.207 in Dda 1993 is preferably substituted with H (e.g., R207H) or I (e.g., R207I).
- the position corresponding to amino acid position T213 in Dda 1993 may be substituted with D, N, E, K, S, G, A, Q, H, P, I, L, M, R, or V.
- the position corresponding to amino acid position T213 in Dda 1993 is preferably substituted with D (e.g., T213D).
- the position corresponding to amino acid position A214 in Dda 1993 may be substituted with G, S, E, K, D, L, N, I, T, V, M, P, Q, R, or H.
- the position corresponding to amino acid position A214 in Dda 1993 is preferably substituted with G (e.g., A214G) or S (e.g., A214S).
- the position corresponding to amino acid position L215 in Dda 1993 may be substituted with Y, I, E, F, D, H, M, Q, V, W, A, T, K, N, P, R, or S.
- the position corresponding to amino acid position L215 in Dda 1993 is preferably substituted with E (e.g., L215E), I (e.g., L215I) or Y (e.g., L215Y).
- the position corresponding to amino acid position M219 in Dda 1993 may be substituted with L, I, V, A, F, Y, K, D, E, S, T, Q, R, or W.
- the position corresponding to amino acid position M219 in Dda 1993 is preferably substituted with L (e.g., M219L).
- the position corresponding to amino acid position V220 in Dda 1993 may be substituted with K, E, N, A, D, R, S, T, Q, L, I, G, H, M, P, F, or Y.
- the position corresponding to amino acid position V220 in Dda 1993 is preferably substituted with K (e.g., V220K).
- the position corresponding to amino acid position 1225 in Dda 1993 may be substituted with V, K, T, D, N, S, E, G, R, Q, P, A, H, L, Y, F, or M.
- the position corresponding to amino acid position 1225 in Dda 1993 is preferably substituted with V (e.g., I225V).
- the position corresponding to amino acid position K227 in Dda 1993 may be substituted with G, A, D, N, S, E, H, M, P, Q, R, or T.
- the position corresponding to amino acid position K227 in Dda 1993 is preferably substituted with G (e.g., K227G).
- the position corresponding to amino acid position L229 in Dda 1993 may be substituted with S, K, V, I, N, E, D, T, G, A, R, Q, H, M, P, F, or Y.
- the position corresponding to amino acid position L229 in Dda 1993 is preferably substituted with K (e.g., L229K) or S (e.g., L229S).
- the position corresponding to amino acid position V237 in Dda 1993 may be substituted with M, I, A, L, F, K, Q, R, S, T, W, or Y.
- the position corresponding to amino acid position V237 in Dda 1993 is preferably substituted with M (e.g., V237M).
- the position corresponding to amino acid position M238 in Dda 1993 may be substituted with I, L, V, F, A, T, Y, K, Q, R, S, or W.
- the position corresponding to amino acid position M238 in Dda 1993 is preferably substituted with I (e.g., M238I).
- the position corresponding to amino acid position K247 in Dda 1993 may be substituted with N, E, R, D, G, H, Q, S, T, A, M, or P.
- the position corresponding to amino acid position K247 in Dda 1993 is preferably substituted with N (e.g., K247N).
- the position corresponding to amino acid position L248 in Dda 1993 may be substituted with F, I, H, M, V, W, Y, A, or T.
- the position corresponding to amino acid position L248 in Dda 1993 is preferably substituted with F (e.g., L248F).
- the position corresponding to amino acid position 1251 in Dda 1993 may be substituted with F, L, E, Y, A, T, H, M, V, or W.
- the position corresponding to amino acid position 1251 in Dda 1993 is preferably substituted with F (e.g., I251F).
- the position corresponding to amino acid position R253 in Dda 1993 may be substituted with Q, A, D, E, H, K, M, N, P, S, T, or Y.
- the position corresponding to amino acid position R253 in Dda 1993 is preferably substituted with Q (e.g., R253Q).
- the position corresponding to amino acid position K254 in Dda 1993 may be substituted with N, S, D, E, G, H, Q, R, T, A, M, or P.
- the position corresponding to amino acid position K254 in Dda 1993 is preferably substituted with N (e.g., K254N) or S (e.g., K254S).
- the position corresponding to amino acid position F257 in Dda 1993 may be substituted with Y, H, I, L, M, Q, V, or W.
- the position corresponding to amino acid position F257 in Dda 1993 is preferably substituted with Y (e.g., F257Y).
- the position corresponding to amino acid position D262 in Dda 1993 may be substituted with P, E, A, K, Q, S, T, G, H, or N.
- the position corresponding to amino acid position D262 in Dda 1993 is preferably substituted with P (e.g., D262P).
- the position corresponding to amino acid position V265 in Dda 1993 may be substituted with E, P, A, K, S, G, D, N, T, H, Q, R, F, I, L, M, or Y.
- the position corresponding to amino acid position V265 in Dda 1993 is preferably substituted with E (e.g., V265E).
- the position corresponding to amino acid position D282 in Dda 1993 may be substituted with P, A, E, K, Q, S, T, G, H, or N.
- the position corresponding to amino acid position D282 in Dda 1993 is preferably substituted with P (e.g., D282P).
- the position corresponding to amino acid position 1289 in Dda 1993 may be substituted with L, V, M, K, E, G, A, F, T, or Y.
- the position corresponding to amino acid position 1289 in Dda 1993 is preferably substituted with L (e.g., I289L).
- the position corresponding to amino acid position Q295 in Dda 1993 may be substituted with D, E, M, S, G, H, K, N, P, T, A, R, or Y.
- the position corresponding to amino acid position Q295 in Dda 1993 is preferably substituted with D (e.g., Q295D).
- the position corresponding to amino acid position Y304 in Dda 1993 may be substituted with K, E, L, P, D, N, S, G, A, R, T, Q, H, V, I, M, F, or W.
- the position corresponding to amino acid position Y304 in Dda 1993 is preferably substituted with K (e.g., Y304K).
- the position corresponding to amino acid position S306 in Dda 1993 may be substituted with E, Y, D, K, T, G, F, V, N, A, H, P, Q, R, or M.
- the position corresponding to amino acid position S306 in Dda 1993 is preferably substituted with E (e.g., S306E).
- the position corresponding to amino acid position V309 in Dda 1993 may be substituted with F, K, I, H, L, M, W, Y, A, or T.
- the position corresponding to amino acid position V309 in Dda 1993 is preferably substituted with F (e.g., V309F).
- the position corresponding to amino acid position V314 in Dda 1993 may be substituted with I, K, E, L, A, F, M, T, or Y.
- the position corresponding to amino acid position V314 in Dda 1993 is preferably substituted with I (e.g., V314I).
- the position corresponding to amino acid position G316 in Dda 1993 may be substituted with E, K, D, V, I, N, L, A, H, P, Q, R, S, or T.
- the position corresponding to amino acid position G316 in Dda 1993 is preferably substituted with E (e.g., G316E).
- the position corresponding to amino acid position L319 in Dda 1993 may be substituted with E, K, I, V, T, N, D, G, A, H, P, Q, R, S, F, M, or Y.
- the position corresponding to amino acid position L319 in Dda 1993 is preferably substituted with E (e.g., L319E).
- the position corresponding to amino acid position R321 in Dda 1993 may be substituted with K, N, E, V, I, D, L, A, H, M, P, Q, S, or T.
- the position corresponding to amino acid position R321 in Dda 1993 is preferably substituted with K (e.g., R321K).
- the position corresponding to amino acid position T329 in Dda 1993 may be substituted with P, S, G, K, A, E, D, I, V, L, N, R, Y, F, Q, or M.
- P e.g., T329P
- the position corresponding to amino acid position G331 in Dda 1993 may be substituted with D, E, K, L, S, I, N, T, V, H, P, Q, or A.
- the position corresponding to amino acid position G331 in Dda 1993 is preferably substituted with D (e.g., G331D).
- the position corresponding to amino acid position D333 in Dda 1993 may be substituted with G, K, E, I, A, N, S, H, P, Q, or T.
- the position corresponding to amino acid position D333 in Dda 1993 is preferably substituted with G (e.g., D333G).
- the position corresponding to amino acid position L349 in Dda 1993 may be substituted with E, N, K, Q, A, T, S, D, V, R, I, H, P, F, M, or Y.
- the position corresponding to amino acid position L349 in Dda 1993 is preferably substituted with E (e.g., L349E).
- the position corresponding to amino acid position F352 in Dda 1993 may be substituted with Y, L, K, E, R, I, A, N, D, H, M, Q, V, or W.
- the position corresponding to amino acid position F352 in Dda 1993 is preferably substituted with Y (e.g., F352Y).
- the position corresponding to amino acid position T362 in Dda 1993 may be substituted with A, I, V, L, E, K, M, D, R, Y, N, Q, S, F, G, or P.
- the position corresponding to amino acid position T362 in Dda 1993 is preferably substituted with A (e.g., T362A).
- the position corresponding to amino acid position N365 in Dda 1993 may be substituted with K, E, A, S, R, D, H, M, P, Q, T, or G.
- the position corresponding to amino acid position N365 in Dda 1993 is preferably substituted with K (e.g., N365K).
- the position corresponding to amino acid position N367 in Dda 1993 may be substituted with S, K, L, E, A, T, I, V, Q, R, Y, F, M, D, G, H, or P.
- the position corresponding to amino acid position N367 in Dda 1993 is preferably substituted with S (e.g., N367S).
- the position corresponding to amino acid position G369 in Dda 1993 may be substituted with E, K, D, N, S, T, R, L, I, A, Y, H, P, or Q.
- the position corresponding to amino acid position G369 in Dda 1993 is preferably substituted with E (e.g., G369E).
- the position corresponding to amino acid position G370 in Dda 1993 may be substituted with K, V, E, I, S, A, N, D, H, M, P, Q, R, or T.
- the position corresponding to amino acid position G370 in Dda 1993 is preferably substituted with K (e.g., G370K).
- the position corresponding to amino acid position P373 in Dda 1993 may be substituted with H, L, N, K, M, Y, F, I, V, S, E, T, A, R, D, G, Q, or W.
- the position corresponding to amino acid position P373 in Dda 1993 is preferably substituted with H (e.g., P373) or L (e.g., P373L).
- the position corresponding to amino acid position A380 in Dda 1993 may be substituted with V, L, T, I, K, M, F, Y, E, G, P, Q, R, or S.
- the position corresponding to amino acid position A380 in Dda 1993 is preferably substituted with V (e.g., A380V).
- the position corresponding to amino acid position Q383 in Dda 1993 may be substituted with L, T, M, Y, K, F, I, V, R, A, E, S, N, D, G, H, or P.
- the position corresponding to amino acid position Q383 in Dda 1993 is preferably substituted with L (e.g., Q383L).
- the position corresponding to amino acid position S385 in Dda 1993 may be substituted with I, T, A, Y, K, H, R, F, L, M, V, D, E, G, N, P, or Q.
- the position corresponding to amino acid position S385 in Dda 1993 is preferably substituted with I (e.g., S385I).
- the position corresponding to amino acid position P391 in Dda 1993 may be substituted with G, Y, F, H, A, V, D, N, S, E, K, Q, or T.
- the position corresponding to amino acid position P391 in Dda 1993 is preferably substituted with G (e.g., P391G).
- the position corresponding to amino acid position S393 in Dda 1993 may be substituted with I, V, L, T, E, D, M, F, A, Y, G, H, K, N, P, Q, or R.
- the position corresponding to amino acid position S393 in Dda 1993 is preferably substituted with I (e.g., S393I).
- the position corresponding to amino acid position F395 in Dda 1993 may be substituted with I, V, Y, L, T, A, M, H, or W.
- the position corresponding to amino acid position F395 in Dda 1993 is preferably substituted with I (e.g., F395I).
- the position corresponding to amino acid position M401 in Dda 1993 may be substituted with L, S, V, I, G, D, E, R, Q, A, W, N, T, F, Y, K, H, or P.
- the position corresponding to amino acid position M401 in Dda 1993 is preferably substituted with L (e.g., M401L), S (e.g., M401S) or V (e.g., M401V).
- the position corresponding to amino acid position S402 in Dda 1993 may be substituted with T, V, R, I, K, L, E, H, Q, M, A, D, N, P, or G.
- the position corresponding to amino acid position S402 in Dda 1993 is preferably substituted with T (e.g., S402T).
- the position corresponding to amino acid position V403 in Dda 1993 may be substituted with F, Y, I, H, L, M, W, A, or T.
- the position corresponding to amino acid position V403 in Dda 1993 is preferably substituted with F (e.g., V403F).
- the position corresponding to amino acid position D404 in Dda 1993 may be substituted with N, K, R, E, G, H, Q, S, T, or P.
- the position corresponding to amino acid position D404 in Dda 1993 is preferably substituted with N (e.g., D404N).
- the position corresponding to amino acid position R405 in Dda 1993 may be substituted with N, K, D, E, G, H, Q, S, T, A, M, or P.
- the position corresponding to amino acid position R405 in Dda 1993 is preferably substituted with K (e.g., R405K) or N (e.g., R405N).
- the position corresponding to amino acid position A406 in Dda 1993 may be substituted with T, V, I, D, E, K, L, M, N, P, Q, R, S, G, F, or Y.
- the position corresponding to amino acid position A406 in Dda 1993 is preferably substituted with T (e.g., A406T) or V (e.g., A406V).
- the position corresponding to amino acid position 1413 in Dda 1993 may be substituted with E, L, K, S, D, G, N, P, Y, F, T, R, A, H, Q, M, or V.
- the position corresponding to amino acid position 1413 in Dda 1993 is preferably substituted with E (e.g., I413E).
- the position corresponding to amino acid position H414 in Dda 1993 may be substituted with Y, D, F, Q, K, N, S, E, G, P, R, T, L, I, A, M, V, or W.
- the position corresponding to amino acid position H414 in Dda 1993 is preferably substituted with D (e.g., H414D) or Y (e.g., H414Y).
- the position corresponding to amino acid position V418 in Dda 1993 may be substituted with K, E, P, R, D, T, A, S, N, L, I, Y, G, H, M, Q, or F.
- the position corresponding to amino acid position V418 in Dda 1993 is preferably substituted with K (e.g., V418K).
- the position corresponding to amino acid position E419 in Dda 1993 may be substituted with Q, D, A, H, K, M, N, P, R, S, T, or Y.
- the position corresponding to amino acid position E419 in Dda 1993 is preferably substituted with E (e.g., E419Q).
- the position corresponding to amino acid position A421 in Dda 1993 may be substituted with Q, L, R, N, S, K, D, E, H, M, P, T, Y, G, I, V, or F.
- the position corresponding to amino acid position A421 in Dda 1993 is preferably substituted with L (e.g., A421L) or Q (e.g., A421Q).
- the position corresponding to amino acid position Q422 in Dda 1993 may be substituted with K, R, L, N, A, D, E, H, M, P, S, T, or Y.
- the position corresponding to amino acid position Q422 in Dda 1993 is preferably substituted with K (e.g., Q422K).
- the position corresponding to amino acid position V427 in Dda 1993 may be substituted with T, E, D, A, I, K, L, M, N, P, Q, R, S, F, or Y.
- the position corresponding to amino acid position V427 in Dda 1993 is preferably substituted with T (e.g., V427T).
- the position corresponding to amino acid position V429 in Dda 1993 may be substituted with I, L, A, F, M, T, or Y.
- the position corresponding to amino acid position V429 in Dda 1993 is preferably substituted with I (e.g., V429I).
- the position corresponding to amino acid position G432 in Dda 1993 may be substituted with P, A, S, T, D, E, K, Q, or N.
- the position corresponding to amino acid position G432 in Dda 1993 is preferably substituted with P (e.g., G432P).
- the position corresponding to amino acid position Y434 in Dda 1993 may be substituted with K, I, V, R, E, T, N, D, L, S, Q, H, G, A, M, P, F, or W.
- the position corresponding to amino acid position Y434 in Dda 1993 is preferably substituted with K (e.g., Y434K).
- the position corresponding to amino acid position D435 in Dda 1993 may be substituted with E, Q, A, H, K, N, P, R, S, T, or G.
- the position corresponding to amino acid position D435 in Dda 1993 is preferably substituted with E (e.g., D435E).
- the position corresponding to amino acid position F437 in Dda 1993 may be substituted with Y, I, H, L, M, Q, V, or W.
- the position corresponding to amino acid position F437 in Dda 1993 is preferably substituted with Y (e.g., F437Y).
- the modified Dda helicase of the invention may further comprise one or more of the 45 or 31 substitutions set out below or one or more of the 45 or 31 substitutions at the position(s) corresponding to the 45 or 31 amino acid positions in Dda 1993 set out below.
- the invention also provides a modified DNA dependent ATPase (Dda) helicase comprising one or more of the following substitutions or one or more of the following substitutions at the position(s) corresponding to the following amino acid positions in Dda 1993:
- Dda DNA dependent ATPase
- T2N - T2N, T2K, T2S, T2D, T2I, T2L, T2E, T2F, T2G, T2H, T2Q, T2R, T2A, T2M, T2P, or T2V, preferably T2N,
- E47D E47G, E47H, E47N, E47P, E47Q, E47S, E47T, E47A, or E47R, preferably E47D,
- E54I E54V, E54K, E54Y, E54T, E54L, E54N, E54A, E54F, E54M, E54D, E54H, E54P, E54Q, E54R, or E54S, preferably E54I,
- S83E, S83D, S83A, S83P, S83G, or S83M preferably S83E, - K86N, K86G, K86D, K86E, K86H, K86Q, K86R, K86S, K86T, K86M, or K86P, preferably K86N,
- N88K preferably N88K
- N88E preferably N88T
- N88S preferably N88V
- N88Q preferably N88A, N88L, N88D, N88G, N88M, N88I, or N88P, preferably N88K
- V90E V90D, V90K, V90Q, V90N, V90S, V90A, V90I, V90L, V90M, V90P, V90R, V90F, V90Y, V90H, or V90G, preferably V90D or V90E,
- T91E - T91E, T91D, T91R, T91K, T91Y, T91H, T91P, T91A, T91V, T91L, T91I, T91S, or T91M, preferably T91E,
- E93P E93D
- E93G E93A
- E93L E93K
- E93Q E93S
- E93T E93H
- E93N E93N
- E93R preferably E93P
- V96K V96K
- V96T V96P
- V96D V96G
- V96S V96I
- V96N V96A
- V96H V96H
- V96M V96M
- K101R K101T, K101S, K101V, K101E, K101I, K101G, K101L, K101M, K101P, K101Q, K101D, K101H, or K101N, preferably K101R,
- V103K V103N, V103L, V103I, V103T, V103R, V103Y, V103S, V103D, V103A, V103E, V103H, V103M, V103P, V103Q, or V103F, preferably V103K,
- G153E preferably G153E, G153S, G153D, G153T, G153N, G153K, G153A, G153Q, G153R, G153L, G153H, or G153P, preferably G153E,
- N155K preferably N155K, N155P, N155S, N155E, N155D, N155G, N155T, N155A, N155V, N155L, N155I, N155H, N155M, N155Q, or N155R, preferably N155K,
- - K177A or K177P - W195F, W195Y, W195P, W195K, W195E, W195I, W195L, W195D, W195S, W195G, W195T, W195V, W195Q, W195N, W195R, W195M, or W195H, preferably W195Y, W195P, W195K, W195E, W195I, W195G, W195T, W195Q, W195N, W195R, W195M, or W195H, more preferably W195F or W195Y,
- I196P I196L, I196D, I196A, I196E, I196K, I196Q, I196S, I196T, I196F, I196M, I196V, or I196Y, preferably I196L or I196P,
- D198N D198P, D198E, D198G, D198H, D198K, D198Q, D198R, D198S, or D198T, preferably D198N, D198P, D198E, D198G, D198H, D198K, D198Q, D198R, or D198T, more preferably D198N,
- T210A T210M, T210P, T210Q, T210I, T210L, T210V, or T210F,
- K243R, K243E, K243D K243A, K243S, K243M, K243N, K243P, K243T, or K243Y, preferably K243R or K243E,
- E258G E258K, E258N, E258P, E258D, E258S, E258A, E258H, E258Q, E258R, or E258T, preferably E258G, E258K, E258N, E258P, E258D, E258H, E258Q, E258R, or E258T, more preferably E258G,
- F276T preferably F276T, F276S, F276L, F276P, F276E, F276D, F276N, or F276Q, preferably F276T,
- I281E I281V, I281K, I281A, I281D, I281H, I281N, I281P, I281Q, I281R, I281S, I281T, I281F, I281L, I281M, or I281Y, preferably I281E,
- N292E N292A, N292S, N292D, N292K, N292H, N292R, N292Q, or N292T, preferably N292E,
- E301K, E301D, E301N, E301S, E301A, E301H, E301M, E301P, E301Q, E301R, or E301T preferably E301K
- Y335P Y335K, Y335E, Y335F, Y335T, Y335Q, Y335S, Y335R, Y335D, Y335V, Y335N, Y335G, Y335H, Y335M, or Y335W, preferably Y335P,
- Y336E Y336H, Y336K, Y336I, Y336G, Y336S, Y336T, Y336D, Y336V, Y336P, Y336N, Y336R, Y336Q, Y336A, Y336F, Y336M, or Y336W, preferably Y336E, R337E, R337K, R337T, R337Q, R337P, R337V, R337F, R337S, R337D, R337G,
- T359V T359M, T359A, T359I, T359Y, T359F, T359E, T359K, T359N, T359D, T359P, T359Q, T359R, T359S, T359W, or T359G, preferably T359A, T359V, T359I or T359M, more preferably T359V,
- K364N K364Q, K364D, K364E, K364G, K364H, K364S, K364T, K364M, K364P, K364Y, K364I, K364L, or K364V, preferably K364N or K364Q,
- W366D W366N, W366K, W366A, W366S, W366I, W366E, W366R, W366M, W366T, W366Q, W366Y, W366G, W366F, W366V, W366P, or W366H, preferably W366D or W366N,
- the modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising one or more of the 45 substitutions set out above.
- the modified Dda helicase of the invention may comprise any number and combination of the substitutions set out above.
- the modified Dda helicase of the invention may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44 or 45 of the substitutions above.
- the modified Dda helicase of the invention may further be one in which one or more of the positions corresponding to the 113, 22, or 21 amino acid positions in Dda 1993 listed above are modified or substituted as described above.
- the invention also provides a modified DNA dependent ATPase (Dda) helicase comprising one or more of the following substitutions or one or more of the following substitutions at the position(s) corresponding to the following amino acid positions in Dda 1993:
- T2N T2K, T2S, T2D, T2I, T2L, T2E, T2F, T2G, T2H, T2Q, T2R, T2A, T2M, T2P, or
- T2V preferably T2N
- E47D E47G, E47H, E47N, E47P, E47Q, E47S, E47T, E47A, or E47R, preferably E47D,
- E54I E54V, E54K, E54Y, E54T, E54L, E54N, E54A, E54F, E54M, E54D, E54H, E54P, E54Q, E54R, or E54S, preferably E54I,
- K86N a K86N, K86G, K86D, K86E, K86H, K86Q, K86R, K86S, K86T, K86M, or K86P, preferably K86N,
- N88K preferably N88K
- N88E preferably N88T
- N88S preferably N88V
- N88Q preferably N88A, N88L, N88D, N88G, N88M, N88I, or N88P, preferably N88K
- V90E V90D, V90K, V90Q, V90N, V90S, V90A, V90I, V90L, V90M, V90P, V90R, V90F, V90Y, V90H, or V90G, preferably V90D or V90E,
- T91E - T91E, T91D, T91R, T91K, T91Y, T91H, T91P, T91A, T91V, T91L, T91I, T91S, or T91M, preferably T91E,
- E93P E93D
- E93G E93A
- E93L E93K
- E93Q E93S
- E93T E93H
- E93N E93N
- E93R preferably E93P
- K101R K101T, K101S, K101V, K101E, K101I, K101G, K101L, K101M, K101P, K101Q, K101D, K101H, or K101N, preferably K101R,
- V103K V103N, V103L, V103I, V103T, V103R, V103Y, V103S, V103D, V103A, V103E, V103H, V103M, V103P, V103Q, or V103F, preferably V103K,
- G153E preferably G153E, G153S, G153D, G153T, G153N, G153K, G153A, G153Q, G153R, G153L, G153H, or G153P, preferably G153E,
- N155K, N155P, N155S, N155E, N155D, N155G, N155T, N155A, N155V, N155L, N155I, N155H, N155M, N155Q, or N155R preferably N155K, - W195F, W195Y, W195P, W195K, W195E, W195I, W195L, W195D, W195S, W195G, W195T, W195V, W195Q, W195N, W195R, W195M, or W195H, preferably W195Y, W195P, W195K, W195E, W195I, W195G, W195T, W195Q, W195N, W195R, W195M, or W195H, more preferably W195F or W195Y,
- I196P I196L, I196D, I196A, I196E, I196K, I196Q, I196S, I196T, I196F, I196M, I196V, or I196Y, preferably I196L or I196P,
- D198N D198P, D198E, D198G, D198H, D198K, D198Q, D198R, D198S, or D198T, preferably D198N, D198P, D198E, D198G, D198H, D198K, D198Q, D198R, or D198T, more preferably D198N,
- K243R, K243E, K243D K243A, K243S, K243M, K243N, K243P, K243T, or K243Y, preferably K243R or K243E,
- E258G E258K, E258N, E258P, E258D, E258S, E258A, E258H, E258Q, E258R, or E258T, preferably E258G, E258K, E258N, E258P, E258D, E258H, E258Q, E258R, or E258T, more preferably E258G,
- F276T preferably F276T, F276S, F276L, F276P, F276E, F276D, F276N, or F276Q, preferably F276T,
- I281E I281V, I281K, I281A, I281D, I281H, I281N, I281P, I281Q, I281R, I281S, I281T, I281F, I281L, I281M, or I281Y, preferably I281E,
- N292E N292A, N292S, N292D, N292K, N292H, N292R, N292Q, or N292T, preferably N292E,
- E301K, E301D, E301N, E301S, E301A, E301H, E301M, E301P, E301Q, E301R, or E301T preferably E301K
- Y335P Y335K, Y335E, Y335F, Y335T, Y335Q, Y335S, Y335R, Y335D, Y335V, Y335N, Y335G, Y335H, Y335M, or Y335W, preferably Y335P,
- Y336E Y336H, Y336K, Y336I, Y336G, Y336S, Y336T, Y336D, Y336V, Y336P, Y336N, Y336R, Y336Q, Y336A, Y336F, Y336M, or Y336W, preferably Y336E,
- L354K, L354E, L354D, L354V, L354Y, L354R, L354S, L354N, L354H, L354M, L354P, L354T, L354F, or L354I preferably L354K, - T359V, T359M, T359A, T359I, T359Y, T359F, T359E, T359K, T359N, T359D, T359P, T359Q, T359R, T359S, T359W, or T359G, preferably T359A, T359V, T359I or T359M, more preferably T359V,
- K364N K364Q, K364D, K364E, K364G, K364H, K364S, K364T, K364M, K364P, K364Y, K364I, K364L, or K364V, preferably K364N or K364Q,
- W366D W366N, W366K, W366A, W366S, W366I, W366E, W366R, W366M, W366T, W366Q, W366Y, W366G, W366F, W366V, W366P, or W366H, preferably W366D or W366N, and
- the modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising one or more of the 31 substitutions set out above.
- the modified Dda helicase of the invention may comprise any number and combination of the substitutions set out above.
- the modified Dda helicase of the invention may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, or 31 of the substitutions above.
- the modified Dda helicase of the invention may further be one in which one or more of the positions corresponding to the 113, 22, or 21 amino acid positions in Dda 1993 listed above are modified or substituted as described above.
- the modified Dda helicase of the invention preferably comprises one or more of the following substitutions or one or more of the following substitutions at the position(s) corresponding to the following amino acid positions in Dda 1993:
- I196P I196L, I196D, I196A, I196E, I196K, I196Q, I196S, I196T, I196F, I196M, I196V, or I196Y, preferably I196L or I196P,
- Y336E Y336H, Y336K, Y336I, Y336G, Y336S, Y336T, Y336D, Y336V, Y336P, Y336N, Y336R, Y336Q, Y336A, Y336F, Y336M, or Y336W, preferably Y336E,
- T359V, T359M, T359A, T359I, T359Y, T359F, T359E, T359K, T359N, T359D, T359P, T359Q, T359R, T359S, T359W, or T359G preferably T359A, T359V, T359I or T359M, more preferably T359V, K364N, K364Q, K364D, K364E, K364G, K364H, K364S, K364T, K364M, K364P,
- K364Y, K364I, K364L, or K364V preferably K364N or K364Q
- the modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising one or more of the five substitutions set out above.
- the modified Dda helicase of the invention may comprise any number and combination of these five substitutions.
- the modified Dda helicase of the invention may comprise 1, 2, 3, 4, or 5 of the substitutions.
- the modified Dda helicase of the invention may further be one in which one or more of the positions corresponding to the 113, 22 or 21 amino acid positions in Dda 1993 listed above are modified or substituted as described above.
- the helicase is preferably a variant of SEQ ID NO: 118 which comprises substitutions at:
- W195/K358/Q422 such as W195F/K358I/Q422K, W195R/K358M/Q422N, W195K/K358P/Q422D, W195Y/K358I/Q422K, W195K/K358I/Q422N, W195F/K358M/Q422K, W195Y/K358G/Q422A, W195I/K358I/Q422K, W195F/K358I/Q422D or W195I/K358G/Q422A;
- W195/K364/K368 such as W195K/K364D/K368Q, W195Y/K364Q/K368A, W195I/K364Q/K368A, W195R/K364D/K368T, W195Y/K364I/K368T, W195F/K364Q/K368Q, W195R/K364Q/K368A, W195I/K364P/K368T, W195F/K364Q/K368A or W195R/K364I/K368Y;
- W195/K364/Q422 such as W195S/K364I/Q422A, W195Y/K364I/Q422A, W195F/K364I/Q422D, W195Y/K364P/Q422N, W195S/K364P/Q422A, W195R/K364I/Q422A, W195Y/K364Q/Q422K, W195F/K364Q/Q422K, W195K/K364P/Q422N or W195R/K364D/Q422D;
- W195/K368/Q422 such as W195Y/K368A/Q422K, W195K/K368Q/Q422N, W195F/K368A/Q422K, W195S/K368A/Q422K, W195F/K368Q/Q422D, W195Y/K368T/Q422K, W195V/K368T/Q422A, W195K/K368V/Q422K, W195V/K368T/Q422D or W195S/K368V/Q422A;
- W195/Y350/K368 such as W195Y/Y350E/K368A, W195Y/Y350D/K368T, W195K/Y350I/K368P, W195R/Y350S/K368Q, W195I/Y350D/K368I, W195S/Y350K/K368T, W195R/Y350F/K368V, W195F/Y350E/K368A W195F/Y350I/K368T or W195K/Y350W/K368Q; K67/M401, such as K67M/M401L, K67A/M401V, K67Q/M401S or K67N/M401W;
- G153/M401 such as G153E/M401S, G153Q/M401G, G153E/M401V or G153L/M401W;
- G153/V403 such as G153E/V403F, G153L/V403L, G153Q/V403M or G153N/V403H;
- M401/V403 such as M401S/V403I, M401L/V403H, M401V/V403F or M401R/V403A;
- K67/G153 such as K67M/G153L, K67A/G153E, K67L/G153H or K67Q/G153A;
- K67/K368 such as K67M/K368Q, K67N/K368E, K67Q/K368R or K67A/K368A;
- K368/M401 such as K368A/M401V, K368E/M401S, K368Q/M401G or K368R/M401W;
- K368/V403 such as K368N/V403A, K368P/V403H, K368A/V403F or K368F/V403L;
- A60/K368 such as A60K/K368N, A60T/K368A, A60L/K368E or A60D/K368R;
- K67/V403 such as K67M/V403H, K67L/V403I, K67N/V403A or K67A/V403F;
- G153/W195/M401 such as G153E/W195Q/M401G, G153N/W195K/M401L, G153Q/W195F/M401S, G153N/W195Q/M401G, G153E/W195Q/M401L, G153L/W195Q/M401G, G153E/W195F/M401S, G153Q/W195Q/M401L, G153L/W195K/M401G or G153N/W195R/M401W;
- K25/P134 such as K25N/P134T, K25D/P134G, K25A/P134D or K25V/P134E;
- P134/W135/C136 such as P134E/W135N/C136M, P134K/W135A/C136F, P134G/W135A/C136T, P134E/W135A/C136F, P134K/W135N/C136F, P134K/W135D/C136M, P134E/W135D/C136V, P134K/W135N/C136T, P134T/W135A/C136M or P134G/W135F/C136F;
- W195/V220 such as W195K/V220A, W195R/V220P, W195F/V220K or W195I/V220Y;
- W195/Q422 such as W195F/Q422K, W195I/Q422N, W195P/Q422A or W195K/Q422D; or
- V220/Q422 such as V220P/Q422A, V220I/Q422D, V220A/Q422T or V220K/Q422K.
- Preferred combinations in SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 include the combinations of amino acids which correspond to the combinations in SEQ ID NO: 118 listed above.
- the invention also provides a modified DNA dependent ATPase (Dda) helicase, wherein the helicase comprises one or more modifications or substitutions at one or more positions in one or more of the (i) tower domain, (ii) 1A domain and (iii) 2A domain and wherein the helicase displays one or more of (a) an increased speed, (b) an improved accuracy and (c) a decreased speed spread ratio when used in methods of controlling the movement of an analyte through a transmembrane pore and/or in methods of characterising an analyte using a transmembrane pore.
- the modified Dda helicase of the invention may comprise any number and combination of the modified or substitutions set out above.
- the modified Dda helicase of the invention may comprise one or more modifications or substitutions at one or more positions in (i), (ii), (iii), (i) and (ii), (i) and (iii), (ii) and (iii) or (i), (ii) and (iii).
- the modified Dda helicase of the invention may display (a), (b), (c), (a) and (b), (a) and (c), (b) and (c) or (a), (b) and (c).
- the one or more positions in the 2A domain may be proximal to motif V/Va.
- the one or more positions in the 1A domain may be in motif la.
- Motif V and motif la are helicase motifs associated with energy transduction from ATP hydrolysis to DNA translocation.
- the domains (i)-(iii) are defined in more detail below.
- the modified Dda helicase of the invention may comprise one or more modifications or substitutions at any of the positions described above or below and/or one or more of the substitutions described above or below.
- Any of the modified Dda helicases of the invention or variants of SEQ ID NO: 118 discussed above may further comprise the substitution MIG.
- modified Dda helicase of the invention or variant of SEQ ID NO: 118 may further comprise any of the modifications, mutations or substitutions discussed below.
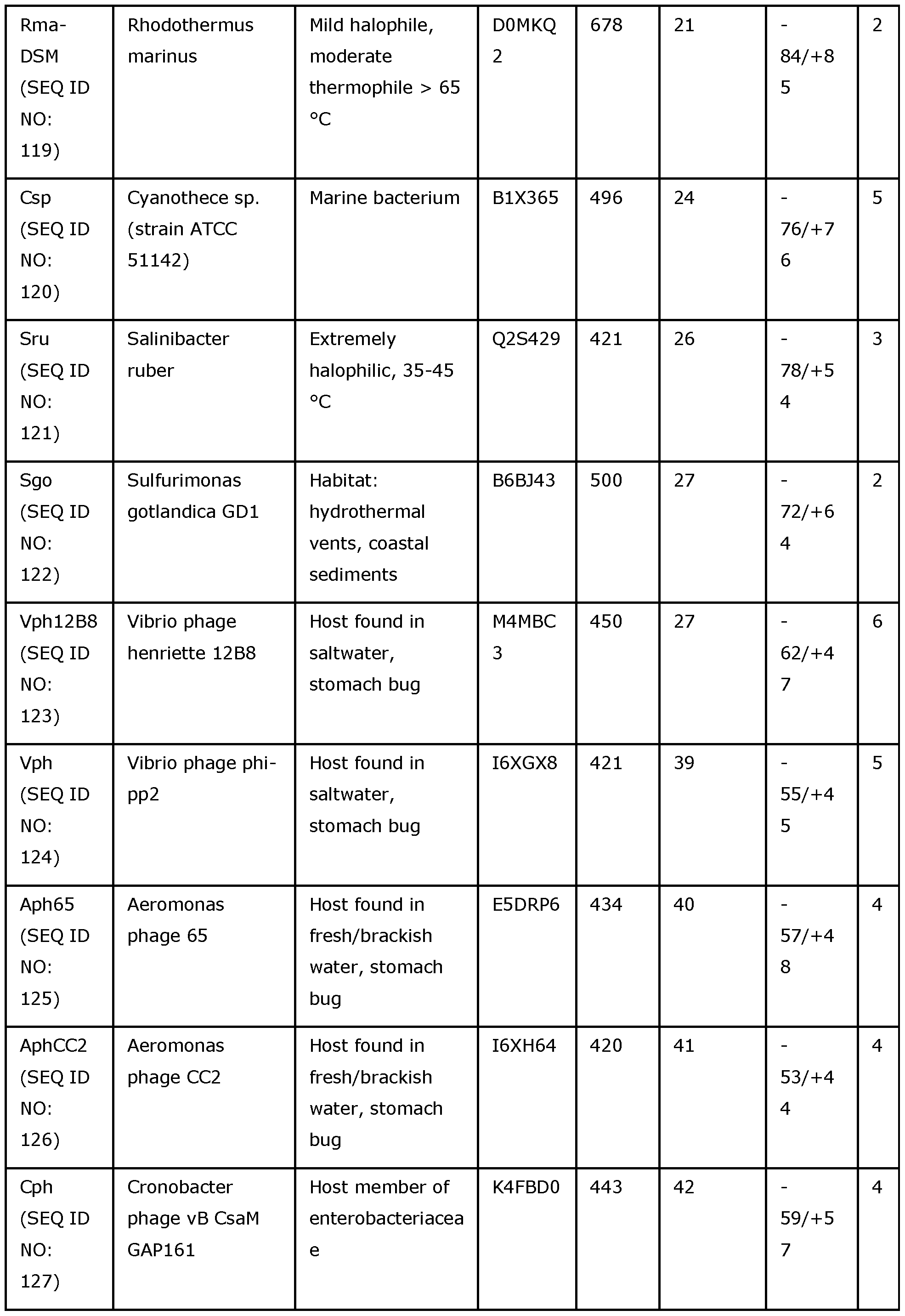
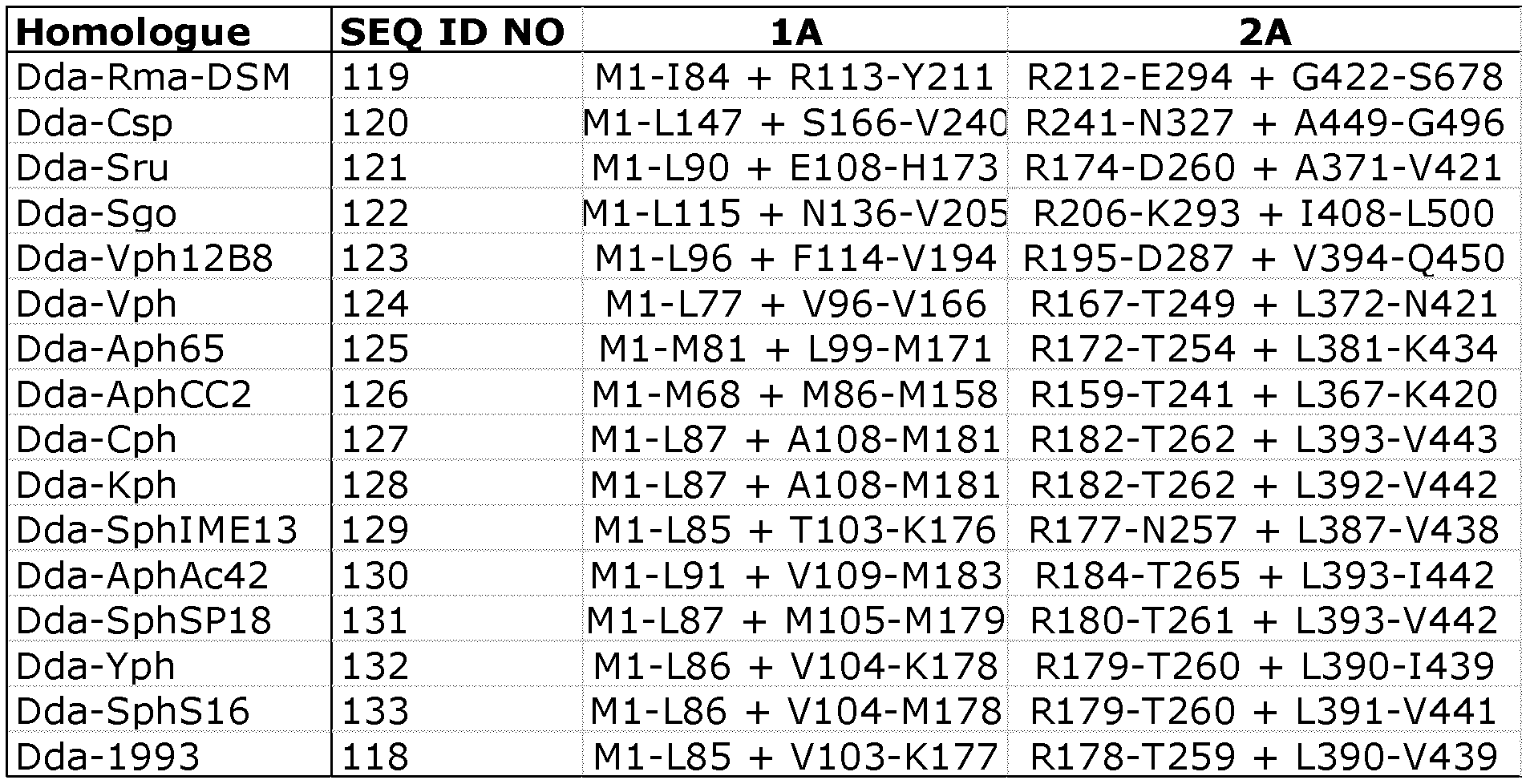
- the Dda helicase that is modified in accordance with the invention may be any of SEQ ID NOs: 118 to 133.
- SEQ ID NO: 118 is Dda 1993.
- the modified Dda helicase preferably comprises a variant of any of SEQ ID NOs: 118 to 133.
- the variant may have any % of the sequence homologies/identities to any of SEQ ID NOs: 118 to 113 set out below.
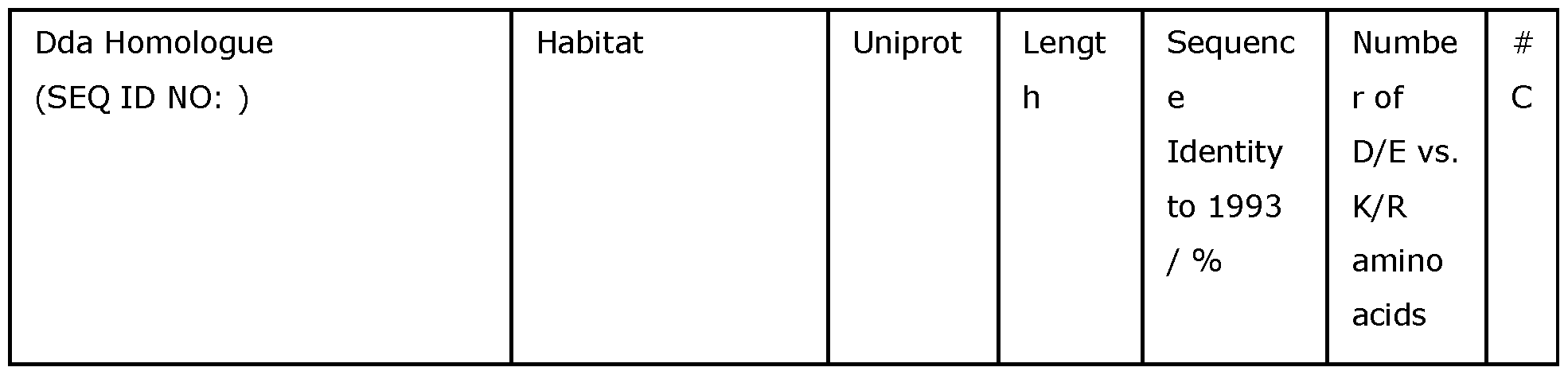
- Table 1 summarises the preferred Dda helicases which may be modified in accordance with the invention.
- the modified helicase of the invention may further be one in which one or more of the positions corresponding to amino acid positions 55, 114, 156, 177, 210, 221, 350 and 358 in Dda 1993 are modified or substituted.
- the positions corresponding to positions 55, 114, 156, 177, 210, 221, 350 and 358 in Dda 1993 are discussed in more detail below.
- Positions 55, 114, 156 and 177 are in the 1A domain of Dda 1993.
- Positions 210 and 221 are in the 2A domain of Dda 1993.
- Positions 350 and 358 are in the tower domain of Dda 1993.
- the modified Dda helicase of the invention may further comprise a modification or substitution at any number and combination of the positions corresponding to amino acid positions (a) 55, (b) 114, (c) 156, (d) 177, (e) 210, (f) 221, (g) 350 and (h) 358, including at (a); (b);
- the modified Dda helicase of the invention may further be one in which one or more of the positions corresponding to amino acid positions 114, 177, 350 and 358 in Dda 1993 are modified or substituted.
- the corresponding to positions 114, 177, 350 and 358 in Dda 1993 are discussed in more detail below.
- Positions 114 and 177 are in the 1A domain of Dda 1993.
- Positions Y350 and K358 are in the tower domain of Dda 1993.
- the modified Dda helicase of the invention may further comprise a modification or substitution at any number and combination of the positions corresponding to amino acid positions (a) 114, (b) 177, (c) 350 and (d) 358 in Dda 1993, including at (a); (b); (c); (d); (a) and (b); (a) and (c); (a) and (d); (b) and (c); (b) and (d); (c) and (d); (a), (b) and (c); (a), (b) and (d); (a), (c) and
- the position corresponding to amino acid position 55 in Dda 1993 is preferably substituted with D, E, K, N or S.
- the position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with K (T55K). These substitutions increase the speed and increase the accuracy when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821). These substitutions also decrease the normalised speed distribution when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
- the position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with A, V, I, L, M, F, Y, W, G, P, S, T, N or Q.
- the position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with A, G, I, L, M, P, S, T or V.
- the position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with G, L, S or T. These substitutions decrease the speed when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821).
- the position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with A, I, M, P, or V.
- substitutions increase the speed when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821).
- the position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with G (C11G). This substitution decreases the speed and increases the accuracy when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821).
- the position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with I or P. These substitutions increase the speed and decrease the accuracy when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821).
- the position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with G, I or P. These substitutions decrease the normalised speed distribution when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821).
- the position corresponding to amino acid position 114 in Dda 1993 is most preferably substituted with I (Cl 141).
- the position corresponding to amino acid position 156 in Dda 1993 is preferably substituted with A, E, F, G, I, L, M, P, S, V, Y, D, K or N.
- the position corresponding to amino acid position 156 in Dda 1993 is preferably substituted with F (T156F).
- F F
- This substitution increases the speed and increases the accuracy when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
- This substitution also decreases the normalised speed distribution when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
- the position corresponding to amino acid position 177 in Dda 1993 is preferably substituted with D, E, F, G, H, I, L, M, N, Q, R, S, T, V, W or Y.
- the position corresponding to amino acid position 177 in Dda 1993 is preferably substituted with F, G, S, V, W or Y. These substitutions decrease the speed when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821).
- the position corresponding to amino acid position 177 in Dda 1993 is preferably substituted with D, E, G, H, I, L, M, N, Q, R, or T.
- substitutions increase the speed when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821).
- the position corresponding to amino acid position 177 in Dda 1993 is preferably substituted with F, H, I, L, M, N or W. These substitutions decrease the accuracy and the normalised speed distribution when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). They have different effects on the speed (Example 2 of PCT/EP2023/059821).
- the position corresponding to amino acid position 177 in Dda 1993 is preferably substituted with N (K177N).
- the position corresponding to amino acid position 210 in Dda 1993 is preferably substituted with D, E, K, S, N, R, H or Y.
- the position corresponding to amino acid position 210 in Dda 1993 is preferably substituted with R (T210R), H (T210H) or K (T210K).
- the position corresponding to amino acid position 210 in Dda 1993 is preferably substituted with K (T210K).
- This substitution increases the speed and increases the accuracy when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
- This substitution also decreases the normalised speed distribution when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
- the position corresponding to amino acid position 221 in Dda 1993 is preferably substituted with D, K, E, Q, R, A, H, L, T or Y.
- the position corresponding to amino acid position 221 in Dda 1993 is preferably substituted with D (N221D) or E (N221E).
- the position corresponding to amino acid position 221 in Dda 1993 is preferably substituted with E (N221E).
- This substitution increases the speed and increases the accuracy when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
- This substitution also decreases the normalised speed distribution when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
- the position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with D, E, A, V, I, L, M, F, W, R, H, K, L, S, T, N or Q.
- the position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with I, F, W or S.
- the position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with I or S (Y350I or Y350S).
- the position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with I (Y350I). This substitution increases the speed and decreases the accuracy and normalised speed distribution when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821).
- the position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with I or S (Y350I or Y350S). These substitutions have the effects shown in Example 4 of PCT/EP2023/059821 when used with a pore complex described therein.
- the position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with A, D, E, G, K, L, N, Q, R, T, V, H or M.
- the position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with D (Y350D) or E (Y350E).
- the position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with E (Y350E).
- the position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with D, E, A, V, I, L, M, F, Y, W, R, H, L, S, T, N or Q.
- the position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with E, I, L or M. These substitutions decrease the speed when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821).
- the position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with I or M. These substitutions decrease the speed and increase the accuracy when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821).
- the position corresponding to amino acid position 358 in Dda 1993 is preferably substituted M (K358M).
- the position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with A, E, F, I, M or S. These substitutions increase the accuracy when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821).
- the position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with A, E, F, I or M. These substitutions decrease the speed when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821).
- the position corresponding to amino acid position 358 in Dda 1993 is preferably substituted S (K358S).
- This substitution increases the speed when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821).
- the position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with A, E, I, M or S. These substitutions decrease the normalised speed distribution when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821).
- the position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with (K358F). These substitutions increase the normalised speed distribution when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821).
- Example 3 uses a CsgG pore complex containing a CsgF peptide.
- the position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with I, L or Q. These substitutions decrease the speed and increase the accuracy and normalised speed distribution when used to characterise a polynucleotide analyte (Example 4 of PCT/EP2023/059821).
- the position corresponding to amino acid position 358 in Dda 1993 is most preferably substituted with I (K358I).
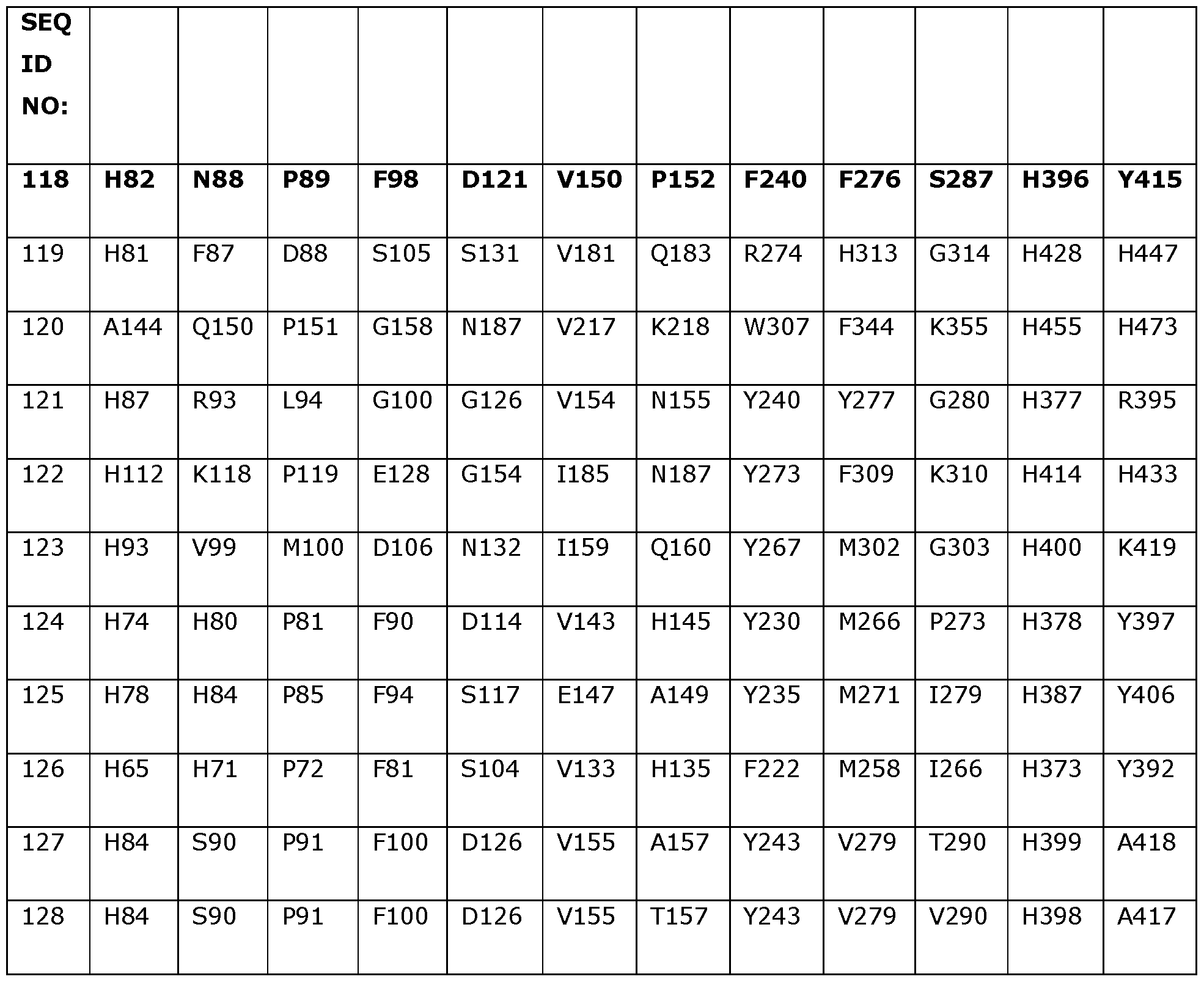
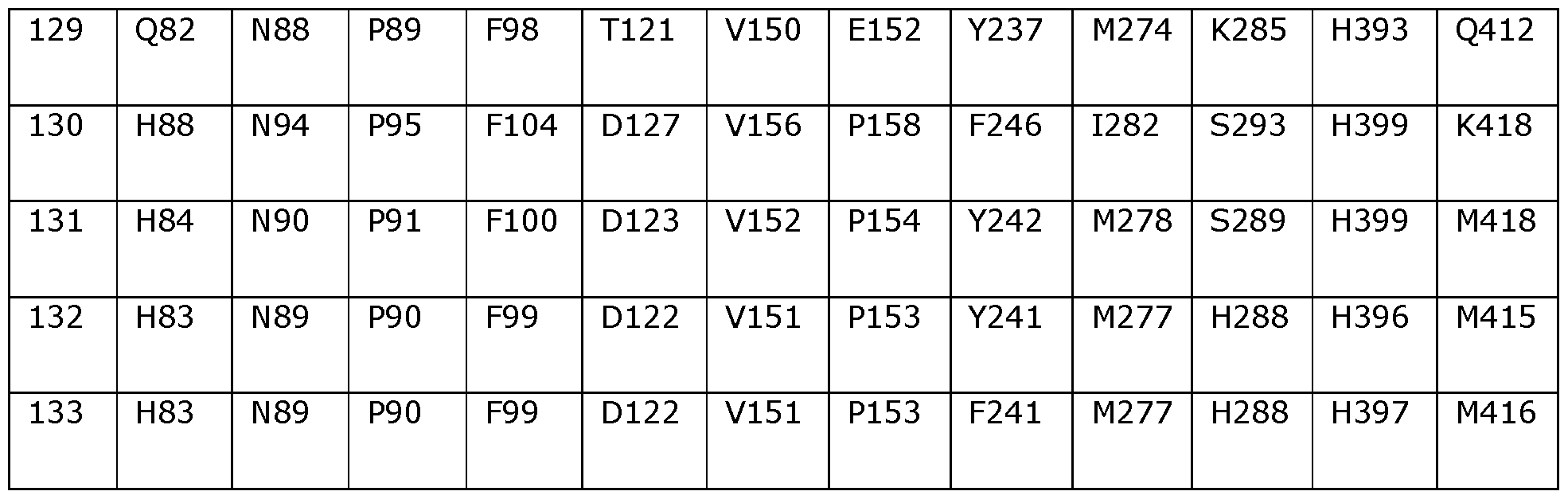
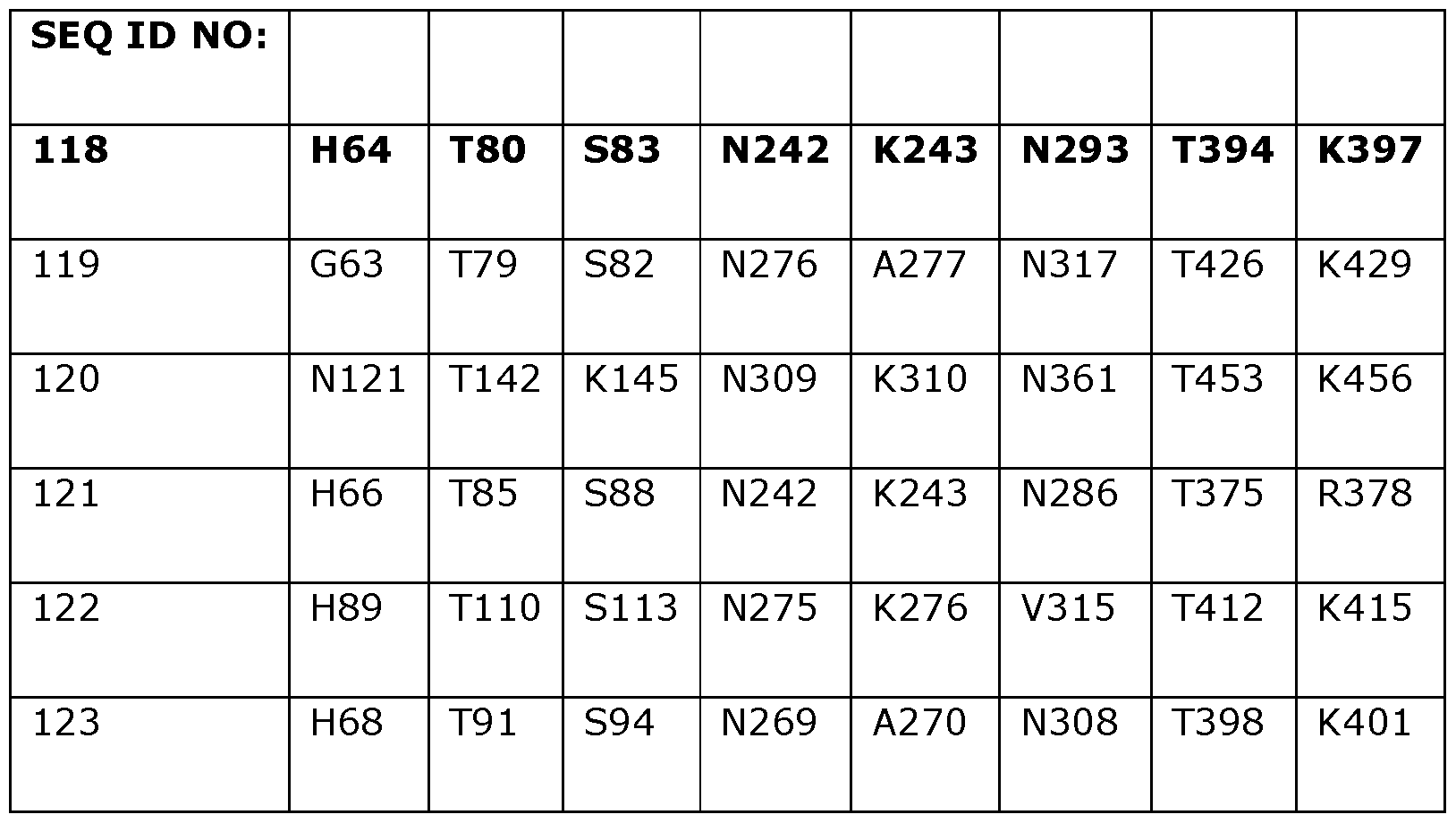
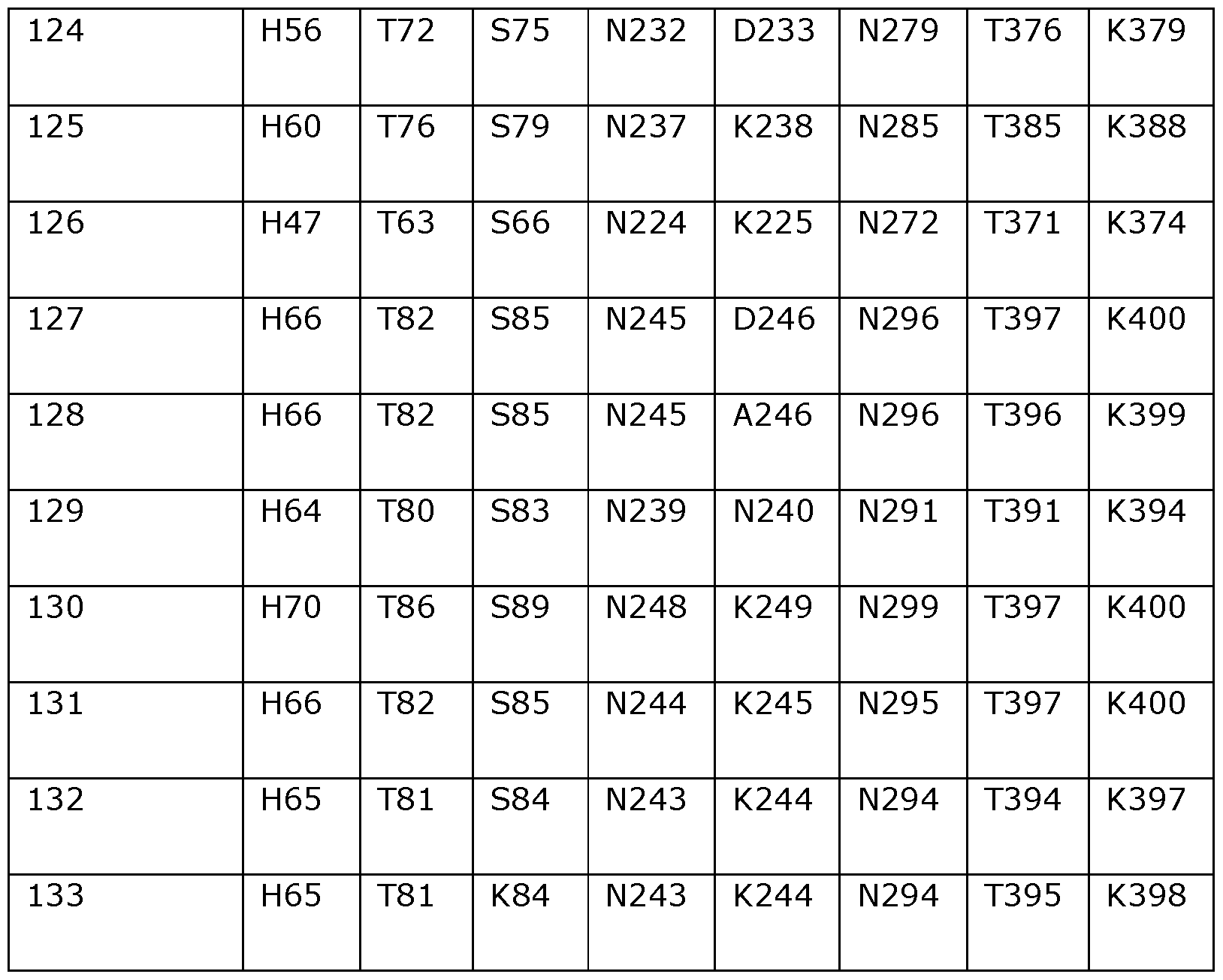
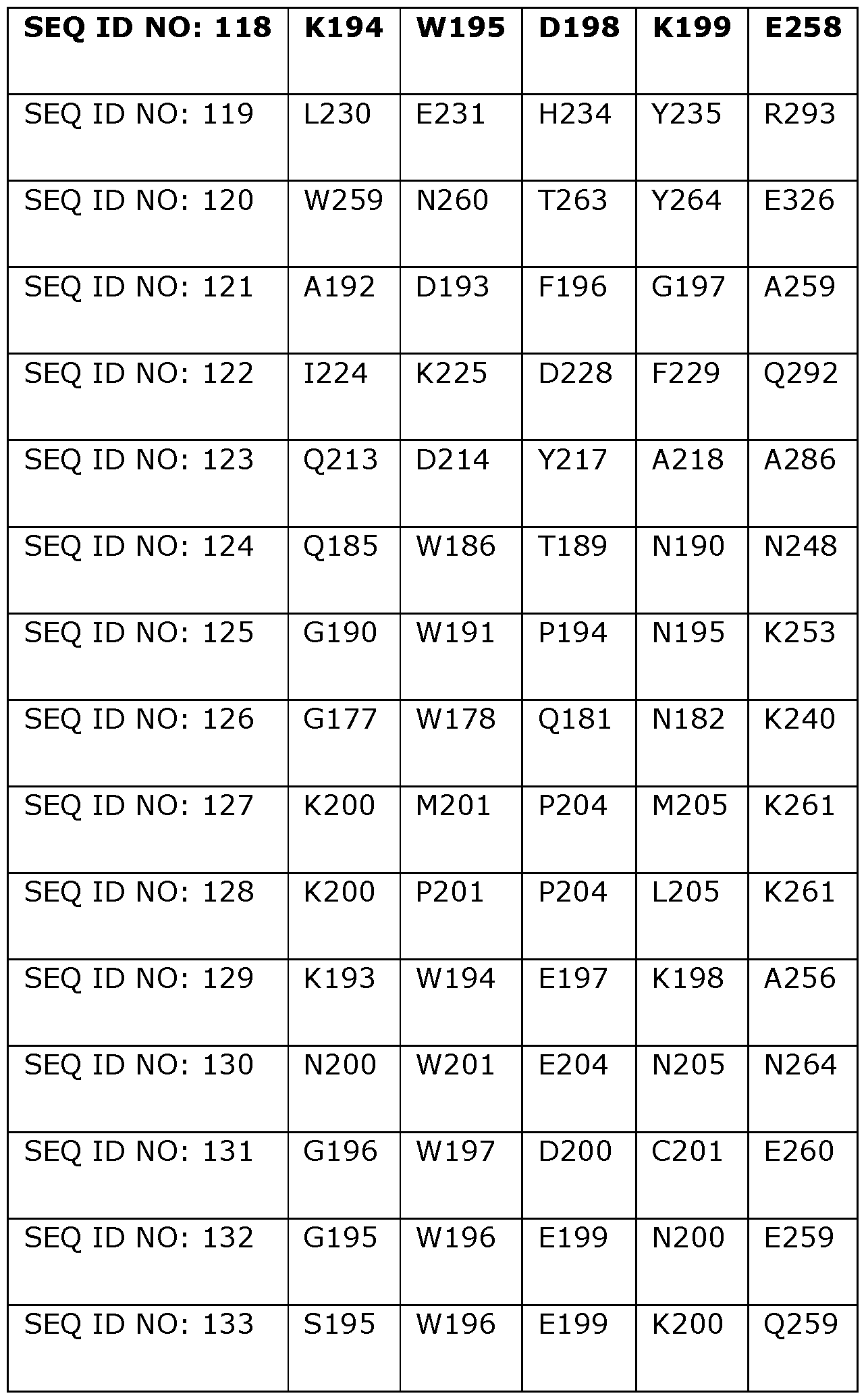
- Table 2 shows the amino acids in SEQ ID NOs: 119 to 133 which correspond to positions 40, 55, 114, 156, 177, 210, 221, 350 and 358 in SEQ ID NO: 118.
- the modified helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising one or more of (a)-(h) as follows:
- K177D, K177E, K177G, K177H, K177I, K177L, K177M, K177N, K177Q, K177R, or K177T
- T210R T210H or T210Y, or
- K358A, K358E, K358F, K358I or K358M are K358A, K358E, K358F, K358I or K358M,
- K358I The variant may include any combination and permutation of (a)-(h) as set out above.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising one or more of (a), (b), (c) and (d) as follows:
- K177D, K177E, K177G, K177H, K177I, K177L, K177M, K177N, K177Q, K177R, or K177T
- K358A, K358E, K358F, K358I or K358M are K358A, K358E, K358F, K358I or K358M,
- the variant may include (a); (b); (c); (d); (a) and (b); (a) and (c); (a) and (d); (b) and
- a preferred variant of SEQ ID NO: 118 comprises: C114I; K177M; Y350I; K358I; C114I and K177M; C114I and Y350I; C114I and K358I; K177M and Y350I; K177M and K358I; Y350I and K358I; C114I, K177M and Y350I; C114I, K177M and K358I; C114I, Y350I and K358I; K177M, Y350I and K358I; or C114I, K177M, Y350I and K358I.
- the helicase preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein one or more of the positions corresponding to amino acid positions 55, 114, 156, 177, 210, 221, 350 and 358 in Dda 1993 are modified or substituted as defined above (including specific substitutions).
- Various combinations and permutations of one or more of positions 55, 114, 156, 177, 210, 221, 350 and 358 in Dda 1993 are defined above with reference to (a)-(h).
- the helicase preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein one or more of the positions corresponding to amino acid positions 114, 177, 350 and 358 in Dda 1993 are modified or substituted as defined above (including specific substitutions).
- Various combinations and permutations of one or more of positions 114, 177, 350 and 358 in Dda 1993 are defined above with reference to (a)-(d).
- the helicase preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein one or more of the positions corresponding to amino acid positions 114, 177 and 358 in Dda 1993 are modified or substituted as defined above (including specific substitutions).
- any of the modified helicases of the invention may further comprise a modification or substitution at the position corresponding to amino acid position 40 in Dda 1993.
- Position 40 or the corresponding position may be substituted with as A, V, I, L, M, F, Y or W.
- Positions which correspond to position T40 in Dda 1993 are shown in Table 2 above.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which, in addition to the modifications/substitution set out above, further comprises a substitution at T40, such as T40A, T40V, T40I, T40L, T40M, T40F, T40Y or T40W.
- the substitution is preferably T40Y.
- the modified Dda helicase of the invention may further be one in which the position corresponding to amino acid position 40 in Dda 1993 is modified or substituted.
- Position T40 is in the tower domain of Dda 1993.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at T40, such as T40A, T40V, T40I, T40L, T40M, T40F, T40Y or T40W.
- the substitution is preferably T40Y.
- the modified Dda helicase of the invention may further comprise a modification or substitution at one or more of the positions corresponding to amino acid positions (a) 55, (b) 114, (c) 156, (d) 177, (e) 210, (f) 221, (g) 350 and (h) 358, including any of the combinations and permutations of (a)-(h) set out above.
- the modified Dda helicase of the invention may further comprise a modification or substitution at one or more of the positions corresponding to amino acid positions (a) 114, (b) 177, (c) 350 and (d) 358 in Dda 1993, including at (a); (b); (c); (d); (a) and (b); (a) and (c); (a) and (d); (b) and (c); (b) and
- the helicase preferably comprises a variant of SEQ ID NO: 119, 120, 121,
- modified helicases of the invention may further comprise any of the modifications, substitutions, combinations of modifications or combination of substitutions discussed below.
- the modified helicases of the invention provide more consistent movement of the target analyte with respect to, such as through, the transmembrane pore leading to improved accuracy.
- the helicases preferably provide more consistent movement from one k-mer to another or from k-mer to k-mer as the target analyte, such as polynucleotide, moves with respect to, such as through, the pore.
- the helicases allow the target analyte, such as target polynucleotide, to move with respect to, such as through, the transmembrane pore more smoothly.
- the helicases preferably provide more regular or less irregular movement of the target analyte, such as target polynucleotide, with respect to, such as through, the transmembrane pore.
- the modification(s) typically increase accuracy by at least 0.1%, at least 0.5%, at least 1%, at least 5%, at least 10%, at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80% or at least 90% compared to a helicase without the modification.
- the modified helicase has the ability to control the movement of a polynucleotide.
- the ability of a helicase to control the movement of a polynucleotide can be assayed using any method known in the art. For instance, the helicase may be contacted with a polynucleotide and the position of the polynucleotide may be determined using standard methods.
- the ability of a modified helicase to control the movement of a polynucleotide is typically assayed in a nanopore system, such as the ones described below and, in particular, as described in the Examples.
- a modified helicase of the invention may be isolated, substantially isolated, purified or substantially purified.
- a helicase is isolated or purified if it is completely free of any other components, such as lipids, polynucleotides, pore monomers or other proteins.
- a helicase is substantially isolated if it is mixed with carriers or diluents which will not interfere with its intended use.
- a helicase is substantially isolated or substantially purified if it is present in a form that comprises less than 10%, less than 5%, less than 2% or less than 1% of other components, such as lipids, polynucleotides, pore monomers or other proteins.
- Dda helicase may be modified in accordance with the invention.
- Preferred Dda helicases are discussed below and described in WO2015/055981, WO2015/166276 and WO2016/055777 (all incorporated by reference).
- Dda helicases typically comprises the following five domains: 1A (RecA-like motor) domain, 2A (RecA-like motor) domain, tower domain, pin domain and hook domain (Xiaoping He et al., 2012, Structure; 20: 1189-1200).
- the domains may be identified using protein modelling, x-ray diffraction measurement of the protein in a crystalline state (Rupp B (2009). Biomolecular Crystallography: Principles, Practice and Application to Structural Biology. New York: Garland Science.), nuclear magnetic resonance (NMR) spectroscopy of the protein in solution (Mark Rance; Cavanagh, John; Wayne J. Fairbrother; Arthur W. Hunt III; Skelton, NNicholas J. (2007).
- modified helicase of the invention preferably further comprises any of the following additional modifications, substitutions, combinations of modifications or combination of substitutions.
- the helicase of the invention may be one in which at least one amino acid which interacts with a transmembrane pore is further substituted. Any number of amino acids may substituted, such as 1 or more, 2 or more, 3 or more, 4 or more, 5 or more or 6 or more amino acids. 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more amino acids may be substituted.
- the amino acids which interact with a transmembrane pore can be identified using protein modelling as discussed above.
- the helicase of the invention is preferably one in which at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in single stranded DNA (ssDNA) is further substituted with an amino acid which comprises a larger side chain (R group). Any number of amino acids may substituted, such as 1 or more, 2 or more, 3 or more, 4 or more, 5 or more or 6 or more amino acids. Each amino acid may interact with the base, the sugar or the base and the sugar.
- the amino acids which interact with the sugar and/or base of one or more nucleotides in single stranded DNA can be identified using protein modelling as discussed above.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 wherein the at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in ssDNA is at least one of H82, N88, P89, F98, D121, V150, P152, F240, F276, S287, H396 and Y415.
- These numbers correspond to the relevant positions in SEQ ID NO: 118 and may need to be altered in the case of variants where one or more amino acids have been inserted or deleted compared with SEQ ID NO: 118.
- a skilled person can determine the corresponding positions in a variant as discussed above.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 wherein the at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in ssDNA is F98 and one or more H82, N88, P89, D121, V150, P152, F240, F276, S287, H396 and Y415, such as F98/H82, F98/N88, F98/P89, F98/D121, F98/V150, F98/ P152, F98/F240, F98/F276, F98/S287 or F98/H396.
- the helicase of the invention is preferably a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein the at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in ssDNA is at least one of the amino acids which correspond to H82, N88, P89, F98, D121, V150, P152, F240, F276, S287, H396 and Y415in SEQ ID NO: 118.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein the at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in ssDNA is the amino acid which corresponds to F98 in SEQ ID NO: 118 and one or more of the amino acids which correspond to H82, N88, P89, D121, V150, P152, F240, F276, S287, H396 and Y415 in SEQ ID NO: 118, such as the amino acids which correspond to F98/H82, F98/N88, F98/P89, F98/D121, F98/V150, F98/ P152, F98/F240, F98/F276, F98/S287 or F98/H396.
- the at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in ssDNA is preferably at least one amino acid which intercalates between the nucleotides in ssDNA.
- Amino acids which intercalate between nucleotides in ssDNA can be modeled as discussed above.
- the at least one amino acid which intercalates between the nucleotides in ssDNA is preferably at least one of P89, F98 and V150 in SEQ ID NO: 118, such as P89, F98, V150, P89/F98, P89/V150, F98/V150 or P89/F98/V150.
- the at least one amino acid which intercalates between the nucleotides in ssDNA in SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 is preferably at least one of the amino acids which correspond to P89, F98 and V150 in SEQ ID NO: 118, such as P89, F98, V150, P89/F98, P89/V150, F98/V150 or P89/F98/V150. Corresponding amino acids are shown in Table 3 above.
- the larger side chain (R group) preferably (a) contains an increased number of carbon atoms, (b) has an increased length, (c) has an increased molecular volume and/or (d) has an increased van der Waals volume.
- the larger side chain (R group) preferably (a); (b); (c); (d); (a) and (b); (a) and (c); (a) and (d); (b) and (c); (b) and (d); (c) and (d); (a), (b) and (c); (a), (b) and (d); (a), (c) and (d); (b), (c) and (d); or (a), (b), (c) and (d).
- Each of (a) to (d) may be measured using standard methods in the art.
- the larger side chain (R group) preferably increases the (i) electrostatic interactions (ii) (ii) hydrogen bonding and/or (iii) cation-pi (cation-n) interactions between the at least one amino acid and the one or more nucleotides in ssDNA, such as increases (i); (ii); (iii); (i) and (ii); (i) and (iii); (ii) and (iii); and (i), (ii) and (iii).
- R group increases any of these interactions. For instance in (i), positively charged amino acids, such as arginine (R), histidine (H) and lysine (K), have R groups which increase electrostatic interactions.
- amino acids such as asparagine (N), serine (S), glutamine (Q), threonine (T) and histidine (H) have R groups which increase hydrogen bonding.
- aromatic amino acids such as phenylalanine (F), tryptophan (Wl), tyrosine (Y) or histidine (H) have R groups which increase cation-pi (cation-n) interactions.
- Specific substitutions below are labelled (i) to (iii) to reflect these changes.
- Other possible substitutions are labelled (iv). These (iv) substitutions typically increase the length of the side chain (R group).
- the amino acid which comprises a larger side chain (R) may be a non-natural amino acid.
- the non-natural amino acid may be any of those discussed below.
- the amino acid which comprises a larger side chain (R group) is preferably not alanine (A), cysteine (C), glycine (G), selenocysteine (U), methionine (M), aspartic acid (D) or glutamic acid (E).
- Histidine (H) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q) or asparagine (N) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W). Histidine (H) is more preferably substituted with (a) N, Q or W or (b) Y, F, Q or K.
- Asparagine (N) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q) or histidine (H) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W). Asparagine (N) is more preferably substituted with R, H, W or Y.
- Proline (P) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N), threonine (T) or histidine (H), (iii) tyrosine (Y), phenylalanine (F) or tryptophan (W) or (iv) leucine (L), valine (V) or isoleucine (I).
- Proline (P) is more preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N), threonine (T) or histidine (H), (iii) phenylalanine (F) or tryptophan (W) or (iv) leucine (L), valine (V) or isoleucine (I).
- Proline (P) is more preferably substituted with (a) F, (b) L, V, I, T or F or (c) W, F, Y, H, I, L or V.
- Valine (V) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H), (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W) or (iv) isoleucine (I) or leucine (L).
- Valine (V) is more preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H), (iii) tyrosine (Y) or tryptophan (W) or (iv) isoleucine (I) or leucine (L).
- Valine (V) is more preferably substituted with I or H or I, L, N, W or H.
- Phenylalanine (F) is preferably substituted with (i) arginine (R) or lysine (K), (ii) histidine (H) or (iii) tyrosine (Y) or tryptophan (W). Phenylalanine (F) is more preferably substituted with (a) W, (b) W, Y or H, (c) W, R or K or (d) K, H, W or R.
- Glutamine (Q) is preferably substituted with (i) arginine (R) or lysine (K) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W).
- Alanine (A) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H), (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W) or (iv) isoleucine (I) or leucine (L).
- Serine (S) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H), (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W) or (iv) isoleucine (I) or leucine (L).
- Serine (S) is preferably substituted with K, R, W or F
- Lysine (K) is preferably substituted with (i) arginine (R) or (iii) tyrosine (Y) or tryptophan (W).
- Arginine (R) is preferably substituted with (iii) tyrosine (Y) or tryptophan (W).
- Methionine (M) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W).
- Leucine (L) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q) or asparagine (N) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W).
- Aspartic acid (D) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W). Aspartic acid (D) is more preferably substituted with H, Y or K.
- Glutamic acid (E) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W).
- Isoleucine (I) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H), (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W) or (iv) leucine (L).
- Tyrosine (Y) is preferably substituted with (i) arginine (R) or lysine (K) or (iii) tryptophan (W). Tyrosine (Y) is more preferably substituted with W or R.
- the helicase more preferably comprises a variant of SEQ ID NO: 118 and comprises (a) P89F, (b) F98W, (c) V150I, (d) V150H, (e) P89F and F98W, (f) P89F and V150I, (g) P89F and V150H, (h) F98W and V150I, (i) F98W and V150H (j) P89F, F98W and V150I or (k) P89F, F98W and V150H.
- the helicase more preferably comprises a variant of SEQ ID NO: 118 which comprises: H82N; H82Q; H82W; N88R; N88H; N88W; N88Y; P89L; P89V; P89I ; P89E; P89T; P89F; D121H; D121Y; D121K; V150I; V150L; V150N; V150W; V150H; P152W; P152F; P152Y; P152H; P152I; P152L; P152V; F240W; F240Y; F240H; F276W; F276R; F276K; F276H; S287K; S287R; S287W; S287F; H396Y; H396F; H396Q; H396K; Y415W; Y415R; F98W/H82N; F98W/H82Q; F98W/H82W; F98W/N88R; F98W
- the helicase of the invention is preferably one in which at least one amino acid which interacts with one or more phosphate groups in one or more nucleotides in ssDNA is further substituted. Any number of amino acids may be substituted, such as 1 or more, 2 or more, 3 or more, 4 or more, 5 or more or 6 or more amino acids. Nucleotides in ssDNA each comprise three phosphate groups. Each amino which is substituted may interact with any number of the phosphate groups at a time, such as one, two or three phosphate groups at a time. The amino acids which interact with one or more phosphate groups can be identified using protein modelling as discussed above.
- substitution preferably increases the (i) electrostatic interactions, (ii) hydrogen bonding and/or (iii) cation-pi (cation-n) interactions between the at least one amino acid and the one or more phosphate groups in ssDNA.
- Preferred substitutions which increase (i), (ii) and (iii) are discussed below using the labelling (i), (ii) and (iii).
- the substitution preferably increases the net positive charge of the position.
- the net charge at any position can be measured using methods known in the art. For instance, the isolectric point may be used to define the net charge of an amino acid. The net charge is typically measured at about 7.5.
- the substitution is preferably the substitution of a negatively charged amino acid with a positively charged, uncharged, non-polar or aromatic amino acid.
- a negatively charged amino acid is an amino acid with a net negative charge.
- Negatively charged amino acids include, but are not limited to, aspartic acid (D) and glutamic acid (E).
- a positively charged amino acid is an amino acid with a net positive charge.
- the positively charged amino acid can be naturally-occurring or non-naturally- occurring.
- the positively charged amino acid may be synthetic or modified.
- modified amino acids with a net positive charge may be specifically designed for use in the invention.
- a number of different types of modification to amino acids are well known in the art.
- Preferred naturally-occurring positively charged amino acids include, but are not limited to, histidine (H), lysine (K) and arginine (R).
- the uncharged amino acid, non-polar amino acid or aromatic amino acid can be naturally occurring or non-naturally-occurring. It may be synthetic or modified.
- Uncharged amino acids have no net charge. Suitable uncharged amino acids include, but are not limited to, cysteine (C), serine (S), threonine (T), methionine (M), asparagines (N) and glutamine (Q).
- Non-polar amino acids have non-polar side chains.
- Suitable non-polar amino acids include, but are not limited to, glycine (G), alanine (A), proline (P), isoleucine (I), leucine (L) and valine (V).
- Aromatic amino acids have an aromatic side chain. Suitable aromatic amino acids include, but are not limited to, histidine (H), phenylalanine (F), tryptophan (W) and tyrosine (Y).
- the helicase preferably comprises a variant of SEQ ID NO: 118 wherein the at least one amino acid which interacts with one or more phosphates in one or more nucleotides in ssDNA is at least one of H64, T80, S83, N242, K243, N293, T394 and K397. These numbers correspond to the relevant positions in SEQ ID NO: 89 and may need to be altered in the case of variants where one or more amino acids have been inserted or deleted compared with SEQ ID NO: 118. A skilled person can determine the corresponding positions in a variant as discussed above.
- the helicase preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 and wherein the at least one amino acid which interacts with one or more phosphates in one or more nucleotides in ssDNA is at least one of the amino acids which correspond to H64, T80, S83, N242, K243, N293, T394 and K397 in SEQ ID NO: 118.
- Table 4 shows the amino acids in SEQ ID NOs: 119 to 133 which correspond to H64, T80, S83, N242, K243, N293, T 394 and K397 in SEQ ID NO: 118.
- Histidine (H) is preferably substituted with (i) arginine (R) or lysine (K), (ii) asparagine (N), serine (S), glutamine (Q) or threonine (T), (iii) phenylalanine (F), tryptophan (W) or tyrosine (Y). Histidine (H) is preferably substituted with (a) N, Q, K or F or (b) N, Q or W.
- Threonine (T) is preferably substituted with (i) arginine (R), histidine (H) or lysine (K), (ii) asparagine (N), serine (S), glutamine (Q) or histidine (H) or (iii) phenylalanine (F), tryptophan (W), tyrosine (Y) or histidine (H).
- Threonine (T) is more preferably substituted with (a) K, Q or N or (b) K, H or N.
- Serine (s) is preferably substituted with (i) arginine (R), histidine (H) or lysine (K), (ii) asparagine (N), glutamine (Q), threonine (T) or histidine (H) or (iii) phenylalanine (F), tryptophan (W), tyrosine (Y) or histidine (H).
- Serine (S) is more preferably substituted with H, N, K, T, R or Q.
- Asparagine (N) is preferably substituted with (i) arginine (R), histidine (H) or lysine (K), (ii) serine (S), glutamine (Q), threonine (T) or histidine (H) or (iii) phenylalanine (F), tryptophan (W), tyrosine (Y) or histidine (H).
- Asparagine (N) is more preferably substituted with (a) H or Q or (b) Q, K or H.
- Lysine (K) is preferably substituted with (i) arginine (R) or histidine (H), (ii) asparagine (N), serine (S), glutamine (Q), threonine (T) or histidine (H) or (iii) phenylalanine (F), tryptophan (W), tyrosine (Y) or histidine (H). Lysine (K) is more preferably substituted with (a) Q or H or (b) R, H or Y.
- the helicase more preferably comprises a variant of SEQ ID NO: 118 and comprises one or more of, such as all of, (a) H64N, H64Q, H64K or H64F, (b) T80K, T80Q or T80N, (c) S83H, S83N, S83K, S83T, S83R, or S83Q (d) N242H or N242Q, (e) K243Q or K243H, (f) N293Q, N293K or N293H, (g) T394K, T394H or T394N or (h) K397R, K397H or K397Y.
- SEQ ID NO: 118 comprises one or more of, such as all of, (a) H64N, H64Q, H64K or H64F, (b) T80K, T80Q or T80N, (c) S83H, S83N, S83K, S83T, S83R, or S83Q (d)
- the helicase is preferably a variant of SEQ ID NO: 118 which comprises substitutions at:
- F98/H64 such as F98W/H64N, F98W/H64Q, F98W/H64K or F98W/H64F;
- F98/T80 such as F98W/T80K, F98W/T80Q, F98W/T80N;
- F98/H82 such as F98W/H82N, F98W/H82Q or F98W/H82W;
- F98/S83 such as F98W/S83H, F98W/S83N, F98W/S83K, F98W/S83T, F98W/S83R or F98W/S83Q;
- F98/N242 such as F98W/N242H, F98W/N242Q, F98W/K243Q or F98W/K243H;
- F98/N293 such as F98W/N293Q, F98W/N293K, F98W/N293H, F98W/T394K, F98W/T394H, F98W/T394N, F98W/H396Y, F98W/H396F, F98W/H396Q or F98W/H396K; or
- F98/K397 such as F98W/K397R, F98W/K397H or F98W/K397Y.
- Preferred combinations in SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 include the combinations of amino acids which correspond to the combinations in SEQ ID NO: 118 listed above.
- the helicase of the invention is further one in which the part of the helicase which interacts with a transmembrane pore further comprises one or more modifications, preferably one or more substitutions.
- the part of the helicase which interacts with a transmembrane pore is typically the part of the helicase which interacts with a transmembrane pore when the helicase is used to control the movement of a polynucleotide through the pore, for instance as discussed in more detail below.
- the part typically comprises the amino acids that interact with or contact the pore when the helicase is used to control the movement of a polynucleotide through the pore, for instance as discussed in more detail below.
- the part typically comprises the amino acids that interact with or contact the pore when the helicase is bound to or attached to an analyte such as polynucleotide which is moving through the pore under an applied potential.
- the part which interacts with the transmembrane pore typically comprises the amino acids at positions 1, 2, 3, 4, 5, 6, 51, 176, 177, 178, 179, 180, 181, 185, 189, 191, 193, 194, 195, 197, 198, 199, 200, 201, 202, 203, 204, 207, 208, 209,
- the part which interacts with the transmembrane pore preferably comprises the amino acids at
- the part which interacts with the transmembrane pore preferably comprises one or more of, such as 2, 3, 4 or 5 of, the amino acids at positions K194, W195, K198, K199 and E258 in SEQ ID NO: 118.
- the variant of SEQ ID NO: 118 preferably comprises a modification at one or more of (a), K194, (b) W195, (c) D198, (d) K199 and (d) E258.
- the variant of SEQ ID NO: 118 preferably comprises a substitution at one or more of (a) K194, such as K194L, (b) W195, such as W195A, (c) D198, such as D198V, (d) K199, such as K199L and (e) E258, such as E258L.
- the variant may comprise ⁇ a ⁇ ; ⁇ b ⁇ ; ⁇ c ⁇ ; ⁇ d ⁇ ; ⁇ e ⁇ ; ⁇ a,b ⁇ ; ⁇ a,c ⁇ ; ⁇ a,d ⁇ ; ⁇ a,e ⁇ ; ⁇ b,c ⁇ ; ⁇ b,d ⁇ ; ⁇ b,e ⁇ ; ⁇ c,d ⁇ ; ⁇ c,e ⁇ ; ⁇ d,e ⁇ ; ⁇ a,b,c ⁇ ; ⁇ a,b,d ⁇ ; ⁇ a,b,e ⁇ ; ⁇ a,c,d ⁇ ; ⁇ a,c,e ⁇ ; ⁇ a,d,e ⁇ ; ⁇ b,c,d ⁇ ; ⁇ b,c,e ⁇ ; ⁇ b,d,e ⁇ ; ⁇ c,d,e ⁇ ; ⁇ a,b,c,d ⁇ ; ⁇ a,b,c,d ⁇ ; ⁇ a,b,c,d ⁇ ; ⁇ a,b,c,d ⁇ ; ⁇ a,b,c,d ⁇ ; ⁇ a,b,c,d ⁇
- the part of the polynucleotide binding protein which interacts with the transmembrane pore preferably comprises the amino acid at position 194 or 199 of SEQ ID NO: 118.
- the variant preferably comprises K194A, K194V, K194F, K194D, K194S, K194W or K194L and/or K199A, K199V, K199F, K199D, K199S, K199W or K199L.
- the part which interacts with the transmembrane pore typically comprises the amino acids at positions which correspond to those in SEQ ID NO: 118 listed above.
- Amino acids in SEQ ID NOs: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 and 133 which correspond to these positions in SEQ ID NO: 118 can be identified using the alignment in Table 5 below.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at F98, such as F98R, F98K, F98Q, F98N, F98H, F98Y, F98F or F98W, and a substitution at K194, such as K194A, K194V, K194F, K194D, K194S, K194W or K194L, and/or K199, such as K199A, K199V, K199F, K199D, K199S, K199W or K199L.
- F98 such as F98R, F98K, F98Q, F98N, F98H, F98Y, F98F or F98W
- K194A such as K194V, K194F, K194D, K194S, K194W or K194L
- K199 such as K199A, K199V, K199F, K199D, K199S, K199W or K199L.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 which further comprises a substitution at the position which corresponds to F98 in SEQ ID NO: 118 and a substitution at the position(s) which correspond to K194 and/or K199 in SEQ ID NO: 118. These corresponding positions may be replaced with any of the amino acids listed above for F98, K194 and K119 in SEQ ID NO: 118.
- the helicase is preferably a variant of SEQ ID NO: 118 which further comprises substitutions at:
- F98/K194/H64 such as F98W/K194L/H64N, F98W/K194L/H64Q, F98W/K194L/H64K or F98W/K194L/H64F;
- F98/K194/T80 such as F98W/K194L/T80K, F98W/K194L/T80Q or F98W/K194L/T80N;
- F98/K194/H82 such as F98W/K194L/H82N, F98W/K194L/H82Q or F98W/K194L/H82W
- F98/S83/K194 such as F98W/S83H/K194L, F98W/S83T/K194L, F98W/S83R/K194L, F98W/S83Q/K194L, F98W/S83N/K194L, F98W/S83K/K194L, F98W/N88R/K194L, F98W/N88H/K194L, F98W/N88W/K194L or F98W/N88Y/K194L;
- F98/P89/K194 such as F98W/P89L/K194L, F98W/P89V/K194L, F98W/P89I/K194L or F98W/P89T/K194L;
- F98/D121/K194 such as F98W/D121H/K194L, F98W/D121Y/K194L or F98W/D121K/K194L;
- F98/V150/K194 such as F98W/V150I/K194L, F98W/V150L/K194L, F98W/V150N/K194L, F98W/V150W/K194L or F98W/V150H/K194L;
- F98/P152/K194 such as F98W/P152W/K194L, F98W/P152F/K194L, F98W/P152Y/K194L, F98W/P152H/K194L, F98W/P152I/K194L, F98W/P152L/K194L or F98W/P152V/K194L;
- F98/F240/K194 such as F98W/F240W/K194L, F98W/F240Y/K194L or F98W/F240H/K194L;
- F98/N242/K194 such as F98W/N242H/K194L or F98W/N242Q/K194L
- F98/K194/S287 such as F98W/K194L/S287K, F98W/K194L/S287R, F98W/K194L/S287W or F98W/K194L/S287F;
- F98/N293/K194 such as F98W/N293Q/K194L, F98W/N293K/K194L or F98W/N293H/K194L;
- F98/T394/K194 such as F98W/T394K/K194L, F98W/T394H/K194L or F98W/T394N/K194L;
- F98/H396/K194 such as F98W/H396Y/K194L, F98W/H396F/K194L, F98W/H396Q/K194L or F98W/H396K/K194L;
- F98/K397/K194 such as F98W/K397R/K194L, F98W/K397H/K194L or F98W/K397Y/K194L; or
- F98/Y415/K194 such as F98W/Y415W/K194L or F98W/Y415R/K194L.
- K194 may be replaced with any of W195, D198, K199 and E258.
- the modified helicase preferably further comprises a modification or substitution at the position(s) corresponding to amino acid positions 98 and/or 194 in Dda 1993.
- Position 98 or the corresponding position may be substituted with R, H, K, S, T, N, Q, A, V, I, L, M, Y or W.
- Position 98 or the corresponding position is preferably substituted with R, K, Q, N, H, Y or W.
- Position 194 or the corresponding position may be substituted with A, V, I, L, M, F, Y, W, D, E, S, T, N or Q.
- Position 194 or the corresponding position is preferably substituted with A, V, F, D, S, W or L.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at F98, such as F98R, F98K, F98Q, F98N, F98H, F98Y or F98W, and/or a substitution at K194, such as K194A, K194V, K194F, K194D, K194S, K194W or K194L.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises F98W and K194L.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 which further comprises a substitution at the position which corresponds to F98 in SEQ ID NO: 118 and/or a substitution at the position which corresponds to K194 in SEQ ID NO: 118.
- K194 may be replaced with any of W195, D198, K199 and E258. Modifications in the tower domain and/or pin domain and/or 1A domain
- the modified helicase preferably further comprises a modification or substitution at the position corresponding to amino acid position 360 in Dda 1993.
- Position 360 or the corresponding position may be substituted with C, G, P, A, V, I, L, M, F, Y or W.
- Position 360 or the corresponding position is preferably substituted with C or Y.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at A360, such as A360C or A360Y.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises K358I and A360C.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 which comprises a substitution at the position which corresponds to A360 in SEQ ID NO: 118.
- the modified helicase preferably further comprises a modification or substitution at one or more of the positions corresponding to amino acid positions 94, 98 and 109 in Dda 1993, such as position(s) 94, 98, 109, 94 and 98, 94 and 109, 98 and 109 and 94, 98 and 109. These positions are all in the pin domain. Position 94 or the corresponding position may be substituted with C, G, P, A, V, I, L, M, F, Y or W. Position 94 or the corresponding position is preferably substituted with C or Y. Position 98 or the corresponding position may be substituted with R, H, K, S, T, N, Q, A, V, I, L, MY or W.
- Position 98 or the corresponding position is preferably substituted with R, K, Q, N, H, Y or W.
- Position 109 or the corresponding position may be substituted with A, V, I, L, M, F, Y or W.
- Position 109 or the corresponding position is preferably substituted with A or V.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at one or more of E94, F98 and C109 (including all the combinations set out above).
- Preferred variants comprise substitutions at:
- E94 and F98 such as E94C or E94Y and F98R, F98K, F98Q, F98N, F98H, F98Y, F98F or F98W;
- E94 and C109 such as E94C or E94Y and C109A or C109V;
- E94, F98 and C109 such as E94C or E94Y and F98R, F98K, F98Q, F98N, F98H, F98Y, F98F or F98W and C109A or C109V.
- More preferred variants comprise: E94C and F98W; E94C and C109A; F98W and C109A; or E94C, F98W and C109A.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 which further comprises a substitution at the position(s) which corresponds to one or more of E94, F98 and C109 in SEQ ID NO: 118.
- Table 6 includes information for E94, C109, C136 and A360 (with reference to modified helicases disclosed above and below).
- the helicase of the invention is preferably one in which at least one cysteine residue (i.e. one or more cysteine residues) and/or at least one non-natural amino acid (i.e. one or more non-natural amino acids) have further been introduced into (i) the tower domain and/or (ii) the pin domain and/or the (iii) 1A (RecA-like motor) domain, wherein the helicase has the ability to control the movement of a polynucleotide.
- cysteine residue i.e. one or more cysteine residues
- non-natural amino acid i.e. one or more non-natural amino acids
- At least one cysteine residue and/or at least one non-natural amino acid may be introduced into the tower domain, the pin domain, the 1A domain, the tower domain and the pin domain, the tower domain and the 1A domain or the tower domain, the pin domain and the 1A domain.
- the helicase of the invention is preferably one in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into each of (i) the tower domain and (ii) the pin domain and/or the 1A (RecA-like motor) domain, i.e. into the tower domain and the pin domain, the tower domain and the 1A domain or the tower domain, the pin domain and the 1A domain. Any number of cysteine residues and/or non-natural amino acids may be introduced into each domain.
- cysteine residues may be introduced and/or 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more non-natural amino acids may be introduced. Only one or more cysteine residues may be introduced. Only one or more non-natural amino acids may be introduced. A combination of one or more cysteine residues and one or more non-natural amino acids may be introduced.
- the at least one cysteine residue and/or at least one non-natural amino acid are/is preferably introduced by substitution. Methods for doing this are known in the art.
- modifications do not prevent the helicase from binding to a polynucleotide. These modifications decrease the ability of the polynucleotide to unbind or disengage from the helicase. In other words, the one or more modifications increase the processivity of the helicase by preventing dissociation from the polynucleotide strand.
- the thermal stability of the enzyme is typically also increased by the one or more modifications giving it an improved structural stability that is beneficial in Strand Sequencing.
- a non-natural amino acid is an amino that is not naturally found in a helicase.
- the non- natural amino acid is preferably not histidine, alanine, isoleucine, arginine, leucine, asparagine, lysine, aspartic acid, methionine, cysteine, phenylalanine, glutamic acid, threonine, glutamine, tryptophan, glycine, valine, proline, serine or tyrosine.
- the non- natural amino acid is more preferably not any of the twenty amino acids in the previous sentence or selenocysteine.
- Preferred non-natural amino acids for use in the invention include, but are not limited, to 4- Azido-L-phenylalanine (Faz), 4-Acetyl-L-phenylalanine, 3-Acetyl-L-phenylalanine, 4- Acetoacetyl-L-phenylalanine, O-Allyl-L-tyrosine, 3-(Phenylselanyl)-L-alanine, O-2-Propyn-l- yl-L-tyrosine, 4-(Dihydroxyboryl)-L-phenylalanine, 4- [(Ethylsulfa nyl)carbonyl]-L- phenylalanine, (2S)-2-amino-3-4-[(propan-2-ylsulfanyl)carbonyl] phenyl; propanoic acid, (2S)-2-a m i no-3-4- [(2-amino-3-sulfanylpropanoy
- Table 7 (which is separated in two parts) identifies the residues making up each domain in each Dda homologue (SEQ ID NOs: 118 to 133).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues D260-P274 and N292-A389) and/or (ii) the pin domain (residues K86-E102) and/or the (iii) 1A domain (residues M1-L85 and V103- K177).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N292-A389 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 119 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues G295-N309 and F316-Y421) and/or (ii) the pin domain (residues Y85-L112) and/or the (iii) 1A domain (residues MI-184 and R.113- Y211).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues F316-Y421 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 120 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues V328-P342 and N360-Y448) and/or (ii) the pin domain (residues K148-N165) and/or the (iii) 1A domain (residues M1-L147 and S166- V240).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N360-Y448 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 121 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues A261-T275 and T285-Y370) and/or (ii) the pin domain (residues G91-E107) and/or the (iii) 1A domain (residues M1-L90 and E108- H173).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues T285-Y370 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 122 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues G294-I307 and T314-Y407) and/or (ii) the pin domain (residues G116-T135) and/or the (iii) 1A domain (residues M1-L115 and N136- V205).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues T314-Y407 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 123 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues V288-E301 and N307-N393) and/or (ii) the pin domain (residues G97-P113) and/or the (iii) 1A domain (residues M1-L96 and F114- V194).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N307-N393 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 124 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues S250-P264 and E278-S371) and/or (ii) the pin domain (residues K78-E95) and/or the (iii) 1A domain (residues M1-L77 and V96-V166).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues E278-S371 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 125 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues K255-P269 and T284-S380) and/or (ii) the pin domain (residues K82-K98) and/or the (iii) 1A domain (residues M1-M81 and L99- M171).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues T284-S380 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 126 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues D242-P256 and T271-S366) and/or (ii) the pin domain (residues K69-K85) and/or the (iii) 1A domain (residues M1-M68 and MSG- MISS).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues T271-S366 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 127 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues T263-P277 and N295-P392) and/or (ii) the pin domain (residues K88-K107) and/or the (iii) 1A domain (residues M1-L87 and A108- M181).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N295-P392 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 128 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues D263-P277 and N295-A391) and/or (ii) the pin domain (residues K88-K107) and/or the (iii) 1A domain (residues M1-L87 and A108- M181).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N295-A391 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 129 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues A258-P272 and N290-P386) and/or (ii) the pin domain (residues K86-G102) and/or the (iii) 1A domain (residues M1-L85 and T103- K176).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N290-P386 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 130 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues L266-P280 and N298-A392) and/or (ii) the pin domain (residues K92-D108) and/or the (iii) 1A domain (residues M1-L91 and V109- M183).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N298-A392 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 131 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues D262-P276 and N294-A392) and/or (ii) the pin domain (residues K88-E104) and/or the (iii) 1A domain (residues M1-L87 and M105- M179).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N294-A392 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 132 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues D261-P275 and N293-A389) and/or (ii) the pin domain (residues K87-E103) and/or the (iii) 1A domain (residues M1-L86 and V104- K178).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N293-A389 of the tower domain.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 133 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues E261-P275 and T293-A390) and/or (ii) the pin domain (residues K87-E103) and/or the (iii) 1A domain (residues M1-L86 and V104- M178).
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues T293-A390 of the tower domain.
- the helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 118 to 133 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into each of (i) the tower domain and (ii) the pin domain and/or the 1A domain.
- the helicase of the invention more preferably comprises a variant of any one of SEQ ID NOs: 118 to 133 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into each of (i) the tower domain, (ii) the pin domain and (iii) the 1A domain. Any number and combination of cysteine residues and non-natural amino acids may be introduced as discussed above.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises (i) E94C and/or A360C; (ii) E93C and/or K358C; (iii) E93C and/or A360C; (iv) E93C and/or E361C; (v) E93C and/or K364C; (vi) E94C and/or L354C; (vii) E94C and/or K358C; (viii) E93C and/or L354C; (ix) E94C and/or E361C; (x) E94C and/or K364C; (xi) L97C and/or L354C; (xii) L97C and/or K358C; (xiii) L97C and/or A360C; (xiv) L97C and/or E361C; (xv) L97C and/or K364C; (xvi) K123C and/
- the helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises a cysteine residue at the positions which correspond to those in SEQ ID NO: 118 as defined in any of (i) to (Ixii). Positions in any one of SEQ ID NOs: 119 to 133 which correspond to those in SEQ ID NO: 118 can be identified using the alignment of SEQ ID NOs: 118 to 133 below.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 92 which further comprises (a) D99C and/or L341C, (b) Q98C and/or L341C or (d) Q98C and/or A340C.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 96 which further comprises D90C and/or A349C.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 102 which further comprises D96C and/or A362C.
- the helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 118 to 133 as defined in any one of (i) to (Ixii) in which Faz is further introduced at one or more of the specific positions instead of cysteine. Faz may be introduced at each specific position instead of cysteine.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises (i) E94Faz and/or A360C; (ii) E94C and/or A360Faz; (iii) E94Faz and/or A360Faz; (iv) Y92L, E94Y, Y350N, A360Faz and Y363N; (v) A360Faz; (vi) E94Y and A360Faz; (vii) Y92L, E94Faz, Y350N, A360Y and Y363N; (viii) Y92L, E94Faz and A360Y; (ix) E94Faz and A360Y; and (x) E94C, G357Faz and A360C.
- the helicase of the invention preferably further comprises one or more single amino acid deletions from the pin domain. Any number of single amino acid deletions may be made, such as 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more.
- the helicase more preferably comprises a variant of SEQ ID NO: 118 which further comprises deletion of E93, deletion of E95 or deletion of E93 and E95.
- the helicase more preferably comprises a variant of SEQ ID NO: 118 which comprises (a) E94C, deletion of N95 and A360C; (b) deletion of E93, deletion of E94, deletion of N95 and A360C; (c) deletion of E93, E94C, deletion of N95 and A360C or (d) E93C, deletion of N95 and A360C.
- the helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises deletion of the position corresponding to E93 in SEQ ID NO: 118, deletion of the position corresponding to E95 in SEQ ID NO: 118 or deletion of the positions corresponding to E93 and E95 in SEQ ID NO: 118.
- the helicase of the invention preferably further comprises one or more single amino acid deletions from the hook domain. Any number of single amino acid deletions may be made, such as 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more.
- the helicase more preferably comprises a variant of SEQ ID NO: 118 which further comprises deletion of any number of positions T278 to S287.
- the helicase more preferably comprises a variant of SEQ ID NO: 118 which further comprises (a) E94C, deletion of Y279 to K284 and A360C, (b) E94C, deletion of T278, Y279, V286 and S287 and A360C, (c) E94C, deletion of 1281 and K284 and replacement with a single G and A360C, (d) E94C, deletion of K280 and P2845 and replacement with a single G and A360C, or (e) deletion of Y279 to K284, E94C, F276A and A230C.
- the helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises deletion of any number of the positions corresponding to 278 to 287 in SEQ ID NO: 118.
- the helicase of the invention preferably further comprises one or more single amino acid deletions from the pin domain and one or more single amino acid deletions from the hook domain.
- the helicase of the invention is preferably one in which at least one cysteine residue and/or at least one non-natural amino acid have been further introduced into the hook domain and/or the 2A (RecA-like) domain. Any number and combination of cysteine residues and non-natural amino acids may be introduced as discussed above for the tower, pin and 1A domains.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L275-F291) and/or the 2A (RecA-like) domain (residues R178-T259 and L390-V439).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 119 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues A310-L315) and/or the 2A (RecA-like) domain (residues R212-E294 and G422-S678).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 120 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues V343-L359) and/or the 2A (RecA-like) domain (residues R241-N327 and A449-G496).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 121 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues W276-L284) and/or the 2A (RecA-like) domain (residues R174-D260 and A371-V421).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 122 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues R308-Y313) and/or the 2A (RecA-like) domain (residues R206-K293 and I408-L500).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 123 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues M302-W306) and/or the 2A (RecA-like) domain (residues R195-D287 and V394-Q450).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 124 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues V265-I277) and/or the 2A (RecA-like) domain (residues R167-T249 and L372-N421).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 125 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues V270-F283) and/or the 2A (RecA-like) domain (residues R172-T254 and L381-K434).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 126 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues V257-F270) and/or the 2A (RecA-like) domain (residues R159-T241 and L367-K420).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 127 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L278-Y294) and/or the 2A (RecA-like) domain (residues R182-T262 and L393-V443).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 128 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L278-Y294) and/or the 2A (RecA-like) domain (residues R182-T262 and L392-V442).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 129 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L273-F289) and/or the 2A (RecA-like) domain (residues R177-N257 and L387-V438).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 130 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L281-F297) and/or the 2A (RecA-like) domain (residues R184-T265 and L393-I442).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 131 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues H277-F293) and/or the 2A (RecA-like) domain (residues R180-T261 and L393-V442).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 132 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L276-F292) and/or the 2A (RecA-like) domain (residues R179-T260 and L390-I439).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 133 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L276-F292) and/or the 2A (RecA-like) domain (residues R179-T260 and L391-V441).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises one or more of (i) I181C; (ii) Y279C; (iii) I281C; and (iv) E288C.
- the helicase may comprise any combination of (i) to (iv), such as (i); (ii); (iii); (iv); (i) and (ii); (i) and (iii); (i) and (iv); (ii) and (iii); (ii) and (iv); (iiii) and (iv); or (i), (ii), (iii) and (iv).
- the helicase more preferably comprises a variant of SEQ ID NO: 118 which comprises (a) E94C, I281C and A360C or (b) E94C, I281C, G357C and A360C.
- the helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises a cysteine residue at one or more of the position(s) which correspond to those in SEQ ID NO: 118 as defined in (i) to (iv), (a) and (b).
- the helicase may comprise any of these variants in which Faz is introduced at one or more of the specific positions (or each specific position) instead of cysteine.
- the helicase of the invention is preferably further modified to reduce its surface negative charge.
- Surface residues can be identified in the same way as the Dda domains disclosed above.
- Surface negative charges are typically surface negatively-charged amino acids, such as aspartic acid (D) and glutamic acid (E).
- the helicase is preferably modified to neutralise one or more surface negative charges by substituting one or more negatively charged amino acids with one or more positively charged amino acids, uncharged amino acids, non-polar amino acids and/or aromatic amino acids or by introducing one or more positively charged amino acids, preferably adjacent to one or more negatively charged amino acids.
- Suitable positively charged amino acids include, but are not limited to, histidine (H), lysine (K) and arginine (R). Uncharged amino acids have no net charge.
- Suitable uncharged amino acids include, but are not limited to, cysteine (C), serine (S), threonine (T), methionine (M), asparagine (N) and glutamine (Q).
- Non-polar amino acids have non-polar side chains.
- Suitable non-polar amino acids include, but are not limited to, glycine (G), alanine (A), proline (P), isoleucine (I), leucine (L) and valine (V).
- Aromatic amino acids have an aromatic side chain. Suitable aromatic amino acids include, but are not limited to, histidine (H), phenylalanine (F), tryptophan (W) and tyrosine (Y).
- the helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 and the one or more negatively charged amino acids correspond to one or more of D5, E8, E23, E47, D167, E172, D202, D212 and E273 in SEQ ID NO: 118.
- Amino acids in SEQ ID NOs: 119 to 133 which correspond to D5, E8, E23, E47, D167, E172, D202, D212 and E273 in SEQ ID NO: 118 can be determined using the alignment in WO2015/055981.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which comprises (a) E94C, E273G and A360C or (b) E94C, E273G, N292G and A360C.
- the helicase of the invention is preferably further modified by the removal of one or more native cysteine residues. Any number of native cysteine residues may be removed.
- the one or more cysteine residues are preferably removed by substitution.
- the one or more cysteine residues are preferably substituted with alanine (A), serine (S) or valine (V).
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 and the one or more native cysteine residues are one or more of C109, C114, C136, C171 and C412. Any number and combination of these cysteine residues may be removed.
- the variant of SEQ ID NO: 118 may comprise C109; C114; C136; C171; C412; C109 and C114; C109 and C136; C109 and C171; C109 and C412; C114 and C136; C114 and C171; C114 and C412; C136 and C171; C136 and C412; C171 and C412; C109, C114 and C136; C109, C114 and C171; C109, C114 and C412; C109, C136 and C171; C109, C136 and C412; C109, C171 and C412; C114, C136 and C171; C114, C136 and C412; C114, C171 and C412; C136, C171 and C412; C109, C114, C136 and C171; C109, C114, C136 and C412; C114, C171 and C412; C109, C114, C136 and C171; C109, C114
- the modified helicase preferably further comprises a modification or substitution at the position(s) corresponding to amino acid position(s) 109 and/or 136 in Dda 1993. This removes one or two cysteine residues. This may be in addition to a modification or substitution at one or more positions corresponding to amino acid positions 55, 114, 156, 177, 210, 221, 350 and 358 in Dda 1993, a modification or substitution at one or more positions corresponding to amino acid positions 114, 177, 350 and 358 in Dda 1993 and/or a modification or substitution at the position corresponding to position 40 in Dda 1993. Position 109 or the corresponding position may be substituted with A, V, I, L, M, F, Y or W.
- Position 109 or the corresponding position is preferably substituted with A or V.
- Position 136 or the corresponding position may be substituted with A, V, I, L, M, F, Y or W.
- Position 136 or the corresponding position is preferably substituted with A or V.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at C109, such as C109A, C109V, C109I, C109L, C109M, C109F, C109Y or C109W and/or at C136, such as C136A, C136V, C136I, C136L, C136T, C136M, C136F, C136Y or C136W.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises C109A and/or C136A.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 which further comprises a substitution at the position(s) which correspond(s) to C109 and/or C136 in SEQ ID NO: 118.
- the helicase of the invention is preferably one in which at least one cysteine residue (i.e. one or more cysteine residues) and/or at least one non-natural amino acid (i.e. one or more non-natural amino acids) have further been introduced into the tower domain only. Suitable modifications are discussed above.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising the following mutations: E93C and K364C; E94C and K364C; E94C and A360C; L97C and E361C; L97C and E361C and C412A; K123C and E361C; K123C, E361C and C412A; N155C and K358C; N155C, K358C and C412A; N155C and L354C; N155C, L354C and C412A; deltaE93, E94C, deltaN95 and A360C; E94C, deltaN95 and A360C; E94C, Q100C, I127C and A360C; L354C; G357C; E94C, G357C and A360C; E94C, Y279C and A360C; E94C, I281C and A360C; E94C, Y279Faz and A
- the helicase of the invention is one in which at least one cysteine residue and/or at least one non-natural amino acid have been further introduced into the hook domain and/or the 2A (RecA-like motor) domain, wherein the helicase has the ability to control the movement of a polynucleotide. At least one cysteine residue and/or at least one non-natural amino acid is preferably introduced into the hook domain and the 2A (RecA-like motor) domain.
- cysteine residues and/or non-natural amino acids may be introduced into each domain. For instance, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more cysteine residues may be introduced and/or 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more non-natural amino acids may be introduced. Only one or more cysteine residues may be introduced. Only one or more non-natural amino acids may be introduced. A combination of one or more cysteine residues and one or more non-natural amino acids may be introduced.
- the at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced by substitution. Methods for doing this are known in the art. Suitable modifications of the hook domain and/or the 2A (RecA-like motor) domain are discussed above.
- the helicase of the invention is preferably a variant of SEQ ID NO: 118 further comprising (a) Y279C, I181C, E288C, Y279C and I181C, (b) Y279C and E288C, (c) I181C and E288C or (d) Y279C, I181C and E288C.
- the helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 199 to 133 which further comprises a mutation at one or more of the position(s) which correspond to those in SEQ ID NO: 118 as defined in (a) to (d).
- the helicase is modified to reduce its surface negative charge, wherein the helicase has the ability to control the movement of a polynucleotide. Suitable modifications are discussed above. Any number of surface negative charges may be neutralised.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising the following mutations: E273G; E8R, E47K and D202K; D5K, E23N, D167K, E172R and D212R; or D5K, E8R, E23N, E47K, D167K, E172R, D202K and D212R.
- Other modified helicases E273G; E8R, E47K and D202K; D5K, E23N, D167K, E172R and D212R; or D5K, E8R, E23N, E47K, D167K, E172R, D202K and D212R.
- the helicase of the invention comprises a variant of SEQ ID NO: 118 further comprising : A360K; Y92L and/or A360Y; Y92L, Y350N and Y363N; Y92L and/or Y363N; or Y92L.
- a variant of SEQ ID NO: 118 may comprise one or more of the following mutations: K38A; T91F; T91N; T91Q; T91W; V96E; V96F; V96L; V96Q; V96R; V96W; V96Y; P274G; V286F; V286W; V286Y; F291G; N292F; N292G; N292P; N292Y; G294Y; G294F; K364A; and W378A.
- a variant of SEQ ID NO: 118 may comprise: K38A, E94C and A360C; H64K; E94C and A360C; H64N; E94C and A360C;
- E94C, A360C and H396W E94C, A360C and Y415F; E94C, A360C and Y415K; E94C, A360C and Y415M; or E94C, A360C and Y415W.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises (a) E94C/A360C/W378A or (b) E94C/A360C/C109A/C136A/W378A or (d) E94C/A360C/C109A/C136A/W378A and then (AM1)G1G2 (i.e. deletion of Ml and then addition G1 and G2).
- Preferred variants of any one of SEQ ID NOs: 118 to 133 have (in addition to the modifications of the invention) the N-terminal methionine (M) replaced with one glycine residue (G). In the examples this is shown as (AMl)Gl. It may also be termed MIG. Any of the variants discussed above may further comprise MIG.
- the most preferred helicases of the invention comprise a variant of SEQ ID NO: 118 which further (a) E94C/F98W/A360C/C109A/C136A/K194L, (b) M1G/E94C/F98W/A360C/C109A/C136A/K194L; (c) E94C/F98W/A360C/C109A/C136A/K199L; or (d) M1G/E94C/F98W/A360C/C109A/C136A/K199L.
- the helicase of the invention may further be one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: 86, 90, 92, 97, 101, 102, 273, 293, 300, 301, 303, 305, 308, 310, 312 317, 323, 328, 332, 334, 335, 336, 337, 339, 351, 354, 359, 361, 364, 366, 368, 371, 374, 376, 377, 379 and 388.
- the helicase of the invention may further be one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: 351, 354 and 361.
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising one or more of the following substitutions: K86A, V90T , Y92A, Y92D, Y92F, Y92G, Y92H, Y92N, Y92Q, Y92S, Y92T, Y92V, Y92W, L97H, K101A, E102G, E273A, N293H, I200A, I300F, E301I, E303I, Y305L, F308I, F308L, K310A, K310I, K310L, R312I, R312L, R312M, E317I, E317Y, W323H, E328I, D332A, D332L, E334A, E334I, E334Y, Y335A, Y335I, Y335L, Y336L, R337I, R337L
- the helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising one or more of the following substitutions: (a) K351I or K351Q, (b) L354A or L354Q and (c) E361I or E361Q.
- the variant may comprise (a), (b), (c), (a) and (b), (a) and (c), (b) and (c) or (a), (b) and (c). These substitutions may be made in combination with any of the modifications or substitutions of the invention above.
- a variant of a helicase is an enzyme that has an amino acid sequence which varies from that of the wild-type helicase and which has polynucleotide binding activity.
- a variant of any one of SEQ ID NOs: 118 to 133 is an enzyme that has an amino acid sequence which varies from that of any one of SEQ ID NOs: 118 to 133 and which has polynucleotide binding activity.
- Polynucleotide binding activity can be determined using methods known in the art. Suitable methods include, but are not limited to, fluorescence anisotropy, tryptophan fluorescence and electrophoretic mobility shift assay (EMSA). For instance, the ability of a variant to bind a single stranded polynucleotide can be determined as described in the Examples.
- the variant has helicase activity. This can be measured in various ways. For instance, the ability of the variant to translocate along a polynucleotide can be measured using electrophysiology, a fluorescence assay or ATP hydrolysis.
- the variant may include modifications that facilitate handling of the polynucleotide encoding the helicase and/or facilitate its activity at high salt concentrations and/or room temperature.
- a variant will preferably be at least 20% homologous to that sequence based on amino acid similarity or identity. More preferably, the variant polypeptide may be at least 30%, at least 40%, at least 45%, at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90% and more preferably at least 95%, 97% or 99% homologous based on amino acid identity to the amino acid sequence of any one of SEQ ID NOs: 118 to 133 over the entire sequence.
- the variant polypeptide may be at least 30%, at least 40%, at least 45%, at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90% and more preferably at least 95%, 97% or 99% identical to the amino acid sequence of any one of SEQ ID NOs: 118 to 133 over the entire sequence.
- the variant of any one of SEQ ID NOs: 118 to 133 may comprise one or more substitutions, one or more deletions and/or one or more additions as discussed below.
- Standard methods in the art may be used to determine homology.
- the UWGCG Package provides the BESTFIT program which can be used to calculate homology, for example used on its default settings (Devereux et al (1984) Nucleic Acids Research 12, p387-395).
- the PILEUP and BLAST algorithms can be used to calculate homology or line up sequences (such as identifying equivalent residues or corresponding sequences (typically on their default settings)), for example as described in Altschul S. F. (1993) J Mol Evol 36:290- 300; Altschul, S.F et al (1990) J Mol Biol 215:403-10.
- Software for performing BLAST analyses is publicly available through the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/).
- Preferred variants of any one of SEQ ID NOs: 118 to 133 have a non-natural amino acid, such as Faz, at the amino- (N-) terminus and/or carboxy (C-) terminus.
- Preferred variants of any one of SEQ ID NOs: 118 to 133 have a cysteine residue at the amino- (N-) terminus and/or carboxy (C-) terminus.
- Preferred variants of any one of SEQ ID NOs: 118 to 133 have a cysteine residue at the amino- (N-) terminus and a non-natural amino acid, such as Faz, at the carboxy (C-) terminus or vice versa.
- Preferred variants of SEQ ID NO: 118 contain one or more of, such as all of, the following modifications E54G, D151E, I196N and G357A.
- none of the introduced cysteines and/or non-natural amino acids in a modified helicase of the invention are connected to one another.
- two more of the introduced cysteines and/or non-natural amino acids in a modified helicase of the invention are connected to one another. This typically reduces the ability of the helicase of the invention to unbind from a polynucleotide.
- Any number and combination of two more of the introduced cysteines and/or non-natural amino acids may be connected to one another. For instance, 3, 4, 5, 6, 7, 8 or more cysteines and/or non-natural amino acids may be connected to one another.
- One or more cysteines may be connected to one or more cysteines.
- One or more cysteines may be connected to one or more non-natural amino acids, such as Faz.
- One or more non-natural amino acids, such as Faz may be connected to one or more non-natural amino acids, such as Faz.
- the two or more cysteines and/or non-natural amino acids may be connected in any way.
- the connection can be transient, for example non-covalent. Even transient connection will reduce unbinding of the polynucleotide from the helicase.
- the two or more cysteines and/or non-natural amino acids are preferably connected by affinity molecules.
- Suitable affinity molecules are known in the art.
- the affinity molecules are preferably (a) complementary polynucleotides (WO 2010/086602 incorporated herein by reference in its entirety), (b) an antibody or a fragment thereof and the complementary epitope (Biochemistry 6thEd, W.H. Freeman and co (2007) pp953-954), (c) peptide zippers (O'Shea et al., Science 254 (5031): 539-544), (d) capable of interacting by -sheet augmentation (Remaut and Waksman Trends Biochem. Sci.
- the two or more parts may be transiently connected by a hexa-his tag or Ni-NTA.
- the two or more cysteines and/or non-natural amino acids are preferably permanently connected.
- a connection is permanent if is not broken while the helicase is used or cannot be broken without intervention on the part of the user, such as using reduction to open -S-S- bonds.
- the two or more cysteines and/or non-natural amino acids are preferably covalently- attached.
- the two or more cysteines and/or non-natural amino acids may be covalently attached using any method known in the art.
- the two or more cysteines and/or non-natural amino acids may be covalently attached via their naturally occurring amino acids, such as cysteines, threonines, serines, aspartates, asparagines, glutamates and glutamines.
- Naturally occurring amino acids may be modified to facilitate attachment.
- the naturally occurring amino acids may be modified by acylation, phosphorylation, glycosylation or farnesylation. Other suitable modifications are known in the art. Modifications to naturally occurring amino acids may be posttranslation modifications.
- the two or more cysteines and/or non-natural amino acids may be attached via amino acids that have been introduced into their sequences. Such amino acids are preferably introduced by substitution.
- the introduced amino acid may be cysteine or a non-natural amino acid that facilitates attachment.
- Suitable non-natural amino acids include, but are not limited to, 4-azido-L-phenylalanine (Faz), any one of the amino acids numbered 1-71 included in figure 1 of Liu C. C. and Schultz P. G., Annu. Rev. Biochem., 2010, 79, 413-444 or any one of the amino acids listed below.
- the introduced amino acids may be modified as discussed above.
- the two or more cysteines and/or non-natural amino acids are connected using linkers.
- Linker molecules are discussed in more detail below.
- One suitable method of connection is cysteine linkage. This is discussed in more detail below.
- the two or more cysteines and/or non-natural amino acids are preferably connected using one or more, such as two or three, linkers.
- the one or more linkers may be designed to reduce the size of, or close, the opening as discussed above. If one or more linkers are being used to close the opening as discussed above, at least a part of the one or more linkers is preferably oriented such that it is not parallel to the polynucleotide when it is bound by the helicase. More preferably, all of the linkers are oriented in this manner.
- At least a part of the one or more linkers preferably crosses the opening in an orientation that is not parallel to the polynucleotide when it bound by the helicase. More preferably, all of the linkers cross the opening in this manner. In these embodiments, at least a part of the one or more linkers may be perpendicular to the polynucleotide. Such orientations effectively close the opening such that the polynucleotide cannot unbind from the helicase through the opening.
- Each linker may have two or more functional ends, such as two, three or four functional ends. Suitable configurations of ends in linkers are well known in the art.
- One or more ends of the one or more linkers are preferably covalently attached to the helicase. If one end is covalently attached, the one or more linkers may transiently connect the two or more cysteines and/or non-natural amino acids as discussed above. If both or all ends are covalently attached, the one or more linkers permanently connect the two or more cysteines and/or non-natural amino acids.
- the one or more linkers are preferably amino acid sequences and/or chemical crosslinkers.
- Suitable amino acid linkers such as peptide linkers, are known in the art.
- the length, flexibility and hydrophilicity of the amino acid or peptide linker are typically designed such that it reduces the size of the opening, but does not to disturb the functions of the helicase.
- Preferred flexible peptide linkers are stretches of 2 to 20, such as 4, 6, 8, 10 or 16, serine and/or glycine amino acids. More preferred flexible linkers include (SG)i, (SG)2, (SG)s, (SG)4, (SG) 5 , (SG)8, (SG) IO, (SG)is or (SG)2o wherein S is serine and G is glycine.
- Preferred rigid linkers are stretches of 2 to 30, such as 4, 6, 8, 16 or 24, proline amino acids. More preferred rigid linkers include (P)i2 wherein P is proline.
- the amino acid sequence of a linker preferably comprises a polynucleotide binding moiety. Such moieties and the advantages associated with their use are discussed
- Suitable chemical crosslinkers are well-known in the art. Suitable chemical crosslinkers include, but are not limited to, those including the following functional groups: maleimide, active esters, succinimide, azide, alkyne (such as dibenzocyclooctynol (DIBO or DBCO), difluoro cycloalkynes and linear alkynes), phosphine (such as those used in traceless and non-traceless Staudinger ligations), haloacetyl (such as iodoacetamide), phosgene type reagents, sulfonyl chloride reagents, isothiocyanates, acyl halides, hydrazines, disulphides, vinyl sulfones, aziridines and photoreactive reagents (such as aryl azides, diaziridines).
- alkyne such as dibenzocyclooctynol (DIBO or DBCO), diflu
- Reactions between amino acids and functional groups may be spontaneous, such as cysteine/maleimide, or may require external reagents, such as Cu(I) for linking azide and linear alkynes.
- Linkers can comprise any molecule that stretches across the distance required. Linkers can vary in length from one carbon (phosgene-type linkers) to many Angstroms. Examples of linear molecules, include but are not limited to, are polyethyleneglycols (PEGs), polypeptides, polysaccharides, deoxyribonucleic acid (DNA), peptide nucleic acid (PNA), threose nucleic acid (TNA), glycerol nucleic acid (GNA), saturated and unsaturated hydrocarbons, polyamides. These linkers may be inert or reactive, in particular they may be chemically cleavable at a defined position, or may be themselves modified with a fluorophore or ligand. The linker is preferably resistant to dithiothreitol (DTT).
- DTT dithiothreitol
- Preferred crosslinkers include 2,5-dioxopyrrolidin-l-yl 3-(pyridin-2-yldisulfanyl)propanoate, 2,5-dioxopyrrolidin-l-yl 4-(pyridin-2-yldisulfanyl)butanoate and 2,5-dioxopyrrolidin-l-yl 8- (pyridin-2-yldisulfanyl)octananoate, di-maleimide PEG lk, di-maleimide PEG 3.4k, di- maleimide PEG 5k, di-maleimide PEG 10k, bis(maleimido)ethane (BMOE), bis- maleimidohexane (BMH), 1,4-bis-maleimidobutane (BMB), 1,4 bis-maleimidyl-2,3- dihydroxybutane (BMDB), BM[PEO]2 (1,8-bis-maleimidodiethylenegly
- the one or more linkers may be cleavable. This is discussed in more detail below.
- the two or more cysteines and/or non-natural amino acids may be connected using two different linkers that are specific for each other. One of the linkers is attached to one part and the other is attached to another part. The linkers should react to form a modified helicase of the invention.
- the two or more cysteines and/or non-natural amino acids may be connected using the hybridization linkers described in WO 2010/086602 (incorporated herein by reference in its entirety).
- the two or more cysteines and/or non- natural amino acids may be connected using two or more linkers each comprising a hybridizable region and a group capable of forming a covalent bond.
- the hybridizable regions in the linkers hybridize and link the two or more cysteines and/or non-natural amino acids.
- the linked cysteines and/or non-natural amino acids are then coupled via the formation of covalent bonds between the groups. Any of the specific linkers disclosed in WO 2010/086602 (incorporated herein by reference in its entirety) may be used in accordance with
- the two or more cysteines and/or non-natural amino acids may be modified and then attached using a chemical crosslinker that is specific for the two modifications. Any of the crosslinkers discussed above may be used.
- the linkers may be labeled. Suitable labels include, but are not limited to, fluorescent molecules (such as Cy3 or AlexaFluor®555), radioisotopes, e.g. 125 I, 35 S, enzymes, antibodies, antigens, polynucleotides and ligands such as biotin. Such labels allow the amount of linker to be quantified.
- the label could also be a cleavable purification tag, such as biotin, or a specific sequence to show up in an identification method, such as a peptide that is not present in the protein itself, but that is released by trypsin digestion.
- a preferred method of connecting two or more cysteines is via cysteine linkage. This can be mediated by a bi-functional chemical crosslinker or by an amino acid linker with a terminal presented cysteine residue.
- any bi-functional linker may be designed to ensure that the size of the opening is reduced sufficiently and the function of the helicase is retained.
- Suitable linkers include bismaleimide crosslinkers, such as 1,4- bis(maleimido)butane (BMB) or bis(maleimido)hexane.
- BMB 1,4- bis(maleimido)butane
- BMB bis(maleimido)hexane
- cysteine residues may be enhanced by modification of the adjacent residues, for example on a peptide linker. For instance, the basic groups of flanking arginine, histidine or lysine residues will change the pKa of the cysteines thiol group to that of the more reactive S’ group.
- cysteine residues may be protected by thiol protective groups such as 5,5'-dithiobis-(2-nitrobenzoic acid) (dTNB). These may be reacted with one or more cysteine residues of the helicase before a linker is attached. Selective deprotection of surface accessible cysteines may be possible using reducing reagents immobilized on beads (for example immobilized tris(2-carboxyethyl) phosphine, TCEP). Cysteine linkage is discussed in more detail below.
- Faz linkage Another preferred method of attachment via Faz linkage. This can be mediated by a bifunctional chemical linker or by a polypeptide linker with a terminal presented Faz residue.
- the helicase of the invention may also be modified to increase the attraction between (i) the tower domain and (ii) the pin domain and/or the 1A domain. Any known chemical modifications can be made in accordance with the invention. These types of modification are disclosed in WO 2015/055981 (incorporated herein by reference in its entirety).
- the invention provides a helicase of the invention in which at least one charged amino acid has further been introduced into (i) the tower domain and/or (ii) the pin domain and/or (iii) the 1A (RecA-like motor) domain, wherein the helicase has the ability to control the movement of a polynucleotide.
- the ability of the helicase to control the movement of a polynucleotide may be measured as discussed above.
- the invention preferably provides a helicase of the invention in which at least one charged amino acid has further been introduced into (i) the tower domain and (ii) the pin domain and/or the 1A domain.
- the at least one charged amino acid may be negatively charged or positively charged.
- the at least one charged amino acid is preferably oppositely charged to any amino acid(s) with which it interacts in the helicase.
- at least one positively charged amino acid may be introduced into the tower domain at a position which interacts with a negatively charged amino acid in the pin domain.
- the at least one charged amino acid is typically introduced at a position which is not charged in the wild-type (i.e. unmodified) helicase.
- the at least one charged amino acid may be used to replace at least one oppositely charged amino acid in the helicase.
- a positively charged amino acid may be used to replace a negatively charged amino acid.
- the at least one charged amino acid may be natural, such as arginine (R), histidine (H), lysine (K), aspartic acid (D) or glutamic acid (D).
- the at least one charged amino acid may be artificial or non-natural. Any number of charged amino acids may be introduced into each domain. For instance, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more charged amino acids may be introduced into each domain.
- the helicase preferably comprises a variant of SEQ ID NO: 118 which comprises a positively charged amino acid at one or more of the following positions: (i) 93; (ii) 354; (iii) 360; (iv) 361; (v) 94; (vi) 97; (vii) 155; (viii) 357; (ix) 100; and (x) 127.
- the helicase preferably comprises a variant of SEQ ID NO: 118 which comprises a negatively charged amino acid at one or more of the following positions: (i) 354; (ii) 358; (iii) 360; (iv) 364; (v) 97; (vi) 123; (vii) 155; (viii); 357; (ix) 100; and (x) 127.
- the helicase preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which comprises a positively charged amino acid or negatively charged amino acid at the positions which correspond to those in SEQ ID NO: 118 as defined in any of (i) to (x). Positions in any one of SEQ ID NOs: 119 to 133 which correspond to those in SEQ ID NO: 118 can be identified using the alignment of SEQ ID NOs: 118 to 133 below.
- the helicase preferably comprises a variant of SEQ ID NO: 118 which is modified by the introduction of at least one charged amino acid such that it comprises oppositely charged amino acid at the following positions: (i) 93 and 354; (ii) 93 and 358; (iii) 93 and 360; (iv) 93 and 361; (v) 93 and 364; (vi) 94 and 354; (vii) 94 and 358; (viii) 94 and 360; (ix) 94 and 361; (x) 94 and 364; (xi) 97 and 354; (xii) 97 and 358; (xiii) 97 and 360; (xiv) 97 and 361; (xv) 97 and 364; (xvi) 123 and 354; (xvii) 123 and 358; (xviii) 123 and 360; (xix) 123 and 361; (xx) 123 and 364; (xxi) 155 and 354; (xxii
- the helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises oppositely charged amino acids at the positions which correspond to those in SEQ ID NO: 118 as defined in any of (i) to (xxv).
- the helicase of the invention may further be one in which (i) at least one charged amino acid has been introduced into the tower domain and (ii) at least one oppositely charged amino acid has been introduced into the pin domain and/or the 1A (RecA-like motor) domain, wherein the helicase has the ability to control the movement of a polynucleotide.
- the at least one charged amino acid may be negatively charged and the at least one oppositely charged amino acid may be positively charged or vice versa.
- Suitable charged amino acids are discussed above. Any number of charged amino acids and any number of oppositely charged amino acids may be introduced. For instance, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more charged amino acids may be introduced and/or 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more oppositely charged amino acids may be introduced.
- the charged amino acids are typically introduced at positions which are not charged in the wild-type helicase.
- One or both of the charged amino acids may be used to replace charged amino acids in the helicase.
- a positively charged amino acid may be used to replace a negatively charged amino acid.
- the charged amino acids may be introduced at any of the positions in the (i) tower domain and (ii) pin domain and/or 1A domain discussed above.
- the oppositely charged amino acids are typically introduced such that they will interact in the resulting helicase.
- the helicase preferably comprises a variant of SEQ ID NO: 118 in which oppositely charged amino acids have been introduced at the following positions: (i) 97 and 354; (ii) 97 and 360; (iii) 155 and 354; or (iv) 155 and 360.
- the helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises oppositely charged amino acids at the positions which correspond to those in SEQ ID NO: 118 as defined in any of (i) to (iv).
- the invention also provides a construct comprising a modified helicase of the invention and an additional polynucleotide binding moiety, wherein the helicase is attached to the polynucleotide binding moiety and the construct has the ability to control the movement of a polynucleotide.
- the construct is artificial or non-natural.
- a construct of the invention is a useful tool for controlling the movement of a polynucleotide during Strand Sequencing.
- a construct of the invention is even less likely than a modified helicase of the invention to disengage from the polynucleotide being sequenced.
- the construct can provide even greater read lengths of the polynucleotide as it controls the translocation of the polynucleotide through a nanopore.
- a targeted construct that binds to a specific polynucleotide sequence can also be designed.
- the polynucleotide binding moiety may bind to a specific polynucleotide sequence and thereby target the helicase portion of the construct to the specific sequence.
- the construct has the ability to control the movement of a polynucleotide. This can be determined as discussed above.
- a construct of the invention may be isolated, substantially isolated, purified or substantially purified.
- a construct is isolated or purified if it is completely free of any other components, such as lipids, polynucleotides or pore monomers.
- a construct is substantially isolated if it is mixed with carriers or diluents which will not interfere with its intended use.
- a construct is substantially isolated or substantially purified if it is present in a form that comprises less than 10%, less than 5%, less than 2% or less than 1% of other components, such as lipids, polynucleotides or pore monomers.
- the helicase may be any of the helicases of the invention discussed above.
- the helicase is preferably covalently attached to the additional polynucleotide binding moiety.
- the helicase may be attached to the moiety at more than one, such as two or three, points.
- the helicase can be covalently attached to the moiety using any method known in the art. Suitable methods are discussed above with reference to connecting the two or more parts.
- the helicase and moiety may be produced separately and then attached together.
- the two components may be attached in any configuration. For instance, they may be attached via their terminal (i.e. amino or carboxy terminal) amino acids. Suitable configurations include, but are not limited to, the amino terminus of the moiety being attached to the carboxy terminus of the helicase and vice versa.
- the two components may be attached via amino acids within their sequences.
- the moiety may be attached to one or more amino acids in a loop region of the helicase.
- terminal amino acids of the moiety are attached to one or more amino acids in the loop region of a helicase.
- the helicase is chemically attached to the moiety, for instance via one or more linker molecules as discussed above.
- the helicase is genetically fused to the moiety.
- a helicase is genetically fused to a moiety if the whole construct is expressed from a single polynucleotide sequence.
- the coding sequences of the helicase and moiety may be combined in any way to form a single polynucleotide sequence encoding the construct. Genetic fusion of a pore to a nucleic acid binding protein is discussed in WO 2010/004265 (incorporated herein by reference in its entirety).
- the helicase and moiety may be genetically fused in any configuration.
- the helicase and moiety may be fused via their terminal amino acids.
- the amino terminus of the moiety may be fused to the carboxy terminus of the helicase and vice versa.
- the amino acid sequence of the moiety is preferably added in frame into the amino acid sequence of the helicase.
- the moiety is preferably inserted within the sequence of the helicase.
- the helicase and moiety are typically attached at two points, i.e. via the amino and carboxy terminal amino acids of the moiety.
- the amino and carboxy terminal amino acids of the moiety are in close proximity and are each attached to adjacent amino acids in the sequence of the helicase or variant thereof.
- the moiety is inserted into a loop region of the helicase.
- the helicase may be attached directly to the moiety.
- the helicase is preferably attached to the moiety using one or more, such as two or three, linkers as discussed above.
- the one or more linkers may be designed to constrain the mobility of the moiety.
- the helicase and/or the moiety may be modified to facilitate attachment of the one or more linker as discussed above.
- Cleavable linkers can be used as an aid to separation of constructs from non-attached components and can be used to further control the synthesis reaction.
- a hetero-bifunctional linker may react with the helicase, but not the moiety. If the free end of the linker can be used to bind the helicase protein to a surface, the unreacted helicases from the first reaction can be removed from the mixture. Subsequently, the linker can be cleaved to expose a group that reacts with the moiety.
- conditions may be optimised first for the reaction to the helicase, then for the reaction to the moiety after cleavage of the linker. The second reaction would also be much more directed towards the correct site of reaction with the moiety because the linker would be confined to the region to which it is already attached.
- the helicase may be covalently attached to the bifunctional crosslinker before the helicase/crosslinker complex is covalently attached to the moiety.
- the moiety may be covalently attached to the bifunctional crosslinker before the bifunctional crosslinker/moiety complex is attached to the helicase.
- the helicase and moiety may be covalently attached to the chemical crosslinker at the same time.
- Preferred methods of attaching the helicase to the moiety are cysteine linkage and Faz linkage as described above.
- a reactive cysteine is presented on a peptide linker that is genetically attached to the moiety. This means that additional modifications will not necessarily be needed to remove other accessible cysteine residues from the moiety.
- Cross-linkage of helicases or moieties to themselves may be prevented by keeping the concentration of linker in a vast excess of the helicase and/or moiety.
- a "lock and key" arrangement may be used in which two linkers are used. Only one end of each linker may react together to form a longer linker and the other ends of the linker each react with a different part of the construct (i.e. helicase or moiety). This is discussed in more detail below.
- the site of attachment is selected such that, when the construct is contacted with a polynucleotide, both the helicase and the moiety can bind to the polynucleotide and control its movement.
- the invention also provides a method of producing a construct of the invention, comprising attaching a helicase of the invention to an additional polynucleotide binding moiety and thereby producing the construct.
- the method preferably further comprises determining whether or not the resulting construct is capable of controlling the movement of a polynucleotide. All of the construct embodiments discussed above equally apply to the method.
- Pores and pore complexes for use in the invention are substantially identical to Pores and pore complexes for use in the invention.
- the invention provides various methods and products in which a modified Dda helicase or a construct of the invention is combined/used with a transmembrane pore.
- the transmembrane pore may be used a solid-state pore.
- the transmembrane pore may be a transmembrane protein pore.
- the transmembrane protein pore may be derived from a hemolysin, such as alpha hemolysin or gamma hemolysin, leukocidin, Mycobacterium smegmatis porin A (MspA), MspB, MspC, MspD, PorARr, PorBRr, PorARc, lysenin, outer membrane porin F (OmpF), outer membrane porin G (OmpG), outer membrane phospholipase A, Neisseria autotransporter lipoprotein (NalP) and WZA, CsgG, CsgG/CsgF, ClyA, Spl, haemolytic protein fragaceatoxin C (FraC), Iota toxin, Anthrax protective antigen, Vibrio cholerae cytolysin, Cytotoxin K (CytK), CELIII, Aerolysin, InvG, GspD, PilQ, necrotic enteritis
- the transmembrane protein pore preferably comprises a CsgG pore or a homologue or mutant thereof, or an isolated pore complex comprising a CsgG pore, or a homologue or mutant thereof, and a modified CsgF peptide, or a homologue or mutant thereof.
- the CsgG pore or a homologue or mutant thereof, the isolated pore complex comprising a CsgG pore, or a homologue or mutant thereof, and a modified CsgF peptide, or a homologue or mutant thereof preferably comprises at least one mutant monomer.
- a mutant CsgG monomer is a monomer whose sequence varies from that of a wild-type CsgG monomer and which retains the ability to form a pore.
- a mutant monomer may also be referred to herein as a variant. Methods for confirming the ability of mutant monomers to form pores are well-known in the art and are discussed in more detail below.
- the at least one mutant monomer or variant may have any of the %s of homology/sequence identity to SEQ ID NO: 117 or SEQ ID NO: 3 set out below.
- the at least one mutant monomer may contain any of the modifications, mutations or substitutions described below, including the types of modifications and substitutions described with reference to the Dda helicases of the invention.
- the at least one mutant monomer may contain any of the additional modifications, mutations or substitutions described in WO2016/034591, WO2017/149316, WO2017/149317 and, WO2017/149318, WO2018/211241, and W02019/002893 (all incorporated by reference herein in their entirety).
- the invention provides a method of determining the presence, absence or one or more characteristics of a target analyte.
- the method involves contacting the target analyte with a transmembrane pore and a helicase of the invention or a construct of the invention such that the helicase or construct controls the movement of the target analyte with respect to, such as into or through, the pore and taking one or more measurements as the analyte moves with respect to the pore and thereby determining the presence, absence or one or more characteristics of the analyte.
- the target analyte may also be called the template analyte or the analyte of interest.
- the transmembrane pore may be any of the pores or complexes discussed above.
- the transmembrane pore typically comprises at least 7, at least 8, at least 9 or at least 10 monomers, such as 7, 8, 9 or 10 CsgG monomers.
- the transmembrane pore preferably comprises eight or nine identical CsgG monomers.
- One or more, such as 2, 3, 4, 5, 6, 7, 8, 9 or 10, of the CsgG monomers is preferably chemically modified, or the CsgF peptide is chemically modified.
- the monomers, such as the CsgG monomers, or homologues or mutants thereof, and the modified CsgF monomers, or homologues or mutants thereof, may be derived from any organism.
- the analyte may pass through the CsgG constriction, followed by the CsgF constriction. In an alternative embodiment the analyte may pass through the CsgF constriction, followed by the CsgG constriction, depending on the orientation of the CsgG/CsgF complex in the membrane.
- the method is for determining the presence, absence or one or more characteristics of a target analyte.
- the method may be for determining the presence, absence or one or more characteristics of at least one analyte.
- the method may concern determining the presence, absence or one or more characteristics of two or more analytes.
- the method may comprise determining the presence, absence or one or more characteristics of any number of analytes, such as 2, 5, 10, 15, 20, 30, 40, 50, 100 or more analytes. Any number of characteristics of the one or more analytes may be determined, such as 1, 2, 3, 4, 5, 10 or more characteristics.
- the degree of reduction in ion flow is related to the size of the obstruction within, or in the vicinity of, the pore. Binding of a molecule of interest, also referred to as an "analyte", in or near the pore therefore provides a detectable and measurable event, thereby forming the basis of a "biological sensor".
- Suitable molecules for nanopore sensing include nucleic acids; proteins; peptides; polysaccharides and small molecules (refers here to a low molecular weight (e.g., ⁇ 900Da or ⁇ 500Da) organic or inorganic compound) such as pharmaceuticals, toxins, cytokines, and pollutants. Detecting the presence of biological molecules finds application in personalised drug development, medicine, diagnostics, life science research, environmental monitoring and in the security and/or the defence industry.
- the transmembrane pore may serve as a molecular or biological sensor.
- the transmembrane pore can be derived or isolated from bacterial proteins (e.g., E. coli, Salmonella typhi).
- the transmembrane pore can be recombinantly produced. Procedures for analyte detection are described in Howorka et al. Nature Biotechnology (2012) Jun 7; 30(6):506-7.
- the analyte molecule that is to be detected may bind to either face of the channel, or within the lumen of the channel itself. The position of binding may be determined by the size of the molecule to be sensed.
- the one or more characteristics of the target analyte are preferably measured by electrical measurement and/or optical measurement.
- the electrical measurement is preferably a current measurement, an impedance measurement, a tunnelling measurement or a field effect transistor (FET) measurement.
- FET field effect transistor
- the method preferably further comprises the step of applying a voltage across the pore to form a complex between the pore and the helicase or construct.
- the target analyte is preferably a metal ion, an inorganic salt, a polymer, an amino acid, a peptide, a polypeptide, a protein, a nucleotide, an oligonucleotide, a polynucleotide, a monosaccharide, an oligosaccharide, a polysaccharide, a dye, a bleach, a pharmaceutical, a diagnostic agent, a recreational drug, an explosive, a toxic compound, or an environmental pollutant.
- the target analyte is preferably a target polynucleotide.
- the method may concern determining the presence, absence or one or more characteristics of two or more analytes of the same type, such as two or more proteins, two or more nucleotides or two or more pharmaceuticals.
- the method may concern determining the presence, absence or one or more characteristics of two or more analytes of different types, such as one or more proteins, one or more nucleotides and one or more pharmaceuticals.
- the target analyte preferably the target polynucleotide, may be modified by methylation, by oxidation, by damage, with one or more proteins or with one or more labels, tags or spacers.
- the target analyte can be secreted from cells.
- the target analyte can be an analyte that is present inside cells such that the analyte must be extracted from the cells before the method can be carried out.
- the target analyte is an amino acid, a peptide, a polypeptides or protein.
- the amino acid, peptide, polypeptide or protein can be naturally-occurring or non- naturally-occurring.
- the polypeptide or protein can include within them synthetic or modified amino acids. Several different types of modification to amino acids are known in the art. Suitable amino acids and modifications thereof are above. It is to be understood that the target analyte can be modified by any method available in the art.
- the target analyte is a polynucleotide, such as a nucleic acid, which is defined as a macromolecule comprising two or more nucleotides.
- Nucleic acids are particularly suitable for nanopore sequencing.
- the naturally-occurring nucleic acid bases in DNA and RNA may be distinguished by their physical size. As a nucleic acid molecule, or individual base, passes through the channel of a nanopore, the size differential between the bases causes a directly correlated reduction in the ion flow through the channel. The variation in ion flow may be recorded. Suitable electrical measurement techniques for recording ion flow variations are discussed above.
- the characteristic reduction in ion flow can be used to identify the particular nucleotide and associated base traversing the channel in real-time.
- the open-channel ion flow is reduced as the individual nucleotides of the nucleic sequence of interest sequentially pass through the channel of the nanopore due to the partial blockage of the channel by the nucleotide. It is this reduction in ion flow that is measured using the suitable recording techniques described above.
- the reduction in ion flow may be calibrated to the reduction in measured ion flow for known nucleotides through the channel resulting in a means for determining which nucleotide is passing through the channel, and therefore, when done sequentially, a way of determining the nucleotide sequence of the nucleic acid passing through the nanopore.
- it has typically required for the reduction in ion flow through the channel to be directly correlated to the size of the individual nucleotide passing through the constriction (or "reading head").
- sequencing may be performed upon an intact nucleic acid polymer that is 'threaded' through the pore via the action of an associated polymerase, for example.
- sequences may be determined by passage of nucleotide triphosphate bases that have been sequentially removed from a target nucleic acid in proximity to the pore (see for example WO 2014/187924 incorporated herein by reference in its entirety).
- the polynucleotide may be single stranded or double stranded. At least a portion of the polynucleotide is preferably double stranded.
- the polynucleotide is most preferably ribonucleic nucleic acid (RIMA) or deoxyribonucleic acid (DNA).
- said method using a polynucleotide as an analyte alternatively comprises determining one or more characteristics selected from (i) the length of the polynucleotide, (ii) the identity of the polynucleotide, (iii) the sequence of the polynucleotide, (iv) the secondary structure of the polynucleotide and (v) whether or not the polynucleotide is modified.
- the polynucleotide can be any length (i).
- the polynucleotide can be at least 10, at least 50, at least 100, at least 150, at least 200, at least 250, at least 300, at least 400 or at least 500 nucleotides or nucleotide pairs in length.
- the polynucleotide can be 1000 or more nucleotides or nucleotide pairs, 5000 or more nucleotides or nucleotide pairs in length or 100000 or more nucleotides or nucleotide pairs in length. Any number of polynucleotides can be investigated. For instance, the method may concern characterising 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 30, 50, 100 or more polynucleotides.
- polynucleotides may be different polynucleotides or two instances of the same polynucleotide.
- the polynucleotide can be naturally occurring or artificial.
- the method may be used to verify the sequence of a manufactured oligonucleotide. The method is typically carried out in vitro.
- Embodiments involving the modified helicases and constructs of the invention The invention provides polynucleotides, vectors and host cells. These are discussed above.
- the invention provides various methods using the Dda helicases of the invention in particular for controlling the movement of an analyte and characterising a target analyte. Such methods are described above and in WO2016/034591, WO2017/149316, WO2017/149317, WO2017/149318, WO2018/211241, W02019/002893, WO2015/055981, WO2015/166276 and WO2016/055777 (all incorporated by reference herein).
- the modified Dda helicases and constructs of the invention may be used to form sensors for characterising target analytes. These sensors may be formed by contacting the pore and the helicase or construct in the presence of the target analyte and applying a potential across the pore.
- the potential may be a voltage potential or a chemical potential.
- the helicase or the construct may be covalently attached to the pore, for instance as described above.
- the modified Dda helicases and constructs of the invention may be combined/used with a solid state pore or a transmembrane protein pore.
- the transmembrane protein pore may be any of those discussed above. Kits
- the invention also provides a kit for characterising a target analyte comprising (a) a pore and a helicase or a construct of the invention or (b) a helicase or construct of the invention and one or more loading moieties.
- a target analyte comprising (a) a pore and a helicase or a construct of the invention or (b) a helicase or construct of the invention and one or more loading moieties.
- the pore may be any of those discussed above. Preferred combinations of pores and helicases are discussed in more detail below.
- the kit preferably further comprises the components of a membrane.
- the kit may comprise components of any type of membranes, such as an amphiphilic layer or a triblock copolymer membrane.
- the kit may further comprise one or more anchors, such as cholesterol, for coupling the target analyte to the membrane.
- the kit may further comprise one or more polynucleotide adaptors that can be attached to a target polynucleotide to facilitate characterisation of the polynucleotide.
- the anchor such as cholesterol, is attached to the polynucleotide adaptor.
- the kit may additionally comprise one or more other reagents or instruments which enable any of the embodiments mentioned above to be carried out.
- reagents or instruments include one or more of the following: suitable buffer(s) (aqueous solutions), means to obtain a sample from a subject (such as a vessel or an instrument comprising a needle), means to amplify and/or express polynucleotides or voltage or patch clamp apparatus.
- Reagents may be present in the kit in a dry state such that a fluid sample resuspends the reagents.
- the kit may also, optionally, comprise instructions to enable the kit to be used in the method of the invention or details regarding for which organism the method may be used.
- the kit may also comprise additional components useful in analyte characterization.
- Preferred polynucleotide binding proteins are polymerases, exonucleases, helicases and topoisomerases, such as gyrases.
- Preferred polynucleotide binding moieties are or are derived from polymerases, exonucleases, helicases and topoisomerases, such as gyrases.
- Suitable enzymes include, but are not limited to, exonuclease I from E. coli, exonuclease III enzyme from E. coli, Red from T. thermophilus and bacteriophage lambda exonuclease, TatD exonuclease and variants thereof. Three subunits comprising the Red sequence from T.
- thermophilus or a variant thereof interact to form a trimer exonuclease.
- the polymerase may be PyroPhage® 3173 DNA Polymerase (which is commercially available from Lucigen® Corporation), SD Polymerase (commercially available from Bioron®) or variants thereof.
- the enzyme may be Phi29 DNA polymerase (SEQ ID NO: 7) or a variant thereof.
- the topoisomerase is preferably a member of any of the Moiety Classification (EC) groups 5.99.1.2 and 5.99.1.3.
- the enzyme is most preferably derived from a helicase, such as Hel308 Mbu, Hel308 Csy, Hel308 Tga, Hel308 Mhu, Tral Eco, XPD Mbu or a variant thereof.
- a helicase such as Hel308 Mbu, Hel308 Csy, Hel308 Tga, Hel308 Mhu, Tral Eco, XPD Mbu or a variant thereof.
- Any helicase may be used in the invention.
- the helicase may be or be derived from a Hel308 helicase, a RecD helicase, such as Tral helicase or a TrwC helicase, a XPD helicase or a Dda helicase.
- the helicase may be any of the helicases, modified helicases or helicase constructs disclosed in WO 2013/057495; WO 2013/098562; WO2013098561; WO 2014/013260; WO 2014/013259; WO 2014/013262 and WO 2015/055981. All of these are incorporated by reference in their entirety.
- the invention also provides an apparatus for characterising target analytes in a sample, comprising (a) a plurality of pores and (b) a plurality of helicases or a plurality of constructs of the invention.
- the plurality of pores may be any of those discussed above. Preferred combinations of pores and helicases are discussed in more detail below.
- the invention may use an array comprising a plurality of membranes. Any of the embodiments discussed above with respect to the membrane equally apply the array of the invention.
- the array may be set up to perform any of the methods described above.
- each membrane in the array comprises one pore. Due to the manner in which the array is formed, for example, the array may comprise one or more membranes that do not comprise a pore, and/or one or more membranes that comprise two or more pores.
- the array may comprise from about 2 to about 1000, such as from about 10 to about 800, from about 20 to about 600 or from about 30 to about 500 membranes.
- the invention may use a system comprising (a) a membrane or an array, (b) means for applying a potential across the membrane(s) and (c) means for detecting electrical or optical signals across the membrane(s).
- the pores and membranes may be any as described above and below.
- the system further comprises a first chamber and a second chamber, wherein the first and second chambers are separated by the membrane(s).
- the system may further comprise a target analyte, wherein the target analyte is transiently located within the continuous channel and wherein one end of the target analyte is located in the first chamber and one end of the target analyte is located in the second chamber.
- the target analyte is preferably a target polypeptide or a target polynucleotide.
- the system further comprises an electrically conductive solution in contact with the pore(s), electrodes providing a voltage potential across the membrane(s), and a measurement system for measuring the current through the pore(s).
- the voltage applied across the membranes and pore is from +5 V to -5 V, such as -600 mV to +600mV or -400 mV to +400 mV.
- the voltage used is preferably in the range 100 mV to 240 mV and more preferably in the range of 120 mV to 220 mV. It is possible to increase discrimination between different amino acids or nucleotides by a pore by using an increased applied potential. Any suitable electrically conductive solution may be used.
- the solution may comprise charge carriers, such as metal salts, for example alkali metal salt, halide salts, for example chloride salts, such as alkali metal chloride salt.
- Charge carriers may include ionic liquids or organic salts, for example tetramethyl ammonium chloride, trimethylphenyl ammonium chloride, phenyltrimethyl ammonium chloride, or l-ethyl-3-methyl imidazolium chloride.
- salt is present in the aqueous solution in the chamber. Potassium chloride (KCI), sodium chloride (NaCI), caesium chloride (CsCI) or a mixture of potassium ferrocyanide and potassium ferricyanide is typically used.
- KCI, NaCI and a mixture of potassium ferrocyanide and potassium ferricyanide are preferred.
- the charge carriers may be asymmetric across the membrane. For instance, the type and/or concentration of the charge carriers may be different on each side of the membrane, e.g., in each chamber.
- the salt concentration may be at saturation.
- the salt concentration may be 3 M or lower and is typically from 0.1 to 2.5 M, from 0.3 to 1.9 M, from 0.5 to 1.8 M, from 0.7 to 1.7 M, from 0.9 to 1.6 M or from 1 M to 1.4 M.
- the salt concentration is preferably from 150 mM to 1 M.
- the method is preferably carried out using a salt concentration of at least 0.3 M, such as at least 0.4 M, at least 0.5 M, at least 0.6 M, at least 0.8 M, at least 1.0 M, at least 1.5 M, at least 2.0 M, at least 2.5 M or at least 3.0 M.
- High salt concentrations provide a high signal to noise ratio and allow for currents indicative of the presence of an amino acid or nucleotide to be identified against the background of normal current fluctuations.
- a buffer may be present in the electrically conductive solution.
- the buffer is phosphate buffer.
- Other suitable buffers are HEPES and Tris-HCI buffer.
- the pH of the electrically conductive solution may be from 4.0 to 12.0, from 4.5 to 10.0, from 5.0 to 9.0, from 5.5 to 8.8, from 6.0 to 8.7 or from 7.0 to 8.8 or 7.5 to 8.5.
- the pH used is preferably about 7.5.
- the system may be comprised in an apparatus.
- the apparatus may be any conventional apparatus for analyte analysis, such as an array or a chip.
- the apparatus is preferably set up to carry out the disclosed method.
- the apparatus may comprise a chamber comprising an aqueous solution and a barrier that separates the chamber into two sections.
- the barrier typically has an aperture in which the membrane(s) containing the pore(s) are formed.
- the barrier forms the membrane in which the pore is present.
- the apparatus may also comprise an electrical circuit capable of applying a potential and measuring an electrical signal across the membrane and pore.
- the apparatus may be any of those described in WO 2008/102120, WO 2009/077734, WO 2010/122293, WO 2011/067559, or WO 00/28312 (all incorporated herein by reference in their entirety).
- the membrane is preferably an amphiphilic layer.
- An amphiphilic layer is a layer formed from amphiphilic molecules, such as phospholipids, which have both hydrophilic and lipophilic properties.
- the amphiphilic molecules may be synthetic or naturally occurring. Non-naturally occurring amphiphiles and amphiphiles which form a monolayer are known in the art and include, for example, block copolymers (Gonzalez-Perez et al., Langmuir, 2009, 25, 10447-10450). Block copolymers are polymeric materials in which two or more monomer sub-units that are polymerized together to create a single polymer chain.
- Block copolymers typically have properties that are contributed by each monomer sub-unit. However, a block copolymer may have unique properties that polymers formed from the individual sub-units do not possess. Block copolymers can be engineered such that one of the monomer sub-units is hydrophobic (i.e., lipophilic), whilst the other sub-unit(s) are hydrophilic whilst in aqueous media. In this case, the block copolymer may possess amphiphilic properties and may form a structure that mimics a biological membrane.
- the block copolymer may be a diblock (consisting of two monomer sub-units) but may also be constructed from more than two monomer sub-units to form more complex arrangements that behave as amphipiles.
- the copolymer may be a triblock, tetrablock or pentablock copolymer.
- the membrane is preferably a triblock copolymer membrane.
- the membrane is most preferably one of the membranes disclosed in International Application No. WO2014/064443 or WO2014/064444.
- the amphiphilic molecules may be chemically modified or functionalised to facilitate coupling of the polynucleotide.
- the amphiphilic layer may be a monolayer or a bilayer.
- the amphiphilic layer is typically planar.
- the amphiphilic layer may be curved.
- the amphiphilic layer may be supported.
- Amphiphilic membranes are typically naturally mobile, essentially acting as two-dimensional fluids with lipid diffusion rates of approximately IO -8 cm s’ 1 . This means that the pore and coupled polynucleotide can typically move within an amphiphilic membrane.
- the membrane may be a lipid bilayer.
- Lipid bilayers are models of cell membranes and serve as excellent platforms for a range of experimental studies.
- lipid bilayers can be used for in vitro investigation of membrane proteins by single-channel recording.
- lipid bilayers can be used as biosensors to detect the presence of a range of substances.
- the lipid bilayer may be any lipid bilayer. Suitable lipid bilayers include, but are not limited to, a planar lipid bilayer, a supported bilayer, or a liposome.
- the lipid bilayer is preferably a planar lipid bilayer. Suitable lipid bilayers are disclosed in WO 2008/102121, WO 2009/077734, and WO 2006/100484 (all incorporated herein by reference in their entirety).
- the membrane comprises a solid-state layer.
- Solid state layers can be formed from both organic and inorganic materials including, but not limited to, microelectronic materials, insulating materials such as SisIX , AI2O3, and SiO, organic and inorganic polymers such as polyamide, plastics such as Teflon® or elastomers such as two-component addition-cure silicone rubber, and glasses.
- the solid-state layer may be formed from graphene. Suitable graphene layers are disclosed in WO 2009/035647 (incorporated herein by reference in its entirety).
- the pore is typically present in an amphiphilic membrane or layer contained within the solid-state layer, for instance within a hole, well, gap, channel, trench or slit within the solid-state layer.
- amphiphilic membrane or layer contained within the solid-state layer for instance within a hole, well, gap, channel, trench or slit within the solid-state layer.
- suitable solid state/amphiphilic hybrid systems are disclosed in WO 2009/020682 and WO 2012/005857 (both incorporated herein by reference in their entirety). Any of the amphiphilic membranes or layers discussed above may be used.
- the method is typically carried out using (i) an artificial amphiphilic layer comprising a pore, (ii) an isolated, naturally occurring lipid bilayer comprising a pore, or (iii) a cell having a pore inserted therein.
- the method is typically carried out using an artificial amphiphilic layer, such as an artificial triblock copolymer layer.
- the layer may comprise other transmembrane and/or intramembrane proteins as well as other molecules in addition to the pore. Suitable apparatus and conditions are discussed below.
- the method of the invention is typically carried out in vitro.
- the modified helicase of the invention may be used with a pore or a transmembrane pore.
- the modified helicase of the invention may be used in combination with any of the pores described in WO2016/034591, WO2017/149316, WO2017/149317, WO2017/149318, WO2018/211241, W02019/002893 and PCT/EP2023/059821 (all incorporated by reference herein in their entirety).
- the modified helicase of the invention is preferably used with a CsgG pore as described in WO2016/034591 (incorporated herein by reference) as in Example 2 or a CsgG:CsgF pore complex as described in W02019/002893 (incorporated herein by reference).
- the helicase of the invention is preferably used with a CsgG pore or a CsgG:CsgF pore complex described above and vice versa.
- the CsgG pore or a CsgG:CsgF pore complex of the invention preferably comprises Q100A and/or N102A or N102S and the helicases preferably comprises any of the substitutions in PCT/EP2023/059821, preferably Y350I and/or K358I.
- SEQ ID NO: 1 shows polynucleotide sequence of wild-type E. coli CsgG from strain K12, including signal sequence (Gene ID: 945619).
- SEQ ID NO:2 shows amino acid sequence of wild-type E. coli CsgG including signal sequence (Uniprot accession number P0AEA2).
- SEQ ID NO:3 shows amino acid sequence of wild-type E. coli CsgG as mature protein (Uniprot accession number P0AEA2).
- SEQ ID NO:4 shows polynucleotide sequence of wild-type E. coli CsgF from strain K12, including signal sequence (Gene ID: 945622).
- SEQ ID NO:5 shows amino acid sequence of wild-type E. coli CsgF including signal sequence (Uniprot accession number P0AE98).
- SEQ ID NO:6 shows amino acid sequence of wild-type E. coli CsgF as mature protein (Uniprot accession number P0AE98).
- SEQ ID N0:7 shows polynucleotide sequence of a fragment of wild-type E. coli CsgF encoding amino acids 1 to 27 and a C-terminal 6 His tag.
- SEQ ID NO:8 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 1 to 27 and a C-terminal 6 His tag.
- SEQ ID NO:9 shows polynucleotide sequence of a fragment of wild-type E. coli CsgF encoding amino acids 1 to 38 and a C-terminal 6 His tag.
- SEQ ID NO: 10 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 1 to 38 and a C-terminal 6 His tag.
- SEQ ID NO: 11 shows polynucleotide sequence of a fragment of wild-type E. coli CsgF encoding amino acids 1 to 48 and a C-terminal 6 His tag.
- SEQ ID NO: 12 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 1 to 48 and a C-terminal 6 His tag.
- SEQ ID NO: 13 shows polynucleotide sequence of a fragment of wild-type E. coli CsgF encoding amino acids 1 to 64 and a C-terminal 6 His tag.
- SEQ ID NO: 14 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 1 to 64 and a C-terminal 6 His tag.
- SEQ ID NO: 15 shows amino acid sequence of a peptide corresponding to residues 20 to 53 of E. coli CsgF
- SEQ ID NO: 16 shows amino acid sequence of a peptide corresponding to residues 20 to 42 of E. coli CsgF, including KD at its C-terminus
- SEQ ID NO: 17 shows amino acid sequence of a peptide corresponding to residues 23 to 55 of CsgF homologue Q88H88
- SEQ ID NO: 18 shows amino acid sequence of a peptide corresponding to residues 25 to 57 of CsgF homologue A0A143HJA0
- SEQ ID NO: 19 shows amino acid sequence of a peptide corresponding to residues 21 to 53 of CsgF homologue Q5E245
- SEQ ID NO:20 shows amino acid sequence of a peptide corresponding to residues 19 to 51 of CsgF homologue Q084E5
- SEQ ID NO:21 shows amino acid sequence of a peptide corresponding to residues 15 to 47 of CsgF homologue F0LZU2
- SEQ ID NO:22 shows amino acid sequence of a peptide corresponding to residues 26 to 58 of CsgF homologue A0A136HQR0
- SEQ ID NO:23 shows amino acid sequence of a peptide corresponding to residues 21 to 53 of CsgF homologue A0A0W1SRL3
- SEQ ID NO:24 shows amino acid sequence of a peptide corresponding to residues 26 to 59 of CsgF homologue B0UH01
- SEQ ID NO:25 shows amino acid sequence of a peptide corresponding to residues 22 to 53 of CsgF homologue Q6NAU5
- SEQ ID NO:26 shows amino acid sequence of a peptide corresponding to residues 7 to 38 of CsgF homologue G8PUY5
- SEQ ID NO:27 shows amino acid sequence of a peptide corresponding to residues 25 to 57 of CsgF homologue A0A0S2ETP7
- SEQ ID NO:28 shows amino acid sequence of a peptide corresponding to residues 19 to 51 of CsgF homologue E3I1Z1
- SEQ ID NO:29 shows amino acid sequence of a peptide corresponding to residues 24 to 55 of CsgF homologue F3Z094
- SEQ ID NO:30 shows amino acid sequence of a peptide corresponding to residues 21 to 53 of CsgF homologue A0A176T7M2
- SEQ ID NO:31 shows amino acid sequence of a peptide corresponding to residues 14 to 45 of CsgF homologue D2QPP8
- SEQ ID NO:32 shows amino acid sequence of a peptide corresponding to residues 28 to 58 of CsgF homologue N2IYT1
- SEQ ID NO:33 shows amino acid sequence of a peptide corresponding to residues 26 to 58 of CsgF homologue W7QHV5
- SEQ ID NO:34 shows amino acid sequence of a peptide corresponding to residues 23 to 55 of CsgF homologue D4ZLW2
- SEQ ID NO:35 shows amino acid sequence of a peptide corresponding to residues 21 to 53 of CsgF homologue D2QT92
- SEQ ID NO:36 shows amino acid sequence of a peptide corresponding to residues 20 to 51 of CsgF homologue A0A167UJA2
- SEQ ID NO:37 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 20 to 27.
- SEQ ID NO:38 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 20 to 38.
- SEQ ID NO:39 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 20 to 48.
- SEQ ID NO:40 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 20 to 64.
- SEQ ID NO:41 shows the nucleotide sequence of primer CsgF_d27_end
- SEQ ID NO:42 shows the nucleotide sequence of primer CsgF_d38_end
- SEQ ID NO:43 shows the nucleotide sequence of primer CsgF_d48_end
- SEQ ID NO:44 shows the nucleotide sequence of primer CsgF_d64_end
- SEQ ID NO:45 shows the nucleotide sequence of primer pNa62_CsgF_histag_Fw
- SEQ ID NO:46 shows the nucleotide sequence of primer CsgF-His_pET22b_FW
- SEQ ID NO:47 shows the nucleotide sequence of primer CsgF-His_pET22b_Rev
- SEQ ID NO:48 shows the nucleotide sequence of primer csgEFG_pDONR221_FW
- SEQ ID NO:49 shows the nucleotide sequence of primer csgEFG_pDONR221_Rev
- SEQ ID NO: 50 shows the nucleotide sequence of primer Mut_csgF_His_FW
- SEQ ID NO:51 shows the nucleotide sequence of primer Mut_csgF_His_Rev
- SEQ ID NO:52 shows the nucleotide sequence of primer DelCsgE_Rev
- SEQ ID NO:53 shows the nucleotide sequence of primer DelCsgE FW
- SEQ ID NO: 54 shows the amino acid sequence of residues 1 to 30 of mature E. coli CsgF
- SEQ ID NO: 55 shows the amino acid sequence of residues 1 to 35 of mature E. coli CsgF
- SEQ ID NO: 56 shows the amino acid sequence of a mutated (T4C/N17S) CsgF sequence with a signal sequence, and a TEV protease cleavage site (ENLYFQS) inserted between residues 35 and 36 of sequence of the mature protein.
- SEQ ID NO: 57 shows the amino acid sequence of a mutated (N17S-Del) CsgF sequence with a signal sequence, and a TEV protease cleavage site (ENLYFQS) inserted between residues 35 and 36 of sequence of the mature protein.
- SEQ ID NO: 58 shows the amino acid sequence of a mutated (G1C/N17S) CsgF sequence with a signal sequence, and a TEV protease cleavage site (ENLYFQS) inserted between residues 35 and 36 of sequence of the mature protein.
- SEQ ID NO: 59 shows the amino acid sequence of a mutated (G1C) CsgF sequence with a signal sequence, and a TEV protease cleavage site (ENLYFQS) inserted between residues 35 and 36 of sequence of the mature protein.
- SEQ ID NO: 60 shows the amino acid sequence of a CsgF sequence with a signal sequence, a TEV protease cleavage site (ENLYFQS) inserted between residues 45 and 46 of sequence of the mature protein, and a Hisio tag at the C-terminus.
- ENLYFQS TEV protease cleavage site
- SEQ ID NO: 61 shows the amino acid sequence of a CsgF sequence with a signal sequence, a TEV protease cleavage site (ENLYFQS) inserted between residues 35 and 36 of sequence of the mature protein, and a Hisio tag at the C-terminus.
- ENLYFQS TEV protease cleavage site
- SEQ ID NO: 62 shows the amino acid sequence of a CsgF sequence with a signal sequence, a TEV protease cleavage site (ENLYFQS) inserted between residues 30 and 31 of sequence of the mature protein, and a Hisio tag at the C-terminus.
- ENLYFQS TEV protease cleavage site
- SEQ ID NO: 63 shows the amino acid sequence of a CsgF sequence with a signal sequence, a TEV protease cleavage site (ENLYFQS) inserted between residues 45 and 51 of sequence of the mature protein, and a Hisio tag at the C-terminus.
- ENLYFQS TEV protease cleavage site
- SEQ ID NO: 64 shows the amino acid sequence of a CsgF sequence with a signal sequence, a TEV protease cleavage site (ENLYFQS) inserted between residues 30 and 37 of sequence of the mature protein, and a Hisio tag at the C-terminus.
- ENLYFQS TEV protease cleavage site
- SEQ ID NO: 65 shows the amino acid sequence of a CsgF sequence with a signal sequence, a HCV C3 protease cleavage site (LEVLFQGP) inserted between residues 34 and 36 of sequence of the mature protein, and a Hisio tag at the C-terminus.
- LCVLFQGP HCV C3 protease cleavage site
- SEQ ID NO: 66 shows the amino acid sequence of a CsgF sequence with a signal sequence, a HCV C3 protease cleavage site (LEVLFQGP) inserted between residues 42 and 43 of sequence of the mature protein, and a Hisio tag at the C-terminus.
- SEQ ID NO: 67 shows the amino acid sequence of a CsgF sequence with a signal sequence, a HCV C3 protease cleavage site (LEVLFQGP) inserted between residues 38 and 47 of sequence of the mature protein, and a Hisio tag at the C-terminus.
- SEQ ID NO: 68 shows the amino acid sequence of YP_001453594.1 : 1-248 of hypothetical protein CKO_02032 [Citrobacter koseri ATCC BAA-895], which is 99% identical to SEQ ID NO: 3.
- SEQ ID NO: 69 shows the amino acid sequence of WP_001787128.1: 16-238 of curli production assembly/transport component CsgG, partial [Salmonella enterica], which is 98% to SEQ ID NO: 3.
- SEQ ID NO: 70 shows the amino acid sequence of KEY44978.1
- SEQ ID NO: 71 shows the amino acid sequence of YP_003364699.1 : 16-277 of curli production assembly/transport component [Citrobacter rodentium ICC168], which is 97% identical to SEQ ID NO: 3.
- SEQ ID NO: 72 shows the amino acid sequence of YP_004828099.1 : 16-277 of curli production assembly/transport component CsgG [Enterobacter asburiae LF7a], which is 94% identical to SEQ ID NO: 3.
- SEQ ID NO: 73 shows the amino acid sequence of WP_006819418.1: 19-280 of transporter [Yokenella regensburgei], which is 91% identical to SEQ ID NO: 3.
- SEQ ID NO: 74 shows the amino acid sequence of WP_024556654.1: 16-277 of curli production assembly/transport protein CsgG [Cronobacter pulveris], which is 89% identical to SEQ ID NO: 3.
- SEQ ID NO: 75 shows the amino acid sequence of YP_005400916.1 : 16-277 of curli production assembly/transport protein CsgG [Rahnella aquatilis HX2], which is 84% identical to SEQ ID NO: 3.
- SEQ ID NO: 76 shows the amino acid sequence of KFC99297.1 : 20-278 of CsgG family curli production assembly/transport component [Kluyvera ascorbata ATCC 33433], which is 82% identical to SEQ ID NO: 3.
- SEQ ID NO: 77 shows the amino acid sequence of KFC86716.11 : 16-274 of CsgG family curli production assembly/transport component [Hafnia alvei ATCC 13337], which is 81% identical to SEQ ID NO: 3.
- SEQ ID NO: 78 shows the amino acid sequence of YP_007340845.11 : 16-270 of uncharacterised protein involved in formation of curli polymers [Enterobacteriaceae bacterium strain FGI 57], which is 76% identical to SEQ ID NO: 3.
- SEQ ID NO: 79 shows the amino acid sequence of WP_010861740.1: 17-274 of curli production assembly/transport protein CsgG [Plesiomonas shigelloides], which is 70% identical to SEQ ID NO: 3.
- SEQ ID NO: 80 shows the amino acid sequence of YP_205788.1 : 23-270 of curli production assembly/transport outer membrane lipoprotein component CsgG [Vibrio fischeri ESI 14], which is 60% identical to SEQ ID NO: 3.
- SEQ ID NO: 81 shows the amino acid sequence of WP_017023479.1: 23-270 of curli production assembly protein CsgG [Aliivibrio logei], which is 59% identical to SEQ ID NO: 3.
- SEQ ID NO: 82 shows the amino acid sequence of WP_007470398.1: 22-275 of Curli production assembly/transport component CsgG [Photobacterium sp. AK15], which is 57% identical to SEQ ID NO: 3.
- SEQ ID NO: 83 shows the amino acid sequence of WP_021231638.1: 17-277 of curli production assembly protein CsgG [Aeromonas veronii], which is 56% identical to SEQ ID NO: 3.
- SEQ ID NO: 84 shows the amino acid sequence of WP_033538267.1: 27-265 of curli production assembly/transport protein CsgG [Shewanella sp. ECSMB14101], which is 56% identical to SEQ ID NO: 3.
- SEQ ID NO: 85 shows the amino acid sequence of WP_003247972.1: 30-262 of curli production assembly protein CsgG [Pseudomonas putida], which is 54% identical to SEQ ID NO: 3.
- SEQ ID NO: 86 shows the amino acid sequence of YP_003557438.1 : 1-234 of curli production assembly/transport component CsgG [Shewanella violacea DSS12], which is 53% identical to SEQ ID NO: 3.
- SEQ ID NO: 87 shows the amino acid sequence of WP_027859066.1: 36-280 of curli production assembly/transport protein CsgG [Marinobacterium jannaschii], which is 53% identical to SEQ ID NO: 3.
- SEQ ID NO: 88 shows the amino acid sequence of CEJ70222.1: 29-262 of Curli production assembly/transport component CsgG [Chryseobacterium oranimense G311], which is 50% identical to SEQ ID NO: 3.
- SEQ ID NOs: 89 to 104 show the sequences in Table 1.
- SEQ ID NO: 1 (> P0AEA2; coding sequence for WT CsgG from E. coli K12)
- SEQ ID NO:2 (> P0AEA2 (1 : 277); WT prepro CsgG from E. coli K12)
- SEQ ID NO:3 (> P0AEA2 (16: 277); mature CsgG from E. coli K12)
- SEQ ID NO:4 (> P0AE98; coding sequence for WT CsgF from E. coli K12)
- SEQ ID NO:6 (>P0AE98 (20: 138); WT mature CsgF from E. coli K12)
- SEQ ID NO:16 (>P0AE98 (20 :42); mature peptide of CsgF 20:42+KD)
Landscapes
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- Health & Medical Sciences (AREA)
- Engineering & Computer Science (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Genetics & Genomics (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Health & Medical Sciences (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Biochemistry (AREA)
- General Engineering & Computer Science (AREA)
- Microbiology (AREA)
- Biotechnology (AREA)
- Molecular Biology (AREA)
- Medicinal Chemistry (AREA)
- Biomedical Technology (AREA)
- Physics & Mathematics (AREA)
- Analytical Chemistry (AREA)
- Biophysics (AREA)
- Immunology (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
The present invention relates to modified Dda helicases which can be used to control the movement of analytes such as polynucleotides. The modified Dda helicases are used in analyte detection and characterisation.
Description
NOVEL ENZYMES
TECHNICAL FIELD
The present invention relates to modified Dda helicases which can be used to control the movement of analytes such as polynucleotides. The modified Dda helicases are used in analyte detection and characterisation.
BACKGROUND
Two of the essential components of analyte, especially polymer, characterization using nanopore sensing are (1) the control of polymer movement through the pore and (2) the discrimination of the composing building blocks as the polymer is moved through the pore. During nanopore sensing, the narrowest part of the pore typically corresponds to the most discriminating part of the nanopore with respect to the change in measurement signal as a function of the analyte moving with respect to the nanopore. WO2015/055981, WO2015/166276, WO2016/055777, and PCT/EP2023/059821, incorporated by reference herein in their entirety, describe polynucleotide binding proteins, specifically Dda helicases, which can be used to control the movement of analytes with respect to a transmembrane protein pore such as the CsgG pores described herein.
SUMMARY OF THE INVENTION
The inventors have surprisingly identified specific Dda mutants (known herein as modified Dda helicases or modified helicases) which have an improved ability to control the movement of an analyte through a pore. When sequencing a polynucleotide using a pore, the system jointly estimates the number and identity of bases/nucleotides passing through the pore. Better control over variability in the speed of movement can reduce one of the sources of statistical noise and simplify the estimation task. Runs of consecutive short dwells of a polynucleotide in the pore may trigger a failure to call the underlying nucleotides/bases resulting in a deletion error. Unusually long dwells may lead to insertion errors. Ensuring that each nucleotide/base spends a sufficient time interval in the pore is helpful for resolving statistical uncertainty in the nucleotide/base identity from noisy signal levels. Further information can be extracted from dependence of dwell times on nucleotide/base identities, for example via interactions with the motor enzyme. Reducing the overall variability in dwell times can help to extract more precise information through this channel. During regions in which signal levels provide limited information about movement (e.g., long homopolymer regions) multi-nucleotide/base dwell times can be used to infer the number of bases traversing the pore. Reducing variability in dwell times can make these inferences more precise.
In some embodiments the modified helicases of the invention display increased speed when used in methods of controlling the movement of an analyte through a transmembrane pore
and in methods of characterising an analyte using a transmembrane pore. The speed at which an analyte passes through/relative to the pore may be increased by at least about 5%, at least about 10%, at least about 15%, at least about 20%, at least about 25%, at least about 30%, at least about 35%, at least about 40%, at least about 45%, at least about 50%, at least about 55%, at least about 60%, at least about 65%, at least about 70%, at least about 75%, at least about 80%, at least about 85%, at least about 90%, at least about 100%, at least about 150%, at least about 200%, at least about 300%, at least about 400%, at least about 500% or greater relative to the speed at which the analyte moves with respect to a pore when using a Dda helicase which does not comprise a mutation of the invention. The inventors have surprisingly found that these alterations in speed, such as increases in speed, have minimal or no effect on accuracy readings. This is particularly advantageous in a method of characterising an analyte wherein an analyte is contacted with the pore and a helicase of the invention such that the polynucleotide binding protein controls the movement of the target analyte through/relative to the pore.
In some embodiments the modified helicases of the invention display improved accuracy when used in methods of controlling the movement of an analyte through a transmembrane pore and in methods of characterising an analyte using a transmembrane pore. In the context of analyte characterisation (particularly polynucleotides), accuracy is interpreted to mean raw read simplex accuracy; that is a single pass of a single molecule through a transmembrane pore. Accuracy is a useful measure to track platform improvements of sequencing devices. Accuracy can also refer to consensus accuracy or to the accuracy in detecting something specific such as a mutation in a polynucleotide analyte for example. Additionally or alternatively, accuracy is interpreted to mean the percentage of bases above a certain confidence level, where the confidence level has been pre-calibrated. In some embodiments the modified helicases of the invention display improved accuracy with minimal to no changes in speed. In some embodiments accuracy is improved to give less than 10% error, less than 5% error, less than 4 % error, less than 3% error, less than 2% error, less than 1% error, less than 0.1% error. The modified helicases identified by the inventors typically comprise a combination of mutations, namely one or more modifications in the part of the modified helicase which interacts with a transmembrane pore. Accuracy may also by influenced by the speed which the polymer translocates the pore under enzyme control and the speed may be altered by altering the concentration of ATP provided to the enzyme. The inventors have surprisingly realised that the enzyme can exhibit changes in speed during successive polymer translocations within the same sequencing run under the same conditions which can give rise to a decrease in accuracy.
Accuracy may be influenced by a number of factors such as the nanopore shape and composition, the enzyme as well as the interaction between the enzyme and nanopore. It is also influenced by the speed at which the polymer translocates the pore under enzyme
control and the translocation speed may be increased or lowered by altering the concentration of ATP provided to the enzyme. The inventors have surprisingly realised that changes in speed occur during successive polymer translocations within the same sequencing run under the same sequencing conditions, which can give rise to a decrease in sequencing accuracy. The variation in sequencing speed for a number of polymers may be measured to obtain a normalised speed distribution and the inventors have surprisingly realised that some modified enzymes can give rise to a lower normalised speed distribution and therefore an increased sequencing accuracy.
The speed spread ratio of a modified helicase is the normalised speed distribution of the modified helicase divided by the normalised speed distribution of the control, unmodified helicase. In some embodiments the modified helicases of the invention display a decreased speed spread ratio when used in methods of controlling the movement of an analyte through a transmembrane pore and in methods of characterising an analyte using a transmembrane pore. The speed spread ratio may be decreased by at least about 21%, at least about 21.8%, at least about 22%, at least about 25%, at least about 30%, at least about 35%, at least about 40%, at least about 45%, at least about 50%, at least about at least about 55%, at least about 60%, at least about 65%, at least about 70%, at least about 75%, at least about 80%, at least about 85%, at least about 90% relative to the spread speed ratio of a Dda helicase which does not comprise a mutation of the invention. A decreased speed spread ratio indicates that the modified helicase of the invention has a narrower distribution of speeds than the control, unmodified helicase.
The invention provides a modified DNA dependent ATPase (Dda) helicase, wherein the helicase (a) is one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: F3, D4, 116, K24, K25, H26, H27, V28, P33, T42, F44, 146, A48, A60, P62, A65, K67, K68, 169, K72, K76, S79, 184, P104, K108, V117, Y120, R122, L124, 1132, P134, W135, T137, N144, K145, E154, A157, Y158, P161, K166, Q170, T174, V176, N180, A181, V190, Y197, V200, V201, R207, T213, A214, L215, M219, V220, 1225, K227, L229, V237, M238, K247, L248, 1251, R253, K254, F257, D262, V265, D282, 1289, Q295, Y304, S306, V309, V314, G316, L319, R321, T329, G331, D333, L349, F352, T362, N365, N367, G369, G370, P373, A380, Q383, S385, P391, S393, F395, M401, S402, V403, D404, R405, A406, 1413, H414, V418, E419, A421, Q422, V427, V429, G432, Y434, D435, and F437 and/or (b) comprises one or more of the following substitutions or one or more of the following substitutions at the position(s) corresponding to the following amino acid positions in Dda 1993:
- T2N, T2K, T2S, T2D, T2I, T2L, T2E, T2F, T2G, T2H, T2Q, T2R, T2A, T2M, T2P, or
E47D, E47G, E47H, E47N, E47P, E47Q, E47S, E47T, E47A, or E47R,
- E54I, E54V, E54K, E54Y, E54T, E54L, E54N, E54A, E54F, E54M, E54D, E54H, E54P, E54Q, E54R, or E54S,
- T55F, T55V, T55H, T55I, T55L, T55M, T55W, T55Y, T55A, T55P, T55Q, or T55R,
- S83E, S83D, S83A, S83P, S83G, or S83M,
- K86N, K86G, K86D, K86E, K86H, K86Q, K86R, K86S, K86T, K86M, or K86P,
- N88K, N88E, N88T, N88S, N88V, N88Q, N88A, N88L, N88D, N88G, N88M, N88I, or N88P,
- P89G, P89K, P89M, P89Q, P89R, or P89D,
- V90E, V90D, V90K, V90Q, V90N, V90S, V90A, V90I, V90L, V90M, V90P, V90R, V90F, V90Y, V90H, or V90G,
- T91E, T91D, T91R, T91K, T91Y, T91H, T91P, T91A, T91V, T91L, T91I, T91S, or T91M,
- Y92K, Y92E, Y92R, Y92P, Y92I, or Y92M,
- E93P, E93D, E93G, E93A, E93L, E93K, E93Q, E93S, E93T, E93H, E93N, or E93R,
- V96K, V96T, V96P, V96D, V96G, V96S, V96I, V96N, V96A, V96H, or V96M,
- F98M or F98I,
- K101R, K101T, K101S, K101V, K101E, K101I, K101G, K101L, K101M, K101P, K101Q, K101D, K101H, or K101N,
- V103K, V103N, V103L, V103I, V103T, V103R, V103Y, V103S, V103D, V103A, V103E, V103H, V103M, V103P, V103Q, or V103F,
- C114K or C114R,
- D151G, D151V, D151N, D151K, D151T, D151A, D151H, D151P, D151Q, D151R, or D151S,
G153E, G153S, G153D, G153T, G153N, G153K, G153A, G153Q, G153R, G153L,
G153H, or G153P,
- N155K, N155P, N155S, N155E, N155D, N155G, N155T, N155A, N155V, N155L, N155I, N155H, N155M, N155Q, or N155R,
- T156H, T156Q, T156R, or T156W,
- K177A or K177P,
- W195F, W195Y, W195P, W195K, W195E, W195I, W195L, W195D, W195S, W195G, W195T, W195V, W195Q, W195N, W195R, W195M, or W195H,
- I196P, I196L, I196D, I196A, I196E, I196K, I196Q, I196S, I196T, I196F, I196M, I196V, or I196Y,
- D198N, D198P, D198E, D198G, D198H, D198K, D198Q, D198R, D198S, or D198T,
- T210A, T210M, T210P, T210Q, T210I, T210L, T210V, or T210F,
- N221P, N221S, or N221G,
- K243R, K243E, K243D, K243A, K243S, K243M, K243N, K243P, K243T, or K243Y,
- E258G, E258K, E258N, E258P, E258D, E258S, E258A, E258H, E258Q, E258R, or E258T,
- F276T, F276S, F276L, F276P, F276E, F276D, F276N, or F276Q,
- I281E, I281V, I281K, I281A, I281D, I281H, I281N, I281P, I281Q, I281R, I281S, I281T, I281F, I281L, I281M, or I281Y,
- N292E, N292A, N292S, N292D, N292K, N292H, N292R, N292Q, or N292T,
- E301K, E301D, E301N, E301S, E301A, E301H, E301M, E301P, E301Q, E301R, or E301T,
- F308K, F308P, F308R, F308T, F308G, F308E, F308S, F308V, F308A, F308Y, F308N, F308M, F308D, F308H, F308Q, or F308W,
- Y335P, Y335K, Y335E, Y335F, Y335T, Y335Q, Y335S, Y335R, Y335D, Y335V, Y335N, Y335G, Y335H, Y335M, or Y335W,
- Y336E, Y336H, Y336K, Y336I, Y336G, Y336S, Y336T, Y336D, Y336V, Y336P, Y336N, Y336R, Y336Q, Y336A, Y336F, Y336M, or Y336W,
- R337E, R337K, R337T, R337Q, R337P, R337V, R337F, R337S, R337D, R337G, R337Y, R337A, R337H, or R337N,
Y350P,
L354K, L354E, L354D, L354V, L354Y, L354R, L354S, L354N, L354H, L354M, L354P,
L354T, L354F, or L354I,
- K358G or K358P,
- T359V, T359M, T359A, T359I, T359Y, T359F, T359E, T359K, T359N, T359D, T359P, T359Q, T359R, T359S, T359W, or T359G,
- K364N, K364Q, K364D, K364E, K364G, K364H, K364S, K364T, K364M, K364P, K364Y, K364I, K364L, or K364V,
- W366D, W366N, W366K, W366A, W366S, W366I, W366E, W366R, W366M, W366T, W366Q, W366Y, W366G, W366F, W366V, W366P, or W366H,
- K368A, K368E, K368S, K368G, K368D, K368F, K368M, K368T, K368V, K368Y, K368H, K368N, K368P, K368Q, or K368R, and
- S382R, S382N, S382K, S382E, S382Q, S382A, S382H, S382M, S382T, S382D, S382G, or S382P.
The invention also provides:
- a construct comprising a helicase of the invention and an additional polynucleotide binding moiety, wherein the helicase is attached to the polynucleotide binding moiety and the construct has the ability to control the movement of an analyte;
- a polynucleotide which comprises a sequence which encodes a helicase of the invention or a construct of the invention;
- a vector which comprises a polynucleotide of the invention operably linked to a promoter;
- a host cell comprising a vector of the invention;
- a method of making a helicase of the invention or a construct of the invention, which comprises expressing a polynucleotide of the invention, transfecting a cell with a vector of the invention or culturing a host cell of the invention;
- a method of controlling the movement of an analyte, comprising contacting the analyte with a helicase of the invention or a construct of the invention and thereby controlling the movement of the analyte;
a method of characterising a target analyte, comprising:
(a) contacting the target analyte with a transmembrane pore and a helicase of the invention or a construct of the invention such that the helicase or construct controls the movement of the target analyte through the pore; and
(b) taking one or more measurements as the target analyte moves with respect to the pore wherein the measurements are indicative of one or more characteristics of the target analyte and thereby characterising the target analyte;
- a method of forming a sensor for characterising a target analyte, comprising forming a complex between (a) a pore and (b) a helicase of the invention or a construct of the invention and thereby forming a sensor for characterising the target analyte;
- a sensor for characterising a target analyte, comprising a complex between (a) a pore and (b) a helicase of the invention or a construct of the invention;
- use of a helicase of the invention or a construct of the invention to control the movement of a target analyte through a pore;
- a kit for characterising a target analyte comprising
(a) a pore and a helicase of the invention or a construct of the invention; or
(b) a helicase of the invention or a construct of the invention and one or more loading moieties;
- an apparatus for characterising target analytes in a sample, comprising (a) a plurality of pores and (b) a plurality of helicases of the invention or a plurality of constructs of the invention;
- a method of producing a helicase of the invention, comprising:
(a) providing a helicase; and
(b) modifying the helicase to produce a helicase of the invention;
- a method of producing a construct of the invention, comprising attaching a helicase of the invention to an additional polynucleotide binding moiety and thereby producing the construct; a series of two or more helicases attached to a polynucleotide, wherein at least one of the two or more helicases is a helicase of the invention; and
a method of improving the movement of a target analyte with respect to a transmembrane pore when the movement is controlled by a DNA dependent ATPase (Dda) helicase, wherein the DNA dependent ATPase (Dda) helicase is modified according to the invention which improves the movement of the target analyte with respect to the transmembrane pore.
DETAILED DESCRIPTION
All publications, patents and patent applications cited herein, whether supra or infra, are hereby incorporated by reference in their entirety. All publications, patents, and patent applications mentioned in this specification are herein incorporated by reference to the same extent as if each individual publication, patent, or patent application was specifically and individually indicated to be incorporated by reference. To the extent publications and patents or patent applications incorporated by reference contradict the disclosure contained in the specification, the specification is intended to supersede and/or take precedence over any such contradictory material.
In all of the discussion herein, the standard one letter codes for amino acids are used. These are as follows: alanine (A), arginine (R), asparagine (N), aspartic acid (D), cysteine (C), glutamic acid (E), glutamine (Q), glycine (G), histidine (H), isoleucine (I), leucine (L), lysine (K), methionine (M), phenylalanine (F), proline (P), serine (S), threonine (T), tryptophan (W), tyrosine (Y) and valine (V). Standard substitution notation is also used, i.e. Q42R means that Q at position 42 is replaced with R.
In the paragraphs herein where different amino acids at a specific position are separated by the I symbol, the I symbol means "or". For instance, Q87R/K means Q87R or Q87K. In the paragraphs herein where different positions are separated by the I symbol, the I symbol means "and" such that Y51/N55 is Y51 and N55.
Definitions
Unless specifically defined herein, all terms used herein have the same meaning as they would to one skilled in the art of the present invention. Practitioners are particularly directed to Sambrook et al., Molecular Cloning: A Laboratory Manual, 4th ed., Cold Spring Harbor Press, Plainsview, New York (2012); and Ausubel et al., Current Protocols in Molecular Biology (Supplement 114), John Wiley & Sons, New York (2016), for definitions and terms of the art. The definitions provided herein should not be construed to have a scope less than understood by a person of ordinary skill in the art.
"About" as used herein when referring to a measurable value such as an amount, a temporal duration, and the like, is meant to encompass variations of ± 20 % or ± 10 %, more preferably ± 5 %, even more preferably ± 1 %, and still more preferably ± 0.1 %
from the specified value, as such variations are appropriate to perform the disclosed methods.
The definitions in WO2015/055981, WO2015/166276, and WO2016/055777, and PCT/EP2023/059821 are incorporated by reference herein in their entirety.
A "homologue" or "homologues" of a protein encompass peptides, oligopeptides, polypeptides, proteins and enzymes having amino acid substitutions, deletions and/or insertions relative to the unmodified or wild-type protein in question and having similar biological and functional activity as the unmodified protein from which they are derived. The term "amino acid identity" as used herein refers to the extent that sequences are identical on an amino acid-by-amino acid basis over a window of comparison. Thus, a "percentage of sequence identity" is calculated by comparing two optimally aligned sequences over the window of comparison, determining the number of positions at which the identical amino acid residue (e.g., Ala, Pro, Ser, Thr, Gly, Vai, Leu, He, Phe, Tyr, Trp, Lys, Arg, His, Asp, Glu, Asn, Gin, Cys and Met) occurs in both sequences to yield the number of matched positions, dividing the number of matched positions by the total number of positions in the window of comparison (i.e., the window size), and multiplying the result by 100 to yield the percentage of sequence identity.
Modified Pda helicases
The present invention provides a modified Dda helicase. The one or more specific modifications are discussed in more detail below. Modifications according to the invention include one or more substitutions as discussed below.
The invention provides a modified DNA dependent ATPase (Dda) helicase in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: F3, D4, 116, K24, K25, H26, H27, V28, P33, T42, F44, 146, A48, A60, P62, A65, K67, K68, 169, K72, K76, S79, 184, P104, K108, V117, Y120, R122, L124, 1132, P134, W135, T137, N144, K145, E154, A157, Y158, P161, K166, Q170, T174, V176,
N180, A181, V190, Y197, V200, V201, R207, T213, A214, L215, M219, V220, 1225, K227,
L229, V237, M238, K247, L248, 1251, R253, K254, F257, D262, V265, D282, 1289, Q295,
Y304, S306, V309, V314, G316, L319, R321, T329, G331, D333, L349, F352, T362, N365,
N367, G369, G370, P373, A380, Q383, S385, P391, S393, F395, M401, S402, V403, D404, R405, A406, 1413, H414, V418, E419, A421, Q422, V427, V429, G432, Y434, D435, and F437. The advantages of substitutions at these positions are explained in Table 8.
In all embodiments described herein, corresponding positions or positions corresponding to specific amino acid positions may be determined by standard techniques in the art. For example, the PILEUP and BLAST algorithms mentioned above can be used to align the
sequence of a Dda helicase with Dda 1993 (SEQ ID NO: 118) and hence to identify corresponding residues.
The modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising a substitution or modification at one or more of the 113 positions set out above (i.e., one or more of positions F3 to F437).
The modified Dda helicase of the invention may comprise a modification or substitution at any number and combination of the 113 positions corresponding to the amino acid positions set out above. The modified helicase of the invention may comprise a modification or substitution at 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22,
23, 24, 25, 26, 27 , 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45,
46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68,
69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91,
92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103, 104, 105, 106, 107, 108, 109, 110, 111, 112, or 113 of the positions corresponding to the amino acid positions set out above (i.e., from F3 to F437).
The modified Dda helicase is preferably one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: A60, P62, K68, K72, K76, N144, R207, V237, Y304, V309, G316, L319, G331, D333, L349, P373, S385, P391, M401, R405, A406, and H414. The modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising a modification or substitution at one or more of these 22 positions.
The modified Dda helicase of the invention may comprise a modification or substitution at any number and combination of the positions corresponding to the 22 amino acid positions set out above. The modified Dda helicase of the invention may comprise a modification or substitution at 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, or 22 of the positions corresponding to the amino acid positions set out above (i.e., from A60 to H414).
The modified Dda helicase is preferably one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: A60, P62, K68, K72, K76, N144, R207, V237, Y304, V309, G316, L319, G331, D333, L349, P373, S385, P391, M401, A406, and H414. The modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising a modification or substitution at one or more of these 21 positions.
The modified Dda helicase of the invention may comprise a modification or substitution at any number and combination of the positions corresponding to the 21 amino acid positions
set out above. The modified Dda helicase of the invention may comprise a modification or substitution at 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, or 21 of the positions corresponding to the amino acid positions set out above (i.e., from A60 to H414).
The modified Dda helicase is preferably one in which the position corresponding to amino acid position R405 in Dda 1993 is modified or substituted. The modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising a modification or substitution at R405.
The position corresponding to amino acid position F3 in Dda 1993 may be substituted with
K, Y, I, E, G, L, A, S, D, R, V, T, N, H, M, P, Q, or W. The position corresponding to amino acid position F3 in Dda 1993 is preferably substituted with K (e.g., F3K) or Y (e.g., F3Y).
The position corresponding to amino acid position D4 in Dda 1993 may be substituted with I, L, K, P, S, E, Y, T, F, M, V, A, G, H, N, or Q. The position corresponding to amino acid position D4 in Dda 1993 is preferably substituted with I (e.g., D4I) or L (e.g., D4L).
The position corresponding to amino acid position 116 in Dda 1993 may be substituted with G, A, E, L, K, D, N, S, F, M, T, V, or Y. The position corresponding to amino acid position 116 in Dda 1993 is preferably substituted with G (e.g., I16G).
The position corresponding to amino acid position K24 in Dda 1993 may be substituted with G, P, A, D, N, S, E, H, M, Q, R, or T. The position corresponding to amino acid position K24 in Dda 1993 is preferably substituted with G (e.g., K24G) or P (e.g., K24P).
The position corresponding to amino acid position K25 in Dda 1993 may be substituted with A, N, G, D, E, I, L, M, P, Q, R, S, T, V, or H. The position corresponding to amino acid position K25 in Dda 1993 is preferably substituted with A (e.g., K25A).
The position corresponding to amino acid position H26 in Dda 1993 may be substituted with
L, K, R, Q, S, M, A, T, N, P, G, E, I, V, F, Y, or D. The position corresponding to amino acid position H26 in Dda 1993 is preferably substituted with K (e.g., H26K) or L (e.g., H26L).
The position corresponding to amino acid position H27 in Dda 1993 may be substituted with F, I, V, L, Y, A, T, S, M, W, D, E, K, N, Q, or R. The position corresponding to amino acid position H27 in Dda 1993 is preferably substituted with F (e.g., H27F).
The position corresponding to amino acid position V28 in Dda 1993 may be substituted with I, F, A, L, M, T, or Y. The position corresponding to amino acid position V28 in Dda 1993 is preferably substituted with I (e.g., V28I).
The position corresponding to amino acid position P33 in Dda 1993 may be substituted with A, Y, F, G, E, I, K, L, M, Q, R, S, T, V, D, H, or W. The position corresponding to amino acid position P33 in Dda 1993 is preferably substituted with A (e.g., P33A) or Y (e.g., P33Y).
The position corresponding to amino acid position T42 in Dda 1993 may be substituted with I, L, A, F, M, V, Y, D, E, K, N, P, Q, R, or S. The position corresponding to amino acid position T42 in Dda 1993 is preferably substituted with I (e.g., T42I).
The position corresponding to amino acid position F44 in Dda 1993 may be substituted with K, A, R, Y, G, H, T, S, Q, L, N, E, M, V, D, P, I, or W. The position corresponding to amino acid position F44 in Dda 1993 is preferably substituted with K (e.g., F44K).
The position corresponding to amino acid position 146 in Dda 1993 may be substituted with V, A, F, L, M, T, or Y. The position corresponding to amino acid position 146 in Dda 1993 is preferably substituted with V (e.g., I46V).
The position corresponding to amino acid position A48 in Dda 1993 may be substituted with K, H, Y, R, E, N, Q, S, T, D, M, P, G, I, L, V, F, or W. The position corresponding to amino acid position A46 in Dda 1993 is preferably substituted with H (e.g., A48H), K (e.g., A48K) or Y (e.g., A48Y).
The position corresponding to amino acid position A60 in Dda 1993 may be substituted with T, D, E, I, K, L, M, N, P, Q, R, S, V, or G. The position corresponding to amino acid position A60 in Dda 1993 is preferably substituted with T (e.g., A60T).
The position corresponding to amino acid position P62 in Dda 1993 may be substituted with S, A, D, E, G, H, K, M, N, Q, R, or T. The position corresponding to amino acid position P62 in Dda 1993 is preferably substituted with S (e.g., P62S).
The position corresponding to amino acid position A65 in Dda 1993 may be substituted with V, Q, K, G, R, Y, L, I, F, T, M, H, E, P, S, D, or N. The position corresponding to amino acid position A65 in Dda 1993 is preferably substituted with Q (e.g., A65Q) or V (e.g., A65V).
The position corresponding to amino acid position K67 in Dda 1993 may be substituted with A, V, T, S, R, M, E, G, I, L, P, Q, D, H, N, F, or Y. The position corresponding to amino acid position K67 in Dda 1993 is preferably substituted with A (e.g., K67A) or V (e.g., K67V).
The position corresponding to amino acid position K68 in Dda 1993 may be substituted with A, N, E, G, I, L, M, P, Q, R, S, T, V, D, or H. The position corresponding to amino acid position K68in Dda 1993 is preferably substituted with A (e.g., K68A).
The position corresponding to amino acid position 169 in Dda 1993 may be substituted with E, V, R, Q, N, A, K, T, S, D, L, H, M, P, F, or Y. The position corresponding to amino acid position 169 in Dda 1993 is preferably substituted with E (e.g., I69E) or L (e.g., I69V).
The position corresponding to amino acid position K72 in Dda 1993 may be substituted with A, E, G, I, L, M, P, Q, R, S, T, V, D, H, or N. The position corresponding to amino acid position K72 in Dda 1993 is preferably substituted with A (e.g., K72A).
The position corresponding to amino acid position K76 in Dda 1993 may be substituted with M, Y, F, I, V, R, A, L, Q, S, T, W, D, E, H, N, or P. The position corresponding to amino acid position K76 in Dda 1993 is preferably substituted with M (e.g., K76M) or Y (e.g., K76Y).
The position corresponding to amino acid position S79 in Dda 1993 may be substituted with T, K, R, V, E, A, D, I, L, M, N, P, Q, G, or H. The position corresponding to amino acid position S79 in Dda 1993 is preferably substituted with T (e.g., S79T).
The position corresponding to amino acid position 184 in Dda 1993 may be substituted with L, V, F, A, M, T, or Y. The position corresponding to amino acid position 184 in Dda 1993 is preferably substituted with L (or e.g., I84L).
The position corresponding to amino acid position P104 in Dda 1993 may be substituted with Y, K, E, F, H, I, L, M, Q, V, W, A, D, S, or T. The position corresponding to amino acid position P104 in Dda 1993 is preferably substituted with Y (e.g., P104Y).
The position corresponding to amino acid position K108 in Dda 1993 may be substituted with Y, E, D, H, N, S, G, F, I, L, M, Q, V, W, A, P, R, or T. The position corresponding to amino acid position K108 in Dda 1993 is preferably substituted with D (e.g., K108D), E (e.g., K108E) or Y (e.g., K108Y).
The position corresponding to amino acid position V117 in Dda 1993 may be substituted with A, S, E, G, I, K, L, M, P, Q, R, T, F, or Y. The position corresponding to amino acid position V117 in Dda 1993 is preferably substituted with A (e.g., V117A).
The position corresponding to amino acid position Y120 in Dda 1993 may be substituted with V, L, I, M, F, A, T, S, H, Q, or W. The position corresponding to amino acid position Y120 in Dda 1993 is preferably substituted with V (e.g., Y120V).
The position corresponding to amino acid position R122 in Dda 1993 may be substituted with E, K, N, D, L, T, S, A, V, I, P, Q, H, or M. The position corresponding to amino acid position R122 in Dda 1993 is preferably substituted with E (e.g., R122E).
The position corresponding to amino acid position L124 in Dda 1993 may be substituted with T, A, D, E, I, K, M, N, P, Q, R, S, V, F, or Y. The position corresponding to amino acid position L124 in Dda 1993 is preferably substituted with T (e.g., L124T).
The position corresponding to amino acid position 1132 in Dda 1993 may be substituted with T, V, L, A, D, E, K, M, N, P, Q, R, S, F, or Y. The position corresponding to amino acid position 1132 in Dda 1993 is preferably substituted with T (e.g., I132T).
The position corresponding to amino acid position P134 in Dda 1993 may be substituted with D, E, K, S, L, A, R, I, N, V, T, G, H, or Q. The position corresponding to amino acid position P134 in Dda 1993 is preferably substituted with D (e.g., P134D).
The position corresponding to amino acid position W135 in Dda 1993 may be substituted with D, H, T, N, E, G, S, K, Y, Q, R, F, A, P, or M. The position corresponding to amino acid position W135 in Dda 1993 is preferably substituted with D (e.g., W135D) or H (e.g., W135H).
The position corresponding to amino acid position T137 in Dda 1993 may be substituted with K, R, I, V, A, D, E, H, M, N, P, Q, S, or L. The position corresponding to amino acid position T137 in Dda 1993 is preferably substituted with K (e.g., T137K)).
The position corresponding to amino acid position N144 in Dda 1993 may be substituted with S, P, K, T, R, E, A, D, I, L, Q, Y, H, V, M, or G. The position corresponding to amino acid position N144 in Dda 1993 is preferably substituted with S (e.g., N144S).
The position corresponding to amino acid position K145 in Dda 1993 may be substituted with D, A, H, R, G, N, E, P, Q, S, T, or M. The position corresponding to amino acid position K145 in Dda 1993 is preferably substituted with D (e.g., K145D).
The position corresponding to amino acid position E154 in Dda 1993 may be substituted with N, S, V, I, T, D, G, H, K, Q, R, A, or P. The position corresponding to amino acid position E154 in Dda 1993 is preferably substituted with N (e.g., K154N).
The position corresponding to amino acid position A157 in Dda 1993 may be substituted with E, D, S, K, P, N, H, Q, R, T, G, I, L, M, or V. The position corresponding to amino acid position A157 in Dda 1993 is preferably substituted with E (e.g., A157E).
The position corresponding to amino acid position Y158 in Dda 1993 may be substituted with Q, S, E, L, D, A, I, K, F, N, G, R, V, T, H, P, M, or W. The position corresponding to amino acid position Y158 in Dda 1993 is preferably substituted with E (e.g., Y158E), Q (e.g., Y158Q) or S (e.g., Y158S).
The position corresponding to amino acid position P161 in Dda 1993 may be substituted with E, D, K, S, N, R, Q, T, A, or H. The position corresponding to amino acid position P161 in Dda 1993 is preferably substituted with E (e.g., P161E).
The position corresponding to amino acid position K166 in Dda 1993 may be substituted with N, P, D, E, G, H, Q, R, S, T, A, or M. The position corresponding to amino acid position K166 in Dda 1993 is preferably substituted with N (e.g., K166N).
The position corresponding to amino acid position Q170 in Dda 1993 may be substituted with K, I, V, T, S, E, R, N, H, Y, F, A, D, M, or P. The position corresponding to amino acid position Q170 in Dda 1993 is preferably substituted with K (e.g., Q170K).
The position corresponding to amino acid position T174 in Dda 1993 may be substituted with S, A, D, E, G, H, K, M, N, P, Q, R, I, L, or V. The position corresponding to amino acid position T174 in Dda 1993 is preferably substituted with S (e.g., T174S).
The position corresponding to amino acid position V176 in Dda 1993 may be substituted with I, G, P, A, T, R, S, Q, K, L, E, F, M, or Y. The position corresponding to amino acid position V176 in Dda 1993 is preferably substituted with I (e.g., V176I).
The position corresponding to amino acid position N180 in Dda 1993 may be substituted with D, E, K, A, G, H, P, Q, S, T, or R. The position corresponding to amino acid position N180 in Dda 1993 is preferably substituted with D (e.g., N180D).
The position corresponding to amino acid position A181 in Dda 1993 may be substituted with G, N, L, S, R, T, D, K, Q, H, E, I, M, P, or V. The position corresponding to amino acid position A181 in Dda 1993 is preferably substituted with G (e.g., A180G) or N (e.g., A180N).
The position corresponding to amino acid position V190 in Dda 1993 may be substituted with I, A, F, L, M, T, or Y. The position corresponding to amino acid position V190 in Dda 1993 is preferably substituted with I (e.g., V190I).
The position corresponding to amino acid position Y197 in Dda 1993 may be substituted with K, P, R, S, E, G, D, T, L, N, I, V, F, A, H, M, Q, or W. The position corresponding to amino acid position Y197 in Dda 1993 is preferably substituted with K (e.g., Y197K).
The position corresponding to amino acid position V200 in Dda 1993 may be substituted with I, T, F, K, D, L, N, E, S, Y, P, A, or M. The position corresponding to amino acid position V200 in Dda 1993 is preferably substituted with I (e.g., V200I).
The position corresponding to amino acid position V201 in Dda 1993 may be substituted with F, Y, I, K, L, H, M, W, A, or T. The position corresponding to amino acid position V201 in Dda 1993 is preferably substituted with F (e.g., V201F).
The position corresponding to amino acid position R.207 in Dda 1993 may be substituted with I, H, K, V, F, Y, L, T, E, A, M, N, Q, S, or D. The position corresponding to amino acid position R.207 in Dda 1993 is preferably substituted with H (e.g., R207H) or I (e.g., R207I).
The position corresponding to amino acid position T213 in Dda 1993 may be substituted with D, N, E, K, S, G, A, Q, H, P, I, L, M, R, or V. The position corresponding to amino acid position T213 in Dda 1993 is preferably substituted with D (e.g., T213D).
The position corresponding to amino acid position A214 in Dda 1993 may be substituted with G, S, E, K, D, L, N, I, T, V, M, P, Q, R, or H. The position corresponding to amino acid position A214 in Dda 1993 is preferably substituted with G (e.g., A214G) or S (e.g., A214S).
The position corresponding to amino acid position L215 in Dda 1993 may be substituted with Y, I, E, F, D, H, M, Q, V, W, A, T, K, N, P, R, or S. The position corresponding to amino acid position L215 in Dda 1993 is preferably substituted with E (e.g., L215E), I (e.g., L215I) or Y (e.g., L215Y).
The position corresponding to amino acid position M219 in Dda 1993 may be substituted with L, I, V, A, F, Y, K, D, E, S, T, Q, R, or W. The position corresponding to amino acid position M219 in Dda 1993 is preferably substituted with L (e.g., M219L).
The position corresponding to amino acid position V220 in Dda 1993 may be substituted with K, E, N, A, D, R, S, T, Q, L, I, G, H, M, P, F, or Y. The position corresponding to amino acid position V220 in Dda 1993 is preferably substituted with K (e.g., V220K).
The position corresponding to amino acid position 1225 in Dda 1993 may be substituted with V, K, T, D, N, S, E, G, R, Q, P, A, H, L, Y, F, or M. The position corresponding to amino acid position 1225 in Dda 1993 is preferably substituted with V (e.g., I225V).
The position corresponding to amino acid position K227 in Dda 1993 may be substituted with G, A, D, N, S, E, H, M, P, Q, R, or T. The position corresponding to amino acid position K227 in Dda 1993 is preferably substituted with G (e.g., K227G).
The position corresponding to amino acid position L229 in Dda 1993 may be substituted with S, K, V, I, N, E, D, T, G, A, R, Q, H, M, P, F, or Y. The position corresponding to amino acid position L229 in Dda 1993 is preferably substituted with K (e.g., L229K) or S (e.g., L229S).
The position corresponding to amino acid position V237 in Dda 1993 may be substituted with M, I, A, L, F, K, Q, R, S, T, W, or Y. The position corresponding to amino acid position V237 in Dda 1993 is preferably substituted with M (e.g., V237M).
The position corresponding to amino acid position M238 in Dda 1993 may be substituted with I, L, V, F, A, T, Y, K, Q, R, S, or W. The position corresponding to amino acid position M238 in Dda 1993 is preferably substituted with I (e.g., M238I).
The position corresponding to amino acid position K247 in Dda 1993 may be substituted with N, E, R, D, G, H, Q, S, T, A, M, or P. The position corresponding to amino acid position K247 in Dda 1993 is preferably substituted with N (e.g., K247N).
The position corresponding to amino acid position L248 in Dda 1993 may be substituted with F, I, H, M, V, W, Y, A, or T. The position corresponding to amino acid position L248 in Dda 1993 is preferably substituted with F (e.g., L248F).
The position corresponding to amino acid position 1251 in Dda 1993 may be substituted with F, L, E, Y, A, T, H, M, V, or W. The position corresponding to amino acid position 1251 in Dda 1993 is preferably substituted with F (e.g., I251F).
The position corresponding to amino acid position R253 in Dda 1993 may be substituted with Q, A, D, E, H, K, M, N, P, S, T, or Y. The position corresponding to amino acid position R253 in Dda 1993 is preferably substituted with Q (e.g., R253Q).
The position corresponding to amino acid position K254 in Dda 1993 may be substituted with N, S, D, E, G, H, Q, R, T, A, M, or P. The position corresponding to amino acid position K254 in Dda 1993 is preferably substituted with N (e.g., K254N) or S (e.g., K254S).
The position corresponding to amino acid position F257 in Dda 1993 may be substituted with Y, H, I, L, M, Q, V, or W. The position corresponding to amino acid position F257 in Dda 1993 is preferably substituted with Y (e.g., F257Y).
The position corresponding to amino acid position D262 in Dda 1993 may be substituted with P, E, A, K, Q, S, T, G, H, or N. The position corresponding to amino acid position D262 in Dda 1993 is preferably substituted with P (e.g., D262P).
The position corresponding to amino acid position V265 in Dda 1993 may be substituted with E, P, A, K, S, G, D, N, T, H, Q, R, F, I, L, M, or Y. The position corresponding to amino acid position V265 in Dda 1993 is preferably substituted with E (e.g., V265E).
The position corresponding to amino acid position D282 in Dda 1993 may be substituted with P, A, E, K, Q, S, T, G, H, or N. The position corresponding to amino acid position D282 in Dda 1993 is preferably substituted with P (e.g., D282P).
The position corresponding to amino acid position 1289 in Dda 1993 may be substituted with L, V, M, K, E, G, A, F, T, or Y. The position corresponding to amino acid position 1289 in Dda 1993 is preferably substituted with L (e.g., I289L).
The position corresponding to amino acid position Q295 in Dda 1993 may be substituted with D, E, M, S, G, H, K, N, P, T, A, R, or Y. The position corresponding to amino acid position Q295 in Dda 1993 is preferably substituted with D (e.g., Q295D).
The position corresponding to amino acid position Y304 in Dda 1993 may be substituted with K, E, L, P, D, N, S, G, A, R, T, Q, H, V, I, M, F, or W. The position corresponding to amino acid position Y304 in Dda 1993 is preferably substituted with K (e.g., Y304K).
The position corresponding to amino acid position S306 in Dda 1993 may be substituted with E, Y, D, K, T, G, F, V, N, A, H, P, Q, R, or M. The position corresponding to amino acid position S306 in Dda 1993 is preferably substituted with E (e.g., S306E).
The position corresponding to amino acid position V309 in Dda 1993 may be substituted with F, K, I, H, L, M, W, Y, A, or T. The position corresponding to amino acid position V309 in Dda 1993 is preferably substituted with F (e.g., V309F).
The position corresponding to amino acid position V314 in Dda 1993 may be substituted with I, K, E, L, A, F, M, T, or Y. The position corresponding to amino acid position V314 in Dda 1993 is preferably substituted with I (e.g., V314I).
The position corresponding to amino acid position G316 in Dda 1993 may be substituted with E, K, D, V, I, N, L, A, H, P, Q, R, S, or T. The position corresponding to amino acid position G316 in Dda 1993 is preferably substituted with E (e.g., G316E).
The position corresponding to amino acid position L319 in Dda 1993 may be substituted with E, K, I, V, T, N, D, G, A, H, P, Q, R, S, F, M, or Y. The position corresponding to amino acid position L319 in Dda 1993 is preferably substituted with E (e.g., L319E).
The position corresponding to amino acid position R321 in Dda 1993 may be substituted with K, N, E, V, I, D, L, A, H, M, P, Q, S, or T. The position corresponding to amino acid position R321 in Dda 1993 is preferably substituted with K (e.g., R321K).
The position corresponding to amino acid position T329 in Dda 1993 may be substituted with P, S, G, K, A, E, D, I, V, L, N, R, Y, F, Q, or M. The position corresponding to amino acid position T329 in Dda 1993 is preferably substituted with P (e.g., T329P).
The position corresponding to amino acid position G331 in Dda 1993 may be substituted with D, E, K, L, S, I, N, T, V, H, P, Q, or A. The position corresponding to amino acid position G331 in Dda 1993 is preferably substituted with D (e.g., G331D).
The position corresponding to amino acid position D333 in Dda 1993 may be substituted with G, K, E, I, A, N, S, H, P, Q, or T. The position corresponding to amino acid position D333 in Dda 1993 is preferably substituted with G (e.g., D333G).
The position corresponding to amino acid position L349 in Dda 1993 may be substituted with E, N, K, Q, A, T, S, D, V, R, I, H, P, F, M, or Y. The position corresponding to amino acid position L349 in Dda 1993 is preferably substituted with E (e.g., L349E).
The position corresponding to amino acid position F352 in Dda 1993 may be substituted with Y, L, K, E, R, I, A, N, D, H, M, Q, V, or W. The position corresponding to amino acid position F352 in Dda 1993 is preferably substituted with Y (e.g., F352Y).
The position corresponding to amino acid position T362 in Dda 1993 may be substituted with A, I, V, L, E, K, M, D, R, Y, N, Q, S, F, G, or P. The position corresponding to amino acid position T362 in Dda 1993 is preferably substituted with A (e.g., T362A).
The position corresponding to amino acid position N365 in Dda 1993 may be substituted with K, E, A, S, R, D, H, M, P, Q, T, or G. The position corresponding to amino acid position N365 in Dda 1993 is preferably substituted with K (e.g., N365K).
The position corresponding to amino acid position N367 in Dda 1993 may be substituted with S, K, L, E, A, T, I, V, Q, R, Y, F, M, D, G, H, or P. The position corresponding to amino acid position N367 in Dda 1993 is preferably substituted with S (e.g., N367S).
The position corresponding to amino acid position G369 in Dda 1993 may be substituted with E, K, D, N, S, T, R, L, I, A, Y, H, P, or Q. The position corresponding to amino acid position G369 in Dda 1993 is preferably substituted with E (e.g., G369E).
The position corresponding to amino acid position G370 in Dda 1993 may be substituted with K, V, E, I, S, A, N, D, H, M, P, Q, R, or T. The position corresponding to amino acid position G370 in Dda 1993 is preferably substituted with K (e.g., G370K).
The position corresponding to amino acid position P373 in Dda 1993 may be substituted with H, L, N, K, M, Y, F, I, V, S, E, T, A, R, D, G, Q, or W. The position corresponding to
amino acid position P373 in Dda 1993 is preferably substituted with H (e.g., P373) or L (e.g., P373L).
The position corresponding to amino acid position A380 in Dda 1993 may be substituted with V, L, T, I, K, M, F, Y, E, G, P, Q, R, or S. The position corresponding to amino acid position A380 in Dda 1993 is preferably substituted with V (e.g., A380V).
The position corresponding to amino acid position Q383 in Dda 1993 may be substituted with L, T, M, Y, K, F, I, V, R, A, E, S, N, D, G, H, or P. The position corresponding to amino acid position Q383 in Dda 1993 is preferably substituted with L (e.g., Q383L).
The position corresponding to amino acid position S385 in Dda 1993 may be substituted with I, T, A, Y, K, H, R, F, L, M, V, D, E, G, N, P, or Q. The position corresponding to amino acid position S385 in Dda 1993 is preferably substituted with I (e.g., S385I).
The position corresponding to amino acid position P391 in Dda 1993 may be substituted with G, Y, F, H, A, V, D, N, S, E, K, Q, or T. The position corresponding to amino acid position P391 in Dda 1993 is preferably substituted with G (e.g., P391G).
The position corresponding to amino acid position S393 in Dda 1993 may be substituted with I, V, L, T, E, D, M, F, A, Y, G, H, K, N, P, Q, or R. The position corresponding to amino acid position S393 in Dda 1993 is preferably substituted with I (e.g., S393I).
The position corresponding to amino acid position F395 in Dda 1993 may be substituted with I, V, Y, L, T, A, M, H, or W. The position corresponding to amino acid position F395 in Dda 1993 is preferably substituted with I (e.g., F395I).
The position corresponding to amino acid position M401 in Dda 1993 may be substituted with L, S, V, I, G, D, E, R, Q, A, W, N, T, F, Y, K, H, or P. The position corresponding to amino acid position M401 in Dda 1993 is preferably substituted with L (e.g., M401L), S (e.g., M401S) or V (e.g., M401V).
The position corresponding to amino acid position S402 in Dda 1993 may be substituted with T, V, R, I, K, L, E, H, Q, M, A, D, N, P, or G. The position corresponding to amino acid position S402 in Dda 1993 is preferably substituted with T (e.g., S402T).
The position corresponding to amino acid position V403 in Dda 1993 may be substituted with F, Y, I, H, L, M, W, A, or T. The position corresponding to amino acid position V403 in Dda 1993 is preferably substituted with F (e.g., V403F).
The position corresponding to amino acid position D404 in Dda 1993 may be substituted with N, K, R, E, G, H, Q, S, T, or P. The position corresponding to amino acid position D404 in Dda 1993 is preferably substituted with N (e.g., D404N).
The position corresponding to amino acid position R405 in Dda 1993 may be substituted with N, K, D, E, G, H, Q, S, T, A, M, or P. The position corresponding to amino acid position R405 in Dda 1993 is preferably substituted with K (e.g., R405K) or N (e.g., R405N).
The position corresponding to amino acid position A406 in Dda 1993 may be substituted with T, V, I, D, E, K, L, M, N, P, Q, R, S, G, F, or Y. The position corresponding to amino acid position A406 in Dda 1993 is preferably substituted with T (e.g., A406T) or V (e.g., A406V).
The position corresponding to amino acid position 1413 in Dda 1993 may be substituted with E, L, K, S, D, G, N, P, Y, F, T, R, A, H, Q, M, or V. The position corresponding to amino acid position 1413 in Dda 1993 is preferably substituted with E (e.g., I413E).
The position corresponding to amino acid position H414 in Dda 1993 may be substituted with Y, D, F, Q, K, N, S, E, G, P, R, T, L, I, A, M, V, or W. The position corresponding to amino acid position H414 in Dda 1993 is preferably substituted with D (e.g., H414D) or Y (e.g., H414Y).
The position corresponding to amino acid position V418 in Dda 1993 may be substituted with K, E, P, R, D, T, A, S, N, L, I, Y, G, H, M, Q, or F. The position corresponding to amino acid position V418 in Dda 1993 is preferably substituted with K (e.g., V418K).
The position corresponding to amino acid position E419 in Dda 1993 may be substituted with Q, D, A, H, K, M, N, P, R, S, T, or Y. The position corresponding to amino acid position E419 in Dda 1993 is preferably substituted with E (e.g., E419Q).
The position corresponding to amino acid position A421 in Dda 1993 may be substituted with Q, L, R, N, S, K, D, E, H, M, P, T, Y, G, I, V, or F. The position corresponding to amino acid position A421 in Dda 1993 is preferably substituted with L (e.g., A421L) or Q (e.g., A421Q).
The position corresponding to amino acid position Q422 in Dda 1993 may be substituted with K, R, L, N, A, D, E, H, M, P, S, T, or Y. The position corresponding to amino acid position Q422 in Dda 1993 is preferably substituted with K (e.g., Q422K).
The position corresponding to amino acid position V427 in Dda 1993 may be substituted with T, E, D, A, I, K, L, M, N, P, Q, R, S, F, or Y. The position corresponding to amino acid position V427 in Dda 1993 is preferably substituted with T (e.g., V427T).
The position corresponding to amino acid position V429 in Dda 1993 may be substituted with I, L, A, F, M, T, or Y. The position corresponding to amino acid position V429 in Dda 1993 is preferably substituted with I (e.g., V429I).
The position corresponding to amino acid position G432 in Dda 1993 may be substituted with P, A, S, T, D, E, K, Q, or N. The position corresponding to amino acid position G432 in Dda 1993 is preferably substituted with P (e.g., G432P).
The position corresponding to amino acid position Y434 in Dda 1993 may be substituted with K, I, V, R, E, T, N, D, L, S, Q, H, G, A, M, P, F, or W. The position corresponding to amino acid position Y434 in Dda 1993 is preferably substituted with K (e.g., Y434K).
The position corresponding to amino acid position D435 in Dda 1993 may be substituted with E, Q, A, H, K, N, P, R, S, T, or G. The position corresponding to amino acid position D435 in Dda 1993 is preferably substituted with E (e.g., D435E).
The position corresponding to amino acid position F437 in Dda 1993 may be substituted with Y, I, H, L, M, Q, V, or W. The position corresponding to amino acid position F437 in Dda 1993 is preferably substituted with Y (e.g., F437Y).
The modified Dda helicase of the invention may further comprise one or more of the 45 or 31 substitutions set out below or one or more of the 45 or 31 substitutions at the position(s) corresponding to the 45 or 31 amino acid positions in Dda 1993 set out below.
The invention also provides a modified DNA dependent ATPase (Dda) helicase comprising one or more of the following substitutions or one or more of the following substitutions at the position(s) corresponding to the following amino acid positions in Dda 1993:
- T2N, T2K, T2S, T2D, T2I, T2L, T2E, T2F, T2G, T2H, T2Q, T2R, T2A, T2M, T2P, or T2V, preferably T2N,
- E47D, E47G, E47H, E47N, E47P, E47Q, E47S, E47T, E47A, or E47R, preferably E47D,
- E54I, E54V, E54K, E54Y, E54T, E54L, E54N, E54A, E54F, E54M, E54D, E54H, E54P, E54Q, E54R, or E54S, preferably E54I,
- T55F, T55V, T55H, T55I, T55L, T55M, T55W, T55Y, T55A, T55P, T55Q, or T55R, preferably T55F or T55V,
- S83E, S83D, S83A, S83P, S83G, or S83M, preferably S83E,
- K86N, K86G, K86D, K86E, K86H, K86Q, K86R, K86S, K86T, K86M, or K86P, preferably K86N,
- N88K, N88E, N88T, N88S, N88V, N88Q, N88A, N88L, N88D, N88G, N88M, N88I, or N88P, preferably N88K,
- P89G, P89K, P89M, P89Q, P89R, or P89D,
- V90E, V90D, V90K, V90Q, V90N, V90S, V90A, V90I, V90L, V90M, V90P, V90R, V90F, V90Y, V90H, or V90G, preferably V90D or V90E,
- T91E, T91D, T91R, T91K, T91Y, T91H, T91P, T91A, T91V, T91L, T91I, T91S, or T91M, preferably T91E,
- Y92K, Y92E, Y92R, Y92P, Y92I, or Y92M,
- E93P, E93D, E93G, E93A, E93L, E93K, E93Q, E93S, E93T, E93H, E93N, or E93R, preferably E93P,
- V96K, V96T, V96P, V96D, V96G, V96S, V96I, V96N, V96A, V96H, or V96M,
- F98M or F98I,
- K101R, K101T, K101S, K101V, K101E, K101I, K101G, K101L, K101M, K101P, K101Q, K101D, K101H, or K101N, preferably K101R,
- V103K, V103N, V103L, V103I, V103T, V103R, V103Y, V103S, V103D, V103A, V103E, V103H, V103M, V103P, V103Q, or V103F, preferably V103K,
- C114K or C114R,
- D151G, D151V, D151N, D151K, D151T, D151A, D151H, D151P, D151Q, D151R, or D151S,
- G153E, G153S, G153D, G153T, G153N, G153K, G153A, G153Q, G153R, G153L, G153H, or G153P, preferably G153E,
- N155K, N155P, N155S, N155E, N155D, N155G, N155T, N155A, N155V, N155L, N155I, N155H, N155M, N155Q, or N155R, preferably N155K,
- T156H, T156Q, T156R, or T156W,
- K177A or K177P,
- W195F, W195Y, W195P, W195K, W195E, W195I, W195L, W195D, W195S, W195G, W195T, W195V, W195Q, W195N, W195R, W195M, or W195H, preferably W195Y, W195P, W195K, W195E, W195I, W195G, W195T, W195Q, W195N, W195R, W195M, or W195H, more preferably W195F or W195Y,
- I196P, I196L, I196D, I196A, I196E, I196K, I196Q, I196S, I196T, I196F, I196M, I196V, or I196Y, preferably I196L or I196P,
- D198N, D198P, D198E, D198G, D198H, D198K, D198Q, D198R, D198S, or D198T, preferably D198N, D198P, D198E, D198G, D198H, D198K, D198Q, D198R, or D198T, more preferably D198N,
- T210A, T210M, T210P, T210Q, T210I, T210L, T210V, or T210F,
- N221P, N221S, or N221G,
- K243R, K243E, K243D, K243A, K243S, K243M, K243N, K243P, K243T, or K243Y, preferably K243R or K243E,
- E258G, E258K, E258N, E258P, E258D, E258S, E258A, E258H, E258Q, E258R, or E258T, preferably E258G, E258K, E258N, E258P, E258D, E258H, E258Q, E258R, or E258T, more preferably E258G,
- F276T, F276S, F276L, F276P, F276E, F276D, F276N, or F276Q, preferably F276T,
- I281E, I281V, I281K, I281A, I281D, I281H, I281N, I281P, I281Q, I281R, I281S, I281T, I281F, I281L, I281M, or I281Y, preferably I281E,
- N292E, N292A, N292S, N292D, N292K, N292H, N292R, N292Q, or N292T, preferably N292E,
- E301K, E301D, E301N, E301S, E301A, E301H, E301M, E301P, E301Q, E301R, or E301T, preferably E301K,
- F308K, F308P, F308R, F308T, F308G, F308E, F308S, F308V, F308A, F308Y, F308N, F308M, F308D, F308H, F308Q, or F308W, preferably F308K or F308P,
- Y335P, Y335K, Y335E, Y335F, Y335T, Y335Q, Y335S, Y335R, Y335D, Y335V, Y335N, Y335G, Y335H, Y335M, or Y335W, preferably Y335P,
- Y336E, Y336H, Y336K, Y336I, Y336G, Y336S, Y336T, Y336D, Y336V, Y336P, Y336N, Y336R, Y336Q, Y336A, Y336F, Y336M, or Y336W, preferably Y336E,
R337E, R337K, R337T, R337Q, R337P, R337V, R337F, R337S, R337D, R337G,
R337Y, R337A, R337H, or R337N,
Y350P,
- L354K, L354E, L354D, L354V, L354Y, L354R, L354S, L354N, L354H, L354M, L354P, L354T, L354F, or L354I, preferably L354K,
- K358G or K358P,
- T359V, T359M, T359A, T359I, T359Y, T359F, T359E, T359K, T359N, T359D, T359P, T359Q, T359R, T359S, T359W, or T359G, preferably T359A, T359V, T359I or T359M, more preferably T359V,
- K364N, K364Q, K364D, K364E, K364G, K364H, K364S, K364T, K364M, K364P, K364Y, K364I, K364L, or K364V, preferably K364N or K364Q,
- W366D, W366N, W366K, W366A, W366S, W366I, W366E, W366R, W366M, W366T, W366Q, W366Y, W366G, W366F, W366V, W366P, or W366H, preferably W366D or W366N,
- K368A, K368E, K368S, K368G, K368D, K368F, K368M, K368T, K368V, K368Y, K368H, K368N, K368P, K368Q, or K368R, and
- S382R, S382N, S382K, S382E, S382Q, S382A, S382H, S382M, S382T, S382D, S382G, or S382P, preferably S382N or S382R.
These are the 45 substitutions discussed above. The advantages of these substitutions are explained in Table 8.
The modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising one or more of the 45 substitutions set out above.
The modified Dda helicase of the invention may comprise any number and combination of the substitutions set out above. The modified Dda helicase of the invention may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44 or 45 of the substitutions above. The modified Dda helicase of the invention may further be one in which one or more of the positions corresponding to the 113, 22, or 21 amino acid positions in Dda 1993 listed above are modified or substituted as described above.
The invention also provides a modified DNA dependent ATPase (Dda) helicase comprising one or more of the following substitutions or one or more of the following substitutions at the position(s) corresponding to the following amino acid positions in Dda 1993:
T2N, T2K, T2S, T2D, T2I, T2L, T2E, T2F, T2G, T2H, T2Q, T2R, T2A, T2M, T2P, or
T2V, preferably T2N,
- E47D, E47G, E47H, E47N, E47P, E47Q, E47S, E47T, E47A, or E47R, preferably E47D,
- E54I, E54V, E54K, E54Y, E54T, E54L, E54N, E54A, E54F, E54M, E54D, E54H, E54P, E54Q, E54R, or E54S, preferably E54I,
- T55F, T55V, T55H, T55I, T55L, T55M, T55W, T55Y, T55A, T55P, T55Q, or T55R, preferably T55F or T55V,
- S83E, S83D, S83A, S83P, S83G, or S83M, preferably S83E,
- K86N, K86G, K86D, K86E, K86H, K86Q, K86R, K86S, K86T, K86M, or K86P, preferably K86N,
- N88K, N88E, N88T, N88S, N88V, N88Q, N88A, N88L, N88D, N88G, N88M, N88I, or N88P, preferably N88K,
- V90E, V90D, V90K, V90Q, V90N, V90S, V90A, V90I, V90L, V90M, V90P, V90R, V90F, V90Y, V90H, or V90G, preferably V90D or V90E,
- T91E, T91D, T91R, T91K, T91Y, T91H, T91P, T91A, T91V, T91L, T91I, T91S, or T91M, preferably T91E,
- E93P, E93D, E93G, E93A, E93L, E93K, E93Q, E93S, E93T, E93H, E93N, or E93R, preferably E93P,
- K101R, K101T, K101S, K101V, K101E, K101I, K101G, K101L, K101M, K101P, K101Q, K101D, K101H, or K101N, preferably K101R,
- V103K, V103N, V103L, V103I, V103T, V103R, V103Y, V103S, V103D, V103A, V103E, V103H, V103M, V103P, V103Q, or V103F, preferably V103K,
- G153E, G153S, G153D, G153T, G153N, G153K, G153A, G153Q, G153R, G153L, G153H, or G153P, preferably G153E,
- N155K, N155P, N155S, N155E, N155D, N155G, N155T, N155A, N155V, N155L, N155I, N155H, N155M, N155Q, or N155R, preferably N155K,
- W195F, W195Y, W195P, W195K, W195E, W195I, W195L, W195D, W195S, W195G, W195T, W195V, W195Q, W195N, W195R, W195M, or W195H, preferably W195Y, W195P, W195K, W195E, W195I, W195G, W195T, W195Q, W195N, W195R, W195M, or W195H, more preferably W195F or W195Y,
- I196P, I196L, I196D, I196A, I196E, I196K, I196Q, I196S, I196T, I196F, I196M, I196V, or I196Y, preferably I196L or I196P,
- D198N, D198P, D198E, D198G, D198H, D198K, D198Q, D198R, D198S, or D198T, preferably D198N, D198P, D198E, D198G, D198H, D198K, D198Q, D198R, or D198T, more preferably D198N,
- K243R, K243E, K243D, K243A, K243S, K243M, K243N, K243P, K243T, or K243Y, preferably K243R or K243E,
- E258G, E258K, E258N, E258P, E258D, E258S, E258A, E258H, E258Q, E258R, or E258T, preferably E258G, E258K, E258N, E258P, E258D, E258H, E258Q, E258R, or E258T, more preferably E258G,
- F276T, F276S, F276L, F276P, F276E, F276D, F276N, or F276Q, preferably F276T,
- I281E, I281V, I281K, I281A, I281D, I281H, I281N, I281P, I281Q, I281R, I281S, I281T, I281F, I281L, I281M, or I281Y, preferably I281E,
- N292E, N292A, N292S, N292D, N292K, N292H, N292R, N292Q, or N292T, preferably N292E,
- E301K, E301D, E301N, E301S, E301A, E301H, E301M, E301P, E301Q, E301R, or E301T, preferably E301K,
- F308K, F308P, F308R, F308T, F308G, F308E, F308S, F308V, F308A, F308Y, F308N, F308M, F308D, F308H, F308Q, or F308W, preferably F308K or F308P,
- Y335P, Y335K, Y335E, Y335F, Y335T, Y335Q, Y335S, Y335R, Y335D, Y335V, Y335N, Y335G, Y335H, Y335M, or Y335W, preferably Y335P,
- Y336E, Y336H, Y336K, Y336I, Y336G, Y336S, Y336T, Y336D, Y336V, Y336P, Y336N, Y336R, Y336Q, Y336A, Y336F, Y336M, or Y336W, preferably Y336E,
- L354K, L354E, L354D, L354V, L354Y, L354R, L354S, L354N, L354H, L354M, L354P, L354T, L354F, or L354I, preferably L354K,
- T359V, T359M, T359A, T359I, T359Y, T359F, T359E, T359K, T359N, T359D, T359P, T359Q, T359R, T359S, T359W, or T359G, preferably T359A, T359V, T359I or T359M, more preferably T359V,
- K364N, K364Q, K364D, K364E, K364G, K364H, K364S, K364T, K364M, K364P, K364Y, K364I, K364L, or K364V, preferably K364N or K364Q,
- W366D, W366N, W366K, W366A, W366S, W366I, W366E, W366R, W366M, W366T, W366Q, W366Y, W366G, W366F, W366V, W366P, or W366H, preferably W366D or W366N, and
- S382R, S382N, S382K, S382E, S382Q, S382A, S382H, S382M, S382T, S382D, S382G, or S382P, preferably S382N or S382R.
These are the 31 substitutions discussed above. The advantages of these substitutions are explained in Table 8.
The modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising one or more of the 31 substitutions set out above.
The modified Dda helicase of the invention may comprise any number and combination of the substitutions set out above. The modified Dda helicase of the invention may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, or 31 of the substitutions above. The modified Dda helicase of the invention may further be one in which one or more of the positions corresponding to the 113, 22, or 21 amino acid positions in Dda 1993 listed above are modified or substituted as described above.
The modified Dda helicase of the invention preferably comprises one or more of the following substitutions or one or more of the following substitutions at the position(s) corresponding to the following amino acid positions in Dda 1993:
- I196P, I196L, I196D, I196A, I196E, I196K, I196Q, I196S, I196T, I196F, I196M, I196V, or I196Y, preferably I196L or I196P,
- Y336E, Y336H, Y336K, Y336I, Y336G, Y336S, Y336T, Y336D, Y336V, Y336P, Y336N, Y336R, Y336Q, Y336A, Y336F, Y336M, or Y336W, preferably Y336E,
- T359V, T359M, T359A, T359I, T359Y, T359F, T359E, T359K, T359N, T359D, T359P, T359Q, T359R, T359S, T359W, or T359G, preferably T359A, T359V, T359I or T359M, more preferably T359V,
K364N, K364Q, K364D, K364E, K364G, K364H, K364S, K364T, K364M, K364P,
K364Y, K364I, K364L, or K364V, preferably K364N or K364Q, and
S382R, S382N, S382K, S382E, S382Q, S382A, S382H, S382M, S382T, S382D, S382G, or S382P, preferably S382N or S382R.
The modified Dda helicase of the invention preferably comprises a variant of SEQ ID NO: 118 comprising one or more of the five substitutions set out above.
The modified Dda helicase of the invention may comprise any number and combination of these five substitutions. The modified Dda helicase of the invention may comprise 1, 2, 3, 4, or 5 of the substitutions. The modified Dda helicase of the invention may further be one in which one or more of the positions corresponding to the 113, 22 or 21 amino acid positions in Dda 1993 listed above are modified or substituted as described above.
The helicase is preferably a variant of SEQ ID NO: 118 which comprises substitutions at:
- W195/K358/Q422, such as W195F/K358I/Q422K, W195R/K358M/Q422N, W195K/K358P/Q422D, W195Y/K358I/Q422K, W195K/K358I/Q422N, W195F/K358M/Q422K, W195Y/K358G/Q422A, W195I/K358I/Q422K, W195F/K358I/Q422D or W195I/K358G/Q422A;
- W195/K364/K368, such as W195K/K364D/K368Q, W195Y/K364Q/K368A, W195I/K364Q/K368A, W195R/K364D/K368T, W195Y/K364I/K368T, W195F/K364Q/K368Q, W195R/K364Q/K368A, W195I/K364P/K368T, W195F/K364Q/K368A or W195R/K364I/K368Y;
- W195/K364/Q422, such as W195S/K364I/Q422A, W195Y/K364I/Q422A, W195F/K364I/Q422D, W195Y/K364P/Q422N, W195S/K364P/Q422A, W195R/K364I/Q422A, W195Y/K364Q/Q422K, W195F/K364Q/Q422K, W195K/K364P/Q422N or W195R/K364D/Q422D;
- W195/K368/Q422, such as W195Y/K368A/Q422K, W195K/K368Q/Q422N, W195F/K368A/Q422K, W195S/K368A/Q422K, W195F/K368Q/Q422D, W195Y/K368T/Q422K, W195V/K368T/Q422A, W195K/K368V/Q422K, W195V/K368T/Q422D or W195S/K368V/Q422A;
- W195/Y350/K368, such as W195Y/Y350E/K368A, W195Y/Y350D/K368T, W195K/Y350I/K368P, W195R/Y350S/K368Q, W195I/Y350D/K368I, W195S/Y350K/K368T, W195R/Y350F/K368V, W195F/Y350E/K368A W195F/Y350I/K368T or W195K/Y350W/K368Q;
K67/M401, such as K67M/M401L, K67A/M401V, K67Q/M401S or K67N/M401W;
- G153/M401, such as G153E/M401S, G153Q/M401G, G153E/M401V or G153L/M401W;
- G153/V403, such as G153E/V403F, G153L/V403L, G153Q/V403M or G153N/V403H;
- M401/V403, such as M401S/V403I, M401L/V403H, M401V/V403F or M401R/V403A;
- K67/G153, such as K67M/G153L, K67A/G153E, K67L/G153H or K67Q/G153A;
- K67/K368, such as K67M/K368Q, K67N/K368E, K67Q/K368R or K67A/K368A;
- K368/M401, such as K368A/M401V, K368E/M401S, K368Q/M401G or K368R/M401W;
- K368/V403, such as K368N/V403A, K368P/V403H, K368A/V403F or K368F/V403L;
- A60/K368, such as A60K/K368N, A60T/K368A, A60L/K368E or A60D/K368R;
- K67/V403, such as K67M/V403H, K67L/V403I, K67N/V403A or K67A/V403F;
- G153/W195/M401, such as G153E/W195Q/M401G, G153N/W195K/M401L, G153Q/W195F/M401S, G153N/W195Q/M401G, G153E/W195Q/M401L, G153L/W195Q/M401G, G153E/W195F/M401S, G153Q/W195Q/M401L, G153L/W195K/M401G or G153N/W195R/M401W;
- K25/P134, such as K25N/P134T, K25D/P134G, K25A/P134D or K25V/P134E;
- P134/W135/C136, such as P134E/W135N/C136M, P134K/W135A/C136F, P134G/W135A/C136T, P134E/W135A/C136F, P134K/W135N/C136F, P134K/W135D/C136M, P134E/W135D/C136V, P134K/W135N/C136T, P134T/W135A/C136M or P134G/W135F/C136F;
- W195/V220, such as W195K/V220A, W195R/V220P, W195F/V220K or W195I/V220Y;
- W195/Q422, such as W195F/Q422K, W195I/Q422N, W195P/Q422A or W195K/Q422D; or
- V220/Q422, such as V220P/Q422A, V220I/Q422D, V220A/Q422T or V220K/Q422K.
Preferred combinations in SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 include the combinations of amino acids which correspond to the combinations in SEQ ID NO: 118 listed above.
The invention also provides a modified DNA dependent ATPase (Dda) helicase, wherein the helicase comprises one or more modifications or substitutions at one or more positions in one or more of the (i) tower domain, (ii) 1A domain and (iii) 2A domain and wherein the helicase displays one or more of (a) an increased speed, (b) an improved accuracy and (c) a decreased speed spread ratio when used in methods of controlling the movement of an analyte through a transmembrane pore and/or in methods of characterising an analyte using a transmembrane pore. The modified Dda helicase of the invention may comprise any number and combination of the modified or substitutions set out above. The modified Dda helicase of the invention may comprise one or more modifications or substitutions at one or more positions in (i), (ii), (iii), (i) and (ii), (i) and (iii), (ii) and (iii) or (i), (ii) and (iii). The modified Dda helicase of the invention may display (a), (b), (c), (a) and (b), (a) and (c), (b) and (c) or (a), (b) and (c). The one or more positions in the 2A domain may be proximal to motif V/Va. The one or more positions in the 1A domain may be in motif la. Motif V and motif la are helicase motifs associated with energy transduction from ATP hydrolysis to DNA translocation. The domains (i)-(iii) are defined in more detail below. The modified Dda helicase of the invention may comprise one or more modifications or substitutions at any of the positions described above or below and/or one or more of the substitutions described above or below.
Any of the modified Dda helicases of the invention or variants of SEQ ID NO: 118 discussed above may further comprise the substitution MIG.
The modified Dda helicase of the invention or variant of SEQ ID NO: 118 may further comprise any of the modifications, mutations or substitutions discussed below.
The Dda helicase that is modified in accordance with the invention may be any of SEQ ID NOs: 118 to 133. SEQ ID NO: 118 is Dda 1993. The modified Dda helicase preferably comprises a variant of any of SEQ ID NOs: 118 to 133. The variant may have any % of the sequence homologies/identities to any of SEQ ID NOs: 118 to 113 set out below.
Table 1 below summarises the preferred Dda helicases which may be modified in accordance with the invention.
One or more of positions 55, 114, 156, 177, 210, 221, 350 and 358 of Dda 1993
The modified helicase of the invention may further be one in which one or more of the positions corresponding to amino acid positions 55, 114, 156, 177, 210, 221, 350 and 358 in Dda 1993 are modified or substituted. The positions corresponding to positions 55, 114, 156, 177, 210, 221, 350 and 358 in Dda 1993 are discussed in more detail below. Positions 55, 114, 156 and 177 are in the 1A domain of Dda 1993. Positions 210 and 221 are in the 2A domain of Dda 1993. Positions 350 and 358 are in the tower domain of Dda 1993. The
modified Dda helicase of the invention may further comprise a modification or substitution at any number and combination of the positions corresponding to amino acid positions (a) 55, (b) 114, (c) 156, (d) 177, (e) 210, (f) 221, (g) 350 and (h) 358, including at (a); (b);
(c); (d); (e); (f); (g); (h); (a) and (b); (a) and (c); (a) and (d); (a) and (e); (a) and (f); (a) and (g); (a) and (h); (b) and (c); (b) and (d); (b) and (e); (b) and (f); (b) and (g); (b) and (h); (c) and (d); (c) and (e); (c) and (f); (c) and (g); (c) and (h); (d) and (e); (d) and (f);
(d) and (g); (d) and (h); (e) and (f); (e) and (g); (e) and (h); (f) and (g); (f) and (h); (g) and (h); (a), (b) and (c); (a), (b) and (d); (a), (b) and (e); (a), (b) and (f); (a), (b) and (g); (a), (b) and (h); (a), (c) and (d); (a), (c) and (e); (a), (c) and (f); (a), (c) and (g);
(a), (c) and (h); (a), (d) and (e); (a), (d) and (f); (a), (d) and (g); (a), (d) and (h); (a), (e) and (f); (a), (e) and (g); (a), (e) and (h); (a), (f) and (g); (a), (f) and (h); (a), (g) and (h);
(b), (c) and (d); (b), (c) and (e); (b), (c) and (f); (b), (c) and (g); (b), (c) and (h); (b), (d) and (e); (b), (d) and (f); (b), (d) and (g); (b), (d) and (h); (b), (e) and (f); (b), (e) and
(g); (b), (e) and (h); (b), (f) and (g); (b), (f) and (h); (b), (g) and (h); (c), (d) and (e);
(c), (d) and (f); (c), (d) and (g); (c), (d) and (h); (c), (e) and (f); (c), (e) and (g); (c), (e) and (h); (c), (f) and (g); (c), (f) and (h); (c), (g) and (h); (d), (e) and (f); (d), (e) and (g);
(d), (e) and (h); (d), (f) and (g); (d), (f) and (h); (d), (g) and (h); (e), (f) and (g); (e), (f) and (h); (e), (g) and (h); (f), (g) and (h); (a), (b), (c) and (d); (a), (b), (c) and (e); (a),
(b), (c and (f); (a), (b), (c) and (g); (a), (b), (c) and (h); (a), (b), (d) and (e); (a), (b), (d) and (f); (a), (b), (d) and (g); (a), (b), (d) and (h); (a), (b), (e) and (f); (a), (b), (e) and
(g); (a), (b), (e) and (h); (a), (b), (f) and (g); (a), (b), (f) and (h); (a), (b), (g) and (h);
(a), (c), (d) and (e); (a), (c), (d) and (f); (a), (c), (d) and (g); (a), (c), (d) and (h); (a),
(c), (e) and (f); (a), (c), (e) and (g); (a), (c), (e) and (h); (a), (c), (f) and (g); (a), (c), (f) and (h); (a), (c), (g) and (h); (a), (d), (e) and (f); (a), (d), (e) and (g); (a), (d), (e) and
(h); (a), (d), (f) and (g); (a), (d), (f) and (h); (a), (d), (g) and (h); (a), (e), (f) and (g);
(a), (e), (f) and (h); (a), (e), (g) and (h); (a), (f), (g) and (h); (b), (c), (d) and (e); (b),
(c), (d) and (f); (b), (c), (d) and (g); (b), (c), (d) and (h); (b), (c), (e) and (f); (b), (c), (e) and (g); (b), (c), (e) and (h); (b), (c), (f) and (g); (b), (c), (f) and (h); (b), (c), (g) and
(h); (b), (d), (e) and (f); (b), (d), (e) and (g); (b), (d), (e) and (h); (b), (d), (f) and (g);
(b), (d), (f) and (h); (b), (d), (g) and (h); (b), (e), (f) and (g); (b), (e), (f) and (h); (b),
(e), (g) and (h); (b), (f), (g) and (h); (c), (d), (e) and (f); (c), (d), (e) and (g); (c), (d), (e) and (h); (c), (d), (f) and (g); (c), (d), (f) and (h); (c), (d), (g) and (h); (c), (e), (f) and (g);
(c), (e), (f) and (h); (c), (e), (g) and (h); (c), (f), (g) and (h); (d), (e), (f) and (g); (d), (e),
(f) and (h); (d), (e), (g) and (h); (d), (f), (g) and (h); (e), (f), (g) and (h); (a), (b), (c), (d) and (e); (a), (b), (c), (d) and (f); (a), (b), (c), (d) and (g); (a), (b), (c), (d) and (h); (a),
(b), (c), (e) and (f); (a), (b), (c), (e) and (g); (a), (b), (c), (e) and (h); (a), (b), (c), (f) and
(g); (a), (b), (c), (f) and (h); (a), (b), (c), (g) and (h); (a), (b), (d), (e) and (f); (a), (b),
(d), (e) and (g); (a), (b), (d), (e) and (h); (a), (b), (d), (f) and (g); (a), (b), (d), (f) and
(h); (a), (b), (d), (g) and (h); (a), (b), (e), (f) and (g); (a), (b), (e), (f) and (h); (a), (b),
(e), (g) and (h); (a), (b), (f), (g) and (h); (a), (c), (d), (e) and (f); (a), (c), (d), (e) and
(g); (a), (c), (d), (e) and (h); (a), (c), (d), (f) and (g); (a), (c), (d), (f) and (h); (a), (c),
(d), (g) and (h); (a), (c), (e), (f) and (g); (a), (c), (e), (f) and (h); (a), (c), (e), (g) and
(h); (a), (c), (f), (g) and (h); (a), (d), (e), (f) and (g); (a), (d), (e), (f) and (h); (a), (d),
(e), (g) and (h); (a), (d), (f), (g) and (h); (a), (e), (f), (g) and (h); (b), (c), (d), (e) and
(f); (b), (c), (d), (e) and (g); (b), (c), (d), (e) and (h); (b), (c), (d), (f) and (g); (b), (c),
(d), (f) and (h); (b), (c), (d), (g) and (h); (b), (c), (e), (f) and (g); (b), (c), (e), (f) and (h);
(b), (c), (e), (g) and (h); (b), (c), (f), (g) and (h); (b), (d), (e), (f) and (g); (b), (d), (e), (f) and (h); (b), (d), (e), (g) and (h); (b), (d), (f), (g) and (h); (b), (e), (f), (g) and (h); (c),
(d), (e), (f) and (g); (c), (d), (e), (f) and (h); (c), (d), (e), (g) and (h); (c), (d), (f), (g) and
(h); (c), (e), (f), (g) and (h); (d), (e), (f), (g) and (h); (a), (b), (c), (d), (e) and (f); (a),
(b), (c), (d), (e) and (g); (a), (b), (c), (d), (e) and (h); (a), (b), (c), (d), (f) and (g); (a),
(b), (c), (d), (f) and (h); (a), (b), (c), (d), (g) and (h); (a), (b), (c), (e), (f) and (g); (a),
(b), (c), (e), (f) and (h); (a), (b), (c), (e), (g) and (h); (a), (b), (c), (f), (g) and (h); (a),
(b), (d), (e), (f) and (g); (a), (b), (d), (e), (f) and (h); (a), (b), (d), (e), (g) and (h); (a),
(b), (d), (f), (g) and (h); (a), (b), (e), (f), (g) and (h); (a), (c), (d), (e), (f) and (g); (a),
(c), (d), (e), (f) and (h); (a), (c), (d), (e), (g) and (h); (a), (c), (d), (f), (g) and (h); (a),
(c), (e), (f), (g) and (h); (a), (d), (e), (f), (g) and (h); (b), (c), (d), (e), (f) and (g); (b),
(c), (d), (e), (f) and (h); (b), (c), (d), (e), (g) and (h); (b), (c), (d), (f), (g) and (h); (b),
(c), (e), (f), (g) and (h); (b), (d), (e), (f), (g) and (h); (c), (d), (e), (f), (g) and (h); (a),
(b), (c), (d), (e), (f) and (g); (a), (b), (c), (d), (e), (f) and (h); (a), (b), (c), (d), (e), (g) and (h); (a), (b), (c), (d), (f), (g) and (h); (a), (b), (c), (e), (f), (g) and (h); (a), (b), (d),
(e), (f), (g) and (h); (a), (c), (d), (e), (f), (g) and (h); (b), (c), (d), (e), (f), (g) and (h); or (a), (b), (c), (d), (e), (f), (g) and (h).
The modified Dda helicase of the invention may further be one in which one or more of the positions corresponding to amino acid positions 114, 177, 350 and 358 in Dda 1993 are modified or substituted. The corresponding to positions 114, 177, 350 and 358 in Dda 1993 are discussed in more detail below. Positions 114 and 177 are in the 1A domain of Dda 1993. Positions Y350 and K358 are in the tower domain of Dda 1993. The modified Dda helicase of the invention may further comprise a modification or substitution at any number and combination of the positions corresponding to amino acid positions (a) 114, (b) 177, (c) 350 and (d) 358 in Dda 1993, including at (a); (b); (c); (d); (a) and (b); (a) and (c); (a) and (d); (b) and (c); (b) and (d); (c) and (d); (a), (b) and (c); (a), (b) and (d); (a), (c) and
(d); (b), (c) and (d); or (a), (b), (c) and (d).
The position corresponding to amino acid position 55 in Dda 1993 is preferably substituted with D, E, K, N or S. The position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with K (T55K). These substitutions increase the speed and increase the accuracy when used to characterise a polynucleotide analyte (Example 5 of
PCT/EP2023/059821). These substitutions also decrease the normalised speed distribution when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
The position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with A, V, I, L, M, F, Y, W, G, P, S, T, N or Q. The position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with A, G, I, L, M, P, S, T or V. The position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with G, L, S or T. These substitutions decrease the speed when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with A, I, M, P, or V. These substitutions increase the speed when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with G (C11G). This substitution decreases the speed and increases the accuracy when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with I or P. These substitutions increase the speed and decrease the accuracy when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 114 in Dda 1993 is preferably substituted with G, I or P. These substitutions decrease the normalised speed distribution when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 114 in Dda 1993 is most preferably substituted with I (Cl 141).
The position corresponding to amino acid position 156 in Dda 1993 is preferably substituted with A, E, F, G, I, L, M, P, S, V, Y, D, K or N. The position corresponding to amino acid position 156 in Dda 1993 is preferably substituted with F (T156F). This substitution increases the speed and increases the accuracy when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821). This substitution also decreases the normalised speed distribution when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
The position corresponding to amino acid position 177 in Dda 1993 is preferably substituted with D, E, F, G, H, I, L, M, N, Q, R, S, T, V, W or Y. The position corresponding to amino acid position 177 in Dda 1993 is preferably substituted with F, G, S, V, W or Y. These substitutions decrease the speed when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 177 in Dda 1993 is preferably substituted with D, E, G, H, I, L, M, N, Q, R, or T. These substitutions increase the speed when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 177 in Dda 1993 is preferably substituted with F, H, I, L, M, N or W. These substitutions
decrease the accuracy and the normalised speed distribution when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). They have different effects on the speed (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 177 in Dda 1993 is preferably substituted with N (K177N). This substitution decreases the accuracy and increases the normalised speed distribution when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 177 in Dda 1993 is most preferably substituted with M (K177M).
The position corresponding to amino acid position 210 in Dda 1993 is preferably substituted with D, E, K, S, N, R, H or Y. The position corresponding to amino acid position 210 in Dda 1993 is preferably substituted with R (T210R), H (T210H) or K (T210K). The position corresponding to amino acid position 210 in Dda 1993 is preferably substituted with K (T210K). This substitution increases the speed and increases the accuracy when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821). This substitution also decreases the normalised speed distribution when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
The position corresponding to amino acid position 221 in Dda 1993 is preferably substituted with D, K, E, Q, R, A, H, L, T or Y. The position corresponding to amino acid position 221 in Dda 1993 is preferably substituted with D (N221D) or E (N221E). The position corresponding to amino acid position 221 in Dda 1993 is preferably substituted with E (N221E). This substitution increases the speed and increases the accuracy when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821). This substitution also decreases the normalised speed distribution when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
The position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with D, E, A, V, I, L, M, F, W, R, H, K, L, S, T, N or Q. The position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with I, F, W or S. The position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with I or S (Y350I or Y350S). The position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with I (Y350I). This substitution increases the speed and decreases the accuracy and normalised speed distribution when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821). The position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with I or S (Y350I or Y350S). These substitutions have the effects shown in Example 4 of PCT/EP2023/059821 when used with a pore complex described therein.
The position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with A, D, E, G, K, L, N, Q, R, T, V, H or M. The position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with D (Y350D) or E (Y350E). The position corresponding to amino acid position 350 in Dda 1993 is preferably substituted with E (Y350E). This substitution increases the speed and increases the accuracy when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821). This substitution also decreases the normalised speed distribution when used to characterise a polynucleotide analyte (Example 5 of PCT/EP2023/059821).
The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with D, E, A, V, I, L, M, F, Y, W, R, H, L, S, T, N or Q.
The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with E, I, L or M. These substitutions decrease the speed when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with I or M. These substitutions decrease the speed and increase the accuracy when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted M (K358M). This substitution decreases the speed and increase the accuracy and the normalised speed distribution when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with I (K358I). This substitution decreases the speed and normalised speed distribution and increases the accuracy when used to characterise a polynucleotide analyte (Example 2 of PCT/EP2023/059821). Example 2 uses a CsgG pore without a CsgF peptide.
The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with A, E, F, I, M or S. These substitutions increase the accuracy when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821). The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with A, E, F, I or M. These substitutions decrease the speed when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821). The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted S (K358S). This substitution increases the speed when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821). The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with A, E, I, M or S. These substitutions decrease the normalised speed distribution when used to characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821). The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with (K358F). These substitutions increase the normalised speed distribution when used to
characterise a polynucleotide analyte (Example 3 of PCT/EP2023/059821). Example 3 uses a CsgG pore complex containing a CsgF peptide.
The position corresponding to amino acid position 358 in Dda 1993 is preferably substituted with I, L or Q. These substitutions decrease the speed and increase the accuracy and normalised speed distribution when used to characterise a polynucleotide analyte (Example 4 of PCT/EP2023/059821).
The position corresponding to amino acid position 358 in Dda 1993 is most preferably substituted with I (K358I).
Table 2 shows the amino acids in SEQ ID NOs: 119 to 133 which correspond to positions 40, 55, 114, 156, 177, 210, 221, 350 and 358 in SEQ ID NO: 118.
The modified helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising one or more of (a)-(h) as follows:
(a)
T55D, T55E, T55K, T55N or T55S, or
T55K.
(b)
C114A, C114V, C114I, C114L, C114M, C114F, C114Y, C114W, C114G, C114P, C114S,
C114T, C114N or C114Q,
C114A, C114G, C114I, C114L, C114M, C114P, C114S, C114T or C114V,
C114G, C114L, C114S or C114T,
C114A, C114I, C114M, C114P, or C114V
C11G,
C114I or C114P
C114G, C114I or C114P, or
C114I;
(c)
T156A, T156E, T156F, T156G, T156I, T156L, T156M, T156P, T156S, T156V, T156Y, T156D,
T156K or T156N, or
T156F;
(d)
K177D, K177E, K177F, K177G, K177H, K177I, K177L, K177M, K177N, K177Q, K177R,
K177S, K177T, K177V, K177W or K177Y,
K177F, K177G, K177S, K177V, K177W or K177Y,
K177D, K177E, K177G, K177H, K177I, K177L, K177M, K177N, K177Q, K177R, or K177T,
K177F, K177H, K177I, K177L, K177M, K177N or K177W,
K177N, or
K177M;
(e)
T210D, T210E, T210K, T210S, T210N, T210R, T210H or T210Y,
T210R, T210H or T210Y, or
T210K;
(f)
N221D, N221K, N221E, N221Q, N221R, N221A, N221H, N221L, N221T or N221Y,
N221D or N221E, or
N221E;
(g)
Y350D, Y350E, Y350A, Y350V, Y350I, Y350L, Y350M, Y350F, Y350W, Y350R, Y350H,
Y350K, Y350L, Y350S, Y350T, Y350N or Y350Q,
Y350I or Y350S,
Y350I,
Y350S,
Y350I, Y350F, Y350W or Y350S,
Y350A, Y350D, Y350E, Y350G, Y350K, Y350L, Y350N, Y350Q, Y350R, Y350T, Y350V, Y350H or Y350M,
Y350D or Y350E, or
Y350E; and
(h)
K358D, K358E, K358A, K358V, K358I, K358L, K358M, K358F, K358Y, K358W, K358R,
K358H, K358L, K358S, K358T, K358N or K358Q,
K358E, K358I, K358L or K358M,
K358I or K358M,
K358M,
K358I,
K358A, K358E, K358F, K358I, K358M or K358S,
K358A, K358E, K358F, K358I or K358M,
K358S,
K358A, K358E, K358I, K358M or K358S,
K358F,
K358I, K358L or K358Q, or
K358I.
The variant may include any combination and permutation of (a)-(h) as set out above.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising one or more of (a), (b), (c) and (d) as follows:
(a)
C114A, C114V, C114I, C114L, C114M, C114F, C114Y, C114W, C114G, C114P, C114S,
C114T, C114N or C114Q,
C114A, C114G, C114I, C114L, C114M, C114P, C114S, C114T or C114V,
C114G, C114L, C114S or C114T,
C114A, C114I, C114M, C114P, or C114V
C11G,
C114I or C114P
C114G, C114I or C114P, or
C114I;
(b)
K177D, K177E, K177F, K177G, K177H, K177I, K177L, K177M, K177N, K177Q, K177R,
K177S, K177T, K177V, K177W or K177Y,
K177F, K177G, K177S, K177V, K177W or K177Y,
K177D, K177E, K177G, K177H, K177I, K177L, K177M, K177N, K177Q, K177R, or K177T,
K177F, K177H, K177I, K177L, K177M, K177N or K177W,
K177N, or
K177M;
(c)
Y350D, Y350E, Y350A, Y350V, Y350I, Y350L, Y350M, Y350F, Y350W, Y350R, Y350H,
Y350K, Y350L, Y350S, Y350T, Y350N or Y350Q,
Y350I or Y350S,
Y350I, or
Y350S; and
(d)
K358D, K358E, K358A, K358V, K358I, K358L, K358M, K358F, K358Y, K358W, K358R,
K358H, K358L, K358S, K358T, K358N or K358Q,
K358E, K358I, K358L or K358M,
K358I or K358M,
K358M,
K358I,
K358A, K358E, K358F, K358I, K358M or K358S,
K358A, K358E, K358F, K358I or K358M,
K358S,
K358A, K358E, K358I, K358M or K358S,
K358F,
K358I, K358L or K358Q, or
K358I.
The variant may include (a); (b); (c); (d); (a) and (b); (a) and (c); (a) and (d); (b) and
(c); (b) and (d); (c) and (d); (a), (b) and (c); (a), (b) and (d); (a), (c) and (d); (b), (c) and
(d); or (a), (b), (c) and (d).
A preferred variant of SEQ ID NO: 118 comprises: C114I; K177M; Y350I; K358I; C114I and K177M; C114I and Y350I; C114I and K358I; K177M and Y350I; K177M and K358I; Y350I and K358I; C114I, K177M and Y350I; C114I, K177M and K358I; C114I, Y350I and K358I; K177M, Y350I and K358I; or C114I, K177M, Y350I and K358I.
The helicase preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein one or more of the positions corresponding to amino acid positions 55, 114, 156, 177, 210, 221, 350 and 358 in Dda 1993 are modified or substituted as defined above (including specific substitutions). Various combinations and permutations of one or more of positions 55, 114, 156, 177, 210, 221, 350 and 358 in Dda 1993 are defined above with reference to (a)-(h).
The helicase preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein one or more of the positions corresponding to amino acid positions 114, 177, 350 and 358 in Dda 1993 are modified or substituted as defined above (including specific substitutions). Various combinations and permutations of one or more of positions 114, 177, 350 and 358 in Dda 1993 are defined above with reference to (a)-(d).
The helicase preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein one or more of the positions corresponding to amino acid positions 114, 177 and 358 in Dda 1993 are modified or substituted as defined above (including specific substitutions).
Position 40 in Dda 1993
Any of the modified helicases of the invention may further comprise a modification or substitution at the position corresponding to amino acid position 40 in Dda 1993. Position 40 or the corresponding position may be substituted with as A, V, I, L, M, F, Y or W.
Positions which correspond to position T40 in Dda 1993 are shown in Table 2 above. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which, in addition to the modifications/substitution set out above, further comprises a substitution at T40, such as T40A, T40V, T40I, T40L, T40M, T40F, T40Y or T40W. The substitution is preferably T40Y.
The modified Dda helicase of the invention may further be one in which the position corresponding to amino acid position 40 in Dda 1993 is modified or substituted. Position T40 is in the tower domain of Dda 1993. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at T40, such as T40A, T40V, T40I, T40L, T40M, T40F, T40Y or T40W. The substitution is preferably T40Y. The modified Dda helicase of the invention may further comprise a modification or substitution at one or more of the positions corresponding to amino acid positions (a) 55, (b) 114, (c) 156, (d) 177, (e) 210, (f) 221, (g) 350 and (h) 358, including any of the combinations and permutations of (a)-(h) set out above. The modified Dda helicase of the invention may further comprise a modification or substitution at one or more of the positions corresponding to amino acid positions (a) 114, (b) 177, (c) 350 and (d) 358 in Dda 1993, including at (a); (b); (c); (d); (a) and (b); (a) and (c); (a) and (d); (b) and (c); (b) and
(d); (c) and (d); (a), (b) and (c); (a), (b) and (d); (a), (c) and (d); (b), (c) and (d); or (a),
(b), (c) and (d). The helicase preferably comprises a variant of SEQ ID NO: 119, 120, 121,
122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein the position corresponding to amino acid position 40 in Dda 1993 is modified or substituted as defined above (including specific substitutions).
The modified helicases of the invention may further comprise any of the modifications, substitutions, combinations of modifications or combination of substitutions discussed below.
Other helicases of the invention
The use of Dda helicases in analyte characterisation are described in WO2015/055981, WO2015/166276 and WO2016/055777 (all incorporated by reference).
The modified helicases of the invention provide more consistent movement of the target analyte with respect to, such as through, the transmembrane pore leading to improved accuracy. The helicases preferably provide more consistent movement from one k-mer to another or from k-mer to k-mer as the target analyte, such as polynucleotide, moves with respect to, such as through, the pore. The helicases allow the target analyte, such as target polynucleotide, to move with respect to, such as through, the transmembrane pore more smoothly. The helicases preferably provide more regular or less irregular movement of the
target analyte, such as target polynucleotide, with respect to, such as through, the transmembrane pore.
The modification(s) typically increase accuracy by at least 0.1%, at least 0.5%, at least 1%, at least 5%, at least 10%, at least 20%, at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80% or at least 90% compared to a helicase without the modification.
The ability of a helicase to control the movement of a polynucleotide can be determined as described in the Examples.
The modified helicase has the ability to control the movement of a polynucleotide. The ability of a helicase to control the movement of a polynucleotide can be assayed using any method known in the art. For instance, the helicase may be contacted with a polynucleotide and the position of the polynucleotide may be determined using standard methods. The ability of a modified helicase to control the movement of a polynucleotide is typically assayed in a nanopore system, such as the ones described below and, in particular, as described in the Examples.
A modified helicase of the invention may be isolated, substantially isolated, purified or substantially purified. A helicase is isolated or purified if it is completely free of any other components, such as lipids, polynucleotides, pore monomers or other proteins. A helicase is substantially isolated if it is mixed with carriers or diluents which will not interfere with its intended use. For instance, a helicase is substantially isolated or substantially purified if it is present in a form that comprises less than 10%, less than 5%, less than 2% or less than 1% of other components, such as lipids, polynucleotides, pore monomers or other proteins.
Dda helicases
Any Dda helicase may be modified in accordance with the invention. Preferred Dda helicases are discussed below and described in WO2015/055981, WO2015/166276 and WO2016/055777 (all incorporated by reference).
Dda helicases typically comprises the following five domains: 1A (RecA-like motor) domain, 2A (RecA-like motor) domain, tower domain, pin domain and hook domain (Xiaoping He et al., 2012, Structure; 20: 1189-1200). The domains may be identified using protein modelling, x-ray diffraction measurement of the protein in a crystalline state (Rupp B (2009). Biomolecular Crystallography: Principles, Practice and Application to Structural Biology. New York: Garland Science.), nuclear magnetic resonance (NMR) spectroscopy of the protein in solution (Mark Rance; Cavanagh, John; Wayne J. Fairbrother; Arthur W. Hunt III; Skelton, NNicholas J. (2007). Protein NMR spectroscopy: principles and practice (2nd
ed.). Boston: Academic Press.) or cryo-electron microscopy of the protein in a frozen- hydrated state (van Heel M, Gowen B, Matadeen R, Orlova EV, Finn R, Pape T, Cohen D, Stark H, Schmidt R, Schatz M, Patwardhan A (2000). "Single-particle electron cryomicroscopy: towards atomic resolution.". Q Rev Biophys. 33: 307-69). Structural information of proteins determined by above mentioned methods are publicly available from the protein bank (PDB) database.
In addition to the modifications or substitutions set out above, the modified helicase of the invention preferably further comprises any of the following additional modifications, substitutions, combinations of modifications or combination of substitutions.
Modifications of the invention
The helicase of the invention may be one in which at least one amino acid which interacts with a transmembrane pore is further substituted. Any number of amino acids may substituted, such as 1 or more, 2 or more, 3 or more, 4 or more, 5 or more or 6 or more amino acids. 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more amino acids may be substituted. The amino acids which interact with a transmembrane pore can be identified using protein modelling as discussed above.
Base and/or sugar interactions
The helicase of the invention is preferably one in which at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in single stranded DNA (ssDNA) is further substituted with an amino acid which comprises a larger side chain (R group). Any number of amino acids may substituted, such as 1 or more, 2 or more, 3 or more, 4 or more, 5 or more or 6 or more amino acids. Each amino acid may interact with the base, the sugar or the base and the sugar. The amino acids which interact with the sugar and/or base of one or more nucleotides in single stranded DNA can be identified using protein modelling as discussed above.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 wherein the at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in ssDNA is at least one of H82, N88, P89, F98, D121, V150, P152, F240, F276, S287, H396 and Y415. These numbers correspond to the relevant positions in SEQ ID NO: 118 and may need to be altered in the case of variants where one or more amino acids have been inserted or deleted compared with SEQ ID NO: 118. A skilled person can determine the corresponding positions in a variant as discussed above. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 wherein the at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in ssDNA is F98 and one or more H82, N88, P89, D121, V150, P152, F240, F276, S287, H396 and Y415,
such as F98/H82, F98/N88, F98/P89, F98/D121, F98/V150, F98/ P152, F98/F240, F98/F276, F98/S287 or F98/H396.
The helicase of the invention is preferably a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein the at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in ssDNA is at least one of the amino acids which correspond to H82, N88, P89, F98, D121, V150, P152, F240, F276, S287, H396 and Y415in SEQ ID NO: 118. The helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 wherein the at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in ssDNA is the amino acid which corresponds to F98 in SEQ ID NO: 118 and one or more of the amino acids which correspond to H82, N88, P89, D121, V150, P152, F240, F276, S287, H396 and Y415 in SEQ ID NO: 118, such as the amino acids which correspond to F98/H82, F98/N88, F98/P89, F98/D121, F98/V150, F98/ P152, F98/F240, F98/F276, F98/S287 or F98/H396. Table 3 shows the amino acids in SEQ ID NOs: 119 to 133 which correspond to H82, N88, P89, F98, D121, V150, P152, F240, F276, S287, H396 and Y415 in SEQ ID NO: 118.
The at least one amino acid which interacts with the sugar and/or base of one or more nucleotides in ssDNA is preferably at least one amino acid which intercalates between the nucleotides in ssDNA. Amino acids which intercalate between nucleotides in ssDNA can be modeled as discussed above. The at least one amino acid which intercalates between the nucleotides in ssDNA is preferably at least one of P89, F98 and V150 in SEQ ID NO: 118, such as P89, F98, V150, P89/F98, P89/V150, F98/V150 or P89/F98/V150.
The at least one amino acid which intercalates between the nucleotides in ssDNA in SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 is preferably at least one of the amino acids which correspond to P89, F98 and V150 in SEQ ID NO: 118, such as P89, F98, V150, P89/F98, P89/V150, F98/V150 or P89/F98/V150. Corresponding amino acids are shown in Table 3 above.
Larger R groups
The larger side chain (R group) preferably (a) contains an increased number of carbon atoms, (b) has an increased length, (c) has an increased molecular volume and/or (d) has an increased van der Waals volume. The larger side chain (R group) preferably (a); (b); (c); (d); (a) and (b); (a) and (c); (a) and (d); (b) and (c); (b) and (d); (c) and (d); (a), (b) and (c); (a), (b) and (d); (a), (c) and (d); (b), (c) and (d); or (a), (b), (c) and (d). Each of (a) to (d) may be measured using standard methods in the art.
The larger side chain (R group) preferably increases the (i) electrostatic interactions (ii) (ii) hydrogen bonding and/or (iii) cation-pi (cation-n) interactions between the at least one amino acid and the one or more nucleotides in ssDNA, such as increases (i); (ii); (iii); (i) and (ii); (i) and (iii); (ii) and (iii); and (i), (ii) and (iii). A skilled person can determine if the R group increases any of these interactions. For instance in (i), positively charged amino acids, such as arginine (R), histidine (H) and lysine (K), have R groups which increase electrostatic interactions. For instance in (ii), amino acids such as asparagine (N), serine (S), glutamine (Q), threonine (T) and histidine (H) have R groups which increase hydrogen bonding. For instance in (iii), aromatic amino acids, such as phenylalanine (F), tryptophan
(Wl), tyrosine (Y) or histidine (H), have R groups which increase cation-pi (cation-n) interactions. Specific substitutions below are labelled (i) to (iii) to reflect these changes. Other possible substitutions are labelled (iv). These (iv) substitutions typically increase the length of the side chain (R group).
The amino acid which comprises a larger side chain (R) may be a non-natural amino acid. The non-natural amino acid may be any of those discussed below.
The amino acid which comprises a larger side chain (R group) is preferably not alanine (A), cysteine (C), glycine (G), selenocysteine (U), methionine (M), aspartic acid (D) or glutamic acid (E).
Histidine (H) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q) or asparagine (N) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W). Histidine (H) is more preferably substituted with (a) N, Q or W or (b) Y, F, Q or K.
Asparagine (N) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q) or histidine (H) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W). Asparagine (N) is more preferably substituted with R, H, W or Y.
Proline (P) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N), threonine (T) or histidine (H), (iii) tyrosine (Y), phenylalanine (F) or tryptophan (W) or (iv) leucine (L), valine (V) or isoleucine (I). Proline (P) is more preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N), threonine (T) or histidine (H), (iii) phenylalanine (F) or tryptophan (W) or (iv) leucine (L), valine (V) or isoleucine (I). Proline (P) is more preferably substituted with (a) F, (b) L, V, I, T or F or (c) W, F, Y, H, I, L or V.
Valine (V) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H), (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W) or (iv) isoleucine (I) or leucine (L). Valine (V) is more preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H), (iii) tyrosine (Y) or tryptophan (W) or (iv) isoleucine (I) or leucine (L). Valine (V) is more preferably substituted with I or H or I, L, N, W or H.
Phenylalanine (F) is preferably substituted with (i) arginine (R) or lysine (K), (ii) histidine (H) or (iii) tyrosine (Y) or tryptophan (W). Phenylalanine (F) is more preferably substituted with (a) W, (b) W, Y or H, (c) W, R or K or (d) K, H, W or R.
Glutamine (Q) is preferably substituted with (i) arginine (R) or lysine (K) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W).
Alanine (A) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H), (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W) or (iv) isoleucine (I) or leucine (L).
Serine (S) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H), (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W) or (iv) isoleucine (I) or leucine (L). Serine (S) is preferably substituted with K, R, W or F
Lysine (K) is preferably substituted with (i) arginine (R) or (iii) tyrosine (Y) or tryptophan (W).
Arginine (R) is preferably substituted with (iii) tyrosine (Y) or tryptophan (W).
Methionine (M) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W).
Leucine (L) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q) or asparagine (N) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W).
Aspartic acid (D) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W). Aspartic acid (D) is more preferably substituted with H, Y or K.
Glutamic acid (E) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H) or (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W).
Isoleucine (I) is preferably substituted with (i) arginine (R) or lysine (K), (ii) glutamine (Q), asparagine (N) or histidine (H), (iii) phenylalanine (F), tyrosine (Y) or tryptophan (W) or (iv) leucine (L).
Tyrosine (Y) is preferably substituted with (i) arginine (R) or lysine (K) or (iii) tryptophan (W). Tyrosine (Y) is more preferably substituted with W or R.
The helicase more preferably comprises a variant of SEQ ID NO: 118 and comprises (a) P89F, (b) F98W, (c) V150I, (d) V150H, (e) P89F and F98W, (f) P89F and V150I, (g) P89F and V150H, (h) F98W and V150I, (i) F98W and V150H (j) P89F, F98W and V150I or (k) P89F, F98W and V150H.
The helicase more preferably comprises a variant of SEQ ID NO: 118 which comprises: H82N; H82Q; H82W; N88R; N88H; N88W; N88Y; P89L; P89V; P89I ; P89E; P89T; P89F; D121H; D121Y; D121K; V150I; V150L; V150N; V150W; V150H; P152W; P152F; P152Y;
P152H; P152I; P152L; P152V; F240W; F240Y; F240H; F276W; F276R; F276K; F276H; S287K; S287R; S287W; S287F; H396Y; H396F; H396Q; H396K; Y415W; Y415R; F98W/H82N; F98W/H82Q; F98W/H82W; F98W/N88R; F98W/N88H; F98W/N88W; F98W/N88Y; F98W/P89L; F98W/P89V; F98W/P89I; F98W/P89T; F98W/P89F; F98W/D121H; F98W/D121Y; F98W/D121K; F98W/V150I; F98W/V150L; F98W/V150N; F98W/V150W; F98W/V150H; F98W/P152W; F98W/P152F; F98W/P152Y; F98W/P152H; F98W/P152I; F98W/P152L; F98W/P152V; F98W/F240W; F98W/F240Y; F98W/F240H; F98W/F276W; F98W/F276R;F98W/F276K; F98W/F276H; F98W/S287K; F98W/S287R; F98W/S287W; F98W/S287F; F98W/H396Y; F98W/H396F; F98W/H396Q; F98W/Y415W; or F98W/Y415R.
Phosphate interactions
The helicase of the invention is preferably one in which at least one amino acid which interacts with one or more phosphate groups in one or more nucleotides in ssDNA is further substituted. Any number of amino acids may be substituted, such as 1 or more, 2 or more, 3 or more, 4 or more, 5 or more or 6 or more amino acids. Nucleotides in ssDNA each comprise three phosphate groups. Each amino which is substituted may interact with any number of the phosphate groups at a time, such as one, two or three phosphate groups at a time. The amino acids which interact with one or more phosphate groups can be identified using protein modelling as discussed above.
The substitution preferably increases the (i) electrostatic interactions, (ii) hydrogen bonding and/or (iii) cation-pi (cation-n) interactions between the at least one amino acid and the one or more phosphate groups in ssDNA. Preferred substitutions which increase (i), (ii) and (iii) are discussed below using the labelling (i), (ii) and (iii).
The substitution preferably increases the net positive charge of the position. The net charge at any position can be measured using methods known in the art. For instance, the isolectric point may be used to define the net charge of an amino acid. The net charge is typically measured at about 7.5. The substitution is preferably the substitution of a negatively charged amino acid with a positively charged, uncharged, non-polar or aromatic amino acid. A negatively charged amino acid is an amino acid with a net negative charge. Negatively charged amino acids include, but are not limited to, aspartic acid (D) and glutamic acid (E). A positively charged amino acid is an amino acid with a net positive charge. The positively charged amino acid can be naturally-occurring or non-naturally- occurring. The positively charged amino acid may be synthetic or modified. For instance, modified amino acids with a net positive charge may be specifically designed for use in the invention. A number of different types of modification to amino acids are well known in the art. Preferred naturally-occurring positively charged amino acids include, but are not limited to, histidine (H), lysine (K) and arginine (R).
The uncharged amino acid, non-polar amino acid or aromatic amino acid can be naturally occurring or non-naturally-occurring. It may be synthetic or modified. Uncharged amino acids have no net charge. Suitable uncharged amino acids include, but are not limited to, cysteine (C), serine (S), threonine (T), methionine (M), asparagines (N) and glutamine (Q). Non-polar amino acids have non-polar side chains. Suitable non-polar amino acids include, but are not limited to, glycine (G), alanine (A), proline (P), isoleucine (I), leucine (L) and valine (V). Aromatic amino acids have an aromatic side chain. Suitable aromatic amino acids include, but are not limited to, histidine (H), phenylalanine (F), tryptophan (W) and tyrosine (Y).
The helicase preferably comprises a variant of SEQ ID NO: 118 wherein the at least one amino acid which interacts with one or more phosphates in one or more nucleotides in ssDNA is at least one of H64, T80, S83, N242, K243, N293, T394 and K397. These numbers correspond to the relevant positions in SEQ ID NO: 89 and may need to be altered in the case of variants where one or more amino acids have been inserted or deleted compared with SEQ ID NO: 118. A skilled person can determine the corresponding positions in a variant as discussed above.
The helicase preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 and wherein the at least one amino acid which interacts with one or more phosphates in one or more nucleotides in ssDNA is at least one of the amino acids which correspond to H64, T80, S83, N242, K243, N293, T394 and K397 in SEQ ID NO: 118.
Table 4 shows the amino acids in SEQ ID NOs: 119 to 133 which correspond to H64, T80, S83, N242, K243, N293, T 394 and K397 in SEQ ID NO: 118.
Histidine (H) is preferably substituted with (i) arginine (R) or lysine (K), (ii) asparagine (N), serine (S), glutamine (Q) or threonine (T), (iii) phenylalanine (F), tryptophan (W) or tyrosine (Y). Histidine (H) is preferably substituted with (a) N, Q, K or F or (b) N, Q or W. Threonine (T) is preferably substituted with (i) arginine (R), histidine (H) or lysine (K), (ii) asparagine (N), serine (S), glutamine (Q) or histidine (H) or (iii) phenylalanine (F), tryptophan (W), tyrosine (Y) or histidine (H). Threonine (T) is more preferably substituted with (a) K, Q or N or (b) K, H or N.
Serine (s) is preferably substituted with (i) arginine (R), histidine (H) or lysine (K), (ii) asparagine (N), glutamine (Q), threonine (T) or histidine (H) or (iii) phenylalanine (F), tryptophan (W), tyrosine (Y) or histidine (H). Serine (S) is more preferably substituted with H, N, K, T, R or Q.
Asparagine (N) is preferably substituted with (i) arginine (R), histidine (H) or lysine (K), (ii) serine (S), glutamine (Q), threonine (T) or histidine (H) or (iii) phenylalanine (F), tryptophan (W), tyrosine (Y) or histidine (H). Asparagine (N) is more preferably substituted with (a) H or Q or (b) Q, K or H.
Lysine (K) is preferably substituted with (i) arginine (R) or histidine (H), (ii) asparagine (N), serine (S), glutamine (Q), threonine (T) or histidine (H) or (iii) phenylalanine (F),
tryptophan (W), tyrosine (Y) or histidine (H). Lysine (K) is more preferably substituted with (a) Q or H or (b) R, H or Y.
The helicase more preferably comprises a variant of SEQ ID NO: 118 and comprises one or more of, such as all of, (a) H64N, H64Q, H64K or H64F, (b) T80K, T80Q or T80N, (c) S83H, S83N, S83K, S83T, S83R, or S83Q (d) N242H or N242Q, (e) K243Q or K243H, (f) N293Q, N293K or N293H, (g) T394K, T394H or T394N or (h) K397R, K397H or K397Y.
Combinations
The helicase is preferably a variant of SEQ ID NO: 118 which comprises substitutions at:
- F98/H64, such as F98W/H64N, F98W/H64Q, F98W/H64K or F98W/H64F;
- F98/T80, such as F98W/T80K, F98W/T80Q, F98W/T80N;
- F98/H82, such as F98W/H82N, F98W/H82Q or F98W/H82W;
- F98/S83, such as F98W/S83H, F98W/S83N, F98W/S83K, F98W/S83T, F98W/S83R or F98W/S83Q;
- F98/N242, such as F98W/N242H, F98W/N242Q, F98W/K243Q or F98W/K243H;
- F98/N293, such as F98W/N293Q, F98W/N293K, F98W/N293H, F98W/T394K, F98W/T394H, F98W/T394N, F98W/H396Y, F98W/H396F, F98W/H396Q or F98W/H396K; or
- F98/K397, such as F98W/K397R, F98W/K397H or F98W/K397Y.
Preferred combinations in SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 include the combinations of amino acids which correspond to the combinations in SEQ ID NO: 118 listed above.
Pore interaction
The helicase of the invention is further one in which the part of the helicase which interacts with a transmembrane pore further comprises one or more modifications, preferably one or more substitutions. The part of the helicase which interacts with a transmembrane pore is typically the part of the helicase which interacts with a transmembrane pore when the helicase is used to control the movement of a polynucleotide through the pore, for instance as discussed in more detail below. The part typically comprises the amino acids that interact with or contact the pore when the helicase is used to control the movement of a
polynucleotide through the pore, for instance as discussed in more detail below. The part typically comprises the amino acids that interact with or contact the pore when the helicase is bound to or attached to an analyte such as polynucleotide which is moving through the pore under an applied potential.
In SEQ ID NO: 118, the part which interacts with the transmembrane pore typically comprises the amino acids at positions 1, 2, 3, 4, 5, 6, 51, 176, 177, 178, 179, 180, 181, 185, 189, 191, 193, 194, 195, 197, 198, 199, 200, 201, 202, 203, 204, 207, 208, 209,
210, 211, 212, 213, 216, 219, 220, 221, 223, 224, 226, 227, 228, 229, 247, 254, 255,
256, 257, 258, 259, 260, 261, 298, 300, 304, 308, 318, 319, 321, 337, 347, 350, 351,
405, 415, 422, 434, 437, 438. These numbers correspond to the relevant positions in SEQ
ID NO: 118 and may need to be altered in the case of variants where one or more amino acids have been inserted or deleted compared with SEQ ID NO: 118. A skilled person can determine the corresponding positions in a variant as discussed above. The part which interacts with the transmembrane pore preferably comprises the amino acids at
(a) positions 1, 2, 4, 51, 177, 178, 179, 180, 185, 193, 195, 197, 198, 199, 200, 202, 203, 204, 207, 208, 209, 210, 211, 212, 216, 221, 223, 224, 226, 227, 228, 229, 254, 255, 256, 257, 258, 260, 304, 318, 321, 347, 350, 351, 405, 415, 422, 434, 437 and 438 in SEQ ID NO: 118; or
(b) positions 1, 2, 178, 179, 180, 185, 195, 197, 198, 199, 200, 202, 203, 207, 209, 210, 212, 216, 221, 223, 226, 227, 255, 258, 260, 304, 350 and 438 in SEQ ID NO: 118.
The part which interacts with the transmembrane pore preferably comprises one or more of, such as 2, 3, 4 or 5 of, the amino acids at positions K194, W195, K198, K199 and E258 in SEQ ID NO: 118. The variant of SEQ ID NO: 118 preferably comprises a modification at one or more of (a), K194, (b) W195, (c) D198, (d) K199 and (d) E258. The variant of SEQ ID NO: 118 preferably comprises a substitution at one or more of (a) K194, such as K194L, (b) W195, such as W195A, (c) D198, such as D198V, (d) K199, such as K199L and (e) E258, such as E258L. The variant may comprise {a}; {b}; {c}; {d}; {e}; {a,b}; {a,c}; {a,d}; {a,e}; {b,c}; {b,d}; {b,e}; {c,d}; {c,e}; {d,e}; {a,b,c}; {a,b,d}; {a,b,e}; {a,c,d}; {a,c,e}; {a,d,e}; {b,c,d}; {b,c,e}; {b,d,e}; {c,d,e}; {a,b,c,d}; {a,b,c,e}; {a,b,d,e}; {a,c,d,e}; {b,c,d,e}; or {a,b,c,d,e}. The modifications or substitutions set out in this paragraph are preferred when the modified polynucleotide binding protein interacts with a pore derived from MspA, particularly any of the modified pores discussed below.
The part of the polynucleotide binding protein which interacts with the transmembrane pore preferably comprises the amino acid at position 194 or 199 of SEQ ID NO: 118. The variant preferably comprises K194A, K194V, K194F, K194D, K194S, K194W or K194L and/or K199A, K199V, K199F, K199D, K199S, K199W or K199L. In SEQ ID NO: 119, 120, 121,
122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 , the part which interacts with the transmembrane pore typically comprises the amino acids at positions which correspond to those in SEQ ID NO: 118 listed above. Amino acids in SEQ ID NOs: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 and 133 which correspond to these positions in SEQ ID NO: 118 can be identified using the alignment in Table 5 below.
Preferred combinations
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at F98, such as F98R, F98K, F98Q, F98N, F98H, F98Y, F98F or F98W, and a substitution at K194, such as K194A, K194V, K194F, K194D, K194S, K194W
or K194L, and/or K199, such as K199A, K199V, K199F, K199D, K199S, K199W or K199L. The helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 which further comprises a substitution at the position which corresponds to F98 in SEQ ID NO: 118 and a substitution at the position(s) which correspond to K194 and/or K199 in SEQ ID NO: 118. These corresponding positions may be replaced with any of the amino acids listed above for F98, K194 and K119 in SEQ ID NO: 118.
The helicase is preferably a variant of SEQ ID NO: 118 which further comprises substitutions at:
- F98/K194/H64, such as F98W/K194L/H64N, F98W/K194L/H64Q, F98W/K194L/H64K or F98W/K194L/H64F;
- F98/K194/T80, such as F98W/K194L/T80K, F98W/K194L/T80Q or F98W/K194L/T80N;
- F98/K194/H82, such as F98W/K194L/H82N, F98W/K194L/H82Q or F98W/K194L/H82W
- F98/S83/K194, such as F98W/S83H/K194L, F98W/S83T/K194L, F98W/S83R/K194L, F98W/S83Q/K194L, F98W/S83N/K194L, F98W/S83K/K194L, F98W/N88R/K194L, F98W/N88H/K194L, F98W/N88W/K194L or F98W/N88Y/K194L;
- F98/S83/K194/F276, such as F98W/S83H/K194L/F276K;
- F98/P89/K194, such as F98W/P89L/K194L, F98W/P89V/K194L, F98W/P89I/K194L or F98W/P89T/K194L;
- F98/D121/K194, such as F98W/D121H/K194L, F98W/D121Y/K194L or F98W/D121K/K194L;
- F98/V150/K194, such as F98W/V150I/K194L, F98W/V150L/K194L, F98W/V150N/K194L, F98W/V150W/K194L or F98W/V150H/K194L;
- F98/P152/K194, such as F98W/P152W/K194L, F98W/P152F/K194L, F98W/P152Y/K194L, F98W/P152H/K194L, F98W/P152I/K194L, F98W/P152L/K194L or F98W/P152V/K194L;
- F98/F240/K194, such as F98W/F240W/K194L, F98W/F240Y/K194L or F98W/F240H/K194L;
- F98/N242/K194, such as F98W/N242H/K194L or F98W/N242Q/K194L;
- F98/K194/F276, such as F98W/K194L/F276K, F98W/K194L/F276H, F98W/K194L/F276W or F98W/K194L/F276R;
- F98/K194/S287, such as F98W/K194L/S287K, F98W/K194L/S287R, F98W/K194L/S287W or F98W/K194L/S287F;
- F98/N293/K194, such as F98W/N293Q/K194L, F98W/N293K/K194L or F98W/N293H/K194L;
- F98/T394/K194, such as F98W/T394K/K194L, F98W/T394H/K194L or F98W/T394N/K194L;
- F98/H396/K194, such as F98W/H396Y/K194L, F98W/H396F/K194L, F98W/H396Q/K194L or F98W/H396K/K194L;
- F98/K397/K194, such as F98W/K397R/K194L, F98W/K397H/K194L or F98W/K397Y/K194L; or
- F98/Y415/K194, such as F98W/Y415W/K194L or F98W/Y415R/K194L.
In any of the above combinations, K194 may be replaced with any of W195, D198, K199 and E258.
The modified helicase preferably further comprises a modification or substitution at the position(s) corresponding to amino acid positions 98 and/or 194 in Dda 1993. Position 98 or the corresponding position may be substituted with R, H, K, S, T, N, Q, A, V, I, L, M, Y or W. Position 98 or the corresponding position is preferably substituted with R, K, Q, N, H, Y or W. Position 194 or the corresponding position may be substituted with A, V, I, L, M, F, Y, W, D, E, S, T, N or Q. Position 194 or the corresponding position is preferably substituted with A, V, F, D, S, W or L. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at F98, such as F98R, F98K, F98Q, F98N, F98H, F98Y or F98W, and/or a substitution at K194, such as K194A, K194V, K194F, K194D, K194S, K194W or K194L. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises F98W and K194L. The helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 which further comprises a substitution at the position which corresponds to F98 in SEQ ID NO: 118 and/or a substitution at the position which corresponds to K194 in SEQ ID NO: 118.
In any of the above combinations, K194 may be replaced with any of W195, D198, K199 and E258.
Modifications in the tower domain and/or pin domain and/or 1A domain
The modified helicase preferably further comprises a modification or substitution at the position corresponding to amino acid position 360 in Dda 1993. Position 360 or the corresponding position may be substituted with C, G, P, A, V, I, L, M, F, Y or W. Position 360 or the corresponding position is preferably substituted with C or Y. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at A360, such as A360C or A360Y. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises K358I and A360C. The helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 which comprises a substitution at the position which corresponds to A360 in SEQ ID NO: 118.
The modified helicase preferably further comprises a modification or substitution at one or more of the positions corresponding to amino acid positions 94, 98 and 109 in Dda 1993, such as position(s) 94, 98, 109, 94 and 98, 94 and 109, 98 and 109 and 94, 98 and 109. These positions are all in the pin domain. Position 94 or the corresponding position may be substituted with C, G, P, A, V, I, L, M, F, Y or W. Position 94 or the corresponding position is preferably substituted with C or Y. Position 98 or the corresponding position may be substituted with R, H, K, S, T, N, Q, A, V, I, L, MY or W. Position 98 or the corresponding position is preferably substituted with R, K, Q, N, H, Y or W. Position 109 or the corresponding position may be substituted with A, V, I, L, M, F, Y or W. Position 109 or the corresponding position is preferably substituted with A or V. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a substitution at one or more of E94, F98 and C109 (including all the combinations set out above). Preferred variants comprise substitutions at:
- E94 and F98, such as E94C or E94Y and F98R, F98K, F98Q, F98N, F98H, F98Y, F98F or F98W;
- E94 and C109, such as E94C or E94Y and C109A or C109V;
- F98 and C109, such as F98R, F98K, F98Q, F98N, F98H, F98Y, F98F or F98W and C109A or C109V; or
- E94, F98 and C109, such as E94C or E94Y and F98R, F98K, F98Q, F98N, F98H, F98Y, F98F or F98W and C109A or C109V.
More preferred variants comprise: E94C and F98W; E94C and C109A; F98W and C109A; or E94C, F98W and C109A. The helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 which
further comprises a substitution at the position(s) which corresponds to one or more of E94, F98 and C109 in SEQ ID NO: 118.
Table 6 includes information for E94, C109, C136 and A360 (with reference to modified helicases disclosed above and below).
The helicase of the invention is preferably one in which at least one cysteine residue (i.e. one or more cysteine residues) and/or at least one non-natural amino acid (i.e. one or more non-natural amino acids) have further been introduced into (i) the tower domain and/or (ii) the pin domain and/or the (iii) 1A (RecA-like motor) domain, wherein the helicase has the ability to control the movement of a polynucleotide. These types of modification are disclosed in WO 2015/055981 (incorporated herein by reference in its entirety). At least one cysteine residue and/or at least one non-natural amino acid may be introduced into the tower domain, the pin domain, the 1A domain, the tower domain and the pin domain, the tower domain and the 1A domain or the tower domain, the pin domain and the 1A domain. The helicase of the invention is preferably one in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into each of (i) the tower domain and (ii) the pin domain and/or the 1A (RecA-like motor) domain, i.e. into the tower domain and the pin domain, the tower domain and the 1A domain or the tower domain, the pin domain and the 1A domain.
Any number of cysteine residues and/or non-natural amino acids may be introduced into each domain. For instance, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more cysteine residues may be introduced and/or 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more non-natural amino acids may be introduced. Only one or more cysteine residues may be introduced. Only one or more non- natural amino acids may be introduced. A combination of one or more cysteine residues and one or more non-natural amino acids may be introduced.
The at least one cysteine residue and/or at least one non-natural amino acid are/is preferably introduced by substitution. Methods for doing this are known in the art.
These modifications do not prevent the helicase from binding to a polynucleotide. These modifications decrease the ability of the polynucleotide to unbind or disengage from the helicase. In other words, the one or more modifications increase the processivity of the helicase by preventing dissociation from the polynucleotide strand. The thermal stability of the enzyme is typically also increased by the one or more modifications giving it an improved structural stability that is beneficial in Strand Sequencing.
A non-natural amino acid is an amino that is not naturally found in a helicase. The non- natural amino acid is preferably not histidine, alanine, isoleucine, arginine, leucine, asparagine, lysine, aspartic acid, methionine, cysteine, phenylalanine, glutamic acid, threonine, glutamine, tryptophan, glycine, valine, proline, serine or tyrosine. The non- natural amino acid is more preferably not any of the twenty amino acids in the previous sentence or selenocysteine.
Preferred non-natural amino acids for use in the invention include, but are not limited, to 4- Azido-L-phenylalanine (Faz), 4-Acetyl-L-phenylalanine, 3-Acetyl-L-phenylalanine, 4- Acetoacetyl-L-phenylalanine, O-Allyl-L-tyrosine, 3-(Phenylselanyl)-L-alanine, O-2-Propyn-l- yl-L-tyrosine, 4-(Dihydroxyboryl)-L-phenylalanine, 4- [(Ethylsulfa nyl)carbonyl]-L- phenylalanine, (2S)-2-amino-3-4-[(propan-2-ylsulfanyl)carbonyl] phenyl; propanoic acid, (2S)-2-a m i no-3-4- [(2-amino-3-sulfanylpropanoyl)a mi no] phenyl; propanoic acid, O-Methyl- L-tyrosine, 4-Amino-L-phenylalanine, 4-Cyano-L-phenylalanine, 3-Cyano-L-phenylalanine, 4-Fluoro-L-phenylalanine, 4-Iodo-L-phenylalanine, 4-Bromo-L-phenylalanine, O- (Trifluoromethyl)tyrosine, 4-Nitro-L-phenylalanine, 3-Hydroxy-L-tyrosine, 3-Amino-L- tyrosine, 3-Iodo-L-tyrosine, 4-Isopropyl-L-phenylalanine, 3-(2-Naphthyl)-L-alanine, 4- Phenyl-L-phenylalanine, (2S)-2-amino-3-(naphthalen-2-ylamino)propanoic acid, 6- (Methylsulfanyl)norleucine, 6-Oxo-L-lysine, D-tyrosine, (2R)-2-Hydroxy-3-(4- hydroxyphenyl) propanoic acid, (2 R)-2-Am mon ioocta noate3-(2,2'-Bipyridin-5-yl)-D-a la nine, 2-amino-3-(8-hydroxy-3-quinolyl)propanoic acid, 4-Benzoyl-L-phenylalanine, S-(2- Nitrobenzyl)cysteine, (2R)-2-amino-3-[(2-nitrobenzyl)sulfanyl]propanoic acid, (2S)-2- amino-3-[(2-nitrobenzyl)oxy] propanoic acid, O-(4,5-Dimethoxy-2-nitrobenzyl)-L-serine,
(2S)-2-amino-6-([(2-nitrobenzyl)oxy]carbonyl;amino)hexanoic acid, O-(2-Nitrobenzyl)-L- tyrosine, 2-Nitrophenylalanine, 4-[(E)-Phenyldiazenyl]-L-phenylalanine, 4-[3- (Trifluoromethyl)-3H-diaziren-3-yl]-D-phenylalanine, 2-amino-3-[[5-(dimethylamino)-l- na phthy I] sulfonyla mi no] propanoic acid, (2S)-2-amino-4-(7-hydroxy-2-oxo-2/7-chromen-4- yl)butanoic acid, (2S)-3-[(6-acetylnaphthalen-2-yl)amino]-2-aminopropanoic acid, 4- (Carboxymethyl)phenylalanine, 3-Nitro-L-tyrosine, O-Sulfo-L-tyrosine, (2R)-6-Acetamido-2- ammoniohexanoate, 1-Methylhistidine, 2-Aminononanoic acid, 2-Aminodecanoic acid, L- Homocysteine, 5-Sulfanylnorvaline, 6-Sulfanyl-L-norleucine, 5-(Methylsulfanyl)-L-norvaline, N6-[(2R,3R)-3-Methyl-3,4-dihydro-2H-pyrrol-2-yl]carbonyl;-L-lysine, N6- [(Benzyloxy)carbony I] lysine, (2S)-2-a mi no-6- [(cyclopentylcarbonyl)amino] hexanoic acid, N6-[(Cyclopentyloxy)carbonyl]-L-lysine, (2S)-2-amino-6-[(2R)-tetra hydrofuran -2- ylcarbonyl] amino; hexanoic acid, (2S)-2-amino-8-[(2R,3S)-3-ethynyltetrahydrofuran-2-yl]- 8-oxooctanoic acid, N6-(tert-Butoxycarbonyl)-L-lysine, (2S)-2-Hydroxy-6-([(2-methyl-2- propanyl)oxy]carbonyl;amino)hexanoic acid, N6-[(Allyloxy)carbonyl]lysine, (2S)-2-amino-6- ([(2-azidobenzyl)oxy]carbonyl;amino)hexanoic acid, N6-L-Prolyl-L-lysine, (2S)-2-amino-6- [(prop-2-yn-l-yloxy)carbonyl]amino;hexanoic acid and N6-[(2-Azidoethoxy)carbonyl]-L- lysine. The most preferred non-natural amino acid is 4-azido-L-phenylalanine (Faz).
Table 7 below (which is separated in two parts) identifies the residues making up each domain in each Dda homologue (SEQ ID NOs: 118 to 133).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues D260-P274 and N292-A389) and/or (ii) the pin domain (residues K86-E102) and/or the (iii) 1A domain (residues M1-L85 and V103- K177). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N292-A389 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 119 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues G295-N309 and F316-Y421) and/or (ii) the pin domain (residues Y85-L112) and/or the (iii) 1A domain (residues MI-184 and R.113- Y211). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues F316-Y421 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 120 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues V328-P342 and N360-Y448) and/or (ii) the pin domain (residues K148-N165) and/or the (iii) 1A domain (residues M1-L147 and S166- V240). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N360-Y448 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 121 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues A261-T275 and T285-Y370) and/or (ii) the pin domain (residues G91-E107) and/or the (iii) 1A domain (residues M1-L90 and E108-
H173). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues T285-Y370 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 122 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues G294-I307 and T314-Y407) and/or (ii) the pin domain (residues G116-T135) and/or the (iii) 1A domain (residues M1-L115 and N136- V205). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues T314-Y407 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 123 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues V288-E301 and N307-N393) and/or (ii) the pin domain (residues G97-P113) and/or the (iii) 1A domain (residues M1-L96 and F114- V194). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N307-N393 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 124 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues S250-P264 and E278-S371) and/or (ii) the pin domain (residues K78-E95) and/or the (iii) 1A domain (residues M1-L77 and V96-V166). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues E278-S371 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 125 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues K255-P269 and T284-S380) and/or (ii) the pin domain (residues K82-K98) and/or the (iii) 1A domain (residues M1-M81 and L99- M171). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues T284-S380 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 126 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues D242-P256 and T271-S366) and/or (ii) the pin domain (residues K69-K85) and/or the (iii) 1A domain (residues M1-M68 and MSG- MISS). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues T271-S366 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 127 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues T263-P277 and N295-P392) and/or (ii) the
pin domain (residues K88-K107) and/or the (iii) 1A domain (residues M1-L87 and A108- M181). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N295-P392 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 128 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues D263-P277 and N295-A391) and/or (ii) the pin domain (residues K88-K107) and/or the (iii) 1A domain (residues M1-L87 and A108- M181). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N295-A391 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 129 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues A258-P272 and N290-P386) and/or (ii) the pin domain (residues K86-G102) and/or the (iii) 1A domain (residues M1-L85 and T103- K176). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N290-P386 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 130 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues L266-P280 and N298-A392) and/or (ii) the pin domain (residues K92-D108) and/or the (iii) 1A domain (residues M1-L91 and V109- M183). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N298-A392 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 131 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues D262-P276 and N294-A392) and/or (ii) the pin domain (residues K88-E104) and/or the (iii) 1A domain (residues M1-L87 and M105- M179). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N294-A392 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 132 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into (i) the tower domain (residues D261-P275 and N293-A389) and/or (ii) the pin domain (residues K87-E103) and/or the (iii) 1A domain (residues M1-L86 and V104- K178). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues N293-A389 of the tower domain.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 133 in which at least one cysteine residue and/or at least one non-natural amino acid have further been
introduced into (i) the tower domain (residues E261-P275 and T293-A390) and/or (ii) the pin domain (residues K87-E103) and/or the (iii) 1A domain (residues M1-L86 and V104- M178). The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced into residues T293-A390 of the tower domain.
The helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 118 to 133 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into each of (i) the tower domain and (ii) the pin domain and/or the 1A domain. The helicase of the invention more preferably comprises a variant of any one of SEQ ID NOs: 118 to 133 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into each of (i) the tower domain, (ii) the pin domain and (iii) the 1A domain. Any number and combination of cysteine residues and non-natural amino acids may be introduced as discussed above.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises (i) E94C and/or A360C; (ii) E93C and/or K358C; (iii) E93C and/or A360C; (iv) E93C and/or E361C; (v) E93C and/or K364C; (vi) E94C and/or L354C; (vii) E94C and/or K358C; (viii) E93C and/or L354C; (ix) E94C and/or E361C; (x) E94C and/or K364C; (xi) L97C and/or L354C; (xii) L97C and/or K358C; (xiii) L97C and/or A360C; (xiv) L97C and/or E361C; (xv) L97C and/or K364C; (xvi) K123C and/or L354C; (xvii) K123C and/or K358C; (xviii) K123C and/or A360C; (xix) K123C and/or E361C; (xx) K123C and/or K364C; (xxi) N155C and/or L354C; (xxii) N155C and/or K358C; (xxiii) N155C and/or A360C; (xxiv) N155C and/or E361C; (xxv) N155C and/or K364C; (xxvi) any of (i) to (xxv) and G357C; (xxvii) any of (i) to (xxv) and Q100C; (xxviii) any of (i) to (xxv) and I127C; (xxix) any of (i) to (xxv) and Q100C and I127C; (xxx) E94C and/or F377C; (xxxi) N95C; (xxxii) T91C; (xxxiii) Y92L, E94Y, Y350N, A360C and Y363N; (xxxiv) E94Y and A360C; (xxxv) A360C; (xxxvi) Y92L, E94C, Y350N, A360Y and Y363N; (xxxvii) Y92L, E94C and A360Y; (xxxviii) E94C and/or A360C and F276A; (xxxix) E94C and/or L356C; (xl) E93C and/or E356C; (xli) E93C and/or G357C; (xlii) E93C and/or A360C; (xliii) N95C and/or W378C; (xliv) T91C and/or S382C; (xlv) T91C and/or W378C; (xlvi) E93C and/or N353C; (xlvii) E93C and/or S382C; (xlviii) E93C and/or K381C; (xlix) E93C and/or D379C; (I) E93C and/or S375C; (Ii) E93C and/or W378C; (Iii) E93C and/or W374C; (liii) E94C and/or N353C; (liv) E94C and/or S382C; (Iv) E94C and/or K381C; (Ivi) E94C and/or D379C; (Ivii) E94C and/or S375C; (Iviii) E94C and/or W378C; (lix) E94C and/or W374C; (Ix) E94C and A360Y; (Ixi) E94C, G357C and A360C or (Ixii) T2C, E94C and A360C. In any one of (i) to (Ixii), and/or is preferably and.
The helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises a cysteine residue at the positions which correspond to those in SEQ ID NO: 118 as defined in any of (i) to (Ixii). Positions in any one of SEQ ID
NOs: 119 to 133 which correspond to those in SEQ ID NO: 118 can be identified using the alignment of SEQ ID NOs: 118 to 133 below. The helicase of the invention preferably comprises a variant of SEQ ID NO: 92 which further comprises (a) D99C and/or L341C, (b) Q98C and/or L341C or (d) Q98C and/or A340C. The helicase of the invention preferably comprises a variant of SEQ ID NO: 96 which further comprises D90C and/or A349C. The helicase of the invention preferably comprises a variant of SEQ ID NO: 102 which further comprises D96C and/or A362C.
The helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 118 to 133 as defined in any one of (i) to (Ixii) in which Faz is further introduced at one or more of the specific positions instead of cysteine. Faz may be introduced at each specific position instead of cysteine. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises (i) E94Faz and/or A360C; (ii) E94C and/or A360Faz; (iii) E94Faz and/or A360Faz; (iv) Y92L, E94Y, Y350N, A360Faz and Y363N; (v) A360Faz; (vi) E94Y and A360Faz; (vii) Y92L, E94Faz, Y350N, A360Y and Y363N; (viii) Y92L, E94Faz and A360Y; (ix) E94Faz and A360Y; and (x) E94C, G357Faz and A360C.
The helicase of the invention preferably further comprises one or more single amino acid deletions from the pin domain. Any number of single amino acid deletions may be made, such as 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more. The helicase more preferably comprises a variant of SEQ ID NO: 118 which further comprises deletion of E93, deletion of E95 or deletion of E93 and E95. The helicase more preferably comprises a variant of SEQ ID NO: 118 which comprises (a) E94C, deletion of N95 and A360C; (b) deletion of E93, deletion of E94, deletion of N95 and A360C; (c) deletion of E93, E94C, deletion of N95 and A360C or (d) E93C, deletion of N95 and A360C. The helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises deletion of the position corresponding to E93 in SEQ ID NO: 118, deletion of the position corresponding to E95 in SEQ ID NO: 118 or deletion of the positions corresponding to E93 and E95 in SEQ ID NO: 118.
The helicase of the invention preferably further comprises one or more single amino acid deletions from the hook domain. Any number of single amino acid deletions may be made, such as 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more. The helicase more preferably comprises a variant of SEQ ID NO: 118 which further comprises deletion of any number of positions T278 to S287. The helicase more preferably comprises a variant of SEQ ID NO: 118 which further comprises (a) E94C, deletion of Y279 to K284 and A360C, (b) E94C, deletion of T278, Y279, V286 and S287 and A360C, (c) E94C, deletion of 1281 and K284 and replacement with a single G and A360C, (d) E94C, deletion of K280 and P2845 and replacement with a single G and A360C, or (e) deletion of Y279 to K284, E94C, F276A and A230C. The helicase of the invention preferably comprises a variant of any one of SEQ ID
NOs: 119 to 133 which further comprises deletion of any number of the positions corresponding to 278 to 287 in SEQ ID NO: 118.
The helicase of the invention preferably further comprises one or more single amino acid deletions from the pin domain and one or more single amino acid deletions from the hook domain.
The helicase of the invention is preferably one in which at least one cysteine residue and/or at least one non-natural amino acid have been further introduced into the hook domain and/or the 2A (RecA-like) domain. Any number and combination of cysteine residues and non-natural amino acids may be introduced as discussed above for the tower, pin and 1A domains.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L275-F291) and/or the 2A (RecA-like) domain (residues R178-T259 and L390-V439).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 119 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues A310-L315) and/or the 2A (RecA-like) domain (residues R212-E294 and G422-S678).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 120 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues V343-L359) and/or the 2A (RecA-like) domain (residues R241-N327 and A449-G496).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 121 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues W276-L284) and/or the 2A (RecA-like) domain (residues R174-D260 and A371-V421).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 122 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues R308-Y313) and/or the 2A (RecA-like) domain (residues R206-K293 and I408-L500).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 123 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues M302-W306) and/or the 2A (RecA-like) domain (residues R195-D287 and V394-Q450).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 124 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues V265-I277) and/or the 2A (RecA-like) domain (residues R167-T249 and L372-N421).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 125 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues V270-F283) and/or the 2A (RecA-like) domain (residues R172-T254 and L381-K434).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 126 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues V257-F270) and/or the 2A (RecA-like) domain (residues R159-T241 and L367-K420).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 127 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L278-Y294) and/or the 2A (RecA-like) domain (residues R182-T262 and L393-V443).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 128 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L278-Y294) and/or the 2A (RecA-like) domain (residues R182-T262 and L392-V442).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 129 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L273-F289) and/or the 2A (RecA-like) domain (residues R177-N257 and L387-V438).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 130 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L281-F297) and/or the 2A (RecA-like) domain (residues R184-T265 and L393-I442).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 131 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues H277-F293) and/or the 2A (RecA-like) domain (residues R180-T261 and L393-V442).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 132 in which at least one cysteine residue and/or at least one non-natural amino acid have further been
introduced into the hook domain (residues L276-F292) and/or the 2A (RecA-like) domain (residues R179-T260 and L390-I439).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 133 in which at least one cysteine residue and/or at least one non-natural amino acid have further been introduced into the hook domain (residues L276-F292) and/or the 2A (RecA-like) domain (residues R179-T260 and L391-V441).
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises one or more of (i) I181C; (ii) Y279C; (iii) I281C; and (iv) E288C. The helicase may comprise any combination of (i) to (iv), such as (i); (ii); (iii); (iv); (i) and (ii); (i) and (iii); (i) and (iv); (ii) and (iii); (ii) and (iv); (iii) and (iv); or (i), (ii), (iii) and (iv). The helicase more preferably comprises a variant of SEQ ID NO: 118 which comprises (a) E94C, I281C and A360C or (b) E94C, I281C, G357C and A360C. The helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises a cysteine residue at one or more of the position(s) which correspond to those in SEQ ID NO: 118 as defined in (i) to (iv), (a) and (b). The helicase may comprise any of these variants in which Faz is introduced at one or more of the specific positions (or each specific position) instead of cysteine.
The helicase of the invention is preferably further modified to reduce its surface negative charge. Surface residues can be identified in the same way as the Dda domains disclosed above. Surface negative charges are typically surface negatively-charged amino acids, such as aspartic acid (D) and glutamic acid (E).
The helicase is preferably modified to neutralise one or more surface negative charges by substituting one or more negatively charged amino acids with one or more positively charged amino acids, uncharged amino acids, non-polar amino acids and/or aromatic amino acids or by introducing one or more positively charged amino acids, preferably adjacent to one or more negatively charged amino acids. Suitable positively charged amino acids include, but are not limited to, histidine (H), lysine (K) and arginine (R). Uncharged amino acids have no net charge. Suitable uncharged amino acids include, but are not limited to, cysteine (C), serine (S), threonine (T), methionine (M), asparagine (N) and glutamine (Q). Non-polar amino acids have non-polar side chains. Suitable non-polar amino acids include, but are not limited to, glycine (G), alanine (A), proline (P), isoleucine (I), leucine (L) and valine (V). Aromatic amino acids have an aromatic side chain. Suitable aromatic amino acids include, but are not limited to, histidine (H), phenylalanine (F), tryptophan (W) and tyrosine (Y).
Preferred substitutions include, but are not limited to, substitution of E with R, substitution of E with K, substitution of E with N, substitution of D with K and substitution of D with R.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 and the one or more negatively charged amino acids are one or more of D5, E8, E23, E47, D167, E172, D202, D212 and E273. Any number of these amino acids may be neutralised, such as 1, 2, 3, 4, 5, 6, 7 or 8 of them. Any combination may be neutralised. The helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 and the one or more negatively charged amino acids correspond to one or more of D5, E8, E23, E47, D167, E172, D202, D212 and E273 in SEQ ID NO: 118. Amino acids in SEQ ID NOs: 119 to 133 which correspond to D5, E8, E23, E47, D167, E172, D202, D212 and E273 in SEQ ID NO: 118 can be determined using the alignment in WO2015/055981. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which comprises (a) E94C, E273G and A360C or (b) E94C, E273G, N292G and A360C.
The helicase of the invention is preferably further modified by the removal of one or more native cysteine residues. Any number of native cysteine residues may be removed. The one or more cysteine residues are preferably removed by substitution. The one or more cysteine residues are preferably substituted with alanine (A), serine (S) or valine (V). The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 and the one or more native cysteine residues are one or more of C109, C114, C136, C171 and C412. Any number and combination of these cysteine residues may be removed. For instance, the variant of SEQ ID NO: 118 may comprise C109; C114; C136; C171; C412; C109 and C114; C109 and C136; C109 and C171; C109 and C412; C114 and C136; C114 and C171; C114 and C412; C136 and C171; C136 and C412; C171 and C412; C109, C114 and C136; C109, C114 and C171; C109, C114 and C412; C109, C136 and C171; C109, C136 and C412; C109, C171 and C412; C114, C136 and C171; C114, C136 and C412; C114, C171 and C412; C136, C171 and C412; C109, C114, C136 and C171; C109, C114, C136 and C412; C109, C114, C171 and C412; C109, C136, C171 and C412; C114, C136, C171 and C412; or C109, C114, C136, C171 and C412.
The modified helicase preferably further comprises a modification or substitution at the position(s) corresponding to amino acid position(s) 109 and/or 136 in Dda 1993. This removes one or two cysteine residues. This may be in addition to a modification or substitution at one or more positions corresponding to amino acid positions 55, 114, 156, 177, 210, 221, 350 and 358 in Dda 1993, a modification or substitution at one or more positions corresponding to amino acid positions 114, 177, 350 and 358 in Dda 1993 and/or a modification or substitution at the position corresponding to position 40 in Dda 1993. Position 109 or the corresponding position may be substituted with A, V, I, L, M, F, Y or W. Position 109 or the corresponding position is preferably substituted with A or V. Position 136 or the corresponding position may be substituted with A, V, I, L, M, F, Y or W. Position 136 or the corresponding position is preferably substituted with A or V. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises a
substitution at C109, such as C109A, C109V, C109I, C109L, C109M, C109F, C109Y or C109W and/or at C136, such as C136A, C136V, C136I, C136L, C136T, C136M, C136F, C136Y or C136W. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises C109A and/or C136A. The helicase of the invention preferably comprises a variant of SEQ ID NO: 119, 120, 121, 122, 123, 124, 125, 126, 127, 128, 129, 130, 131, 132 or 133 which further comprises a substitution at the position(s) which correspond(s) to C109 and/or C136 in SEQ ID NO: 118. The helicase of the invention is preferably one in which at least one cysteine residue (i.e. one or more cysteine residues) and/or at least one non-natural amino acid (i.e. one or more non-natural amino acids) have further been introduced into the tower domain only. Suitable modifications are discussed above.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising the following mutations: E93C and K364C; E94C and K364C; E94C and A360C; L97C and E361C; L97C and E361C and C412A; K123C and E361C; K123C, E361C and C412A; N155C and K358C; N155C, K358C and C412A; N155C and L354C; N155C, L354C and C412A; deltaE93, E94C, deltaN95 and A360C; E94C, deltaN95 and A360C; E94C, Q100C, I127C and A360C; L354C; G357C; E94C, G357C and A360C; E94C, Y279C and A360C; E94C, I281C and A360C; E94C, Y279Faz and A360C; Y279C and G357C; I281C and G357C; E94C, Y279C, G357C and A360C; E94C, I281C, G357C and A360C; E8R, E47K, E94C, D202K and A360C; D5K, E23N, E94C, D167K, E172R, D212R and A360C; D5K, E8R, E23N, E47K, E94C, D167K, E172R, D202K, D212R and A360C; E94C, C114A, C171A, A360C and C412D; E94C, C114A, C171A, A360C and C412S; E94C, C109A, C136A and A360C; E94C, C109A, C114A, C136A, C171A, A360C and C412S; E94C, C109V, C114V, C171A, A360C and C412S; C109A, C114A, C136A, G153C, C171A, E361C and C412A; C109A, C114A, C136A, G153C, C171A, E361C and C412D; C109A, C114A, C136A, G153C, C171A, E361C and C412S; C109A, C114A, C136A, G153C, C171A, K358C and C412A; C109A, C114A, C136A, G153C, C171A, K358C and C412D; C109A, C114A, C136A, G153C, C171A, K358C and C412S; C109A, C114A, C136A, N155C, C171A, K358C and C412A; C109A, C114A, C136A, N155C, C171A, K358C and C412D; C109A, C114A, C136A, N155C, C171A, K358C and C412S; C109A, C114A, C136A, N155C, C171A, L354C and C412A; C109A, C114A, C136A, N155C, C171A, L354C and C412D; C109A, C114A, C136A, N155C, C171A, L354C and C412S; C109A, C114A, K123C, C136A, C171A, E361C and C412A; C109A, C114A, K123C, C136A, C171A, E361C and C412D; C109A, C114A, K123C, C136A, C171A, E361C and C412S; C109A, C114A, K123C, C136A, C171A, K358C and C412A; C109A, C114A, K123C, C136A, C171A, K358C and C412D; C109A, C114A, K123C, C136A, C171A, K358C and C412S; C109A, C114A, C136A, G153C, C171A, E361C and C412A; E94C, C109A, C114A, C136A, C171A, A360C and C412D; E94C, C109A, C114V, C136A, C171A, A360C and C412D; E94C, C109V, C114A, C136A, C171A, A360C and C412D; L97C, C109A,
C114A, C136A, C171A, E361C and C412A; L97C, C109A, C114A, C136A, C171A, E361C and C412D; or L97C, C109A, C114A, C136A, C171A, E361C and C412S.
Modifications in the hook domain and/or 2A domain
In one embodiment, the helicase of the invention is one in which at least one cysteine residue and/or at least one non-natural amino acid have been further introduced into the hook domain and/or the 2A (RecA-like motor) domain, wherein the helicase has the ability to control the movement of a polynucleotide. At least one cysteine residue and/or at least one non-natural amino acid is preferably introduced into the hook domain and the 2A (RecA-like motor) domain.
Any number of cysteine residues and/or non-natural amino acids may be introduced into each domain. For instance, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more cysteine residues may be introduced and/or 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more non-natural amino acids may be introduced. Only one or more cysteine residues may be introduced. Only one or more non- natural amino acids may be introduced. A combination of one or more cysteine residues and one or more non-natural amino acids may be introduced.
The at least one cysteine residue and/or at least one non-natural amino acid are preferably introduced by substitution. Methods for doing this are known in the art. Suitable modifications of the hook domain and/or the 2A (RecA-like motor) domain are discussed above.
The helicase of the invention is preferably a variant of SEQ ID NO: 118 further comprising (a) Y279C, I181C, E288C, Y279C and I181C, (b) Y279C and E288C, (c) I181C and E288C or (d) Y279C, I181C and E288C. The helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 199 to 133 which further comprises a mutation at one or more of the position(s) which correspond to those in SEQ ID NO: 118 as defined in (a) to (d).
Surface modification
In one embodiment, the helicase is modified to reduce its surface negative charge, wherein the helicase has the ability to control the movement of a polynucleotide. Suitable modifications are discussed above. Any number of surface negative charges may be neutralised.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising the following mutations: E273G; E8R, E47K and D202K; D5K, E23N, D167K, E172R and D212R; or D5K, E8R, E23N, E47K, D167K, E172R, D202K and D212R.
Other modified helicases
In one embodiment, the helicase of the invention comprises a variant of SEQ ID NO: 118 further comprising : A360K; Y92L and/or A360Y; Y92L, Y350N and Y363N; Y92L and/or Y363N; or Y92L.
Other modifications
In addition to the specific mutations disclosed above, a variant of SEQ ID NO: 118 may comprise one or more of the following mutations: K38A; T91F; T91N; T91Q; T91W; V96E; V96F; V96L; V96Q; V96R; V96W; V96Y; P274G; V286F; V286W; V286Y; F291G; N292F; N292G; N292P; N292Y; G294Y; G294F; K364A; and W378A.
In addition to the specific mutations disclosed above, a variant of SEQ ID NO: 118 may comprise: K38A, E94C and A360C; H64K; E94C and A360C; H64N; E94C and A360C;
H64Q; E94C and A360C; H64S; E94C and A360C; H64W, E94C and A360C; T80K, E94C and A360C; T80K, S83K, E94C, N242K, N293K and A360C; T80K, S83K, E94C, N242K, N293K, A360C and T394K; T80K, S83K, E94C, N293K and A360C; T80K, S83K, E94C, A360C and T394K; T80K, S83K, E94C, A360C and T394N; T80K, E94C, N242K and A360C; T80K, E94C, N242K, N293K and A360C; T80K, E94C, N293K and A360C; T80N, E94C and A360C; H82A, E94C and A360C; H82A, P89A, E94C, F98A and A360C; H82F, E94C and A360C;
H82Q, E94C, A360C; H82R, E94C and A360C; H82W, E94C and A360C; H82W, P89W, E94C, F98W and A360C; H82Y, E94C and A360C; S83K, E94C and A360C; S83K, T80K, E94C, A360C and T394K; S83N, E94C and A360C; S83T, E94C and A360C; N88H, E94C and A360C; N88Q, E94C and A360C; P89A, E94C and A360C; P89A, F98W, E94C and A360C;
P89A, E94C, F98Y and A360C; P89A, E94C, F98A and A360C; P89F, E94C and A360C;
P89S, E94C and A360C; P89T, E94C and A360C; P89W, E94C, F98W and A360C; P89Y, E94C and A360C; T91F, E94C and A360C; T91N, E94C and A360C; T91Q, E94C and A360C; T91W, E94C and A360C; E94C, V96E and A360C; E94C, V96F and A360C; E94C, V96L and A360C; E94C, V96Q and A360C; E94C, V96R and A360C; E94C, V96W and A360C; E94C, V96Y and A360C; E94C, F98A and A360C; E94C, F98L and A360C; E94C, F98V and A360C; E94C, F98Y and A360C; E94C; F98W and A360C; E94C, V150A and A360C; E94C, V150F and A360C; E94C, V150I and A360C; E94C, V150K and A360C; E94C, V150L and A360C;
E94C, V150S and A360C; E94C, V150T and A360C; E94C, V150W and A360C; E94C, V150Y and A360C; E94C, F240Y and A360C; E94C, F240W and A360C; E94C, N242K and A360C; E94C, N242K, N293K and A360C; E94C, P274G and A360C; E94C, L275G and A360C;
E94C, F276A and A360C; E94C, F276I and A360C; E94C, F276M and A360C; E94C, F276V and A360C; E94C, F276W and A360C; E94C, F276Y and A360C; E94C, V286F and A360C; E94C, V286W and A360C; E94C, V286Y and A360C; E94C, S287F and A360C; E94C, S287W and A360C; E94C, S287Y and A360C; E94C, F291G and A360C; E94C, N292F and A360C;
E94C, N292G and A360C; E94C, N292P and A360C; E94C, N292Y and A360C; E94C, N293F and A360C; E94C, N293K and A360C; E94C, N293Q and A360C; E94C, N293Y and A360C; E94C, G294F and A360C; E94C, G294Y and A360C; E94C, A36C and K364A; E94C, A360C, W378A; E94C, A360C and T394K; E94C, A360C and H396Q; E94C, A360C and H396S;
E94C, A360C and H396W; E94C, A360C and Y415F; E94C, A360C and Y415K; E94C, A360C and Y415M; or E94C, A360C and Y415W.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 which further comprises (a) E94C/A360C/W378A or (b) E94C/A360C/C109A/C136A/W378A or (d) E94C/A360C/C109A/C136A/W378A and then (AM1)G1G2 (i.e. deletion of Ml and then addition G1 and G2).
Preferred variants of any one of SEQ ID NOs: 118 to 133 have (in addition to the modifications of the invention) the N-terminal methionine (M) replaced with one glycine residue (G). In the examples this is shown as (AMl)Gl. It may also be termed MIG. Any of the variants discussed above may further comprise MIG.
The most preferred helicases of the invention comprise a variant of SEQ ID NO: 118 which further (a) E94C/F98W/A360C/C109A/C136A/K194L, (b) M1G/E94C/F98W/A360C/C109A/C136A/K194L; (c) E94C/F98W/A360C/C109A/C136A/K199L; or (d) M1G/E94C/F98W/A360C/C109A/C136A/K199L.
Additional modifications/substitutions
The helicase of the invention may further be one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: 86, 90, 92, 97, 101, 102, 273, 293, 300, 301, 303, 305, 308, 310, 312 317, 323, 328, 332, 334, 335, 336, 337, 339, 351, 354, 359, 361, 364, 366, 368, 371, 374, 376, 377, 379 and 388. The helicase of the invention may further be one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: 351, 354 and 361. Any number and combination of these modifications/substitutions may be made, including at 351, 354, 361, 351 and 354, 351 and 361, 354 and 361 or 351, 354 and 361. These positions may be modified or substituted in combination with any of the modifications or substitutions of the invention above.
The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising one or more of the following substitutions: K86A, V90T , Y92A, Y92D, Y92F, Y92G, Y92H, Y92N, Y92Q, Y92S, Y92T, Y92V, Y92W, L97H, K101A, E102G, E273A, N293H, I200A, I300F, E301I, E303I, Y305L, F308I, F308L, K310A, K310I, K310L, R312I, R312L,
R312M, E317I, E317Y, W323H, E328I, D332A, D332L, E334A, E334I, E334Y, Y335A, Y335I, Y335L, Y336L, R337I, R337L, R337M, K339A, K339I, K339L, K351I, K351Q, L354Q, L354A, T359L, E361T, E361I, E361Q, K364A, K364R, W366L, K368A, K368I, K368L, K371I, K371L, K371M, W374A, W374L, W374L, D376S, F377A, D379A, D379L or K388R. The helicase of the invention preferably comprises a variant of SEQ ID NO: 118 further comprising one or more of the following substitutions: (a) K351I or K351Q, (b) L354A or L354Q and (c) E361I or E361Q. The variant may comprise (a), (b), (c), (a) and (b), (a) and (c), (b) and (c) or (a), (b) and (c). These substitutions may be made in combination with any of the modifications or substitutions of the invention above.
Variants
A variant of a helicase is an enzyme that has an amino acid sequence which varies from that of the wild-type helicase and which has polynucleotide binding activity. In particular, a variant of any one of SEQ ID NOs: 118 to 133 is an enzyme that has an amino acid sequence which varies from that of any one of SEQ ID NOs: 118 to 133 and which has polynucleotide binding activity. Polynucleotide binding activity can be determined using methods known in the art. Suitable methods include, but are not limited to, fluorescence anisotropy, tryptophan fluorescence and electrophoretic mobility shift assay (EMSA). For instance, the ability of a variant to bind a single stranded polynucleotide can be determined as described in the Examples.
The variant has helicase activity. This can be measured in various ways. For instance, the ability of the variant to translocate along a polynucleotide can be measured using electrophysiology, a fluorescence assay or ATP hydrolysis.
The variant may include modifications that facilitate handling of the polynucleotide encoding the helicase and/or facilitate its activity at high salt concentrations and/or room temperature.
Over the entire length of the amino acid sequence of any one of SEQ ID NOs: 118 to 133, a variant will preferably be at least 20% homologous to that sequence based on amino acid similarity or identity. More preferably, the variant polypeptide may be at least 30%, at least 40%, at least 45%, at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90% and more preferably at least 95%, 97% or 99% homologous based on amino acid identity to the amino acid sequence of any one of SEQ ID NOs: 118 to 133 over the entire sequence. More preferably, the variant polypeptide may be at least 30%, at least 40%, at least 45%, at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90% and more preferably at least 95%, 97% or 99% identical to the amino acid sequence of any one of SEQ ID NOs: 118 to 133 over the entire sequence. There may be at least
70%, for example at least 80%, at least 85%, at least 90% or at least 95%, amino acid identity over a stretch of 100 or more, for example 150, 200, 300, 400 or 500 or more, contiguous amino acids ("hard homology"). In particular, in addition to the specific modifications discussed above, the variant of any one of SEQ ID NOs: 118 to 133 may comprise one or more substitutions, one or more deletions and/or one or more additions as discussed below.
Standard methods in the art may be used to determine homology. For example the UWGCG Package provides the BESTFIT program which can be used to calculate homology, for example used on its default settings (Devereux et al (1984) Nucleic Acids Research 12, p387-395). The PILEUP and BLAST algorithms can be used to calculate homology or line up sequences (such as identifying equivalent residues or corresponding sequences (typically on their default settings)), for example as described in Altschul S. F. (1993) J Mol Evol 36:290- 300; Altschul, S.F et al (1990) J Mol Biol 215:403-10. Software for performing BLAST analyses is publicly available through the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/).
Preferred variants of any one of SEQ ID NOs: 118 to 133 have a non-natural amino acid, such as Faz, at the amino- (N-) terminus and/or carboxy (C-) terminus. Preferred variants of any one of SEQ ID NOs: 118 to 133 have a cysteine residue at the amino- (N-) terminus and/or carboxy (C-) terminus. Preferred variants of any one of SEQ ID NOs: 118 to 133 have a cysteine residue at the amino- (N-) terminus and a non-natural amino acid, such as Faz, at the carboxy (C-) terminus or vice versa.
Preferred variants of SEQ ID NO: 118 contain one or more of, such as all of, the following modifications E54G, D151E, I196N and G357A.
No connection
In one preferred embodiment, none of the introduced cysteines and/or non-natural amino acids in a modified helicase of the invention are connected to one another.
Connecting two more of the introduced cysteines and/or non-natural amino acids
In another preferred embodiment, two more of the introduced cysteines and/or non-natural amino acids in a modified helicase of the invention are connected to one another. This typically reduces the ability of the helicase of the invention to unbind from a polynucleotide.
Any number and combination of two more of the introduced cysteines and/or non-natural amino acids may be connected to one another. For instance, 3, 4, 5, 6, 7, 8 or more cysteines and/or non-natural amino acids may be connected to one another. One or more cysteines may be connected to one or more cysteines. One or more cysteines may be
connected to one or more non-natural amino acids, such as Faz. One or more non-natural amino acids, such as Faz, may be connected to one or more non-natural amino acids, such as Faz.
The two or more cysteines and/or non-natural amino acids may be connected in any way. The connection can be transient, for example non-covalent. Even transient connection will reduce unbinding of the polynucleotide from the helicase.
The two or more cysteines and/or non-natural amino acids are preferably connected by affinity molecules. Suitable affinity molecules are known in the art. The affinity molecules are preferably (a) complementary polynucleotides (WO 2010/086602 incorporated herein by reference in its entirety), (b) an antibody or a fragment thereof and the complementary epitope (Biochemistry 6thEd, W.H. Freeman and co (2007) pp953-954), (c) peptide zippers (O'Shea et al., Science 254 (5031): 539-544), (d) capable of interacting by -sheet augmentation (Remaut and Waksman Trends Biochem. Sci. (2006) 31 436-444), (e) capable of hydrogen bonding, pi-stacking or forming a salt bridge, (f) rotaxanes (Xiang Ma and He Tian Chem. Soc. Rev., 2010,39, 70-80), (g) an aptamer and the complementary protein (James, W. in Encyclopedia of Analytical Chemistry, R.A. Meyers (Ed.) pp. 4848- 4871 John Wiley & Sons Ltd, Chichester, 2000) or (h) half-chelators (Hammerstein et al. J Biol Chem. 2011 April 22; 286(16): 14324-14334). For (e), hydrogen bonding occurs between a proton bound to an electronegative atom and another electronegative atom. Pi- stacking requires two aromatic rings that can stack together where the planes of the rings are parallel. Salt bridges are between groups that can delocalize their electrons over several atoms, e. g. between aspartate and arginine.
The two or more parts may be transiently connected by a hexa-his tag or Ni-NTA.
The two or more cysteines and/or non-natural amino acids are preferably permanently connected. In the context of the invention, a connection is permanent if is not broken while the helicase is used or cannot be broken without intervention on the part of the user, such as using reduction to open -S-S- bonds.
The two or more cysteines and/or non-natural amino acids are preferably covalently- attached. The two or more cysteines and/or non-natural amino acids may be covalently attached using any method known in the art.
The two or more cysteines and/or non-natural amino acids may be covalently attached via their naturally occurring amino acids, such as cysteines, threonines, serines, aspartates, asparagines, glutamates and glutamines. Naturally occurring amino acids may be modified to facilitate attachment. For instance, the naturally occurring amino acids may be modified by acylation, phosphorylation, glycosylation or farnesylation. Other suitable modifications
are known in the art. Modifications to naturally occurring amino acids may be posttranslation modifications. The two or more cysteines and/or non-natural amino acids may be attached via amino acids that have been introduced into their sequences. Such amino acids are preferably introduced by substitution. The introduced amino acid may be cysteine or a non-natural amino acid that facilitates attachment. Suitable non-natural amino acids include, but are not limited to, 4-azido-L-phenylalanine (Faz), any one of the amino acids numbered 1-71 included in figure 1 of Liu C. C. and Schultz P. G., Annu. Rev. Biochem., 2010, 79, 413-444 or any one of the amino acids listed below. The introduced amino acids may be modified as discussed above.
In a preferred embodiment, the two or more cysteines and/or non-natural amino acids are connected using linkers. Linker molecules are discussed in more detail below. One suitable method of connection is cysteine linkage. This is discussed in more detail below. The two or more cysteines and/or non-natural amino acids are preferably connected using one or more, such as two or three, linkers. The one or more linkers may be designed to reduce the size of, or close, the opening as discussed above. If one or more linkers are being used to close the opening as discussed above, at least a part of the one or more linkers is preferably oriented such that it is not parallel to the polynucleotide when it is bound by the helicase. More preferably, all of the linkers are oriented in this manner. If one or more linkers are being used to close the opening as discussed above, at least a part of the one or more linkers preferably crosses the opening in an orientation that is not parallel to the polynucleotide when it bound by the helicase. More preferably, all of the linkers cross the opening in this manner. In these embodiments, at least a part of the one or more linkers may be perpendicular to the polynucleotide. Such orientations effectively close the opening such that the polynucleotide cannot unbind from the helicase through the opening.
Each linker may have two or more functional ends, such as two, three or four functional ends. Suitable configurations of ends in linkers are well known in the art.
One or more ends of the one or more linkers are preferably covalently attached to the helicase. If one end is covalently attached, the one or more linkers may transiently connect the two or more cysteines and/or non-natural amino acids as discussed above. If both or all ends are covalently attached, the one or more linkers permanently connect the two or more cysteines and/or non-natural amino acids.
The one or more linkers are preferably amino acid sequences and/or chemical crosslinkers.
Suitable amino acid linkers, such as peptide linkers, are known in the art. The length, flexibility and hydrophilicity of the amino acid or peptide linker are typically designed such that it reduces the size of the opening, but does not to disturb the functions of the helicase. Preferred flexible peptide linkers are stretches of 2 to 20, such as 4, 6, 8, 10 or 16, serine
and/or glycine amino acids. More preferred flexible linkers include (SG)i, (SG)2, (SG)s, (SG)4, (SG)5, (SG)8, (SG) IO, (SG)is or (SG)2o wherein S is serine and G is glycine. Preferred rigid linkers are stretches of 2 to 30, such as 4, 6, 8, 16 or 24, proline amino acids. More preferred rigid linkers include (P)i2 wherein P is proline. The amino acid sequence of a linker preferably comprises a polynucleotide binding moiety. Such moieties and the advantages associated with their use are discussed below.
Suitable chemical crosslinkers are well-known in the art. Suitable chemical crosslinkers include, but are not limited to, those including the following functional groups: maleimide, active esters, succinimide, azide, alkyne (such as dibenzocyclooctynol (DIBO or DBCO), difluoro cycloalkynes and linear alkynes), phosphine (such as those used in traceless and non-traceless Staudinger ligations), haloacetyl (such as iodoacetamide), phosgene type reagents, sulfonyl chloride reagents, isothiocyanates, acyl halides, hydrazines, disulphides, vinyl sulfones, aziridines and photoreactive reagents (such as aryl azides, diaziridines).
Reactions between amino acids and functional groups may be spontaneous, such as cysteine/maleimide, or may require external reagents, such as Cu(I) for linking azide and linear alkynes.
Linkers can comprise any molecule that stretches across the distance required. Linkers can vary in length from one carbon (phosgene-type linkers) to many Angstroms. Examples of linear molecules, include but are not limited to, are polyethyleneglycols (PEGs), polypeptides, polysaccharides, deoxyribonucleic acid (DNA), peptide nucleic acid (PNA), threose nucleic acid (TNA), glycerol nucleic acid (GNA), saturated and unsaturated hydrocarbons, polyamides. These linkers may be inert or reactive, in particular they may be chemically cleavable at a defined position, or may be themselves modified with a fluorophore or ligand. The linker is preferably resistant to dithiothreitol (DTT).
Preferred crosslinkers include 2,5-dioxopyrrolidin-l-yl 3-(pyridin-2-yldisulfanyl)propanoate, 2,5-dioxopyrrolidin-l-yl 4-(pyridin-2-yldisulfanyl)butanoate and 2,5-dioxopyrrolidin-l-yl 8- (pyridin-2-yldisulfanyl)octananoate, di-maleimide PEG lk, di-maleimide PEG 3.4k, di- maleimide PEG 5k, di-maleimide PEG 10k, bis(maleimido)ethane (BMOE), bis- maleimidohexane (BMH), 1,4-bis-maleimidobutane (BMB), 1,4 bis-maleimidyl-2,3- dihydroxybutane (BMDB), BM[PEO]2 (1,8-bis-maleimidodiethyleneglycol), BM[PEO]3 (1,11- bis-maleimidotriethylene glycol), tris[2-maleimidoethyl]amine (TMEA), DTME dithiobismaleimidoethane, bis-maleimide PEG3, bis-maleimide PEGU, DBCO-maleimide, DBCO-PEG4-maleimide, DBCO-PEG4-NH2, DBCO-PEG4-NHS, DBCO-NHS, DBCO-PEG-DBCO 2.8kDa, DBCO-PEG-DBCO 4.0kDa, DBCO-15 atoms-DBCO, DBCO-26 atoms-DBCO, DBCO- 35 atoms-DBCO, DBCO-PEG4-S-S-PEG3-biotin, DBCO-S-S-PEG3-biotin, DBCO-S-S-PEG11- biotin, (succinimidyl 3-(2-pyridyldithio)propionate (SPDP) and maleimide-PEG(2kDa)-
maleimide (ALPHA, OMEGA-BIS-MALEIMIDO POLYETHYLENE GLYCOL)). The most preferred crosslinker is maleimide-propyl-SRDFWRS-(l,2-diaminoethane)-propyl-maleimide.
The one or more linkers may be cleavable. This is discussed in more detail below.
The two or more cysteines and/or non-natural amino acids may be connected using two different linkers that are specific for each other. One of the linkers is attached to one part and the other is attached to another part. The linkers should react to form a modified helicase of the invention. The two or more cysteines and/or non-natural amino acids may be connected using the hybridization linkers described in WO 2010/086602 (incorporated herein by reference in its entirety). In particular, the two or more cysteines and/or non- natural amino acids may be connected using two or more linkers each comprising a hybridizable region and a group capable of forming a covalent bond. The hybridizable regions in the linkers hybridize and link the two or more cysteines and/or non-natural amino acids. The linked cysteines and/or non-natural amino acids are then coupled via the formation of covalent bonds between the groups. Any of the specific linkers disclosed in WO 2010/086602 (incorporated herein by reference in its entirety) may be used in accordance with the invention.
The two or more cysteines and/or non-natural amino acids may be modified and then attached using a chemical crosslinker that is specific for the two modifications. Any of the crosslinkers discussed above may be used.
The linkers may be labeled. Suitable labels include, but are not limited to, fluorescent molecules (such as Cy3 or AlexaFluor®555), radioisotopes, e.g. 125I, 35S, enzymes, antibodies, antigens, polynucleotides and ligands such as biotin. Such labels allow the amount of linker to be quantified. The label could also be a cleavable purification tag, such as biotin, or a specific sequence to show up in an identification method, such as a peptide that is not present in the protein itself, but that is released by trypsin digestion.
A preferred method of connecting two or more cysteines is via cysteine linkage. This can be mediated by a bi-functional chemical crosslinker or by an amino acid linker with a terminal presented cysteine residue.
The length, reactivity, specificity, rigidity and solubility of any bi-functional linker may be designed to ensure that the size of the opening is reduced sufficiently and the function of the helicase is retained. Suitable linkers include bismaleimide crosslinkers, such as 1,4- bis(maleimido)butane (BMB) or bis(maleimido)hexane. One drawback of bi-functional linkers is the requirement of the helicase to contain no further surface accessible cysteine residues if attachment at specific sites is preferred, as binding of the bi-functional linker to surface accessible cysteine residues may be difficult to control and may affect substrate
binding or activity. If the helicase does contain several accessible cysteine residues, modification of the helicase may be required to remove them while ensuring the modifications do not affect the folding or activity of the helicase. This is discussed in WO 2010/086603 (incorporated herein by reference in its entirety). The reactivity of cysteine residues may be enhanced by modification of the adjacent residues, for example on a peptide linker. For instance, the basic groups of flanking arginine, histidine or lysine residues will change the pKa of the cysteines thiol group to that of the more reactive S’ group. The reactivity of cysteine residues may be protected by thiol protective groups such as 5,5'-dithiobis-(2-nitrobenzoic acid) (dTNB). These may be reacted with one or more cysteine residues of the helicase before a linker is attached. Selective deprotection of surface accessible cysteines may be possible using reducing reagents immobilized on beads (for example immobilized tris(2-carboxyethyl) phosphine, TCEP). Cysteine linkage is discussed in more detail below.
Another preferred method of attachment via Faz linkage. This can be mediated by a bifunctional chemical linker or by a polypeptide linker with a terminal presented Faz residue.
Other modifications
The helicase of the invention may also be modified to increase the attraction between (i) the tower domain and (ii) the pin domain and/or the 1A domain. Any known chemical modifications can be made in accordance with the invention. These types of modification are disclosed in WO 2015/055981 (incorporated herein by reference in its entirety).
In particular, the invention provides a helicase of the invention in which at least one charged amino acid has further been introduced into (i) the tower domain and/or (ii) the pin domain and/or (iii) the 1A (RecA-like motor) domain, wherein the helicase has the ability to control the movement of a polynucleotide. The ability of the helicase to control the movement of a polynucleotide may be measured as discussed above. The invention preferably provides a helicase of the invention in which at least one charged amino acid has further been introduced into (i) the tower domain and (ii) the pin domain and/or the 1A domain.
The at least one charged amino acid may be negatively charged or positively charged. The at least one charged amino acid is preferably oppositely charged to any amino acid(s) with which it interacts in the helicase. For instance, at least one positively charged amino acid may be introduced into the tower domain at a position which interacts with a negatively charged amino acid in the pin domain. The at least one charged amino acid is typically introduced at a position which is not charged in the wild-type (i.e. unmodified) helicase. The at least one charged amino acid may be used to replace at least one oppositely charged
amino acid in the helicase. For instance, a positively charged amino acid may be used to replace a negatively charged amino acid.
Suitable charged amino acids are discussed above. The at least one charged amino acid may be natural, such as arginine (R), histidine (H), lysine (K), aspartic acid (D) or glutamic acid (D). Alternatively, the at least one charged amino acid may be artificial or non-natural. Any number of charged amino acids may be introduced into each domain. For instance, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more charged amino acids may be introduced into each domain.
The helicase preferably comprises a variant of SEQ ID NO: 118 which comprises a positively charged amino acid at one or more of the following positions: (i) 93; (ii) 354; (iii) 360; (iv) 361; (v) 94; (vi) 97; (vii) 155; (viii) 357; (ix) 100; and (x) 127. The helicase preferably comprises a variant of SEQ ID NO: 118 which comprises a negatively charged amino acid at one or more of the following positions: (i) 354; (ii) 358; (iii) 360; (iv) 364; (v) 97; (vi) 123; (vii) 155; (viii); 357; (ix) 100; and (x) 127. The helicase preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which comprises a positively charged amino acid or negatively charged amino acid at the positions which correspond to those in SEQ ID NO: 118 as defined in any of (i) to (x). Positions in any one of SEQ ID NOs: 119 to 133 which correspond to those in SEQ ID NO: 118 can be identified using the alignment of SEQ ID NOs: 118 to 133 below.
The helicase preferably comprises a variant of SEQ ID NO: 118 which is modified by the introduction of at least one charged amino acid such that it comprises oppositely charged amino acid at the following positions: (i) 93 and 354; (ii) 93 and 358; (iii) 93 and 360; (iv) 93 and 361; (v) 93 and 364; (vi) 94 and 354; (vii) 94 and 358; (viii) 94 and 360; (ix) 94 and 361; (x) 94 and 364; (xi) 97 and 354; (xii) 97 and 358; (xiii) 97 and 360; (xiv) 97 and 361; (xv) 97 and 364; (xvi) 123 and 354; (xvii) 123 and 358; (xviii) 123 and 360; (xix) 123 and 361; (xx) 123 and 364; (xxi) 155 and 354; (xxii) 155 and 358; (xxiii) 155 and 360; (xxiv) 155 and 361; (xxv) 155 and 364. The helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises oppositely charged amino acids at the positions which correspond to those in SEQ ID NO: 118 as defined in any of (i) to (xxv).
The helicase of the invention may further be one in which (i) at least one charged amino acid has been introduced into the tower domain and (ii) at least one oppositely charged amino acid has been introduced into the pin domain and/or the 1A (RecA-like motor) domain, wherein the helicase has the ability to control the movement of a polynucleotide. The at least one charged amino acid may be negatively charged and the at least one oppositely charged amino acid may be positively charged or vice versa. Suitable charged amino acids are discussed above. Any number of charged amino acids and any number of
oppositely charged amino acids may be introduced. For instance, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more charged amino acids may be introduced and/or 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 or more oppositely charged amino acids may be introduced.
The charged amino acids are typically introduced at positions which are not charged in the wild-type helicase. One or both of the charged amino acids may be used to replace charged amino acids in the helicase. For instance, a positively charged amino acid may be used to replace a negatively charged amino acid. The charged amino acids may be introduced at any of the positions in the (i) tower domain and (ii) pin domain and/or 1A domain discussed above. The oppositely charged amino acids are typically introduced such that they will interact in the resulting helicase. The helicase preferably comprises a variant of SEQ ID NO: 118 in which oppositely charged amino acids have been introduced at the following positions: (i) 97 and 354; (ii) 97 and 360; (iii) 155 and 354; or (iv) 155 and 360. The helicase of the invention preferably comprises a variant of any one of SEQ ID NOs: 119 to 133 which further comprises oppositely charged amino acids at the positions which correspond to those in SEQ ID NO: 118 as defined in any of (i) to (iv).
Construct
The invention also provides a construct comprising a modified helicase of the invention and an additional polynucleotide binding moiety, wherein the helicase is attached to the polynucleotide binding moiety and the construct has the ability to control the movement of a polynucleotide. The construct is artificial or non-natural.
A construct of the invention is a useful tool for controlling the movement of a polynucleotide during Strand Sequencing. A construct of the invention is even less likely than a modified helicase of the invention to disengage from the polynucleotide being sequenced. The construct can provide even greater read lengths of the polynucleotide as it controls the translocation of the polynucleotide through a nanopore.
A targeted construct that binds to a specific polynucleotide sequence can also be designed. As discussed in more detail below, the polynucleotide binding moiety may bind to a specific polynucleotide sequence and thereby target the helicase portion of the construct to the specific sequence.
The construct has the ability to control the movement of a polynucleotide. This can be determined as discussed above.
A construct of the invention may be isolated, substantially isolated, purified or substantially purified. A construct is isolated or purified if it is completely free of any other components, such as lipids, polynucleotides or pore monomers. A construct is substantially isolated if it is mixed with carriers or diluents which will not interfere with its intended use. For instance, a
construct is substantially isolated or substantially purified if it is present in a form that comprises less than 10%, less than 5%, less than 2% or less than 1% of other components, such as lipids, polynucleotides or pore monomers.
The helicase may be any of the helicases of the invention discussed above.
The helicase is preferably covalently attached to the additional polynucleotide binding moiety. The helicase may be attached to the moiety at more than one, such as two or three, points.
The helicase can be covalently attached to the moiety using any method known in the art. Suitable methods are discussed above with reference to connecting the two or more parts.
The helicase and moiety may be produced separately and then attached together. The two components may be attached in any configuration. For instance, they may be attached via their terminal (i.e. amino or carboxy terminal) amino acids. Suitable configurations include, but are not limited to, the amino terminus of the moiety being attached to the carboxy terminus of the helicase and vice versa. Alternatively, the two components may be attached via amino acids within their sequences. For instance, the moiety may be attached to one or more amino acids in a loop region of the helicase. In a preferred embodiment, terminal amino acids of the moiety are attached to one or more amino acids in the loop region of a helicase.
In a preferred embodiment, the helicase is chemically attached to the moiety, for instance via one or more linker molecules as discussed above. In another preferred embodiment, the helicase is genetically fused to the moiety. A helicase is genetically fused to a moiety if the whole construct is expressed from a single polynucleotide sequence. The coding sequences of the helicase and moiety may be combined in any way to form a single polynucleotide sequence encoding the construct. Genetic fusion of a pore to a nucleic acid binding protein is discussed in WO 2010/004265 (incorporated herein by reference in its entirety).
The helicase and moiety may be genetically fused in any configuration. The helicase and moiety may be fused via their terminal amino acids. For instance, the amino terminus of the moiety may be fused to the carboxy terminus of the helicase and vice versa. The amino acid sequence of the moiety is preferably added in frame into the amino acid sequence of the helicase. In other words, the moiety is preferably inserted within the sequence of the helicase. In such embodiments, the helicase and moiety are typically attached at two points, i.e. via the amino and carboxy terminal amino acids of the moiety. If the moiety is inserted within the sequence of the helicase, it is preferred that the amino and carboxy terminal amino acids of the moiety are in close proximity and are each attached to adjacent
amino acids in the sequence of the helicase or variant thereof. In a preferred embodiment, the moiety is inserted into a loop region of the helicase.
The helicase may be attached directly to the moiety. The helicase is preferably attached to the moiety using one or more, such as two or three, linkers as discussed above. The one or more linkers may be designed to constrain the mobility of the moiety. The helicase and/or the moiety may be modified to facilitate attachment of the one or more linker as discussed above.
Cleavable linkers can be used as an aid to separation of constructs from non-attached components and can be used to further control the synthesis reaction. For example, a hetero-bifunctional linker may react with the helicase, but not the moiety. If the free end of the linker can be used to bind the helicase protein to a surface, the unreacted helicases from the first reaction can be removed from the mixture. Subsequently, the linker can be cleaved to expose a group that reacts with the moiety. In addition, by following this sequence of linkage reactions, conditions may be optimised first for the reaction to the helicase, then for the reaction to the moiety after cleavage of the linker. The second reaction would also be much more directed towards the correct site of reaction with the moiety because the linker would be confined to the region to which it is already attached.
The helicase may be covalently attached to the bifunctional crosslinker before the helicase/crosslinker complex is covalently attached to the moiety. Alternatively, the moiety may be covalently attached to the bifunctional crosslinker before the bifunctional crosslinker/moiety complex is attached to the helicase. The helicase and moiety may be covalently attached to the chemical crosslinker at the same time.
Preferred methods of attaching the helicase to the moiety are cysteine linkage and Faz linkage as described above. In a preferred embodiment, a reactive cysteine is presented on a peptide linker that is genetically attached to the moiety. This means that additional modifications will not necessarily be needed to remove other accessible cysteine residues from the moiety.
Cross-linkage of helicases or moieties to themselves may be prevented by keeping the concentration of linker in a vast excess of the helicase and/or moiety. Alternatively, a "lock and key" arrangement may be used in which two linkers are used. Only one end of each linker may react together to form a longer linker and the other ends of the linker each react with a different part of the construct (i.e. helicase or moiety). This is discussed in more detail below.
The site of attachment is selected such that, when the construct is contacted with a polynucleotide, both the helicase and the moiety can bind to the polynucleotide and control its movement.
The invention also provides a method of producing a construct of the invention, comprising attaching a helicase of the invention to an additional polynucleotide binding moiety and thereby producing the construct. The method preferably further comprises determining whether or not the resulting construct is capable of controlling the movement of a polynucleotide. All of the construct embodiments discussed above equally apply to the method.
Pores and pore complexes for use in the invention
The invention provides various methods and products in which a modified Dda helicase or a construct of the invention is combined/used with a transmembrane pore. The transmembrane pore may be used a solid-state pore. The transmembrane pore may be a transmembrane protein pore. The transmembrane protein pore may be derived from a hemolysin, such as alpha hemolysin or gamma hemolysin, leukocidin, Mycobacterium smegmatis porin A (MspA), MspB, MspC, MspD, PorARr, PorBRr, PorARc, lysenin, outer membrane porin F (OmpF), outer membrane porin G (OmpG), outer membrane phospholipase A, Neisseria autotransporter lipoprotein (NalP) and WZA, CsgG, CsgG/CsgF, ClyA, Spl, haemolytic protein fragaceatoxin C (FraC), Iota toxin, Anthrax protective antigen, Vibrio cholerae cytolysin, Cytotoxin K (CytK), CELIII, Aerolysin, InvG, GspD, PilQ, necrotic enteritis B-like toxin (NetB), portal proteins including G20c, P23_45, T4, SPP1, P22 and Phi29, Monalysin, Clostridium perfringens beta toxin, parasporin-2, epsilon toxin, lectin from the parasitic mushroom Laetiporus sulphureus (LSL), volvatoxin, Cry toxins, CytlAa or Cyt2Aa.
The transmembrane protein pore preferably comprises a CsgG pore or a homologue or mutant thereof, or an isolated pore complex comprising a CsgG pore, or a homologue or mutant thereof, and a modified CsgF peptide, or a homologue or mutant thereof. The CsgG pore or a homologue or mutant thereof, the isolated pore complex comprising a CsgG pore, or a homologue or mutant thereof, and a modified CsgF peptide, or a homologue or mutant thereof preferably comprises at least one mutant monomer.
A mutant CsgG monomer is a monomer whose sequence varies from that of a wild-type CsgG monomer and which retains the ability to form a pore. A mutant monomer may also be referred to herein as a variant. Methods for confirming the ability of mutant monomers to form pores are well-known in the art and are discussed in more detail below. The at least one mutant monomer or variant may have any of the %s of homology/sequence identity to
SEQ ID NO: 117 or SEQ ID NO: 3 set out below. The at least one mutant monomer may contain any of the modifications, mutations or substitutions described below, including the types of modifications and substitutions described with reference to the Dda helicases of the invention. The at least one mutant monomer may contain any of the additional modifications, mutations or substitutions described in WO2016/034591, WO2017/149316, WO2017/149317 and, WO2017/149318, WO2018/211241, and W02019/002893 (all incorporated by reference herein in their entirety).
Methods of characterising an analyte
The invention provides a method of determining the presence, absence or one or more characteristics of a target analyte. The method involves contacting the target analyte with a transmembrane pore and a helicase of the invention or a construct of the invention such that the helicase or construct controls the movement of the target analyte with respect to, such as into or through, the pore and taking one or more measurements as the analyte moves with respect to the pore and thereby determining the presence, absence or one or more characteristics of the analyte. The target analyte may also be called the template analyte or the analyte of interest. The transmembrane pore may be any of the pores or complexes discussed above. The transmembrane pore typically comprises at least 7, at least 8, at least 9 or at least 10 monomers, such as 7, 8, 9 or 10 CsgG monomers. The transmembrane pore preferably comprises eight or nine identical CsgG monomers. One or more, such as 2, 3, 4, 5, 6, 7, 8, 9 or 10, of the CsgG monomers is preferably chemically modified, or the CsgF peptide is chemically modified. The monomers, such as the CsgG monomers, or homologues or mutants thereof, and the modified CsgF monomers, or homologues or mutants thereof, may be derived from any organism. The analyte may pass through the CsgG constriction, followed by the CsgF constriction. In an alternative embodiment the analyte may pass through the CsgF constriction, followed by the CsgG constriction, depending on the orientation of the CsgG/CsgF complex in the membrane.
The method is for determining the presence, absence or one or more characteristics of a target analyte. The method may be for determining the presence, absence or one or more characteristics of at least one analyte. The method may concern determining the presence, absence or one or more characteristics of two or more analytes. The method may comprise determining the presence, absence or one or more characteristics of any number of analytes, such as 2, 5, 10, 15, 20, 30, 40, 50, 100 or more analytes. Any number of characteristics of the one or more analytes may be determined, such as 1, 2, 3, 4, 5, 10 or more characteristics.
The binding of a molecule in the channel of the pore complex, or in the vicinity of either opening of the channel will have an effect on the open-channel ion flow through the pore, which is the essence of "molecular sensing" of pore channels. In a similar manner to the
nucleic acid sequencing application, variation in the open-channel ion flow can be measured using suitable measurement techniques by the change in electrical current (for example, WO 2000/28312 and D. Stoddart et al., Proc. Natl. Acad. Sci., 2010, 106, 7702-7 or WO 2009/077734; all incorporated herein by reference in their entirety). The degree of reduction in ion flow, as measured by the reduction in electrical current, is related to the size of the obstruction within, or in the vicinity of, the pore. Binding of a molecule of interest, also referred to as an "analyte", in or near the pore therefore provides a detectable and measurable event, thereby forming the basis of a "biological sensor". Suitable molecules for nanopore sensing include nucleic acids; proteins; peptides; polysaccharides and small molecules (refers here to a low molecular weight (e.g., < 900Da or < 500Da) organic or inorganic compound) such as pharmaceuticals, toxins, cytokines, and pollutants. Detecting the presence of biological molecules finds application in personalised drug development, medicine, diagnostics, life science research, environmental monitoring and in the security and/or the defence industry.
The transmembrane pore may serve as a molecular or biological sensor. In some embodiments, the transmembrane pore can be derived or isolated from bacterial proteins (e.g., E. coli, Salmonella typhi). In some embodiments, the transmembrane pore can be recombinantly produced. Procedures for analyte detection are described in Howorka et al. Nature Biotechnology (2012) Jun 7; 30(6):506-7. The analyte molecule that is to be detected may bind to either face of the channel, or within the lumen of the channel itself. The position of binding may be determined by the size of the molecule to be sensed.
The one or more characteristics of the target analyte are preferably measured by electrical measurement and/or optical measurement. The electrical measurement is preferably a current measurement, an impedance measurement, a tunnelling measurement or a field effect transistor (FET) measurement. The method preferably further comprises the step of applying a voltage across the pore to form a complex between the pore and the helicase or construct.
The target analyte is preferably a metal ion, an inorganic salt, a polymer, an amino acid, a peptide, a polypeptide, a protein, a nucleotide, an oligonucleotide, a polynucleotide, a monosaccharide, an oligosaccharide, a polysaccharide, a dye, a bleach, a pharmaceutical, a diagnostic agent, a recreational drug, an explosive, a toxic compound, or an environmental pollutant. The target analyte is preferably a target polynucleotide. The method may concern determining the presence, absence or one or more characteristics of two or more analytes of the same type, such as two or more proteins, two or more nucleotides or two or more pharmaceuticals. Alternatively, the method may concern determining the presence, absence or one or more characteristics of two or more analytes of different types, such as one or more proteins, one or more nucleotides and one or more pharmaceuticals.
The target analyte, preferably the target polynucleotide, may be modified by methylation, by oxidation, by damage, with one or more proteins or with one or more labels, tags or spacers.
The target analyte can be secreted from cells. Alternatively, the target analyte can be an analyte that is present inside cells such that the analyte must be extracted from the cells before the method can be carried out.
In one embodiment, the target analyte is an amino acid, a peptide, a polypeptides or protein. The amino acid, peptide, polypeptide or protein can be naturally-occurring or non- naturally-occurring. The polypeptide or protein can include within them synthetic or modified amino acids. Several different types of modification to amino acids are known in the art. Suitable amino acids and modifications thereof are above. It is to be understood that the target analyte can be modified by any method available in the art.
In another embodiment, the target analyte is a polynucleotide, such as a nucleic acid, which is defined as a macromolecule comprising two or more nucleotides. Nucleic acids are particularly suitable for nanopore sequencing. The naturally-occurring nucleic acid bases in DNA and RNA may be distinguished by their physical size. As a nucleic acid molecule, or individual base, passes through the channel of a nanopore, the size differential between the bases causes a directly correlated reduction in the ion flow through the channel. The variation in ion flow may be recorded. Suitable electrical measurement techniques for recording ion flow variations are discussed above. Through suitable calibration, the characteristic reduction in ion flow can be used to identify the particular nucleotide and associated base traversing the channel in real-time. In typical nanopore nucleic acid sequencing, the open-channel ion flow is reduced as the individual nucleotides of the nucleic sequence of interest sequentially pass through the channel of the nanopore due to the partial blockage of the channel by the nucleotide. It is this reduction in ion flow that is measured using the suitable recording techniques described above. The reduction in ion flow may be calibrated to the reduction in measured ion flow for known nucleotides through the channel resulting in a means for determining which nucleotide is passing through the channel, and therefore, when done sequentially, a way of determining the nucleotide sequence of the nucleic acid passing through the nanopore. For the accurate determination of individual nucleotides, it has typically required for the reduction in ion flow through the channel to be directly correlated to the size of the individual nucleotide passing through the constriction (or "reading head"). It will be appreciated that sequencing may be performed upon an intact nucleic acid polymer that is 'threaded' through the pore via the action of an associated polymerase, for example. Alternatively, sequences may be determined by passage of nucleotide triphosphate bases that have been sequentially removed from a
target nucleic acid in proximity to the pore (see for example WO 2014/187924 incorporated herein by reference in its entirety).
The polynucleotide may be single stranded or double stranded. At least a portion of the polynucleotide is preferably double stranded. The polynucleotide is most preferably ribonucleic nucleic acid (RIMA) or deoxyribonucleic acid (DNA). In particular, said method using a polynucleotide as an analyte alternatively comprises determining one or more characteristics selected from (i) the length of the polynucleotide, (ii) the identity of the polynucleotide, (iii) the sequence of the polynucleotide, (iv) the secondary structure of the polynucleotide and (v) whether or not the polynucleotide is modified.
The polynucleotide can be any length (i). For example, the polynucleotide can be at least 10, at least 50, at least 100, at least 150, at least 200, at least 250, at least 300, at least 400 or at least 500 nucleotides or nucleotide pairs in length. The polynucleotide can be 1000 or more nucleotides or nucleotide pairs, 5000 or more nucleotides or nucleotide pairs in length or 100000 or more nucleotides or nucleotide pairs in length. Any number of polynucleotides can be investigated. For instance, the method may concern characterising 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 30, 50, 100 or more polynucleotides. If two or more polynucleotides are characterised, they may be different polynucleotides or two instances of the same polynucleotide. The polynucleotide can be naturally occurring or artificial. For instance, the method may be used to verify the sequence of a manufactured oligonucleotide. The method is typically carried out in vitro.
Embodiments involving the modified helicases and constructs of the invention The invention provides polynucleotides, vectors and host cells. These are discussed above.
The invention provides various methods using the Dda helicases of the invention in particular for controlling the movement of an analyte and characterising a target analyte. Such methods are described above and in WO2016/034591, WO2017/149316, WO2017/149317, WO2017/149318, WO2018/211241, W02019/002893, WO2015/055981, WO2015/166276 and WO2016/055777 (all incorporated by reference herein).
The modified Dda helicases and constructs of the invention may be used to form sensors for characterising target analytes. These sensors may be formed by contacting the pore and the helicase or construct in the presence of the target analyte and applying a potential across the pore. The potential may be a voltage potential or a chemical potential. The helicase or the construct may be covalently attached to the pore, for instance as described above.
In these methods and sensors, the modified Dda helicases and constructs of the invention may be combined/used with a solid state pore or a transmembrane protein pore. The transmembrane protein pore may be any of those discussed above.
Kits
The invention also provides a kit for characterising a target analyte comprising (a) a pore and a helicase or a construct of the invention or (b) a helicase or construct of the invention and one or more loading moieties. The pore may be any of those discussed above. Preferred combinations of pores and helicases are discussed in more detail below.
The kit preferably further comprises the components of a membrane. The kit may comprise components of any type of membranes, such as an amphiphilic layer or a triblock copolymer membrane. The kit may further comprise one or more anchors, such as cholesterol, for coupling the target analyte to the membrane. The kit may further comprise one or more polynucleotide adaptors that can be attached to a target polynucleotide to facilitate characterisation of the polynucleotide. In one embodiment, the anchor, such as cholesterol, is attached to the polynucleotide adaptor.
The kit may additionally comprise one or more other reagents or instruments which enable any of the embodiments mentioned above to be carried out. Such reagents or instruments include one or more of the following: suitable buffer(s) (aqueous solutions), means to obtain a sample from a subject (such as a vessel or an instrument comprising a needle), means to amplify and/or express polynucleotides or voltage or patch clamp apparatus. Reagents may be present in the kit in a dry state such that a fluid sample resuspends the reagents. The kit may also, optionally, comprise instructions to enable the kit to be used in the method of the invention or details regarding for which organism the method may be used. Finally, the kit may also comprise additional components useful in analyte characterization.
Polynucleotide binding proteins and moieties
Preferred polynucleotide binding proteins are polymerases, exonucleases, helicases and topoisomerases, such as gyrases. Preferred polynucleotide binding moieties are or are derived from polymerases, exonucleases, helicases and topoisomerases, such as gyrases. Suitable enzymes include, but are not limited to, exonuclease I from E. coli, exonuclease III enzyme from E. coli, Red from T. thermophilus and bacteriophage lambda exonuclease, TatD exonuclease and variants thereof. Three subunits comprising the Red sequence from T. thermophilus or a variant thereof interact to form a trimer exonuclease. The polymerase may be PyroPhage® 3173 DNA Polymerase (which is commercially available from Lucigen® Corporation), SD Polymerase (commercially available from Bioron®) or variants thereof. The enzyme may be Phi29 DNA polymerase (SEQ ID NO: 7) or a variant thereof. The topoisomerase is preferably a member of any of the Moiety Classification (EC) groups 5.99.1.2 and 5.99.1.3.
The enzyme is most preferably derived from a helicase, such as Hel308 Mbu, Hel308 Csy, Hel308 Tga, Hel308 Mhu, Tral Eco, XPD Mbu or a variant thereof. Any helicase may be used in the invention. The helicase may be or be derived from a Hel308 helicase, a RecD helicase, such as Tral helicase or a TrwC helicase, a XPD helicase or a Dda helicase. The helicase may be any of the helicases, modified helicases or helicase constructs disclosed in WO 2013/057495; WO 2013/098562; WO2013098561; WO 2014/013260; WO 2014/013259; WO 2014/013262 and WO 2015/055981. All of these are incorporated by reference in their entirety.
Apparatus
The invention also provides an apparatus for characterising target analytes in a sample, comprising (a) a plurality of pores and (b) a plurality of helicases or a plurality of constructs of the invention. The plurality of pores may be any of those discussed above. Preferred combinations of pores and helicases are discussed in more detail below.
Any of the specific embodiments discussed above, especially in relation to the pores, pore complexes, helicases of the invention, and constructs of the invention, are equally applicable to the apparatuses of the invention.
Arrays
The invention may use an array comprising a plurality of membranes. Any of the embodiments discussed above with respect to the membrane equally apply the array of the invention. The array may be set up to perform any of the methods described above.
In a preferred embodiment, each membrane in the array comprises one pore. Due to the manner in which the array is formed, for example, the array may comprise one or more membranes that do not comprise a pore, and/or one or more membranes that comprise two or more pores. The array may comprise from about 2 to about 1000, such as from about 10 to about 800, from about 20 to about 600 or from about 30 to about 500 membranes.
System
The invention may use a system comprising (a) a membrane or an array, (b) means for applying a potential across the membrane(s) and (c) means for detecting electrical or optical signals across the membrane(s).
The pores and membranes may be any as described above and below.
In one embodiment, the system further comprises a first chamber and a second chamber, wherein the first and second chambers are separated by the membrane(s). When used to characterise a target analyte, the system may further comprise a target analyte, wherein the target analyte is transiently located within the continuous channel and wherein one end
of the target analyte is located in the first chamber and one end of the target analyte is located in the second chamber. The target analyte is preferably a target polypeptide or a target polynucleotide.
In one embodiment, the system further comprises an electrically conductive solution in contact with the pore(s), electrodes providing a voltage potential across the membrane(s), and a measurement system for measuring the current through the pore(s). In one embodiment, the voltage applied across the membranes and pore is from +5 V to -5 V, such as -600 mV to +600mV or -400 mV to +400 mV. The voltage used is preferably in the range 100 mV to 240 mV and more preferably in the range of 120 mV to 220 mV. It is possible to increase discrimination between different amino acids or nucleotides by a pore by using an increased applied potential. Any suitable electrically conductive solution may be used. For example, the solution may comprise charge carriers, such as metal salts, for example alkali metal salt, halide salts, for example chloride salts, such as alkali metal chloride salt. Charge carriers may include ionic liquids or organic salts, for example tetramethyl ammonium chloride, trimethylphenyl ammonium chloride, phenyltrimethyl ammonium chloride, or l-ethyl-3-methyl imidazolium chloride. In an exemplary system, salt is present in the aqueous solution in the chamber. Potassium chloride (KCI), sodium chloride (NaCI), caesium chloride (CsCI) or a mixture of potassium ferrocyanide and potassium ferricyanide is typically used. KCI, NaCI and a mixture of potassium ferrocyanide and potassium ferricyanide are preferred. The charge carriers may be asymmetric across the membrane. For instance, the type and/or concentration of the charge carriers may be different on each side of the membrane, e.g., in each chamber.
The salt concentration may be at saturation. The salt concentration may be 3 M or lower and is typically from 0.1 to 2.5 M, from 0.3 to 1.9 M, from 0.5 to 1.8 M, from 0.7 to 1.7 M, from 0.9 to 1.6 M or from 1 M to 1.4 M. The salt concentration is preferably from 150 mM to 1 M. The method is preferably carried out using a salt concentration of at least 0.3 M, such as at least 0.4 M, at least 0.5 M, at least 0.6 M, at least 0.8 M, at least 1.0 M, at least 1.5 M, at least 2.0 M, at least 2.5 M or at least 3.0 M. High salt concentrations provide a high signal to noise ratio and allow for currents indicative of the presence of an amino acid or nucleotide to be identified against the background of normal current fluctuations.
A buffer may be present in the electrically conductive solution. Typically, the buffer is phosphate buffer. Other suitable buffers are HEPES and Tris-HCI buffer. The pH of the electrically conductive solution may be from 4.0 to 12.0, from 4.5 to 10.0, from 5.0 to 9.0, from 5.5 to 8.8, from 6.0 to 8.7 or from 7.0 to 8.8 or 7.5 to 8.5. The pH used is preferably about 7.5.
The system may be comprised in an apparatus. The apparatus may be any conventional apparatus for analyte analysis, such as an array or a chip. The apparatus is preferably set up to carry out the disclosed method. For example, the apparatus may comprise a chamber comprising an aqueous solution and a barrier that separates the chamber into two sections. The barrier typically has an aperture in which the membrane(s) containing the pore(s) are formed. Alternatively, the barrier forms the membrane in which the pore is present.
The apparatus may also comprise an electrical circuit capable of applying a potential and measuring an electrical signal across the membrane and pore.
The apparatus may be any of those described in WO 2008/102120, WO 2009/077734, WO 2010/122293, WO 2011/067559, or WO 00/28312 (all incorporated herein by reference in their entirety).
Membrane
Any suitable membrane may be used in the invention. The membrane is preferably an amphiphilic layer. An amphiphilic layer is a layer formed from amphiphilic molecules, such as phospholipids, which have both hydrophilic and lipophilic properties. The amphiphilic molecules may be synthetic or naturally occurring. Non-naturally occurring amphiphiles and amphiphiles which form a monolayer are known in the art and include, for example, block copolymers (Gonzalez-Perez et al., Langmuir, 2009, 25, 10447-10450). Block copolymers are polymeric materials in which two or more monomer sub-units that are polymerized together to create a single polymer chain. Block copolymers typically have properties that are contributed by each monomer sub-unit. However, a block copolymer may have unique properties that polymers formed from the individual sub-units do not possess. Block copolymers can be engineered such that one of the monomer sub-units is hydrophobic (i.e., lipophilic), whilst the other sub-unit(s) are hydrophilic whilst in aqueous media. In this case, the block copolymer may possess amphiphilic properties and may form a structure that mimics a biological membrane. The block copolymer may be a diblock (consisting of two monomer sub-units) but may also be constructed from more than two monomer sub-units to form more complex arrangements that behave as amphipiles. The copolymer may be a triblock, tetrablock or pentablock copolymer. The membrane is preferably a triblock copolymer membrane.
The membrane is most preferably one of the membranes disclosed in International Application No. WO2014/064443 or WO2014/064444.
The amphiphilic molecules may be chemically modified or functionalised to facilitate coupling of the polynucleotide. The amphiphilic layer may be a monolayer or a bilayer. The amphiphilic layer is typically planar. The amphiphilic layer may be curved. The amphiphilic layer may be supported.
Amphiphilic membranes are typically naturally mobile, essentially acting as two-dimensional fluids with lipid diffusion rates of approximately IO-8 cm s’1. This means that the pore and coupled polynucleotide can typically move within an amphiphilic membrane.
The membrane may be a lipid bilayer. Lipid bilayers are models of cell membranes and serve as excellent platforms for a range of experimental studies. For example, lipid bilayers can be used for in vitro investigation of membrane proteins by single-channel recording. Alternatively, lipid bilayers can be used as biosensors to detect the presence of a range of substances. The lipid bilayer may be any lipid bilayer. Suitable lipid bilayers include, but are not limited to, a planar lipid bilayer, a supported bilayer, or a liposome. The lipid bilayer is preferably a planar lipid bilayer. Suitable lipid bilayers are disclosed in WO 2008/102121, WO 2009/077734, and WO 2006/100484 (all incorporated herein by reference in their entirety).
In another preferred embodiment, the membrane comprises a solid-state layer. Solid state layers can be formed from both organic and inorganic materials including, but not limited to, microelectronic materials, insulating materials such as SisIX , AI2O3, and SiO, organic and inorganic polymers such as polyamide, plastics such as Teflon® or elastomers such as two-component addition-cure silicone rubber, and glasses. The solid-state layer may be formed from graphene. Suitable graphene layers are disclosed in WO 2009/035647 (incorporated herein by reference in its entirety). If the membrane comprises a solid-state layer, the pore is typically present in an amphiphilic membrane or layer contained within the solid-state layer, for instance within a hole, well, gap, channel, trench or slit within the solid-state layer. The skilled person can prepare suitable solid state/amphiphilic hybrid systems. Suitable systems are disclosed in WO 2009/020682 and WO 2012/005857 (both incorporated herein by reference in their entirety). Any of the amphiphilic membranes or layers discussed above may be used.
The method is typically carried out using (i) an artificial amphiphilic layer comprising a pore, (ii) an isolated, naturally occurring lipid bilayer comprising a pore, or (iii) a cell having a pore inserted therein. The method is typically carried out using an artificial amphiphilic layer, such as an artificial triblock copolymer layer. The layer may comprise other transmembrane and/or intramembrane proteins as well as other molecules in addition to the pore. Suitable apparatus and conditions are discussed below. The method of the invention is typically carried out in vitro.
Preferred combinations of pores and helicases
In several embodiments above, the modified helicase of the invention may be used with a pore or a transmembrane pore.
The modified helicase of the invention may be used in combination with any of the pores described in WO2016/034591, WO2017/149316, WO2017/149317, WO2017/149318, WO2018/211241, W02019/002893 and PCT/EP2023/059821 (all incorporated by reference herein in their entirety). The modified helicase of the invention is preferably used with a CsgG pore as described in WO2016/034591 (incorporated herein by reference) as in Example 2 or a CsgG:CsgF pore complex as described in W02019/002893 (incorporated herein by reference).
The helicase of the invention is preferably used with a CsgG pore or a CsgG:CsgF pore complex described above and vice versa. The CsgG pore or a CsgG:CsgF pore complex of the invention preferably comprises Q100A and/or N102A or N102S and the helicases preferably comprises any of the substitutions in PCT/EP2023/059821, preferably Y350I and/or K358I.
It is to be understood that although particular embodiments, specific configurations as well as materials and/or molecules, have been discussed herein for engineered cells and methods according to the present invention, various changes or modifications in form and detail may be made without departing from the scope and spirit of this invention. The following examples are provided to better illustrate particular embodiments, and they should not be considered limiting the application. The application is limited only by the claims.
Sequences
SEQ ID NO: 1 shows polynucleotide sequence of wild-type E. coli CsgG from strain K12, including signal sequence (Gene ID: 945619).
SEQ ID NO:2 shows amino acid sequence of wild-type E. coli CsgG including signal sequence (Uniprot accession number P0AEA2).
SEQ ID NO:3 shows amino acid sequence of wild-type E. coli CsgG as mature protein (Uniprot accession number P0AEA2).
SEQ ID NO:4 shows polynucleotide sequence of wild-type E. coli CsgF from strain K12, including signal sequence (Gene ID: 945622).
SEQ ID NO:5 shows amino acid sequence of wild-type E. coli CsgF including signal sequence (Uniprot accession number P0AE98).
SEQ ID NO:6 shows amino acid sequence of wild-type E. coli CsgF as mature protein (Uniprot accession number P0AE98).
SEQ ID N0:7 shows polynucleotide sequence of a fragment of wild-type E. coli CsgF encoding amino acids 1 to 27 and a C-terminal 6 His tag.
SEQ ID NO:8 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 1 to 27 and a C-terminal 6 His tag.
SEQ ID NO:9 shows polynucleotide sequence of a fragment of wild-type E. coli CsgF encoding amino acids 1 to 38 and a C-terminal 6 His tag.
SEQ ID NO: 10 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 1 to 38 and a C-terminal 6 His tag.
SEQ ID NO: 11 shows polynucleotide sequence of a fragment of wild-type E. coli CsgF encoding amino acids 1 to 48 and a C-terminal 6 His tag.
SEQ ID NO: 12 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 1 to 48 and a C-terminal 6 His tag.
SEQ ID NO: 13 shows polynucleotide sequence of a fragment of wild-type E. coli CsgF encoding amino acids 1 to 64 and a C-terminal 6 His tag.
SEQ ID NO: 14 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 1 to 64 and a C-terminal 6 His tag.
SEQ ID NO: 15 shows amino acid sequence of a peptide corresponding to residues 20 to 53 of E. coli CsgF
SEQ ID NO: 16 shows amino acid sequence of a peptide corresponding to residues 20 to 42 of E. coli CsgF, including KD at its C-terminus
SEQ ID NO: 17 shows amino acid sequence of a peptide corresponding to residues 23 to 55 of CsgF homologue Q88H88
SEQ ID NO: 18 shows amino acid sequence of a peptide corresponding to residues 25 to 57 of CsgF homologue A0A143HJA0
SEQ ID NO: 19 shows amino acid sequence of a peptide corresponding to residues 21 to 53 of CsgF homologue Q5E245
SEQ ID NO:20 shows amino acid sequence of a peptide corresponding to residues 19 to 51 of CsgF homologue Q084E5
SEQ ID NO:21 shows amino acid sequence of a peptide corresponding to residues 15 to 47 of CsgF homologue F0LZU2
SEQ ID NO:22 shows amino acid sequence of a peptide corresponding to residues 26 to 58 of CsgF homologue A0A136HQR0
SEQ ID NO:23 shows amino acid sequence of a peptide corresponding to residues 21 to 53 of CsgF homologue A0A0W1SRL3
SEQ ID NO:24 shows amino acid sequence of a peptide corresponding to residues 26 to 59 of CsgF homologue B0UH01
SEQ ID NO:25 shows amino acid sequence of a peptide corresponding to residues 22 to 53 of CsgF homologue Q6NAU5
SEQ ID NO:26 shows amino acid sequence of a peptide corresponding to residues 7 to 38 of CsgF homologue G8PUY5
SEQ ID NO:27 shows amino acid sequence of a peptide corresponding to residues 25 to 57 of CsgF homologue A0A0S2ETP7
SEQ ID NO:28 shows amino acid sequence of a peptide corresponding to residues 19 to 51 of CsgF homologue E3I1Z1
SEQ ID NO:29 shows amino acid sequence of a peptide corresponding to residues 24 to 55 of CsgF homologue F3Z094
SEQ ID NO:30 shows amino acid sequence of a peptide corresponding to residues 21 to 53 of CsgF homologue A0A176T7M2
SEQ ID NO:31 shows amino acid sequence of a peptide corresponding to residues 14 to 45 of CsgF homologue D2QPP8
SEQ ID NO:32 shows amino acid sequence of a peptide corresponding to residues 28 to 58 of CsgF homologue N2IYT1
SEQ ID NO:33 shows amino acid sequence of a peptide corresponding to residues 26 to 58 of CsgF homologue W7QHV5
SEQ ID NO:34 shows amino acid sequence of a peptide corresponding to residues 23 to 55 of CsgF homologue D4ZLW2
SEQ ID NO:35 shows amino acid sequence of a peptide corresponding to residues 21 to 53 of CsgF homologue D2QT92
SEQ ID NO:36 shows amino acid sequence of a peptide corresponding to residues 20 to 51 of CsgF homologue A0A167UJA2
SEQ ID NO:37 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 20 to 27.
SEQ ID NO:38 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 20 to 38.
SEQ ID NO:39: shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 20 to 48.
SEQ ID NO:40 shows amino acid sequence of a fragment of wild-type E. coli CsgF encompassing amino acids 20 to 64.
SEQ ID NO:41 shows the nucleotide sequence of primer CsgF_d27_end
SEQ ID NO:42 shows the nucleotide sequence of primer CsgF_d38_end
SEQ ID NO:43 shows the nucleotide sequence of primer CsgF_d48_end
SEQ ID NO:44 shows the nucleotide sequence of primer CsgF_d64_end
SEQ ID NO:45 shows the nucleotide sequence of primer pNa62_CsgF_histag_Fw
SEQ ID NO:46 shows the nucleotide sequence of primer CsgF-His_pET22b_FW
SEQ ID NO:47 shows the nucleotide sequence of primer CsgF-His_pET22b_Rev
SEQ ID NO:48 shows the nucleotide sequence of primer csgEFG_pDONR221_FW
SEQ ID NO:49 shows the nucleotide sequence of primer csgEFG_pDONR221_Rev
SEQ ID NO: 50 shows the nucleotide sequence of primer Mut_csgF_His_FW
SEQ ID NO:51 shows the nucleotide sequence of primer Mut_csgF_His_Rev
SEQ ID NO:52 shows the nucleotide sequence of primer DelCsgE_Rev
SEQ ID NO:53 shows the nucleotide sequence of primer DelCsgE FW
SEQ ID NO: 54 shows the amino acid sequence of residues 1 to 30 of mature E. coli CsgF
SEQ ID NO: 55 shows the amino acid sequence of residues 1 to 35 of mature E. coli CsgF
SEQ ID NO: 56 shows the amino acid sequence of a mutated (T4C/N17S) CsgF sequence with a signal sequence, and a TEV protease cleavage site (ENLYFQS) inserted between residues 35 and 36 of sequence of the mature protein.
SEQ ID NO: 57 shows the amino acid sequence of a mutated (N17S-Del) CsgF sequence with a signal sequence, and a TEV protease cleavage site (ENLYFQS) inserted between residues 35 and 36 of sequence of the mature protein.
SEQ ID NO: 58 shows the amino acid sequence of a mutated (G1C/N17S) CsgF sequence with a signal sequence, and a TEV protease cleavage site (ENLYFQS) inserted between residues 35 and 36 of sequence of the mature protein.
SEQ ID NO: 59 shows the amino acid sequence of a mutated (G1C) CsgF sequence with a signal sequence, and a TEV protease cleavage site (ENLYFQS) inserted between residues 35 and 36 of sequence of the mature protein.
SEQ ID NO: 60 shows the amino acid sequence of a CsgF sequence with a signal sequence, a TEV protease cleavage site (ENLYFQS) inserted between residues 45 and 46 of sequence of the mature protein, and a Hisio tag at the C-terminus.
SEQ ID NO: 61 shows the amino acid sequence of a CsgF sequence with a signal sequence, a TEV protease cleavage site (ENLYFQS) inserted between residues 35 and 36 of sequence of the mature protein, and a Hisio tag at the C-terminus.
SEQ ID NO: 62 shows the amino acid sequence of a CsgF sequence with a signal sequence, a TEV protease cleavage site (ENLYFQS) inserted between residues 30 and 31 of sequence of the mature protein, and a Hisio tag at the C-terminus.
SEQ ID NO: 63 shows the amino acid sequence of a CsgF sequence with a signal sequence, a TEV protease cleavage site (ENLYFQS) inserted between residues 45 and 51 of sequence of the mature protein, and a Hisio tag at the C-terminus.
SEQ ID NO: 64 shows the amino acid sequence of a CsgF sequence with a signal sequence, a TEV protease cleavage site (ENLYFQS) inserted between residues 30 and 37 of sequence of the mature protein, and a Hisio tag at the C-terminus.
SEQ ID NO: 65 shows the amino acid sequence of a CsgF sequence with a signal sequence, a HCV C3 protease cleavage site (LEVLFQGP) inserted between residues 34 and 36 of sequence of the mature protein, and a Hisio tag at the C-terminus.
SEQ ID NO: 66 shows the amino acid sequence of a CsgF sequence with a signal sequence, a HCV C3 protease cleavage site (LEVLFQGP) inserted between residues 42 and 43 of sequence of the mature protein, and a Hisio tag at the C-terminus.
SEQ ID NO: 67 shows the amino acid sequence of a CsgF sequence with a signal sequence, a HCV C3 protease cleavage site (LEVLFQGP) inserted between residues 38 and 47 of sequence of the mature protein, and a Hisio tag at the C-terminus.
SEQ ID NO: 68 shows the amino acid sequence of YP_001453594.1 : 1-248 of hypothetical protein CKO_02032 [Citrobacter koseri ATCC BAA-895], which is 99% identical to SEQ ID NO: 3.
SEQ ID NO: 69 shows the amino acid sequence of WP_001787128.1: 16-238 of curli production assembly/transport component CsgG, partial [Salmonella enterica], which is 98% to SEQ ID NO: 3.
SEQ ID NO: 70 shows the amino acid sequence of KEY44978.1 | : 16-277 of curli production assembly/transport protein CsgG [Citrobacter amalonaticus], which is 98% identical to SEQ ID NO: 3.
SEQ ID NO: 71 shows the amino acid sequence of YP_003364699.1 : 16-277 of curli production assembly/transport component [Citrobacter rodentium ICC168], which is 97% identical to SEQ ID NO: 3.
SEQ ID NO: 72 shows the amino acid sequence of YP_004828099.1 : 16-277 of curli production assembly/transport component CsgG [Enterobacter asburiae LF7a], which is 94% identical to SEQ ID NO: 3.
SEQ ID NO: 73 shows the amino acid sequence of WP_006819418.1: 19-280 of transporter [Yokenella regensburgei], which is 91% identical to SEQ ID NO: 3.
SEQ ID NO: 74 shows the amino acid sequence of WP_024556654.1: 16-277 of curli production assembly/transport protein CsgG [Cronobacter pulveris], which is 89% identical to SEQ ID NO: 3.
SEQ ID NO: 75 shows the amino acid sequence of YP_005400916.1 : 16-277 of curli production assembly/transport protein CsgG [Rahnella aquatilis HX2], which is 84% identical to SEQ ID NO: 3.
SEQ ID NO: 76 shows the amino acid sequence of KFC99297.1 : 20-278 of CsgG family curli production assembly/transport component [Kluyvera ascorbata ATCC 33433], which is 82% identical to SEQ ID NO: 3.
SEQ ID NO: 77 shows the amino acid sequence of KFC86716.11 : 16-274 of CsgG family curli production assembly/transport component [Hafnia alvei ATCC 13337], which is 81% identical to SEQ ID NO: 3.
SEQ ID NO: 78 shows the amino acid sequence of YP_007340845.11 : 16-270 of uncharacterised protein involved in formation of curli polymers [Enterobacteriaceae bacterium strain FGI 57], which is 76% identical to SEQ ID NO: 3.
SEQ ID NO: 79 shows the amino acid sequence of WP_010861740.1: 17-274 of curli production assembly/transport protein CsgG [Plesiomonas shigelloides], which is 70% identical to SEQ ID NO: 3.
SEQ ID NO: 80 shows the amino acid sequence of YP_205788.1 : 23-270 of curli production assembly/transport outer membrane lipoprotein component CsgG [Vibrio fischeri ESI 14], which is 60% identical to SEQ ID NO: 3.
SEQ ID NO: 81 shows the amino acid sequence of WP_017023479.1: 23-270 of curli production assembly protein CsgG [Aliivibrio logei], which is 59% identical to SEQ ID NO: 3.
SEQ ID NO: 82 shows the amino acid sequence of WP_007470398.1: 22-275 of Curli production assembly/transport component CsgG [Photobacterium sp. AK15], which is 57% identical to SEQ ID NO: 3.
SEQ ID NO: 83 shows the amino acid sequence of WP_021231638.1: 17-277 of curli production assembly protein CsgG [Aeromonas veronii], which is 56% identical to SEQ ID NO: 3.
SEQ ID NO: 84 shows the amino acid sequence of WP_033538267.1: 27-265 of curli production assembly/transport protein CsgG [Shewanella sp. ECSMB14101], which is 56% identical to SEQ ID NO: 3.
SEQ ID NO: 85 shows the amino acid sequence of WP_003247972.1: 30-262 of curli production assembly protein CsgG [Pseudomonas putida], which is 54% identical to SEQ ID NO: 3.
SEQ ID NO: 86 shows the amino acid sequence of YP_003557438.1 : 1-234 of curli production assembly/transport component CsgG [Shewanella violacea DSS12], which is 53% identical to SEQ ID NO: 3.
SEQ ID NO: 87 shows the amino acid sequence of WP_027859066.1: 36-280 of curli production assembly/transport protein CsgG [Marinobacterium jannaschii], which is 53% identical to SEQ ID NO: 3.
SEQ ID NO: 88 shows the amino acid sequence of CEJ70222.1: 29-262 of Curli production assembly/transport component CsgG [Chryseobacterium oranimense G311], which is 50% identical to SEQ ID NO: 3.
SEQ ID NOs: 89 to 104 show the sequences in Table 1.
SEQ ID NO: 1 (> P0AEA2; coding sequence for WT CsgG from E. coli K12)
ATGCAGCGCTTATTTCTTTTGGTTGCCGTCATGTTACTGAGCGGATGCTTAACCGCCCCGCCTAAAG
AAGCCGCCAGACCGACATTAATGCCTCGTGCTCAGAGCTACAAAGATTTGACCCATCTGCCAGCGCC
GACGGGTAAAATCTTTGTTTCGGTATACAACATTCAGGACGAAACCGGGCAATTTAAACCCTACCCG
GCAAGTAACTTCTCCACTGCTGTTCCGCAAAGCGCCACGGCAATGCTGGTCACGGCACTGAAAGATT
CTCGCTGGTTTATACCGCTGGAGCGCCAGGGCTTACAAAACCTGCTTAACGAGCGCAAGATTATTCG
TGCGGCACAAGAAAACGGCACGGTTGCCATTAATAACCGAATCCCGCTGCAATCTTTAACGGCGGCA
AATATCATGGTTGAAGGTTCGATTATCGGTTATGAAAGCAACGTCAAATCTGGCGGGGTTGGGGCAA
GATATTTTGGCATCGGTGCCGACACGCAATACCAGCTCGATCAGATTGCCGTGAACCTGCGCGTCGT
CAATGTGAGTACCGGCGAGATCCTTTCTTCGGTGAACACCAGTAAGACGATACTTTCCTATGAAGTT
CAGGCCGGGGTTTTCCGCTTTATTGACTACCAGCGCTTGCTTGAAGGGGAAGTGGGTTACACCTCGA
ACGAACCTGTTATGCTGTGCCTGATGTCGGCTATCGAAACAGGGGTCATTTTCCTGATTAATGATGG
TATCGACCGTGGTCTGTGGGATTTGCAAAATAAAGCAGAACGGCAGAATGACATTCTGGTGAAATAC
CGCCATATGTCGGTTCCACCGGAATCCTGA
SEQ ID NO:2 (> P0AEA2 (1 : 277); WT prepro CsgG from E. coli K12)
MQRLFLLVAVMLLSGCLTAPPKEAARPTLMPRAQSYKDLTHLPAPTGKIFVSVYNIQDETGQFKPYPASNF
STAVPQSATAMLVTALKDSRWFIPLERQGLQNLLNERKIIRAAQENGTVAINNRIPLQSLTAANIMVEGSI
IGYESNVKSGGVGARYFGIGADTQYQLDQIAVNLRVVNVSTGEILSSVNTSKTILSYEVQAGVFRFIDYQ
RLLEGEVGYTSNEPVMLCLMSAIETGVIFLINDGIDRGLWDLQNKAERQNDILVKYRHMSVPPES
SEQ ID NO:3 (> P0AEA2 (16: 277); mature CsgG from E. coli K12)
CLTAPPKEAARPTLMPRAQSYKDLTHLPAPTGKIFVSVYNIQDETGQFKPYPASNFSTAVPQSATAMLVT
ALKDSRWFIPLERQGLQNLLNERKIIRAAQENGTVAINNRIPLQSLTAANIMVEGSIIGYESNVKSGGVG
ARYFGIGADTQYQLDQIAVNLRVVNVSTGEILSSVNTSKTILSYEVQAGVFRFIDYQRLLEGEVGYTSNEP
VMLCLMSAIETGVIFLINDGIDRGLWDLQNKAERQNDILVKYRHMSVPPES
SEQ ID NO:4 (> P0AE98; coding sequence for WT CsgF from E. coli K12)
ATGCGTGTCAAACATGCAGTAGTTCTACTCATGCTTATTTCGCCATTAAGTTGGGCTGGAACCATGAC
TTTCCAGTTCCGTAATCCAAACTTTGGTGGTAACCCAAATAATGGCGCTTTTTTATTAAATAGCGCTC
AGGCCCAAAACTCTTATAAAGATCCGAGCTATAACGATGACTTTGGTATTGAAACACCCTCAGCGTTA
GATAACTTTACTCAGGCCATCCAGTCACAAATTTTAGGTGGGCTACTGTCGAATATTAATACCGGTAA
ACCGGGCCGCATGGTGACCAACGATTATATTGTCGATATTGCCAACCGCGATGGTCAATTGCAGTTG
AACGTGACAGATCGTAAAACCGGACAAACCTCGACCATCCAGGTTTCGGGTTTACAAAATAACTCAA
CCGATTTT
SEQ ID NO:5 (>P0AE98 (1 : 138); WT pre CsgF from E. coli K12)
MRVKHAVVLLMLISPLSWAGTMTFQFRNPNFGGNPNNGAFLLNSAQAQNSYKDPSYNDDFGIETPSAL
DNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQVSGLQNNSTD
F
SEQ ID NO:6 (>P0AE98 (20: 138); WT mature CsgF from E. coli K12)
GTMTFQFRNPNFGGNPNNGAFLLNSAQAQNSYKDPSYNDDFGIETPSALDNFTQAIQSQILGGLLSNIN
TGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQVSGLQNNSTDF
SEQ ID NO:7 (>P0AE98; coding sequence for CsgF l :27_6His)
ATGCGTGTCAAACATGCAGTAGTTCTACTCATGCTTATTTCGCCATTAAGTTGGGCTGGAACCATGAC TTTCCAGTTCCGTCATCACCATCACCATCACTAAGCCC
SEQ ID NO:8 (>P0AE98 (1 :28); preprotein of CsgF 20:27_6His)
MRVKHAVVLLMLISPLSWA GTMTFQFR HHHHHH
SEQ ID NO:9 (>P0AE98; coding sequence for CsgF l :38_6His)
ATGCGTGTCAAACATGCAGTAGTTCTACTCATGCTTATTTCGCCATTAAGTTGGGCTGGAACCATGAC
TTTCCAGTTCCGTAATCCAAACTTTGGTGGTAACCCAAATAATGGCCATCACCATCACCATCACTAAG CCC
SEQ ID NO:10 (> P0AE98 (1 :39); preprotein of CsgF 20:38_6His)
MRVKHAVVLLMLISPLSWAGTMTFQFRNPNFGGNPNNG HHHHHH
SEQ ID NO:11 (> P0AE98; coding sequence for CsgF l :48_6His)
ATGCGTGTCAAACATGCAGTAGTTCTACTCATGCTTATTTCGCCATTAAGTTGGGCTGGAACCATGAC
TTTCCAGTTCCGTAATCCAAACTTTGGTGGTAACCCAAATAATGGCGCTTTTTTATTAAATAGCGCTC AGGCCCAACATCACCATCACCATCACTAAGCCC
SEQ ID NO:12 (> P0AE98 (1 :49); preprotein of CsgF 20:48_6His)
MRVKHAVVLLMLISPLSWAGTMTFQFRNPNFGGNPNNGAFLLNSAQAQ HHHHHH
SEQ ID NO:13 (> P0AE98; coding sequence for CsgF l :64_6His)
ATGCGTGTCAAACATGCAGTAGTTCTACTCATGCTTATTTCGCCATTAAGTTGGGCTGGAACCATGAC
TTTCCAGTTCCGTAATCCAAACTTTGGTGGTAACCCAAATAATGGCGCTTTTTTATTAAATAGCGCTC
AGGCCCAAAACTCTTATAAAGATCCGAGCTATAACGATGACTTTGGTATTGAAACA
CATCACCATCACCATCACTAAGCCC
SEQ ID NO:14 (> P0AE98 (1 :65); preprotein of CsgF 20:64_6His)
MRVKHAVVLLMLISPLSWAGTMTFQFRNPNFGGNPNNGAFLLNSAQAQNSYKDPSYNDDFGIETHHHH HH
SEQ ID NO:15 (>P0AE98 (20 : 53); mature peptide of CsgF 20: 53)
GTMTFQFRNPNFGGNPNNGAFLLNSAQAQNSYKD
SEQ ID NO:16 (>P0AE98 (20 :42); mature peptide of CsgF 20:42+KD)
GTMTFQFRNPNFGGNPNNGAFLLKD
SEQ ID NO:17 (>Q88H88_PSEPK (23 : 55))
TELVYTPVNPAFGGNPLNGTWLLNNAQAQNDY
SEQ ID NO:18 (>A0A143HJA0_9GAMM (25: 57))
TELIYEPVNPNFGGNPLNGSYLLNNAQAQDRH
SEQ ID NO:19 (>Q5E245_VIBF1 (21 : 53))
SELVYTPVNPNFGGNPLNTSHLFGGANAINDY
SEQ ID NO:20 (>Q084E5_SHEFN (19: 51))
TQLVYTPVNPAFGGSYLNGSYLLANASAQNEH
SEQ ID NO:21 (>F0LZU2_VIBFN (15 :47))
SSLVYEPVNPTFGGNPLNTTHLFSRAEAINDY
SEQ ID NO:22 (>A0A136HQR0_9ALTE (26: 58))
TELVYEPINPSFGGNPLNGSFLLSKANSQNAH
SEQ ID NO:23 (>AOAOW1SRL3_9GAMM (21 : 53))
TEIVYQPINPSFGGNPMNGSFLLQKAQSQNAH
SEQ ID NO:24 (> B0UH01_METS4 (26: 59))
SSLVYQPVNPAFGGPQLNGSWLQAEANAQNIPQ
SEQ ID NO:25 (>Q6NAU5_RHOPA (22: 53))
GSLVYTPTNPAFGGSPLNGSWQMQQATAGNH
SEQ ID NO:26 (>G8PUY5_PSEUV (7 :38))
QQLIYQPTNPSFGGYAANTTHLFATANAQKTA
SEQ ID NO:27 (>A0A0S2ETP7_9RHIZ (25 : 57))
GDLVYTPVNPSFGGSPLNSAHLLSIAGAQKNA
SEQ ID NO:28 (>E3I1Z1_RHOVT (19 : 51))
AELGYTPVNPSFGGSPLNGSTLLSEASAQKPN
SEQ ID NO:29 (>F3Z094_DESAF (24: 55))
TELVFSFTNPSFGGDPMIGNFLLNKADSQKR
SEQ ID NO:30 (>AOA176T7M2_9FLAO (21 : 53))
QQLVYKSINPFFGGGDSFAYQQLLASANAQND
SEQ ID NO:31 (> D2QPP8_SPILD (14:45))
QALVYHPNNPAFGGNTFNYQWMLSSAQAQDR
SEQ ID NO:32 (> N2IYT1_9PSED (26: 58))
TELVYTPKNPAFGGSPLNGSYLLGNAQAQNDY
SEQ ID NO:33 (> W7QHV5_9GAMM (26: 58))
GQLIYQPINPSFGGDPLLGNHLLNKAQAQDTK
SEQ ID NO:34 (> D4ZLW2_SHEVD (23: 55))
TQLIYTPVNPNFGGSYLNGSYLLANASVQNDH
SEQ ID NO:35 (> D2QT92_SPILD (21 : 53))
QAFVYHPNNPNFGGNTFNYSWMLSSAQAQDRT
SEQ ID NO:36 (>AOA167UJA2_9FLAO (20 : 51))
QGLIYKPKNPAFGGDTFNYQWLASSAESQNK
SEQ ID NO:37(>POAE98 (20:28); mature peptide of CsgF 20 :27)
GTMTFQFR
SEQ ID NO:38(>POAE98 (20:39); mature peptide of CsgF 20 :38)
GTMTFQFRNPNFGGNPNNG
SEQ ID NO:39(>POAE98 (20:49); mature peptide of CsgF 20 :48)
GTMTFQFRNPNFGGNPNNGAFLLNSAQAQ
SEQ ID NO:40(>P0AE98 (20:65); mature peptide of CsgF 20 :64)
GTMTFQFRNPNFGGNPNNGAFLLNSAQAQNSYKDPSYNDDFGIET
SEQ ID NO:41 (CsgF_d27_end)
ACGGAACTGGAAAGTCATGGTTCC
SEQ ID NO:42 (CsgF_d38_end)
GCCATTATTTGGGTTACCACCAAAGTTTGG
SEQ ID NO:43 (CsgF_d48_end)
TTGGGCCTGAGCGCTATTTAATAAAAAAGC
SEQ ID NO:44 (CsgF_d64_end)
TGTTTCAATACCAAAGTCATCGTTATAGCTCGG
SEQ ID NO:45 (pNa62_CsgF_histag_Fw)
CATCACCATCACCATCACTAAGCCC
SEQ ID NO:46 (CsgF-His_pET22b_FW) cccccatatgGGAACCATGACTTTCCAGTTCC
SEQ ID NO:47: (CsgF-His_pET22b_Rev) ccccGAATTCCTAatggtgatggtgatggtgGTAAAAATCGGTTGAGTTATTTTG
SEQ ID NO:48: (csgEFG_pDONR221_FW)
GGGGACAAGTTTGTACAAAAAAGCAGGCTACCTCAGGCGATAAAGCCATGAAACGTTA
SEQ ID NO:49: (csgEFG_pDONR221_Rev)
GGGGACCACTTTGTACAAGAAAGCTGGGTGTTTAAACTCATTTTTCGAACTGCGGGTGGCTCCAAGC
GCTGG
SEQ ID NO:50: (Mut_csgF_His_FW) CAAAATAACTCAACCGATTTTcatcaccatcaccatcacTAAGCCCCAGCTTCATAAGG
SEQ ID NO:51: (Mut_csgF_His_Rev)
CCTTATGAAGCTGGGGCTTAgtgatggtgatggtgatgAAAATCGGTTGAGTTATTTTG
SEQ ID NO:52: (DelCsgE_Rev)
AGCCTGC I I I I I I GTACAAAC
SEQ ID NO:53: (DelCsgE FW)
ATAAAAAATTGTTCGGAGGCTGC
SEQ ID NO:54 (>P0AE98 (20 : 50); mature peptide of CsgF 1 :30)
GTMTFQFRNPNFGGNPNNGAFLLNSAQAQN
SEQ ID NO:55 (>P0AE98 (20 : 54); mature peptide of CsgF 1 :35)
GTMTFQFRNPNFGGNPNNGAFLLNSAQAQNSYKDP
Examples of CsgF sequences with protease cleavage sites made into proteins. Signal peptide is shown in bold TEV protease cleavage site in bold and underline and HCV C3 protease cleavage site in underline. StrepII indicate the Strep tag at the C terminus, H10 indicates the lOxHistidine tag at the C terminus and ** indicates STOP codons.
SEQ ID NO:56 Pro-CsgF-Eco-(WT-T4C/N17S/P35-TEV-S36)-StrepII
MRVKHAVVLLMLISPLSWAGTMCFOFRNPNFGGNPSNGAFLLNSAOAONSYKDPENLYFOSSYND DFGIETPSALDNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQV SG LQ N N STD FS A WS H PQ FE K * *
SEQ ID NO:57 Pro-CsgF-Eco-(WT-N17S-Del(P35-[TEV]-S36)-StrepII
MRVKHAWLLMLISPLSWAGTMTFOFRNPNFGGNPSNGAFLLNSAQAQNSYKDPENLYFOSSYND DFGIETPSALDNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQV SG LQ N N STD FS A WS H PQ FE K * *
SEQ ID NO:58 Pro-CsgF-Eco-(WT-GlC/N17S/P35-[TEV]-S36)-StrepII
MRVKHAWLLMLISPLSWACTMTFOFRNPNFGGNPSNGAFLLNSAQAQNSYKDPENLYFOSSYND
DFGIETPSALDNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQV SGLQNNSTDFSAWSHPQFEK**
SEQ ID NO:59 Pro-CsgF-Eco-(WT-GlC/P35-[TEV]-S36)-StrepII
MRVKHAWLLMLISPLSWACTMTFOFRNPNFGGNPNNGAFLLNSAQAQNSYKDPENLYFOSSYND
DFGIETPSALDNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQV SGLQNNSTDFSAWSHPQFEK**
SEQ ID NO:60 Pro-CsgF-Eco-(WT-T45-TEV-P46)-H10
VSGLQNNSTDFHHHHHHHHHH**
SEQ ID NO:61 Pro-CsgF-Eco-(WT-P35-TEV-S36)-H10
MRVKHAVVLLMLISPLSWAGTMTFOFRNPNFGGNPNNGAFLLNSAOAONSYKDPENLYFOSSYND
DFGIETPSALDNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQV SGLQNNSTDFHHHHHHHHHH**
SEQ ID NO:62 Pro-CsgF-Eco-(WT-N30-TEV-S31)-H10
MRVKHAVVLLMLISPLSWAGTMTFOFRNPNFGGNPNNGAFLLNSAOAONENLYFOSSYKDPSYND
DFGIETPSALDNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQV SGLQNNSTDFHHHHHHHHHH**
SEQ ID NO:63 Pro-CsgF-Eco-(WT-T45-TEV-F51)-H10
NNSTDFHHHHHHHHHH**
SEQ ID NO:64 Pro-CsgF-Eco-(WT-N30-TEV-Y37)-H10
MRVKHAWLLMLISPLSWAGTMTFOFRNPNFGGNPNNGAFLLNSAQAQNENLYFOSYNDDFGIET
PSALDNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQVSGLQN
NSTDFHHHHHHHHHH**
SEQ ID NO:65 Pro-CsgF-Eco-(WT-D34-[C3]-S36)
MRVKHAVVLLMLISPLSWACTMTFQFRNPNFGGNPNNGAFLLNSAQAQNSYKDLEVLFQGPSYND DFGIETPSALDNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQV SGLQNNSTDFSAWSHPQFEK**
SEQ ID NO:66 Pro-CsgF-Eco-(WT-I42-[C3]-E43)
MRVKHAVVLLMLISPLSWACTMTFQFRNPNFGGNPNNGAFLLNSAQAQNSYKDPSYNDDFGILEVL FQGPETPSALDNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQ VSGLQNNSTDFSAWSHPQFEK**
SEQ ID NO:67 Pro-CsgF-Eco-(WT-N38-[C3]-S47)
MRVKHAVVLLMLISPLSWACTMTFQFRNPNFGGNPNNGAFLLNSAQAQNSYKDPSYNLEVLFQGPS ALDNFTQAIQSQILGGLLSNINTGKPGRMVTNDYIVDIANRDGQLQLNVTDRKTGQTSTIQVSGLQNNS TDFSAWSHPQFEK**
SEQ ID NO: 68
MPRAQSYKDLTHLPMPTGKIFVSVYNIQDETGQFKPYPASNFSTAVPQSATAMLVTALKDSRWFIPLERQ GLQNLLNERKIIRAAQENGTVAINNRIPLQSLTAANIMVEGSIIGYESNVKSGGVGARYFGIGADTQYQL DQIAVNLRVVNVSTGEILSSVNTSKTILSYEVQAGVFRFIDYQRLLEGEIGYTSNEPVMLCLMSAIETGVI FLINDGIDRGLWDLQNKAERQNDILVKYRHMSVPPES
SEQ ID NO: 69
CLTAPPKQAAKPTLMPRAQSYKDLTHLPAPTGKIFVSVYNIQDETGQFKPYPASNFSTAVPQSATAMLVT ALKDSRWFIPLERQGLQNLLNERKIIRAAQENGTVAMNNRIPLQSLTAANIMVEGSIIGYESNVKSGGVG ARYFGIGADTQYQLDQIAVNLRVVNVSTGEILSSVNTSKTILSYEVQAGVFRFIDYQRLLEGEIGYTSNEP VMLCLMSAIETG
SEQ ID NO: 70
CLTAPPKEAAKPTLMPRAQSYKDLTHLPIPTGKIFVSVYNIQDETGQFKPYPASNFSTAVPQSATAMLVTA LKDSRWFVPLERQGLQNLLNERKIIRAAQENGTVAINNRIPLQSLTAANIMVEGSIIGYESNVKSGGVGA RYFGIGADTQYQLDQIAVNLRVVNVSTGEILSSVNTSKTILSYEVQAGVFRFIDYQRLLEGEIGYTSNEPV MLCLMSAIETGVIFLINDGIDRGLWDLQNKADRQNDILVKYRHMSVPPES
SEQ ID NO: 71
CLTTPPKEAAKPTLMPRAQSYKDLTHLPVPTGKIFVSVYNIQDETGQFKPYPASNFSTAVPQSATAMLVTA LKDSRWFIPLERQGLQNLLNERKIIRAAQENGTVAINNRIPLPSLTAANIMVEGSIIGYESNVKSGGAGA RYFGIGADTQYQLDQIAVNLRVVNVSTGEILSSVNTSKTILSYEVQAGVFRFIDYQRLLEGEIGYTSNEPV MLCLMSAIETGVIFLINDGIDRGLWDLQNKADRQNDILVKYRQMSVPPES
SEQ ID NO: 72
CLTAPPKEAAKPTLMPRAQSYRDLTHLPAPTGKIFVSVYNIQDETGQFKPYPASNFSTAVPQSATAMLVT
ALKDSHWFIPLERQGLQNLLNERKIIRAAQENGTVANNNRMPLQSLAAANVMIEGSIIGYESNVKSGGV
GARYFGIGADTQYQLDQIAVNLRVVNVSTGEVLSSVNTSKTILSYEVQAGVFRFIDYQRLLEGEIGYTSN EPVMMCLMSAIETGVIFLINDGIDRGLWDLQNKADAQNPVLVKYRDMSVPPES
SEQ ID NO: 73
CLTAPPKEAAKPTLMPRAQSYRDLTHLPLPSGKVFVSVYNIQDETGQFKPYPASNFSTAVPQSATAMLVT
ALKDSRWFVPLERQGLQNLLNERKIIRAAQENGTVADNNRIPLQSLTAANVMIEGSIIGYESNVKSGGV GARYFGIGADTQYQLDQIAVNLRVVNVSTGEVLSSVNTSKTILSYEVQAGVFRFVDYQRLLEGEIGYTSN EPVMLCLMSAIETGVIYLINDGIERGLWDLQQKADVDNPILARYRNMSAPPES
SEQ ID NO: 74
CLTAPPKEAAKPTLMPRAQSYRDLTNLPDPKGKLFVSVYNIQDETGQFKPYPASNFSTAVPQSATSMLVT
ALKDSRWFIPLERQGLQNLLNERKIIRAAQENGTVAENNRMPLQSLVAANVMIEGSIIGYESNVKSGGV GARYFGIGGDTQYQLDQIAVNLRVVNVSTGEVLSSVNTSKTILSYEVQAGVFRFIDYQRLLEGEIGYTAN EPVMLCLMSAIETGVIHLINDGINRGLWELKNKGDAKNTILAKYRSMAVPPES
SEQ ID NO: 75
CLTAAPKEAARPTLLPRAPSYTDLTHLPSPQGRIFVSVYNIQDETGQFKPYPACNFSTAVPQSATAMLVSA
LKDSKWFIPLERQGLQNLLNERKIIRAAQENGSVAINNQRPLSSLVAANILIEGSIIGYESNVKSGGVGA
RYFGIGASTQYQLDQIAVNLRAVDVNTGEVLSSVNTSKTILSYEVQAGVFRFIDYQRLLEGELGYTTNEP VMLCLMSAIESGVIYLVNDGIERNLWQLQNPSEINSPILQRYKNNIVPAES
SEQ ID NO: 76
CITSPPKQAAKPTLLPRSQSYQDLTHLPEPQGRLFVSVYNISDETGQFKPYPASNFSTSVPQSATAMLVS
ALKDSNWFIPLERQGLQNLLNERKIIRAAQENGTVAVNNRTQLPSLVAANILIEGSIIGYESNVKSGGAG
ARYFGIGASTQYQLDQIAVNLRVVNVSTGEVLSSVNTSKTILSYEFQAGVFRYIDYQRLLEGEVGYTVNE PVMLCLMSAIETGVIYLVNDGISRNLWQLKNASDINSPVLEKYKSIIVP
SEQ ID NO: 77
CLTAPPKQAAKPTLMPRAQSYQDLTHLPEPAGKLFVSVYNIQDETGQFKPYPASNFSTAVPQSATAMLVS
ALKDSGWFIPLERQGLQNLLNERKIIRAAQENGTAAVNNQHQLSSLVAANVLVEGSIIGYESNVKSGGA
GARFFGIGASTQYQLDQIAVNLRVVDVNTGQVLSSVNTSKTILSYEVQAGVFRYIDYQRLLEGEIGYTTN EPVMLCVMSAIETGVIYLVNDGINRNLWTLKNPQDAKSSVLERYKSTIVP
SEQ ID NO: 78
CITTPPQEAAKPTLLPRDATYKDLVSLPQPRGKIYVAVYNIQDETGQFQPYPASNFSTSVPQSATAMLVSS LKDSRWFVPLERQGLNNLLNERKIIRAAQQNGTVGDNNASPLPSLYSANVIVEGSIIGYASNVKTGGFG ARYFGIGGSTQYQLDQVAVNLRIVNVHTGEVLSSVNTSKTILSYEIQAGVFRFIDYQRLLEGEAGFTTNEP VMTCLMSAIEEGVIHLINDGINKKLWALSNAADINSEVLTRYRK
SEQ ID NO: 79
ITEVPKEAAKPTLMPRASTYKDLVALPKPNGKIIVSVYSVQDETGQFKPLPASNFSTAVPQSGNAMLTSAL KDSGWFVPLEREGLQNLLNERKIIRAAQENGTVAANNQQPLPSLLSANVVIEGAIIGYDSDIKTGGAGAR YFGIGADGKYRVDQVAVNLRAVDVRTGEVLLSVNTSKTILSSELSAGVFRFIEYQRLLELEAGYTTNEPV MMCMMSALEAGVAHLIVEGIRQNLWSLQNPSDINNPIIQRYMKEDVP
SEQ ID NO: 80
PETSESPTLMQRGANYIDLISLPKPQGKIFVSVYDFRDQTGQYKPQPNSNFSTAVPQGGTALLTMALLDS EWFYPLERQGLQNLLTERKIIRAAQKKQESISNHGSTLPSLLSANVMIEGGIVAYDSNIKTGGAGARYLG IGGSGQYRADQVTVNIRAVDVRSGKILTSVTTSKTILSYEVSAGAFRFVDYKELLEVELGYTNNEPVNIAL MSAIDSAVIHLIVKGVQQGLWRPANLDTRNNPIFKKY
SEQ ID NO: 81
PDASESPTLMQRGATYLDLISLPKPQGKIYVSVYDFRDQTGQYKPQPNSNFSTAVPQGGTALLTMALLD SEWFYPLERQGLQNLLTERKIIRAAQKKQESISNHGSTLPSLLSANVMIEGGIVAYDSNIKTGGAGARYL GIGGSGQYRADQVTVNIRAVDVRSGKILTSVTTSKTILSYELSAGAFRFVDYKELLEVELGYTNNEPVNIA LMSAIDSAVIHLIVKGIEEGLWRPENQNGKENPIFRKY
SEQ ID NO: 82
PETSKEPTLMARGTAYQDLVSLPLPKGKVYVSVYDFRDQTGQYKPQPNSNFSTAVPQGGAALLTTALLD SRWFMPLEREGLQNLLTERKIIRAAQKKDEIPTNHGVHLPSLASANIMVEGGIVAYDTNIQTGGAGARYL GVGASGQYRTDQVTVNIRAVDVRTGRILLSVTTSKTILSKELQTGVFKFVDYKDLLEAELGYTTNEPVNL AVMSAIDAAVVHVIVDGIKTGLWEPLRGEDLQHPIIQEYMNRSKP
SEQ ID NO: 83
CATHIGSPVADEKATLMPRSVSYKELISLPKPKGKIVAAVYDFRDQTGQYLPAPASNFSTAVTQGGVAML STALWDSQWFVPLEREGLQNLLTERKIVRAAQNKPNVPGNNANQLPSLVAANILIEGGIVAYDSNVRTG GAGAKYFGIGASGEYRVDQVTVNLRAVDIRSGRILNSVTTSKTVMSQQVQAGVFRFVEYKRLLEAEAGF STNEPVQMCVMSAIESGVIRLIANGVRDNLWQLADQRDIDNPILQEYLQDNAP
SEQ ID NO: 84
ASSSLMPKGESYYDLINLPAPQGVMLAAVYDFRDQTGQYKPIPSSNFSTAVPQSGTAFLAQALNDSSWF IPVEREGLQNLLTERKIVRAGLKGDANKLPQLNSAQILMEGGIVAYDTNVRTGGAGARYLGIGAATQFRV DTVTVNLRAVDIRTGRLLSSVTTTKSILSKEITAGVFKFIDAQELLESELGYTSNEPVSLCVASAIESAVVH MIADGIWKGAWNLADQASGLRSPVLQKY
SEQ ID NO: 85
QDSETPTLTPRASTYYDLINMPRPKGRLMAVVYGFRDQTGQYKPTPASSFSTSVTQGAASMLMDALSAS GWFVVLEREGLQNLLTERKIIRASQKKPDVAENIMGELPPLQAANLMLEGGIIAYDTNVRSGGEGARYLG IDISREYRVDQVTVNLRAVDVRTGQVLANVMTSKTIYSVGRSAGVFKFIEFKKLLEAEVGYTTNEPAQLC VLSAIESAVGHLLAQGIEQRLWQV
SEQ ID NO: 86
MPKSDTYYDLIGLPHPQGSMLAAVYDFRDQTGQYKAIPSSNFSTAVPQSGTAFLAQALNDSSWFVPVER EGLQNLLTERKIVRAGLKGEANQLPQLSSAQILMEGGIVAYDTNIKTGGAGARYLGIGVNSKFRVDTVTV NLRAVDIRTGRLLSSVTTTKSILSKEVSAGVFKFIDAQDLLESELGYTSNEPVSLCVAQAIESAVVHMIAD GIWKRAWNLADTASGLNNPVLQKY
SEQ ID NO: 87
LTRRMSTYQDLIDMPAPRGKIVTAVYSFRDQSGQYKPAPSSSFSTAVTQGAAAMLVNVLNDSGWFIPLE
REGLQNILTERKIIRAALKKDNVPVNNSAGLPSLLAANIMLEGGIVGYDSNIHTGGAGARYFGIGASEKY RVDEVTVNLRAIDIRTGRILHSVLTSKKILSREIRSDVYRFIEFKHLLEMEAGITTNDPAQLCVLSAIESAV AHLIVDGVIKKSWSLADPNELNSPVIQAYQQQRI
SEQ ID NO: 88
PSDPERSTMGELTPSTAELRNLPLPNEKIVIGVYKFRDQTGQYKPSENGNNWSTAVPQGTTTILIKALED SRWFIPIERENIANLLNERQIIRSTRQEYMKDADKNSQSLPPLLYAGILLEGGVISYDSNTMTGGFGARYF GIGASTQYRQDRITIYLRAVSTLNGEILKTVYTSKTILSTSVNGSFFRYIDTERLLEAEVGLTQNEPVQLAV TEAIEKAVRSLIIEGTRDKIW
SEQ ID NOs: 89 to 116 are CsgF peptides that may be used in the invention.
SEQ ID NO: 117 is SEQ ID NO: 3 with a W at position 97.
CLTAPPKEAARPTLMPRAQSYKDLTHLPAPTGKIFVSVYNIQDETGQFKPYPASNFSTAVPQSATAMLVT
ALKDSRWFIPLERQGLQNLLNERKIIWAAQENGTVAINNRIPLQSLTAANIMVEGSIIGYESNVKSGGVG ARYFGIGADTQYQLDQIAVNLRVVNVSTGEILSSVNTSKTILSYEVQAGVFRFIDYQRLLEGEVGYTSNEP VMLCLMSAIETGVIFLINDGIDRGLWDLQNKAERQNDILVKYRHMSVPPES
SEQ ID NO: 118 (>P32270; Enterobacteria phage T4)
MTFDDLTEGQKNAFNIVMKAIKEKKHHVTINGPAGTGKTTLTKFIIEALISTGGTGIILA
APTHAAKKILSKLSGKEASTIHSILKINPVTYEENVLFEQKEVPDLAKCRVLICDEVSMY DRKLFKILLSTIPPWCTIIGIGDNKQIRPVEPGENTAYISPFFTHKDFYQCELTEVKRSN APIIDVATDVRNGKWNYDKVVDGHGVRGFTGDTALRDFMVNYFSIVKSLDDLFENRVMAF
TNKSVDKLNSIIRKKIFETDKDFIVGEIIVMQEPLFKTYKIDGKPVSEIIFNNGQLVRII
EAEYTSTFVKARGVPGEYLIRHWDLTVETYGDDEYYREKIKIISSDEELYKFNLFLAKTA
ETYKNWNKGGKAPWSDFWDAKSQFSKVKALPASTFHKAQGMSVDRAFIYTPCIHYADVEL AQQLLYVGVTRGRYDVFYV
SEQ ID NO: 119 (>D0MKQ2; Rhodothermus marinus)
MEELSNEQQRVLDHVLAWLERNDAPPIFILTGSAGTGKTLLIRHLVRALQDRRIHYALAA
PTGRAARILSERTGDHARTLHSLIYIFDRYQLVEEADRQTDEPLSLQLHFALRSAEHDAR
LIIVDEASMVSDTAGEEELYRFGSGRLLNDLLTFARLIPKRDRPPTTRLLFVGDPAQLPP
VGQSVSPALSAQYLRDTFGLSAETAHLRSVYRQRKGHPILETATALRNALEKGHYHTFRL
PEQPPDLRPVGLEEAIETTATDFRRQNPSVLLCRTNALARKLNAAVRARLWGREGLPPQP
GDLLLVNRNAPLHGLFNGDLVLVETVGPLEHRRVGRRGRPPVDLYFRDVELLYPHEKPRN
RIRCKLLENLLESPDGQLSPDIIQALLIDFYRRHPSLKHGSSEFRLMLANDAYFNALHVR
YGYAMTVHKAQGGEWKRATVVFNDWRHFRHAEFFRWAYTAITRAREELLTIGAPSFEALS
DMRWQPAPSVPAPEQAAENATRFPLKALETYHQRLSEALTAAGIETTGVELLQYAVRYHL
ARADRTTRIQYYYRGDGQISRIVTLGGADDPELTQQAYALFERILSEPPADSGELPENPL LREFLERAHLRLEGSGIRIVHWKEMPYALRLYFSADGENVTIDFYYNRRGVWTHAQEVGR SSSGALFARIQSLLQADS
SEQ ID NO: 120 (>B1X365; Cyanothece ATCC51142)
MSQSVVVPDELGEHTAVIEFYQDAVDKIEPKIVFLELRKNVVDWVSRTQLKIEEKEIQA
TGLTRQQQTAYKEMINFIENSSEQYFRLSGYAGTGKSFLMAKVIEWLKQEDYKYSVAAPT
NKAAKNLTQIARSQGIKIEATTVAKLLKLQPTIDVDTGQQSFEFNSEKELELKDYDVIII
DEYSMLNKDNFRDLQQAVKGGESKFIFVGDSSQLPPVKEKEPIVANHPDIRKSANLTQIV
RYDGEIVKVAESIRRNPRWNHQTYPFETVADGTIIKLNTEDWLQQALSHFEKEDWLSNPD
YVRMITWRNKTADKYNQAIREALYGENVEQLVVGDRLIAKKPVFRSLPGGKKKEKKIILN
NSEECKVIETPKINYNEKYKWEFYQVKVRTDEGGMIELRILTSESEEKRQKKLKELAKRA REEENYSEKKKQWAIYYELDELFDNMAYAYALTCHKAQGSSIDNVFLLVSDMHYCRDKTK MIYTGLTRAKKCCYVG
SEQ ID NO: 121 (>Q2S429; Salinibacter ruber)
MSTFADAPFTEDQEEAYDHVYDRLAQGERFTGLRGYAGTGKTYLVSRLVEQLLDEDCTVT
VCAPTHKAVQVLSDELGDAPVQMQTLHSFLGLRLQPKQDGEYELVAEEERNFAEGVVIVD EASMIGREEWSHIQDAPFWVQWLFVGDPAQLPPVNEDPSPALDVPGPTLETIHRQAADNP ILELATKIRTGADGRFGSTFEDGKGVAVTRNREEFLDSILRAFDADAFAEDATHARVLAY
RNKTVRRYNREIRAERYGADADRFVEGEWLVGTETWYYDGVQRLTNSEEVRVKKAQVETF EADDQSEWTVWELKIRTPGRGLTRTIHVLHEEERERYENALERRRGKAEDDPSKWDRFFE LRERFARVDYAYATTVHRAQGSTYDTVFVDHRDLRVCRGEERGALLYVAVTRPSRRLALLV
SEQ ID NO: 122 (>B6BJ43 (UPI000183B2F5); Sullfurimonas gotlandica GDI) MKILNKETYKLSLHQEEVFTQIVSQLDTKVSSILKSTNIEDYLLSLTGPAGTGKTFLTTQ IAKYLVEKRKESEYPMSSDFDFTITAPTHKAVGVLSKLLRENNIQSSCKTIHSFLGIKPF IDYTTGEEKFVVDKTNKRKDRTSILIVDESSMIGNTLYEYILEAIEDKRVNVVLFIGDPY QLLPIENSKNEIYDLPNRFFLSEVVRQAENSYIIRVATKLRERIKNQDFISLQQFFQENM EDEITFFHNKEAFLEDFYKEEEWYKENKILATYKNKDVDAFNKIIRNKFWEQKGNTTPST LLAGDMIRFKDAYTVGDITIYHNGQELQLGSTEVKYHDSLHIEYWECKSIYALEQQVFRV VNPDSEAVFNQKLQSLATKAKQAKFPDNKKLWKLYYETRNMFANVQYIHASTIHKLQGST YDVSYIDIFSLVHNHYMSDEEKYRLLYVAITRASKDIKIFMSAFDRTSDEKVIINNQNSE TMNTLKQLHDIDIILKDLDL
SEQ ID NO: 123 (>M4MBC3; Vibrio phage henriette 12B8)
MADFELTLGQKTVLGEVISTILKPVNLNDTSRFHTMHGPAGSGKTTVLQRIISQIPAYKT IGFCSPTHKSVKVIRRMAREAGISHRVDIRTIHSALGLVMKPVRGDEVLVKEPFAEERIY DVLIIDEAGMLNDELIMYILESQSSKVIFVGDMCQIGPIQSNLPEEDGYTPTSTDDVSKV FTEVEMMSALTEVVRQAEGSPIIQLATEFRLAQDDIYADLPRIVTNTTPDGNGIITMPNG NWVDSAVARFQSDQFKEDPDHCRIVCYTNAMVDLCNDLVRKRLFGADVPEWLEDEILVAQ EMGSTWNNADELRIVSIDDHFDQQYEVPCWRMQLESVEDHKLHNALVVKGDYIEDFKFRL NAIAERANTDKNMSGMHWKEFWGMRKKFNTFKNVYAGTAHKSQGSTFDYTYVFTPDFYKF GATMTIKRLLYTAITRSRYTTYFAMNTGAQ
SEQ ID NO: 124 (>I6XGX8; Vibrio phage phi-pp2)
MGLTNCQQGAMDAFLESDGHMTISGPAGSGKTFLMKSILEALESKGKNVTMVTPTHQAKN VLHKATGQEVSTIHSLLKIHPDTYEDQKHFTQSGEVEGLDEIDVLVVEEASMVDEELFQI TGRTMPRKCRILAVGDKYQLQPVKHDPGVISPFFTKFTTFEMNEVVRQAKDNPLIQVATE VRNGQWLRTNWSKERRQGVLHVPNVNKMLDTYLSKVNSPEDLLDYRILAYTNDCVDTFNG IIREHVYNTSEPFIPGEYLVTQMPVMVSNGKYPVCVIENGEVVKILDVRQKTIDGMLPKV DNEAFDVAVLTVEKEDGNVYEFTVLWDDLQKERFARYLSVAAGTYKSMRGNTKRYWRAFW GLKEQMIETKSLGASTVHKSQGTTVKGVCLYTQDMGYAEPEILQQLVYVGLTRPTDWALY N
SEQ ID NO: 125 (>E5DRP6; Aeromonas phage 65)
MSESEITLTPSQNMAVNEVKNGTGHITISGPPGSGKTFLVKYLIKMLGDELGTVLAAPTH QAKIVLTEMSGIEACTIHSLMKIHPETLEDIQIFDQSKLPDLSNIRYLIVEEASMHSKTL FKITMKSIPPTCRIIAIGDKDQIQPEEHAQGELSPYFTDPRFSQIRLTDIMRQSLDNPII
QVATKIREGGWIEPNWNRDTKTGVYKVSGITDLVNSYLRAVKTPEDLTKYRFLAYTNKVV
NKVNSIVREHVYKTKLPFIEGEKIVLQEPVMVEHEDDTIETIFTNGEVVTINEIEVFDRT
IRIDGSPEFKVNAAKLSVSSDYSGIEHDFCVLYGSESRLEFEYQLSESAGNIKQMGKGGN QRSAWKSFWAAKKMFIETKSLGASTIHKSQGSTVKGVWLALHDIHYADEELKQQLVYVGV TRPTDFCLYFDGTK
SEQ ID NO: 126 (>I6XH64; Aeromonas phage CC2)
MAVDAVQSGTGHITISGPPGSGKTFLVKYIIKMLGDELGTVLAAPTHQAKIVLTEMSGIE
ACTIHSLMKIHPETLEDIQIFDQSKMPDLSTVRYLIIEEASMHSKALFNITMKSIPPTCR
IIAIGDKDQIQPVDHAPGELSPYFTDSRFTQIRMTDIMRQSLDNPIIQVATTIREGGWIY
QNWNKEKKSGVYKVKSITDLINSYLRVVKTPEDLTKYRFLAFTNKVVDKVNSIVRKHVYK
TDLPFIEGEKLVLQEPVMVEYDDDTIETIFTNGEVVTVDEIEVSDMNIRIDGSPAFSISV AKLKVTSDFSGVTHDIMSVYGEDSKAEFNYQLSEAAAVIKQMQRGQTKAAWASFWDAKKT FTETKSLGACTIHKSQGSTVKGVWLGLHDISYADTDLQQQLVYVGVTRPTDFCLYFDGSK
SEQ ID NO: 127 (>K4FBD0; Cronobacter phage vB CsaM GAP161)
MSELTFDDLSDDQKSAHDRVIHNIQNAIHTTITGGPGVGKTTLVKFVFNTLKGLGISGIW
LTAPTHQAKNVLAAATGMDATTIHSALKISPVTNEELRVFEQQKGKKAPDLSTCRVFVVE
EVSMVDMDLFRIIRRSIPSNAVILGLGDKDQIRPVNADGRVELSPFFDEEIFDVIRMDKI
MRQAEGNPIIQVSRAVRDGKMLKPMSVGDLGVFQHANAVDFLRQYFRRVKTPDDLIENRM
FAYTNDNVDKLNATIRKHLYKTTEPFILDEVIVMQEPLVQEMRLNGQIFTEIVYNNNEKI
RVLEIIPRREVIKAEKCDEKIEIEFYLLKTVSLEEETEAQIQVVVDPVMKDRLGNYLAYV
ASTYKRIKQQTGYKAPWHSFWAIKNKFQDVKPLPVCTYHKSQGSTYDHAYMYTRDAYAFA DYDLCKQLIYVGVTRARYTVDYV
SEQ ID NO: 128 (>D5JF67; Klebsiella phage KP15)
MSELTFDDLSEDQKNAHDRVIKNIRNKIHTTITGGPGVGKTTLVKFVFETLKKLGISGIW
LTAPTHQAKNVLSEAVGMDATTIHSALKISPVTNEELRVFEQQKGKKAADLSECRVFVVE
EVSMVDKELFRIIKRTIPSCAVILGLGDKDQIRPVNTEGITELSPFFDEEIFDVIRMDKI
MRQAEGNPIIQVSRAIRDGKPLMPLMNGELGVMKHENASDFLRRYFSRVKTPDDLNNNRM
FAYTNANVDKLNAVIRKHLYKTDQPFIVGEVVVMQEPLVTEGRVNGVSFVEVIYNNNEQI
KILEIIPRSDTIKADRCDPVQIDYFLMKTESMFEDTKADIQVIADPVMQERLGDYLNYVA
FQYKKMKQETGYKAPWYSFWQIKNKFQTVKALPVCTYHKGQGSTYDHSYMYTRDAYAYAD YELCKQLLYVGTTRARFTVDYV
SEQ ID NO: 129 (>J7HXT5; Stenotrophomonas phage IME13)
MVTYDDLTVGQKDAIEKALQAMRTKRHITIRGPAGSGKTTMTRFLLERLFQTGQQGIVLT
APTHQAKKELSKHALRKSYTIQSVLKINPSTLEENQIFEQKGTPDFSKTRVLICDEVSFY
TRKLFDILMRNVPSHCVVIGIGDKAQIRGVSEDDTHELSPFFTDNRFEQVELTEVKRHQG
PIIEVATDIRNGKWIYEKLDDSGNGVKQFHTVKDFLSKYFERTKTPNDLLENRIMAYTNN SVDKLNSVIRKQLYGANAAPFLPDEILVMQEPLMFDIDIGGQTLKEVIFNNGQNVRVINV KPSRKTLKAKGVGEIEVECTMLECESYEEDEDDYRRAWFTVVHDQNTQYAINEFLSIIAE KYRSREVFPNWKDFWAIRNTFVKVRPLGAMTFHKSQGSTFDNAYLFTPCLHQYCRDPDVA
QELIYVGNTRARKNVCFV
SEQ ID NO: 130 (>E5EYE6; Acinetobacter phage Ac42)
MNFEDLTEGQKNAYTAAIKAIETVPSSSAEKRHLTINGPAGTGKTTLTKFLIAELIRRGE
RGVYLAAPTHQAKKVLSQHAGMEASTIHSLLKINPTTYEDSTTFEQKDVPDMSECRVLIC
DEASMYDLKLFQILMSSIPLCCTVIALGDIAQIRPVEPGAFEGQVSPFFTYEKFEQVSLT EVMRSNAPIIDVATSIRTGNWIYENVIDGAGVHNLTSERSVKSFMEKYFSIVKTPEDLFE NRLLAFTNKSVDDLNKIVRKKIYNTLEPFIDGEVLVMQEPLIKSYTYEGKKVSEIVFNNG EMVKVLCCSQTSDEISVRGCSTKYMVRYWQLDLQSLDDPDLTGSINVIVDEAEINKLNLV
LGKSAEQFKSGAVKAAWADWWKLKRNFHKVKALPCSTIHKSQGTSVDNVFLYTPCIHKAD SQLAQQLLYVGATRARH N VYYI
SEQ ID NO: 131 (>E3SFA5; Shigella phage SP18)
MIKFEDLNTGQKEAFDYITEAIQRRSGECITLNGPAGTGKTTLTKFVIDHLVRNGVMGIV
LAAPTHQAKKVLSKLSGQTANTIHSILKINPTTYEDQNIFEQREMPDMSKCNVLVCDEAS MYDGSLFKIICNSVPEWCTILGIGDMHQLQPVDPGSTQQKISPFFTHPKFKQIHLTEVMR SNAPIIEVATEIRNGGWFRDCMYDGHGVQGFTSQTALKDFMVNYFGIVKDADMLMENRMY AYTNKSVEKLNNIIRRKLYETDKAFLPYEVLVMQEPHMKELEFEGKKFSETIFNNGQLVR IKDCKYTSTILRCKGESHQLVINYWDLEVESIDEDEEYQVDRIKVLPEDQQPKFQAYLAK VADTYKQMKAAGKRPEWKDFWKARRTFLKVRALPVSTIHKAQGVSVDKAFIYTPCIHMAE
AS LASQ LAYVG ITRA RYD AYYV
SEQ ID NO: 132 (>I7J3V8; Yersinia phage phiRl-RT)
MITYDDLTDGQKSAFDNTMEAIKNKKGHITINGPAGTGKTTLTKFIIDHLIKTGEAGIIL
CAPTHQAKKVLSKLSGMDASTIHSVLKINPTTYEENQIFEQREVPDLAACRVLICDEASF
YDRKLFGIILATVPSWCTVIALGDKDQLRPVTPGESEQQLSPFFSHAKFKQVHLTEIKRS NGPIIQVATDIRNGGWLSENIVDGEGVHAFNSNTALKDFMIRYFDVVKTADDLIESRMLA YTNKSVDKLNGIIRRKLYETDKPFINGEVLVMQEPLMKELEFDGKKFHEIVFNNGQLVKI LYASETSTFISARNVPGEYMIRYWNLEVETADSDDDYATSQIQVICDPAEMTKFQMFLAK
TADTYKNSGVKAYWKDFWSVKNKFKKVKALPVSTIHKSQGCTVNNTFLYTPCIHMADAQL AKQLLYVG ATRARTN LYYI
SEQ ID NO: 133 (>M1EA88; Salmonella phage S16)
MITFEQLTSGQKLAFDETIRAIKEKKNHVTINGPAGTGKTTLTKFIMEHLVSTGETGIIL TAPTHAAKKVLTKLSGMEANTIHKILKINPTTYEESMLFEQKEVPDLASCRVLICDEASM
WDRKLFKILMASIPKWCTIVAIGDVAQIRPVDPGETEAHISPFFIHKDFKQLNLTEVMRS NAPIIDVATDIRNGSWIYEKTVDGHGVHGFTSTTALKDFMMQYFSIVKSPEDLFENRMLA FTNKSVDKLNSIIRRRLYQTEEAFVVGEVIVMQEPLMRELVFEGKKFHETLFTNGQYVRI LSADYTSSFLGAKGVSGEHLIRHWVLDVETYDDEEYAREKINVISDEQEMNKFQFFLAKT ADTYKNWNKGGKAPWSEFWDAKRKFHKVKALPCSTFHKAQGISVDSSFIYTPCIHVSSDN KFKLELLYVGATRGRHDVFFV
The following Examples illustrate the invention.
EXAMPLES
Detailed methods for making and testing mutant CsG pores and mutant CsgG/CsGF complexes are described in WO2016/034591, WO2017/149316, WO2017/149317, WO2017/149318, WO2018/211241 and W02019/002893 (all incorporated by reference herein in their entirety). Detailed methods for making modified Dda helicases are described in WO2015/055981, WO2015/166276 and WO2016/055777 (all incorporated herein by reference in their entirety).
Example 1
This example describes a method of comparing speed, accuracy and speed spread ratio properties of variant polynucleotide binding proteins controlling the movement of a polynucleotide against a control variant polynucleotide binding protein controlling the movement of a polynucleotide using a nanopore described in WO 2019/002893.
Barcoded 3.6 kb Analyte Preparation
A double stranded 3.6 kb DNA analyte was prepared using specific primers and PCR. The PCR product was subjected to NEBNext end repair and NEBNext dA-tailing modules (New England Biolabs (NEB)), to generate 3' dA overhangs. Barcodes were introduced into the analyte using EXP-PBC001 (Oxford Nanopore Technologies), following the manufacturer's guidelines.
Variant sequencing adaptor preparation
Recombinant expression vectors encoding the variants of polynucleotide binding proteins described in WO2016/055777 and PCT/EP2023/059821, with an N-terminal affinity and solubility tag were transformed into chemically competent E.coli cells. The cells were plated onto an LB agar plate containing antibiotics. Colonies from the agar plate were inoculated into LB growth media, grown to OD 0.400 - 0.800 and induced, then grown for a further 16 hours at 18°C. The cells were either lysed with Bugbuster extraction reagent (Merck 70921) in the presence of lysozyme, benzonase and protease inhibitors or lysed by sonication in the presence of benzonase and protease inhibitors. The supernatant was further purified using affinity chromatography. Purified variant polynucleotide binding protein was bound to
sequencing Y adaptors described in WO 2015/110813A1 (herein incorporated by reference in its entirety) in 50 mM Hepes pH8.0, lOOmM Potassium acetate, ImM EDTA for 10 minutes at ambient temperature. TMAD (SIGMA) was added to a final concentration of 100 pM and incubated for an hour at 34°C. 10 mM ATP, 10 mM MgCI? and 0.5 M NaCI were added and incubated for 10 minutes at ambient temperature. The variant polynucleotide binding protein bound sequencing adaptors were purified using Sera-Mag SpeedBeads (Thermo Scientific). These were the variant sequencing adaptors.
Control sequencing adaptor preparation
A variant of the polynucleotide binding protein described in WO2016/055777 was used as a control for each of the variant positions. The control polynucleotide binding proteins were purified and loaded onto sequencing adaptors as described above. These were the control sequencing adaptors.
Library preparation
The variant sequencing adaptors and control sequencing adaptors were ligated to barcoded 3.6 kb analytes, each variant was ligated to a different barcode, and each control was ligated to a different barcode. The variant libraries and control libraries were pooled. The pooled library was prepared for sequencing and run on a MinlON flow cell following the manufacturer's guidelines (Oxford Nanopore Technologies). Up to 6 variants were run on a single MinlON flow cell with their control, this control was the internal flow cell control.
Electrical Measurements
Electrical measurements were acquired on FL0-PRO114M PromethlON Flow Cells and PromethlON (Oxford Nanopore Technologies). A standard sequencing script was run and raw data collected in a bulk FAST5 file using MinKNOW software (Oxford Nanopore Technologies).
Data Analysis
The DNA library was sequenced using a standard basecalling algorithm from Guppy (Oxford Nanopore Technologies). The sequenced reads were de-multiplexed using Guppy (Oxford Nanopore Technologies). Greater than or equal to 500 reads for each variant and control were mapped to the 3.6 kb analyte reference sequence.
The speed of an individual DNA strand as it translocates through the pore was calculated by dividing the number of bases mapped to the reference by the duration of the read (measured in bases per second). The median speed (bases per second) was the median speed of multiple individual DNA strands as they translocated through the pore. The median
speed of the variant was divided by the median speed of the control. This was the Speed ratio. The Speed ratio was subtracted from 1 and multiplied by 100. This was the Speed % change. Hence, a positive Speed % change indicated that the variant translocated DNA more quickly than the control.
The variation of speed of multiple individual DNA strands as they translocated through the pore was measured by calculating the interquartile range. Hence, a small interquartile range implied a narrow distribution of speeds, and a large interquartile range implied a broad distribution of speeds. The normalised speed distribution was calculated by dividing the interquartile range of the speed by the median speed. The normalised speed distribution of the variant was divided by the normalised speed distribution of the control, this was the Speed Spread ratio. The Speed Spread ratio was subtracted from 1 and multiplied by 100. This was the Speed Spread % change. Hence, a negative Speed Spread % change indicated that the variant has a narrower distribution of speeds than the control.
The accuracy of an individual DNA strand was calculated by mapping the basecalled sequence to the reference. The median accuracy was the median accuracy of multiple individual DNA strands. The median accuracy of the variant was divided by the median accuracy of the control. This was the Accuracy ratio. The Accuracy ratio was subtracted from 1 and multiplied by 100. This was the Accuracy % change. Hence, a positive Accuracy % change indicated that the variant was more accurate than the control.
The results are shown in Table 8 below. Column 1 identifies the variant based on SEQ ID NO: 118. Columns 2-4 show the % change relative to the control Dda (not having the mutations).
Table 8: Speed, Accuracy and Speed Spread Ratio changes of the modified helicases of the invention.
Claims
1. A modified DNA dependent ATPase (Dda) helicase, wherein the helicase (a) is one in which one or more of the positions corresponding to the following amino acid positions in Dda 1993 are modified or substituted: F3, D4, 116, K24, K25, H26, H27, V28, P33, T42, F44, 146, A48, A60, P62, A65, K67, K68, 169, K72, K76, S79, 184, P104, K108, V117, Y120, R122, L124, 1132, P134, W135, T137, N144, K145, E154, A157, Y158, P161, K166, Q170, T174, V176, N180, A181, V190, Y197, V200, V201, R207, T213, A214, L215, M219, V220, 1225, K227, L229, V237, M238, K247, L248, 1251, R253, K254, F257, D262, V265, D282, 1289, Q295, Y304, S306, V309, V314, G316, L319, R321, T329, G331, D333, L349, F352, T362, N365, N367, G369, G370, P373, A380, Q383, S385, P391, S393, F395, M401, S402, V403, D404, R405, A406, 1413, H414, V418, E419, A421, Q422, V427, V429, G432, Y434, D435, and F437 and/or (b) comprises one or more of the following substitutions or one or more of the following substitutions at the position(s) corresponding to the following amino acid positions in Dda 1993:
- T2N, T2K, T2S, T2D, T2I, T2L, T2E, T2F, T2G, T2H, T2Q, T2R, T2A, T2M, T2P, or T2V,
- E47D, E47G, E47H, E47N, E47P, E47Q, E47S, E47T, E47A, or E47R,
- E54I, E54V, E54K, E54Y, E54T, E54L, E54N, E54A, E54F, E54M, E54D, E54H, E54P, E54Q, E54R, or E54S,
- T55F, T55V, T55H, T55I, T55L, T55M, T55W, T55Y, T55A, T55P, T55Q, or T55R,
- S83E, S83D, S83A, S83P, S83G, or S83M,
- K86N, K86G, K86D, K86E, K86H, K86Q, K86R, K86S, K86T, K86M, or K86P,
- N88K, N88E, N88T, N88S, N88V, N88Q, N88A, N88L, N88D, N88G, N88M, N88I, or N88P,
- P89G, P89K, P89M, P89Q, P89R, or P89D,
- V90E, V90D, V90K, V90Q, V90N, V90S, V90A, V90I, V90L, V90M, V90P, V90R, V90F, V90Y, V90H, or V90G,
- T91E, T91D, T91R, T91K, T91Y, T91H, T91P, T91A, T91V, T91L, T91I, T91S, or T91M,
- Y92K, Y92E, Y92R, Y92P, Y92I, or Y92M,
- E93P, E93D, E93G, E93A, E93L, E93K, E93Q, E93S, E93T, E93H, E93N, or E93R,
V96K, V96T, V96P, V96D, V96G, V96S, V96I, V96N, V96A, V96H, or V96M,
- F98M or F98I,
- K101R, K101T, K101S, K101V, K1O1E, K1O1I, K101G, K1O1L, K101M, K1O1P, K1O1Q, K101D, K1O1H, or K1O1N,
- V103K, V103N, V103L, V103I, V103T, V103R, V103Y, V103S, V103D, V103A, V103E, V103H, V103M, V103P, V103Q, or V103F,
- C114K or C114R,
- D151G, D151V, D151N, D151K, D151T, D151A, D151H, D151P, D151Q, D151R, or D151S,
- G153E, G153S, G153D, G153T, G153N, G153K, G153A, G153Q, G153R, G153L, G153H, or G153P,
- N155K, N155P, N155S, N155E, N155D, N155G, N155T, N155A, N155V, N155L, N155I, N155H, N155M, N155Q, or N155R,
- T156H, T156Q, T156R, or T156W,
- K177A or K177P,
- W195F, W195Y, W195P, W195K, W195E, W195I, W195L, W195D, W195S, W195G, W195T, W195V, W195Q, W195N, W195R, W195M, or W195H,
- I196P, I196L, I196D, I196A, I196E, I196K, I196Q, I196S, I196T, I196F, I196M, I196V, or I196Y,
- D198N, D198P, D198E, D198G, D198H, D198K, D198Q, D198R, D198S, or D198T,
- T210A, T210M, T210P, T210Q, T210I, T210L, T210V, or T210F,
- N221P, N221S, or N221G,
- K243R, K243E, K243D, K243A, K243S, K243M, K243N, K243P, K243T, or K243Y,
- E258G, E258K, E258N, E258P, E258D, E258S, E258A, E258H, E258Q, E258R, or E258T,
- F276T, F276S, F276L, F276P, F276E, F276D, F276N, or F276Q,
- I281E, I281V, I281K, I281A, I281D, I281H, I281N, I281P, I281Q, I281R, I281S, I281T, I281F, I281L, I281M, or I281Y,
- N292E, N292A, N292S, N292D, N292K, N292H, N292R, N292Q, or N292T,
- E301K, E301D, E301N, E301S, E301A, E301H, E301M, E301P, E301Q, E301R, or E301T,
- F308K, F308P, F308R, F308T, F308G, F308E, F308S, F308V, F308A, F308Y, F308N, F308M, F308D, F308H, F308Q, or F308W,
- Y335P, Y335K, Y335E, Y335F, Y335T, Y335Q, Y335S, Y335R, Y335D, Y335V, Y335N, Y335G, Y335H, Y335M, or Y335W,
- Y336E, Y336H, Y336K, Y336I, Y336G, Y336S, Y336T, Y336D, Y336V, Y336P, Y336N, Y336R, Y336Q, Y336A, Y336F, Y336M, or Y336W,
- R337E, R337K, R337T, R337Q, R337P, R337V, R337F, R337S, R337D, R337G, R337Y, R337A, R337H, or R337N,
- Y350P,
- L354K, L354E, L354D, L354V, L354Y, L354R, L354S, L354N, L354H, L354M, L354P, L354T, L354F, or L354I,
- K358G or K358P,
- T359V, T359M, T359A, T359I, T359Y, T359F, T359E, T359K, T359N, T359D, T359P, T359Q, T359R, T359S, T359W, or T359G,
- K364N, K364Q, K364D, K364E, K364G, K364H, K364S, K364T, K364M, K364P, K364Y, K364I, K364L, or K364V,
- W366D, W366N, W366K, W366A, W366S, W366I, W366E, W366R, W366M, W366T, W366Q, W366Y, W366G, W366F, W366V, W366P, or W366H,
- K368A, K368E, K368S, K368G, K368D, K368F, K368M, K368T, K368V, K368Y, K368H, K368N, K368P, K368Q, or K368R, and
- S382R, S382N, S382K, S382E, S382Q, S382A, S382H, S382M, S382T, S382D, S382G, or S382P.
2. A modified DNA dependent ATPase (Dda) helicase according to claim 1, wherein
the position corresponding to amino acid position F3 in Dda 1993 is substituted with K, Y, I, E, G, L, A, S, D, R, V, T, N, H, M, P, Q, or W, the position corresponding to amino acid position D4 in Dda 1993 is substituted with I, L, K, P, S, E, Y, T, F, M, V, A, G, H, N, or Q, the position corresponding to amino acid position 116 in Dda 1993 is substituted with G, A, E, L, K, D, N, S, F, M, T, V, or Y, the position corresponding to amino acid position K24 in Dda 1993 is substituted with G, P, A, D, N, S, E, H, M, Q, R, or T, the position corresponding to amino acid position K25 in Dda 1993 is substituted with A, N, G, D, E, I, L, M, P, Q, R, S, T, V, or H, the position corresponding to amino acid position H26 in Dda 1993 is substituted with L, K, R, Q, S, M, A, T, N, P, G, E, I, V, F, Y, or D, the position corresponding to amino acid position H27 in Dda 1993 is substituted with F, I, V, L, Y, A, T, S, M, W, D, E, K, N, Q, or R, the position corresponding to amino acid position V28 in Dda 1993 is substituted with I, F, A, L, M, T, or Y, the position corresponding to amino acid position P33 in Dda 1993 is substituted with A, Y, F, G, E, I, K, L, M, Q, R, S, T, V, D, H, or W, the position corresponding to amino acid position T42 in Dda 1993 is substituted with I, L, A, F, M, V, Y, D, E, K, N, P, Q, R, or S, the position corresponding to amino acid position F44 in Dda 1993 is substituted with K, A, R, Y, G, H, T, S, Q, L, N, E, M, V, D, P, I, or W, the position corresponding to amino acid position 146 in Dda 1993 is substituted with V, A, F, L, M, T, or Y, the position corresponding to amino acid position A48 in Dda 1993 is substituted with K, H, Y, R, E, N, Q, S, T, D, M, P, G, I, L, V, F, or W, the position corresponding to amino acid position A60 in Dda 1993 is substituted with T, D, E, I, K, L, M, N, P, Q, R, S, V, or G, the position corresponding to amino acid position P62 in Dda 1993 is substituted with S, A, D, E, G, H, K, M, N, Q, R, or T,
the position corresponding to amino acid position A65 in Dda 1993 is substituted with V, Q, K, G, R, Y, L, I, F, T, M, H, E, P, S, D, or N, the position corresponding to amino acid position K67 in Dda 1993 is substituted with A, V, T, S, R, M, E, G, I, L, P, Q, D, H, N, F, or Y, the position corresponding to amino acid position K68 in Dda 1993 is substituted with A, N, E, G, I, L, M, P, Q, R, S, T, V, D, or H, the position corresponding to amino acid position 169 in Dda 1993 is substituted with E, V, R, Q, N, A, K, T, S, D, L, H, M, P, F, or Y, position corresponding to amino acid position K72 in Dda 1993 is substituted with A, E, G, I, L, M, P, Q, R, S, T, V, D, H, or N, the position corresponding to amino acid position K76 in Dda 1993 is substituted with M, Y, F, I, V, R, A, L, Q, S, T, W, D, E, H, N, or P, the position corresponding to amino acid position S79 in Dda 1993 is substituted with T, K, R, V, E, A, D, I, L, M, N, P, Q, G, or H, the position corresponding to amino acid position 184 in Dda 1993 is substituted with L, V, F, A, M, T, or Y, the position corresponding to amino acid position P104 in Dda 1993 is substituted with Y, K, E, F, H, I, L, M, Q, V, W, A, D, S, or T, the position corresponding to amino acid position K108 in Dda 1993 is substituted with Y, E, D, H, N, S, G, F, I, L, M, Q, V, W, A, P, R, or T, the position corresponding to amino acid position V117 in Dda 1993 is substituted with A, S, E, G, I, K, L, M, P, Q, R, T, F, or Y, the position corresponding to amino acid position Y120 in Dda 1993 is substituted with V, L, I, M, F, A, T, S, H, Q, or W, the position corresponding to amino acid position R122 in Dda 1993 is substituted with E, K, N, D, L, T, S, A, V, I, P, Q, H, or M, the position corresponding to amino acid position L124 in Dda 1993 is substituted with T, A, D, E, I, K, M, N, P, Q, R, S, V, F, or Y, the position corresponding to amino acid position 1132 in Dda 1993 is substituted with T, V, L, A, D, E, K, M, N, P, Q, R, S, F, or Y,
he position corresponding to amino acid position P134 in Dda 1993 is substituted with D, E, K, S, L, A, R, I, N, V, T, G, H, or Q, the position corresponding to amino acid position W135 in Dda 1993 is substituted with D, H, T, N, E, G, S, K, Y, Q, R, F, A, P, or M, the position corresponding to amino acid position T137 in Dda 1993 is substituted with K, R, I, V, A, D, E, H, M, N, P, Q, S, or L, the position corresponding to amino acid position N144 in Dda 1993 is substituted with S, P, K, T, R, E, A, D, I, L, Q, Y, H, V, M, or G, the position corresponding to amino acid position K145 in Dda 1993 is substituted with D, A, H, R, G, N, E, P, Q, S, T, or M, the position corresponding to amino acid position E154 in Dda 1993 is substituted with N, S, V, I, T, D, G, H, K, Q, R, A, or P, the position corresponding to amino acid position A157 in Dda 1993 is substituted with E, D, S, K, P, N, H, Q, R, T, G, I, L, M, or V, the position corresponding to amino acid position Y158 in Dda 1993 is substituted with Q, S, E, L, D, A, I, K, F, N, G, R, V, T, H, P, M, or W, the position corresponding to amino acid position P161 in Dda 1993 is substituted with E, D, K, S, N, R, Q, T, A, or H, the position corresponding to amino acid position K166 in Dda 1993 is substituted with N, P, D, E, G, H, Q, R, S, T, A, or M, the position corresponding to amino acid position Q170 in Dda 1993 is substituted with K, I, V, T, S, E, R, N, H, Y, F, A, D, M, or P, the position corresponding to amino acid position T174 in Dda 1993 is substituted with S, A, D, E, G, H, K, M, N, P, Q, R, I, L, or V, the position corresponding to amino acid position V176 in Dda 1993 is substituted with I, G, P, A, T, R, S, Q, K, L, E, F, M, or Y, the position corresponding to amino acid position N180 in Dda 1993 is substituted with D, E, K, A, G, H, P, Q, S, T, or R, the position corresponding to amino acid position A181 in Dda 1993 is substituted with G, N, L, S, R, T, D, K, Q, H, E, I, M, P, or V,
the position corresponding to amino acid position V190 in Dda 1993 is substituted with I, A, F, L, M, T, or Y, the position corresponding to amino acid position Y197 in Dda 1993 is substituted with K, P, R, S, E, G, D, T, L, N, I, V, F, A, H, M, Q, or W, the position corresponding to amino acid position V200 in Dda 1993 is substituted with I, T, F, K, D, L, N, E, S, Y, P, A, or M, the position corresponding to amino acid position V201 in Dda 1993 is substituted with F, Y, I, K, L, H, M, W, A, or T, the position corresponding to amino acid position R207 in Dda 1993 is substituted with I, H, K, V, F, Y, L, T, E, A, M, N, Q, S, or D, the position corresponding to amino acid position T213 in Dda 1993 is substituted with D, N, E, K, S, G, A, Q, H, P, I, L, M, R, or V, the position corresponding to amino acid position A214 in Dda 1993 is substituted with G, S, E, K, D, L, N, I, T, V, M, P, Q, R, or H, the position corresponding to amino acid position L215 in Dda 1993 is substituted with Y, I, E, F, D, H, M, Q, V, W, A, T, K, N, P, R, or S, the position corresponding to amino acid position M219 in Dda 1993 is substituted with L, I, V, A, F, Y, K, D, E, S, T, Q, R, or W, the position corresponding to amino acid position V220 in Dda 1993 is substituted with K, E, N, A, D, R, S, T, Q, L, I, G, H, M, P, F, or Y, the position corresponding to amino acid position 1225 in Dda 1993 is substituted with V, K, T, D, N, S, E, G, R, Q, P, A, H, L, Y, F, or M, the position corresponding to amino acid position K227 in Dda 1993 is substituted with G, A, D, N, S, E, H, M, P, Q, R, or T, the position corresponding to amino acid position L229 in Dda 1993 is substituted with S, K, V, I, N, E, D, T, G, A, R, Q, H, M, P, F, or Y, the position corresponding to amino acid position V237 in Dda 1993 is substituted with M, I, A, L, F, K, Q, R, S, T, W, or Y, the position corresponding to amino acid position M238 in Dda 1993 is substituted with I, L, V, F, A, T, Y, K, Q, R, S, or W,
the position corresponding to amino acid position K247 in Dda 1993 is substituted with N, E, R, D, G, H, Q, S, T, A, M, or P, the position corresponding to amino acid position L248 in Dda 1993 is substituted with F, I, H, M, V, W, Y, A, or T, the position corresponding to amino acid position 1251 in Dda 1993 is substituted with F, L, E, Y, A, T, H, M, V, or W, the position corresponding to amino acid position R253 in Dda 1993 is substituted with Q, A, D, E, H, K, M, N, P, S, T, or Y, the position corresponding to amino acid position K254 in Dda 1993 is substituted with N, S, D, E, G, H, Q, R, T, A, M, or P, the position corresponding to amino acid position F257 in Dda 1993 is substituted with Y, H, I, L, M, Q, V, or W, the position corresponding to amino acid position D262 in Dda 1993 is substituted with P, E, A, K, Q, S, T, G, H, or N, the position corresponding to amino acid position V265 in Dda 1993 is substituted with E, P, A, K, S, G, D, N, T, H, Q, R, F, I, L, M, or Y, the position corresponding to amino acid position D282 in Dda 1993 is substituted with P, A, E, K, Q, S, T, G, H, or N, the position corresponding to amino acid position 1289 in Dda 1993 is substituted with L, V, M, K, E, G, A, F, T, or Y, the position corresponding to amino acid position Q295 in Dda 1993 is substituted with D, E, M, S, G, H, K, N, P, T, A, R, or Y, the position corresponding to amino acid position Y304 in Dda 1993 is substituted with K, E, L, P, D, N, S, G, A, R, T, Q, H, V, I, M, F, or W, the position corresponding to amino acid position S306 in Dda 1993 is substituted with E, Y, D, K, T, G, F, V, N, A, H, P, Q, R, or M, the position corresponding to amino acid position V309 in Dda 1993 is substituted with F, K, I, H, L, M, W, Y, A, or T, the position corresponding to amino acid position V314 in Dda 1993 is substituted with I, K, E, L, A, F, M, T, or Y,
the position corresponding to amino acid position G316 in Dda 1993 is substituted with E, K, D, V, I, N, L, A, H, P, Q, R, S, or T, the position corresponding to amino acid position L319 in Dda 1993 is substituted with E, K, I, V, T, N, D, G, A, H, P, Q, R, S, F, M, or Y, the position corresponding to amino acid position R321 in Dda 1993 is substituted with K, N, E, V, I, D, L, A, H, M, P, Q, S, or T, the position corresponding to amino acid position T329 in Dda 1993 is substituted with P, S, G, K, A, E, D, I, V, L, N, R, Y, F, Q, or M, the position corresponding to amino acid position G331 in Dda 1993 is substituted with D, E, K, L, S, I, N, T, V, H, P, Q, or A, the position corresponding to amino acid position D333 in Dda 1993 is substituted with G, K, E, I, A, N, S, H, P, Q, or T, the position corresponding to amino acid position L349 in Dda 1993 is substituted with E, N, K, Q, A, T, S, D, V, R, I, H, P, F, M, or Y, the position corresponding to amino acid position F352 in Dda 1993 is substituted with Y, L, K, E, R, I, A, N, D, H, M, Q, V, or W, the position corresponding to amino acid position T362 in Dda 1993 is substituted with A, I, V, L, E, K, M, D, R, Y, N, Q, S, F, G, or P, the position corresponding to amino acid position N365 in Dda 1993 is substituted with K, E, A, S, R, D, H, M, P, Q, T, or G, the position corresponding to amino acid position N367 in Dda 1993 is substituted with S, K, L, E, A, T, I, V, Q, R, Y, F, M, D, G, H, or P, the position corresponding to amino acid position G369 in Dda 1993 is substituted with E, K, D, N, S, T, R, L, I, A, Y, H, P, or Q, the position corresponding to amino acid position G370 in Dda 1993 is substituted with K, V, E, I, S, A, N, D, H, M, P, Q, R, or T, the position corresponding to amino acid position P373 in Dda 1993 is substituted with H, L, N, K, M, Y, F, I, V, S, E, T, A, R, D, G, Q, or W, the position corresponding to amino acid position A380 in Dda 1993 is substituted with V, L, T, I, K, M, F, Y, E, G, P, Q, R, or S,
the position corresponding to amino acid position Q383 in Dda 1993 is substituted with L, T, M, Y, K, F, I, V, R, A, E, S, N, D, G, H, or P, the position corresponding to amino acid position S385 in Dda 1993 is substituted with I, T, A, Y, K, H, R, F, L, M, V, D, E, G, N, P, or Q, the position corresponding to amino acid position P391 in Dda 1993 is substituted with G, Y, F, H, A, V, D, N, S, E, K, Q, or T, the position corresponding to amino acid position S393 in Dda 1993 is substituted with I, V, L, T, E, D, M, F, A, Y, G, H, K, N, P, Q, or R, the position corresponding to amino acid position F395 in Dda 1993 is substituted with I, V, Y, L, T, A, M, H, or W, the position corresponding to amino acid position M401 in Dda 1993 is substituted with L, S, V, I, G, D, E, R, Q, A, W, N, T, F, Y, K, H, or P, the position corresponding to amino acid position S402 in Dda 1993 is substituted with T, V, R, I, K, L, E, H, Q, M, A, D, N, P, or G, the position corresponding to amino acid position V403 in Dda 1993 is substituted with F, Y, I, H, L, M, W, A, or T, the position corresponding to amino acid position D404 in Dda 1993 is substituted with N, K, R, E, G, H, Q, S, T, or P, the position corresponding to amino acid position R405 in Dda 1993 is substituted with N, K, D, E, G, H, Q, S, T, A, M, or P, the position corresponding to amino acid position A406 in Dda 1993 is substituted with T, V, I, D, E, K, L, M, N, P, Q, R, S, G, F, or Y, the position corresponding to amino acid position 1413 in Dda 1993 is substituted with E, L, K, S, D, G, N, P, Y, F, T, R, A, H, Q, M, or V, the position corresponding to amino acid position H414 in Dda 1993 is substituted with Y, D, F, Q, K, N, S, E, G, P, R, T, L, I, A, M, V, or W, the position corresponding to amino acid position V418 in Dda 1993 is substituted with K, E, P, R, D, T, A, S, N, L, I, Y, G, H, M, Q, or F, the position corresponding to amino acid position E419 in Dda 1993 is substituted with Q, D, A, H, K, M, N, P, R, S, T, or Y,
the position corresponding to amino acid position A421 in Dda 1993 is substituted with Q, L, R, N, S, K, D, E, H, M, P, T, Y, G, I, V, or F, the position corresponding to amino acid position Q422 in Dda 1993 is substituted with K, R, L, N, A, D, E, H, M, P, S, T, or Y, the position corresponding to amino acid position V427 in Dda 1993 is substituted with T, E, D, A, I, K, L, M, N, P, Q, R, S, F, or Y, the position corresponding to amino acid position V429 in Dda 1993 is substituted with I, L, A, F, M, T, or Y, the position corresponding to amino acid position G432 in Dda 1993 is substituted with P, A, S, T, D, E, K, Q, or N, the position corresponding to amino acid position Y434 in Dda 1993 is substituted with K, I, V, R, E, T, N, D, L, S, Q, H, G, A, M, P, F, or W, the position corresponding to amino acid position D435 in Dda 1993 is substituted with E, Q, A, H, K, N, P, R, S, T, or G, and/or the position corresponding to amino acid position F437 in Dda 1993 is substituted with Y, I, H, L, M, Q, V, or W.
3. A construct comprising a helicase according to claim 1 or 2 and an additional polynucleotide binding moiety, wherein the helicase is attached to the polynucleotide binding moiety and the construct has the ability to control the movement of an analyte.
4. A construct according to claim 3, wherein the construct comprises two or more helicases according to claim 1 or 2.
5. A polynucleotide which comprises a sequence which encodes a helicase according to claim 1 or 2 or a construct according to claim 3 or 4.
6. A vector which comprises a polynucleotide according to claim 5 operably linked to a promoter.
7. A host cell comprising a vector according to claim 6.
8. A method of making a helicase according to claim 1 or 2 or a construct according to claim 3 or 4, which comprises expressing a polynucleotide according to claim 5, transfecting a cell with a vector according to claim 6 or culturing a host cell according to claim 7.
9. A method of controlling the movement of an analyte, comprising contacting the analyte with a helicase according to claim 1 or 2 or a construct according to claim 3 or 4 and thereby controlling the movement of the analyte.
10. A method according to claim 9, wherein the method is for controlling the movement of an analyte through a transmembrane pore.
11. A method of characterising a target analyte, comprising:
(a) contacting the target analyte with a transmembrane pore and a helicase according to claim 1 or 2 or a construct according to claim 3 or 4 such that the helicase or construct controls the movement of the target analyte through the pore; and
(b) taking one or more measurements as the target analyte moves with respect to the pore wherein the measurements are indicative of one or more characteristics of the target analyte and thereby characterising the target analyte.
12. A method according to claim 11, wherein the method comprises:
(a) contacting the target analyte with a transmembrane pore and a helicase according to claim 1 or 2 or a construct according to claim 3 or 4 such that the helicase or the construct controls the movement of the target analyte through the pore; and
(b) measuring the current passing through the pore as the target analyte moves with respect to the pore wherein the current is indicative of one or more characteristics of the target analyte and thereby characterising the target analyte.
13. A method according to claim 11 or 12, wherein the target analyte is a polynucleotide.
14. A method according to any one of claims 11 to 23, wherein the pore is a transmembrane protein pore or a solid state pore.
15. A method of forming a sensor for characterising a target analyte, comprising forming a complex between (a) a pore and (b) a helicase according to claim 1 or 2 or a construct according to claim 3 or 4 and thereby forming a sensor for characterising the target analyte.
16. A method according to claim 15, wherein the complex is formed by covalently attaching the pore to the helicase or construct.
17. A sensor for characterising a target analyte, comprising a complex between (a) a pore and (b) a helicase according to claim 1 or 2 or a construct according to claim 3 or 4.
18. Use of a helicase according to claim 1 or 2 or a construct according to claim 3 or 4 to control the movement of a target analyte through a pore.
19. A kit for characterising a target analyte comprising
(a) a pore and a helicase according to claim 1 or 2 or a construct according to claim 3 or 4; or
(b) a helicase according to claim 1 or 2 or a construct according to claim 3 or 4 and one or more loading moieties.
20. An apparatus for characterising target analytes in a sample, comprising (a) a plurality of pores and (b) a plurality of helicases according to claim 1 or 2 or a plurality of constructs according to claim 3 or 4.
21. A method of producing a helicase according to claim 1 or 2, comprising:
(a) providing a helicase; and
(b) modifying the helicase to produce a helicase according to claim 1 or 2.
22. A method according to claim 21, wherein the method further comprises (c) determining whether or not the resulting helicase is capable of controlling the movement of a polynucleotide.
23. A method of producing a construct according to claim 3 or 4, comprising attaching a helicase according to claim 1 or 2 to an additional polynucleotide binding moiety and thereby producing the construct.
24. A series of two or more helicases attached to a polynucleotide, wherein at least one of the two or more helicases is a helicase according to claim 1 or 2.
25. A method of improving the movement of a target analyte with respect to a transmembrane pore when the movement is controlled by a DNA dependent ATPase (Dda) helicase, wherein the DNA dependent ATPase (Dda) helicase is modified as defined in claim 1 or 2 which improves the movement of the target analyte with respect to the transmembrane pore.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| GB2315935.3 | 2023-10-18 | ||
| GBGB2315935.3A GB202315935D0 (en) | 2023-10-18 | 2023-10-18 | Novel enzymes |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025083247A1 true WO2025083247A1 (en) | 2025-04-24 |
Family
ID=88863866
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/EP2024/079566 Pending WO2025083247A1 (en) | 2023-10-18 | 2024-10-18 | Novel enzymes |
Country Status (2)
| Country | Link |
|---|---|
| GB (1) | GB202315935D0 (en) |
| WO (1) | WO2025083247A1 (en) |
Citations (30)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2000028312A1 (en) | 1998-11-06 | 2000-05-18 | The Regents Of The University Of California | A miniature support for thin films containing single channels or nanopores and methods for using same |
| WO2006100484A2 (en) | 2005-03-23 | 2006-09-28 | Isis Innovation Limited | Deliver of molecules to a li id bila |
| WO2008102121A1 (en) | 2007-02-20 | 2008-08-28 | Oxford Nanopore Technologies Limited | Formation of lipid bilayers |
| WO2009020682A2 (en) | 2007-05-08 | 2009-02-12 | The Trustees Of Boston University | Chemical functionalization of solid-state nanopores and nanopore arrays and applications thereof |
| WO2009035647A1 (en) | 2007-09-12 | 2009-03-19 | President And Fellows Of Harvard College | High-resolution molecular graphene sensor comprising an aperture in the graphene layer |
| WO2009077734A2 (en) | 2007-12-19 | 2009-06-25 | Oxford Nanopore Technologies Limited | Formation of layers of amphiphilic molecules |
| WO2010004265A1 (en) | 2008-07-07 | 2010-01-14 | Oxford Nanopore Technologies Limited | Enzyme-pore constructs |
| WO2010086602A1 (en) | 2009-01-30 | 2010-08-05 | Oxford Nanopore Technologies Limited | Hybridization linkers |
| WO2010122293A1 (en) | 2009-04-20 | 2010-10-28 | Oxford Nanopore Technologies Limited | Lipid bilayer sensor array |
| WO2011067559A1 (en) | 2009-12-01 | 2011-06-09 | Oxford Nanopore Technologies Limited | Biochemical analysis instrument |
| WO2012005857A1 (en) | 2010-06-08 | 2012-01-12 | President And Fellows Of Harvard College | Nanopore device with graphene supported artificial lipid membrane |
| WO2013057495A2 (en) | 2011-10-21 | 2013-04-25 | Oxford Nanopore Technologies Limited | Enzyme method |
| WO2013098561A1 (en) | 2011-12-29 | 2013-07-04 | Oxford Nanopore Technologies Limited | Method for characterising a polynucelotide by using a xpd helicase |
| WO2013098562A2 (en) | 2011-12-29 | 2013-07-04 | Oxford Nanopore Technologies Limited | Enzyme method |
| WO2014013262A1 (en) | 2012-07-19 | 2014-01-23 | Oxford Nanopore Technologies Limited | Enzyme construct |
| WO2014013259A1 (en) | 2012-07-19 | 2014-01-23 | Oxford Nanopore Technologies Limited | Ssb method |
| WO2014013260A1 (en) | 2012-07-19 | 2014-01-23 | Oxford Nanopore Technologies Limited | Modified helicases |
| WO2014064444A1 (en) | 2012-10-26 | 2014-05-01 | Oxford Nanopore Technologies Limited | Droplet interfaces |
| WO2014064443A2 (en) | 2012-10-26 | 2014-05-01 | Oxford Nanopore Technologies Limited | Formation of array of membranes and apparatus therefor |
| WO2014187924A1 (en) | 2013-05-24 | 2014-11-27 | Illumina Cambridge Limited | Pyrophosphorolytic sequencing |
| WO2015055981A2 (en) | 2013-10-18 | 2015-04-23 | Oxford Nanopore Technologies Limited | Modified enzymes |
| WO2015110813A1 (en) | 2014-01-22 | 2015-07-30 | Oxford Nanopore Technologies Limited | Method for attaching one or more polynucleotide binding proteins to a target polynucleotide |
| WO2015166276A1 (en) | 2014-05-02 | 2015-11-05 | Oxford Nanopore Technologies Limited | Method of improving the movement of a target polynucleotide with respect to a transmembrane pore |
| WO2016034591A2 (en) | 2014-09-01 | 2016-03-10 | Vib Vzw | Mutant pores |
| WO2016055777A2 (en) | 2014-10-07 | 2016-04-14 | Oxford Nanopore Technologies Limited | Modified enzymes |
| WO2017149316A1 (en) | 2016-03-02 | 2017-09-08 | Oxford Nanopore Technologies Limited | Mutant pore |
| WO2017203268A1 (en) * | 2016-05-25 | 2017-11-30 | Oxford Nanopore Technologies Limited | Method |
| WO2018211241A1 (en) | 2017-05-04 | 2018-11-22 | Oxford Nanopore Technologies Limited | Transmembrane pore consisting of two csgg pores |
| WO2019002893A1 (en) | 2017-06-30 | 2019-01-03 | Vib Vzw | Novel protein pores |
| WO2024138631A1 (en) * | 2022-12-30 | 2024-07-04 | 深圳华大生命科学研究院 | Mutant of dda helicase, and preparation method therefor and use thereof in sequencing |
-
2023
- 2023-10-18 GB GBGB2315935.3A patent/GB202315935D0/en not_active Ceased
-
2024
- 2024-10-18 WO PCT/EP2024/079566 patent/WO2025083247A1/en active Pending
Patent Citations (34)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2000028312A1 (en) | 1998-11-06 | 2000-05-18 | The Regents Of The University Of California | A miniature support for thin films containing single channels or nanopores and methods for using same |
| WO2006100484A2 (en) | 2005-03-23 | 2006-09-28 | Isis Innovation Limited | Deliver of molecules to a li id bila |
| WO2008102121A1 (en) | 2007-02-20 | 2008-08-28 | Oxford Nanopore Technologies Limited | Formation of lipid bilayers |
| WO2008102120A1 (en) | 2007-02-20 | 2008-08-28 | Oxford Nanopore Technologies Limited | Lipid bilayer sensor system |
| WO2009020682A2 (en) | 2007-05-08 | 2009-02-12 | The Trustees Of Boston University | Chemical functionalization of solid-state nanopores and nanopore arrays and applications thereof |
| WO2009035647A1 (en) | 2007-09-12 | 2009-03-19 | President And Fellows Of Harvard College | High-resolution molecular graphene sensor comprising an aperture in the graphene layer |
| WO2009077734A2 (en) | 2007-12-19 | 2009-06-25 | Oxford Nanopore Technologies Limited | Formation of layers of amphiphilic molecules |
| WO2010004265A1 (en) | 2008-07-07 | 2010-01-14 | Oxford Nanopore Technologies Limited | Enzyme-pore constructs |
| WO2010086602A1 (en) | 2009-01-30 | 2010-08-05 | Oxford Nanopore Technologies Limited | Hybridization linkers |
| WO2010086603A1 (en) | 2009-01-30 | 2010-08-05 | Oxford Nanopore Technologies Limited | Enzyme mutant |
| WO2010122293A1 (en) | 2009-04-20 | 2010-10-28 | Oxford Nanopore Technologies Limited | Lipid bilayer sensor array |
| WO2011067559A1 (en) | 2009-12-01 | 2011-06-09 | Oxford Nanopore Technologies Limited | Biochemical analysis instrument |
| WO2012005857A1 (en) | 2010-06-08 | 2012-01-12 | President And Fellows Of Harvard College | Nanopore device with graphene supported artificial lipid membrane |
| WO2013057495A2 (en) | 2011-10-21 | 2013-04-25 | Oxford Nanopore Technologies Limited | Enzyme method |
| WO2013098561A1 (en) | 2011-12-29 | 2013-07-04 | Oxford Nanopore Technologies Limited | Method for characterising a polynucelotide by using a xpd helicase |
| WO2013098562A2 (en) | 2011-12-29 | 2013-07-04 | Oxford Nanopore Technologies Limited | Enzyme method |
| WO2014013262A1 (en) | 2012-07-19 | 2014-01-23 | Oxford Nanopore Technologies Limited | Enzyme construct |
| WO2014013259A1 (en) | 2012-07-19 | 2014-01-23 | Oxford Nanopore Technologies Limited | Ssb method |
| WO2014013260A1 (en) | 2012-07-19 | 2014-01-23 | Oxford Nanopore Technologies Limited | Modified helicases |
| WO2014064444A1 (en) | 2012-10-26 | 2014-05-01 | Oxford Nanopore Technologies Limited | Droplet interfaces |
| WO2014064443A2 (en) | 2012-10-26 | 2014-05-01 | Oxford Nanopore Technologies Limited | Formation of array of membranes and apparatus therefor |
| WO2014187924A1 (en) | 2013-05-24 | 2014-11-27 | Illumina Cambridge Limited | Pyrophosphorolytic sequencing |
| WO2015055981A2 (en) | 2013-10-18 | 2015-04-23 | Oxford Nanopore Technologies Limited | Modified enzymes |
| WO2015110813A1 (en) | 2014-01-22 | 2015-07-30 | Oxford Nanopore Technologies Limited | Method for attaching one or more polynucleotide binding proteins to a target polynucleotide |
| WO2015166276A1 (en) | 2014-05-02 | 2015-11-05 | Oxford Nanopore Technologies Limited | Method of improving the movement of a target polynucleotide with respect to a transmembrane pore |
| WO2016034591A2 (en) | 2014-09-01 | 2016-03-10 | Vib Vzw | Mutant pores |
| WO2016055777A2 (en) | 2014-10-07 | 2016-04-14 | Oxford Nanopore Technologies Limited | Modified enzymes |
| WO2017149316A1 (en) | 2016-03-02 | 2017-09-08 | Oxford Nanopore Technologies Limited | Mutant pore |
| WO2017149317A1 (en) | 2016-03-02 | 2017-09-08 | Oxford Nanopore Technologies Limited | Mutant pore |
| WO2017149318A1 (en) | 2016-03-02 | 2017-09-08 | Oxford Nanopore Technologies Limited | Mutant pores |
| WO2017203268A1 (en) * | 2016-05-25 | 2017-11-30 | Oxford Nanopore Technologies Limited | Method |
| WO2018211241A1 (en) | 2017-05-04 | 2018-11-22 | Oxford Nanopore Technologies Limited | Transmembrane pore consisting of two csgg pores |
| WO2019002893A1 (en) | 2017-06-30 | 2019-01-03 | Vib Vzw | Novel protein pores |
| WO2024138631A1 (en) * | 2022-12-30 | 2024-07-04 | 深圳华大生命科学研究院 | Mutant of dda helicase, and preparation method therefor and use thereof in sequencing |
Non-Patent Citations (18)
| Title |
|---|
| "Uniprot", Database accession no. POAE98 |
| ALTSCHUL S. F., J MOL EVOL, vol. 36, 1993, pages 290 - 300 |
| ALTSCHUL, S.F ET AL., J MOL BIOL, vol. 215, 1990, pages 403 - 10 |
| AUSUBEL ET AL.: "Current Protocols in Molecular Biology", 2016, JOHN WILEY & SONS |
| D. STODDART ET AL., PROC. NATL. ACAD. SCI., vol. 106, 2010, pages 7702 - 7 |
| DEVEREUX ET AL., NUCLEIC ACIDS RESEARCH, vol. 12, 1984, pages 387 - 395 |
| GONZALEZ-PEREZ ET AL., LANGMUIR, vol. 25, 2009, pages 10447 - 10450 |
| HAMMERSTEIN ET AL., J BIOL CHEM., vol. 286, no. 16, 22 April 2011 (2011-04-22), pages 14324 - 14334 |
| HOWORKA ET AL., NATURE BIOTECHNOLOGY, vol. 30, no. 6, 7 June 2012 (2012-06-07), pages 506 - 7 |
| JAMES, W: "Encyclopedia of Analytical Chemistry", 2000, JOHN WILEY & SONS LTD, pages: 4848 - 4871 |
| LIU C. C.SCHULTZ P. G., ANNU. REV. BIOCHEM., vol. 79, 2010, pages 413 - 444 |
| MARK RANCECAVANAGH, JOHNWAYNE J. FAIRBROTHERARTHUR W: "Hunt III; Skelton, NNicholas J.", 2007, FREEMAN AND CO, pages: 953 - 954 |
| O'SHEA ET AL., SCIENCE, vol. 254, no. 5031, pages 539 - 544 |
| REMAUTWAKSMAN TRENDS, BIOCHEM. SCI., vol. 31, 2006, pages 436 - 444 |
| RUPP B: "Biomolecular Crystallography: Principles, Practice and Application to Structural Biology", NEW YORK: GARLAND SCIENCE, 2009 |
| VAN HEEL MGOWEN BMATADEEN RORLOVA EVFINN RPAPE TCOHEN DSTARK HSCHMIDT RSCHATZ M: "Single-particle electron cryo-microscopy: towards atomic resolution", Q REV BIOPHYS., vol. 33, 2000, pages 307 - 69, XP001106808, DOI: 10.1017/S0033583500003644 |
| XIANG MAHE TIAN, CHEM. SOC. REV., vol. 39, 2010, pages 70 - 80 |
| XIAOPING HE ET AL., STRUCTURE, vol. 20, 2012, pages 1189 - 1200 |
Also Published As
| Publication number | Publication date |
|---|---|
| GB202315935D0 (en) | 2023-11-29 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US11739377B2 (en) | Method of improving the movement of a target polynucleotide with respect to a transmembrane pore | |
| US11945840B2 (en) | Protein pores | |
| JP7499761B2 (en) | pore | |
| KR102436445B1 (en) | Modified enzymes | |
| KR102472805B1 (en) | mutant pore | |
| US20180037874A9 (en) | Modified helicases | |
| US20250137046A1 (en) | Pore | |
| WO2025083247A1 (en) | Novel enzymes | |
| US20250188533A1 (en) | Nanopore | |
| AU2023253169A1 (en) | Novel modified protein pores and enzymes | |
| US20220162568A1 (en) | Pore |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24798746 Country of ref document: EP Kind code of ref document: A1 |