WO2025049858A1 - Molecules for treatment of cancer - Google Patents
Molecules for treatment of cancer Download PDFInfo
- Publication number
- WO2025049858A1 WO2025049858A1 PCT/US2024/044603 US2024044603W WO2025049858A1 WO 2025049858 A1 WO2025049858 A1 WO 2025049858A1 US 2024044603 W US2024044603 W US 2024044603W WO 2025049858 A1 WO2025049858 A1 WO 2025049858A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- cdr
- seq
- amino acid
- acid sequence
- antigen
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/2878—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the NGF-receptor/TNF-receptor superfamily, e.g. CD27, CD30, CD40, CD95
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/2803—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily
- C07K16/2827—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against the immunoglobulin superfamily against B7 molecules, e.g. CD80, CD86
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/505—Medicinal preparations containing antigens or antibodies comprising antibodies
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/20—Immunoglobulins specific features characterized by taxonomic origin
- C07K2317/21—Immunoglobulins specific features characterized by taxonomic origin from primates, e.g. man
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/31—Immunoglobulins specific features characterized by aspects of specificity or valency multispecific
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/33—Crossreactivity, e.g. for species or epitope, or lack of said crossreactivity
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/34—Identification of a linear epitope shorter than 20 amino acid residues or of a conformational epitope defined by amino acid residues
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/56—Immunoglobulins specific features characterized by immunoglobulin fragments variable (Fv) region, i.e. VH and/or VL
- C07K2317/569—Single domain, e.g. dAb, sdAb, VHH, VNAR or nanobody®
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/75—Agonist effect on antigen
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/76—Antagonist effect on antigen, e.g. neutralization or inhibition of binding
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
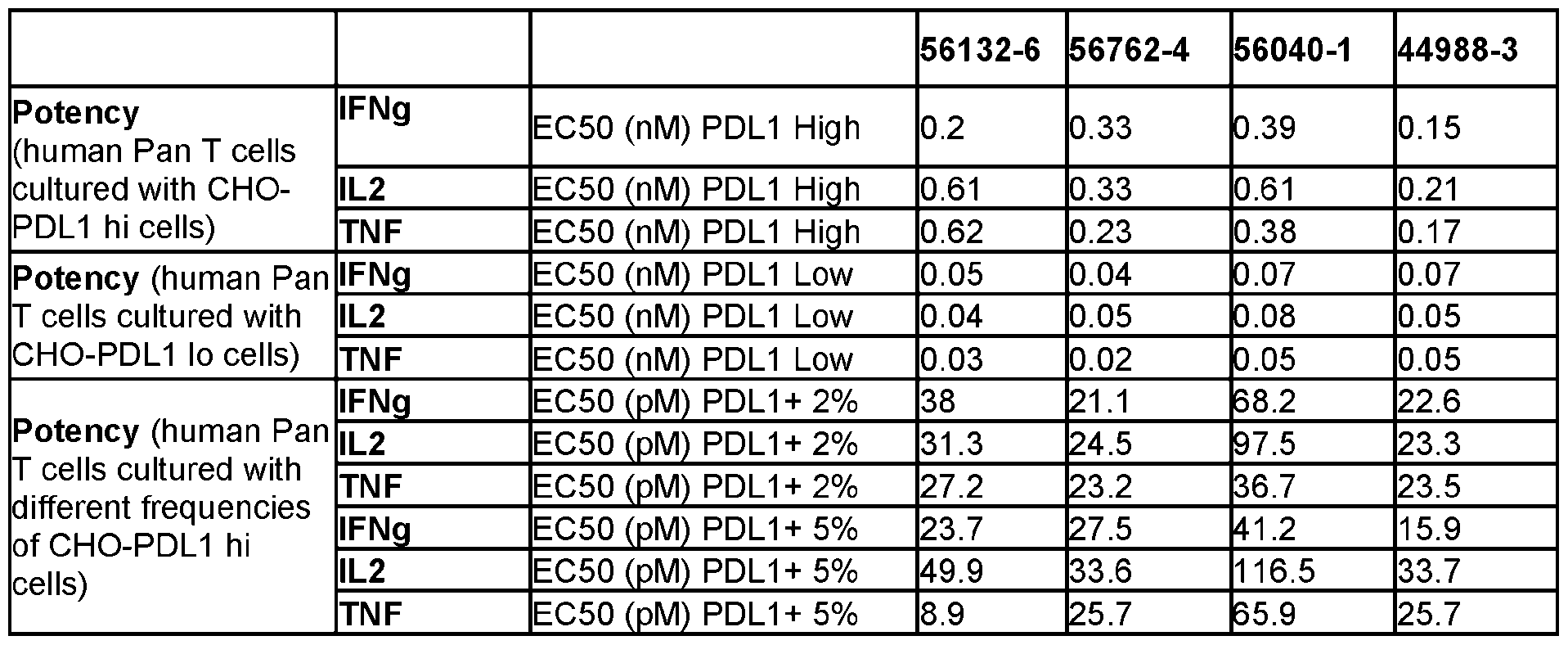
- C07K2317/92—Affinity (KD), association rate (Ka), dissociation rate (Kd) or EC50 value
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/94—Stability, e.g. half-life, pH, temperature or enzyme-resistance
Definitions
- the present invention relates to antigen binding proteins, for example, bispecific molecules, for the treatment of cancer.
- BACKGROUND [4] The PD-1/PD-L1 axis is involved in the suppression of T cell immune responses in cancer. Antagonists of this pathway have been clinically validated across a number of solid tumor indications. Nivolumab and pembrolizumab are two such inhibitors that target the PD-1 pathway, and each has been approved by the U.S. Food and Drug Administration (FDA) for the treatment of metastatic melanoma.
- FDA U.S. Food and Drug Administration
- 4-1BB which is also known as CD137 or TNFRSF9, is a member of the TNF receptor superfamily. 4-1BB was first identified as a molecule whose expression is induced by T-cell activation (Kwon Y.H. and Weissman S.M. (1989), Proc. Natl. Acad. Sci. USA 86, 1963-1967).
- 4-1BB in T- and B-lymphocytes, NK-cells, NKT-cells, monocytes, neutrophils, and dendritic cells as well as cells of non- hematopoietic origin such as endothelial and smooth muscle cells.
- Expression of 4-1BB in different cell types is mostly inducible and driven by various stimulatory signals, such as T-cell receptor (TCR) or B-cell receptor triggering, as well as signaling induced through co- stimulatory molecules or receptors of pro-inflammatory cytokines.
- TCR T-cell receptor
- B-cell receptor triggering as well as signaling induced through co- stimulatory molecules or receptors of pro-inflammatory cytokines.
- 4-1BB signaling is known to stimulate IFN ⁇ secretion and proliferation of NK cells, as well as to promote DC activation as indicated by their increased survival and capacity to secret cytokines and upregulate co- stimulatory molecules.
- 4-1BB is best characterized as a co-stimulatory molecule which modulates TCR- induced activation in both the CD4+ and CD8+ subsets of T-cells.
- agonistic 4-1BB-specific antibodies enhance proliferation of T-cells, stimulate lymphokine secretion and decrease sensitivity of T-lymphocytes to activation-induced cells death (Snell L.M. et al. (2011) Immunol. Rev.244, 197-217).
- 4- 1BB agonists can also induce infiltration and retention of activated T-cells in the tumor through 4-1BB -mediated upregulation of intercellular adhesion molecule 1 (ICAM1) and vascular cell adhesion molecule 1 (VCAM1) on tumor vascular endothelium.4-1BB triggering may also reverse the state of T-cell anergy induced by exposure to soluble antigen that may contribute to disruption of immunological tolerance in the tumor micro- environment or during chronic infections.
- IAM1 intercellular adhesion molecule 1
- VCAM1 vascular cell adhesion molecule 1
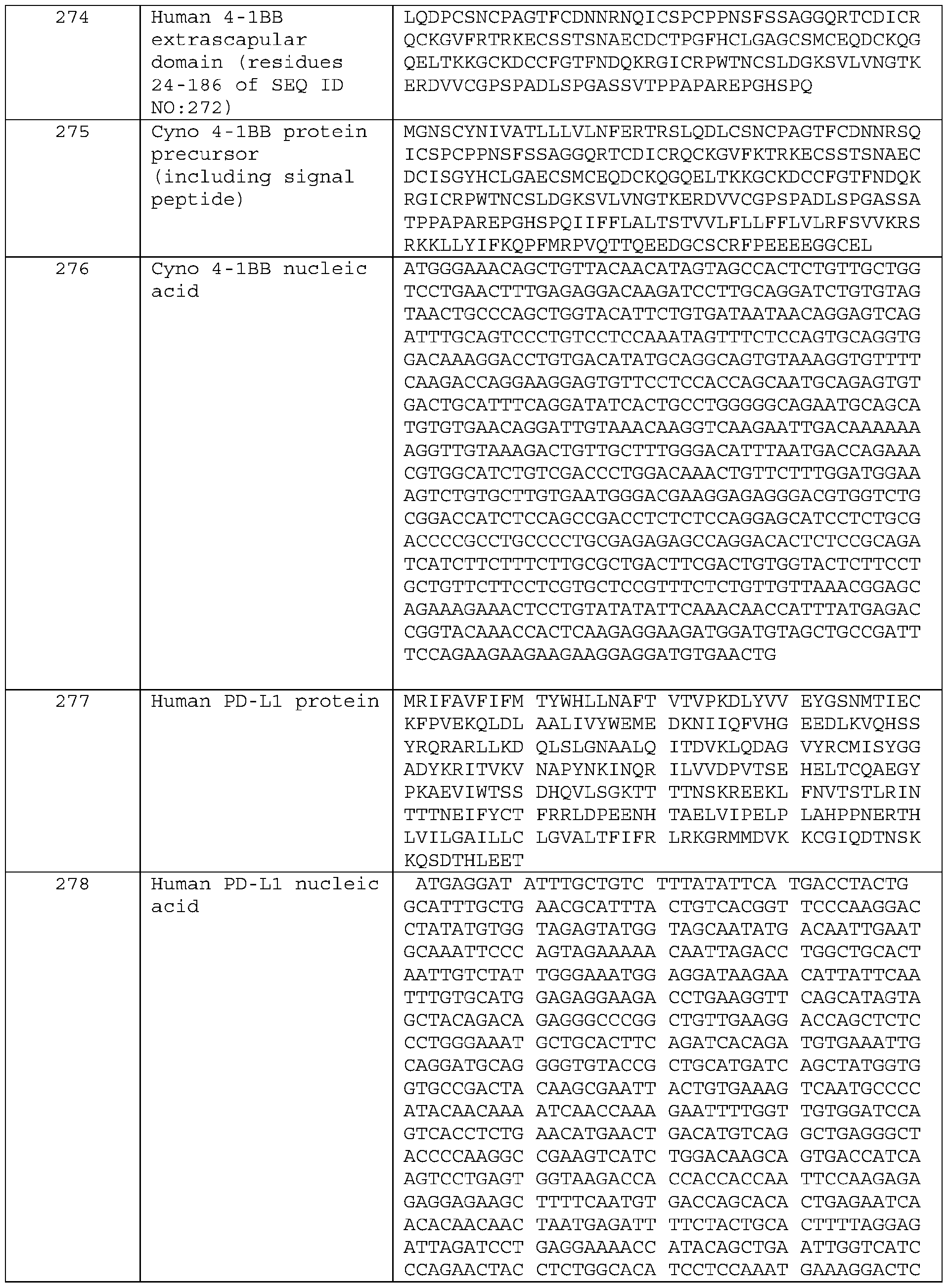
- a 4-1BB antigen-binding protein comprising a heavy chain variable domain (VH) and a light chain variable domain (VL), wherein said protein binds to the Cysteine-rich pseudo repeat 1 (CRD1) of human 4-1BB (corresponding to residues 24-45 of SEQ ID NO: 272), and is a crosslinking-dependent agonist.
- the 4-1BB antigen-binding protein of E1 comprising (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 151, 159, 324 or 167; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 152, 160, 325, or 168.
- the 4-1BB antigen-binding protein of E1, comprising: (i) a CDR-H1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 55, 79 or 103; (ii) a CDR-H2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 56, 80, or 104; (iii) a CDR-H3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical
- E4 The 4-1BB antigen-binding protein of any one of E1-E3, comprising: (1) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.55-60, respectively (14A5); (2) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.79-84, respectively (14A5.002); or (3) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.103-108, respectively (16D1.001).
- a 4-1BB antigen-binding protein comprising a heavy chain variable domain (VH) and a light chain variable domain (VL), wherein said protein binds to the Cysteine-rich pseudo repeat 2 (CRD2) of human 4-1BB (corresponding to residues 47-86 of SEQ ID NO: 272), and is a crosslinking-dependent agonist.
- VH heavy chain variable domain
- VL light chain variable domain
- CCD2 Cysteine-rich pseudo repeat 2
- the 4-1BB antigen-binding protein of E5, comprising (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 149, 161, 157, 169, 147, 163, 165, 153, 171, 175, 155, 404, 406, 408, or 410; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 150, 162, 158, 170, 148, 164, 166, 154, 172, 176, 156, 405, 407, 409, or 411. E7.
- the 4-1BB antigen-binding protein of E5, comprising: (i) a CDR-H1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 49, 85, 73, 109, 43, 91, 97, 61, 115, 127, 67, 386, or 392; (ii) a CDR-H2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 50, 86, 74, 110, 44, 92, 98, 62, 116, 128, 68, 387, or 393; (iii) a CDR-H3 comprising
- E8 The 4-1BB antigen-binding protein of any one of E5-E7, comprising: (1) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.49-54, respectively (6F9); (2) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.85-90, respectively (6F9.009); (3) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.73-78, respectively (14G12.017); (4) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR
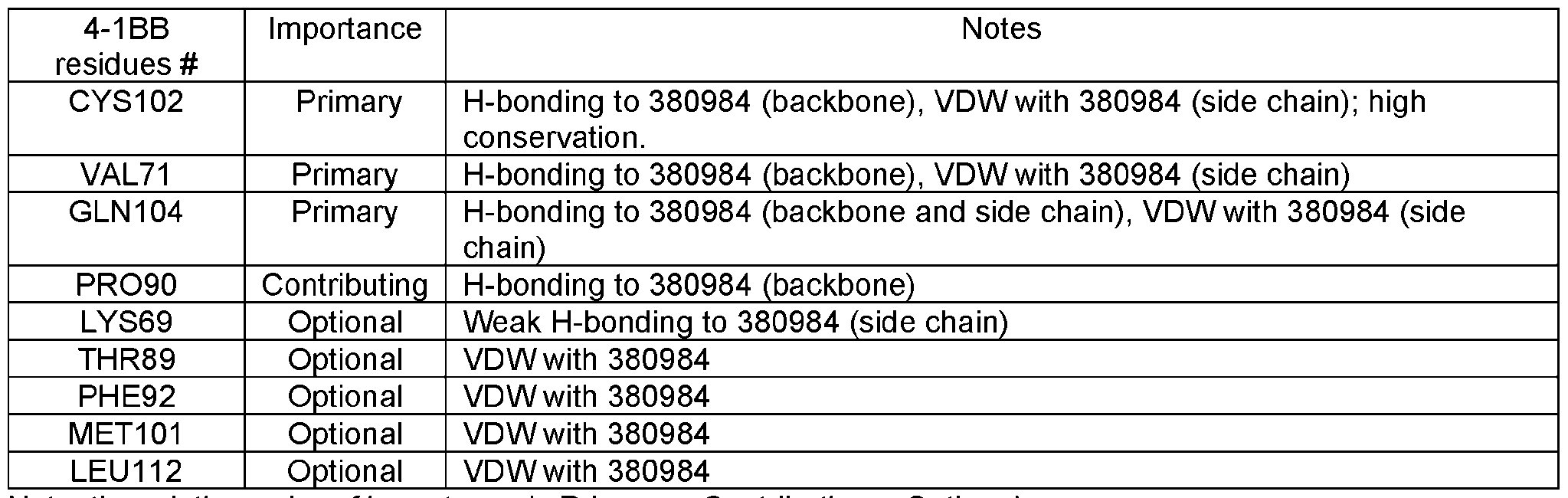
- a 4-1BB antigen-binding protein comprising a heavy chain variable domain and a light chain variable domain, wherein said protein binds to the Cysteine-rich pseudo repeat 3 (CRD3) of human 4- 1BB (corresponding to residues 87-118 of SEQ ID NO: 272), and is a crosslinking-dependent agonist.
- CCD3 Cysteine-rich pseudo repeat 3
- the 4-1BB antigen-binding protein of E9 comprising (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 173; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 174.
- the 4-1BB antigen-binding protein of E9 comprising: (i) a CDR-H1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 121; (ii) a CDR-H2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 122; (iii) a CDR-H3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 123; (
- E12 The 4-1BB antigen-binding protein of any one of E9-E11, comprising: a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs: 121-126, respectively (15A6.011).
- E13 The 4-1BB antigen-binding protein of any one of E9-E11, comprising: a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs: 121-126, respectively (15A6.011).
- E21. The 4-1BB antigen-binding protein of any one of E1-E15 and E18-E19, comprising a VL framework derived from a human germline V ⁇ framework sequence.
- E22. The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH1, VH2, VH3, VH4, or VH5 framework sequence.
- E24 The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH3 framework sequence.
- E25 The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH4 framework sequence.
- E26. The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH2 framework sequence.
- E27 The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH5 framework sequence.
- E28 The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH5 framework sequence.
- the 4-1BB antigen-binding protein of any one of E1-E29 comprising: a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 147, 149, 151, 153, 155, 157, 159, 324, 161, 163, 165, 167, 169, 171, 173, 175, 404, 406, 408, and 410.
- E31 E31.
- the 4-1BB antigen-binding protein of any one of E1-E30 comprising: a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 148, 150, 152, 154, 156, 158, 160, 325, 162, 164, 166, 168, 170, 172, 174, 176, 405, 407, 409, and 411. E32.
- the 4-1BB antigen-binding protein of any one of E1-E31 comprising: (a) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.147, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.148; (b) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.149, and a VL comprising an amino acid sequence at least 90%,
- a 4-1BB antigen-binding protein that comprises a heavy chain variable domain (VH) and does not comprise a light chain variable domain (VL), wherein said protein comprises the CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, or 385.
- VH heavy chain variable domain
- VL light chain variable domain
- a 4-1BB antigen-binding protein that comprises a heavy chain variable domain (VH) and does not comprise a light chain variable domain (VL), wherein said VH comprises: (i) a CDR-H1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 326, 329, 332, 335, 338, 341, 344, 347, 350, 353, 356, 359, 362, 365, or 368; (ii) a CDR-H2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 327, 330, 333, 336, 339, 342, 345,
- E35 The 4-1BB antigen-binding protein of E33 or E34, comprising: (1) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.326-328, respectively; (2) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.329-331, respectively; (3) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.332-334, respectively; (4) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.335-337, respectively; (5) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.338-340, respectively; (6) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.341-343, respectively; (7)
- E37 The 4-1BB antigen-binding protein of any one of E33-E36, comprising a VH framework derived from a human germline VH1, VH2, VH3, VH4, or VH5 framework sequence.
- E38. The 4-1BB antigen-binding protein of any one of E33-E37, comprising a VH framework derived from a human germline VH1 framework sequence.
- E39. The 4-1BB antigen-binding protein of any one of E33-E37, comprising a VH framework derived from a human germline VH3 framework sequence.
- E40 The 4-1BB antigen-binding protein of any one of E33-E37, comprising a VH framework derived from a human germline VH4 framework sequence.
- the 4-1BB antigen-binding protein of any one of E33-E37 comprising a VH framework derived from a human germline VH2 framework sequence.
- E42. The 4-1BB antigen-binding protein of any one of E33-E37, comprising a VH framework derived from a human germline VH5 framework sequence.
- E43. The 4-1BB antigen-binding protein of any one of E33-E42, comprising a VH framework sequence is at least 90% identical to the human germline framework sequence from which it is derived.
- the 4-1BB antigen-binding protein of any one of E33-E43, comprising a VH framework sequence is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to the human germline framework sequence from which it is derived.
- the 4-1BB antigen-binding protein of any one of E33-E44 comprising: a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, and 385.
- a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, and 385.
- the 4-1BB antigen-binding protein of any one of E33-E45 comprising: (a) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.371; (b) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.372; (c) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.373; (d) a VH comprising an
- E47 The 4-1BB antigen-binding protein of any one of E1-E46 and E315-E342, further comprising a heavy chain CH1 domain.
- E48. The 4-1BB antigen-binding protein of E47, wherein said CH1 domain is the CH1 domain of an IgG (for example IgG1, lgG2, lgG3, or lgG4).
- E49. The 4-1BB antigen-binding protein of E47 or E48, wherein said CH1 domain is the CH1 domain of a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4).
- E53. The 4-1BB antigen-binding protein of E52, wherein the Fc region is the Fc region of an IgA (for example IgA1 or lgA2), IgD, IgE, IgM, or IgG (for example IgG1, lgG2, lgG3, or lgG4).
- E54 The 4-1BB antigen-binding protein of E52 or E53, wherein the Fc region is the Fc region of an IgG.
- the 4-1BB antigen-binding protein of E54 wherein the IgG is selected from the group consisting of IgG1, lgG2, lgG3, and lgG4.
- E56 The 4-1BB antigen-binding protein of E55, wherein the IgG is IgG1, IgG2, or IgG4.
- E57 The 4-1BB antigen-binding protein of any one of E52-E56, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: L234A, L235A, L235E, G237A, and combination thereof (numbering according to the EU index).
- E58 The 4-1BB antigen-binding protein of E54, wherein the IgG is selected from the group consisting of IgG1, lgG2, lgG3, and lgG4.
- the 4-1BB antigen-binding protein of E57 comprising L234A and L235A mutations.
- E59 The 4-1BB antigen-binding protein of any one of E52-E58, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: V259C, A287C, R292C, V302C, L306C, V323C, I332C, and a combination thereof (numbering according to the EU index).
- E61. The 4-1BB antigen-binding protein of E60, comprising a N297G mutation.
- E62. The 4-1BB antigen-binding protein of E60, comprising A287C, N297G, and L306C mutations.
- the 4-1BB antigen-binding protein of E60 comprising R292C, N297G, and V302C mutations.
- E64. The 4-1BB antigen-binding protein of any one of E52-E63, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: M252Y, S254T, T256E, and a combination thereof.
- E65 The 4-1BB antigen-binding protein of E64, comprising M252Y, S254T, T256E mutations.
- E67. The 4-1BB antigen-binding protein of any one of E52-E65, wherein the lysine residue (K) at the C-terminus of the Fc region is present.
- E68. The 4-1BB antigen-binding protein of any one of E52-E65, wherein the glycine and lysine residues (GK) at the C-terminus of the Fc region are present.
- E70. The 4-1BB antigen-binding protein of any one of E52-E69, wherein said Fc region comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, or 426.
- E71 The 4-1BB antigen-binding protein of any one of E52-E69, wherein said Fc region comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 263, 267, or 483.
- E72 The 4-1BB antigen-binding protein of any one of E52-E69, wherein said Fc region comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 91%, at
- E73. The 4-1BB antigen-binding protein of E72, wherein said constant domain is the constant domain of an IgA (for example IgA1 or lgA2), IgD, IgE, IgM, or IgG (for example IgG1, lgG2, lgG3, or lgG4).
- the 4-1BB antigen-binding protein of E72 or E73 wherein said constant domain is the constant domain of an IgG (for example IgG1, IgG2, IgG3, or IgG4), preferably a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4).
- IgG for example IgG1, IgG2, IgG3, or IgG4
- human IgG for example, human IgG1, human IgG2, human IgG3, or human IgG4
- E78. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E77, further comprising a kappa light chain constant domain that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428.
- E79 The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E77, further comprising a lambda light chain constant domain that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424. E80.
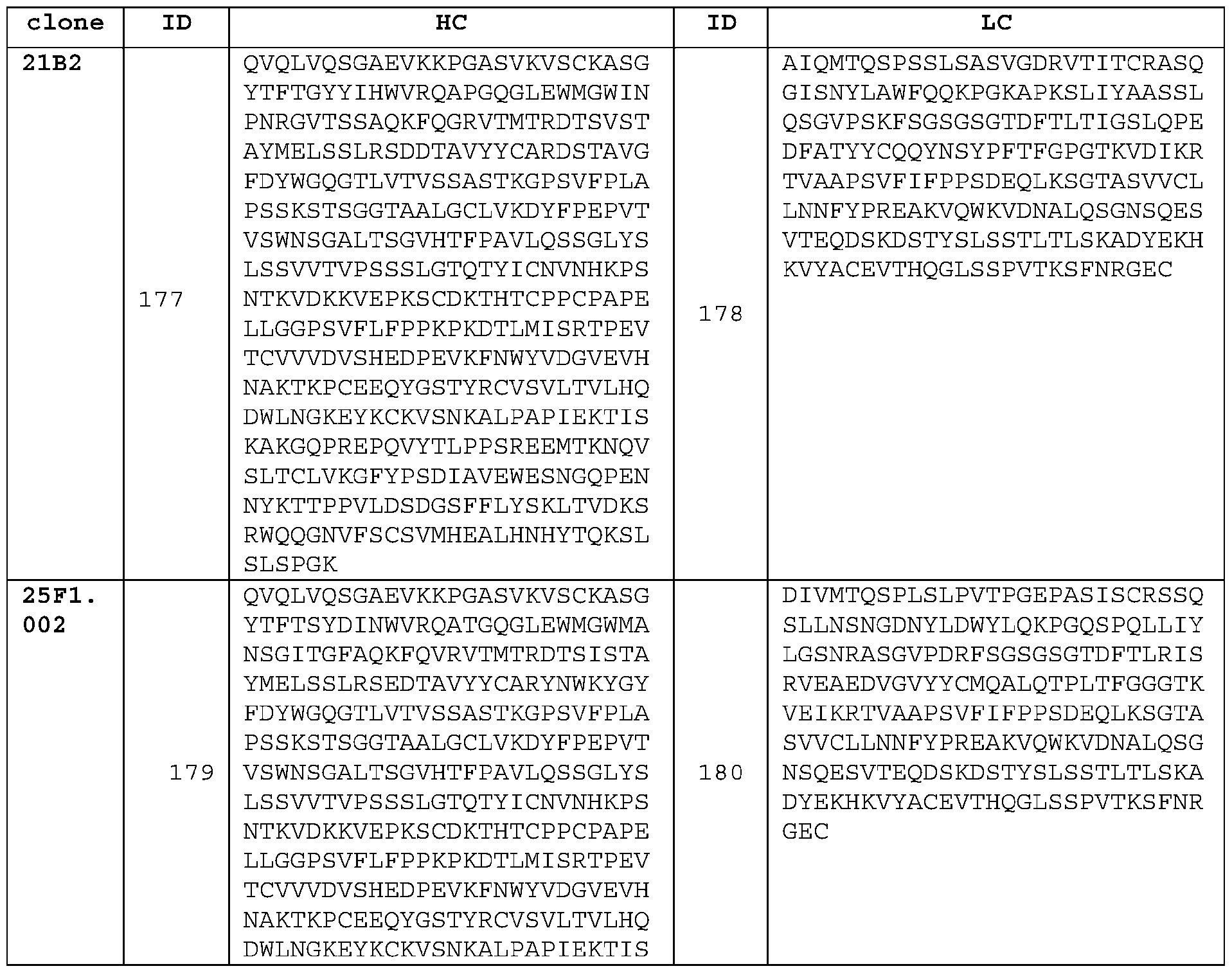
- the 4-1BB antigen-binding protein of any one of E1-E32 and E47-E79 comprising: (i) a heavy chain (HC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 191, 193, 195, 197, 199, 201, 203, 205, 207, 209, 211, 213, 215, 217, and 219; and (ii) a light chain (LC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 192, 194, 196, 198, 200, 202, 204, 206, 208,
- E81 The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E80, comprising: (a) a heavy chain (HC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.191, and a light chain (LC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.192; (b) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.193,
- E82 The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E81, which is an antibody.
- E83 The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E81, which is an antigen- binding fragment of an antibody, such as a Fab fragment.
- E84. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E81, which is an scFv.
- E85 The 4-1BB antigen-binding protein of E84, wherein said scFv comprises a first linker between VH and VL.
- the 4-1BB antigen-binding protein of E85 wherein said first linker comprises: (a) a glycine rich peptide; (b) a peptide comprising glycine and serine; (c) a peptide comprising (Gly-Gly-Ser) n , wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 234); (d) a peptide comprising (Gly-Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 232); (e) a peptide comprising (Gly-Gly-Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 233); (f) a peptide comprising (Gly-Gly-Gly-Gly-Gln)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 235), or (g
- E88. The 4-1BB antigen-binding protein of any one of E1-E32 and E84-E87, comprising a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 291 (14A5.002 scFv).
- E89 The 4-1BB antigen-binding protein of any one of E1-E32 and E84-E87, comprising a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 445 (14A5.002 scFv #2).
- E90 The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E81, which is a Fab.
- the 4-1BB antigen-binding protein of E90 comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 298, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 300 (6F
- the 4-1BB antigen-binding protein of E90 comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 305, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 307
- the 4-1BB antigen-binding protein of E90 comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 316, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 317
- the 4-1BB antigen-binding protein of E90 comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-227 of SEQ ID NO: 434, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to S
- the 4-1BB antigen-binding protein of E90 comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-227 of SEQ ID NO: 449, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to S
- E96 The 4-1BB antigen-binding protein of any one of E1-E95, wherein the antigen-binding protein binds to 4-1BB with a KD value of or less than: about 500nM, about 400nM, about 300nM, about 200nM, about 150nM, about 100nM, about 90nM, about 80nM, about 70nM, about 60nM, about 50nM, about 40nM, about 30nM, about 25nM, about 20nM, about 15nM, about 10nM, about 9nM, about 8nM, about 7nM, about 6nM, about 5nM, about 4nM, about 3nM, about 2nM, about 1 nM, about 900pM, about 800pM, about 700pM, about 600pM, about 500pM, about 400pM, about 300pM, about 250pM, about 200pM, about 150pM, about 100pM, about 50pM, about 40pM, about 30pM, about 25p
- E97 A 4-1BB antigen-binding protein that competes for binding to 4-1BB with any one of the 4-1BB antigen binding protein of E1-E96 and E315-E342.
- E98 A 4-1BB antigen-binding protein that binds to substantially the same epitope as any one of the 4- 1BB antigen binding protein of E1-E96 and E315-E342.
- E99 A bispecific molecule that comprises the 4-1BB antigen-binding protein of any one of E1-E98 and E315-E342, and further comprises a second antigen-binding moiety.
- E100 The bispecific molecule of E99, wherein said second antigen-binding moiety binds to a protein of the immune checkpoint pathway.
- the bispecific molecule of E100, wherein the protein of the immune checkpoint pathway is CTLA- 4, PD-1, PD-L1, PD-L2, B7-H3, B7-H4, CEACAM-1, TIGIT, LAG3, CD112, CD112R, CD96, TIM3, or BTLA.
- the bispecific molecule of E101, wherein the protein of the immune checkpoint pathway is PD- L1.
- a PD-L1 antigen-binding protein comprising (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 133, 135, 137, 139, 141, 143, 322, 145, 398, 400, or 402; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 134, 136, 138, 140, 142, 144, 323, 146, 399, 401, or 403. E104.
- a PD-L1 antigen-binding protein comprising: (i) a CDR-H1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 1, 7, 13, 19, 25, 31, or 37; (ii) a CDR-H2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 2, 8, 14, 20, 26, 32, or 38; (iii) a CDR-H3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or
- E105 The PD-L1 antigen-binding protein of E103 or E104, comprising: (1) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.1-6, respectively; (2) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.7-12, respectively; (3) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.13-18, respectively; (4) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.19-24, respectively; (5)
- E106 The PD-L1 antigen-binding protein of any one of E103-E105, comprising a VL framework derived from a human germline V ⁇ framework sequence.
- E107. The PD-L1 antigen-binding protein of any one of E103-E105, comprising a VL framework derived from a human germline V ⁇ framework sequence.
- E108. The PD-L1 antigen-binding protein of any one of E103-E107, comprising a VH framework derived from a human germline VH1, VH2, VH3, VH4, or VH5 framework sequence.
- E109 The PD-L1 antigen-binding protein of any one of E103-E105, comprising a VL framework derived from a human germline VH1, VH2, VH3, VH4, or VH5 framework sequence.
- the PD-L1 antigen-binding protein of any one of E103-E107 comprising a VH framework derived from a human germline VH1 framework sequence.
- E110. The PD-L1 antigen-binding protein of any one of E103-E107, comprising a VH framework derived from a human germline VH2 framework sequence.
- E111. The PD-L1 antigen-binding protein of any one of E103-E107, comprising a VH framework derived from a human germline VH3 framework sequence.
- E112 The PD-L1 antigen-binding protein of any one of E103-E107, comprising a VH framework derived from a human germline VH4 framework sequence.
- the PD-L1 antigen-binding protein of any one of E103-E107 comprising a VH framework derived from a human germline VH5 framework sequence.
- E114. The PD-L1 antigen-binding protein of any one of E103-E113, comprising a VL framework sequence and a VH framework sequence, and wherein one or both of the VL framework sequence or VH framework sequence is at least 90% identical to the human germline framework sequence from which it is derived.
- the PD-L1 antigen-binding protein of any one of E103-E114 comprising a VL framework sequence and a VH framework sequence, and wherein one or both of the VL framework sequence or VH framework sequence is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to the human germline framework sequence from which it is derived.
- the PD-L1 antigen-binding protein of any one of E103-E115 comprising a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs:133, 135, 137, 139, 141, 143, 322, 145, 398, 400, and 402 E117.
- the PD-L1 antigen-binding protein of any one of E103-E116 comprising a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 134, 136, 138, 140, 142, 323144, 146, 399, 401, and 403. E118.
- the PD-L1 antigen-binding protein of any one of E103-E117 comprising: (1) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.133, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.134; (2) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.135, and a VL comprising an amino acid sequence at least 90%, at least
- E120. The PD-L1 antigen-binding protein of E119, wherein said CH1 domain is the CH1 domain of an IgG (for example IgG1, lgG2, lgG3, or lgG4).
- E121. The PD-L1 antigen-binding protein of E119 or E120, wherein said CH1 domain is the CH1 domain of a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4).
- the PD-L1 antigen-binding protein of any one of E119-E121 wherein said CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 422, or 425.
- E124. The PD-L1 antigen-binding protein of any one of E103-E123, further comprising an Fc region.
- E125 The PD-L1 antigen-binding protein of E124, wherein the Fc region is the Fc region of an IgA (for example IgA1 or lgA2), IgD, IgE, IgM, or IgG (for example IgG1, lgG2, lgG3, or lgG4).
- E126 The PD-L1 antigen-binding protein of E124 or E125, wherein the Fc region is the Fc region of an IgG.
- E127 The PD-L1 antigen-binding protein of E126, wherein the IgG is selected from the group consisting of IgG1, lgG2, lgG3, and lgG4. E128.
- the PD-L1 antigen-binding protein of E126 wherein the IgG is IgG1, IgG2, or IgG4.
- E129. The PD-L1 antigen-binding protein of any one of E124-E128, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: L234A, L235A, L235E, G237A, and combination thereof (numbering according to the EU index).
- E130. The PD-L1 antigen-binding protein of E129, comprising L234A and L235A mutations. E131.
- said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: V259C, A287C, R292C, V302C, L306C, V323C, I332C, and a combination thereof (numbering according to the EU index).
- E132. The PD-L1 antigen-binding protein of E131, comprising a N297G mutation.
- the PD-L1 antigen-binding protein of E131 comprising R292C, N297G, and V302C mutations.
- E135. The PD-L1 antigen-binding protein of any one of E124-E134, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: M252Y, S254T, T256E, and a combination thereof.
- E136. The PD-L1 antigen-binding protein of E135, comprising M252Y, S254T, T256E mutations.
- E138. The PD-L1 antigen-binding protein of any one of E124-E136, wherein the lysine residue (K) at the C-terminus of the Fc region is present.
- E139. The PD-L1 antigen-binding protein of any one of E124-E136, wherein the glycine and lysine residues (GK) at the C-terminus of the Fc region are present.
- E141. The PD-L1 antigen-binding protein of any one of E124-E140, wherein said Fc region comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, or 426.
- E142 The PD-L1 antigen-binding protein of any one of E124-E140, wherein said Fc region comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 263, 267, or 483.
- E143 E143.
- E144. The PD-L1 antigen-binding protein of E143, wherein said constant domain is the constant domain of an IgA (for example IgA1 or lgA2), IgD, IgE, IgM, or IgG (for example IgG1, lgG2, lgG3, or lgG4).
- the PD-L1 antigen-binding protein of E143 or E144 wherein said constant domain is the constant domain of an IgG (for example IgG1, IgG2, IgG3, or IgG4), preferably a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4).
- IgG for example IgG1, IgG2, IgG3, or IgG4
- human IgG for example, human IgG1, human IgG2, human IgG3, or human IgG4
- E150 The PD-L1 antigen-binding protein of any one of E103-E148, further comprising a lambda light chain constant domain that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424. E151.
- the PD-L1 antigen-binding protein of any one of E103-E150 comprising: (i) a heavy chain (HC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 177, 179, 181, 183, 185, 187, and 189; and (ii) a light chain (LC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 178, 180, 182, 184, 186, 188, and 190.
- HC heavy chain
- LC light chain
- E152 The PD-L1 antigen-binding protein of any one of E103-E151, comprising: (a) a heavy chain (HC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.177, and a light chain (LC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.178; (b) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.179, and a
- E153 The PD-L1 antigen-binding protein of any one of E103-E152, which is an antibody.
- E154. PD-L1 antigen-binding protein of any one of E103-E152, which is an antigen-binding fragment of an antibody, such as a Fab fragment.
- E155. The PD-L1 antigen-binding protein of any one of E103-E152, which is an scFv.
- E156 The PD-L1 antigen-binding protein of E155, wherein said scFv comprises a first linker between VH and VL.
- the PD-L1 antigen-binding protein of E156 wherein said first linker comprises: (a) a glycine rich peptide; (b) a peptide comprising glycine and serine; (c) a peptide comprising (Gly-Gly-Ser) n , wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 234); (d) a peptide comprising (Gly-Gly-Gly-Ser) n , wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 232); (e) a peptide comprising (Gly-Gly-Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 233); (f) a peptide comprising (Gly-Gly-Gly-Gly-Gln)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 235
- E158 The PD-L1 antigen-binding protein of E156, wherein said first linker comprises the amino acid sequence of (Gly-Gly-Gly-Gly-Ser)3 (SEQ ID NO: 230).
- E159. The PD-L1 antigen-binding protein of any one of E155-E158, comprising a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 284 (26F6.002.009 scFv).
- E160 The PD-L1 antigen-binding protein of any one of E155-E158, comprising a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 436 (scFv from 56039).
- E161 The PD-L1 antigen-binding protein of any one of E103-E152, which is a Fab. E162.
- the PD-L1 antigen-binding protein of E161, comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-221 SEQ ID NO: 451, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID
- a PD-L1 antigen-binding protein that competes for binding to PD-L1 with any one of the PD-L1 antigen binding protein of E103-E163.
- E165. A PD-L1 antigen-binding protein that binds to substantially the same epitope as any one of the PD-L1 antigen binding protein of E103-E163.
- E166. A bispecific molecule comprising the PD-L1 antigen-binding protein of any one of E103-E165, and further comprising a second antigen-binding moiety.
- the bispecific molecule of E166 wherein the second antigen-binding moiety binds to a T-cell co- stimulatory molecule (such as: CD28, Inducible Co-Stimulator (ICOS), CTLA4 (Cytotoxic T-Lymphocyte- Associated protein 4), 4-1BB (also known as CD137), OX40 (also known as CD134), CD27, CD30, DR3, Glucocorticoid-Induced TNFR family Related (GITR), or Herpes Virus Entry Mediator (HVEM)).
- a T-cell co- stimulatory molecule such as: CD28, Inducible Co-Stimulator (ICOS), CTLA4 (Cytotoxic T-Lymphocyte- Associated protein 4), 4-1BB (also known as CD137), OX40 (also known as CD134), CD27, CD30, DR3, Glucocorticoid-Induced TNFR family Related (GITR), or Herpes Virus Entry Mediator (HVEM
- a bispecific molecule comprising (i) a 4-1BB antigen-binding moiety comprising any one of E1- E98 and E315-E342, and (ii) a PD-L1 antigen-binding moiety of any one of embodiments E103-E165. E170.
- a bispecific molecule comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker - scFv, wherein said scFv comprises a VH B and a VL B , and wherein said VH B and VL B are connected via a second linker; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said scFv binds to 4-1BB, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region.
- a bispecific molecule comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker - scFv, wherein said scFv comprises a VHB and a VL B , and wherein said VH B and VL B are connected via a second linker; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VL A – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said scFv binds to PD-L1, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region.
- a bispecific molecule comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – first linker – scFv – third linker – monomeric CH2 – monomeric CH3, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a second linker; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VL A – CL; wherein said VH A -CH1 and said VL A -CL form a Fab that binds to PD-L1, said scFv binds to 4-1BB, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region.
- a bispecific molecule comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – first linker – scFv – third linker – monomeric CH2 – monomeric CH3, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a second linker; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VH A -CH1 and said VL A -CL form a Fab that binds to 4-1BB, said scFv binds to PD-L1, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region.
- a bispecific molecule comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker – VH B – CH1’; and (ii) two copies of a first light chain that comprises, from N-terminus to C-terminus: VL A – CL; (iii) two copies of a second light chain that comprises, from N-terminus to C-terminus: VLB – CL’; wherein said VHA-CH1 and VLA-CL form a first Fab that binds to PD-L1, said VHB-CH1’ and VLB-CL’ form a second Fab that binds to 4-1BB, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region.
- a bispecific molecule comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – monomeric CH2 – monomeric CH3 – first linker – VH B – CH1’; and (ii) two copies of a first light chain that comprises, from N-terminus to C-terminus: VLA – CL; (iii) two copies of a second light chain that comprises, from N-terminus to C-terminus: VLB – CL’; wherein said VH A -CH1 and VL A -CL form a first Fab that binds to 4-1BB, said VH B -CH1’ and VL B -CL’ form a second Fab that binds to PD-L1, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region.
- E176 The bispecific molecule of E174 or E175, wherein (i) said CH1 comprises a mutation to a positively charged residue and said CL comprises a mutation to a negatively charged residue, such that said CH1 and CL form a first charge pair, and (ii) said CH1’ comprises a mutation to a negatively charged residue and said CL’ comprises a mutation to a positively charged residue, such that said CH1’ and CL’ form a second charge pair.
- said CH1 comprises a mutation to K or R (preferably K) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246.
- E181. The bispecific molecule of E174 or E175, wherein (i) said CH1 comprises a mutation to a negatively charged residue and said CL comprises a mutation to a positively charged residue, such that said CH1 and CL form a first charge pair, and (ii) said CH1’ comprises a mutation to a positively charged residue and said CL’ comprises a mutation to a negatively charged residue, such that said CH1’ and CL’ form a second charge pair.
- said CH1 comprises a mutation to a negatively charged residue and said CL comprises a mutation to a negatively charged residue, such that said CH1’ and CL’ form a second charge pair.
- E184. The bispecific molecule of any one of E181- E183, wherein said CH1’ comprises a mutation to K or R (preferably K) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246.
- a bispecific molecule comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – first linker – VHB – second linker – monomeric CH2 – monomeric CH3; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VH A -CH1 and said VL A -CL form a Fab that binds to PD-L1, said VH B binds to 4-1BB, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (Fab-VH(M2)-Fc, Fig.14D). E187.
- a bispecific molecule comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – first linker – VH B – second linker – monomeric CH2 – monomeric CH3; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VL A – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said VHB binds to PD-L1, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (Fab-VH(M2)-Fc, Fig.14D). E188.
- a bispecific molecule comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – monomeric CH2 – monomeric CH3 – first linker - VHB; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VH A -CH1 and said VL A -CL form a Fab that binds to PD-L1, said VH B binds to 4-1BB, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (IgG-VH(C2), Fig.14C) E189.
- a bispecific molecule comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker - VH B ; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VL A – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said VHB binds to PD-L1, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (IgG-VH(C2), Fig.14C) E190.
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – monomeric CH2 – monomeric CH3 – first linker – scFv; wherein said scFv comprises a VH B and a VL B , and wherein said VHB and VLB are connected via a second linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VL A – CL; wherein said VHA-CH1 and said VLA-CL form two Fab domains that binds to 4-1BB, said scFv binds to PD-L1, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker – scFv; wherein said scFv comprises a VHB and a VLB, and wherein said VH B and VL B are connected via a second linker (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VH A -CH1 and said VL A -CL form two Fab domains that binds to PD-L1, said scFv binds to 4- 1BB, and said monomeric CH2
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – first linker – scFv – third linker – monomeric CH2 – monomeric CH3; wherein said scFv comprises a VH B and a VLB, and wherein said VHB and VLB are connected via a second linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VL A – CL; wherein said VHA-CH1 and said VLA-CL form two Fab domains that binds to 4-1BB, said scFv binds to PD-L1, and said monomeric CH
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – first linker – scFv – third linker – monomeric CH2 – monomeric CH3; wherein said scFv comprises a VHB and a VL B , and wherein said VH B and VL B are connected via a second linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VL A – CL; wherein said VHA-CH1 and said VLA-CL form two Fab domains that binds to PD-L1, said scFv binds to 4- 1BB, and said
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – monomeric CH2’ – monomeric CH3’, wherein said scFv comprises a VH B and a VL B , and wherein said VHB and VLB are connected via a first linker; and (iii) a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VH A -CH1 and said VL A -CL form a Fab that binds to PD-L1, said scFv binds to 4-1BB, and said monomeric CH2 – monomeric CH3 from (i) and
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – monomeric CH2’ – monomeric CH3’, wherein said scFv comprises a VHB and a VLB, and wherein said VH B and VL B are connected via a first linker; and (iii) a light chain that comprises, from N-terminus to C-terminus: VL A – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said scFv binds to PD-L1, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VH B – first linker – monomeric CH2’ – monomeric CH3’; and (iii) a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said VHB binds to 4-1BB, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region.
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHB – first linker – monomeric CH2’ – monomeric CH3’; and (iii) a light chain that comprises, from N-terminus to C-terminus: VL A – CL; wherein said VH A -CH1 and said VL A -CL form a Fab that binds to 4-1BB, said VH B binds to PD-L1, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region.
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – VH A – CH1 – monomeric CH2 – monomeric CH3, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a first linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said scFv binds to 4-1BB, and said monomeric CH2 – monomeric CH3 from (
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – VHA – CH1 – monomeric CH2 – monomeric CH3, wherein said scFv comprises a VH B and a VL B , and wherein said VH B and VL B are connected via a first linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VL A – CL; wherein said VH A -CH1 and said VL A -CL form a Fab that binds to 4-1BB, said scFv binds to PD-L1, and said monomeric CH
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – monomeric CH2 (1) – monomeric CH3 (1) , wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a first linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – monomeric CH2 (2) – monomeric CH3 (2) – monomeric CH2 (3) – monomeric CH3 (3) ; (iii) a light chain that comprises, from N-terminus to C-terminus: VLA – CL; and (iv) a fourth chain comprising monomeric CH2 (4) – monomeric CH3 (4) ; wherein said VH A -CH1 and said VL A -CL form a Fab
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – monomeric CH2 (1) – monomeric CH3 (1) , wherein said scFv comprises a VHB and a VLB, and wherein said VH B and VL B are connected via a first linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – monomeric CH2 (2) – monomeric CH3 (2) – monomeric CH2 (3) – monomeric CH3 (3) ; (iii) a light chain that comprises, from N-terminus to C-terminus: VLA – CL; and (iv) a fourth chain comprising monomeric CH2 (4) – monomeric CH3 (4) ; wherein said VH A -CH1 and said VL A
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VH B – CH1’ – monomeric CH2’ – monomeric CH3’; (iii) a first light chain that comprises, from N-terminus to C-terminus: VLA – CL; and (iv) a second light chain that comprises, from N-terminus to C-terminus: VLB – CL’; wherein said VH A -CH1 and said VL A -CL form a first Fab that binds to 4-1BB, wherein said VH B -CH1’ and said VL B -CL’ form a second Fab that to PD-L1, and wherein said mono
- a bispecific molecule comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VH A – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHB – CH1’ – monomeric CH2’ – monomeric CH3’; (iii) a first light chain that comprises, from N-terminus to C-terminus: VL A – CL; and (iv) a second light chain that comprises, from N-terminus to C-terminus: VL B – CL’; wherein said VHA-CH1 and said VLA-CL form a first Fab that binds to PD-L1, wherein said VHB-CH1’ and said VL B -CL’ form a second Fab that to 4-1BB, and wherein said monomeric CH2 – monomeric CH3 from (i)
- E204 The bispecific molecule of any one of E202 or E203, wherein (i) said CH1 comprises a mutation to a positively charged residue and said CL comprises a mutation to a negatively charged residue, such that said CH1 and CL form a first charge pair, and (ii) said CH1’ comprises a mutation to a negatively charged residue and said CL’ comprises a mutation to a positively charged residue, such that said CH1’ and CL’ form a second charge pair.
- E205 The bispecific molecule of E204, wherein said CH1 comprises a mutation to K or R (preferably K) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246.
- E207. The bispecific molecule of any one of E204- E206, wherein said CH1’ comprises a mutation to E or D (preferably E) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246.
- said CH1 comprises a mutation to a negatively charged residue and said CL comprises a mutation to a positively charged residue, such that said CH1’ and CL’ form a second charge pair.
- E212. The bispecific molecule of any one of E209- E211, wherein said CH1’ comprises a mutation to K or R (preferably K) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246.
- E213. The bispecific molecule of any one of E209- E212, wherein said CL’ comprises a mutation to E or D (preferably E) at position 176 (Kabat numbering), or at a position that corresponds to residue 69 of SEQ ID NO:237 or 239.
- E214. The bispecific molecule of any one of E190- E213, wherein said CH2, or CH2’, or CH2 (1) , or CH2 (2) , or CH2 (3) , or CH2 (4) comprises a mutation wherein a positively charged residue is mutated to a negatively charge residue, or a mutation wherein a negatively charged residue is mutated to a positively charge residue.
- the bispecific molecule of any one of E190- E214, wherein said CH3, or CH3’, or CH3 (1) , or CH3 (2) , or CH3 (3) , or CH3 (4) comprises a mutation wherein a positively charged residue is mutated to a negatively charge residue, or a mutation wherein a negatively charged residue is mutated to a positively charge residue.
- E216. The bispecific molecule of any one of E190- E215, wherein said CH3, or CH3’, or CH3 (1) , or CH3 (2) , or CH3 (3) , or CH3 (4) comprises a mutation from K to E or D (preferably D) at position 392 (EU index numbering), or at a position that corresponds to residue 275 of SEQ ID NO:246.
- E217 The bispecific molecule of any one of E190- E216, wherein said CH3, or CH3’, or CH3 (1) , or CH3 (2) , or CH3 (3) , or CH3 (4) comprises a mutation from K to D or E (preferably D) at position 409 (EU index numbering), or at a position that corresponds to residue 292 of SEQ ID NO:246. E218.
- the bispecific molecule of any one of E170- E219, wherein said CH1 or CH1’ is the CH1 domain of a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4).
- E220 is the CH1 domain of a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4).
- E222. The bispecific molecule of any one of E170-E221, wherein said Fc is the Fc region of an IgG.
- E223. The bispecific molecule of E222, wherein the IgG is selected from the group consisting of IgG1, lgG2, lgG3, and lgG4.
- E224. The bispecific molecule of E223, wherein the IgG is IgG1, IgG2, or IgG4.
- E225. The bispecific molecule of any one of E170-E224, wherein said Fc is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: L234A, L235A, L235E, G237A, and combination thereof (numbering according to the EU index).
- E226. The bispecific molecule of E225, comprising L234A and L235A mutations.
- E229. The bispecific molecule of E228, comprising a N297G mutation.
- E230. The bispecific molecule of E228, comprising A287C, N297G, and L306C mutations.
- E235 The bispecific molecule of any one of E170-E233, wherein the lysine residue (K) at the C- terminus of the Fc region is present.
- E237. The bispecific molecule of any one of E170-E233, wherein the glycine and lysine residues (GK) at the C-terminus of the Fc region are deleted.
- E240 The bispecific molecule of any one of E170-E239, wherein said CL or CL’ is a kappa or lambda light chain constant domain.
- E241 The bispecific molecule of any one of E170-E240, wherein said CL or CL’ is a kappa light chain constant domain that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428. E242.
- E246 The bispecific molecule of any one of E170-E245, wherein said first linker, second linker, or third linker, each independently comprises: (a) GGGG (SEQ ID NO: 222); (b) GGGGSGGGGSGGGGS (SEQ ID NO: 230); (c) GGGGQGGGGQ (SEQ ID NO: 504); or (d) GGGGSGGGGS (SEQ ID NO: 229).
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 283, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 220.
- E248. A bispecific molecule comprising two copies of SEQ ID NO: 283, and two copies of SEQ ID NO: 220.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 287, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 206. E250.
- a bispecific molecule comprising two copies of SEQ ID NO: 287, and two copies of SEQ ID NO: 206. (clone 11252) E251.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 290, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 184.
- a bispecific molecule comprising two copies of SEQ ID NO: 290, and two copies of SEQ ID NO: 184. (clone 11253) E253.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 294, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 186.
- a bispecific molecule comprising two copies of SEQ ID NO: 294, and two copies of SEQ ID NO: 186. (clone 11255).
- E255. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 297; (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 299; and (iii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at
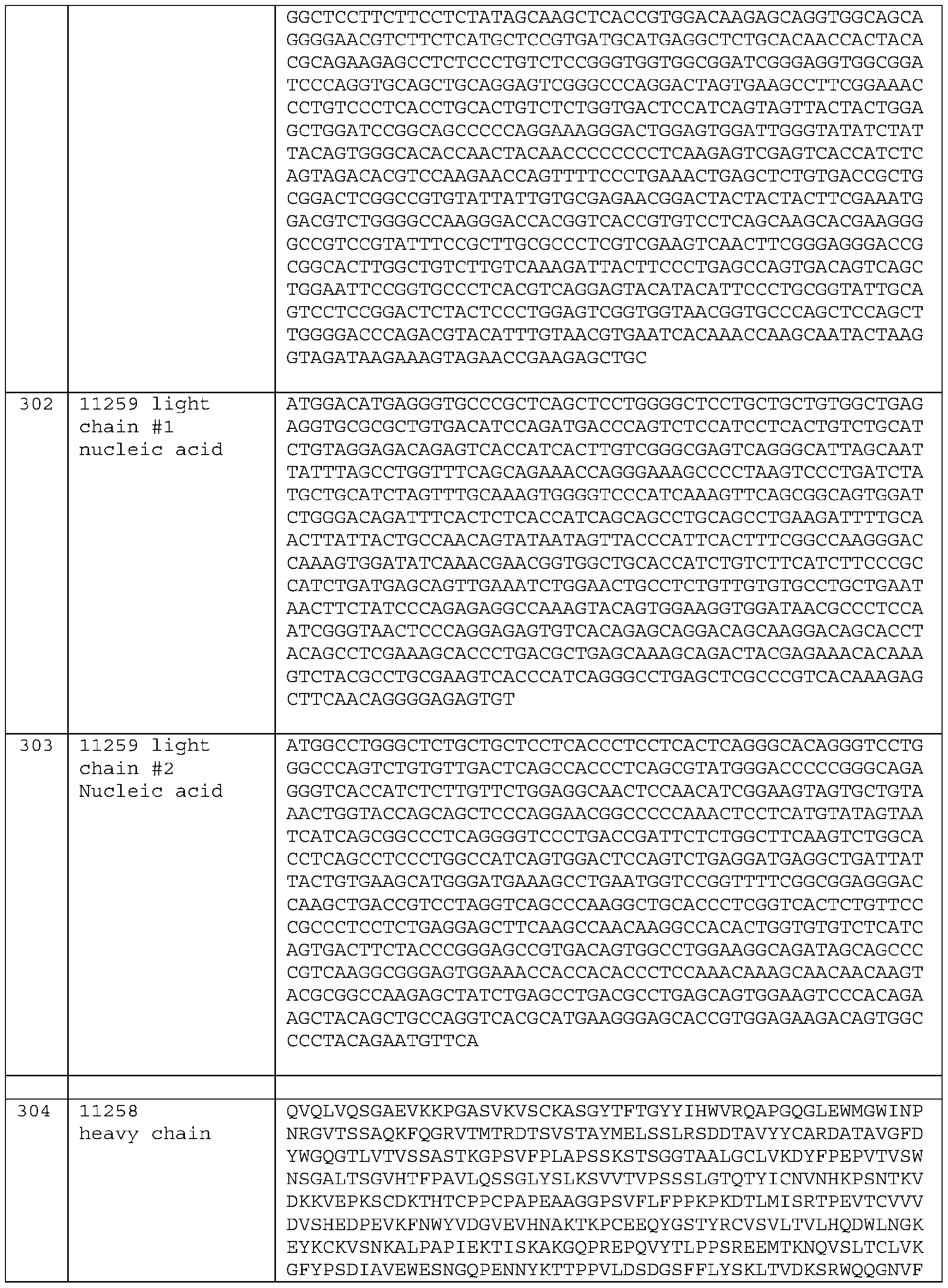
- a bispecific molecule comprising two copies of SEQ ID NO: 297, two copies of SEQ ID NO: 299, and two copies of SEQ ID NO: 300. (clone 11259) E257.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 304; (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 306; and (iii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%
- a bispecific molecule comprising two copies of SEQ ID NO: 304, two copies of SEQ ID NO: 306, and two copies of SEQ ID NO: 307. (clone 11258) E259.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 311; (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 312; and (iii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%,
- a bispecific molecule comprising two copies of SEQ ID NO: 311, two copies of SEQ ID NO: 312, and two copies of SEQ ID NO: 307. (clone 11262) E261.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 315; (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 312; and (iii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least
- a bispecific molecule comprising two copies of SEQ ID NO: 315, two copies of SEQ ID NO: 312, and two copies of SEQ ID NO: 317. (clone 11264) E263.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 320; (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 312; and (iii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%,
- a bispecific molecule comprising two copies of SEQ ID NO: 320, two copies of SEQ ID NO: 312, and two copies of SEQ ID NO: 300. (clone 11265) E265.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 443, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 444.
- a bispecific molecule comprising two copies of SEQ ID NO: 443, and two copies of SEQ ID NO: 444. (clone 44988) E267.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 434, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 435.
- a bispecific molecule comprising two copies of SEQ ID NO: 434, and two copies of SEQ ID NO: 435. (clone 56039) E269.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 449, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 450.
- a bispecific molecule comprising two copies of SEQ ID NO: 449, and two copies of SEQ ID NO: 450. (clone 56040) E271.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 451, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 452.
- a bispecific molecule comprising two copies of SEQ ID NO: 451, and two copies of SEQ ID NO: 452. (clone 56041) E273.
- a bispecific molecule comprising (i) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 437, (ii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 438, (iii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO
- a bispecific molecule comprising: one copy of SEQ ID NO: 449, one copy of SEQ ID NO: 450, one copy of SEQ ID NO:439, and one copy of SEQ ID NO: 440. (clone 56042) E275.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 432, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 433.
- E276 A bispecific molecule comprising two copies of SEQ ID NO: 432, and two copies of SEQ ID NO: 433.
- a bispecific molecule comprising (i) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 446, (ii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 447, and (iii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 448.
- a bispecific molecule comprising: one copy of SEQ ID NO: 446, one copy of SEQ ID NO: 447, and one copy of SEQ ID NO: 448. (clone 56639) E279.
- a bispecific molecule comprising (i) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 429, (ii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 430, and (iii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least
- a bispecific molecule comprising: one copy of SEQ ID NO: 429, one copy of SEQ ID NO: 430, and one copy of SEQ ID NO: 431. (clone 56761) E281.
- a bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 442, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 441.
- a bispecific molecule comprising two copies of SEQ ID NO: 442, and two copies of SEQ ID NO: 441. (clone 56762) E283.
- the bispecific molecule of any one of E99-E102 and E166-E282, wherein said bispecific molecule activates 4-1BB upon binding to PD-L1.
- a nucleic acid comprising a nucleotide sequence encoding the 4-1BB antigen-binding protein of any one of E1-E98 and E319-E346, the PD-L1 antigen-binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166-E284, or one of the polypeptide chains of any one of the foregoing. E286.
- the nucleic acid of E285, comprising a nucleotide sequence that is at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical any one of SEQ ID NOs: 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465.
- E287 A vector comprising the nucleic acid of E285 or E286.
- E288 A host cell comprising the nucleic acid of E285 or E286, or the vector of E287.
- E289. The host cell of E288, wherein said host cell is a mammalian cell.
- E290. The host cell of E289, wherein said host cell is a CHO cell or a HEK-293 cell, or an Sp2.0 cell.
- a kit comprising (i) the 4-1BB antigen-binding protein of any one of E1-E98 and E319-E346, the PD-L1 antigen-binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166-E284, the nucleic acid of E285 or E286, the vector of E287, the host cell of E288-E290, or a combination thereof; and (ii) instructions for use. E292.
- a pharmaceutical composition comprising (i) the 4-1BB antigen-binding protein of any one of E1- E98 and E319-E346, the PD-L1 antigen-binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166-E284, the nucleic acid of E285 or E286, the vector of E287, the host cell of E288-E290, or a combination thereof; and (ii) a pharmaceutically acceptable carrier, excipient, or diluent. E293.
- E295. A method of treating cancer, comprising administering to a subject in need thereof a therapeutically effective amount of the 4-1BB antigen-binding protein of any one of E1-E98 and E319- E346, the PD-L1 antigen-binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166-E284, or the pharmaceutical composition of E292.
- E296 The method of E295, wherein cancer is solid tumor.
- E297 The method of E295 or E296, wherein said subject is a human.
- E298 The method of any one of E295-E297, wherein said cancer comprises cells that express PD-L1. E299.
- the cancer is brain cancer, bladder cancer, breast cancer, clear cell kidney cancer, cervical cancer, colon cancer, rectal cancer, endometrial cancer, gastric cancer, head/neck squamous cell carcinoma, lip cancer, oral cancer, liver cancer, lung squamous cell carcinoma, melanoma, meso
- the cancer is adrenocortical tumor, alveolar soft part sarcoma, carcinoma, chondrosarcoma, desmoid tumors
- E301 The method of any one of E295-E298, wherein the cancer is acute lymphoblastic leukemia (ALL), acute myeloid leukemia (AML), chronic lymphocytic leukemia (CLL), or chronic myeloid leukemia (CML).
- ALL acute lymphoblastic leukemia
- AML acute myeloid leukemia
- CLL chronic lymphocytic leukemia
- CML chronic myeloid leukemia
- E302 The method of any one of E295-E298, wherein the cancer is diffuse large B-cell lymphoma (DLBCL), follicular lymphoma, Hodgkin's lymphoma (HL), mantle cell lymphoma (MCL), multiple myeloma (MM), myelodysplastic syndrome (MDS), non-Hodgkin's lymphoma (NHL), or small lymphocytic lymphoma (SLL).
- DLBCL diffuse large B-cell lymphoma
- HL
- any one of E295-E302, wherein said 4-1BB antigen-binding protein, PD-L1 antigen-binding protein, bispecific molecule, fusion protein, polypeptide, or pharmaceutical composition is administered about twice a week, once a week, once every two weeks, once every three weeks, once every four weeks, once every five weeks, once every six weeks, once every seven weeks, once every eight weeks, once every nine weeks, once every ten weeks, twice a month, once a month, once every two months, once every three months, or once every four months.
- a method of assessing immunogenicity of a bispecific molecule wherein said immunogenicity is attributed to a T-cell epitope, and wherein said bispecific molecule comprises two domains: (1) a first domain that binds to a Dendritic Cell (DC) surface antigen; and (2) a second domain that binds to a T cell co-stimulatory molecule, the method comprises: (a) obtaining a first protein that comprises said first domain but does not comprise said second domain; (b) obtaining a second protein that comprises said second domain but does not comprise said first domain; (c) incubating said first protein and second protein with a cell culture that comprises DCs and T cells; and (d) assessing the activation or proliferation of T cells, wherein the activation or proliferation of T cells is indicative that said bispecific molecule comprises an immunogenic T cell epitope.
- DC Dendritic Cell
- E311 The method of E310, wherein said DC surface antigen is PD-L1.
- E312 The method of E311, wherein said DC surface antigen is CD8A, CLEC9A, ITGAE, ITGAX, THBD (CD141), XCR, CD1C, CD207, ITGAM, NOTCH2, SIRPA, CLEC4C, LILRB4, NRP1, CCR7, CD14, MRC1 (CD206), CD209, or CD1A.
- T-cell co-stimulatory molecule is CD28, Inducible Co-Stimulator (ICOS), CTLA4 (Cytotoxic T-Lymphocyte-Associated protein 4), 4-1BB (also known as CD137), OX40 (also known as CD134), CD27, CD30, DR3, Glucocorticoid-Induced TNFR family Related (GITR), or Herpes Virus Entry Mediator (HVEM).
- T-cell co-stimulatory molecule is CD28, Inducible Co-Stimulator (ICOS), CTLA4 (Cytotoxic T-Lymphocyte-Associated protein 4), 4-1BB (also known as CD137), OX40 (also known as CD134), CD27, CD30, DR3, Glucocorticoid-Induced TNFR family Related (GITR), or Herpes Virus Entry Mediator (HVEM).
- E314 The method of any one of E310-E313, wherein said T-cell co-stimulatory molecule is 4
- E317 The method of any one of E310-E315, wherein said second protein further comprise an IgG Fc domain.
- said Fc domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, or 426.
- E318 The method of E315 or E316, wherein said Fc domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 263, 267, or 483. E319.
- a 4-1BB antigen-binding protein comprising a heavy chain variable domain (VH) and does not comprise light chain variable domain (VL), wherein said protein binds to an epitope that comprises residues C102, V71, and Q104, according to the numbering of SEQ ID NO:272.
- E320 The 4-1BB antigen-binding protein of E319, wherein said epitope further comprises residue P90, according to the numbering of SEQ ID NO:272.
- the 4-1BB antigen-binding protein of E319 or E320, wherein said epitope further comprises one or more residues selected from the group consisting of: K69, T89, F92, M101, and L112, according to the numbering of SEQ ID NO:272. E322.
- E323 The 4-1BB antigen-binding protein of any one of E319-E322, wherein one or more of the following substitutions substantially disrupts the binding of said antigen-binding protein to said epitope: (1) C102 is replaced by A; (2) Q104 is replaced by A; or (3) K69 is replaced by A. E324.
- KD binding affinity
- E327. The 4-1BB antigen-binding protein of any one of E319-E326, wherein said 4-1BB is a human 4- 1BB.
- E328. The 4-1BB antigen-binding protein of E327, wherein said 4-1BB comprises SEQ ID NO:566. E329.
- E330. The 4-1BB antigen-binding protein of any one of E319-E329, wherein said VH comprises (VH numbering according to Kabat): (1) H45 is Leu, Phe, Ile, or Tyr; (2) H47 is Trp, Phe, Leu, or Tyr; (3) H100B is Tyr, Arg, His, Lys, or Met; (4) H100D is Thr, Ala, Asn, Cys, Gln, Lys, Met, or Val; and (5) H100F is Phe, Trp or Tyr.
- VH comprises (VH numbering according to Kabat): (1) H45 is Leu, Phe, Ile, or Tyr; (2) H47 is Trp, Phe, Leu, or Tyr; (3) H100B is Tyr, Arg, His, Lys, or Met; (4) H100D is Thr
- E332 The 4-1BB antigen-binding protein of any one of E319-E331, wherein said VH comprises (VH numbering according to Kabat): (1) H45 is Leu; (2) H47 is Trp; (3) H100B is Tyr; (4) H100D is Thr; and (5) H100F is Phe.
- E333 comprises (VH numbering according to Kabat): (1) H45 is Leu; (2) H47 is Trp; (3) H100B is Tyr; (4) H100D is Thr; and (5) H100F is Phe.
- VH comprises (VH numbering according to Kabat): (6) H97 is Ser, Arg, Asn, Gln, Glu, His,,Leu, Lys, Met, Phe, Thr, Trp, Tyr, or Val; (7) H100E is Ser, Ala, Asn, Asp, Cys, His, Trp, Tyr, or Val; and (9) H102 is Tyr, Ile, Lys, or Val. E334.
- the 4-1BB antigen-binding protein of any one of E319-E333 wherein said VH comprises (VH numbering according to Kabat): (6) H97 is Ser or Leu; (7) H100E is Ser or Val; and (9) H102 is Tyr or Lys.
- E340. The 4-1BB antigen-binding protein of any one of E310-E338, comprising a VH framework derived from a human germline VH1 framework sequence.
- E341. The 4-1BB antigen-binding protein of any one of E319-E338, comprising a VH framework derived from a human germline VH3 framework sequence.
- E342 The 4-1BB antigen-binding protein of any one of E319-E338, comprising a VH framework derived from a human germline VH4 framework sequence.
- the 4-1BB antigen-binding protein of any one of E319-E344, comprising a VH framework sequence is at least 90% identical to the human germline framework sequence from which it is derived. E346.
- the 4-1BB antigen-binding protein of any one of E319-E345, comprising a VH framework sequence is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to the human germline framework sequence from which it is derived.
- FIG.1A is a schematic of the steps taken to generate 4-1BB mAbs and PD-L1 mAbs of the present disclosure.
- FIG.1B is a schematic of the steps taken to generate bispecific molecules comprising a 4-1BB binding protein and PD-L1 binding protein of the present disclosure.
- FIG.2A is an illustration of a bispecific molecule comprising an IgG moiety and a scFv moiety.
- FIG. 2B is an illustration of a bispecific molecule comprising an IgG moiety and a Fab moiety.
- FIG.3 is a table listing engineered binding proteins and characteristics thereof.
- FIG.4 is a table listing bispecific molecules generated herein and characteristics thereof.
- FIG.5 is a schematic representation of bispecific molecules of interest (dotted lines showing where the IdeS enzyme cleaves), and 2 representative LC-MS spectra of the bispecific molecules. Representative graphs from clones 11259 and 11262 are shown.
- FIG.6A is a graph of the IL-2 produced by T-cells activated by artificial APCs plotted as a function of bispecific molecule concentration.
- FIG.6B is a graph of the IL-2 produced by T-cells activated by artificial APCs plotted as a function of bispecific molecule concentration.
- FIGs.7A-7D are graphs of the tumor volume plotted as a function of time following tumor implant for mice treated with the active agent indicated at the top of each graph.
- FIG.8A is a graph of the tumor volume plotted as a function of time following tumor implant for mice treated with the active agents indicated in the key.
- FIG.8B is a graph of the percent survival plotted as a function of time following tumor implant for mice treated with the active agents indicated in the key.
- FIG.9 is a table listing characteristics of 4-1BB antigen-binding proteins described herein.
- FIGs.10A-10B are graphs showing crosslinking dependent activity of 4-1BB UniAbs compared to crosslinker independent control.
- FIGs.11A-11D are graphs showing cell binding dose curves for anti-41BB UniAbs using activated T-cells, CHO-human 4-1BB, CHO cyno 4-1BB, off target cell lines compared to a crosslinker independent control.
- FIG.12 a schematic of the steps taken to generate bispecific molecules comprising a 4-1BB binding protein and PD-L1 binding protein of the present disclosure.
- FIGs.13A-13K are illustrations of bispecific molecule format used in second round of screening. Letters “a” and “b” are used to indicate the first antigen (“a”) or the second antigen (“b”) that the bispecific molecule binds to.
- N means that the second antigen-binding moiety is fused at the N-terminus of the first antigen-binding molecule, and the number refers to the number of antigen-binding moieties.
- N2 two “b” antigen-binding moieties fused at the N-termini of the “a’ antigen-binding molecule.
- B means the second antigen-binding moiety is fused at the hinge region, and
- C means the second antigen-binding moiety is fused at the C-terminus.
- FIGs.14A-14D provides domain information of certain bispecific formats, including IgG-scFv, Fab- scFv-Fc, IgG-VH, and Fab-VH-Fc.
- FIGs.15A-15B summarizes the biophysical properties of the bispecific molecules disclosed herein.
- FIGs.16A-16C show the potencies of the bispecific molecules disclosed herein.
- FIG.17 shows that the activation of 4-1BB by the bispecific molecule is dependent on the binding of PDL1.
- FIGs.18A-18B show the pre-ADA assessment of the bispecific molecules disclosed herein.
- FIGs.19A-19B shows the immunogenicity assessment of the bispecific molecules disclosed herein.
- FIG.20A shows the overall structure of human 4-1BB and molecule 380984.
- the crystal structure of human 4-1BB in complex with 380984 is shown as cartoon.4-1BB is in light gray color, and Amgen- 380984 is in dark gray color.
- the N-, C- terminus, and CDR-3 are labeled.
- the residues engaged in ligand 4-1BBL binding are marked as sphere.
- FIG.20B shows the close-up view of the contact between human 4-1BB and 380984.
- Human 4-1BB (colored in light gray) and Amgen-380984 (colored in dark gray) are illustrated as transparent cartoon. All interacting residues are shown as sticks; the back-bone nitrogen atoms are shown as sphere. Hydrogen bonds are represented as gray dashed lines.
- FIGs 21A-21G are graphs showing the comparison of 4-1BB binders and their potential orientation on cell membrane.
- the structure of human 4-1BB is shown as surface in color light gray; its binders are shown as cartoon in color dark gray.
- A Three human 4-1BB receptors bind to a trimeric ligand 4-1BBL, each protomer of human 4-1BB are labeled as number 1, 2, 3. PDB: 6mgp.
- B Protomer 2 of the 4-1BB- 4-1BBL complex is shown in the same orientation as in A. (C).
- the structure of human 4-1BB and Urelumab complex (BMS, PDB:6MHR) is superimposed to the protomer 2 of 4-1BB-4-BBL complex by aligning on the CRDs 3 and 4. The location of Fc is labeled.
- D Protomer 1 of the 4-1BB-4-1BBL complex is shown in the same orientation as in A.
- E The structure of human 4-1BB and 380984 is superimposed to the protomer 1 of 4-1BB-4-BBL complex by aligning the CRDs 3 and 4. The location of Fc is labeled.
- F The structure of human 4-1BB and Utomilumac (Pfizer, PDB: 6MI2) is superimposed to the promotor by aligning the CRDs 3 and 4.
- 4-1BB is an immune co-stimulatory protein expressed on activated T cells.4-1BB agonist antibodies have demonstrated efficacy in prophylactic and therapeutic settings in both monotherapy and combination therapy tumor models, and have established protective & durable anti-tumor T-cell memory responses. Agonizing 4-1BB by crosslinking antibody enhances T cell proliferation, survival and cytokine production upon TCR engagement.
- phase I and II data from Urelumab revealed a liver toxicity that appeared to be on target and dose dependent, halting initial clinical development of Urelumab.
- the anti-hu4-1BB huIgG2 utomilumab (PF-05082566) displays a better safety profile but lower agonistic potency. [34] Seeking to overcome this challenge, the inventors designed bispecific molecules to promote target- mediated clustering of 4-1BB.
- 4-1BB undergoes trimerization upon binding to its ligand (4-1BBL); and 4-1BB multimerization and clustering (“crosslinking”) is a prerequisite for its signaling pathway. Therefore, the 4-1BB antibodies were specifically selected as “crosslinking dependent,” meaning that the agonistic activity of the antibody is dependent upon the crosslinking of 4- 1BB. Without 4-1BB crosslinking, the binding of the antibody to 4-1BB leads to minimal agonist activity, thereby avoiding toxicities exhibited by Urelumab. Urelumab is believed to be a crosslinking independent agonist antibody, meaning the binding of Urelumab to 4-1BB is sufficient to trigger 4-1BB activation.
- PD-L1 which is also known as B7-H1 or CD274, is expressed in multiple types of cancers (e.g., breast cancer, lung cancer, melanoma).
- PD-L1 expressed on tumor cells, binds to the inhibitory checkpoint receptor, PD-1, on activated tumor infiltrating lymphocytes.
- the interaction between PD-L1 and PD-1 delivers an inhibitory signal to T cells and ultimately prevents tumor elimination.
- the data also demonstrate the dual activity upon administration of a combination of a monospecific 4-1BB binding protein and a monospecific PD-L1 binding protein.
- agonizing 4-1BB enhances T-cell proliferation, survival & cytokine production upon TCR engagement; and
- simultaneously blocking PD-L1 and agonizing 4-1BB can have synergistic anti-tumor activities.
- the bispecific molecules described herein address the dose-limiting hepatotoxicity of systemic 4-1BB agonism, because 4-1BB activation is crosslinking dependent. For normal tissues that do not express PD-L1 or express low levels of PD-L1 (such as hepatic cell), minimal crosslinking of 4-1BB will occur, and 4-1BB activation will be limited. In contrast, in cancer cells that express high level of PD- L1, through PD-L1-binding, the bispecific molecules are brought to the proximity of each other, thus promoting 4-1BB crosslinking and T-cell co-stimulation. Systemic toxicities should be limited because 4- 1BB activation will be largely confined to tumors that express high level of PD-L1.
- the present disclosure provides bispecific molecules comprising a 4-1BB binding protein and a PD-L1 binding protein.
- Related antigen binding proteins including, for instance, monospecific 4-1BB binding proteins and monospecific PD-L1 binding proteins, are further provided.
- Antigen Binding Proteins 2.1 Antigen Binding Protein Types
- the antigen-binding proteins of the present disclosure can take any one of many forms of antigen- binding proteins known in the art.
- the antigen-binding protein is an antibody or immunoglobulin, or an antigen binding fragment of an antibody or immunoglobulin, or an antibody protein product.
- antibodies form a family of plasma proteins known as immunoglobulins and comprise of immunoglobulin domains.
- an antibody refers to a protein having a conventional immunoglobulin format, comprising heavy and light chains, and comprising variable and constant regions.
- an antibody may be an IgG which is a “Y-shaped” structure of two identical pairs of polypeptide chains, each pair having one “light” (typically having a molecular weight of about 25 kDa) and one “heavy” chain (typically having a molecular weight of about 50-70 kDa).
- An antibody has a variable region and a constant region.
- variable region is generally about 100-110 or more amino acids, comprises three complementarity determining regions (CDRs), is primarily responsible for antigen recognition, and substantially varies among other antibodies that bind to different antigens.
- the constant region allows the antibody to recruit cells and molecules of the immune system.
- the variable region is made of the N-terminal regions of each light chain and heavy chain, while the constant region is made of the C-terminal portions of each of the heavy and light chains.
- Antibodies can comprise any constant region known in the art. Human light chains are classified as kappa and lambda light chains. Heavy chains are classified as mu, delta, gamma, alpha, or epsilon, and define the antibody's isotype as IgM, IgD, IgG, IgA, and IgE, respectively.
- IgG has several subclasses, including, but not limited to IgG1, IgG2, IgG3, and IgG4.
- IgM has subclasses, including, but not limited to, IgM1 and IgM2.
- Embodiments of the present disclosure include all such classes or isotypes of antibodies.
- the light chain constant region can be, for example, a kappa- or lambda-type light chain constant region, e.g., a human kappa- or lambda-type light chain constant region.
- the heavy chain constant region can be, for example, an alpha-, delta-, epsilon-, gamma-, or mu-type heavy chain constant regions, e.g., a human alpha-, delta-, epsilon-, gamma-, or mu-type heavy chain constant region.
- the antibody is an antibody of isotype IgA, IgD, IgE, IgG, or IgM, including any one of IgG1, IgG2, IgG3 or IgG4.
- the antibody can be a monoclonal antibody or a polyclonal antibody.
- the antibody comprises a sequence that is substantially similar to a naturally-occurring antibody produced by a mammal, e.g., mouse, rabbit, goat, horse, chicken, hamster, human, and the like.
- the antibody can be considered as a mammalian antibody, e.g., a mouse antibody, rabbit antibody, goat antibody, horse antibody, chicken antibody, hamster antibody, human antibody, and the like.
- the antibody is a human antibody.
- the antibody is a chimeric antibody or a humanized antibody.
- the term "chimeric antibody" refers to an antibody containing domains from two or more different antibodies.
- a chimeric antibody can, for example, contain the constant domains from one species and the variable domains from a second, or more generally, can contain stretches of amino acid sequence from at least two species.
- a chimeric antibody also can contain domains of two or more different antibodies within the same species.
- humanized when used in relation to antibodies refers to antibodies having at least CDR regions from a non-human source which are engineered to have a structure and immunological function more similar to true human antibodies than the original source antibodies.
- humanizing can involve grafting a CDR from a non-human antibody, such as a mouse antibody, into a human antibody.
- Humanizing also can involve select amino acid substitutions to make a non-human sequence more similar to a human sequence.
- An antibody can be cleaved into fragments by enzymes, such as, e.g., papain and pepsin. Papain cleaves an antibody to produce two Fab fragments and a single Fc fragment.
- the antigen binding protein of the present disclosure comprises an antigen binding fragment of an antibody.
- antigen binding fragment of an antibody refers to a portion of an antibody molecule that retains the ability to specifically bind to an antigen (preferably with substantially the same binding affinity).
- an antigen-binding fragment examples include but not limited to: (i) a Fab fragment, a monovalent fragment consisting of the VL, VH, CL and CH1 domains; (ii) a F(ab')2 fragment, a bivalent fragment comprising two Fab fragments linked by a disulfide bridge at the hinge region; (iii) a Fd fragment consisting of the VH and CH1 domains; (iv) a Fv fragment consisting of the VL and VH domains of a single arm of an antibody, (v) a dAb fragment (Ward et al., 1989 Nature 341 :544-546), which consists of a VH domain.
- a Fab fragment a monovalent fragment consisting of the VL, VH, CL and CH1 domains
- a F(ab')2 fragment a bivalent fragment comprising two Fab fragments linked by a disulfide bridge at the hinge region
- a Fd fragment consisting of the VH and
- VL and VH are coded for by separate genes, they can be joined, using recombinant methods, by a synthetic linker that enables them to be made as a single protein chain in which the VL and VH regions pair to form monovalent molecules (see e.g., Bird et al. Science 242:423- 426 (1988) and Huston et al., 1988, Proc. Natl. Acad. Sci. USA 85:5879-5883).
- Other forms of single chain antibody protein products, such as diabodies are also encompassed.
- Diabodies are bivalent, bispecific antibody protein products in which VH and VL domains are expressed on a single polypeptide chain, but using a linker that is too short to allow for pairing between the two domains on the same chain, thereby forcing the domains to pair with complementary domains of another chain and creating two antigen-binding sites (see e.g., Holliger et al, 1993, Proc. Natl. Acad. Sci. USA 90:6444-6448; Poljak et al., 1994, Structure 2:1121 -1123).
- antibody protein products include disulfide-bond stabilized scFv (ds-scFv), single chain Fab (scFab), as well as di- and multimeric antibody formats like dia-, tria- and tetra-bodies, or minibodies (miniAbs) that comprise different formats consisting of scFvs linked to oligomerization domains.
- minibodies minibodies that comprise different formats consisting of scFvs linked to oligomerization domains.
- the smallest fragments are VHH/VH of camelid heavy chain Abs as well as single domain Abs (sdAb).
- a peptibody or peptide-Fc fusion is yet another antibody protein product.
- the structure of a peptibody consists of a biologically active peptide grafted onto an Fc domain.
- Peptibodies are well-described in the art.
- Antigen-binding proteins disclosed herein also include “heavy chain only” molecules. This type of antigen-binding proteins lack the light chain of a conventional antibody. Examples of such “heavy chain only” molecules include, for example, single domain molecules such as UniDabs® (VH only); and homodimeric molecules comprising the VH antigen-binding domain and the CH2 and CH3 constant domains, in the absence of the CH1 domain (e.g., UniAb®). Non-limiting examples of “heavy chain only” antigen binding proteins are described, for example, in WO2018/039180.
- Such heavy chain only molecules can be produced by UniRat®, which is a triple knockout rat wherein the expressions of the native variable coding sequences and the heavy and light chain constant regions have been inactivated.
- the UniRat has been genetically modified to exclusively express the full human VDJ repertoire (all VH families), with transgenes of human heavy chain variable domains linked to a conserved rat Fc. Immunization of the UniRat elicits a normal antibody response that results in the expression of UniAbs, human heavy-chain-only antibodies of approximately 80 kDa, contrasting with the standard ⁇ 150 kDa human IgG.
- VH domains from the UniRat can be assembled as modular domains of multispecific antigen binding proteins.
- Bispecific formats can generally be divided into five major classes: BsIgG, appended IgG, BsAb fragments, bispecific fusion proteins and BsAb conjugates. See, e.g., Spiess et al., Molecular Immunology 67(2) Part A: 97-106 (2015).
- the antigen binding protein of the present disclosure comprises any one of these antibody protein products.
- the antigen binding protein of the present disclosure comprises any one of an scFv, Fab, VHH, VH, Fv fragment, ds-scFv, scFab, UniDab, dimeric antibody, multimeric antibody (e.g., a diabody, triabody, tetrabody), miniAb, peptibody VHH/VH of camelid heavy chain antibody, sdAb, diabody; UniAb; a triabody; a tetrabody; a bispecific or trispecific antibody, BsIgG, appended IgG, BsAb fragment, bispecific fusion protein, and BsAb conjugate.
- the antigen binding protein of the present disclosure comprises an antibody protein product in monomeric form, or polymeric, oligomeric, or multimeric form.
- the antibody comprises two or more distinct antigen binding regions fragments
- the antibody is considered bispecific, trispecific, or multi-specific, or bivalent, trivalent, or multivalent, depending on the number of distinct epitopes that are recognized and bound by the antibody.
- Many of the antigen-binding proteins disclosed herein comprise two different chains, one derived from the heavy chain of an antibody, and one derived from the light chain of an antibody.
- the heavy/light chain has been modified and is no longer the classical immunoglobulin heavy/light chain, for convenience, it is still generally called “heavy chain” or “HC” if it is based on heavy chain backbone, and “light chain” or “LC” if it is based on light chain backbone.
- HC heavy chain
- LC light chain backbone
- the “HC” comprises an IgG heavy chain fused to an scFv. It would be apparent to a skilled artisan whether HC is a traditional immunoglobulin heavy chain or a modified version based on immunoglobulin heavy chain backbone.
- Some exemplary antigen binding proteins disclosed herein are characterized by the location or epitopes they bind to, or by the paratopes that they comprise.
- An “epitope” refers to the area or region of an antigen to which an antigen binding protein specifically binds, e.g., an area or region comprising residues that interacts with the antigen-binding protein.
- Epitopes can be linear or conformational. Epitopes can be determined by any method well known in the art. For example, epitopes can be determined by conventional immunoassays. Alternatively, one may competitively screen antigen-binding proteins for binding to the same epitope.
- the term “paratope” is derived from the above definition of “epitope” by reversing the perspective, and refers to the area or region of an antigen-binding protein which is involved in binding of an antigen, e.g., an area or region comprising residues that interacts with the antigen.
- a paratope may be linear or conformational (such as discontinuous residues in CDRs).
- the epitope/paratope can be defined and characterized at different levels of detail using a variety of experimental and computational epitope mapping methods.
- the experimental methods include mutagenesis, X-ray crystallography, Nuclear Magnetic Resonance (NMR) spectroscopy, Hydrogen/deuterium exchange Mass Spectrometry (HX-MS), cryo-EM, and various competition binding methods.
- NMR Nuclear Magnetic Resonance
- HX-MS Hydrogen/deuterium exchange Mass Spectrometry
- cryo-EM cryo-EM
- various competition binding methods As each method relies on a unique principle, the description of an epitope is linked to the method by which it has been determined. Thus, the epitope/paratope for a given binding pair will be defined differently depending on the mapping method employed.
- the epitope/paratope for the interaction between the antigen and the antigen-binding protein can be defined by the spatial coordinates defining the atomic contacts present in the interaction, as well as information about their relative contributions to the binding thermodynamics.
- an epitope/paratope residue can be characterized by the spatial coordinates defining the atomic contacts between the binding pair.
- the epitope/paratope residue can be defined by a specific criterion, e.g., distance between atoms in the antigen and the antigen-binding protein (e.g., a distance of equal to or less than 4.5 ⁇ from a heavy atom of the antigen and a heavy atom of the antigen- binding protein ("contact" residues)).
- an epitope/paratope residue can be characterized as participating in a hydrogen bond interaction with the cognate antibody/antigen, or with a water molecule that is also hydrogen bonded to the antigen/antigen-binding protein (water-mediated hydrogen bonding).
- an epitope/paratope residue can be characterized as forming a salt bridge with a residue of the cognate antibody/antigen.
- an epitope/paratope residue can be characterized as a residue having a non-zero change in buried surface area (BSA) due to the interaction between the antigen and the antigen-binding protein.
- BSA buried surface area
- epitope/paratope can be characterized through function, e.g., by competition binding with other antigen-binding molecules.
- the epitope/paratope can also be defined more generically as comprising amino acid residues for which substitution by another amino acid will alter the characteristics of the interaction between the binding pair (e.g., alanine scanning).
- a 4-1BB epitope residue refers to a 4-1BB residue: (i) having a heavy atom (i.e., a non-hydrogen atom) that is within a distance of 4.5 ⁇ from a heavy atom of the antigen-binding protein (also called “contact” residues); (ii) participating in a hydrogen bond with a residue of the antigen-binding protein, or with a water molecule that is also hydrogen bonded to the antigen-binding protein (water- mediated hydrogen bonding), (iii) participating in a salt bridge to a residue of the antigen-binding protein, and/or (iv) having a non-zero change in buried surface area (BSA) due to interaction with the antigen-binding protein.
- a heavy atom i.e., a non-hydrogen atom
- BSA buried surface area
- epitope residues under category (iv) are selected if it has a BSA of 20 ⁇ 2 or greater, or is involved in electrostatic interactions when the antigen-binding protein binds to 4-1BB.
- a paratope residue refers to an antigen-binding protein residue (i) having a heavy atom (i.e., a non-hydrogen atom) that is within a distance of 4.5 ⁇ from a heavy atom of 4-1BB (also called “contact” residues), (ii) participating in a hydrogen bond with an antigen residue, or with a water molecule that is also hydrogen bonded to 4-1BB (water-mediated hydrogen bonding), (iii) participating in a salt bridge to a residue of 4-1BB, and/or (iv) having a non-zero change in buried surface area due to interaction with 4- 1BB.
- a heavy atom i.e., a non-hydrogen atom
- 4-1BB also called “contact” residues
- participating in a hydrogen bond with an antigen residue or with a water molecule that is also hydrogen bonded to 4-1BB (water-mediated hydrogen bonding)
- (iii) participating in a salt bridge to a residue of 4-1BB and/
- paratope residues under category (iv) are selected if it has a BSA of 20 ⁇ 2 or greater, or is involved in electrostatic interactions when the antigen-binding protein binds to 4- 1BB.
- BSA BSA of 20 ⁇ 2 or greater
- paratope residues under category (iv) are selected if it has a BSA of 20 ⁇ 2 or greater, or is involved in electrostatic interactions when the antigen-binding protein binds to 4- 1BB.
- Epitopes are said to be separate (unique) if no amino acid residue is shared by the epitopes.
- Epitopes characterized by competition binding are said to be overlapping if the binding of two molecules are mutually exclusive, i.e., binding of one molecule excludes simultaneous or consecutive binding of the other molecule; and epitopes are said to be separate (unique) if the antigen is able to accommodate binding of both molecules simultaneously.
- 4-1BB is a glycosylated type I membrane protein comprises four cysteine-rich pseudo repeats (CRDs) forming the extracellular domain, a short helical transmembrane domain, and a cytoplasmic signaling domain.
- Human 4-1BB (SEQ ID NO: 272; UniProtKB - Q07011) comprises 255 amino acid residues: residues 1-23 forms signal peptide that is cleaved before mature protein is produced, residues 24 – 186 forms extracellular domain, residues 187 – 213 forms transmembrane helical structure, and residues 214 – 255 forms cytoplasmic domain.
- the four CRD domains of human 4-1BB are defined as follows: CRD1: residues 24-45; CDR2, residues 47-86; CRD3: residues 87-118, and CRD4: residues 119- 159.
- the 4-1BB antigen-binding proteins described herein bind to an epitope in CRD1 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to an epitope in CRD2 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to an epitope in CRD3 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to an epitope in CRD4 of 4-1BB. [62] In certain aspects, the 4-1BB antigen-binding proteins described herein bind to CRD1 of 4-1BB.
- the 4-1BB antigen-binding proteins described herein bind to CRD2 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to CRD3 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to CRD4 of 4-1BB.
- FIGs.21A-21G illustrate the advantageous of the binding location as exemplified in Example 14. As shown in FIGs.21C, Urelumab binds to CRD1, and is orientated away from cell surface.
- FIGs.21F and 21G show the binding of Utomilumab to 4-1BB.
- Utomilumab binds to the CDR4 domain. The antibody is close to cell surface and oriented parallel to the cell surface. This orientation results in a large distance between 4-1BB monomers, making 4-1BB crosslinking more difficult (Fig.21F).
- Molecule 380984 binds to CDR3 and is orientated vertically to the cell surface, which is much more desirable than parallel orientation. Compared with Utomilumab, the distance between 4-1BB monomers is much smaller due to its vertical orientation (Fig.21E).
- the term “bind” here refers to specific binding, a term well understood in the art, and methods to determine such specific binding are also well known in the art. A molecule is said to exhibit “binding” if it reacts or associates more frequently, more rapidly, with greater duration and/or with greater affinity with a particular cell or substance than it does with alternative cells or substances.
- an antigen- binding protein “binds” to a target if it binds with greater affinity, avidity, more readily, and/or with greater duration than it binds to other substances.
- a 4-1BB binding protein that binds to the CRD1 domain is a protein that binds this domain with greater affinity, avidity, more readily, and/or with greater duration than it binds to other domains and regions of 4-1BB, or other proteins that are not 4-1BB. It is also understood by reading this definition that, for example, an antigen-binding protein which specifically or preferentially binds to a first target may or may not specifically bind to a second target.
- binding does not necessarily require (although it can include) exclusive binding.
- an antigen-binding protein may bind with a KD value that is numerically less than 1x10 -6 M, 1x10 -7 M, 1x10 -8 M, or 1x10 -9 M.
- the present disclosure provides a 4-1BB antigen-binding protein that binds to an epitope that comprises residues C102, V71, and Q104.
- the epitope may further comprise P90.
- the epitope may further comprise one or more residues selected from the group consisting of: K69, T89, F92, M101, and L112.
- epitope residues are numbered according to the numbering of SEQ ID NO:272 (human 4-1BB).
- Corresponding residues from other 4-1BB homologs, isoforms, variants, or fragments can be identified according to sequence alignment or structural alignment that is known in the art. For example, alignments can be done by hand or by using well-known sequence alignment programs such as ClustalW2, or "BLAST 2 Sequences" using default parameters.
- C102, V71, and Q104 were found to be “primary” residues for molecule 380984 binding; P90 was found to be “contributing” residues for scFv32211 binding; and K69, T89, F92, M101, and L112, while involved in H-bonding and Van Der Waals interactions with 380984, were found to be “optional” residue for 380984 binding. [66] Based on the structural studies, one or more of the following substitutions would likely substantially disrupt the binding of 380984 binding to 4-1BB: (i) C102 is replaced with A; (ii) A104 is replaced with A; or(iii) K69 is replaced with A.
- CDRs Complementarity Determining Regions
- CDRs The “conformational” definition of CDRs is based on residues that make enthalpic contributions to antigen binding (see, e.g., Makabe et al., 2008, J. Biol. Chem., 283:1156-1166). North has identified canonical CDR conformations using a different preferred set of CDR definitions (North et al., 2011, J. Mol. Biol.406: 228-256). In another approach, referred to herein as the “conformational definition” of CDRs, the positions of the CDRs may be identified as the residues that make enthalpic contributions to antigen binding (Makabe et al., 2008, J Biol. Chem.283:1156-1166).
- Martin definition (also called enhanced Chothia definition) combines the Kabat and Chothia definitions and differs from them only in the heavy chain, where CDR- H1 includes all residues of Kabat and Chothia while CDR-H2 is seven residues shorter than that defined by Kabat (Martin, Bioinformatics tools for antibody engineering. Handbook of Therapeutic Antibodies. Weinheim: Wiley-VCH Verlag GmbH; (2008). p.95–117; see also the database maintained by the Institute of Structural and Molecular Biology at the University College London, http://www.bioinf.org.uk/abs/#cdrid).
- CDR boundary definitions may not strictly follow one of the above approaches, but will nonetheless overlap with at least a portion of the Kabat CDRs, although they may be shortened or lengthened in light of prediction or experimental findings that particular residues or groups of residues or even entire CDRs do not significantly impact antigen binding.
- “combined” CDRs may also be used. Therefore, a CDR may refer to CDRs defined by any approach known in the art, including combinations of approaches.
- the CDRs (or other residue of the antibody) may be defined in accordance with any of Kabat, Chothia, North, AbM, Contact, IMGT, Martin, combined Kabat and Chothia, and/or conformational definitions.
- Table N1 shows several commonly used definitions of CDRs: Table N1. Definitions of CDRs. Loop Kabat AbM Chothia 1 Contact 2 IMGT L1 L24-L34 L24-L34 L26-L32 L30-L36 L27-L32 7; C - : 6-3 ; C - : 5 -56; C - 3: 95- 0 . abe s a consensus o C ot a de nton based upon Chothia & Lesk (1987) (e.g., CDR-H3) and Chothia et al., 1989, Nature 342:877-883 (e.g., CDR-H2). 2.
- the 4-1BB antigen- binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:147, 149, 151, 153, 155, 157, 159, 324, 161, 163, 165, 167, 169, 171, 173, 175, 404, 406, 408, or 410; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 148, 150, 152, 154, 156, 158, 160, 325, 162, 164, 166, 168, 170, 172, 174, 176, 405, 407, 409, or 411.
- the three heavy chain CDRs and three light chain CDRs come from the same clone as shown in Sequence Table B and Table K1.
- the CDRs are defined according to Kabat, Chothia, AbM, contact, or IMGT.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:147; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:148.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:149; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:150.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:151; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:152.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:153; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:154.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:155; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:156.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:157; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:158.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:159 or 324; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:160 or 325.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:161; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:162.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:163; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:164.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:165; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:166.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:167; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:168.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:169; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:170.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:171; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:172.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:173; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:174.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:175; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:176.
- the antigen binding protein may further bind to its target 4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:404; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:405.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:406; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:407.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:408; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:409.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:410; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:411.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 147, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 148.
- the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 149, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 150. [91] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 151, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 152.
- the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 153, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 154. [93] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 155, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 156.
- the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 157, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 158. [95] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 159 or 324, and (ii) the light chain CDR-L1, CDR- L2, and CDR-L3 of SEQ ID NO: 160 or 325.
- the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 161, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 162. [97] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 163, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 164.
- the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 165, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 166. [99] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 167, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 168.
- the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 169, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 170. [101] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 171, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 172.
- the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 173, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 174. [103] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 175, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 176.
- the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 404, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 405. [105] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 406, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 407.
- the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 408, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 409. [107] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 410, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 411.
- the 4-1BB antigen binding proteins comprise (a) CDR-H1 amino acid sequence set forth in Sequence Table B and Table K1, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set for in Table B or Table K1; (b) CDR-H2 amino acid sequence set forth in Sequence Table B and Table K1, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Table B or Table K1; (c) a CDR-H3 amino acid sequence set forth in Sequence Table B and Table K1, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 1-4 amino acids (
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises 3, 4, 5, or all 6 of the amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table B and Table K1.
- the antigen binding protein comprises each of the LC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table B and Table K1 and at least 1 or 2 of the HC CDR amino acid sequences designated by the SEQ ID NOs in under the same clone name of Sequence Table B and Table K1.
- the antigen binding protein comprises each of the HC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table B and Table K1 and at least 1 or 2 of the LC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table B and Table K1.
- the antigen binding protein comprises six CDR amino acid sequences listed under the same clone name in Sequence Tables or comprising six CDR amino acid sequences selected from the group consisting of: (1) SEQ ID NOs: 43-48, (2) SEQ ID NOs: 49-54, (3) SEQ ID NOs: 55-60, (4) SEQ ID NOs: 61-66, (5) SEQ ID NOs: 67-72, (6) SEQ ID NOs: 73-78, (7) SEQ ID NOs: 79-84, (8) SEQ ID NOs: 85-90, (9) SEQ ID NOs: 91-96, (10) SEQ ID NOs: 97-102, (11) SEQ ID NOs: 103-108, (12) SEQ ID NOs: 109-114, (13) SEQ ID NOs: 115-120, (14) SEQ ID NOs: 121-126, (15) SEQ ID NOs: 127-132, (16) SEQ ID NOs: 386-391, (17) SEQ ID NOs: 121
- the CDR sequences of the exemplary 4-1BB antigen-binding proteins disclosed herein can be aligned to identify consensus sequences.
- the CDR sequences are aligned according to Kabat numbering (i.e., residues with the same Kabat numbering are aligned), and the results are shown in Tables N2 & N3. Table N2.
- VH3 X2 A S or G 17 X1 X2 X3 X4 G X5 X6 X7 Y Y Y G M D
- V X1 G or S 1 BB
- VH-only 4-1BB antigen-binding proteins in particular, UniDabs that are derived from UniRat technology.
- exemplified herein are 4-1BB antigen-binding protein that comprises a heavy chain variable domain (VH) and does not comprise a light chain variable domain (VL), wherein said protein comprises the CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, or 385.
- CDRs can be identified according to the definitions of the Kabat, Chothia, the accumulation of both Kabat and Chothia, AbM, contact, North, and/or conformational definitions or any method of CDR determination well known in the art.
- the present disclosure provides a 4-1BB antigen binding protein comprising a heavy chain variable domain (VH) but does not comprise a light chain variable domain (VL), wherein said 4-1BB antigen binding protein comprises the following paratope residues (VH numbering according to Kabat): (1) H45 is Leu, Phe, Ile, or Tyr; (2) H47 is Trp, Phe, Leu, or Tyr; (3) H100B is Tyr, Arg, His, Lys, or Met; (4) H100D is Thr, Ala, Asn, Cys, Gln, Lys, Met, or Val; and (5) H100F is Phe, Trp or Tyr.
- VH numbering according to Kabat (1) H45 is Leu, Phe, Ile, or Tyr; (2) H47 is Trp, Phe, Leu, or Tyr; (3) H100B is Tyr, Arg, His, Lys, or Met; (4) H100D is Thr, Ala, Asn, Cys, Gln, Lys, Met, or Val; and
- H45 is Leu or Phe
- H47 is Trp or Leu
- H100B is Tyr or Met
- H100D is Thr or Val
- H100F is Phe or Trp.
- H45 is Leu; (2) H47 is Trp; (3) H100B is Tyr; (4) H100D is Thr; and (5) H100F is Phe.
- the 4-1BB antigen binding protein may further comprise the following paratope residues: (VH according to Kabat): (6) H97 is Ser, Arg, Asn, Gln, Glu, His,,Leu, Lys, Met, Phe, Thr, Trp, Tyr, or Val; (7) H100E is Ser, Ala, Asn, Asp, Cys, His, Trp, Tyr, or Val; and (9) H102 is Tyr, Ile, Lys, or Val. Based on structural studies, these paratope residues were found to be “optional” residues from clone 380984that bind to 4-1BB.
- ((6) H97 is Ser or Leu; (7) H100E is Ser or Val; and (9) H102 is Tyr or Lys. More preferably, (6) H97 is Ser; (7) H100E is Ser; and (9) H102 is Tyr.
- CDR-H3 is responsible for contacting FAP residues.
- Leu45 and Trp47 are characterize as framework residues under Kabat, AbM, Chothia, and IMGT definitions. Contact definition characterizes Trp47 as part of CDR-H2.
- the 4-1BB antigen-binding protein comprises (a) a VH that comprises: (i) a CDR-H3 comprising any one of SEQ ID NOs:576-578 and 580; and (ii) framework residue H45 is Leu and framework residue H47 is Trp; or (b) a VH that comprises: (i) a CDR-H2 comprising SEQ ID NO:574, (ii) a CDR-H3 comprising SEQ ID NO: 579; and (iii) framework residue H45 is Leu.
- said framework residues, CDR-H2, or CDR-H3 contact 4-1BB residues C102, V71, Q104, P90, K69, T89, F92, M101, or L112, according to the numbering of SEQ ID NO:272.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:371.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:372.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:373.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:374.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:375.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:376.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:377.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:378.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:379.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:380.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:381.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:382.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:383.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:384.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:385.
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 371.
- the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 372.
- the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 373. [137] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 374. [138] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 375.
- the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 376.
- the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 377.
- the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 378.
- the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 379. [143] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 380. [144] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 381.
- the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 382. [146] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 383. [147] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 384.
- the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 385.
- the 4-1BB antigen binding proteins comprise (a) CDR-H1 amino acid sequence set forth in Sequence Table I, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table I; (b) CDR-H2 amino acid sequence set forth in Sequence Table I, or a variant sequence thereof which differs by only 1- 4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table I; (c) a CDR-H3 amino acid sequence set forth in
- the antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
- the 4-1BB antigen binding protein comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising: (1) SEQ ID NOs: 326-328, respectively (2) SEQ ID NOs: 329-331, respectively, (3) SEQ ID NOs: 332-334, respectively, (4) SEQ ID NOs: 335-337, respectively, (5) SEQ ID NOs: 338-340, respectively, (6) SEQ ID NOs: 341-343, respectively, (7) SEQ ID NOs: 344-346, respectively, (8) SEQ ID NOs: 347-349, respectively, (9) SEQ ID NOs: 350-352, respectively, (10) SEQ ID NOs: 353-355, respectively, (11) SEQ ID NOs: 356-358, respectively, (12) SEQ ID NOs: 359-3
- the CDR sequences of these VH-only 4-1BB antigen-binding proteins can be aligned to identify consensus sequences.
- the CDR sequences are aligned according to Kabat numbering (i.e., residues with the same Kabat numbering are aligned), and the results are shown in Table N9.
- Gaps represent absent residues in Kabat CDR-H2, between H52 and H53 (H52A-H52C), and in Kabat CDR- H3, between H100 and H101 (H100A-H100K).
- Table N10 summarizes the consensus sequences derived from this method. Table N9.
- a VH or VL domain framework comprises four framework sub-regions, FR1, FR2, FR3 and FR4, interspersed with CDRs in the following structure: FR1 - CDR1 - FR2 - CDR2 - FR3 - CDR3 - FR4.
- the antigen binding proteins described herein may comprise a VH framework, such as a human germline VH framework sequence, and a VL framework, such as human germline VL framework sequences.
- VH framework such as a human germline VH framework sequence
- VL framework such as human germline VL framework sequences.
- Preferred human germline light chain frameworks are frameworks derived from V ⁇ or V ⁇ germlines.
- a sequence is “derived from” one or more germlines, what is referred to is a structural relationship, in which the features of a sequence correspond to the noted germline sequences, but may comprise somatic mutations or other amino acid differences relative to the noted germline sequence.
- a sequence to be “derived from” a germline an actual process of deriving that sequence from a germline sequence (either via molecular biology or computational analysis) is not necessarily required.
- VL frameworks may be derived from one of the framework of the following germlines: DPK9 (IMGT name: IGKV1-39), DPK12 (IMGT name: IGKV2D-29), DPK18 (IMGT name: IGKV2-30), DPK24 (IMGT name: IGKV4-1), HK102_V1 (IMGT name: IGKV1-5), DPK1 (IMGT name: IGKV1-33), DPK8 (IMGT name: IGKV1-9), DPK3 (IMGT name: IGKV1-6), DPK21 (IMGT name: IGKV3- 15), Vg_38K (IMGT name: IGKV3-11 ), DPK22 (IMGT name: IGKV3-20), DPK15 (IMGT name: IGKV2- 28), DPL16 (IMGT name: IGLV3-19), DPL8 (IMGT name: IGLV1-40), V1-22 (IMGT name: IGLV6-57).
- the framework sequence may be derived from a human germline consensus framework sequence, such as the framework of human V ⁇ 1 consensus sequence, V ⁇ 3 consensus sequence, V ⁇ 1 consensus sequence, V ⁇ 2 consensus sequence, V ⁇ 3 consensus sequence. Sequences of human germline frameworks are available from various public databases, such as V-base, IMGT, NCBI, or Abysis.
- the 4-1BB antigen binding proteins described herein may comprise a VL framework, wherein the framework may comprise one or more amino acid substitutions, additions, or deletions, while still retaining functional and structural similarity with the germline from which it was derived.
- the VL framework is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to a human germline VL framework sequence.
- the antigen binding protein, antibody, or antigen binding fragment thereof comprises a VL framework comprising 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 amino acid substitutions, additions or deletions relative to the human germline VL framework sequence.
- the VH framework sequence can be derived from a human VH3 germline, a VH1 germline, a VH5 germline, a human VH2 germline, or a VH4 germline.
- Preferred human germline heavy chain frameworks are frameworks derived from VH1, VH2, VH3, or VH4 germlines.
- VH frameworks may be derived from the framework of one of the following germlines: DP54 or IGHV3-7, DP47 or IGHV3-23, DP71 or IGHV4-59, DP75 or IGHV1-2_02, DP10 or IGHV1-69, DP7 or IGHV1-46, DP49 or IGHV3-30, DP51 or IGHV3-48, DP38 or IGHV3-15, DP79 or IGHV4-39, DP78 or IGHV4-30-4, DP73 or IGHV5-51, DP50 or IGHV3-33, DP46 or IGHV3-30-3, DP31 or IGHV3-9.
- the framework sequence may be derived from the framework of a consensus sequence, such as: VH3 germline consensus sequence, VH1 germline consensus sequence, VH5 germline consensus sequence, VH2 germline consensus sequence, or VH4 germline consensus sequence.
- the antigen binding proteins described herein may comprise a VH framework, wherein the framework may comprise one or more amino acid substitutions, additions, or deletions, while still retaining functional and structural similarity with the germline from which it was derived.
- the VH framework is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to a human germline VH framework sequence.
- the antigen binding protein, antibody, or antigen binding fragment thereof comprises a VH framework comprising 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 amino acid substitutions, additions or deletions relative to the human germline VH framework sequence.
- the 4-1BB antigen-binding protein comprises a VH that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 147, 149, 151, 153, 155, 157, 159, 324, 161, 163, 165, 167, 169, 171, 173, 175, 404, 406, 408, or 410.
- the 4-1BB antigen-binding protein comprises a VL that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 148, 150, 152, 154, 156, 158, 160, 325, 162, 164, 166, 168, 170, 172, 174, 176, 405, 407, 409, or 411.
- the 4-1BB antigen binding protein comprises a pair of VH and VL sequences listed under the same clone name in Table B and Table K1.
- the 4-1BB antigen-binding protein comprises a VH but does not comprise a VL, wherein said VH comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, or 385.
- the 4-1BB antigen binding protein comprises a CH1 domain, preferably a human CH1 domain (such as a human IgG1 CH1, a human IgG2 CH1, a human IgG3 CH1, or a human IgG4 CH1).
- a human CH1 domain such as a human IgG1 CH1, a human IgG2 CH1, a human IgG3 CH1, or a human IgG4 CH1.
- Non-limiting examples of human CH1 sequences are provided in the Sequence Tables.
- the CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 261, 265, 269, 422, or 425.
- the 4-1BB antigen binding proteins described herein comprises an Fc domain.
- the Fc domain can be derived from IgA (e.g., IgA1 or lgA2), IgG, IgE, or IgG (e.g., IgG1, lgG2, lgG3, or lgG4).
- the Fc domain comprises wild type sequence of a human Fc domain.
- Non-limiting examples of human Fc domain sequences are provided in the Sequence Table.
- the Fc domain comprises one or more mutations resulting in altered biological activity, such as to improve half-life/stability or to render the antibody more suitable for expression/manufacturability.
- mutations may be introduced into the Fc domain to reduce the effector activity (e.g., WO 2005/063815), and/or to increase the homogeneity during the production of the recombinant protein.
- amino acid residues in the IgG heavy constant domain of an antibody are numbered according the EU index of Edelman et al., 1969, Proc. Natl. Acad. Sci. USA 63(1):78-85 as described in Kabat et al., 1991, referred to herein as the “EU index numbering.”
- the constant domain comprises from residue 118 to 447
- the Fc domain comprises from residue 236 to 447 of the human lgG1 constant domain.
- the Fc domain is the Fc domain of human lgG1 and comprises one or more of the following effector-null mutations: L234A, L235A, and G237A (numbering according to the EU index), often referred as “LALA” mutations.
- L234A L235A
- G237A numbering according to the EU index
- LALA L235E
- the Fc region comprises a Stable Effector Functionless (SEFL) mutation to inhibit or reduce the ability to interact with Fc ⁇ receptors, sequence.
- SEFL mutations are known in the art. See, e.g., Liu et al., J Biol Chem 292: 1876-1883 (2016); and Jacobsen et al., J. Biol.
- the SEFL mutation comprises one or more of the following mutations, numbered according to the EU system: L242C, A287C, R292C, N297G, V302C, L306C, and/or K334C.
- the SEFL mutation comprises N297G.
- the SEFL mutation comprises A287C, N297G, and L306C. In other exemplary aspects, the SEFL mutation comprises R292C, N297G, and V302C (i.e., SEFL2-2).
- the Fc region comprises a YTE mutation.
- the M252Y/S254T/T256E (EU index numbering, referred to “YTE”) triple mutation have been shown to increase IgG half-life in cynomolgus monkeys by an approximate 4-fold increase.
- C-terminal lysine clipping is a common phenomenon occurring during the bioproduction of monoclonal antibodies.
- CpD carboxypeptidase D
- PAM peptidylglycine ⁇ -amidating monooxygenase
- the product is often a mixture of C-terminal processing variants, with heavy chain C-terminus ends at (amidated) proline, glycine, or lysine.
- the terminal lysine may be absent; in some embodiments, the terminal lysine may be present; in some embodiments, the terminal glycine-lysine may be absent; in some embodiments, the terminal glycine-lysine may be present.
- a pharmaceutically suitable composition may comprise a mixture of species that do and do not comprise the terminal lysine and/or glycine-lysine.
- the 4-1BB antigen binding proteins described herein comprise Fc that is derived from an IgG1.
- the Fc comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, 426, 263, 267, or 483.
- the 4-1BB antigen binding proteins described herein comprise an IgG1 heavy chain constant domain.
- the heavy chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 244, 245, 246, 247, 248, 249, 256, 257, 258, 260, 412, 415, 417, 418, 427, 264, 268, or 271.
- the 4-1BB antigen-binding protein described herein comprising a kappa or lambda light chain constant domain.
- the kappa light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428.
- the lambda light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424.
- the 4-1BB antigen-binding protein may be a full-length immunoglobulin, a Fab, or an scFv.
- Exemplary full length 4-1BB binding immunoglobulins are shown as Sequence Table F.
- Exemplary Fab and scFV domains are shown in Sequence Table H and Table K6.
- the scFv may comprises a linker between VH and VL.
- Exemplary linker sequences, such as GS-based linkers, are provided in the Sequence Table G.
- the scFv described herein comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 291.
- the scFv described herein comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 445 (14A5.002 scFv #2).
- the Fab described herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 298, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 300 (6F9.009 Fab
- the Fab described herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 305, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 307 (19G1.016
- the Fab described herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 316, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 317 (6C7.018
- the Fab described herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-227 of SEQ ID NO: 434, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:
- the Fab described herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-227 of SEQ ID NO: 449, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:
- PD-L1 antigen-binding proteins are provided herein.
- Examples of PD-L1 antigen-binding proteins are provided in the Sequence Tables. As discussed in detail above, the CDR sequences provided in the Sequence Tables are based on the Kabat definition. However, other definitions for CDRs may also be used.
- the PD-L1 antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 133, 135, 137, 139, 141, 143, 322, 145, 398, 400, or 402; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 134, 136, 138, 140, 142, 144, 323, 146, 399, 401, or 403.
- the three heavy chain CDRs and three light chain CDRs come from the same clone as shown in Sequence Table C.
- the CDRs are defined according to Kabat, Chothia, AbM, contact, or IMGT [182]
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:133; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:134.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:135; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:136.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:137; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:138.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:139; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:140.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:141; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:142.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:143; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:144.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:322; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:323.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:145; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:146.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:398; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:399.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:400; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO: 401.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:402; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:403.
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:133; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 134. [194] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:1353; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 136.
- the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:137; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 138. [196] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:1393; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 140.
- the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:141; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 142. [198] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:143; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 144.
- the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:322; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 323. [200] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:145; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 146.
- the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:398; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 399.
- the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:400; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 401.
- the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:402; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 403.
- the PD-L1 antigen binding proteins comprise (a) CDR-H1 amino acid sequence set forth in Sequence Table A and Table K2, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table A or Table K2; (b) CDR-H2 amino acid sequence set forth in Sequence Table A and Table K2, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table A or Table K2; (c) a CDR-H3 amino acid sequence set forth in Sequence Table A and Table K2, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least
- the antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
- the PD-L1 antigen binding protein comprises 3, 4, 5, or all 6 of the amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table A and Table K2.
- the antigen binding protein comprises each of the LC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table A and Table K2 and at least 1 or 2 of the HC CDR amino acid sequences designated by the SEQ ID NOs in under the same clone name of Sequence Table A and Table K2.
- the antigen binding protein comprises each of the HC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table A and Table K2, and at least 1 or 2 of the LC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table A and Table K2.
- the antigen binding protein comprises six CDR amino acid sequences listed under the same clone name in Sequence Table A and Table K2, or comprising six CDR amino acid sequences selected from the group consisting of: (1) SEQ ID NOs: 1-6, (2) SEQ ID NOs: 7- 12, (3) SEQ ID NOs: 13-18, (4) SEQ ID NOs: 19-24, (5) SEQ ID NOs: 25-30, (6) SEQ ID NOs: 31-36, and (7) SEQ ID NOs: 37-42.
- Preferred VH and VL framework sequences for PD-L1 antigen-binding proteins disclosed herein are human framework sequences described in detail above.
- the PD-L1 antigen-binding protein comprises a VH that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 133, 135, 137, 139, 141, 143, 322, 145, 398, 400, or 402.
- the PD-L1 antigen-binding protein comprises a VL that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 134, 136, 138, 140, 142, 323144, 146, 399, 401, or 403.
- the PD-L1 antigen binding protein comprises sequences or variants of sequences from a pair of VH and VL sequences listed under the same clone name in Table C and Table K3.
- the PD-L1 antigen-binding proteins disclosed herein may further comprises a CL, a CH1, and/or a Fc region as described in detail above.
- the PD-L1 antigen binding protein comprises a CH1 domain, preferably a human CH1 domain (such as a human IgG1 CH1, a human IgG2 CH1, a human IgG3 CH1, or a human IgG4 CH1).
- a human CH1 domain such as a human IgG1 CH1, a human IgG2 CH1, a human IgG3 CH1, or a human IgG4 CH1.
- Non-limiting examples of human CH1 sequences are provided in the Sequence Tables.
- the CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 261, 265, 269, 422, or 425.
- the PD-L1 antigen binding proteins described herein comprises an Fc domain.
- the Fc domain can be derived from IgA (e.g., IgA1 or lgA2), IgG, IgE, or IgG (e.g., IgG1, lgG2, lgG3, or lgG4).
- the Fc domain comprises wild type sequence of a human Fc domain.
- Non-limiting examples of human Fc domain sequences are provided in the Sequence Table.
- the Fc domain is the Fc domain of human lgG1 and comprises one or more of the following effector-null mutations: L234A, L235A, and G237A (numbering according to the EU index), often referred as “LALA” mutations.
- the Fc region comprises a Stable Effector Functionless (SEFL) mutation to reduce the ability to interact with Fc ⁇ receptors.
- SEFL mutation comprises one or more of the following mutations, numbered according to the EU system: L242C, A287C, R292C, N297G, V302C, L306C, and/or K334C.
- the SEFL mutation comprises N297G.
- the SEFL mutation comprises A287C, N297G, and L306C.
- the SEFL mutation comprises R292C, N297G, and V302C (i.e., SEFL2-2).
- the Fc region comprises a YTE mutation.
- the M252Y/S254T/T256E (EU index numbering, referred to “YTE”) triple mutation have been shown to increase IgG half-life in cynomolgus monkeys by an approximate 4-fold increase.
- the PD-L1 antigen binding proteins described herein comprise Fc that is derived from an IgG1.
- the Fc comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, 426, 263, 267, or 483.
- the PD-L1 antigen binding proteins described herein comprise an IgG1 heavy chain constant domain.
- the heavy chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 244, 245, 246, 247, 248, 249, 256, 257, 258, 260, 412, 415, 417, 418, 427, 264, 268, or 271.
- the PD-L1 antigen-binding protein described herein comprising a kappa or lambda light chain constant domain.
- the kappa light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428.
- the lambda light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424.
- the PD-L1 antigen-binding protein may be a full-length immunoglobulin, a Fab, or an scFv.
- Exemplary full length 4-1BB binding immunoglobulins are shown as Sequence Table E.
- Exemplary Fab and scFV domains are shown in Sequence Table H and Table K6.
- the scFv may comprises a linker between VH and VL.
- Exemplary linker sequences, such as GS-based linkers, are provided in the Sequence Table G.
- the scFv disclosed herein comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 284 (26F6.002.009 scFv).
- the scFv disclosed herein comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 436 (scFv from 56039).
- the Fab disclosed herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-221 SEQ ID NO: 451, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 45
- the antigen binding protein is a bispecific molecule which binds to two different antigens or targets. In various instances, the bispecific molecule binds to both 4-1BB and PD-L1.
- the bispecific molecule binds to both 4-1BB and PD-L1.
- FIGs.2A-2B, FIGs.13A-13K, and FIGs.14A-14D Many different formats of bispecific molecules have been exemplified, some of which are depicted in FIGs.2A-2B, FIGs.13A-13K, and FIGs.14A-14D.
- VHA VH chains and VL chains are generally referred to as VHA, VHB, VLA, or VLB, indicating that they bind to two different antigens, a or b.
- VHA and VLA in general indicates that the variable domain binds to antigen a; VH B and VL B generally indicate that the variable domain binds to antigen b.
- the constant domain may also need to be engineered to ensure correct heavy chain and light chain pairing. Since the engineering in constant domains generally involve limited number of mutations, different versions of constant domains are generally distinguished by CH1 versus CH1’, CH2 vs CH2’, CH3 vs CH3’, or CL vs CL’. Occasionally, complex structures such as the one that is illustrated in FIG.13K may require designations such as CH2 (1) , CH2 (2) , CH2 (3) , CH2 (4) , etc. to illustrate different constant domains that are used.
- some of the bispecific molecule comprises disclosed herein have 4 antigen binding sites, 2 of which bind to 4-1BB protein and 2 of which bind to PD-L1.
- each 4-1BB binding site is identical to the other and/or each PD-L1 binding site is identical to the other.
- the bispecific molecule comprises an IgG moiety and a scFv moiety. As shown in FIG.14A, there are two Fab moieties that bind to one antigen (e.g., PD-L1 or 4-1BB).
- Each Fab moiety comprises two chains: a heavy chain comprising a heavy chain variable domain A (VH A ) and a CH1 domain, and a light chain comprising a light chain variable domain A (VL A ) and a CL domain.
- Each Fab is connected to one chain of Fc (monomeric CH2- monomeric CH3) to form an antibody (IgG). Because this part of the structure is essentially an IgG, there is no new linker between Fab and Fc (Fab and Fc are connected through “hinge” sequence just like a wildtype IgG).
- scFv moieties that bind to the other antigen (e.g., 4-1BB or PD-L1).
- a first linker then connects the C-terminus of one CH3 domain to the N-terminus of one scFv.
- Each scFv comprises a heavy chain variable domain B (VHB) and a light chain variable domain B (VLB); and the VHB and VLB are connected via a second linker.
- VHB heavy chain variable domain B
- VLB light chain variable domain B
- This structure is sometimes referred to as “IgG-scFv” format (one or more scFv moieties attached to an IgG molecule).
- each target (PD-L1, 4-1BB) has two binding domains
- the bispecific molecules exemplified in FIG.14A is often referred herein to as “bivalent” bispecific molecules; nonetheless, it should be noted that it is also acceptable in the art to refer to such kind of molecule as “tetravalent,” as altogether there are four binding domains.
- the configuration depicted in FIG.14A is also referred to IgG- scFv(C2) because two copies of scFv are attached at the C-termini of the IgG molecule.
- the bispecific molecule comprises a scFv moiety that is inserted between the Fab and Fc (hinge) region of an immunoglobulin (sometimes referred to as “Fab-scFv-Fc”).
- the bispecific molecule may comprises: (i) two Fab moieties that bind to one antigen (e.g., 4-1BB or PD-L1), wherein each Fab moiety comprises two chains: a heavy chain comprising a heavy chain variable domain A (VHA) and a CH1 domain, and a light chain comprising a light chain variable domain A (VLA) and a CL domain; (ii) two scFv moieties that bind to another antigen (e.g., PD-L1 or 4-1BB), wherein each scFv comprises a heavy chain variable domain B (VH B ) and a light chain variable domain B (VL B ); and (iii) one Fc region that comprises two chains, each chain comprising a monomeric CH2 domain and a monomeric CH3 domain.
- VHA heavy chain variable domain A
- VLA light chain variable domain A
- CL domain CL domain
- two scFv moieties that bind to another antigen e.g.,
- a first linker connects the C-terminus of one CH1 domain to the N-terminus of one scFv
- a second linker links the VHB and VLB of the scFv moiety
- a third linker connects the C-terminus of one scFv to the N-terminus of one chain of the Fc region.
- the molecule exemplified in FIG.14B is essentially a tetravalent molecule, with two binding moieties for PD-L1 and two binding moieties for 4-1BB, but often called bivalent bispecific molecules.
- FIG.2B Another bispecific configuration is depicted in FIG.2B.
- the bispecific molecule comprises an antibody (IgG) moiety and a Fab moiety.
- IgG antibody
- Fab Fab
- Each Fab moiety comprises two chains: a heavy chain comprising a heavy chain variable domain A (VHA) and a CH1 domain, and a light chain comprising a light chain variable domain A (VLA) and a CL domain.
- VHA heavy chain variable domain A
- VLA light chain variable domain A
- Each Fab is connected to one chain of Fc (monomeric CH2- monomeric CH3) to form an antibody (IgG).
- Fc monomeric CH2- monomeric CH3
- a first linker then connects the C-terminus of one CH3 domain to the N- terminus of one Fab.
- Each Fab comprises a heavy chain variable domain B (VHB) and a CH1’, and a light chain variable domain B (VLB) and a CL’ domain.
- VHB heavy chain variable domain B
- VLB light chain variable domain B
- CL CL’ domain
- CH1 may comprises a mutation to a positively charged residue and CL may comprise a mutation to a negatively charged residue, such that said CH1 and CL form a first charge pair (top part of Fig.2B), and (ii) CH1’ may comprise a mutation to a negatively charged residue and CL’ may comprise a mutation to a positively charged residue, such that said CH1’ and CL’ form a second charge pair (bottom part of Fig. 2B).
- the positively charged residues and negatively charged residues can be introduced at multiple positions in CH1 and CL domain, as long as the residue is present on the surface of the constant domain (such that it can interact with the corresponding residue to form a charge pair), and that the mutation does not generally disrupt the conformation of the constant domain.
- the mutation is introduced at position 183 of the CH domain (EU index numbering), and position 176 of the CL domain (Kabat numbering).
- amino acid residues in the IgG heavy constant domain of an antibody are numbered according the EU index of Edelman et al., 1969, Proc. Natl. Acad. Sci. USA 63(1):78-85 as described in Kabat et al., 1991, referred to herein as the “EU index numbering.”
- the constant domain comprises from residue 118 to 447
- the Fc domain comprises from residue 236 to 447 of the human lgG1 constant domain.
- Comparison between EU numbering and other numbering systems can be found, e.g., at IGMT database.
- the amino acid sequence of human IgG constant region is provided below (SEQ ID NO:246), with position 183 underlined.
- Position 183 corresponds to residue 66 of SEQ ID NO: 265.
- CH1 domain comprises residues 1-98
- hinge region comprises residues 99-110
- CH2 domain comprises residues 111- 223, and CH3 domain comprises residues 224-330.
- ASTKGPSVFP LAPSSKSTSG GTAALGCLVK DYFPEPVTVS WNSGALTSGV HTFPAVLQSS
- CH3 domain comprises residues 224-330.
- amino acid residue of a query sequence "corresponds to" a designated position of a reference sequence (e.g., position 66 of SEQ ID NO:246) when, by aligning the query amino acid sequence with the reference sequence, the position of the residue matches the designated position.
- sequence alignment programs such as ClustalW2, or "BLAST 2 Sequences” using default parameters.
- Amino acid residues of the light chain constant domain are numbered according to Kabat et al., 1991, “Sequences of Proteins of Immunological Interest 5th Ed.”, 1991, NATIONAL INSTITUTES OF HEALTH.
- Kappa light chain also has EU index numbering, and the EU index and Kabat numbering are identical.
- Lambda light chain does not have EU index numbering.
- the amino acid sequences of example human kappa and lambda constant regions are provided below (SEQ ID NOs:237 and 239), with position 176 underlined. Position 176 corresponds to residue 69 of SEQ ID NO: 237, and residue 69 of SEQ ID NO: 239.
- the first residue (“R” in SEQ ID NO:237 and “G” in SEQ ID NO:239) is considered an optional junction residue. This residue sometimes is shown as the first residue in the CL domain, and sometimes as the last residue in the VL domain; both are accepted in the art.
- An amino acid residue of a query sequence "corresponds to" a designated position of a reference sequence (e.g., position 69 of SEQ ID NO:237 or 239) when, by aligning the query amino acid sequence with the reference sequence, the position of the residue matches the designated position.
- Such alignments can be done by hand or by using well-known sequence alignment programs such as ClustalW2, or "BLAST 2 Sequences" using default parameters.
- Positively charged residues include lysine, arginine and histidine.
- lysine (K) is used.
- arginine (R) is used.
- Negatively charged residues include aspartic acid and glutamic acid.
- glutamic acid (E) is used.
- Other positions may also be used to introduce charge pairs.
- positions 123 and 124 of the CL domains and positions 147 and 213 of CH1 domains may be replaced by positively charged residues or negatively charged residues. Further two or more residues in a constant domain may be changed to introduce additional charge pairs.
- the bispecific molecule comprises two VH-only binding domains inserted between the Fab and Fc (hinge) region of an immunoglobulin (FIG.14D), sometime referred to as Fab-VH-Fc or Fab-VH(M2)-Fc.
- a first linker links the C-terminus of the Fab with the N- terminus of the VH, and a second linker linkers the C-terminus of the VH with the N-terminus of the Fc.
- the bispecific molecule comprises two VH-only binding domains, each linked to one C-terminus of an IgG heavy chain (FIG.14C). This configuration is also referred to as IgG-VH or IgG-VH(C2).
- the bispecific molecule comprises one scFv attached to the C- terminus of one of the two IgG heavy chains (FIG.13C).
- the bispecific molecule comprises one scFv inserted between the CH1 domain and CH2 domain (hinge region) of one of the two IgG heavy chains (FIG.13D).
- This configuration involves bivalent binding of one antigen, and monovalent binding of the second antigen, and referred to as Fab-scFv(M1)-Fc.
- the two light chains are identical, whereas the two heavy chains are asymmetric.
- the bispecific molecule is a hetero-IG, comprising one scFv linked to one CH2-CH3 chain, and one Fab linked to the other CH2-CH3 chain (FIG.13G).
- This configuration involves monovalent binding of one antigen through one Fab, and monovalent binding of the second antigen through one scFv.
- the two heavy chains are asymmetric and there is only one light chain.
- the configuration is also referred to as [Fab*scFv] hetero-Fc.
- the bispecific molecule comprises one VH-only binding domain linked to one CH2-CH3 chain, and one Fab linked to the other CH2-CH3 chain (FIG.13H).
- the bispecific molecule comprises one scFv attached to the N- terminus of one of the two IgG heavy chains (FIG.13J).
- This configuration involves bivalent binding of one antigen, and monovalent binding of the second antigen, and is referred to as scFv(N1)-IgG because only one copy of the scFv is attached to the N-terminus of the IgG.
- FIG.13K illustrates another configuration of a bispecific molecule, referred to as ([scFv*Fab] hetero-Fc)-Fc.
- the molecule comprises four chains, and is essentially a [scFv*Fab] hetero-Fc (Fig.13K) structure linked to a second Fc domain.
- the molecule comprises (i) a first heavy chain that comprises a scFv linked to a CH2 (1) -CH3 (1) chain, (ii) a second heavy chain that comprise an IgG heavy chain (with CH2 (2) -CH3 (2) ) linked to a CH2 (3) -CH3 (3) (i.e., VH-CH1-CH2 (2) -CH3 (2) -CH2 (3) -CH3 (3) ), (iii) a light chain, and (iv) a CH2 (4) -CH3 (4) chain.
- FIG.13I illustrates another configuration, essentially a hetero-IgG molecule.
- one heavy chain and one light chain form a Fab that binds to one antigen
- a second heavy chain and a second light chain form a second Fab that binds to a second antigen.
- the molecule is an asymmetric molecule with one 4-1BB binding domain and one PD-L1 binding domain.
- charge pairs may need to be created in CH1, CH1’, CL, and CL’, as disclosed in detail above.
- additional mutations need to be created such as two identical heavy chains do not pair with each other in the Fc region, such that an asymmetric IgG can be created.
- one of the heavy chains uses wild type CH3 sequences, and the other heavy chain comprises mutations in which a positively charged residue in the original wild type is mutated to a negatively charged residue to promote heterologous chain pairing.
- CH3 domain comprises residues 224-330.
- a mutation from K to E or D preferably D
- at position 392 EU index numbering
- a mutation from K to D or E at position 409 (EU index numbering), or at a position that corresponds to residue 292 of SEQ ID NO:246 if EU index is not available, is introduced.
- a mutation from K to E or D (preferably D) at position 439 (EU index numbering), or at a position that corresponds to residue 322 of SEQ ID NO:246 if EU index is not available, is introduced.
- K residues are underline below.
- a “linker” is a molecule or group of molecules that connects two separate entities (e.g., 4-1BB binding protein and PD-
- Protein linkers are particularly preferred, and they may be expressed as a component of the recombinant protein using standard recombinant DNA techniques well-known in the art.
- the linkers may all be the same, or some or all of the linkers may be different from each other.
- the linker is a peptidyl linker.
- the peptidyl linker comprises about 1 to 30 amino acid residues.
- Exemplary linkers include, e.g., a glycine rich peptide; a peptide comprising glycine and serine; a peptide having a sequence [Gly-Gly-Ser]n (SEQ ID NO:436), wherein n is 1, 2, 3, 4, 5, or 6; or a peptide having a sequence [Gly-Gly-Gly-Gly-Ser]n (SEQ ID NO: 389), wherein n is 1, 2, 3, 4, 5, or 6.
- a glycine rich peptide linker comprises a peptide linker, wherein at least 25% of the residues are glycine.
- Glycine rich peptide linkers are well known in the art (e.g., Chichili et al. Protein Sci.2013 February; 22(2): 153-167).
- the peptidyl linker may also be a proline-threonine rich peptide linker.
- FIGs.2A-2B, FIGs.13A-13K, and FIGs.14A-14D when bispecific molecule comprises a scFv moiety, mutations may be introduced to scFv to further improve stability. For example, it has been reported that insufficient interface stability between the heavy and light chains of scFv fragments could be the main cause of irreversible scFv inactivation.
- Fv fragments have been reported to dissociate into heavy-chain variable domains (VH) and light-chain variable domains (VL) with K D values ranging from 10 ⁇ 9 to 10 ⁇ 6 M.
- An interdomain disulfide bond have been used to further improve scFv stability. For example, mutation to Cys at the site of H44 (Kabat numbering), and mutation to Cys at L100 (Kabat numbering) would not significantly affect the domain folding. The two cysteines can then form an intramolecular disulfide bond to further stabilize the scFv.
- cysteine clamp Such mutation is sometimes referred to as “cysteine clamp.”
- C-C Specific examples of scFv comprising cysteine clamps are shown in Sequence Tables C and D, where mutations at H44 (Kabat numbering) and at L100 (Kabat numbering) were used to create disulfide bonds (referred to as “C-C”).
- C-C Specific examples of scFv comprising cysteine clamp in a bispecific format are shown in Sequence Table H and K6.
- a cysteine clamp has been introduced in some of the scFv for IgG-scFv bispecific molecules exemplified herein. In some of the Fab-scFv-Fc bispecific molecules, cysteine clamp is not present.
- cysteine clamp may require evaluation of stability and biologically activities of the scFv.
- the removal of the constant domain (CH1 and C ⁇ or C ⁇ ) lowers the stability of the Fv domain.
- This may require the addition of a linker fusion between the VH and VL domains to avoid molecule dissociation.
- some Fv domains may have an increased probability of being in a dissociated state, exposing their hydrophobic VH and VL interfaces. This could cause increased aggregation, and require additional stability using a disulfide bond or cysteine clamp that covalently links the VH to the VL.
- cysteine clamp tends to create a stabilized product post purification, it could also lead to other issues.
- a poorly positioned cys-clamp can alter the orientation of the VH and VL domains such that it exposes new interfaces or cause a loss in antigen binding due to a new paratope interface. If possible, a scFv domain lacking a cys-clamp with good biophysical properties could be preferential.
- the bispecific molecule disclosed herein comprises a CH1 domain, preferably a human CH1 domain (such as a human IgG1 CH1, a human IgG2 CH1, a human IgG3 CH1, or a human IgG4 CH1), as discussed in detail above.
- the CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 261, 265, 269, 422, or 425.
- the bispecific molecule disclosed herein comprises an Fc domain, as discussed in detail above.
- the Fc domain can be derived from IgA (e.g., IgA1 or lgA2), IgG, IgE, or IgG (e.g., IgG1, lgG2, lgG3, or lgG4).
- the Fc domain comprises wild type sequence of a human Fc domain.
- Non-limiting examples of human Fc domain sequences are provided in the Sequence Table.
- the Fc domain is the Fc domain of human lgG1 and comprises one or more of the following effector-null mutations: L234A, L235A, and G237A (numbering according to the EU index), often referred as “LALA” mutations.
- the Fc region comprises a Stable Effector Functionless (SEFL) mutation to reduce the ability to interact with Fc ⁇ receptors.
- SEFL mutation comprises one or more of the following mutations, numbered according to the EU system: L242C, A287C, R292C, N297G, V302C, L306C, and/or K334C.
- the SEFL mutation comprises N297G. In exemplary aspects, the SEFL mutation comprises A287C, N297G, and L306C. In other exemplary aspects, the SEFL mutation comprises R292C, N297G, and V302C (i.e., SEFL2-2).
- the Fc region comprises a YTE mutation.
- the M252Y/S254T/T256E (EU index numbering, referred to “YTE”) triple mutation have been shown to increase IgG half-life in cynomolgus monkeys by an approximate 4-fold increase.
- the bispecific molecule disclosed herein comprise Fc that is derived from an IgG1.
- the Fc comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, 426, 263, 267, or 483.
- the bispecific molecule disclosed herein comprise an IgG1 heavy chain constant domain.
- the heavy chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 244, 245, 246, 247, 248, 249, 256, 257, 258, 260, 412, 415, 417, 418, 427, 264, 268, or 271.
- the bispecific molecule disclosed herein comprising a kappa or lambda light chain constant domain.
- the kappa light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428.
- the lambda light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:283; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:220.
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:287; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:206.
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:290; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:184.
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:294; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:186.
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:297; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:299; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:304; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:306; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%,
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:311; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:312; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:315; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:312; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:320; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:312; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:443; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:444.
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:434; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:435.
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:449; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:450.
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:451; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:452.
- the bispecific molecule may comprise two copies of each sequence.
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:449; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:450; (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:429; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:430; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 9
- the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:442; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:441.
- the bispecific molecule may comprise two copies of each sequence.
- the antigen binding proteins provided herein bind to their respective targets or antigens in a non- covalent and reversible manner.
- the binding strength of the antigen binding proteins to their targets or antigens may be described in terms of its affinity, a measure of the strength of interaction between the binding site of the antigen binding protein and the target or antigen (e.g., 4-1BB or PD-L1).
- the antigen binding proteins provided herein have high-affinity for their target or antigen (e.g., 4-1BB or PD-L1) and thus will bind a greater amount of the target or antigen (e.g., 4-1BB or PD-L1) in a shorter period of time than low-affinity antigen binding proteins.
- the antigen binding proteins provided herein have low-affinity for 4-1BB or PD-L1 and thus will bind a lesser amount of 4-1BB or PD-L1 in a longer period of time than high-affinity antigen binding proteins.
- the antigen binding protein has an equilibrium association constant, KA, which is at least 10 5 M -1 , at least 10 6 M -1 , at least 10 7 M -1 , at least 10 8 M -1 , at least 10 9 M -1 , at least 10 10 M -1 , at least 10 11 M -1 , at least 10 12 M -1 , at least 10 13 M -1 , or at least 10 14 M -1 .
- KA can be influenced by factors including pH, temperature and buffer composition.
- the binding strength of the antigen binding protein to its target or antigen e.g., 4-1BB or PD-L1
- the binding strength of the antigen binding protein to its target or antigen may be described in terms of its sensitivity.
- KD is the equilibrium dissociation constant, a ratio of k off /k on , between the antigen binding protein and its target or antigen (e.g., 4-1BB or PD-L1).
- K D and KA are inversely related.
- the K D value relates to the concentration of the antigen binding protein (the amount of antigen binding protein needed for a particular experiment) and so the lower the K D value (lower concentration needed) the higher the affinity of the antigen binding protein.
- the binding strength of the antigen binding protein to its target e.g., 4-1BB or PD- L1 may be described in terms of KD.
- the KD value of the antigen binding proteins provided herein is about 10 -1 M or less, about 10 -2 M or less, about 10 -3 M or less, about 10 -4 M or less, about 10 -5 M or less, about 10 -6 M or less, about 10 -7 M or less, about 10 -8 M or less, about 10 -9 M or less, about 10 -10 M or less, about 10 -11 M or less, about 10 -12 M or less, about 10 -13 M or less, about 10 -14 M or less, from about 10 -5 M to about 10 -15 M, from about 10 -6 M to about 10 -15 M, from about 10 -7 M to about 10 -15 M, from about 10 -8 M to about 10 -15 M, from about 10 -9 M to about 10 -15 M, from about 10 -10 M to about 10 -15 M, from about 10 -5 M to about 10 -14 M, from about 10 -6 M to about 10 -14 M, from about 10 -7 M to about 10 -14 M, from about 10 -8 M to
- the K D of the antigen binding proteins provided herein is micromolar, nanomolar, picomolar or femtomolar. In exemplary aspects, the KD of the antigen binding proteins provided herein is within a range of about 10 -4 to 10 -6 M, or 10 -7 to 10 -9 M, or 10 -10 to 10 -12 M, or 10 -13 to 10 -15 M. In exemplary aspects, the antigen binding protein binds to the human 4-1BB or PD-L1 with a K D value that is from about 0.07 nM to about 4 nM.
- the antigen binding protein binds to the human 4-1BB or PD-L1 with a KD of from about 0.01 nM to about 50 nM, from about 0.02 nM to about 50 nM, from about 0.05 nM to about 50 nM, from about 0.05 nM to about 45 nM, from 0.05 nM to about 40 nM, from about 0.05 nM to about 35 nM, from about 0.05 nM to about 30 nM, from about 0.05 nM to about 25 nM, from about 0.05 nM to about 20 nM, from about 0.05 nM to about 15 nM, or from about 0.05 nM to about 10 nM.
- the antigen binding protein binds to the cynomolgus monkey 4-1BB or PD-L1 with a KD that is from about 0.05 nM to about 4 nM. In exemplary aspects, the antigen binding protein binds to the cynomolgus monkey 4-1BB or PD-L1 with a K D of from about 0.01 nM to about 50 nM, from about 0.02 nM to about 50 nM, from about 0.05 nM to about 50 nM, from about 0.05 nM to about 45 nM, from 0.05 nM to about 40 nM, from about 0.05 nM to about 35 nM, from about 0.05 nM to about 30 nM, from about 0.05 nM to about 25 nM, from about 0.05 nM to about 20 nM, from about 0.05 nM to about 15 nM, or from about 0.05 nM to about 10 nM.
- the 4-1BB antigen binding protein binds to human 4-1BB with a K D value of from about 0.05 nM to about 5 nM. In exemplary embodiments, the 4-1BB antigen binding protein binds to cynomolgus monkey 4-1BB with a KD value of from about 0.05 nM to about 5 nM. In exemplary embodiments, the PD-L1 antigen binding protein binds to human PD-L1 with a K D value of from about 0.05 nM to about 5 nM.
- the PD-L1 antigen binding protein binds to cynomolgus monkey PD-L1 with a K D value of from about 0.05 nM to about 5 nM.
- KD values can be determined using methods well established in the art.
- One exemplary method for measuring K D is surface plasmon resonance (SPR), a method well-known in the art (e.g., Nguyen et al. Sensors (Basel).2015 May 5; 15(5):10481-510).
- K D value may be measured by SPR using a biosensor system such as a BIACORE® system.
- BIAcore kinetic analysis comprises analyzing the binding and dissociation of an antigen from chips with immobilized molecules (e.g.
- K D of a protein is by using Bio-Layer Interferometry (e.g., Shah et al. J Vis Exp.2014; (84): 51383). K D value may be measured by Bio-Layer Interferometry using OCTET® technology (Octet QKe system, ForteBio). Alternatively or in addition, a KinExA® (Kinetic Exclusion Assay) assay, available from Sapidyne Instruments (Boise, Id.) can also be used. Any method known in the art for assessing the binding affinity between two binding partners is encompassed herein.
- the KD value is measured by surface plasmon resonance (SPR).
- Antigen e.g., 4- 1BB or PD-L1
- the antigen may be immobilized, e.g., on a solid surface.
- the antigen may be immobilized to a chip, for example by covalent coupling (such as amine coupling).
- the chip may be a CM5 sensor chip.
- This refractive index change is measured in real time (sampling in a kinetic analysis experiment is taken every 0.1 s), and the result plotted as response units (RU) versus time (termed a sensorgram).
- a response will also be generated if there is a difference in the refractive indices of the running and sample buffers.
- This background response must be subtracted from the sensorgram to obtain the actual binding response.
- the background response is recorded by injecting the analyte through a control or reference flow cell, which has no ligand or an irrelevant ligand immobilized to the sensor surface.
- the real-time measurement of association and dissociation of a binding interaction allows for the calculation of association and dissociation rate constants and the corresponding affinity constants.
- One RU represents the binding of 1 pg of protein per square mm. More than 50 pg per square mm of analyte binding is generally needed in practice to generate good reproducible responses.
- Dissociation of the antigen-binding protein from the antigen may be monitored for about 3600 seconds.
- the SPR analysis may be conducted, and the data collected at between about 15°C and about 37°C.
- the SPR analysis may be conducted, and the data collected at between about 25°C and 37°C.
- the SPR analysis may be conducted, and the data collected at about 37°C.
- the SPR analysis may be conducted, and the data collected at 37°C.
- the K D value may be measured by SPR using a BIAcore T200 instrument.
- the SPR rates and affinities may be determined by fitting resulting sensorgram data to a 1:1 model in BIAcore T200 Evaluation software version 1.0.
- the collection rate may be about 1 Hz.
- Another method for determining the K D of an antibody is by using Bio-Layer Interferometry (BLI), typically using OCTET® technology (Octet QKe system, ForteBio).
- BBI Bio-Layer Interferometry
- OCTET® technology Octet QKe system, ForteBio
- biosensor analysis is used.
- one interactant is immobilized on the surface of the biosensor ("ligand,” such as an antigen-binding protein) and the other remains in solution (“analyte”, such as an antigen).
- ligand such as an antigen-binding protein
- an antigen such as an antigen
- the assay begins with an initial baseline or equilibration step using assay buffer.
- a ligand such as an antigen-binding protein
- loading either by direct immobilization or capture-based method.
- biosensors are dipped into buffer solution for a baseline step to assess assay drift and determine loading level of ligand.
- biosensors are dipped into a solution containing the ligand's binding partner, the analyte (association).
- association the binding interaction of the analyte to the immobilized ligand is measured.
- the biosensor is dipped into buffer solution without analyte, and the bound analyte is allowed to come off the ligand (dissociation).
- the series of assay steps is then repeated on new or regenerated biosensors for each analyte being tested. Each binding response is measured and reported in real time on a sensorgram trace.
- the 4-1BB binding protein binds human 4-1BB with a K D value of or less than: about 200nM, about 150nM, about 100nM, about 90nM, about 80nM, about 70nM, about 60nM, about 50nM, about 40nM, about 30nM, about 25nM, about 20nM, about 15nM, about 10nM, about 9nM, about 8nM, about 7nM, about 6nM, about 5nM, about 4nM, about 3nM, about 2nM, about 1 nM, about 900pM, about 800pM, about 700pM, about 600pM, about 500pM, about 400pM, about 300pM, about 250pM, about 200pM, about 150pM, about 100pM, about 50pM, about 40pM, about 30pM
- KD value may be measured by surface plasmon resonance (SPR) (e.g., a Biacore T200 instrument); or it may be measured by bio-layer interferometry (BLI) (e.g., a ForteBio Octet instrument).
- SPR surface plasmon resonance
- BLI bio-layer interferometry
- the PD-L1 binding protein binds human PD-L1 with a K D value of or less than: about 200nM, about 150nM, about 100nM, about 90nM, about 80nM, about 70nM, about 60nM, about 50nM, about 40nM, about 30nM, about 25nM, about 20nM, about 15nM, about 10nM, about 9nM, about 8nM, about 7nM, about 6nM, about 5nM, about 4nM, about 3nM, about 2nM, about 1 nM, about 900pM, about 800pM, about 700pM, about 600pM, about 500pM, about 400pM, about 300pM, about 250pM, about 200pM, about 150pM, about 100pM, about 50pM, about 40pM, about 30pM, about 25pM, about 20pM, about 15pM, about 10pM, about 5pM, or about 1pM.
- KD value may be measured by surface plasmon resonance (SPR) (e.g., a Biacore T200 instrument); or it may be measured by bio-layer interferometry (BLI) (e.g., a ForteBio Octet instrument).
- SPR surface plasmon resonance
- BBI bio-layer interferometry
- the 4-1BB agonists disclosed herein need to be able to induce 4-1BB clustering in addition to specific target engagement. It is possible that lower affinity binders can trigger sufficient agonism but not negative feedback mechanisms.
- Exemplary method to measure binding affinity is also provided in Examples. 2.7 Cross-Reactivity
- the antigen binding protein binds to human 4-1BB.
- a reference amino acid sequence of human 4-1BB is provided herein as SEQ ID NO: 272 (full length precursor) and SEQ ID NO: 274 (extracapsular domain).
- the antigen binding protein binds to cynomolgus monkey (cyno) 4-1BB.
- the amino acid sequence of cyno 4-1BB is provided herein as SEQ ID NO: 275 (precursor).
- the antigen binding protein binds with high affinity to both human 4- 1BB and cyno 4-1BB.
- the antigen-binding proteins of the present disclosure bind to human 4-1BB and cyno 4-1BB but do not cross-react with any other 4-1BB orthologs.
- the antigen binding protein binds with high affinity to both human 4-1BB and cyno 4-1BB and does not bind to any other 4-1BB ortholog, e.g., does not bind to mouse 4-1BB, rat 4-1BB, canine 4-1BB, bovine 4-1BB, and the like.
- the antigen-binding proteins of the present disclosure have a selectivity for human and cyno 4-1BB which is at least 10-fold, 5-fold, 4-fold, 3-fold, 2- fold greater than the selectivity of the antigen-binding protein for any other 4-1BB ortholog.
- the antigen-binding proteins of the present disclosure have a K D for human and cyno 4- 1BB which is at least 10-fold, 5-fold, 4-fold, 3-fold, 2-fold less than the KD of the antigen-binding protein for any other 4-1BB ortholog.
- the antigen binding protein binds to human 4-1BB but does not bind to cyno 4-1BB.
- the antigen binding protein binds to human PD-L1.
- the amino acid sequence of human PD-L1 is provided herein as SEQ ID NO: 277.
- the antigen binding protein binds to cyno PD-L1.
- the amino acid sequence of cyno PD-L1 is provided herein as SEQ ID NO: 279.
- the antigen binding protein binds with high affinity to both human PD-L1and cyno PD- L1.
- the antigen-binding proteins of the present disclosure bind to human PD-L1 and cyno PD-L1 do not cross-react with any other PD-L1 orthologs.
- the antigen binding protein binds with high affinity to both human PD-L1and cyno PD-L1 and does not bind to any other PD-L1 ortholog, e.g., does not bind to mouse PD-L1, rat PD-L1, canine PD-L1, or bovine PD-L1.
- the antigen-binding proteins of the present disclosure have a K D value for human and cyno PD-L1 which is numerically at least 10-fold, 5-fold, 4-fold, 3-fold, 2-fold less than the KD of the antigen-binding protein for any other PD-L1 ortholog.
- the antigen binding protein binds to human PD-L1 but does not bind to cyno PD-L1.
- 2.8 Competition assays [293] Also encompassed herein are antigen-binding proteins that compete with the 4-1BB antigen binding proteins disclosed herein (such as those exemplified in the sequence tables) for the same target, and antigen-binding proteins that compete with the PD-L1 antigen binding proteins disclosed herein (such as those exemplified in the sequence tables) for the same target.
- the antigen-binding protein inhibits a binding interaction between human 4-1BB and a reference antibody, which reference antibody is known to bind to 4-1BB.
- the antigen-binding proteins exhibit an IC 50 of less than about 2000 nM, less than about 1500 nM, less than about 1000 nM, less than about 900 nm, less than about 800 nm, less than about 700 nm, less than about 600 nm, less than about 500 nm, less than about 400 nm, less than about 300 nm, less than about 200 nm, or less than about 100 nm.
- the antigen-binding proteins exhibit an IC 50 of less than about 90 nM, less than about 80 nM, less than about 70 nM, less than about 60 nM, less than about 50 nM, less than about 40 nM, less than about 30 nM, less than about 20 nM, or less than about 10 nM.
- the 4-1BB antigen- binding protein competes with the reference antibody for binding to human 4-1BB and thereby reduce the amount of human 4-1BB bound to the reference antibody as determined by a FACS-based assay in which the fluorescence of a fluorophore-conjugated secondary antibody which binds to the Fc of the reference antibody is measured in the absence or presence of a particular amount of the 4-1BB antigen- binding protein.
- the FACS-based assay is carried out with the reference antibody, fluorophore-conjugated secondary antibody and cells which express 4-1BB.
- the cells are genetically-engineered to overexpress 4-1BB.
- the cells are HEK293T cells transduced with a viral vector to express 4-1BB.
- the cells endogenously express 4- 1BB.
- the cells which endogenously express 4-1BB are pre-determined as low 4-1BB-expressing cells or high 4-1BB-expressing cells.
- the antigen-binding protein inhibits a binding interaction between human PD-L1 and a reference antibody, which reference antibody is known to bind to PD-L1.
- the reference antibody may be a PD-L1 antigen-binding protein disclosed herein, such as those disclosed in the Sequence Tables.
- a PD-L1 antigen-binding protein competes with the reference antibody for binding to human PD-L1 and thereby reduce the amount of human PD-L1 bound to the reference antibody as determined by an in vitro competitive binding assay.
- the PD-L1 antigen-binding protein inhibits the binding interaction between human PD-L1 and the reference antibody and the inhibition is characterized by an IC50.
- the PD-L1 antigen-binding protein exhibits an IC50 of less than about 2500 nM for inhibiting the binding interaction between human PD-L1 and the reference antibody.
- the antigen-binding proteins exhibit an IC 50 of less than about 2000 nM, less than about 1500 nM, less than about 1000 nM, less than about 900 nm, less than about 800 nm, less than about 700 nm, less than about 600 nm, less than about 500 nm, less than about 400 nm, less than about 300 nm, less than about 200 nm, or less than about 100 nm.
- the antigen-binding proteins exhibit an IC 50 of less than about 90 nM, less than about 80 nM, less than about 70 nM, less than about 60 nM, less than about 50 nM, less than about 40 nM, less than about 30 nM, less than about 20 nM, or less than about 10 nM.
- the PD-L1 antigen- binding protein competes with the reference antibody for binding to human PD-L1 and thereby reduce the amount of human PD-L1 bound to the reference antibody as determined by a FACS-based assay in which the fluorescence of a fluorophore-conjugated secondary antibody which binds to the Fc of the reference antibody is measured in the absence or presence of a particular amount of the PD-L1 antigen- binding protein.
- the FACS-based assay is carried out with the reference antibody, fluorophore-conjugated secondary antibody and cells which express PD-L1.
- the cells are genetically-engineered to overexpress PD-L1.
- the cells are HEK293T cells transduced with a viral vector to express PD-L1.
- the cells endogenously express PD-L1.
- the cells which endogenously express PD-L1 are pre-determined as low PD-L1 -expressing cells or high PD-L1-expressing cells.
- Other binding assays e.g., competitive binding assays or competition assays, which test the ability of one antigen-binding molecule (such as an antibody) to compete with a second antigen binding molecule (such as a second antibody) for binding to an antigen, or to an epitope thereof, are known in the art.
- the present disclosure further provides nucleic acids comprising a nucleotide sequence encoding the antigen-binding proteins disclosed herein.
- the nucleic acid may comprise a single nucleic acid molecule, or two or more nucleic acid molecules (for example, a first nucleic acid molecule encoding a heavy chain amino acid sequence and a second nucleic acid molecule encoding a light chain amino acid sequence).
- the nucleic acids of the present disclosure are recombinant.
- the nucleic acid comprises a nucleotide sequence that has at least about 50%, at least about 60%, at least about 70%, at least about 80%, at least about 85%, at least about 90%, or has greater than about 90% (e.g., about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or 100%) sequence identity to any one of SEQ ID NOs: 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465.
- SEQ ID NOs 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310
- nucleic acid sequence that encoding a signal peptide may be added to the 5’ of 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465.
- Recombinant expression of the bispecific molecules, as well as various antigen-binding proteins (or antigen-binding moieties) disclosed herein often require that the molecules be secreted.
- Translocation of a nascent protein from the cytosol into the ER mediated by its signal peptide is an important step in protein secretion. It is understood that the signal peptide is present (and often critical) during the initial synthesis of a nascent protein, but then, signal peptide is cleaved during secretion process. Therefore, while the mature protein no longer has the signal peptide; having the signal peptide coding sequence in the nucleic acid is generally necessary to recombinantly express the protein.
- the present disclosure further provides nucleic acids that are capable of hybridizing to any one of SEQ ID NOs: 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465, or a complimentary sequence of any one of SEQ ID NOs: 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465, under a moderately string
- a “moderately stringent condition” includes prewashing in a solution of 5X SSC, 0.5% SDS, 1.0 mM EDTA (pH 8.0); hybridizing at 50 °C-65 °C, 5X SSC, overnight; followed by washing twice at 65°C for 20 minutes with each of 2X, 0.5X and 0.2X SSC containing 0.1 % SDS.
- a “highly stringent condition” includes, for example, (1) employ low ionic strength and high temperature for washing, for example 0.015M sodium chloride/0.0015M sodium citrate/0.5 % sodium dodecyl sulfate at 50°C; (2) employ during hybridization a denaturing agent, such as formamide, for example, 50% (v/v) formamide with 0.1% bovine serum albumin/0.1% Ficoll/0.1% polyvinylpyrrolidone/50 mM sodium phosphate buffer at pH 6.5 with 750 mM sodium chloride, 75 mM sodium citrate at 42 °C; or (3) employ 50% formamide, 5XSSC (0.75 M NaCI, 0.075 M sodium citrate), 50 mM sodium phosphate (pH 6.8), 0.1% sodium pyrophosphate, 5X Denhardt's solution, sonicated salmon sperm DNA (50 ⁇ g/ml), 0.1% SDS, and 10% dextran sulfate at 42°C, with washes at
- nucleic acids of the present disclosure in some aspects are incorporated into a vector.
- the present disclosure provides vectors comprising any of the presently disclosed nucleic acids.
- the vector is a recombinant expression vector.
- the term "recombinant expression vector” means a genetically-modified oligonucleotide or polynucleotide construct that permits the expression of an mRNA, protein, polypeptide, or peptide by a host cell, when the construct comprises a nucleotide sequence encoding the mRNA, protein, polypeptide, or peptide, and the vector is contacted with the cell under conditions sufficient to have the mRNA, protein, polypeptide, or peptide expressed within the cell.
- the vectors of the present disclosure are not naturally-occurring as a whole. However, parts of the vectors can be naturally-occurring.
- the presently disclosed vectors can comprise any type of nucleotides, including, but not limited to DNA and RNA, which can be single- stranded or double-stranded, synthesized or obtained in part from natural sources, and which can contain natural, non-natural or altered nucleotides.
- the vectors can comprise naturally-occurring or non-naturally- occurring internucleotide linkages, or both types of linkages. In some aspects, the altered nucleotides or non-naturally occurring internucleotide linkages do not hinder the transcription or replication of the vector.
- the vector of the present disclosure can be any suitable vector, and can be used to transform or transfect any suitable host.
- Suitable vectors include those designed for propagation and expansion or for expression or both, such as plasmids and viruses.
- the vector can be selected from the group consisting of the pUC series (Fermentas Life Sciences), the pBluescript series (Stratagene, LaJoIIa, CA), the pET series (Novagen, Madison, WI), the pGEX series (Pharmacia Biotech, Uppsala, Sweden), and the pEX series (Clontech, Palo Alto, CA).
- Bacteriophage vectors such as ⁇ GTIO, ⁇ GTl 1, ⁇ ZapII (Stratagene), ⁇ EMBL4, and ⁇ NMl 149, also can be used.
- the vector is a viral vector, e.g., a retroviral vector.
- the vectors of the present disclosure can be prepared using standard recombinant DNA techniques described in, for example, Sambrook et al., infra, and Ausubel et al., infra.
- Constructs of expression vectors which are circular or linear, can be prepared to contain a replication system functional in a prokaryotic or eukaryotic host cell.
- Replication systems can be derived, e.g., from CoIEl, 2 ⁇ plasmid, ⁇ , SV40, bovine papilloma virus, and the like.
- the vector comprises regulatory sequences, such as transcription and translation initiation and termination codons, which are specific to the type of host (e.g., bacterium, fungus, plant, or animal) into which the vector is to be introduced, as appropriate and taking into consideration whether the vector is DNA- or RNA- based.
- the vector can include one or more marker genes, which allow for selection of transformed or transfected hosts.
- Marker genes include biocide resistance, e.g., resistance to antibiotics, heavy metals, etc., complementation in an auxotrophic host to provide prototrophy, and the like.
- Suitable marker genes for the presently disclosed expression vectors include, for instance, neomycin/G418 resistance genes, hygromycin resistance genes, histidinol resistance genes, tetracycline resistance genes, and ampicillin resistance genes.
- the vector can comprise a native or normative promoter operably linked to the nucleotide sequence encoding the polypeptide (including functional portions and functional variants thereof), or to the nucleotide sequence which is complementary to or which hybridizes to the nucleotide sequence encoding the antigen-binding protein.
- promoters e.g., strong, weak, inducible, tissue- specific and developmental- specific.
- the selection of promoters e.g., strong, weak, inducible, tissue- specific and developmental- specific, is within the ordinary skill of the artisan.
- the combining of a nucleotide sequence with a promoter is also within the skill of the artisan.
- the promoter can be a non-viral promoter or a viral promoter, e.g., a cytomegalovirus (CMV) promoter, an SV40 promoter, an RSV promoter, and a promoter found in the long-terminal repeat of the murine stem cell virus.
- CMV cytomegalovirus
- SV40 promoter SV40 promoter
- RSV promoter a promoter found in the long-terminal repeat of the murine stem cell virus.
- Certain exemplary antigen-binding proteins disclosed herein comprises two different polypeptide chains. To further improve the expression and pairing of the two chains, different promoters of different strengths may be operably linked to coding sequences under different MXS selection conditions, such that the expression levels of the two chains may be adjusted to ensure optimal expression and chain pairing. See, e.g., Example 8 disclosed below.
- host cells comprising a nucleic acid or vector of the present disclosure.
- the term "host cell” refers to any type of cell that can contain the presently disclosed vector and is capable of producing an expression product encoded by the nucleic acid (e.g., mRNA, protein).
- the host cell in some aspects is an adherent cell or a suspended cell, i.e., a cell that grows in suspension.
- the host cell in exemplary aspects is a cultured cell or a primary cell, i.e., isolated directly from an organism, e.g., a human.
- the host cell can be of any cell type, can originate from any type of tissue, and can be of any developmental stage.
- the cell is a eukaryotic cell, including, but not limited to, a yeast cell, filamentous fungi cell, protozoa cell, algae cell, insect cell, or mammalian cell.
- a yeast cell filamentous fungi cell
- protozoa cell algae cell
- insect cell insect cell
- mammalian cell Such host cells are described in the art. See, e.g., Kunert et al., Appl. Microbiol Biotechnol.100: 3451-61 (2016).
- the eukaryotic cells are mammalian cells.
- the mammalian cells are non-human mammalian cells.
- the cells are Chinese Hamster Ovary (CHO) cells and derivatives thereof (e.g., CHO-K1, CHO pro-3, CS9), mouse myeloma cells (e.g., NS0, GS-NS0, Sp2/0), cells engineered to be deficient in dihydrofolatereductase (DHFR) activity (e.g., DUKX-X11, DG44), human embryonic kidney 293 (HEK293) cells or derivatives thereof (e.g., HEK293T, HEK293-EBNA), green African monkey kidney cells (e.g., COS cells, VERO cells), human cervical cancer cells (e.g., HeLa), human bone osteosarcoma epithelial cells U2-OS, adenocarcinomic human alveolar basal epithelial cells A549, human fibrosarcoma cells HT1080, mouse brain tumor cells CAD, embryonic carcinoma cells P19, mouse embryo fibroblast cells NIH 3
- the host cell is CS9 (a CHO cell line).
- the host cell is in some aspects is a prokaryotic cell, e.g., a bacterial cell.
- a population of cells comprising at least one host cell described herein.
- the population of cells in some aspects is a heterogeneous population comprising the host cell comprising vectors described, in addition to at least one other cell, which does not comprise any of the vectors.
- the population of cells is a substantially homogeneous population, in which the population comprises mainly host cells (e.g., consisting essentially of) comprising the vector.
- the population in some aspects is a clonal population of cells, in which all cells of the population are clones of a single host cell comprising a vector, such that all cells of the population comprise the vector.
- the population of cells is a clonal population comprising host cells comprising a vector as described herein.
- Methods of Manufacture [312]
- the antigen-binding proteins disclosed herein may be obtained by methods known in the art. Suitable methods of de novo synthesizing polypeptides are described in, for example, Chan et al., Fmoc Solid Phase Peptide Synthesis, Oxford University Press, Oxford, United Kingdom, 2005; Peptide and Protein Drug Analysis, ed.
- the antigen-binding proteins disclosed herein are recombinantly produced using a nucleic acid encoding the amino acid sequence of the molecule using standard recombinant methods.
- a cell line producing recombinant antigen-binding protein e.g., a protein derived from an antibody and comprises two chains, one based on antibody heavy chain and one based on antibody light chain
- GS glutamine synthetase
- MSX l-methionine sulfoximine
- the composition comprises agents which enhance the chemico-physico features of the antigen-binding molecule, nucleic acid, vector, or host cell, or a combination thereof, e.g., via stabilizing, for example, at certain temperatures (e.g., room temperature), increasing shelf life, reducing degradation, e.g., oxidation protease mediated degradation, increasing half-life of the antigen- binding protein, etc.
- the composition additionally comprises a pharmaceutically acceptable carrier, diluents, or excipient.
- the pharmaceutical composition can comprise any pharmaceutically acceptable ingredients, including, for example, acidifying agents, additives, adsorbents, aerosol propellants, air displacement agents, alkalizing agents, anticaking agents, anticoagulants, antimicrobial preservatives, antioxidants, antiseptics, bases, binders, buffering agents, chelating agents, coating agents, coloring agents, desiccants, detergents, diluents, disinfectants, disintegrants, dispersing agents, dissolution enhancing agents, dyes, emollients, emulsifying agents, emulsion stabilizers, fillers, film forming agents, flavor enhancers, flavoring agents, flow enhancers, gelling agents, granulating agents, humectants, lubricants, mucoadhesives, ointment bases, ointments, oleaginous vehicles, organic bases, pastille bases, pigments, plasticizers, polishing agents, preservatives, sequestering agents, skin penet
- compositions comprising an active agent and one or more pharmaceutically acceptable salts; polyols; surfactants; osmotic balancing agents; tonicity agents; anti-oxidants; antibiotics; antimycotics; bulking agents; lyoprotectants; anti-foaming agents; chelating agents; preservatives; colorants; analgesics; or additional pharmaceutical agents.
- the pharmaceutical composition comprises one or more polyols and/or one or more surfactants, optionally, in addition to one or more excipients, including but not limited to, pharmaceutically acceptable salts; osmotic balancing agents (tonicity agents); anti-oxidants; antibiotics; antimycotics; bulking agents; lyoprotectants; anti- foaming agents; chelating agents; preservatives; colorants; and analgesics.
- pharmaceutically acceptable salts including but not limited to, pharmaceutically acceptable salts; osmotic balancing agents (tonicity agents); anti-oxidants; antibiotics; antimycotics; bulking agents; lyoprotectants; anti- foaming agents; chelating agents; preservatives; colorants; and analgesics.
- the pharmaceutical composition can contain formulation materials for modifying, maintaining or preserving, for example, the pH, osmolarity, viscosity, clarity, color, isotonicity, odor, sterility, stability, rate of dissolution or release, adsorption or penetration of the composition.
- formulation materials for modifying, maintaining or preserving for example, the pH, osmolarity, viscosity, clarity, color, isotonicity, odor, sterility, stability, rate of dissolution or release, adsorption or penetration of the composition.
- suitable formulation materials include, but are not limited to, amino acids (such as glycine, glutamine, asparagine, arginine or lysine); antimicrobials; antioxidants (such as ascorbic acid, sodium sulfite or sodium hydrogen-sulfite); buffers (such as borate, bicarbonate, Tris-HCl, citrates, phosphates or other organic acids); bulking agents (such as mannitol or glycine); chelating agents (such as ethylenediamine tetraacetic acid (EDTA)); complexing agents (such as caffeine, polyvinylpyrrolidone, beta-cyclodextrin or hydroxypropyl-beta-cyclodextrin); fillers; monosaccharides; disaccharides; and other carbohydrates (such as glucose, mannose or dextrins); proteins (such as serum albumin, gelatin or immunoglobulins); coloring, flavoring and diluting agents; emulsifying agents;
- amino acids
- the pharmaceutical compositions can be formulated to achieve a physiologically compatible pH.
- the pH of the pharmaceutical composition can be for example between about 4 or about 5 and about 8.0 or about 4.5 and about 7.5 or about 5.0 to about 7.5.
- the pH of the pharmaceutical composition is between 5.5 and 7.5.
- the method in exemplary embodiments, is a method of treating a subject in need thereof, comprising administering to the subject in need thereof a pharmaceutical composition of the present disclosure in an amount effective to treat the subject.
- an antigen binding protein or pharmaceutical compositions of the present disclosure may be for medical use in a human subject.
- the antigen binding protein or pharmaceutical compositions of the present disclosure may for medical use that does not comprise systemic immune activation or liver toxicity in the human subject.
- the bispecific molecules disclosed herein is a cross-linking dependent agonist for 4-1BB. Therefore, the bispecific molecules induce 4-1BB activation upon PD-L1 binding and subsequent clustering (crosslinking) of 4-1BB.
- compositions of the present disclosure are useful for activating 4-1BB signaling.
- the 4-1BB agonist activity of the compositions provided herein allow such entities to be useful in methods of enhancing T cell activity and enhancing an immune response, and, in particular, an immune response against a tumor or cancer.
- provided herein are methods of enhancing T cell activity in a subject, enhancing T cell survival and effector function, restricting terminal differentiation and loss of replicative potential, promoting T cell longevity, and enhancing cytotoxicity against target (e.g., cancer) cells.
- the methods comprise administering to the subject the pharmaceutical composition of the present disclosure in an effective amount.
- the T cell activity or immune response is directed against a cancer cell or cancer tissue or a tumor cell or tumor.
- the immune response is a humoral immune response.
- the immune response is an innate immune response.
- the immune response which is enhanced is a T-cell mediated immune response.
- the term “enhance” and words stemming therefrom may not be a 100% or complete enhancement or increase. Rather, there are varying degrees of enhancement of which one of ordinary skill in the art recognizes as having a potential benefit or therapeutic effect.
- the pharmaceutical compositions of the present disclosure may enhance, e.g., T cell activity or enhance an immune response, to any amount or level.
- the enhancement provided by the methods of the present disclosure is at least or about a 10% enhancement (e.g., at least or about a 20% enhancement, at least or about a 30% enhancement, at least or about a 40% enhancement, at least or about a 50% enhancement, at least or about a 60% enhancement, at least or about a 70% enhancement, at least or about a 80% enhancement, at least or about a 90% enhancement, at least or about a 95% enhancement, at least or about a 98% enhancement).
- a 10% enhancement e.g., at least or about a 20% enhancement, at least or about a 30% enhancement, at least or about a 40% enhancement, at least or about a 50% enhancement, at least or about a 60% enhancement, at least or about a 70% enhancement, at least or about a 80% enhancement, at least or about a 90% enhancement, at least or about a 95% enhancement, at least or about a 98% enhancement.
- T cell activity assays are described in Bercovici et al., Clin Diagn Lab Immunol. 7(6): 859–864 (2000). Methods of measuring immune responses are described in e.g., Macatangay et al., Clin Vaccine Immunol 17(9): 1452-1459 (2010), and Clay et al., Clin Cancer Res.7(5):1127-35 (2001). [330] Additionally provided herein are methods of treating a subject with cancer and methods of treating a subject with a solid tumor. In exemplary embodiments, the method comprises administering to the subject the pharmaceutical composition of the present disclosure in an amount effective for treating the cancer or the solid tumor in the subject.
- the cancer treatable by the methods disclosed herein can be any cancer, e.g., any malignant growth or tumor caused by abnormal and uncontrolled cell division that may spread to other parts of the body through the lymphatic system or the blood stream.
- the cancer in some aspects is one selected from the group consisting of acute lymphocytic cancer, acute myeloid leukemia, alveolar rhabdomyosarcoma, bone cancer, brain cancer, breast cancer, cancer of the anus, anal canal, or anorectum, cancer of the eye, cancer of the intrahepatic bile duct, cancer of the joints, cancer of the neck, gallbladder, or pleura, cancer of the nose, nasal cavity, or middle ear, cancer of the oral cavity, cancer of the vulva, chronic lymphocytic leukemia, chronic myeloid cancer, colon cancer, esophageal cancer, cervical cancer, gastrointestinal carcinoid tumor, Hodgkin lymphoma, hypopharynx cancer, kidney cancer, larynx cancer, liver
- the cancer is selected from the group consisting of: head and neck, ovarian, cervical, bladder and oesophageal cancers, pancreatic, gastrointestinal cancer, gastric, breast, endometrial and colorectal cancers, hepatocellular carcinoma, glioblastoma, bladder, lung cancer, e.g., non-small cell lung cancer (NSCLC), bronchioloalveolar carcinoma.
- the tumor is non-small cell lung cancer (NSCLC), head and neck cancer, renal cancer, triple negative breast cancer, and gastric cancer.
- the subject has a tumor (e.g., a solid tumor, a hematological malignancy, or a lymphoid malignancy) and the pharmaceutical composition is administered to the subject in an amount effective to treat the tumor in the subject.
- the tumor is non-small cell lung cancer (NSCLC), small cell lung cancer (SCLC), head and neck cancer, renal cancer, breast cancer, melanoma, ovarian cancer, liver cancer, pancreatic cancer, colon cancer, prostate cancer, gastric cancer, lymphoma or leukemia, and the pharmaceutical composition is administered to the subject in an amount effective to treat the tumor in the subject.
- the cancer comprises a solid tumor.
- the term “treat,” as well as words related thereto, do not necessarily imply 100% or complete treatment. Rather, there are varying degrees of treatment of which one of ordinary skill in the art recognizes as having a potential benefit or therapeutic effect.
- the methods of treating cancer of the present disclosure can provide any amount or any level of treatment.
- the treatment provided by the method of the present disclosure can include treatment of one or more conditions or symptoms or signs of the cancer being treated. Also, the treatment provided by the methods of the present disclosure can encompass slowing the progression of the cancer.
- the methods can treat cancer by virtue of enhancing the T cell activity or an immune response against the cancer, reducing tumor or cancer growth, reducing metastasis of tumor cells, increasing cell death of tumor or cancer cells, and the like.
- the methods treat by way of delaying the onset or recurrence of the cancer by 1 day, 2 days, 4 days, 6 days, 8 days, 10 days, 15 days, 30 days, two months, 4 months, 6 months, 1 year, 2 years, 4 years, or more.
- the methods treat by way increasing the survival of the subject.
- the antigen-binding proteins disclosed herein target cancer-associated fibroblasts that is present in tumor stroma.
- Tumor stroma broadly defined as the non-cancer cell and non-immune cell components of tumors, is viewed traditionally as the structural components holding tumor tissues together.
- Tumor stroma is composed of extracellular matrix and specialized connective tissue cells, including fibroblasts and mesenchymal stromal cells. Tumors generally need stroma for nutritional support and the removal of waste products, but stromal content can vary markedly in different types of cancers. For example, many lymphomas have minimal stroma whereas the stroma may make up 90% of other solid tumors.
- the antigen-binding proteins disclosed herein in particular target PD-L1-expressing tumors.
- Fibroblasts are capable of infiltrating tumors and PD-L1-expressing cells can be easily identified by methods well known in the art, such as immunostaining.
- the active agents described herein are administered alone, and in alternative embodiments, are administered in combination with another therapeutic agent, e.g., another active agent of the present disclosure of a different type (e.g., structure). Accordingly, the present disclosure provides a combination comprising a first antigen binding protein which targets PD-L1 and a second antigen binding protein which targets 4-1BB, each of which is an antigen binding protein according to the present disclosures.
- the first antigen binding protein is any one of the PD-L1 binding proteins disclosed herein (such as those in Tables A and K2), and optionally, the second antigen binding protein is any one of the 4-1BB binding proteins disclosed herein (such as those disclosed in Tables B and K1).
- the present disclosure provides the combination as a composition, e.g., pharmaceutical composition, in various instances. Accordingly, the present disclosure provides a composition, e.g., pharmaceutical composition, comprising the first antigen binding protein and the second antigen binding protein. In exemplary instances, the first antigen binding protein and the second antigen binding protein are present in the composition at a ratio of about 1:1. In some aspects, the combination or composition, further comprises a third antigen binding protein.
- the disclosure provides a method of treating cancer, comprising administering to a subject in need thereof a therapeutically effective amount of an antigen-binding molecule disclosed herein, and a therapeutically effective amount of PD-1 inhibitor.
- the bispecific molecule disclosed herein is administered in combination with a PD-1 inhibitor.
- the PD-1 inhibitor and the antigen-binding molecule disclosed herein may be administered concurrently or sequentially.
- the PD-1 inhibitor is an antibody, or antigen-binding fragment thereof.
- the antibody, or antigen-binding portion thereof binds to human PD-1.
- Examples of antibodies that bind to human PD-1 are described, e.g, in US7488802, US7521051, US8008449, US8354509, US8168757, WO2004/004771, WO2004/072286, WO2004/056875, and US2011/0271358.
- Specific anti-human PD-1 antibodies useful for the invention described herein include, for example: K- 3945 (Pembrolizumab, Keytruda ® ; U.S.
- Patent No.8,952,136 M-3475, a humanized IgG4 mAb with the structure described in WHO Drug Information, Vol.27, No.2, pages 161-162 (2013); nivoiumab (BMS- 936558), a human IgG4 mAb with the structure described in WHO Drug Information, Vol.27, No.1, pages 68-69 (2013); the humanized antibodies h409A11, h409A16 and h409A17, which are described in WO2008/156712; AMP-514, which is being developed by Medlmmune; humanized antibody CT-011 (Pidilizumab) a monoclonal antibody being developed by Medivation, and anti-PD-1 antibodies disclosed in WO2015/119923 (the heavy and light chains comprise SEQ ID NO: 21 and SEQ ID NO: 22, respectively).
- Additional PD-1 inhibitors include Cemiplimab (Libtayo, approved for the treatment of cutaneous squamous cell carcinoma (CSCC) or locally advanced CSCC who are not candidates for curative surgery or curative radiation); and Dostarlimab (Jemperli, approved for the treatment of mismatch repair deficient (dMMR) recurrent or advanced endometrial cancer and mismatch repair deficient (dMMR) recurrent or advanced solid tumors).
- the other therapeutic aims to treat or prevent cancer.
- the other therapeutic is a chemotherapeutic agent.
- the other therapeutic is an agent used in radiation therapy for the treatment of cancer.
- the active agents described herein are administered in combination with one or more of platinum coordination compounds, topoisomerase inhibitors, antibiotics, antimitotic alkaloids and difluoronucleosides.
- the subject is a mammal, including, but not limited to, mammals of the order Rodentia, such as mice and hamsters, and mammals of the order Logomorpha, such as rabbits, mammals from the order Carnivora, including Felines (cats) and Canines (dogs), mammals from the order Artiodactyla, including Bovines (cows) and Swines (pigs) or of the order Perssodactyla, including Equines (horses).
- mammals of the order Rodentia such as mice and hamsters
- mammals of the order Logomorpha such as rabbits, mammals from the order Carnivora, including Felines (cats) and Canines (dogs), mammals from the order Artiodactyla, including Bovines (cows) and Swines (pigs) or of
- kits comprising an antigen binding protein (including a bispecific molecule), nucleic acid, vector, or host cell of the present disclosure, or a combination thereof.
- the antigen binding protein, nucleic acid, vector, or host cell is provided in the kit as a unit dose.
- unit dose refers to a discrete amount dispersed in a suitable carrier.
- the unit dose is the amount sufficient to provide a subject with a desired effect, e.g., treatment of cancer.
- the kit comprises several unit doses, e.g., a week or month supply of unit doses, optionally, each of which is individually packaged or otherwise separated from other unit doses.
- the components of the kit/unit dose are packaged with instructions for administration to a patient.
- the kit comprises one or more devices for administration to a patient, e.g., a needle and syringe, and the like.
- the antigen binding protein, nucleic acid, vector, host cell, or a combination thereof is/are pre-packaged in a ready to use form, e.g., a syringe, an intravenous bag, etc.
- the ready to use form is for a single use.
- the kit comprises multiple single use, ready to use forms of the antigen binding protein, nucleic acid, vector, or host cell of the present disclosure.
- the kit further comprises other therapeutic or diagnostic agents or pharmaceutically acceptable carriers (e.g., solvents, buffers, diluents, etc.), including any of those described herein. 6.
- T cell epitope content (which is typically a 9-mer peptide) is one of the factors that contributes to antigenicity.
- MHC or HLA major histocompatibility complex
- T cell epitope immunogenicity This allows the epitopes with higher binding affinities to be more likely to be displayed on the surface of the cell (in the context of MHC molecules) where they are recognized by their corresponding T cell receptor (TCR).
- Antigen-presenting cells typically dendritic cells (DCs) can process antigens and present peptide epitopes in conjugation with human leukocyte antigen (HLA) class II molecules to specific na ⁇ ve helper-T (Th) cells, which results in the activation of Th cells.
- HLA human leukocyte antigen
- Th na ⁇ ve helper-T
- the activated Th cells can stimulate B cells to produce antibodies against antigens.
- anti-drug antibodies ADAs
- detection of T cell responses is frequently employed to monitor the ability of biotherapeutic candidates to elicit immune system activation, i.e., the immunogenicity.
- the most widely used method is DC:T cell proliferation assay.
- the PD-L1-targeting arm of the molecule can bind to the PD-L1 expressed on the surface of DC, and the 4-1BB targeting arm of the molecule can bind to the 4-1BB on the surface of T cells.
- the molecule is designed to activates 4-1BB signal pathway through PD-L1 binding and subsequent clustering of 4-1BB. The activation of the 4-1BB pathway in turn causes T cell activation and proliferation. Therefore, in a typical DC:T assay, such type of molecules will always produce a false positive result, due to the underlying mechanism of action.
- the inventors solved the problem by dividing the bispecific molecules into different domains, for example, by converting the bispecific molecule into a PD-L1 binding molecule (fused with Fc), a 4-1BB binding molecule (fused with Fc), and subjecting these individual domains into a DC:T cell proliferation assay, and comparing the results with an Fc domain alone (control).
- bispecific molecules could also receive false positives produced by a conventional DC:T cell assay, if such bispecific molecule comprises two domains: a first domain that binds to a DC surface antigen, and a second domain that binds to a T cell co-stimulatory molecule.
- the molecule itself can bridge the DC and T cells, causing the activation and proliferation of T cells. Therefore, principles described here are applicable to these molecules, and the two domains should be separately expressed as two different proteins to detect true T-cell epitopes.
- DC surface antigens are known.
- prominent conventional DC (cDC, sometimes also referred to as classical DC or myeloid DC) surface antigens include CD8A, CLEC9A, ITGAE, ITGAX, THBD (CD141), XCR1, CD1C, CD207, ITGAM, NOTCH2, and SIRPA.
- Common surface antigens for plasmacytoid DCs (pDCs) include CLEC4C, LILRB4, NRP1, CCR7, and B220.
- Surface markers reported for monocyte-derived DCs (mo-DCs) include CD14, MRC1 (CD206), CD209, SIRPA, ITGAM (CD11b), and CD1A.
- Langherans surface markers include CD1A, CD207 (Langerin), and ID2.
- AXL+Siglec6+ dendritic cells (AS DC) surface markers include AXL and Siglec6 (CD327).
- common T-cell co-stimulatory molecules include CD28, Inducible Co- Stimulator (ICOS), CTLA4 (Cytotoxic T-Lymphocyte-Associated protein 4), 4-1BB (also known as CD137), OX40 (also known as CD134), CD27, CD30, DR3, Glucocorticoid-Induced TNFR family Related (GITR), or Herpes Virus Entry Mediator (HVEM).
- the disclosure provides a method of assessing immunogenicity of a bispecific molecule, wherein said immunogenicity is attributed to a T-cell epitope, and wherein said bispecific molecule comprises two domains: (1) a first domain that binds to a Dendritic Cell (DC) surface antigen; and (2) a second domain that binds to a T cell co-stimulatory molecule, the method comprises: (a) obtaining a first protein that comprises said first domain but does not comprise said second domain; (b) obtaining a second protein that comprises said second domain but does not comprise said first domain; (c) incubating said first protein and second protein with a cell culture that comprises DCs and T cells; (d) assessing the activation or proliferation of T cells.
- DC Dendritic Cell
- the activation or proliferation of T cells is indicative that said bispecific molecule comprises an immunogenic T cell epitope.
- the first and second proteins can be fused to an Fc domain, as exemplified in the Examples, and the results can be compared with a parallel assay using Fc domain alone as a negative control.
- the following examples are given merely to illustrate the present invention and not in any way to limit its scope.
- EXAMPLE 1 This example describes the generation and characterization of antibodies to human PD-L1 and antibodies to 4-1BB.
- a panel of lead 4-1BB monoclonal antibodies and PD-L1 monoclonal antibodies was identified by carrying out the steps schematically described in Figure 1A.
- Immunization Campaign 1 Mouse Strains [353] Fully human antibodies to human PD-L1 and 4-1BB were generated by immunizing XENOMOUSE® transgenic mice (U.S. Pat. NOs.6,114,598; 6,162,963;6,833,268; 7,049,426; 7,064,244, which are incorporated herein by reference in their entirety; Green et al., 1994, Nature Genetics 7:13-21; Mendez et al., 1997, Nature Genetics 15:146-156; Green and Jakobovits, 1998, J. Ex. Med, 188:483-495; Kellerman and Green, Current Opinion in Biotechnology 13, 593-597, 2002).
- mice were immunized 14-18 times over 6-10 weeks using the Helios Gene Gun system according to the manufacturer’s instructions (BioRad, Hercules, California). Briefly, expression vectors encoding wild type human or rhesus GIPR were coated onto gold beads (BioRad, Hercules, California) and delivered to the epidermis of a shaved mouse or rat abdomen.
- mice were immunized with a human PD-L1 or 4-1BB recombinant protein representing the N-terminal extracellular domain.
- Animals were immunized with recombinant protein mixed with Alum and CpG-ODN 12 times over 4-6 weeks using sub-cutaneous injections.
- the initial boost was 5 ⁇ g (4-1BB) or 10 ⁇ g (PD-L1) and subsequent boosts were 5 ⁇ g for both programs.
- PD- L1 or 4-1BB-specific serum titers were monitored by live-cell FACS analysis on an Accuri flow cytometer (BD Biosciences) using transiently transfected 293T cells.
- luciferase produced was measured by Bio-Glo Luciferase Assay System reagent (Promega), after which the plates were incubated for 20 minutes at room temperature, and luminescence detected with EnVision plate reader (PerkinElmer).
- ESN samples were tested at 100-fold dilution.
- the ESN were quantitated and normalized prior to potency screening.
- ESN samples or purified antibodies were serially titrated in assay media and used to block CHO-PD-L1 stimulation of human PD-1 reporter cells. The number of antibodies showing desired activity during single concentration screening are shown in Table 2. TABLE 2.
- Immature dendritic cells and Pan T cells were purchased from Astarte (Bothel, Washington; currently known as Cellero). Immature dendritic cells were further differentiated into mature dendritic cells using recombinant human IL-4, GM-CSF, and TNF-a (CellXVivoTM Human Monocyte-Derived Dendritic Cell Differentiation Kit R&D Systems #CDK004).
- the hybridoma supernatants and control antibodies were prepared to desired concentration in ICM media, and then were transferred to 96-well high binding plates contained the pre-activated T-cells. Assay plates were incubated in 37 o C, 5%CO2 incubator for 72 hours. On Day 5 of assay, cells were transferred to a v-bottom plate for proliferation readout, using Click-iT Plus EdU Alexa Fluor 488 Flow Cytometry Assay Kit (Molecular Probes, Cat# C10633). Read out the EdU positive cells by flow cytometry analyzer (BD FACSCanto). [363] The hybridoma supernatants were screened at single point of 1ug/mL. The individual value of percentage of EdU positive cells for select antibodies is shown in Table 6. 6.
- Pre-activated primary human T-cells proliferation with crosslinking Frozen purified human T-cells were purchased from Biospecialty. They were added to 6-well TC plates pre-coated with 1 ⁇ g/mL of anti-huCD3 (Clone OKT3, eBioscience, cat#16-0037), incubated overnight at 37 o C, 5%CO2 for pre-activation.
- the hybridoma supernatants and control antibodies were prepared to desired concentration in ICM media, and then were added to 96- well high binding plates (Costar, cat#3369) pre-coated with 200 ng/mL of anti-huCD3 and 1 ⁇ g/mL of goat anti-human IgG Fc antibody (Jackson, cat#31125) for crosslinking of testing samples. Incubated at 37 o C, 5%CO2 for 1 hr, followed by PBS wash to remove the unbound samples. The pre-activated T-cells were collected and washed with ICM.
- T cells were stimulated with 5 ⁇ g/mL of anti-human CD3 clone OKT3 (eBioscience) and 1 ⁇ g/mL of anti-human CD28 (BD Pharmingen) for 72 hours at 37 °C/5% CO2 in a plate that had been pre-coated with 5 ⁇ g/mL anti mouse IgG Fc (Pierce). After 72 hours, cells were removed, washed and suspended at a concentration of 0.5x10 6 cells/mL with 10 ng/mL of IL-2 (Pepro Tech). Cells were then incubated for another 48 to 72 hours at 37 °C/5 % CO2.
- cynomolgus primary cell binding assay For cynomolgus primary cell binding assay, cynomolgus PBMCs (SNBL) were thawed and suspended in a concentration between 4x10 6 and 5x10 6 cells/mL. PBMCs were stimulated with 1 ⁇ g/mL of anti-human CD3 clone SP34 (BD Pharmingen) and 1 ⁇ g/mL of anti-human CD28 (BD Pharmingen) for 72 hours at 37 °C/5 % CO 2 in a plate that had been pre-coated with 5 ⁇ g/mL anti-mouse IgG Fc (Pierce).
- BD Pharmingen anti-human CD3 clone SP34
- BD Pharmingen anti-human CD28
- PBMCs were stimulated with 1 ⁇ g/mL of anti-human CD3 clone SP34 (BD Pharmingen) and 1 ⁇ g/mL of anti-human CD28 (BD Pharmingen) for 72 hours at 37 °C/5 % CO2 in a plate that had been pre-coated with 5 ⁇ g/mL anti-mouse IgG Fc (Pierce). After 72 hours, cells were removed, washed and suspended at a concentration of 0.5x10 6 cells/mL with 20 ng/mL of IL-2 (Pepro Tech). Cells were then incubated for another 72 hours.
- IL-2 Pepro Tech
- HEK293 cells were transiently transfected with the mammalian expression constructs described above. The following day, 15 ⁇ L of exhausted hybridoma media was added to cells expressing each construct. Then, the transfected HEK293 cells were washed and incubated with the nuclear stain Hoechst 33342 and a secondary detection antibody (Goat anti Human IgG Fc Alexa 647 (Jackson ImmunoResearch)).
- the capture surface was first prepared through amine coupling of an anti-human Fc goat pAb (Thermo-Fisher, Cat. #31125) onto a Xantec 200m (CM5 analog) chip.
- the unpurified mAbs at 10 ug/ml, were cycled over the prepared chip to allow capture of approximately 450 RUs.
- a six-point, 2-fold serial dilution of the human PD-L1 (1- 239)E3KMut1His(PL39847) protein from 50 nM was injected from the lowest to highest concentration. Following regeneration of the chip, and recapture of the mAb panel, the same method was repeated using the cyno PD-L1 (1-238)-Flag8xHis protein.
- RNA total or mRNA was purified from wells containing the PD-1 antagonist antibody-producing hybridoma cells using a Qiagen RNeasy mini or the Invitrogen mRNA catcher plus kit. Purified RNA was used to amplify the antibody heavy and light chain variable region (V) genes using cDNA synthesis via reverse transcription, followed by a polymerase chain reaction (RT-PCR). The fully human antibody gamma heavy chain was obtained using the Qiagen One Step Reverse Transcriptase PCR kit (Qiagen).
- This method was used to generate the first strand cDNA from the RNA template and then to amplify the variable region of the gamma heavy chain using multiplex PCR.
- the 5’ gamma chain-specific primer annealed to the signal sequence of the antibody heavy chain, while the 3’ primer annealed to a region of the gamma constant domain.
- the fully human kappa light chain was obtained using the Qiagen One Step Reverse Transcriptase PCR kit (Qiagen). This method was used to generate the first strand cDNA from the RNA template and then to amplify the variable region of the kappa light chain using multiplex PCR.
- the 5’ kappa light chain-specific primer annealed to the signal sequence of the antibody light chain while the 3’ primer annealed to a region of the kappa constant domain.
- the fully human lambda light chain was obtained using the Qiagen One Step Reverse Transcriptase PCR kit (Qiagen). This method was used to generate the first strand cDNA from the RNA template and then to amplify the variable region of the lambda light chain using multiplex PCR.
- the 5’ lambda light chain-specific primer annealed to the signal sequence of light chain while the 3’ primer annealed to a region of the lambda constant domain.
- the amplified cDNA was purified enzymatically using exonuclease I and alkaline phosphatase and the purified PCR product was sequenced directly. Amino acid sequences were deduced from the corresponding nucleic acid sequences bioinformatically. Two additional, independent RT-PCR amplification and sequencing cycles were completed for each hybridoma sample in order to confirm that any mutations observed were not a consequence of the PCR. The derived amino acid sequences were then analyzed to determine the germline sequence origin of the antibodies and to identify deviations from the germline sequence. A comparison of each of the heavy and light chain sequences to their original germline sequences are indicated.
- IC50 transit refers to the concentration of molecule that gives a 50% of max activity.
- Relative potency compares the potency of the clone being tested, relative to the original parental clone, expressed as ratios.
- the degenerate codon NNK was introduced at an isomerization site DGGF (SEQ ID NO: 348) to cover all 20 amino acids. This allowed for selection of molecules based on number of Fabs displayed on the cell surface and amount of binding. Scanning in this region only produced a single weaker variant sequence DGGF (SEQ ID NO: 348) to DGYV (SEQ ID NO: 349). Both sequences were explored as recombinant IgG-scFv molecules with the DGYV (SEQ ID NO: 349) advancing.
- EXAMPLE 3 [378] This example describes the construction of bivalent bispecific molecules to 4-1BB and PD-L1. A schematic of the steps carried out to produce bispecific molecules are shown in Figure 1B.
- Molecular modeling was conducted as a quality control check on designs. The final panel resulted in 12 sequences which had lower liability risk, retention of function, and crosslinking dependent. The 12 mAbs were stressed to evaluate if the remaining predicted liabilities were true liabilities.
- a yeast display method looking for a viable replacement of the DGGF site (SEQ ID NO: 348) using 4 serial NNK codons (SEQ ID NO: 350) resulted in a new variant which mutated residues to DGYV (SEQ ID NO: 349). The theory is that the Y could inhibit the isomerization more since it is less flexible than a G. This substitution also appeared to be more potent.
- Bivalent Bispecific Panel In parallel to the yeast display selection described in Example 2, an optimized bivalent bispecific panel was designed. Briefly, optimized Fv sequences (described in Example 2) were used to create two types of bivalent bispecifics. The first type is the fusion of a single chain Fv domain containing a 3xG4S linker (SEQ ID NO: 346) and an engineering disulfide bond to the c-terminus of the CH3 domain of a full antibody, after removal of the GK sequence. The first type is referred to as an IgG-scFv and a schematic is shown in Figure 2A.
- the second type of bivalent bispecific utilizes the charge pair technology to fuse an additional Fab domain to the c-terminus of the CH3 domain of a full antibody, after removal of the GK sequence.
- the charge pair mutations hinder the formation of incorrect heavy chain/light chain pairing.
- the second type is referred to as an IgG-Fab and a schematic is shown in Figure 2B. Both molecules use an IgG1 SEFL2 Fc domain.
- the anti-4-1BB and anti-PD-L1 Fv domains were positioned in combinations in both orientations in each format. [382]
- These molecules were produced using standard cloning, expression, and purification technologies. Purification was done in 1 of 2 methods depending on the modality.
- the IgG-Fab molecules were purified by standard ProA LFAS, whereas the IgG-scFvs were purified through a triple tandem LFAS system (ProA, In Line Dilution, SEC).
- ProA In Line Dilution, SEC
- the functional activity in a co-culture CHO (+/- huPD-L1) artificial APC with PDMCs assay was the primary selection criterion. Other selection experiments were the yield (>80m/mL), Quality, (MCE/SEC), qualitative viscosity readout, and MSQC.
- the IgG-Fabs were further selected using non-reduced MSQC to evaluate correct chain pairing ratios. Additional selections were conducted based on purification quality, production yields, and functional activity.
- Figure 3 provides a table listing exemplary bispecific molecules constructed and characterized. 2. Additional selection of lead molecules.
- the initial selection criteria for the 14 (7 IgG-Fab/7 IgG-scFv) molecules were expanded to 18 to include the yeast display fix resulting in anti 4-1BB 19G1.016.001 and two new variants anti 4-1BB 14G12.017.001 and anti 4-1BB 6C7.018 along with one new format, referred to as scFab-Fab hetero-IgG.
- a number of bispecific molecules were not selected due to purification, viscosity, aggregation, stability, and heavy;light chain pairing issues.
- the final 10 molecules included 4 IgG-scFv, 1 scFab-Fc- Fab, 5 IgG-Fabs and were subjected to further analysis. Briefly, the conformational integrity of 10 bispecific molecules was validated by LC-MS analysis of non-reduced materials (Agilent 6224 TOF MS). Bispecific molecules with engineered charge pair mutations (CPMs) to drive correct heavy/light chain pairing were also analyzed by LC-MS after proteolytic cleavage below the hinge disulfides with IdeS protease (Genovis) to confirm correct heavy/light chain pairing. The biophysical stability of the molecules at 70 mg/mL total protein concentration were also validated.
- CCMs charge pair mutations
- thermostability of the molecules were assessed by differential scanning calorimetry at 1 mg/ml (Malvern MicroCal System). Tryptophan oxidation, Asparagine deamidation, and Aspartic acid isomerization in CDR regions were also assessed by peptide mapping analysis of both 2 week 400C and 192 klux.hr stressed materials (Thermo Q-Exactive LC-MS System). The results are summarized in the table of Figure 4.
- Figure 5 provides LC-MS spectra of IdeS-protease cleaved samples confirming heavy-light chain pairing in molecules with engineered charge pair mutations.
- CHO/ ⁇ CD3scFv cells were also sequentially transfected with pcDNA3-1-Zeo_HuPD-L1 and under Hygromycin B and zeocin selection, sorted for high PD-L1 expression and then transduced with GFP retroviral expression vector containing Hu CD32a/FcgRII.
- Final aAPCs were sorted for Low ⁇ CD3scFv, high human PD-L1 and CD32a expression.
- aAPC and human primary T cell coculture assay were also sequentially transfected with pcDNA3-1-Zeo_HuPD-L1 and under Hygromycin B and zeocin selection, sorted for high PD-L1 expression and then transduced with GFP retroviral expression vector containing Hu CD32a/FcgRII.
- Final aAPCs were sorted for Low ⁇ CD3scFv, high human PD-L1 and CD32a expression.
- CHO cells expressing anti-humanCD3scFv, human PD-L1 and human Cd32a were used as aAPCs in co-culture assay with human Pan T Cells to test PDL1-41BB bispecific molecules in comparison to anti-PD-L1 and anti-4-1BB mAbs as single agents or in combination.
- PD-L1x4-1BB bispecific molecule, anti-PD-L1, and anti-4-1BB antibodies were serially diluted from 50nM (final in assay) and added in triplicate to co-culture assay with aAPCs and Pan T-cells (isolated from human PBMC using Miltenyi Pan T cell isolation Kit # 130-096-535) at 1:2 ratio (50,000 aAPCs + 100,000 Pan T cells) per well in 96-well tissue culture plate.
- IL-2 released in supernatants were quantitated in MesoScale Discovery IL-2 V-Plex assay (#K151QQD). IL-2 concentrations were analyzed using GraphPad Prism 4 parameter curve fit.
- mice were sourced from Charles River Laboratories (Hollister, CA site) and were provided water and chow ad libitum and maintained in a pathogen-free facility. Mice used in syngeneic tumor experiments were 6-8 weeks of age at the time implant. MC38 cells were inoculated in the right flank at 3 ⁇ 105 cells per implant and allowed to grow for 8 days.
- mice were then randomized by tumor volume (50-100mm3) and treated with anti-PD-L1 blocking mAb (MIH5), anti-4-1BB agnostic mAb (LOB12.3) in mouse IgG1 backbone, anti-4-1BB agonistic mAb (LOB12.3) in mouse IgG2a LALA-PG mutant backbone (incapable of Fc receptor binding), 4-1BB-PD-L1 surrogate bispecific antibody (in mouse IgG2a LALA-PG backbone) or isotype control mAb (mouse IgG2a LALA-PG). All the mAb were dosed i.p.
- chimeric 4-1BB constructs of interest were transiently expressed on 293T cells and antibody binding was detected by flow cytometry.
- 10 million 293T cells were diluted to 1 million cells per mL in 10mL of FreeStyle 293T expression medium and allowed to incubate on a shaker at 37°C for 3 hours.10 ug of DNA construct (see Table 15 for construct details) and 10uL of 293Fectin were each added to 500uL OptiMEM separately and incubated at room temperature for 5 minutes.
- Transfected 293T cells were diluted to 30,000 cells per 15 uL in FACS buffer (1X PBS + 2% FBS), and 15 uL of cell suspension was added to the wells containing supernatant (1:2 dilution of antibodies). The primary antibodies were incubated on cells for 1 hour at 4°C.
- CRD1 also named pre-ligand-binding assembly domain (PLAD)
- PAD pre-ligand-binding assembly domain
- Utomilumab a ligand-blocking antibody, binds 4-1BB between CRDs 3 and 4.
- urelumab binds 4-1BB CRD1, away from the ligand binding site.
- these antibodies demonstrate distinct differences in their agonist activity and reported toxicity. Chin reports that antibodies bind 4-1BB in markedly different modes. In view of these differences, antibodies that bind to different CRDs of 4-1BB were selected and investigate.
- any given antibody binds to the same regions would likely trigger similar conformational changes as ligand binding and therefore be agonistic.
- region binding will also likely block the natural receptor/ligand binding, and the impact of such interference in vivo is unknown. Therefore, identifying an agonistic antibody that binds to a region outside of receptor/ligand interaction sites, such as CRD1 of 4-1BB receptor, could be beneficial to potentially leave the natural receptor/ligand interactions largely intact while still be able to trigger robust receptor signaling. Accordingly, antibodies that bind to CRD1 were identified and characterized ( Figure 9).
- FIG. 9 summarizes epitope regions of certain 4-1BB antibodies disclosed herein.
- EXAMPLE 7 This example describes the generation of UniDab® anti-4-1BB binding domains.
- 18 UniRat® animals were immunized with human 4-1BB protein (Sino Bio 10041-H08H). Animals were immunized with Titermax/Ribi adjuvant and Freunds adjuvant. Pre-harvest bleeds were collected and serum titers against the immunogen (human 4-1BB) was completed. The draining lymph nodes of immunized animals were harvested post-immunization and RNA was extracted from the B-cells in the lymph nodes. Left and right lymph nodes for each animal were analyzed separately.
- cDNA samples containing the full VH region of the heavy chains were sequenced using NGS. Sequences from the immunizations was analyzed and the sequences showing the greatest evidence for antigen-specific affinity maturation were selected high throughput cloning and expression.673 VH genes representing 202 CDR3 families, were identified as top candidates for assembly and expression in HEK 293 cells. These were cloned into a proprietary expression vector containing the human IgG1 constant region with the CH1 domain removed. Subsequently, these UniAb® containing heavy chain only expression vectors were expressed individually in HEK293 cells to produce fully human heavy chain only UniAb®. These UniAb® were functionally evaluated in a high throughput manner for both on and off target cell binding and protein binding.
- the top 25 candidates were identified based on desirable binding to 41BB and not to any off-target cell line. The top candidates were then evaluated by 24 well transient expression after Protein A purification. These molecules were assessed for the following parameters: (1) cell binding dose curves with on-target (CHO-41BB) and off-target CHO-S cells; (2) cross-linker dependent activity; (3) assessing developability through measurements of aggregation propensity by SEC; (4) performing kinetic analysis by Octet to determine KD measurements; and (5) competition group assessment by Octet with benchmark antibodies.
- NF- ⁇ B reporter activity of 4-1BB UniAb® in the presence and absence of cross-linker.4-1BB UniAb® dilutions and cross-linker solution (Goat F(ab)2 anti-human IgG Fc, Abcam, Cat # ab98526) were prepared. On the day of the assay, assay buffer (RPMI-1640 + 1% FBS) was prepared using FBS and RPMI-1640 provided in the reporter assay kit (Promega, Cat# CS196005). NF- ⁇ B-luc2/4-1BB Jurkat cells were thawed in a water bath and removed immediately prior to a complete thaw.
- Cells from each thawed vial were added to 9.5 mL assay buffer.
- the Jurkat cells and CD137 UniAb® /cross-linker or CD137 UniAb® or assay buffer (blank) were added to the plate.
- the plates were incubated at 37 °C, 8% CO 2 for 6 hours.
- the reporter substrate reagent was prepared by reconstituting one vial of Bio-GloTM Luciferase Assay Substrate with the Bio-GloTM Luciferase Assay Buffer. After 6 hours of incubation, plates were removed from the incubator and allowed to equilibrate to room temperature for 10 minutes.
- Bio-GloTM Reagent were at ambient temperature prior to addition to the plates.75 ⁇ L of reconstituted substrate was added to each well and the reaction was allowed to stabilize for 10 minutes at room temperature. Reporter signal was detected using a plate luminometer and fold induction was calculated using the average signal from “blank” wells on each plate as the denominator (background). In this assay, in the absence of cross-linker, there is no activation of CD137 receptor and luminescence signal is absent. In the presence of cross-linker, it induces the crosslinking of CD137 and a dose dependent luminescence. Emax is reported as fold induction over background baseline NF- ⁇ B induction in no antibody (blank) treatment condition (target cells + reporter cells alone).
- FIGs.10A-10B Results from this assay are provided in FIGs.10A-10B.
- FIGs.10A-10B Cell binding dose curves for CD137 UniAb® molecules using primary human and Cyno cells.
- Human PBMCs were activated with 2 ng/mL of recombinant human IL-2 (Rnd Systems, Cat# 202- IL-010), and seeded onto OKT3-coated petri dishes. After 3 days of activation at 37oC, 8% CO2, PBMCs were harvested and enriched for T-cells by negative selection (Miltenyi Biotec, Cat# 130-096-535).
- Human PBMCs were activated with 2 ng/mL of recombinant human IL-2 (Rnd Systems, Cat# 202-IL- 010), and seeded onto OKT3-coated petri dishes. After 3 days of activation at 37oC, 8% CO2, PBMCs were harvested and enriched for T-cells by negative selection (Miltenyi Biotec, Cat# 130-096-535). Increasing concentrations of purified CD137 UniAb® molecules, or heavy chain only isotype control, were incubated with various 4-1BB-expressing or 41-BB-negative cell lines.
- the table summarizes the Emax and EC50 values for cell binding to CHO_Human 4-1BB cells, CHO_Cyno 4-1BB cells, and CHO-S (off-target) cells. All EC50 values are shown in units of ng/mL. ND indicates weak or no detectable binding, so EC50 values could not be calculated.
- Table 17 Activated T-cells CHO-hu4-1BB CHO-cy4-1BB CHO-S Clone IDs Emax EC50 Emax EC50 Emax EC50 Emax EC50 [413] Epitope binning data from CD137 UniAb® molecules.
- the engineered 6 anti-PD-L1 and 14 anti-4-1BB binders were combined in 11 bispecific formats as described in the bispecific formatting section of FIG.12 and illustrated in FIGs.13A-13K and 14A-14D. Resulting molecules were generated by recombinant cloning, expression, and purification.13 molecules in 7 formats with favorable manufacturability, stability, and function were advanced to large scale production for more extensive analysis. Of those, 10 molecules passed large scale production criteria and were advanced to protein characterization.
- EXAMPLE 10 [419] This example describes the biological activities and properties of exemplary bispecific molecules. Table 20 summarizes the structures of these molecules and the sequences can be found in the Sequence Table. “CC” refers to cysteine clamps introduced into the scFv domain.
- Lability of predicted hotspots was measured by peptide mapping after stress at 40 o C for 4 weeks and also after exposure to visible light. Viscosity of molecules concentrated at 70 mg/mL and at 140 mg/mL was measured using cone and plate methods with a target of less than 10 cP at 70 mg/mL. Melting temperature was measured by Differential Scanning Calorimetry (DSC). Clipping after stress of 20 o C and 40 o C incubation for 2 weeks and 4 weeks was measured by reduced capillary electrophoresis in a sodium dodecyl sulfide gel (rCE-SDS).
- DSC Differential Scanning Calorimetry
- Molecules with a change of less a than 2% change in molecular weight species during the stress were ranked more highly than molecules with greater than or equal to a 2% change in molecular weight species.
- Thermal stability after stress of 20 o C and 40 o C incubation for 2 weeks and 4 weeks, after exposure to visible light, and after repeated freeze thaw cycles was measured by size exclusion chromatography (SEC).
- Molecules with a change of less a than 2% change in molecular weight species during the stress were ranked more highly than molecules with greater than or equal to a 2% change in molecular weight species.
- FIG.16A shows the pan T cell response to the bispecific molecules when co-cultured with CHO-K1 artificial APCs that express aCD3scFv and PD-L1. PDL1x41BB bispecific molecules are then titrated in to enhance T cell activity. The aAPCs also express CD32 to enable cross-linking with the 4-1BB 14A5 control mAb.
- FIG.16B shows the ability of the bispecific molecules to restore T cell activation in a mixed lymphocyte reaction (MLR).
- MLR assays use T cells from one individual and MHC class II-expressing cells from a second individual. Interaction of the alloreactive TCR with the MHC class II molecule induces T cell activation resulting in T cell proliferation and IL-2 production. Interaction of PD-1 on the T cells with PD-L1 on the dendritic cells decreases T cell activation and the concomitant IL-2 production.
- FIG.16C shows the activity of the anti-PDL1 arms of PDL1x41BB bispecific molecules tested in the PD1/PDL1 reporter assay developed by Promega.
- Table 22 summarizes the binding affinities of the bispecific molecules to its two targets.
- FIG.17 shows the result of one exemplary bispecific moelcules.
- the bispecific molecules in FIG.17 and anti- PD-L1 mAb required the PD-L1 expression in target cells for their activities, whereas anti-4-1BB mAb activates T cells regardless of PD-L1 expression.
- 4-1BB agonists low affinity (rather than high affinity) binders delivered greater activity through increased clustering.
- molecule 56762 has lower affinity to human 41BB ( ⁇ 1000-fold lower affinity) compared to a reference molecule (data not shown).
- molecule 56762 shows higher activity (about 2-fold higher potency) compared to the reference molecule in vitro.
- EXAMPLE 11 This example describes the serum reactivity (serum immunogenicity) of the bispecific molecules. 11.1. Materials and Methods [429] Meso Scale Discovery (MSD) based immunoassay bridging method: Serum from forty-nine healthy donors (twenty males and twenty females) was diluted 1:30 in 1% BSA in PBS. Diluted samples were then incubated with a mixture consisting of biotin and ruthenium conjugated PDL1x41BB molecules.
- Biotin or ruthenium were covalently attached to the PDL1x41BB bispecific molecules. This process includes an initial buffer- exchange, a conjugation reaction, and a post-conjugation buffer exchange to remove free labels from the conjugated PDL1x41BB molecules. 11.2 Results [431] To evaluate the serum reactivity for the bispecific molecules, forty-nine na ⁇ ve human serum donors were tested in bridging assays. FIG.18A-18B. Fig.18A shows the % ECL inhibition in the MSD bridging assay for the four PDL1x41BB molecules. Candidates 1, 2, and 4 had less than 40% of donors depleting over 40%, passing assay acceptance criteria for acceptable pre-existing reactivity.
- Non-compartmental analysis was performed on the individual serum concentration-nominal time data for each capture reagent method and presented in Table 23. Each molecule had similar PK parameters with both assays, and the PK of all molecules resembled expected characteristics for a human IgG in mouse serum.
- Male and female cynomolgus monkeys were injected intravenously with 4.05 ⁇ M/kg of each PDL1x41BB bispecific molecule. Serum samples were collected and quantitation of PDL1x41BB bispecific molecules was conducted using electrochemiluminescent immunoassays with biotinylated human PDL1 as the capture reagent and a ruthenylated mouse anti-human IgG Fc as the detection reagent.
- Non-human primate PK Dose CL (mL/kg/hr) 44988-3 0.8 mg/kg; 4.05 ⁇ M/kg 0.439 EXAMPLE 13
- This example describes DC:T assays that are used to evaluate T-cell based immunogenicity (T cell epitope) of a molecule.
- PBMC were thawed and CD14 + cells were isolated by positive magnetic bead selection according to the manufacturer’s protocol. Isolated CD4 + monocytes were then differentiated into immature dendritic cells (iDC) for 5 days with GM-CSF and IL-4.
- iDC immature dendritic cells
- iDC were washed on the Curiox and treated with media (negative control), a molecule with low clinical immunogenicity (low risk benchmark) at 0.3 ⁇ M, KLH (positive control) at 50 ⁇ g/mL, or test molecules at 0.3 ⁇ M for 3 hours.
- iDC were washed on the Curiox and matured for 2 days with TNF ⁇ and IL-1 ⁇ .
- mature DC mDC
- a stimulation index was calculated for each donor by taking the mean percent EdU + of the CD3 + CD4 + population of (6-8) replicate wells treated with control or test molecules and dividing by the mean percent EdU + of the CD3 + CD4 + population of (6-8) replicates treated with media alone.
- SI value >2 is considered a positive response, meaning the donor demonstrated an immune response to the test or control molecule. This SI metric is a well-accepted industry standard for establishing responses.
- the Fc domain alone served as a baseline for the experiment since a human IgG1 Fc domain is not expected to elicit a robust T-dependent antibody response, given the abundance of this domain in human serum. Furthermore, the Fc domain of the bispecific molecule disclosed herein has been incorporated into other molecules that have been tested in clinical studies with acceptable immunogenicity. By comparing the anti-PD-L1 mAb and the anti-41BB-Fc fusion to the Fc alone, the contribution of the anti-PD-L1 fragment antigen-binding (Fab) and the anti-41BB domain to sequence- based immunogenic risk could be assessed. A total of 30 donors were tested, and 5 were excluded due to a lack of antigen specificity.
- Fab fragment antigen-binding
- the 8xHistidine tag at C-terminus of 4-1BB was removed via proteolytic cleavage. 1.8 mg purified untagged 4-1BB was mixed with 3 mg 380984 to form a complex. The complex was obtained by removing excess 380984 using size-exclusion chromatography and followed by SDS-PAGE to confirm the purified complex. The purified 4-1BB with 380984 complex was concentrated to 9 mg/ml for crystallization setup. [442] Crystallization. Crystals of the 4-1BB with 380984 complex were obtained in the following condition: 0.1M MES pH 5.6, 1.5M Ammonium sulfate, and the sitting drop vapor diffusion technique by mixing 300 nL protein with 300 nL crystallization buffer.
- the structure was solved by molecular replacement using the published 4-1BB structure (PDB:6mhr) and a homology model of the 380984 as templates.
- the molecule replacement solution suggested that one 4-1BB monomer and one UniDab_VH molecule were packed into the asymmetric unit with high confidence.
- the final refinement Rwork/Rfree factors are 0.225/0.255.
- the structure is in good geometry with RMSD of bonds of 0.008 ⁇ and RMSD angles of 1.011o; the Ramachandran plot shows 95.80% residues are in favorable backbone dihedral angles, and 4.20% residues are allowed.
- Epitope and Paratope analysis [445]
- the asymmetric unit contains one 4-1BB monomer and one 380984.
- the overall crystal structure of human 4-1BB in complex with 380984 is shown in FIG.
- the human 4-1BB folds into a V-shape molecule with four CRDs in a linear arrangement and a significant bend within CRD3.
- the epitopes-paratopes interface partially overlaps with the 4-1BBL ligand binding residues (FIG.20A).
- Table 25A shows the 380984 residues which make contact with 4-1BB. As shown in the table, Kabat CDR-3 of the V H residues, S103, Y108, T110, S111, F112, and Y114 contact 4-1BB residues K69, V71, Q104, M101, and L112; Kabat framework residue L45 and W47 contact 4-1BB residues P90 and Q104. Table 25A.
- Trp47 While the majority of the paratope residues are in CDR-H3, Leu45 and Trp47 are characterize as framework residues under Kabat, AbM, Chothia, and IMGT definitions. Contact definition characterizes Trp47 as part of CDR-H2. Table 25B. Ranking of the 4-1BB residue importance in the binding interface 4-1BB Importance Notes residues # CYS102 Primar H-bondin to 380984 (backbone) VDW with 380984 (side chain); hi h Table 25C.
- Ligand 4-1BBL induced receptor 4-1BB clustering is prerequisite for the downstream signaling.
- a published 4-1BB-4-1BBL complex structure has shown that three receptor 4-1BB parallelly bind to a bell- shaped trimer ligand 4-1BBL(FIG.3A).
- the center axis of the receptor-ligand complex should be vertically oriented towards the cell membrane, like other TNF receptor-ligand complexes.
- the 380984 binds along the side of 4-1BB at CRDs 2 and 3, with its N-terminus, where Fc of the antibody connects, pointing away from the cell membrane (FIG.3B).
- the bulk of the antibody is also away from the receptor and the cell membrane, which increase the capability of the Fc ⁇ R engagement and clustering.
- the Fabs are orientated parallel to the membrane, where engagement of Fc ⁇ R may be more restricted (FIG.3C and D). Table 25D.
- the center axis of the receptor-ligand complex should be vertically oriented towards the cell membrane, like other TNF receptor-ligand complexes.
- the 380984 binds along the side of 4-1BB at CRDs 2 and 3, with its N-terminus, where Fc of the molecule connects, pointing away from the cell membrane (FIG.21E).
- Fc Fc of the molecule connects
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Immunology (AREA)
- Organic Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- Medicinal Chemistry (AREA)
- General Health & Medical Sciences (AREA)
- Biophysics (AREA)
- Biochemistry (AREA)
- Genetics & Genomics (AREA)
- Molecular Biology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Pharmacology & Pharmacy (AREA)
- Animal Behavior & Ethology (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
- General Chemical & Material Sciences (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Peptides Or Proteins (AREA)
Abstract
Bispecific molecules comprising a 4-1BB binding protein and a PD-L1 binding protein are provided herein. Related monospecific molecules binding to either 4-1BB or PD-L1 are also provided herein. Methods of treatment comprising administering a pharmaceutical composition comprising a bispecific or monospecific molecule as described herein are further provided.
Description
MOLECULES FOR TREATMENT OF CANCER CROSS REFERENCE TO RELATED APPLICATIONS [1] This application claims the benefit of U.S. Provisional Application Serial No.63/580,194, filed September 01, 2023, and U.S. Provisional Application Serial No.63/606,175, filed December 05, 2023. The foregoing applications are incorporated by reference in its entirety. INCORPORATION BY REFERENCE OF MATERIAL SUBMITTED ELECTRONICALLY [2] Incorporated by reference in its entirety is a computer-readable nucleotide/amino acid sequence listing submitted concurrently herewith and identified as follows: 10669-US02-PRI_SeqListing; created November 30, 2023. FIELD OF THE INVENTION [3] The present invention relates to antigen binding proteins, for example, bispecific molecules, for the treatment of cancer. BACKGROUND [4] The PD-1/PD-L1 axis is involved in the suppression of T cell immune responses in cancer. Antagonists of this pathway have been clinically validated across a number of solid tumor indications. Nivolumab and pembrolizumab are two such inhibitors that target the PD-1 pathway, and each has been approved by the U.S. Food and Drug Administration (FDA) for the treatment of metastatic melanoma. Recently, researchers have tested the paradigm of checkpoint inhibition in the setting of other tumor types. While some advances have been made, checkpoint inhibition therapy still remains in the shadows of other cancer treatment options. [5] Studies of checkpoint inhibitors in combination with other agents are underway or recently have been completed. The combination of nivolumab and ipilimumab, a CTLA-4 receptor blocking antibody, for example, was tested in a Phase III clinical trial on patients with unresectable stage III or IV melanoma. In this study, the percentage of patients achieving a complete response was the highest among those that received the combination of nivolumab and ipilimumab, beating the outcome exhibited by those in the group receiving either drug alone. However, the response to immunotherapies that block CTLA-4 and PD- 1 checkpoint receptors is not universal, and multiple mechanisms by which tumors evade response have been identified. As an approach to enhance the overall efficacy and to limit tumor resistance, combination therapies targeting multiple pathways represent a rational next step. [6] 4-1BB, which is also known as CD137 or TNFRSF9, is a member of the TNF receptor superfamily. 4-1BB was first identified as a molecule whose expression is induced by T-cell activation (Kwon Y.H. and
Weissman S.M. (1989), Proc. Natl. Acad. Sci. USA 86, 1963-1967). Subsequent studies demonstrated expression of 4-1BB in T- and B-lymphocytes, NK-cells, NKT-cells, monocytes, neutrophils, and dendritic cells as well as cells of non- hematopoietic origin such as endothelial and smooth muscle cells. Expression of 4-1BB in different cell types is mostly inducible and driven by various stimulatory signals, such as T-cell receptor (TCR) or B-cell receptor triggering, as well as signaling induced through co- stimulatory molecules or receptors of pro-inflammatory cytokines. [7] 4-1BB signaling is known to stimulate IFNγ secretion and proliferation of NK cells, as well as to promote DC activation as indicated by their increased survival and capacity to secret cytokines and upregulate co- stimulatory molecules. However, 4-1BB is best characterized as a co-stimulatory molecule which modulates TCR- induced activation in both the CD4+ and CD8+ subsets of T-cells. In combination with TCR triggering, agonistic 4-1BB-specific antibodies enhance proliferation of T-cells, stimulate lymphokine secretion and decrease sensitivity of T-lymphocytes to activation-induced cells death (Snell L.M. et al. (2011) Immunol. Rev.244, 197-217). In line with these co-stimulatory effects of 4-1BB antibodies on T-cells in vitro, their administration to tumor bearing mice leads to potent anti-tumor effects in many experimental tumor models (Melero I. et al. (1997), Nat. Med.3, 682-685; Narazaki H. et al. (2010), Blood 115, 1941-1948). In vivo depletion experiments demonstrated that CD8+ T-cells play the most critical role in anti-tumoral effect of 4-1BB-specific antibodies. However, depending on the tumor model or combination therapy, which includes anti-4-1BB, contributions of other types of cells such as DCs, NK-cells or CD4+ T-cells have been reported (MuriUo O. et al. (2009), Eur. J. Immunol.39, 2424- 2436; Stagg J. et al. (2011), Proc. Natl. Acad. Sci. USA 108, 7142-7147). [8] In addition to their direct effects on different lymphocyte subsets, 4- 1BB agonists can also induce infiltration and retention of activated T-cells in the tumor through 4-1BB -mediated upregulation of intercellular adhesion molecule 1 (ICAM1) and vascular cell adhesion molecule 1 (VCAM1) on tumor vascular endothelium.4-1BB triggering may also reverse the state of T-cell anergy induced by exposure to soluble antigen that may contribute to disruption of immunological tolerance in the tumor micro- environment or during chronic infections. [9] It has been reported that systemic administration of 4-1BB-specific agonistic antibodies induces expansion of CD8+ T-cells associated with liver toxicity (Dubrot J. et al. (2010), Cancer Immunol. Immunother.59, 1223-1233). In human clinical trials (ClinicalTrials.gov, NCT00309023), 4-1BB agonistic antibodies (BMS-663513) administered once every three weeks for 12 weeks induced stabilization of the disease in patients with melanoma, ovarian or renal cell carcinoma. However, the same antibody given in another trial (NCT00612664) caused grade 4 hepatitis leading to termination of the trial (Simeone E. and Ascierto P.A. (2012), J. Immunotoxicology 9, 241-247). [10] Thus, there is a need for new generation agonists that effectively engage 4-1BB while avoiding undesired side effects. There also is a need for therapies which target multiple pathways including the
PD-1 pathway in order to enhance the overall efficacy and to limit the tumor resistance of checkpoint inhibitor therapy. SUMMARY [11] Based on the disclosure provided herein, those skilled in the art will recognize, or be able to ascertain using no more than routine experimentation, many equivalents to the specific embodiments of the invention described herein. Such equivalents are intended to be encompassed by the following embodiments (E). E1. A 4-1BB antigen-binding protein comprising a heavy chain variable domain (VH) and a light chain variable domain (VL), wherein said protein binds to the Cysteine-rich pseudo repeat 1 (CRD1) of human 4-1BB (corresponding to residues 24-45 of SEQ ID NO: 272), and is a crosslinking-dependent agonist. E2. The 4-1BB antigen-binding protein of E1, comprising (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 151, 159, 324 or 167; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 152, 160, 325, or 168. E3. The 4-1BB antigen-binding protein of E1, comprising: (i) a CDR-H1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 55, 79 or 103; (ii) a CDR-H2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 56, 80, or 104; (iii) a CDR-H3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 57, 81, or 105; (iv) a CDR-L1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 58, 82, or 106; (v) a CDR-L2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 59, 83, or 107; and (vi) a CDR-L3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 60, 84, or 108. E4. The 4-1BB antigen-binding protein of any one of E1-E3, comprising:
(1) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.55-60, respectively (14A5); (2) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.79-84, respectively (14A5.002); or (3) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.103-108, respectively (16D1.001). E5. A 4-1BB antigen-binding protein comprising a heavy chain variable domain (VH) and a light chain variable domain (VL), wherein said protein binds to the Cysteine-rich pseudo repeat 2 (CRD2) of human 4-1BB (corresponding to residues 47-86 of SEQ ID NO: 272), and is a crosslinking-dependent agonist. E6. The 4-1BB antigen-binding protein of E5, comprising (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 149, 161, 157, 169, 147, 163, 165, 153, 171, 175, 155, 404, 406, 408, or 410; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 150, 162, 158, 170, 148, 164, 166, 154, 172, 176, 156, 405, 407, 409, or 411. E7. The 4-1BB antigen-binding protein of E5, comprising: (i) a CDR-H1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 49, 85, 73, 109, 43, 91, 97, 61, 115, 127, 67, 386, or 392; (ii) a CDR-H2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 50, 86, 74, 110, 44, 92, 98, 62, 116, 128, 68, 387, or 393; (iii) a CDR-H3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 51, 87, 75, 111, 45, 93, 99, 63, 117, 129, 69, 388, or 394; (iv) a CDR-L1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 52, 88, 76, 112, 46, 94, 100, 64, 118, 130, 70, 389, or 395; (v) a CDR-L2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 53, 89, 77, 113, 47, 95, 101, 65, 119, 131, 71, 390, or 396; and (vi) a CDR-L3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 54, 90, 78, 114, 48, 96, 102, 66, 120, 132, 72, 391, or 397. E8. The 4-1BB antigen-binding protein of any one of E5-E7, comprising: (1) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.49-54, respectively (6F9);
(2) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.85-90, respectively (6F9.009); (3) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.73-78, respectively (14G12.017); (4) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.109-114, respectively (17H1.009); (5) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.43-48, respectively (6C7); (6) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.91-96, respectively (6C7.018); (7) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.97-102, respectively (4E9.020); (8) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.61-66, respectively (19G1); (9) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.115-120, respectively (19G1.016); (10) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.127-132, respectively (19G1.016.001); (11) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.67-72, respectively (15A12.012); (12) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.386-391, respectively (56039); or (13) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.392-397, respectively (56040). E9. A 4-1BB antigen-binding protein comprising a heavy chain variable domain and a light chain variable domain, wherein said protein binds to the Cysteine-rich pseudo repeat 3 (CRD3) of human 4- 1BB (corresponding to residues 87-118 of SEQ ID NO: 272), and is a crosslinking-dependent agonist. E10. The 4-1BB antigen-binding protein of E9, comprising (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 173; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 174. E11. The 4-1BB antigen-binding protein of E9, comprising:
(i) a CDR-H1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 121; (ii) a CDR-H2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 122; (iii) a CDR-H3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 123; (iv) a CDR-L1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 124; (v) a CDR-L2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 125; and (vi) a CDR-L3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 126. E12. The 4-1BB antigen-binding protein of any one of E9-E11, comprising: a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs: 121-126, respectively (15A6.011). E13. A 4-1BB antigen-binding protein comprising a heavy chain variable domain and a light chain variable domain, wherein said VH comprises: (i) a CDR-H1 comprising X1 Y X2 X3 X4 (SEQ ID NO: 474), wherein: X1 = R, S, T, N, or G; X2 = Y, A, S, or G; X3 = W, M, or L; and X4 = S, N, or H; (ii) a CDR-H2 comprising X1 I X2 X3 X4 X5 X6 X7 X8 Y X9 X10 X11 X12 X13 X14 K X15 (SEQ ID NO:475), wherein X1 = Y, R, A, V, or L; X2 = D, Y, S, W, or G; X3 = G, S, Y, I, or absent; X4 = D, Y, T, or S; X5 = S, G, A, or T; X6 = G or S; X7 = N, H, S, T, Q, or A; X8 = T, I, E, or K; X9 = N or Y; X10 = Y or D; X11 = N, A, or V; X12 = P, D, or E; X13 = S or P; X14 = L or V; and X15 = S or G; and (iii) a CDR-H3 comprising X1 X2 X3 X4 X5 X6 X7 X8 Y X9 X10 X11 X12 X13 X14 X15 X16 X17 D X18 (SEQ ID NO: 476), wherein X1 = G, T, E, or D; X2 = V, D, T, G, L, A, E, or H; X3 = A, Y, G, I, N, V, I, or L; X4 = A, Y, F, T, D, L, or V; X5 = G, Y, or A; X6 = R, F, D, E, T, G, N, or Y; X7 = G or absent; X8 = K or absent; X9 = T or absent; X10 = T, A, P, or absent; X11 = S, L, T, A, S, Y, L, or absent; X12 = G, L, Y, or absent; X13 = Y, H, or absent; X14 = Y, H, or absent; X15 = D, Y, F, R, or absent; X16 = E, W, G, A, Y, or absent; X17 = I, M, F, or L; and X18 = P, V, Y, or F; and
wherein said VL comprises: (i) a CDR-L1 comprising X1 X2 X3 X4 X5 X6 X7 X8 Y X9 X10 X11 X12 X13 X14 X15 X16 (SEQ ID NO: 477), wherein X1 = G, S, R, or K; X2 = G, A, or S; X3 = N, S, D, or G; X4 = N, Q, K, or A; X5 = S or absent; X6 = N, L, or absent; X7 = L or absent; X8 = H, R, or absent; X9 = T, S, or absent; X10 = I, S, L, G, or D; X11 = G, I, or P; X12 = S, K, D, or G; X13 = K, S, R, N, or T; X14 = N, A, or Y; X15 = V, L, or A; and X16 = H, N, C, Y, F, A, or S; (ii) a CDR-L2 comprising X1 X2 X3 X4 X5 X6 S (SEQ ID NO:478), wherein X1 = D, S, A, Q, or E; X2 = D, N, A, or L; X3 = S, H, or A; X4 = D, Q, S, R, K, N, V, or T; X5 = R, or L; and X6 = P, Q, or F; and (iii) a CDR-L3 comprising X1 X2 X3 X4 X5 X6 X7 X8 Y X9 X10 X11 (SEQ ID NO: 479), wherein X1 = Q, E, L, Y, or M; X2 = V, A, Q, T, S, or K; X3 = W, R, T, Y, or S; X4 = D, F, Y, I, or N; X5 = S, D, E, H, or Q; X6 = S, I, A, L, V, T, or R; X7 = S, L, P, A, or absent; X8 = D or N; X9 = H or G; X10 = V, P, F, L, G, or W ; and X11 = V, T, M, or A. E14. A 4-1BB antigen-binding protein comprising a heavy chain variable domain and a light chain variable domain, wherein said VH is a VH3 that comprises: (i) a CDR-H1 comprising X1 Y X2 X3 X4 (SEQ ID NO:484), wherein X1 = S, T, N, or G, X2 = A, S, or G; X3 = M or L; and X4 = N, or H; or a CDR-H1 comprising X1 X2 A M S (SEQ IDNO:485), wherein X1 = H, Y, or S; and X2 = Y or P; (ii) a CDR-H2 comprising X1 I X2 X3 X4 X5 X6 X7 X8 Y X9 X10 X11 S V K G (SEQ ID NO:486), X1 = X2 = = X4 = D = =
H; X3 = I, N, G, D, A, I, V, or L; X4 = F, I, D, L, T, Y or A; X5 = G, Y, L, or A; X6 = V, F, G, T, E, Y, or absent; X7 = V, Y, A, G, T, L, or absent; X8 = K, Y, L, or absent; X9 = T, Y, or absent; X10 = N, P, or absent; X11 = Y or absent; X12 = Y or absent; X13 = Y, H, or absent; X14 = Y, H, A, I, or absent; X15 = I, F, Y, E, R, or absent; X16 = Y, G, A, N, or absent; X17 = M, F, or L; X18 = D or Q; and X19 = Y, V, H, or F; or a CDR-H3 comprising X1 X2 X3 X4 G X5 X6 X7 Y Y Y G M D V (SEQ ID NO:488), wherein X1 = G or S; X2 = Y or G; X3 = S or Y; X4 = Y or V; X5 = Y or E; X6 = Y or L; and X7 = Y or L. E15. A 4-1BB antigen-binding protein comprising a heavy chain variable domain and a light chain variable domain, wherein said VH is a VH4 that comprises: (i) a CDR-H1 comprising X1 Y Y W S (SEQ IDNO:489), wherein X1 = S or R; (ii) a CDR-H2 comprising X1 I X2 X3 S G X4 T N Y N P X5 L K S (SEQ ID NO: 490), wherein X1 = Y or R; X2 = Y or D; X3 = Y, D or T; X4 = S, N, Q, or H; and X5 = S or P; and
(iii) a CDR-H3 comprising X1 X2 X3 X4 X5 X6 X7 X8 Y X9 X10 X11 X12 X13 X14 D X15 (SEQ ID NO:491), wherein X1 = G, R, T, L, or E; X2 = I, V, D, or T; X3 = A, G, or Y; X4 = A, Y, or N; X5 = A, G, N, Y, or absent; X6 = G, Y, D, or absent; X7 = Y, T, or absent; X8 = Y, S, or absent; X9 = G or absent; X10 = Y or absent; X11 = Y or absent; X12 = N, Y, F, D, or absent; X13 = W, R, G, or E; X14 = F, I, or M; and X15 = P or V. E16. The 4-1BB antigen-binding protein of E14 or E15, wherein said VL is a Vκ2 that comprises: (i) a CDR-L1 comprising K S S Q S L L X1 X2 X3 G K T Y L X4 (SEQ ID NO: 492), wherein X1 = R or H; X2 = S or T; X3 = D or S; and X4 = Y or F; (ii) a CDR-L2 comprising X1 X2 S N R F S (SEQ ID NO: 493), wherein X1 = E or D; and X2 = V or L; and (iii) a CDR-L3 comprising M Q X1 I X2 X3 P X4 T (SEQ ID NO: 494), wherein X1 = S or T; X2 = Q or H; X3 = L or R; and X4 = W or F. E17. The 4-1BB antigen-binding protein of E14 or E15, wherein said VL is a Vκ1 that comprises: (i) a CDR-L1 comprising R A S Q X1 I X2 X3 Y L N (SEQ ID NO: 495), wherein X1 = S, G, or T; X2 = S, G, or K; and X3 = S, T, N, or R; (ii) a CDR-L2 comprising A X1 S X2 L Q S (SEQ ID NO: 496), wherein X1 = A or I; and X2 = S, T, or N; and (iii) a CDR-L3 comprising X1 X2 X3 X4 X5 X6 X7 X8 T (SEQ ID NO: 497), wherein X1 = Q or L; X2 = Q or K; X3 = S, T, Y, or R; X4 = Y, N, or F; X5 = S or absent; X6 = T, S, V, or I; X7 = P or A; and X8 = L or F. E18. The 4-1BB antigen-binding protein of E14 or E15, wherein said VL is a Vλ3 that comprises: (i) a CDR-L1 comprising X1 G X2 X3 X4 X5 X6 X7 X8 X9 X10 (SEQ ID NO: 498), wherein X1 = G or S; X2 = N or D; X3 = N, A, or K; X4 = I or L; X5 = G or P; X6 = S, K, or D; X7 = K or N; X8 = S,
= S or A; and X3 = D, K, or R; and (iii) a CDR-L3 comprising X1 X2 X3 D S X4 X5 X6 X7 X8 X9 (SEQ ID NO: 500), wherein X1 = Q or Y; X2 = V, S, A, or T; X3 = W, T, or R; X4 = S or absent; X5 = S, G, or absent; X6 = D, N, T, or S; X7 = H or A; X8 = V or G; and X9 = V, A, or M. E19. The 4-1BB antigen-binding protein of E14 or E15, wherein said VL is a Vλ1 that comprises: (i) a CDR-L1 comprising S G X1 X2 S N I G S X3 X4 V N (SEQ ID NO: 501), wherein X1 = G or S; X2 = S or N; X3 = N (ii) a CDR-L2
(iii) a CDR-L3 comprising X1 A W D D S L N G X2 V (SEQ ID NO: 503), wherein X1 = A or E; and X2 = P or V. E20. The 4-1BB antigen-binding protein of any one of E1-E17, comprising a VL framework derived from a human germline Vκ framework sequence. E21. The 4-1BB antigen-binding protein of any one of E1-E15 and E18-E19, comprising a VL framework derived from a human germline Vλ framework sequence. E22. The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH1, VH2, VH3, VH4, or VH5 framework sequence. E23. The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH1 framework sequence. E24. The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH3 framework sequence. E25. The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH4 framework sequence. E26. The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH2 framework sequence. E27. The 4-1BB antigen-binding protein of any one of E1-E21, comprising a VH framework derived from a human germline VH5 framework sequence. E28. The 4-1BB antigen-binding protein of any one of E14-E27, comprising a VL framework sequence and a VH framework sequence, and wherein one or both of the VL framework sequence or VH framework sequence is at least 90% identical to the human germline framework sequence from which it is derived. E29. The 4-1BB antigen-binding protein of any one of E14-E28, comprising a VL framework sequence and a VH framework sequence, and wherein one or both of the VL framework sequence or VH framework sequence is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to the human germline framework sequence from which it is derived. E30. The 4-1BB antigen-binding protein of any one of E1-E29, comprising: a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 147, 149, 151, 153, 155, 157, 159, 324, 161, 163, 165, 167, 169, 171, 173, 175, 404, 406, 408, and 410.
E31. The 4-1BB antigen-binding protein of any one of E1-E30, comprising: a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 148, 150, 152, 154, 156, 158, 160, 325, 162, 164, 166, 168, 170, 172, 174, 176, 405, 407, 409, and 411. E32. The 4-1BB antigen-binding protein of any one of E1-E31, comprising: (a) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.147, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.148; (b) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.149, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.150; (c) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.151, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.152; (d) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.153, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.154; (e) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.155, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.156; (f) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.157, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.158;
(g) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.159 or 324, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.160 or 325; (h) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.161, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.162; (i) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.163, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.164; (j) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.165, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.166; (k) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.167, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.168; (l) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.169, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.170; (m) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.171, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.172;
(n) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.173, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.174; (o) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.175, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.176; (p) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.404, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.405; (q) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.406, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.407; (r) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.408, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.409; or (s) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.410, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.411. E33. A 4-1BB antigen-binding protein that comprises a heavy chain variable domain (VH) and does not comprise a light chain variable domain (VL), wherein said protein comprises the CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, or 385.
E34. A 4-1BB antigen-binding protein that comprises a heavy chain variable domain (VH) and does not comprise a light chain variable domain (VL), wherein said VH comprises: (i) a CDR-H1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 326, 329, 332, 335, 338, 341, 344, 347, 350, 353, 356, 359, 362, 365, or 368; (ii) a CDR-H2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 327, 330, 333, 336, 339, 342, 345, 348, 351, 354, 357, 360, 363, 366, 369; and (iii) a CDR-H3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 328, 331, 334, 337, 340, 343, 346, 349, 352, 355, 358, 361, 364, 367, 370. E35. The 4-1BB antigen-binding protein of E33 or E34, comprising: (1) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.326-328, respectively; (2) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.329-331, respectively; (3) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.332-334, respectively; (4) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.335-337, respectively; (5) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.338-340, respectively; (6) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.341-343, respectively; (7) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.344-346, respectively; (8) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.347-349, respectively; (9) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.350-352, respectively; (10) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.353-355, respectively; (11) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.356-358, respectively; (12) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.359-361, respectively; (13) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.362-364, respectively; (14) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.365-367, respectively; or (15) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.368-370, respectively; E36. A 4-1BB antigen-binding protein that comprises a heavy chain variable domain (VH) and does not comprise a light chain variable domain (VL), wherein said VH comprises: (i) a CDR-H1 comprising X1 X2 X3 M X4 (SEQ ID NO: 480), wherein X1 = S, D, T, or N; X2 = F, Y, or S; X3 = A, W, G, Y, N, or V; and X4 = T, S, H, N, or I; (ii) a CDR-H2 comprising X1 I X2 X3 X4 X5 X6 G X7 X8 Y X9 X10 X11 X12 X13 X14 X15 X16 G (SEQ ID NO: 481), wherein X1 = A, F, N, E, V, G, or Y; X2 = S, R, N, T, or H; X3 = G, S, Q, Y, D,
or W; X4 = K or absent; X5 = A, T, or absent; X6 = S, N, D, G, or Y; X7 = G, S, D, T, or E; X8 = S, T, E, K, or D; X9 = T, K, or I; X10 = Y, E, F, G, or S; X11 = Y or S; X12 = A, V, or P; X13 = G, A, D, or E; X14 = S or A; X15 = V, E, or M; and (ii) a CDR-H3 comprising X1 X2 X3 X4 X5 X6 ID NO:482), wherein X1 = E, D, G, L, M, F, or
M, G, P, R, Y, L, or T; X4 = E, P, T, D, G, R, S, or L; X5 = S, P, A, E, V, I, H, or Y; X6 = S, L, P, T, V, I, G, or F; X7 = L, A, or absent; X8 = V, M, A, G, I, or absent; X9 = Y, L, A, T, P, G, W, S, or absent; X10 = Y, R, N, T, or absent; X11 = T, G, S, V, R, H, P, E, A, or absent; X12 = T, Y, S, G, or A; X13 = S, H, N, T, G, or A; X14 = F or L; X15 = D or E; and X16 = Y or I. E37. The 4-1BB antigen-binding protein of any one of E33-E36, comprising a VH framework derived from a human germline VH1, VH2, VH3, VH4, or VH5 framework sequence. E38. The 4-1BB antigen-binding protein of any one of E33-E37, comprising a VH framework derived from a human germline VH1 framework sequence. E39. The 4-1BB antigen-binding protein of any one of E33-E37, comprising a VH framework derived from a human germline VH3 framework sequence. E40. The 4-1BB antigen-binding protein of any one of E33-E37, comprising a VH framework derived from a human germline VH4 framework sequence. E41. The 4-1BB antigen-binding protein of any one of E33-E37, comprising a VH framework derived from a human germline VH2 framework sequence. E42. The 4-1BB antigen-binding protein of any one of E33-E37, comprising a VH framework derived from a human germline VH5 framework sequence. E43. The 4-1BB antigen-binding protein of any one of E33-E42, comprising a VH framework sequence is at least 90% identical to the human germline framework sequence from which it is derived. E44. The 4-1BB antigen-binding protein of any one of E33-E43, comprising a VH framework sequence is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to the human germline framework sequence from which it is derived. E45. The 4-1BB antigen-binding protein of any one of E33-E44, comprising: a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, and 385.
E46. The 4-1BB antigen-binding protein of any one of E33-E45, comprising: (a) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.371; (b) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.372; (c) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.373; (d) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.374; (e) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.375; (f) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.376; (g) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.377; (h) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.378; (i) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.379; (j) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.380; (k) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.381; (l) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.382;
(m) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.383; (n) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.384; or (o) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.385. E47. The 4-1BB antigen-binding protein of any one of E1-E46 and E315-E342, further comprising a heavy chain CH1 domain. E48. The 4-1BB antigen-binding protein of E47, wherein said CH1 domain is the CH1 domain of an IgG (for example IgG1, lgG2, lgG3, or lgG4). E49. The 4-1BB antigen-binding protein of E47 or E48, wherein said CH1 domain is the CH1 domain of a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4). E50. The 4-1BB antigen-binding protein of any one of E47-E49, wherein said CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 261, 265, 269, 422, or 425. E51. The 4-1BB antigen-binding protein of any one of E47-E50, wherein said CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 422, or 425. E52. The 4-1BB antigen-binding protein of any one of E1-E51 and E315-E342, further comprising an Fc region. E53. The 4-1BB antigen-binding protein of E52, wherein the Fc region is the Fc region of an IgA (for example IgA1 or lgA2), IgD, IgE, IgM, or IgG (for example IgG1, lgG2, lgG3, or lgG4). E54. The 4-1BB antigen-binding protein of E52 or E53, wherein the Fc region is the Fc region of an IgG.
E55. The 4-1BB antigen-binding protein of E54, wherein the IgG is selected from the group consisting of IgG1, lgG2, lgG3, and lgG4. E56. The 4-1BB antigen-binding protein of E55, wherein the IgG is IgG1, IgG2, or IgG4. E57. The 4-1BB antigen-binding protein of any one of E52-E56, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: L234A, L235A, L235E, G237A, and combination thereof (numbering according to the EU index). E58. The 4-1BB antigen-binding protein of E57, comprising L234A and L235A mutations. E59. The 4-1BB antigen-binding protein of any one of E52-E58, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: V259C, A287C, R292C, V302C, L306C, V323C, I332C, and a combination thereof (numbering according to the EU index). E60. The 4-1BB antigen-binding protein of any one of E52-E59, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: L242C, A287C, R292C, N297G, V302C, L306C, K334C, and a combination thereof (numbering according to the EU index). E61. The 4-1BB antigen-binding protein of E60, comprising a N297G mutation. E62. The 4-1BB antigen-binding protein of E60, comprising A287C, N297G, and L306C mutations. E63. The 4-1BB antigen-binding protein of E60, comprising R292C, N297G, and V302C mutations. E64. The 4-1BB antigen-binding protein of any one of E52-E63, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: M252Y, S254T, T256E, and a combination thereof. E65. The 4-1BB antigen-binding protein of E64, comprising M252Y, S254T, T256E mutations. E66. The 4-1BB antigen-binding protein of any one of E52-E65, wherein the lysine residue (K) at the C-terminus of the Fc region is deleted. E67. The 4-1BB antigen-binding protein of any one of E52-E65, wherein the lysine residue (K) at the C-terminus of the Fc region is present. E68. The 4-1BB antigen-binding protein of any one of E52-E65, wherein the glycine and lysine residues (GK) at the C-terminus of the Fc region are present.
E69. The 4-1BB antigen-binding protein of any one of E52-E65, wherein the glycine and lysine residues (GK) at the C-terminus of the Fc region are deleted. E70. The 4-1BB antigen-binding protein of any one of E52-E69, wherein said Fc region comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, or 426. E71. The 4-1BB antigen-binding protein of any one of E52-E69, wherein said Fc region comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 263, 267, or 483. E72. The 4-1BB antigen-binding protein of any one of E1-E71 and E315-E342, further comprising a heavy chain constant domain. E73. The 4-1BB antigen-binding protein of E72, wherein said constant domain is the constant domain of an IgA (for example IgA1 or lgA2), IgD, IgE, IgM, or IgG (for example IgG1, lgG2, lgG3, or lgG4). E74. The 4-1BB antigen-binding protein of E72 or E73, wherein said constant domain is the constant domain of an IgG (for example IgG1, IgG2, IgG3, or IgG4), preferably a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4). E75. The 4-1BB antigen-binding protein of any one of E72-E74, wherein said constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 244, 245, 246, 247, 248, 249, 256, 257, 258, 260, 412, 415, 417, 418, or 427. E76. The 4-1BB antigen-binding protein of any one of E72-E74, wherein said constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 264, 268, or 271. E77. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E76, further comprising a kappa or lambda light chain constant domain.
E78. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E77, further comprising a kappa light chain constant domain that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428. E79. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E77, further comprising a lambda light chain constant domain that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424. E80. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E79, comprising: (i) a heavy chain (HC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 191, 193, 195, 197, 199, 201, 203, 205, 207, 209, 211, 213, 215, 217, and 219; and (ii) a light chain (LC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 192, 194, 196, 198, 200, 202, 204, 206, 208, 210, 212, 214, 216, 218, and 220. E81. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E80, comprising: (a) a heavy chain (HC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.191, and a light chain (LC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.192; (b) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.193, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.194; (c) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.195, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.196;
(d) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.197, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.198; (e) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.199, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.200; (f) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.201, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.202; (g) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.203, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.204; (h) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.205, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.206; (i) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.207, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.208; (j) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.209, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.210; (k) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100%
identical to SEQ ID NO.211, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.212; (l) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.213, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.214; (m) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.215, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.216; (n) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.217, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.218; or (o) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.219, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.220. E82. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E81, which is an antibody. E83. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E81, which is an antigen- binding fragment of an antibody, such as a Fab fragment. E84. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E81, which is an scFv. E85. The 4-1BB antigen-binding protein of E84, wherein said scFv comprises a first linker between VH and VL. E86. The 4-1BB antigen-binding protein of E85, wherein said first linker comprises: (a) a glycine rich peptide; (b) a peptide comprising glycine and serine; (c) a peptide comprising (Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 234); (d) a peptide comprising (Gly-Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 232); (e) a peptide comprising (Gly-Gly-Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6
(SEQ ID NO: 233); (f) a peptide comprising (Gly-Gly-Gly-Gly-Gln)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 235), or (g) a peptide comprising any one of SEQ ID NOs.221-235 and 504. E87. The 4-1BB antigen-binding protein of E85, wherein said first linker comprises the amino acid sequence of (Gly-Gly-Gly-Gly-Ser)3 (SEQ ID NO: 230). E88. The 4-1BB antigen-binding protein of any one of E1-E32 and E84-E87, comprising a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 291 (14A5.002 scFv). E89. The 4-1BB antigen-binding protein of any one of E1-E32 and E84-E87, comprising a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 445 (14A5.002 scFv #2). E90. The 4-1BB antigen-binding protein of any one of E1-E32 and E47-E81, which is a Fab. E91. The 4-1BB antigen-binding protein of E90, comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 298, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 300 (6F9.009 Fab). E92. The 4-1BB antigen-binding protein of E90, comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 305, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 307 (19G1.016 Fab). E93. The 4-1BB antigen-binding protein of E90, comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least
87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 316, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 317 (6C7.018 Fab). E94. The 4-1BB antigen-binding protein of E90, comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-227 of SEQ ID NO: 434, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 435 (4-1BB Fab of 56039). E95. The 4-1BB antigen-binding protein of E90, comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-227 of SEQ ID NO: 449, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 450 (4-1BB Fab of 56040). E96. The 4-1BB antigen-binding protein of any one of E1-E95, wherein the antigen-binding protein binds to 4-1BB with a KD value of or less than: about 500nM, about 400nM, about 300nM, about 200nM, about 150nM, about 100nM, about 90nM, about 80nM, about 70nM, about 60nM, about 50nM, about 40nM, about 30nM, about 25nM, about 20nM, about 15nM, about 10nM, about 9nM, about 8nM, about 7nM, about 6nM, about 5nM, about 4nM, about 3nM, about 2nM, about 1 nM, about 900pM, about 800pM, about 700pM, about 600pM, about 500pM, about 400pM, about 300pM, about 250pM, about 200pM, about 150pM, about 100pM, about 50pM, about 40pM, about 30pM, about 25pM, about 20pM, about 15pM, about 10pM, about 5pM, or about 1pM. E97. A 4-1BB antigen-binding protein that competes for binding to 4-1BB with any one of the 4-1BB antigen binding protein of E1-E96 and E315-E342. E98. A 4-1BB antigen-binding protein that binds to substantially the same epitope as any one of the 4- 1BB antigen binding protein of E1-E96 and E315-E342.
E99. A bispecific molecule that comprises the 4-1BB antigen-binding protein of any one of E1-E98 and E315-E342, and further comprises a second antigen-binding moiety. E100. The bispecific molecule of E99, wherein said second antigen-binding moiety binds to a protein of the immune checkpoint pathway. E101. The bispecific molecule of E100, wherein the protein of the immune checkpoint pathway is CTLA- 4, PD-1, PD-L1, PD-L2, B7-H3, B7-H4, CEACAM-1, TIGIT, LAG3, CD112, CD112R, CD96, TIM3, or BTLA. E102. The bispecific molecule of E101, wherein the protein of the immune checkpoint pathway is PD- L1. E103. A PD-L1 antigen-binding protein, comprising (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 133, 135, 137, 139, 141, 143, 322, 145, 398, 400, or 402; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 134, 136, 138, 140, 142, 144, 323, 146, 399, 401, or 403. E104. A PD-L1 antigen-binding protein, comprising: (i) a CDR-H1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 1, 7, 13, 19, 25, 31, or 37; (ii) a CDR-H2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 2, 8, 14, 20, 26, 32, or 38; (iii) a CDR-H3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 3, 9, 15, 21, 27, 33, or 39; (iv) a CDR-L1 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 4, 10, 16, 22, 28, 34, or 40; (v) a CDR-L2 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 5, 11, 17, 23, 29, 35, or 41; and (vi) a CDR-L3 comprising a sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 6, 12, 18, 24, 30, 36, or 42. E105. The PD-L1 antigen-binding protein of E103 or E104, comprising:
(1) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.1-6, respectively; (2) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.7-12, respectively; (3) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.13-18, respectively; (4) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.19-24, respectively; (5) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.25-30, respectively; (6) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.31-36, respectively; or (7) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.37-42, respectively. E106. The PD-L1 antigen-binding protein of any one of E103-E105, comprising a VL framework derived from a human germline Vκ framework sequence. E107. The PD-L1 antigen-binding protein of any one of E103-E105, comprising a VL framework derived from a human germline Vλ framework sequence. E108. The PD-L1 antigen-binding protein of any one of E103-E107, comprising a VH framework derived from a human germline VH1, VH2, VH3, VH4, or VH5 framework sequence. E109. The PD-L1 antigen-binding protein of any one of E103-E107, comprising a VH framework derived from a human germline VH1 framework sequence. E110. The PD-L1 antigen-binding protein of any one of E103-E107, comprising a VH framework derived from a human germline VH2 framework sequence. E111. The PD-L1 antigen-binding protein of any one of E103-E107, comprising a VH framework derived from a human germline VH3 framework sequence. E112. The PD-L1 antigen-binding protein of any one of E103-E107, comprising a VH framework derived from a human germline VH4 framework sequence. E113. The PD-L1 antigen-binding protein of any one of E103-E107, comprising a VH framework derived from a human germline VH5 framework sequence.
E114. The PD-L1 antigen-binding protein of any one of E103-E113, comprising a VL framework sequence and a VH framework sequence, and wherein one or both of the VL framework sequence or VH framework sequence is at least 90% identical to the human germline framework sequence from which it is derived. E115. The PD-L1 antigen-binding protein of any one of E103-E114, comprising a VL framework sequence and a VH framework sequence, and wherein one or both of the VL framework sequence or VH framework sequence is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to the human germline framework sequence from which it is derived. E116. The PD-L1 antigen-binding protein of any one of E103-E115, comprising a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs:133, 135, 137, 139, 141, 143, 322, 145, 398, 400, and 402 E117. The PD-L1 antigen-binding protein of any one of E103-E116, comprising a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 134, 136, 138, 140, 142, 323144, 146, 399, 401, and 403. E118. The PD-L1 antigen-binding protein of any one of E103-E117, comprising: (1) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.133, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.134; (2) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.135, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.136; (3) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.137, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.138; (4) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100%
identical to SEQ ID NO.139, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.140; (5) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.141, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.142; (6) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.143, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.144; (7) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.322, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.323; (8) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.145, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.146; (9) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.398, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.399; (10) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.400, and a VL comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.401; or (11) a VH comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.402, and a VL comprising an amino acid sequence at least 90%, at least
91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.403. E119. The PD-L1 antigen-binding protein of any one of E103-E118, further comprising a heavy chain CH1 domain. E120. The PD-L1 antigen-binding protein of E119, wherein said CH1 domain is the CH1 domain of an IgG (for example IgG1, lgG2, lgG3, or lgG4). E121. The PD-L1 antigen-binding protein of E119 or E120, wherein said CH1 domain is the CH1 domain of a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4). E122. The PD-L1 antigen-binding protein of any one of E119-E121, wherein said CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 261, 265, 269, 422, or 425. E123. The PD-L1 antigen-binding protein of any one of E119-E121, wherein said CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 422, or 425. E124. The PD-L1 antigen-binding protein of any one of E103-E123, further comprising an Fc region. E125. The PD-L1 antigen-binding protein of E124, wherein the Fc region is the Fc region of an IgA (for example IgA1 or lgA2), IgD, IgE, IgM, or IgG (for example IgG1, lgG2, lgG3, or lgG4). E126. The PD-L1 antigen-binding protein of E124 or E125, wherein the Fc region is the Fc region of an IgG. E127. The PD-L1 antigen-binding protein of E126, wherein the IgG is selected from the group consisting of IgG1, lgG2, lgG3, and lgG4. E128. The PD-L1 antigen-binding protein of E126, wherein the IgG is IgG1, IgG2, or IgG4. E129. The PD-L1 antigen-binding protein of any one of E124-E128, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: L234A, L235A, L235E, G237A, and combination thereof (numbering according to the EU index).
E130. The PD-L1 antigen-binding protein of E129, comprising L234A and L235A mutations. E131. The PD-L1 antigen-binding protein of any one of E124-E130, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: V259C, A287C, R292C, V302C, L306C, V323C, I332C, and a combination thereof (numbering according to the EU index). E131. The PD-L1 antigen-binding protein of any one of E124-E130, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: L242C, A287C, R292C, N297G, V302C, L306C, K334C, and a combination thereof (numbering according to the EU index). E132. The PD-L1 antigen-binding protein of E131, comprising a N297G mutation. E133. The PD-L1 antigen-binding protein of E131, comprising A287C, N297G, and L306C mutations. E134. The PD-L1 antigen-binding protein of E131, comprising R292C, N297G, and V302C mutations. E135. The PD-L1 antigen-binding protein of any one of E124-E134, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: M252Y, S254T, T256E, and a combination thereof. E136. The PD-L1 antigen-binding protein of E135, comprising M252Y, S254T, T256E mutations. E137. The PD-L1 antigen-binding protein of any one of E124-E136, wherein the lysine residue (K) at the C-terminus of the Fc region is deleted. E138. The PD-L1 antigen-binding protein of any one of E124-E136, wherein the lysine residue (K) at the C-terminus of the Fc region is present. E139. The PD-L1 antigen-binding protein of any one of E124-E136, wherein the glycine and lysine residues (GK) at the C-terminus of the Fc region are present. E140. The PD-L1 antigen-binding protein of any one of E124-E136, wherein the glycine and lysine residues (GK) at the C-terminus of the Fc region are deleted. E141. The PD-L1 antigen-binding protein of any one of E124-E140, wherein said Fc region comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, or 426.
E142. The PD-L1 antigen-binding protein of any one of E124-E140, wherein said Fc region comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 263, 267, or 483. E143. The PD-L1 antigen-binding protein of any one of E103-E142, further comprising a heavy chain constant domain. E144. The PD-L1 antigen-binding protein of E143, wherein said constant domain is the constant domain of an IgA (for example IgA1 or lgA2), IgD, IgE, IgM, or IgG (for example IgG1, lgG2, lgG3, or lgG4). E145. The PD-L1 antigen-binding protein of E143 or E144, wherein said constant domain is the constant domain of an IgG (for example IgG1, IgG2, IgG3, or IgG4), preferably a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4). E146. The PD-L1 antigen-binding protein of any one of E143-E145, wherein said constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 244, 245, 246, 247, 248, 249, 256, 257, 258, 260, 412, 415, 417, 418, or 427. E147. The PD-L1 antigen-binding protein of any one of E143-E145, wherein said constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 264, 268, or 271. E148. The PD-L1 antigen-binding protein of any one of E103-E147, further comprising a kappa or lambda light chain constant domain. E149. The PD-L1 antigen-binding protein of any one of E103-E148, further comprising a kappa light chain constant domain that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428. E150. The PD-L1 antigen-binding protein of any one of E103-E148, further comprising a lambda light chain constant domain that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least
85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424. E151. The PD-L1 antigen-binding protein of any one of E103-E150, comprising: (i) a heavy chain (HC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 177, 179, 181, 183, 185, 187, and 189; and (ii) a light chain (LC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of SEQ ID NOs: 178, 180, 182, 184, 186, 188, and 190. E152. The PD-L1 antigen-binding protein of any one of E103-E151, comprising: (a) a heavy chain (HC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.177, and a light chain (LC) comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.178; (b) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.179, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.180; (c) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.181, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.182; (d) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.183, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.184; (e) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.185, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.186;
(f) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.187, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.188; or (g) a HC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.189, and a LC comprising an amino acid sequence at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO.190. E153. The PD-L1 antigen-binding protein of any one of E103-E152, which is an antibody. E154. PD-L1 antigen-binding protein of any one of E103-E152, which is an antigen-binding fragment of an antibody, such as a Fab fragment. E155. The PD-L1 antigen-binding protein of any one of E103-E152, which is an scFv. E156. The PD-L1 antigen-binding protein of E155, wherein said scFv comprises a first linker between VH and VL. E157. The PD-L1 antigen-binding protein of E156, wherein said first linker comprises: (a) a glycine rich peptide; (b) a peptide comprising glycine and serine; (c) a peptide comprising (Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 234); (d) a peptide comprising (Gly-Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 232); (e) a peptide comprising (Gly-Gly-Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 233); (f) a peptide comprising (Gly-Gly-Gly-Gly-Gln)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 235), or (g) a peptide comprising any one of SEQ ID NOs.221-235 and 504. E158. The PD-L1 antigen-binding protein of E156, wherein said first linker comprises the amino acid sequence of (Gly-Gly-Gly-Gly-Ser)3 (SEQ ID NO: 230). E159. The PD-L1 antigen-binding protein of any one of E155-E158, comprising a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 284 (26F6.002.009 scFv). E160. The PD-L1 antigen-binding protein of any one of E155-E158, comprising a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at
least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 436 (scFv from 56039). E161. The PD-L1 antigen-binding protein of any one of E103-E152, which is a Fab. E162. The PD-L1 antigen-binding protein of E161, comprising a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-221 SEQ ID NO: 451, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 452 (Fab from 56041). E163. The PD-L1 antigen-binding protein of any one of E103-E162, wherein the protein binds PD-L1 with a KD value of or less than: about 10nM, about 5nM, about 2nM, about 1 nM, about 900pM, about 800pM, about 700pM, about 600pM, about 500pM, about 400pM, about 300pM, about 250pM, about 200pM, about 150pM, about 100pM, about 50pM, about 40pM, about 30pM, about 25pM, about 20pM, about 15pM, about 10pM, about 5pM, or about 1 pM. E164. A PD-L1 antigen-binding protein that competes for binding to PD-L1 with any one of the PD-L1 antigen binding protein of E103-E163. E165. A PD-L1 antigen-binding protein that binds to substantially the same epitope as any one of the PD-L1 antigen binding protein of E103-E163. E166. A bispecific molecule comprising the PD-L1 antigen-binding protein of any one of E103-E165, and further comprising a second antigen-binding moiety. E167. The bispecific molecule of E166, wherein the second antigen-binding moiety binds to a T-cell co- stimulatory molecule (such as: CD28, Inducible Co-Stimulator (ICOS), CTLA4 (Cytotoxic T-Lymphocyte- Associated protein 4), 4-1BB (also known as CD137), OX40 (also known as CD134), CD27, CD30, DR3, Glucocorticoid-Induced TNFR family Related (GITR), or Herpes Virus Entry Mediator (HVEM)). E168. The bispecific molecule of E167, wherein the T-cell co-stimulatory molecule is 4-1BB. E169. A bispecific molecule comprising (i) a 4-1BB antigen-binding moiety comprising any one of E1- E98 and E315-E342, and (ii) a PD-L1 antigen-binding moiety of any one of embodiments E103-E165. E170. A bispecific molecule, comprising:
(i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker - scFv, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a second linker; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said scFv binds to 4-1BB, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (IgG-scFv(C2), Fig.14A) E171. A bispecific molecule, comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker - scFv, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a second linker; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said scFv binds to PD-L1, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (IgG-scFv(C2), Fig.14A) E172. A bispecific molecule, comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – first linker – scFv – third linker – monomeric CH2 – monomeric CH3, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a second linker; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said scFv binds to 4-1BB, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (Fab-scFv(M2)-Fc, Fig.14B) E173. A bispecific molecule, comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – first linker – scFv – third linker – monomeric CH2 – monomeric CH3, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a second linker; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said scFv binds to PD-L1, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (Fab-scFv(M2)-Fc, Fig.14B) E174. A bispecific molecule, comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker – VHB – CH1’; and (ii) two copies of a first light chain that comprises, from N-terminus to C-terminus: VLA – CL; (iii) two copies of a second light chain that comprises, from N-terminus to C-terminus: VLB – CL’; wherein said VHA-CH1 and VLA-CL form a first Fab that binds to PD-L1, said VHB-CH1’ and VLB-CL’ form a second Fab that binds to 4-1BB, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (IgG-Fab, Fig.2B)
E175. A bispecific molecule, comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker – VHB – CH1’; and (ii) two copies of a first light chain that comprises, from N-terminus to C-terminus: VLA – CL; (iii) two copies of a second light chain that comprises, from N-terminus to C-terminus: VLB – CL’; wherein said VHA-CH1 and VLA-CL form a first Fab that binds to 4-1BB, said VHB-CH1’ and VLB-CL’ form a second Fab that binds to PD-L1, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (IgG-Fab, Fig.2B) E176. The bispecific molecule of E174 or E175, wherein (i) said CH1 comprises a mutation to a positively charged residue and said CL comprises a mutation to a negatively charged residue, such that said CH1 and CL form a first charge pair, and (ii) said CH1’ comprises a mutation to a negatively charged residue and said CL’ comprises a mutation to a positively charged residue, such that said CH1’ and CL’ form a second charge pair. E177. The bispecific molecule of E176, wherein said CH1 comprises a mutation to K or R (preferably K) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246. E178. The bispecific molecule of E176 or E177, wherein said CL comprises mutation to E or D (preferably E) at position 176 (Kabat numbering), or at a position that corresponds to residue 69 of SEQ ID NO:237 or 239. E179. The bispecific molecule of any one of E176- E178, wherein said CH1’ comprises a mutation to E or D (preferably E) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246. E180. The bispecific molecule of any one of E176- E179, wherein said CL’ comprises a mutation to K or R (preferably K) at position 176 (Kabat numbering), or at a position that corresponds to residue 69 of SEQ ID NO:237 or 239. E181. The bispecific molecule of E174 or E175, wherein (i) said CH1 comprises a mutation to a negatively charged residue and said CL comprises a mutation to a positively charged residue, such that said CH1 and CL form a first charge pair, and (ii) said CH1’ comprises a mutation to a positively charged residue and said CL’ comprises a mutation to a negatively charged residue, such that said CH1’ and CL’ form a second charge pair. E182. The bispecific molecule of E181, wherein said CH1 comprises a mutation to E or D (preferably E) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246.
E183. The bispecific molecule of E181 or E182, wherein said CL comprises mutation to K or R (preferably R) at position 176 (Kabat numbering), or at a position that corresponds to residue 69 of SEQ ID NO:237 or 239. E184. The bispecific molecule of any one of E181- E183, wherein said CH1’ comprises a mutation to K or R (preferably K) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246. E185. The bispecific molecule of any one of E181- E184, wherein said CL’ comprises a mutation to E or D (preferably E) at position 176 (Kabat numbering), or at a position that corresponds to residue 69 of SEQ ID NO:237 or 239. E186. A bispecific molecule, comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – first linker – VHB – second linker – monomeric CH2 – monomeric CH3; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said VHB binds to 4-1BB, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (Fab-VH(M2)-Fc, Fig.14D). E187. A bispecific molecule, comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – first linker – VHB – second linker – monomeric CH2 – monomeric CH3; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said VHB binds to PD-L1, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (Fab-VH(M2)-Fc, Fig.14D). E188. A bispecific molecule, comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker - VHB; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said VHB binds to 4-1BB, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (IgG-VH(C2), Fig.14C) E189. A bispecific molecule, comprising: (i) two copies of a heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker - VHB; and (ii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said VHB binds to PD-L1, and said two copies of monomeric CH2 – monomeric CH3 form a Fc region. (IgG-VH(C2), Fig.14C)
E190. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker – scFv; wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a second linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form two Fab domains that binds to 4-1BB, said scFv binds to PD-L1, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. (IgG-scFv(C1), Fig.13C) E191. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3 – first linker – scFv; wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a second linker (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form two Fab domains that binds to PD-L1, said scFv binds to 4- 1BB, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. (IgG-scFv(C1), Fig.13C) E192. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – first linker – scFv – third linker – monomeric CH2 – monomeric CH3; wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a second linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form two Fab domains that binds to 4-1BB, said scFv binds to PD-L1, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. (Fab-scFv(M1)-Fc, Fig.13D) E193. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – first linker – scFv – third linker – monomeric CH2 – monomeric CH3; wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a second linker;
(ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form two Fab domains that binds to PD-L1, said scFv binds to 4- 1BB, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. (Fab-scFv(M1)-Fc, Fig.13D) E194. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – monomeric CH2’ – monomeric CH3’, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a first linker; and (iii) a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said scFv binds to 4-1BB, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. ([Fab*scFv] hetero-Fc, Fig.13G) E195. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – monomeric CH2’ – monomeric CH3’, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a first linker; and (iii) a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said scFv binds to PD-L1, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. ([Fab*scFv] hetero-Fc, Fig.13G) E196. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHB – first linker – monomeric CH2’ – monomeric CH3’; and (iii) a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said VHB binds to 4-1BB, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. ([Fab*VH] hetero-Fc, Fig.13H)
E197. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHB – first linker – monomeric CH2’ – monomeric CH3’; and (iii) a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said VHB binds to PD-L1, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. ([Fab*VH] hetero-Fc, Fig.13H) E198. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – VHA – CH1 – monomeric CH2 – monomeric CH3, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a first linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said scFv binds to 4-1BB, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. (scFv(N1)-IgG, Fig.13J) E199. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – VHA – CH1 – monomeric CH2 – monomeric CH3, wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a first linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2’ – monomeric CH3’; and (iii) two copies of a light chain that comprises, from N-terminus to C-terminus: VLA – CL; wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said scFv binds to PD-L1, and said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. (scFv(N1)-IgG, Fig.13J) E200. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – monomeric CH2(1) – monomeric CH3(1), wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a first linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2(2) – monomeric CH3(2) – monomeric CH2(3) – monomeric CH3(3);
(iii) a light chain that comprises, from N-terminus to C-terminus: VLA – CL; and (iv) a fourth chain comprising monomeric CH2(4) – monomeric CH3(4); wherein said VHA-CH1 and said VLA-CL form a Fab that binds to PD-L1, said scFv binds to 4-1BB, said monomeric CH2(1) – monomeric CH3(1) from (i) and monomeric CH2(2) – monomeric CH3(2) from (ii) form a first Fc region, and said monomeric CH2(3) – monomeric CH3(3) from (ii) and monomeric CH2(4) – monomeric CH3(4) from (iv) form a second Fc region. ([scFv*Fab] hetero-Fc)-Fc, (Fig.13K) E201. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: scFv – second linker – monomeric CH2(1) – monomeric CH3(1), wherein said scFv comprises a VHB and a VLB, and wherein said VHB and VLB are connected via a first linker; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2(2) – monomeric CH3(2) – monomeric CH2(3) – monomeric CH3(3); (iii) a light chain that comprises, from N-terminus to C-terminus: VLA – CL; and (iv) a fourth chain comprising monomeric CH2(4) – monomeric CH3(4); wherein said VHA-CH1 and said VLA-CL form a Fab that binds to 4-1BB, said scFv binds to PD-L1, said monomeric CH2(1) – monomeric CH3(1) from (i) and monomeric CH2(2) – monomeric CH3(2) from (ii) form a first Fc region, and said monomeric CH2(3) – monomeric CH3(3) from (ii) and monomeric CH2(4) – monomeric CH3(4) from (iv) form a second Fc region. ([scFv*Fab] hetero-Fc)-Fc, (Fig.13K) E202. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHB – CH1’ – monomeric CH2’ – monomeric CH3’; (iii) a first light chain that comprises, from N-terminus to C-terminus: VLA – CL; and (iv) a second light chain that comprises, from N-terminus to C-terminus: VLB – CL’; wherein said VHA-CH1 and said VLA-CL form a first Fab that binds to 4-1BB, wherein said VHB-CH1’ and said VLB-CL’ form a second Fab that to PD-L1, and wherein said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. (hetero-IgG, Fig.13I) E203. A bispecific molecule, comprising: (i) a first heavy chain that comprises, from N-terminus to C-terminus: VHA – CH1 – monomeric CH2 – monomeric CH3; (ii) a second heavy chain that comprises, from N-terminus to C-terminus: VHB – CH1’ – monomeric CH2’ – monomeric CH3’; (iii) a first light chain that comprises, from N-terminus to C-terminus: VLA – CL; and (iv) a second light chain that comprises, from N-terminus to C-terminus: VLB – CL’;
wherein said VHA-CH1 and said VLA-CL form a first Fab that binds to PD-L1, wherein said VHB-CH1’ and said VLB-CL’ form a second Fab that to 4-1BB, and wherein said monomeric CH2 – monomeric CH3 from (i) and monomeric CH2’ – monomeric CH3’ from (ii) form a Fc region. (hetero-IgG, Fig.13I) E204. The bispecific molecule of any one of E202 or E203, wherein (i) said CH1 comprises a mutation to a positively charged residue and said CL comprises a mutation to a negatively charged residue, such that said CH1 and CL form a first charge pair, and (ii) said CH1’ comprises a mutation to a negatively charged residue and said CL’ comprises a mutation to a positively charged residue, such that said CH1’ and CL’ form a second charge pair. E205. The bispecific molecule of E204, wherein said CH1 comprises a mutation to K or R (preferably K) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246. E206. The bispecific molecule of E204 or E205, wherein said CL comprises mutation to E or D (preferably E) at position 176 (Kabat numbering), or at a position that corresponds to residue 69 of SEQ ID NO:237 or 239. E207. The bispecific molecule of any one of E204- E206, wherein said CH1’ comprises a mutation to E or D (preferably E) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246. E208. The bispecific molecule of any one of E204- E207, wherein said CL’ comprises a mutation to K or R (preferably K) at position 176 (Kabat numbering), or at a position that corresponds to residue 69 of SEQ ID NO:237 or 239. E209. The bispecific molecule of E202 or E203, wherein (i) said CH1 comprises a mutation to a negatively charged residue and said CL comprises a mutation to a positively charged residue, such that said CH1 and CL form a first charge pair, and (ii) said CH1’ comprises a mutation to a positively charged residue and said CL’ comprises a mutation to a negatively charged residue, such that said CH1’ and CL’ form a second charge pair. E210. The bispecific molecule of E209, wherein said CH1 comprises a mutation to E or D (preferably E) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246. E211. The bispecific molecule of E209 or E210, wherein said CL comprises mutation to K or R (preferably R) at position 176 (Kabat numbering), or at a position that corresponds to residue 69 of SEQ ID NO:237 or 239.
E212. The bispecific molecule of any one of E209- E211, wherein said CH1’ comprises a mutation to K or R (preferably K) at position 183 (EU index numbering), or at a position that corresponds to residue 66 of SEQ ID NO:246. E213. The bispecific molecule of any one of E209- E212, wherein said CL’ comprises a mutation to E or D (preferably E) at position 176 (Kabat numbering), or at a position that corresponds to residue 69 of SEQ ID NO:237 or 239. E214. The bispecific molecule of any one of E190- E213, wherein said CH2, or CH2’, or CH2(1), or CH2(2), or CH2(3), or CH2(4) comprises a mutation wherein a positively charged residue is mutated to a negatively charge residue, or a mutation wherein a negatively charged residue is mutated to a positively charge residue. E215. The bispecific molecule of any one of E190- E214, wherein said CH3, or CH3’, or CH3(1), or CH3(2), or CH3(3), or CH3(4) comprises a mutation wherein a positively charged residue is mutated to a negatively charge residue, or a mutation wherein a negatively charged residue is mutated to a positively charge residue. E216. The bispecific molecule of any one of E190- E215, wherein said CH3, or CH3’, or CH3(1), or CH3(2), or CH3(3), or CH3(4) comprises a mutation from K to E or D (preferably D) at position 392 (EU index numbering), or at a position that corresponds to residue 275 of SEQ ID NO:246. E217. The bispecific molecule of any one of E190- E216, wherein said CH3, or CH3’, or CH3(1), or CH3(2), or CH3(3), or CH3(4) comprises a mutation from K to D or E (preferably D) at position 409 (EU index numbering), or at a position that corresponds to residue 292 of SEQ ID NO:246. E218. The bispecific molecule of any one of E190- E217, wherein said CH3, or CH3’, or CH3(1), or CH3(2), or CH3(3), or CH3(4) comprises a mutation from K to E or D (preferably D) at position 439 (EU index numbering), or at a position that corresponds to residue 322 of SEQ ID NO:246. E219. The bispecific molecule of any one of E170- E219, wherein said CH1 or CH1’ is the CH1 domain of a human IgG (for example, human IgG1, human IgG2, human IgG3, or human IgG4). E220. The PD-L1 antigen-binding protein of any one of E170-E219, wherein said CH1 or CH1’ domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 261, 265, 269, 422, or 425.
E221. The bispecific molecule of any one of E170-E220, wherein said CH1 or CH1’ domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 422, or 425. E222. The bispecific molecule of any one of E170-E221, wherein said Fc is the Fc region of an IgG. E223. The bispecific molecule of E222, wherein the IgG is selected from the group consisting of IgG1, lgG2, lgG3, and lgG4. E224. The bispecific molecule of E223, wherein the IgG is IgG1, IgG2, or IgG4. E225. The bispecific molecule of any one of E170-E224, wherein said Fc is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: L234A, L235A, L235E, G237A, and combination thereof (numbering according to the EU index). E226. The bispecific molecule of E225, comprising L234A and L235A mutations. E227. The bispecific molecule of any one of E170-E226, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: V259C, A287C, R292C, V302C, L306C, V323C, I332C, and a combination thereof (numbering according to the EU index). E228. The bispecific molecule of any one of E170-E227, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: L242C, A287C, R292C, N297G, V302C, L306C, K334C, and a combination thereof (numbering according to the EU index). E229. The bispecific molecule of E228, comprising a N297G mutation. E230. The bispecific molecule of E228, comprising A287C, N297G, and L306C mutations. E231. The bispecific molecule of E228, comprising R292C, N297G, and V302C mutations. E232. The bispecific molecule of any one of E170-E231, wherein said Fc region is derived from an IgG Fc, and further comprises one or more mutations selection from the group consisting of: M252Y, S254T, T256E, and a combination thereof. E233. The bispecific molecule of E232, comprising M252Y, S254T, T256E mutations.
E234. The bispecific molecule of any one of E170-E233, wherein the lysine residue (K) at the C- terminus of the Fc region is deleted. E235. The bispecific molecule of any one of E170-E233, wherein the lysine residue (K) at the C- terminus of the Fc region is present. E236. The bispecific molecule of any one of E170-E233, wherein the glycine and lysine residues (GK) at the C-terminus of the Fc region are present. E237. The bispecific molecule of any one of E170-E233, wherein the glycine and lysine residues (GK) at the C-terminus of the Fc region are deleted. E238. The bispecific molecule of any one of E170-E237, wherein said Fc comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, or 426. E239. The bispecific molecule of any one of E170-E237, wherein said Fc region comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 263, 267, or 483. E240. The bispecific molecule of any one of E170-E239, wherein said CL or CL’ is a kappa or lambda light chain constant domain. E241. The bispecific molecule of any one of E170-E240, wherein said CL or CL’ is a kappa light chain constant domain that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428. E242. The bispecific molecule of any one of E170-E241, wherein said CL or CL’ a lambda light chain constant domain that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424.
E243. The bispecific molecule of any one of E170-E242, comprising a 4-1BB antigen-binding protein of any one of E1-E98 and E315-E342. E244. The bispecific molecule of any one of E170-E243, comprising PD-L1 antigen-binding protein of any one of E103-E165. E245. The bispecific molecule of any one of E170-E244, wherein said first linker, second linker, or third linker, each independently comprises: (a) a glycine rich peptide; (b) a peptide comprising glycine and serine; (c) a peptide comprising (Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 234); (d) a peptide comprising (Gly-Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 232); (e) a peptide comprising (Gly-Gly-Gly-Gly-Ser)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 233); (f) a peptide comprising (Gly-Gly-Gly-Gly-Gln)n, wherein n is 1 , 2, 3, 4, 5, or 6 (SEQ ID NO: 235), or (g) a peptide comprising any one of SEQ ID NOs.221-235 and 504. E246. The bispecific molecule of any one of E170-E245, wherein said first linker, second linker, or third linker, each independently comprises: (a) GGGG (SEQ ID NO: 222); (b) GGGGSGGGGSGGGGS (SEQ ID NO: 230); (c) GGGGQGGGGQ (SEQ ID NO: 504); or (d) GGGGSGGGGS (SEQ ID NO: 229). E247. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 283, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 220. E248. A bispecific molecule comprising two copies of SEQ ID NO: 283, and two copies of SEQ ID NO: 220. (clone 11250) E249. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 287, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 206. E250. A bispecific molecule comprising two copies of SEQ ID NO: 287, and two copies of SEQ ID NO: 206. (clone 11252) E251. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 290, and (ii) two copies of an amino acid sequence
that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 184. E252. A bispecific molecule comprising two copies of SEQ ID NO: 290, and two copies of SEQ ID NO: 184. (clone 11253) E253. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 294, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 186. E254. A bispecific molecule comprising two copies of SEQ ID NO: 294, and two copies of SEQ ID NO: 186. (clone 11255). E255. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 297; (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 299; and (iii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 300. E256. A bispecific molecule comprising two copies of SEQ ID NO: 297, two copies of SEQ ID NO: 299, and two copies of SEQ ID NO: 300. (clone 11259) E257. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 304; (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 306; and (iii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 307. E258. A bispecific molecule comprising two copies of SEQ ID NO: 304, two copies of SEQ ID NO: 306, and two copies of SEQ ID NO: 307. (clone 11258) E259. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 311; (ii) two copies of an amino acid sequence that
is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 312; and (iii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 307. E260. A bispecific molecule comprising two copies of SEQ ID NO: 311, two copies of SEQ ID NO: 312, and two copies of SEQ ID NO: 307. (clone 11262) E261. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 315; (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 312; and (iii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 317. E262. A bispecific molecule comprising two copies of SEQ ID NO: 315, two copies of SEQ ID NO: 312, and two copies of SEQ ID NO: 317. (clone 11264) E263. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 320; (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 312; and (iii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 300. E264. A bispecific molecule comprising two copies of SEQ ID NO: 320, two copies of SEQ ID NO: 312, and two copies of SEQ ID NO: 300. (clone 11265) E265. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 443, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 444. E266. A bispecific molecule comprising two copies of SEQ ID NO: 443, and two copies of SEQ ID NO: 444. (clone 44988)
E267. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 434, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 435. E268. A bispecific molecule comprising two copies of SEQ ID NO: 434, and two copies of SEQ ID NO: 435. (clone 56039) E269. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 449, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 450. E270. A bispecific molecule comprising two copies of SEQ ID NO: 449, and two copies of SEQ ID NO: 450. (clone 56040) E271. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 451, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 452. E272. A bispecific molecule comprising two copies of SEQ ID NO: 451, and two copies of SEQ ID NO: 452. (clone 56041) E273. A bispecific molecule comprising (i) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 437, (ii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 438, (iii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 439, and (iv) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 440. E274. A bispecific molecule comprising: one copy of SEQ ID NO: 449, one copy of SEQ ID NO: 450, one copy of SEQ ID NO:439, and one copy of SEQ ID NO: 440. (clone 56042)
E275. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 432, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 433. E276. A bispecific molecule comprising two copies of SEQ ID NO: 432, and two copies of SEQ ID NO: 433. (clone 56132) E277. A bispecific molecule comprising (i) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 446, (ii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 447, and (iii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 448. E278. A bispecific molecule comprising: one copy of SEQ ID NO: 446, one copy of SEQ ID NO: 447, and one copy of SEQ ID NO: 448. (clone 56639) E279. A bispecific molecule comprising (i) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 429, (ii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 430, and (iii) an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 431. E280. A bispecific molecule comprising: one copy of SEQ ID NO: 429, one copy of SEQ ID NO: 430, and one copy of SEQ ID NO: 431. (clone 56761) E281. A bispecific molecule comprising (i) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 442, and (ii) two copies of an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical SEQ ID NO: 441. E282. A bispecific molecule comprising two copies of SEQ ID NO: 442, and two copies of SEQ ID NO: 441. (clone 56762)
E283. The bispecific molecule of any one of E99-E102 and E166-E282, wherein said bispecific molecule is a crosslinking dependent agonist of 4-1BB. E284. The bispecific molecule of any one of E99-E102 and E166-E282, wherein said bispecific molecule activates 4-1BB upon binding to PD-L1. E285. A nucleic acid comprising a nucleotide sequence encoding the 4-1BB antigen-binding protein of any one of E1-E98 and E319-E346, the PD-L1 antigen-binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166-E284, or one of the polypeptide chains of any one of the foregoing. E286. The nucleic acid of E285, comprising a nucleotide sequence that is at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical any one of SEQ ID NOs: 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465. E287. A vector comprising the nucleic acid of E285 or E286. E288. A host cell comprising the nucleic acid of E285 or E286, or the vector of E287. E289. The host cell of E288, wherein said host cell is a mammalian cell. E290. The host cell of E289, wherein said host cell is a CHO cell or a HEK-293 cell, or an Sp2.0 cell. E291. A kit comprising (i) the 4-1BB antigen-binding protein of any one of E1-E98 and E319-E346, the PD-L1 antigen-binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166-E284, the nucleic acid of E285 or E286, the vector of E287, the host cell of E288-E290, or a combination thereof; and (ii) instructions for use. E292. A pharmaceutical composition comprising (i) the 4-1BB antigen-binding protein of any one of E1- E98 and E319-E346, the PD-L1 antigen-binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166-E284, the nucleic acid of E285 or E286, the vector of E287, the host cell of E288-E290, or a combination thereof; and (ii) a pharmaceutically acceptable carrier, excipient, or diluent. E293. A method of making the 4-1BB antigen-binding protein of any one of E1-E98 and E319-E346, the PD-L1 antigen-binding protein of any one of E103-E165, or the bispecific molecule of any one of E99- E102 and E166-E284, comprising culturing the host cell of any one of E288-E290, under a condition
wherein the 4-1BB antigen-binding protein, the PD-L1 antigen-binding protein, or the bispecific molecule, is expressed. E294. The method of E293, further comprising harvesting the expressed the 4-1BB antigen-binding protein of any one of E1-E98 and E319-E346, the PD-L1 antigen-binding protein of any one of E103- E165, or the bispecific molecule of any one of E99-E102 and E166-E284. E295. A method of treating cancer, comprising administering to a subject in need thereof a therapeutically effective amount of the 4-1BB antigen-binding protein of any one of E1-E98 and E319- E346, the PD-L1 antigen-binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166-E284, or the pharmaceutical composition of E292. E296. The method of E295, wherein cancer is solid tumor. E297. The method of E295 or E296, wherein said subject is a human. E298. The method of any one of E295-E297, wherein said cancer comprises cells that express PD-L1. E299. The method of any one of E295-E298, wherein the cancer is brain cancer, bladder cancer, breast cancer, clear cell kidney cancer, cervical cancer, colon cancer, rectal cancer, endometrial cancer, gastric cancer, head/neck squamous cell carcinoma, lip cancer, oral cancer, liver cancer, lung squamous cell carcinoma, melanoma, mesothelioma, non-small-cell lung cancer (NSCLC), non-melanoma skin cancer, ovarian cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, sarcoma, small-cell lung cancer (SCLC), Squamous Cell Carcinoma of the Head and Neck (SCCHN), triple negative breast cancer, renal cell carcinoma, or thyroid cancer. E300. The method of any one of E295-E298, wherein the cancer is adrenocortical tumor, alveolar soft part sarcoma, carcinoma, chondrosarcoma, desmoid tumors, desmoplastic small round cell tumor, endocrine tumors, endodermal sinus tumor, epithelioid hemangioendothelioma, Ewing sarcoma, germ cell tumor, hepatoblastoma, hepatocellular carcinoma, melanoma, nephroma, neuroblastoma, non- rhabdomyosarcoma soft tissue sarcoma (NRSTS), osteosarcoma, paraspinal sarcoma, retinoblastoma, rhabdomyosarcoma, synovial sarcoma, or Wilms tumor. E301. The method of any one of E295-E298, wherein the cancer is acute lymphoblastic leukemia (ALL), acute myeloid leukemia (AML), chronic lymphocytic leukemia (CLL), or chronic myeloid leukemia (CML). E302. The method of any one of E295-E298, wherein the cancer is diffuse large B-cell lymphoma (DLBCL), follicular lymphoma, Hodgkin's lymphoma (HL), mantle cell lymphoma (MCL), multiple myeloma (MM), myelodysplastic syndrome (MDS), non-Hodgkin's lymphoma (NHL), or small lymphocytic lymphoma (SLL).
E303. The method of any one of E295-E302, wherein said 4-1BB antigen-binding protein, PD-L1 antigen-binding protein, bispecific molecule, or pharmaceutical composition is administered intravenously. E304. The method of any one of E295-E302, wherein said 4-1BB antigen-binding protein, PD-L1 antigen-binding protein, bispecific molecule, or pharmaceutical composition is administered subcutaneously. E305. The method of any one of E295-E302, wherein said 4-1BB antigen-binding protein, PD-L1 antigen-binding protein, bispecific molecule, fusion protein, polypeptide, or pharmaceutical composition is administered about twice a week, once a week, once every two weeks, once every three weeks, once every four weeks, once every five weeks, once every six weeks, once every seven weeks, once every eight weeks, once every nine weeks, once every ten weeks, twice a month, once a month, once every two months, once every three months, or once every four months. E306. The 4-1BB antigen-binding protein of any one of E1-E98 and E319-E346, the PD-L1 antigen- binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166- E284, or the pharmaceutical composition of E292 for use as a medicament. E307. The 4-1BB antigen-binding protein of any one of E1-E98 and E319-E346, the PD-L1 antigen- binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166- E284, or the pharmaceutical composition of E292 for use in treating cancer in a subject. E308. Use of the 4-1BB antigen-binding protein of any one of E1-E98 and E319-E346, the PD-L1 antigen-binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166-E284, or the pharmaceutical composition of E292 in the manufacture of a medicament for treating cancer in a subject. E309. Use of the 4-1BB antigen-binding protein of any one of E1-E98 and E319-E346, the PD-L1 antigen-binding protein of any one of E103-E165, the bispecific molecule of any one of E99-E102 and E166-E284, or the pharmaceutical composition of E292 for treating cancer in a subject. E310. A method of assessing immunogenicity of a bispecific molecule, wherein said immunogenicity is attributed to a T-cell epitope, and wherein said bispecific molecule comprises two domains: (1) a first domain that binds to a Dendritic Cell (DC) surface antigen; and (2) a second domain that binds to a T cell co-stimulatory molecule, the method comprises: (a) obtaining a first protein that comprises said first domain but does not comprise said second domain; (b) obtaining a second protein that comprises said second domain but does not comprise said first domain;
(c) incubating said first protein and second protein with a cell culture that comprises DCs and T cells; and (d) assessing the activation or proliferation of T cells, wherein the activation or proliferation of T cells is indicative that said bispecific molecule comprises an immunogenic T cell epitope. E311. The method of E310, wherein said DC surface antigen is PD-L1. E312. The method of E311, wherein said DC surface antigen is CD8A, CLEC9A, ITGAE, ITGAX, THBD (CD141), XCR, CD1C, CD207, ITGAM, NOTCH2, SIRPA, CLEC4C, LILRB4, NRP1, CCR7, CD14, MRC1 (CD206), CD209, or CD1A. E313. The method of any one of E310-E312, wherein said T-cell co-stimulatory molecule is CD28, Inducible Co-Stimulator (ICOS), CTLA4 (Cytotoxic T-Lymphocyte-Associated protein 4), 4-1BB (also known as CD137), OX40 (also known as CD134), CD27, CD30, DR3, Glucocorticoid-Induced TNFR family Related (GITR), or Herpes Virus Entry Mediator (HVEM). E314. The method of any one of E310-E313, wherein said T-cell co-stimulatory molecule is 4-1BB. E315. The method of any one of E310-E314, wherein said first protein further comprise an IgG Fc domain. E316. The method of any one of E310-E315, wherein said second protein further comprise an IgG Fc domain. E317. The method of E315 or E316, wherein said Fc domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, or 426. E318. The method of E315 or E316, wherein said Fc domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 263, 267, or 483. E319. A 4-1BB antigen-binding protein, comprising a heavy chain variable domain (VH) and does not comprise light chain variable domain (VL), wherein said protein binds to an epitope that comprises residues C102, V71, and Q104, according to the numbering of SEQ ID NO:272.
E320. The 4-1BB antigen-binding protein of E319, wherein said epitope further comprises residue P90, according to the numbering of SEQ ID NO:272. E321. The 4-1BB antigen-binding protein of E319 or E320, wherein said epitope further comprises one or more residues selected from the group consisting of: K69, T89, F92, M101, and L112, according to the numbering of SEQ ID NO:272. E322. The 4-1BB antigen-binding protein of any one of E319-E321, wherein said epitope further comprises residues K69, T89, F92, M101, and L112, according to the numbering of SEQ ID NO:272. E323. The 4-1BB antigen-binding protein of any one of E319-E322, wherein one or more of the following substitutions substantially disrupts the binding of said antigen-binding protein to said epitope: (1) C102 is replaced by A; (2) Q104 is replaced by A; or (3) K69 is replaced by A. E324. The 4-1BB antigen-binding protein of any one of E319-E313, wherein said antigen-binding protein binds to said epitope with a binding affinity (KD) value that is at least 100-fold less, at least 200-fold less, at least 300-fold less, at least 400-fold less, at least 500-fold less, at least 600-fold less, at least 700-fold less, at least 800-fold less, at least 900-fold less, or at least 1000-fold less, than its KD value for an epitope comprising one or more of the following substitutions: (1) C102 is replaced by A; (2) Q104 is replaced by A; or (3) K69 is replaced by A. E325. The 4-1BB antigen-binding protein of E323 or E324, wherein said KD value is measured by surface plasmon resonance (SPR), optionally using a Biacore T200 instrument. E326. The 4-1BB antigen-binding protein of E323 or E324, wherein said KD value is measured by bio- layer interferometry (BLI), optionally using a ForteBio Octet instrument. E327. The 4-1BB antigen-binding protein of any one of E319-E326, wherein said 4-1BB is a human 4- 1BB. E328. The 4-1BB antigen-binding protein of E327, wherein said 4-1BB comprises SEQ ID NO:566. E329. The 4-1BB antigen-binding protein of any one of E319-E328, wherein said epitope is determined by X-ray crystallography or cryoEM. E330. The 4-1BB antigen-binding protein of any one of E319-E329, wherein said VH comprises (VH numbering according to Kabat): (1) H45 is Leu, Phe, Ile, or Tyr; (2) H47 is Trp, Phe, Leu, or Tyr; (3) H100B is Tyr, Arg, His, Lys, or Met; (4) H100D is Thr, Ala, Asn, Cys, Gln, Lys, Met, or Val; and (5) H100F is Phe, Trp or Tyr.
E331. The 4-1BB antigen-binding protein of any one of E319-E330, wherein said VH comprises (VH numbering according to Kabat): (1) H45 is Leu or Phe; (2) H47 is Trp or Leu; (3) H100B is Tyr or Met; (4) H100D is Thr or Val; and (5) H100F is Phe or Trp. E332. The 4-1BB antigen-binding protein of any one of E319-E331, wherein said VH comprises (VH numbering according to Kabat): (1) H45 is Leu; (2) H47 is Trp; (3) H100B is Tyr; (4) H100D is Thr; and (5) H100F is Phe. E333. The 4-1BB antigen-binding protein of any one of E319-E332, wherein said VH comprises (VH numbering according to Kabat): (6) H97 is Ser, Arg, Asn, Gln, Glu, His,,Leu, Lys, Met, Phe, Thr, Trp, Tyr, or Val; (7) H100E is Ser, Ala, Asn, Asp, Cys, His, Trp, Tyr, or Val; and (9) H102 is Tyr, Ile, Lys, or Val. E334. The 4-1BB antigen-binding protein of any one of E319-E333, wherein said VH comprises (VH numbering according to Kabat): (6) H97 is Ser or Leu; (7) H100E is Ser or Val; and (9) H102 is Tyr or Lys. E335. The 4-1BB antigen-binding protein of any one of E319-E334, wherein said VH comprises (VH numbering according to Kabat): (6) H97 is Ser; (7) H100E is Ser; and (9) H102 is Tyr. E336. The 4-1BB antigen-binding protein of any one of E319-E335, wherein: (a) said VH comprises: (i) a CDR-H3 comprising any one of SEQ ID NOs:576-578 and 580; and (ii) framework residue H45 is Leu and framework residue H47 is Trp; or (b) said VH comprise: (i) a CDR-H2 comprising SEQ ID NO:574, (ii) a CDR-H3 comprising SEQ ID NO: 579; and (iii) framework residue H45 is Leu. E337. The 4-1BB antigen-binding protein of E336, wherein said CDR-H3 and framework residues contact one or more 4-1BB residues selected from the group consisting of: C102, V71, Q104, P90, K69, T89, F92, M101, and L112 (numbering according to SEQ ID NO:272). E338. The 4-1BB antigen-binding protein of E337, wherein said contacting is defined as within 4.5Å distance between a heavy atom in 4-1BB and a heavy atom in the 4-1BB antigen-binding protein, as determined by X-ray crystallography or cyroEM. E339. The 4-1BB antigen-binding protein of any one of E319-E338, comprising a VH framework derived from a human germline VH1, VH2, VH3, VH4, or VH5 framework sequence. E340. The 4-1BB antigen-binding protein of any one of E310-E338, comprising a VH framework derived from a human germline VH1 framework sequence. E341. The 4-1BB antigen-binding protein of any one of E319-E338, comprising a VH framework derived from a human germline VH3 framework sequence.
E342. The 4-1BB antigen-binding protein of any one of E319-E338, comprising a VH framework derived from a human germline VH4 framework sequence. E343. The 4-1BB antigen-binding protein of any one of E319-E338, comprising a VH framework derived from a human germline VH2 framework sequence. E344. The 4-1BB antigen-binding protein of any one of E319-E338, comprising a VH framework derived from a human germline VH5 framework sequence. E345. The 4-1BB antigen-binding protein of any one of E319-E344, comprising a VH framework sequence is at least 90% identical to the human germline framework sequence from which it is derived. E346. The 4-1BB antigen-binding protein of any one of E319-E345, comprising a VH framework sequence is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to the human germline framework sequence from which it is derived. BRIEF DESCRIPTION OF THE DRAWINGS [12] FIG.1A is a schematic of the steps taken to generate 4-1BB mAbs and PD-L1 mAbs of the present disclosure. FIG.1B is a schematic of the steps taken to generate bispecific molecules comprising a 4-1BB binding protein and PD-L1 binding protein of the present disclosure. [13] FIG.2A is an illustration of a bispecific molecule comprising an IgG moiety and a scFv moiety. FIG. 2B is an illustration of a bispecific molecule comprising an IgG moiety and a Fab moiety. [14] FIG.3 is a table listing engineered binding proteins and characteristics thereof. [15] FIG.4 is a table listing bispecific molecules generated herein and characteristics thereof. [16] FIG.5 is a schematic representation of bispecific molecules of interest (dotted lines showing where the IdeS enzyme cleaves), and 2 representative LC-MS spectra of the bispecific molecules. Representative graphs from clones 11259 and 11262 are shown. All IgG-Fabs (11258, 11259, 11262, 11264, and 11265) have been confirmed to have correct HC-LC Pairs. [17] FIG.6A is a graph of the IL-2 produced by T-cells activated by artificial APCs plotted as a function of bispecific molecule concentration. FIG.6B is a graph of the IL-2 produced by T-cells activated by artificial APCs plotted as a function of bispecific molecule concentration. [18] FIGs.7A-7D are graphs of the tumor volume plotted as a function of time following tumor implant for mice treated with the active agent indicated at the top of each graph.
[19] FIG.8A is a graph of the tumor volume plotted as a function of time following tumor implant for mice treated with the active agents indicated in the key. FIG.8B is a graph of the percent survival plotted as a function of time following tumor implant for mice treated with the active agents indicated in the key. [20] FIG.9 is a table listing characteristics of 4-1BB antigen-binding proteins described herein. [21] FIGs.10A-10B are graphs showing crosslinking dependent activity of 4-1BB UniAbs compared to crosslinker independent control. [22] FIGs.11A-11D are graphs showing cell binding dose curves for anti-41BB UniAbs using activated T-cells, CHO-human 4-1BB, CHO cyno 4-1BB, off target cell lines compared to a crosslinker independent control. [23] FIG.12 a schematic of the steps taken to generate bispecific molecules comprising a 4-1BB binding protein and PD-L1 binding protein of the present disclosure. [24] FIGs.13A-13K are illustrations of bispecific molecule format used in second round of screening. Letters “a” and “b” are used to indicate the first antigen (“a”) or the second antigen (“b”) that the bispecific molecule binds to. Letters “N” means that the second antigen-binding moiety is fused at the N-terminus of the first antigen-binding molecule, and the number refers to the number of antigen-binding moieties. N2 = two “b” antigen-binding moieties fused at the N-termini of the “a’ antigen-binding molecule. “B” means the second antigen-binding moiety is fused at the hinge region, and “C” means the second antigen-binding moiety is fused at the C-terminus. [25] FIGs.14A-14D provides domain information of certain bispecific formats, including IgG-scFv, Fab- scFv-Fc, IgG-VH, and Fab-VH-Fc. [26] FIGs.15A-15B summarizes the biophysical properties of the bispecific molecules disclosed herein. [27] FIGs.16A-16C show the potencies of the bispecific molecules disclosed herein. [28] FIG.17 shows that the activation of 4-1BB by the bispecific molecule is dependent on the binding of PDL1. [29] FIGs.18A-18B show the pre-ADA assessment of the bispecific molecules disclosed herein. [30] FIGs.19A-19B shows the immunogenicity assessment of the bispecific molecules disclosed herein. [31] FIG.20A shows the overall structure of human 4-1BB and molecule 380984. The crystal structure of human 4-1BB in complex with 380984 is shown as cartoon.4-1BB is in light gray color, and Amgen- 380984 is in dark gray color. The N-, C- terminus, and CDR-3 are labeled. The residues engaged in
ligand 4-1BBL binding are marked as sphere. FIG.20B shows the close-up view of the contact between human 4-1BB and 380984. Human 4-1BB (colored in light gray) and Amgen-380984 (colored in dark gray) are illustrated as transparent cartoon. All interacting residues are shown as sticks; the back-bone nitrogen atoms are shown as sphere. Hydrogen bonds are represented as gray dashed lines. [32] FIGs 21A-21G are graphs showing the comparison of 4-1BB binders and their potential orientation on cell membrane. The structure of human 4-1BB is shown as surface in color light gray; its binders are shown as cartoon in color dark gray. (A), Three human 4-1BB receptors bind to a trimeric ligand 4-1BBL, each protomer of human 4-1BB are labeled as number 1, 2, 3. PDB: 6mgp. (B). Protomer 2 of the 4-1BB- 4-1BBL complex is shown in the same orientation as in A. (C). The structure of human 4-1BB and Urelumab complex (BMS, PDB:6MHR) is superimposed to the protomer 2 of 4-1BB-4-BBL complex by aligning on the CRDs 3 and 4. The location of Fc is labeled. (D). Protomer 1 of the 4-1BB-4-1BBL complex is shown in the same orientation as in A. (E). The structure of human 4-1BB and 380984 is superimposed to the protomer 1 of 4-1BB-4-BBL complex by aligning the CRDs 3 and 4. The location of Fc is labeled. (F). The structure of human 4-1BB and Utomilumac (Pfizer, PDB: 6MI2) is superimposed to the promotor by aligning the CRDs 3 and 4. The location of Fc are labeled. (G). The structure of Utomilumac and 380984 are superimposed to the same 4-1BB protomer. The sketch models of the full antibody in complex with its binders are shown on the left of each structure. The relative location of cell membrane is also shown at the bottom of each structure. DETAILED DESCRIPTION 1. Overview [33] 4-1BB is an immune co-stimulatory protein expressed on activated T cells.4-1BB agonist antibodies have demonstrated efficacy in prophylactic and therapeutic settings in both monotherapy and combination therapy tumor models, and have established protective & durable anti-tumor T-cell memory responses. Agonizing 4-1BB by crosslinking antibody enhances T cell proliferation, survival and cytokine production upon TCR engagement. However, clinical development of 4-1BB agonistic antibodies has been hampered by dose-limiting hepatotoxicity. For example, phase I and II data from Urelumab (BMS- 663513) revealed a liver toxicity that appeared to be on target and dose dependent, halting initial clinical development of Urelumab. The anti-hu4-1BB huIgG2 utomilumab (PF-05082566) displays a better safety profile but lower agonistic potency. [34] Seeking to overcome this challenge, the inventors designed bispecific molecules to promote target- mediated clustering of 4-1BB. It is understood that 4-1BB undergoes trimerization upon binding to its ligand (4-1BBL); and 4-1BB multimerization and clustering (“crosslinking”) is a prerequisite for its signaling pathway. Therefore, the 4-1BB antibodies were specifically selected as “crosslinking dependent,” meaning that the agonistic activity of the antibody is dependent upon the crosslinking of 4-
1BB. Without 4-1BB crosslinking, the binding of the antibody to 4-1BB leads to minimal agonist activity, thereby avoiding toxicities exhibited by Urelumab. Urelumab is believed to be a crosslinking independent agonist antibody, meaning the binding of Urelumab to 4-1BB is sufficient to trigger 4-1BB activation. The bispecific molecules disclosed herein take advantage of this crosslinking dependent effect, and the activation of 4-1BB is controlled through a PD-L1 binding moiety, which its expression further limits where the crosslinking events can be happening. [35] PD-L1, which is also known as B7-H1 or CD274, is expressed in multiple types of cancers (e.g., breast cancer, lung cancer, melanoma). In the tumor microenvironment, PD-L1, expressed on tumor cells, binds to the inhibitory checkpoint receptor, PD-1, on activated tumor infiltrating lymphocytes. The interaction between PD-L1 and PD-1 delivers an inhibitory signal to T cells and ultimately prevents tumor elimination. [36] Presented herein are data demonstrating simultaneous activation of the 4-1BB co-stimulatory pathway and blockade of the PD-L1/PD-1 inhibitory pathway upon administration of bispecific molecules comprising a 4-1BB binding protein and a PD-L1 binding protein. The data also demonstrate the dual activity upon administration of a combination of a monospecific 4-1BB binding protein and a monospecific PD-L1 binding protein. Without being bound to a particular theory, it is believed that (i) agonizing 4-1BB enhances T-cell proliferation, survival & cytokine production upon TCR engagement; and (ii) simultaneously blocking PD-L1 and agonizing 4-1BB can have synergistic anti-tumor activities. [37] In addition, the bispecific molecules described herein address the dose-limiting hepatotoxicity of systemic 4-1BB agonism, because 4-1BB activation is crosslinking dependent. For normal tissues that do not express PD-L1 or express low levels of PD-L1 (such as hepatic cell), minimal crosslinking of 4-1BB will occur, and 4-1BB activation will be limited. In contrast, in cancer cells that express high level of PD- L1, through PD-L1-binding, the bispecific molecules are brought to the proximity of each other, thus promoting 4-1BB crosslinking and T-cell co-stimulation. Systemic toxicities should be limited because 4- 1BB activation will be largely confined to tumors that express high level of PD-L1. [38] Accordingly, the present disclosure provides bispecific molecules comprising a 4-1BB binding protein and a PD-L1 binding protein. Related antigen binding proteins, including, for instance, monospecific 4-1BB binding proteins and monospecific PD-L1 binding proteins, are further provided. 2. Antigen Binding Proteins 2.1 Antigen Binding Protein Types [39] The antigen-binding proteins of the present disclosure can take any one of many forms of antigen- binding proteins known in the art. In exemplary aspects, the antigen-binding protein is an antibody or
immunoglobulin, or an antigen binding fragment of an antibody or immunoglobulin, or an antibody protein product. [40] Collectively, antibodies form a family of plasma proteins known as immunoglobulins and comprise of immunoglobulin domains. (Janeway et al., Immunobiology: The Immune System in Health and Disease, 4th ed., Elsevier Science Ltd./Garland Publishing, 1999). As used herein, the term “antibody” refers to a protein having a conventional immunoglobulin format, comprising heavy and light chains, and comprising variable and constant regions. For example, an antibody may be an IgG which is a “Y-shaped” structure of two identical pairs of polypeptide chains, each pair having one “light” (typically having a molecular weight of about 25 kDa) and one “heavy” chain (typically having a molecular weight of about 50-70 kDa). An antibody has a variable region and a constant region. In IgG formats, the variable region is generally about 100-110 or more amino acids, comprises three complementarity determining regions (CDRs), is primarily responsible for antigen recognition, and substantially varies among other antibodies that bind to different antigens. The constant region allows the antibody to recruit cells and molecules of the immune system. The variable region is made of the N-terminal regions of each light chain and heavy chain, while the constant region is made of the C-terminal portions of each of the heavy and light chains. (Janeway et al., “Structure of the Antibody Molecule and the Immunoglobulin Genes”, Immunobiology: The Immune System in Health and Disease, 4th ed. Elsevier Science Ltd./Garland Publishing, (1999)). [41] Antibodies can comprise any constant region known in the art. Human light chains are classified as kappa and lambda light chains. Heavy chains are classified as mu, delta, gamma, alpha, or epsilon, and define the antibody's isotype as IgM, IgD, IgG, IgA, and IgE, respectively. IgG has several subclasses, including, but not limited to IgG1, IgG2, IgG3, and IgG4. IgM has subclasses, including, but not limited to, IgM1 and IgM2. Embodiments of the present disclosure include all such classes or isotypes of antibodies. The light chain constant region can be, for example, a kappa- or lambda-type light chain constant region, e.g., a human kappa- or lambda-type light chain constant region. The heavy chain constant region can be, for example, an alpha-, delta-, epsilon-, gamma-, or mu-type heavy chain constant regions, e.g., a human alpha-, delta-, epsilon-, gamma-, or mu-type heavy chain constant region. Accordingly, in exemplary embodiments, the antibody is an antibody of isotype IgA, IgD, IgE, IgG, or IgM, including any one of IgG1, IgG2, IgG3 or IgG4. [42] The antibody can be a monoclonal antibody or a polyclonal antibody. In some embodiments, the antibody comprises a sequence that is substantially similar to a naturally-occurring antibody produced by a mammal, e.g., mouse, rabbit, goat, horse, chicken, hamster, human, and the like. In this regard, the antibody can be considered as a mammalian antibody, e.g., a mouse antibody, rabbit antibody, goat antibody, horse antibody, chicken antibody, hamster antibody, human antibody, and the like. In certain aspects, the antibody is a human antibody. In certain aspects, the antibody is a chimeric antibody or a humanized antibody. The term "chimeric antibody" refers to an antibody containing domains from two or more different antibodies. A chimeric antibody can, for example, contain the constant domains from one
species and the variable domains from a second, or more generally, can contain stretches of amino acid sequence from at least two species. A chimeric antibody also can contain domains of two or more different antibodies within the same species. The term "humanized" when used in relation to antibodies refers to antibodies having at least CDR regions from a non-human source which are engineered to have a structure and immunological function more similar to true human antibodies than the original source antibodies. For example, humanizing can involve grafting a CDR from a non-human antibody, such as a mouse antibody, into a human antibody. Humanizing also can involve select amino acid substitutions to make a non-human sequence more similar to a human sequence. [43] An antibody can be cleaved into fragments by enzymes, such as, e.g., papain and pepsin. Papain cleaves an antibody to produce two Fab fragments and a single Fc fragment. Pepsin cleaves an antibody to produce a F(ab’)2 fragment and a pFc’ fragment. In exemplary aspects of the present disclosure, the antigen binding protein of the present disclosure comprises an antigen binding fragment of an antibody. As used herein, “antigen binding fragment” of an antibody refers to a portion of an antibody molecule that retains the ability to specifically bind to an antigen (preferably with substantially the same binding affinity). Examples of an antigen-binding fragment include but not limited to: (i) a Fab fragment, a monovalent fragment consisting of the VL, VH, CL and CH1 domains; (ii) a F(ab')2 fragment, a bivalent fragment comprising two Fab fragments linked by a disulfide bridge at the hinge region; (iii) a Fd fragment consisting of the VH and CH1 domains; (iv) a Fv fragment consisting of the VL and VH domains of a single arm of an antibody, (v) a dAb fragment (Ward et al., 1989 Nature 341 :544-546), which consists of a VH domain. [44] The architecture of antibodies has been exploited to create a growing range of alternative formats that span a molecular-weight range of at least about 12–150 kDa and has a valency (n) range from monomeric (n = 1), to dimeric (n = 2), to trimeric (n = 3), to tetrameric (n = 4), and potentially higher; such alternative formats are referred to herein as “antibody protein products.” [45] The building block that is most frequently used to create novel antibody-based formats is the single-chain variable (V)-domain antibody fragment (scFv). Although the two domains of the Fv fragment, VL and VH, are coded for by separate genes, they can be joined, using recombinant methods, by a synthetic linker that enables them to be made as a single protein chain in which the VL and VH regions pair to form monovalent molecules (see e.g., Bird et al. Science 242:423- 426 (1988) and Huston et al., 1988, Proc. Natl. Acad. Sci. USA 85:5879-5883). [46] Other forms of single chain antibody protein products, such as diabodies are also encompassed. Diabodies are bivalent, bispecific antibody protein products in which VH and VL domains are expressed on a single polypeptide chain, but using a linker that is too short to allow for pairing between the two domains on the same chain, thereby forcing the domains to pair with complementary domains of another
chain and creating two antigen-binding sites (see e.g., Holliger et al, 1993, Proc. Natl. Acad. Sci. USA 90:6444-6448; Poljak et al., 1994, Structure 2:1121 -1123). [47] Other antibody protein products include disulfide-bond stabilized scFv (ds-scFv), single chain Fab (scFab), as well as di- and multimeric antibody formats like dia-, tria- and tetra-bodies, or minibodies (miniAbs) that comprise different formats consisting of scFvs linked to oligomerization domains. The smallest fragments are VHH/VH of camelid heavy chain Abs as well as single domain Abs (sdAb). A peptibody or peptide-Fc fusion is yet another antibody protein product. The structure of a peptibody consists of a biologically active peptide grafted onto an Fc domain. Peptibodies are well-described in the art. See, e.g., Shimamoto et al., mAbs 4(5): 586-591 (2012). [48] Antigen-binding proteins disclosed herein also include “heavy chain only” molecules. This type of antigen-binding proteins lack the light chain of a conventional antibody. Examples of such “heavy chain only” molecules include, for example, single domain molecules such as UniDabs® (VH only); and homodimeric molecules comprising the VH antigen-binding domain and the CH2 and CH3 constant domains, in the absence of the CH1 domain (e.g., UniAb®). Non-limiting examples of “heavy chain only” antigen binding proteins are described, for example, in WO2018/039180. Such heavy chain only molecules can be produced by UniRat®, which is a triple knockout rat wherein the expressions of the native variable coding sequences and the heavy and light chain constant regions have been inactivated. The UniRat has been genetically modified to exclusively express the full human VDJ repertoire (all VH families), with transgenes of human heavy chain variable domains linked to a conserved rat Fc. Immunization of the UniRat elicits a normal antibody response that results in the expression of UniAbs, human heavy-chain-only antibodies of approximately 80 kDa, contrasting with the standard ~150 kDa human IgG. In particular, VH domains from the UniRat, called UniDab, approximately 12.5 kDa (~100 amino acids), can be assembled as modular domains of multispecific antigen binding proteins. [49] Bispecific formats can generally be divided into five major classes: BsIgG, appended IgG, BsAb fragments, bispecific fusion proteins and BsAb conjugates. See, e.g., Spiess et al., Molecular Immunology 67(2) Part A: 97-106 (2015). [50] In exemplary aspects, the antigen binding protein of the present disclosure comprises any one of these antibody protein products. In exemplary aspects, the antigen binding protein of the present disclosure comprises any one of an scFv, Fab, VHH, VH, Fv fragment, ds-scFv, scFab, UniDab, dimeric antibody, multimeric antibody (e.g., a diabody, triabody, tetrabody), miniAb, peptibody VHH/VH of camelid heavy chain antibody, sdAb, diabody; UniAb; a triabody; a tetrabody; a bispecific or trispecific antibody, BsIgG, appended IgG, BsAb fragment, bispecific fusion protein, and BsAb conjugate. [51] In exemplary instances, the antigen binding protein of the present disclosure comprises an antibody protein product in monomeric form, or polymeric, oligomeric, or multimeric form. In certain
embodiments in which the antibody comprises two or more distinct antigen binding regions fragments, the antibody is considered bispecific, trispecific, or multi-specific, or bivalent, trivalent, or multivalent, depending on the number of distinct epitopes that are recognized and bound by the antibody. [52] Many of the antigen-binding proteins disclosed herein comprise two different chains, one derived from the heavy chain of an antibody, and one derived from the light chain of an antibody. Although the heavy/light chain has been modified and is no longer the classical immunoglobulin heavy/light chain, for convenience, it is still generally called “heavy chain” or “HC” if it is based on heavy chain backbone, and “light chain” or “LC” if it is based on light chain backbone. For example, for tetravalent bispecific molecule IgG-scFv, the “HC” comprises an IgG heavy chain fused to an scFv. It would be apparent to a skilled artisan whether HC is a traditional immunoglobulin heavy chain or a modified version based on immunoglobulin heavy chain backbone. 2.2 Binding Location [53] Some exemplary antigen binding proteins disclosed herein are characterized by the location or epitopes they bind to, or by the paratopes that they comprise. An “epitope” refers to the area or region of an antigen to which an antigen binding protein specifically binds, e.g., an area or region comprising residues that interacts with the antigen-binding protein. Epitopes can be linear or conformational. Epitopes can be determined by any method well known in the art. For example, epitopes can be determined by conventional immunoassays. Alternatively, one may competitively screen antigen-binding proteins for binding to the same epitope. An approach to achieve this is to conduct competition and cross- competition studies to find antigen-binding proteins that compete or cross-compete with one another for binding to an antigen (such as 4-1BB). [54] The term “paratope” is derived from the above definition of “epitope” by reversing the perspective, and refers to the area or region of an antigen-binding protein which is involved in binding of an antigen, e.g., an area or region comprising residues that interacts with the antigen. A paratope may be linear or conformational (such as discontinuous residues in CDRs). [55] The epitope/paratope can be defined and characterized at different levels of detail using a variety of experimental and computational epitope mapping methods. The experimental methods include mutagenesis, X-ray crystallography, Nuclear Magnetic Resonance (NMR) spectroscopy, Hydrogen/deuterium exchange Mass Spectrometry (HX-MS), cryo-EM, and various competition binding methods. As each method relies on a unique principle, the description of an epitope is linked to the method by which it has been determined. Thus, the epitope/paratope for a given binding pair will be defined differently depending on the mapping method employed.
[56] At its most detailed level, the epitope/paratope for the interaction between the antigen and the antigen-binding protein can be defined by the spatial coordinates defining the atomic contacts present in the interaction, as well as information about their relative contributions to the binding thermodynamics. At one level, an epitope/paratope residue can be characterized by the spatial coordinates defining the atomic contacts between the binding pair. In one aspect, the epitope/paratope residue can be defined by a specific criterion, e.g., distance between atoms in the antigen and the antigen-binding protein (e.g., a distance of equal to or less than 4.5 Å from a heavy atom of the antigen and a heavy atom of the antigen- binding protein ("contact" residues)). In another aspect, an epitope/paratope residue can be characterized as participating in a hydrogen bond interaction with the cognate antibody/antigen, or with a water molecule that is also hydrogen bonded to the antigen/antigen-binding protein (water-mediated hydrogen bonding). In another aspect, an epitope/paratope residue can be characterized as forming a salt bridge with a residue of the cognate antibody/antigen. In yet another aspect, an epitope/paratope residue can be characterized as a residue having a non-zero change in buried surface area (BSA) due to the interaction between the antigen and the antigen-binding protein. [57] At a further less detailed level, epitope/paratope can be characterized through function, e.g., by competition binding with other antigen-binding molecules. The epitope/paratope can also be defined more generically as comprising amino acid residues for which substitution by another amino acid will alter the characteristics of the interaction between the binding pair (e.g., alanine scanning). [58] In the context of an X-ray derived crystal structure or cryo-EM structure, as exemplified herein with respect to 4-1BB antigen-binding proteins, unless otherwise specified, a 4-1BB epitope residue refers to a 4-1BB residue: (i) having a heavy atom (i.e., a non-hydrogen atom) that is within a distance of 4.5 Å from a heavy atom of the antigen-binding protein (also called “contact” residues); (ii) participating in a hydrogen bond with a residue of the antigen-binding protein, or with a water molecule that is also hydrogen bonded to the antigen-binding protein (water- mediated hydrogen bonding), (iii) participating in a salt bridge to a residue of the antigen-binding protein, and/or (iv) having a non-zero change in buried surface area (BSA) due to interaction with the antigen-binding protein. In general, a cutoff is imposed for BSA to avoid inclusion of residues that have minimal interactions. Therefore, unless otherwise specified, epitope residues under category (iv) are selected if it has a BSA of 20 Å2 or greater, or is involved in electrostatic interactions when the antigen-binding protein binds to 4-1BB. Similarly, in the context of an X-ray derived crystal structure or cryo-EM structure, unless otherwise specified or contradicted by context, a paratope residue, refers to an antigen-binding protein residue (i) having a heavy atom (i.e., a non-hydrogen atom) that is within a distance of 4.5 Å from a heavy atom of 4-1BB (also called “contact” residues), (ii) participating in a hydrogen bond with an antigen residue, or with a water molecule that is also hydrogen bonded to 4-1BB (water-mediated hydrogen bonding), (iii) participating in a salt bridge to a residue of 4-1BB, and/or (iv) having a non-zero change in buried surface area due to interaction with 4- 1BB. Again, unless otherwise specified, paratope residues under category (iv) are selected if it has a BSA
of 20 Å2 or greater, or is involved in electrostatic interactions when the antigen-binding protein binds to 4- 1BB. [59] Dependent on the epitope mapping method used, and obtained at different levels of detail, it follows that comparison of epitopes for different antigen-binding protein on the same antigen can similarly be conducted at different levels of detail. For example, epitopes described on the amino acid level, e.g., determined from an X-ray or cryo-EM structure, are said to be identical if they contain the same set of amino acid residues. Epitopes are said to be separate (unique) if no amino acid residue is shared by the epitopes. Epitopes characterized by competition binding are said to be overlapping if the binding of two molecules are mutually exclusive, i.e., binding of one molecule excludes simultaneous or consecutive binding of the other molecule; and epitopes are said to be separate (unique) if the antigen is able to accommodate binding of both molecules simultaneously. [60] 4-1BB is a glycosylated type I membrane protein comprises four cysteine-rich pseudo repeats (CRDs) forming the extracellular domain, a short helical transmembrane domain, and a cytoplasmic signaling domain. Human 4-1BB (SEQ ID NO: 272; UniProtKB - Q07011) comprises 255 amino acid residues: residues 1-23 forms signal peptide that is cleaved before mature protein is produced, residues 24 – 186 forms extracellular domain, residues 187 – 213 forms transmembrane helical structure, and residues 214 – 255 forms cytoplasmic domain. The four CRD domains of human 4-1BB are defined as follows: CRD1: residues 24-45; CDR2, residues 47-86; CRD3: residues 87-118, and CRD4: residues 119- 159. [61] In certain aspects, the 4-1BB antigen-binding proteins described herein bind to an epitope in CRD1 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to an epitope in CRD2 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to an epitope in CRD3 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to an epitope in CRD4 of 4-1BB. [62] In certain aspects, the 4-1BB antigen-binding proteins described herein bind to CRD1 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to CRD2 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to CRD3 of 4-1BB. In certain aspects, the 4-1BB antigen-binding proteins described herein bind to CRD4 of 4-1BB. [63] In particular, as exemplified in Example 14, certain epitope residues within CRD3 were found to be particularly advantageous. Although not wishing to be bound by a particular theory, FIGs.21A-21G illustrate the advantageous of the binding location as exemplified in Example 14. As shown in FIGs.21C, Urelumab binds to CRD1, and is orientated away from cell surface. Because Urelumab does not compete with 4-1BBL for binding, it is believed that simultaneous binding of Urelumab and 4-1BBL may cause synergistic or additive effect, resulting in undesired activation of 4-1BB pathway in normal tissues. This
may help explain the observation that Urelumab activates 4-1BB pathway in hepatic cells, resulting in liver toxicity. FIGs.21F and 21G show the binding of Utomilumab to 4-1BB. Utomilumab binds to the CDR4 domain. The antibody is close to cell surface and oriented parallel to the cell surface. This orientation results in a large distance between 4-1BB monomers, making 4-1BB crosslinking more difficult (Fig.21F). That may explain the weak activity of Utomilumab. Molecule 380984, on the other hand, binds to CDR3 and is orientated vertically to the cell surface, which is much more desirable than parallel orientation. Compared with Utomilumab, the distance between 4-1BB monomers is much smaller due to its vertical orientation (Fig.21E). [64] The term “bind” here refers to specific binding, a term well understood in the art, and methods to determine such specific binding are also well known in the art. A molecule is said to exhibit “binding” if it reacts or associates more frequently, more rapidly, with greater duration and/or with greater affinity with a particular cell or substance than it does with alternative cells or substances. For example, an antigen- binding protein “binds” to a target if it binds with greater affinity, avidity, more readily, and/or with greater duration than it binds to other substances. For example, a 4-1BB binding protein that binds to the CRD1 domain is a protein that binds this domain with greater affinity, avidity, more readily, and/or with greater duration than it binds to other domains and regions of 4-1BB, or other proteins that are not 4-1BB. It is also understood by reading this definition that, for example, an antigen-binding protein which specifically or preferentially binds to a first target may or may not specifically bind to a second target. As such, “binding,” “specific binding” or “preferential binding” does not necessarily require (although it can include) exclusive binding. If numerical examples are of interest, an antigen-binding protein may bind with a KD value that is numerically less than 1x10-6 M, 1x10-7 M, 1x10-8 M, or 1x10-9 M. [65] In some embodiments, the present disclosure provides a 4-1BB antigen-binding protein that binds to an epitope that comprises residues C102, V71, and Q104. In certain embodiments, the epitope may further comprise P90. In certain embodiments, the epitope may further comprise one or more residues selected from the group consisting of: K69, T89, F92, M101, and L112. These epitope residues are numbered according to the numbering of SEQ ID NO:272 (human 4-1BB). Corresponding residues from other 4-1BB homologs, isoforms, variants, or fragments can be identified according to sequence alignment or structural alignment that is known in the art. For example, alignments can be done by hand or by using well-known sequence alignment programs such as ClustalW2, or "BLAST 2 Sequences" using default parameters. As exemplified by the structural data, C102, V71, and Q104 were found to be “primary” residues for molecule 380984 binding; P90 was found to be “contributing” residues for scFv32211 binding; and K69, T89, F92, M101, and L112, while involved in H-bonding and Van Der Waals interactions with 380984, were found to be “optional” residue for 380984 binding. [66] Based on the structural studies, one or more of the following substitutions would likely substantially disrupt the binding of 380984 binding to 4-1BB: (i) C102 is replaced with A; (ii) A104 is replaced with A;
or(iii) K69 is replaced with A. An Alanine-scan shows that none of the epitope residues potentially abrogate the VH binding because the hydrogen bonds are mostly thought backbone. The C102 is highly conserved disulfide within 4-1BB CRD3 domain, maintaining its structural stability, which mutation would potentially break the fold and also affect the VH binding. 2.3 Structure of 4-1BB Antigen Binding Proteins [67] As used herein, “Complementarity Determining Regions” (CDRs) can be identified according to the definitions of the Kabat, Chothia, the accumulation of both Kabat and Chothia, AbM, contact, North, and/or conformational definitions or any method of CDR determination well known in the art. See, e.g., Kabat et al., 1991, Sequences of Proteins of Immunological Interest, 5th ed. (hypervariable regions); Chothia & Lesk, 1987, J Mol Biol., 196:901-917 (structural loop structures). The identity of the amino acid residues in a particular antibody that make up a CDR can be determined using methods well known in the art. AbM definition of CDRs is a compromise between Kabat and Chothia and uses Oxford Molecular’s AbM antibody modeling software (Accelrys®). The “contact” definition of CDRs is based on observed antigen contacts, set forth in MacCallum et al., 1996, J. Mol. Biol., 262:732-745. The “conformational” definition of CDRs is based on residues that make enthalpic contributions to antigen binding (see, e.g., Makabe et al., 2008, J. Biol. Chem., 283:1156-1166). North has identified canonical CDR conformations using a different preferred set of CDR definitions (North et al., 2011, J. Mol. Biol.406: 228-256). In another approach, referred to herein as the “conformational definition” of CDRs, the positions of the CDRs may be identified as the residues that make enthalpic contributions to antigen binding (Makabe et al., 2008, J Biol. Chem.283:1156-1166). Martin definition (also called enhanced Chothia definition) combines the Kabat and Chothia definitions and differs from them only in the heavy chain, where CDR- H1 includes all residues of Kabat and Chothia while CDR-H2 is seven residues shorter than that defined by Kabat (Martin, Bioinformatics tools for antibody engineering. Handbook of Therapeutic Antibodies. Weinheim: Wiley-VCH Verlag GmbH; (2008). p.95–117; see also the database maintained by the Institute of Structural and Molecular Biology at the University College London, http://www.bioinf.org.uk/abs/#cdrid). Still other CDR boundary definitions may not strictly follow one of the above approaches, but will nonetheless overlap with at least a portion of the Kabat CDRs, although they may be shortened or lengthened in light of prediction or experimental findings that particular residues or groups of residues or even entire CDRs do not significantly impact antigen binding. For example, “combined” CDRs may also be used. Therefore, a CDR may refer to CDRs defined by any approach known in the art, including combinations of approaches. For any given embodiment containing more than one CDR, the CDRs (or other residue of the antibody) may be defined in accordance with any of Kabat, Chothia, North, AbM, Contact, IMGT, Martin, combined Kabat and Chothia, and/or conformational definitions.
[68] For example, Table N1 shows several commonly used definitions of CDRs: Table N1. Definitions of CDRs. Loop Kabat AbM Chothia1 Contact2 IMGT L1 L24-L34 L24-L34 L26-L32 L30-L36 L27-L32 7;
C - : 6-3 ; C - : 5 -56; C - 3: 95- 0 . abe s a consensus o C ot a de nton based upon Chothia & Lesk (1987) (e.g., CDR-H3) and Chothia et al., 1989, Nature 342:877-883 (e.g., CDR-H2). 2. Any of the numbering schemes can be used for these CDR definitions, except the contact definition uses the Chothia or Martin (Enhanced Chothia) definition. 3. The end of the Chothia CDR-H1 loop when numbered using the Kabat numbering convention varies between H32 and H34 depending on the length of the loop. (This is because the Kabat numbering scheme places the insertions at H35A and H35B.) [69] The CDR sequences provided in the Sequence Tables are based on the Kabat definition. However, other definitions for CDRs may also be used. Accordingly, in some embodiments, the 4-1BB antigen- binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:147, 149, 151, 153, 155, 157, 159, 324, 161, 163, 165, 167, 169, 171, 173, 175, 404, 406, 408, or 410; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 148, 150, 152, 154, 156, 158, 160, 325, 162, 164, 166, 168, 170, 172, 174, 176, 405, 407, 409, or 411. In some embodiments, the three heavy chain CDRs and three light chain CDRs come from the same clone as shown in Sequence Table B and Table K1. In exemplary embodiments, the CDRs are defined according to Kabat, Chothia, AbM, contact, or IMGT. [70] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:147; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:148. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
[71] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:149; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:150. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [72] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:151; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:152. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [73] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:153; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:154. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [74] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:155; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:156. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [75] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:157; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:158. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [76] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:159 or 324; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:160 or 325. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [77] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy
chain CDRs in SEQ ID NO:161; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:162. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [78] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:163; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:164. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [79] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:165; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:166. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [80] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:167; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:168. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [81] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:169; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:170. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [82] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:171; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:172. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [83] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:173; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:174. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less.
[84] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:175; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:176. The antigen binding protein may further bind to its target 4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [85] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:404; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:405. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [86] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:406; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:407. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [87] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:408; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:409. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [88] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:410; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:411. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [89] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 147, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 148. [90] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 149, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 150.
[91] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 151, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 152. [92] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 153, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 154. [93] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 155, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 156. [94] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 157, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 158. [95] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 159 or 324, and (ii) the light chain CDR-L1, CDR- L2, and CDR-L3 of SEQ ID NO: 160 or 325. [96] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 161, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 162. [97] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 163, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 164. [98] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 165, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 166. [99] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 167, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 168. [100] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 169, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 170.
[101] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 171, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 172. [102] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 173, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 174. [103] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 175, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 176. [104] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 404, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 405. [105] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 406, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 407. [106] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 408, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 409. [107] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 410, and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 411. [108] In exemplary aspects, the 4-1BB antigen binding proteins comprise (a) CDR-H1 amino acid sequence set forth in Sequence Table B and Table K1, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set for in Table B or Table K1; (b) CDR-H2 amino acid sequence set forth in Sequence Table B and Table K1, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Table B or Table K1; (c) a CDR-H3 amino acid sequence set forth in Sequence Table B and Table K1, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Table B or Table K1; (d) a CDR-L1 amino acid sequence set forth in Sequence Table B and Table K1, or a variant sequence thereof which differs by only 1-4 amino acids
(e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Table B or Table K1; (e) a CDR-L2 amino acid sequence set forth in Sequence Table B and Table K1, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Table B or Table K1; (f) a CDR-L3 amino acid sequence set forth in Sequence Table B and Table K1, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Table B or Table K1; or (g) a combination of any two, three, four, five, or six of (a)-(f). The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [109] In exemplary embodiments, the 4-1BB antigen binding protein comprises 3, 4, 5, or all 6 of the amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table B and Table K1. In exemplary embodiments, the antigen binding protein comprises each of the LC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table B and Table K1 and at least 1 or 2 of the HC CDR amino acid sequences designated by the SEQ ID NOs in under the same clone name of Sequence Table B and Table K1. In exemplary embodiments, the antigen binding protein comprises each of the HC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table B and Table K1 and at least 1 or 2 of the LC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table B and Table K1. In exemplary embodiments, the antigen binding protein comprises six CDR amino acid sequences listed under the same clone name in Sequence Tables or comprising six CDR amino acid sequences selected from the group consisting of: (1) SEQ ID NOs: 43-48, (2) SEQ ID NOs: 49-54, (3) SEQ ID NOs: 55-60, (4) SEQ ID NOs: 61-66, (5) SEQ ID NOs: 67-72, (6) SEQ ID NOs: 73-78, (7) SEQ ID NOs: 79-84, (8) SEQ ID NOs: 85-90, (9) SEQ ID NOs: 91-96, (10) SEQ ID NOs: 97-102, (11) SEQ ID NOs: 103-108, (12) SEQ ID NOs: 109-114, (13) SEQ ID NOs: 115-120, (14) SEQ ID NOs: 121-126, (15) SEQ ID NOs: 127-132, (16) SEQ ID NOs: 386-391, (17) SEQ ID NOs.: 392-397. [110] The CDR sequences of the exemplary 4-1BB antigen-binding proteins disclosed herein can be aligned to identify consensus sequences. In one alignment, the CDR sequences are aligned according to Kabat numbering (i.e., residues with the same Kabat numbering are aligned), and the results are shown in Tables N2 & N3. Table N2. Light chain CDR alignment based on Kabat numbering (SEQ ID numbers of these CDR sequences can be found in Table B) Clone name CDR-L1 CDR-L2 CDR-L3 7 NN I KNVH DDDRP VWD DHVV
14A5 RASQ-----SIKRYLN AASSLQS QQRFSIP--FT 19G1 SGDK-----LGDNYAC QDSRRPS QTWDSSA--VM 15A12.012 RASQ-----SISTYLN AASSLQS LQTYSA---LT
(SEQ ID numbers of these sequences can be found in Table B) Clone name CDR-H1 CDR-H2 CDR-H3 6C7 RYYWS YID-DSGNTNYNPSLKS GVAAGR----------IDP
[111] For Kabat CDR-H2, additional residues are assigned between H52 and H53 (e.g., H52A, H52B, H52C), those positions are represented by a gap if a particular CDR-H2 does not have a corresponding residue. Similarly, gaps are used to align Kabat CDR-H3, between H100 and H101 (H100A-H100K according to Kabat numbering), Kabat CDR-L1 between L27 and L28 (L27A-L27F according to Kabat numbering), and Kabat CDR-L3 between L95 and L96 (L95A-L95F according to Kabat numbering). Table N4 summarizes the consensus sequences derived from this method. Table N4. Consensus Sequences of CDRs Seq No. Name Sequence
4-1 BB CDR- X1 I X2 X3 X4 X5 X6 X7 X8 Y X9 X10 X11 X12 X13 X14 K X15 H2 X1 = Y, R, A, V, or L X2 = D, Y, S, W, or G 17
X4 = D, Q, S, R, K, N, V, or T X5 = R, or L X6 = P, Q, or F
[112] Alternatively, the CDR sequences can be aligned according to the germline sequences those clones are derived from. Table N5-N7 summarize such alignment. Germline sequences are in bold. Table N5. Heavy chain germline alignment (SEQ ID numbers of these sequences can be found in Table B and Table M1) H_CDR1 H_CDR2 H_CDR3 VH3|3-33/D3|3-3|RF3/JH4 SYGMH VIWYDGSNKYYADSVKG ITIFGVV IIYFDY Y V V V V V V V V H F V V V V V V
VH4|4-59/D6|6-13|RF2/JH5 SYYWS YIYY-SGSTNYNPSLKS GIAAAG--------NWFDP 6C7 RYYWS YIDD-SGNTNYNPSLKS GVAAG----------RIDP P V V P P
(SEQ ID numbers of these sequences can be found in Table B and Table M2) K_CDR1 K_CDR2 K_CDR3 VK2|A2/JK1 KSSQSLLHSDGKTYLY EVSNRFS MQSIQLPWT
. (SEQ ID numbers of these sequences can be found in Table B and Table M3) L_CDR1 L_CDR2 L_CDR3
19G1 SGD--KLGDNYAC QDSRRPS QTWDS--SAVM 19G1.016 SGD--KLGDNYAS QDSRRPS QTWDS--SAVM
, le N8. Table N8. Consensus CDR sequences based on germline alignment X1 Y X2 X3 X4 4-1 BB CDR- X1 = S, T, N, or G 484 H1 (VH3 X2 = A S or G 17
X1 X2 X3 X4 G X5 X6 X7 Y Y Y G M D V X1 = G or S 1 BB CDR X2 = Y or G
X4 = Y, N, or F X5 = S or absent X6 = T, S, V, or I
[114] Also provided herein are VH-only 4-1BB antigen-binding proteins, in particular, UniDabs that are derived from UniRat technology. [115] For example, exemplified herein are 4-1BB antigen-binding protein that comprises a heavy chain variable domain (VH) and does not comprise a light chain variable domain (VL), wherein said protein comprises the CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, or 385. As explained in detail above, CDRs can be identified according to the definitions of the Kabat, Chothia, the accumulation of both Kabat and Chothia, AbM, contact, North, and/or conformational definitions or any method of CDR determination well known in the art. The CDR
sequences shown in Table I are defined according to Kabat. However, other definitions for CDRs may also be used. [116] In some embodiments, the present disclosure provides a 4-1BB antigen binding protein comprising a heavy chain variable domain (VH) but does not comprise a light chain variable domain (VL), wherein said 4-1BB antigen binding protein comprises the following paratope residues (VH numbering according to Kabat): (1) H45 is Leu, Phe, Ile, or Tyr; (2) H47 is Trp, Phe, Leu, or Tyr; (3) H100B is Tyr, Arg, His, Lys, or Met; (4) H100D is Thr, Ala, Asn, Cys, Gln, Lys, Met, or Val; and (5) H100F is Phe, Trp or Tyr. Based on structural studies, these paratope residues were found to be “primary” residues from clone 380984 that bind to 4-1BB. Preferably, (1) H45 is Leu or Phe; (2) H47 is Trp or Leu; (3) H100B is Tyr or Met; (4) H100D is Thr or Val; and (5) H100F is Phe or Trp. More preferably, (1) H45 is Leu; (2) H47 is Trp; (3) H100B is Tyr; (4) H100D is Thr; and (5) H100F is Phe. [117] The 4-1BB antigen binding protein may further comprise the following paratope residues: (VH according to Kabat): (6) H97 is Ser, Arg, Asn, Gln, Glu, His,,Leu, Lys, Met, Phe, Thr, Trp, Tyr, or Val; (7) H100E is Ser, Ala, Asn, Asp, Cys, His, Trp, Tyr, or Val; and (9) H102 is Tyr, Ile, Lys, or Val. Based on structural studies, these paratope residues were found to be “optional” residues from clone 380984that bind to 4-1BB. Preferably, ((6) H97 is Ser or Leu; (7) H100E is Ser or Val; and (9) H102 is Tyr or Lys. More preferably, (6) H97 is Ser; (7) H100E is Ser; and (9) H102 is Tyr. [118] Based on structural studies, it was discovered that, for 380984, among the three complementarity determining regions (CDRs), CDR-H3 is responsible for contacting FAP residues. In addition, Leu45 and Trp47 are characterize as framework residues under Kabat, AbM, Chothia, and IMGT definitions. Contact definition characterizes Trp47 as part of CDR-H2. Accordingly, in certain embodiments, the 4-1BB antigen-binding protein comprises (a) a VH that comprises: (i) a CDR-H3 comprising any one of SEQ ID NOs:576-578 and 580; and (ii) framework residue H45 is Leu and framework residue H47 is Trp; or (b) a VH that comprises: (i) a CDR-H2 comprising SEQ ID NO:574, (ii) a CDR-H3 comprising SEQ ID NO: 579; and (iii) framework residue H45 is Leu. In certain embodiments, said framework residues, CDR-H2, or CDR-H3 contact 4-1BB residues C102, V71, Q104, P90, K69, T89, F92, M101, or L112, according to the numbering of SEQ ID NO:272. [119] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:371. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [120] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy
chain CDRs in SEQ ID NO:372. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [121] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:373. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [122] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:374. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [123] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:375. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [124] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:376. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [125] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:377. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [126] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:378. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [127] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:379. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [128] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy
chain CDRs in SEQ ID NO:380. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [129] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:381. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [130] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:382. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [131] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:383. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [132] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:384. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [133] In various embodiments, the 4-1BB antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:385. The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [134] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 371. [135] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 372. [136] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 373. [137] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 374.
[138] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 375. [139] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 376. [140] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 377. [141] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 378. [142] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 379. [143] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 380. [144] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 381. [145] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 382. [146] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 383. [147] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 384. [148] In some embodiments, the 4-1BB antigen-binding protein disclosed herein comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 385. [149] In exemplary aspects, the 4-1BB antigen binding proteins comprise (a) CDR-H1 amino acid sequence set forth in Sequence Table I, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table I; (b) CDR-H2 amino acid sequence set forth in Sequence Table I, or a variant sequence thereof which differs by only 1- 4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table I; (c) a CDR-H3 amino acid sequence set forth in Sequence Table I, or a variant sequence thereof which differs
by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table I; or (d) a combination of any two, or three (a)-(c). The antigen binding protein may further bind to its target (4-1BB) with a KD value of 100 nM or less, or a KD value of 500 nM or less. [150] In exemplary embodiments, the 4-1BB antigen binding protein comprises a CDR-H1, a CDR-H2, and a CDR-H3 comprising: (1) SEQ ID NOs: 326-328, respectively (2) SEQ ID NOs: 329-331, respectively, (3) SEQ ID NOs: 332-334, respectively, (4) SEQ ID NOs: 335-337, respectively, (5) SEQ ID NOs: 338-340, respectively, (6) SEQ ID NOs: 341-343, respectively, (7) SEQ ID NOs: 344-346, respectively, (8) SEQ ID NOs: 347-349, respectively, (9) SEQ ID NOs: 350-352, respectively, (10) SEQ ID NOs: 353-355, respectively, (11) SEQ ID NOs: 356-358, respectively, (12) SEQ ID NOs: 359-361, respectively, (13) SEQ ID NOs: 362-364, respectively, (14) SEQ ID NOs: 365-367, respectively, or (15) SEQ ID NOs: 368-370, respectively. [151] The CDR sequences of these VH-only 4-1BB antigen-binding proteins can be aligned to identify consensus sequences. In one alignment, the CDR sequences are aligned according to Kabat numbering (i.e., residues with the same Kabat numbering are aligned), and the results are shown in Table N9. Gaps represent absent residues in Kabat CDR-H2, between H52 and H53 (H52A-H52C), and in Kabat CDR- H3, between H100 and H101 (H100A-H100K). Table N10 summarizes the consensus sequences derived from this method. Table N9. UniDab CDR alignment based on Kabat numbering clone CDR1 CDR2 CDR3 386340 VH SFAMT AISG--SGGSTYYAGSVKG EAYESS-GYYTTSFDY Y Y Y Y Y Y Y I I I Y I I Y
Table N10. UniDab CDR consensus sequences
Seq No. Name Sequence X1 X2 X3 M X4 16
[152] In general, CDRs are separated by “framework” (FR) residues. A VH or VL domain framework comprises four framework sub-regions, FR1, FR2, FR3 and FR4, interspersed with CDRs in the following structure: FR1 - CDR1 - FR2 - CDR2 - FR3 - CDR3 - FR4. Accordingly, the antigen binding proteins described herein may comprise a VH framework, such as a human germline VH framework sequence, and a VL framework, such as human germline VL framework sequences. [153] Preferred human germline light chain frameworks are frameworks derived from Vκ or Vλ germlines. It will be understood that if a sequence is “derived from” one or more germlines, what is referred to is a
structural relationship, in which the features of a sequence correspond to the noted germline sequences, but may comprise somatic mutations or other amino acid differences relative to the noted germline sequence. For a sequence to be “derived from” a germline, an actual process of deriving that sequence from a germline sequence (either via molecular biology or computational analysis) is not necessarily required. For example, VL frameworks may be derived from one of the framework of the following germlines: DPK9 (IMGT name: IGKV1-39), DPK12 (IMGT name: IGKV2D-29), DPK18 (IMGT name: IGKV2-30), DPK24 (IMGT name: IGKV4-1), HK102_V1 (IMGT name: IGKV1-5), DPK1 (IMGT name: IGKV1-33), DPK8 (IMGT name: IGKV1-9), DPK3 (IMGT name: IGKV1-6), DPK21 (IMGT name: IGKV3- 15), Vg_38K (IMGT name: IGKV3-11 ), DPK22 (IMGT name: IGKV3-20), DPK15 (IMGT name: IGKV2- 28), DPL16 (IMGT name: IGLV3-19), DPL8 (IMGT name: IGLV1-40), V1-22 (IMGT name: IGLV6-57). Alternatively, or in addition, the framework sequence may be derived from a human germline consensus framework sequence, such as the framework of human Vλ1 consensus sequence, Vλ3 consensus sequence, Vκ1 consensus sequence, Vκ2 consensus sequence, Vκ3 consensus sequence. Sequences of human germline frameworks are available from various public databases, such as V-base, IMGT, NCBI, or Abysis. [154] The 4-1BB antigen binding proteins described herein may comprise a VL framework, wherein the framework may comprise one or more amino acid substitutions, additions, or deletions, while still retaining functional and structural similarity with the germline from which it was derived. In some aspects, the VL framework is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to a human germline VL framework sequence. In some aspects, the antigen binding protein, antibody, or antigen binding fragment thereof, comprises a VL framework comprising 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 amino acid substitutions, additions or deletions relative to the human germline VL framework sequence. [155] The VH framework sequence can be derived from a human VH3 germline, a VH1 germline, a VH5 germline, a human VH2 germline, or a VH4 germline. Preferred human germline heavy chain frameworks are frameworks derived from VH1, VH2, VH3, or VH4 germlines. For example, VH frameworks may be derived from the framework of one of the following germlines: DP54 or IGHV3-7, DP47 or IGHV3-23, DP71 or IGHV4-59, DP75 or IGHV1-2_02, DP10 or IGHV1-69, DP7 or IGHV1-46, DP49 or IGHV3-30, DP51 or IGHV3-48, DP38 or IGHV3-15, DP79 or IGHV4-39, DP78 or IGHV4-30-4, DP73 or IGHV5-51, DP50 or IGHV3-33, DP46 or IGHV3-30-3, DP31 or IGHV3-9. Alternatively, or in addition, the framework sequence may be derived from the framework of a consensus sequence, such as: VH3 germline consensus sequence, VH1 germline consensus sequence, VH5 germline consensus sequence, VH2 germline consensus sequence, or VH4 germline consensus sequence.
[156] The antigen binding proteins described herein may comprise a VH framework, wherein the framework may comprise one or more amino acid substitutions, additions, or deletions, while still retaining functional and structural similarity with the germline from which it was derived. In some aspects, the VH framework is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to a human germline VH framework sequence. In some aspects, the antigen binding protein, antibody, or antigen binding fragment thereof, comprises a VH framework comprising 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 amino acid substitutions, additions or deletions relative to the human germline VH framework sequence. [157] In exemplary embodiments, the 4-1BB antigen-binding protein comprises a VH that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 147, 149, 151, 153, 155, 157, 159, 324, 161, 163, 165, 167, 169, 171, 173, 175, 404, 406, 408, or 410. In exemplary embodiments, the 4-1BB antigen-binding protein comprises a VL that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 148, 150, 152, 154, 156, 158, 160, 325, 162, 164, 166, 168, 170, 172, 174, 176, 405, 407, 409, or 411. Preferably, the 4-1BB antigen binding protein comprises a pair of VH and VL sequences listed under the same clone name in Table B and Table K1. [158] In exemplary embodiments, the 4-1BB antigen-binding protein comprises a VH but does not comprise a VL, wherein said VH comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, or 385. [159] In some embodiments, the 4-1BB antigen binding protein comprises a CH1 domain, preferably a human CH1 domain (such as a human IgG1 CH1, a human IgG2 CH1, a human IgG3 CH1, or a human IgG4 CH1). Non-limiting examples of human CH1 sequences are provided in the Sequence Tables. In some embodiments, the CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 261, 265, 269, 422, or 425. [160] In certain embodiments, the 4-1BB antigen binding proteins described herein comprises an Fc domain. The Fc domain can be derived from IgA (e.g., IgA1 or lgA2), IgG, IgE, or IgG (e.g., IgG1, lgG2,
lgG3, or lgG4). In some embodiments, the Fc domain comprises wild type sequence of a human Fc domain. Non-limiting examples of human Fc domain sequences are provided in the Sequence Table. [161] In some embodiments, the Fc domain comprises one or more mutations resulting in altered biological activity, such as to improve half-life/stability or to render the antibody more suitable for expression/manufacturability. For example, mutations may be introduced into the Fc domain to reduce the effector activity (e.g., WO 2005/063815), and/or to increase the homogeneity during the production of the recombinant protein. [162] In general, amino acid residues in the IgG heavy constant domain of an antibody are numbered according the EU index of Edelman et al., 1969, Proc. Natl. Acad. Sci. USA 63(1):78-85 as described in Kabat et al., 1991, referred to herein as the “EU index numbering.” Typically, the constant domain comprises from residue 118 to 447, and the Fc domain comprises from residue 236 to 447 of the human lgG1 constant domain. Comparison between EU numbering and other numbering systems can be found, e.g., at IGMT database. [163] Amino acid residues of the light chain constant domain are numbered according to Kabat et al., 1991, "Sequences of Proteins of Immunological Interest 5th Ed.", 1991, NATIONAL INSTITUTES OF HEALTH. Kappa light chain also has EU index numbering, and the EU index and Kabat numbering are identical. Lambda light chain does not have EU index numbering. [164] In some embodiments, the Fc domain is the Fc domain of human lgG1 and comprises one or more of the following effector-null mutations: L234A, L235A, and G237A (numbering according to the EU index), often referred as “LALA” mutations. [165] It has been reported that a single mutation of L235E was sufficient for knocking out binding to Fc receptors on U937 cells. Furthermore, the 100-fold reduction in binding to FcγR also resulted in lower T cell activation and proliferation in the presence of the L235E Fc mutant IgG1. Building upon this initial mutation it was found that the combination of L234A and L235A (commonly called LALA mutations) eliminated FcγRIIa binding. These two mutations were later shown to eliminate detectable binding to FcγRI, IIa, and IIIa for both IgG1 and IgG4. Other sites have been reported to knockout Fc receptor binding, such as Gly237Ala, Glu318Ala, Asp265Ala and Glu233Pro mutations. [166] In exemplary embodiments, the Fc region comprises a Stable Effector Functionless (SEFL) mutation to inhibit or reduce the ability to interact with Fcγ receptors, sequence. SEFL mutations are known in the art. See, e.g., Liu et al., J Biol Chem 292: 1876-1883 (2016); and Jacobsen et al., J. Biol. Chem.292: 1865-1875 (2017), and Estes et al., iScience, Volume 24, Issue 12, 2021,103447. Further, US US9546203 discloses a Fc region comprising a N297G mutation, and one or more substitutions at position V259, A287, R292, V302, L306, V323, or I332, using EU numbering scheme, with a cysteine
amino acid residue. In exemplary aspects, the SEFL mutation comprises one or more of the following mutations, numbered according to the EU system: L242C, A287C, R292C, N297G, V302C, L306C, and/or K334C. In exemplary aspects, the SEFL mutation comprises N297G. In exemplary aspects, the SEFL mutation comprises A287C, N297G, and L306C. In other exemplary aspects, the SEFL mutation comprises R292C, N297G, and V302C (i.e., SEFL2-2). [167] In exemplary embodiments, the Fc region comprises a YTE mutation. The M252Y/S254T/T256E (EU index numbering, referred to “YTE”) triple mutation have been shown to increase IgG half-life in cynomolgus monkeys by an approximate 4-fold increase. [168] C-terminal lysine clipping is a common phenomenon occurring during the bioproduction of monoclonal antibodies. Often, the lysine residue is removed via carboxypeptidase D (CpD), which results in generation of a mixture of antibody isoforms bearing zero or one C-terminal lysine residues on each heavy chain. Further, following C-terminal lysine cleavage, peptidylglycine α-amidating monooxygenase (PAM) catalyzes the hydroxylation of glycine and removal of the glyoxylate from the glycine residue, leaving an amidated C-terminal proline. Therefore, during recombinant production of a monoclonal antibody, the product is often a mixture of C-terminal processing variants, with heavy chain C-terminus ends at (amidated) proline, glycine, or lysine. Sometimes, it may be desirable to delete the C-terminal lysine of the Fc domain to increase the homogeneity during the production of the recombinant protein. [169] In some embodiments, the terminal lysine may be absent; in some embodiments, the terminal lysine may be present; in some embodiments, the terminal glycine-lysine may be absent; in some embodiments, the terminal glycine-lysine may be present. It will be appreciated that in some embodiments, a pharmaceutically suitable composition may comprise a mixture of species that do and do not comprise the terminal lysine and/or glycine-lysine. [170] In exemplary embodiments, the 4-1BB antigen binding proteins described herein comprise Fc that is derived from an IgG1. In some embodiments, the Fc comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, 426, 263, 267, or 483. [171] In exemplary embodiments, the 4-1BB antigen binding proteins described herein comprise an IgG1 heavy chain constant domain. In some embodiments, the heavy chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at
least 99%, or 100% identical to SEQ ID NO: 244, 245, 246, 247, 248, 249, 256, 257, 258, 260, 412, 415, 417, 418, 427, 264, 268, or 271. [172] In some embodiments, the 4-1BB antigen-binding protein described herein comprising a kappa or lambda light chain constant domain. In some embodiments, the kappa light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428. In some embodiments, the lambda light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424. [173] The 4-1BB antigen-binding protein may be a full-length immunoglobulin, a Fab, or an scFv. Exemplary full length 4-1BB binding immunoglobulins are shown as Sequence Table F. Exemplary Fab and scFV domains are shown in Sequence Table H and Table K6. The scFv may comprises a linker between VH and VL. Exemplary linker sequences, such as GS-based linkers, are provided in the Sequence Table G. [174] In some embodiments, the scFv described herein comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 291. [175] In some embodiments, the scFv described herein comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 445 (14A5.002 scFv #2). [176] In some embodiments, the Fab described herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 298, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 300 (6F9.009 Fab).
[177] In some embodiments, the Fab described herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 305, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 307 (19G1.016 Fab). [178] In some embodiments, the Fab described herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 316, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 317 (6C7.018 Fab). [179] In some embodiments, the Fab described herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-227 of SEQ ID NO: 434, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 435 (4-1BB Fab of 56039). [180] In some embodiments, the Fab described herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-227 of SEQ ID NO: 449, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 450 (4-1BB Fab of 56040). 2.4 Structure of PD-L1 Antigen Binding Proteins
[181] Also provided herein are PD-L1 antigen-binding proteins. Examples of PD-L1 antigen-binding proteins are provided in the Sequence Tables. As discussed in detail above, the CDR sequences provided in the Sequence Tables are based on the Kabat definition. However, other definitions for CDRs may also be used. Accordingly, in some embodiments, the PD-L1 antigen-binding protein disclosed herein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 133, 135, 137, 139, 141, 143, 322, 145, 398, 400, or 402; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 134, 136, 138, 140, 142, 144, 323, 146, 399, 401, or 403. In some embodiments, the three heavy chain CDRs and three light chain CDRs come from the same clone as shown in Sequence Table C. In exemplary embodiments, the CDRs are defined according to Kabat, Chothia, AbM, contact, or IMGT [182] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:133; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:134. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [183] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:135; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:136. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [184] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:137; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:138. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [185] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:139; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:140. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [186] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:141; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:142. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less.
[187] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:143; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:144. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [188] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:322; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:323. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [189] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:145; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:146. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [190] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:398; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:399. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [191] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:400; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO: 401. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [192] In various embodiments, the PD-L1 antigen binding protein comprises a VH comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% identical to the three heavy chain CDRs in SEQ ID NO:402; and a VL comprising three CDRs that in combination are at least 85%, at least 90%, or at least 95% to the three light chain CDRs in SEQ ID NO:403. The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [193] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:133; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 134.
[194] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:1353; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 136. [195] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:137; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 138. [196] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:1393; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 140. [197] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:141; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 142. [198] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:143; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 144. [199] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:322; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 323. [200] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:145; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 146. [201] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:398; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 399. [202] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:400; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 401. [203] In some embodiments, the PD-L1 antigen-binding protein comprises: (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:402; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 403.
[204] In exemplary aspects, the PD-L1 antigen binding proteins comprise (a) CDR-H1 amino acid sequence set forth in Sequence Table A and Table K2, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table A or Table K2; (b) CDR-H2 amino acid sequence set forth in Sequence Table A and Table K2, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table A or Table K2; (c) a CDR-H3 amino acid sequence set forth in Sequence Table A and Table K2, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table A or Table K2; (d) a CDR-L1 amino acid sequence set forth in Sequence Table A and Table K2, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table A or Table K2; (e) a CDR-L2 amino acid sequence set forth in Sequence Table A and Table K2, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table A or Table K2; (f) a CDR-L3 amino acid sequence set forth in Sequence Table A and Table K2, or a variant sequence thereof which differs by only 1-4 amino acids (e.g., 1, 2, 3, 4 amino acids) or which has at least about 80%, at least about 85%, at least about 90%, or at least about 95% sequence identity to the sequence set forth in Sequence Table A or Table K2; or (g) a combination of any two, three, four, five, or six of (a)-(f). The antigen binding protein may further bind to its target (PD-L1) with a KD value of 100 nM or less, or a KD value of 50 nM or less. [205] In exemplary embodiments, the PD-L1 antigen binding protein comprises 3, 4, 5, or all 6 of the amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table A and Table K2. In exemplary embodiments, the antigen binding protein comprises each of the LC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table A and Table K2 and at least 1 or 2 of the HC CDR amino acid sequences designated by the SEQ ID NOs in under the same clone name of Sequence Table A and Table K2. In exemplary embodiments, the antigen binding protein comprises each of the HC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table A and Table K2, and at least 1 or 2 of the LC CDR amino acid sequences designated by the SEQ ID NOs under the same clone name in Sequence Table A and Table K2. In exemplary embodiments, the antigen binding protein comprises six CDR amino acid sequences listed under the same clone name in Sequence Table A and Table K2, or comprising six CDR amino acid sequences selected from the group consisting of: (1) SEQ ID NOs: 1-6, (2) SEQ ID NOs: 7- 12, (3) SEQ ID NOs: 13-18, (4) SEQ ID NOs: 19-24, (5) SEQ ID NOs: 25-30, (6) SEQ ID NOs: 31-36, and (7) SEQ ID NOs: 37-42.
[206] Preferred VH and VL framework sequences for PD-L1 antigen-binding proteins disclosed herein are human framework sequences described in detail above. [207] In exemplary embodiments, the PD-L1 antigen-binding protein comprises a VH that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 133, 135, 137, 139, 141, 143, 322, 145, 398, 400, or 402. In exemplary embodiments, the PD-L1 antigen-binding protein comprises a VL that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 134, 136, 138, 140, 142, 323144, 146, 399, 401, or 403. Preferably, the PD-L1 antigen binding protein comprises sequences or variants of sequences from a pair of VH and VL sequences listed under the same clone name in Table C and Table K3. [208] The PD-L1 antigen-binding proteins disclosed herein may further comprises a CL, a CH1, and/or a Fc region as described in detail above. [209] In some embodiments, the PD-L1 antigen binding protein comprises a CH1 domain, preferably a human CH1 domain (such as a human IgG1 CH1, a human IgG2 CH1, a human IgG3 CH1, or a human IgG4 CH1). Non-limiting examples of human CH1 sequences are provided in the Sequence Tables. In some embodiments, the CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 261, 265, 269, 422, or 425. [210] In certain embodiments, the PD-L1 antigen binding proteins described herein comprises an Fc domain. The Fc domain can be derived from IgA (e.g., IgA1 or lgA2), IgG, IgE, or IgG (e.g., IgG1, lgG2, lgG3, or lgG4). In some embodiments, the Fc domain comprises wild type sequence of a human Fc domain. Non-limiting examples of human Fc domain sequences are provided in the Sequence Table. [211] In some embodiments, the Fc domain is the Fc domain of human lgG1 and comprises one or more of the following effector-null mutations: L234A, L235A, and G237A (numbering according to the EU index), often referred as “LALA” mutations. [212] In exemplary embodiments, the Fc region comprises a Stable Effector Functionless (SEFL) mutation to reduce the ability to interact with Fcγ receptors. In exemplary aspects, the SEFL mutation comprises one or more of the following mutations, numbered according to the EU system: L242C, A287C, R292C, N297G, V302C, L306C, and/or K334C. In exemplary aspects, the SEFL mutation
comprises N297G. In exemplary aspects, the SEFL mutation comprises A287C, N297G, and L306C. In other exemplary aspects, the SEFL mutation comprises R292C, N297G, and V302C (i.e., SEFL2-2). [213] In exemplary embodiments, the Fc region comprises a YTE mutation. The M252Y/S254T/T256E (EU index numbering, referred to “YTE”) triple mutation have been shown to increase IgG half-life in cynomolgus monkeys by an approximate 4-fold increase. [214] In exemplary embodiments, the PD-L1 antigen binding proteins described herein comprise Fc that is derived from an IgG1. In some embodiments, the Fc comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, 426, 263, 267, or 483. [215] In exemplary embodiments, the PD-L1 antigen binding proteins described herein comprise an IgG1 heavy chain constant domain. In some embodiments, the heavy chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 244, 245, 246, 247, 248, 249, 256, 257, 258, 260, 412, 415, 417, 418, 427, 264, 268, or 271. [216] In some embodiments, the PD-L1 antigen-binding protein described herein comprising a kappa or lambda light chain constant domain. In some embodiments, the kappa light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428. In some embodiments, the lambda light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424. [217] The PD-L1 antigen-binding protein may be a full-length immunoglobulin, a Fab, or an scFv. Exemplary full length 4-1BB binding immunoglobulins are shown as Sequence Table E. Exemplary Fab and scFV domains are shown in Sequence Table H and Table K6. The scFv may comprises a linker between VH and VL. Exemplary linker sequences, such as GS-based linkers, are provided in the Sequence Table G.
[218] In some embodiments, the scFv disclosed herein comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 284 (26F6.002.009 scFv). [219] In some embodiments, the scFv disclosed herein comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 436 (scFv from 56039). [220] In some embodiments, the Fab disclosed herein comprises a Fab heavy chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to residues 1-221 SEQ ID NO: 451, and a Fab light chain sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 452 (Fab from 56041). 2.5 4-1BB x PD-L1 Bispecific Molecules. [221] In various aspects, the antigen binding protein is a bispecific molecule which binds to two different antigens or targets. In various instances, the bispecific molecule binds to both 4-1BB and PD-L1. [222] Many different formats of bispecific molecules have been exemplified, some of which are depicted in FIGs.2A-2B, FIGs.13A-13K, and FIGs.14A-14D. [223] In these exemplary configurations, different VH chains and VL chains are generally referred to as VHA, VHB, VLA, or VLB, indicating that they bind to two different antigens, a or b. VHA and VLA in general indicates that the variable domain binds to antigen a; VHB and VLB generally indicate that the variable domain binds to antigen b. Sometimes, the constant domain may also need to be engineered to ensure correct heavy chain and light chain pairing. Since the engineering in constant domains generally involve limited number of mutations, different versions of constant domains are generally distinguished by CH1 versus CH1’, CH2 vs CH2’, CH3 vs CH3’, or CL vs CL’. Occasionally, complex structures such as the one that is illustrated in FIG.13K may require designations such as CH2(1), CH2(2), CH2(3), CH2(4), etc. to illustrate different constant domains that are used.
[224] In exemplary aspects, some of the bispecific molecule comprises disclosed herein have 4 antigen binding sites, 2 of which bind to 4-1BB protein and 2 of which bind to PD-L1. Optionally, each 4-1BB binding site is identical to the other and/or each PD-L1 binding site is identical to the other. [225] In one particular example, the bispecific molecule comprises an IgG moiety and a scFv moiety. As shown in FIG.14A, there are two Fab moieties that bind to one antigen (e.g., PD-L1 or 4-1BB). Each Fab moiety comprises two chains: a heavy chain comprising a heavy chain variable domain A (VHA) and a CH1 domain, and a light chain comprising a light chain variable domain A (VLA) and a CL domain. Each Fab is connected to one chain of Fc (monomeric CH2- monomeric CH3) to form an antibody (IgG). Because this part of the structure is essentially an IgG, there is no new linker between Fab and Fc (Fab and Fc are connected through “hinge” sequence just like a wildtype IgG). In addition, there are two scFv moieties that bind to the other antigen (e.g., 4-1BB or PD-L1). A first linker then connects the C-terminus of one CH3 domain to the N-terminus of one scFv. Each scFv comprises a heavy chain variable domain B (VHB) and a light chain variable domain B (VLB); and the VHB and VLB are connected via a second linker. This structure is sometimes referred to as “IgG-scFv” format (one or more scFv moieties attached to an IgG molecule). Because each target (PD-L1, 4-1BB) has two binding domains, the bispecific molecules exemplified in FIG.14A is often referred herein to as “bivalent” bispecific molecules; nonetheless, it should be noted that it is also acceptable in the art to refer to such kind of molecule as “tetravalent,” as altogether there are four binding domains. The configuration depicted in FIG.14A is also referred to IgG- scFv(C2) because two copies of scFv are attached at the C-termini of the IgG molecule. [226] In another particular example, the bispecific molecule comprises a scFv moiety that is inserted between the Fab and Fc (hinge) region of an immunoglobulin (sometimes referred to as “Fab-scFv-Fc”). For example, as shown in FIG.14B, the bispecific molecule may comprises: (i) two Fab moieties that bind to one antigen (e.g., 4-1BB or PD-L1), wherein each Fab moiety comprises two chains: a heavy chain comprising a heavy chain variable domain A (VHA) and a CH1 domain, and a light chain comprising a light chain variable domain A (VLA) and a CL domain; (ii) two scFv moieties that bind to another antigen (e.g., PD-L1 or 4-1BB), wherein each scFv comprises a heavy chain variable domain B (VHB) and a light chain variable domain B (VLB); and (iii) one Fc region that comprises two chains, each chain comprising a monomeric CH2 domain and a monomeric CH3 domain. A first linker connects the C-terminus of one CH1 domain to the N-terminus of one scFv, a second linker links the VHB and VLB of the scFv moiety, and a third linker connects the C-terminus of one scFv to the N-terminus of one chain of the Fc region. Similar to the IgG-scFv structure, the molecule exemplified in FIG.14B is essentially a tetravalent molecule, with two binding moieties for PD-L1 and two binding moieties for 4-1BB, but often called bivalent bispecific molecules. The configuration depicted in FIG.14B is also referred to Fab-scFv(M2)-Fc because two copies of scFv are inserted in the middle of the IgG molecule.
[227] Another bispecific configuration is depicted in FIG.2B. The bispecific molecule comprises an antibody (IgG) moiety and a Fab moiety. As shown in FIG.2B, there are two Fab moieties that bind to one antigen (e.g., PD-L1 or 4-1BB). Each Fab moiety comprises two chains: a heavy chain comprising a heavy chain variable domain A (VHA) and a CH1 domain, and a light chain comprising a light chain variable domain A (VLA) and a CL domain. Each Fab is connected to one chain of Fc (monomeric CH2- monomeric CH3) to form an antibody (IgG). In addition, there are two Fab moieties that bind to the other antigen (e.g., 4-1BB or PD-L1). A first linker then connects the C-terminus of one CH3 domain to the N- terminus of one Fab. Each Fab comprises a heavy chain variable domain B (VHB) and a CH1’, and a light chain variable domain B (VLB) and a CL’ domain. This particular structure is sometimes referred to as “IgG-Fab” format (one or more Fab moieties attached to an IgG molecule). [228] To ensure that the heavy chain and light chain are paired correctly for an IgG-Fab (i.e., CH1 pairs with CL, and CH1’ pairs with CL’), charge pairs may need to be created. For example, in some aspects, (i) CH1 may comprises a mutation to a positively charged residue and CL may comprise a mutation to a negatively charged residue, such that said CH1 and CL form a first charge pair (top part of Fig.2B), and (ii) CH1’ may comprise a mutation to a negatively charged residue and CL’ may comprise a mutation to a positively charged residue, such that said CH1’ and CL’ form a second charge pair (bottom part of Fig. 2B). Because the CL comprises a negatively charged residue, and the CH1’ also comprises a negatively charged residue, the likelihood of CL and CH1’ mis-pairing is unlikely. That way, mixed pairing of 4-1BB light and PD-L1 heavy chain, or vice versa, can be reduced or inhibited. [229] The positively charged residues and negatively charged residues can be introduced at multiple positions in CH1 and CL domain, as long as the residue is present on the surface of the constant domain (such that it can interact with the corresponding residue to form a charge pair), and that the mutation does not generally disrupt the conformation of the constant domain. In an exemplary embodiment, the mutation is introduced at position 183 of the CH domain (EU index numbering), and position 176 of the CL domain (Kabat numbering). [230] In general, amino acid residues in the IgG heavy constant domain of an antibody are numbered according the EU index of Edelman et al., 1969, Proc. Natl. Acad. Sci. USA 63(1):78-85 as described in Kabat et al., 1991, referred to herein as the “EU index numbering.” Typically, the constant domain comprises from residue 118 to 447, and the Fc domain comprises from residue 236 to 447 of the human lgG1 constant domain. Comparison between EU numbering and other numbering systems can be found, e.g., at IGMT database. [231] The amino acid sequence of human IgG constant region is provided below (SEQ ID NO:246), with position 183 underlined. Position 183 corresponds to residue 66 of SEQ ID NO: 265. CH1 domain comprises residues 1-98, hinge region comprises residues 99-110, CH2 domain comprises residues 111- 223, and CH3 domain comprises residues 224-330.
ASTKGPSVFP LAPSSKSTSG GTAALGCLVK DYFPEPVTVS WNSGALTSGV HTFPAVLQSS GLYSLSSVVT VPSSSLGTQT YICNVNHKPS NTKVDKKVEP KSCDKTHTCP PCPAPELLGG PSVFLFPPKP KDTLMISRTP EVTCVVVDVS HEDPEVKFNW YVDGVEVHNA KTKPREEQYN STYRVVSVLT VLHQDWLNGK EYKCKVSNKA LPAPIEKTIS KAKGQPREPQ VYTLPPSRDE LTKNQVSLTC LVKGFYPSDI AVEWESNGQP ENNYKTTPPV LDSDGSFFLY SKLTVDKSRW QQGNVFSCSV MHEALHNHYT QKSLSLSPGK [232] If the constant domain is derived from an immunoglobulin other than human IgG1, the sequence may be aligned against SEQ ID NO:246, and a mutation may be introduced at a position that corresponds to residue 66 of SEQ ID NO:246. An amino acid residue of a query sequence "corresponds to" a designated position of a reference sequence (e.g., position 66 of SEQ ID NO:246) when, by aligning the query amino acid sequence with the reference sequence, the position of the residue matches the designated position. Such alignments can be done by hand or by using well-known sequence alignment programs such as ClustalW2, or "BLAST 2 Sequences" using default parameters. [233] Amino acid residues of the light chain constant domain are numbered according to Kabat et al., 1991, "Sequences of Proteins of Immunological Interest 5th Ed.", 1991, NATIONAL INSTITUTES OF HEALTH. Kappa light chain also has EU index numbering, and the EU index and Kabat numbering are identical. Lambda light chain does not have EU index numbering. [234] The amino acid sequences of example human kappa and lambda constant regions are provided below (SEQ ID NOs:237 and 239), with position 176 underlined. Position 176 corresponds to residue 69 of SEQ ID NO: 237, and residue 69 of SEQ ID NO: 239. Note that the first residue (“R” in SEQ ID NO:237 and “G” in SEQ ID NO:239) is considered an optional junction residue. This residue sometimes is shown as the first residue in the CL domain, and sometimes as the last residue in the VL domain; both are accepted in the art. (R)TVAAPSVFI FPPSDEQLKS GTASVVCLLN NFYPREAKVQ WKVDNALQSG NSQESVTEQD SKDSTYSLSS TLTLSKADYE KHKVYACEVT HQGLSSPVTK SFNRGEC (kappa constant, SEQ ID NO:237) (G)QPKANPTVT LFPPSSEELQ ANKATLVCLI SDFYPGAVTV AWKADGSPVK AGVETTKPSK QSNNKYAASS YLSLTPEQWK SHRSYSCQVT HEGSTVEKTV APTECS (lambda constant, SEQ ID NO:239) [235] If the constant domain is derived from an immunoglobulin other than human kappa and lambda, the sequence may be aligned against SEQ ID NO:237 or 239, and a mutation may be introduced at a position that corresponds to residue 69 of SEQ ID NO:237 or 239. An amino acid residue of a query sequence "corresponds to" a designated position of a reference sequence (e.g., position 69 of SEQ ID NO:237 or 239) when, by aligning the query amino acid sequence with the reference sequence, the position of the
residue matches the designated position. Such alignments can be done by hand or by using well-known sequence alignment programs such as ClustalW2, or "BLAST 2 Sequences" using default parameters. [236] Positively charged residues include lysine, arginine and histidine. In an exemplary embodiment, lysine (K) is used. In an exemplary embodiment, arginine (R) is used. Negatively charged residues include aspartic acid and glutamic acid. In an exemplary embodiment, glutamic acid (E) is used. [237] Other positions may also be used to introduce charge pairs. For example, positions 123 and 124 of the CL domains and positions 147 and 213 of CH1 domains may be replaced by positively charged residues or negatively charged residues. Further two or more residues in a constant domain may be changed to introduce additional charge pairs. [238] In another exemplary configuration, the bispecific molecule comprises two VH-only binding domains inserted between the Fab and Fc (hinge) region of an immunoglobulin (FIG.14D), sometime referred to as Fab-VH-Fc or Fab-VH(M2)-Fc. A first linker links the C-terminus of the Fab with the N- terminus of the VH, and a second linker linkers the C-terminus of the VH with the N-terminus of the Fc. [239] In another exemplary configuration, the bispecific molecule comprises two VH-only binding domains, each linked to one C-terminus of an IgG heavy chain (FIG.14C). This configuration is also referred to as IgG-VH or IgG-VH(C2). [240] In another exemplary configuration, the bispecific molecule comprises one scFv attached to the C- terminus of one of the two IgG heavy chains (FIG.13C). This configuration involves bivalent binding of one antigen, and monovalent binding of the second antigen, and referred to as IgG-scFv(C1). The two light chains are identical, whereas the two heavy chains are asymmetric. [241] In another exemplary configuration, the bispecific molecule comprises one scFv inserted between the CH1 domain and CH2 domain (hinge region) of one of the two IgG heavy chains (FIG.13D). This configuration involves bivalent binding of one antigen, and monovalent binding of the second antigen, and referred to as Fab-scFv(M1)-Fc. The two light chains are identical, whereas the two heavy chains are asymmetric. [242] In another exemplary configuration, the bispecific molecule is a hetero-IG, comprising one scFv linked to one CH2-CH3 chain, and one Fab linked to the other CH2-CH3 chain (FIG.13G). This configuration involves monovalent binding of one antigen through one Fab, and monovalent binding of the second antigen through one scFv. The two heavy chains are asymmetric and there is only one light chain. The configuration is also referred to as [Fab*scFv] hetero-Fc. [243] In another exemplary configuration, the bispecific molecule comprises one VH-only binding domain linked to one CH2-CH3 chain, and one Fab linked to the other CH2-CH3 chain (FIG.13H). This
configuration involves monovalent binding of one antigen through one Fab, and monovalent binding of the second antigen through one VH-only binding domain. The two heavy chains are asymmetric and there is only one light chain. The configuration is also referred to as [Fab*VH] hetero-Fc. [244] In another exemplary configuration, the bispecific molecule comprises one scFv attached to the N- terminus of one of the two IgG heavy chains (FIG.13J). This configuration involves bivalent binding of one antigen, and monovalent binding of the second antigen, and is referred to as scFv(N1)-IgG because only one copy of the scFv is attached to the N-terminus of the IgG. The two light chains are identical, whereas the two heavy chains are asymmetric. [245] FIG.13K illustrates another configuration of a bispecific molecule, referred to as ([scFv*Fab] hetero-Fc)-Fc. The molecule comprises four chains, and is essentially a [scFv*Fab] hetero-Fc (Fig.13K) structure linked to a second Fc domain. The molecule comprises (i) a first heavy chain that comprises a scFv linked to a CH2(1)-CH3(1) chain, (ii) a second heavy chain that comprise an IgG heavy chain (with CH2(2)-CH3(2)) linked to a CH2(3)-CH3(3) (i.e., VH-CH1-CH2(2)-CH3(2)-CH2(3)-CH3(3)), (iii) a light chain, and (iv) a CH2(4)-CH3(4) chain. The CH2(1)-CH3(1) from (i) and the CH2(2)-CH3(2) from (ii) form a first Fc domain, and the CH2(3)-CH3(3) from (ii) and the CH2(4)-CH3(4) from (iv) form a second Fc domain. Each CH2-CH3 chain would require certain mutations to create certain charge pairs, such as the two Fc domains are paired correctly. [246] FIG.13I illustrates another configuration, essentially a hetero-IgG molecule. In this configuration, one heavy chain and one light chain form a Fab that binds to one antigen, and a second heavy chain and a second light chain form a second Fab that binds to a second antigen. The molecule is an asymmetric molecule with one 4-1BB binding domain and one PD-L1 binding domain. To ensure that the heavy chain and light chain are paired correctly (i.e., CH1 pairs with CL, and CH1’ pairs with CL’), charge pairs may need to be created in CH1, CH1’, CL, and CL’, as disclosed in detail above. [247] In addition, additional mutations need to be created such as two identical heavy chains do not pair with each other in the Fc region, such that an asymmetric IgG can be created. In some embodiments, one of the heavy chains uses wild type CH3 sequences, and the other heavy chain comprises mutations in which a positively charged residue in the original wild type is mutated to a negatively charged residue to promote heterologous chain pairing. [248] Referring to SEQ ID NO:246 again, CH3 domain comprises residues 224-330. In certain embodiments, a mutation from K to E or D (preferably D) at position 392 (EU index numbering), or at a position that corresponds to residue 275 of SEQ ID NO:246 if EU index is not available, is introduced. In certain embodiments, a mutation from K to D or E (preferably D) at position 409 (EU index numbering), or at a position that corresponds to residue 292 of SEQ ID NO:246 if EU index is not available, is introduced. In certain embodiments, a mutation from K to E or D (preferably D) at position 439 (EU index numbering),
or at a position that corresponds to residue 322 of SEQ ID NO:246 if EU index is not available, is introduced. These K residues are underline below. ASTKGPSVFP LAPSSKSTSG GTAALGCLVK DYFPEPVTVS WNSGALTSGV HTFPAVLQSS GLYSLSSVVT VPSSSLGTQT YICNVNHKPS NTKVDKKVEP KSCDKTHTCP PCPAPELLGG PSVFLFPPKP KDTLMISRTP EVTCVVVDVS HEDPEVKFNW YVDGVEVHNA KTKPREEQYN STYRVVSVLT VLHQDWLNGK EYKCKVSNKA LPAPIEKTIS KAKGQPREPQ VYTLPPSRDE LTKNQVSLTC LVKGFYPSDI AVEWESNGQP ENNYKTTPPV LDSDGSFFLY SKLTVDKSRW QQGNVFSCSV MHEALHNHYT QKSLSLSPGK [249] A “linker” is a molecule or group of molecules that connects two separate entities (e.g., 4-1BB binding protein and PD-L1 binding protein) to one another and can provide spacing and flexibility between the two entities such that they are able to achieve a conformation in which they, e.g., specifically bind their respective targets (e.g., 4-1BB and PD-L1). Protein linkers are particularly preferred, and they may be expressed as a component of the recombinant protein using standard recombinant DNA techniques well-known in the art. For recombinant proteins described herein comprising two or more linkers (for example IgG-scFv and Fab-scFv-Fc formats), the linkers may all be the same, or some or all of the linkers may be different from each other. [250] In some embodiments, the linker is a peptidyl linker. In some embodiments, the peptidyl linker comprises about 1 to 30 amino acid residues. Exemplary linkers include, e.g., a glycine rich peptide; a peptide comprising glycine and serine; a peptide having a sequence [Gly-Gly-Ser]n (SEQ ID NO:436), wherein n is 1, 2, 3, 4, 5, or 6; or a peptide having a sequence [Gly-Gly-Gly-Gly-Ser]n (SEQ ID NO: 389), wherein n is 1, 2, 3, 4, 5, or 6. A glycine rich peptide linker comprises a peptide linker, wherein at least 25% of the residues are glycine. Glycine rich peptide linkers are well known in the art (e.g., Chichili et al. Protein Sci.2013 February; 22(2): 153-167). The peptidyl linker may also be a proline-threonine rich peptide linker. [251] As shown in FIGs.2A-2B, FIGs.13A-13K, and FIGs.14A-14D, when bispecific molecule comprises a scFv moiety, mutations may be introduced to scFv to further improve stability. For example, it has been reported that insufficient interface stability between the heavy and light chains of scFv fragments could be the main cause of irreversible scFv inactivation. Fv fragments have been reported to dissociate into heavy-chain variable domains (VH) and light-chain variable domains (VL) with KD values ranging from 10−9 to 10−6 M. An interdomain disulfide bond have been used to further improve scFv stability. For example, mutation to Cys at the site of H44 (Kabat numbering), and mutation to Cys at L100 (Kabat numbering) would not significantly affect the domain folding. The two cysteines can then form an intramolecular disulfide bond to further stabilize the scFv. Such mutation is sometimes referred to as “cysteine clamp.”
[252] Specific examples of scFv comprising cysteine clamps are shown in Sequence Tables C and D, where mutations at H44 (Kabat numbering) and at L100 (Kabat numbering) were used to create disulfide bonds (referred to as “C-C”). Specific examples of scFv comprising cysteine clamp in a bispecific format are shown in Sequence Table H and K6. [253] A cysteine clamp has been introduced in some of the scFv for IgG-scFv bispecific molecules exemplified herein. In some of the Fab-scFv-Fc bispecific molecules, cysteine clamp is not present. Thus, the use (or non-use) of cysteine clamp may require evaluation of stability and biologically activities of the scFv. In general, it is believed that the removal of the constant domain (CH1 and Cλ or Cκ) lowers the stability of the Fv domain. This may require the addition of a linker fusion between the VH and VL domains to avoid molecule dissociation. With the decreased interface between the HC and LC, some Fv domains may have an increased probability of being in a dissociated state, exposing their hydrophobic VH and VL interfaces. This could cause increased aggregation, and require additional stability using a disulfide bond or cysteine clamp that covalently links the VH to the VL. On the other hand, although cysteine clamp tends to create a stabilized product post purification, it could also lead to other issues. A poorly positioned cys-clamp can alter the orientation of the VH and VL domains such that it exposes new interfaces or cause a loss in antigen binding due to a new paratope interface. If possible, a scFv domain lacking a cys-clamp with good biophysical properties could be preferential. In IgG-scFv format, because scFv is located at the C-terminus and more exposed, it appears that a cys-clamp can improve the biophysical properties of scFv, whereas in the Fab-scFv-Fc format, it appears that the scFv without cysteine clamp may be feasible under some circumstances because the scFv is sandwiched between Fab and Fc and thus more protected. [254] In some embodiments, the bispecific molecule disclosed herein comprises a CH1 domain, preferably a human CH1 domain (such as a human IgG1 CH1, a human IgG2 CH1, a human IgG3 CH1, or a human IgG4 CH1), as discussed in detail above. Non-limiting examples of human CH1 sequences are provided in the Sequence Tables. In some embodiments, the CH1 domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 242, 259, 261, 265, 269, 422, or 425. [255] In certain embodiments, the bispecific molecule disclosed herein comprises an Fc domain, as discussed in detail above. The Fc domain can be derived from IgA (e.g., IgA1 or lgA2), IgG, IgE, or IgG (e.g., IgG1, lgG2, lgG3, or lgG4). In some embodiments, the Fc domain comprises wild type sequence of a human Fc domain. Non-limiting examples of human Fc domain sequences are provided in the Sequence Table.
[256] In some embodiments, the Fc domain is the Fc domain of human lgG1 and comprises one or more of the following effector-null mutations: L234A, L235A, and G237A (numbering according to the EU index), often referred as “LALA” mutations. [257] In exemplary embodiments, the Fc region comprises a Stable Effector Functionless (SEFL) mutation to reduce the ability to interact with Fcγ receptors. In exemplary aspects, the SEFL mutation comprises one or more of the following mutations, numbered according to the EU system: L242C, A287C, R292C, N297G, V302C, L306C, and/or K334C. In exemplary aspects, the SEFL mutation comprises N297G. In exemplary aspects, the SEFL mutation comprises A287C, N297G, and L306C. In other exemplary aspects, the SEFL mutation comprises R292C, N297G, and V302C (i.e., SEFL2-2). [258] In exemplary embodiments, the Fc region comprises a YTE mutation. The M252Y/S254T/T256E (EU index numbering, referred to “YTE”) triple mutation have been shown to increase IgG half-life in cynomolgus monkeys by an approximate 4-fold increase. [259] In exemplary embodiments, the bispecific molecule disclosed herein comprise Fc that is derived from an IgG1. In some embodiments, the Fc comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 243, 250, 251, 252, 253, 254, 255, 413, 423, 426, 263, 267, or 483. [260] In exemplary embodiments, the bispecific molecule disclosed herein comprise an IgG1 heavy chain constant domain. In some embodiments, the heavy chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 90%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 244, 245, 246, 247, 248, 249, 256, 257, 258, 260, 412, 415, 417, 418, 427, 264, 268, or 271. [261] In some embodiments, the bispecific molecule disclosed herein comprising a kappa or lambda light chain constant domain. In some embodiments, the kappa light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 236, 237, 240, 414, 416, 419, 420, 421, or 428. In some embodiments, the lambda light chain constant domain comprises a sequence that is at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 86%, at least 87%, at least 88%, at least 89%, at least 90%, at
least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO: 238, 239, 241, or 424. [262] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:283; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:220. The bispecific molecule may comprise two copies of each sequence. [263] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:287; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:206. The bispecific molecule may comprise two copies of each sequence. [264] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:290; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:184. The bispecific molecule may comprise two copies of each sequence. [265] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:294; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:186. The bispecific molecule may comprise two copies of each sequence. [266] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:297; (ii) an amino acid sequence at least 60%, at least 65%, at
least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:299; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:300. The bispecific molecule may comprise two copies of each sequence. [267] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:304; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:306; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:307. The bispecific molecule may comprise two copies of each sequence. [268] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:311; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:312; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:307. The bispecific molecule may comprise two copies of each sequence. [269] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:315; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:312; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at
least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:317. The bispecific molecule may comprise two copies of each sequence. [270] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:320; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:312; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:300. The bispecific molecule may comprise two copies of each sequence. [271] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:443; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:444. The bispecific molecule may comprise two copies of each sequence. [272] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:434; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:435. The bispecific molecule may comprise two copies of each sequence. [273] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:449; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:450. The bispecific molecule may comprise two copies of each sequence.
[274] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:451; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:452. The bispecific molecule may comprise two copies of each sequence. [275] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:449; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:450; (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:439; and (iv) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:440. [276] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:432; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:433. The bispecific molecule may comprise two copies of each sequence. [277] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:446; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:447; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:448.
[278] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:429; (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:430; and (iii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:431. [279] In exemplary embodiments, the bispecific molecule comprises: (i) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:442; and (ii) an amino acid sequence at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:441. The bispecific molecule may comprise two copies of each sequence. 2.6 Binding Characteristics [280] The antigen binding proteins provided herein bind to their respective targets or antigens in a non- covalent and reversible manner. In exemplary embodiments, the binding strength of the antigen binding proteins to their targets or antigens (e.g., 4-1BB or PD-L1) may be described in terms of its affinity, a measure of the strength of interaction between the binding site of the antigen binding protein and the target or antigen (e.g., 4-1BB or PD-L1). In exemplary aspects, the antigen binding proteins provided herein have high-affinity for their target or antigen (e.g., 4-1BB or PD-L1) and thus will bind a greater amount of the target or antigen (e.g., 4-1BB or PD-L1) in a shorter period of time than low-affinity antigen binding proteins. In exemplary aspects, the antigen binding proteins provided herein have low-affinity for 4-1BB or PD-L1 and thus will bind a lesser amount of 4-1BB or PD-L1 in a longer period of time than high-affinity antigen binding proteins. In exemplary aspects, the antigen binding protein has an equilibrium association constant, KA, which is at least 105 M-1, at least 106 M-1, at least 107 M-1, at least 108 M-1, at least 109 M-1, at least 1010 M-1, at least 1011 M-1, at least 1012 M-1, at least 1013 M-1, or at least 1014 M-1. As understood by the artisan of ordinary skill, KA can be influenced by factors including pH, temperature and buffer composition. [281] In exemplary embodiments, the binding strength of the antigen binding protein to its target or antigen (e.g., 4-1BB or PD-L1) may be described in terms of its sensitivity. KD is the equilibrium dissociation constant, a ratio of koff/kon, between the antigen binding protein and its target or antigen (e.g., 4-1BB or PD-L1). KD and KA are inversely related. The KD value relates to the concentration of the
antigen binding protein (the amount of antigen binding protein needed for a particular experiment) and so the lower the KD value (lower concentration needed) the higher the affinity of the antigen binding protein. In exemplary aspects, the binding strength of the antigen binding protein to its target (e.g., 4-1BB or PD- L1) may be described in terms of KD. In exemplary aspects, the KD value of the antigen binding proteins provided herein is about 10-1 M or less, about 10-2 M or less, about 10-3 M or less, about 10-4 M or less, about 10-5 M or less, about 10-6 M or less, about 10-7 M or less, about 10-8 M or less, about 10-9 M or less, about 10-10 M or less, about 10-11 M or less, about 10-12 M or less, about 10-13 M or less, about 10-14 M or less, from about 10-5 M to about 10-15 M, from about 10-6 M to about 10-15 M, from about 10-7 M to about 10-15 M, from about 10-8 M to about 10-15 M, from about 10-9 M to about 10-15 M, from about 10-10 M to about 10-15 M, from about 10-5 M to about 10-14 M, from about 10-6 M to about 10-14 M, from about 10-7 M to about 10-14 M, from about 10-8 M to about 10-14 M, from about 10-9 M to about 10-14 M, from about 10-10 M to about 10-14 M, from about 10-5 M to about 10-13 M, from about 10-6 M to about 10-13 M, from about 10- 7 M to about 10-13 M, from about 10-8 M to about 10-13 M, from about 10-9 M to about 10-13 M, or from about 10-10 M to about 10-13 M. [282] In exemplary aspects, the KD of the antigen binding proteins provided herein is micromolar, nanomolar, picomolar or femtomolar. In exemplary aspects, the KD of the antigen binding proteins provided herein is within a range of about 10-4 to 10-6 M, or 10-7 to 10-9 M, or 10-10 to 10-12 M, or 10-13 to 10-15 M. In exemplary aspects, the antigen binding protein binds to the human 4-1BB or PD-L1 with a KD value that is from about 0.07 nM to about 4 nM. In exemplary aspects, the antigen binding protein binds to the human 4-1BB or PD-L1 with a KD of from about 0.01 nM to about 50 nM, from about 0.02 nM to about 50 nM, from about 0.05 nM to about 50 nM, from about 0.05 nM to about 45 nM, from 0.05 nM to about 40 nM, from about 0.05 nM to about 35 nM, from about 0.05 nM to about 30 nM, from about 0.05 nM to about 25 nM, from about 0.05 nM to about 20 nM, from about 0.05 nM to about 15 nM, or from about 0.05 nM to about 10 nM. In exemplary aspects, the antigen binding protein binds to the cynomolgus monkey 4-1BB or PD-L1 with a KD that is from about 0.05 nM to about 4 nM. In exemplary aspects, the antigen binding protein binds to the cynomolgus monkey 4-1BB or PD-L1 with a KD of from about 0.01 nM to about 50 nM, from about 0.02 nM to about 50 nM, from about 0.05 nM to about 50 nM, from about 0.05 nM to about 45 nM, from 0.05 nM to about 40 nM, from about 0.05 nM to about 35 nM, from about 0.05 nM to about 30 nM, from about 0.05 nM to about 25 nM, from about 0.05 nM to about 20 nM, from about 0.05 nM to about 15 nM, or from about 0.05 nM to about 10 nM. In exemplary embodiments, the 4-1BB antigen binding protein binds to human 4-1BB with a KD value of from about 0.05 nM to about 5 nM. In exemplary embodiments, the 4-1BB antigen binding protein binds to cynomolgus monkey 4-1BB with a KD value of from about 0.05 nM to about 5 nM. In exemplary embodiments, the PD-L1 antigen binding protein binds to human PD-L1 with a KD value of from about 0.05 nM to about 5 nM. In exemplary embodiments, the PD-L1 antigen binding protein binds to cynomolgus monkey PD-L1 with a KD value of from about 0.05 nM to about 5 nM.
[283] KD values can be determined using methods well established in the art. One exemplary method for measuring KD is surface plasmon resonance (SPR), a method well-known in the art (e.g., Nguyen et al. Sensors (Basel).2015 May 5; 15(5):10481-510). KD value may be measured by SPR using a biosensor system such as a BIACORE® system. BIAcore kinetic analysis comprises analyzing the binding and dissociation of an antigen from chips with immobilized molecules (e.g. molecules comprising epitope binding domains), on their surface. Another well-known method in the art for determining the KD of a protein is by using Bio-Layer Interferometry (e.g., Shah et al. J Vis Exp.2014; (84): 51383). KD value may be measured by Bio-Layer Interferometry using OCTET® technology (Octet QKe system, ForteBio). Alternatively or in addition, a KinExA® (Kinetic Exclusion Assay) assay, available from Sapidyne Instruments (Boise, Id.) can also be used. Any method known in the art for assessing the binding affinity between two binding partners is encompassed herein. [284] In some aspects, the KD value is measured by surface plasmon resonance (SPR). Antigen (e.g., 4- 1BB or PD-L1) may be immobilized, e.g., on a solid surface. The antigen may be immobilized to a chip, for example by covalent coupling (such as amine coupling). The chip may be a CM5 sensor chip. As the analyte binds to the ligand the accumulation of protein on the sensor surface causes an increase in refractive index. This refractive index change is measured in real time (sampling in a kinetic analysis experiment is taken every 0.1 s), and the result plotted as response units (RU) versus time (termed a sensorgram). A response (background response) will also be generated if there is a difference in the refractive indices of the running and sample buffers. This background response must be subtracted from the sensorgram to obtain the actual binding response. The background response is recorded by injecting the analyte through a control or reference flow cell, which has no ligand or an irrelevant ligand immobilized to the sensor surface. The real-time measurement of association and dissociation of a binding interaction allows for the calculation of association and dissociation rate constants and the corresponding affinity constants. One RU represents the binding of 1 pg of protein per square mm. More than 50 pg per square mm of analyte binding is generally needed in practice to generate good reproducible responses. [285] Dissociation of the antigen-binding protein from the antigen may be monitored for about 3600 seconds. The SPR analysis may be conducted, and the data collected at between about 15°C and about 37°C. The SPR analysis may be conducted, and the data collected at between about 25°C and 37°C. The SPR analysis may be conducted, and the data collected at about 37°C. The SPR analysis may be conducted, and the data collected at 37°C. The KD value may be measured by SPR using a BIAcore T200 instrument. The SPR rates and affinities may be determined by fitting resulting sensorgram data to a 1:1 model in BIAcore T200 Evaluation software version 1.0. The collection rate may be about 1 Hz. [286] Another method for determining the KD of an antibody is by using Bio-Layer Interferometry (BLI), typically using OCTET® technology (Octet QKe system, ForteBio). In some embodiments, biosensor
analysis is used. Typically, one interactant is immobilized on the surface of the biosensor ("ligand," such as an antigen-binding protein) and the other remains in solution (“analyte”, such as an antigen). The assay begins with an initial baseline or equilibration step using assay buffer. Next, a ligand (such as an antigen-binding protein) is immobilized on the surface of the biosensor (loading), either by direct immobilization or capture-based method. After ligand immobilization, biosensors are dipped into buffer solution for a baseline step to assess assay drift and determine loading level of ligand. After the baseline step, biosensors are dipped into a solution containing the ligand's binding partner, the analyte (association). In this step, the binding interaction of the analyte to the immobilized ligand is measured. Following analyte association, the biosensor is dipped into buffer solution without analyte, and the bound analyte is allowed to come off the ligand (dissociation). The series of assay steps is then repeated on new or regenerated biosensors for each analyte being tested. Each binding response is measured and reported in real time on a sensorgram trace. The instrument may be Octet QKe system, Octet RED96 system, Octet QK384 system, or RED384 system. [287] In certain embodiments, the 4-1BB binding protein binds human 4-1BB with a KD value of or less than: about 200nM, about 150nM, about 100nM, about 90nM, about 80nM, about 70nM, about 60nM, about 50nM, about 40nM, about 30nM, about 25nM, about 20nM, about 15nM, about 10nM, about 9nM, about 8nM, about 7nM, about 6nM, about 5nM, about 4nM, about 3nM, about 2nM, about 1 nM, about 900pM, about 800pM, about 700pM, about 600pM, about 500pM, about 400pM, about 300pM, about 250pM, about 200pM, about 150pM, about 100pM, about 50pM, about 40pM, about 30pM, about 25pM, about 20pM, about 15pM, about 10pM, about 5pM, or about 1pM. KD value may be measured by surface plasmon resonance (SPR) (e.g., a Biacore T200 instrument); or it may be measured by bio-layer interferometry (BLI) (e.g., a ForteBio Octet instrument). [288] In certain embodiments, the PD-L1 binding protein binds human PD-L1 with a KD value of or less than: about 200nM, about 150nM, about 100nM, about 90nM, about 80nM, about 70nM, about 60nM, about 50nM, about 40nM, about 30nM, about 25nM, about 20nM, about 15nM, about 10nM, about 9nM, about 8nM, about 7nM, about 6nM, about 5nM, about 4nM, about 3nM, about 2nM, about 1 nM, about 900pM, about 800pM, about 700pM, about 600pM, about 500pM, about 400pM, about 300pM, about 250pM, about 200pM, about 150pM, about 100pM, about 50pM, about 40pM, about 30pM, about 25pM, about 20pM, about 15pM, about 10pM, about 5pM, or about 1pM. KD value may be measured by surface plasmon resonance (SPR) (e.g., a Biacore T200 instrument); or it may be measured by bio-layer interferometry (BLI) (e.g., a ForteBio Octet instrument). [289] One particular observation made by the inventors is that for 4-1BB agonists, low affinity (rather than high affinity) binders delivered greater activity through increased clustering. For example, our internal data demonstrated that one of the UniDab-based 4-1BB agonist has much lower affinity to human 41BB (~1000-fold lower affinity) compared to a reference molecule. Surprisingly, this molecule shows
higher activity (about 2-fold higher potency) compared to the reference molecule in vitro. Unlike natural and direct-targeting antibodies, the 4-1BB agonists disclosed herein need to be able to induce 4-1BB clustering in addition to specific target engagement. It is possible that lower affinity binders can trigger sufficient agonism but not negative feedback mechanisms. [290] Exemplary method to measure binding affinity is also provided in Examples. 2.7 Cross-Reactivity [291] In various aspects, the antigen binding protein binds to human 4-1BB. A reference amino acid sequence of human 4-1BB is provided herein as SEQ ID NO: 272 (full length precursor) and SEQ ID NO: 274 (extracapsular domain). In various aspects, the antigen binding protein binds to cynomolgus monkey (cyno) 4-1BB. The amino acid sequence of cyno 4-1BB is provided herein as SEQ ID NO: 275 (precursor). In exemplary aspects, the antigen binding protein binds with high affinity to both human 4- 1BB and cyno 4-1BB. In various embodiments, the antigen-binding proteins of the present disclosure bind to human 4-1BB and cyno 4-1BB but do not cross-react with any other 4-1BB orthologs. In various instances, the antigen binding protein binds with high affinity to both human 4-1BB and cyno 4-1BB and does not bind to any other 4-1BB ortholog, e.g., does not bind to mouse 4-1BB, rat 4-1BB, canine 4-1BB, bovine 4-1BB, and the like. In various embodiments, the antigen-binding proteins of the present disclosure have a selectivity for human and cyno 4-1BB which is at least 10-fold, 5-fold, 4-fold, 3-fold, 2- fold greater than the selectivity of the antigen-binding protein for any other 4-1BB ortholog. In various embodiments, the antigen-binding proteins of the present disclosure have a KD for human and cyno 4- 1BB which is at least 10-fold, 5-fold, 4-fold, 3-fold, 2-fold less than the KD of the antigen-binding protein for any other 4-1BB ortholog. In some embodiments, the antigen binding protein binds to human 4-1BB but does not bind to cyno 4-1BB. [292] In various instances, the antigen binding protein binds to human PD-L1. The amino acid sequence of human PD-L1 is provided herein as SEQ ID NO: 277. In various aspects, the antigen binding protein binds to cyno PD-L1. The amino acid sequence of cyno PD-L1 is provided herein as SEQ ID NO: 279. In exemplary aspects, the antigen binding protein binds with high affinity to both human PD-L1and cyno PD- L1. In various embodiments, the antigen-binding proteins of the present disclosure bind to human PD-L1 and cyno PD-L1 do not cross-react with any other PD-L1 orthologs. In various instances, the antigen binding protein binds with high affinity to both human PD-L1and cyno PD-L1 and does not bind to any other PD-L1 ortholog, e.g., does not bind to mouse PD-L1, rat PD-L1, canine PD-L1, or bovine PD-L1. In various embodiments, the antigen-binding proteins of the present disclosure have a KD value for human and cyno PD-L1 which is numerically at least 10-fold, 5-fold, 4-fold, 3-fold, 2-fold less than the KD of the antigen-binding protein for any other PD-L1 ortholog. In some embodiments, the antigen binding protein binds to human PD-L1 but does not bind to cyno PD-L1.
2.8 Competition assays [293] Also encompassed herein are antigen-binding proteins that compete with the 4-1BB antigen binding proteins disclosed herein (such as those exemplified in the sequence tables) for the same target, and antigen-binding proteins that compete with the PD-L1 antigen binding proteins disclosed herein (such as those exemplified in the sequence tables) for the same target. [294] In various embodiments, the antigen-binding protein inhibits a binding interaction between human 4-1BB and a reference antibody, which reference antibody is known to bind to 4-1BB. By way of example, the reference antibody may be a 4-1BB antigen-binding protein disclosed herein, such as those disclosed in the Sequence Tables. In various instances, a 4-1BB antigen-binding protein competes with the reference antibody for binding to human 4-1BB and thereby reduce the amount of human 4-1BB bound to the reference antibody as determined by an in vitro competitive binding assay. In various aspects, the 4- 1BB antigen-binding protein inhibits the binding interaction between human 4-1BB and the reference antibody and the inhibition is characterized by an IC50. In various aspects, the 4-1BB antigen-binding protein exhibits an IC50 of less than about 2500 nM for inhibiting the binding interaction between human 4- 1BB and the reference antibody. In various aspects, the antigen-binding proteins exhibit an IC50 of less than about 2000 nM, less than about 1500 nM, less than about 1000 nM, less than about 900 nm, less than about 800 nm, less than about 700 nm, less than about 600 nm, less than about 500 nm, less than about 400 nm, less than about 300 nm, less than about 200 nm, or less than about 100 nm. In various aspects, the antigen-binding proteins exhibit an IC50 of less than about 90 nM, less than about 80 nM, less than about 70 nM, less than about 60 nM, less than about 50 nM, less than about 40 nM, less than about 30 nM, less than about 20 nM, or less than about 10 nM. In various instances, the 4-1BB antigen- binding protein competes with the reference antibody for binding to human 4-1BB and thereby reduce the amount of human 4-1BB bound to the reference antibody as determined by a FACS-based assay in which the fluorescence of a fluorophore-conjugated secondary antibody which binds to the Fc of the reference antibody is measured in the absence or presence of a particular amount of the 4-1BB antigen- binding protein. In various aspects, the FACS-based assay is carried out with the reference antibody, fluorophore-conjugated secondary antibody and cells which express 4-1BB. In various aspects, the cells are genetically-engineered to overexpress 4-1BB. In some aspects, the cells are HEK293T cells transduced with a viral vector to express 4-1BB. In alternative aspects, the cells endogenously express 4- 1BB. Before the FACS-based assay is carried out, in some aspects, the cells which endogenously express 4-1BB are pre-determined as low 4-1BB-expressing cells or high 4-1BB-expressing cells. [295] In various embodiments, the antigen-binding protein inhibits a binding interaction between human PD-L1 and a reference antibody, which reference antibody is known to bind to PD-L1. By way of example, the reference antibody may be a PD-L1 antigen-binding protein disclosed herein, such as those disclosed in the Sequence Tables. In various instances, a PD-L1 antigen-binding protein competes with the
reference antibody for binding to human PD-L1 and thereby reduce the amount of human PD-L1 bound to the reference antibody as determined by an in vitro competitive binding assay. In various aspects, the PD-L1 antigen-binding protein inhibits the binding interaction between human PD-L1 and the reference antibody and the inhibition is characterized by an IC50. In various aspects, the PD-L1 antigen-binding protein exhibits an IC50 of less than about 2500 nM for inhibiting the binding interaction between human PD-L1 and the reference antibody. In various aspects, the antigen-binding proteins exhibit an IC50 of less than about 2000 nM, less than about 1500 nM, less than about 1000 nM, less than about 900 nm, less than about 800 nm, less than about 700 nm, less than about 600 nm, less than about 500 nm, less than about 400 nm, less than about 300 nm, less than about 200 nm, or less than about 100 nm. In various aspects, the antigen-binding proteins exhibit an IC50 of less than about 90 nM, less than about 80 nM, less than about 70 nM, less than about 60 nM, less than about 50 nM, less than about 40 nM, less than about 30 nM, less than about 20 nM, or less than about 10 nM. In various instances, the PD-L1 antigen- binding protein competes with the reference antibody for binding to human PD-L1 and thereby reduce the amount of human PD-L1 bound to the reference antibody as determined by a FACS-based assay in which the fluorescence of a fluorophore-conjugated secondary antibody which binds to the Fc of the reference antibody is measured in the absence or presence of a particular amount of the PD-L1 antigen- binding protein. In various aspects, the FACS-based assay is carried out with the reference antibody, fluorophore-conjugated secondary antibody and cells which express PD-L1. In various aspects, the cells are genetically-engineered to overexpress PD-L1. In some aspects, the cells are HEK293T cells transduced with a viral vector to express PD-L1. In alternative aspects, the cells endogenously express PD-L1. Before the FACS-based assay is carried out, in some aspects, the cells which endogenously express PD-L1 are pre-determined as low PD-L1 -expressing cells or high PD-L1-expressing cells. [296] Other binding assays, e.g., competitive binding assays or competition assays, which test the ability of one antigen-binding molecule (such as an antibody) to compete with a second antigen binding molecule (such as a second antibody) for binding to an antigen, or to an epitope thereof, are known in the art. See, e.g., Trikha et al., Int J Cancer 110: 326-335 (2004); Tam et al., Circulation 98(11): 1085-1091 (1998). U.S. Patent Application Publication No. US20140178905, Chand et al., Biologicals 46: 168-171 (2017); Liu et al., Anal Biochem 525: 89-91 (2017); and Goolia et al., J Vet Diagn Invest 29(2): 250-253 (2017). Also, other methods of comparing two antigen binding molecules are known in the art, and include, for example, surface plasmon resonance (SPR). SPR can be used to determine the binding constants of the two binding molecules, and the two binding constants can be compared. 3. Nucleic Acids, Vectors, and Host Cells 3.1 Nucleic acids [297] The present disclosure further provides nucleic acids comprising a nucleotide sequence encoding the antigen-binding proteins disclosed herein. The nucleic acid may comprise a single nucleic acid
molecule, or two or more nucleic acid molecules (for example, a first nucleic acid molecule encoding a heavy chain amino acid sequence and a second nucleic acid molecule encoding a light chain amino acid sequence). In some aspects, the nucleic acids of the present disclosure are recombinant. [298] In exemplary embodiments, the nucleic acid comprises a nucleotide sequence that has at least about 50%, at least about 60%, at least about 70%, at least about 80%, at least about 85%, at least about 90%, or has greater than about 90% (e.g., about 91%, about 92%, about 93%, about 94%, about 95%, about 96%, about 97%, about 98%, about 99%, or 100%) sequence identity to any one of SEQ ID NOs: 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465. It would be immediately apparent to a skill artisan which part of SEQ ID NOs: 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465 encode the amino acid sequences corresponding to the fragments listed in the sequence tables. [299] In addition, nucleic acid sequence that encoding a signal peptide may be added to the 5’ of 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465. Recombinant expression of the bispecific molecules, as well as various antigen-binding proteins (or antigen-binding moieties) disclosed herein often require that the molecules be secreted. Translocation of a nascent protein from the cytosol into the ER mediated by its signal peptide is an important step in protein secretion. It is understood that the signal peptide is present (and often critical) during the initial synthesis of a nascent protein, but then, signal peptide is cleaved during secretion process. Therefore, while the mature protein no longer has the signal peptide; having the signal peptide coding sequence in the nucleic acid is generally necessary to recombinantly express the protein. [300] The present disclosure further provides nucleic acids that are capable of hybridizing to any one of SEQ ID NOs: 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465, or a complimentary sequence of any one of SEQ ID NOs: 285, 286, 288, 289, 292, 293, 295, 296, 301, 302, 303, 308, 309, 310, 313, 314, 318, 319, 321, 466, 467, 458, 459, 471, 472, 473, 505, 460, 462, 461, 463, 456, 457, 468, 469, 470, 453, 454, 455, 464, and 465, under a moderately stringent condition, or under a highly stringent condition. A “moderately stringent condition” includes prewashing in a solution of 5X SSC, 0.5% SDS, 1.0 mM EDTA (pH 8.0); hybridizing at 50 °C-65 °C, 5X SSC, overnight; followed by washing twice at 65°C for 20 minutes with each of 2X, 0.5X and 0.2X SSC containing 0.1 % SDS. A “highly stringent condition” includes, for example, (1) employ low ionic strength and high temperature for washing, for example 0.015M sodium chloride/0.0015M sodium citrate/0.5 % sodium dodecyl sulfate at 50°C; (2) employ during hybridization a denaturing agent, such as formamide, for
example, 50% (v/v) formamide with 0.1% bovine serum albumin/0.1% Ficoll/0.1% polyvinylpyrrolidone/50 mM sodium phosphate buffer at pH 6.5 with 750 mM sodium chloride, 75 mM sodium citrate at 42 °C; or (3) employ 50% formamide, 5XSSC (0.75 M NaCI, 0.075 M sodium citrate), 50 mM sodium phosphate (pH 6.8), 0.1% sodium pyrophosphate, 5X Denhardt's solution, sonicated salmon sperm DNA (50 μg/ml), 0.1% SDS, and 10% dextran sulfate at 42°C, with washes at 42°C in 0.2X SSC (sodium chloride/sodium citrate) and 50% formamide at 55°C, followed by a high-stringency wash consisting of 0.1X SSC containing EDTA at 55°C. The skilled artisan will recognize how to adjust the temperature, ionic strength, etc. as necessary to accommodate factors such as probe length and the like. 3.2 Vectors [301] The nucleic acids of the present disclosure in some aspects are incorporated into a vector. In this regard, the present disclosure provides vectors comprising any of the presently disclosed nucleic acids. In exemplary aspects, the vector is a recombinant expression vector. For purposes herein, the term "recombinant expression vector" means a genetically-modified oligonucleotide or polynucleotide construct that permits the expression of an mRNA, protein, polypeptide, or peptide by a host cell, when the construct comprises a nucleotide sequence encoding the mRNA, protein, polypeptide, or peptide, and the vector is contacted with the cell under conditions sufficient to have the mRNA, protein, polypeptide, or peptide expressed within the cell. The vectors of the present disclosure are not naturally-occurring as a whole. However, parts of the vectors can be naturally-occurring. The presently disclosed vectors can comprise any type of nucleotides, including, but not limited to DNA and RNA, which can be single- stranded or double-stranded, synthesized or obtained in part from natural sources, and which can contain natural, non-natural or altered nucleotides. The vectors can comprise naturally-occurring or non-naturally- occurring internucleotide linkages, or both types of linkages. In some aspects, the altered nucleotides or non-naturally occurring internucleotide linkages do not hinder the transcription or replication of the vector. [302] The vector of the present disclosure can be any suitable vector, and can be used to transform or transfect any suitable host. Suitable vectors include those designed for propagation and expansion or for expression or both, such as plasmids and viruses. The vector can be selected from the group consisting of the pUC series (Fermentas Life Sciences), the pBluescript series (Stratagene, LaJoIIa, CA), the pET series (Novagen, Madison, WI), the pGEX series (Pharmacia Biotech, Uppsala, Sweden), and the pEX series (Clontech, Palo Alto, CA). Bacteriophage vectors, such as λGTIO, λGTl 1, λZapII (Stratagene), λEMBL4, and λNMl 149, also can be used. Examples of plant expression vectors include pBI101, pBI101.2, pBI101.3, pBI121 and pBIN19 (Clontech). Examples of animal expression vectors include pEUK-Cl, pMAM and pMAMneo (Clontech). In some aspects, the vector is a viral vector, e.g., a retroviral vector.
[303] The vectors of the present disclosure can be prepared using standard recombinant DNA techniques described in, for example, Sambrook et al., infra, and Ausubel et al., infra. Constructs of expression vectors, which are circular or linear, can be prepared to contain a replication system functional in a prokaryotic or eukaryotic host cell. Replication systems can be derived, e.g., from CoIEl, 2 μ plasmid, λ, SV40, bovine papilloma virus, and the like. [304] In some aspects, the vector comprises regulatory sequences, such as transcription and translation initiation and termination codons, which are specific to the type of host (e.g., bacterium, fungus, plant, or animal) into which the vector is to be introduced, as appropriate and taking into consideration whether the vector is DNA- or RNA- based. [305] The vector can include one or more marker genes, which allow for selection of transformed or transfected hosts. Marker genes include biocide resistance, e.g., resistance to antibiotics, heavy metals, etc., complementation in an auxotrophic host to provide prototrophy, and the like. Suitable marker genes for the presently disclosed expression vectors include, for instance, neomycin/G418 resistance genes, hygromycin resistance genes, histidinol resistance genes, tetracycline resistance genes, and ampicillin resistance genes. [306] The vector can comprise a native or normative promoter operably linked to the nucleotide sequence encoding the polypeptide (including functional portions and functional variants thereof), or to the nucleotide sequence which is complementary to or which hybridizes to the nucleotide sequence encoding the antigen-binding protein. The selection of promoters, e.g., strong, weak, inducible, tissue- specific and developmental- specific, is within the ordinary skill of the artisan. Similarly, the combining of a nucleotide sequence with a promoter is also within the skill of the artisan. The promoter can be a non-viral promoter or a viral promoter, e.g., a cytomegalovirus (CMV) promoter, an SV40 promoter, an RSV promoter, and a promoter found in the long-terminal repeat of the murine stem cell virus. [307] Certain exemplary antigen-binding proteins disclosed herein comprises two different polypeptide chains. To further improve the expression and pairing of the two chains, different promoters of different strengths may be operably linked to coding sequences under different MXS selection conditions, such that the expression levels of the two chains may be adjusted to ensure optimal expression and chain pairing. See, e.g., Example 8 disclosed below. 3.3 Host cells [308] Provided herein are host cells comprising a nucleic acid or vector of the present disclosure. As used herein, the term "host cell" refers to any type of cell that can contain the presently disclosed vector and is capable of producing an expression product encoded by the nucleic acid (e.g., mRNA, protein). The host cell in some aspects is an adherent cell or a suspended cell, i.e., a cell that grows in
suspension. The host cell in exemplary aspects is a cultured cell or a primary cell, i.e., isolated directly from an organism, e.g., a human. The host cell can be of any cell type, can originate from any type of tissue, and can be of any developmental stage. [309] In exemplary aspects, the cell is a eukaryotic cell, including, but not limited to, a yeast cell, filamentous fungi cell, protozoa cell, algae cell, insect cell, or mammalian cell. Such host cells are described in the art. See, e.g., Kunert et al., Appl. Microbiol Biotechnol.100: 3451-61 (2016). In exemplary aspects, the eukaryotic cells are mammalian cells. In exemplary aspects, the mammalian cells are non-human mammalian cells. In some aspects, the cells are Chinese Hamster Ovary (CHO) cells and derivatives thereof (e.g., CHO-K1, CHO pro-3, CS9), mouse myeloma cells (e.g., NS0, GS-NS0, Sp2/0), cells engineered to be deficient in dihydrofolatereductase (DHFR) activity (e.g., DUKX-X11, DG44), human embryonic kidney 293 (HEK293) cells or derivatives thereof (e.g., HEK293T, HEK293-EBNA), green African monkey kidney cells (e.g., COS cells, VERO cells), human cervical cancer cells (e.g., HeLa), human bone osteosarcoma epithelial cells U2-OS, adenocarcinomic human alveolar basal epithelial cells A549, human fibrosarcoma cells HT1080, mouse brain tumor cells CAD, embryonic carcinoma cells P19, mouse embryo fibroblast cells NIH 3T3, mouse fibroblast cells L929, mouse neuroblastoma cells N2a, human breast cancer cells MCF-7, retinoblastoma cells Y79, human retinoblastoma cells SO-Rb50, human liver cancer cells Hep G2, mouse B myeloma cells J558L, or baby hamster kidney (BHK) cells (Gaillet et al.2007; Khan, Adv Pharm Bull 3(2): 257-263 (2013)). In a particular embodiment, the host cell is CS9 (a CHO cell line). [310] For purposes of amplifying or replicating the vector, the host cell is in some aspects is a prokaryotic cell, e.g., a bacterial cell. [311] Also provided by the present disclosure is a population of cells comprising at least one host cell described herein. The population of cells in some aspects is a heterogeneous population comprising the host cell comprising vectors described, in addition to at least one other cell, which does not comprise any of the vectors. Alternatively, in some aspects, the population of cells is a substantially homogeneous population, in which the population comprises mainly host cells (e.g., consisting essentially of) comprising the vector. The population in some aspects is a clonal population of cells, in which all cells of the population are clones of a single host cell comprising a vector, such that all cells of the population comprise the vector. In exemplary embodiments of the present disclosure, the population of cells is a clonal population comprising host cells comprising a vector as described herein. 3.4. Methods of Manufacture [312] The antigen-binding proteins disclosed herein may be obtained by methods known in the art. Suitable methods of de novo synthesizing polypeptides are described in, for example, Chan et al., Fmoc Solid Phase Peptide Synthesis, Oxford University Press, Oxford, United Kingdom, 2005; Peptide and
Protein Drug Analysis, ed. Reid, R., Marcel Dekker, Inc., 2000; Epitope Mapping, ed. Westwood et al., Oxford University Press, Oxford, United Kingdom, 2000; and U.S. Patent No.5,449,752. Additional exemplary methods of making the peptides of the invention are set forth herein. [313] Also, in some aspects, the antigen-binding proteins disclosed herein are recombinantly produced using a nucleic acid encoding the amino acid sequence of the molecule using standard recombinant methods. See, for instance, Sambrook et al., Molecular Cloning: A Laboratory Manual.3rd ed., Cold Spring Harbor Press, Cold Spring Harbor, NY 2001; and Ausubel et al., Current Protocols in Molecular Biology, Greene Publishing Associates and John Wiley & Sons, NY, 1994. [314] Methods of making antigen-binding proteins disclosed herein are further provided herein. In exemplary embodiments, the method comprises culturing a presently disclosed host cell so as to express the antigen-binding protein and harvesting the expressed antigen-binding protein. The host cell can be any of the host cells described herein. In exemplary aspects, the host cell is selected from the group consisting of: CHO cells, NS0 cells, COS cells, VERO cells, and BHK cells. In exemplary aspects, the step of culturing a host cell comprises culturing the host cell in a growth medium to support the growth and expansion of the host cell. In exemplary aspects, the growth medium increases cell density, culture viability and productivity in a timely manner. In exemplary aspects, the growth medium comprises amino acids, vitamins, inorganic salts, glucose, and serum as a source of growth factors, hormones, and attachment factors. In exemplary aspects, the growth medium is a fully chemically defined media consisting of amino acids, vitamins, trace elements, inorganic salts, lipids and insulin or insulin-like growth factors. In addition to nutrients, the growth medium also helps maintain pH and osmolality. Several growth media are commercially available and are described in the art. See, e.g., Arora, “Cell Culture Media: A Review” MATER METHODS 3:175 (2013). [315] In exemplary aspects, the method of making antigen-binding proteins disclosed herein comprises culturing the host cell in a feed medium. In exemplary aspects, the method comprises culturing in a feed medium in a fed-batch mode. Methods of recombinant protein production are known in the art. See, e.g., Li et al., “Cell culture processes for monoclonal antibody production” MAbs 2(5): 466–477 (2010). [316] Typically, to establish a cell line producing recombinant antigen-binding protein (e.g., a protein derived from an antibody and comprises two chains, one based on antibody heavy chain and one based on antibody light chain), it is often necessary to integrate the heavy and light chain coding sequences from a single or separate vector into the genome followed by a stringency and efficiency metabolic selection to find high producing cell lines. In glutamine synthetase (GS)-CHO expression system, selection of high-producing cell lines is based on controlling the balance between the expression level of GS and the concentration of its specific inhibitor, l-methionine sulfoximine (MSX). As disclosed and exemplified in Example 8, one method of expressing an antigen-binding protein disclosed herein is to optimize the expression of the heavy chain and/or light chain using promoters with different strengths
under different MSX concentrations. As GS acts both as a selectable marker and a means to achieve gene amplification, the optimal selection stringency for each promoter was determined using various MSX concentrations. Results from these experiments established the preferred vector design plus selection stringency for approximately seven different biologic modalities (including multispecific proteins). As such, provided herein are vector designs strategies for efficient expression of antigen-binding proteins. [317] The method of making antigen-binding proteins disclosed herein can comprise one or more steps for purifying the molecule from a cell culture or the supernatant thereof and preferably recovering the purified protein. In exemplary aspects, the method comprises one or more chromatography steps, e.g., affinity chromatography (e.g., protein A affinity chromatography), ion exchange chromatography, hydrophobic interaction chromatography. In exemplary aspects, the method comprises purifying the protein using a Protein A affinity chromatography resin. [318] In exemplary embodiments, the method further comprises steps for formulating the purified protein, etc., thereby obtaining a formulation comprising the purified protein. Such steps are described, for example, in Formulation and Process Development Strategies for Manufacturing, eds. Jameel and Hershenson, John Wiley & Sons, Inc. (Hoboken, NJ), 2010. 4. Pharmaceutical Compositions and Method of Treatment 4.1 Pharmaceutical Compositions [319] Compositions comprising an antigen-binding protein, a nucleic acid, a vector, a host cell, or a combination thereof, are provided herein. The compositions may comprise the antigen-binding protein, nucleic acid, vector, or host cell, or a combination thereof, in isolated and/or purified form. [320] In exemplary aspects, the composition comprises agents which enhance the chemico-physico features of the antigen-binding molecule, nucleic acid, vector, or host cell, or a combination thereof, e.g., via stabilizing, for example, at certain temperatures (e.g., room temperature), increasing shelf life, reducing degradation, e.g., oxidation protease mediated degradation, increasing half-life of the antigen- binding protein, etc. [321] In exemplary aspects of the present disclosure, the composition additionally comprises a pharmaceutically acceptable carrier, diluents, or excipient. The pharmaceutical composition can comprise any pharmaceutically acceptable ingredients, including, for example, acidifying agents, additives, adsorbents, aerosol propellants, air displacement agents, alkalizing agents, anticaking agents, anticoagulants, antimicrobial preservatives, antioxidants, antiseptics, bases, binders, buffering agents, chelating agents, coating agents, coloring agents, desiccants, detergents, diluents, disinfectants, disintegrants, dispersing agents, dissolution enhancing agents, dyes, emollients, emulsifying agents, emulsion stabilizers, fillers, film forming agents, flavor enhancers, flavoring agents, flow enhancers,
gelling agents, granulating agents, humectants, lubricants, mucoadhesives, ointment bases, ointments, oleaginous vehicles, organic bases, pastille bases, pigments, plasticizers, polishing agents, preservatives, sequestering agents, skin penetrants, solubilizing agents, solvents, stabilizing agents, suppository bases, surface active agents, surfactants, suspending agents, sweetening agents, therapeutic agents, thickening agents, tonicity agents, toxicity agents, viscosity-increasing agents, water- absorbing agents, water-miscible cosolvents, water softeners, or wetting agents. See, e.g., the Handbook of Pharmaceutical Excipients, Third Edition, A. H. Kibbe (Pharmaceutical Press, London, UK, 2000), which is incorporated by reference in its entirety. Remington’s Pharmaceutical Sciences, Sixteenth Edition, E. W. Martin (Mack Publishing Co., Easton, Pa., 1980), which is incorporated by reference in its entirety. [322] In exemplary aspects, the pharmaceutical composition comprises formulation materials that are nontoxic to recipients at the dosages and concentrations employed. In specific embodiments, pharmaceutical compositions comprising an active agent and one or more pharmaceutically acceptable salts; polyols; surfactants; osmotic balancing agents; tonicity agents; anti-oxidants; antibiotics; antimycotics; bulking agents; lyoprotectants; anti-foaming agents; chelating agents; preservatives; colorants; analgesics; or additional pharmaceutical agents. In exemplary aspects, the pharmaceutical composition comprises one or more polyols and/or one or more surfactants, optionally, in addition to one or more excipients, including but not limited to, pharmaceutically acceptable salts; osmotic balancing agents (tonicity agents); anti-oxidants; antibiotics; antimycotics; bulking agents; lyoprotectants; anti- foaming agents; chelating agents; preservatives; colorants; and analgesics. [323] In certain embodiments, the pharmaceutical composition can contain formulation materials for modifying, maintaining or preserving, for example, the pH, osmolarity, viscosity, clarity, color, isotonicity, odor, sterility, stability, rate of dissolution or release, adsorption or penetration of the composition. In such embodiments, suitable formulation materials include, but are not limited to, amino acids (such as glycine, glutamine, asparagine, arginine or lysine); antimicrobials; antioxidants (such as ascorbic acid, sodium sulfite or sodium hydrogen-sulfite); buffers (such as borate, bicarbonate, Tris-HCl, citrates, phosphates or other organic acids); bulking agents (such as mannitol or glycine); chelating agents (such as ethylenediamine tetraacetic acid (EDTA)); complexing agents (such as caffeine, polyvinylpyrrolidone, beta-cyclodextrin or hydroxypropyl-beta-cyclodextrin); fillers; monosaccharides; disaccharides; and other carbohydrates (such as glucose, mannose or dextrins); proteins (such as serum albumin, gelatin or immunoglobulins); coloring, flavoring and diluting agents; emulsifying agents; hydrophilic polymers (such as polyvinylpyrrolidone); low molecular weight polypeptides; salt-forming counterions (such as sodium); preservatives (such as benzalkonium chloride, benzoic acid, salicylic acid, thimerosal, phenethyl alcohol, methylparaben, propylparaben, chlorhexidine, sorbic acid or hydrogen peroxide); solvents (such as glycerin, propylene glycol or polyethylene glycol); sugar alcohols (such as mannitol or sorbitol); suspending agents; surfactants or wetting agents (such as pluronics, PEG, sorbitan esters, polysorbates
such as polysorbate 20, polysorbate, triton, tromethamine, lecithin, cholesterol, tyloxapal); stability enhancing agents (such as sucrose or sorbitol); tonicity enhancing agents (such as alkali metal halides, preferably sodium or potassium chloride, mannitol sorbitol); delivery vehicles; diluents; excipients and/or pharmaceutical adjuvants. See, REMINGTON'S PHARMACEUTICAL SCIENCES, 18″ Edition, (A. R. Genrmo, ed.), 1990, Mack Publishing Company. [324] The pharmaceutical compositions can be formulated to achieve a physiologically compatible pH. In some embodiments, the pH of the pharmaceutical composition can be for example between about 4 or about 5 and about 8.0 or about 4.5 and about 7.5 or about 5.0 to about 7.5. In exemplary embodiments, the pH of the pharmaceutical composition is between 5.5 and 7.5. 4.2 Methods of Treatment [325] Methods of treatment are additionally provided by the present disclosure. The method, in exemplary embodiments, is a method of treating a subject in need thereof, comprising administering to the subject in need thereof a pharmaceutical composition of the present disclosure in an amount effective to treat the subject. As such, an antigen binding protein or pharmaceutical compositions of the present disclosure may be for medical use in a human subject. By way of example, the antigen binding protein or pharmaceutical compositions of the present disclosure may for medical use that does not comprise systemic immune activation or liver toxicity in the human subject. For example, as discussed in detail above, the bispecific molecules disclosed herein is a cross-linking dependent agonist for 4-1BB. Therefore, the bispecific molecules induce 4-1BB activation upon PD-L1 binding and subsequent clustering (crosslinking) of 4-1BB. [326] The pharmaceutical compositions of the present disclosure are useful for activating 4-1BB signaling. Without being bound to a particular theory, the 4-1BB agonist activity of the compositions provided herein allow such entities to be useful in methods of enhancing T cell activity and enhancing an immune response, and, in particular, an immune response against a tumor or cancer. [327] Accordingly, provided herein are methods of enhancing T cell activity in a subject, enhancing T cell survival and effector function, restricting terminal differentiation and loss of replicative potential, promoting T cell longevity, and enhancing cytotoxicity against target (e.g., cancer) cells. In exemplary embodiments, the methods comprise administering to the subject the pharmaceutical composition of the present disclosure in an effective amount. In exemplary aspects, the T cell activity or immune response is directed against a cancer cell or cancer tissue or a tumor cell or tumor. In exemplary aspects, the immune response is a humoral immune response. In exemplary aspects, the immune response is an innate immune response. In exemplary aspects, the immune response which is enhanced is a T-cell mediated immune response.
[328] As used herein, the term “enhance” and words stemming therefrom may not be a 100% or complete enhancement or increase. Rather, there are varying degrees of enhancement of which one of ordinary skill in the art recognizes as having a potential benefit or therapeutic effect. In this respect, the pharmaceutical compositions of the present disclosure may enhance, e.g., T cell activity or enhance an immune response, to any amount or level. In exemplary embodiments, the enhancement provided by the methods of the present disclosure is at least or about a 10% enhancement (e.g., at least or about a 20% enhancement, at least or about a 30% enhancement, at least or about a 40% enhancement, at least or about a 50% enhancement, at least or about a 60% enhancement, at least or about a 70% enhancement, at least or about a 80% enhancement, at least or about a 90% enhancement, at least or about a 95% enhancement, at least or about a 98% enhancement). [329] Methods of measuring T cell activity and immune responses are known in the art. T cell activity can be measured by, for example, a cytotoxicity assay, such as those described in Fu et al., PLoS ONE 5(7): e11867 (2010). Other T cell activity assays are described in Bercovici et al., Clin Diagn Lab Immunol. 7(6): 859–864 (2000). Methods of measuring immune responses are described in e.g., Macatangay et al., Clin Vaccine Immunol 17(9): 1452-1459 (2010), and Clay et al., Clin Cancer Res.7(5):1127-35 (2001). [330] Additionally provided herein are methods of treating a subject with cancer and methods of treating a subject with a solid tumor. In exemplary embodiments, the method comprises administering to the subject the pharmaceutical composition of the present disclosure in an amount effective for treating the cancer or the solid tumor in the subject. [331] The cancer treatable by the methods disclosed herein can be any cancer, e.g., any malignant growth or tumor caused by abnormal and uncontrolled cell division that may spread to other parts of the body through the lymphatic system or the blood stream. The cancer in some aspects is one selected from the group consisting of acute lymphocytic cancer, acute myeloid leukemia, alveolar rhabdomyosarcoma, bone cancer, brain cancer, breast cancer, cancer of the anus, anal canal, or anorectum, cancer of the eye, cancer of the intrahepatic bile duct, cancer of the joints, cancer of the neck, gallbladder, or pleura, cancer of the nose, nasal cavity, or middle ear, cancer of the oral cavity, cancer of the vulva, chronic lymphocytic leukemia, chronic myeloid cancer, colon cancer, esophageal cancer, cervical cancer, gastrointestinal carcinoid tumor, Hodgkin lymphoma, hypopharynx cancer, kidney cancer, larynx cancer, liver cancer, lung cancer, malignant mesothelioma, melanoma, multiple myeloma, nasopharynx cancer, non-Hodgkin lymphoma, ovarian cancer, pancreatic cancer, peritoneum, omentum, and mesentery cancer, pharynx cancer, prostate cancer, rectal cancer, renal cancer (e.g., renal cell carcinoma (RCC)), small intestine cancer, soft tissue cancer, stomach cancer, testicular cancer, thyroid cancer, ureter cancer, and urinary bladder cancer. In particular aspects, the cancer is selected from the group consisting of: head and neck, ovarian, cervical, bladder and oesophageal cancers, pancreatic, gastrointestinal cancer, gastric, breast, endometrial and colorectal cancers, hepatocellular carcinoma, glioblastoma,
bladder, lung cancer, e.g., non-small cell lung cancer (NSCLC), bronchioloalveolar carcinoma. In particular embodiments, the tumor is non-small cell lung cancer (NSCLC), head and neck cancer, renal cancer, triple negative breast cancer, and gastric cancer. In exemplary aspects, the subject has a tumor (e.g., a solid tumor, a hematological malignancy, or a lymphoid malignancy) and the pharmaceutical composition is administered to the subject in an amount effective to treat the tumor in the subject. In other exemplary aspects, the tumor is non-small cell lung cancer (NSCLC), small cell lung cancer (SCLC), head and neck cancer, renal cancer, breast cancer, melanoma, ovarian cancer, liver cancer, pancreatic cancer, colon cancer, prostate cancer, gastric cancer, lymphoma or leukemia, and the pharmaceutical composition is administered to the subject in an amount effective to treat the tumor in the subject. In some exemplary aspects, the cancer comprises a solid tumor. [332] As used herein, the term “treat,” as well as words related thereto, do not necessarily imply 100% or complete treatment. Rather, there are varying degrees of treatment of which one of ordinary skill in the art recognizes as having a potential benefit or therapeutic effect. In this respect, the methods of treating cancer of the present disclosure can provide any amount or any level of treatment. Furthermore, the treatment provided by the method of the present disclosure can include treatment of one or more conditions or symptoms or signs of the cancer being treated. Also, the treatment provided by the methods of the present disclosure can encompass slowing the progression of the cancer. For example, the methods can treat cancer by virtue of enhancing the T cell activity or an immune response against the cancer, reducing tumor or cancer growth, reducing metastasis of tumor cells, increasing cell death of tumor or cancer cells, and the like. In exemplary aspects, the methods treat by way of delaying the onset or recurrence of the cancer by 1 day, 2 days, 4 days, 6 days, 8 days, 10 days, 15 days, 30 days, two months, 4 months, 6 months, 1 year, 2 years, 4 years, or more. In exemplary aspects, the methods treat by way increasing the survival of the subject. [333] In particular, the antigen-binding proteins disclosed herein target cancer-associated fibroblasts that is present in tumor stroma. Tumor stroma, broadly defined as the non-cancer cell and non-immune cell components of tumors, is viewed traditionally as the structural components holding tumor tissues together. Tumor stroma is composed of extracellular matrix and specialized connective tissue cells, including fibroblasts and mesenchymal stromal cells. Tumors generally need stroma for nutritional support and the removal of waste products, but stromal content can vary markedly in different types of cancers. For example, many lymphomas have minimal stroma whereas the stroma may make up 90% of other solid tumors. The antigen-binding proteins disclosed herein in particular target PD-L1-expressing tumors. Fibroblasts are capable of infiltrating tumors and PD-L1-expressing cells can be easily identified by methods well known in the art, such as immunostaining. 4.3 Combination Therapy
[334] In some embodiments, the active agents described herein are administered alone, and in alternative embodiments, are administered in combination with another therapeutic agent, e.g., another active agent of the present disclosure of a different type (e.g., structure). Accordingly, the present disclosure provides a combination comprising a first antigen binding protein which targets PD-L1 and a second antigen binding protein which targets 4-1BB, each of which is an antigen binding protein according to the present disclosures. In various instances, the first antigen binding protein is any one of the PD-L1 binding proteins disclosed herein (such as those in Tables A and K2), and optionally, the second antigen binding protein is any one of the 4-1BB binding proteins disclosed herein (such as those disclosed in Tables B and K1). [335] The present disclosure provides the combination as a composition, e.g., pharmaceutical composition, in various instances. Accordingly, the present disclosure provides a composition, e.g., pharmaceutical composition, comprising the first antigen binding protein and the second antigen binding protein. In exemplary instances, the first antigen binding protein and the second antigen binding protein are present in the composition at a ratio of about 1:1. In some aspects, the combination or composition, further comprises a third antigen binding protein. [336] In some embodiments, the disclosure provides a method of treating cancer, comprising administering to a subject in need thereof a therapeutically effective amount of an antigen-binding molecule disclosed herein, and a therapeutically effective amount of PD-1 inhibitor. In certain embodiments, the bispecific molecule disclosed herein is administered in combination with a PD-1 inhibitor. The PD-1 inhibitor and the antigen-binding molecule disclosed herein may be administered concurrently or sequentially. [337] In certain embodiments, the PD-1 inhibitor is an antibody, or antigen-binding fragment thereof. In certain embodiments, the antibody, or antigen-binding portion thereof, binds to human PD-1. Examples of antibodies that bind to human PD-1, are described, e.g, in US7488802, US7521051, US8008449, US8354509, US8168757, WO2004/004771, WO2004/072286, WO2004/056875, and US2011/0271358. Specific anti-human PD-1 antibodies useful for the invention described herein include, for example: K- 3945 (Pembrolizumab, Keytruda®; U.S. Patent No.8,952,136); M-3475, a humanized IgG4 mAb with the structure described in WHO Drug Information, Vol.27, No.2, pages 161-162 (2013); nivoiumab (BMS- 936558), a human IgG4 mAb with the structure described in WHO Drug Information, Vol.27, No.1, pages 68-69 (2013); the humanized antibodies h409A11, h409A16 and h409A17, which are described in WO2008/156712; AMP-514, which is being developed by Medlmmune; humanized antibody CT-011 (Pidilizumab) a monoclonal antibody being developed by Medivation, and anti-PD-1 antibodies disclosed in WO2015/119923 (the heavy and light chains comprise SEQ ID NO: 21 and SEQ ID NO: 22, respectively). Additional PD-1 inhibitors include Cemiplimab (Libtayo, approved for the treatment of cutaneous squamous cell carcinoma (CSCC) or locally advanced CSCC who are not candidates for
curative surgery or curative radiation); and Dostarlimab (Jemperli, approved for the treatment of mismatch repair deficient (dMMR) recurrent or advanced endometrial cancer and mismatch repair deficient (dMMR) recurrent or advanced solid tumors). [338] In some aspects, the other therapeutic aims to treat or prevent cancer. In some embodiments, the other therapeutic is a chemotherapeutic agent. In some embodiments, the other therapeutic is an agent used in radiation therapy for the treatment of cancer. Accordingly, in some aspects, the active agents described herein are administered in combination with one or more of platinum coordination compounds, topoisomerase inhibitors, antibiotics, antimitotic alkaloids and difluoronucleosides. 4.4 Subjects [339] In some embodiments of the present disclosure, the subject is a mammal, including, but not limited to, mammals of the order Rodentia, such as mice and hamsters, and mammals of the order Logomorpha, such as rabbits, mammals from the order Carnivora, including Felines (cats) and Canines (dogs), mammals from the order Artiodactyla, including Bovines (cows) and Swines (pigs) or of the order Perssodactyla, including Equines (horses). In some aspects, the mammals are of the order Primates, Ceboids, or Simoids (monkeys) or of the order Anthropoids (humans and apes). In some aspects, the mammal is a human. 5. Kits [340] The present disclosure additionally provides kits comprising an antigen binding protein (including a bispecific molecule), nucleic acid, vector, or host cell of the present disclosure, or a combination thereof. In exemplary aspects, the antigen binding protein, nucleic acid, vector, or host cell is provided in the kit as a unit dose. For purposes herein “unit dose" refers to a discrete amount dispersed in a suitable carrier. In exemplary aspects, the unit dose is the amount sufficient to provide a subject with a desired effect, e.g., treatment of cancer. In exemplary aspects, the kit comprises several unit doses, e.g., a week or month supply of unit doses, optionally, each of which is individually packaged or otherwise separated from other unit doses. In some embodiments, the components of the kit/unit dose are packaged with instructions for administration to a patient. In some embodiments, the kit comprises one or more devices for administration to a patient, e.g., a needle and syringe, and the like. In some aspects, the antigen binding protein, nucleic acid, vector, host cell, or a combination thereof, is/are pre-packaged in a ready to use form, e.g., a syringe, an intravenous bag, etc. In exemplary aspects, the ready to use form is for a single use. In exemplary aspects, the kit comprises multiple single use, ready to use forms of the antigen binding protein, nucleic acid, vector, or host cell of the present disclosure. In some aspects, the kit further comprises other therapeutic or diagnostic agents or pharmaceutically acceptable carriers (e.g., solvents, buffers, diluents, etc.), including any of those described herein.
6. Immunogenicity Assays [341] Protein-based drug candidates can be immunogenic for reasons including route of administration, dose frequency and the underlying antigenicity of the therapeutic protein. T cell epitope content (which is typically a 9-mer peptide) is one of the factors that contributes to antigenicity. The binding strength of T cell epitopes to major histocompatibility complex (MHC or HLA) molecules is a key determinant in T cell epitope immunogenicity. This allows the epitopes with higher binding affinities to be more likely to be displayed on the surface of the cell (in the context of MHC molecules) where they are recognized by their corresponding T cell receptor (TCR). [342] Antigen-presenting cells (APCs), typically dendritic cells (DCs), can process antigens and present peptide epitopes in conjugation with human leukocyte antigen (HLA) class II molecules to specific naïve helper-T (Th) cells, which results in the activation of Th cells. The activated Th cells can stimulate B cells to produce antibodies against antigens. For instance, anti-drug antibodies (ADAs) are generated in this way. Therefore, detection of T cell responses is frequently employed to monitor the ability of biotherapeutic candidates to elicit immune system activation, i.e., the immunogenicity. The most widely used method is DC:T cell proliferation assay. [343] The inventors discovered that the 41BB x PD-L1 bispecific molecules disclosed herein posed a unique problem for DC:T assays. The PD-L1-targeting arm of the molecule can bind to the PD-L1 expressed on the surface of DC, and the 4-1BB targeting arm of the molecule can bind to the 4-1BB on the surface of T cells. The molecule is designed to activates 4-1BB signal pathway through PD-L1 binding and subsequent clustering of 4-1BB. The activation of the 4-1BB pathway in turn causes T cell activation and proliferation. Therefore, in a typical DC:T assay, such type of molecules will always produce a false positive result, due to the underlying mechanism of action. Indeed experimental results confirmed that typical DC:T cell proliferation assay had produced flawed results. The assay could not be used to assess T cell epitopes that are driven by certain unique amino acid sequence (sequence-based immunogenicity, typically attributed to 9-mer peptides). [344] The inventors solved the problem by dividing the bispecific molecules into different domains, for example, by converting the bispecific molecule into a PD-L1 binding molecule (fused with Fc), a 4-1BB binding molecule (fused with Fc), and subjecting these individual domains into a DC:T cell proliferation assay, and comparing the results with an Fc domain alone (control). This way, the contribution of the anti- PD-L1 portion and the anti-41BB portion to sequence-based immunogenic risk could be assessed. [345] In addition to 41BB x PD-L1 bispecific molecules disclosed herein, one can envision that other bispecific molecules could also receive false positives produced by a conventional DC:T cell assay, if such bispecific molecule comprises two domains: a first domain that binds to a DC surface antigen, and a second domain that binds to a T cell co-stimulatory molecule. The molecule itself can bridge the DC and
T cells, causing the activation and proliferation of T cells. Therefore, principles described here are applicable to these molecules, and the two domains should be separately expressed as two different proteins to detect true T-cell epitopes. [346] Many DC surface antigens (or surface markers) are known. For example, prominent conventional DC (cDC, sometimes also referred to as classical DC or myeloid DC) surface antigens include CD8A, CLEC9A, ITGAE, ITGAX, THBD (CD141), XCR1, CD1C, CD207, ITGAM, NOTCH2, and SIRPA. Common surface antigens for plasmacytoid DCs (pDCs) include CLEC4C, LILRB4, NRP1, CCR7, and B220. Surface markers reported for monocyte-derived DCs (mo-DCs) include CD14, MRC1 (CD206), CD209, SIRPA, ITGAM (CD11b), and CD1A. Langherans surface markers include CD1A, CD207 (Langerin), and ID2. AXL+Siglec6+ dendritic cells (AS DC) surface markers include AXL and Siglec6 (CD327). [347] In addition to 4-1BB, common T-cell co-stimulatory molecules include CD28, Inducible Co- Stimulator (ICOS), CTLA4 (Cytotoxic T-Lymphocyte-Associated protein 4), 4-1BB (also known as CD137), OX40 (also known as CD134), CD27, CD30, DR3, Glucocorticoid-Induced TNFR family Related (GITR), or Herpes Virus Entry Mediator (HVEM). [348] Accordingly, the disclosure provides a method of assessing immunogenicity of a bispecific molecule, wherein said immunogenicity is attributed to a T-cell epitope, and wherein said bispecific molecule comprises two domains: (1) a first domain that binds to a Dendritic Cell (DC) surface antigen; and (2) a second domain that binds to a T cell co-stimulatory molecule, the method comprises: (a) obtaining a first protein that comprises said first domain but does not comprise said second domain; (b) obtaining a second protein that comprises said second domain but does not comprise said first domain; (c) incubating said first protein and second protein with a cell culture that comprises DCs and T cells; (d) assessing the activation or proliferation of T cells. The activation or proliferation of T cells is indicative that said bispecific molecule comprises an immunogenic T cell epitope. [349] If desired, the first and second proteins can be fused to an Fc domain, as exemplified in the Examples, and the results can be compared with a parallel assay using Fc domain alone as a negative control. [350] The following examples are given merely to illustrate the present invention and not in any way to limit its scope. EXAMPLES EXAMPLE 1 [351] This example describes the generation and characterization of antibodies to human PD-L1 and antibodies to 4-1BB.
[352] A panel of lead 4-1BB monoclonal antibodies and PD-L1 monoclonal antibodies was identified by carrying out the steps schematically described in Figure 1A. More details of each step are provided below. Immunization Campaign 1. Mouse Strains [353] Fully human antibodies to human PD-L1 and 4-1BB were generated by immunizing XENOMOUSE® transgenic mice (U.S. Pat. NOs.6,114,598; 6,162,963;6,833,268; 7,049,426; 7,064,244, which are incorporated herein by reference in their entirety; Green et al., 1994, Nature Genetics 7:13-21; Mendez et al., 1997, Nature Genetics 15:146-156; Green and Jakobovits, 1998, J. Ex. Med, 188:483-495; Kellerman and Green, Current Opinion in Biotechnology 13, 593-597, 2002). Animals from the XMG4 and XMG2 XENOMOUSE® strains were used for these immunizations. 2. Immunizations [354] Multiple immunogens and routes of immunization were used to generate anti-human PD-L1 and 4- 1BB immune responses. For genetic immunizations, mice were immunized 14-18 times over 6-10 weeks using the Helios Gene Gun system according to the manufacturer’s instructions (BioRad, Hercules, California). Briefly, expression vectors encoding wild type human or rhesus GIPR were coated onto gold beads (BioRad, Hercules, California) and delivered to the epidermis of a shaved mouse or rat abdomen. For soluble protein immunizations, mice were immunized with a human PD-L1 or 4-1BB recombinant protein representing the N-terminal extracellular domain. Animals were immunized with recombinant protein mixed with Alum and CpG-ODN 12 times over 4-6 weeks using sub-cutaneous injections. The initial boost was 5 µg (4-1BB) or 10 µg (PD-L1) and subsequent boosts were 5 µg for both programs. PD- L1 or 4-1BB-specific serum titers were monitored by live-cell FACS analysis on an Accuri flow cytometer (BD Biosciences) using transiently transfected 293T cells. Animals with the highest antigen-specific serum titers against human PD-L1 or 4-1BB were sacrificed and used for hybridoma generation (Kohler and Milstein, 1975). 3. Hybridoma Generation [355] Animals exhibiting suitable serum titers were identified and lymphocytes were obtained from spleen and/or draining lymph nodes. Pooled lymphocytes (from each harvest) were dissociated from lymphoid tissue by grinding in a suitable medium. B cells were selected and/or expanded using standard methods, and fused with a suitable fusion partner using techniques that were known in the art. Selection and Characterization of Monoclonal Antibodies
1. Initial Selection of PD-L1 and 4-1BB Specific Binding Antibodies [356] Exhausted hybridoma supernatants were tested for binding to human PD-L1 or human or cyno 4- 1BB transiently expressed on HEK293 cells by Cell Insight. Briefly, HEK293 cells were transiently transfected with a mammalian expression construct encoding either human PD-L1 or human/cynomolgus 4-1BB using 293Fectin. The following day, 15 µL of exhausted hybridoma media was added to each well of a 384 well FMAT plate. Then, the transfected HEK293 cells, the nuclear stain Hoechst 33342 and a secondary detection antibody (Goat anti Human IgG Fc Alexa 488 (Jackson ImmunoResearch) were mixed and 30 µL of this mixture was added to each well of a 384 well FMAT plate. The supernatant was then aspirated using an AquaMax plate reader and 30 µL of FACS buffer was added to each well using a multidrop instrument. The plates were then read on the Cell Insight platform using the Cell Health Profiling Bio-App. Tables 1A and 1B summarize the antigen-specific hits identified in the PD-L1 harvests and the 4-1BB harvests, respectively. TABLE 1A (PD-1) Harvest Antigen-specific binders 1 1032
TABLE 1B (4-1BB) Harvest Antigen-specific binders
2. Jurkat human PD-1/NFAT-luciferase reporter assay [357] Jurkat cells stably expressing human PD-1 and NFAT-luciferase reporter (Promega) and Chinese Hamster Ovary (CHO) stably expressing human PD-L1 (Promega) were used. Test molecules were diluted and titrated using the assay buffer in 384-well black/clear bottom assay plates (Corning). The prepared cells were seeded at 40,000 cells/well total by first mixing the prepared cells at a 1:1 ratio, and then adding the cell mixture to the assay plates. The amount of luciferase produced was measured by Bio-Glo Luciferase Assay System reagent (Promega), after which the plates were incubated for 20 minutes at room temperature, and luminescence detected with EnVision plate reader (PerkinElmer). For single point assay, ESN samples were tested at 100-fold dilution. For positive wells, the ESN were
quantitated and normalized prior to potency screening. For potency determination, ESN samples or purified antibodies were serially titrated in assay media and used to block CHO-PD-L1 stimulation of human PD-1 reporter cells. The number of antibodies showing desired activity during single concentration screening are shown in Table 2. TABLE 2. Functional binders from the panel of PD-L1 specific binders Harvest Functional binders Ag-specific binders 1 347 1032 [358] The a
c e a o es ee u a o s o ee e e poe cy o e a - -L1 antibodies. The activity of select anti-PD-L1 antibodies in shown in Table 3. TABLE 3 Potency (pM) Ab ID n 1 n 2 3.
y p y y [359] The potency of anti-PD-L1 antibody, an immune checkpoint inhibitor, can be measured in vitro in a MLR assay. This is a co-culture assay using monocyte derived mature human dendritic cells mixed with allogenic human CD3+ Pan T cells. Immature dendritic cells and Pan T cells were purchased from Astarte (Bothel, Washington; currently known as Cellero). Immature dendritic cells were further differentiated into mature dendritic cells using recombinant human IL-4, GM-CSF, and TNF-a (CellXVivo™ Human Monocyte-Derived Dendritic Cell Differentiation Kit R&D Systems #CDK004). After 3 days, suspended and loosely attached mature dendritic cells were harvested for MLR, washed, and resuspended in X- Vivo15 (Lonza BE02-054Q) assay media. Pan T cells were thawed, washed, and resuspended with X- Vivo 15 media. Antibodies were resuspended and serially diluted 3-fold serial 10 dilutions in X-Vivo15 media. In a 96-well round bottom sterile tissue culture plate, 200,000 Pan T cells, 20,000 mature dendritic cells, and anti-PD-L1 antibody were plated in 200μL final volume per well. Initial concentration of antibodies in plate was at 150 nM, 3-fold serial for 10 dilutions in triplicate to determine the potency. Cells
with no antibody served as negative control. MLR assay incubated at 37oC/5% CO2 for 3 days (~72 hours) and supernatant harvested and frozen. Supernatant was tested for Il-2 release using Mesoscale Discovery (MSD) human IL-2 V-Plex assay (MSD# K151QQD). The potency of select anti-PD-L1 antibodies shown in Table 4. TABLE 4 Potency (pM) Ab ID n=1 n=2
4. Granzyme B release with crosslinking in pre-activated primary human T-cells [360] Frozen purified human T-cells were purchased from Biospecialty. They were added to 6-well TC plates pre-coated with 1ug/mL of anti-huCD3 (Clone OKT3, eBioscience, cat#16-0037), incubated overnight at 37oC, 5%CO2 for pre-activation. On the second day of assay, the hybridoma exhausted supernatants and control antibodies were prepared to desired concentration in ICM media, and then were added to 96-well high binding plates (Costar, cat#3369) pre-coated with 200 ng/mL of anti-huCD3 and 1µg/mL of goat anti-human IgG Fc antibody (Jackson, cat#31125) for crosslinking of testing samples. The pre-activated T-cells were collected and washed. They were counted and resuspended to 0.4x106/mL in ICM, and then transferred to high binding assay plates contained testing samples, at the final of 40,000 cells in 100uL volume per well. Assay plates were incubated in 37oC, 5%CO2 incubator overnight. On Day 3 of assay, assay supernatants were transferred to 96-well high binding plates for Granzyme B detection, using Granzyme B ELISA kit (MabTech, cat# 3485-1H6). OD (optical density) value of each well was obtained using a microplate reader (Tecan GENios) set to 450nm. [361] The individual OD value from Granzyme B ELISA for select antibodies run at 500 ng/mL (6C7 and 6F9) or ~85 ng/mL (14A5 and 19G1) are shown in the Table 5. TABLE 5. Granzyme B release results for select anti-4-1BB antibodies. Antibody ID Single Point Single Point activit (n=1) activit (n=2)
19G1 1.307 1.282 Control 0.220 0.206 5. Pre-activated prim
inking [362] Frozen purified human T-cells were purchased from Biospecialty. They were added to 6-well TC plates pre-coated with 1 µg/mL of anti-huCD3 (Clone OKT3, eBioscience, cat#16-0037), incubated overnight at 37oC, 5%CO2 for pre-activation. On the second day of assay, the pre-activated T-cells were collected and washed with ICM. They were counted and resuspended to 1x106/mL in ICM contained final of 200 ng/mL of anti-huCD3, and then transferred to high binding assay plates, at the final of 50,000 cells per well. The hybridoma supernatants and control antibodies were prepared to desired concentration in ICM media, and then were transferred to 96-well high binding plates contained the pre-activated T-cells. Assay plates were incubated in 37oC, 5%CO2 incubator for 72 hours. On Day 5 of assay, cells were transferred to a v-bottom plate for proliferation readout, using Click-iT Plus EdU Alexa Fluor 488 Flow Cytometry Assay Kit (Molecular Probes, Cat# C10633). Read out the EdU positive cells by flow cytometry analyzer (BD FACSCanto). [363] The hybridoma supernatants were screened at single point of 1ug/mL. The individual value of percentage of EdU positive cells for select antibodies is shown in Table 6. 6. Pre-activated primary human T-cells proliferation with crosslinking [364] Frozen purified human T-cells were purchased from Biospecialty. They were added to 6-well TC plates pre-coated with 1 µg/mL of anti-huCD3 (Clone OKT3, eBioscience, cat#16-0037), incubated overnight at 37oC, 5%CO2 for pre-activation. On the second day of assay, the hybridoma supernatants and control antibodies were prepared to desired concentration in ICM media, and then were added to 96- well high binding plates (Costar, cat#3369) pre-coated with 200 ng/mL of anti-huCD3 and 1 µg/mL of goat anti-human IgG Fc antibody (Jackson, cat#31125) for crosslinking of testing samples. Incubated at 37oC, 5%CO2 for 1 hr, followed by PBS wash to remove the unbound samples. The pre-activated T-cells were collected and washed with ICM. They were counted and resuspended to 0.4x106/mL in ICM, and then transferred to high binding assay plates contained testing samples, at the final of 40,000 cells in 100 µL volume per well. Assay plates were incubated in 37oC, 5%CO2 incubator for 96 hours. On Day 5 of assay, T-cells proliferation were determined with CellTiter GloTM (Promega, CXat#G755B). Luminescence value of each well was obtained using a microplate reader (Tecan). [365] The proliferation was calculated as fold increase over anti-huCD3 alone and plotted against the antibody log concentration using GraphPad Prism (Version 7.04) log (Agonist) vs. response -- Variable
slope (four parameters). The transit EC50 value and fold induction for select antibodies are shown in Table 6. TABLE 6. Functional Activity of select antibodies against 4-1BB Proliferation Antibody ID Single Point without Potency with Potency with crosslinking
. rmary e n ng ssays or - [366] The binding of hybridoma supernatants to PDL1 expressed by primary human and cynomolgus monkey cells were tested by flow cytometry. For human primary cell binding assay, purified human T cells (Biological Specialty Corp.) were suspended at a concentration of 2.5x106 cells/mL. T cells were stimulated with 5 µg/mL of anti-human CD3 clone OKT3 (eBioscience) and 1 µg/mL of anti-human CD28 (BD Pharmingen) for 72 hours at 37 °C/5% CO2 in a plate that had been pre-coated with 5 µg/mL anti mouse IgG Fc (Pierce). After 72 hours, cells were removed, washed and suspended at a concentration of 0.5x106 cells/mL with 10 ng/mL of IL-2 (Pepro Tech). Cells were then incubated for another 48 to 72 hours at 37 °C/5 % CO2. For cynomolgus primary cell binding assay, cynomolgus PBMCs (SNBL) were thawed and suspended in a concentration between 4x106 and 5x106 cells/mL. PBMCs were stimulated with 1 µg/mL of anti-human CD3 clone SP34 (BD Pharmingen) and 1 µg/mL of anti-human CD28 (BD Pharmingen) for 72 hours at 37 °C/5 % CO2 in a plate that had been pre-coated with 5 µg/mL anti-mouse IgG Fc (Pierce). After 72 hours, cells were removed, washed and suspended at a concentration of 0.5x106 cells/mL with 20 ng/mL of IL-2 (Pepro Tech). Cells were then incubated for another 48 to 72 hours. After the final incubation, cells were prepared for flow cytometry by incubation with normalized hybridoma supernatants, positive control antibodies and isotype control antibodies at 1 µg/mL final concentration. Alexa Fluor 647 AffiniPure F(ab’)2 Fragment Goat Anti-Human IgG (H+L) (Jackson ImmunoResearch) was used for secondary detection and YoPro1 (Invitrogen) was used for a live/dead cell stain. Cells were then run on BD FACSCanto II flow cytometer to detect anti-PDL1 antibody binding. The results for select antibodies are expressed as FACS geomean of PDL1 expressing cells and data are shown in Table 7. TABLE 7. Binding of select anti-PD-L1 antibodies to primary human and cynomolgus T cells
Antibody ID Primary Cyno (FACS Geomean) Primary Human (FACS Geomean)
[367] The binding of hybridoma supernatants to 4-1BB expressed by cynomolgus monkey cells were tested by flow cytometry. For primary cell binding assay, cynomolgus PBMCs (SNBL) were suspended in a concentration between 4x106 and 5x106 cells/mL. PBMCs were stimulated with 1 µg/mL of anti-human CD3 clone SP34 (BD Pharmingen) and 1 µg/mL of anti-human CD28 (BD Pharmingen) for 72 hours at 37 °C/5 % CO2 in a plate that had been pre-coated with 5 µg/mL anti-mouse IgG Fc (Pierce). After 72 hours, cells were removed, washed and suspended at a concentration of 0.5x106 cells/mL with 20 ng/mL of IL-2 (Pepro Tech). Cells were then incubated for another 72 hours. After the final incubation, cells were prepared for flow cytometry by incubation with normalized hybridoma supernatants, positive control antibodies and isotype control antibodies at 1 µg/mL final concentration. Alexa Fluor 647 AffiniPure F(ab’)2 Fragment Goat Anti-Human IgG (H+L) (Jackson ImmunoReserach) at was used for secondary detection and YoPro1 (Invitrogen) was used for a live/dead cell stain. Cells were then run on BD FACSCanto II flow cytometer to detect anti-4-1BB antibody binding. Results are expressed as % positive cells and FACS geomean of 4-1BB expressing cells and data are shown in Table 8. TABLE 8 Antibody binding to primary cynomolgus T cells Antibody ID Primary Cyno (% Positive cells) Primary Cyno (FACS Geomean)
9. Receptor – Ligand Competition Assay [368] PD-1-binding hybridoma supernatants were then tested for their ability to block PD-1 from binding ligand. Competitive binding assays were performed on the antigen-specific hybridoma supernatant samples using FACS on HEK293 cells transiently expressing human PD-1 as follows. HEK293 cells
expressing human PD-1 were mixed with the antibody sample (hybridoma supernatants specific for PD-1) and incubated for 1 hour at 4°C, and then washed. Cells with bound sample were then incubated with huPD-L1-Fc-Alexa647 or huPD-L2-Fc-Alexa 647 (R&D systems, Minneapolis, MN) for 45 minutes at 4°C. The 7-AAD cell viability stain was then added and the cells incubated for a further 15 minutes at 4°C, washed and resuspended in FACS buffer. Samples were analyzed using a BD Accuri™ Flow Cytometer and an Intellicyt HyperCyt autoSampler. The data in the Table 9 reflects that percent inhibition of human PD-L1 of PD-L2 binding to human PD-1 at 1 µg/mL. The data in the Table 10 reflects that percent inhibition of 4-1BBL binding to 4-1BB with anti-4-1BB antibodies. TABLE 9. Competition analysis of anti-PD-L1 antibodies with PD-1 or B7-1 binding to PD-L1 Antibody ID PD-1 Receptor – Ligand assay (% B7-1 Receptor – Ligand assay (% inhibition) inhibition)
Antibody ID Receptor – Ligand assay (% inhibition) Determination based on 80% cut-off 6C7 59% Non-blocker
. pp g - . [369] Exhausted hybridoma supernatants were tested for binding to human 4-1BB, mouse 4-1BB or chimeric proteins containing one domain of human 4-1BB replaced with the corresponding domain of mouse 4-1BB. Briefly, HEK293 cells were transiently transfected with the mammalian expression constructs described above. The following day, 15 µL of exhausted hybridoma media was added to cells expressing each construct. Then, the transfected HEK293 cells were washed and incubated with the nuclear stain Hoechst 33342 and a secondary detection antibody (Goat anti Human IgG Fc Alexa 647 (Jackson ImmunoResearch)). After 15 minutes, the cells were washed, and the cells were analyzed on a BD Accuri flow cytometry machine with Intellicyte autosampler. Based on the binding profiles to the different chimeric molecules, an assessment of the antibody binding domain was determined and is listed in Table 11 for select antibodies. TABLE 11. Mouse cross-reactivity and domain mapping of select anti-4-1BB antibodies.
Antibody ID Domain Mapping Mouse 4-1BB cross-reactive? 6C7 CRD2 Yes
[370] Affinity determination on a panel of 91 anti-PD-L1 mAbs was performed using a IBIS MX96 SPRi instrument (WasatchMicrofluidics, Utah, USA-now Carterra). The capture surface was first prepared through amine coupling of an anti-human Fc goat pAb (Thermo-Fisher, Cat. #31125) onto a Xantec 200m (CM5 analog) chip. The unpurified mAbs, at 10 ug/ml, were cycled over the prepared chip to allow capture of approximately 450 RUs. A six-point, 2-fold serial dilution of the human PD-L1 (1- 239)E3KMut1His(PL39847) protein from 50 nM was injected from the lowest to highest concentration. Following regeneration of the chip, and recapture of the mAb panel, the same method was repeated using the cyno PD-L1 (1-238)-Flag8xHis protein. The data was pre-processed in SPRINT X (IBIS, NL), while the kinetic fits, affinity determination and ranking were performed in Scrubber 2 (Biologic Software, AU) and MS Office 365 Excel (Microsoft, US). Table 12 summarizes the affinity to human and cynomolgus PD-L1. TABLE 12 Affinity to human and cynomolgus PD-L1 Antibody ID Affinity to Human PD-L1 Affinity to Cyno PD-L1
12. Molecular Rescue and Sequencing of PD-L1 and 4-1BB Antibodies [371] RNA (total or mRNA) was purified from wells containing the PD-1 antagonist antibody-producing hybridoma cells using a Qiagen RNeasy mini or the Invitrogen mRNA catcher plus kit. Purified RNA was used to amplify the antibody heavy and light chain variable region (V) genes using cDNA synthesis via reverse transcription, followed by a polymerase chain reaction (RT-PCR). The fully human antibody gamma heavy chain was obtained using the Qiagen One Step Reverse Transcriptase PCR kit (Qiagen). This method was used to generate the first strand cDNA from the RNA template and then to amplify the variable region of the gamma heavy chain using multiplex PCR. The 5’ gamma chain-specific primer
annealed to the signal sequence of the antibody heavy chain, while the 3’ primer annealed to a region of the gamma constant domain. The fully human kappa light chain was obtained using the Qiagen One Step Reverse Transcriptase PCR kit (Qiagen). This method was used to generate the first strand cDNA from the RNA template and then to amplify the variable region of the kappa light chain using multiplex PCR. The 5’ kappa light chain-specific primer annealed to the signal sequence of the antibody light chain while the 3’ primer annealed to a region of the kappa constant domain. The fully human lambda light chain was obtained using the Qiagen One Step Reverse Transcriptase PCR kit (Qiagen). This method was used to generate the first strand cDNA from the RNA template and then to amplify the variable region of the lambda light chain using multiplex PCR. The 5’ lambda light chain-specific primer annealed to the signal sequence of light chain while the 3’ primer annealed to a region of the lambda constant domain. [372] The amplified cDNA was purified enzymatically using exonuclease I and alkaline phosphatase and the purified PCR product was sequenced directly. Amino acid sequences were deduced from the corresponding nucleic acid sequences bioinformatically. Two additional, independent RT-PCR amplification and sequencing cycles were completed for each hybridoma sample in order to confirm that any mutations observed were not a consequence of the PCR. The derived amino acid sequences were then analyzed to determine the germline sequence origin of the antibodies and to identify deviations from the germline sequence. A comparison of each of the heavy and light chain sequences to their original germline sequences are indicated. The amino acid sequences corresponding to complementary determining regions (CDRs) of the sequenced antibodies were aligned and these alignments were used to group the clones by similarity. EXAMPLE 2 [373] This example describes the engineering of Anti-PD-L1 and Anti 4-1BB Monoclonal Antibodies 1. Fv Engineering [374] Monoclonal antibody sequences where analyzed specifically in the Fv domains for structural and chemical liabilities which could cause stability issues. Mutation sequences in the framework regions of the molecule were analyzed for potentially detrimental mutations occurring during somatic hypermutations. Fixes for these residues were to back mutate to the germline residue. Chemical liabilities were identified through sequence analysis for free cysteines, N-linked glycosylation, tryptophan oxidation, asparagine deamination, and aspartic acid isomerization. Replacement residues for those positions were selected after antibody homology modeling through MOE (Molecular Operating Environment by Chemical Computing Group) for best fit residues. Recombinant variants of the parental molecule were made with various combinations of the mentioned fixes. The backbone was switched from the hybridoma IgG2 or IgG4 to IgG1 or IgG1 SEFL2. Detailed information on SEFL2 can be found, e.g., in Jacobsen et al., Engineering an IgG scaffold lacking effector function with optimized developability J. Biol. Chem., 292
(2017), pp.1865-1875; and Estes et al., Next generation Fc scaffold for multispecific antibodies, iScience, Volume 24, Issue 12, 2021,103447. These were produced using standard cloning, expression, and purification technologies. Selected mutational variants were selected based on purification quality, production yields, and functional activity. [375] Table 13 summarizes the engineered 4-1BB antibodies and their characteristics including the % inhibition exhibited in a receptor-ligand competition assay and purification yield. [376] Table 14 summarizes the engineered PD-L1 antibodies and their characteristics including their IC50 and relative potency. Assays were conducted following the protocols described above (“Jurkat human PD-1/NFAT-luciferase reporter assay”), using Jurkat cells stably expressing human PD-1 and NFAT-luciferase reporter and CHO cells stably expressing human PD-L1. “IC50 transit” refers to the concentration of molecule that gives a 50% of max activity. “Relative potency” compares the potency of the clone being tested, relative to the original parental clone, expressed as ratios. Table 13 Characterization of selected 4-1BB antibodies Anti-41BB Antibody CRD R/L % Purification Yield N=1 AC50 IP N=1 Max Name Domain Inhibition mg/L [pM] Activity
Table 14 Characterization of selected PD-L1 antibodies Anti-PDL1 Ab Name IC50 transit IC50 transit Relative Relative / L 1 / L 2 P t P t 2
21B2.011 22 41 0.72 0.85 25F1.002.005 53 66 1.63 1.52
2. Yeast Display Engineering of 41BB Variant 19G1.016.001 From 19G1.016. [377] The 19G1.016 variant Fv sequence was displayed as a Fab on the surface of yeast through a fusion to alpha agglutinin. The degenerate codon NNK was introduced at an isomerization site DGGF (SEQ ID NO: 348) to cover all 20 amino acids. This allowed for selection of molecules based on number of Fabs displayed on the cell surface and amount of binding. Scanning in this region only produced a single weaker variant sequence DGGF (SEQ ID NO: 348) to DGYV (SEQ ID NO: 349). Both sequences were explored as recombinant IgG-scFv molecules with the DGYV (SEQ ID NO: 349) advancing. EXAMPLE 3 [378] This example describes the construction of bivalent bispecific molecules to 4-1BB and PD-L1. A schematic of the steps carried out to produce bispecific molecules are shown in Figure 1B. Details of steps are provided below. 1. Molecule design and generation of lead panel [379] Selection of anti-PD-L1 leads. As described in Example 1, XENOMOUSE® campaigns for PD-L1 were run with a variety of immunogens. There were 13 parental anti-PD-L1 mAbs selected to move into antibody optimization. Sequence analysis was run on all 13 looking for covariance violations, tryptophan oxidation sites in CDR3s, Isomerization sites, and Deamidation sites. These were substituted to low liability risk residues which could retain functional activity. Molecular modeling was conducted as a quality control check on designs. The final panel resulted in 15 sequences which had lower liability risk and retention of function. These 15 sequences were then converted to scFvs and fused to the control mAb 655-314 to assess functional conversion to a scFv format. From the 15 sequences 3 appeared viable candidates to move forward into 4-1BB fusions. [380] Selection of anti-4-1BB leads. As described in Example 1, XENOMOUSE® campaigns for 4-1BB were run with a variety of immunogens. There were 16 parental anti-PD-L1 mAbs selected to move into antibody optimization. Sequence analysis was run on all 16 looking for covariance violations, tryptophan oxidation sites in CDR3s, Isomerization sites, and Deamidation sites. These were substituted to low liability risk residues which could retain functional activity. Molecular modeling was conducted as a quality control check on designs. The final panel resulted in 12 sequences which had lower liability risk, retention of function, and crosslinking dependent. The 12 mAbs were stressed to evaluate if the remaining
predicted liabilities were true liabilities. One mAb, 19G1.016, was shown to have isomerization DG. A yeast display method looking for a viable replacement of the DGGF site (SEQ ID NO: 348) using 4 serial NNK codons (SEQ ID NO: 350) resulted in a new variant which mutated residues to DGYV (SEQ ID NO: 349). The theory is that the Y could inhibit the isomerization more since it is less flexible than a G. This substitution also appeared to be more potent. [381] Bivalent Bispecific Panel. In parallel to the yeast display selection described in Example 2, an optimized bivalent bispecific panel was designed. Briefly, optimized Fv sequences (described in Example 2) were used to create two types of bivalent bispecifics. The first type is the fusion of a single chain Fv domain containing a 3xG4S linker (SEQ ID NO: 346) and an engineering disulfide bond to the c-terminus of the CH3 domain of a full antibody, after removal of the GK sequence. The first type is referred to as an IgG-scFv and a schematic is shown in Figure 2A. The second type of bivalent bispecific utilizes the charge pair technology to fuse an additional Fab domain to the c-terminus of the CH3 domain of a full antibody, after removal of the GK sequence. The charge pair mutations hinder the formation of incorrect heavy chain/light chain pairing. The second type is referred to as an IgG-Fab and a schematic is shown in Figure 2B. Both molecules use an IgG1 SEFL2 Fc domain. The anti-4-1BB and anti-PD-L1 Fv domains were positioned in combinations in both orientations in each format. [382] These molecules were produced using standard cloning, expression, and purification technologies. Purification was done in 1 of 2 methods depending on the modality. The IgG-Fab molecules were purified by standard ProA LFAS, whereas the IgG-scFvs were purified through a triple tandem LFAS system (ProA, In Line Dilution, SEC). [383] The functional activity in a co-culture CHO (+/- huPD-L1) artificial APC with PDMCs assay was the primary selection criterion. Other selection experiments were the yield (>80m/mL), Quality, (MCE/SEC), qualitative viscosity readout, and MSQC. The IgG-Fabs were further selected using non-reduced MSQC to evaluate correct chain pairing ratios. Additional selections were conducted based on purification quality, production yields, and functional activity. [384] Figure 3 provides a table listing exemplary bispecific molecules constructed and characterized. 2. Additional selection of lead molecules. [385] The initial selection criteria for the 14 (7 IgG-Fab/7 IgG-scFv) molecules were expanded to 18 to include the yeast display fix resulting in anti 4-1BB 19G1.016.001 and two new variants anti 4-1BB 14G12.017.001 and anti 4-1BB 6C7.018 along with one new format, referred to as scFab-Fab hetero-IgG. [386] A number of bispecific molecules were not selected due to purification, viscosity, aggregation, stability, and heavy;light chain pairing issues. The final 10 molecules included 4 IgG-scFv, 1 scFab-Fc- Fab, 5 IgG-Fabs and were subjected to further analysis. Briefly, the conformational integrity of 10
bispecific molecules was validated by LC-MS analysis of non-reduced materials (Agilent 6224 TOF MS). Bispecific molecules with engineered charge pair mutations (CPMs) to drive correct heavy/light chain pairing were also analyzed by LC-MS after proteolytic cleavage below the hinge disulfides with IdeS protease (Genovis) to confirm correct heavy/light chain pairing. The biophysical stability of the molecules at 70 mg/mL total protein concentration were also validated. The thermostability of the molecules were assessed by differential scanning calorimetry at 1 mg/ml (Malvern MicroCal System). Tryptophan oxidation, Asparagine deamidation, and Aspartic acid isomerization in CDR regions were also assessed by peptide mapping analysis of both 2 week 40⁰C and 192 klux.hr stressed materials (Thermo Q-Exactive LC-MS System). The results are summarized in the table of Figure 4. Figure 5 provides LC-MS spectra of IdeS-protease cleaved samples confirming heavy-light chain pairing in molecules with engineered charge pair mutations. [387] Four preferred candidates were selected based on these analyses: 3 IgG-scFv (11250, 11252, 11253), and one IgG-Fab (11259). EXAMPLE 4 [388] This example demonstrates PDL1-41BB bispecific antibodies can simultaneously block PD-L1/PD1 inhibitory pathway and activate 4-1BB costimulatory pathway in primary T cells in a PD-L1 dependent fashion. [389] Generation of artificial antigen presenting cells (aAPC). CHOK1 expressing αCD3scFv clones under Hygromycin B selection were generated using transient transfection of αCD3scFv in pcDNA3.1- HygroB. CHO/αCD3scFv cells were also sequentially transfected with pcDNA3-1-Zeo_HuPD-L1 and under Hygromycin B and zeocin selection, sorted for high PD-L1 expression and then transduced with GFP retroviral expression vector containing Hu CD32a/FcgRII. Final aAPCs were sorted for Low αCD3scFv, high human PD-L1 and CD32a expression. [390] aAPC and human primary T cell coculture assay. CHO cells expressing anti-humanCD3scFv, human PD-L1 and human Cd32a were used as aAPCs in co-culture assay with human Pan T Cells to test PDL1-41BB bispecific molecules in comparison to anti-PD-L1 and anti-4-1BB mAbs as single agents or in combination. PD-L1x4-1BB bispecific molecule, anti-PD-L1, and anti-4-1BB antibodies were serially diluted from 50nM (final in assay) and added in triplicate to co-culture assay with aAPCs and Pan T-cells (isolated from human PBMC using Miltenyi Pan T cell isolation Kit # 130-096-535) at 1:2 ratio (50,000 aAPCs + 100,000 Pan T cells) per well in 96-well tissue culture plate. IL-2 released in supernatants were quantitated in MesoScale Discovery IL-2 V-Plex assay (#K151QQD). IL-2 concentrations were analyzed using GraphPad Prism 4 parameter curve fit. [391] By engineering a PD-L1-4-1BB bispecific molecules on a FcR-binding inert backbone , it is believed that robust anti-tumor response is induced through activation of 4-1BB and inhibition of PDL1
simultaneously, with limited systemic hepatotoxicity effects by requiring PD-L1 expression for 4-1BB crosslinking. [392] Indeed, it was observed in an in vitro coculture assay, when primary T cells were activated with artificial antigen presenting cells in the presence of PDL1-41BB bispecific molecules, the molecule induced robust T cell activation, greater than either 4-1BB or PD-L1 mAb alone or in combination (Fig. 6A). Moreover, when the PD-L1 expression was absent from the surface of artificial antigen presenting cells, PD-L1-4-1BB bispecific molecules were unable to induce any T cell activation upon coculture (Fig. 6B), demonstrating the requirement of PDL1 expression for the bispecific molecules to activate 4-1BB costimulatory pathway in T cells. EXAMPLE 5 [393] This example demonstrates an in vivo tumor study to evaluate a bispecific molecule of the present disclosure. [394] Results from an in vitro assay demonstrated that the potency of the bispecific antibody was at least as good as the combination of individual mAb used to construct the bispecific molecule. To evaluate the bispecific molecule’s efficacy in vivo, syngeneic murine MC38 colon adenocarcinoma models were used to assay the in vivo effects of a bispecific molecule of the present disclosure. [395] All mice were sourced from Charles River Laboratories (Hollister, CA site) and were provided water and chow ad libitum and maintained in a pathogen-free facility. Mice used in syngeneic tumor experiments were 6-8 weeks of age at the time implant. MC38 cells were inoculated in the right flank at 3 × 105 cells per implant and allowed to grow for 8 days. For treatment, the mice were then randomized by tumor volume (50-100mm3) and treated with anti-PD-L1 blocking mAb (MIH5), anti-4-1BB agnostic mAb (LOB12.3) in mouse IgG1 backbone, anti-4-1BB agonistic mAb (LOB12.3) in mouse IgG2a LALA-PG mutant backbone (incapable of Fc receptor binding), 4-1BB-PD-L1 surrogate bispecific antibody (in mouse IgG2a LALA-PG backbone) or isotype control mAb (mouse IgG2a LALA-PG). All the mAb were dosed i.p. at 200 µg per mouse, while 4-1BB x PD-L1 bispecific molecules were dosed i.p. at 267 µg per mouse. Tumor were measured with a digital caliper twice every week until the end of study. Animals are removed from study when tumor volume exceeds 800mm3. [396] The results are shown in Figures 7A to 8B. Individual tumor growth curves (Fig.7A-7D), average tumor volumes (Fig.8A) and cumulative survival curves (Fig.8B) of MC38 tumor bearing mice following treatment as indicated. Mice received the indicated treatments once the tumors were randomized. Tumor were measured with a digital caliper twice every week until the end of study. Survival curves were analyzed for statistical significance using the Kaplan-Meier estimator with Mantel-Cox log rank test to compare curves. As shown in these figures, 41BB-PDL1 surrogate bispecific antibody achieved similar efficacy as the combination of blocking anti-PDL1 and agonistic anti-41BB mAbs, where both treatment
groups saw 8 out 10 mice were tumor free at the end of study. At the meantime, mice treated with 41BB mAb on a LALA-PG backbone that couldn’t bind to Fc receptor for crosslinking had uncontrollable tumor growth, even in combination with anti-PDL1 blocking mAb. EXAMPLE 6 [397] This example describes epitope selection of anti-4-1BB antibodies. [398] To screen Cyno 4-1BB cross reactive samples for binding on five Human-Mouse 4-1BB chimera constructs and better characterize the extracellular domain binding characteristics of this antibody panel, chimeric 4-1BB constructs of interest were transiently expressed on 293T cells and antibody binding was detected by flow cytometry. For transfection, 10 million 293T cells were diluted to 1 million cells per mL in 10mL of FreeStyle 293T expression medium and allowed to incubate on a shaker at 37°C for 3 hours.10 ug of DNA construct (see Table 15 for construct details) and 10uL of 293Fectin were each added to 500uL OptiMEM separately and incubated at room temperature for 5 minutes. The 2 solutions in OptiMEM were then mixed together by inversion and allowed to incubate for 20 minutes. The DNA, 293Fectin, and OptiMEM solution was then added drop-wise to the cells. After 24 hours, the cells were tested for expression of target protein using flow cytometry. The E3K immunogenic peptide was fused to the c-terminus of 4-1BB to increase the immunogenic response. Table 15 Construct Swapped Region
[399] 15 uL of each antibody of interest in hybridoma supernatant was added to plates at 2 ug/mL antibody concentration. Transfected 293T cells were diluted to 30,000 cells per 15 uL in FACS buffer (1X PBS + 2% FBS), and 15 uL of cell suspension was added to the wells containing supernatant (1:2 dilution of antibodies). The primary antibodies were incubated on cells for 1 hour at 4°C. Following 2 washes with 75 uL PBS per well, 15 uL of secondary antibody at a concentration of 10 ug/mL (Goat anti-Human AlexaFluor 647, Goat anti-Mouse Dylight, or Goat anti-Amermenian Hamster Alexa 647 (Jackson ImmunoResearch) and 7-AAD (Sigma) at 5 ug/mL was added to the appropriate wells containing 15uL of cell suspension (1:2 dilution) and allowed to incubate for 15 minutes at 4 degrees Celsius. After a wash with 75 uL of FACS buffer and another wash with 75 uL of PBS, cells were resuspended in 30uL of FACS buffer. Spherotech beads were prepared by diluting them 1:160 in FACS buffer to be used for fluorescence compensation. Samples and beads were read out on the Accuri Flow Cytometer with Intellicyte autosampler. [400] It has been reported that the second and third CRDs (CRD2 and CRD3) appear to play a major role in modulating the binding affinity and specificity. Therefore, agonist antibodies that bind to CRD2 and CRD3 would likely mimic 4-1BB ligand binding and activate 4-1BB signaling pathway. Accordingly, a number of antibodies that bind to CRD2 and CRD3 were identified and characterized (Figure 12). [401] CRD1, also named pre-ligand-binding assembly domain (PLAD), is physically distinct from the domain that forms the major contact with ligand, and there is increasing evidence for PLAD-mediated receptor association and trimerization (Yi et al., PLoS One.2014; 9(1): e86337). Further, Chin et. al. (Nature Communications 9, Article number: 4679 (2018)) reported the structures of the human 4-1BB/4- 1BBL complex, the 4-1BBL trimer alone, and 4-1BB bound to utomilumab (Pfizer) or urelumab (BMS). The 4-1BB/4-1BBL complex displays a unique interaction between receptor and ligand when compared with other TNF family members. Utomilumab, a ligand-blocking antibody, binds 4-1BB between CRDs 3 and 4. In contrast, urelumab binds 4-1BB CRD1, away from the ligand binding site. [402] Despite that utomilumab and urelumab show similar binding affinities, these antibodies demonstrate distinct differences in their agonist activity and reported toxicity. Chin reports that antibodies bind 4-1BB in markedly different modes. In view of these differences, antibodies that bind to different CRDs of 4-1BB were selected and investigate. [403] As the 4-1BBL binds to the 4-1BB receptor around the entire CRD2 and part of CRD3 regions, any given antibody binds to the same regions would likely trigger similar conformational changes as ligand binding and therefore be agonistic. However, the above-mentioned region binding will also likely block the natural receptor/ligand binding, and the impact of such interference in vivo is unknown. Therefore, identifying an agonistic antibody that binds to a region outside of receptor/ligand interaction sites, such as CRD1 of 4-1BB receptor, could be beneficial to potentially leave the natural receptor/ligand interactions
largely intact while still be able to trigger robust receptor signaling. Accordingly, antibodies that bind to CRD1 were identified and characterized (Figure 9). [404] Figure 9 summarizes epitope regions of certain 4-1BB antibodies disclosed herein. EXAMPLE 7 [405] This example describes the generation of UniDab® anti-4-1BB binding domains. [406] 18 UniRat® animals were immunized with human 4-1BB protein (Sino Bio 10041-H08H). Animals were immunized with Titermax/Ribi adjuvant and Freunds adjuvant. Pre-harvest bleeds were collected and serum titers against the immunogen (human 4-1BB) was completed. The draining lymph nodes of immunized animals were harvested post-immunization and RNA was extracted from the B-cells in the lymph nodes. Left and right lymph nodes for each animal were analyzed separately. cDNA samples containing the full VH region of the heavy chains were sequenced using NGS. Sequences from the immunizations was analyzed and the sequences showing the greatest evidence for antigen-specific affinity maturation were selected high throughput cloning and expression.673 VH genes representing 202 CDR3 families, were identified as top candidates for assembly and expression in HEK 293 cells. These were cloned into a proprietary expression vector containing the human IgG1 constant region with the CH1 domain removed. Subsequently, these UniAb® containing heavy chain only expression vectors were expressed individually in HEK293 cells to produce fully human heavy chain only UniAb®. These UniAb® were functionally evaluated in a high throughput manner for both on and off target cell binding and protein binding. Ultimately, 25 leads representing 21 CDR3 sequences were scaled up by 24 well transfection and assessed for crosslinker dependent activity, developability, epitope, and binding affinity. Subsequence characterization of representative family members revealed. EXAMPLE 8 [407] This example describes the characterization of UniAb® anti-4-1BB binding domains. The 4-1BB molecules are also referred to as CD137 in the Tables and Figures. Table 16 Unique CDR3 Goal antibodies Families
[408] The goal of NGS and rank analysis was to identify 420 unique UniAb® sequence candidates and maximize sequence diversity. In primary screening, gene assembly, expression and binding assays were performed. UniAb® were assessed for protein binding by ELISA and cell binding was flow cytometry. The top 25 candidates were identified based on desirable binding to 41BB and not to any off-target cell line. The top candidates were then evaluated by 24 well transient expression after Protein A purification. These molecules were assessed for the following parameters: (1) cell binding dose curves with on-target (CHO-41BB) and off-target CHO-S cells; (2) cross-linker dependent activity; (3) assessing developability through measurements of aggregation propensity by SEC; (4) performing kinetic analysis by Octet to determine KD measurements; and (5) competition group assessment by Octet with benchmark antibodies. [409] NF-κB reporter activity of 4-1BB UniAb® in the presence and absence of cross-linker.4-1BB UniAb® dilutions and cross-linker solution (Goat F(ab)2 anti-human IgG Fc, Abcam, Cat # ab98526) were prepared. On the day of the assay, assay buffer (RPMI-1640 + 1% FBS) was prepared using FBS and RPMI-1640 provided in the reporter assay kit (Promega, Cat# CS196005). NF-κB-luc2/4-1BB Jurkat cells were thawed in a water bath and removed immediately prior to a complete thaw. Cells from each thawed vial (0.5 mL) were added to 9.5 mL assay buffer. The Jurkat cells and CD137 UniAb® /cross-linker or CD137 UniAb® or assay buffer (blank) were added to the plate. The plates were incubated at 37 °C, 8% CO2 for 6 hours. The reporter substrate reagent was prepared by reconstituting one vial of Bio-Glo™ Luciferase Assay Substrate with the Bio-Glo™ Luciferase Assay Buffer. After 6 hours of incubation, plates were removed from the incubator and allowed to equilibrate to room temperature for 10 minutes. Bio-Glo™ Reagent were at ambient temperature prior to addition to the plates.75 µL of reconstituted substrate was added to each well and the reaction was allowed to stabilize for 10 minutes at room temperature. Reporter signal was detected using a plate luminometer and fold induction was calculated using the average signal from “blank” wells on each plate as the denominator (background). In this assay, in the absence of cross-linker, there is no activation of CD137 receptor and luminescence signal is absent. In the presence of cross-linker, it induces the crosslinking of CD137 and a dose dependent luminescence. Emax is reported as fold induction over background baseline NF-κB induction in no antibody (blank) treatment condition (target cells + reporter cells alone). Results from this assay are provided in FIGs.10A-10B. [410] Cell binding dose curves for CD137 UniAb® molecules using primary human and Cyno cells. Human PBMCs were activated with 2 ng/mL of recombinant human IL-2 (Rnd Systems, Cat# 202- IL-010), and seeded onto OKT3-coated petri dishes. After 3 days of activation at 37ºC, 8% CO2, PBMCs were harvested and enriched for T-cells by negative selection (Miltenyi Biotec, Cat# 130-096-535). [411] Cell binding dose curves with activated T-cells, CHO_huCD137, CHO_cynoCD137 and CHO- S. Human PBMCs were activated with 2 ng/mL of recombinant human IL-2 (Rnd Systems, Cat# 202-IL-
010), and seeded onto OKT3-coated petri dishes. After 3 days of activation at 37ºC, 8% CO2, PBMCs were harvested and enriched for T-cells by negative selection (Miltenyi Biotec, Cat# 130-096-535). Increasing concentrations of purified CD137 UniAb® molecules, or heavy chain only isotype control, were incubated with various 4-1BB-expressing or 41-BB-negative cell lines. Stained cells were assayed using a standard flow cytometry protocol on a BD FACS Celesta HTS plate reader with a goat F(ab')2 anti-human IgG-PE secondary detection antibody (Southern Biotech, Cat# 2042-09). MFI from the PE channel was plotted against the log of the concentration of the respective molecules. Dose curves were fitted to a 4- parameter logistic regression and EC50 values were calculated where applicable. [412] EC50 and Emax from cell binding data. EC50 and Emax values from the cell binding data described above are summarized in Table 17. The table summarizes the Emax and EC50 values for cell binding to CHO_Human 4-1BB cells, CHO_Cyno 4-1BB cells, and CHO-S (off-target) cells. All EC50 values are shown in units of ng/mL. ND indicates weak or no detectable binding, so EC50 values could not be calculated. Table 17 Activated T-cells CHO-hu4-1BB CHO-cy4-1BB CHO-S Clone IDs Emax EC50 Emax EC50 Emax EC50 Emax EC50
[413] Epitope binning data from CD137 UniAb® molecules. An Octet QK 384 system equipped with HIS1K - Anti-Penta-HIS sensors was used for epitope binning experiments. The binning experiments were performed using an in-tandem competition assay format. Briefly, recombinant huCD137-his protein was coated on the sensors, then the molecule (indicated by the rows in Table 18) was loaded to saturate the immobilized antigen. Then the next molecule was added and its ability to bind to the pre-saturated antigen was evaluated. The data was analyzed using Fortebio data analysis HT software. Four main bins were identified by epitope binning analysis and are summarized in Table 18. Table 18 Clone ID Epitope Bins
386334 2 386346 1AB
[414] Kinetics of binding for lead UniAb® to human 41BB recombinant protein were evaluated on a Biacore 4000 instrument using a Protein A coated chip. Tm and Tagg were measured using UNcle platform and protein aggregation was measured by SE-HPLC. For Tm and Tagg measurements, higher Tms and Taggs are generally preferred, and protein samples were heated from 250C to 700C at 10C ramp rate. The results are summarized in Table 19. Table 19 Clone ID Avg Tm1 (°C) Avg Tagg 266 (°C) KD (nM) 386352 538 522 142
EXAMPLE 9 [415] This example describes the construction of additional bivalent bispecific molecules to 4-1BB and PD-L1. A schematic of the steps carried out to produce bispecific molecules are shown in FIG.12. 1. Molecule design and generation of lead panel
[416] Selection of anti-PD-L1 leads. As described in Example 1, XENOMOUSE® campaigns for PD-L1 were run with a variety of immunogens. There were 13 parental anti-PD-L1 mAbs selected to move into antibody optimization. Sequence analysis was run on all 13 looking for covariance violations, tryptophan oxidation sites in CDR3s, Isomerization sites, and Deamidation sites. These were substituted to low liability risk residues which could retain functional activity.92 engineered anti-PD-L1 mAbs were generated and screened for retained function in the absence of liabilities. From this screen, 9 variants of anti-PDL1 antibodies were advanced to the construction of bispecific modalities. [417] Selection of anti-4-1BB leads. From the 4-1BB immunization panels, there were 16 parental anti- 4-1BB antibodies and VH domains were selected to move into antibody optimization. Sequence analysis was run on all 16 looking for covariance violations, tryptophan oxidation sites in CDR3s, Isomerization sites, and Deamidation sites. These were substituted to low liability risk residues which could retain functional activity.249 engineered anti-4-1BB binders were generated and screened for retained function in the absence of liabilities. From this screen, 14 variants of anti-4-1BB antibodies and VH domains were advanced to the construction of bispecific modalities. [418] Selection of anti-PD-L1 anti-4-1BB bispecific binder combinations and modalities. The engineered 6 anti-PD-L1 and 14 anti-4-1BB binders were combined in 11 bispecific formats as described in the bispecific formatting section of FIG.12 and illustrated in FIGs.13A-13K and 14A-14D. Resulting molecules were generated by recombinant cloning, expression, and purification.13 molecules in 7 formats with favorable manufacturability, stability, and function were advanced to large scale production for more extensive analysis. Of those, 10 molecules passed large scale production criteria and were advanced to protein characterization. EXAMPLE 10 [419] This example describes the biological activities and properties of exemplary bispecific molecules. Table 20 summarizes the structures of these molecules and the sequences can be found in the Sequence Table. “CC” refers to cysteine clamps introduced into the scFv domain. Table 20 name 41BB parental PDL1 Format Constant Additional Modifications parental
56761 386354 VH 21B2 [Fab*VH]-heteroFc huIgG1 SEFL2.2, YTE 56762 380984 VH 21B2 IgG-VH (C2) huIgG1 SEFL2.2, YTE
[420] Biophysical properties of the bispecific molecules. FIGs.15A-15B summarizes biophysical properties of exemplary bispecific molecules, titled 56040-1, 44988-3, 56132-6, 56762.4, 56639-3, 56761- 4, 56042-1, and 56039-1. Purity and identity confirmation of purified samples were measured by mass spectrometry with each measured molecular weight matching the calculated molecular weight. Lability of predicted hotspots was measured by peptide mapping after stress at 40oC for 4 weeks and also after exposure to visible light. Viscosity of molecules concentrated at 70 mg/mL and at 140 mg/mL was measured using cone and plate methods with a target of less than 10 cP at 70 mg/mL. Melting temperature was measured by Differential Scanning Calorimetry (DSC). Clipping after stress of 20oC and 40oC incubation for 2 weeks and 4 weeks was measured by reduced capillary electrophoresis in a sodium dodecyl sulfide gel (rCE-SDS). Molecules with a change of less a than 2% change in molecular weight species during the stress were ranked more highly than molecules with greater than or equal to a 2% change in molecular weight species. Thermal stability after stress of 20oC and 40oC incubation for 2 weeks and 4 weeks, after exposure to visible light, and after repeated freeze thaw cycles was measured by size exclusion chromatography (SEC). Molecules with a change of less a than 2% change in molecular weight species during the stress were ranked more highly than molecules with greater than or equal to a 2% change in molecular weight species. The presence of structural isoforms of equal molecular weight and lack of solubility was tested for after stress at 40oC for 4 weeks by analytical hydrophobic interaction chromatography (aHIC). [421] Biological activities of the bispecific molecules. Table 21 summarizes potency of the bispecific molecules disclosed herein. Table 21 56132-6 56762-4 56040-1 44988-3
IFNg EC50 (pM) PDL1+ 20% 88.7 62.6 94.6 50.3 IL2 EC50 (pM) PDL1+ 20% 104.5 94.8 152.9 69.6 TNF EC50 ( M) PDL1+ 20% 837 629 936 618
[422] FIG.16A shows the pan T cell response to the bispecific molecules when co-cultured with CHO-K1 artificial APCs that express aCD3scFv and PD-L1. PDL1x41BB bispecific molecules are then titrated in to enhance T cell activity. The aAPCs also express CD32 to enable cross-linking with the 4-1BB 14A5 control mAb. Activity of the molecules was assessed using aAPCs that express high levels of PDL1. T cell response were determined by the amount of IFN-g secreted into the co-culture supernatant. [423] FIG.16B shows the ability of the bispecific molecules to restore T cell activation in a mixed lymphocyte reaction (MLR). MLR assays use T cells from one individual and MHC class II-expressing cells from a second individual. Interaction of the alloreactive TCR with the MHC class II molecule induces T cell activation resulting in T cell proliferation and IL-2 production. Interaction of PD-1 on the T cells with PD-L1 on the dendritic cells decreases T cell activation and the concomitant IL-2 production. Blocking the PD-1/PD-L1 interaction restores T cell activation and the concomitant IL-2 production. [424] FIG.16C shows the activity of the anti-PDL1 arms of PDL1x41BB bispecific molecules tested in the PD1/PDL1 reporter assay developed by Promega. [425] Table 22 summarizes the binding affinities of the bispecific molecules to its two targets. Table 22 Affinity Binding (1:1) (SPR-based) Test Binding to human 4-1BB Binding to cyno 4-1BB Human Articles KD M Rm x C mm nt KD M Rm x C mm nt Vs Cyno
11253 2.70E-08 199 KD ~30 nM >1.0E- 123 KD >100nM <10 07 Test Binding to human PD-L1 Binding to cyno PD-L1 Human o
[426] Finally, FIG.17 shows the PD-L1 Dependent 4-1BB Agonism of the bispecific molecules. There was no activity of T cells driven by the bispecific molecules for PDL1-negative target cells (FIG.17, showing the result of one exemplary bispecific moelcules.). The bispecific molecules in FIG.17 and anti- PD-L1 mAb required the PD-L1 expression in target cells for their activities, whereas anti-4-1BB mAb activates T cells regardless of PD-L1 expression. [427] One particular observation from our data is that for 4-1BB agonists, low affinity (rather than high affinity) binders delivered greater activity through increased clustering. For example, molecule 56762 has lower affinity to human 41BB (~1000-fold lower affinity) compared to a reference molecule (data not shown). Surprisingly, molecule 56762 shows higher activity (about 2-fold higher potency) compared to the reference molecule in vitro. EXAMPLE 11 [428] This example describes the serum reactivity (serum immunogenicity) of the bispecific molecules. 11.1. Materials and Methods [429] Meso Scale Discovery (MSD) based immunoassay bridging method: Serum from forty-nine healthy donors (twenty males and twenty females) was diluted 1:30 in 1% BSA in PBS. Diluted samples were then incubated with a mixture consisting of biotin and ruthenium conjugated PDL1x41BB molecules. During this incubation the two antigen binding sites of anti-drug antibodies formed a bridge between the conjugated PDL1x41BB bispecific molecules. The sample mixture was then added to a blocked MSD streptavidin standard bind microtiter plate (MSD, # L15SA-1). The biotinylated-molecule bound to the streptavidin-coated surface of the well resulting in the immobilization of the bridged complex. The plate was washed to remove any unbound complexes and tripropylamine gold read buffer (MSD, # R92TG-2) was added to each well. Using the MSD Quickplex SQ 120, an electrical current was placed across the plate-associated electrodes. The result was a series of electrically induced oxidation-reduction reactions
involving ruthenium (from the captured complex) and tripropylamine. The subsequent luminescent signal was quantified as electrochemiluminescence (ECL) units. When unlabeled PDL1x41BB bispecific molecule was added to the conjugation mixture, the unlabeled material competed with the labeled PDL1x41BB bispecific molecules for binding, leading to a drop in ECL signal and demonstrating the specificity of the anti-drug antibody for the drug. The %ECL inhibition was calculated to represent the level of signal drop. [430] Generation of biotinylated and ruthenylated PDL1x41BB molecules. Biotin or ruthenium were covalently attached to the PDL1x41BB bispecific molecules. This process includes an initial buffer- exchange, a conjugation reaction, and a post-conjugation buffer exchange to remove free labels from the conjugated PDL1x41BB molecules. 11.2 Results [431] To evaluate the serum reactivity for the bispecific molecules, forty-nine naïve human serum donors were tested in bridging assays. FIG.18A-18B. Fig.18A shows the % ECL inhibition in the MSD bridging assay for the four PDL1x41BB molecules. Candidates 1, 2, and 4 had less than 40% of donors depleting over 40%, passing assay acceptance criteria for acceptable pre-existing reactivity. The data for Candidate #3 shown in Fig.18A was not used due to the conjugates bridging together and cross reacting in the bridging assay. Biotinylated Candidate #3 was subsequently run in the universal pre-existing reactivity assay (shown in Fig 18B) that has a universal detector. Figure 18B shows the % ECL inhibition for Candidate #3 in the HUDA (human universal indirect assay). Here, Candidate #3 had less than 40% of donors depleting over 40%, passing assay criteria for acceptable pre-existing reactivity. EXAMPLE 12 [432] This example describes PK profiles of the bispecific molecules. [433] Male human FcRn homozygous knock-in mice (B6.Cg-Fcgrttm1Dcr Tg(FCGRT)32Dcr/DcrJ) were injected intravenously with 1 mg/kg of each PDL1x41BB bispecific molecule. Serum samples were collected and quantitation of PDL1x41BB bispecific molecules was conducted using electrochemiluminescent immunoassays with either biotinylated human PDL1 or biotinylated human 41BB as the capture reagent and a ruthenylated mouse anti-human IgG Fc as the detection reagent. Non-compartmental analysis was performed on the individual serum concentration-nominal time data for each capture reagent method and presented in Table 23. Each molecule had similar PK parameters with both assays, and the PK of all molecules resembled expected characteristics for a human IgG in mouse serum. [434] Male and female cynomolgus monkeys were injected intravenously with 4.05 µM/kg of each PDL1x41BB bispecific molecule. Serum samples were collected and quantitation of PDL1x41BB
bispecific molecules was conducted using electrochemiluminescent immunoassays with biotinylated human PDL1 as the capture reagent and a ruthenylated mouse anti-human IgG Fc as the detection reagent. Non-compartmental analysis was performed on the individual serum concentration-nominal time data from 0 to 168 hours and presented in Table 24. The clearance values resembled expected characteristics for a human IgG in non-human primate serum, with roughly equivalent values for all molecules. [435] Tables 23-24 summarizes the pharmacokinetic profile in Human FcRn transgenic mice and cyno monkey Table 23. Mouse PK (huFcRn transgenic mice*) Dose 41BB CL PDL1 CL 41BB T1/2 PDL1 T1/2 (mg/kg) (ml/kg/hr) (ml/kg/hr) (Day) (Day)
Table 24. Non-human primate PK Dose CL (mL/kg/hr) 44988-3 0.8 mg/kg; 4.05 µM/kg 0.439
EXAMPLE 13 [436] This example describes DC:T assays that are used to evaluate T-cell based immunogenicity (T cell epitope) of a molecule. [437] PBMC were thawed and CD14+ cells were isolated by positive magnetic bead selection according to the manufacturer’s protocol. Isolated CD4+ monocytes were then differentiated into immature dendritic cells (iDC) for 5 days with GM-CSF and IL-4. On day 5, iDC were washed on the Curiox and treated with media (negative control), a molecule with low clinical immunogenicity (low risk benchmark) at 0.3 μM, KLH (positive control) at 50 μg/mL, or test molecules at 0.3 μM for 3 hours. Following the incubation, iDC were washed on the Curiox and matured for 2 days with TNFα and IL-1β. Following maturation, mature DC (mDC) were washed on the Curiox and then co-cultured with autologous CD4+ T cells freshly isolated from PBMC at a ratio of 1:10 (DC:T). After 5 days of co-culture, 7.5 μM EdU was added to the cultured
cells to allow for DNA incorporation into proliferating cells. On day 7 of co-culture, cells were stained for CD3, CD4, and EdU incorporation using the Click-iT kit following the manufacturer’s protocol. A stimulation index (SI) was calculated for each donor by taking the mean percent EdU+ of the CD3+ CD4+ population of (6-8) replicate wells treated with control or test molecules and dividing by the mean percent EdU+ of the CD3+ CD4+ population of (6-8) replicates treated with media alone. A SI value >2 is considered a positive response, meaning the donor demonstrated an immune response to the test or control molecule. This SI metric is a well-accepted industry standard for establishing responses. Donors that responded to all proteins tested (including controls) were considered non-antigen specific and excluded from the analysis. [438] The bispecific molecules disclosed herein could not be directly tested in the DC:T assay because high expression of PD-L1 on monocyte-derived DCs enabled the bispecific molecules to be retained on the surface of the dendritic cells throughout the maturation and washing process. This retention could potentially result in direct stimulation of T cells via 41BB, confounding assessment of sequence-based immunogenic risk. To circumvent this problem, for one bispecific molecule, 3 proteins were independently expressed: the anti-PD-L1 domain (in mAb format), the anti-41BB domain fused with Fc, and the Fc domain alone. The Fc domain alone served as a baseline for the experiment since a human IgG1 Fc domain is not expected to elicit a robust T-dependent antibody response, given the abundance of this domain in human serum. Furthermore, the Fc domain of the bispecific molecule disclosed herein has been incorporated into other molecules that have been tested in clinical studies with acceptable immunogenicity. By comparing the anti-PD-L1 mAb and the anti-41BB-Fc fusion to the Fc alone, the contribution of the anti-PD-L1 fragment antigen-binding (Fab) and the anti-41BB domain to sequence- based immunogenic risk could be assessed. A total of 30 donors were tested, and 5 were excluded due to a lack of antigen specificity. As shown in Fig.19A, 23 out of 25 (92%) donors showed a response to KLH, and 3 of 25 (12%) donors showed a response to the low-risk benchmark mAb protein. These results are within the expected response range and qualify this assay for us in screening test molecules. As shown in Figure 19B, there were no significant differences in stimulation index responses when the Fc domain was compared to the PD-L1 mAb and 41BB Fc-VH domains using the Mann-Whitney test (p=0.7111 and 0.8891, respectively). EXAMPLE 14. Identification of epitope and paratope residues of 4-1BB-binding proteins [439] This example describes structural studies that identified the epitope and paratope residues of 4- 1BB-binding proteins disclosed herein. 1. Materials and Methods [440] Purification of 4-1BB in complex with clone 380984. The sequence of clone 380984 is shown as SEQ ID NO: 372. Human soluble 4-1BB fragment is shown as residues 24-162 of SEQ ID NO: 272.
[441] Human 4-1BB was transiently expressed in Expi293FTM GnTI- with an 8xHistidine tag at C-terminus (SEQ ID NO: 566) and purified through the Ni-NTA column with imidazole gradient elution. The protein was further purified to homogeneity via size-exclusion chromatography using HiLoad 26/200 Superdex 200 (GE Healthcare). The 8xHistidine tag at C-terminus of 4-1BB was removed via proteolytic cleavage. 1.8 mg purified untagged 4-1BB was mixed with 3 mg 380984 to form a complex. The complex was obtained by removing excess 380984 using size-exclusion chromatography and followed by SDS-PAGE to confirm the purified complex. The purified 4-1BB with 380984 complex was concentrated to 9 mg/ml for crystallization setup. [442] Crystallization. Crystals of the 4-1BB with 380984 complex were obtained in the following condition: 0.1M MES pH 5.6, 1.5M Ammonium sulfate, and the sitting drop vapor diffusion technique by mixing 300 nL protein with 300 nL crystallization buffer. This condition yielded cube-like crystals that diffracted to 2.62 Å. [443] X-ray Crystallography data collection. Crystals were transiently cryo-protected using 80% crystallization buffer with 20% glycerol. The ALS staff performed the synchrotron data collection at the ALS 5.0.2 beamline. Image frames were processed using HKL2000 software (HKL Research Inc.). The crystal belongs to space group P3121 with unit cell: a=86.00 Å, b=86.00 Å, c=163.79 Å; α=β=90º, γ=120º. [444] Structural determination and refinement. The structure was solved by molecular replacement using the published 4-1BB structure (PDB:6mhr) and a homology model of the 380984 as templates. The molecule replacement solution suggested that one 4-1BB monomer and one UniDab_VH molecule were packed into the asymmetric unit with high confidence. After several rounds of iterative model building, manual adjustment, and structural refinement using Coot (Acta Cryst. D. and Phenix (Acta Cryst. D66, 213-221, 2010). The final refinement Rwork/Rfree factors are 0.225/0.255. The structure is in good geometry with RMSD of bonds of 0.008 Å and RMSD angles of 1.011º; the Ramachandran plot shows 95.80% residues are in favorable backbone dihedral angles, and 4.20% residues are allowed. 2. Epitope and Paratope analysis [445] In the crystal structure, the asymmetric unit contains one 4-1BB monomer and one 380984. The overall crystal structure of human 4-1BB in complex with 380984 is shown in FIG. The human 4-1BB folds into a V-shape molecule with four CRDs in a linear arrangement and a significant bend within CRD3. The epitopes-paratopes interface partially overlaps with the 4-1BBL ligand binding residues (FIG.20A). The structure was visualized using ChimeraX (UCSF Resource for Biocomputing, Visualization, and Informatics, supported in part by the National Institutes of Health). [446] Close-up analysis revealed that the 380984 binds to 4-1BB CRD2 and 3 domains primarily through its CDR-3 (FIG.20B). The contact consists of 4-1BB residues K69, V71, T89, P90, F92, M101, C102, and
Q104, which make hydrogen bonds and Van der Waals force interactions with the 380984 residues L45, W47, S103, Y108, T110, S111, and F112. Most of the hydrogen bonds are through the main chain/backbone. The numbering is based on SEQ ID NO.272 for 4-1BB and SEQ ID NO.372 for 380984. [447] Table 25A shows the 380984 residues which make contact with 4-1BB. As shown in the table, Kabat CDR-3 of the VH residues, S103, Y108, T110, S111, F112, and Y114 contact 4-1BB residues K69, V71, Q104, M101, and L112; Kabat framework residue L45 and W47 contact 4-1BB residues P90 and Q104. Table 25A. Summary of 4-1BB epitope residues interacting with 380984 paratope residues 4-1BB atoms 380984 atoms Distance (Å) (residue number based on SEQ (residue number based on SEQ
[448] We further characterized the importance of the epitope and paratope residues identified in Tables 25B and 25C. For example, when contacts are primarily with the backbone, or when the contacts are energetically neutral, the residue may tolerate a wide range of mutations. Significant contacts of an entire side chain with the other binding partner, or high conservation in a computational evolution simulation, are evidence that a particular residue may have increased importance relative to other residues. While the
majority of the paratope residues are in CDR-H3, Leu45 and Trp47 are characterize as framework residues under Kabat, AbM, Chothia, and IMGT definitions. Contact definition characterizes Trp47 as part of CDR-H2. Table 25B. Ranking of the 4-1BB residue importance in the binding interface 4-1BB Importance Notes residues # CYS102 Primar H-bondin to 380984 (backbone) VDW with 380984 (side chain); hi h
Table 25C. Ranking of the 380984 residues importance in the binding interface 380984 Position Importance Notes residues # according
[449] Additionally, we analyzed whether certain epitope and paratope residues can tolerate or not tolerate certain mutations based on structural data, and these analyses are summarized as Tables 25D and 25E. [450] Ligand 4-1BBL induced receptor 4-1BB clustering is prerequisite for the downstream signaling. A published 4-1BB-4-1BBL complex structure has shown that three receptor 4-1BB parallelly bind to a bell- shaped trimer ligand 4-1BBL(FIG.3A). The center axis of the receptor-ligand complex should be vertically
oriented towards the cell membrane, like other TNF receptor-ligand complexes. In our structure, the 380984 binds along the side of 4-1BB at CRDs 2 and 3, with its N-terminus, where Fc of the antibody connects, pointing away from the cell membrane (FIG.3B). Thus, the bulk of the antibody is also away from the receptor and the cell membrane, which increase the capability of the FcγR engagement and clustering. As a comparison, when 4-1BB complexed with the other two other clinical agonist antibodies, utomilumab (PF-05082566, PDB 6MI2) and urelumab (BMS663513, PDB 6MHR), the Fabs are orientated parallel to the membrane, where engagement of FcγR may be more restricted (FIG.3C and D). Table 25D. Point mutations of 4-1BB epitope residues that could potentially abrogate the 380984 recognition Mutations Descriptions CYS102 to ALA Disruption of a highly conserved intra-disulfide linkage and the l ili f RD i f 41BB i ll k
380984 Position Acceptable substitutions Preferred substitutions residues # according
3. Relative orientation of the antibody. [451] Ligand 4-1BBL induced receptor 4-1BB clustering is prerequisite for the downstream signaling. A published 4-1BB-4-1BBL complex structure has shown that three receptor 4-1BB parallelly bind to a bell- shaped trimer ligand 4-1BBL (FIGs.21A and 21B). The center axis of the receptor-ligand complex should be vertically oriented towards the cell membrane, like other TNF receptor-ligand complexes. In our structure, the 380984 binds along the side of 4-1BB at CRDs 2 and 3, with its N-terminus, where Fc of the molecule connects, pointing away from the cell membrane (FIG.21E). Thus, the bulk of the antibody is also away from the receptor and the cell membrane, which increase the capability of the FcγR engagement and clustering. As a comparison, when 4-1BB complexed with the other two other clinical agonist antibodies, utomilumab (PF-05082566, PDB 6MI2) and urelumab (BMS663513, PDB 6MHR), the
Fabs are orientated parallel to the membrane, where engagement of FcγR may be more restricted (FIGs. 21C and 21F). Table 25F.380984 CDRs Region Definition Sequence Fragment SEQ ID CDR-H1 Chothia GFTFGDY--- 566 FTFDYAM
SEQUENCE TALBES Table A: Round 1 PD-L1 binder CDR sequences clone ID CDR1 ID CDR2 ID CDR3 21B2 H 1 GYYIH 2 WINPNRGVTSSAQKFQG 3 DSTAVGFDY Y Y
clone ID CDR1 ID CDR2 ID CDR3 6C7 H 43 RYYWS 44 YIDDSGNTNYNPSLKS 45 GVAAGRIDP P P G
clone ID HC V-region (VH) ID LC V-region (VL) 1B2 133 QVQLVQSGAEVKKPGASVKVSCKASGYT 134 AIQMTQSPSSLSASVGDRVTITCRASQ FTGYYIHWVRQAPGQGLEWMGWINPNRG GISNYLAWFQQKPGKAPKSLIYAASSL E Q Y S K Q L E Q L E Q Y S K Q L E Q L E R Q L E
Clone HC V-region LC V-region I P D L S Q S
VTAADSAVYYCARTDYYYFEMDVWGQGT EDEADYYCEAWDDSLNGPVFGGGTKLT TVTVSS VL Q L E L P D Q L E L P D L Q L E Q L E S Q S T I P D L Q S S K Q L E
17H1. 169 EVQLVESGGGLVQPGGSLTLSCAASGFT 170 DIQMTQSPSSLSASVGDRVTITCRASQ 009 FSGYSMNWVRQAPGKGLEWVSYIGISTS SISNYLNWYQQQPGKAPKFLIYAASTL AIYYAESVKGRFTISRDNAKNSLFLQMN QSGVPSRFSGSGSGTDFTLTISSLQPE L P D Q Y S K L P D s
clone ID HC ID LC 21B2 QVQLVQSGAEVKKPGASVKVSCKASG AIQMTQSPSSLSASVGDRVTITCRASQ L E R L S H Q Y S K A G A R
KAKGQPREPQVYTLPPSREEMTKNQV SLTCLVKGFYPSDIAVEWESNGQPEN NYKTTPPVLDSDGSFFLYSKLTVDKS Q L E R L S H Q L E R L S H Q Y S K A G A R
NYKTTPPVLDSDGSFFLYSKLTVDKS RWQQGNVFSCSVMHEALHNHYTQKSL SLSPGK Q L E R L S H Q L E R L S H
s clone HC LC NI P D L V E S
NKALPAPIEKTISKAKGQPREPQVYTL PPSREEMTKNQVSLTCLVKGFYPSDIA VEWESNGQPENNYKTTPPVLDSDGSFF S Q S T T G W S Q L E R L S H L P D Q L T R
15A12 EVQLVESGGGLVQPGGSLRLSCAASGF DIQMTQSPSSLSASVGDRVTITCRASQ .012 TFSSYSMNWVRQAPGKGLEWVSYISSS SISTYLNWYQQKPGKAPKLLIYAASSL GSTIYYAESVKGRFTISRDNAKNSLYL QSGVPSRFSGSESGTDFTLTISSLQPE T L V K L P D L V E S Q L E R L S H S Q S T T
TSGGTAALGCLVKDYFPEPVTVSWNSG LVCLISDFYPGAVTVAWKADSSPVKAG ALTSGVHTFPAVLQSSGLYSLSSVVTV VETTTPSKQSNNKYAASSYLSLTPEQW PSSSLGTQTYICNVNHKPSNTKVDKKV KSHRSYSCQVTHEGSTVEKTVAPTECS I P D L V E S Q S S K A G A R Q L E R L S H
TCVVVDVSHEDPEVKFNWYVDGVEVHN AKTKPREEQYNSTYRVVSVLTVLHQDW LNGKEYKCKVSNKALPAPIEKTISKAK Q L E R L S H L P D Q L T R Q Y S K A G A R
SFFLYSKLTVDKSRWQQGNVFSCSVMH EALHNHYTQKSLSLSPGK L P D Q L T R
SEQ ID Name Sequence 221 Linker GGS Q E Q E A C K S D C
(Kappa) with charge pair Human Light chain QPKAAPSVTLFPPSSEELQANKATLVCLISDFYPGAVTVAWKA C S N C S Q N H S N K T I S Q S N K T I S Q S N K T I S Q S N K T I S Q S N K T I S
DIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQ QGNVFSCSVMHEALHNHYTQKSLSLSPG Human IGG1 HC ASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNS N K T I S Q C S Q N H C S Q N H C S Q N H C S Q N H C S Q N H C S Q N H S N K T I S Q S N
with SELF2 LALA HKPSNTKVDKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPK mutation and PKDTLYITREPEVTCVVVDVSHEDPEVKFNWYVDGVEVHNAKT terminal G KPCEEQYGSTYRCVSVLTVLHQDWLNGKEYKCKVSNKALPAPI S Q S N K T I S Q S N S N K T I S Q S D V G T L S D D R T A L S N C V G T L
Human IgG3 HC C- ASTKGPSVFP LAPCSRSTSG GTAALGCLVK DYFPEPVTVS domain WNSGALTSGV HTFPAVLQSS GLYSLSSVVT VPSSSLGTQT YTCNVNHKPS NTKVDKRVEL KTPLGDTTHT CPRCPEPKSC P D H T N E S D Q K K L S T F G C N D L N S C A L E g g g g t t a a a a g g a t g a g
Human 4-1BB LQDPCSNCPAGTFCDNNRNQICSPCPPNSFSSAGGQRTCDICR extrascapular QCKGVFRTRKECSSTSNAECDCTPGFHCLGAGCSMCEQDCKQG domain (residues QELTKKGCKDCCFGTFNDQKRGICRPWTNCSLDGKSVLVNGTK Q C K A S G G G G T T A A A A G G A T C C T C S G Y N H K C T T A A C G G C A T A A A G C C
ACTTGGTAAT TCTGGGAGCC ATCTTATTAT GCCTTGGTGT AGCACTGACA TTCATCTTCC GTTTAAGAAA AGGGAGAATG ATGGATGTGA AAAAATGTGG CATCCAAGAT ACAAACTCAA C N G Y N H K c a t t a a g a a t g a c a g t g g t a a a Q C K V R F g g g g t t a a a a g g a t g
ggcagaaagaaactcctgtatatattcaaacaaccatttatga gaccagtacaaactactcaagaggaagatggctgtagctgccg atttccagaagaagaagaaggaggatgtgaactggccagcttt t S S
u qu p u (nucleic acid sequences all encoding signal peptide that is clipped in mature proteins) SEQ Description sequence ID G L E K E Q S W G V G G T P D S Q R L V S G C C C C C T
GGGGGAGTTATTATACTACTACGGTATGGACGTCTGGGGCCAAGGGACCACGG TCACCGTGTCCTCAGCCTCCACCAAGGGCCCATCGGTCTTCCCCCTGGCACCC TCCTCCAAGAGCACCTCTGGGGGCACAGCGGCCCTGGGCTGCCTGGTCAAGGA G C A T G C G A C A G A G T A G C T T G T G T C A C A C T A C G A G G G T T G C G A C G Y D W V
DKKVEPKSCDKTHTCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVV DVSHEDPEVKFNWYVDGVEVHNAKTKPCEEQYGSTYRCVSVLTVLHQDWLNGK EYKCKVSNKALPAPIEKTISKAKGQPREPQVYTLPPSREEMTKNQVSLTCLVK F S T G L C R L V S H K V P G C T G G T A T T T G C G A C C T G G C G C A C A A A C A C T T T C C A
AGTCAGACCATTAGCAGCTATTTAAATTGGTATCAGCAGAAACCAGGGAAAGC CCCTAAACTCCTGATCTATGCTATCTCCAATTTGCAAAGTGGGGTCCCATCAA GGTTCAGTGGCAGTGGATCTGGGACAGATTTCACTCTCACCATCAGCAGTCTG T G A A A A T C C C C T A C P D W V V K K F T S G G I T G S L S I S G G C C G A C C T T T
GACGGTGTCGTGGAACTCAGGCGCCCTGACCAGCGGCGTGCACACCTTCCCGG CTGTCCTACAGTCCTCAGGACTCTACTCCCTCAGCAGCGTGGTGACCGTGCCC TCCAGCAGCTTGGGCACCCAGACCTACATCTGCAACGTGAATCACAAGCCCAG A C C T G G C G C A C A A A C A T C G T C A C A C T A C G T T A T A C C T A T A G N D W V V K K
GFYPSDIAVEWESNGQPENNYKTTPPVLDSDGSFFLYSKLTVDKSRWQQGNVF SCSVMHEALHNHYTQKSLSLSPGGGGSGGGGSQVQLQESGPGLVKPSETLSLT CTVSGGSISSYYWSWIRQPAGKCLEWIGRIYTSGSTNYNPSLKSRVTMSVDTS G G I T G S L I G L K G C T G G C A T T T G C G A C C T G G C G C A C A A A C A T C G T C A C A C T
CTCACCATCACCAGTCTGCAACCTGAAGATTTTGCAACTTACTATTGTCAACA GAGATTTAGTATCCCATTTACTTTCGGCTGCGGGACCAAAGTGGATGTCAAAC GA G A T C T G C G T T A C C G C C G A C C T T T G C G A C C T G G C G C A C A A A C A T C G G G G C A T
TGGGGACCCAGACGTACATTTGTAACGTGAATCACAAACCAAGCAATACTAAG GTAGATAAGAAAGTAGAACCGAAGAGCTGC P D W V V K K F T S F L Y D W V S I S G H K V P G C C G A C C T T T G C G A C C T G G C G C A C
GGCTCCTTCTTCCTCTATAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCA GGGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCACTACA CGCAGAAGAGCCTCTCCCTGTCTCCGGGTGGTGGCGGATCGGGAGGTGGCGGA C A T C G G G G C A T G G T T A T A C C T A T A G G A A A A T C C C C T A C P D W V V K K F
SCSVMHEALHNHYTQKSLSLSPGGGGSGGGGSEEQLSESGGGLVQPGGSLRLS CAASGFTFSSYAMSWVRQAPGKGLEWVSAISGSGGSTYYADSVKGRFTISRDK SKNTLYLQMNSLRAEDTAVYYCAKDGGFGELLYYYGMDVWGQGTTVTVSSAST V G L E K S I S G R L V S G C C G A C C T T T G C G A C C T G G C G C A C A A A C A T T G A C T T C
ATTCCCTGCGGTATTGCAGTCCTCCGGACTCTACTCCCTGGAGTCGGTGGTAA CGGTGCCCAGCTCCAGCTTGGGGACCCAGACGTACATTTGTAACGTGAATCAC AAACCAAGCAATACTAAGGTAGATAAGAAAGTAGAACCGAAGAGCTGC G T T A T A C C T A T A G G A G G G T T G C G A C G P D W V V K K F S K T V G L E K N I S
QESVTEQDSKDSTYSLESTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRG EC 11262 SFELTQPPSVSVSPGQTARITCSGDKLGDNYASWYQQKPGQSPVLVIYQDSRR L V S G C T G A C C T T T G C G A C C T G G C G C A C A A A C A T T G A C T T C A C G T C A T A C C T
AACTTCTATCCCAGAGAGGCCAAAGTACAGTGGAAGGTGGATAACGCCCTCCA ATCGGGTAACTCCCAGGAGAGTGTCACAGAGCAGGACAGCAAGGACAGCACCT ACAGCCTCGAAAGCACCCTGACGCTGAGCAAAGCAGACTACGAGAAACACAAA G G A G G G T T G C G A C G P D W V V K K F T S P Y P D W V N I S G R T A E G C T G A
GAGTCACCATGACCAGGGACACCTCCATCAGTACAGCCTACATGGAGCTGAGC AGCCTGAGATCTGAGGACACGGCCGTGTATTTCTGTGCGAGAGATTTTTTCAC TGGTTATTTTGACTACTGGGGCCAGGGAACCCTGGTCACCGTGTCCTCAGCCT T T G C G A C C T G G C G C A C A A A C A T C G C C G G C G A G T C A T A C C T A T A G G A G A G T
CAGGTGTGGGATAGTAGTAGTGATCATGTGGTATTCGGCGGAGGGACCAAGCT GACCGTCCTAGGTCAGCCCAAGGCTGCACCCTCGGTCACTCTGTTCCCGCCCT CCTCTGAGGAGCTTCAAGCCAACAAGGCCACACTGGTGTGTCTCATCAGTGAC A G C C P D W V V K K F T S F L Y D W V N I S G H K V P G C T G A C C T T T G C G A C C T
ACGTGGACGGCGTGGAGGTGCATAATGCCAAGACAAAGCCGTGCGAGGAGCAG TACGGCAGCACGTACCGTTGCGTCAGCGTCCTCACCGTCCTGCACCAGGACTG GCTGAATGGCAAGGAGTACAAGTGCAAGGTGTCCAACAAAGCCCTCCCAGCCC G C A C A A A C A T C G G G G C A T G G T C A T A C C T A T A G G A A A A T C C C C T A C
y
clone ID CDR1 ID CDR2 ID CDR3 386340 VH 326 SFAMT 327 AISGSGGSTYYAGSVKG 328 EAYESSGYYTTSFDY Y Y
Clone ID VH EVQLLESGGGLVQPGGSLRLSCAASGFTFSSFAMTWLRQAPGKGLEWVSAISGSGGSTYYAGSV AA SE AV SV SV SV SV SV SM SV
380969 382 EVQLVESGGGLVNPGRSLRLSCTASGFTFGDYAMSWFRQAPGKGLEWVGFIRSKTYGGTTEYAA SVKGRFTISRDDSKSIAYLQMNSLKTEDTAVYYCAREGLDSGISYATSFDYRGQGTLVTVSS SV SV SV
Name Seq Seq Seq No. CDR1 No. CDR2 No. CDR3 56039 4-1BB V V DP DY
Name Seq CDR1 Seq Seq No No CDR2 No CDR3
56639, 44988, 56041, 56042, 56132, 56761, 19 GYYIH 20 WINPNRGVTSSAQKFQG 21 DATAVGFDY
Seq No. Name Sequence 56639 56041 V V G V V G G V V
Seq No. Name Sequence D S S D S S S S F E
11 56042 VL DIQMTQSPSSLSASVGDRVTITCRASQSISNYLNWYQQQPGKAPKFLIYAASTLQSGVPSRF SGSGSGTDFTLTISSLQPEDFATYYCQQSYSTPLTFGGGTKVEIKR 44988 VH VLESGPGLVKPSETLSLTCTVSGGSISSYYWSWIRPAGKCLEWIGRIYTSGSTNYNPS S F E S D
Seq Name Sequence No. SS GG YG EE RW DS VD AK KS DS SS GG YG EE RW SS GG YG EE RW
19 56042 light TVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDS chain #1 KDSTYSLESTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC SS GG YG KE RW DS DS SS VD AK DS KQ SS VD AK DS DS SS GG YG EE RW
Seq No. Name Sequence SS AS GL PS ST MT
KNQVSLTCLVKGFYPSDIAVEWESNGQPENNYDTTPPVLDSDGSFFLYSDLTVDKSRWQQ GNVFSCSVMHEALHNHYTQDSLSLSPGK DIQMTQSPSSLSASVGDRVTITCRASQGISNYLAWFQQKPGKAPKSLIYAASSLQSGVPS PP LT YY TL DP AP NY PS PP LT SS AS GL GG GR GG VD AK DS ER PP SL YY TV AV VQ QK GS YA GG YV KA LD GF GG LL IK SS AS GL PS ST MT QQ PS PP
SDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLESTLT LSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC EVQLVESGGGLVQPGGSLTLSCAASGFTFSGYSMNWVRQAPGKGLEWVSYIGISTSAIYY AS GL PS ST MT QQ PS PP LT PS PP LT SS AS GL PS ST MT QQ TA LQ PS PP LT SS AS GL PS ST MT QQ TV SS MT GS YN LV PG CG SS AS GL PS ST MT QQ
DIQMTQSPSSLSASVGDRVTITCRASQGISNYLAWFQQKPGKAPKSLIYAASSLQSGVPS 48 56639 light RFSGSGSGTDFTLTISSLQPEDFATYYCQQYNSYPFTFGQGTKVDIKRTVAAPSVFIFPP chain SDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGNSQESVTEQDSKDSTYSLSSTLT YY VT EV IE KT ER PP SL YY TV AV VQ QK GS YA GG YV KA LD PS PP LT SS AS GL GP TI GS FA GG YV KA LD
s Seq No. Name Sequence TC CC CT AC CT CC CA AC TC TC
TGCAACGTGAATCACAAGCCCAGCAACACCAAGGTGGACAAGAAAGTTGAGCCCAAATCT TGTGACAAAACTCACACATGCCCACCGTGCCCAGCACCTGAACTCCTGGGGGGACCGTCA GTCTTCCTCTTCCCCCCAAAACCCAAGGACACCCTCTACATCACCCGGGAACCTGAGGTC TG CG AC CC CC TG AC AG AC CC CA CA CT AA CA AT AG CG GC TC CT AC AT CC TG CT TG CT CC AG CC TG GC AC CC CT CC CA CA CT AA CA AT AG CG GC TC CC CT AC
ATGGAGCTGAGCAGTCTGAGATCTGACGACACGGCCGTGTATTACTGTGCGAGAGATGCT ACAGCAGTAGGCTTTGACTACTGGGGCCAGGGAACCCTGGTCACCGTGTCCTCAGCCTCC ACCAAGGGCCCATCGGTCTTCCCCCTGGCACCCTCCTCCAAGAGCACCTCTGGGGGCACA AC TC TC CT GA TT GT GA AA GG GA TC CA AC AC AG AA AG AG CC GG GC TC GC GA TG GG CC AC AA TG GC TC CT AC AT GT TC AG CG TC TG AG AG AA AA AG TG GT CT CC GC
AGCTATTTAAATTGGTATCAGCAGAAACCAGGGAAAGCCCCTAAACTCCTGATCTATGCT ATCTCCAATTTGCAAAGTGGGGTCCCATCAAGGTTCAGTGGCAGTGGATCTGGGACAGAT TTCACTCTCACCATCAGCAGTCTGCAACCTGAAGATTTTGCAACTTACTACTGTCAACAG GT CA TC TG CG AC CC CC TG AC AG AG TC CC CT AC CT CC CA AC TC TC CT CA TC TG CG AC CC CC TG AC AG AC CC CA CA CT AA CA AT AG CG GC TC CT AC TT AT CC CA
GCCGCTCTGGGCTGTCTGGTGAAGGACTACTTTCCTGAGCCCGTCACCGTGTCCTGGAAC TCCGGCGCCCTGACAAGCGGCGTGCATACCTTCCCCGCCGTGCTCCAAAGCTCCGGACTG TACAGCCTGGAGAGCGTGGTGACCGTCCCCTCCAGCAGCCTGGGAACCCAGACCTACATC GC CC TG TT CA AC CC CC TG AG AG AA CC CA CA CT GC CA AT AG CG GC CC CA CA CT AA CA AT AG CG GC TC CC CT AC CT CC CA AC TC TC CT CA TC TG CG AC CC CC TG
GAGTGGGAGAGCAATGGGCAGCCGGAGAACAACTACAAGACCACGCCTCCCGTGCTGGAC TCCGACGGCTCCTTCTTCCTCTATAGCAAGCTCACCGTGGACAAGAGCAGGTGGCAGCAG GGGAACGTCTTCTCATGCTCCGTGATGCATGAGGCTCTGCACAACCACTACACGCAGAAG TG CT GG CG AA CG CC CC CA CA CT AA CA AT AG CG GC TC CC CT AC CT CC CA AC TC TC CT CA TC TG CG AC CC CC TG AC AG AG TG TC GC AG CT CT CA CC CA AG GT CT
TACTATTGTCAACAGAGATTTAGTATCCCATTTACTTTCGGCTGCGGGACCAAAGTGGAT GTCAAATAG CAGGTGCAGCTGGTGCAGTCTGGGGCTGAGGTGAAGAAGCCTGGGGCCTCAGTGAAAGTC CC CT AC CT CC CA AC TC TC CT CA TC TG CG AC CC CC TG AC AG AC CC CA CA CT AA CA AT AG CG GC TC CT AT AT AC CC AG TC TC AG GG AG CC AC CC AC AC TC GC GA
TTCTCTGGCTCCAACTCTGGGAACACAGCCACTCTGACCATCAGCGGGACCCAGGCTATG GATGAGGCTGACTATTACTGTCAGACGTGGCACAGCAGCGCTACCGTGTTCGGCGGAGGG ACCAAGCTGACCGTCCTAGGTCAGCCCAAGGCTGCACCCTCGGTCACTCTGTTCCCGCCC AC AA TG GC TC CT AC AT GT TC AG CG TC TG AG AG AA AA AG TG GT CT CC GC CT AT AG GT CA TC TG CG AC CC CC TG AC AG AG CC CA CA CT AA CA AT AG CG GC TC CC
CCTGGACAAGGGCTTGAGTGGATGGGATGGATCAACCCTAACAGAGGTGTCACAAGCTCT GCACAGAAGTTTCAGGGCAGGGTCACCATGACCAGGGACACGTCCGTCAGCACAGCCTAC ATGGAGCTGAGCAGTCTGAGATCTGACGACACGGCCGTGTATTACTGTGCGAGAGATGCT CC CA AC TC TC CT CA TC GG TC AC AC CT CC AG CT AT AG GT CA TC TG CG AC CC CC TG AC AG AG
Germline name H_CDR1 H_CDR2 H_CDR3 Y V V V V H
SEQ ID NO 521 522 523 VH3|3-23/D3|3-10|RF1/JH6 SYAMS AISGSGGSTYYADSVKG VLLWFGELL*YYYYYGMDV V P V P
K_CDR1 K_CDR2 K_CDR3 VK2|A2/JK1 KSSQSLLHSDGKTYLY EVSNRFS MQSIQLPWT
L_CDR1 L_CDR2 L_CDR3
[452] All references, including publications, patent applications, and patents, cited herein are hereby incorporated by reference to the same extent as if each reference were individually and specifically indicated to be incorporated by reference and were set forth in its entirety herein. [453] The use of the terms “a” and “an” and “the” and similar referents in the context of describing the disclosure (especially in the context of the following claims) are to be construed to cover both the singular and the plural, unless otherwise indicated herein or clearly contradicted by context. The terms “comprising,” “having,” “including,” and “containing” are to be construed as open-ended terms (i.e., meaning “including, but not limited to,”) unless otherwise noted. [454] Recitation of ranges of values herein are merely intended to serve as a shorthand method of referring individually to each separate value falling within the range and each endpoint, unless otherwise indicated herein, and each separate value and endpoint is incorporated into the specification as if it were individually recited herein. [455] All methods described herein can be performed in any suitable order unless otherwise indicated herein or otherwise clearly contradicted by context. The use of any and all examples, or exemplary language (e.g., “such as”) provided herein, is intended merely to better illuminate the disclosure and does not pose a limitation on the scope of the disclosure unless otherwise claimed. No language in the specification should be construed as indicating any non-claimed element as essential to the practice of the disclosure. [456] Preferred embodiments of this disclosure are described herein, including the best mode known to the inventors for carrying out the disclosure. Variations of those preferred embodiments may become apparent to those of ordinary skill in the art upon reading the foregoing description. The inventors expect skilled artisans to employ such variations as appropriate, and the inventors intend for the disclosure to be practiced otherwise than as specifically described herein. Accordingly, this disclosure includes all modifications and equivalents of the subject matter recited in the claims appended hereto as permitted by applicable law. Moreover, any combination of the above-described elements in all possible variations thereof is encompassed by the disclosure unless otherwise indicated herein or otherwise clearly contradicted by context.
Claims
WHAT IS CLAIMED: 1. A 4-1BB antigen-binding protein comprising a heavy chain variable domain (VH) and a light chain variable domain (VL), wherein (i) said VH comprises the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO:147, 149, 151, 153, 155, 157, 159, 324, 161, 163, 165, 167, 169, 171, 173, 175, 404, 406, 408, or 410; and (ii) said VL comprises the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 148, 150, 152, 154, 156, 158, 160, 325, 162, 164, 166, 168, 170, 172, 174, 176, 405, 407, 409, or 411. 2. The 4-1BB antigen-binding protein of claim 1, comprising: (1) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.43-48, respectively; (2) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.49-54, respectively; (3) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.55-60, respectively; (4) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.61-66, respectively; (5) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.67-72, respectively; (6) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.73-78, respectively; (7) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.79-84, respectively; (8) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.85-90, respectively; (9) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.91-96, respectively; (10) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.97-102, respectively; (11) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.103-108, respectively; (12) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.109-114, respectively; (13) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.115-120, respectively; (14) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.121-126, respectively;
(15) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.127-132, respectively; (16) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.386-391, respectively; or (17) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.392-397, respectively. 3. The 4-1BB antigen-binding protein of claim 1 or 2, comprising a VL framework sequence and a VH framework sequence, and (i) wherein the VL framework sequence and/or the VH framework sequence is at least 90% identical to a human VL germline framework sequence or VH germline sequence, respectively, (ii) wherein said human germline VL framework sequence is the framework sequence of: DPK9, DPK12, DPK18, DPK24, HK102_V1 , DPK1 , DPK8, DPK3, DPK21 , Vg_38K, DPK22, DPK15, DPL16, DPL8, V1-22, Vλ consensus, Vλ1 consensus, Vλ3 consensus, Vκ consensus, Vκ1 consensus, Vκ2 consensus, or Vκ3 consensus, and (iii) wherein said human germline VH framework sequence is the framework sequence of: DP54, DP47, DP50, DP31 , DP46, DP71 , DP75, DP10, DP7, DP49, DP51 , DP38, DP79, DP78, DP73, VH3 consensus, VH5 consensus, VH1 consensus, VH2 consensus, or VH4 consensus. 4. The 4-1BB antigen-binding protein of any one of claims 1-3, comprising: (a) a heavy chain variable region (VH) that is at least 90% identical to SEQ ID NO: 147, 149, 151, 153, 155, 157, 159, 324, 161, 163, 165, 167, 169, 171, 173, 175, 404, 406, 408, or 410; (b) a light chain variable region (VL) that is at least 90% identical to SEQ ID NO: 148, 150, 152, 154, 156, 158, 160, 325, 162, 164, 166, 168, 170, 172, 174, 176, 405, 407, 409, or 411. 5. The 4-1BB antigen-binding protein of any one of claims 1-4, comprising: (1) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.147, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.148; (2) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.149, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.150; (3) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.151, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.152; (4) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.153, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.154; (5) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.155, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.156; (6) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.157, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.158;
(7) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.159 or 324, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.160 or 325; (8) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.161, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.162; (9) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.163, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.164 (or 336); (10) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.165, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.166; (11) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.167, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.168; (12) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.169, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.170; (13) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.171, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.172 (or 337); (14) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.173, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.174; (15) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.175, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.176; (15) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.404, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.405; (16) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.406, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.407; (17) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.408, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.409; or (15) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.410, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.411. 6. A 4-1BB antigen-binding protein that comprises a heavy chain variable domain (VH) and does not comprise a light chain variable domain (VL), wherein said protein comprises the CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, or 385. 7. The 4-1BB antigen-binding protein of claim 6, comprising: (1) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.326-328, respectively; (2) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.329-331, respectively; (3) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.332-334, respectively; (4) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.335-337, respectively; (5) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.338-340, respectively; (6) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.341-343, respectively;
(7) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.344-346, respectively; (8) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.347-349, respectively; (9) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.350-352, respectively; (10) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.353-355, respectively; (11) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.356-358, respectively; (12) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.359-361, respectively; (13) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.362-364, respectively; (14) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.365-367, respectively; or (15) a CDR-H1, a CDR-H2, and a CDR-H3 comprising SEQ ID NOs.368-370, respectively; 8. A 4-1BB antigen-binding protein that comprises a heavy chain variable domain (VH) and does not comprise a light chain variable domain (VL), wherein said protein binds to an epitope that comprises residues C102, V71, and Q104, according to the numbering of SEQ ID NO:272. 9. The 4-1BB antigen-binding protein of claim 8, wherein said epitope further comprises residue P90, according to the numbering of SEQ ID NO:272. 10. The 4-1BB antigen-binding protein of claims 8 or 9, wherein said epitope further comprises one or more residues selected from the group consisting of: K69, T89, F92, M101, and L112, according to the numbering of SEQ ID NO:272. 11. The 4-1BB antigen-binding protein of any one of claims 6-10, comprising a VH framework sequence, and (i) wherein said VH framework sequence is at least 90% identical to a human germline framework sequence, and (ii) wherein said human germline VH framework sequence is the framework sequence of: DP54, DP47, DP50, DP31 , DP46, DP71 , DP75, DP10, DP7, DP49, DP51 , DP38, DP79, DP78, DP73, VH3 consensus, VH5 consensus, VH1 consensus, VH2 consensus, or VH4 consensus. 12. The 4-1BB antigen-binding protein of any one of claims 6-11, comprising: a heavy chain variable region (VH) that is at least 90% identical to any one of SEQ ID NOs.371, 372, 373, 374, 375, 376, 377, 378, 379, 380, 381, 382, 383, 384, and 385. 13. The 4-1BB antigen-binding protein of any one of 6-12, comprising: (a) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.371; (b) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.372; (c) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.373; (d) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.374; (e) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.375; (f) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.376; (g) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.377;
(h) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.378; (i) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.379; (j) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.380; (k) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.381; (l) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.382; (m) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.383; (n) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.384; or (o) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.385. 14. The 4-1BB antigen-binding protein of any one of claims 1-13, wherein said 4-1BB antigen binding protein is a crosslinking dependent agonist of 4-1BB. 15. A PD-L1 antigen-binding protein, comprising (i) the heavy chain CDR-H1, CDR-H2, and CDR-H3 of SEQ ID NO: 133, 135, 137, 139, 141, 143, 322, 145, 398, 400, or 402; and (ii) the light chain CDR-L1, CDR-L2, and CDR-L3 of SEQ ID NO: 134, 136, 138, 140, 142, 144, 323, 146, 399, 401, or 403. 16. The PD-L1 antigen-binding protein of claim 15, comprising: (1) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.1-6, respectively; (2) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.7-12, respectively; (3) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.13-18, respectively; (4) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.19-24, respectively; (5) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.25-30, respectively; (6) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.31-36, respectively; or (7) a CDR-H1, a CDR-H2, a CDR-H3, a CDR-L1, a CDR-L2, and a CDR-L3 comprising SEQ ID NOs.37-42, respectively. 17. The PD-L1 antigen-binding protein of claim 15 or 16, comprising a VL framework sequence and a VH framework sequence, and (i) wherein the VL framework sequence and/or the VH framework sequence is at least 90% identical to a human VL germline framework sequence or VH germline framework sequence, respectively, (ii) wherein said human germline VL framework sequence is the framework sequence of: DPK9, DPK12, DPK18, DPK24, HK102_V1 , DPK1 , DPK8, DPK3, DPK21 , Vg_38K, DPK22, DPK15, DPL16, DPL8, V1-22, Vλ consensus, Vλ1 consensus, Vλ3 consensus, Vκ consensus, Vκ1 consensus, Vκ2 consensus, or Vκ3 consensus, and (iii) wherein said human germline
VH framework sequence is the framework sequence of: DP54, DP47, DP50, DP31 , DP46, DP71 , DP75, DP10, DP7, DP49, DP51 , DP38, DP79, DP78, DP73, VH3 consensus, VH5 consensus, VH1 consensus, VH2 consensus, or VH4 consensus. 18. The PD-L1 antigen-binding protein of any one of claims 15-17, comprising: (1) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.133, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.134; (2) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.135, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.136; (3) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.137, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.138; (4) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.139, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.140; (5) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.141, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.142; (6) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.143, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.144; (7) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.322, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.323; (8) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.145, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.146; (9) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.398, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.399; (10) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.400, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.401; or (11) a VH comprising an amino acid sequence at least 90% identical to SEQ ID NO.402, and a VL comprising an amino acid sequence at least 90% identical to SEQ ID NO.403. 19. The 4-1BB binding protein of any one of claims 1-14, or the PD-L1 binding protein of any one of claims 15-18, further comprising a CH1 domain, a CH2 domain, a CH3 domain, a CL domain, or any combination thereof. 20. A bispecific molecule, comprising a 4-1BB binding protein of any one of claims 1-14 and a PD-L1 binding protein of any one of claims 15-18. 21. The bispecific molecule of claim 20, wherein the bispecific molecule is one of the following formats: Fab-scFv(M2)-Fc, Fab-VH(M2)-Fc, IgG-scFv(C1), Fab-scFv(M1)-Fc, IgG-scFv(C2), IgG-VH(C2), [Fab*scFv]hetero-Fc, [Fab*VH]hetero-Fc, hetero-IgG, scFv(N1)-IgG, ([scFv*Fab]hetero-Fc)-Fc, and IgG- Fab.
2. The bispecific molecule of claim 20 or 21, comprising: (1) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 283, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 220; (2) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 287, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 206; (3) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 290, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 184; (4) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 294, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 186; (5) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 297, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 300; (6) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 304, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 307; (7) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 311, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 307; (8) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO:312, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 317; (9) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 312, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 300; (10) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 443, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 444; (11) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 434, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 435; (12) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 449, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 450; (13) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 451, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 452; (14) (i) one copy of an amino acid sequence that is at least 90% identical to SEQ ID NO: 449, (ii) one copy of an amino acid sequence that is at least 90% identical SEQ ID NO: 450, (iii) one copy of an amino acid sequence that is at least 90% identical SEQ ID NO: 439; and (iv) one copy of an amino acid sequence that is at least 90% identical SEQ ID NO: 440; (15) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 432, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 433; (16) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 447, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 448;
(17) (i) one copy of an amino acid sequence that is at least 90% identical to SEQ ID NO: 429, (ii) one copy of an amino acid sequence that is at least 90% identical SEQ ID NO: 430, and (iii) one copy of an amino acid sequence that is at least 90% identical SEQ ID NO: 431. (18) (i) two copies of an amino acid sequence that is at least 90% identical to SEQ ID NO: 442, and (ii) two copies of an amino acid sequence that is at least 90% identical SEQ ID NO: 441. 23. The bispecific molecule of any one of claims 20-22, wherein said bispecific molecule a crosslinking dependent agonist of 4-1BB. 24 A nucleic acid comprising a nucleotide sequence encoding the 4-1BB antigen-binding protein of any one of claims 1-14 or 19, the PD-L1 antigen-binding protein of any one of claims 15-19, the bispecific molecule of any one of claims 20-23, or a polypeptide chain of any one of the foregoing. 25. A vector comprising the nucleic acid of claim 24. 26. A host cell comprising the nucleic acid of claim 24 or the vector of claim 25. 27. A pharmaceutical composition comprising (i) the 4-1BB antigen-binding protein of any one of claims 1-14 or 19, the PD-L1 antigen-binding protein of any one of claims 15-19, or the bispecific molecule of any one of claims 20-23, and (ii) a pharmaceutically acceptable carrier, excipient, or diluent. 28. A method of making the 4-1BB antigen-binding protein of the 4-1BB antigen-binding protein of any one of claims 1-14 or 19, the PD-L1 antigen-binding protein of any one of claims 15-19, or the bispecific molecule of any one of claims 20-23, comprising culturing the host cell of claim 26, under a condition wherein the 4-1BB antigen-binding protein, the PD-L1 antigen-binding protein, or the bispecific molecule is expressed. 29. A method of treating cancer, comprising administering to a subject in need thereof a therapeutically effective amount of the 4-1BB antigen-binding protein of any one of claims 1-14 or 19, the PD-L1 antigen-binding protein of any one of claims 15-19, the bispecific molecule of any one of claims 20-23, or the pharmaceutical composition claim 27.
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363580194P | 2023-09-01 | 2023-09-01 | |
| US63/580,194 | 2023-09-01 | ||
| US202363606175P | 2023-12-05 | 2023-12-05 | |
| US63/606,175 | 2023-12-05 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025049858A1 true WO2025049858A1 (en) | 2025-03-06 |
Family
ID=92816687
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/044603 Pending WO2025049858A1 (en) | 2023-09-01 | 2024-08-30 | Molecules for treatment of cancer |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025049858A1 (en) |
Citations (19)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5449752A (en) | 1991-05-02 | 1995-09-12 | Seikagaku Kogyo K.K. | Polypeptides with affinity to lipopolysaccharides and their uses |
| US6114598A (en) | 1990-01-12 | 2000-09-05 | Abgenix, Inc. | Generation of xenogeneic antibodies |
| US6162963A (en) | 1990-01-12 | 2000-12-19 | Abgenix, Inc. | Generation of Xenogenetic antibodies |
| WO2004004771A1 (en) | 2002-07-03 | 2004-01-15 | Ono Pharmaceutical Co., Ltd. | Immunopotentiating compositions |
| WO2004056875A1 (en) | 2002-12-23 | 2004-07-08 | Wyeth | Antibodies against pd-1 and uses therefor |
| WO2004072286A1 (en) | 2003-01-23 | 2004-08-26 | Ono Pharmaceutical Co., Ltd. | Substance specific to human pd-1 |
| US6833268B1 (en) | 1999-06-10 | 2004-12-21 | Abgenix, Inc. | Transgenic animals for producing specific isotypes of human antibodies via non-cognate switch regions |
| WO2005063815A2 (en) | 2003-11-12 | 2005-07-14 | Biogen Idec Ma Inc. | Fcϝ receptor-binding polypeptide variants and methods related thereto |
| US7064244B2 (en) | 1996-12-03 | 2006-06-20 | Abgenix, Inc. | Transgenic mammals having human Ig loci including plural VH and VK regions and antibodies produced therefrom |
| WO2008156712A1 (en) | 2007-06-18 | 2008-12-24 | N. V. Organon | Antibodies to human programmed death receptor pd-1 |
| US8008449B2 (en) | 2005-05-09 | 2011-08-30 | Medarex, Inc. | Human monoclonal antibodies to programmed death 1 (PD-1) and methods for treating cancer using anti-PD-1 antibodies alone or in combination with other immunotherapeutics |
| US20110271358A1 (en) | 2008-09-26 | 2011-11-03 | Dana-Farber Cancer Institute, Inc. | Human anti-pd-1, pd-l1, and pd-l2 antibodies and uses therefor |
| US8168757B2 (en) | 2008-03-12 | 2012-05-01 | Merck Sharp & Dohme Corp. | PD-1 binding proteins |
| US20140178905A1 (en) | 2001-12-03 | 2014-06-26 | Amgen Inc. | Antibody categorization based on binding characteristics |
| WO2015119923A1 (en) | 2014-02-04 | 2015-08-13 | Pfizer Inc. | Combination of a pd-1 antagonist and a 4-abb agonist for treating cancer |
| WO2016134358A1 (en) * | 2015-02-22 | 2016-08-25 | Sorrento Therapeutics, Inc. | Antibody therapeutics that bind cd137 |
| US9546203B2 (en) | 2013-03-14 | 2017-01-17 | Amgen Inc. | Aglycosylated Fc-containing polypeptides with cysteine substitutions |
| WO2018039180A1 (en) | 2016-08-24 | 2018-03-01 | Teneobio, Inc. | Transgenic non-human animals producing modified heavy chain-only antibodies |
| EP3470428A1 (en) * | 2017-10-10 | 2019-04-17 | Numab Innovation AG | Antibodies targeting cd137 and methods of use thereof |
-
2024
- 2024-08-30 WO PCT/US2024/044603 patent/WO2025049858A1/en active Pending
Patent Citations (24)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US6114598A (en) | 1990-01-12 | 2000-09-05 | Abgenix, Inc. | Generation of xenogeneic antibodies |
| US6162963A (en) | 1990-01-12 | 2000-12-19 | Abgenix, Inc. | Generation of Xenogenetic antibodies |
| US5449752A (en) | 1991-05-02 | 1995-09-12 | Seikagaku Kogyo K.K. | Polypeptides with affinity to lipopolysaccharides and their uses |
| US7064244B2 (en) | 1996-12-03 | 2006-06-20 | Abgenix, Inc. | Transgenic mammals having human Ig loci including plural VH and VK regions and antibodies produced therefrom |
| US7049426B2 (en) | 1999-06-10 | 2006-05-23 | Abgenix, Inc. | Transgenic animals for producing specific isotypes of human antibodies via non-cognate switch regions |
| US6833268B1 (en) | 1999-06-10 | 2004-12-21 | Abgenix, Inc. | Transgenic animals for producing specific isotypes of human antibodies via non-cognate switch regions |
| US20140178905A1 (en) | 2001-12-03 | 2014-06-26 | Amgen Inc. | Antibody categorization based on binding characteristics |
| WO2004004771A1 (en) | 2002-07-03 | 2004-01-15 | Ono Pharmaceutical Co., Ltd. | Immunopotentiating compositions |
| US7521051B2 (en) | 2002-12-23 | 2009-04-21 | Medimmune Limited | Methods of upmodulating adaptive immune response using anti-PD-1 antibodies |
| WO2004056875A1 (en) | 2002-12-23 | 2004-07-08 | Wyeth | Antibodies against pd-1 and uses therefor |
| US7488802B2 (en) | 2002-12-23 | 2009-02-10 | Wyeth | Antibodies against PD-1 |
| WO2004072286A1 (en) | 2003-01-23 | 2004-08-26 | Ono Pharmaceutical Co., Ltd. | Substance specific to human pd-1 |
| WO2005063815A2 (en) | 2003-11-12 | 2005-07-14 | Biogen Idec Ma Inc. | Fcϝ receptor-binding polypeptide variants and methods related thereto |
| US8008449B2 (en) | 2005-05-09 | 2011-08-30 | Medarex, Inc. | Human monoclonal antibodies to programmed death 1 (PD-1) and methods for treating cancer using anti-PD-1 antibodies alone or in combination with other immunotherapeutics |
| US8952136B2 (en) | 2007-06-18 | 2015-02-10 | Merck Sharp & Dohme B.V. | Antibodies to human programmed death receptor PD-1 |
| US8354509B2 (en) | 2007-06-18 | 2013-01-15 | Msd Oss B.V. | Antibodies to human programmed death receptor PD-1 |
| WO2008156712A1 (en) | 2007-06-18 | 2008-12-24 | N. V. Organon | Antibodies to human programmed death receptor pd-1 |
| US8168757B2 (en) | 2008-03-12 | 2012-05-01 | Merck Sharp & Dohme Corp. | PD-1 binding proteins |
| US20110271358A1 (en) | 2008-09-26 | 2011-11-03 | Dana-Farber Cancer Institute, Inc. | Human anti-pd-1, pd-l1, and pd-l2 antibodies and uses therefor |
| US9546203B2 (en) | 2013-03-14 | 2017-01-17 | Amgen Inc. | Aglycosylated Fc-containing polypeptides with cysteine substitutions |
| WO2015119923A1 (en) | 2014-02-04 | 2015-08-13 | Pfizer Inc. | Combination of a pd-1 antagonist and a 4-abb agonist for treating cancer |
| WO2016134358A1 (en) * | 2015-02-22 | 2016-08-25 | Sorrento Therapeutics, Inc. | Antibody therapeutics that bind cd137 |
| WO2018039180A1 (en) | 2016-08-24 | 2018-03-01 | Teneobio, Inc. | Transgenic non-human animals producing modified heavy chain-only antibodies |
| EP3470428A1 (en) * | 2017-10-10 | 2019-04-17 | Numab Innovation AG | Antibodies targeting cd137 and methods of use thereof |
Non-Patent Citations (57)
| Title |
|---|
| "Acta Cryst. D. and Phenix", ACTA CRYST, vol. D66, 2010, pages 213 - 221 |
| "REMINGTON'S PHARMACEUTICAL SCIENCES", 1990, MACK PUBLISHING COMPANY |
| ARORA: "Cell Culture Media: A Review", MATER METHODS, vol. 3, 2013, pages 175 |
| BERCOVICI ET AL., CLIN DIAGN LAB IMMUNOL, vol. 7, no. 6, 2000, pages 859 - 864 |
| BIRD ET AL., SCIENCE, vol. 242, 1988, pages 423 - 426 |
| CHAN ET AL.: "Fmoc Solid Phase Peptide Synthesis", 2005, OXFORD UNIVERSITY PRESS |
| CHAND ET AL., BIOLOGICALS, vol. 46, 2017, pages 168 - 171 |
| CHICHILI ET AL., PROTEIN SCI, vol. 22, no. 2, February 2013 (2013-02-01), pages 153 - 167 |
| CHIN, NATURE COMMUNICATIONS, vol. 9, 2018 |
| CHOTHIA ET AL., NATURE, vol. 342, 1989, pages 877 - 883 |
| CHOTHIALESK, J MOL BIOL., vol. 196, 1987, pages 901 - 917 |
| CLARKE STARLYNN C ET AL: "Multispecific Antibody Development Platform Based on Human Heavy Chain Antibodies", FRONTIERS IN IMMUNOLOGY, FRONTIERS MEDIA, LAUSANNE, CH, vol. 9, 1 January 2019 (2019-01-01), pages 3037 - 1, XP009512208, ISSN: 1664-3224, [retrieved on 20190107], DOI: 10.3389/FIMMU.2018.03037 * |
| CLAY ET AL., CLIN CANCER RES, vol. 7, no. 5, 2001, pages 1127 - 35 |
| DUBROT J ET AL., CANCER IMMUNOL., vol. 59, 2010, pages 1221 - 1233 |
| E. W. MARTIN: "Remington's Pharmaceutical Sciences", 1980, MACK PUBLISHING CO. |
| EDELMAN ET AL., PROC. NATL. ACAD. SCI. USA, vol. 63, no. 1, 1969, pages 78 - 85 |
| ESTES ET AL.: "Next generation Fc scaffold for multispecific antibodies", ISCIENCE, vol. 24, 2021, pages 103447, XP055932216, DOI: 10.1016/j.isci.2021.103447 |
| FU ET AL., PLOS ONE, vol. 5, no. 7, 2010, pages e11867 |
| GOOLIA ET AL., J VET DIAGN INVEST, vol. 29, no. 2, 2017, pages 250 - 253 |
| GREEN ET AL., NATURE GENETICS, vol. 7, 1994, pages 13 - 21 |
| GREENJAKOBOVITS, J. EX. MED, vol. 188, 1998, pages 483 - 495 |
| HOLLIGER ET AL., PROC. NATL. ACAD. SCI. USA, vol. 90, 1993, pages 6444 - 6448 |
| HUSTON ET AL., PROC. NATL. ACAD. SCI. USA, vol. 85, 1988, pages 5879 - 5883 |
| JACOBSEN ET AL.: "Engineering an IgG scaffold lacking effector function with optimized developability", J. BIOL. CHEM., vol. 292, 2017, pages 1865 - 1875, XP055604093, DOI: 10.1074/jbc.M116.748525 |
| JANEWAY ET AL.: "Immunobiology: The Immune System in Health and Disease", 1999, ELSEVIER SCIENCE LTD./GARLAND PUBLISHING, article "Structure of the Antibody Molecule and the Immunoglobulin Genes" |
| KABAT ET AL.: "Sequences of Proteins of Immunological Interest", 1991, NATIONAL INSTITUTES OF HEALTH |
| KELLERMANGREEN, CURRENT OPINION IN BIOTECHNOLOGY, vol. 13, 2002, pages 593 - 597 |
| KHAN, ADV PHARM BULL, vol. 3, no. 2, 2013, pages 257 - 263 |
| KUNERT ET AL., APPL. MICROBIOL BIOTECHNOL., vol. 100, 2016, pages 3451 - 61 |
| KWON Y.H.WEISSMAN S.M., PROC. NATL. ACAD. SCI. USA, vol. 86, 1989, pages 1963 - 1967 |
| LI ET AL.: "Cell culture processes for monoclonal antibody production", MABS, vol. 2, no. 5, 2010, pages 466 - 477, XP055166177, DOI: 10.4161/mabs.2.5.12720 |
| LIU ET AL., ANAL BIOCHEM, vol. 525, 2017, pages 89 - 91 |
| LIU ET AL., J BIOL CHEM, vol. 292, 2016, pages 1876 - 1883 |
| MACATANGAY ET AL., CLIN VACCINE IMMUNOL, vol. 17, no. 9, 2010, pages 1452 - 1459 |
| MACCALLUM ET AL., J. MOL. BIOL., vol. 262, 1996, pages 732 - 745 |
| MAKABE ET AL., J BIOL. CHEM., vol. 283, 2008, pages 1156 - 1166 |
| MAKABE ET AL., J. BIOL. CHEM., vol. 283, 2008, pages 1156 - 1166 |
| MARTIN: "Handbook of Therapeutic Antibodies", 2008, WILEY-VCH VERLAG GMBH, article "Bioinformatics tools for antibody engineering", pages: 95 - 117 |
| MELERO I ET AL., NAT. MED., vol. 3, 1997, pages 682 - 685 |
| MENDEZ ET AL., NATURE GENETICS, vol. 15, 1997, pages 146 - 156 |
| MURIUO O ET AL., EUR. J. IMMUNOL., vol. 39, 2009, pages 2424 - 2436 |
| NARAZAKI H, BLOOD, vol. 115, 2010, pages 1941 - 1948 |
| NGUYEN ET AL., SENSORS (BASEL, vol. 15, no. 5, 5 May 2015 (2015-05-05), pages 10481 - 510 |
| NORTH ET AL., J. MOL. BIOL., vol. 406, 2011, pages 228 - 256 |
| POLJAK ET AL., STRUCTURE, vol. 2, 1994, pages 1121 - 1123 |
| RUDIKOFF S ET AL: "Single amino acid substitution altering antigen-binding specificity", PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES (PNAS), NATIONAL ACADEMY OF SCIENCES, vol. 79, no. 6, 1 March 1982 (1982-03-01), pages 1979 - 1983, XP002986051, ISSN: 0027-8424, DOI: 10.1073/PNAS.79.6.1979 * |
| SAMBROOK ET AL.: "A Laboratory Manual", 2001, COLD SPRING HARBOR PRESS, article "Molecular Cloning" |
| SHAH ET AL., J VIS EXP, no. 84, 2014, pages 51383 |
| SHIMAMOTO ET AL., MABS, vol. 4, no. 5, 2012, pages 586 - 591 |
| SIMEONE E.ASCIERTO P.A., J. IMMUNOTOXICOLOGY, vol. 9, 2012, pages 241 - 247 |
| SNELL L.M. ET AL., IMMUNOL. REV., vol. 244, 2011, pages 197 - 217 |
| SPIESS ET AL., MOLECULAR IMMUNOLOGY, vol. 67, no. 2, 2015, pages 97 - 106 |
| STAGG J ET AL., PROC. NATL. ACAD. SCI. USA, vol. 108, 2011, pages 7142 - 7147 |
| TAM ET AL., CIRCULATION, vol. 98, no. 11, pages 1085 - 1091 |
| TRIKHA ET AL., INT J CANCER, vol. 110, 2004, pages 326 - 335 |
| WHO DRUG INFORMATION, vol. 27, no. 1, 2013, pages 161 - 162 |
| YI ET AL., PLOS ONE, vol. 9, no. 1, 2014, pages e86337 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US20220387586A1 (en) | Bispecific antibodies specifically binding to pd1 and lag3 | |
| JP7654539B2 (en) | Anti-αvβ8 antibodies, compositions and uses thereof | |
| TWI781098B (en) | Novel b7-h3-binding molecules, antibody drug conjugates thereof and methods of use thereof | |
| US20240301087A1 (en) | Anti-steap1 antigen-binding protein | |
| AU2019411511B2 (en) | Anti-BTLA antibodies | |
| EP4217391A1 (en) | Novel human antibodies binding to human cd3 epsilon | |
| AU2025275198A1 (en) | BTLA antibodies | |
| WO2025049858A1 (en) | Molecules for treatment of cancer | |
| AU2024330458A1 (en) | Molecules for treatment of cancer | |
| US20240269276A1 (en) | Specific binding protein targeting pd-l1 and cd73 | |
| CA3098096C (en) | Antibody against tim-3 and application thereof | |
| RU2812478C2 (en) | ANTI-αvβ8 ANTIBODIES AND COMPOSITIONS AND THEIR USE | |
| WO2025259515A2 (en) | Combination treatment | |
| WO2025096842A2 (en) | Fibroblast targeting molecules | |
| TW202305007A (en) | Specific binding protein targeting pd-l1 and cd73 | |
| WO2025235357A1 (en) | Bispecific antibodies that bind to cd28 and nectin-4 | |
| JP2026021532A (en) | BTLA antibody | |
| AU2024226156A1 (en) | C-kit binding proteins, chimeric antigen receptors, and uses thereof | |
| HK40042102A (en) | Anti-avb8 antibodies and compositions and uses thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24773246 Country of ref document: EP Kind code of ref document: A1 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: AU2024330458 Country of ref document: AU |