WO2025042806A2 - C-met binding antibodies, nucleic acids encoding same, and methods of use - Google Patents
C-met binding antibodies, nucleic acids encoding same, and methods of use Download PDFInfo
- Publication number
- WO2025042806A2 WO2025042806A2 PCT/US2024/042856 US2024042856W WO2025042806A2 WO 2025042806 A2 WO2025042806 A2 WO 2025042806A2 US 2024042856 W US2024042856 W US 2024042856W WO 2025042806 A2 WO2025042806 A2 WO 2025042806A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- vhh
- seq
- amino acid
- acid sequence
- set forth
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/28—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants
- C07K16/2863—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against receptors, cell surface antigens or cell surface determinants against receptors for growth factors, growth regulators
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P35/00—Antineoplastic agents
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/505—Medicinal preparations containing antigens or antibodies comprising antibodies
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/20—Immunoglobulins specific features characterized by taxonomic origin
- C07K2317/22—Immunoglobulins specific features characterized by taxonomic origin from camelids, e.g. camel, llama or dromedary
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/20—Immunoglobulins specific features characterized by taxonomic origin
- C07K2317/24—Immunoglobulins specific features characterized by taxonomic origin containing regions, domains or residues from different species, e.g. chimeric, humanized or veneered
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/31—Immunoglobulins specific features characterized by aspects of specificity or valency multispecific
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/30—Immunoglobulins specific features characterized by aspects of specificity or valency
- C07K2317/33—Crossreactivity, e.g. for species or epitope, or lack of said crossreactivity
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/56—Immunoglobulins specific features characterized by immunoglobulin fragments variable (Fv) region, i.e. VH and/or VL
- C07K2317/567—Framework region [FR]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/56—Immunoglobulins specific features characterized by immunoglobulin fragments variable (Fv) region, i.e. VH and/or VL
- C07K2317/569—Single domain, e.g. dAb, sdAb, VHH, VNAR or nanobody®
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/60—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments
- C07K2317/64—Immunoglobulins specific features characterized by non-natural combinations of immunoglobulin fragments comprising a combination of variable region and constant region components
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/73—Inducing cell death, e.g. apoptosis, necrosis or inhibition of cell proliferation
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/76—Antagonist effect on antigen, e.g. neutralization or inhibition of binding
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/77—Internalization into the cell
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/92—Affinity (KD), association rate (Ka), dissociation rate (Kd) or EC50 value
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/94—Stability, e.g. half-life, pH, temperature or enzyme-resistance
Definitions
- the polypeptide or VHH is conjugated, fused, or linked to an amino acid sequence that is at least 80%, at least 85%, at least 90%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or 100% identical to the human IgG4 PAA hinge and Fc domain sequence set forth in SEQ ID NO:643.
- the polypeptide or VHH is conjugated, fused, or linked to an amino acid sequence that is identical to the human IgG4 PAA hinge and Fc domain sequence set forth in SEQ ID NO:643 except for 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 amino acid substitutions.
- the Fc domain includes mutations that promote heterodimerization (e.g., knob-into hole mutations, electrostatic steering mutations).
- the disclosure features a polypeptide comprising a variable domain of the heavy chain of a heavy chain-only antibody (VHH) that specifically binds c-MET (human and/or cyno c-MET), the VHH comprising VHH complementarity determining region (CDR) 1, VHH CDR2, and VHH CDR3 of any of the amino acid sequences set forth in SEQ ID NOs.: 1 to 40, wherein 1 to 20 amino acids in the framework regions of the VHH are humanized, wherein the polypeptide is linked to a half-life extending moiety, and optionally, wherein the polypeptide is conjugated, fused, or linked to an amino acid sequence that is at least 80%, at least 85%, at least 90%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%,
- the polypeptide competes with HGF for binding to c-MET. In other cases, the polypeptide does not compete with HGF for binding to c- MET.
- the polypeptide or VHH comprises a second VHH that specifically binds to an antigen selected from the group consisting of: CD3, EGFR, VEGF, VEGFR, HGF, CXCR2, CXCR4, CXCR7, CXCL11, CXCL12, CEA, PSMA, MMR, PD-1, PDL-1, survivin, HER2, HGFR, P2X7, death receptor 5 CTLA4, CD7, CD8, CD11b, CD20, CD38, CD45, M HC-II, fibronectin, TUFM, CapG, CAIX, CD33, CD47, ARTC22, B3GAT1 (CD57), CCR7 (CD197), CD16, CD16a, CD2 CD226, CD244, CD27, CD3, CD300A, CD34, CD58,
- the VHH CDR1, VHH CDR2, and VHH CDR3 comprises the corresponding VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of any one of the forty clones set forth in Tables 3 to 7.
- the VHH specifically binds c-Met (human and/or cyno) and comprises: (a) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:41, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:42, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:43; (b) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:44, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:45, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:46; (c) a VHH CDR1 comprising the amino acid sequence set forth
- the VHH binds to human c-MET expressed on CHO cells with an EC 50 of about 0.1 to about 250 nM when the VHH is in monovalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC 50 of about 0.1 to about 100 nM when the VHH is in monovalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC 50 of about 0.1 to about 75 nM when the VHH is in monovalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC 50 of about 0.1 to about 50 nM when the VHH is in monovalent format.
- the VHH binds to human c-MET expressed on CHO cells with an EC 50 of about 0.1 to about 25 nM when the VHH is in bivalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC 50 of about 0.1 to about 15 nM when the VHH is in bivalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC 50 of about 0.1 to about 10 nM when the VHH is in bivalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC 50 of about 0.1 to about 5 nM when the VHH is in bivalent format.
- the polypeptide is a Fab.
- the means for binding human and cynomolgus c- MET is linked to the polypeptide via a peptide linker (e.g., glycine linker, serine linker, glycine-serine linker).
- the linker is G4S (SEQ ID NO: 806), (G4S)3 (SEQ ID NO: 649), or (G4S)5 (SEQ ID NO: 807).
- the different antigen is an antigen on a T cell.
- the different antigen is an antigen on an NK cell. In some cases, the different antigen is an antigen on a tumor cell. In some instances, the different antigen is an antigen selected from the group consisting of: CD3, EGFR, VEGF, VEGFR, HGF, CXCR2, CXCR4, CXCR7, CXCL11, CXCL12, CEA, PSMA, MMR, PD-1, PDL-1, survivin, HER2, HGFR, P2X7, death receptor 5 CTLA4, CD7, CD8, CD11b, CD20, CD38, CD45, M HC-II, fibronectin, TUFM, CapG, CAIX, CD33, CD47, ARTC22, B3GAT1 (CD57), CCR7 (CD197), CD16, CD16a, CD16b, CD2 CD226, CD244, CD27, CD3, CD300A, CD34, CD58, CD59, CD69, CSF2, CX3CR1, CXCR1
- the first human Ig and second human Ig are a human IgG1, human IgG2, human IgG3, or human IgG4, or variants thereof.
- the variant differs from the native human IgG hinge and/or Fc sequence by 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 amino acids.
- the C-terminal of the polypeptide and the C-terminal of the binding molecule are each linked to a sequence comprising an amino acid sequence that is at least 80%, at least 85%, at least 90%, at least 95%, or 100% identical to the IgG4 PAA hinge and Fc domain set forth in SEQ ID NO:643.
- the disclosure features a c-MET binding chimeric antigen receptor (CAR) comprising any one or more of the above-described polypeptides or VHHs.
- the CAR comprises a c-MET binding domain, a transmembrane domain, and an intracellular signaling domain.
- the transmembrane domain is from a CD3, CD4, CD8, or CD28 molecule.
- the intracellular signaling domain is from CD3-zeta.
- the c-MET binding CAR further comprises CD28 and CD137 signaling domains and CD3 ⁇ (c-Met-28-137-3 ⁇ ).
- the c-MET CAR comprises a CD28 hinge region, a CD28 transmembrane domain, and a FC ⁇ R1 ⁇ intracellular T cell signaling domain, wherein the CD28 transmembrane domain is connected to a FC ⁇ R1 ⁇ intracellular T cell signaling domain.
- the c-MET binding CAR is expressed on the surface of a T cell. In other cases, the c-MET binding CAR is expressed on the surface of an NK cell.
- the CAR comprises a means for binding human and cynomolgus c-MET, a transmembrane domain, and an intracellular signaling domain.
- the disclosure features a nucleic acid or nucleic acids encoding the polypeptide or VHH described above, the bispecific antibody described above, or the c-MET binding CAR described above.
- a vector or vectors comprising the nucleic acid or nucleic acids described above.
- a host cell comprising the nucleic acid or nucleic acids of claim 8, or the vector or vectors described above.
- the disclosure features a method of making a polypeptide, a bispecific antibody, or the c-MET binding CAR.
- the method comprises culturing a host cell described above under conditions that facilitate expression of the polypeptide, the bispecific antibody, or the c-MET binding CAR. In some cases, the method further involves isolating the polypeptide, the bispecific antibody, or the c-MET binding CAR. In certain cases, the method further comprises formulating the polypeptide, the bispecific antibody, or the c-MET binding CAR as a sterile pharmaceutical composition. In another aspect, provided herein is a pharmaceutical composition comprising the polypeptide, the bispecific antibody, or the c-MET binding CAR described herein, and a pharmaceutically acceptable carrier.
- the disclosure relates to a pharmaceutical composition
- a pharmaceutical composition comprising a pharmaceutically acceptable carrier and a means for binding human and cynomolgus c-MET.
- the disclosure features a method of treating a c-MET expressing cancer in a human subject in need thereof, or killing a tumor cell in a human subject in need thereof, or decreasing the rate of tumor growth in a human subject in need thereof.
- the method comprises administering to the human subject a therapeutically effective amount of the polypeptide o, the bispecific antibody, or a T cell or NK cell expressing the c-MET binding CAR described herein
- the cancer is a solid tumor.
- the cancer is selected from the group consisting of gastric cancer, esophageal cancer, medulloblastoma, glioma, colon cancer, head and neck cancer, lung cancer, kidney cancer, thyroid cancer, colorectal cancer, pancreatic cancer, ovarian cancer, and breast cancer.
- the disclosure features a mRNA comprising an open reading frame (ORF) encoding a VHH, a polypeptide, a bispecific antibody, or a c-MET binding CAR described herein.
- the mRNA comprises a 5’terminal cap, a 5’UTR, a 3’UTR, and a poly A region.
- the disclosure features a polynucleotide comprising an mRNA comprising: (i) a 5' UTR; (ii) an open reading frame (ORF) encoding a polypeptide, the bispecific antibody, or the c-MET binding CAR described herein; (iii) a stop codon; and (iv) a 3' UTR.
- the mRNA comprises a microRNA (miR) binding site.
- the microRNA is expressed in an immune cell of hematopoietic lineage or a cell that expresses TLR7 and/or TLR8 and secretes pro-inflammatory cytokines and/or chemokines.
- the microRNA binding site is for a microRNA selected from miR-126, miR-142, miR-144, miR-146, miR-150, miR-155, miR-16, miR-21, miR- 223, miR-24, miR-27, miR-26a, or any combination thereof.
- the microRNA binding site is for a microRNA selected from miR126-3p, miR-142-3p, miR- 142-5p, miR-155, or any combination thereof.
- the microRNA binding site is located in the 3' UTR of the mRNA.
- the 5' UTR comprises a nucleic acid sequence at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:654.
- the 3' UTR comprises a nucleic acid sequence at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:674.
- wherein the mRNA comprises a 5' terminal cap.
- the sterol is cholesterol, adosterol, agosterol A, atheronals, avenasterol, azacosterol, blazein, cerevisterol, colestolone, cycloartenol, daucosterol, 7- dehydrocholesterol, 5-dehydroepisterol, 7-dehydrositosterol, 20 ⁇ ,22R- dihydroxycholesterol, dinosterol, epibrassicasterol, episterol, ergosterol, ergosterol, fecosterol, fucosterol, fungisterol, ganoderenic acid, ganoderic acid, ganoderiol, ganodermadiol, 7 ⁇ -hydroxycholesterol, 22R-hydroxycholesterol, 27-hydroxycholesterol, inotodiol, lanosterol, lathosterol, lichesterol, lucidadiol, lumisterol, oxycholesterol, oxysterol, parkeo
- the c-MET may be from human or cynomolgus monkey.
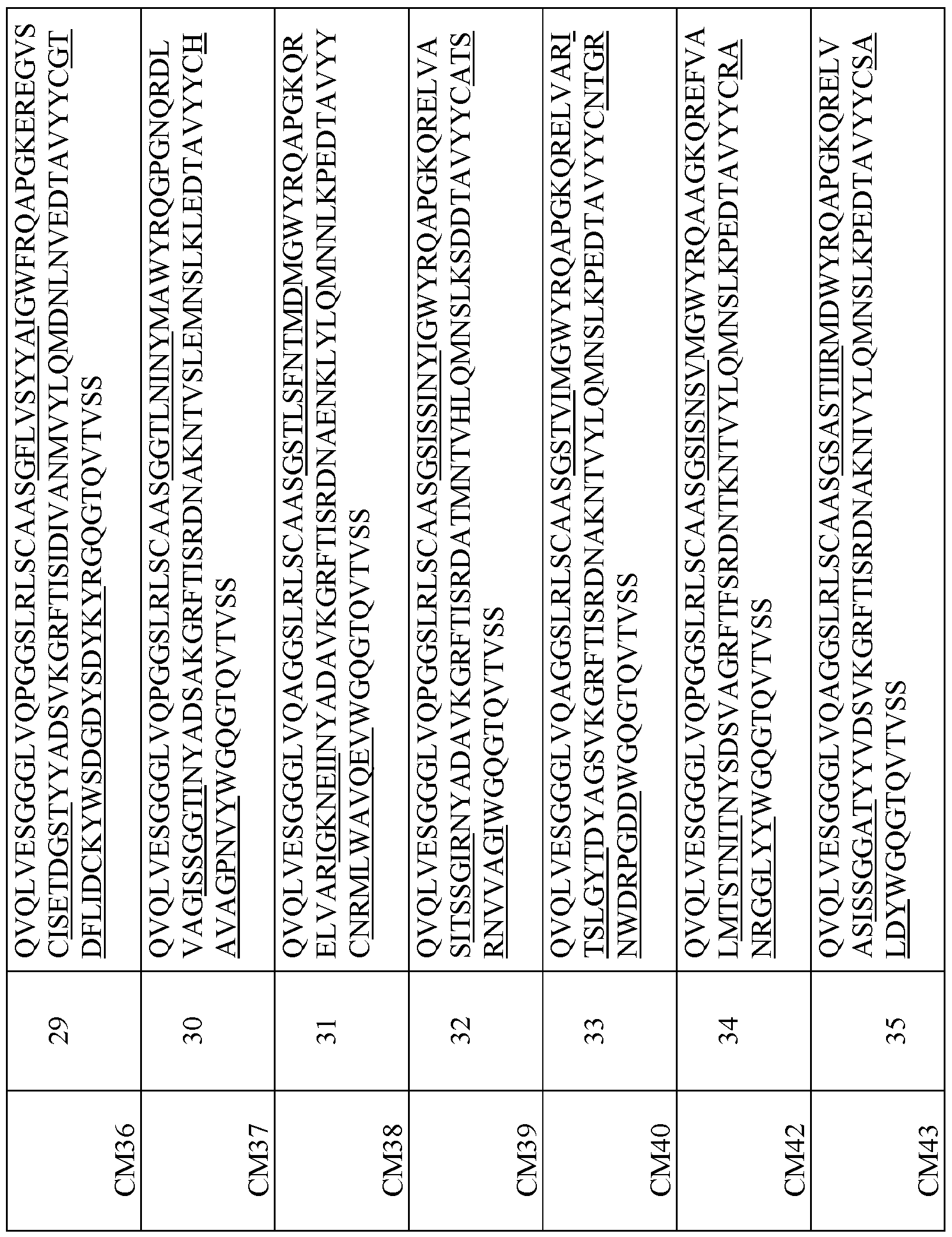
- Table 1 provides the amino acid sequences of several novel VHHs.
- This disclosure features a binding molecule or polypeptide that comprises the VHH amino acid sequence of any of the clones listed in Table 1.
- the binding molecule or polypeptide comprises a VHH that specifically binds c-MET (e.g., human and/or cyno) and comprises a sequence that is at least 80%, at least 85%, at least 90%, or at least 95% identical to an amino acid sequence set forth in any one of SEQ ID NOs.: 1 to 40.
- the binding molecule or polypeptide comprises a VHH that specifically binds c-MET (e.g., human and cyno) and comprises an amino acid sequence set forth in any one of SEQ ID NO:1 to 40 except for having 1, 2, 34, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 457, 48, 49, or 50 amino acid substitutions.
- the variability of the above sequences from an amino acid sequence set forth in any one of SEQ ID NO:1 to 40 may be due to humanization of the VHH.
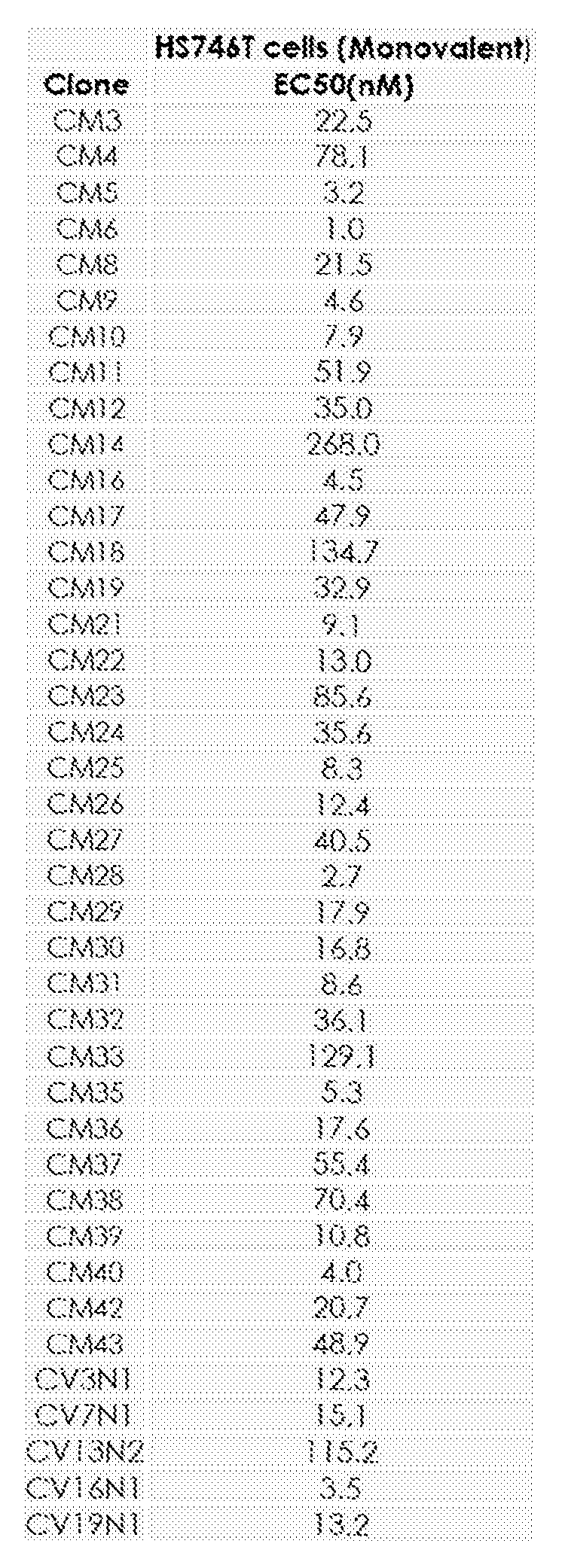
- Binding molecules or polypeptides comprising the amino acid sequence set forth in any one of SEQ ID NOs.: 1 to 40 in a monovalent format bind to a c-MET expressing cancer cell such as HS746T.
- the binding molecule or polypeptide comprises the VHH in a bivalent format.
- a bivalent format of the VHH is produced by linking the VHH to the same or another VHH.
- the amino acid sequence of human IgG4PAA is provided below (the hinge is shown in bold; the CH2 domain is italicized; and the CH3 domain is underlined; the CH2 and CH3 domains together form the Fc): ESKYGPPCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQ FNWYVDGVEVHNAKTKPREEQFNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSS IEKTISKAKGQPREPQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKSL SLSLGK (SEQ ID NO:643)
- the hinge and Fc domain comprises an amino acid sequence set forth in SEQ ID NO:643 except for 1, 2, 34, 5, 6, 7, 8, 9, or 10 amino acid substitutions.
- substitutions can be made to e.g., alter effector function (e.g., increasing or decreasing effector function), or to promote heterodimerization.
- Such substitutions made as part of Fc domain engineering are well known in the art. See e.g., Liu H, Saxena A, Sidhu SS and Wu D (2017) Fc Engineering for Developing Therapeutic Bispecific Antibodies and Novel Scaffolds. Front. Immunol. 8:38. doi: 10.3389/fimmu.2017.00038; Wilkinson I, Anderson S, Fry J, Julien LA, Neville D, Qureshi O, et al. (2021) Fc-engineered antibodies with immune effector functions completely abolished.
- the binding molecule or polypeptide comprising a VHH in bivalent format binds a cell expressing human or cyno c-MET (e.g., a cell line such as CHO) with an EC50 of about 0.1 nM to about 10 nM. In some cases, the binding molecule or polypeptide comprising a VHH in bivalent format binds a cell expressing human or cyno c-MET (e.g., a cell line such as CHO) with an EC50 of about 0.1 nM to about 5 nM. All of the anti-c-MET VHHs disclosed herein bind to c-MET when the VHH is in a bivalent format.
- VHH constructs are competing – i.e., compete with HGF for binding c-MET (clones CM3, CM4, CM8, CM10, CM11, CM16, CM19, CM23, CM25, CM30, CM32, CM33, CM37, CM38, CM39, CM42); others are semi-competitive for binding to c-MET (CM12, CM14, CM17, CM18, CM21, CM24, CM31, CM40, CV7N1, CV13N2); while still others do not compete with HGF for binding c-MET (CM5, CM6, CM9, CM22, CM26, CM27, CM28, CM29, CM35, CM36, CM43).
- c-MET clones CM3, CM4, CM8, CM10, CM11, CM16, CM19, CM23, CM25, CM30, CM32, CM33,
- binding molecules or polypeptides that comprise a VHH that specifically binds c-MET (human and/or cyno), wherein the VHH comprises the VHH CDR1, VHH CDR2, and VHH CDR3 of the VHH of any one of the clones listed in Table 1.
- the VHH CDRs of the VHH can be based on any CDR definition. Exemplary CDR numbering systems are provided below as Table 2 (see, e.g., Lafrance et al,. Dev. Comp. Immunol., 27(1):55-77 (2003)).
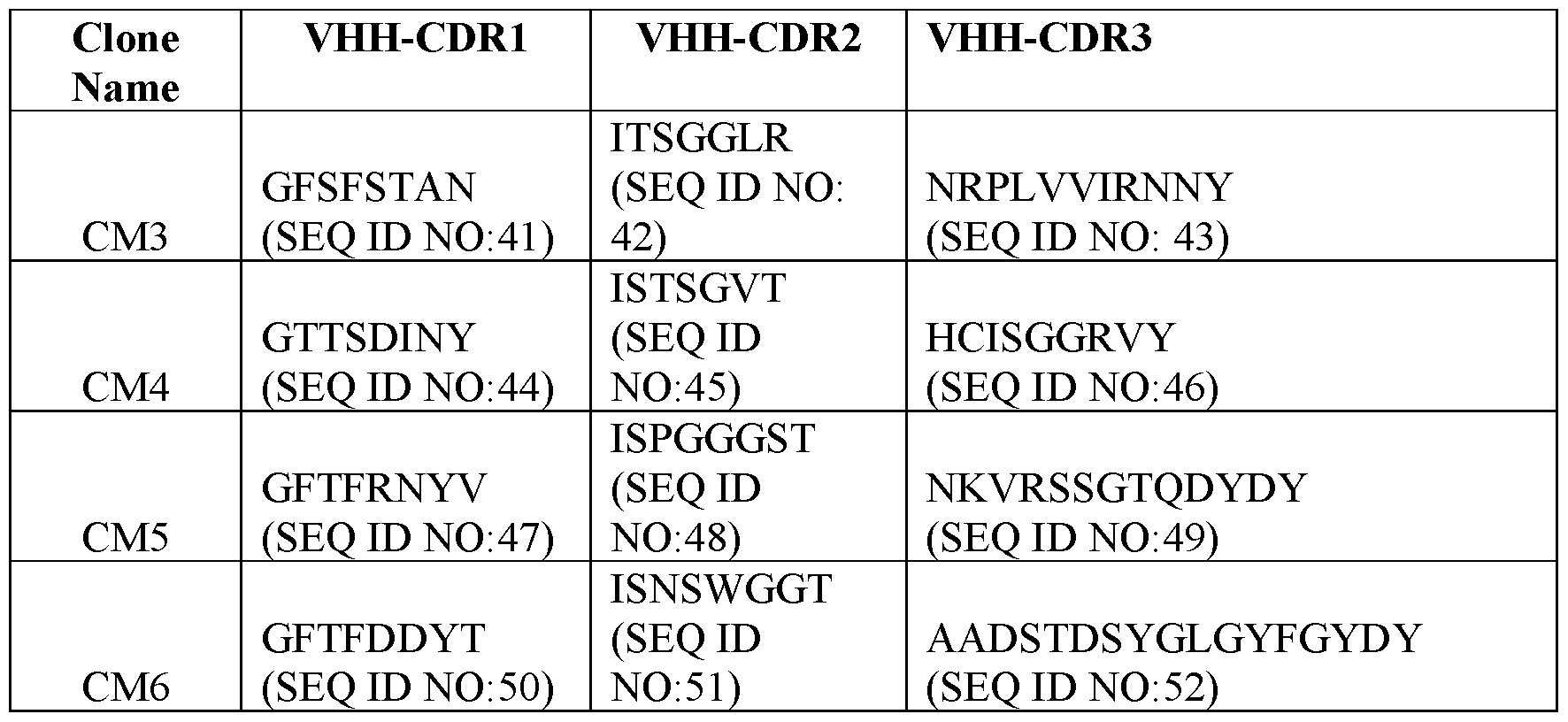
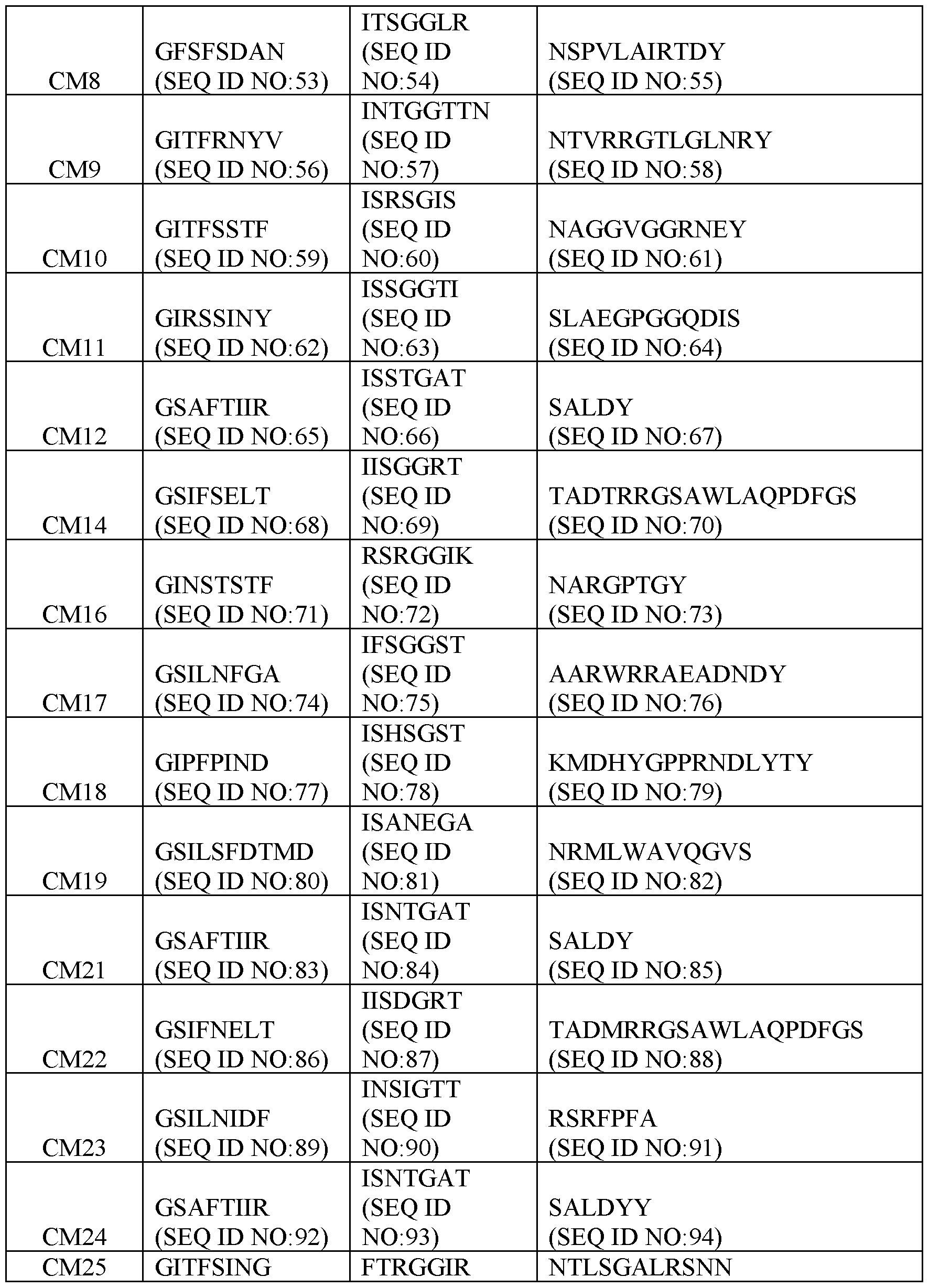
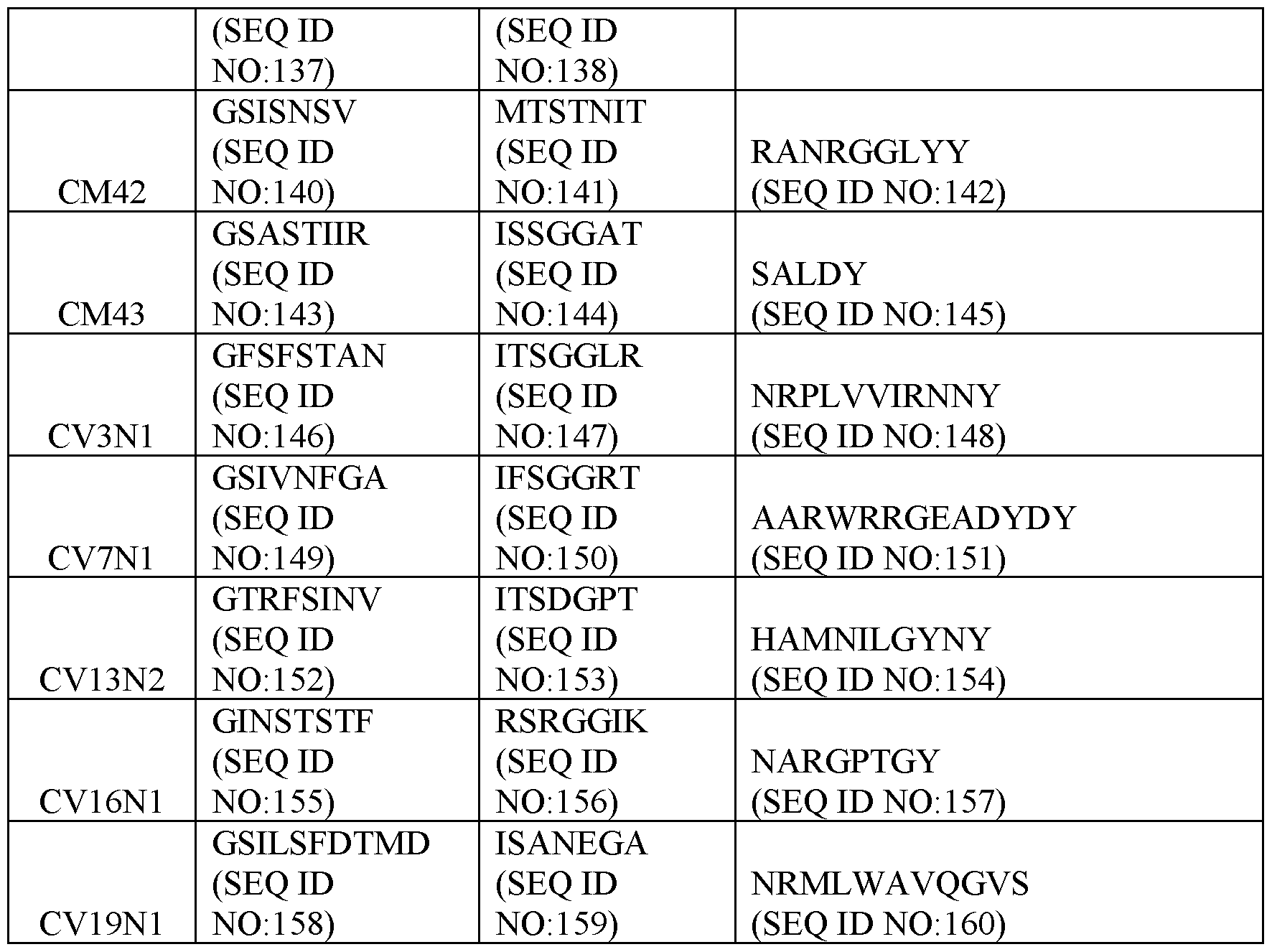
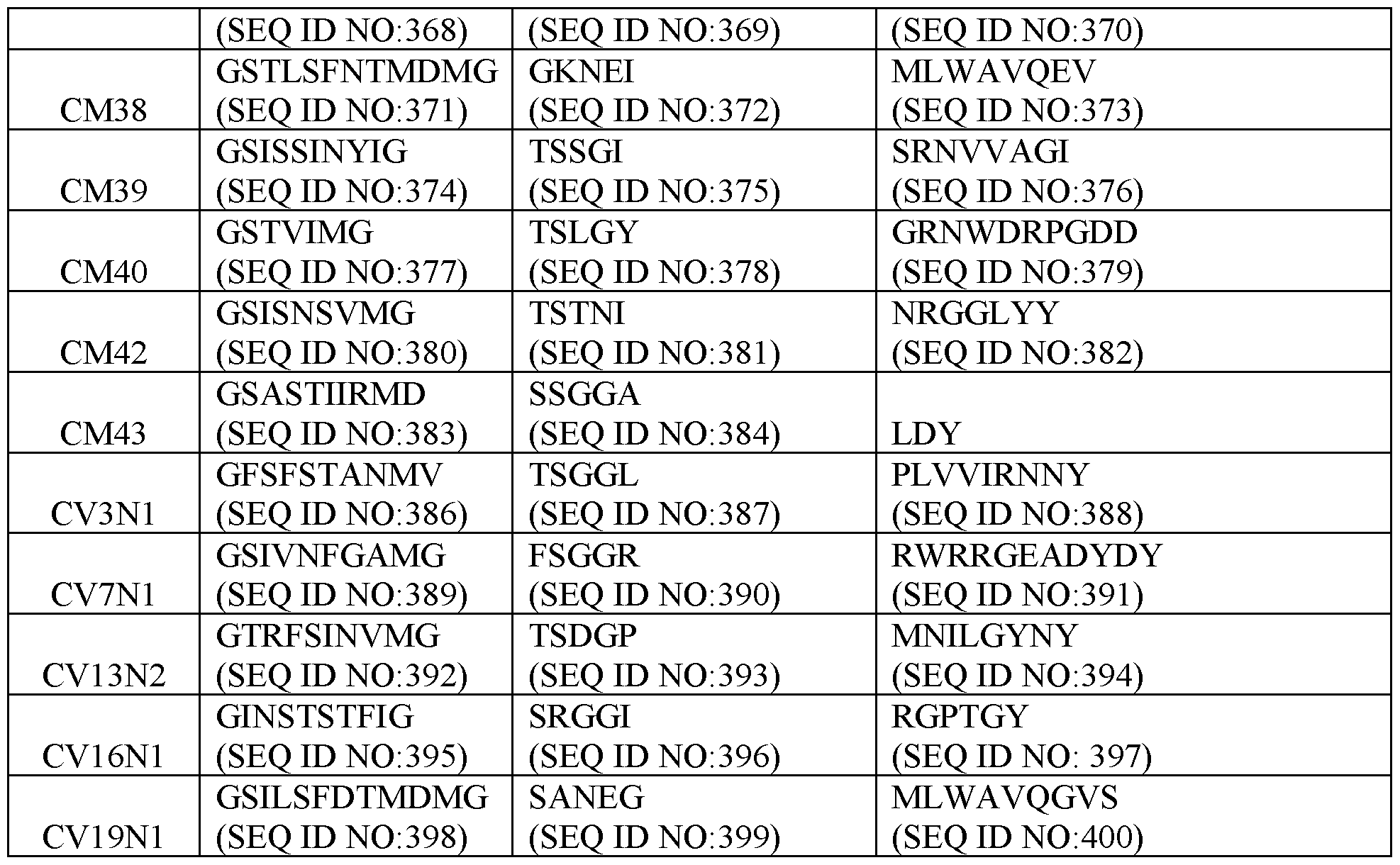
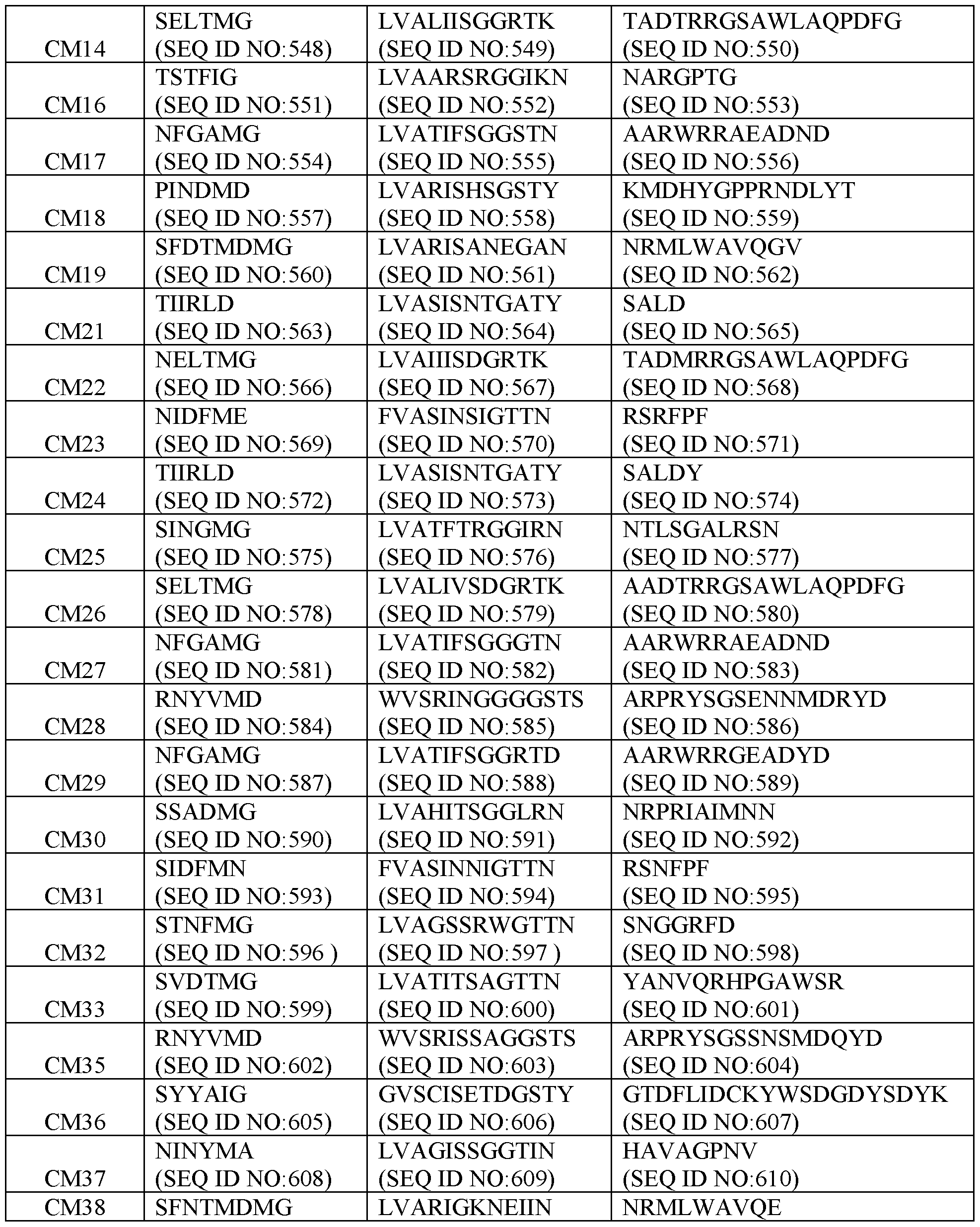
- Table 2 Exemplary CDR Definitions Provided in Tables 3 to 7 below are the VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of the clones according to IMGT, Kabat, Chothia, enhanced Chothia, and contact definitions.
- Table 3 VHH CDRs Based on IMGT CDR Definition
- Table 4 VHH CDRs Based on Kabat CDR Definition
- the disclosure provides a binding molecule or polypeptide comprising a VHH that specifically binds c-MET (human and/or cyno), wherein the VHH comprises the VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of any one clone listed in Table 3.
- the disclosure provides a binding molecule or polypeptide comprising a VHH that specifically binds c-MET (human and cyno), wherein the VHH comprises: (a) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:41, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:42, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:43; (b) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:44, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:45, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:46; (c) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:47, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:48, and a VHH CDR3 comprising the amino acid sequence
- the disclosure provides a binding molecule or polypeptide comprising a VHH that specifically binds c-MET (human and cyno), wherein the VHH comprises the VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of any one clone listed in Table 6.
- the disclosure provides a binding molecule or polypeptide comprising a VHH that specifically binds c-MET (human and cyno), wherein the VHH comprises the VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of any one clone listed in Table 7.
- the above VHHs are not humanized. In some instances, the above VHHs are humanized.
- 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, or 50 framework amino acids of the VHH are substituted as a result of humanization. In some instances, full humanization is avoided. In some cases, less than 20, 15, 10, 8, 7, 6, 5, 4, or 3 amino acids, or 1 to 25, 1 to 20, 1 to 15, 1 to 10, or 1 to 5 amino acids in the VHH framework regions are humanized. In some cases, 1 to 15 amino acids in the VHH framework regions of the sequences in Table 1 are humanized.
- the above VHHs are in a monovalent format. In some instances, the above VHHs are in a bivalent format (e.g., by linking the VHH to a hinge- Ig Fc domain). In some instances, the above VHHs are linked to a hinge-Fc of a human Ig (e.g., IgA, IgG1, IgG2, IgG3, or IgG4). In some instances, the above VHHs are linked to a hinge from human IgA.
- a human Ig e.g., IgA, IgG1, IgG2, IgG3, or IgG4
- the above VHHs are linked to a hinge from human IgG1 or IgG4. In some instances, the above VHHs comprise a Fc region from human IgG1 or human IgG4. In some instances, any of the above VHHs can be linked to a sequence that is at least 80%, at least 85%, at least 90%, or at least 95% identical to the amino acid sequence of human IgG4PAA (SEQ ID NO:643). In some instances, any of the above VHHs can be linked to a sequence that is identical to the amino acid sequence of human IgG4PAA (SEQ ID NO:643).
- the human Ig Fc region is modified to e.g., alter (increase or decrease) the effector function and/or to promote heterodimerization.
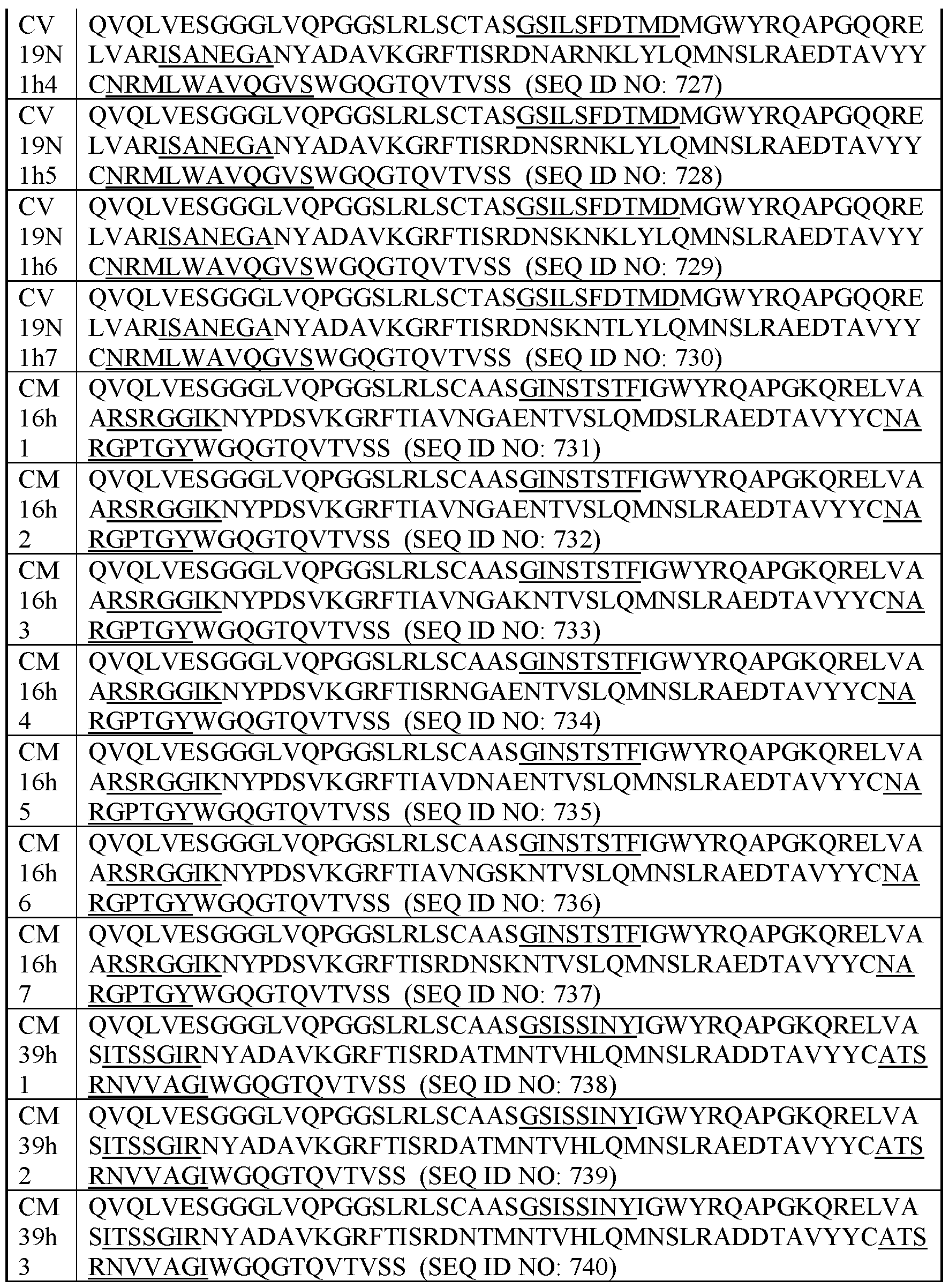
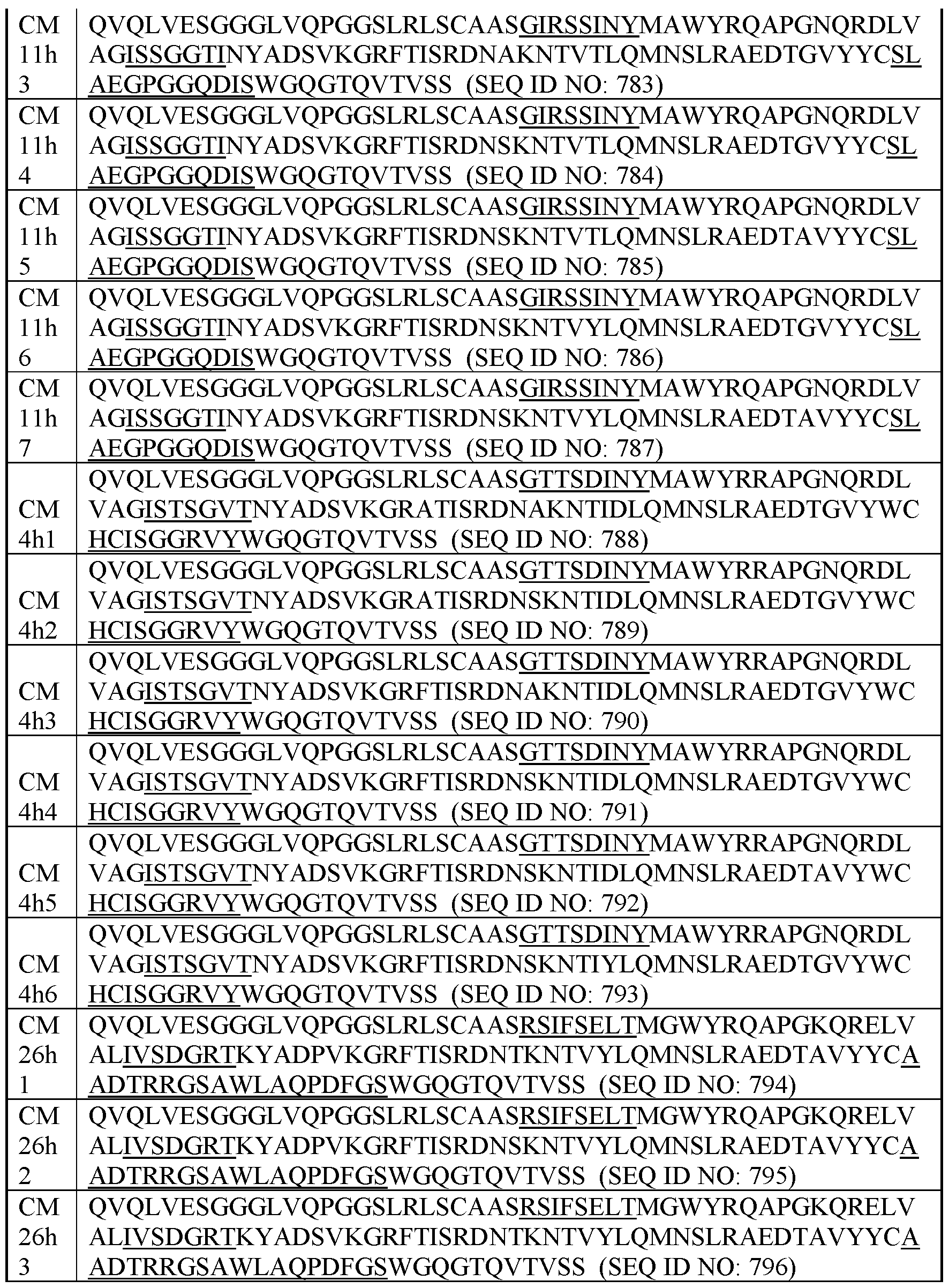
- exemplary Humanized C-MET VHHs This disclosure also encompasses humanized VHHs that specifically bind C- MET (human and/or cyno C-MET). Table I below provides the amino acid sequences of exemplary humanized VHHs that specifically bind C-MET (human and/or cyno C-MET). Table I.
- the Fc region is also engineered to improve its therapeutic activity. See e.g., Mimoto et al., Curr Pharm Biotechnol., 2016;17(15):1298-1314; Chen et al., Theranostics, 2021; 11(4): 1901–1917; Delidakis et al., Ann Rev. Biomed Engg., 24:249-274 (2022) (all incorporated by reference herein).
- Multispecific Constructs Another binding molecule encompassed by this disclosure is a multispecific construct. These multispecific constructs bind to three or more different epitopes. In one instance, three or more different VHHs are linked together. In some instances, one VHH is linked by its C-terminal to the N-terminal of the heavy chain of a whole antibody. In another instance, two VHHs are linked by their C-terminals to the N-terminals of the two heavy chains of a whole antibody. In some cases, the two VHHs are identical. In other cases, the two VHHs are different. In yet another instance, one VHH is linked by its C- terminal to the C-terminal of the heavy chain of a whole antibody.
- two VHHs are linked by their C-terminals to the C-terminals of the two heavy chains of a whole antibody.
- the two VHHs are identical.
- the two VHHs are different.
- a multispecific construct is based on a bispecific construct described above, wherein the bispecific construct is conjugated at the C- terminal of one or both of its heavy chains to one or two VHHs.
- the C-terminally conjugated VHH or VHHs can bind to a different epitope than the VHHs of the bispecific construct.
- at least one antigen binding domain specifically binds to c- MET.
- Chimeric Antigen Receptors Also encompassed by this disclosure are chimeric antigen receptors that include one or more VHHs described herein.
- a typical chimeric antigen receptor (CAR) includes four domains, each of which serves a distinct purpose; the antigen recognition domain at the exterior of the cell is responsible for antigen recognition, the hinge region and transmembrane domain provides stability whereas, the intracellular domain in the endodomain of the receptor plays a crucial role in transmitting signals to activate the effector function of CAR T or NK cells.
- the CAR comprises a VHH of this disclosure that specifically binds c-MET, a hinge region, a transmembrane domain, and an endodomain.
- Any hinge region, transmembrane domain, and endodomain that are employed in CARs can be used. See e.g., Ahmad et al., Advances in Cancer Biology – Metastasis, Vol.4, 100035 (2022); Feins et al., Am. J Hemaol., 94, S1:S3-S9, 2019; Hajari et al., IUBMB Life 71(2019) 1259-1267; De Munter et al., Int J Mol Sci 19(2018) 1-11; Xie et al., PNAS 116 (2019):7624-7631; Xie et al., Cancer Immunol.
- CARs of the present disclosure comprise an antigen binding domain (e.g., anti-c- MET VHH), a hinge domain, a transmembrane domain, and an intracellular signaling domain.
- the CARs comprise an antigen binding domain, a spacer domain, a transmembrane domain, a costimulatory domain, and a signaling domain.
- the antigen binding domain (e.g., anti-c-MET VHH) may be operably linked to another domain of the CAR, such as the hinge domain, the transmembrane domain and/or the intracellular domain, both described elsewhere herein, for expression in an immune cells (e.g., a T cell).
- a first nucleic acid sequence encoding the antigen binding domain (e.g., anti-c-MET VHH) is operably linked to a second nucleic acid encoding a hinge and transmembrane domain, and further operably linked to a third a nucleic acid sequence encoding an intracellular domain.
- the nucleic acid is mRNA.
- a polypeptide, binding molecule, or VHH is linked to a drug.
- a tumor targeted drug/therapy is conjugated to the polypeptide, binding molecule, or VHH of this disclosure to create an ADC.
- ADCs exploit the targeted efficiency of antibodies combined with the action of the cytotoxic payload conjugated to it. This results in specific targeting of cancer cells and thus lessens off-target side effects.
- the payload can be any drug that kills or inhibits the growth of the c-Met related/expressing cancer that is being targeted.
- the payload is multikinase inhibitor 178 or Mertansine (DM1).
- a polypeptide, binding molecule, or VHH is linked to a photosensitizer.
- a collagen trimerization moiety particularly a human collagen moiety, and a lung surfactant protein D moiety.
- a polypeptide, binding molecule, or VHH is linked to a moiety that facilitates the polypeptide crossing the blood brain barrier (e.g., anti-transferrin antibody, FC5, FC44, a VHH that binds IGF-1R, VHHs E9, ni3a, and pa2H).
- a polypeptide, binding molecule, or VHH is linked to a half-life extension moiety (e.g., PEG (linear and branched), human serum albumin, a VHH that binds human serum albumin, PASylation (PA600), human Ig Fc).
- the disclosure provides a nucleic acid or nucleic acids that encode the polypeptides, binding molecules, and VHHs of this disclosure.
- the nucleic acid may be DNA or mRNA.
- the nucleic acid may be modified.
- the coding sequence may be modified to take into account the preferred codon usage of the expression host.
- vectors that comprise the above nucleic acids. In some instances the vector is an expression vector. In other instances, the vector is a viral vector.
- host cells that comprise the nucleic acid or vectors above. In some instances, the host cells are bacterial cells, yeast cells, insect cells, or mammalian cells. In some instances, the host cell is E. coli.
- the host cell is Pichia pastoris or Saccharomyces cerevisiae. In some instances, the host cell is a CHO cell, a COS cell, a HEK293T cell, or a NIH3T3 cell.
- the disclosure also features methods of making the polypeptides, binding molecules, or VHHs described herein. The method involves culturing the host cell under conditions that promote the expression of the polypeptide, binding molecule, or VHH. In some instances, the polypeptide, binding molecule, or VHH is purified. In certain instances, the polypeptide, binding molecule, or VHH is formulated as a sterile pharmaceutical composition.
- compositions comprising the polypeptides, binding molecules, or VHHs described herein, and a pharmaceutically acceptable carrier.
- the pharmaceutically acceptable carrier is phosphate buffered saline.
- the pharmaceutically acceptable carrier is sterile distilled water.
- mRNA Constructs In some aspects, the polynucleotides disclosed herein are or function as a messenger RNA (mRNA).
- mRNA messenger RNA
- the term “messenger RNA” (mRNA) refers to any polynucleotide which encodes at least one peptide or polypeptide of interest and which is capable of being translated to produce the encoded peptide polypeptide of interest in vitro, in vivo, in situ or ex vivo.
- the disclosure provides a polynucleotide comprising an mRNA comprising: (i) a 5' UTR; (ii) an open reading frame (ORF) encoding a polypeptide, VHH, bispecific antibody, multispecific antibody, or CAR of claim described herein; (iii) a stop codon; and (iv) a 3' UTR.
- the mRNA can include a 5’-cap.
- the mRNA can also include a poly-A region.
- compositions including an mRNA encoding a polypeptide described herein (e.g., a polypeptide comprising an anti-c-MET VHH, a bivalent c-MET VHH-IgG4PAA, a bispecific antibody comprising an anti-C-MET VHH, a multispecific antibody comprising an anti-C-MET VHH, a CAR comprising an anti-C-MET VHH).
- the mRNAs include (i) a 5’-cap structure; (ii) a 5’-UTR; (iii) an open reading frame encoding the polypeptide; (iv) a 3’-untranslated region (3’-UTR); and (v) a poly-A region.
- mRNA: 5’-cap The 5'-cap structure of an mRNA is involved in nuclear export, increasing mRNA stability and binds the mRNA Cap Binding Protein (CBP), which is responsible for mRNA stability in the cell and translation competency through the association of CBP with poly(A) binding protein to form the mature cyclic mRNA species.
- CBP mRNA Cap Binding Protein
- the cap further assists the removal of 5' proximal introns removal during mRNA splicing.
- mRNA molecules may be 5'-end capped generating a 5'-ppp-5'-triphosphate linkage between a terminal guanosine cap residue and the 5'-terminal transcribed sense nucleotide of the mRNA.
- This 5'-guanylate cap may then be methylated to generate an N7-methyl-guanylate residue.
- nucleotides may be used during the capping reaction.
- a Vaccinia Capping Enzyme from New England Biolabs (Ipswich, MA) may be used with ⁇ -thio-guanosine nucleotides according to the manufacturer’s instructions to create a phosphorothioate linkage in the 5'-ppp-5' cap.
- Additional alternative guanosine nucleotides may be used such as ⁇ -methyl-phosphonate and seleno-phosphate nucleotides.
- the Anti-Reverse Cap Analog (ARCA) cap contains two guanosines linked by a 5'-5'-triphosphate group, wherein one guanosine contains an N7 methyl group as well as a 3'-O-methyl group (i.e., N7,3'-O-dimethyl-guanosine-5'-triphosphate-5'- guanosine (m 7 G-3'mppp-G; which may equivalently be designated 3' O-Me- m7G(5')ppp(5')G)).
- N7,3'-O-dimethyl-guanosine-5'-triphosphate-5'- guanosine m 7 G-3'mppp-G; which may equivalently be designated 3' O-Me- m7G(5')ppp(5')G
- Non- limiting examples of a N7-(4-chlorophenoxyethyl) substituted dinucleotide form of a cap analog include a N7-(4-chlorophenoxyethyl)-G(5’)ppp(5’)G and a N7-(4- chlorophenoxyethyl)-m 3’-O G(5’)ppp(5’)G cap analog (see, e.g., the various cap analogs and the methods of synthesizing cap analogs described in Kore et al. Bioorganic & Medicinal Chemistry 21:4570-4574 (2013); the contents of which are herein incorporated by reference in its entirety).
- a 5' terminal cap may include a guanosine analog.
- Useful guanosine analogs include inosine, N1-methyl- guanosine, 2'-fluoro-guanosine, 7-deaza-guanosine, 8-oxo-guanosine, 2-amino- guanosine, LNA-guanosine, and 2-azido-guanosine.
- the 5’ terminal cap is or comprises m7GpppGm.
- the nucleic acids described herein may contain a modified 5’- cap. A modification on the 5’-cap may increase the stability of mRNA, increase the half- life of the mRNA, and could increase the mRNA translational efficiency.
- the modified 5’-cap may include, but is not limited to, one or more of the following modifications: modification at the 2’ and/or 3’ position of a capped guanosine triphosphate (GTP), a replacement of the sugar ring oxygen (that produced the carbocyclic ring) with a methylene moiety (CH 2 ), a modification at the triphosphate bridge moiety of the cap structure, or a modification at the nucleobase (G) moiety.
- GTP capped guanosine triphosphate
- CH 2 methylene moiety
- G nucleobase
- UTRs Untranslated Regions
- UTRs Untranslated regions
- UTRs are nucleic acid sections of a polynucleotide before a start codon (5' UTR) and after a stop codon (3' UTR) that are not translated.
- a polynucleotide e.g., a ribonucleic acid (RNA), e.g., a messenger RNA (mRNA)
- RNA ribonucleic acid
- mRNA messenger RNA
- ORF open reading frame
- UTR e.g., a 5'UTR or functional fragment thereof, a 3'UTR or functional fragment thereof, or a combination thereof.
- Translation of a polynucleotide comprising an open reading frame encoding a polypeptide can be controlled and regulated by a variety of mechanisms that are provided by various cis-acting nucleic acid structures.
- RNA elements that form hairpins or other higher-order (e.g., pseudoknot) intramolecular mRNA secondary structures can provide a translational regulatory activity to a polynucleotide, wherein the RNA element influences or modulates the initiation of polynucleotide translation, particularly when the RNA element is positioned in the 5' UTR close to the 5’-cap structure (Pelletier and Sonenberg (1985) Cell 40(3):515-526; Kozak (1986) Proc Natl Acad Sci 83:2850-2854).
- Cis-acting RNA elements can also affect translation elongation, being involved in numerous frameshifting events (Namy et al., (2004) Mol Cell 13(2):157-168).
- Internal ribosome entry sequences represent another type of cis-acting RNA element that are typically located in 5' UTRs, but have also been reported to be found within the coding region of naturally-occurring mRNAs (Holcik et al. (2000) Trends Genet 16(10):469-473).
- Naturally-occurring uORFs occur singularly or multiply within the 5' UTRs of numerous mRNAs and influence the translation of the downstream major ORF, usually negatively (with the notable exception of GCN4 mRNA in yeast and ATF4 mRNA in mammals, where uORFs serve to promote the translation of the downstream major ORF under conditions of increased eIF2 phosphorylation (Hinnebusch (2005) Annu Rev Microbiol 59:407-450)).
- exemplary translational regulatory activities provided by components, structures, elements, motifs, and/or specific sequences comprising polynucleotides (e.g., mRNA) include, but are not limited to, mRNA stabilization or destabilization (Baker & Parker (2004) Curr Opin Cell Biol 16(3):293-299), translational activation (Villalba et al., (2011) Curr Opin Genet Dev 21(4):452-457), and translational repression (Blumer et al., (2002) Mech Dev 110(1-2):97-112).
- RNA elements can confer their respective functions when used to modify, by incorporation into, heterologous polynucleotides (Goldberg-Cohen et al., (2002) J Biol Chem 277(16):13635-13640).
- the present disclosure provides synthetic polynucleotides comprising a modification (e.g., an RNA element), wherein the modification provides a desired translational regulatory activity.
- the disclosure provides a polynucleotide comprising a 5’ untranslated region (UTR), an initiation codon, a full open reading frame encoding a polypeptide, a 3’ UTR, and at least one modification, wherein the at least one modification provides a desired translational regulatory activity, for example, a modification that promotes and/or enhances the translational fidelity of mRNA translation.
- the desired translational regulatory activity is a cis-acting regulatory activity.
- the desired translational regulatory activity is an increase in the residence time of the 43 S pre-initiation complex (PIC) or ribosome at, or proximal to, the initiation codon.
- the desired translational regulatory activity is an increase in the initiation of polypeptide synthesis at or from the initiation codon. In some instances, the desired translational regulatory activity is an increase in the amount of polypeptide translated from the full open reading frame. In some instances, the desired translational regulatory activity is an increase in the fidelity of initiation codon decoding by the PIC or ribosome. In some instances, the desired translational regulatory activity is inhibition or reduction of leaky scanning by the PIC or ribosome. In some instances, the desired translational regulatory activity is a decrease in the rate of decoding the initiation codon by the PIC or ribosome.
- the desired translational regulatory activity is inhibition or reduction in the initiation of polypeptide synthesis at any codon within the mRNA other than the initiation codon. In some instances, the desired translational regulatory activity is inhibition or reduction of the amount of polypeptide translated from any open reading frame within the mRNA other than the full open reading frame. In some instances, the desired translational regulatory activity is inhibition or reduction in the production of aberrant translation products. In some instances, the desired translational regulatory activity is a combination of one or more of the foregoing translational regulatory activities.
- the present disclosure provides a polynucleotide, e.g., an mRNA, comprising an RNA element that comprises a sequence and/or an RNA secondary structure(s) that provides a desired translational regulatory activity as described herein.
- the mRNA comprises an RNA element that comprises a sequence and/or an RNA secondary structure(s) that promotes and/or enhances the translational fidelity of mRNA translation.
- the mRNA comprises an RNA element that comprises a sequence and/or an RNA secondary structure(s) that provides a desired translational regulatory activity, such as inhibiting and/or reducing leaky scanning.
- the disclosure provides an mRNA that comprises an RNA element that comprises a sequence and/or an RNA secondary structure(s) that inhibits and/or reduces leaky scanning thereby promoting the translational fidelity of the mRNA.
- the RNA element comprises natural and/or modified nucleotides. In some instances, the RNA element comprises of a sequence of linked nucleotides, or derivatives or analogs thereof that provides a desired translational regulatory activity as described herein. In some instances, the RNA element comprises a sequence of linked nucleotides, or derivatives or analogs thereof that forms or folds into a stable RNA secondary structure, wherein the RNA secondary structure provides a desired translational regulatory activity as described herein.
- RNA elements can be identified and/or characterized based on the primary sequence of the element (e.g., GC-rich element), by RNA secondary structure formed by the element (e.g.
- RNA molecules e.g., located within the 5’ UTR of an mRNA
- RNA molecule e.g., located within the 5’ UTR of an mRNA
- biological function and/or activity of the element e.g., “translational enhancer element”
- the GC- rich RNA element is located about 30, about 25, about 20, about 15, about 10, about 5, about 4, about 3, about 2, or about 1 nucleotide(s) upstream of a Kozak consensus sequence in the 5’ UTR of the mRNA. In another embodiment, the GC-rich RNA element is located 15-30, 15-20, 15-25, 10-15, or 5-10 nucleotides upstream of a Kozak consensus sequence. In another embodiment, the GC-rich RNA element is located immediately adjacent to a Kozak consensus sequence in the 5’ UTR of the mRNA.
- the disclosure provides a GC-rich RNA element which comprises a sequence of 3-30, 5-25, 10-20, 15-20, about 20, about 15, about 12, about 10, about 7, about 6 or about 3 nucleotides, derivatives or analogs thereof, linked in any order, wherein the sequence composition is 70-80% cytosine, 60- 70% cytosine, 50%-60% cytosine, 40-50% cytosine, 30-40% cytosine bases.
- the disclosure provides a GC-rich RNA element which comprises a sequence of 3-30, 5-25, 10-20, 15-20, about 20, about 15, about 12, about 10, about 7, about 6 or about 3 nucleotides, derivatives or analogs thereof, linked in any order, wherein the sequence composition is about 80% cytosine, about 70% cytosine, about 60% cytosine, about 50% cytosine, about 40% cytosine, or about 30% cytosine.
- the disclosure provides a GC-rich RNA element which comprises a sequence of 20, 19, 18, 17, 16, 15, 14, 13, 12, 11, 10, 9, 8, 7, 6, 5, 4, or 3 nucleotides, or derivatives or analogs thereof, linked in any order, wherein the sequence composition is 70-80% cytosine, 60-70% cytosine, 50%-60% cytosine, 40- 50% cytosine, or 30-40% cytosine.
- the disclosure provides a GC-rich RNA element which comprises a sequence of 20, 19, 18, 17, 16, 15, 14, 13, 12, 11, 10, 9, 8, 7, 6, 5, 4, or 3 nucleotides, or derivatives or analogs thereof, linked in any order, wherein the sequence composition is about 80% cytosine, about 70% cytosine, about 60% cytosine, about 50% cytosine, about 40% cytosine, or about 30% cytosine.
- the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising a sequence of linked nucleotides, or derivatives or analogs thereof, preceding a Kozak consensus sequence in a 5’ UTR of the mRNA, wherein the GC-rich RNA element is located about 30, about 25, about 20, about 15, about 10, about 5, about 4, about 3, about 2, or about 1 nucleotide(s) upstream of a Kozak consensus sequence in the 5’ UTR of the mRNA, and wherein the GC-rich RNA element comprises a sequence of 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 nucleotides, or derivatives or analogs thereof, linked in any order, wherein the sequence composition is >50% cytosine.
- at least one modification is a GC-rich RNA element comprising a sequence of linked nucleotides, or derivatives or analogs thereof, preceding a Kozak consensus sequence in
- the sequence composition is >55% cytosine, >60% cytosine, >65% cytosine, >70% cytosine, >75% cytosine, >80% cytosine, >85% cytosine, or >90% cytosine.
- the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising a sequence of linked nucleotides, or derivatives or analogs thereof, preceding a Kozak consensus sequence in a 5’ UTR of the mRNA, wherein the GC-rich RNA element comprises any one of the sequences set forth in Table 8.
- the GC-rich RNA element is located about 30, about 25, about 20, about 15, about 10, about 5, about 4, about 3, about 2, or about 1 nucleotide(s) upstream of a Kozak consensus sequence in the 5’ UTR of the mRNA. In another embodiment, the GC-rich RNA element is located about 15-30, 15-20, 15-25, 10-15, or 5-10 nucleotides upstream of a Kozak consensus sequence. In another embodiment, the GC-rich RNA element is located immediately adjacent to a Kozak consensus sequence in the 5’ UTR of the mRNA.
- the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising the sequence V1 [CCCCGGCGCC (SEQ ID NO:650)] as set forth in Table 8, or derivatives or analogs thereof, preceding a Kozak consensus sequence in the 5’ UTR of the mRNA.
- the GC-rich element comprises the sequence V1 as set forth in Table 8 located immediately adjacent to and upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA.
- the GC-rich element comprises the sequence V1 as set forth in Table 8 located 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10 bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA. In other instances, the GC-rich element comprises the sequence V1 as set forth in Table 8 located 1-3, 3-5, 5-7, 7-9, 9-12, or 12-15 bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA.
- the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising the sequence V2 [CCCCGGC] as set forth in Table 8, or derivatives or analogs thereof, preceding a Kozak consensus sequence in the 5’ UTR of the mRNA.
- the GC-rich element comprises the sequence V2 as set forth in Table 8 located immediately adjacent to and upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA.
- the GC-rich element comprises the sequence V2 as set forth in Table 8 located 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10 bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA.
- the GC-rich element comprises the sequence V2 as set forth in Table 8 located 1-3, 3-5, 5-7, 7-9, 9-12, or 12- 15 bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA.
- the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising the sequence EK [GCCGCC] as set forth in Table 8, or derivatives or analogs thereof, preceding a Kozak consensus sequence in the 5’ UTR of the mRNA.
- the GC-rich element comprises the sequence EK as set forth in Table 8 located immediately adjacent to and upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA.
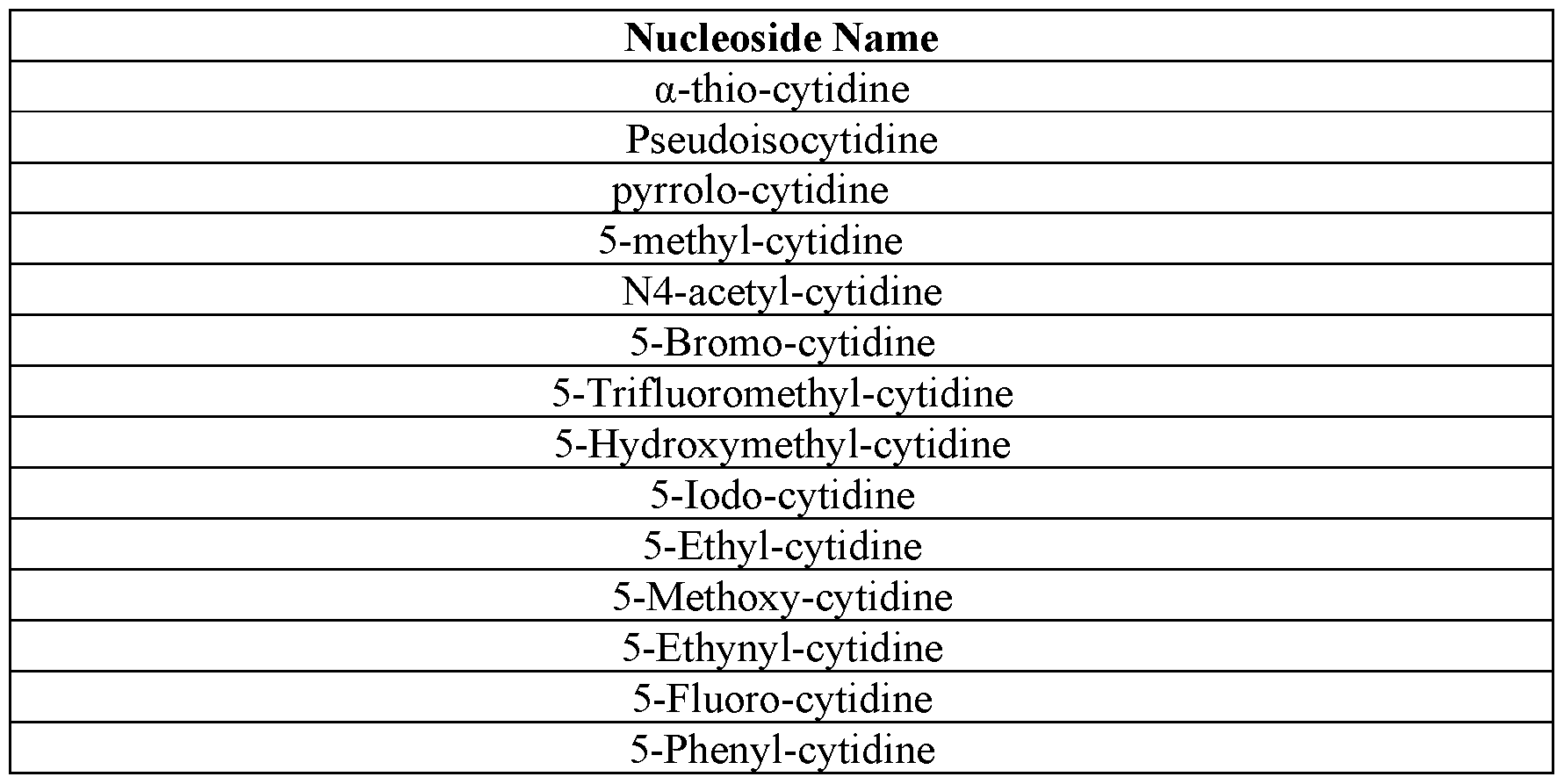
- the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5- hydroxymethyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5- bromo-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5-iodo-cytosine, and cytosine as the only uracils and cytosines.
- the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5-methoxy-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5- methoxy-uracil, uracil, 5-ethyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5- phenyl-cytosine, and cytosine as the only uracils and cytosines.
- the polynucleotides of the disclosure contain 1-methyl- pseudouracil, uracil, 5-phenyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-ethnyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, N4-methyl-cytosine, and cytosine as the only uracils and cytosines.
- the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-fluoro- cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, N4-acetyl- cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, pseudoisocytosine, and cytosine as the only uracils and cytosines.
- the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-formyl- cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-aminoallyl- cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-carboxy- cytosine, and cytosine as the only uracils and cytosines.
- the polynucleotides of the disclosure contain 5-methoxy- uridine, uridine, 5-methyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-trifluoromethyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5- hydroxymethyl-cytidine, and cytidine as the only uridines and cytidines.
- the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5- bromo-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-iodo-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-methoxy-cytidine, and cytidine as the only uridines and cytidines.
- the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-ethyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5- methoxy-uridine, uridine, 5-phenyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy- uridine, uridine, 5-ethnyl-cytidine, and cytidine as the only uridines and cytidines.
- the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, N4-methyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-fluoro- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, N4-acetyl-cytidine, and cytidine as the only uridines and cytidines.
- the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, pseudoisocytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-formyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5- methoxy-uridine, uridine, 5-aminoallyl-cytidine, and cytidine as the only uridines and cytidines.
- the polynucleotides of the disclosure contain 5-methoxy- uridine, uridine, 5-carboxy-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl- pseudouridine, uridine, 5-methyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-trifluoromethyl-cytidine, and cytidine as the only uridines and cytidines.
- the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-hydroxymethyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-bromo-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-iodo-cytidine, and cytidine as the only uridines and cytidines.
- the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-ethnyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, N4-methyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-fluoro- cytidine, and cytidine as the only uridines and cytidines.
- the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, N4-acetyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, pseudoisocytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-formyl- cytidine, and cytidine as the only uridines and cytidines.
- the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-aminoallyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-carboxy- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain the uracil of one of the nucleosides of Table 9 and uracil as the only uracils.
- the polynucleotides of the disclosure contain a uridine of Table 9 and uridine as the only uridines.
- Table 9 – Exemplary Uracil-containing Nucleosides the polynucleotides of the disclosure contain the cytosine of one of the nucleosides of Table 10 and cytosine as the only cytosines.
- the polynucleotides of the disclosure contain a cytidine of Table 10 and cytidine as the only cytidines.
- the alternative nucleotides which may be incorporated into a polynucleotide molecule, can be altered on the internucleoside linkage (e.g., phosphate backbone).
- the phrases “phosphate” and “phosphodiester” are used interchangeably.
- Backbone phosphate groups can be altered by replacing one or more of the oxygen atoms with a different substituent.
- the alternative nucleosides and nucleotides can include the wholesale replacement of an unaltered phosphate moiety with another internucleoside linkage as described herein.
- alternative phosphate groups include, but are not limited to, phosphorothioate, phosphoroselenates, boranophosphates, boranophosphate esters, hydrogen phosphonates, phosphoramidates, phosphorodiamidates, alkyl or aryl phosphonates, and phosphotriesters.
- Phosphorodithioates have both non-linking oxygens replaced by sulfur.
- the phosphate linker can also be altered by the replacement of a linking oxygen with nitrogen (bridged phosphoramidates), sulfur (bridged phosphorothioates), and carbon (bridged methylene-phosphonates).
- the alternative nucleosides and nucleotides can include the replacement of one or more of the non-bridging oxygens with a borane moiety (BH3), sulfur (thio), methyl, ethyl and/or methoxy.
- BH3 borane moiety
- sulfur (thio) sulfur (thio)
- methyl ethyl
- methoxy ethoxy of two non-bridging oxygens at the same position
- two non-bridging oxygens at the same position e.g., the alpha ( ⁇ ), beta ( ⁇ ) or gamma ( ⁇ ) position
- a sulfur (thio) and a methoxy e.g., the alpha ( ⁇ ), beta ( ⁇ ) or gamma ( ⁇ ) position
- the replacement of one or more of the oxygen atoms at the ⁇ position of the phosphate moiety is provided to confer stability (such as against exonucleases and endonucleases) to RNA and DNA through the unnatural phosphorothioate backbone linkages.
- Phosphorothioate DNA and RNA have increased nuclease resistance and subsequently a longer half-life in a cellular environment. While not wishing to be bound by theory, phosphorothioate linked polynucleotide molecules are expected to also reduce the innate immune response through weaker binding/activation of cellular innate immune molecules.
- an alternative nucleoside includes an alpha-thio-nucleoside (e.g., 5'-O-(1-thiophosphate)-adenosine, 5'-O-(1-thiophosphate)-cytidine ( ⁇ -thio- cytidine), 5'-O-(1-thiophosphate)-guanosine, 5'-O-(1-thiophosphate)-uridine, or 5'-O-(1- thiophosphate)-pseudouridine).
- alpha-thio-nucleoside e.g., 5'-O-(1-thiophosphate)-adenosine, 5'-O-(1-thiophosphate)-cytidine ( ⁇ -thio- cytidine), 5'-O-(1-thiophosphate)-guanosine, 5'-O-(1-thiophosphate)-uridine, or 5'-O-(1- thiophosphate
- the polynucleotides of the disclosure can include a combination of alterations to the sugar, the nucleobase, and/or the internucleoside linkage. These combinations can include any one or more alterations described herein.
- mRNA Poly-A tail During RNA processing, a long chain of adenosine nucleotides (poly(A) tail) is normally added to mRNA molecules to increase the stability of the mRNA. Immediately after transcription, the 3' end of the transcript is cleaved to free a 3' hydroxyl. Then poly(A) polymerase adds a chain of adenosine nucleotides to the RNA.
- polyadenylation adds a poly-A tail that is between 100 and 250 residues long.
- Methods for the stabilization of RNA by incorporation of chain-terminating nucleosides at the 3 ’-terminus include those described in International Patent Publication No. WO2013/103659, incorporated herein in its entirety.
- Poly (A) tail deadenylation by 3' exonucleases is a key step in cellular mRNA degradation in eukaryotes. By blocking 3' exonucleases, the functional half-life of mRNA can be increased, resulting in increased protein expression.
- Chemical and enzymatic ligation strategies to modify the 3' end of mRNA with reverse chirality adenosine (LA10) and/or inverted deoxythymidine (IdT) are known to those of skill in the art and have been demonstrated to extend mRNA half-life in cellular and in vivo studies.
- the poly(A)tail of the mRNA includes a 3’ LA10 or IdT modification. For example, as described in International Patent Publication No. WO2017/049275, the tail modifications of which are incorporated by reference in their entirety.
- Additional strategies have been explored to further stabilize mRNA, including: chemical modification of the 3’ nucleotide (e.g., conjugation of a morpholino to the 3’ end of the poly(A)tail); incorporation of stabilizing sequences after the poly(A) tail (e.g., a co-polymer, a stem-loop, or a triple helix); and/or annealing of structured oligos to the 3' end of an mRNA, as described, for example, in International Patent Publication No. WO2017/049286, the stabilized linkages of which are incorporated by reference in their entirety.
- chemical modification of the 3’ nucleotide e.g., conjugation of a morpholino to the 3’ end of the poly(A)tail
- stabilizing sequences after the poly(A) tail e.g., a co-polymer, a stem-loop, or a triple helix
- annealing of structured oligos to the 3' end
- Annealing an oligonucleotide e.g., an oligonucleotide conjugate
- a complex secondary structure e.g., a triple-helix structure or a stem-loop structure
- the length of a poly(A) tail of the present disclosure is greater than 30 nucleotides in length. In some instances, the poly(A) tail is greater than 35 nucleotides in length. In some instances, the length is at least 40 nucleotides. In another embodiment, the length is at least 45 nucleotides. In some instances, the length is at least 50 nucleotides. In some instances, the length is at least 55 nucleotides. In another embodiment, the length is at least 60 nucleotides. In another embodiment, the length is at least 65 nucleotides. In another embodiment, the length is at least 70 nucleotides.
- the length is at least 80 nucleotides. In some instances, the length is at least 90 nucleotides. In some instances, the length is at least 100 nucleotides. In some instances, the length is at least 120 nucleotides. In some instances, the length is at least 140 nucleotides. In some instances, the length is at least 160 nucleotides. In some instances, the length is at least 180 nucleotides. In some instances, the length is at least 200 nucleotides. In some instances, the length is at least 250 nucleotides. In some instances, the length is at least 300 nucleotides. In some instances, the length is at least 350 nucleotides. In some instances, the length is at least 400 nucleotides.
- the length is at least 450 nucleotides. In some instances, the length is at least 500 nucleotides. In some instances, the length is at least 600 nucleotides. In some instances, the length is at least 700 nucleotides. In some instances, the length is at least 800 nucleotides. In some instances, the length is at least 900 nucleotides. In some instances, the length is at least 1000 nucleotides. In some instances, the length is at least 1100 nucleotides. In some instances, the length is at least 1200 nucleotides. In some instances, the length is at least 1300 nucleotides. In some instances, the length is at least 1400 nucleotides.
- the length is at least 1500 nucleotides. In some instances, the length is at least 1600 nucleotides. In some instances, the length is at least 1700 nucleotides. In some instances, the length is at least 1800 nucleotides. In some instances, the length is at least 1900 nucleotides. In some instances, the length is at least 2000 nucleotides. In some instances, the length is at least 2500 nucleotides. In some instances, the length is at least 3000 nucleotides.
- the poly(A) tail may be 80 nucleotides, 120 nucleotides, or 160 nucleotides in length. In some instances, the poly(A) tail may be 20, 40, 80, 100, 120, 140 or 160 nucleotides in length. In some instances, the poly(A) tail is designed relative to the length of the mRNA. This design may be based on the length of the coding region of the mRNA, the length of a particular feature or region of the mRNA, or based on the length of the ultimate product expressed from the RNA.
- poly(A) tail When relative to any additional feature of the RNA (e.g., other than the mRNA portion which includes the poly(A) tail), poly(A) tail may be 10, 20, 30, 40, 50, 60, 70, 80, 90 or 100% greater in length than the additional feature.
- the poly(A) tail may also be designed as a fraction of the mRNA to which it belongs. In this context, the poly(A) tail may be 10, 20, 30, 40, 50, 60, 70, 80, or 90% or more of the total length of the construct or the total length of the construct minus the poly(A) tail.
- engineered binding sites and/or the conjugation of nucleic acids or mRNA for poly(A) binding protein may be used to enhance expression.
- the mRNA which includes a poly(A) tail or a poly-A-G quartet may be stabilized by an alteration to the 3’region of the nucleic acid that can prevent and/or inhibit the addition of oligo(U) (see, e.g., International Patent Publication No. WO2013/103659, incorporated herein by reference in its entirety).
- the mRNA which includes a poly(A) tail or a poly- A-G quartet, may be stabilized by the addition of an oligonucleotide that terminates in a 3’-deoxynucleoside, 2’,3’-dideoxynucleoside 3'-O-methylnucleosides, 3'-O- ethylnucleosides, 3'-arabinosides, and other alternative nucleosides known in the art and/or described herein Synthesis of Polynucleotides
- the polynucleotide molecules e.g., mRNA
- for use in accordance with the disclosure may be prepared according to any useful technique, as described herein.
- product formation can be monitored by spectroscopic means, such as nuclear magnetic resonance spectroscopy (e.g., 1 H or 13 C) infrared spectroscopy, spectrophotometry (e.g., UV-visible), or mass spectrometry, or by chromatography (e.g., high performance liquid chromatography (HPLC) or thin layer chromatography).
- spectroscopic means such as nuclear magnetic resonance spectroscopy (e.g., 1 H or 13 C) infrared spectroscopy, spectrophotometry (e.g., UV-visible), or mass spectrometry, or by chromatography (e.g., high performance liquid chromatography (HPLC) or thin layer chromatography).
- spectroscopic means such as nuclear magnetic resonance spectroscopy (e.g., 1 H or 13 C) infrared spectroscopy, spectrophotometry (e.g., UV-visible), or mass spectrometry, or by chromatography (e.g., high performance
- the reactions of the processes described herein can be carried out in suitable solvents, which can be readily selected by one of skill in the art of organic synthesis. Suitable solvents can be substantially nonreactive with the starting materials (reactants), the intermediates, or products at the temperatures at which the reactions are carried out (i.e., temperatures which can range from the solvent’s freezing temperature to the solvent’s boiling temperature). A given reaction can be carried out in one solvent or a mixture of more than one solvent. Depending on the particular reaction step, suitable solvents for a particular reaction step can be selected.
- Resolution of racemic mixtures of alternative polynucleotides or nucleic acids can be carried out by any of numerous methods known in the art.
- An example method includes fractional recrystallization using a “chiral resolving acid” which is an optically active, salt-forming organic acid.
- Suitable resolving agents for fractional recrystallization methods are, for example, optically active acids, such as the D and L forms of tartaric acid, acid, dibenzoyltartaric acid, mandelic acid, malic acid, lactic acid or the various optically active camphorsulfonic acids.
- Resolution of racemic mixtures can also be carried out by elution on a column packed with an optically active resolving agent (e.g., dinitrobenzoylphenylglycine).
- an optically active resolving agent e.g., dinitrobenzoylphenylglycine
- Suitable elution solvent composition can be determined by one skilled in the art.
- Alternative nucleosides and nucleotides e.g., building block molecules
- nucleosides and nucleotides can be prepared according to the synthetic methods described in Ogata et al., J. Org. Chem. 74:2585-2588 (2009); Purmal et al., Nucl.
- polynucleotide includes one or more alternative nucleosides or nucleotides, the polynucleotides of the disclosure may or may not be uniformly altered along the entire length of the molecule.
- nucleotide e.g., purine or pyrimidine, or any one or more or all of A, G, U, C
- nucleotide may or may not be uniformly altered in a polynucleotide of the disclosure, or in a given predetermined sequence region thereof.
- all nucleotides X in a polynucleotide of the disclosure (or in a given sequence region thereof) are altered, wherein X may any one of nucleotides A, G, U, C, or any one of the combinations A+G, A+U, A+C, G+U, G+C, U+C, A+G+U, A+G+C, G+U+C or A+G+C.
- nucleotide analogs or other alteration(s) may be located at any position(s) of a polynucleotide such that the function of the polynucleotide is not substantially decreased.
- An alteration may also be a 5' or 3' terminal alteration.
- the polynucleotide may contain from 1% to 100% alternative nucleosides, nucleotides, or internucleoside linkages (either in relation to overall nucleotide content, or in relation to one or more types of nucleotide, i.e.
- any one or more of A, G, U or C) or any intervening percentage e.g., from 1% to 20%, from 1% to 25%, from 1% to 50%, from 1% to 60%, from 1% to 70%, from 1% to 80%, from 1% to 90%, from 1% to 95%, from 10% to 20%, from 10% to 25%, from 10% to 50%, from 10% to 60%, from 10% to 70%, from 10% to 80%, from 10% to 90%, from 10% to 95%, from 10% to 100%, from 20% to 25%, from 20% to 50%, from 20% to 60%, from 20% to 70%, from 20% to 80%, from 20% to 90%, from 20% to 95%, from 20% to 100%, from 50% to 60%, from 50% to 70%, from 50% to 80%, from 50% to 90%, from 50% to 95%, from 50% to 100%, from 70% to 80%, from 70% to 90%, from 70% to 95%, from 70% to 100%, from 80% to 90%, from 80% to 95%, from 90% to 100%, and from 95% to 100.
- any intervening percentage e.g.,
- the remaining percentage necessary to total 100% is accounted for by the corresponding natural nucleoside, nucleotide, or internucleoside linkage. In other embodiments, the remaining percentage necessary to total 100% is accounted for by a second alternative nucleoside, nucleotide, or internucleoside linkage.
- the present disclosure also provides LNP compositions.
- the lipid nanoparticle compositions described herein may be used for the delivery of therapeutic and/or prophylactic agents, e.g., mRNAs, to mammalian cells or organs.
- the lipid nanoparticles described herein have little or no immunogenicity.
- the lipid compounds disclosed herein have a lower immunogenicity as compared to a reference lipid (e.g, MC3, KC2, or DLinDMA).
- a formulation comprising a lipid disclosed herein and a therapeutic or prophylactic agent, e.g, mRNA, has an increased therapeutic index as compared to a corresponding formulation which comprises a reference lipid (e.g, MC3, KC2, or DLinDMA) and the same therapeutic or prophylactic agent.
- a reference lipid e.g, MC3, KC2, or DLinDMA
- compositions comprising: (a) a delivery agent comprising a lipid nanoparticle; and (b) a polynucleotide encoding a VHH, polypeptide, bispecific antibody, multispecific antibody, or CAR of the disclosure.
- polynucleotides of the present disclosure are included in a lipid nanoparticle (LNP).
- Lipid nanoparticles according to the present disclosure may comprise: (i) an ionizable lipid (e.g, an ionizable amino lipid; (ii) a sterol or other structural lipid; (iii) a non-cationic helper lipid or phospholipid; and (iv) a PEG- modified lipid.
- lipid nanoparticles according to the present disclosure further comprise one or more polynucleotides of the present disclosure (e.g., mRNA).
- the lipid nanoparticles according to the present disclosure can be generated using components, compositions, and methods as are generally known in the art, see, for example PCT/US2016/052352; PCT/US2016/068300; PCT/US2017/037551; PCT/US2015/027400; PCT/US2016/047406; PCT/US2016000129; PCT/US2016/014280; PCT/US2016/014280; PCT/US2017/038426; PCT/US2014/027077; PCT/US2014/055394; PCT/US2016/52117; PCT/US2012/069610; PCT/US2017/027492; PCT/US2016/059575 and PCT/US2016/069491 all of which are incorporated by reference herein in their entirety.
- the lipid nanoparticle comprises: (i) 20 to 60 mol.% ionizable cationic lipid (e.g., ionizable amino lipid), (ii) 25 to 55 mol.% sterol or other structural lipid, (iii) 5 to 25 mol.% non-cationic lipid (e.g., phospholipid), and (iv) 0.5 to 15 mol.% PEG-modified lipid.
- ionizable cationic lipid e.g., ionizable amino lipid
- sterol or other structural lipid e.g., sterol or other structural lipid
- non-cationic lipid e.g., phospholipid
- iv 0.5 to 15 mol.% PEG-modified lipid.
- the lipid nanoparticle comprises: (i) 40 to 50 mol.% ionizable cationic lipid (e.g., ionizable amino lipid), (ii) 30 to 45 mol.% sterol or other structural lipid, (iii) 5 to 15 mol.% non-cationic lipid (e.g., phospholipid), and (iv) 1 to 5 mol.% PEG-modified lipid.
- ionizable cationic lipid e.g., ionizable amino lipid
- sterol or other structural lipid e.g., sterol or other structural lipid
- 5 to 15 mol.% non-cationic lipid e.g., phospholipid
- 1 to 5 mol.% PEG-modified lipid e.g., PEG-modified lipid.
- the lipid nanoparticle comprises: (i) 45 to 50 mol.% ionizable cationic lipid (e.g., ionizable amino lipid), (ii) 35 to 45 mol.% sterol or other structural lipid, (iii) 8 to 12 mol.% non-cationic lipid (e.g., phospholipid), and (iv) 1.5 to 3.5 mol.% PEG-modified lipid.
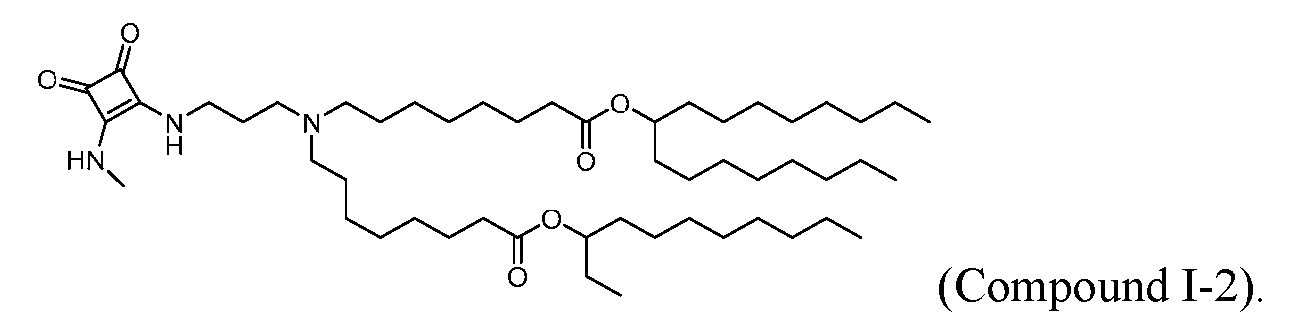
- “Compounds” numbered with an “I-” prefix e.g., “Compound I-1,” “Compound I-2,” “Compound I-3,” “Compound I-VI,” etc., indicate specific ionizable lipid compounds.
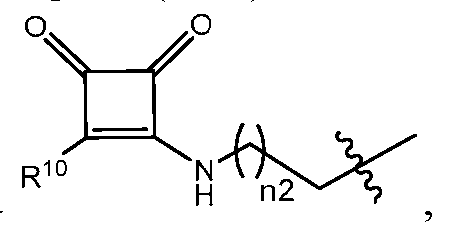
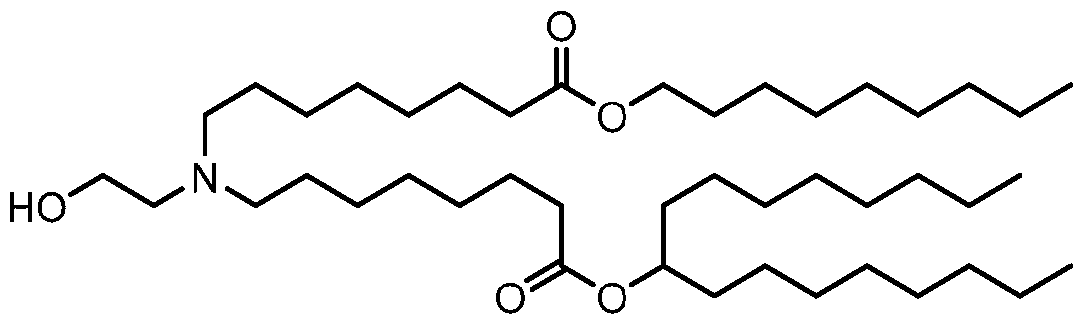
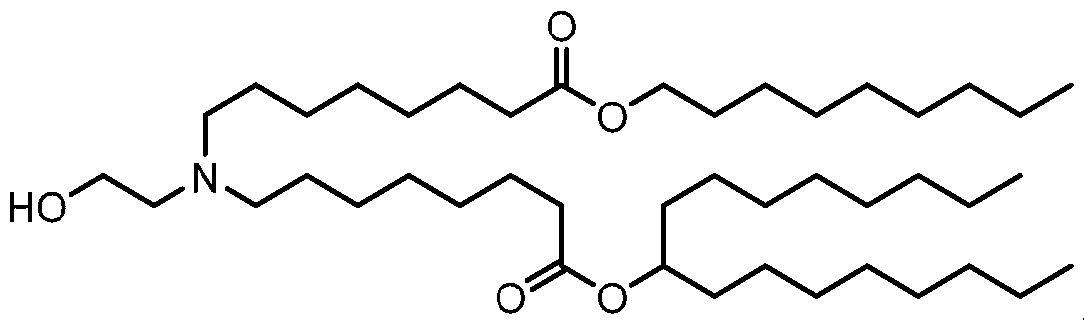
- the lipid nanoparticle of the present disclosure comprises an ionizable cationic lipid (e.g., an ionizable amino lipid) that is a compound of Formula (I): or its N-oxide, or a salt or isomer thereof, wherein R’ a is R’ branched ; wherein R’ branched is: wherein denotes a point of attachment; wherein R a ⁇ , R a ⁇ , R a ⁇ , and R a ⁇ are each independently selected from the group consisting of H, C 2-12 alkyl, and C 2-12 alkenyl; R 2 and R 3 are each independently selected from the group consisting of C 1-14 alkyl and C 2-14 alkenyl; R 4 is selected from the group consisting of -(CH
- R’ a is R’ branched ;
- R’ branched is denotes a point of attachment;
- R a ⁇ , R a ⁇ , R a ⁇ , and R a ⁇ are each H;
- R 2 and R 3 are each C 1-14 alkyl;
- R 4 is -(CH 2 ) n OH; n is 2;
- each R 5 is H;
- each R 6 is H;
- M and M’ are each - C(O)O-;
- R’ is a C 1-12 alkyl; l is 5; and
- m is 7.
- R’ a is R’ branched ;
- R’ branched is ; denotes a point a ⁇ a ⁇ a ⁇ a ⁇ of attachment;
- R , R , R , and R are each H;
- R 2 and R 3 are each C 1-14 alkyl;
- R 4 is -(CH 2 ) n OH; n is 2;
- each R 5 is H;
- each R 6 is H;
- M and M’ are each -C(O)O-;
- R’ is a C 1-12 alkyl; l is 3; and m is 7.
- R’ a is R’ branched ;
- R’ branched is a ⁇ denotes a point of attachment;
- R is C 2-12 alkyl;
- R a ⁇ , R a ⁇ , and R a ⁇ are each H;
- R 2 and R 3 are each C 1-14 alkyl;
- R 4 is R 10 is NH(C 1-6 alkyl);
- n2 is 2;
- R 5 is H; each R 6 is H;
- M and M’ are each -C(O)O-;
- R’ is a C 1-12 alkyl; l is 5; and
- m is 7.
- R’ a is R’ branched ;
- R’ branched is denotes a point of attachment a ⁇ a ⁇ ;
- R , R , and R a ⁇ are each H;
- R a ⁇ is C 2-12 alkyl;
- R 2 and R 3 are each C 1-14 alkyl;
- R 4 is -(CH 2 )nOH; n is 2;
- each R 5 is H;
- each R 6 is H;
- M and M’ are each -C(O)O-;
- R’ is a C 1-12 alkyl; l is 5; and m is 7.
- the compound of Formula (I) is selected from:
- the compound of Formula (I) is:
- the compound of Formula (I) is:
- the compound of Formula (I) is:
- the lipid composition of the lipid nanoparticle composition disclosed herein can comprise one or more phospholipids, for example, one or more saturated or (poly)unsaturated phospholipids or a combination thereof.
- phospholipids comprise a phospholipid moiety and one or more fatty acid moieties.
- a phospholipid moiety can be selected, for example, from the non-limiting group consisting of phosphatidyl choline, phosphatidyl ethanolamine, phosphatidyl glycerol, phosphatidyl serine, phosphatidic acid, 2-lysophosphatidyl choline, and a sphingomyelin.
- a fatty acid moiety can be selected, for example, from the non-limiting group consisting of lauric acid, myristic acid, myristoleic acid, palmitic acid, palmitoleic acid, stearic acid, oleic acid, linoleic acid, alpha-linolenic acid, erucic acid, phytanoic acid, arachidic acid, arachidonic acid, eicosapentaenoic acid, behenic acid, docosapentaenoic acid, and docosahexaenoic acid.
- Particular phospholipids can facilitate fusion to a membrane.
- a cationic phospholipid can interact with one or more negatively charged phospholipids of a membrane (e.g., a cellular or intracellular membrane). Fusion of a phospholipid to a membrane can allow one or more elements (e.g., a therapeutic agent) of a lipid- containing composition (e.g., LNPs) to pass through the membrane permitting, e.g., delivery of the one or more elements to a target tissue.
- elements e.g., a therapeutic agent
- a lipid- containing composition e.g., LNPs
- Non-natural phospholipid species including natural species with modifications and substitutions including branching, oxidation, cyclization, and alkynes are also contemplated.
- a phospholipid can be functionalized with or cross-linked to one or more alkynes (e.g., an alkenyl group in which one or more double bonds is replaced with a triple bond).
- alkynes e.g., an alkenyl group in which one or more double bonds is replaced with a triple bond.
- an alkyne group can undergo a copper-catalyzed cycloaddition upon exposure to an azide.
- Such reactions can be useful in functionalizing a lipid bilayer of a nanoparticle composition to facilitate membrane permeation or cellular recognition or in conjugating a nanoparticle composition to a useful component such as a targeting or imaging moiety (e.g., a dye).
- Phospholipids include, but are not limited to, glycerophospholipids such as phosphatidylcholines, phosphatidylethanolamines, phosphatidylserines, phosphatidylinositols, phosphatidy glycerols, and phosphatidic acids. Phospholipids also include phosphosphingolipid, such as sphingomyelin.
- a phospholipid of the present disclosure comprises 1,2- distearoyl-sn-glycero-3-phosphocholine (DSPC), 1,2-dioleoyl-sn-glycero-3- phosphoethanolamine (DOPE), 1,2-dilinoleoyl-sn-glycero-3-phosphocholine (DLPC), 1,2-dimyristoyl-sn-gly cero-phosphocholine (DMPC), 1,2-dioleoyl-sn-glycero-3- phosphocholine (DOPC), l,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC), 1,2- diundecanoyl-sn-glycero-phosphocholine (DUPC), 1-palmitoyl-2-oleoyl-sn-glycero-3- phosphocholine (POPC), 1,2-di-O-octadecenyl-sn-glycero-3-phosphocholine (PO
- a phospholipid useful or potentially useful in the present disclosure is an analog or variant of DSPC.
- Alternative Lipids In certain instances, a phospholipid useful or potentially useful in the present disclosure comprises a modified phosphocholine moiety, wherein the alkyl chain linking the quaternary amine to the phosphoryl group is not ethylene (e.g., n is not 2). Therefore, in certain instances, a phospholipid useful.
- an alternative lipid is used in place of a phospholipid of the present disclosure.
- an alternative lipid of the present disclosure is oleic acid. In certain instances, the alternative lipid is one of the following:
- the lipid composition of a pharmaceutical composition disclosed herein can comprise one or more structural lipids.
- structural lipid refers to sterols and also to lipids containing sterol moieties. Incorporation of structural lipids in the lipid nanoparticle may help mitigate aggregation of other lipids in the particle.
- Structural lipids can be selected from the group including but not limited to, cholesterol, fecosterol, sitosterol, ergosterol, campesterol, stigmasterol, brassicasterol, tomatidine, tomatine, ursolic acid, alpha-tocopherol, hopanoids, phytosterols, steroids, and mixtures thereof.
- the structural lipid is a sterol.
- sterols are a subgroup of steroids consisting of steroid alcohols.
- the structural lipid is a steroid.
- the structural lipid is cholesterol.
- the structural lipid is an analog of cholesterol.
- the structural lipid is alpha-tocopherol.
- the structural lipids may be one or more of the structural lipids described in U.S. Application No. 62/520,530.
- Polyethylene Glycol (PEG)-Lipids The lipid composition of a pharmaceutical composition disclosed herein can comprise one or more a polyethylene glycol (PEG) lipid.
- PEG-lipid refers to polyethylene glycol (PEG)- modified lipids.
- PEG-lipids include PEG-modified phosphatidylethanolamine and phosphatidic acid, PEG-ceramide conjugates (e.g., PEG- CerC14 or PEG-CerC20), PEG-modified dialkylamines and PEG-modified 1,2- diacyloxypropan-3-amines.
- PEGylated lipids PEGylated lipids.
- a PEG lipid can be PEG-c-DOMG, PEG-DMG, PEG-DLPE, PEG-DMPE, PEG-DPPC, or a PEG-DSPE lipid.
- the PEG-lipid includes, but not limited to 1,2-dimyristoyl-sn- glycerol methoxypolyethylene glycol (PEG-DMG), 1,2-distearoyl-sn-glycero-3- phosphoethanolamine-N-[amino(polyethylene glycol)] (PEG-DSPE), PEG-disteryl glycerol (PEG-DSG), PEG-dipalmetoleyl, PEG-dioleyl, PEG-distearyl, PEG- diacylglycamide (PEG-DAG), PEG-dipalmitoyl phosphatidylethanolamine (PEG-DPPE), or PEG-l,2-dimyristyloxlpropyl-3-amine (PEG-c-DMA).
- PEG-DMG 1,2-dimyristoyl-sn- glycerol methoxypolyethylene glycol
- PEG-DSPE 1,2-distearoyl-s
- the PEG-lipid is selected from the group consisting of a PEG- modified phosphatidylethanolamine, a PEG-modified phosphatidic acid, a PEG-modified ceramide, a PEG-modified dialkylamine, a PEG-modified diacylglycerol, a PEG- modified dialkylglycerol, and mixtures thereof.
- the lipid moiety of the PEG-lipids includes those having lengths of from about C 14 to about C 22 , preferably from about C 14 to about C 16 .
- a PEG moiety for example an mPEG-NH 2 , has a size of about 1000, 2000, 5000, 10,000, 15,000 or 20,000 daltons.
- the PEG-lipid is PEG 2k -DMG.
- the lipid nanoparticles described herein can comprise a PEG lipid which is a non-diffusible PEG.
- Non-limiting examples of non-diffusible PEGs include PEG-DSG and PEG-DSPE.
- PEG-lipids are known in the art, such as those described in U.S. Patent No. 8,158,601 and International Publ. No. WO 2015/130584 A2, which are incorporated herein by reference in their entirety.
- lipid component of a lipid nanoparticle composition may include one or more molecules comprising polyethylene glycol, such as PEG or PEG-modified lipids. Such species may be alternately referred to as PEGylated lipids.
- a PEG lipid is a lipid modified with polyethylene glycol.
- a PEG lipid may be selected from the non-limiting group including PEG-modified phosphatidylethanolamines, PEG-modified phosphatidic acids, PEG-modified ceramides, PEG-modified dialkylamines, PEG-modified diacylglycerols, PEG-modified dialkylglycerols, and mixtures thereof.
- a PEG lipid may be PEG-c-DOMG, PEG-DMG, PEG-DLPE, PEG-DMPE, PEG-DPPC, or a PEG-DSPE lipid.
- PEG lipids useful in the present disclosure can be PEGylated lipids described in International Publication No.
- PEG lipid is a PEG-OH lipid.
- a “PEG- OH lipid” is a PEGylated lipid having one or more hydroxyl (–OH) groups on the lipid.
- the PEG- OH lipid includes one or more hydroxyl groups on the PEG chain.
- a PEG-OH or hydroxy-PEGylated lipid comprises an —OH group at the terminus of the PEG chain.
- the lipid composition of the pharmaceutical compositions disclosed herein does not comprise a PEG-lipid.
- the PEG-lipids may be one or more of the PEG lipids described in U.S. Application No. 62/520,530.
- a PEG lipid of the present disclosure comprises a PEG- modified phosphatidylethanolamine, a PEG-modified phosphatidic acid, a PEG-modified ceramide, a PEG-modified dialkylamine, a PEG-modified diacylglycerol, a PEG- modified dialkylglycerol, and mixtures thereof.
- the PEG-modified lipid is PEG-DMG, PEG-c-DOMG (also referred to as PEG-DOMG), PEG-DSG and/or PEG-DPG.
- a LNP of the present disclosure comprises an ionizable cationic lipid of Formula I, a phospholipid comprising DSPC, a structural lipid, and a PEG lipid (e.g., 134-hydroxy- 3,6,9,12,15,18,21,24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96,99,102,105,108,111,114,117,120,123,126,129,132- tetratetracontaoxatetratriacontahectyl stearate).
- a PEG lipid e.g., 134-hydroxy- 3,6,9,12,15,18,21,24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96
- a LNP of the present disclosure comprises an ionizable cationic lipid of . In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of , and an alternative lipid comprising oleic acid. In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of , an alternative lipid comprising oleic acid, a structural lipid comprising cholesterol, and a PEG lipid. In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of a phospholipid comprising DOPE, a structural lipid comprising cholesterol, and a PEG lipid. In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of
- a phospholipid comprising DOPE a structural lipid comprising cholesterol, and a PEG lipid.
- a LNP of the present disclosure comprises an N:P ratio of from about 2:1 to about 30:1. In some instances, a LNP of the present disclosure comprises an N:P ratio of about 6:1. In some instances, a LNP of the present disclosure comprises an N:P ratio of about 3:1. In some instances, a LNP of the present disclosure comprises a wt/wt ratio of the ionizable cationic lipid component to the RNA of from about 10:1 to about 100:1.
- a LNP of the present disclosure comprises a wt/wt ratio of the ionizable cationic lipid component to the RNA of about 20:1. In some instances, a LNP of the present disclosure comprises a wt/wt ratio of the ionizable cationic lipid component to the RNA of about 10:1. In some instances, a LNP of the present disclosure has a mean diameter from about 50nm to about 150nm. In some instances, a LNP of the present disclosure has a mean diameter from about 70nm to about 120nm.
- Other Lipid Composition Components The lipid composition of a pharmaceutical composition disclosed herein can include one or more components in addition to those described above.
- the lipid composition can include one or more permeability enhancer molecules, carbohydrates, polymers, surface altering agents (e.g., surfactants), or other components.
- a permeability enhancer molecule can be a molecule described by U.S. Patent Application Publication No.2005/0222064.
- Carbohydrates can include simple sugars (e.g., glucose) and polysaccharides (e.g., glycogen and derivatives and analogs thereof).
- a polymer can be included in and/or used to encapsulate or partially encapsulate a pharmaceutical composition disclosed herein (e.g., a pharmaceutical composition in lipid nanoparticle form).
- a polymer can be biodegradable and/or biocompatible.
- a polymer can be selected from, but is not limited to, polyamines, polyethers, polyamides, polyesters, polycarbamates, polyureas, polycarbonates, polystyrenes, polyimides, polysulfones, polyurethanes, polyacetylenes, polyethylenes, polyethyleneimines, polyisocyanates, polyacrylates, polymethacrylates, polyacrylonitriles, and polyarylates.
- the ratio between the lipid composition and the polynucleotide range can be from about 10:1 to about 60:1 (wt/wt).
- the ratio between the lipid composition and the polynucleotide can be about 10:1, 11:1, 12:1, 13:1, 14:1, 15:1, 16:1, 17:1, 18:1, 19:1, 20:1, 21:1, 22:1, 23:1, 24:1, 25:1, 26:1, 27:1, 28:1, 29:1, 30:1, 31:1, 32:1, 33:1, 34:1, 35:1, 36:1, 37:1, 38:1, 39:1, 40:1, 41:1, 42:1, 43:1, 44:1, 45:1, 46:1, 47:1, 48:1, 49:1, 50:1, 51:1, 52:1, 53:1, 54:1, 55:1, 56:1, 57:1, 58:1, 59:1 or 60:1 (wt/wt).
- the wt/wt ratio of the lipid composition to the polynucleotide encoding a therapeutic agent is about 20:1 or about 15:1.
- the pharmaceutical composition disclosed herein can contain more than one polypeptides.
- a pharmaceutical composition disclosed herein can contain two or more polynucleotides (e.g., RNA, e.g., mRNA).
- the lipid nanoparticles described herein can comprise polynucleotides (e.g., mRNA) in a lipid:polynucleotide weight ratio of 5:1, 10:1, 15:1, 20:1, 25:1, 30:1, 35:1, 40:1, 45:1, 50:1, 55:1, 60:1 or 70:1, or a range or any of these ratios such as, but not limited to, 5:1 to about 10:1, from about 5:1 to about 15:1, from about 5:1 to about 20:1, from about 5:1 to about 25:1, from about 5:1 to about 30:1, from about 5:1 to about 35:1, from about 5:1 to about 40:1, from about 5:1 to about 45:1, from about 5:1 to about 50:1, from about 5:1 to about 55:1, from about 5:1 to about 60:1, from about 5:1 to about 70:1, from about 10:1 to about 15:1, from about 10:1 to about 20:1, from about 10:1 to about 25:
- the lipid nanoparticles described herein can comprise the polynucleotide in a concentration from approximately 0.1 mg/ml to 2 mg/ml such as, but not limited to, 0.1 mg/ml, 0.2 mg/ml, 0.3 mg/ml, 0.4 mg/ml, 0.5 mg/ml, 0.6 mg/ml, 0.7 mg/ml, 0.8 mg/ml, 0.9 mg/ml, 1.0 mg/ml, 1.1 mg/ml, 1.2 mg/ml, 1.3 mg/ml, 1.4 mg/ml, 1.5 mg/ml, 1.6 mg/ml, 1.7 mg/ml, 1.8 mg/ml, 1.9 mg/ml, 2.0 mg/ml or greater than 2.0 mg/ml.
- Nanoparticle Compositions are Formulated as lipid nanoparticles (LNP). Accordingly, the present disclosure also provides nanoparticle compositions comprising (i) a lipid composition comprising a delivery agent such as compound as described herein, and (ii) a polynucleotide encoding a polypeptide. In such nanoparticle composition, the lipid composition disclosed herein can encapsulate the polynucleotide encoding a polypeptide. Nanoparticle compositions are typically sized on the order of micrometers or smaller and can include a lipid bilayer.
- Nanoparticle compositions encompass lipid nanoparticles (LNPs), liposomes (e.g., lipid vesicles), and lipoplexes.
- a nanoparticle composition can be a liposome having a lipid bilayer with a diameter of 500 nm or less.
- Nanoparticle compositions include, for example, lipid nanoparticles (LNPs), liposomes, and lipoplexes.
- nanoparticle compositions are vesicles including one or more lipid bilayers.
- a nanoparticle composition includes two or more concentric bilayers separated by aqueous compartments. Lipid bilayers can be functionalized and/or crosslinked to one another.
- Lipid bilayers can include one or more ligands, proteins, or channels.
- a lipid nanoparticle comprises an ionizable amino lipid, a structural lipid, a phospholipid, and mRNA.
- the LNP comprises an ionizable amino lipid, a PEG-modified lipid, a sterol and a structural lipid.
- the LNP has a molar ratio of about 40-50% ionizable amino lipid; about 5-15% structural lipid; about 30-45% sterol; and about 1-5% PEG-modified lipid.
- the lipid nanoparticle comprises 47-49 mol.% ionizable cationic lipid (e.g.
- ionizable amino lipid e.g., Compound I-1, Compound I-2, or Compound I-3
- 10-12 mol.% non-cationic lipid e.g., phospholipid, e.g., DSPC
- 38-40 mol.% sterol e.g., cholesterol
- 1-3 mol.% PEG-modified lipid e.g., PEG-DMG or Compound P-I
- the lipid nanoparticle (“LNP-1”) may comprise the following components at the following molar ratios: (i) 45-50 mol.% Compound I-1 (ii) 35-45 mol.% sterol (e.g., cholesterol); (iii) 8-12 mol.% phospholipid (e.g., DSPC or DOPE); and (iv) 1.5-3.5 mol.% PEG-lipid.
- the lipid nanoparticle (“LNP-1A”) may comprise the following components at the following molar ratios: (i) 45-50 mol.% Compound I-1 (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid.
- the lipid nanoparticle (“LNP-1B”) may comprise the following components at the following molar ratios: (i) 45-50 mol.% Compound I-1 (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid.
- the lipid nanoparticle (“LNP-2”) may comprise the following: (i) 45-50 mol.% Compound I-2; (ii) 35-45 mol.% sterol (e.g., Cholesterol); (iii) 8-12 mol.% phospholipid (e.g., DSPC or DOPE); and (iv) 1.5-3.5 mol.% PEG-lipid.
- the lipid nanoparticle (“LNP-2A”) may comprise the following: (i) 45-50 mol.% Compound I-2; (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid.
- the lipid nanoparticle (“LNP-2B”) may comprise the following components at the following molar ratios: (i) 45-50 mol.% Compound I-2; (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid.
- the lipid nanoparticle (“LNP-3”) may comprise the following: (i) 45-50 mol.% Compound I-3; (ii) 35-45 mol.% sterol (e.g., Cholesterol); (iii) 8-12 mol.% phospholipid (e.g., DSPC or DOPE); and (iv) 1.5-3.5 mol.% PEG-lipid.
- the lipid nanoparticle (“LNP-3A”) may comprise the following: (i) 45-50 mol.% Compound I-3; (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid.
- the lipid nanoparticle (“LNP-3B”) may comprise the following: (i) 45-50 mol.% Compound I-3; (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid.
- the LNP has a polydispersity value of less than 0.4.
- the LNP has a net neutral charge at a neutral pH.
- the LNP has a mean diameter of 50-150 nm. In some instances, the LNP has a mean diameter of 80-100 nm.
- lipid refers to a small molecule that has hydrophobic or amphiphilic properties. Lipids may be naturally occurring or synthetic. Examples of classes of lipids include, but are not limited to, fats, waxes, sterol-containing metabolites, vitamins, fatty acids, glycerolipids, glycerophospholipids, sphingolipids, saccharolipids, and polyketides, and prenol lipids. In some instances, the amphiphilic properties of some lipids lead them to form liposomes, vesicles, or membranes in aqueous media. In some instances, a lipid nanoparticle (LNP) may comprise an ionizable amino lipid.
- LNP lipid nanoparticle
- the charged moiety may be anionic (i.e., negatively charged) or cationic (i.e., positively charged).
- positively-charged moieties include amine groups (e.g., primary, secondary, and/or tertiary amines), ammonium groups, pyridinium group, guanidine groups, and imidizolium groups.
- the charged moieties comprise amine groups.
- negatively- charged groups or precursors thereof include carboxylate groups, sulfonate groups, sulfate groups, phosphonate groups, phosphate groups, hydroxyl groups, and the like.
- the charge of the charged moiety may vary, in some cases, with the environmental conditions, for example, changes in pH may alter the charge of the moiety, and/or cause the moiety to become charged or uncharged. In general, the charge density of the molecule may be selected as desired. It should be understood that the terms “charged” or “charged moiety” does not refer to a “partial negative charge” or “partial positive charge” on a molecule. The terms “partial negative charge” and “partial positive charge” are given their ordinary meaning in the art. A “partial negative charge” may result when a functional group comprises a bond that becomes polarized such that electron density is pulled toward one atom of the bond, creating a partial negative charge on the atom.
- the ionizable amino lipid may be selected from, but not limited to, Formula CLI-CLXXXII of US Patent No. 7,404,969; each of which is herein incorporated by reference in their entirety.
- the lipid may be a cleavable lipid such as those described in International Publication No. WO2012170889, herein incorporated by reference in its entirety.
- the lipid may be synthesized by methods known in the art and/or as described in International Publication Nos. WO2013086354; the contents of each of which are herein incorporated by reference in their entirety.
- Nanoparticle compositions can be characterized by a variety of methods. For example, microscopy (e.g., transmission electron microscopy or scanning electron microscopy) can be used to examine the morphology and size distribution of a nanoparticle composition. Dynamic light scattering or potentiometry (e.g., potentiometric titrations) can be used to measure zeta potentials. Dynamic light scattering can also be utilized to determine particle sizes. Instruments such as the Zetasizer Nano ZS (Malvern Instruments Ltd, Malvern, Worcestershire, UK) can also be used to measure multiple characteristics of a nanoparticle composition, such as particle size, polydispersity index, and zeta potential.
- microscopy e.g., transmission electron microscopy or scanning electron microscopy
- Dynamic light scattering or potentiometry e.g., potentiometric titrations
- Dynamic light scattering can also be utilized to determine particle sizes.
- Instruments such as the Ze
- the size of the nanoparticles can help counter biological reactions such as, but not limited to, inflammation, or can increase the biological effect of the polynucleotide.
- size or “mean size” in the context of nanoparticle compositions refers to the mean diameter of a nanoparticle composition.
- the polynucleotide encoding a polypeptide(s) described herein are formulated in lipid nanoparticles having a diameter from about 10 to about 100 nm such as, but not limited to, about 10 to about 20 nm, about 10 to about 30 nm, about 10 to about 40 nm, about 10 to about 50 nm, about 10 to about 60 nm, about 10 to about 70 nm, about 10 to about 80 nm, about 10 to about 90 nm, about 20 to about 30 nm, about 20 to about 40 nm, about 20 to about 50 nm, about 20 to about 60 nm, about 20 to about 70 nm, about 20 to about 80 nm, about 20 to about 90 nm, about 20 to about 100 nm, about 30 to about 40 nm, about 30 to about 50 nm, about 30 to about 60 nm, about 30 to about 70 nm, about 30 to about 80 nm, about 30 to about 90 nm, about 20 to
- the nanoparticles have a diameter from about 10 to 500 nm. In some instances, the nanoparticle has a diameter greater than 100 nm, greater than 150 nm, greater than 200 nm, greater than 250 nm, greater than 300 nm, greater than 350 nm, greater than 400 nm, greater than 450 nm, greater than 500 nm, greater than 550 nm, greater than 600 nm, greater than 650 nm, greater than 700 nm, greater than 750 nm, greater than 800 nm, greater than 850 nm, greater than 900 nm, greater than 950 nm or greater than 1000 nm.
- the largest dimension of a nanoparticle composition is 1 ⁇ m or shorter (e.g., 1 ⁇ m, 900 nm, 800 nm, 700 nm, 600 nm, 500 nm, 400 nm, 300 nm, 200 nm, 175 nm, 150 nm, 125 nm, 100 nm, 75 nm, 50 nm, or shorter).
- a nanoparticle composition can be relatively homogenous.
- a polydispersity index can be used to indicate the homogeneity of a nanoparticle composition, e.g., the particle size distribution of the nanoparticle composition.
- a small (e.g., less than 0.3) polydispersity index generally indicates a narrow particle size distribution.
- a nanoparticle composition can have a polydispersity index from about 0 to about 0.25, such as 0.01, 0.02, 0.03, 0.04, 0.05, 0.06, 0.07, 0.08, 0.09, 0.10, 0.11, 0.12, 0.13, 0.14, 0.15, 0.16, 0.17, 0.18, 0.19, 0.20, 0.21, 0.22, 0.23, 0.24, or 0.25.
- the polydispersity index of a nanoparticle composition disclosed herein can be from about 0.10 to about 0.20.
- the amount of a polynucleotide present in a pharmaceutical composition disclosed herein can depend on multiple factors such as the size of the polynucleotide, desired target and/or application, or other properties of the nanoparticle composition as well as on the properties of the polynucleotide.
- the amount of an mRNA useful in a nanoparticle composition can depend on the size (expressed as length, or molecular mass), sequence, and other characteristics of the mRNA.
- the relative amounts of a polynucleotide in a nanoparticle composition can also vary.
- the relative amounts of the lipid composition and the polynucleotide present in a lipid nanoparticle composition of the present disclosure can be optimized according to considerations of efficacy and tolerability.
- the N:P ratio can serve as a useful metric.
- the N:P ratio of a nanoparticle composition controls both expression and tolerability, nanoparticle compositions with low N:P ratios and strong expression are desirable.
- N:P ratios vary according to the ratio of lipids to RNA in a nanoparticle composition. In general, a lower N:P ratio is preferred.
- the one or more RNA, lipids, and amounts thereof can be selected to provide an N:P ratio from about 2:1 to about 30:1, such as 2:1, 3:1, 4:1, 5:1, 6:1, 7:1, 8:1, 9:1, 10:1, 12:1, 14:1, 16:1, 18:1, 20:1, 22:1, 24:1, 26:1, 28:1, or 30:1.
- the N:P ratio can be from about 2:1 to about 8:1.
- the N:P ratio is from about 5: 1 to about 8: 1.
- the N:P ratio is between 5: 1 and 6: 1.
- the N:P ratio is about is about 5.67:1.
- the polynucleotides described herein can be formulated for controlled release and/or targeted delivery.
- controlled release refers to a pharmaceutical composition or compound release profile that conforms to a particular pattern of release to effect a therapeutic outcome.
- the polynucleotides can be encapsulated into a delivery agent described herein and/or known in the art for controlled release and/or targeted delivery.
- encapsulate means to enclose, surround or encase. As it relates to the formulation of the compounds of the present disclosure, encapsulation can be substantial, complete or partial.
- substantially encapsulated means that at least greater than 50, 60, 70, 80, 85, 90, 95, 96, 97, 98, 99, or greater than 99% of the pharmaceutical composition or compound of the present disclosure can be enclosed, surrounded or encased within the delivery agent.
- Partial encapsulation or “partially encapsulate” means that less than 10, 10, 20, 30, 40 50 or less of the pharmaceutical composition or compound of the present disclosure can be enclosed, surrounded or encased within the delivery agent.
- the therapeutic nanoparticle polynucleotide can be formulated for sustained release.
- sustained release refers to a pharmaceutical composition or compound that conforms to a release rate over a specific period of time. The period of time can include, but is not limited to, hours, days, weeks, months and years.
- the sustained release nanoparticle of the polynucleotides described herein can be formulated as disclosed in Inti. Pub. No. W02010075072 and U.S. Pub. Nos. US20100216804, US20110217377, US20120201859 and US20130150295, each of which is herein incorporated by reference in their entirety.
- the therapeutic nanoparticle polynucleotide can be formulated to be target specific, such as those described in Inti. Pub. Nos. WO2008121949, WO2010005726, WO2010005725, WO2011084521 and WO2011084518; and U.S. Pub. Nos. US20100069426, US20120004293 and US20100104655, each of which is herein incorporated by reference in its entirety.
- the LNPs can be prepared using microfluidic mixers or micromixers.
- Exemplary microfluidic mixers can include, but are not limited to, a slit interdigital micromixer including, but not limited to those manufactured by Microinnova (Allerheiligen bei Wildon, Austria) and/or a staggered herringbone micromixer (SHM) (see, Zhigaltsev et al., Langmuir.28:3633-40 (2012); Belliveau et al., Molecular Therapy-Nucleic Acids. 1:e37 (2012); Chen et al., J. Am. Chem. Soc. 134(16):6948-51 (2012); each of which is herein incorporated by reference in its entirety).
- SHM herringbone micromixer
- Exemplary micromixers include Slit Interdigital Microstructured Mixer (SIMM-V2) or a Standard Slit Interdigital Micro Mixer (SSIMM) or Caterpillar (CPMM) or Impinging-jet (IJMM,) from the Institut für Mikrotechnik Mainz GmbH, Mainz Germany.
- methods of making LNP using SHM further comprise mixing at least two input streams wherein mixing occurs by microstructure-induced chaotic advection (MICA).
- MICA microstructure-induced chaotic advection
- fluid streams flow through channels present in a herringbone pattern causing rotational flow and folding the fluids around each other.
- This method can also comprise a surface for fluid mixing wherein the surface changes orientations during fluid cycling.
- Methods of generating LNPs using SHM include those disclosed in U.S. Pub. Nos.
- the polynucleotides described herein can be Formulated in lipid nanoparticles using microfluidic technology (see, Whitesides, George M., Nature 442: 368-373 (2006); and Abraham et al., Science 295: 647-651 (2002); each of which is herein incorporated by reference in its entirety).
- the polynucleotides can be Formulated in lipid nanoparticles using a micromixer chip such as, but not limited to, those from Harvard Apparatus (Holliston, MA) or Dolomite Microfluidics (Royston, UK).
- a micromixer chip can be used for rapid mixing of two or more fluid streams with a split and recombine mechanism.
- the nanoparticles described herein are stealth nanoparticles or target-specific stealth nanoparticles such as, but not limited to, those described in U.S. Pub. No. US20130172406, herein incorporated by reference in its entirety.
- the stealth or target-specific stealth nanoparticles can comprise a polymeric matrix, which can comprise two or more polymers such as, but not limited to, polyethylenes, polycarbonates, polyanhydrides, polyhydroxyacids, polypropylfumerates, polycaprolactones, polyamides, polyacetals, polyethers, polyesters, poly(orthoesters), polycyanoacrylates, polyvinyl alcohols, polyurethanes, polyphosphazenes, polyacrylates, polymethacrylates, polycyanoacrylates, polyureas, polystyrenes, polyamines, polyesters, polyanhydrides, polyethers, polyurethanes, polymethacrylates, polyacrylates, polycyanoacrylates, or combinations thereof.
- polymers such as, but not limited to, polyethylenes, polycarbonates, polyanhydrides, polyhydroxyacids, polypropylfumerates, polycaprolactones, polyamides, polyacetals, polyethers, polyester
- VHH containing polypeptides described herein can be used in the treatment of any disease associated the dysregulation of the c-MET/HGF pathway.
- the disease shows an increased activation of the c-MET signaling relative to age-matched human subjects without cancer.
- these VHH containing polypeptides are used in the treatment of cancers with one or more c-MET mutations.
- these VHH containing polypeptides are used in the treatment of cancers with one or more a c- MET amplifications.
- these VHH containing polypeptides are used in the treatment of cancers with c-MET overexpression relative to age-matched human subjects without cancer.
- the methods involve administering to the human subject in need thereof a therapeutically effective amount of a VHH containing polypeptide herein.
- the VHH containing polypeptide is a monovalent polypeptide.
- the VHH containing polypeptide is a bivalent polypeptide.
- the VHH containing polypeptide is a bivalent polypeptide (e.g., a c-MET binding VHH-linked to IgG4PAA).
- the VHH containing polypeptide is a bispecific antibody (e.g., a bispecific T cell engager (BiTE), a bispecific NK cell engager (BiKE); a bispecific antibody that binds both c-MET and another tumor associated/related antigen).
- the VHH containing polypeptide is a multispecific antibody construct. In some cases, the VHH containing polypeptide is a conjugate (e.g., an antibody drug conjugate). In some cases, the human is administered a T cell comprising a CAR-T cell wherein the T cell expresses a CAR comprising a VHH that specifically binds c-MET. In some cases, the human is administered an NK cell comprising a CAR-NK cell wherein the NK cell is engineered to express a CAR comprising a VHH that specifically binds c-MET.
- the polypeptides of this disclosure may be administered subcutaneously or intravenously.
- kits that contain a VHH, a polypeptide, a bispecific antibody, a VHH-conjugate, or a CAR of this disclosure, and/or nucleic acids (e.g., mRNA) encoding the same.
- the kits provided herein contain one or more cells engineered to express and secrete a polypeptide(s) comprising an anti-c- MET VHH of the disclosure, such as a cell containing a nucleic acid molecule(s) of the disclosure.
- kits described herein may include reagents that can be used to produce a pharmaceutical composition of the invention.
- kits described herein may include reagents that can induce the expression of a polypeptide(s) comprising an anti-c- MET VHH of the disclosure within cells (e.g., bacterial, yeast, insect, or mammalian cells).
- Other kits described herein may include tools for engineering a prokaryotic or eukaryotic cell (e.g., a CHO cell or a BL21(DE3) E. Coli cell or an immune cell) so as to express and secrete a polypeptide(s) comprising an anti-c-MET VHH of the disclosure.
- kits may contain CHO cells stored in an appropriate media and optionally frozen according to methods known in the art.
- the kit may also contain a nucleic acid encoding the desired VHH antibody as well as reagents for expressing the VHH antibody or binding protein in the cell.
- a kit described herein may also provide a polypeptide(s) comprising an anti-c- MET VHH of the disclosure, a nucleic acid encoding the same (e.g., mRNA), or a LNP that encapsulates a nucleic acid (e.g., mRNA) that encodes an anti-c-MET VHH described herein, in combination with a package insert describing how the VHH antibody, or nucleic acid may be administered to a subject, for example, for the treatment of a disease, disorder and/or condition (e.g., cancer).
- a disease, disorder and/or condition e.g., cancer
- Llamas were immunized with mRNA encoding full length c-MET, the sequence of which is provided below: ATGAAGGCCCCTGCTGTGCTGGCCCCTGGCATACTGGTGTTGCTGTTCACACT GGTGCAACGGAGTAACGGCGAGTGCAAGGAGGCCCTGGCCAAGAGCGAGAT GAACGTGAATATGAAGTATCAACTGCCCAACTTCACTGCCGAGACCCCTATC CAGAACGTGATCCTTCACGAACACCACATCTTCCTGGGAGCCACCAACTACA TCTACGTCCTAAACGAGGAGGACCTGCAGAAGGTAGCCGAGTACAAGACAG GCCCCGTGCTTGAGCACCCCGACTGCTTCCCCTGCCAGGACTGCAGCAGCAA GGCCAACCTTTCGGGCGGCGTGTGGAAGGACAACATCAACATGGCACTGGTG GTGGACACCTACTACGACGACCAGCTCATCAGCTGCGGCAGCGTGAACAGAG GCACTTGCCAGCGGCACGTTTCCACACAACC
- boost immunizations (alternative human and cyno mRNA at 500 ⁇ g) were given approximately every two weeks intramuscularly and a final fifth boost (day 77) was given intravenously (IV).
- the bleed samples were tested on CHO cells expressing Hu c- MET protein via FACS to check the titers.
- a production bleed of 600ml was taken and used to isolate and cryopreserve PBMCs. 2.
- B-cells FACS Sorting of B-cells and Culturing of Antigen Positive B-cells: PBMCs were incubated with biotinylated c-MET protein and antigen positive VHH B-cells were selected by Fluorescence Activated Cell Sorting (FACS) using fluorescently labeled rabbit anti-camelid VHH antibody AF-647 (Genscript) and anti- biotin-PE antibody (Biolegend). B-cells were sorted at a concentration of 5 x 10 6 cells/mL and cells staining positive with both the antibodies (antigen positive VHH cells) were collected into 96 well plates with 3 cells/well cell density.
- FACS Fluorescence Activated Cell Sorting
- the culture plates were incubated for up to 7-10 days in a CO2 incubator at 37 o C under 5% CO2. Typically, between days 8-11 Proliferating B cell cultures were processed to harvest culture supernatants for screening the secreted antibodies against the target antigen (c-MET protein), and the cell pellets were frozen for isolation of RNA and antibody sequencing. 3. Screening for Antibody Hits via FACS: B cell culture supernatant from each wells were screened for antibody binding to the CHO cells expressing hu c-MET protein via FACS.
- CHO cells expressing Human c- MET were resuspended in FACS buffer (PBS pH 7.4 + 2% FBS +1 mM EDTA) at 1,000,000 cells/ml, and 80 ⁇ l of the cell suspension was added in each well in a 96-well V bottom polypropylene plate (Nunc). The 40 ⁇ l of B-cell supernatants were added in each well.
- FACS buffer PBS pH 7.4 + 2% FBS +1 mM EDTA
- the IgG positive wells were subjected to VHH positive screening using goat anti-llama IgG 2+3-APC secondary antibody from Jackson at 1:300 dilution in FCAS buffer.
- the VHH positive wells were used for mRNA isolation and cDNA synthesis as discussed below. 4. Sequencing Antibodies from Target Positive B-cells: To sequence antibodies, from VHH positive wells, cDNA was generated and sequenced using Sanger sequencing and NGS. The frozen B cell pellets are lysed by adding the lysis buffer.
- PCR was performed according to the standard procedures with 2-5 ⁇ l of cDNA reaction product and with historical primers used in Llama antibody discovery (see, e.g., Conrath et al., Antimicro.-Agents Chemother. v45(10):2807-2812 (2001); Els Pardon, Nat. Protoc. 9(3):674-93 (2014), herein incorporated by reference in their entirety).
- the PCR amplified VHH genes were then sequenced using one of two methods Sanger sequencing or Next Generation Sequencing (NGS). In both cases the PCR used CALL001 forward primer and CALL002 reverse primers that anneal to the leader sequence and 5 ⁇ end of CH2 domain of the heavy chain respectively. The resulting PCR product was sequenced directly via Sanger sequencing and NGS (Azenta, MA). 5. Purification of Single Domain Antibodies: Recombinant antibodies were expressed at Biointron Biologics Inc., (China) in the IgG and VHH-His formats in the CHO cells. The antibodies were purified from 30mL CHO using Ni-NTA affinity chromatography.
- the purified proteins were run on size exclusion chromatography (SEC) to check their size and homogeneity.
- the Antibodies were further analyzed by SDS-PAGE under reducing and non-reducing conditions to check for purity and validation of molecular weight. An average of 2-5 mg protein was purified from 30 mL CHO culture supernatants. 6.
- FACS Cell Binding Assays CHO cell lines that stably express Human and Cyno c-MET were resuspended in FACS buffer (PBS pH7.4 + 2% FBS +2mM EDTA) at 1,000,000 cells/mL, and 80 ⁇ l of the cell suspension was added in each well in a 96-well V bottom polypropylene plate (Nunc).
- the 80 ⁇ l of testing VHH antibodies were mixed to each well at starting concentration of 1000 nM with 12 step 1:3 dilutions. ⁇ After one-hour incubation at 4°C, pelleted cells were washed 2x by ice cold FACS buffer and then resuspended in 80 ⁇ l anti-His-iFluor 647 from Biolegend at 1:500 dilution in FACS buffer, incubated 15min at 4°C, centrifuged at 1500 rpm for 3 min; cell pellets were washed twice and resuspended in 100 ⁇ l FACS buffer for analysis. ⁇ Cellular fluorescence was determined on BD FACSCantoII (Becton-Dickinson, Franklin Lakes, NJ). ⁇ The data was processed with FlowJo software (10.8.1).
- HGF Hepatocyte Growth Factor
- CHO cells expressing Human c-MET were resuspended in FACS buffer (PBS pH7.4 + 2% FBS +1 mM EDTA) at 1,000,000 cells/ml, and 60 ⁇ l of the cell suspension was added in each well in a 96-well V bottom polypropylene plate (Nunc).
- the testing VHH antibodies (IgG format) were added at a fixed 100 nM concentration in each well in the absence (0 nM HGF) and in the presence (100 nM and 1000 nM) of HGF in a final reaction volume of 150 ⁇ l.
- pelleted cells were washed 2x by ice cold FACS buffer and then resuspended in 80 ⁇ l goat anti-human Fc antibody from Abcam at 1:500 dilution in FACS buffer, incubated 15min at 4°C, centrifuged at 1500 rpm for 3 min; cell pellets were washed twice and resuspended in 100 ⁇ l FACS buffer for analysis.
- Cellular fluorescence was determined on BD FACSCantoII (Becton-Dickinson, Franklin Lakes, NJ). ⁇ The data was processed with FlowJo software (10.8.1) and the MFI values were exported and compared. The percentage reduction in MFI data was plotted using R in RStudio.
- VHH Antibodies The VHH antibodies were humanized similar to the method of Hanf et al., Methods.2014 Jan 1;65(1):68-76. doi: 10.1016/j.ymeth.2013.06.024. Epub 2013 Jun 28. Briefly, the complementarity-determining regions (CDRs) sequences of anti-C-MET VHH Abs (Table 1) were annotated using the IMGT numbering scheme. Each VHH nucleotide sequence was generated and used to identify the nearest human germline VH sequences by searching for similar sequences with the NCBI IgBLAST program. Common J and D gene sequences were attached to the VH as the acceptor.
- CDRs complementarity-determining regions
- the framework residues that were critical for huVH/VL interactions are back mutated to llama sequence canonical llama residues, also potentially structural defects due to mismatches at the graft interface can be fixed by mutating some framework residues to llama, or by mutating some residues on the CDRs’ backside to human or to a de novo designed sequence.
- CDR stabilizing or overall fold stabilizing sequences were then back-mutated to the corresponding llama sequence to maintain the biophysical properties and target binding affinity. ⁇ 9.
- Example 2 Generation and Selection of VHH antibodies against c-MET A total of 40 single domain antibodies have been discovered against c-MET protein. Figures 1 and 2 show the monovalent binding of these VHH antibodies to CHO cells expressing human c-MET and Cyno c-MET proteins, respectively, while Figure 5 shows monovalent binding to primary cancer cells (HS746T) expressing c-MET. For 35 of these VHH Abs, bivalent binding was also tested ( Figures 3 and 4).
- VHH Abs For the remaining five VHH Abs (namely CV3N1, CV7N1, CV13N2, CV16N1 & CV19N1), bivalent binding will be tested once these antibodies are available in the bivalent format. Overall, these 40 antibodies showed strong binding to CHO cells expressing human and cyno c- MET and to HS746T c-MET cancer cells. These antibodies are cross reactive with human and cyno based on their bivalent binding. Furthermore, 16 VHH antibodies competed with HGF for c-MET binding (over 50% reduction in MFI when HGF is present) based on the HGF competition assay (Figure 6).
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- Immunology (AREA)
- General Health & Medical Sciences (AREA)
- Medicinal Chemistry (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Biochemistry (AREA)
- Animal Behavior & Ethology (AREA)
- General Chemical & Material Sciences (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Pharmacology & Pharmacy (AREA)
- Biophysics (AREA)
- Genetics & Genomics (AREA)
- Molecular Biology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Peptides Or Proteins (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
Abstract
Described herein are anti-c-MET binding polypeptides. These polypeptides may comprise a monovalent or bivalent VHH that specifically binds c-MET. In some cases, these polypeptides comprise bispecific antibodies, multispecific antibodies, antibody conjugates, or CARs that comprise one or more VHHs that specifically binds c-MET. The disclosure features nucleic acids, vectors, and host cells comprising these polypeptides as well as methods of making these polypeptides. Also provided are methods of using these polypeptides to treat diseases associated with aberrant c-MET expression and/or signaling such as cancer.
Description
C-MET BINDING ANTIBODIES, NUCLEIC ACIDS ENCODING SAME, AND METHODS OF USE CROSS-REFERENCE TO RELATED APPLICATIONS This application claims the benefit of priority of U.S. Provisional Application No. 63/533,759, filed on August 21, 2023, and U.S. Provisional Application No.63/546,326 filed on October 30, 2023, the contents of both of which are incorporated by reference in their entirety herein. SEQUENCE LISTING The instant application contains a Sequence Listing which has been submitted electronically in XML format and is hereby incorporated by reference in its entirety. Said XML copy, created on June 28, 2024, is named 45817-0153WO1_SL.xml and is 726,262 bytes in size. BACKGROUND Hepatocyte growth factor receptor (HGF receptor) also known as c-MET is a protein that is encoded by the MET gene. c-MET is normally expressed by cells of epithelial origin. c-MET is a receptor tyrosine kinase that is expressed on the cell surface where it can bind to its ligand, HGF. Upon HGF binding to c-MET, it induces c-MET dimerization leading to its activation which initiates a series of intracellular signals that mediate embryogenesis and wound healing in normal cells. However, in cancer cells, aberrant HGF/c-MET activation, usually related to c-MET gene mutations, c-Met overexpression, and/or c-MET amplification, promotes tumor development and progression by stimulating the PI3K/AKT, Ras/MAPK, JAK/STAT, SRC, Wnt/β-catenin, and other signaling pathways. Abnormal MET activation in cancer is associated with poor prognosis, where aberrantly active MET triggers tumor growth, angiogenesis, and metastasis. MET is deregulated in many types of human malignancies, including cancers
of kidney, liver, stomach, breast, and brain. Thus, c-MET is a clinically important therapeutic target. SUMMARY This disclosure provides binding agents comprising a c-MET binding polypeptide. The c-MET binding polypeptide can comprise a single domain antibody. The binding agent can be monovalent, bivalent, multivalent, bispecific, or multispecific. The binding agent can also be an antibody conjugate. In addition, the binding agent can be a chimeric antigen receptor that binds to c-MET. Also featured are nucleic acids such as mRNA that encode these binding agents. These binding agents and mRNA can be used to treat diseases associated with aberrant HGF/c-MET activation. In one aspect, the disclosure features a polypeptide comprising a variable domain of the heavy chain of a heavy chain-only antibody (VHH) that specifically binds c-MET (human and/or cyno c-MET), the VHH comprising VHH complementarity determining region (CDR) 1, VHH CDR2, and VHH CDR3 of any of the amino acid sequences set forth in SEQ ID NOs.: 1 to 40. In another aspect, the disclosure features a variable domain of the heavy chain of a heavy chain-only antibody (VHH) that specifically binds c-MET (human and/or cyno c- MET), the VHH comprising VHH complementarity determining region (CDR) 1, VHH CDR2, and VHH CDR3 of any of the amino acid sequences set forth in SEQ ID NOs.: 1 to 40. In some instances, the VHH CDR1, VHH CDR2, and VHH CDR3 are based on CDR definitions according to Kabat, Chothia, enhanced Chothia, contact, IMGT, or AbM CDR definitions. In some instances, the polypeptide or VHH inhibits the binding of Human Growth Factor (HGF) to human c-Met.
In some instances, the polypeptide or VHH does not inhibit the binding of Human Growth Factor (HGF) to human c-Met. In some instances, the VHH is humanized. In some cases, 1 to 60, 1 to 50, 1 to 40, 1 to 30, 1 to 25, 1 to 20, 1 to 15, 1 to 10, 1 to 5, 1 to 4, 5 to 15, 6 to 12, or 20, 19, 18, 17, 16, 15, 14, 13, 12, 11, 10, 9, 8, 7, 6, 5, 4, 3, 2, or 1 amino acid(s) of the VHH are substituted for humanization. In some cases, humanization of the VHH is by resurfacing/veneering. In some cases, humanization of the VHH is by CDR grafting. In some cases, one or more of positions 42, 49, 50, and 52 (numbering based on IMGT numbering) are humanized. In other cases one or more of positions 42, 49, 50, and 52 (numbering based on IMGT numbering) are not humanized. In yet other cases none of positions 42, 49, 50, and 52 (numbering based on IMGT numbering) are humanized. In certain cases, positions 42 and/or 52 (numbering based on IMGT numbering) are not humanized. In other cases, positions 49 and/or 50 (numbering based on IMGT numbering) are not humanized. In certain cases, positions 49 and/or 50 (numbering based on IMGT numbering) are humanized. In some cases, the polypeptide or VHH is conjugated, fused, or linked to an agent selected from the group consisting of a purification tag, a fluorophore, a drug, a photosensitizer, a nanoparticle, a toxic agent, a radionuclide, a VHH, an Fab, a scFv, a multimerization module, a moiety that facilitates the polypeptide crossing the blood brain barrier, and a half-life extension moiety. In some cases, the agent is a half-life extension moiety. In some instances, the agent is conjugated, attached, or linked to the C-terminus of the VHH. In some cases, the polypeptide or VHH is conjugated, fused, or linked to a human Ig Fc domain, optionally, wherein the human Ig Fc domain is conjugated, fused, or linked to the polypeptide or VHH at the C-terminus of the polypeptide or VHH. In some cases, the human Ig Fc domain further comprises a human Ig hinge domain, wherein the hinge domain is attached or linked at the N-terminus of the human Ig Fc domain. In some cases, the human Ig is a human IgG1, human IgG2, human IgG3, or human IgG4. In one
instance, the human Ig is human IgG1. In another instance, the human Ig is human IgG4. In certain cases, the polypeptide or VHH is conjugated, fused, or linked to an amino acid sequence that is at least 80%, at least 85%, at least 90%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or 100% identical to the human IgG4 PAA hinge and Fc domain sequence set forth in SEQ ID NO:643. In certain cases, the polypeptide or VHH is conjugated, fused, or linked to an amino acid sequence that is identical to the human IgG4 PAA hinge and Fc domain sequence set forth in SEQ ID NO:643 except for 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 amino acid substitutions. In some cases, the Fc domain includes mutations that promote heterodimerization (e.g., knob-into hole mutations, electrostatic steering mutations). In one aspect, the disclosure features a polypeptide comprising a variable domain of the heavy chain of a heavy chain-only antibody (VHH) that specifically binds c-MET (human and/or cyno c-MET), the VHH comprising VHH complementarity determining region (CDR) 1, VHH CDR2, and VHH CDR3 of any of the amino acid sequences set forth in SEQ ID NOs.: 1 to 40, wherein 1 to 20 amino acids in the framework regions of the VHH are humanized, wherein the polypeptide is linked to a half-life extending moiety, and optionally, wherein the polypeptide is conjugated, fused, or linked to an amino acid sequence that is at least 80%, at least 85%, at least 90%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, or 100% identical to the human IgG4 PAA hinge and Fc domain sequence set forth in SEQ ID NO:643, or to an amino acid sequence that is identical to the human IgG4 PAA hinge and Fc domain sequence set forth in SEQ ID NO:643 with 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 amino acid substitutions. In some cases, the polypeptide competes with HGF for binding to c-MET. In other cases, the polypeptide does not compete with HGF for binding to c- MET. In some instances, the polypeptide or VHH comprises a second VHH that specifically binds to an antigen selected from the group consisting of: CD3, EGFR, VEGF, VEGFR, HGF, CXCR2, CXCR4, CXCR7, CXCL11, CXCL12, CEA, PSMA, MMR, PD-1, PDL-1, survivin, HER2, HGFR, P2X7, death receptor 5 CTLA4, CD7,
CD8, CD11b, CD20, CD38, CD45, M HC-II, fibronectin, TUFM, CapG, CAIX, CD33, CD47, ARTC22, B3GAT1 (CD57), CCR7 (CD197), CD16, CD16a, CD2 CD226, CD244, CD27, CD3, CD300A, CD34, CD58, CD59, CD69, CSF2, CX3CR1, CXCR1 (CD128), CXCR3 (CD183), CXCR4, EOMES, GZMB, ICAM1 (CD54), IFNG, IL-15R, IL-1R, IL22, IL-2RB (CD122), IL-7R (CD127), ITGA1 (CD49a), ITGA2 (CD49b), ITGAL (CD11a), ITGAM (CD11b), ITGB2 (CD18), KIR, KIR2DL1, KIR2DL2, KIT (CD117), KLRB1C, KLRC1, KLRC2, KLRD1 (CD94), KLRF1, KLRG1, KLRK1, LILRB1, KLRA4, KLRA8, MICA/BNCAM1 (CD56), NK2D, NKP46 (NCR1, CD335), NCR2, NCR3 (CD337), PRF1, SELL (CD62L), SIGLEC7, SLAMF6, SPN, TBX21, and TNFα. In some instances, the VHH CDR1, VHH CDR2, and VHH CDR3 comprises the corresponding VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of any one of the forty clones set forth in Tables 3 to 7. In certain instances, the VHH specifically binds c-Met (human and/or cyno) and comprises: (a) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:41, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:42, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:43; (b) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:44, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:45, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:46; (c) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:47, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:48, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:49: (d) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:50, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:51, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:52;
(e) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:53, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:54, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:55; (f) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:56, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:57, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:58; (g) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:59, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:60, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:61; (h) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:62, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:63, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:64; (i) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:65, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:66, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:67; (j) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:68, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:69, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:70; (k) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:71, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:72, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:73; (l) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:74, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:75, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:76;
(m) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:77, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:78, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO79 ; (n) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:80, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:81, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:82; (o) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:83, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:84, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:85; (p) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:86, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:87, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:88; (q) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:89, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:90, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:91; (r) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:92, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:93, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:94; (s) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:95, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:96, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:97; (t) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:98, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:99, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:100;
(u) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:101, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:102, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:103; (v) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:104, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:105, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:106; (w) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:107, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:108, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:109; (x) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:110, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:111, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:112; (y) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:113, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:114, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:115 ; (z) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:116, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:117, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:118; (aa) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:119, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:120, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:121; (ab) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:122, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:123, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:124;
(ac) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:125, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:126, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:127; (ad) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:128, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:129, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:130; (ae) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:131, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:132, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:133; (af) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:134, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:135, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:136; (ag) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:137, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:138, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:139; (ah) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:140, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:141, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:142; (ai) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:143, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:144, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:145; (aj) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:146, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:147, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:148;
(ak) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:149, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:150, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:151; (al) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:152, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:153, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:154; (am) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:155, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:156, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:157; or (an) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:158, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:159, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:160. In some instances, the VHH specifically binds human c-MET and comprises a sequence that is at least 80%, at least 85%, at least 90%, at least 95%, or 100% identical to any VHH amino acid sequence set forth in Table 1. In certain cases, the VHH comprises a sequence that is set forth in any one of SEQ ID NOs.: 1 to 40. In some instances, the VHH specifically binds human c-MET and comprises a sequence that is at least 80%, at least 85%, at least 90%, at least 95%, or 100% identical to any amino acid sequence set forth in Table I. In certain cases, the VHH comprises a sequence that is set forth in any one of SEQ ID NOs.: 687 to 804, 811, or 813. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC50 of about 0.1 to about 250 nM when the VHH is in monovalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC50 of about 0.1 to about 100 nM when the VHH is in monovalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC50 of about 0.1 to about 75 nM when the VHH is in monovalent format. In some instances, the VHH binds
to human c-MET expressed on CHO cells with an EC50 of about 0.1 to about 50 nM when the VHH is in monovalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC50 of about 0.1 to about 25 nM when the VHH is in bivalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC50 of about 0.1 to about 15 nM when the VHH is in bivalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC50 of about 0.1 to about 10 nM when the VHH is in bivalent format. In some instances, the VHH binds to human c-MET expressed on CHO cells with an EC50 of about 0.1 to about 5 nM when the VHH is in bivalent format. In another aspect, the disclosure features a bispecific antibody comprising the polypeptide or VHH described above; and a binding molecule that binds to a different epitope of c-MET than the polypeptide or VHH described above, or binds to a different antigen. In a further aspect, the disclosure relates to a binding polypeptide that binds to two, or at least two, different epitopes or antigens. The binding polypeptide comprises a means for binding human and cynomolgus c-MET. The binding polypeptide also comprises a polypeptide that binds to a different epitope or antigen. In some cases, the polypeptide is a VHH. In other cases, the polypeptide is a scFv. In yet other cases, the polypeptide is a Fab. In some cases, the means for binding human and cynomolgus c- MET is linked to the polypeptide via a peptide linker (e.g., glycine linker, serine linker, glycine-serine linker). In some cases the linker is (G4S)n where n = 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 (SEQ ID NO: 648). In some cases the linker is G4S (SEQ ID NO: 806), (G4S)3 (SEQ ID NO: 649), or (G4S)5 (SEQ ID NO: 807). In some cases, the different antigen is an antigen on a T cell. In some cases, the different antigen is an antigen on an NK cell. In some cases, the different antigen is an antigen on a tumor cell.
In some instances, the different antigen is an antigen selected from the group consisting of: CD3, EGFR, VEGF, VEGFR, HGF, CXCR2, CXCR4, CXCR7, CXCL11, CXCL12, CEA, PSMA, MMR, PD-1, PDL-1, survivin, HER2, HGFR, P2X7, death receptor 5 CTLA4, CD7, CD8, CD11b, CD20, CD38, CD45, M HC-II, fibronectin, TUFM, CapG, CAIX, CD33, CD47, ARTC22, B3GAT1 (CD57), CCR7 (CD197), CD16, CD16a, CD16b, CD2 CD226, CD244, CD27, CD3, CD300A, CD34, CD58, CD59, CD69, CSF2, CX3CR1, CXCR1 (CD128), CXCR3 (CD183), CXCR4, EOMES, GZMB, ICAM1 (CD54), IFNG, IL-15R, IL-1R, IL22, IL-2RB (CD122), IL-7R (CD127), ITGA1 (CD49a), ITGA2 (CD49b), ITGAL (CD11a), ITGAM (CD11b), ITGB2 (CD18), KIR, KIR2DL1, KIR2DL2, KIT (CD117), KLRB1C, KLRC1, KLRC2, KLRD1 (CD94), KLRF1, KLRG1, KLRK1, LILRB1, KLRA4, KLRA8, MICA/BNCAM1 (CD56), NK2D, NKP46 (NCR1, CD335), NCR2, NCR3 (CD337), PRF1, SELL (CD62L), SIGLEC7, SLAMF6, SPN, TBX21, and TNFα. In some instances, the binding molecule is a VHH. In other instances, the binding molecule is a scFv. In some instances, a first human Ig Fc domain is directly or indirectly linked to the C-terminus of the polypeptide and a second human Ig Fc domain is directly or indirectly linked to the C-terminus of the binding molecule. In some cases, a first human Ig hinge domain is linked to the C-terminus of the polypeptide and a second human Ig hinge domain linked to the C-terminus of the binding molecule, and a first human Ig Fc domain linked to the C-terminus of the human Ig hinge domain and a second human Ig Fc domain to the C-terminus of the human Ig hinge domain. In certain cases, the first human Ig and second human Ig are a human IgG1, human IgG2, human IgG3, or human IgG4, or variants thereof. In some instances, the variant differs from the native human IgG hinge and/or Fc sequence by 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10 amino acids. In some cases, the C-terminal of the polypeptide and the C-terminal of the binding molecule are each linked to a sequence comprising an amino acid sequence that is at least 80%, at least 85%, at least 90%, at least 95%, or 100% identical to the IgG4 PAA hinge and Fc domain set forth in SEQ ID NO:643. In some cases, the first human Ig Fc domain and the second
human Ig Fc domain include mutations that promote heterodimerization between the first human Ig Fc domain and the second human Ig Fc domain (e.g., knob-into hole, electrostatic steering). In another aspect, the disclosure features a c-MET binding chimeric antigen receptor (CAR) comprising any one or more of the above-described polypeptides or VHHs. In some cases, the CAR comprises a c-MET binding domain, a transmembrane domain, and an intracellular signaling domain. In some cases, the transmembrane domain is from a CD3, CD4, CD8, or CD28 molecule. In some cases, the intracellular signaling domain is from CD3-zeta. In some cases, the c-MET binding CAR, further comprises CD28 and CD137 signaling domains and CD3ζ (c-Met-28-137-3ζ). In certain cases, the c-MET CAR comprises a CD28 hinge region, a CD28 transmembrane domain, and a FCεR1γ intracellular T cell signaling domain, wherein the CD28 transmembrane domain is connected to a FCεR1γ intracellular T cell signaling domain. In some cases, the c-MET binding CAR is expressed on the surface of a T cell. In other cases, the c-MET binding CAR is expressed on the surface of an NK cell. In some instances, the CAR comprises a means for binding human and cynomolgus c-MET, a transmembrane domain, and an intracellular signaling domain. In yet another aspect, the disclosure features a nucleic acid or nucleic acids encoding the polypeptide or VHH described above, the bispecific antibody described above, or the c-MET binding CAR described above. In another aspect, provided are a vector or vectors comprising the nucleic acid or nucleic acids described above. Also provided is a host cell comprising the nucleic acid or nucleic acids of claim 8, or the vector or vectors described above. In yet another aspect, the disclosure features a method of making a polypeptide, a bispecific antibody, or the c-MET binding CAR. The method comprises culturing a host cell described above under conditions that facilitate expression of the polypeptide, the bispecific antibody, or the c-MET binding CAR. In some cases, the method further
involves isolating the polypeptide, the bispecific antibody, or the c-MET binding CAR. In certain cases, the method further comprises formulating the polypeptide, the bispecific antibody, or the c-MET binding CAR as a sterile pharmaceutical composition. In another aspect, provided herein is a pharmaceutical composition comprising the polypeptide, the bispecific antibody, or the c-MET binding CAR described herein, and a pharmaceutically acceptable carrier. In yet another aspect, the disclosure relates to a pharmaceutical composition comprising a pharmaceutically acceptable carrier and a means for binding human and cynomolgus c-MET. In another aspect, the disclosure features a method of treating a c-MET expressing cancer in a human subject in need thereof, or killing a tumor cell in a human subject in need thereof, or decreasing the rate of tumor growth in a human subject in need thereof. The method comprises administering to the human subject a therapeutically effective amount of the polypeptide o, the bispecific antibody, or a T cell or NK cell expressing the c-MET binding CAR described herein In some cases, the cancer is a solid tumor. In certain cases, the cancer is selected from the group consisting of gastric cancer, esophageal cancer, medulloblastoma, glioma, colon cancer, head and neck cancer, lung cancer, kidney cancer, thyroid cancer, colorectal cancer, pancreatic cancer, ovarian cancer, and breast cancer. In one aspect, the disclosure features a mRNA comprising an open reading frame (ORF) encoding a VHH, a polypeptide, a bispecific antibody, or a c-MET binding CAR described herein. In some cases, the mRNA comprises a 5’terminal cap, a 5’UTR, a 3’UTR, and a poly A region. In another aspect, the disclosure features a polynucleotide comprising an mRNA comprising: (i) a 5' UTR; (ii) an open reading frame (ORF) encoding a polypeptide, the bispecific antibody, or the c-MET binding CAR described herein; (iii) a stop codon; and (iv) a 3' UTR.
In some cases, the mRNA comprises a microRNA (miR) binding site. In some cases, the microRNA is expressed in an immune cell of hematopoietic lineage or a cell that expresses TLR7 and/or TLR8 and secretes pro-inflammatory cytokines and/or chemokines. In certain cases, the microRNA binding site is for a microRNA selected from miR-126, miR-142, miR-144, miR-146, miR-150, miR-155, miR-16, miR-21, miR- 223, miR-24, miR-27, miR-26a, or any combination thereof. In some cases, the microRNA binding site is for a microRNA selected from miR126-3p, miR-142-3p, miR- 142-5p, miR-155, or any combination thereof. In certain cases, the microRNA binding site is located in the 3' UTR of the mRNA. In some instances, the 5' UTR comprises a nucleic acid sequence at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:654. In some instances, the 3' UTR comprises a nucleic acid sequence at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:674. In some instances, wherein the mRNA comprises a 5' terminal cap. In some cases, the 5' terminal cap comprises a Cap0, Cap1, ARCA, inosine, N1-methyl- guanosine, 2'-fluoro-guanosine, 7-deaza-guanosine, 8-oxo-guanosine, 2-amino- guanosine, LNA-guanosine, 2-azidoguanosine, Cap2, Cap4, 5' methylG cap, or an analog thereof. In some instances, the 5’cap is or comprises m7GpppGm. In certain instances, the mRNA comprises a poly-A region. In some cases, the poly-A region is at least about 10, at least about 20, at least about 30, at least about 40, at least about 50, at least about 60, at least about 70, at least about 80, at least about 90 nucleotides in length, or at least about 100 nucleotides in length. In some cases, the poly- A region is about 10 to about 200, about 20 to about 180, about 50 to about 160, about 70 to about 140, or about 80 to about 120 nucleotides in length. In some instances, the mRNA comprises at least one chemically modified nucleobase, sugar, backbone, or any combination thereof. In certain cases, the at least
one chemically modified nucleobase is selected from the group consisting of pseudouracil (Ψ), N1-methylpseudouracil (m1Ψ), 1-ethylpseudouracil, 2-thiouracil (s2U), 4’-thiouracil, 5-methylcytosine, 5-methyluracil, 5-methoxyuracil, and any combination thereof. In some cases, at least about 25%, at least about 30%, at least about 40%, at least about 50%, at least about 60%, at least about 70%, at least about 80%, at least about 90%, at least about 95%, at least about 99%, or 100% of the uracils are N1-methylpseudouracils. In some instances, the open reading frame consists of nucleosides selected from the group consisting of (i) uridine or a modified uridine, (ii) cytidine or a modified cytidine, (iii) adenosine or a modified adenosine, and (iv) guanosine or a modified guanosine. In some cases, the modified uridine is 1-methylpseudouridine. In some instances, the mRNA comprises a 5’terminal cap comprising m7GpppGm and a poly-A region 100 nucleotides in length. In some instances, all uracils of the polynucleotide are N1-methylpseudouracils. In one aspect, the disclosure provides a delivery agent and a polynucleotide(s) described herein. In some cases, the polynucleotide(s) are encapsulated in a delivery agent. In some cases, the delivery agent comprises a lipid nanoparticle. In some cases, the lipid nanoparticle has a mean particle size of from 80 nm to 160 nm. In certain cases, the lipid nanoparticle has a polydispersity index (PDI) of from 0.02 to 0.2 and/or a lipid:nucleic acid ratio of from 10 to 20. In some cases, the lipid nanoparticle comprises a neutral lipid, an ionizable amino lipid, a polyethyleneglycol (PEG) lipid, and/or a sterol. In certain cases, the neutral lipid is 1,2-distearoyl-sn-glycero-3-phosphocholine. In some cases, the ionizable amino lipid is a compound of Formula (I). In certain cases, the PEG lipid is PEG 2000 dimyristoyl glycerol or (134-hydroxy- 3,6,9,12,15,18,21,24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96,99,102,105,108,111,114,117,120,123,126,129,132- tetratetracontaoxatetratriacontahectyl stearate.
In some cases, the sterol is cholesterol, adosterol, agosterol A, atheronals, avenasterol, azacosterol, blazein, cerevisterol, colestolone, cycloartenol, daucosterol, 7- dehydrocholesterol, 5-dehydroepisterol, 7-dehydrositosterol, 20α,22R- dihydroxycholesterol, dinosterol, epibrassicasterol, episterol, ergosterol, ergosterol, fecosterol, fucosterol, fungisterol, ganoderenic acid, ganoderic acid, ganoderiol, ganodermadiol, 7α-hydroxycholesterol, 22R-hydroxycholesterol, 27-hydroxycholesterol, inotodiol, lanosterol, lathosterol, lichesterol, lucidadiol, lumisterol, oxycholesterol, oxysterol, parkeol, saringosterol, spinasterol, sterol ester, trametenolic acid, zhankuic acid, or zymosterol. In some cases, the sterol is cholesterol. In some cases, the lipid nanoparticle used herein comprises Compound I-1, 134- hydroxy- 3,6,9,12,15,18,21,24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96,99,102,105,108,111,114,117,120,123,126,129,132- tetratetracontaoxatetratriacontahectyl stearate, cholesterol, and ,2-distearoyl-sn-glycero- 3-phosphocholine (DSPC). In another aspect, the disclosure features a pharmaceutical composition comprising a polynucleotide described herein, and a delivery agent. In some cases, the delivery agent comprises a lipid nanoparticle. In some cases, the lipid nanoparticle has a mean particle size of from 80 nm to 160 nm. In certain cases, the lipid nanoparticle has a polydispersity index (PDI) of from 0.02 to 0.2 and/or a lipid:nucleic acid ratio of from 10 to 20. In some cases, the lipid nanoparticle comprises a neutral lipid, an ionizable amino lipid, a polyethyleneglycol (PEG) lipid, and/or a sterol. In certain cases, the neutral lipid is 1,2-distearoyl-sn-glycero-3-phosphocholine. In some cases, the ionizable amino lipid is a compound of Formula (I) (e.g., Compound I-1).
In certain cases, the PEG lipid is PEG 2000 dimyristoyl glycerol or 134-hydroxy- 3,6,9,12,15,18,21,24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96,99,102,105,108,111,114,117,120,123,126,129,132- tetratetracontaoxatetratriacontahectyl stearate. In some cases, the sterol is cholesterol, adosterol, agosterol A, atheronals, avenasterol, azacosterol, blazein, cerevisterol, colestolone, cycloartenol, daucosterol, 7- dehydrocholesterol, 5-dehydroepisterol, 7-dehydrositosterol, 20α,22R- dihydroxycholesterol, dinosterol, epibrassicasterol, episterol, ergosterol, ergosterol, fecosterol, fucosterol, fungisterol, ganoderenic acid, ganoderic acid, ganoderiol, ganodermadiol, 7α-hydroxycholesterol, 22R-hydroxycholesterol, 27-hydroxycholesterol, inotodiol, lanosterol, lathosterol, lichesterol, lucidadiol, lumisterol, oxycholesterol, oxysterol, parkeol, saringosterol, spinasterol, sterol ester, trametenolic acid, zhankuic acid, or zymosterol. In some cases, the sterol is cholesterol. In some cases, the lipid nanoparticle used herein comprises Compound I-1, 134- hydroxy- 3,6,9,12,15,18,21,24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96,99,102,105,108,111,114,117,120,123,126,129,132- tetratetracontaoxatetratriacontahectyl stearate, cholesterol, and ,2-distearoyl-sn-glycero- 3-phosphocholine (DSPC). In another aspect, the disclosure features a method of treating a c-MET expressing cancer in a human subject in need thereof, killing a tumor cell in a human subject in need thereof, or decreasing the rate of tumor growth in a human subject in need thereof. The method comprises administering to the human subject a therapeutically effective amount of the mRNA or pharmaceutical composition comprising the mRNA described herein. The mRNA or pharmaceutical composition comprising the mRNA described herein may be encapsulated in a LNP. In some cases, the cancer is a solid tumor. In certain cases, the cancer is selected from the group consisting of gastric cancer, esophageal cancer, medulloblastoma, glioma,
colon cancer, head and neck cancer, lung cancer, kidney cancer, thyroid cancer, colorectal cancer, pancreatic cancer, ovarian cancer, and breast cancer. In some cases, the lipid nanoparticle used herein comprises Compound I-1, 134- hydroxy- 3,6,9,12,15,18,21,24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96,99,102,105,108,111,114,117,120,123,126,129,132- tetratetracontaoxatetratriacontahectyl stearate, cholesterol, and ,2-distearoyl-sn-glycero-3- phosphocholine (DSPC). In another aspect, the disclosure features a kit comprising (i) the polypeptide; the bispecific antibody; the c-MET binding CAR; the pharmaceutical composition; or the polynucleotide described herein, and (ii) a package insert instructing a user of the kit to administer the polypeptide, bispecific antibody, c-Met binding CAR, pharmaceutical composition, or polynucleotide to a human subject in need thereof. BRIEF DESCRIPTION OF THE DRAWINGS FIGURE 1 provides FACS binding curves for anti-c-MET VHH-His (in the monovalent format) to determine EC50 values against CHO Human c-MET cells. FIGURE 2 provides FACS binding curves for anti-c-MET VHH-His (in the monovalent format) to determine EC50 values against CHO Cyno c-MET cells. FIGURE 3 provides FACS binding curves for anti-c-MET VHH-IgG4PAA-His (in the bivalent format) to determine EC50 values against CHO Human c-MET cells. FIGURE 4 provides FACS binding curves for anti-c-MET VHH-IgG4PAA-His (in the bivalent format) to determine EC50 values against CHO Cyno c-MET cells. FIGURE 5 provides FACS binding curves for anti-c-MET VHH-His (in the monovalent format) to determine EC50 values against HS746T c-MET cancer cells. FIGURE 6 provides FACS HGF competition assay for anti-c-MET-VHH Abs (in the bivalent format) using CHO Human c-MET cells.
FIGURES 7A to 7C provides sequence alignments to highlight residues in the llama anti-c-MET VHHs which were modified during the humanization process. Figure 7A discloses SEQ ID NOS 4, 687-692, 22, 693-698, 813, 33, 699-703, 28, 704-709, 7, 710-716, 3, 717-723, and 811 respectively, in order of appearance. Figure 7B discloses SEQ ID NOS 40, 724-730, 11, 731-737, 32, 738-743, 37, 744-748, 9, 751-755, 24, 756- 760, 20, and 794-797, respectively, in order of appearance. Figure 7C discloses SEQ ID NOS 10, 761-766, 5, 767-773, 30, 774-780, 8, 781-787, 2, 788-793, 16, and 799-804, respectively, in order of appearance. FIGURES 8A to 8C provides FACS binding curves for humanized anti-c-MET VHH- His to determine EC50 values against CHO Hu c-MET cells. FIGURE 9 provides tables summarizing the monovalent EC50 (nM) values for humanized constructs of anti-c-MET VHH clones on CHO Hu c-MET cells. DETAILED DESCRIPTION Featured herein are novel binding polypeptides and antibodies that specifically bind c-MET (e.g., human and/or cyno), polypeptides, nucleic acids, and host cells comprising these c-Met antibodies, and methods of using these antibodies in the treatment of diseases such as cancer. More specifically, the c-MET antibodies described herein are variable domains of the heavy chain of heavy chain-only antibodies (VHHs), also known as single domain antibodies or nanobodies. These antibodies bind with high affinity to cells expressing c-MET. Most of these antibodies bind to c-MET when the VHH is in a monovalent format. All of the anti-c-MET antibodies disclosed herein bind to c-MET when the VHH is in a bivalent format. Some of these bivalent c-MET VHHs compete with HGF for binding c-MET; others are semi-competitive for binding to c- MET; while still others do not compete with HGF for binding c-MET. These VHHs either unmodified or modified (e.g., by humanization) can be used as a component of monovalent, bivalent, bispecific, or multispecific antibodies. In addition, they can be conjugated to other agents (e.g., detectable agents, drugs, radionuclides, antibodies, antigen-binding fragments, Fc domains, half-life extending moieties). For example, the
VHHs of this disclosure may be part of an antibody drug conjugate (ADC). They may also be used as the antigen-binding component of a chimeric antigen receptor (CAR). These VHH containing polypeptides are used in the treatment of cancers having dysregulation of the c-MET/HGF pathway. In some cases, these VHH containing polypeptides are used in the treatment of cancers with one or more of a c-MET mutation(s), a c-MET amplification(s), or c-MET overexpression. Definitions “VHH” means a variable domain of the heavy chain of heavy chain-only antibodies that are produced by camelids. They are monovalent and do not include a light chain. The VHH can be used as is or humanized. The VHH can also be made bivalent or multivalent by e.g., fusing to sequences that facilitate dimerization, trimerization, tetramerization, pentamerization, or by being linked to a whole antibody or an antigen- binding fragment. VHHs are also referred to as single domain antibodies and nanobodies. “Monovalent” means a binding molecule with a single antigen binding domain (e.g., a single VHH). “Bivalent” means a binding molecule with two antigen binding domains (e.g., two VHHs linked together or brought together by a dimerization domain such as a polypeptide comprising a human Ig Fc). “% identity” is the percentage of identical residues between two or more sequences and is determined over the full length of the sequences being compared. It is a function of the number of amino acids or nucleotide residues that are identical in the sequences being compared. Identical residues are defined as residues that are the same in the two sequences in a given position of the alignment of the sequences. The percentage of sequence identity is calculated from the optimal alignment by taking the number of residues identical between two sequences dividing it by the total number of residues in the shortest sequence and multiplying by 100. The optimal alignment is the alignment in which the percentage of identity is the highest possible. Gaps may be introduced into one or both sequences in one or more positions of the alignment to obtain the optimal
alignment. These gaps are then taken into account as non-identical residues for calculation of the percentage of sequence identity. Alignment for purposes of determining the percentage of amino acid or nucleic acid sequence identity can be achieved in various ways using computer programs. For example, the BLAST program (Tatiana t al., FEMS Microbiol. Lett., 174:247-250 (1999)) set to the default parameters, available from the National Center for Biotechnology Information (NCBI) at ncbi.nlm.nih.gov/BLAST/bl2seq/wblast2.cgi, can be used to obtain an optimal alignment of amino acid or nuclic acid sequences to calculate the percentage of sequence identity. “about” means + or – 10% of the recited value, so “about 250 nM” means 225 nM to 275 nM. c-Met Mesenchymal-epithelial transition factor (c-MET) function is vital during various morphogenetic events in both embryonic and adult stages of development. c-MET is a tyrosine kinase receptor which belongs to the MET (MNNG HOS transforming gene) family, and is usually expressed on the surface of various epithelial cells. After binding its ligand, hepatocyte growth factor (HGF), c-MET activates a wide range of different cellular signaling pathways, including those involved in proliferation, motility, migration and invasion. Although c-MET is important in the control of tissue homeostasis under normal physiological conditions, it has also been found to be aberrantly activated in human cancers via mutation, amplification, or protein overexpression. Expression of the c-MET receptor tyrosine kinase and its ligand, hepatocyte growth factor (HGF), has been observed in tumor biopsies of most solid tumors and c-MET signaling has been documented in a wide range of human malignancies (e.g., mesenchyme-derived tumors such as gastric cancer and papillary renal cell carcinoma; metastatic head and neck squamous cell carcinoma, hepatocellular carcinoma, ovarian cancer, lung cancer, and glioma).
The c-MET receptor is formed by proteolytic processing of a common precursor into a single-pass, disulfide-linked α/β heterodimer. The extracellular portion of c-MET is composed of three domain types. The N-terminal 500 residues fold to form a large semaphorin (Sema) domain, which encompasses the whole α-subunit and part of the β- subunit. The Sema domain shares sequence homology with domains found in the semaphorin and plexin families. The PSI domain (found in plexins, semaphorins and integrins) follows the Sema domain, spans approximately 50 residues and includes four disulphide bonds. This domain is connected to the transmembrane helix via four immunoglobulin–plexin–transcription (IPT) domains, which are related to immunoglobulin-like domains and are found in integrins, plexins and transcription factors. Intracellularly, the c-MET receptor contains a tyrosine kinase catalytic domain flanked by distinctive juxtamembrane and carboxy-terminal sequences. When HGF is recognized by c-MET immunoglobulin-like domains and binds the extracellular portion of MET ȕ domain, two c-MET heterodimers dimerize, leading to self-phosphorylation of two tyrosine residues within the kinase catalytic domain (Tyr1234, Tyr1235). The heterodimerization leads to the assembling of SEMA domain, the main binding site for the HGF ligand. The polypeptide sequence of full length human c-MET is provided below: MKAPAVLAPGILVLLFTLVQRSNGECKEALAKSEMNVNMKYQLPNFTAETPIQN VILHEHHIFLGATNYIYVLNEEDLQKVAEYKTGPVLEHPDCFPCQDCSSKANLSG GVWKDNINMALVVDTYYDDQLISCGSVNRGTCQRHVFPHNHTADIQSEVHCIFS PQIEEPSQCPDCVVSALGAKVLSSVKDRFINFFVGNTINSSYFPDHPLHSISVRRLK ETKDGFMFLTDQSYIDVLPEFRDSYPIKYVHAFESNNFIYFLTVQRETLDAQTFHT RIIRFCSINSGLHSYMEMPLECILTEKRKKRSTKKEVFNILQAAYVSKPGAQLARQ IGASLNDDILFGVFAQSKPDSAEPMDRSAMCAFPIKYVNDFFNKIVNKNNVRCLQ HFYGPNHEHCFNRTLLRNSSGCEARRDEYRTEFTTALQRVDLFMGQFSEVLLTSI STFIKGDLTIANLGTSEGRFMQVVVSRSGPSTPHVNFLLDSHPVSPEVIVEHTLNQ NGYTLVITGKKITKIPLNGLGCRHFQSCSQCLSAPPFVQCGWCHDKCVRSEECLS GTWTQQICLPAIYKVFPNSAPLEGGTRLTICGWDFGFRRNNKFDLKKTRVLLGNE SCTLTLSESTMNTLKCTVGPAMNKHFNMSIIISNGHGTTQYSTFSYVDPVITSISPK YGPMAGGTLLTLTGNYLNSGNSRHISIGGKTCTLKSVSNSILECYTPAQTISTEFA VKLKIDLANRETSIFSYREDPIVYEIHPTKSFISGGSTITGVGKNLNSVSVPRMVIN
VHEAGRNFTVACQHRSNSEIICCTTPSLQQLNLQLPLKTKAFFMLDGILSKYFDLI YVHNPVFKPFEKPVMISMGNENVLEIKGNDIDPEAVKGEVLKVGNKSCENIHLHS EAVLCTVPNDLLKLNSELNIEWKQAISSTVLGKVIVQPDQNFTGLIAGVVSISTAL LLLLGFFLWLKKRKQIKDLGSELVRYDARVHTPHLDRLVSARSVSPTTEMVSNE SVDYRATFPEDQFPNSSQNGSCRQVQYPLTDMSPILTSGDSDISSPLLQNTVHIDL SALNPELVQAVQHVVIGPSSLIVHFNEVIGRGHFGCVYHGTLLDNDGKKIHCAVK SLNRITDIGEVSQFLTEGIIMKDFSHPNVLSLLGICLRSEGSPLVVLPYMKHGDLRN FIRNETHNPTVKDLIGFGLQVAKGMKYLASKKFVHRDLAARNCMLDEKFTVKV ADFGLARDMYDKEYYSVHNKTGAKLPVKWMALESLQTQKFTTKSDVWSFGVL LWELMTRGAPPYPDVNTFDITVYLLQGRRLLQPEYCPDPLYEVMLKCWHPKAE MRPSFSELVSRISAIFSTFIGEHYVHVNATYVNVKCVAPYPSLLSSEDNADDEVDT RPASFWETS (SEQ ID NO:641) The polypeptide sequence of full length cynomolgus c-MET is provided below: MKSKSKGLAECLTHDKPLTMKAPAVLVPGILVLLFTLVQRSNGECKEALAKSEM NVNMKYQLPNFTAETAIQNVILHEHHIFLGATNYIYVLNEEDLQKVAEYKTGPVL EHPDCFPCQDCSSKANLSGGVWKDNINMALVVDTYYDDQLISCGSVNRGTCQR HVFPHNHTADIQSEVHCIFSPQIEEPNQCPDCVVSALGAKVLSSVKDRFINFFVGN TINSSYFPHHPLHSISVRRLKETKDGFMFLTDQSYIDVLPEFRDSYPIKYIHAFESN NFIYFLTVQRETLNAQTFHTRIIRFCSLNSGLHSYMEMPLECILTEKRKKRSTKKE VFNILQAAYVSKPGAQLARQIGASLNDDILFGVFAQSKPDSAEPMDRSAMCAFPI KYVNDFFNKIVNKNNVRCLQHFYGPNHEHCFNRTLLRNSSGCEARRDEYRAEFT TALQRVDLFMGQFSEVLLTSISTFVKGDLTIANLGTSEGRFMQVVVSRSGPSTPH VNFLLDSHPVSPEVIVEHPLNQNGYTLVVTGKKITKIPLNGLGCRHFQSCSQCLSA PPFVQCGWCHDKCVRSEECPSGTWTQQICLPAIYKVFPTSAPLEGGTRLTICGWD FGFRRNNKFDLKKTRVLLGNESCTLTLSESTMNTLKCTVGPAMNKHFNMSIIISN GHGTTQYSTFSYVDPIITSISPKYGPMAGGTLLTLTGNYLNSGNSRHISIGGKTCTL KSVSNSILECYTPAQTISTEFAVKLKIDLANRETSIFSYREDPIVYEIHPTKSFISGGS TITGVGKNLHSVSVPRMVINVHEAGRNFTVACQHRSNSEIICCTTPSLQQLNLQLP LKTKAFFMLDGILSKYFDLIYVHNPVFKPFEKPVMISMGNENVLEIKGNDIDPEA VKGEVLKVGNKSCENIHLHSEAVLCTVPNDLLKLNSELNIEWKQAISSTVLGKVI VQPDQNFTGLIAGVVSISIALLLLLGLFLWLKKRKQIKDLGSELVRYDARVHTPH LDRLVSARSVSPTTEMVSNESVDYRATFPEDQFPNSSQNGSCRQVQYPLTDMSPI LTSGDSDISSPLLQNTVHIDLSALNPELVQAVQHVVIGPSSLIVHFNEVIGRGHFGC VYHGTLLDNDGKKIHCAVKSLNRITDIGEVSQFLTEGIIMKDFSHPNVLSLLGICL RSEGSPLVVLPYMKHGDLRNFIRNETHNPTVKDLIGFGLQVAKGMKYLASKKFV HRDLAARNCMLDEKFTVKVADFGLARDMYDKEYYSVHNKTGAKLPVKWMAL ESLQTQKFTTKSDVWSFGVLLWELMTRGAPPYPDVNTFDITVYLLQGRRLLQPE YCPDPLYEVMLKCWHPKAEMRPSFSELVSRISAIFSTFIGEHYVHVNATYVNVKC VAPYPSLLSSEDNADDEVDT (SEQ ID NO:642)
VHHs that Specifically Bind c-Met Provided herein are novel VHHs that specifically bind to c-MET. In some cases, the c-MET may be from human or cynomolgus monkey. Table 1 provides the amino acid sequences of several novel VHHs. This disclosure features a binding molecule or polypeptide that comprises the VHH amino acid sequence of any of the clones listed in Table 1. In some instances, the binding molecule or polypeptide comprises a VHH that specifically binds c-MET (e.g., human and/or cyno) and comprises a sequence that is at least 80%, at least 85%, at least 90%, or at least 95% identical to an amino acid sequence set forth in any one of SEQ ID NOs.: 1 to 40. In some instances, the binding molecule or polypeptide comprises a VHH that specifically binds c-MET (e.g., human and cyno) and comprises an amino acid sequence set forth in any one of SEQ ID NO:1 to 40 except for having 1, 2, 34, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 457, 48, 49, or 50 amino acid substitutions. The variability of the above sequences from an amino acid sequence set forth in any one of SEQ ID NO:1 to 40 may be due to humanization of the VHH. The skilled person could use the alignments of Figures 7A, 7B, and 7C to determine which VHH residue or residues to substitute with a different amino acid(s). These variant VHHs still specifically bind to c-MET (e.g., human and/or cyno). In some instances, the binding molecule or polypeptide specifically binds c-MET (e.g., human and/or cyno) and comprises a sequence that is identical to an amino acid sequence set forth in any one of SEQ ID NOs.: 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, or 40. In some instances, the binding molecule or polypeptide is in a monovalent format. In some cases, the binding molecule or polypeptide binds a cell expressing human or cyno c-MET (e.g., a cell line such as CHO or a cancer cell such as HS746T) with an EC50 of about 0.1 nM to about 250 nM. In some cases, the binding molecule or polypeptide binds a cell expressing human or cyno c-MET (e.g., a cell line such as CHO or a cancer cell such as HS746T) with an EC50 of about 0.5 nM to about 100 nM. In
some cases, the binding molecule or polypeptide binds a cell expressing human or cyno c-MET (e.g., a cell line such as CHO or a cancer cell such as HS746T) with an EC50 of about 0.5 nM to about 50 nM. In some cases, the binding molecule or polypeptide binds a cell expressing human or cyno c-MET (e.g., a cell line such as CHO or a cancer cell such as HS746T) with an EC50 of about 0.5 nM to about 25 nM. In some cases, the binding molecule or polypeptide binds a cell expressing human or cyno c-MET (e.g., a cell line such as CHO or a cancer cell such as HS746T) with an EC50 of about 0.5 nM to about 10 nM. In some cases, the binding molecule or polypeptide does not bind, or binds weakly (e.g., where the EC50 cannot be measured), a cell expressing human or cyno c- MET in a monovalent format. For example, a binding molecule or polypeptide comprising the variable domain amino acid sequences in a monovalent format of clones CM18, CM23, and CM34 do not bind, or binds weakly, to human c-MET expressed on CHO cells. In addition, binding molecule or polypeptide comprising the variable domain amino acid sequences in a monovalent format of clones CM18, CM29, and CM36 do not bind, or binds weakly, to cyno c-MET expressed on CHO cells. Binding molecules or polypeptides comprising the amino acid sequence set forth in any one of SEQ ID NOs.: 1 to 40 in a monovalent format bind to a c-MET expressing cancer cell such as HS746T. In some instances, the binding molecule or polypeptide comprises the VHH in a bivalent format. In some cases, a bivalent format of the VHH is produced by linking the VHH to the same or another VHH. In some cases, a bivalent format of the VHH is arrived at by linking the C-terminus of the VHH of the binding molecule or polypeptide to the N-terminus of a hinge and Fc domain of a human immunoglobulin (e.g., IgA, IgG1, IgG2, IgG3, IgG4). In one case, the hinge and Fc domain is from human IgA. In another case, the hinge and Fc domain is from human IgG4. In some cases, the hinge and Fc domain comprises an amino acid sequence that is at least 80%, at least 85%, at least 90%, at least 95%, or 100% identical to the amino acid sequence of human IgG4PAA (SEQ ID NO:643). The amino acid sequence of human IgG4PAA is provided below (the hinge is shown in bold; the CH2 domain is italicized; and the CH3 domain is underlined; the CH2 and CH3 domains together form the Fc):
ESKYGPPCPPCPAPEAAGGPSVFLFPPKPKDTLMISRTPEVTCVVVDVSQEDPEVQ FNWYVDGVEVHNAKTKPREEQFNSTYRVVSVLTVLHQDWLNGKEYKCKVSNKGLPSS IEKTISKAKGQPREPQVYTLPPSQEEMTKNQVSLTCLVKGFYPSDIAVEWESNGQP ENNYKTTPPVLDSDGSFFLYSRLTVDKSRWQEGNVFSCSVMHEALHNHYTQKSL SLSLGK (SEQ ID NO:643) In some cases, the hinge and Fc domain comprises an amino acid sequence set forth in SEQ ID NO:643 except for 1, 2, 34, 5, 6, 7, 8, 9, or 10 amino acid substitutions. These substitutions can be made to e.g., alter effector function (e.g., increasing or decreasing effector function), or to promote heterodimerization. Such substitutions made as part of Fc domain engineering are well known in the art. See e.g., Liu H, Saxena A, Sidhu SS and Wu D (2017) Fc Engineering for Developing Therapeutic Bispecific Antibodies and Novel Scaffolds. Front. Immunol. 8:38. doi: 10.3389/fimmu.2017.00038; Wilkinson I, Anderson S, Fry J, Julien LA, Neville D, Qureshi O, et al. (2021) Fc-engineered antibodies with immune effector functions completely abolished. PLoS ONE 16(12): e0260954. doi.org/10.1371/journal.pone.0260954; Delidakis et al., Wilkinson I, Anderson S, Fry J, Julien LA, Neville D, Qureshi O, et al. (2021) Fc-engineered antibodies with immune effector functions completely abolished. PLoS ONE 16(12): e0260954. //doi.org/10.1371/journal.pone.0260954, all of which are incorporated by reference herein. In some cases, the binding molecule or polypeptide comprising a VHH linked to a human immunoglobulin hinge and a human immunoglobulin Fc - e.g., human IgA hinge + human IgG Fc or human IgG hinge and Fc (i.e., wherein the VHH is to be used in a bivalent format) binds a cell expressing human or cyno c-MET (e.g., a cell line such as CHO) with an EC50 of about 0.1 nM to about 25 nM. In some cases, the binding molecule or polypeptide comprising a VHH in bivalent format binds a cell expressing human or cyno c-MET (e.g., a cell line such as CHO) with an EC50 of about 0.1 nM to about 10 nM. In some cases, the binding molecule or polypeptide comprising a VHH in bivalent format binds a cell expressing human or cyno c-MET (e.g., a cell line such as CHO) with an EC50 of about 0.1 nM to about 5 nM. All of the anti-c-MET VHHs disclosed herein bind to c-MET when the VHH is in a bivalent format. Some of these
bivalent VHH constructs are competing – i.e., compete with HGF for binding c-MET (clones CM3, CM4, CM8, CM10, CM11, CM16, CM19, CM23, CM25, CM30, CM32, CM33, CM37, CM38, CM39, CM42); others are semi-competitive for binding to c-MET (CM12, CM14, CM17, CM18, CM21, CM24, CM31, CM40, CV7N1, CV13N2); while still others do not compete with HGF for binding c-MET (CM5, CM6, CM9, CM22, CM26, CM27, CM28, CM29, CM35, CM36, CM43). By “competes with HGF” means a more than 50% drop in MFI in presence of HGF. By “semi-competitive with HGF” means Less than 50% drop in MFI in presence of HGF.
Also encompassed by this disclosure are binding molecules or polypeptides that comprise a VHH that specifically binds c-MET (human and/or cyno), wherein the VHH comprises the VHH CDR1, VHH CDR2, and VHH CDR3 of the VHH of any one of the clones listed in Table 1. The VHH CDRs of the VHH can be based on any CDR definition. Exemplary CDR numbering systems are provided below as Table 2 (see, e.g., Lafrance et al,. Dev. Comp. Immunol., 27(1):55-77 (2003)). Table 2: Exemplary CDR Definitions
Provided in Tables 3 to 7 below are the VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of the clones according to IMGT, Kabat, Chothia, enhanced Chothia, and contact definitions. Table 3: VHH CDRs Based on IMGT CDR Definition
Table 4: VHH CDRs Based on Kabat CDR Definition
Table 5: VHH CDRs Based on Chothia CDR Definition
Table 6: VHH CDRs Based on Enhanced Chothia CDR Definition
Table 7: VHH CDRs Based on Contact CDR Definition
^ In some instances, the disclosure provides a binding molecule or polypeptide comprising a VHH that specifically binds c-MET (human and/or cyno), wherein the VHH comprises the VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of any one clone listed in Table 3. For example, the disclosure provides a binding molecule or polypeptide comprising a VHH that specifically binds c-MET (human and cyno), wherein the VHH comprises: (a) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:41, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:42, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:43; (b) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:44, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:45, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:46;
(c) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:47, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:48, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:49: (d) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:50, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:51, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:52; (e) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:53, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:54, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:55; (f) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:56, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:57, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:58; (g) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:59, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:60, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:61; (h) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:62, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:63, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:64; (i) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:65, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:66, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:67; (j) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:68, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:69, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:70;
(k) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:71, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:72, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:73; (l) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:74, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:75, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:76; (m) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:77, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:78, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO79 ; (n) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:80, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:81, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:82; (o) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:83, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:84, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:85; (p) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:86, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:87, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:88; (q) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:89, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:90, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:91; (r) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:92, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:93, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:94;
(s) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:95, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:96, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:97; (t) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:98, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:99, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:100; (u) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:101, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:102, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:103; (v) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:104, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:105, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:106; (w) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:107, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:108, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:109; (x) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:110, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:111, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:112; (y)^a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:113, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:114, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:115 ; (z) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:116, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:117, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:118;
(aa) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:119, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:120, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:121; (ab) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:122, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:123, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:124; (ac) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:125, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:126, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:127; (ad) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:128, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:129, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:130; (ae) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:131, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:132, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:133; (af) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:134, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:135, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:136; (ag) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:137, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:138, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:139; (ah) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:140, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:141, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:142;
(ai) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:143, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:144, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:145; (aj) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:146, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:147, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:148; (ak) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:149, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:150, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:151; (al) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:152, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:153, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:154; (am) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:155, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:156, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:157; or (an) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:158, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:159, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:160. In some instances, the disclosure provides a binding molecule or polypeptide comprising a VHH that specifically binds c-MET (human and cyno), wherein the VHH comprises the VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of any one clone listed in Table 4. In some instances, the disclosure provides a binding molecule or polypeptide comprising a VHH that specifically binds c-MET (human and cyno), wherein the VHH comprises the VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of any one clone listed in Table 5.
In some instances, the disclosure provides a binding molecule or polypeptide comprising a VHH that specifically binds c-MET (human and cyno), wherein the VHH comprises the VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of any one clone listed in Table 6. In some instances, the disclosure provides a binding molecule or polypeptide comprising a VHH that specifically binds c-MET (human and cyno), wherein the VHH comprises the VHH CDR1, VHH CDR2, and VHH CDR3 amino acid sequences of any one clone listed in Table 7. In some instances, the above VHHs are not humanized. In some instances, the above VHHs are humanized. Humanization may be by, e.g. resurfacing/veneering or CDR grafting. Given the high sequence identity of most VHHs with human IGHV3 family a comparison between the human IGHV3 framework sequence (e.g., from IGHV3-23) and the VHH can be used to make amino acid substitutions to replace the camelid amino acid with the amino acid in the corresponding human framework. AlphaFold2 and/or NanoNet can, for example, be used as tools to predict a structural model of the VHH from its sequence. See e.g., Vishwakarma et al., Intl. J. Mol. Sci., 2022, 23, 3721 (incorporated by reference herein). iCAN, Llamanade, and/or INDI can also be for assisting with VHH humanization. See, e.g., Zuo et al., BMC Genom., 2017, 18, 797; Sang et al., Structure, 2022, 30, 418-429.e3; Deszynski et al., Nucleic Acids Res., 2022, 50, D1273-D1281 (all of which are incorporated by reference herein). In some instances, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, or 50 framework amino acids of the VHH are substituted as a result of humanization. In some instances, full humanization is avoided. In some cases, less than 20, 15, 10, 8, 7, 6, 5, 4, or 3 amino acids, or 1 to 25, 1 to 20, 1 to 15, 1 to 10, or 1 to 5 amino acids in the VHH framework regions are humanized. In some cases, 1 to 15 amino acids in the VHH framework regions of the sequences in Table 1 are humanized. In some instances, one or more of IMGT positions 42, 49, 50, and 52 of the VHH are not humanized. In some instances, one or more of IMGT positions 42 and 52 of the VHH are
not humanized. In some instances, one or more of IMGT positions 42, 49, 50, and 52 of the VHH are humanized. In some instances, one or more of IMGT positions 49 and 50 of the VHH are humanized. In some instances, none of IMGT positions 42, 49, 50, and 52 of the VHH are humanized. In some instances, the VHH CDR1, VHH CDR2, and VHH CDR3 are grafted onto a universal humanized VHH scaffold, h-NbBCII10FGLA. See, e.g., Vincke et al., J. Biol. Chem., 284, 3273-3284 (2009), which is incorporated by reference herein. In some instances, the above VHHs are in a monovalent format. In some instances, the above VHHs are in a bivalent format (e.g., by linking the VHH to a hinge- Ig Fc domain). In some instances, the above VHHs are linked to a hinge-Fc of a human Ig (e.g., IgA, IgG1, IgG2, IgG3, or IgG4). In some instances, the above VHHs are linked to a hinge from human IgA. In some instances, the above VHHs are linked to a hinge from human IgG1 or IgG4. In some instances, the above VHHs comprise a Fc region from human IgG1 or human IgG4. In some instances, any of the above VHHs can be linked to a sequence that is at least 80%, at least 85%, at least 90%, or at least 95% identical to the amino acid sequence of human IgG4PAA (SEQ ID NO:643). In some instances, any of the above VHHs can be linked to a sequence that is identical to the amino acid sequence of human IgG4PAA (SEQ ID NO:643). In some instances, the human Ig Fc region is modified to e.g., alter (increase or decrease) the effector function and/or to promote heterodimerization. Exemplary Humanized C-MET VHHs This disclosure also encompasses humanized VHHs that specifically bind C- MET (human and/or cyno C-MET). Table I below provides the amino acid sequences of exemplary humanized VHHs that specifically bind C-MET (human and/or cyno C-MET). Table I. Amino Acid Sequences of Humanized VHHs that Bind C-MET
^ ^ In some instances, the disclosure provides a humanized VHH that specifically binds to C-MET (human and/or cyno) and which comprises an amino acid sequence that is at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to any one of the sequences of SEQ ID NOs.: 687 to 804, 811 or 813. In some cases, these humanized VHHs comprise the VHH CDR-1, VHH-CDR2, and VHH-CDR3 of the parental llama antibody. ^ In some instances, the disclosure provides a humanized VHH that specifically binds to C-MET (human and/or cyno) and which comprises an amino acid sequence that is identical to any one of the sequences of SEQ ID NOs.: 687 to 804, 811 or 813 except for 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 amino acid substitutions within the VHH sequence from Table I. In some cases, these humanized VHHs comprise the VHH CDR-1, VHH-CDR2,
and VHH-CDR3 of the parental llama antibody. In some cases, the substitutions are in one or more of the framework regions of the VHH. The disclosure also encompasses a binding molecule comprising a humanized VHH that specifically binds to C-MET (human and/or cyno) described above. In some cases, the binding molecule comprises a humanized VHH that has an amino acid sequence set out in any one of SEQ ID NOs.: 687 to 804, 811 or 813. In some cases, the binding molecule is multispecific. In certain cases, the binding molecule is bispecific and comprises a second binding moiety (e.g., a second VHH) that binds a different antigen. In some cases, the different antigen is a tumor associated antigen. In some cases, the different antigen is a molecule expressed on T cells or NK cells. In some instances, the above humanized VHHs are in a monovalent format. In some instances, the above humanized VHHs are in a bivalent format (e.g., by linking the VHH to a hinge-Ig Fc domain). In some instances, the above humanized VHHs are linked to a hinge-Fc of a human Ig (e.g., IgA, IgG1, IgG2, IgG3, or IgG4). In some instances, the above humanized VHHs are linked to a hinge from human IgA. In some instances, the above humanized VHHs are linked to a hinge from human IgG1 or IgG4. In some instances, the above humanized VHHs comprise a Fc region from human IgG1 or human IgG4. In some instances, any of the above humanized VHHs can be linked to a sequence that is at least 80%, at least 85%, at least 90%, or at least 95% identical to the amino acid sequence of human IgG4PAA (SEQ ID NO:643). In some instances, any of the above humanized VHHs can be linked to a sequence that is identical to the amino acid sequence of human IgG4PAA (SEQ ID NO:643). In some instances, the human Ig Fc region is modified to e.g., alter (increase or decrease) the effector function and/or to promote heterodimerization. Bispecific Constructs Other binding molecules are also encompassed by this disclosure. For example, the c-MET binding VHHs of this disclosure may be part of a bispecific construct (i.e., a construct that binds to two different epitopes). AVHH of this disclosure can be linked to
itself or to one or more VHHs of this disclosure. In some instances, a VHH of this disclosure may be linked to a VHH that binds to a different epitope of c-MET and/or to one or more other antigens. See, e.g., Verhaar et al., Semin. Immunol., 2021; 52:101425 (incorporated by reference herein). For example the other antigen may be a T cell antigen (e.g., CD3), a NK cell antigen (e.g., B3GAT1 (CD57), CCR7 (CD197), CD16, CD16a, CD16b, CD2 CD226, CD244, CD27, CD3, CD300A, CD34, CD58, CD59, CD69, CSF2, CX3CR1, CXCR1 (CD128), CXCR3 (CD183), CXCR4, EOMES, GZMB, ICAM1 (CD54), IFNG, IL-15R, IL-1R, IL22, IL-2RB (CD122), IL-7R (CD127), ITGA1 (CD49a), ITGA2 (CD49b), ITGAL (CD11a), ITGAM (CD11b), ITGB2 (CD18), KIR, KIR2DL1, KIR2DL2, KIT (CD117), KLRB1C, KLRC1, KLRC2, KLRD1 (CD94), KLRF1, KLRG1, KLRK1, LILRB1, KLRA4, KLRA8, MICA/BNCAM1 (CD56), NK2D, NKP46 (NCR1, CD335), NCR2, NCR3 (CD337), PRF1, SELL (CD62L), SIGLEC7, SLAMF6, SPN, TBX21, and TNFα), a tumor cell membrane protein (e.g., EGFR, PSMA, CAIX, CEA, CD33, CD47, US28, CXCR2, CXCR4, CXCR7, MMR), a tumor microenvironment target ( e.g., VEGF, EIIIB, CapG, TUFM), a ligand, cytokine or chemokine (e.g., EGF, HGF, TNFα, CXCL12, CXCL11), or immune cell marker (e.g., MHC-II, PD-L1, CTLA-4, CD11b, CD7, CD8, CD20, CD38, CD45, Ly-6C/G). In some instances, a VHH of this disclosure may be linked to a VHH that binds to albumin (e.g., human serum albumin). The linker used to join the VHHs preserves the functionality of the different VHH domains. Non-limiting examples of linkers suitable for linking the VHH domains in the polypeptides of this disclosure include, but are not limited to, (GS)n (SEQ ID NO: 808), (GGS)n (SEQ ID NO: 809), (GGGS)n (SEQ ID NO:645), (GGSG)n (SEQ ID NO:646), (GGSGG)n (SEQ ID NO:647), or (GGGGS)n (SEQ ID NO:648), wherein n is 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10. In some instances the length of the linker can be about 15 amino acids. In one instance the linker is (G4S)3 (SEQ ID NO:649). In some instances, the bispecific construct comprises a first VHH of this disclosure (that specifically binds c-MET) linked to a first human Ig hinge-Fc region (e.g., from IgG1, IgG2, IgG3, IgG4, or IgA) and a second VHH (that binds to a different epitope of c-MET than the first VHH, or that specifically binds to a different antigen)
linked to a second human Ig hinge-Fc region (e.g., from IgG1, IgG2, IgG3, IgG4, or IgA). In certain instances, the human Ig hinge-Fc region is at least 80%, at least 85%, at least 90%, at least 95%, or 100% identical to the amino acid sequence of IgG4PAA set forth in SEQ ID NO:643. In certain instances, the human Ig hinge-Fc region used in the bispecific constructs have 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 amino acid substitutions relative to SEQ ID NO:643. In certain instances, the Fc region is engineered to promote heterodimerization. This can be done by any method known in the art such as the use of knob-into-hole, electrostatic steering, or any other design strategies for heterodimerization of Fc. See, Ha et al., Front Immunol. 2016; 7: 394; Moore et al., Methods, 154:38-50 (2019); WO 2017/106462; and Yu et al., Nat Methods, 16, 1095- 1100 (2019) (all incorporated by reference herein). In certain instances, the Fc region is also engineered to improve its therapeutic activity. See e.g., Mimoto et al., Curr Pharm Biotechnol., 2016;17(15):1298-1314; Chen et al., Theranostics, 2021; 11(4): 1901–1917; Delidakis et al., Ann Rev. Biomed Engg., 24:249-274 (2022) (all incorporated by reference herein). Multispecific Constructs Another binding molecule encompassed by this disclosure is a multispecific construct. These multispecific constructs bind to three or more different epitopes. In one instance, three or more different VHHs are linked together. In some instances, one VHH is linked by its C-terminal to the N-terminal of the heavy chain of a whole antibody. In another instance, two VHHs are linked by their C-terminals to the N-terminals of the two heavy chains of a whole antibody. In some cases, the two VHHs are identical. In other cases, the two VHHs are different. In yet another instance, one VHH is linked by its C- terminal to the C-terminal of the heavy chain of a whole antibody. In another instance, two VHHs are linked by their C-terminals to the C-terminals of the two heavy chains of a whole antibody. In some cases, the two VHHs are identical. In other cases, the two VHHs are different. In some instances, a multispecific construct is based on a bispecific construct described above, wherein the bispecific construct is conjugated at the C- terminal of one or both of its heavy chains to one or two VHHs. The C-terminally
conjugated VHH or VHHs can bind to a different epitope than the VHHs of the bispecific construct. In all of the above, at least one antigen binding domain specifically binds to c- MET. Chimeric Antigen Receptors Also encompassed by this disclosure are chimeric antigen receptors that include one or more VHHs described herein. A typical chimeric antigen receptor (CAR) includes four domains, each of which serves a distinct purpose; the antigen recognition domain at the exterior of the cell is responsible for antigen recognition, the hinge region and transmembrane domain provides stability whereas, the intracellular domain in the endodomain of the receptor plays a crucial role in transmitting signals to activate the effector function of CAR T or NK cells. In one instance, the CAR comprises a VHH of this disclosure that specifically binds c-MET, a hinge region, a transmembrane domain, and an endodomain. Any hinge region, transmembrane domain, and endodomain that are employed in CARs can be used. See e.g., Ahmad et al., Advances in Cancer Biology – Metastasis, Vol.4, 100035 (2022); Feins et al., Am. J Hemaol., 94, S1:S3-S9, 2019; Hajari et al., IUBMB Life 71(2019) 1259-1267; De Munter et al., Int J Mol Sci 19(2018) 1-11; Xie et al., PNAS 116 (2019):7624-7631; Xie et al., Cancer Immunol. Res.8(2020) 518-530 (all incorporated by reference herein). In one instance, a VHH that specifically binds c-MET is linked to CD28 and CD137 signaling domains and CD3ζ (c-Met-28-137- 3ζ). See, Huang et al., J Biomed Res.2022 Jan; 36(1): 10–21 (incorporated by reference herein). CARs of the present disclosure comprise an antigen binding domain (e.g., anti-c- MET VHH), a hinge domain, a transmembrane domain, and an intracellular signaling domain. In some instances, the CARs comprise an antigen binding domain, a spacer domain, a transmembrane domain, a costimulatory domain, and a signaling domain. In some aspects, the antigen binding domain (e.g., anti-c-MET VHH) may be operably linked to another domain of the CAR, such as the hinge domain, the transmembrane domain and/or the intracellular domain, both described elsewhere herein,
for expression in an immune cells (e.g., a T cell). In some aspects, a first nucleic acid sequence encoding the antigen binding domain (e.g., anti-c-MET VHH) is operably linked to a second nucleic acid encoding a hinge and transmembrane domain, and further operably linked to a third a nucleic acid sequence encoding an intracellular domain. In some cases, the nucleic acid is mRNA. Conjugates The polypeptides, binding molecules, and VHHs of this disclosure can be fused, linked, or conjugated to other moieties. These moieties may be for detection or purification, or to add another therapeutic agent, radionuclide, or toxin to the polypeptides, binding molecules, and VHHs. In certain instances, the moiety is linked or conjugated to the C-terminal of the polypeptide, binding molecule, or VHH. In some instances, the linker can be any of the Gly Ser linkers discussed above (e.g., (G4S)3 (SEQ ID NO:649)) or the natural hinge of human IgA. In some instances a radionuclide may be directly conjugated to the polypeptide, binding molecule, or VHH, or conjugated via a small chelator. In some instances, a polypeptide, binding molecule, or VHH is linked to a detection or purification tag. In some instances, the tag moiety is His6 (SEQ ID NO:644), c-myc, hemagglutinin (HA), Flag, headlock tag, C-tag, ALFA tag, Avi tag, alkaline phosphatase or horseradish peroxidase. In some instances, a polypeptide, binding molecule, or VHH is linked to a fluorophore (GFP, RFP. In some instances, a polypeptide, binding molecule, or VHH is linked to a drug. In some instances a tumor targeted drug/therapy is conjugated to the polypeptide, binding molecule, or VHH of this disclosure to create an ADC. ADCs exploit the targeted efficiency of antibodies combined with the action of the cytotoxic payload conjugated to it. This results in specific targeting of cancer cells and thus lessens off-target side effects. The payload can be any drug that kills or inhibits the growth of the c-Met related/expressing cancer that is being targeted. In some instances the payload is multikinase inhibitor 178 or Mertansine (DM1). In some instances, a polypeptide, binding molecule, or VHH is linked to a
photosensitizer. In some instances, a polypeptide, binding molecule, or VHH is linked to a nanoparticle (e.g., liposomes, micelles, albumin-based NPs, polymer based polymerosomes). In some instances, a polypeptide, binding molecule, or VHH is linked to a toxic agent. In some instances, a polypeptide, binding molecule, or VHH is linked to a radionuclide (e.g., 89Zr, 68/67Ga, 18F, 99mTc). In some instances, a polypeptide, binding molecule, or VHH is linked to a VHH, a Fab, a scFv, or a whole antibody. In some instances, a polypeptide, binding molecule, or VHH is linked to a multimerization module (e.g. dimerization, trimerization, tetramerization, or pentamerization module). See e.g., Sadeghnezhad, G. et al., Int J Mol Sci., 20, 4818, 2019; Zhang et al., J. Mol Biol., 335, 49-56, 2004 (both incorporated by reference herein). In some cases, the dimerization module is a human Ig hinge-Fc domain (e.g., from IgG1, IgG4, IgA). In some cases, the dimerization module is IgG4PAA. In some cases, the trimerization module, is e.g. a collagen trimerization moiety, particularly a human collagen moiety, and a lung surfactant protein D moiety. In some instances, a polypeptide, binding molecule, or VHH is linked to a moiety that facilitates the polypeptide crossing the blood brain barrier (e.g., anti-transferrin antibody, FC5, FC44, a VHH that binds IGF-1R, VHHs E9, ni3a, and pa2H). In some instances, a polypeptide, binding molecule, or VHH is linked to a half-life extension moiety (e.g., PEG (linear and branched), human serum albumin, a VHH that binds human serum albumin, PASylation (PA600), human Ig Fc). Methods of Making and Pharmaceutical Compositions The disclosure provides a nucleic acid or nucleic acids that encode the polypeptides, binding molecules, and VHHs of this disclosure. The nucleic acid may be DNA or mRNA. The nucleic acid may be modified. The coding sequence may be modified to take into account the preferred codon usage of the expression host. Also provided are vectors that comprise the above nucleic acids. In some instances the vector is an expression vector. In other instances, the vector is a viral vector. Also provided are host cells that comprise the nucleic acid or vectors above. In some instances, the host cells are bacterial cells, yeast cells, insect cells, or mammalian
cells. In some instances, the host cell is E. coli. In some instances, the host cell is Pichia pastoris or Saccharomyces cerevisiae. In some instances, the host cell is a CHO cell, a COS cell, a HEK293T cell, or a NIH3T3 cell. The disclosure also features methods of making the polypeptides, binding molecules, or VHHs described herein. The method involves culturing the host cell under conditions that promote the expression of the polypeptide, binding molecule, or VHH. In some instances, the polypeptide, binding molecule, or VHH is purified. In certain instances, the polypeptide, binding molecule, or VHH is formulated as a sterile pharmaceutical composition. Also provided are pharmaceutical compositions comprising the polypeptides, binding molecules, or VHHs described herein, and a pharmaceutically acceptable carrier. In some instances, the pharmaceutically acceptable carrier is phosphate buffered saline. In some instances, the pharmaceutically acceptable carrier is sterile distilled water. mRNA Constructs In some aspects, the polynucleotides disclosed herein are or function as a messenger RNA (mRNA). As used herein, the term “messenger RNA” (mRNA) refers to any polynucleotide which encodes at least one peptide or polypeptide of interest and which is capable of being translated to produce the encoded peptide polypeptide of interest in vitro, in vivo, in situ or ex vivo. The disclosure provides a polynucleotide comprising an mRNA comprising: (i) a 5' UTR; (ii) an open reading frame (ORF) encoding a polypeptide, VHH, bispecific antibody, multispecific antibody, or CAR of claim described herein; (iii) a stop codon; and (iv) a 3' UTR. In certain instances, the mRNA can include a 5’-cap. In certain instances, the mRNA can also include a poly-A region. The disclosure features compositions including an mRNA encoding a polypeptide described herein (e.g., a polypeptide comprising an anti-c-MET VHH, a bivalent c-MET VHH-IgG4PAA, a bispecific antibody comprising an anti-C-MET VHH, a multispecific
antibody comprising an anti-C-MET VHH, a CAR comprising an anti-C-MET VHH). In one instance, the mRNAs include (i) a 5’-cap structure; (ii) a 5’-UTR; (iii) an open reading frame encoding the polypeptide; (iv) a 3’-untranslated region (3’-UTR); and (v) a poly-A region. mRNA: 5’-cap The 5'-cap structure of an mRNA is involved in nuclear export, increasing mRNA stability and binds the mRNA Cap Binding Protein (CBP), which is responsible for mRNA stability in the cell and translation competency through the association of CBP with poly(A) binding protein to form the mature cyclic mRNA species. The cap further assists the removal of 5' proximal introns removal during mRNA splicing. mRNA molecules may be 5'-end capped generating a 5'-ppp-5'-triphosphate linkage between a terminal guanosine cap residue and the 5'-terminal transcribed sense nucleotide of the mRNA. This 5'-guanylate cap may then be methylated to generate an N7-methyl-guanylate residue. The ribose sugars of the terminal and/or anteterminal transcribed nucleotides of the 5' end of the mRNA may optionally also be 2'-O- methylated.5'-decapping through hydrolysis and cleavage of the guanylate cap structure may target a nucleic acid molecule, such as an mRNA molecule, for degradation. Alterations to the nucleic acids of the present disclosure may generate a non- hydrolyzable cap structure preventing decapping and thus increasing mRNA half-life. Because cap structure hydrolysis requires cleavage of 5'-ppp-5' phosphorodiester linkages, alternative nucleotides may be used during the capping reaction. For example, a Vaccinia Capping Enzyme from New England Biolabs (Ipswich, MA) may be used with α-thio-guanosine nucleotides according to the manufacturer’s instructions to create a phosphorothioate linkage in the 5'-ppp-5' cap. Additional alternative guanosine nucleotides may be used such as α-methyl-phosphonate and seleno-phosphate nucleotides.
Additional alterations include, but are not limited to, 2'-O-methylation of the ribose sugars of 5'-terminal and/or 5'-anteterminal nucleotides of the mRNA (as mentioned above) on the 2'-hydroxyl group of the sugar ring. Multiple distinct 5'-cap structures can be used to generate the 5'-cap of a nucleic acid molecule, such as an mRNA molecule. 5’-cap structures include those described in International Patent Publication Nos. WO2008/127688, WO2008/016473, and WO2011/015347, each of which is incorporated herein by reference in its entirety. Cap analogs, which herein are also referred to as synthetic cap analogs, chemical caps, chemical cap analogs, or structural or functional cap analogs, differ from natural (i.e., endogenous, wild-type or physiological) 5'-caps in their chemical structure, while retaining cap function. Cap analogs may be chemically (i.e., non-enzymatically) or enzymatically synthesized and/linked to a nucleic acid molecule. For example, the Anti-Reverse Cap Analog (ARCA) cap contains two guanosines linked by a 5'-5'-triphosphate group, wherein one guanosine contains an N7 methyl group as well as a 3'-O-methyl group (i.e., N7,3'-O-dimethyl-guanosine-5'-triphosphate-5'- guanosine (m7G-3'mppp-G; which may equivalently be designated 3' O-Me- m7G(5')ppp(5')G)). The 3'-O atom of the other, unaltered, guanosine becomes linked to the 5'-terminal nucleotide of the capped nucleic acid molecule (e.g., an mRNA or mmRNA). The N7- and 3'-O-methlyated guanosine provides the terminal moiety of the capped nucleic acid molecule (e.g., mRNA or mmRNA). Another exemplary cap is mCAP, which is similar to ARCA but has a 2'-O- methyl group on guanosine (i.e., N7,2'-O-dimethyl-guanosine-5'-triphosphate-5'- guanosine, m7Gm-ppp-G). In some instances, the cap is a dinucleotide cap analog. As a non-limiting example, the dinucleotide cap analog may be modified at different phosphate positions with a boranophosphate group or a phophoroselenoate group such as the dinucleotide cap
analogs described in US Patent No. US 8,519,110, the contents of which are herein incorporated by reference in its entirety. In some instances, the cap analog is a N7-(4-chlorophenoxyethyl) substituted dicnucleotide form of a cap analog known in the art and/or described herein. Non- limiting examples of a N7-(4-chlorophenoxyethyl) substituted dinucleotide form of a cap analog include a N7-(4-chlorophenoxyethyl)-G(5’)ppp(5’)G and a N7-(4- chlorophenoxyethyl)-m3’-OG(5’)ppp(5’)G cap analog (see, e.g., the various cap analogs and the methods of synthesizing cap analogs described in Kore et al. Bioorganic & Medicinal Chemistry 21:4570-4574 (2013); the contents of which are herein incorporated by reference in its entirety). In some instances, a cap analog of the present disclosure is a 4-chloro/bromophenoxyethyl analog. While cap analogs allow for the concomitant capping of a nucleic acid molecule in an in vitro transcription reaction, up to 20% of transcripts remain uncapped. This, as well as the structural differences of a cap analog from endogenous 5'-cap structures of nucleic acids produced by the endogenous, cellular transcription machinery, may lead to reduced translational competency and reduced cellular stability. Nucleic acids of the disclosure (e.g., mRNAs of the disclosure) may also be capped post-transcriptionally, using enzymes.5’ cap structures produced by enzymatic capping may enhance binding of cap binding proteins, increase half-life, reduce susceptibility to 5' endonucleases and/or reduce 5' decapping, as compared to synthetic 5'-cap structures known in the art (or to a wild-type, natural or physiological 5'-cap structure). For example, recombinant Vaccinia Virus Capping Enzyme and recombinant 2'-O-methyltransferase enzyme can create a canonical 5'-5'-triphosphate linkage between the 5'-terminal nucleotide of an mRNA and a guanosine cap nucleotide wherein the cap guanosine contains an N7 methylation and the 5'-terminal nucleotide of the mRNA contains a 2'-O-methyl. Such a structure is termed the Cap1 structure. This cap results in a higher translational-competency and cellular stability and a reduced activation of cellular pro-inflammatory cytokines, as compared, e.g., to other 5'cap analog structures
known in the art. Cap structures include 7mG(5')ppp(5')N,pN2p (cap 0), 7mG(5')ppp(5')NlmpNp (cap 1), 7mG(5')-ppp(5')NlmpN2mp (cap 2), and m(7)Gpppm(3)(6,6,2')Apm(2')Apm(2')Cpm(2)(3,2')Up (cap 4). 5' terminal caps may include endogenous caps or cap analogs. A 5' terminal cap may include a guanosine analog. Useful guanosine analogs include inosine, N1-methyl- guanosine, 2'-fluoro-guanosine, 7-deaza-guanosine, 8-oxo-guanosine, 2-amino- guanosine, LNA-guanosine, and 2-azido-guanosine. In some cases, the 5’ terminal cap is or comprises m7GpppGm. In some instances, the nucleic acids described herein may contain a modified 5’- cap. A modification on the 5’-cap may increase the stability of mRNA, increase the half- life of the mRNA, and could increase the mRNA translational efficiency. The modified 5’-cap may include, but is not limited to, one or more of the following modifications: modification at the 2’ and/or 3’ position of a capped guanosine triphosphate (GTP), a replacement of the sugar ring oxygen (that produced the carbocyclic ring) with a methylene moiety (CH2), a modification at the triphosphate bridge moiety of the cap structure, or a modification at the nucleobase (G) moiety. Untranslated Regions (UTRs) Untranslated regions (UTRs) are nucleic acid sections of a polynucleotide before a start codon (5' UTR) and after a stop codon (3' UTR) that are not translated. In some instances, a polynucleotide (e.g., a ribonucleic acid (RNA), e.g., a messenger RNA (mRNA)) of the disclosure comprising an open reading frame (ORF) encoding an antibody further comprises UTR (e.g., a 5'UTR or functional fragment thereof, a 3'UTR or functional fragment thereof, or a combination thereof). Translation of a polynucleotide comprising an open reading frame encoding a polypeptide can be controlled and regulated by a variety of mechanisms that are provided by various cis-acting nucleic acid structures. For example, naturally-occurring, cis-acting RNA elements that form hairpins or other higher-order (e.g., pseudoknot) intramolecular
mRNA secondary structures can provide a translational regulatory activity to a polynucleotide, wherein the RNA element influences or modulates the initiation of polynucleotide translation, particularly when the RNA element is positioned in the 5' UTR close to the 5’-cap structure (Pelletier and Sonenberg (1985) Cell 40(3):515-526; Kozak (1986) Proc Natl Acad Sci 83:2850-2854). Cis-acting RNA elements can also affect translation elongation, being involved in numerous frameshifting events (Namy et al., (2004) Mol Cell 13(2):157-168). Internal ribosome entry sequences (IRES) represent another type of cis-acting RNA element that are typically located in 5' UTRs, but have also been reported to be found within the coding region of naturally-occurring mRNAs (Holcik et al. (2000) Trends Genet 16(10):469-473). In cellular mRNAs, IRES often coexist with the 5'-cap structure and provide mRNAs with the functional capacity to be translated under conditions in which cap-dependent translation is compromised (Gebauer et al., (2012) Cold Spring Harb Perspect Biol 4(7):a012245). Another type of naturally-occurring cis-acting RNA element comprises upstream open reading frames (uORFs). Naturally-occurring uORFs occur singularly or multiply within the 5' UTRs of numerous mRNAs and influence the translation of the downstream major ORF, usually negatively (with the notable exception of GCN4 mRNA in yeast and ATF4 mRNA in mammals, where uORFs serve to promote the translation of the downstream major ORF under conditions of increased eIF2 phosphorylation (Hinnebusch (2005) Annu Rev Microbiol 59:407-450)). Additional exemplary translational regulatory activities provided by components, structures, elements, motifs, and/or specific sequences comprising polynucleotides (e.g., mRNA) include, but are not limited to, mRNA stabilization or destabilization (Baker & Parker (2004) Curr Opin Cell Biol 16(3):293-299), translational activation (Villalba et al., (2011) Curr Opin Genet Dev 21(4):452-457), and translational repression (Blumer et al., (2002) Mech Dev 110(1-2):97-112). Studies have shown that naturally-occurring, cis- acting RNA elements can confer their respective functions when used to modify, by incorporation into, heterologous polynucleotides (Goldberg-Cohen et al., (2002) J Biol Chem 277(16):13635-13640).
The present disclosure provides synthetic polynucleotides comprising a modification (e.g., an RNA element), wherein the modification provides a desired translational regulatory activity. In some instances, the disclosure provides a polynucleotide comprising a 5’ untranslated region (UTR), an initiation codon, a full open reading frame encoding a polypeptide, a 3’ UTR, and at least one modification, wherein the at least one modification provides a desired translational regulatory activity, for example, a modification that promotes and/or enhances the translational fidelity of mRNA translation. In some instances, the desired translational regulatory activity is a cis-acting regulatory activity. In some instances, the desired translational regulatory activity is an increase in the residence time of the 43 S pre-initiation complex (PIC) or ribosome at, or proximal to, the initiation codon. In some instances, the desired translational regulatory activity is an increase in the initiation of polypeptide synthesis at or from the initiation codon. In some instances, the desired translational regulatory activity is an increase in the amount of polypeptide translated from the full open reading frame. In some instances, the desired translational regulatory activity is an increase in the fidelity of initiation codon decoding by the PIC or ribosome. In some instances, the desired translational regulatory activity is inhibition or reduction of leaky scanning by the PIC or ribosome. In some instances, the desired translational regulatory activity is a decrease in the rate of decoding the initiation codon by the PIC or ribosome. In some instances, the desired translational regulatory activity is inhibition or reduction in the initiation of polypeptide synthesis at any codon within the mRNA other than the initiation codon. In some instances, the desired translational regulatory activity is inhibition or reduction of the amount of polypeptide translated from any open reading frame within the mRNA other than the full open reading frame. In some instances, the desired translational regulatory activity is inhibition or reduction in the production of aberrant translation products. In some instances, the desired translational regulatory activity is a combination of one or more of the foregoing translational regulatory activities.
Accordingly, the present disclosure provides a polynucleotide, e.g., an mRNA, comprising an RNA element that comprises a sequence and/or an RNA secondary
structure(s) that provides a desired translational regulatory activity as described herein. In some aspects, the mRNA comprises an RNA element that comprises a sequence and/or an RNA secondary structure(s) that promotes and/or enhances the translational fidelity of mRNA translation. In some aspects, the mRNA comprises an RNA element that comprises a sequence and/or an RNA secondary structure(s) that provides a desired translational regulatory activity, such as inhibiting and/or reducing leaky scanning. In some aspects, the disclosure provides an mRNA that comprises an RNA element that comprises a sequence and/or an RNA secondary structure(s) that inhibits and/or reduces leaky scanning thereby promoting the translational fidelity of the mRNA.
In some instances, the RNA element comprises natural and/or modified nucleotides. In some instances, the RNA element comprises of a sequence of linked nucleotides, or derivatives or analogs thereof that provides a desired translational regulatory activity as described herein. In some instances, the RNA element comprises a sequence of linked nucleotides, or derivatives or analogs thereof that forms or folds into a stable RNA secondary structure, wherein the RNA secondary structure provides a desired translational regulatory activity as described herein. RNA elements can be identified and/or characterized based on the primary sequence of the element (e.g., GC-rich element), by RNA secondary structure formed by the element (e.g. stem-loop), by the location of the element within the RNA molecule (e.g., located within the 5’ UTR of an mRNA), by the biological function and/or activity of the element (e.g., “translational enhancer element”), and any combination thereof.
In some aspects, the disclosure provides an mRNA having one or more structural modifications that inhibits leaky scanning and/or promotes the translational fidelity of mRNA translation, wherein at least one of the structural modifications is a GC-rich RNA element. In some aspects, the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising a sequence of linked nucleotides, or derivatives or analogs thereof, preceding a Kozak consensus sequence in a 5’ UTR of the mRNA. In one embodiment, the GC- rich RNA element is located about 30, about 25, about 20, about 15, about 10, about 5,
about 4, about 3, about 2, or about 1 nucleotide(s) upstream of a Kozak consensus sequence in the 5’ UTR of the mRNA. In another embodiment, the GC-rich RNA element is located 15-30, 15-20, 15-25, 10-15, or 5-10 nucleotides upstream of a Kozak consensus sequence. In another embodiment, the GC-rich RNA element is located immediately adjacent to a Kozak consensus sequence in the 5’ UTR of the mRNA. In any of the foregoing or related aspects, the disclosure provides a GC-rich RNA element which comprises a sequence of 3-30, 5-25, 10-20, 15-20, about 20, about 15, about 12, about 10, about 7, about 6 or about 3 nucleotides, derivatives or analogs thereof, linked in any order, wherein the sequence composition is 70-80% cytosine, 60- 70% cytosine, 50%-60% cytosine, 40-50% cytosine, 30-40% cytosine bases. In any of the foregoing or related aspects, the disclosure provides a GC-rich RNA element which comprises a sequence of 3-30, 5-25, 10-20, 15-20, about 20, about 15, about 12, about 10, about 7, about 6 or about 3 nucleotides, derivatives or analogs thereof, linked in any order, wherein the sequence composition is about 80% cytosine, about 70% cytosine, about 60% cytosine, about 50% cytosine, about 40% cytosine, or about 30% cytosine. In any of the foregoing or related aspects, the disclosure provides a GC-rich RNA element which comprises a sequence of 20, 19, 18, 17, 16, 15, 14, 13, 12, 11, 10, 9, 8, 7, 6, 5, 4, or 3 nucleotides, or derivatives or analogs thereof, linked in any order, wherein the sequence composition is 70-80% cytosine, 60-70% cytosine, 50%-60% cytosine, 40- 50% cytosine, or 30-40% cytosine. In any of the foregoing or related aspects, the disclosure provides a GC-rich RNA element which comprises a sequence of 20, 19, 18, 17, 16, 15, 14, 13, 12, 11, 10, 9, 8, 7, 6, 5, 4, or 3 nucleotides, or derivatives or analogs thereof, linked in any order, wherein the sequence composition is about 80% cytosine, about 70% cytosine, about 60% cytosine, about 50% cytosine, about 40% cytosine, or about 30% cytosine. In some instances, the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising a sequence of linked nucleotides, or derivatives or analogs thereof, preceding
a Kozak consensus sequence in a 5’ UTR of the mRNA, wherein the GC-rich RNA element is located about 30, about 25, about 20, about 15, about 10, about 5, about 4, about 3, about 2, or about 1 nucleotide(s) upstream of a Kozak consensus sequence in the 5’ UTR of the mRNA, and wherein the GC-rich RNA element comprises a sequence of 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 nucleotides, or derivatives or analogs thereof, linked in any order, wherein the sequence composition is >50% cytosine. In some instances, the sequence composition is >55% cytosine, >60% cytosine, >65% cytosine, >70% cytosine, >75% cytosine, >80% cytosine, >85% cytosine, or >90% cytosine. In other aspects, the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising a sequence of linked nucleotides, or derivatives or analogs thereof, preceding a Kozak consensus sequence in a 5’ UTR of the mRNA, wherein the GC-rich RNA element is located about 30, about 25, about 20, about 15, about 10, about 5, about 4, about 3, about 2, or about 1 nucleotide(s) upstream of a Kozak consensus sequence in the 5’ UTR of the mRNA, and wherein the GC-rich RNA element comprises a sequence of about 3-30, 5-25, 10-20, 15-20 or about 20, about 15, about 12, about 10, about 6 or about 3 nucleotides, or derivatives or analogues thereof, wherein the sequence comprises a repeating GC-motif, wherein the repeating GC-motif is [CCG]n, wherein n = 1 to 10 (SEQ ID NO: 810), n= 2 to 8, n= 3 to 6, or n= 4 to 5. In some instances, the sequence comprises a repeating GC-motif [CCG]n, wherein n = 1, 2, 3, 4 or 5. In some instances, the sequence comprises a repeating GC-motif [CCG]n, wherein n = 1, 2, or 3. In some instances, the sequence comprises a repeating GC-motif [CCG]n, wherein n = 1. In some instances, the sequence comprises a repeating GC-motif [CCG]n, wherein n = 2. In some instances, the sequence comprises a repeating GC-motif [CCG]n, wherein n = 3. In some instances, the sequence comprises a repeating GC-motif [CCG]n, wherein n = 4. In some instances, the sequence comprises a repeating GC-motif [CCG]n, wherein n = 5. In another aspect, the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element
comprising a sequence of linked nucleotides, or derivatives or analogs thereof, preceding a Kozak consensus sequence in a 5’ UTR of the mRNA, wherein the GC-rich RNA element comprises any one of the sequences set forth in Table 8. In one embodiment, the GC-rich RNA element is located about 30, about 25, about 20, about 15, about 10, about 5, about 4, about 3, about 2, or about 1 nucleotide(s) upstream of a Kozak consensus sequence in the 5’ UTR of the mRNA. In another embodiment, the GC-rich RNA element is located about 15-30, 15-20, 15-25, 10-15, or 5-10 nucleotides upstream of a Kozak consensus sequence. In another embodiment, the GC-rich RNA element is located immediately adjacent to a Kozak consensus sequence in the 5’ UTR of the mRNA. In other aspects, the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising the sequence V1 [CCCCGGCGCC (SEQ ID NO:650)] as set forth in Table 8, or derivatives or analogs thereof, preceding a Kozak consensus sequence in the 5’ UTR of the mRNA. In some instances, the GC-rich element comprises the sequence V1 as set forth in Table 8 located immediately adjacent to and upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA. In some instances, the GC-rich element comprises the sequence V1 as set forth in Table 8 located 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10 bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA. In other instances, the GC-rich element comprises the sequence V1 as set forth in Table 8 located 1-3, 3-5, 5-7, 7-9, 9-12, or 12-15 bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA. In other aspects, the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising the sequence V2 [CCCCGGC] as set forth in Table 8, or derivatives or analogs thereof, preceding a Kozak consensus sequence in the 5’ UTR of the mRNA. In some instances, the GC-rich element comprises the sequence V2 as set forth in Table 8 located immediately adjacent to and upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA. In some instances, the GC-rich element comprises the sequence V2 as set forth in Table 8 located 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10 bases upstream of the Kozak
consensus sequence in the 5’ UTR of the mRNA. In other instances, the GC-rich element comprises the sequence V2 as set forth in Table 8 located 1-3, 3-5, 5-7, 7-9, 9-12, or 12- 15 bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA. In other aspects, the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising the sequence EK [GCCGCC] as set forth in Table 8, or derivatives or analogs thereof, preceding a Kozak consensus sequence in the 5’ UTR of the mRNA. In some instances, the GC-rich element comprises the sequence EK as set forth in Table 8 located immediately adjacent to and upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA. In some instances, the GC-rich element comprises the sequence EK as set forth in Table 8 located 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10 bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA. In other instances, the GC-rich element comprises the sequence EK as set forth in Table 8 located 1-3, 3-5, 5-7, 7-9, 9-12, or 12- 15 bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA. In yet other aspects, the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising the sequence V1 [CCCCGGCGCC (SEQ ID NO: 650)] as set forth in Table 8, or derivatives or analogs thereof, preceding a Kozak consensus sequence in the 5’ UTR of the mRNA, wherein the 5’ UTR comprises the following sequence shown in Table 8: GGGAAAUAAGAGAGAAAAGAAGAGUAAGAAGAAAUAUAAGA (SEQ ID NO: 653). The skilled artisan will of course recognize that all Us in the RNA sequences described herein will be Ts in a corresponding template DNA sequence, for example, in DNA templates or constructs from which mRNAs of the disclosure are transcribed, e.g., via IVT. In some instances, the GC-rich element comprises the sequence V1 as set forth in Table 8 located immediately adjacent to and upstream of the Kozak consensus sequence in the 5’ UTR sequence shown in Table 8. In some instances, the GC-rich element comprises the sequence V1 as set forth in Table 8 located 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10
bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA, wherein the 5’ UTR comprises the following sequence shown in Table 8: GGGAAAUAAGAGAGAAAAGAAGAGUAAGAAGAAAUAUAAGA (SEQ ID NO: 653). In other instances, the GC-rich element comprises the sequence V1 as set forth in Table 8 located 1-3, 3-5, 5-7, 7-9, 9-12, or 12-15 bases upstream of the Kozak consensus sequence in the 5’ UTR of the mRNA, wherein the 5’ UTR comprises the following sequence shown in Table 8: GGGAAAUAAGAGAGAAAAGAAGAGUAAGAAGAAAUAUAAGA (SEQ ID NO: 653). In some instances, the 5’ UTR comprises the following sequence set forth in Table 8: GGGAAAUAAGAGAGAAAAGAAGAGUAAGAAGAAAUAUAAGACCCCGGCGC CGCCACC (SEQ ID NO: 654) Table 8
EK [GCCGCC] V1 [CCCCGGCGCC] (SEQ ID NO:650) V2 [CCCCGGC] (CCG)n, where n=1-10 (SEQ ID NO: 810) [CCG]n (SEQ ID NO: 810) (GCC)n, where n=1-10 (SEQ ID NO: 812) [GCC]n (SEQ ID NO: 812) In another aspect, the disclosure provides a modified mRNA comprising at least one modification, wherein at least one modification is a GC-rich RNA element comprising a stable RNA secondary structure comprising a sequence of nucleotides, or derivatives or analogs thereof, linked in an order which forms a hairpin or a stem-loop. In one embodiment, the stable RNA secondary structure is upstream of the Kozak consensus sequence. In another embodiment, the stable RNA secondary structure is located about 30, about 25, about 20, about 15, about 10, or about 5 nucleotides upstream of the Kozak consensus sequence. In another embodiment, the stable RNA secondary structure is located about 20, about 15, about 10 or about 5 nucleotides upstream of the Kozak consensus sequence. In another embodiment, the stable RNA secondary structure is located about 5, about 4, about 3, about 2, about 1 nucleotides upstream of the Kozak consensus sequence. In another embodiment, the stable RNA secondary structure is located about 15-30, about 15-20, about 15-25, about 10-15, or about 5-10 nucleotides upstream of the Kozak consensus sequence. In another embodiment, the stable RNA secondary structure is located 12-15 nucleotides upstream of the Kozak consensus sequence. In another embodiment, the stable RNA secondary structure has a deltaG of about -30 kcal/mol, about -20 to -30 kcal/mol, about -20 kcal/mol, about -10 to -20 kcal/mol, about -10 kcal/mol, about -5 to -10 kcal/mol.
In another embodiment, the modification is operably linked to an open reading frame encoding a polypeptide and wherein the modification and the open reading frame are heterologous.
In another embodiment, the sequence of the GC-rich RNA element is comprised exclusively of guanine (G) and cytosine (C) nucleobases.
RNA elements that provide a desired translational regulatory activity as described herein can be identified and characterized using known techniques, such as ribosome profiling. Ribosome profiling is a technique that allows the determination of the positions of PICs and/or ribosomes bound to mRNAs (see e.g., Ingolia et al., (2009) Science 324(5924):218-23, incorporated herein by reference). The technique is based on protecting a region or segment of mRNA, by the PIC and/or ribosome, from nuclease digestion. Protection results in the generation of a 30-bp fragment of RNA termed a ‘footprint’. The sequence and frequency of RNA footprints can be analyzed by methods known in the art (e.g., RNA-seq). The footprint is roughly centered on the A-site of the ribosome. If the PIC or ribosome dwells at a particular position or location along an mRNA, footprints generated at these position would be relatively common. Studies have shown that more footprints are generated at positions where the PIC and/or ribosome exhibits decreased processivity and fewer footprints where the PIC and/or ribosome exhibits increased processivity (Gardin et al., (2014) eLife 3:e03735). In some instances, residence time or the time of occupancy of the PIC or ribosome at a discrete position or location along an polynucleotide comprising any one or more of the RNA elements described herein is determined by ribosome profiling.
A UTR can be homologous or heterologous to the coding region in a polynucleotide. In some instances, the UTR is homologous to the ORF encoding the antibody. In some instances, the UTR is heterologous to the ORF encoding the antibody. In some instances, the polynucleotide comprises two or more 5'UTRs or functional fragments thereof, each of which have the same or different nucleotide sequences. In
some instances, the polynucleotide comprises two or more 3'UTRs or functional fragments thereof, each of which have the same or different nucleotide sequences. In some instances, the 5'UTR or functional fragment thereof, 3' UTR or functional fragment thereof, or any combination thereof is sequence optimized. In some instances, the 5'UTR or functional fragment thereof, 3' UTR or functional fragment thereof, or any combination thereof comprises at least one chemically modified nucleobase, e.g., N1 methylpseudouracil or 5-methoxyuracil. UTRs can have features that provide a regulatory role, e.g., increased or decreased stability, localization and/or translation efficiency. A polynucleotide comprising a UTR can be administered to a cell, tissue, or organism, and one or more regulatory features can be measured using routine methods. In some instances, a functional fragment of a 5'UTR or 3'UTR comprises one or more regulatory features of a full length 5' or 3' UTR, respectively. Natural 5'UTRs bear features that play roles in translation initiation. They harbor signatures like Kozak sequences that are commonly known to be involved in the process by which the ribosome initiates translation of many genes. Kozak sequences have the consensus CCR(A/G)CCAUGG, where R is a purine (adenine or guanine) three bases upstream of the start codon (AUG), which is followed by another 'G'.5'UTRs also have been known to form secondary structures that are involved in elongation factor binding. By engineering the features typically found in abundantly expressed genes of specific target organs, one can enhance the stability and protein production of a polynucleotide. For example, introduction of 5'UTR of liver-expressed mRNA, such as albumin, serum amyloid A, Apolipoprotein A/B/E, transferrin, alpha fetoprotein, erythropoietin, or antibody, can enhance expression of polynucleotides in hepatic cell lines or liver. Likewise, use of 5'UTR from other tissue-specific mRNA to improve expression in that tissue is possible for muscle (e.g., MyoD, Myosin, Myoglobin, Myogenin, Herculin), for endothelial cells (e.g., Tie-1, CD36), for myeloid cells (e.g., C/EBP, AML1, G-CSF, GM-CSF, CD11b, MSR, Fr-1, i-NOS), for leukocytes (e.g.,
CD45, CD18), for adipose tissue (e.g., CD36, GLUT4, ACRP30, adiponectin) and for lung epithelial cells (e.g., SP-A/B/C/D). In some instances, UTRs are selected from a family of transcripts whose proteins share a common function, structure, feature or property. For example, an encoded polypeptide can belong to a family of proteins (i.e., that share at least one function, structure, feature, localization, origin, or expression pattern), which are expressed in a particular cell, tissue or at some time during development. The UTRs from any of the genes or mRNA can be swapped for any other UTR of the same or different family of proteins to create a new polynucleotide. In some instances, the 5’UTR and the 3’UTR can be heterologous. In some instances, the 5'UTR can be derived from a different species than the 3'UTR. In some instances, the 3'UTR can be derived from a different species than the 5'UTR. Co-owned International Patent Application No. PCT/US2014/021522 (Publ. No. WO/2014/164253, incorporated herein by reference in its entirety) provides a listing of exemplary UTRs that can be utilized in the polynucleotide of the present disclosure as flanking regions to an ORF. Exemplary UTRs of the application include, but are not limited to, one or more 5'UTR and/or 3'UTR derived from the nucleic acid sequence of: a globin, such as an α- or β-globin (e.g., a Xenopus, mouse, rabbit, or human globin); a strong Kozak translational initiation signal; a CYBA (e.g., human cytochrome b-245 α polypeptide); an albumin (e.g., human albumin7); a HSD17B4 (hydroxysteroid (17-β) dehydrogenase); a virus (e.g., a tobacco etch virus (TEV), a Venezuelan equine encephalitis virus (VEEV), a Dengue virus, a cytomegalovirus (CMV) (e.g., CMV immediate early 1 (IE1)), a hepatitis virus (e.g., hepatitis B virus), a sindbis virus, or a PAV barley yellow dwarf virus); a heat shock protein (e.g., hsp70); a translation initiation factor (e.g., elF4G); a glucose transporter (e.g., hGLUT1 (human glucose transporter 1)); an actin (e.g., human α or β actin); a GAPDH; a tubulin; a histone; a citric acid cycle enzyme; a topoisomerase (e.g., a 5'UTR of a TOP gene lacking the 5' TOP motif (the oligopyrimidine tract)); a ribosomal
protein Large 32 (L32); a ribosomal protein (e.g., human or mouse ribosomal protein, such as, for example, rps9); an ATP synthase (e.g., ATP5A1 or the β subunit of mitochondrial H+-ATP synthase); a growth hormone e (e.g., bovine (bGH) or human (hGH)); an elongation factor (e.g., elongation factor 1 α1 (EEF1A1)); a manganese superoxide dismutase (MnSOD); a myocyte enhancer factor 2A (MEF2A); a β-F1- ATPase, a creatine kinase, a myoglobin, a granulocyte-colony stimulating factor (G- CSF); a collagen (e.g., collagen type I, alpha 2 (Col1A2), collagen type I, alpha 1 (Col1A1), collagen type VI, alpha 2 (Col6A2), collagen type VI, alpha 1 (Col6A1)); a ribophorin (e.g., ribophorin I (RPNI)); a low density lipoprotein receptor-related protein (e.g., LRP1); a cardiotrophin-like cytokine factor (e.g., Nnt1); calreticulin (Calr); a procollagen-lysine, 2-oxoglutarate 5-dioxygenase 1 (Plod1); and a nucleobindin (e.g., Nucb1). In some instances, the 5'UTR is selected from the group consisting of a β-globin 5’UTR; a 5'UTR containing a strong Kozak translational initiation signal; a cytochrome b-245 α polypeptide (CYBA) 5'UTR; a hydroxysteroid (17-β) dehydrogenase (HSD17B4) 5'UTR; a Tobacco etch virus (TEV) 5'UTR; a Venezuelen equine encephalitis virus (TEEV) 5'UTR; a 5' proximal open reading frame of rubella virus (RV) RNA encoding nonstructural proteins; a Dengue virus (DEN) 5'UTR; a heat shock protein 70 (Hsp70) 5'UTR; a eIF4G 5'UTR; a GLUT15'UTR; functional fragments thereof and any combination thereof. In some instances, the 3'UTR is selected from the group consisting of a β-globin 3’UTR; a CYBA 3'UTR; an albumin 3'UTR; a growth hormone (GH) 3'UTR; a VEEV 3'UTR; a hepatitis B virus (HBV) 3'UTR; α-globin 3'UTR; a DEN 3'UTR; a PAV barley yellow dwarf virus (BYDV-PAV) 3'UTR; an elongation factor 1 α1 (EEF1A1) 3'UTR; a manganese superoxide dismutase (MnSOD) 3'UTR; a β subunit of mitochondrial H(+)- ATP synthase (β-mRNA) 3'UTR; a GLUT13'UTR; a MEF2A 3'UTR; a β-F1-ATPase 3'UTR; functional fragments thereof and combinations thereof.
Wild-type UTRs derived from any gene or mRNA can be incorporated into the polynucleotides of the present disclosure. In some instances, a UTR can be altered relative to a wild type or native UTR to produce a variant UTR, e.g., by changing the orientation or location of the UTR relative to the ORF; or by inclusion of additional nucleotides, deletion of nucleotides, swapping or transposition of nucleotides. In some instances, variants of 5' or 3' UTRs can be utilized, for example, mutants of wild type UTRs, or variants wherein one or more nucleotides are added to or removed from a terminus of the UTR. Additionally, one or more synthetic UTRs can be used in combination with one or more non-synthetic UTRs. See, e.g., Mandal and Rossi, Nat. Protoc. 20138(3):568-82, the contents of which are incorporated herein by reference in their entirety. UTRs or portions thereof can be placed in the same orientation as in the transcript from which they were selected or can be altered in orientation or location. Hence, a 5' and/or 3' UTR can be inverted, shortened, lengthened, or combined with one or more other 5' UTRs or 3' UTRs. In some instances, the polynucleotide comprises multiple UTRs, e.g., a double, a triple or a quadruple 5’UTR or 3’UTR. For example, a double UTR comprises two copies of the same UTR either in series or substantially in series. For example, a double beta- globin 3'UTR can be used (see US2010/0129877, the contents of which are incorporated herein by reference in its entirety). In certain instances, the polynucleotides of the disclosure comprise a 5' UTR and/or a 3' UTR selected from any of the UTRs disclosed herein. In some instances, the 5' UTR comprises: 5' UTR-001 (Upstream UTR) (GGGAAAUAAGAGAGAAAAGAAGAGUAAGAAGAAAUAUAAGAGCCACC) (SEQ ID NO:655);
5' UTR-002 (Upstream UTR) (GGGAGAUCAGAGAGAAAAGAAGAGUAAGAAGAAAUAUAAGAGCCACC) (SEQ ID NO:659); 5' UTR-004 (Upstream UTR) (GGGAGACAAGCUUGGCAUUCCGGUACUGUUGGUAAAGCCACC) (SEQ ID NO:660); 5' UTR-005 (Upstream UTR) (GGGAGAUCAGAGAGAAAAGAAGAGUAAGAAGAAAUAUAAGAGCCACC) (SEQ ID NO:661); 5' UTR-007 (Upstream UTR) (GGGAGACAAGCUUGGCAUUCCGGUACUGUUGGUAAAGCCACC) (SEQ ID NO:662); 5' UTR-008 (Upstream UTR) (GGGAAUUAACAGAGAAAAGAAGAGUAAGAAGAAAUAUAAGAGCCACC) (SEQ ID NO:663); 5' UTR-009 (Upstream UTR) (GGGAAAUUAGACAGAAAAGAAGAGUAAGAAGAAAUAUAAGAGCCACC) (SEQ ID NO:664); 5' UTR-010, Upstream (GGGAAAUAAGAGAGUAAAGAACAGUAAGAAGAAAUAUAAGAGCCACC) (SEQ ID NO:665); 5' UTR-011 (Upstream UTR) (GGGAAAAAAGAGAGAAAAGAAGACUAAGAAGAAAUAUAAGAGCCACC) (SEQ ID NO:666); 5' UTR-012 (Upstream UTR) (GGGAAAUAAGAGAGAAAAGAAGAGUAAGAAGAUAUAUAAGAGCCACC) (SEQ ID NO:667);
5' UTR-013 (Upstream UTR) (GGGAAAUAAGAGACAAAACAAGAGUAAGAAGAAAUAUAAGAGCCACC) (SEQ ID NO:668); 5' UTR-014 (Upstream UTR) (GGGAAAUUAGAGAGUAAAGAACAGUAAGUAGAAUUAAAAGAGCCACC) (SEQ ID NO:669); 5' UTR-015 (Upstream UTR) (GGGAAAUAAGAGAGAAUAGAAGAGUAAGAAGAAAUAUAAGAGCCACC) (SEQ ID NO:670); 5' UTR-016 (Upstream UTR) (GGGAAAUAAGAGAGAAAAGAAGAGUAAGAAGAAAAUUAAGAGCCACC) (SEQ ID NO:671); 5' UTR-017 (Upstream UTR); or (GGGAAAUAAGAGAGAAAAGAAGAGUAAGAAGAAAUUUAAGAGCCACC) (SEQ ID NO:672); 5' UTR-018 (Upstream UTR) 5' UTR (UCAAGCUUUUGGACCCUCGUACAGAAGCUAAUACGACUCACUAUAGGG AAAUAAGAGAGAAAAGAAGAGUAAGAAGAAAUAUAAGAGCCACC) (SEQ ID NO:673). In some instances, the 3' UTR comprises: 3'UTR: UGAUAAUAGGCUGGAGCCUCGGUGGCCUAGCUUCUUGCCCCU UGGGCCUCCCCCCAGCCCCUCCUCCCCUUCCUGCACCCGUACC CCCGUGGUCUUUGAAUAAAGUCUGAGUGGGCGGC (SEQ ID NO:674)
142-3p 3' UTR (UTR including miR142-3p binding site) (UGAUAAUAGUCCAUAAAGUAGGAAACACUACAGCUGGAGCCUCGGUGG CCAUGCUUCUUGCCCCUUGGGCCUCCCCCCAGCCCCUCCUCCCCUUCCUG CACCCGUACCCCCGUGGUCUUUGAAUAAAGUCUGAGUGGGCGGC) (SEQ ID NO:675); 142-3p 3' UTR (UTR including miR142-3p binding site) (UGAUAAUAGGCUGGAGCCUCGGUGGCUCCAUAAAGUAGGAAACACUAC ACAUGCUUCUUGCCCCUUGGGCCUCCCCCCAGCCCCUCCUCCCCUUCCUG CACCCGUACCCCCGUGGUCUUUGAAUAAAGUCUGAGUGGGCGGC) (SEQ ID NO:676); or 142-3p 3' UTR (UTR including miR142-3p binding site) (UGAUAAUAGGCUGGAGCCUCGGUGGCCAUGCUUCUUGCCCCUUCCAUA AAGUAGGAAACACUACAUGGGCCUCCCCCCAGCCCCUCCUCCCCUUCCUG CACCCGUACCCCCGUGGUCUUUGAAUAAAGUCUGAGUGGGCGGC) (SEQ ID NO:677); 142-3p 3' UTR (UTR including miR142-3p binding site) (UGAUAAUAGGCUGGAGCCUCGGUGGCCAUGCUUCUUGCCCCUUGGGCC UCCCCCCAGUCCAUAAAGUAGGAAACACUACACCCCUCCUCCCCUUCCUG CACCCGUACCCCCGUGGUCUUUGAAUAAAGUCUGAGUGGGCGGC) (SEQ ID NO:678); 142-3p 3' UTR (UTR including miR142-3p binding site) (UGAUAAUAGGCUGGAGCCUCGGUGGCCAUGCUUCUUGCCCCUUGGGCC UCCCCCCAGCCCCUCCUCCCCUUCUCCAUAAAGUAGGAAACACUACACUG CACCCGUACCCCCGUGGUCUUUGAAUAAAGUCUGAGUGGGCGGC) (SEQ ID NO:679); 142-3p 3' UTR (UTR including miR142-3p binding site) (UGAUAAUAGGCUGGAGCCUCGGUGGCCAUGCUUCUUGCCCCUUGGGCC
UCCCCCCAGCCCCUCCUCCCCUUCCUGCACCCGUACCCCCUCCAUAAAGU AGGAAACACUACAGUGGUCUUUGAAUAAAGUCUGAGUGGGCGGC) (SEQ ID NO:680). 142-3p 3' UTR (UTR including miR142-3p binding site) (UGAUAAUAGGCUGGAGCCUCGGUGGCCAUGCUUCUUGCCCCUUGGGCC UCCCCCCAGCCCCUCCUCCCCUUCCUGCACCCGUACCCCCGUGGUCUUUG AAUAAAGUUCCAUAAAGUAGGAAACACUACACUGAGUGGGCGGC) (SEQ ID NO:681); 3’UTR-018 GACAGUGCAGUCACCCAUAAAGUAGAAAGCACUACUAACAGCACUGGAG GGUGUAGUGUUUCCUACUUUAUGGAUGAGUGUACUGUG (SEQ ID NO:682) 3' UTR (miR142 and miR126 binding sites variant 1) (UGAUAAUAGUCCAUAAAGUAGGAAACACUACAGCUGGAGCCUCGGUGG CCAUGCUUCUUGCCCCUUGGGCCUCCCCCCAGCCCCUCCUCCCCUUCCUG CACCCGUACCCCCCGCAUUAUUACUCACGGUACGAGUGGUCUUUGAAUA AAGUCUGAGUGGGCGGC) (SEQ ID NO:683) 3' UTR (miR142 and miR126 binding sites variant 2) (UGAUAAUAGUCCAUAAAGUAGGAAACACUACAGCUGGAGCCUCGGUGG CCUAGCUUCUUGCCCCUUGGGCCUCCCCCCAGCCCCUCCUCCCCUUCCUG CACCCGUACCCCCCGCAUUAUUACUCACGGUACGAGUGGUCUUUGAAUA AAGUCUGAGUGGGCGGC) (SEQ ID NO:684); or 3’UTR (miR142-3p binding site variant 3) UGAUAAUAGGCUGGAGCCUCGGUGGCCUAGCUUCUUGCCCCUUGGGCCU CCCCCCAGCCCCUCCUCCCCUUCCUGCACCCGUACCCCCUCCAUAAAGUA
GGAAACACUACAGUGGUCUUUGAAUAAAGUCUGAGUGGGCGGC (SEQ ID NO:685). 3’UTR (miR142-3p binding site variant 3, DNA sequence) TGATAATAGGCTGGAGCCTCGGTGGCCTAGCTTCTTGCCCCTTGGGCCTCC CCCCAGCCCCTCCTCCCCTTCCTGCACCCGTACCCCCTCCATAAAGTAGGA AACACTACAGTGGTCTTTGAATAAAGTCTGAGTGGGCGGC (SEQ ID NO:686). In certain instances, the 5'UTR and/or 3'UTR sequence of the present disclosure comprises a nucleotide sequence at least about 60%, at least about 70%, at least about 80%, at least about 90%, at least about 95%, at least about 96%, at least about 97%, at least about 98%, at least about 99%, or about 100% identical to a sequence selected from the group consisting of 5'UTR sequences comprising any of SEQ ID NOs: 654, 655 and 659-673 and/or 3'UTR sequences comprises any of SEQ ID NOs: 674-686, and any combination thereof. The polynucleotides of the present disclosure can comprise combinations of features. For example, the ORF can be flanked by a 5'UTR that comprises a strong Kozak translational initiation signal and/or a 3'UTR comprising an oligo(dT) sequence for templated addition of a poly-A tail. A 5'UTR can comprise a first polynucleotide fragment and a second polynucleotide fragment from the same and/or different UTRs (see, e.g., US2010/0293625, herein incorporated by reference in its entirety). Other non-UTR sequences can be used as regions or subregions within the polynucleotides of the present disclosure. For example, introns or portions of intron sequences can be incorporated into the polynucleotides of the present disclosure. Incorporation of intronic sequences can increase protein production as well as polynucleotide expression levels. In some instances, the polynucleotide of the present disclosure comprises an internal ribosome entry site (IRES) instead of or in addition to a UTR (see, e.g., Yakubov et al., Biochem. Biophys. Res. Commun.2010394(1):189-193,
the contents of which are incorporated herein by reference in their entirety). In some instances, the polynucleotide comprises an IRES instead of a 5’UTR sequence. In some instances, the polynucleotide comprises an ORF and a viral capsid sequence. In some instances, the polynucleotide comprises a synthetic 5'UTR in combination with a non- synthetic 3'UTR. In some instances, the UTR can also include at least one translation enhancer polynucleotide, translation enhancer element, or translational enhancer elements (collectively, "TEE," which refers to nucleic acid sequences that increase the amount of polypeptide or protein produced from a polynucleotide. As a non-limiting example, the TEE can be located between the transcription promoter and the start codon. In some instances, the 5'UTR comprises a TEE. In one aspect, a TEE is a conserved element in a UTR that can promote translational activity of a nucleic acid such as, but not limited to, cap-dependent or cap- independent translation. In some instances, a 5'UTR and/or 3'UTR comprising at least one TEE described herein can be incorporated in a monocistronic sequence such as, but not limited to, a vector system or a nucleic acid vector. In some instances, a 5'UTR and/or 3'UTR of a polynucleotide of the present disclosure comprises a TEE or portion thereof described herein. In some instances, the TEEs in the 3'UTR can be the same and/or different from the TEE located in the 5'UTR. In some instances, a 5'UTR and/or 3'UTR of a polynucleotide of the present disclosure can include at least 1, at least 2, at least 3, at least 4, at least 5, at least 6, at least 7, at least 8, at least 9, at least 10, at least 11, at least 12, at least 13, at least 14, at least 15, at least 16, at least 17, at least 18 at least 19, at least 20, at least 21, at least 22, at least 23, at least 24, at least 25, at least 30, at least 35, at least 40, at least 45, at least 50, at least 55 or more than 60 TEE sequences. In one embodiment, the 5'UTR of a polynucleotide of the present disclosure can include 1-60, 1-55, 1-50, 1-45, 1-40, 1-35, 1- 30, 1-25, 1-20, 1-15, 1-10, 9, 8, 7, 6, 5, 4, 3, 2, or 1 TEE sequences. The TEE sequences
in the 5'UTR of the polynucleotide of the present disclosure can be the same or different TEE sequences. A combination of different TEE sequences in the 5'UTR of the polynucleotide of the present disclosure can include combinations in which more than one copy of any of the different TEE sequences are incorporated. In some instances, the 5'UTR and/or 3'UTR comprises a spacer to separate two TEE sequences. As a non-limiting example, the spacer can be a 15 nucleotide spacer and/or other spacers known in the art. As another non-limiting example, the 5'UTR and/or 3'UTR comprises a TEE sequence-spacer module repeated at least once, at least twice, at least 3 times, at least 4 times, at least 5 times, at least 6 times, at least 7 times, at least 8 times, at least 9 times, at least 10 times, or more than 10 times in the 5'UTR and/or 3'UTR, respectively. In some instances, the 5'UTR and/or 3'UTR comprises a TEE sequence-spacer module repeated 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 times. In some instances, the spacer separating two TEE sequences can include other sequences known in the art that can regulate the translation of the polynucleotide of the present disclosure, e.g., miR binding site sequences described herein (e.g., miR binding sites and miR seeds). As a non-limiting example, each spacer used to separate two TEE sequences can include a different miR binding site sequence or component of a miR sequence (e.g., miR seed sequence). In some instances, a polynucleotide of the present disclosure comprises a miR and/or TEE sequence. In some instances, the incorporation of a miR sequence and/or a TEE sequence into a polynucleotide of the present disclosure can change the shape of the stem loop region, which can increase and/or decrease translation. See e.g., Kedde et al., Nature Cell Biology 201012(10):1014-20, herein incorporated by reference in its entirety). mRNA: Coding region Provided herein are mRNA that encode the coding regions of the VHHs, polypeptides, bispecific antibodies, multispecific antibodies, and CARs of the disclosure.
In some instances, the disclosure provides mRNA that encode coding regions of a polypeptide which is at least 50%, at least 60%, at least 70%, at least 80%, at least 85%, at least 90%, at least 95%, or 100% identical to any of the VHH sequences described herein. In certain instances, the disclosure provides coding regions that encode a polypeptide which includes 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20 or more mutations of any of the sequences provided or referenced herein. In some instances, the mRNA comprises a codon optimized nucleic acid sequence, wherein the open reading frame (ORF) of the codon optimized nucleic acid sequence is derived from a polypeptide comprising an anti-c-MET VHH. For example, the polynucleotides can comprise a codon optimized ORF encoding a polypeptide comprising an anti-c-MET VHH. The nucleic acid molecules of the disclosure may include one or more alterations. Herein, in a nucleotide, nucleoside, or polynucleotide (such as the mRNA of the disclosure), the terms “alteration” or, as appropriate, “alternative” refer to alteration with respect to A, G, U or C ribonucleotides. The alterations may be various distinct alterations. In some instances, where the nucleic acid is an mRNA, the coding region, the flanking regions, and/or the terminal regions may contain one, two, or more (optionally different) nucleoside or nucleotide alterations. In some instances, an alternative polynucleotide introduced to a cell may exhibit reduced degradation in the cell, as compared to an unaltered polynucleotide. The polynucleotides can include any useful alteration, such as to the sugar, the nucleobase, or the internucleoside linkage (e.g., to a linking phosphate, to a phosphodiester linkage, or to the phosphodiester backbone). In certain instances, alterations (e.g., one or more alterations) are present in each of the sugar and the internucleoside linkage. Alterations according to the present disclosure may be alterations of ribonucleic acids (RNAs) to deoxyribonucleic acids (DNAs) (e.g., the substitution of the 2’OH of the ribofuranosyl ring to 2’H), threose nucleic acids (TNAs), glycol nucleic acids (GNAs), peptide nucleic acids (PNAs), locked nucleic acids (LNAs) or hybrids thereof. Additional alterations are described herein.
Modified Nucleic Acids According to Aduri et al., (Aduri, R. et al., Journal of Chemical Theory and Computation. 3(4):1464-75(2006)), there are 107 naturally occurring nucleosides, including 1-methyladenosine, 2-methylthio-N6-hydroxynorvalyl carbamoyladenosine, 2- methyladenosine, 2-O-ribosylphosphate adenosine, N6-methyl-N6- threonylcarbamoyladenosine, N6-acetyladenosine, N6-glycinylcarbamoyladenosine, N6- isopentenyladenosine, N6-methyladenosine, N6-threonylcarbamoyladenosine, N6,N6- dimethyladenosine, N6-(cis-hydroxyisopentenyl)adenosine, N6- hydroxynorvalylcarbamoyladenosine, 1,2-O-dimethyladenosine, N6,2-O- dimethyladenosine, 2-O-methyladenosine, N6,N6,O-2-trimethyladenosine, 2-methylthio- N6-(cis-hydroxyisopentenyl) adenosine, 2-methylthio-N6-methyladenosine, 2- methylthio-N6-isopentenyladenosine, 2-methylthio-N6-threonyl carbamoyladenosine, 2- thiocytidine, 3-methylcytidine , N4-acetylcytidine, 5-formylcytidine, N4-methylcytidine, 5-methylcytidine, 5-hydroxymethylcytidine, lysidine, N4-acetyl-2-O-methylcytidine, 5- formyl-2-O-methylcytidine, 5,2-O-dimethylcytidine, 2-O-methylcytidine, N4,2-O- dimethylcytidine, N4,N4,2-O-trimethylcytidine, 1-methylguanosine, N2,7- dimethylguanosine, N2-methylguanosine, 2-O-ribosylphosphate guanosine, 7- methylguanosine, under modified hydroxywybutosine, 7-aminomethyl-7- deazaguanosine, 7-cyano-7-deazaguanosine, N2,N2-dimethylguanosine, 4- demethylwyosine, epoxyqueuosine, hydroxywybutosine, isowyosine, N2,7,2-O- trimethylguanosine, N2,2-O-dimethylguanosine, 1,2-O-dimethylguanosine, 2-O- methylguanosine, N2,N2,2-O-trimethylguanosine, N2,N2,7-trimethylguanosine, peroxywybutosine, galactosyl-queuosine, mannosyl-queuosine, queuosine, archaeosine, wybutosine, methylwyosine, wyosine, 2-thiouridine, 3-(3-amino-3- carboxypropyl)uridine, 3-methyluridine, 4-thiouridine, 5-methyl-2-thiouridine, 5- methylaminomethyluridine, 5-carboxymethyluridine, 5- carboxymethylaminomethyluridine, 5-hydroxyuridine, 5-methyluridine, 5- taurinomethyluridine, 5-carbamoylmethyluridine, 5-(carboxyhydroxymethyl)uridine methyl ester, dihydrouridine, 5-methyldihydrouridine, 5-methylaminomethyl-2-
thiouridine, 5-(carboxyhydroxymethyl)uridine, 5-(isopentenylaminomethyl)uridine, 5- (isopentenylaminomethyl)-2-thiouridine, 3,2-O-dimethyluridine, 5- carboxymethylaminomethyl-2-O-methyluridine, 5-carbamoylmethyl-2-O-methyluridine, 5-methoxycarbonylmethyl-2-O-methyluridine, 5-(isopentenylaminomethyl)-2-O- methyluridine, 5,2-O-dimethyluridine, 2-O-methyluridine, 2-thio-2-O-methyluridine, uridine 5-oxyacetic acid, 5-methoxycarbonylmethyluridine, uridine 5-oxyacetic acid methyl ester, 5-methoxyuridine, 5-aminomethyl-2-thiouridine, 5- carboxymethylaminomethyl-2-thiouridine, 5-methylaminomethyl-2-selenouridine, 5- methoxycarbonylmethyl-2-thiouridine, 5-taurinomethyl-2-thiouridine, pseudouridine, 1- methyl-3-(3-amino-3-carboxypropyl)pseudouridine, 1-methylpseudouridine, 3- methylpseudouridine, 2-O-methylpseudouridine, inosine, 1-methylinosine, 1,2-O- dimethylinosine, and 2-O-methylinosine. Each of these may be components of nucleic acids (e.g., mRNA) of the present disclosure. Nucleosides Containing Modified Sugars The alternative nucleosides and nucleotides (e.g., building block molecules), which may be incorporated into a polynucleotide (e.g., RNA or mRNA, as described herein), can be altered on the sugar of the ribonucleic acid. For example, the 2' hydroxyl group (OH) can be modified or replaced with a number of different substituents. Exemplary substitutions at the 2'-position include, but are not limited to, H, halo, optionally substituted C1-6 alkyl; optionally substituted C1-6 alkoxy; optionally substituted C6-10 aryloxy; optionally substituted C3-8 cycloalkyl; optionally substituted C3-8 cycloalkoxy; optionally substituted C6-10 aryloxy; optionally substituted C6-10 aryl-C1-6 alkoxy, optionally substituted C1-12 (heterocyclyl)oxy; a sugar (e.g., ribose, pentose, or any described herein); a polyethyleneglycol (PEG), -O(CH2CH2O)nCH2CH2OR, where R is H or optionally substituted alkyl, and n is an integer from 0 to 20 (e.g., from 0 to 4, from 0 to 8, from 0 to 10, from 0 to 16, from 1 to 4, from 1 to 8, from 1 to 10, from 1 to 16, from 1 to 20, from 2 to 4, from 2 to 8, from 2 to 10, from 2 to 16, from 2 to 20, from 4 to 8, from 4 to 10, from 4 to 16, and from 4 to 20); “locked” nucleic acids (LNA) in which the 2'-hydroxyl is connected by a C1-6 alkylene or C1-6 heteroalkylene bridge to the
4’-carbon of the same ribose sugar, where exemplary bridges included methylene, propylene, ether, or amino bridges; aminoalkyl, as defined herein; aminoalkoxy, as defined herein; amino as defined herein; and amino acid, as defined herein Generally, RNA includes the sugar group ribose, which is a 5-membered ring having an oxygen. Exemplary, non-limiting alternative nucleotides include replacement of the oxygen in ribose (e.g., with S, Se, or alkylene, such as methylene or ethylene); addition of a double bond (e.g., to replace ribose with cyclopentenyl or cyclohexenyl); ring contraction of ribose (e.g., to form a 4-membered ring of cyclobutane or oxetane); ring expansion of ribose (e.g., to form a 6- or 7-membered ring having an additional carbon or heteroatom, such as for anhydrohexitol, altritol, mannitol, cyclohexanyl, cyclohexenyl, and morpholino that also has a phosphoramidate backbone); multicyclic forms (e.g., tricyclo; and “unlocked” forms, such as glycol nucleic acid (GNA) (e.g., R- GNA or S-GNA, where ribose is replaced by glycol units attached to phosphodiester bonds), threose nucleic acid (TNA, where ribose is replace with α-L-threofuranosyl- (3'ĺ2')), and peptide nucleic acid (PNA, where 2-amino-ethyl-glycine linkages replace the ribose and phosphodiester backbone). The sugar group can also contain one or more carbons that possess the opposite stereochemical configuration than that of the corresponding carbon in ribose. Thus, a polynucleotide molecule can include nucleotides containing, e.g., arabinose, as the sugar. Alterations on the Nucleobase The present disclosure provides for alternative nucleosides and nucleotides. As described herein “nucleoside” is defined as a compound containing a sugar molecule (e.g., a pentose or ribose) or derivative thereof in combination with an organic base (e.g., a purine or pyrimidine) or a derivative thereof (also referred to herein as “nucleobase”). As described herein, “nucleotide” is defined as a nucleoside including a phosphate group. Exemplary non-limiting alterations include an amino group, a thiol group, an alkyl group, a halo group, or any described herein. The alternative nucleotides may by
synthesized by any useful method, as described herein (e.g., chemically, enzymatically, or recombinantly to include one or more alternative or alternative nucleosides). In some instances, a nucleic acid of the disclosure (e.g., an mRNA) includes one or more 2’-OMe nucleotides, 2’-methoxyethyl nucleotides (2’-MOE nucleotides), 2’-F nucleotide, 2’-NH2 nucleotide, 2’fluoroarabino nucleotides (FANA nucleotides), locked nucleic acid nucleotides (LNA nucleotides), or 4’-S nucleotides. The alternative nucleotide base pairing encompasses not only the standard adenosine-thymine, adenosine-uracil, and guanosine-cytosine base pairs, but also base pairs formed between nucleotides and/or alternative nucleotides including non-standard or alternative bases, wherein the arrangement of hydrogen bond donors and hydrogen bond acceptors permits hydrogen bonding between a non-standard base and a standard base or between two complementary non-standard base structures. One example of such non-standard base pairing is the base pairing between the alternative nucleotide inosine and adenine, cytosine, or uracil. The alternative nucleosides and nucleotides can include an alternative nucleobase. Examples of nucleobases found in RNA include, but are not limited to, adenine, guanine, cytosine, and uracil. Examples of nucleobase found in DNA include, but are not limited to, adenine, guanine, cytosine, and thymine. These nucleobases can be altered or wholly replaced to provide polynucleotide molecules having enhanced properties (e.g., resistance to nucleases and stability), and these properties may manifest through disruption of the binding of a major groove binding partner. In some instances, the alternative nucleobase is an alternative uracil. Exemplary nucleobases and nucleosides having an alternative uracil include pseudouridine (Ψ), pyridin-4-one ribonucleoside, 5-aza-uridine, 6-aza-uridine, 2-thio-5-aza-uridine, 2-thio- uridine (s2U), 4-thio-uridine (s4U), 4-thio-pseudouridine, 2-thio-pseudouridine, 5- hydroxy-uridine (ho5U), 5-aminoallyl-uridine, 5-halo-uridine (e.g., 5-iodo-uridineor 5- bromo-uridine), 3-methyl-uridine (m3U), 5-methoxy-uridine (mo5U), uridine 5-oxyacetic
acid (cmo5U), uridine 5-oxyacetic acid methyl ester (mcmo5U), 5-carboxymethyl-uridine (cm5U), 1-carboxymethyl-pseudouridine, 5-carboxyhydroxymethyl-uridine (chm5U), 5- carboxyhydroxymethyl-uridine methyl ester (mchm5U), 5-methoxycarbonylmethyl- uridine (mcm5U), 5-methoxycarbonylmethyl-2-thio-uridine (mcm5s2U), 5-aminomethyl- 2-thio-uridine (nm5s2U), 5-methylaminomethyl-uridine (mnm5U), 5-methylaminomethyl- 2-thio-uridine (mnm5s2U), 5-methylaminomethyl-2-seleno-uridine (mnm5se2U), 5- carbamoylmethyl-uridine (ncm5U), 5-carboxymethylaminomethyl-uridine (cmnm5U), 5- carboxymethylaminomethyl-2-thio-uridine (cmnm5s2U), 5-propynyl-uridine, 1-propynyl- pseudouridine, 5-taurinomethyl-uridine (IJm5U), 1-taurinomethyl-pseudouridine, 5- taurinomethyl-2-thio-uridine(IJm5s2U), 1-taurinomethyl-4-thio-pseudouridine, 5-methyl- uridine (m5U, i.e., having the nucleobase deoxythymine), 1-methyl-pseudouridine (m1Ψ), 5-methyl-2-thio-uridine (m5s2U), 1-methyl-4-thio-pseudouridine (m1s4Ψ), 4-thio-1- methyl-pseudouridine, 3-methyl-pseudouridine (m3Ψ), 2-thio-1-methyl-pseudouridine, 1- methyl-1-deaza-pseudouridine, 2-thio-1-methyl-1-deaza-pseudouridine, dihydrouridine (D), dihydropseudouridine, 5,6-dihydrouridine, 5-methyl-dihydrouridine (m5D), 2-thio- dihydrouridine, 2-thio-dihydropseudouridine, 2-methoxy-uridine, 2-methoxy-4-thio- uridine, 4-methoxy-pseudouridine, 4-methoxy-2-thio-pseudouridine, N1-methyl- pseudouridine, 3-(3-amino-3-carboxypropyl)uridine (acp3U), 1-methyl-3-(3-amino-3- carboxypropyl)pseudouridine (acp3 Ψ), 5-(isopentenylaminomethyl)uridine (inm5U), 5- (isopentenylaminomethyl)-2-thio-uridine (inm5s2U), α-thio-uridine, 2'-O-methyl-uridine (Um), 5,2'-O-dimethyl-uridine (m5Um), 2'-O-methyl-pseudouridine (Ψm), 2-thio-2'-O- methyl-uridine (s2Um), 5-methoxycarbonylmethyl-2'-O-methyl-uridine (mcm5Um), 5- carbamoylmethyl-2'-O-methyl-uridine (ncm5Um), 5-carboxymethylaminomethyl-2'-O- methyl-uridine (cmnm5Um), 3,2'-O-dimethyl-uridine (m3Um), and 5- (isopentenylaminomethyl)-2'-O-methyl-uridine (inm5Um), 1-thio-uridine, deoxythymidine, 2’-F-ara-uridine, 2’-F-uridine, 2’-OH-ara-uridine, 5-(2- carbomethoxyvinyl) uridine, and 5-[3-(1-E-propenylamino)uridine. In one instance, the nucleic acid is modified to contain 1-methylpseudouridine (m1Ψ) in lieu of uridine at each instance.
In some instances, the alternative nucleobase is an alternative cytosine. Exemplary nucleobases and nucleosides having an alternative cytosine include 5-aza- cytidine, 6-aza-cytidine, pseudoisocytidine, 3-methyl-cytidine (m3C), N4-acetyl-cytidine (ac4C), 5-formyl-cytidine (f5C), N4-methyl-cytidine (m4C), 5-methyl-cytidine (m5C), 5- halo-cytidine (e.g., 5-iodo-cytidine), 5-hydroxymethyl-cytidine (hm5C), 1-methyl- pseudoisocytidine, pyrrolo-cytidine, pyrrolo-pseudoisocytidine, 2-thio-cytidine (s2C), 2- thio-5-methyl-cytidine, 4-thio-pseudoisocytidine, 4-thio-1-methyl-pseudoisocytidine, 4- thio-1-methyl-1-deaza-pseudoisocytidine, 1-methyl-1-deaza-pseudoisocytidine, zebularine, 5-aza-zebularine, 5-methyl-zebularine, 5-aza-2-thio-zebularine, 2-thio- zebularine, 2-methoxy-cytidine, 2-methoxy-5-methyl-cytidine, 4-methoxy- pseudoisocytidine, 4-methoxy-1-methyl-pseudoisocytidine, lysidine (k2C), α-thio- cytidine, 2'-O-methyl-cytidine (Cm), 5,2'-O-dimethyl-cytidine (m5Cm), N4-acetyl-2'-O- methyl-cytidine (ac4Cm), N4,2'-O-dimethyl-cytidine (m4Cm), 5-formyl-2'-O-methyl- cytidine (f5Cm), N4,N4,2'-O-trimethyl-cytidine (m4 2Cm), 1-thio-cytidine, 2’-F-ara- cytidine, 2’-F-cytidine, and 2’-OH-ara-cytidine. In some instances, the alternative nucleobase is an alternative adenine. Exemplary nucleobases and nucleosides having an alternative adenine include 2-amino-purine, 2, 6- diaminopurine, 2-amino-6-halo-purine (e.g., 2-amino-6-chloro-purine), 6-halo-purine (e.g., 6-chloro-purine), 2-amino-6-methyl-purine, 8-azido-adenosine, 7-deaza-adenine, 7- deaza-8-aza-adenine, 7-deaza-2-amino-purine, 7-deaza-8-aza-2-amino-purine, 7-deaza- 2,6-diaminopurine, 7-deaza-8-aza-2,6-diaminopurine, 1-methyl-adenosine (m1A), 2- methyl-adenine (m2A), N6-methyl-adenosine (m6A), 2-methylthio-N6-methyl-adenosine (ms2m6A), N6-isopentenyl-adenosine (i6A), 2-methylthio-N6-isopentenyl-adenosine (ms2i6A), N6-(cis-hydroxyisopentenyl)adenosine (io6A), 2-methylthio-N6-(cis- hydroxyisopentenyl)adenosine (ms2io6A), N6-glycinylcarbamoyl-adenosine (g6A), N6- threonylcarbamoyl-adenosine (t6A), N6-methyl-N6-threonylcarbamoyl-adenosine (m6t6A), 2-methylthio-N6-threonylcarbamoyl-adenosine (ms2g6A), N6,N6-dimethyl- adenosine (m62A), N6-hydroxynorvalylcarbamoyl-adenosine (hn6A), 2-methylthio-N6- hydroxynorvalylcarbamoyl-adenosine (ms2hn6A), N6-acetyl-adenosine (ac6A), 7-methyl-
adenine, 2-methylthio-adenine, 2-methoxy-adenine, α-thio-adenosine, 2'-O-methyl- adenosine (Am), N6,2'-O-dimethyl-adenosine (m6Am), N6,N6,2'-O-trimethyl-adenosine (m6 2Am), 1,2'-O-dimethyl-adenosine (m1Am), 2'-O-ribosyladenosine (phosphate) (Ar(p)), 2-amino-N6-methyl-purine, 1-thio-adenosine, 8-azido-adenosine, 2’-F-ara- adenosine, 2’-F-adenosine, 2’-OH-ara-adenosine, and N6-(19-amino- pentaoxanonadecyl)-adenosine. In some instances, the alternative nucleobase is an alternative guanine. Exemplary nucleobases and nucleosides having an alternative guanine include inosine (I), 1-methyl- inosine (m1I), wyosine (imG), methylwyosine (mimG), 4-demethyl-wyosine (imG-14), isowyosine (imG2), wybutosine (yW), peroxywybutosine (o2yW), hydroxywybutosine (OhyW), undermodified hydroxywybutosine (OhyW*), 7-deaza-guanosine, queuosine (Q), epoxyqueuosine (oQ), galactosyl-queuosine (galQ), mannosyl-queuosine (manQ), 7- cyano-7-deaza-guanosine (preQ0), 7-aminomethyl-7-deaza-guanosine (preQ1), archaeosine (G+), 7-deaza-8-aza-guanosine, 6-thio-guanosine, 6-thio-7-deaza-guanosine, 6-thio-7-deaza-8-aza-guanosine, 7-methyl-guanosine (m7G), 6-thio-7-methyl-guanosine, 7-methyl-inosine, 6-methoxy-guanosine, 1-methyl-guanosine (m1G), N2-methyl- guanosine (m2G), N2,N2-dimethyl-guanosine (m2 2G), N2,7-dimethyl-guanosine (m2,7G), N2, N2,7-dimethyl-guanosine (m2,2,7G), 8-oxo-guanosine, 7-methyl-8-oxo-guanosine, 1- methyl-6-thio-guanosine, N2-methyl-6-thio-guanosine, N2,N2-dimethyl-6-thio- guanosine, α-thio-guanosine, 2'-O-methyl-guanosine (Gm), N2-methyl-2'-O-methyl- guanosine (m2Gm), N2,N2-dimethyl-2'-O-methyl-guanosine (m22Gm), 1-methyl-2'-O- methyl-guanosine (m1Gm), N2,7-dimethyl-2'-O-methyl-guanosine (m2,7Gm), 2'-O- methyl-inosine (Im), 1,2'-O-dimethyl-inosine (m1Im), 2'-O-ribosylguanosine (phosphate) (Gr(p)) , 1-thio-guanosine, O6-methyl-guanosine, 2’-F-ara-guanosine, and 2’-F- guanosine. The nucleobase of the nucleotide can be independently selected from a purine, a pyrimidine, a purine, or pyrimidine analog. For example, the nucleobase can each be independently selected from adenine, cytosine, guanine, uracil, or hypoxanthine. In some
instances, the nucleobase can also include, for example, naturally-occurring and synthetic derivatives of a base, including pyrazolo[3,4-d]pyrimidines, 5-methylcytosine (5-me-C), 5-hydroxymethyl cytosine, xanthine, hypoxanthine, 2-aminoadenine, 6-methyl, and other alkyl derivatives of adenine and guanine, 2-propyl and other alkyl derivatives of adenine and guanine, 2-thiouracil, 2-thiothymine and 2-thiocytosine, 5-propynyl uracil and cytosine, 6-azo uracil, cytosine and thymine, 5-uracil (pseudouracil), 4-thiouracil, 8-halo (e.g., 8-bromo), 8-amino, 8-thiol, 8-thioalkyl, 8-hydroxyl and other 8-substituted adenines and guanines, 5-halo particularly 5-bromo, 5-trifluoromethyl and other 5- substituted uracils and cytosines, 7-methylguanine and 7-methyladenine, 8-azaguanine and 8-azaadenine, deazaguanine, 7-deazaguanine, 3-deazaguanine, deazaadenine, 7- deazaadenine, 3-deazaadenine, pyrazolo[3,4-d]pyrimidine, imidazo[1,5-a]1,3,5 triazinones, 9-deazapurines, imidazo[4,5-d]pyrazines, thiazolo[4,5-d]pyrimidines, pyrazin-2-ones, 1,2,4-triazine, pyridazine; and 1,3,5 triazine. When the nucleotides are depicted using the shorthand A, G, C, T or U, each letter refers to the representative base and/or derivatives thereof (e.g., A includes adenine or adenine analogs (e.g., 7-deaza adenine)). In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5-methyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5- trifluoromethyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5- hydroxymethyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5- bromo-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5-iodo-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5-methoxy-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5- methoxy-uracil, uracil, 5-ethyl-cytosine, and cytosine as the only uracils and cytosines. In
some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5- phenyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5-ethnyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, N4-methyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5- methoxy-uracil, uracil, 5-fluoro-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, N4-acetyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, pseudoisocytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5-formyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uracil, uracil, 5-aminoallyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5- methoxy-uracil, uracil, 5-carboxy-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl- pseudouracil, uracil, 5-methyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-trifluoromethyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5- hydroxymethyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5- bromo-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-iodo-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-methoxy-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure
contain 1-methyl-pseudouracil, uracil, 5-ethyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl- pseudouracil, uracil, 5-phenyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-ethnyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, N4-methyl-cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-fluoro- cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, N4-acetyl- cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, pseudoisocytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-formyl- cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-aminoallyl- cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouracil, uracil, 5-carboxy- cytosine, and cytosine as the only uracils and cytosines. In some instances, the polynucleotides of the disclosure contain 5-methoxy- uridine, uridine, 5-methyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-trifluoromethyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5- hydroxymethyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5- bromo-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-iodo-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of
the disclosure contain 5-methoxy-uridine, uridine, 5-methoxy-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-ethyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5- methoxy-uridine, uridine, 5-phenyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy- uridine, uridine, 5-ethnyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, N4-methyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-fluoro- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, N4-acetyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, pseudoisocytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy-uridine, uridine, 5-formyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5- methoxy-uridine, uridine, 5-aminoallyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 5-methoxy- uridine, uridine, 5-carboxy-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl- pseudouridine, uridine, 5-methyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-trifluoromethyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-hydroxymethyl-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-bromo-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine,
5-iodo-cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-methoxy- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-ethyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-phenyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-ethnyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, N4-methyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-fluoro- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, N4-acetyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, pseudoisocytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-formyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-aminoallyl- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain 1-methyl-pseudouridine, uridine, 5-carboxy- cytidine, and cytidine as the only uridines and cytidines. In some instances, the polynucleotides of the disclosure contain the uracil of one of the nucleosides of Table 9 and uracil as the only uracils. In other instances, the polynucleotides of the disclosure contain a uridine of Table 9 and uridine as the only uridines.
Table 9 – Exemplary Uracil-containing Nucleosides
In some instances, the polynucleotides of the disclosure contain the cytosine of one of the nucleosides of Table 10 and cytosine as the only cytosines. In other instances, the polynucleotides of the disclosure contain a cytidine of Table 10 and cytidine as the only cytidines. Table 10 – Exemplary Cytosine Containing Nucleosides
Alterations on the Internucleoside Linkage The alternative nucleotides, which may be incorporated into a polynucleotide molecule, can be altered on the internucleoside linkage (e.g., phosphate backbone). Herein, in the context of the polynucleotide backbone, the phrases “phosphate” and “phosphodiester” are used interchangeably. Backbone phosphate groups can be altered by replacing one or more of the oxygen atoms with a different substituent. The alternative nucleosides and nucleotides can include the wholesale replacement of an unaltered phosphate moiety with another internucleoside linkage as described herein. Examples of alternative phosphate groups include, but are not limited to, phosphorothioate, phosphoroselenates, boranophosphates, boranophosphate esters, hydrogen phosphonates, phosphoramidates, phosphorodiamidates, alkyl or aryl phosphonates, and phosphotriesters. Phosphorodithioates have both non-linking oxygens replaced by sulfur. The phosphate linker can also be altered by the replacement of a linking oxygen with nitrogen (bridged phosphoramidates), sulfur (bridged phosphorothioates), and carbon (bridged methylene-phosphonates). The alternative nucleosides and nucleotides can include the replacement of one or more of the non-bridging oxygens with a borane moiety (BH3), sulfur (thio), methyl, ethyl and/or methoxy. As a non-limiting example, two non-bridging oxygens at the same position (e.g., the alpha (α), beta (β) or gamma (γ) position) can be replaced with a sulfur (thio) and a methoxy.
The replacement of one or more of the oxygen atoms at the α position of the phosphate moiety (e.g., α-thio phosphate) is provided to confer stability (such as against exonucleases and endonucleases) to RNA and DNA through the unnatural phosphorothioate backbone linkages. Phosphorothioate DNA and RNA have increased nuclease resistance and subsequently a longer half-life in a cellular environment. While not wishing to be bound by theory, phosphorothioate linked polynucleotide molecules are expected to also reduce the innate immune response through weaker binding/activation of cellular innate immune molecules. In specific instances, an alternative nucleoside includes an alpha-thio-nucleoside (e.g., 5'-O-(1-thiophosphate)-adenosine, 5'-O-(1-thiophosphate)-cytidine (α-thio- cytidine), 5'-O-(1-thiophosphate)-guanosine, 5'-O-(1-thiophosphate)-uridine, or 5'-O-(1- thiophosphate)-pseudouridine). Other internucleoside linkages that may be employed according to the present disclosure, including internucleoside linkages which do not contain a phosphorous atom, are described herein below. Combinations of Alternative Sugars, Nucleobases, and Internucleoside Linkages The polynucleotides of the disclosure can include a combination of alterations to the sugar, the nucleobase, and/or the internucleoside linkage. These combinations can include any one or more alterations described herein. mRNA: Poly-A tail During RNA processing, a long chain of adenosine nucleotides (poly(A) tail) is normally added to mRNA molecules to increase the stability of the mRNA. Immediately after transcription, the 3' end of the transcript is cleaved to free a 3' hydroxyl. Then poly(A) polymerase adds a chain of adenosine nucleotides to the RNA. The process, called polyadenylation, adds a poly-A tail that is between 100 and 250 residues long.
Methods for the stabilization of RNA by incorporation of chain-terminating nucleosides at the 3 ’-terminus include those described in International Patent Publication No. WO2013/103659, incorporated herein in its entirety.
Poly (A) tail deadenylation by 3' exonucleases is a key step in cellular mRNA degradation in eukaryotes. By blocking 3' exonucleases, the functional half-life of mRNA can be increased, resulting in increased protein expression. Chemical and enzymatic ligation strategies to modify the 3' end of mRNA with reverse chirality adenosine (LA10) and/or inverted deoxythymidine (IdT) are known to those of skill in the art and have been demonstrated to extend mRNA half-life in cellular and in vivo studies. In some instances, the poly(A)tail of the mRNA includes a 3’ LA10 or IdT modification. For example, as described in International Patent Publication No. WO2017/049275, the tail modifications of which are incorporated by reference in their entirety.
Additional strategies have been explored to further stabilize mRNA, including: chemical modification of the 3’ nucleotide (e.g., conjugation of a morpholino to the 3’ end of the poly(A)tail); incorporation of stabilizing sequences after the poly(A) tail (e.g., a co-polymer, a stem-loop, or a triple helix); and/or annealing of structured oligos to the 3' end of an mRNA, as described, for example, in International Patent Publication No. WO2017/049286, the stabilized linkages of which are incorporated by reference in their entirety.
Annealing an oligonucleotide (e.g., an oligonucleotide conjugate) with a complex secondary structure (e.g., a triple-helix structure or a stem-loop structure) at the 3 ’end may provide nuclease resistance and increase half-life of mRNA.
Unique poly(A) tail lengths may provide certain advantages to the RNAs of the present disclosure. Generally, the length of a poly(A) tail of the present disclosure is greater than 30 nucleotides in length. In some instances, the poly(A) tail is greater than 35 nucleotides in length. In some instances, the length is at least 40 nucleotides. In another embodiment, the length is at least 45 nucleotides. In some instances, the length is
at least 50 nucleotides. In some instances, the length is at least 55 nucleotides. In another embodiment, the length is at least 60 nucleotides. In another embodiment, the length is at least 65 nucleotides. In another embodiment, the length is at least 70 nucleotides. In some instances, the length is at least 80 nucleotides. In some instances, the length is at least 90 nucleotides. In some instances, the length is at least 100 nucleotides. In some instances, the length is at least 120 nucleotides. In some instances, the length is at least 140 nucleotides. In some instances, the length is at least 160 nucleotides. In some instances, the length is at least 180 nucleotides. In some instances, the length is at least 200 nucleotides. In some instances, the length is at least 250 nucleotides. In some instances, the length is at least 300 nucleotides. In some instances, the length is at least 350 nucleotides. In some instances, the length is at least 400 nucleotides. In some instances, the length is at least 450 nucleotides. In some instances, the length is at least 500 nucleotides. In some instances, the length is at least 600 nucleotides. In some instances, the length is at least 700 nucleotides. In some instances, the length is at least 800 nucleotides. In some instances, the length is at least 900 nucleotides. In some instances, the length is at least 1000 nucleotides. In some instances, the length is at least 1100 nucleotides. In some instances, the length is at least 1200 nucleotides. In some instances, the length is at least 1300 nucleotides. In some instances, the length is at least 1400 nucleotides. In some instances, the length is at least 1500 nucleotides. In some instances, the length is at least 1600 nucleotides. In some instances, the length is at least 1700 nucleotides. In some instances, the length is at least 1800 nucleotides. In some instances, the length is at least 1900 nucleotides. In some instances, the length is at least 2000 nucleotides. In some instances, the length is at least 2500 nucleotides. In some instances, the length is at least 3000 nucleotides.
In some instances, the poly(A) tail may be 80 nucleotides, 120 nucleotides, or 160 nucleotides in length. In some instances, the poly(A) tail may be 20, 40, 80, 100, 120, 140 or 160 nucleotides in length.
In some instances, the poly(A) tail is designed relative to the length of the mRNA. This design may be based on the length of the coding region of the mRNA, the length of a particular feature or region of the mRNA, or based on the length of the ultimate product expressed from the RNA. When relative to any additional feature of the RNA (e.g., other than the mRNA portion which includes the poly(A) tail), poly(A) tail may be 10, 20, 30, 40, 50, 60, 70, 80, 90 or 100% greater in length than the additional feature. The poly(A) tail may also be designed as a fraction of the mRNA to which it belongs. In this context, the poly(A) tail may be 10, 20, 30, 40, 50, 60, 70, 80, or 90% or more of the total length of the construct or the total length of the construct minus the poly(A) tail. In some instances, engineered binding sites and/or the conjugation of nucleic acids or mRNA for poly(A) binding protein may be used to enhance expression. The engineered binding sites may be sensor sequences which can operate as binding sites for ligands of the local microenvironment of the nucleic acids and/or mRNA. As a non- limiting example, the nucleic acids and/or mRNA may include at least one engineered binding site to alter the binding affinity of poly(A) binding protein (PABP) and analogs thereof. The incorporation of at least one engineered binding site may increase the binding affinity of the PABP and analogs thereof. Additionally, multiple distinct nucleic acids or mRNA may be linked together to the PABP (poly(A) binding protein) through the 3'-end using nucleotides at the 3'- terminus of the poly(A) tail. Transfection experiments can be conducted in relevant cell lines and protein production can be assayed by ELISA at 12hr, 24hr, 48hr, 72hr, and day 7 post-transfection. As a non-limiting example, the transfection experiments may be used to evaluate the effect on PABP or analogs thereof binding affinity as a result of the addition of at least one engineered binding site. In some instances, a poly(A) tail may be used to modulate translation initiation. While not wishing to be bound by theory, the poly-A tail recruits PABP which in turn can interact with translation initiation complex and thus may be essential for protein synthesis.
In some instances, a poly(A) tail may also be used in the present disclosure to protect against 3’-5’ exonuclease digestion. In some instances, the nucleic acids or mRNA of the present disclosure are designed to include a poly-A-G Quartet. The G-quartet is a cyclic hydrogen bonded array of four guanosine nucleotides that can be formed by G-rich sequences in both DNA and RNA. In this embodiment, the G-quartet is incorporated at the end of the poly-A tail. The resultant nucleic acid or mRNA may be assayed for stability, protein production and other parameters including half-life at various time points. It has been discovered that the poly-A-G quartet results in protein production equivalent to at least 75% of that seen using a poly-A tail of 120 nucleotides alone. In some instances, the nucleic acids or mRNA of the present disclosure may include a poly(A) tail and may be stabilized by the addition of a chain terminating nucleoside. The nucleic acids and/or mRNA with a poly(A) tail may further include a 5’cap structure. In some instances, the nucleic acids or mRNA of the present disclosure may include a poly-A-G Quartet. The nucleic acids and/or mRNA with a poly-A-G Quartet may further include a 5’cap structure. In some instances, the chain terminating nucleoside which may be used to stabilize the nucleic acid or mRNA including a poly(A) tail or poly-A-G Quartet may be, but is not limited to, those described in International Patent Publication No. WO2013103659, incorporated herein by reference in its entirety. In some instances, the chain terminating nucleosides which may be used with the present disclosure includes, but is not limited to, 3'-deoxyadenosine (cordycepin), 3'-deoxyuridine, 3'-deoxycytosine, 3'-deoxyguanosine, 3'-deoxythymine, 2',3'-dideoxynucleosides, such as 2',3'- dideoxyadenosine, 2',3'-dideoxyuridine, 2',3'-dideoxycytosine, 2',3'- dideoxyguanosine, 2',3'-dideoxythymine, a 2'-deoxynucleoside, or a -O- methylnucleoside.
In some instances, the mRNA which includes a poly(A) tail or a poly-A-G Quartet may be stabilized by an alteration to the 3’region of the nucleic acid that can prevent and/or inhibit the addition of oligo(U) (see, e.g., International Patent Publication No. WO2013/103659, incorporated herein by reference in its entirety). In yet another embodiment, the mRNA, which includes a poly(A) tail or a poly- A-G Quartet, may be stabilized by the addition of an oligonucleotide that terminates in a 3’-deoxynucleoside, 2’,3’-dideoxynucleoside 3'-O-methylnucleosides, 3'-O- ethylnucleosides, 3'-arabinosides, and other alternative nucleosides known in the art and/or described herein Synthesis of Polynucleotides The polynucleotide molecules (e.g., mRNA) for use in accordance with the disclosure may be prepared according to any useful technique, as described herein. The alternative nucleosides and nucleotides used in the synthesis of polynucleotide molecules disclosed herein can be prepared from readily available starting materials using the following general methods and procedures. Where typical or preferred process conditions (e.g., reaction temperatures, times, mole ratios of reactants, solvents, pressures, etc.) are provided, a skilled artisan would be able to optimize and develop additional process conditions. Optimum reaction conditions may vary with the particular reactants or solvent used, but such conditions can be determined by one skilled in the art by routine optimization procedures. The processes described herein can be monitored according to any suitable method known in the art. For example, product formation can be monitored by spectroscopic means, such as nuclear magnetic resonance spectroscopy (e.g., 1H or 13C) infrared spectroscopy, spectrophotometry (e.g., UV-visible), or mass spectrometry, or by chromatography (e.g., high performance liquid chromatography (HPLC) or thin layer chromatography).
Preparation of polynucleotide molecules of the present disclosure can involve the protection and deprotection of various chemical groups. The need for protection and deprotection, and the selection of appropriate protecting groups can be readily determined by one skilled in the art. The chemistry of protecting groups can be found, for example, in Greene, et al., Protective Groups in Organic Synthesis, 2d. Ed., Wiley & Sons, (1991), which is incorporated herein by reference in its entirety. The reactions of the processes described herein can be carried out in suitable solvents, which can be readily selected by one of skill in the art of organic synthesis. Suitable solvents can be substantially nonreactive with the starting materials (reactants), the intermediates, or products at the temperatures at which the reactions are carried out (i.e., temperatures which can range from the solvent’s freezing temperature to the solvent’s boiling temperature). A given reaction can be carried out in one solvent or a mixture of more than one solvent. Depending on the particular reaction step, suitable solvents for a particular reaction step can be selected. Resolution of racemic mixtures of alternative polynucleotides or nucleic acids (e.g., polynucleotides or mRNA molecules) can be carried out by any of numerous methods known in the art. An example method includes fractional recrystallization using a “chiral resolving acid” which is an optically active, salt-forming organic acid. Suitable resolving agents for fractional recrystallization methods are, for example, optically active acids, such as the D and L forms of tartaric acid, acid, dibenzoyltartaric acid, mandelic acid, malic acid, lactic acid or the various optically active camphorsulfonic acids. Resolution of racemic mixtures can also be carried out by elution on a column packed with an optically active resolving agent (e.g., dinitrobenzoylphenylglycine). Suitable elution solvent composition can be determined by one skilled in the art. Alternative nucleosides and nucleotides (e.g., building block molecules) can be prepared according to the synthetic methods described in Ogata et al., J. Org. Chem. 74:2585-2588 (2009); Purmal et al., Nucl. Acids Res.22(1): 72-78 (1994); Fukuhara et
al., Biochemistry, 1(4): 563-568 (1962); and Xu et al., Tetrahedron, 48(9): 1729-1740 (1992), each of which are incorporated by reference in their entirety. If the polynucleotide includes one or more alternative nucleosides or nucleotides, the polynucleotides of the disclosure may or may not be uniformly altered along the entire length of the molecule. For example, one or more or all types of nucleotide (e.g., purine or pyrimidine, or any one or more or all of A, G, U, C) may or may not be uniformly altered in a polynucleotide of the disclosure, or in a given predetermined sequence region thereof. In some embodiments, all nucleotides X in a polynucleotide of the disclosure (or in a given sequence region thereof) are altered, wherein X may any one of nucleotides A, G, U, C, or any one of the combinations A+G, A+U, A+C, G+U, G+C, U+C, A+G+U, A+G+C, G+U+C or A+G+C. Different sugar alterations, nucleotide alterations, and/or internucleoside linkages (e.g., backbone structures) may exist at various positions in the polynucleotide. One of ordinary skill in the art will appreciate that the nucleotide analogs or other alteration(s) may be located at any position(s) of a polynucleotide such that the function of the polynucleotide is not substantially decreased. An alteration may also be a 5' or 3' terminal alteration. The polynucleotide may contain from 1% to 100% alternative nucleosides, nucleotides, or internucleoside linkages (either in relation to overall nucleotide content, or in relation to one or more types of nucleotide, i.e. any one or more of A, G, U or C) or any intervening percentage (e.g., from 1% to 20%, from 1% to 25%, from 1% to 50%, from 1% to 60%, from 1% to 70%, from 1% to 80%, from 1% to 90%, from 1% to 95%, from 10% to 20%, from 10% to 25%, from 10% to 50%, from 10% to 60%, from 10% to 70%, from 10% to 80%, from 10% to 90%, from 10% to 95%, from 10% to 100%, from 20% to 25%, from 20% to 50%, from 20% to 60%, from 20% to 70%, from 20% to 80%, from 20% to 90%, from 20% to 95%, from 20% to 100%, from 50% to 60%, from 50% to 70%, from 50% to 80%, from 50% to 90%, from 50% to 95%, from 50% to 100%, from 70% to 80%, from 70% to 90%, from 70% to 95%, from 70% to 100%, from 80% to 90%, from 80% to 95%, from 80% to 100%, from 90% to 95%, from 90% to 100%,
and from 95% to 100. In some embodiments, the remaining percentage is accounted for by the presence of A, G, U, or C.
When referring to percentage incorporation by an alternative nucleoside, nucleotide, or internucleoside linkage, in some embodiments the remaining percentage necessary to total 100% is accounted for by the corresponding natural nucleoside, nucleotide, or internucleoside linkage. In other embodiments, the remaining percentage necessary to total 100% is accounted for by a second alternative nucleoside, nucleotide, or internucleoside linkage.
Nanoparticles
The present disclosure also provides LNP compositions. The lipid nanoparticle compositions described herein may be used for the delivery of therapeutic and/or prophylactic agents, e.g., mRNAs, to mammalian cells or organs. For example, the lipid nanoparticles described herein have little or no immunogenicity. For example, the lipid compounds disclosed herein have a lower immunogenicity as compared to a reference lipid (e.g, MC3, KC2, or DLinDMA). For example, a formulation comprising a lipid disclosed herein and a therapeutic or prophylactic agent, e.g, mRNA, has an increased therapeutic index as compared to a corresponding formulation which comprises a reference lipid (e.g, MC3, KC2, or DLinDMA) and the same therapeutic or prophylactic agent.
In some instances, the present application provides pharmaceutical compositions comprising: (a) a delivery agent comprising a lipid nanoparticle; and (b) a polynucleotide encoding a VHH, polypeptide, bispecific antibody, multispecific antibody, or CAR of the disclosure.
Lipid Nanoparticles
In some instances, polynucleotides of the present disclosure (e.g., mRNA) are included in a lipid nanoparticle (LNP). Lipid nanoparticles according to the present disclosure may comprise: (i) an ionizable lipid (e.g, an ionizable amino lipid; (ii) a sterol
or other structural lipid; (iii) a non-cationic helper lipid or phospholipid; and (iv) a PEG- modified lipid. In some instances, lipid nanoparticles according to the present disclosure further comprise one or more polynucleotides of the present disclosure (e.g., mRNA). The lipid nanoparticles according to the present disclosure can be generated using components, compositions, and methods as are generally known in the art, see, for example PCT/US2016/052352; PCT/US2016/068300; PCT/US2017/037551; PCT/US2015/027400; PCT/US2016/047406; PCT/US2016000129; PCT/US2016/014280; PCT/US2016/014280; PCT/US2017/038426; PCT/US2014/027077; PCT/US2014/055394; PCT/US2016/52117; PCT/US2012/069610; PCT/US2017/027492; PCT/US2016/059575 and PCT/US2016/069491 all of which are incorporated by reference herein in their entirety. In some instances, the lipid nanoparticle comprises: (i) 20 to 60 mol.% ionizable cationic lipid (e.g., ionizable amino lipid), (ii) 25 to 55 mol.% sterol or other structural lipid, (iii) 5 to 25 mol.% non-cationic lipid (e.g., phospholipid), and (iv) 0.5 to 15 mol.% PEG-modified lipid. In some instances, the lipid nanoparticle comprises: (i) 40 to 50 mol.% ionizable cationic lipid (e.g., ionizable amino lipid), (ii) 30 to 45 mol.% sterol or other structural lipid, (iii) 5 to 15 mol.% non-cationic lipid (e.g., phospholipid), and (iv) 1 to 5 mol.% PEG-modified lipid. In some instances, the lipid nanoparticle comprises: (i) 45 to 50 mol.% ionizable cationic lipid (e.g., ionizable amino lipid), (ii) 35 to 45 mol.% sterol or other structural lipid, (iii) 8 to 12 mol.% non-cationic lipid (e.g., phospholipid), and (iv) 1.5 to 3.5 mol.% PEG-modified lipid. In the following sections, “Compounds” numbered with an “I-” prefix (e.g., “Compound I-1,” “Compound I-2,” “Compound I-3,” “Compound I-VI,” etc., indicate specific ionizable lipid compounds. Likewise, compounds numbered with a “P-” prefix (e.g., “Compound P-I,” etc.) indicate a specific PEG-modified lipid compound.
Ionizable Amino Lipids In some instances, the lipid nanoparticle of the present disclosure comprises an ionizable cationic lipid (e.g., an ionizable amino lipid) that is a compound of Formula (I):
or its N-oxide, or a salt or isomer thereof, wherein R’a is R’branched; wherein R’branched is: wherein
denotes a point of attachment;
wherein Raα, Raβ, Raγ, and Raδ are each independently selected from the group consisting of H, C2-12 alkyl, and C2-12 alkenyl; R2 and R3 are each independently selected from the group consisting of C1-14 alkyl and C2-14 alkenyl; R4 is selected from the group consisting of -(CH2)nOH, wherein n is selected from the group consisting of 1, 2, 3, 4, and 5, and
wherein
denotes a point of attachment; wherein R10 is N(R)2; each R is independently selected from the group consisting of C1-6 alkyl, C2-3 alkenyl, and H; and n2 is selected from the group consisting of 1, 2, 3, 4, 5, 6, 7, 8, 9, and 10; each R5 is independently selected from the group consisting of C1-3 alkyl, C2-3 alkenyl, and H; each R6 is independently selected from the group consisting of C1-3 alkyl, C2-3 alkenyl, and H;
M and M’ are each independently selected from the group consisting of -C(O)O- and -OC(O)-; R’ is a C1-12 alkyl or C2-12 alkenyl; l is selected from the group consisting of 1, 2, 3, 4, and 5; and m is selected from the group consisting of 5, 6, 7, 8, 9, 10, 11, 12, and 13. In some instances, in Formula (I), R’a is R’branched; R’branched is
denotes a point of attachment; Raα, Raβ, Raγ, and Raδ are each H; R2 and R3 are each C1-14 alkyl; R4 is -(CH2)nOH; n is 2; each R5 is H; each R6 is H; M and M’ are each - C(O)O-; R’ is a C1-12 alkyl; l is 5; and m is 7. In some instances, in Formula (I), R’a is R’branched; R’branched is ; denotes a point aα aβ aγ aδ
of attachment; R , R , R , and R are each H; R2 and R3 are each C1-14 alkyl; R4 is -(CH2)nOH; n is 2; each R5 is H; each R6 is H; M and M’ are each -C(O)O-; R’ is a C1-12 alkyl; l is 3; and m is 7. In some instances of the compounds of Formula (I), R’a is R’branched; R’branched is aα
denotes a point of attachment; R is C2-12 alkyl; Raβ, Raγ, and Raδ are each H; R2 and R3 are each C1-14 alkyl; R4 is R10 is NH(C1-6 alkyl); n2 is 2; R5 is H; each R6 is H; M and M’ are
each -C(O)O-; R’ is a C1-12 alkyl; l is 5; and m is 7. In some instances of the compounds of Formula (I), R’a is R’branched;
R’branched is denotes a point of attachment aα aβ
; R , R , and Raδ are each H; Raγ is C2-12 alkyl; R2 and R3 are each C1-14 alkyl; R4 is -(CH2)nOH; n is 2; each R5 is H; each R6 is H; M and M’ are each -C(O)O-; R’ is a C1-12 alkyl; l is 5; and m is 7. In some instances, the compound of Formula (I) is selected from:
In some instances, the compound of Formula (I) is:
In some instances, the compound of Formula (I) is:
Phospholipids
The lipid composition of the lipid nanoparticle composition disclosed herein can comprise one or more phospholipids, for example, one or more saturated or (poly)unsaturated phospholipids or a combination thereof. In general, phospholipids comprise a phospholipid moiety and one or more fatty acid moieties.
A phospholipid moiety can be selected, for example, from the non-limiting group consisting of phosphatidyl choline, phosphatidyl ethanolamine, phosphatidyl glycerol, phosphatidyl serine, phosphatidic acid, 2-lysophosphatidyl choline, and a sphingomyelin.
A fatty acid moiety can be selected, for example, from the non-limiting group consisting of lauric acid, myristic acid, myristoleic acid, palmitic acid, palmitoleic acid, stearic acid, oleic acid, linoleic acid, alpha-linolenic acid, erucic acid, phytanoic acid, arachidic acid, arachidonic acid, eicosapentaenoic acid, behenic acid, docosapentaenoic acid, and docosahexaenoic acid.
Particular phospholipids can facilitate fusion to a membrane. For example, a cationic phospholipid can interact with one or more negatively charged phospholipids of
a membrane (e.g., a cellular or intracellular membrane). Fusion of a phospholipid to a membrane can allow one or more elements (e.g., a therapeutic agent) of a lipid- containing composition (e.g., LNPs) to pass through the membrane permitting, e.g., delivery of the one or more elements to a target tissue. Non-natural phospholipid species including natural species with modifications and substitutions including branching, oxidation, cyclization, and alkynes are also contemplated. For example, a phospholipid can be functionalized with or cross-linked to one or more alkynes (e.g., an alkenyl group in which one or more double bonds is replaced with a triple bond). Under appropriate reaction conditions, an alkyne group can undergo a copper-catalyzed cycloaddition upon exposure to an azide. Such reactions can be useful in functionalizing a lipid bilayer of a nanoparticle composition to facilitate membrane permeation or cellular recognition or in conjugating a nanoparticle composition to a useful component such as a targeting or imaging moiety (e.g., a dye). Phospholipids include, but are not limited to, glycerophospholipids such as phosphatidylcholines, phosphatidylethanolamines, phosphatidylserines, phosphatidylinositols, phosphatidy glycerols, and phosphatidic acids. Phospholipids also include phosphosphingolipid, such as sphingomyelin. In some instances, a phospholipid of the present disclosure comprises 1,2- distearoyl-sn-glycero-3-phosphocholine (DSPC), 1,2-dioleoyl-sn-glycero-3- phosphoethanolamine (DOPE), 1,2-dilinoleoyl-sn-glycero-3-phosphocholine (DLPC), 1,2-dimyristoyl-sn-gly cero-phosphocholine (DMPC), 1,2-dioleoyl-sn-glycero-3- phosphocholine (DOPC), l,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC), 1,2- diundecanoyl-sn-glycero-phosphocholine (DUPC), 1-palmitoyl-2-oleoyl-sn-glycero-3- phosphocholine (POPC), 1,2-di-O-octadecenyl-sn-glycero-3-phosphocholine (18:0 Diether PC), 1-oleoyl-2 cholesterylhemisuccinoyl-sn-glycero-3-phosphocholine (OChemsPC), 1-hexadecyl-sn-glycero-3-phosphocholine (C16 Lyso PC), 1,2- dilinolenoyl-sn-glycero-3-phosphocholine,1,2-diarachidonoyl-sn-glycero-3- phosphocholine, 1,2-didocosahexaenoyl-sn-glycero-3-phosphocholine, 1,2-diphytanoyl-
sn-glycero-3-phosphoethanolamine (ME 16.0 PE), 1,2-distearoyl-sn-glycero-3- phosphoethanolamine, 1,2-dilinoleoyl-sn-glycero-3-phosphoethanolamine, 1,2- dilinolenoyl-sn-glycero-3-phosphoethanolamine, 1,2-diarachidonoyl-sn-glycero-3- phosphoethanolamine, 1,2-didocosahexaenoyl-sn-glycero-3-phosphoethanolamine, 1,2- dioleoyl-sn-glycero-3-phospho-rac-(1-glycerol) sodium salt (DOPG), sphingomyelin, and mixtures thereof. In certain instances, a phospholipid useful or potentially useful in the present disclosure is an analog or variant of DSPC. Alternative Lipids In certain instances, a phospholipid useful or potentially useful in the present disclosure comprises a modified phosphocholine moiety, wherein the alkyl chain linking the quaternary amine to the phosphoryl group is not ethylene (e.g., n is not 2). Therefore, in certain instances, a phospholipid useful. In certain instances, an alternative lipid is used in place of a phospholipid of the present disclosure. In certain instances, an alternative lipid of the present disclosure is oleic acid. In certain instances, the alternative lipid is one of the following:
Structural Lipids
The lipid composition of a pharmaceutical composition disclosed herein can comprise one or more structural lipids. As used herein, the term "structural lipid" refers to sterols and also to lipids containing sterol moieties. Incorporation of structural lipids in the lipid nanoparticle may help mitigate aggregation of other lipids in the particle. Structural lipids can be selected from the group
including but not limited to, cholesterol, fecosterol, sitosterol, ergosterol, campesterol, stigmasterol, brassicasterol, tomatidine, tomatine, ursolic acid, alpha-tocopherol, hopanoids, phytosterols, steroids, and mixtures thereof. In some instances, the structural lipid is a sterol. As defined herein, "sterols" are a subgroup of steroids consisting of steroid alcohols. In certain instances, the structural lipid is a steroid. In certain instances, the structural lipid is cholesterol. In certain instances, the structural lipid is an analog of cholesterol. In certain instances, the structural lipid is alpha-tocopherol. In some instances, the structural lipids may be one or more of the structural lipids described in U.S. Application No. 62/520,530. Polyethylene Glycol (PEG)-Lipids The lipid composition of a pharmaceutical composition disclosed herein can comprise one or more a polyethylene glycol (PEG) lipid. As used herein, the term “PEG-lipid” refers to polyethylene glycol (PEG)- modified lipids. Non-limiting examples of PEG-lipids include PEG-modified phosphatidylethanolamine and phosphatidic acid, PEG-ceramide conjugates (e.g., PEG- CerC14 or PEG-CerC20), PEG-modified dialkylamines and PEG-modified 1,2- diacyloxypropan-3-amines. Such lipids are also referred to as PEGylated lipids. For example, a PEG lipid can be PEG-c-DOMG, PEG-DMG, PEG-DLPE, PEG-DMPE, PEG-DPPC, or a PEG-DSPE lipid. In some instances, the PEG-lipid includes, but not limited to 1,2-dimyristoyl-sn- glycerol methoxypolyethylene glycol (PEG-DMG), 1,2-distearoyl-sn-glycero-3- phosphoethanolamine-N-[amino(polyethylene glycol)] (PEG-DSPE), PEG-disteryl glycerol (PEG-DSG), PEG-dipalmetoleyl, PEG-dioleyl, PEG-distearyl, PEG- diacylglycamide (PEG-DAG), PEG-dipalmitoyl phosphatidylethanolamine (PEG-DPPE), or PEG-l,2-dimyristyloxlpropyl-3-amine (PEG-c-DMA).
In some instances, the PEG-lipid is selected from the group consisting of a PEG- modified phosphatidylethanolamine, a PEG-modified phosphatidic acid, a PEG-modified ceramide, a PEG-modified dialkylamine, a PEG-modified diacylglycerol, a PEG- modified dialkylglycerol, and mixtures thereof. In some instances, the lipid moiety of the PEG-lipids includes those having lengths of from about C14 to about C22, preferably from about C14 to about C16. In some instances, a PEG moiety, for example an mPEG-NH2, has a size of about 1000, 2000, 5000, 10,000, 15,000 or 20,000 daltons. In some instances, the PEG-lipid is PEG2k-DMG. In some instances, the lipid nanoparticles described herein can comprise a PEG lipid which is a non-diffusible PEG. Non-limiting examples of non-diffusible PEGs include PEG-DSG and PEG-DSPE. PEG-lipids are known in the art, such as those described in U.S. Patent No. 8,158,601 and International Publ. No. WO 2015/130584 A2, which are incorporated herein by reference in their entirety. In general, some of the other lipid components (e.g., PEG lipids) of various Formulae, described herein may be synthesized as described International Patent Application No. PCT/US2016/000129, filed December 10, 2016, entitled “Compositions and Methods for Delivery of Therapeutic Agents,” which is incorporated by reference in its entirety. The lipid component of a lipid nanoparticle composition may include one or more molecules comprising polyethylene glycol, such as PEG or PEG-modified lipids. Such species may be alternately referred to as PEGylated lipids. A PEG lipid is a lipid modified with polyethylene glycol. A PEG lipid may be selected from the non-limiting group including PEG-modified phosphatidylethanolamines, PEG-modified phosphatidic acids, PEG-modified ceramides, PEG-modified dialkylamines, PEG-modified diacylglycerols, PEG-modified dialkylglycerols, and mixtures thereof. For example, a
PEG lipid may be PEG-c-DOMG, PEG-DMG, PEG-DLPE, PEG-DMPE, PEG-DPPC, or a PEG-DSPE lipid. In some instances, PEG lipids useful in the present disclosure can be PEGylated lipids described in International Publication No. WO2012099755, the contents of which is herein incorporated by reference in its entirety. Any of these exemplary PEG lipids described herein may be modified to comprise a hydroxyl group on the PEG chain. In certain instances, the PEG lipid is a PEG-OH lipid. As generally defined herein, a “PEG- OH lipid” (also referred to herein as “hydroxy-PEGylated lipid”) is a PEGylated lipid having one or more hydroxyl (–OH) groups on the lipid. In certain instances, the PEG- OH lipid includes one or more hydroxyl groups on the PEG chain. In certain instances, a PEG-OH or hydroxy-PEGylated lipid comprises an –OH group at the terminus of the PEG chain. Each possibility represents a separate embodiment of the present disclosure. In some aspects, the lipid composition of the pharmaceutical compositions disclosed herein does not comprise a PEG-lipid. In some instances, the PEG-lipids may be one or more of the PEG lipids described in U.S. Application No. 62/520,530. In some instances, a PEG lipid of the present disclosure comprises a PEG- modified phosphatidylethanolamine, a PEG-modified phosphatidic acid, a PEG-modified ceramide, a PEG-modified dialkylamine, a PEG-modified diacylglycerol, a PEG- modified dialkylglycerol, and mixtures thereof. In some instances, the PEG-modified lipid is PEG-DMG, PEG-c-DOMG (also referred to as PEG-DOMG), PEG-DSG and/or PEG-DPG. In some instances, a PEG lipid of this disclosure comprises 134-hydroxy- 3,6,9,12,15,18,21,24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96,99,102,105,108,111,114,117,120,123,126,129,132- tetratetracontaoxatetratriacontahectyl stearate. In one instance the PEG lipid has the following structure:
wherein R = 45. In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of Formula I, a phospholipid comprising DSPC, a structural lipid, and a PEG lipid comprising PEG-DMG or 134-hydroxy- 3,6,9,12,15,18,21,24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96,99,102,105,108,111,114,117,120,123,126,129,132- tetratetracontaoxatetratriacontahectyl stearate. In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of Formula I, a phospholipid comprising DSPC, a structural lipid, and a PEG lipid (e.g., 134-hydroxy- 3,6,9,12,15,18,21,24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96,99,102,105,108,111,114,117,120,123,126,129,132- tetratetracontaoxatetratriacontahectyl stearate). In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of
. In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of
, and an alternative lipid comprising oleic acid. In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of
, an alternative lipid comprising oleic acid, a structural lipid comprising cholesterol, and a PEG lipid. In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of
a phospholipid comprising DOPE, a structural lipid comprising cholesterol, and a PEG lipid. In some instances, a LNP of the present disclosure comprises an ionizable cationic lipid of
, a phospholipid comprising DOPE, a structural lipid comprising cholesterol, and a PEG lipid. In some instances, a LNP of the present disclosure comprises an N:P ratio of from about 2:1 to about 30:1.In some instances, a LNP of the present disclosure comprises an N:P ratio of about 6:1. In some instances, a LNP of the present disclosure comprises an N:P ratio of about 3:1. In some instances, a LNP of the present disclosure comprises a wt/wt ratio of the ionizable cationic lipid component to the RNA of from about 10:1 to about 100:1. In some instances, a LNP of the present disclosure comprises a wt/wt ratio of the ionizable cationic lipid component to the RNA of about 20:1. In some instances, a LNP of the present disclosure comprises a wt/wt ratio of the ionizable cationic lipid component to the RNA of about 10:1. In some instances, a LNP of the present disclosure has a mean diameter from about 50nm to about 150nm. In some instances, a LNP of the present disclosure has a mean diameter from about 70nm to about 120nm. Other Lipid Composition Components The lipid composition of a pharmaceutical composition disclosed herein can include one or more components in addition to those described above. For example, the lipid composition can include one or more permeability enhancer molecules, carbohydrates, polymers, surface altering agents (e.g., surfactants), or other components. For example, a permeability enhancer molecule can be a molecule described by U.S. Patent Application Publication No.2005/0222064. Carbohydrates can include simple sugars (e.g., glucose) and polysaccharides (e.g., glycogen and derivatives and analogs thereof).
A polymer can be included in and/or used to encapsulate or partially encapsulate a pharmaceutical composition disclosed herein (e.g., a pharmaceutical composition in lipid nanoparticle form). A polymer can be biodegradable and/or biocompatible. A polymer can be selected from, but is not limited to, polyamines, polyethers, polyamides, polyesters, polycarbamates, polyureas, polycarbonates, polystyrenes, polyimides, polysulfones, polyurethanes, polyacetylenes, polyethylenes, polyethyleneimines, polyisocyanates, polyacrylates, polymethacrylates, polyacrylonitriles, and polyarylates. The ratio between the lipid composition and the polynucleotide range can be from about 10:1 to about 60:1 (wt/wt). In some instances, the ratio between the lipid composition and the polynucleotide can be about 10:1, 11:1, 12:1, 13:1, 14:1, 15:1, 16:1, 17:1, 18:1, 19:1, 20:1, 21:1, 22:1, 23:1, 24:1, 25:1, 26:1, 27:1, 28:1, 29:1, 30:1, 31:1, 32:1, 33:1, 34:1, 35:1, 36:1, 37:1, 38:1, 39:1, 40:1, 41:1, 42:1, 43:1, 44:1, 45:1, 46:1, 47:1, 48:1, 49:1, 50:1, 51:1, 52:1, 53:1, 54:1, 55:1, 56:1, 57:1, 58:1, 59:1 or 60:1 (wt/wt). In some instances, the wt/wt ratio of the lipid composition to the polynucleotide encoding a therapeutic agent is about 20:1 or about 15:1. In some instances, the pharmaceutical composition disclosed herein can contain more than one polypeptides. For example, a pharmaceutical composition disclosed herein can contain two or more polynucleotides (e.g., RNA, e.g., mRNA). In some instances, the lipid nanoparticles described herein can comprise polynucleotides (e.g., mRNA) in a lipid:polynucleotide weight ratio of 5:1, 10:1, 15:1, 20:1, 25:1, 30:1, 35:1, 40:1, 45:1, 50:1, 55:1, 60:1 or 70:1, or a range or any of these ratios such as, but not limited to, 5:1 to about 10:1, from about 5:1 to about 15:1, from about 5:1 to about 20:1, from about 5:1 to about 25:1, from about 5:1 to about 30:1, from about 5:1 to about 35:1, from about 5:1 to about 40:1, from about 5:1 to about 45:1, from about 5:1 to about 50:1, from about 5:1 to about 55:1, from about 5:1 to about 60:1, from about 5:1 to about 70:1, from about 10:1 to about 15:1, from about 10:1 to about 20:1,
from about 10:1 to about 25:1, from about 10:1 to about 30:1, from about 10:1 to about 35:1, from about 10:1 to about 40:1, from about 10:1 to about 45:1, from about 10:1 to about 50:1, from about 10:1 to about 55:1, from about 10:1 to about 60:1, from about 10:1 to about 70:1, from about 15:1 to about 20:1, from about 15:1 to about 25:1,from about 15:1 to about 30:1, from about 15:1 to about 35:1, from about 15:1 to about 40:1, from about 15:1 to about 45:1, from about 15:1 to about 50:1, from about 15:1 to about 55:1, from about 15:1 to about 60:1 or from about 15:1 to about 70:1. In some instances, the lipid nanoparticles described herein can comprise the polynucleotide in a concentration from approximately 0.1 mg/ml to 2 mg/ml such as, but not limited to, 0.1 mg/ml, 0.2 mg/ml, 0.3 mg/ml, 0.4 mg/ml, 0.5 mg/ml, 0.6 mg/ml, 0.7 mg/ml, 0.8 mg/ml, 0.9 mg/ml, 1.0 mg/ml, 1.1 mg/ml, 1.2 mg/ml, 1.3 mg/ml, 1.4 mg/ml, 1.5 mg/ml, 1.6 mg/ml, 1.7 mg/ml, 1.8 mg/ml, 1.9 mg/ml, 2.0 mg/ml or greater than 2.0 mg/ml. Nanoparticle Compositions In some instances, the pharmaceutical compositions disclosed herein are Formulated as lipid nanoparticles (LNP). Accordingly, the present disclosure also provides nanoparticle compositions comprising (i) a lipid composition comprising a delivery agent such as compound as described herein, and (ii) a polynucleotide encoding a polypeptide. In such nanoparticle composition, the lipid composition disclosed herein can encapsulate the polynucleotide encoding a polypeptide. Nanoparticle compositions are typically sized on the order of micrometers or smaller and can include a lipid bilayer. Nanoparticle compositions encompass lipid nanoparticles (LNPs), liposomes (e.g., lipid vesicles), and lipoplexes. For example, a nanoparticle composition can be a liposome having a lipid bilayer with a diameter of 500 nm or less. Nanoparticle compositions include, for example, lipid nanoparticles (LNPs), liposomes, and lipoplexes. In some instances, nanoparticle compositions are vesicles
including one or more lipid bilayers. In certain instances, a nanoparticle composition includes two or more concentric bilayers separated by aqueous compartments. Lipid bilayers can be functionalized and/or crosslinked to one another. Lipid bilayers can include one or more ligands, proteins, or channels. In some instances, a lipid nanoparticle comprises an ionizable amino lipid, a structural lipid, a phospholipid, and mRNA. In some instances, the LNP comprises an ionizable amino lipid, a PEG-modified lipid, a sterol and a structural lipid. In some instances, the LNP has a molar ratio of about 40-50% ionizable amino lipid; about 5-15% structural lipid; about 30-45% sterol; and about 1-5% PEG-modified lipid. In some instances, the lipid nanoparticle comprises 47-49 mol.% ionizable cationic lipid (e.g. ionizable amino lipid, e.g., Compound I-1, Compound I-2, or Compound I-3), 10-12 mol.% non-cationic lipid (e.g., phospholipid, e.g., DSPC), 38-40 mol.% sterol (e.g., cholesterol) or other structural lipid, and 1-3 mol.% PEG-modified lipid (e.g., PEG-DMG or Compound P-I). For instance, in some instances, the lipid nanoparticle (“LNP-1”) may comprise the following components at the following molar ratios: (i) 45-50 mol.% Compound I-1 (ii) 35-45 mol.% sterol (e.g., cholesterol); (iii) 8-12 mol.% phospholipid (e.g., DSPC or DOPE); and (iv) 1.5-3.5 mol.% PEG-lipid. For instance, in some instances, the lipid nanoparticle (“LNP-1A”) may comprise the following components at the following molar ratios: (i) 45-50 mol.% Compound I-1 (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid.
For instance, in some instances, the lipid nanoparticle (“LNP-1B”) may comprise the following components at the following molar ratios: (i) 45-50 mol.% Compound I-1 (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid. In some instances, the lipid nanoparticle (“LNP-2”) may comprise the following: (i) 45-50 mol.% Compound I-2; (ii) 35-45 mol.% sterol (e.g., Cholesterol); (iii) 8-12 mol.% phospholipid (e.g., DSPC or DOPE); and (iv) 1.5-3.5 mol.% PEG-lipid. In some instances, the lipid nanoparticle (“LNP-2A”) may comprise the following: (i) 45-50 mol.% Compound I-2; (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid. For instance, in some instances, the lipid nanoparticle (“LNP-2B”) may comprise the following components at the following molar ratios: (i) 45-50 mol.% Compound I-2; (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid. In some instances, the lipid nanoparticle (“LNP-3”) may comprise the following: (i) 45-50 mol.% Compound I-3; (ii) 35-45 mol.% sterol (e.g., Cholesterol); (iii) 8-12 mol.% phospholipid (e.g., DSPC or DOPE); and
(iv) 1.5-3.5 mol.% PEG-lipid. In some instances, the lipid nanoparticle (“LNP-3A”) may comprise the following: (i) 45-50 mol.% Compound I-3; (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid. In some instances, the lipid nanoparticle (“LNP-3B”) may comprise the following: (i) 45-50 mol.% Compound I-3; (ii) 35-45 mol.% Cholesterol; (iii) 8-12 mol.% DSPC; and (iv) 1.5-3.5 mol.% PEG-lipid. In some instances, the LNP has a polydispersity value of less than 0.4. In some instances, the LNP has a net neutral charge at a neutral pH. In some instances, the LNP has a mean diameter of 50-150 nm. In some instances, the LNP has a mean diameter of 80-100 nm. As generally defined herein, the term “lipid” refers to a small molecule that has hydrophobic or amphiphilic properties. Lipids may be naturally occurring or synthetic. Examples of classes of lipids include, but are not limited to, fats, waxes, sterol-containing metabolites, vitamins, fatty acids, glycerolipids, glycerophospholipids, sphingolipids, saccharolipids, and polyketides, and prenol lipids. In some instances, the amphiphilic properties of some lipids lead them to form liposomes, vesicles, or membranes in aqueous media. In some instances, a lipid nanoparticle (LNP) may comprise an ionizable amino lipid. As used herein, the term “ionizable amino lipid” has its ordinary meaning in the art and may refer to a lipid comprising one or more charged moieties. In some instances, an
ionizable amino lipid may be positively charged or negatively charged. An ionizable amino lipid may be positively charged, in which case it can be referred to as “cationic lipid”. In certain instances, an ionizable amino lipid molecule may comprise an amine group, and can be referred to as an ionizable amino lipid. As used herein, a “charged moiety” is a chemical moiety that carries a formal electronic charge, e.g., monovalent (+1, or -1), divalent (+2, or -2), trivalent (+3, or -3), etc. The charged moiety may be anionic (i.e., negatively charged) or cationic (i.e., positively charged). Examples of positively-charged moieties include amine groups (e.g., primary, secondary, and/or tertiary amines), ammonium groups, pyridinium group, guanidine groups, and imidizolium groups. In a particular embodiment, the charged moieties comprise amine groups. Examples of negatively- charged groups or precursors thereof, include carboxylate groups, sulfonate groups, sulfate groups, phosphonate groups, phosphate groups, hydroxyl groups, and the like. The charge of the charged moiety may vary, in some cases, with the environmental conditions, for example, changes in pH may alter the charge of the moiety, and/or cause the moiety to become charged or uncharged. In general, the charge density of the molecule may be selected as desired. It should be understood that the terms “charged” or “charged moiety” does not refer to a “partial negative charge" or “partial positive charge" on a molecule. The terms “partial negative charge" and “partial positive charge" are given their ordinary meaning in the art. A “partial negative charge" may result when a functional group comprises a bond that becomes polarized such that electron density is pulled toward one atom of the bond, creating a partial negative charge on the atom. Those of ordinary skill in the art will, in general, recognize bonds that can become polarized in this way. The ionizable amino lipid is sometimes referred to in the art as an “ionizable cationic lipid”. In some instances, the ionizable amino lipid may have a positively charged hydrophilic head and a hydrophobic tail that are connected via a linker structure. In addition to these, an ionizable amino lipid may also be a lipid including a cyclic amine group.
In some instances, the ionizable amino lipid may be selected from, but not limited to, an ionizable amino lipid described in International Publication Nos. WO2013086354 and WO2013116126; the contents of each of which are herein incorporated by reference in their entirety. In yet another embodiment, the ionizable amino lipid may be selected from, but not limited to, Formula CLI-CLXXXXII of US Patent No. 7,404,969; each of which is herein incorporated by reference in their entirety. In some instances, the lipid may be a cleavable lipid such as those described in International Publication No. WO2012170889, herein incorporated by reference in its entirety. In some instances, the lipid may be synthesized by methods known in the art and/or as described in International Publication Nos. WO2013086354; the contents of each of which are herein incorporated by reference in their entirety. Nanoparticle compositions can be characterized by a variety of methods. For example, microscopy (e.g., transmission electron microscopy or scanning electron microscopy) can be used to examine the morphology and size distribution of a nanoparticle composition. Dynamic light scattering or potentiometry (e.g., potentiometric titrations) can be used to measure zeta potentials. Dynamic light scattering can also be utilized to determine particle sizes. Instruments such as the Zetasizer Nano ZS (Malvern Instruments Ltd, Malvern, Worcestershire, UK) can also be used to measure multiple characteristics of a nanoparticle composition, such as particle size, polydispersity index, and zeta potential. The size of the nanoparticles can help counter biological reactions such as, but not limited to, inflammation, or can increase the biological effect of the polynucleotide. As used herein, “size” or “mean size” in the context of nanoparticle compositions refers to the mean diameter of a nanoparticle composition.
In some instances, the polynucleotide encoding a polypeptide(s) described herein are formulated in lipid nanoparticles having a diameter from about 10 to about 100 nm such as, but not limited to, about 10 to about 20 nm, about 10 to about 30 nm, about 10 to about 40 nm, about 10 to about 50 nm, about 10 to about 60 nm, about 10 to about 70 nm, about 10 to about 80 nm, about 10 to about 90 nm, about 20 to about 30 nm, about 20 to about 40 nm, about 20 to about 50 nm, about 20 to about 60 nm, about 20 to about 70 nm, about 20 to about 80 nm, about 20 to about 90 nm, about 20 to about 100 nm, about 30 to about 40 nm, about 30 to about 50 nm, about 30 to about 60 nm, about 30 to about 70 nm, about 30 to about 80 nm, about 30 to about 90 nm, about 30 to about 100 nm, about 40 to about 50 nm, about 40 to about 60 nm, about 40 to about 70 nm, about 40 to about 80 nm, about 40 to about 90 nm, about 40 to about 100 nm, about 50 to about 60 nm, about 50 to about 70 nm, about 50 to about 80 nm, about 50 to about 90 nm, about 50 to about 100 nm, about 60 to about 70 nm, about 60 to about 80 nm, about 60 to about 90 nm, about 60 to about 100 nm, about 70 to about 80 nm, about 70 to about 90 nm, about 70 to about 100 nm, about 80 to about 90 nm, about 80 to about 100 nm and/or about 90 to about 100 nm. In some instances, the nanoparticles have a diameter from about 10 to 500 nm. In some instances, the nanoparticle has a diameter greater than 100 nm, greater than 150 nm, greater than 200 nm, greater than 250 nm, greater than 300 nm, greater than 350 nm, greater than 400 nm, greater than 450 nm, greater than 500 nm, greater than 550 nm, greater than 600 nm, greater than 650 nm, greater than 700 nm, greater than 750 nm, greater than 800 nm, greater than 850 nm, greater than 900 nm, greater than 950 nm or greater than 1000 nm. In some instances, the largest dimension of a nanoparticle composition is 1 μm or shorter (e.g., 1 μm, 900 nm, 800 nm, 700 nm, 600 nm, 500 nm, 400 nm, 300 nm, 200 nm, 175 nm, 150 nm, 125 nm, 100 nm, 75 nm, 50 nm, or shorter). A nanoparticle composition can be relatively homogenous. A polydispersity index can be used to indicate the homogeneity of a nanoparticle composition, e.g., the particle
size distribution of the nanoparticle composition. A small (e.g., less than 0.3) polydispersity index generally indicates a narrow particle size distribution. A nanoparticle composition can have a polydispersity index from about 0 to about 0.25, such as 0.01, 0.02, 0.03, 0.04, 0.05, 0.06, 0.07, 0.08, 0.09, 0.10, 0.11, 0.12, 0.13, 0.14, 0.15, 0.16, 0.17, 0.18, 0.19, 0.20, 0.21, 0.22, 0.23, 0.24, or 0.25. In some instances, the polydispersity index of a nanoparticle composition disclosed herein can be from about 0.10 to about 0.20. The amount of a polynucleotide present in a pharmaceutical composition disclosed herein can depend on multiple factors such as the size of the polynucleotide, desired target and/or application, or other properties of the nanoparticle composition as well as on the properties of the polynucleotide. For example, the amount of an mRNA useful in a nanoparticle composition can depend on the size (expressed as length, or molecular mass), sequence, and other characteristics of the mRNA. The relative amounts of a polynucleotide in a nanoparticle composition can also vary. The relative amounts of the lipid composition and the polynucleotide present in a lipid nanoparticle composition of the present disclosure can be optimized according to considerations of efficacy and tolerability. For compositions including an mRNA as a polynucleotide, the N:P ratio can serve as a useful metric. As the N:P ratio of a nanoparticle composition controls both expression and tolerability, nanoparticle compositions with low N:P ratios and strong expression are desirable. N:P ratios vary according to the ratio of lipids to RNA in a nanoparticle composition. In general, a lower N:P ratio is preferred. The one or more RNA, lipids, and amounts thereof can be selected to provide an N:P ratio from about 2:1 to about 30:1, such as 2:1, 3:1, 4:1, 5:1, 6:1, 7:1, 8:1, 9:1, 10:1, 12:1, 14:1, 16:1, 18:1, 20:1, 22:1, 24:1, 26:1, 28:1, or 30:1. In certain instances, the N:P ratio can be from about 2:1 to about 8:1.
In other instances, the N:P ratio is from about 5: 1 to about 8: 1. In certain instances, the N:P ratio is between 5: 1 and 6: 1. In one specific aspect, the N:P ratio is about is about 5.67:1.
In some instances, the polynucleotides described herein can be formulated for controlled release and/or targeted delivery. As used herein, "controlled release" refers to a pharmaceutical composition or compound release profile that conforms to a particular pattern of release to effect a therapeutic outcome. In some instances, the polynucleotides can be encapsulated into a delivery agent described herein and/or known in the art for controlled release and/or targeted delivery. As used herein, the term "encapsulate" means to enclose, surround or encase. As it relates to the formulation of the compounds of the present disclosure, encapsulation can be substantial, complete or partial. The term "substantially encapsulated" means that at least greater than 50, 60, 70, 80, 85, 90, 95, 96, 97, 98, 99, or greater than 99% of the pharmaceutical composition or compound of the present disclosure can be enclosed, surrounded or encased within the delivery agent. "Partial encapsulation" or “partially encapsulate” means that less than 10, 10, 20, 30, 40 50 or less of the pharmaceutical composition or compound of the present disclosure can be enclosed, surrounded or encased within the delivery agent.
In some instances, the therapeutic nanoparticle polynucleotide can be formulated for sustained release. As used herein, "sustained release" refers to a pharmaceutical composition or compound that conforms to a release rate over a specific period of time. The period of time can include, but is not limited to, hours, days, weeks, months and years. As a non-limiting example, the sustained release nanoparticle of the polynucleotides described herein can be formulated as disclosed in Inti. Pub. No. W02010075072 and U.S. Pub. Nos. US20100216804, US20110217377, US20120201859 and US20130150295, each of which is herein incorporated by reference in their entirety.
In some instances, the therapeutic nanoparticle polynucleotide can be formulated to be target specific, such as those described in Inti. Pub. Nos. WO2008121949,
WO2010005726, WO2010005725, WO2011084521 and WO2011084518; and U.S. Pub. Nos. US20100069426, US20120004293 and US20100104655, each of which is herein incorporated by reference in its entirety. The LNPs can be prepared using microfluidic mixers or micromixers. Exemplary microfluidic mixers can include, but are not limited to, a slit interdigital micromixer including, but not limited to those manufactured by Microinnova (Allerheiligen bei Wildon, Austria) and/or a staggered herringbone micromixer (SHM) (see, Zhigaltsev et al., Langmuir.28:3633-40 (2012); Belliveau et al., Molecular Therapy-Nucleic Acids. 1:e37 (2012); Chen et al., J. Am. Chem. Soc. 134(16):6948-51 (2012); each of which is herein incorporated by reference in its entirety). Exemplary micromixers include Slit Interdigital Microstructured Mixer (SIMM-V2) or a Standard Slit Interdigital Micro Mixer (SSIMM) or Caterpillar (CPMM) or Impinging-jet (IJMM,) from the Institut für Mikrotechnik Mainz GmbH, Mainz Germany. In some instances, methods of making LNP using SHM further comprise mixing at least two input streams wherein mixing occurs by microstructure-induced chaotic advection (MICA). According to this method, fluid streams flow through channels present in a herringbone pattern causing rotational flow and folding the fluids around each other. This method can also comprise a surface for fluid mixing wherein the surface changes orientations during fluid cycling. Methods of generating LNPs using SHM include those disclosed in U.S. Pub. Nos. US20040262223 and US20120276209, each of which is incorporated herein by reference in their entirety. In some instances, the polynucleotides described herein can be Formulated in lipid nanoparticles using microfluidic technology (see, Whitesides, George M., Nature 442: 368-373 (2006); and Abraham et al., Science 295: 647-651 (2002); each of which is herein incorporated by reference in its entirety). In some instances, the polynucleotides can be Formulated in lipid nanoparticles using a micromixer chip such as, but not limited to, those from Harvard Apparatus (Holliston, MA) or Dolomite Microfluidics (Royston,
UK). A micromixer chip can be used for rapid mixing of two or more fluid streams with a split and recombine mechanism. In some embodiment, the nanoparticles described herein are stealth nanoparticles or target-specific stealth nanoparticles such as, but not limited to, those described in U.S. Pub. No. US20130172406, herein incorporated by reference in its entirety. The stealth or target-specific stealth nanoparticles can comprise a polymeric matrix, which can comprise two or more polymers such as, but not limited to, polyethylenes, polycarbonates, polyanhydrides, polyhydroxyacids, polypropylfumerates, polycaprolactones, polyamides, polyacetals, polyethers, polyesters, poly(orthoesters), polycyanoacrylates, polyvinyl alcohols, polyurethanes, polyphosphazenes, polyacrylates, polymethacrylates, polycyanoacrylates, polyureas, polystyrenes, polyamines, polyesters, polyanhydrides, polyethers, polyurethanes, polymethacrylates, polyacrylates, polycyanoacrylates, or combinations thereof. Methods of Treatment The VHH containing polypeptides described herein can be used in the treatment of any disease associated the dysregulation of the c-MET/HGF pathway. In some cases, the disease shows an increased activation of the c-MET signaling relative to age-matched human subjects without cancer. In some cases, these VHH containing polypeptides are used in the treatment of cancers with one or more c-MET mutations. In some cases, these VHH containing polypeptides are used in the treatment of cancers with one or more a c- MET amplifications. In some cases, these VHH containing polypeptides are used in the treatment of cancers with c-MET overexpression relative to age-matched human subjects without cancer. The methods involve administering to the human subject in need thereof a therapeutically effective amount of a VHH containing polypeptide herein. In some cases, the VHH containing polypeptide is a monovalent polypeptide. In some cases, the VHH containing polypeptide is a bivalent polypeptide. In some cases, the VHH containing polypeptide is a bivalent polypeptide (e.g., a c-MET binding VHH-linked to IgG4PAA).
In some cases, the VHH containing polypeptide is a bispecific antibody (e.g., a bispecific T cell engager (BiTE), a bispecific NK cell engager (BiKE); a bispecific antibody that binds both c-MET and another tumor associated/related antigen). In some cases, the VHH containing polypeptide is a multispecific antibody construct. In some cases, the VHH containing polypeptide is a conjugate (e.g., an antibody drug conjugate). In some cases, the human is administered a T cell comprising a CAR-T cell wherein the T cell expresses a CAR comprising a VHH that specifically binds c-MET. In some cases, the human is administered an NK cell comprising a CAR-NK cell wherein the NK cell is engineered to express a CAR comprising a VHH that specifically binds c-MET. The polypeptides of this disclosure may be administered subcutaneously or intravenously. In some instances the antibodies are administered to a subject as a bolus IV injection or bolus. This form of delivery can produce high levels of expressed antibody. Kits Also included herein are kits that contain a VHH, a polypeptide, a bispecific antibody, a VHH-conjugate, or a CAR of this disclosure, and/or nucleic acids (e.g., mRNA) encoding the same. In some embodiments, the kits provided herein contain one or more cells engineered to express and secrete a polypeptide(s) comprising an anti-c- MET VHH of the disclosure, such as a cell containing a nucleic acid molecule(s) of the disclosure. A kit described herein may include reagents that can be used to produce a pharmaceutical composition of the invention. Optionally, kits described herein may include reagents that can induce the expression of a polypeptide(s) comprising an anti-c- MET VHH of the disclosure within cells (e.g., bacterial, yeast, insect, or mammalian cells). Other kits described herein may include tools for engineering a prokaryotic or eukaryotic cell (e.g., a CHO cell or a BL21(DE3) E. Coli cell or an immune cell) so as to
express and secrete a polypeptide(s) comprising an anti-c-MET VHH of the disclosure. For example, a kit may contain CHO cells stored in an appropriate media and optionally frozen according to methods known in the art. The kit may also contain a nucleic acid encoding the desired VHH antibody as well as reagents for expressing the VHH antibody or binding protein in the cell. A kit described herein may also provide a polypeptide(s) comprising an anti-c- MET VHH of the disclosure, a nucleic acid encoding the same (e.g., mRNA), or a LNP that encapsulates a nucleic acid (e.g., mRNA) that encodes an anti-c-MET VHH described herein, in combination with a package insert describing how the VHH antibody, or nucleic acid may be administered to a subject, for example, for the treatment of a disease, disorder and/or condition (e.g., cancer). EXAMPLES Example 1: Materials & Methods 1. Immunization of Llamas: Llamas were immunized with mRNA encoding full length c-MET, the sequence of which is provided below: ATGAAGGCCCCTGCTGTGCTGGCCCCTGGCATACTGGTGTTGCTGTTCACACT GGTGCAACGGAGTAACGGCGAGTGCAAGGAGGCCCTGGCCAAGAGCGAGAT GAACGTGAATATGAAGTATCAACTGCCCAACTTCACTGCCGAGACCCCTATC CAGAACGTGATCCTTCACGAACACCACATCTTCCTGGGAGCCACCAACTACA TCTACGTCCTAAACGAGGAGGACCTGCAGAAGGTAGCCGAGTACAAGACAG GCCCCGTGCTTGAGCACCCCGACTGCTTCCCCTGCCAGGACTGCAGCAGCAA GGCCAACCTTTCGGGCGGCGTGTGGAAGGACAACATCAACATGGCACTGGTG GTGGACACCTACTACGACGACCAGCTCATCAGCTGCGGCAGCGTGAACAGAG GCACTTGCCAGCGGCACGTGTTTCCACACAACCACACCGCCGACATCCAGAG CGAGGTGCACTGCATATTCTCCCCTCAGATCGAAGAGCCAAGCCAGTGTCCA GACTGCGTGGTGTCAGCCCTGGGCGCTAAGGTCCTCTCTAGTGTGAAGGACC GGTTCATCAACTTCTTCGTGGGAAATACCATCAACTCCAGCTACTTCCCCGAC CACCCACTGCATTCTATCAGCGTGCGGCGGCTGAAGGAGACAAAGGACGGCT TCATGTTCCTGACTGACCAGAGCTACATCGACGTGCTGCCCGAGTTCCGGGA CAGCTACCCCATTAAATACGTGCACGCCTTTGAGAGCAACAACTTCATTTACT
TCCTGACCGTGCAACGAGAGACCCTGGACGCCCAGACATTCCATACCCGGAT CATCCGGTTCTGCAGCATCAACAGCGGTCTGCACAGCTACATGGAGATGCCC TTAGAGTGTATCCTCACCGAGAAGAGGAAGAAGCGGAGCACCAAGAAGGAA GTGTTTAACATCTTGCAGGCCGCCTACGTTAGCAAGCCTGGCGCCCAATTGGC CCGGCAGATCGGCGCCAGCCTGAACGACGACATCCTGTTTGGCGTGTTCGCC CAGAGCAAGCCCGACAGCGCCGAGCCCATGGACCGGAGCGCCATGTGCGCC TTCCCCATCAAGTACGTGAACGACTTCTTCAACAAGATCGTGAACAAGAACA ACGTGCGGTGCCTGCAGCACTTCTACGGCCCCAACCACGAGCACTGCTTCAA CCGGACCCTGCTGCGGAATAGCAGCGGCTGTGAAGCCCGGAGGGACGAGTA CCGGACCGAATTCACCACCGCCTTGCAGCGGGTGGACCTGTTCATGGGCCAG TTCAGCGAGGTACTGCTGACCTCCATTTCTACCTTCATCAAGGGCGACCTCAC CATCGCCAACCTCGGCACCAGCGAGGGCCGGTTTATGCAGGTGGTGGTGAGC AGAAGCGGCCCCTCAACGCCTCACGTGAACTTCCTGCTGGACAGCCACCCCG TGAGCCCCGAGGTTATCGTGGAGCACACCCTGAATCAGAACGGGTACACCCT GGTCATTACTGGTAAGAAGATTACCAAGATCCCTCTGAACGGCCTGGGCTGT CGGCACTTTCAGAGCTGCTCCCAGTGTCTGAGCGCACCTCCTTTCGTGCAGTG CGGGTGGTGCCACGACAAGTGCGTACGGAGCGAGGAGTGCCTGAGCGGCAC CTGGACCCAGCAGATTTGCCTGCCCGCCATCTACAAGGTGTTTCCCAACTCTG CCCCTCTGGAAGGCGGAACCCGGCTGACCATCTGCGGCTGGGACTTTGGCTT CCGGCGGAATAACAAGTTCGACCTGAAGAAGACCCGGGTGCTGCTCGGCAAC GAAAGCTGCACACTGACTCTGAGCGAGAGCACCATGAATACCCTGAAGTGTA CCGTGGGCCCCGCTATGAACAAGCATTTTAACATGAGCATTATCATCAGCAA CGGCCACGGAACAACCCAGTACAGCACCTTTAGCTACGTTGACCCCGTCATC ACCAGCATCAGCCCCAAGTACGGTCCCATGGCTGGCGGTACCCTGTTGACCC TGACAGGCAACTATCTGAACAGCGGCAACAGCCGGCACATCAGCATCGGCG GCAAGACCTGCACTCTCAAGAGCGTGTCAAACAGCATCCTGGAGTGCTACAC CCCTGCCCAGACCATCAGCACCGAGTTCGCCGTCAAACTGAAGATCGACCTA GCCAACCGGGAGACCTCTATCTTCAGCTACCGGGAGGACCCCATCGTGTACG AGATCCACCCCACCAAGAGCTTCATCAGCGGCGGCAGCACCATCACCGGCGT GGGCAAGAACCTGAACTCCGTGAGCGTGCCCCGGATGGTGATCAACGTTCAC GAGGCTGGGCGCAACTTTACCGTGGCCTGCCAGCACCGGAGCAACAGCGAG ATCATCTGCTGTACCACCCCTAGCCTGCAACAGCTGAACCTGCAGCTGCCCCT GAAGACCAAGGCCTTCTTCATGCTCGACGGCATCCTGAGCAAGTACTTCGAC CTTATCTACGTACATAACCCCGTGTTCAAGCCCTTCGAGAAGCCCGTGATGAT CAGCATGGGTAACGAGAACGTGTTAGAGATCAAAGGCAACGACATCGACCC CGAGGCCGTTAAGGGTGAGGTGCTGAAGGTGGGCAACAAGAGCTGCGAGAA CATCCATTTGCACAGCGAGGCGGTGCTGTGCACCGTGCCCAACGACCTGCTG AAACTGAATAGCGAGCTGAACATCGAGTGGAAGCAGGCCATCAGCTCTACCG TGCTGGGCAAGGTGATCGTCCAGCCCGACCAGAACTTCACCGGCCTGATCGC CGGCGTCGTGTCCATCTCTACAGCCCTGCTCCTGCTTCTGGGGTTCTTCCTGTG GCTGAAGAAACGGAAGCAGATCAAGGACCTGGGCAGCGAGCTCGTGCGGTA
CGACGCCCGGGTCCACACCCCTCATCTGGACCGGCTGGTGTCTGCCCGGTCTG TGTCACCCACCACCGAGATGGTGTCTAACGAGAGCGTGGACTACCGGGCCAC CTTTCCCGAGGACCAGTTCCCCAACAGCAGCCAGAACGGCAGTTGCCGGCAG GTGCAGTACCCTCTGACCGACATGAGCCCCATCCTGACCAGCGGCGACAGCG ACATTAGCAGCCCTCTGCTCCAGAACACCGTGCACATCGATTTGAGCGCCCT GAACCCCGAGCTGGTGCAGGCAGTTCAACACGTGGTGATCGGGCCCAGCAGC CTCATCGTGCACTTTAACGAAGTGATCGGCCGGGGCCACTTCGGCTGCGTGTA CCACGGCACCCTCCTGGACAACGACGGCAAGAAGATCCACTGCGCCGTGAAG AGCCTGAATCGGATCACCGACATCGGCGAAGTGAGCCAGTTCCTTACCGAGG GCATCATCATGAAGGACTTCTCACACCCCAACGTGCTGAGCCTGTTGGGCAT CTGCCTGCGGTCTGAGGGCAGCCCGCTGGTGGTACTGCCCTATATGAAGCAC GGCGACCTGCGGAACTTCATCCGGAACGAGACCCACAACCCCACCGTGAAAG ACCTGATCGGCTTCGGCTTGCAAGTGGCCAAGGGCATGAAGTACCTGGCCAG CAAGAAGTTCGTGCACCGGGACCTTGCCGCACGGAACTGCATGCTTGACGAG AAATTTACAGTGAAAGTGGCCGACTTCGGCCTGGCTCGGGACATGTACGACA AGGAGTACTACAGCGTGCACAACAAGACCGGCGCCAAGCTGCCCGTCAAGT GGATGGCCCTGGAGAGCCTGCAGACCCAGAAGTTTACTACCAAGTCTGACGT GTGGAGCTTCGGCGTGCTGCTGTGGGAACTGATGACCCGAGGAGCCCCTCCT TACCCCGACGTGAACACCTTCGACATCACCGTGTACCTGTTACAGGGCCGGC GGCTACTGCAGCCCGAGTACTGTCCCGACCCGCTGTACGAGGTGATGTTGAA GTGTTGGCACCCCAAGGCCGAGATGCGGCCCAGCTTCAGCGAACTGGTGAGC CGGATCAGCGCCATCTTCTCAACCTTTATCGGCGAGCACTACGTCCACGTAAA CGCCACCTACGTCAACGTGAAGTGCGTGGCCCCATACCCCAGCCTGCTCAGC AGCGAGGACAACGCCGACGACGAGGTGGATACCCGGCCCGCATCCTTCTGGG AGACCAGC (SEQ ID NO:805) For primary immunization 1000 μg mRNA was formulated in complete Freund’s Adjuvant (CFA) and was injected intramuscularly (IM). Following the initial immunizations, four boost immunizations (alternative human and cyno mRNA at 500 μg) were given approximately every two weeks intramuscularly and a final fifth boost (day 77) was given intravenously (IV). One prebleed (day 1) and two test bleed samples (day 42 and day 70, respectively) were collected to determine antibody titers against the c- MET protein in the serum. The bleed samples were tested on CHO cells expressing Hu c- MET protein via FACS to check the titers. On day 88 a production bleed of 600ml was taken and used to isolate and cryopreserve PBMCs. 2. FACS Sorting of B-cells and Culturing of Antigen Positive B-cells:
PBMCs were incubated with biotinylated c-MET protein and antigen positive VHH B-cells were selected by Fluorescence Activated Cell Sorting (FACS) using fluorescently labeled rabbit anti-camelid VHH antibody AF-647 (Genscript) and anti- biotin-PE antibody (Biolegend). B-cells were sorted at a concentration of 5 x 106 cells/mL and cells staining positive with both the antibodies (antigen positive VHH cells) were collected into 96 well plates with 3 cells/well cell density. FACS selected B-cells distributed into 96 well plates were co-cultured with irradiated CD40L expressing feeder cells and the appropriate cytokines. Mammalian B- cell culture methods are described in detail by Kwakkenbos et al.2010, Nat. Med. 16(1):123-128 (2010). Sorting methods using fluorescently labeled immunogen and culture of sorted Llama B cells were based on methods available on mammalian B cell culture (see, e.g., Kwakkenbos et al.,; WO2013076139 A1; Carbonetti et al., JIM 448: 66-73 (2017), herein incorporated by reference in their entirety). The culture plates were incubated for up to 7-10 days in a CO2 incubator at 37oC under 5% CO2. Typically, between days 8-11 Proliferating B cell cultures were processed to harvest culture supernatants for screening the secreted antibodies against the target antigen (c-MET protein), and the cell pellets were frozen for isolation of RNA and antibody sequencing. 3. Screening for Antibody Hits via FACS: B cell culture supernatant from each wells were screened for antibody binding to the CHO cells expressing hu c-MET protein via FACS. CHO cells expressing Human c- MET were resuspended in FACS buffer (PBS pH 7.4 + 2% FBS +1 mM EDTA) at 1,000,000 cells/ml, and 80 μl of the cell suspension was added in each well in a 96-well V bottom polypropylene plate (Nunc). The 40 μl of B-cell supernatants were added in each well. After one-hour incubation at 4°C, cells were pelleted and washed 2x by ice cold FACS buffer and then resuspended in 80 μl goat anti-llama IgG(H+L)-PE secondary
antibody from Jackson at 1:300 dilution in FACS buffer, incubated 15 min at 4°C, centrifuged at 1500 rpm for 3 min; cell pellets were washed twice and resuspended in 100 μl FACS buffer for analysis.ௗ Cellular fluorescence was determined on BD FACSCantoII (Becton-Dickinson, Franklin Lakes, NJ).ௗ The data was processed with FlowJo software (10.8.1). The IgG positive wells were subjected to VHH positive screening using goat anti-llama IgG 2+3-APC secondary antibody from Jackson at 1:300 dilution in FCAS buffer. The VHH positive wells were used for mRNA isolation and cDNA synthesis as discussed below. 4. Sequencing Antibodies from Target Positive B-cells: To sequence antibodies, from VHH positive wells, cDNA was generated and sequenced using Sanger sequencing and NGS. The frozen B cell pellets are lysed by adding the lysis buffer. mRNA was isolated from the lysates according to the manufacturer’s instructions and cDNA synthesis was performed with random hexamers and oligodT reverse primers using Superscript IV first strand synthesis kit (Invitrogen, 18091050). To amplify the variable domain of the heavy chain only antibodies, PCR was performed according to the standard procedures with 2-5 μl of cDNA reaction product and with historical primers used in Llama antibody discovery (see, e.g., Conrath et al., Antimicro.-Agents Chemother. v45(10):2807-2812 (2001); Els Pardon, Nat. Protoc. 9(3):674-93 (2014), herein incorporated by reference in their entirety). The PCR amplified VHH genes were then sequenced using one of two methods Sanger sequencing or Next Generation Sequencing (NGS). In both cases the PCR used CALL001 forward primer and CALL002 reverse primers that anneal to the leader sequence and 5` end of CH2 domain of the heavy chain respectively. The resulting PCR product was sequenced directly via Sanger sequencing and NGS (Azenta, MA). 5. Purification of Single Domain Antibodies: Recombinant antibodies were expressed at Biointron Biologics Inc., (China) in the IgG and VHH-His formats in the CHO cells. The antibodies were purified from 30mL
CHO using Ni-NTA affinity chromatography. The purified proteins were run on size exclusion chromatography (SEC) to check their size and homogeneity. The Antibodies were further analyzed by SDS-PAGE under reducing and non-reducing conditions to check for purity and validation of molecular weight. An average of 2-5 mg protein was purified from 30 mL CHO culture supernatants. 6. FACS Cell Binding Assays: CHO cell lines that stably express Human and Cyno c-MET were resuspended in FACS buffer (PBS pH7.4 + 2% FBS +2mM EDTA) at 1,000,000 cells/mL, and 80 μl of the cell suspension was added in each well in a 96-well V bottom polypropylene plate (Nunc). The 80 μl of testing VHH antibodies were mixed to each well at starting concentration of 1000 nM with 12 step 1:3 dilutions.ௗ After one-hour incubation at 4°C, pelleted cells were washed 2x by ice cold FACS buffer and then resuspended in 80 μ l anti-His-iFluor 647 from Biolegend at 1:500 dilution in FACS buffer, incubated 15min at 4°C, centrifuged at 1500 rpm for 3 min; cell pellets were washed twice and resuspended in 100 μl FACS buffer for analysis.ௗ Cellular fluorescence was determined on BD FACSCantoII (Becton-Dickinson, Franklin Lakes, NJ).ௗ The data was processed with FlowJo software (10.8.1). The EC50 values were determined by fitting the data with nonlinear regression model using GraphPad Prism 9.4.1 (La Jolla, CA). The primary cancer cell line (HS746T) expresses high levels of c-MET protein, and it was used for cell binding assay with the VHH antibodies as well. The assay was performed using the similar protocol as mentioned above. 7. Hepatocyte Growth Factor (HGF) Competition Assay: Hepatocyte Growth Factor (HGF) is a ligand for c-MET receptor, and it has been shown to bind to the c-MET SEMA domain. HGF binding triggers the c-MET dimerization which initiates the downstream signaling pathways. Here, the binding of VHH antibodies was tested in the presence of HGF protein. CHO cells expressing Human c-MET were resuspended in FACS buffer (PBS pH7.4 + 2% FBS +1 mM EDTA) at
1,000,000 cells/ml, and 60 μl of the cell suspension was added in each well in a 96-well V bottom polypropylene plate (Nunc). The testing VHH antibodies (IgG format) were added at a fixed 100 nM concentration in each well in the absence (0 nM HGF) and in the presence (100 nM and 1000 nM) of HGF in a final reaction volume of 150 μl. After one- hour incubation at 4°C, pelleted cells were washed 2x by ice cold FACS buffer and then resuspended in 80 μl goat anti-human Fc antibody from Abcam at 1:500 dilution in FACS buffer, incubated 15min at 4°C, centrifuged at 1500 rpm for 3 min; cell pellets were washed twice and resuspended in 100 μl FACS buffer for analysis. Cellular fluorescence was determined on BD FACSCantoII (Becton-Dickinson, Franklin Lakes, NJ).ௗ The data was processed with FlowJo software (10.8.1) and the MFI values were exported and compared. The percentage reduction in MFI data was plotted using R in RStudio. 8. Humanization of VHH Antibodies: The VHH antibodies were humanized similar to the method of Hanf et al., Methods.2014 Jan 1;65(1):68-76. doi: 10.1016/j.ymeth.2013.06.024. Epub 2013 Jun 28. Briefly, the complementarity-determining regions (CDRs) sequences of anti-C-MET VHH Abs (Table 1) were annotated using the IMGT numbering scheme. Each VHH nucleotide sequence was generated and used to identify the nearest human germline VH sequences by searching for similar sequences with the NCBI IgBLAST program. Common J and D gene sequences were attached to the VH as the acceptor. Next the most similar human VH sequences were identified using BLASTp and used to choose the nearest framework sequences into which the CDR sequences were grafted replacing the human CDRs. Rosetta/Alpha fold was used to create the structural 3D homology model the of the appropriate CDRs that were grafted into the acceptor framework. The framework residues that were critical for huVH/VL interactions are back mutated to llama sequence
canonical llama residues, also potentially structural defects due to mismatches at the graft interface can be fixed by mutating some framework residues to llama, or by mutating some residues on the CDRs’ backside to human or to a de novo designed sequence.ௗௗCDR stabilizing or overall fold stabilizing sequences were then back-mutated to the corresponding llama sequence to maintain the biophysical properties and target binding affinity.ௗௗௗ 9. FACS Cell Binding Assays of Humanized Constructs: CHO cell lines that stably express Human c-MET protein were resuspended in FACS buffer (PBS pH 7.4 + 2% FBS +2 mM EDTA) at 1,000,000 cells/mL, and 80 μl of the cell suspension was added in each well in a 96-well V bottom polypropylene plate (Nunc). The 80 μl of each humanized VHH antibodies were mixed to each well at starting concentration of 1000 nM with 12 step 1:3 dilutions.ௗ After one-hour incubation at 4°C, pelleted cells were washed 2x by ice cold FACS buffer and then resuspended in 80 μl anti-His-iFluor 647 from Biolegend at 1:500 dilution in FACS buffer, incubated 15 min at 4°C, centrifuged at 1500 rpm for 3 min; cell pellets were washed twice and resuspended in 100 μl FACS buffer for analysis.ௗ Cellular fluorescence was determined on BD FACSCantoII (Becton-Dickinson, Franklin Lakes, NJ).ௗ The data was processed with FlowJo software (10.8.1). The EC50 values were determined by fitting the data with nonlinear regression model using GraphPad Prism 9.4.1 (La Jolla, CA). Example 2: Generation and Selection of VHH antibodies against c-MET A total of 40 single domain antibodies have been discovered against c-MET protein. Figures 1 and 2 show the monovalent binding of these VHH antibodies to CHO cells expressing human c-MET and Cyno c-MET proteins, respectively, while Figure 5 shows monovalent binding to primary cancer cells (HS746T) expressing c-MET. For 35 of these VHH Abs, bivalent binding was also tested (Figures 3 and 4). For the remaining five VHH Abs (namely CV3N1, CV7N1, CV13N2, CV16N1 & CV19N1), bivalent binding will be tested once these antibodies are available in the bivalent format. Overall,
these 40 antibodies showed strong binding to CHO cells expressing human and cyno c- MET and to HS746T c-MET cancer cells. These antibodies are cross reactive with human and cyno based on their bivalent binding. Furthermore, 16 VHH antibodies competed with HGF for c-MET binding (over 50% reduction in MFI when HGF is present) based on the HGF competition assay (Figure 6). There are 11 VHH antibodies that did not compete (no % reduction in MFI in the presence of HGF) and 10 that showed semi- competition (less than 50% reduction in MFI in the presence of HGF). The remaining three VHH Abs (CV3N1, CV16N1 and CV19N1) will be tested in future HGF competition assays since the anti-VHH secondary antibody did not bind to them. A summary of this data is presented in Tables A through D below. Table A. The monovalent EC50 (nM) values for anti-c-MET VHH clones on CHO human c-MET and CHO cyno c-MET cells.
Table B. The bivalent EC50 (nM) values for anti-c-MET VHH clones on CHO human c-MET and CHO cyno c-MET cells.
NA = data not available. Table C. The monovalent EC50 (nM) values for anti-c-MET VHH clones on HS746T cancer cells.
Table D. Hepatocyte Growth Factor (HGF) competition for anti-c-MET VHH clones.
Example 3. Generation of Humanized VHH antibodies against c-MET A total of 19 VHH Abs were humanized (see, Figures.7A, 7B, and 7C) and evaluated for binding on CHO-Hu c-MET cells (see, Figure 8A, 8B, and 8C). Most humanized constructs retain binding and show very similar binding as the WT llama VHHs (see, Figure 9).
Claims
WHAT IS CLAIMED IS: 1. A polypeptide comprising a variable domain of the heavy chain of a heavy chain-only antibody (VHH) that specifically binds human and cynomolgus c-MET, the VHH comprising VHH complementarity determining region (CDR) 1, VHH CDR2, and VHH CDR3 of any one of the amino acid sequences set forth in any one of SEQ ID NOs.: 4, 22, 37, 1 to 3, 5 to 21, 23 to 36, and 38 to 40, optionally wherein the VHH CDR1, VHH CDR2, and VHH CDR3 are based on CDR definitions according to Kabat, Chothia, enhanced Chothia, contact, IMGT, or AbM; and further optionally wherein (i) the polypeptide inhibits the binding of Human Growth Factor (HGF) to human c-MET, or (ii) the polypeptide does not inhibit the binding of Human Growth Factor (HGF) to human c- MET.
2. The polypeptide of claim 1, further comprising (i) an agent selected from the group consisting of a purification tag, a fluorophore, a drug, a photosensitizer, a nanoparticle, a toxic agent, a radionuclide, a VHH, an Fab, a scFv, a multimerization module, a moiety that facilitates the polypeptide crossing the blood brain barrier, and a half-life extension moiety, optionally wherein the agent is attached or linked to the C-terminus of the VHH; or (ii) a human Ig Fc domain, optionally further comprising a human Ig hinge domain, and further optionally comprising an amino acid sequence that is at least 80%, at least 85%, at least 90%, at least 95%, or 100% identical to the human IgG4 PAA hinge and Fc domain sequence (SEQ ID NO:643).
3. The polypeptide of claim 1 or 2, further comprising a second VHH that specifically binds to an antigen selected from the group consisting of: CD3, EGFR, VEGF, VEGFR, HGF, CXCR2, CXCR4, CXCR7, CXCL11, CXCL12, CEA, PSMA, MMR, PD-1, PDL- 1, survivin, HER2, HGFR, P2X7, death receptor 5 CTLA4, CD7, CD8, CD11b, CD20, CD38, CD45, M HC-II, fibronectin, TUFM, CapG, CAIX, CD33, CD47, ARTC22, B3GAT1 (CD57), CCR7 (CD197), CD16, CD16a, CD2 CD226, CD244, CD27, CD3, CD300A, CD34, CD58, CD59, CD69, CSF2, CX3CR1, CXCR1 (CD128), CXCR3 (CD183), CXCR4, EOMES, GZMB, ICAM1 (CD54), IFNG, IL-15R, IL-1R, IL22, IL-
2RB (CD122), IL-7R (CD127), ITGA1 (CD49a), ITGA2 (CD49b), ITGAL (CD11a), ITGAM (CD11b), ITGB2 (CD18), KIR, KIR2DL1, KIR2DL2, KIT (CD117), KLRB1C, KLRC1, KLRC2, KLRD1 (CD94), KLRF1, KLRG1, KLRK1, LILRB1, KLRA4, KLRA8, MICA/BNCAM1 (CD56), NK2D, NKP46 (NCR1, CD335), NCR2, NCR3 (CD337), PRF1, SELL (CD62L), SIGLEC7, SLAMF6, SPN, TBX21, and TNFα.
4. The polypeptide of any one of claims 1 to 3, wherein the VHH comprises: (a) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:50, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:51, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:52; (b) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:104, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:105, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:106; (c) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:149, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:150, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:151; (d) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:41, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:42, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:43; (e) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:44, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:45, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:46; (f) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:47, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:48, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:49:
(g) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:53, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:54, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:55; (h) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:56, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:57, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:58; (i) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:59, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:60, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:61; (j) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:62, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:63, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:64; (k) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:65, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:66, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:67; (l) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:68, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:69, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:70; (m) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:71, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:72, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:73; (n) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:74, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:75, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:76;
(o) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:77, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:78, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO79 ; (p) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:80, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:81, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:82; (q) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:83, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:84, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:85; (r) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:86, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:87, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:88; (s) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:89, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:90, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:91; (t) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:92, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:93, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:94; (u) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:95, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:96, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:97; (v) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:98, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:99, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:100;
(w) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:101, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:102, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:103; (x) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:107, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:108, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:109; (y) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:110, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:111, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:112; (z) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:113, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:114, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:115; (aa) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:116, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:117, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:118; (ab) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:119, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:120, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:121; (ac) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:122, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:123, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:124; (ad) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:125, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:126, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:127;
(ae) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:128, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:129, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:130; (af) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:131, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:132, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:133; (ag) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:134, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:135, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:136; (ah) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:137, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:138, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:139; (ai) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:140, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:141, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:142; (aj) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:143, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:144, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:145; (ak) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:146, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:147, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:148; (al) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:152, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:153, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:154;
(am) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:155, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:156, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:157; or (an) a VHH CDR1 comprising the amino acid sequence set forth in SEQ ID NO:158, a VHH CDR2 comprising the amino acid sequence set forth in SEQ ID NO:159, and a VHH CDR3 comprising the amino acid sequence set forth in SEQ ID NO:160.
5. The polypeptide of any one of claims 1 to 4, wherein the VHH comprises a sequence that is at least 80%, at least 85%, at least 90%, at least 95%, or 100% identical to any amino acid sequence set forth in Table I, optionally wherein the VHH comprises a sequence that is set forth in any one of SEQ ID NOs.: 687 to 698 or 813, 744 to 750, 699 to 743 or 811, or 751 to 804.
6. A bispecific antibody comprising (i) the polypeptide of any one of claims 1 to 5; and (ii) a binding molecule that binds to a different epitope of c-MET than the polypeptide or binds to a different antigen, optionally wherein the different antigen is an antigen on a T cell, an NK cell, or a tumor cell, and further optionally wherein the different antigen is an antigen selected from the group consisting of: CD3, EGFR, VEGF, VEGFR, HGF, CXCR2, CXCR4, CXCR7, CXCL11, CXCL12, CEA, PSMA, MMR, PD-1, PDL-1, survivin, HER2, HGFR, P2X7, death receptor 5 CTLA4, CD7, CD8, CD11b, CD20, CD38, CD45, M HC-II, fibronectin, TUFM, CapG, CAIX, CD33, CD47, ARTC22, B3GAT1 (CD57), CCR7 (CD197), CD16, CD16a, CD16b, CD2 CD226, CD244, CD27, CD3, CD300A, CD34, CD58, CD59, CD69, CSF2, CX3CR1, CXCR1 (CD128), CXCR3 (CD183), CXCR4, EOMES, GZMB, ICAM1 (CD54), IFNG, IL-15R, IL-1R, IL22, IL- 2RB (CD122), IL-7R (CD127), ITGA1 (CD49a), ITGA2 (CD49b), ITGAL (CD11a), ITGAM (CD11b), ITGB2 (CD18), KIR, KIR2DL1, KIR2DL2, KIT (CD117), KLRB1C, KLRC1, KLRC2, KLRD1 (CD94), KLRF1, KLRG1, KLRK1, LILRB1, KLRA4, KLRA8, MICA/BNCAM1 (CD56), NK2D, NKP46 (NCR1, CD335), NCR2, NCR3 (CD337), PRF1, SELL (CD62L), SIGLEC7, SLAMF6, SPN, TBX21, and TNFα.
7. A c-MET binding chimeric antigen receptor (CAR) comprising the polypeptide of any one of claims 1 to 5, optionally further comprising CD28 and CD137 signaling domains and CD3ζ (c-MET-28-137-3ζ).
8. A nucleic acid or nucleic acids encoding the polypeptide of any one of claims 1 to 5, the bispecific antibody of claim 6, or the c-MET binding CAR of claim 7.
9. A vector or vectors comprising the nucleic acid or nucleic acids of claim 8.
10. A host cell comprising the nucleic acid or nucleic acids of claim 8, or the vector or vectors of claim 9.
11. A method of making a polypeptide or a bispecific antibody, the method comprising culturing the host cell of claim 10 under conditions that facilitate expression of the polypeptide or the bispecific antibody, and isolating the polypeptide or the bispecific antibody, optionally further comprising formulating the polypeptide or the bispecific antibody as a sterile pharmaceutical composition.
12. A pharmaceutical composition comprising the polypeptide of any one of claims 1 to 5, the bispecific antibody of claim 6, or a T cell or NK cell expressing the c-MET binding CAR of claim 7, and a pharmaceutically acceptable carrier.
13. A method of treating a c-MET expressing cancer in a human subject in need thereof, killing a tumor cell in a human subject in need thereof, or decreasing the rate of tumor growth in a human subject in need thereof, the method comprising administering to the human subject a therapeutically effective amount of the polypeptide of any one of claims 1 to 5, the bispecific antibody of claim 6, or a T cell or NK cell expressing the c-MET binding CAR of claim 7, optionally wherein the cancer is a solid tumor, and further optionally wherein the cancer is selected from the group consisting of gastric cancer, esophageal cancer, medulloblastoma, glioma, colon cancer, head and neck cancer, lung cancer, kidney cancer, thyroid cancer, colorectal cancer, pancreatic cancer, ovarian cancer, and breast cancer.
14. A polynucleotide comprising an mRNA comprising: (i) a 5' UTR; (ii) an open reading frame (ORF) encoding a polypeptide of any one of claims 1 to 5, the bispecific antibody of claim 6, or the c-MET binding CAR of claim 7; (iii) a stop codon; and (iv) a 3' UTR.
15. The polynucleotide of claim 14, wherein the mRNA comprises a microRNA (miR) binding site.
16. The polynucleotide of any one of claim 14 or 15, wherein the 5' UTR comprises a nucleic acid sequence at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:654.
17. The polynucleotide of any one of claims 14 to 16, wherein the 3' UTR comprises a nucleic acid sequence at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or 100% identical to SEQ ID NO:674.
18. The polynucleotide of any one of claims 14 to 17, wherein the mRNA comprises a 5' terminal cap.
19. The polynucleotide of any one of claims 14 to 18, wherein the mRNA comprises a poly-A region.
20. The polynucleotide of any one of claims 14 to 19, wherein the mRNA comprises at least one chemically modified nucleobase, sugar, backbone, or any combination thereof, optionally wherein the at least one chemically modified nucleobase is selected from the group consisting of pseudouracil (Ψ), N1-methylpseudouracil (m1Ψ), 1- ethylpseudouracil, 2-thiouracil (s2U), 4’-thiouracil, 5-methylcytosine, 5-methyluracil, 5- methoxyuracil, and any combination thereof.
21. The polynucleotide of claim 20, wherein at least about 25%, at least about 30%, at least about 40%, at least about 50%, at least about 60%, at least about 70%, at least about 80%, at least about 90%, at least about 95%, at least about 99%, or 100% of the uracils are N1-methylpseudouracils.
22. The polynucleotide of any one of claims 14 to 19, wherein the open reading frame consists of nucleosides selected from the group consisting of (i) uridine or a modified uridine, (ii) cytidine or a modified cytidine, (iii) adenosine or a modified adenosine, and (iv) guanosine or a modified guanosine, optionally wherein the modified uridine is 1- methylpseudouridine.
23. The polynucleotide of any one of claims 14 to 22, wherein the mRNA comprises a 5’terminal cap comprising m7GpppGm and a poly-A region 100 nucleotides in length, optionally wherein all uracils of the polynucleotide are N1-methylpseudouracils. 24. A pharmaceutical composition comprising the polynucleotide of any one of claims 14 to 23, and a delivery agent, optionally wherein the delivery agent comprises a lipid nanoparticle, and further optionally wherein the lipid nanoparticle comprises a neutral lipid, an ionizable amino lipid, a polyethyleneglycol (PEG) lipid, and/or a sterol, further optionally wherein the neutral lipid is 1,2-distearoyl-sn-glycero-3-phosphocholine; wherein the ionizable amino lipid is a compound of Formula (I); wherein the PEG lipid is PEG 2000 dimyristoyl glycerol or 134-hydroxy- 3,6,9,12,15,18,21,
24,27,30,33,36,39,42,45,48,51,54,57,60,63,66,69,72,75,78,81,84,87,90, 93,96,99,102,105,108,111,114,117,120,123,126,129,132- tetratetracontaoxatetratriacontahectyl stearate, and wherein the sterol is cholesterol.
25. A method of treating a c-MET expressing cancer in a human subject in need thereof, killing a tumor cell in a human subject in need thereof, or decreasing the rate of tumor growth in a human subject in need thereof, the method comprising administering to the human subject a therapeutically effective amount of the pharmaceutical composition of claim 24, optionally wherein the cancer is a solid tumor, and further optionally wherein the cancer is selected from the group consisting of gastric cancer, esophageal cancer,
medulloblastoma, glioma, colon cancer, head and neck cancer, lung cancer, kidney cancer, thyroid cancer, colorectal cancer, pancreatic cancer, ovarian cancer, and breast cancer.
26. A kit comprising (i) the polypeptide of any one of claims 1 to 5; the bispecific antibody of any one of claim 6; a T cell or NK cell expressing the c-MET binding CAR of claim 7; the pharmaceutical composition of claim 12 or 24; or the polynucleotide of any one of claims 14-23, and (ii) a package insert instructing a user of the kit to administer the polypeptide, bispecific antibody, c-MET binding CAR, pharmaceutical composition, or polynucleotide to a human subject in need thereof.
Applications Claiming Priority (4)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363533759P | 2023-08-21 | 2023-08-21 | |
| US63/533,759 | 2023-08-21 | ||
| US202363546326P | 2023-10-30 | 2023-10-30 | |
| US63/546,326 | 2023-10-30 |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| WO2025042806A2 true WO2025042806A2 (en) | 2025-02-27 |
| WO2025042806A3 WO2025042806A3 (en) | 2025-05-30 |
Family
ID=92672258
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/042856 Pending WO2025042806A2 (en) | 2023-08-21 | 2024-08-19 | C-met binding antibodies, nucleic acids encoding same, and methods of use |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025042806A2 (en) |
Citations (30)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20040262223A1 (en) | 2001-07-27 | 2004-12-30 | President And Fellows Of Harvard College | Laminar mixing apparatus and methods |
| US20050222064A1 (en) | 2002-02-20 | 2005-10-06 | Sirna Therapeutics, Inc. | Polycationic compositions for cellular delivery of polynucleotides |
| WO2008016473A2 (en) | 2006-07-28 | 2008-02-07 | Applera Corporation | Dinucleotide mrna cap analogs |
| US7404969B2 (en) | 2005-02-14 | 2008-07-29 | Sirna Therapeutics, Inc. | Lipid nanoparticle based compositions and methods for the delivery of biologically active molecules |
| WO2008121949A1 (en) | 2007-03-30 | 2008-10-09 | Bind Biosciences, Inc. | Cancer cell targeting using nanoparticles |
| WO2008127688A1 (en) | 2007-04-13 | 2008-10-23 | Hart Communication Foundation | Synchronizing timeslots in a wireless communication protocol |
| WO2010005725A2 (en) | 2008-06-16 | 2010-01-14 | Bind Biosciences, Inc. | Therapeutic polymeric nanoparticles comprising vinca alkaloids and methods of making and using same |
| WO2010005726A2 (en) | 2008-06-16 | 2010-01-14 | Bind Biosciences Inc. | Therapeutic polymeric nanoparticles with mtor inhibitors and methods of making and using same |
| US20100129877A1 (en) | 2005-09-28 | 2010-05-27 | Ugur Sahin | Modification of RNA, Producing an Increased Transcript Stability and Translation Efficiency |
| WO2010075072A2 (en) | 2008-12-15 | 2010-07-01 | Bind Biosciences | Long circulating nanoparticles for sustained release of therapeutic agents |
| US20100293625A1 (en) | 2007-09-26 | 2010-11-18 | Interexon Corporation | Synthetic 5'UTRs, Expression Vectors, and Methods for Increasing Transgene Expression |
| WO2011015347A1 (en) | 2009-08-05 | 2011-02-10 | Biontech Ag | Vaccine composition comprising 5'-cap modified rna |
| WO2011084518A2 (en) | 2009-12-15 | 2011-07-14 | Bind Biosciences, Inc. | Therapeutic polymeric nanoparticles comprising corticosteroids and methods of making and using same |
| WO2011084521A2 (en) | 2009-12-15 | 2011-07-14 | Bind Biosciences, Inc. | Therapeutic polymeric nanoparticles comprising epothilone and methods of making and using same |
| US8158601B2 (en) | 2009-06-10 | 2012-04-17 | Alnylam Pharmaceuticals, Inc. | Lipid formulation |
| WO2012099755A1 (en) | 2011-01-11 | 2012-07-26 | Alnylam Pharmaceuticals, Inc. | Pegylated lipids and their use for drug delivery |
| US20120201859A1 (en) | 2002-05-02 | 2012-08-09 | Carrasquillo Karen G | Drug Delivery Systems and Use Thereof |
| US20120276209A1 (en) | 2009-11-04 | 2012-11-01 | The University Of British Columbia | Nucleic acid-containing lipid particles and related methods |
| WO2012170889A1 (en) | 2011-06-08 | 2012-12-13 | Shire Human Genetic Therapies, Inc. | Cleavable lipids |
| WO2013076139A1 (en) | 2011-11-23 | 2013-05-30 | F. Hoffmann-La Roche Ag | Cd40l expressing mammalian cells and their use |
| US20130150295A1 (en) | 2006-12-21 | 2013-06-13 | Stryker Corporation | Sustained-Release Formulations Comprising Crystals, Macromolecular Gels, and Particulate Suspensions of Biologic Agents |
| WO2013086354A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| WO2013103659A1 (en) | 2012-01-04 | 2013-07-11 | Board Of Supervisors Of Louisiana State University And Agricultural And Mechanical College | Stabilizing rna by incorporating chain-terminating nucleosides at the 3'-terminus |
| WO2013116126A1 (en) | 2012-02-01 | 2013-08-08 | Merck Sharp & Dohme Corp. | Novel low molecular weight, biodegradable cationic lipids for oligonucleotide delivery |
| US8519110B2 (en) | 2008-06-06 | 2013-08-27 | Board Of Supervisors Of Louisiana State University And Agricultural And Mechanical College | mRNA cap analogs |
| WO2014164253A1 (en) | 2013-03-09 | 2014-10-09 | Moderna Therapeutics, Inc. | Heterologous untranslated regions for mrna |
| WO2015130584A2 (en) | 2014-02-25 | 2015-09-03 | Merck Sharp & Dohme Corp. | Lipid nanoparticle vaccine adjuvants and antigen delivery systems |
| WO2017049275A2 (en) | 2015-09-17 | 2017-03-23 | Moderna Therapeutics, Inc. | Polynucleotides containing a stabilizing tail region |
| WO2017049286A1 (en) | 2015-09-17 | 2017-03-23 | Moderna Therapeutics, Inc. | Polynucleotides containing a morpholino linker |
| WO2017106462A1 (en) | 2015-12-18 | 2017-06-22 | Biogen Ma Inc. | Bispecific antibody platform |
Family Cites Families (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US9777064B2 (en) * | 2015-03-17 | 2017-10-03 | Chimera Bioengineering, Inc. | Smart CAR devices, DE CAR polypeptides, side CARs and uses thereof |
| TW202515920A (en) * | 2017-04-11 | 2025-04-16 | 美商因荷布瑞克斯生物科學公司 | Multispecific polypeptide constructs having constrained cd3 binding and methods of using the same |
| WO2022104236A2 (en) * | 2020-11-16 | 2022-05-19 | Ab Therapeutics, Inc. | Multispecific antibodies and uses thereof |
-
2024
- 2024-08-19 WO PCT/US2024/042856 patent/WO2025042806A2/en active Pending
Patent Citations (36)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20040262223A1 (en) | 2001-07-27 | 2004-12-30 | President And Fellows Of Harvard College | Laminar mixing apparatus and methods |
| US20050222064A1 (en) | 2002-02-20 | 2005-10-06 | Sirna Therapeutics, Inc. | Polycationic compositions for cellular delivery of polynucleotides |
| US20120201859A1 (en) | 2002-05-02 | 2012-08-09 | Carrasquillo Karen G | Drug Delivery Systems and Use Thereof |
| US7404969B2 (en) | 2005-02-14 | 2008-07-29 | Sirna Therapeutics, Inc. | Lipid nanoparticle based compositions and methods for the delivery of biologically active molecules |
| US20100129877A1 (en) | 2005-09-28 | 2010-05-27 | Ugur Sahin | Modification of RNA, Producing an Increased Transcript Stability and Translation Efficiency |
| WO2008016473A2 (en) | 2006-07-28 | 2008-02-07 | Applera Corporation | Dinucleotide mrna cap analogs |
| US20130150295A1 (en) | 2006-12-21 | 2013-06-13 | Stryker Corporation | Sustained-Release Formulations Comprising Crystals, Macromolecular Gels, and Particulate Suspensions of Biologic Agents |
| WO2008121949A1 (en) | 2007-03-30 | 2008-10-09 | Bind Biosciences, Inc. | Cancer cell targeting using nanoparticles |
| WO2008127688A1 (en) | 2007-04-13 | 2008-10-23 | Hart Communication Foundation | Synchronizing timeslots in a wireless communication protocol |
| US20100293625A1 (en) | 2007-09-26 | 2010-11-18 | Interexon Corporation | Synthetic 5'UTRs, Expression Vectors, and Methods for Increasing Transgene Expression |
| US20130172406A1 (en) | 2007-09-28 | 2013-07-04 | Bind Biosciences, Inc. | Cancer Cell Targeting Using Nanoparticles |
| US20120004293A1 (en) | 2007-09-28 | 2012-01-05 | Zale Stephen E | Cancer Cell Targeting Using Nanoparticles |
| US8519110B2 (en) | 2008-06-06 | 2013-08-27 | Board Of Supervisors Of Louisiana State University And Agricultural And Mechanical College | mRNA cap analogs |
| US20100069426A1 (en) | 2008-06-16 | 2010-03-18 | Zale Stephen E | Therapeutic polymeric nanoparticles with mTor inhibitors and methods of making and using same |
| US20100104655A1 (en) | 2008-06-16 | 2010-04-29 | Zale Stephen E | Therapeutic Polymeric Nanoparticles Comprising Vinca Alkaloids and Methods of Making and Using Same |
| WO2010005725A2 (en) | 2008-06-16 | 2010-01-14 | Bind Biosciences, Inc. | Therapeutic polymeric nanoparticles comprising vinca alkaloids and methods of making and using same |
| WO2010005726A2 (en) | 2008-06-16 | 2010-01-14 | Bind Biosciences Inc. | Therapeutic polymeric nanoparticles with mtor inhibitors and methods of making and using same |
| US20100216804A1 (en) | 2008-12-15 | 2010-08-26 | Zale Stephen E | Long Circulating Nanoparticles for Sustained Release of Therapeutic Agents |
| US20110217377A1 (en) | 2008-12-15 | 2011-09-08 | Zale Stephen E | Long Circulating Nanoparticles for Sustained Release of Therapeutic Agents |
| WO2010075072A2 (en) | 2008-12-15 | 2010-07-01 | Bind Biosciences | Long circulating nanoparticles for sustained release of therapeutic agents |
| US8158601B2 (en) | 2009-06-10 | 2012-04-17 | Alnylam Pharmaceuticals, Inc. | Lipid formulation |
| WO2011015347A1 (en) | 2009-08-05 | 2011-02-10 | Biontech Ag | Vaccine composition comprising 5'-cap modified rna |
| US20120276209A1 (en) | 2009-11-04 | 2012-11-01 | The University Of British Columbia | Nucleic acid-containing lipid particles and related methods |
| WO2011084521A2 (en) | 2009-12-15 | 2011-07-14 | Bind Biosciences, Inc. | Therapeutic polymeric nanoparticles comprising epothilone and methods of making and using same |
| WO2011084518A2 (en) | 2009-12-15 | 2011-07-14 | Bind Biosciences, Inc. | Therapeutic polymeric nanoparticles comprising corticosteroids and methods of making and using same |
| WO2012099755A1 (en) | 2011-01-11 | 2012-07-26 | Alnylam Pharmaceuticals, Inc. | Pegylated lipids and their use for drug delivery |
| WO2012170889A1 (en) | 2011-06-08 | 2012-12-13 | Shire Human Genetic Therapies, Inc. | Cleavable lipids |
| WO2013076139A1 (en) | 2011-11-23 | 2013-05-30 | F. Hoffmann-La Roche Ag | Cd40l expressing mammalian cells and their use |
| WO2013086354A1 (en) | 2011-12-07 | 2013-06-13 | Alnylam Pharmaceuticals, Inc. | Biodegradable lipids for the delivery of active agents |
| WO2013103659A1 (en) | 2012-01-04 | 2013-07-11 | Board Of Supervisors Of Louisiana State University And Agricultural And Mechanical College | Stabilizing rna by incorporating chain-terminating nucleosides at the 3'-terminus |
| WO2013116126A1 (en) | 2012-02-01 | 2013-08-08 | Merck Sharp & Dohme Corp. | Novel low molecular weight, biodegradable cationic lipids for oligonucleotide delivery |
| WO2014164253A1 (en) | 2013-03-09 | 2014-10-09 | Moderna Therapeutics, Inc. | Heterologous untranslated regions for mrna |
| WO2015130584A2 (en) | 2014-02-25 | 2015-09-03 | Merck Sharp & Dohme Corp. | Lipid nanoparticle vaccine adjuvants and antigen delivery systems |
| WO2017049275A2 (en) | 2015-09-17 | 2017-03-23 | Moderna Therapeutics, Inc. | Polynucleotides containing a stabilizing tail region |
| WO2017049286A1 (en) | 2015-09-17 | 2017-03-23 | Moderna Therapeutics, Inc. | Polynucleotides containing a morpholino linker |
| WO2017106462A1 (en) | 2015-12-18 | 2017-06-22 | Biogen Ma Inc. | Bispecific antibody platform |
Non-Patent Citations (57)
| Title |
|---|
| ABRAHAM ET AL., SCIENCE, vol. 295, 2002, pages 647 - 651 |
| ADURI, R ET AL., JOURNAL OF CHEMICAL THEORY AND COMPUTATION, vol. 3, no. 4, 2006, pages 1464 - 75 |
| AHMAD ET AL., ADVANCES IN CANCER BIOLOGY - METASTASIS, vol. 4, 2022, pages 100035 |
| BAKER, PARKER, CURR OPIN CELL BIOL, vol. 16, no. 3, 2004, pages 293 - 299 |
| BELLIVEAU ET AL., MOLECULAR THERAPY-NUCLEIC ACIDS, vol. 1, 2012, pages e37 |
| BLUMER ET AL., MECH DEV, vol. 110, no. 1-2, 2002, pages 97 - 112 |
| CARBONETTI ET AL., JIM, vol. 448, 2017, pages 66 - 73 |
| CHEN ET AL., J. AM. CHEM. SOC., vol. 134, no. 16, 2012, pages 6948 - 51 |
| CHEN ET AL., THERANOSTICS, vol. 11, no. 4, 2021, pages 1901 - 1917 |
| CONRATH ET AL., ANTIMICRO. AGENTS CHEMOTHER., vol. 45, no. 10, 2001, pages 2807 - 2812 |
| DE MUNTER ET AL., INT J MOL SCI, vol. 19, 2018, pages 1 - 11 |
| DELIDAKIS ET AL., ANN REV. BIOMED ENGG., vol. 24, 2022, pages 249 - 274 |
| DESZYNSKI ET AL., NUCLEIC ACIDS RES., vol. 50, 2022, pages D1273 - D1281 |
| ELS PARDON, NAT. PROTOC., vol. 9, no. 3, 2014, pages 674 - 93 |
| FEINS ET AL., AM. J HEMAOL., vol. 94, no. S1, 2019, pages S3 - S9 |
| FUKUHARA ET AL., BIOCHEMISTRY, vol. 1, no. 4, 1962, pages 563 - 568 |
| GARDIN ET AL., ELIFE, vol. 3, 2014, pages e03735 |
| GEBAUER ET AL., COLD SPRING HARB PERSPECT BIOL, vol. 4, no. 7, 2012, pages a012245 |
| GOLDBERG-COHEN ET AL., J BIOL CHEM, vol. 277, no. 16, 2002, pages 13635 - 13640 |
| GREENE ET AL.: "Protective Groups in Organic Synthesis", 1991, WILEY & SONS |
| HA ET AL., FRONT IMMUNOL, vol. 7, 2016, pages 394 |
| HAJARI ET AL., IUBMB LIFE, vol. 71, 2019, pages 1259 - 1267 |
| HANF ET AL., METHODS, vol. 65, no. 1, 28 June 2013 (2013-06-28), pages 68 - 76 |
| HINNEBUSCH, ANNU REV MICROBIOL, vol. 59, 2005, pages 407 - 450 |
| HOLCIK ET AL., TRENDS GENET, vol. 16, no. 10, 2000, pages 469 - 473 |
| HUANG ET AL., J BIOMED RES., vol. 36, no. 1, January 2022 (2022-01-01), pages 10 - 21 |
| INGOLIA ET AL., SCIENCE, vol. 324, no. 5924, 2009, pages 218 - 23 |
| KEDDE ET AL., NATURE CELL BIOLOGY, vol. 12, no. 10, 2010, pages 1014 - 20 |
| KORE ET AL., BIOORGANIC & MEDICINAL CHEMISTRY, vol. 21, 2013, pages 4570 - 4574 |
| KOZAK, PROC NATL ACAD SCI, vol. 83, 1986, pages 2850 - 2854 |
| KWAKKENBOS ET AL., NAT. MED., vol. 16, no. 1, 2010, pages 123 - 128 |
| LAFRANCE ET AL., DEV. COMP. IMMUNOL., vol. 27, no. 1, 2003, pages 55 - 77 |
| LIU 1-1S-AXETIA ASIDHU SSWU D: "Fc Engineering for Developing Therapeutic Bispecific Antibodies and Novel Scaffolds", FRONT. IMMUNOL, vol. 8, 2017, pages 38 |
| MANDALROSSI, NAT. PROTOC., vol. 8, no. 3, 2013, pages 568 - 82 |
| MIMOTO ET AL., CURR PHARM BIOTECHNOL., vol. 17, no. 15, 2016, pages 1298 - 1314 |
| MOORE ET AL., METHODS, vol. 154, 2019, pages 38 - 50 |
| NAMY ET AL., MOL CELL, vol. 13, no. 2, 2004, pages 157 - 168 |
| OGATA ET AL., J. ORG. CHEM., vol. 74, 2009, pages 2585 - 2588 |
| PELLETIERSONENBERG, CELL, vol. 40, no. 3, 1985, pages 515 - 526 |
| PURMAL ET AL., NUCL. ACIDS RES., vol. 22, no. 1, 1994, pages 72 - 78 |
| SADEGHNEZHAD, G ET AL., INT J MOL SCI., vol. 20, 2019, pages 4818 |
| SANG ET AL., STRUCTURE, vol. 30, 2022, pages 418 - 429 |
| TATIANA, FEMS MICROBIOL. LETT., vol. 174, 1999, pages 247 - 250 |
| VERHAAR ET AL., SEMIN. IMMUNOL., vol. 52, 2021, pages 101425 |
| VILLALBA, CURR OPIN GENET DEV, vol. 21, no. 4, 2011, pages 452 - 457 |
| VINCKE ET AL., J. BIOL. CHEM., vol. 284, 2009, pages 3273 - 3284 |
| VISHWAKARMA ET AL., INTL. J. MOL. SCI., vol. 23, 2022, pages 3721 |
| WHITESIDESGEORGE M., NATURE, vol. 442, 2006, pages 368 - 373 |
| WILKINSON 1ANDERSON SFRY JJULIEN LANEVILLE DQURESHI O ET AL.: "Fe-engineered antibodies with immune effector functions completely abolished", PLOS ONE, vol. 16, no. 12, 2021, pages e0260954, XP093016187, DOI: 10.1371/journal.pone.0260954 |
| XIE ET AL., CANCER IMMUNOL. RES., vol. 8, 2020, pages 518 - 530 |
| XIE ET AL., PNAS, vol. 116, 2019, pages 7624 - 7631 |
| XU ET AL., TETRAHEDRON, vol. 48, no. 9, 1992, pages 1729 - 1740 |
| YAKUBOV ET AL., BIOCHEM. BIOPHYS. RES. COMMUN., vol. 394, no. 1, 2010, pages 189 - 193 |
| YU ET AL., NAT METHODS, vol. 16, 2019, pages 1095 - 1100 |
| ZHANG ET AL., J. MOL BIOL., vol. 335, 2004, pages 49 - 56 |
| ZHIGALTSEV ET AL., LANGMUIR, vol. 28, 2012, pages 3633 - 40 |
| ZUO ET AL., BMC GENOM., vol. 18, 2017, pages 797 |
Also Published As
| Publication number | Publication date |
|---|---|
| WO2025042806A3 (en) | 2025-05-30 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US11802144B2 (en) | Circular RNA compositions and methods | |
| US20240209068A1 (en) | Mucosal expression of antibody structures and isotypes by mrna | |
| JP2024160243A (en) | mRNA combination therapy for the treatment of cancer | |
| KR20230069042A (en) | Circular RNA compositions and methods | |
| WO2021236855A1 (en) | Circular rna compositions and methods | |
| IL302641A (en) | Genetically engineered cells and methods for their preparation | |
| US20210123075A1 (en) | Compositions and methods for immunooncology | |
| CN113891718B (en) | Artificial immune surveillance chimeric antigen receptor (AI-CAR) and its expressing cells | |
| WO2020061397A1 (en) | Compositions and methods for delivery of nucleic acids | |
| TW202246504A (en) | Ror1 targeting chimeric antigen receptor | |
| KR20250006275A (en) | Acetylated ribonucleic acid and uses thereof | |
| JP2025535029A (en) | Novel anti-mesothelin chimeric antigen receptors and modified immune cells | |
| WO2024118866A1 (en) | Gpc3-specific antibodies, binding domains, and related proteins and uses thereof | |
| WO2025042806A2 (en) | C-met binding antibodies, nucleic acids encoding same, and methods of use | |
| WO2024259531A1 (en) | Lipid nanoparticles containing disulfide lipids and formulations thereof | |
| WO2025064369A1 (en) | Humanized anti-cd3 binding molecules and uses thereof | |
| WO2024206126A1 (en) | Cd16-binding antibodies and uses thereof | |
| WO2024206209A1 (en) | Anti-cd38, anti-bcma, anti-gprc5d, and anti-fcrh5 antibodies and uses to treat various diseases such as cancer | |
| WO2024097639A1 (en) | Hsa-binding antibodies and binding proteins and uses thereof | |
| WO2026006203A2 (en) | Compositions and methods for making circular rna | |
| CA3216051A1 (en) | Glutamic acid-based lipids, lipid nanoparticle containing glutamic acid-based lipids, and formulations thereof | |
| CA3215927A1 (en) | Lipid nanoparticles containing disulfide lipids and formulations thereof | |
| CA3204195A1 (en) | Lipid nanoparticles containing disulfide lipids and formulations thereof | |
| CA3207463A1 (en) | Lipid nanoparticles containing disulfide lipids and formulations thereof | |
| WO2024206329A1 (en) | Nucleic acid molecules encoding bi-specific secreted engagers and uses thereof |