WO2025040598A2 - Peptides displayed by mhc for use in immunotherapy against different types of cancer - Google Patents
Peptides displayed by mhc for use in immunotherapy against different types of cancer Download PDFInfo
- Publication number
- WO2025040598A2 WO2025040598A2 PCT/EP2024/073138 EP2024073138W WO2025040598A2 WO 2025040598 A2 WO2025040598 A2 WO 2025040598A2 EP 2024073138 W EP2024073138 W EP 2024073138W WO 2025040598 A2 WO2025040598 A2 WO 2025040598A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- peptide
- cancer
- salt
- cells
- peptides
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K39/0005—Vertebrate antigens
- A61K39/0011—Cancer antigens
- A61K39/001184—Cancer testis antigens, e.g. SSX, BAGE, GAGE or SAGE
- A61K39/001189—PRAME
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
- C07K14/4701—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals not used
- C07K14/4702—Regulators; Modulating activity
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
- C07K14/4701—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals not used
- C07K14/4748—Tumour specific antigens; Tumour rejection antigen precursors [TRAP], e.g. MAGE
Definitions
- peptides that are derived from the Preferentially Expressed Antigen in Melanoma (PRAME) protein, that can be used in immunotherapeutic methods.
- PRAME Preferentially Expressed Antigen in Melanoma
- the peptides are tumor-associated T cell peptide epitopes, that can serve as an active pharmaceutical ingredient of a vaccine composition that stimulates an anti-tumor immune response.
- the peptides disclosed herein can also be used to bind or stimulate T cells in vitro that may be transferred into a subject.
- Such a peptide bound to a molecule of the major histocompatibility complex (MHC), or a peptide as such can also be the target of an antibody, a soluble T cell receptor, bispecific T cell engager, and another binding molecule, e.g. a proteinaceous binding molecule.
- binding molecules can be used in immunotherapeutic methods, wherein soluble binding molecules can be administered directly to a subject to be treated, whereas membrane-bound binding molecules can be expressed in suitable cells that can be used for immunotherapy.
- peptide sequences derived from peptides from the PRAME protein which were isolated from complexes with MHC class I molecules of human tumor cells. Said peptide sequences can be used in vaccine compositions for eliciting anti-tumor immune responses, or as targets for the development of pharmaceutically / immunologically active compounds and cells.
- the presentation of peptides on almost any cell is to be distinguished from the presentation of peptides by specialized antigen-presenting cells, such as dendritic cells, via crosspresentation.
- dendritic cells specialized antigen-presenting cells
- these cells are also capable of taking up extracellular proteins and presenting fragments thereof on their cell surface.
- the present invention can be taken to generally be based on the identification of peptides that have been found on the extracellular surface of cancer cells, where they are bound to molecules of the major histocompatibility complex (MHC).
- MHC major histocompatibility complex
- MHC molecules possess a peptide-binding groove, within which a matching peptide can be bound.
- Different MHC variants have different amino acid residues at specific positions in the binding groove, which are called anchor residues (Murphy, 2022).
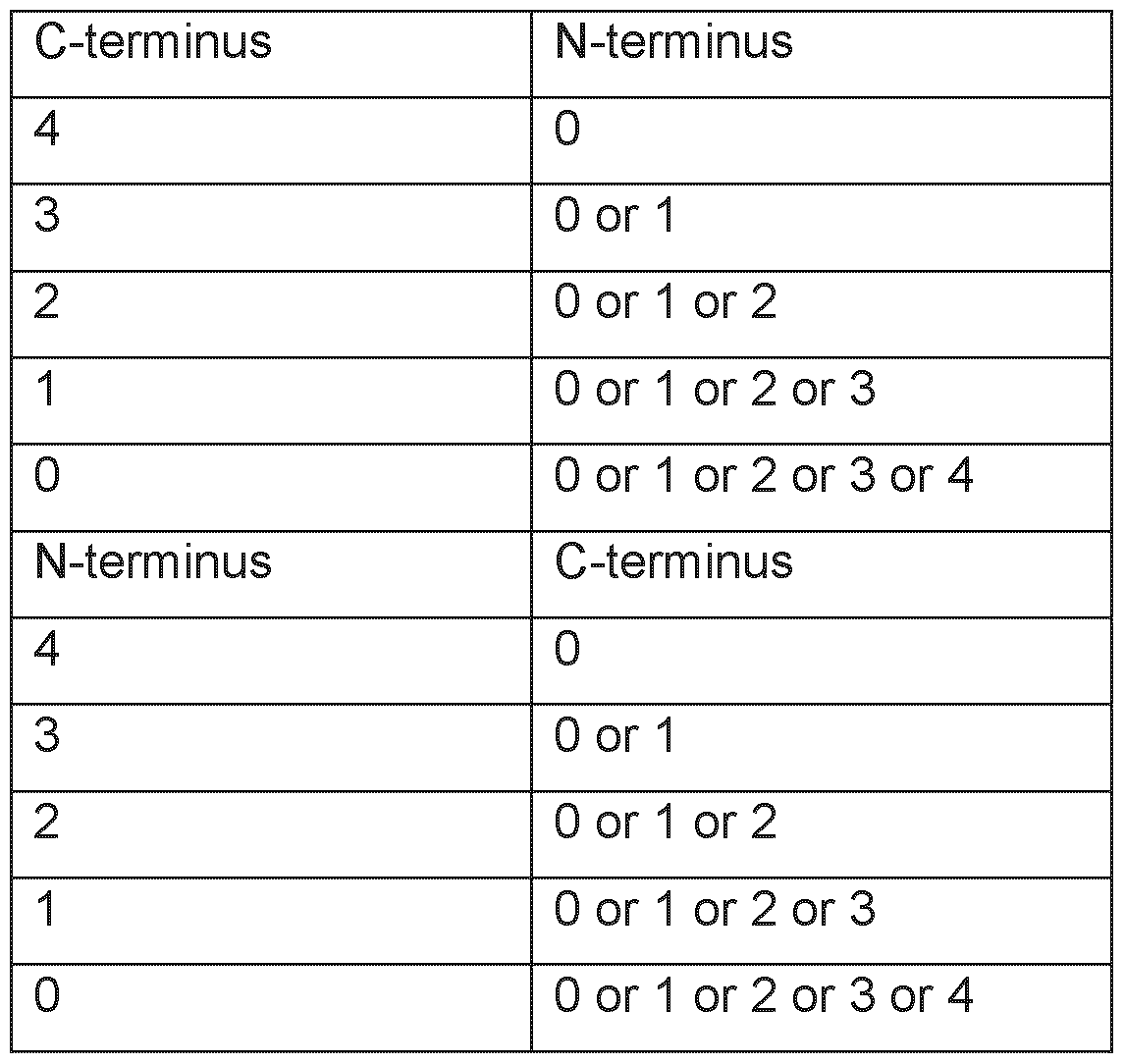
- a respective binding groove has two, in some cases three, pockets that undergo binding with amino acid side chains. Some of the bound peptides are too long to be encompassed in the cleft, which then typically protrude outside of the same (La Gruta, 2018).
- T cell receptors mediates the detection of cancerspecific peptides and the activation of T killer cells.
- Activated killer T cells can destroy antigen-presenting cells by perforins and granzymes.
- the T cell mediated immune response thus differs from the immunoglobulin mediated immune response. While antibodies can also recognize linear epitopes, they mainly recognize conformational epitopes exposed on the surfaces of proteins with tertiary structure. In contrast, CD8 + T cells recognize short linear peptide fragments, typically 8 to around 13 amino acids in length, which are presented in an extended conformation. MHC molecules can thus be taken to present a short linear epitope in a sterically fixed form (Murphy, 2022). The repertoire of T-cell receptor specificity may have both a genetic basis and a basis in selection in the thymus (La Gruta, 2018).
- Immunotherapy of cancer represents an option of specific targeting of cancer cells while minimizing side effects. Cancer immunotherapy makes use of the existence of tumor associated antigens.
- T cell-based immunotherapy targets peptide epitopes derived from tumor-associated or tumor-specific proteins, which are presented by MHC molecules.
- the antigens that are recognized by the tumor-specific T lymphocytes, that is, the epitopes thereof, can be molecules derived from all protein classes, such as enzymes, receptors, transcription factors, etc. which are expressed and, as compared to unaltered cells of the same origin, usually up-regulated in cells of the respective tumor.
- MHC class I There are two classes of MHC molecules, MHC class I and MHC class ii.
- MHC class I molecules are composed of an alpha heavy chain and beta-2-microglobulin
- MHC class ii molecules of an alpha and a beta chain. Their three-dimensional conformation results in a binding groove, which engages in non-covalent interactions with peptides.
- MHC class I molecules can be found on most nucleated cells. They present peptides that result from proteolytic cleavage of predominantly endogenous proteins, defective ribosomal products (DRIPs) and larger peptides. Also, peptides derived from endosomal compartments or exogenous sources are also frequently found on MHC class I molecules. This non-classical way of class I presentation is referred to as crosspresentation in the literature (Brossart and Bevan, 1997; Rock et al., 1990). MHC class ii molecules can be found predominantly on professional antigen-presenting cells (APCs), and primarily present peptides of exogenous or transmembrane proteins that are taken up by APCs e.g. during endocytosis and are subsequently processed.
- APCs professional antigen-presenting cells
- TCR T cell receptor
- CD4-positive helper T cells bearing the appropriate TCR. It is well known that the TCR, the peptide and the MHC are thereby present in a stoichiometric amount of 1 : 1 : 1 .
- MHC class I peptide For an MHC class I peptide to be able to trigger (elicit) a cellular immune response, it must first of all be bound to an MHC molecule. The binding is dependent on the allele of the MHC molecule and specific polymorphisms of the amino acid sequence of the peptide. MHC class I binding peptides are typically 8-13 amino acid residues in length and usually contain two conserved residues ("anchors") in their sequence that interact with the corresponding binding groove of the MHC molecule. In this way each MHC allele has a “binding motif” determining which peptides can bind specifically to the binding groove.
- MHC class I dependent immune reaction peptides bound to MHC class I molecules expressed by cells, e.g., tumor cells, can be recognized by T cells bearing specific TCRs which in turn initiate the immune reaction.
- peptides to be recognized by T-lymphocytes as tumor-specific or associated antigenic epitopes and to be used in a therapy, particular prerequisites must be fulfilled.
- the antigenic epitope should be over-presented on tumor cells in comparison to normal healthy cells.
- the expression level of the source protein of the peptide does not necessarily directly correlate with the presentation level of a peptide. It may be advantageous that a peptide is mainly displayed, or presented, on tumor cells and not, or in comparably small amounts, on normal healthy cells, for example cells of normal or healthy tissues.
- a peptide that is over-presented on the surface of a tumor cell can be derived from any protein found in the respective tumor cell.
- the therapeutic effect of the binder does not depend on the TCR repertoire of the person receiving said binding molecule. In these cases, the presentation is the determining factor.
- a peptide comprising an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59, or a pharmaceutically acceptable salt thereof, wherein the peptide has an overall length of up to 16 amino acids.
- the peptide, or the salt thereof provided herein has the ability to bind to an MHC class I molecule, and/or said peptide, when bound to said MHC, is capable of being recognized by CD8 T cells.
- the pharmaceutically acceptable salt provided herein is one of a chloride salt, a bromide salt, an iodide salt, a thiocyanate salt, a phosphate salt, a phosphonium salt, a nitrate salt, a sulfate salt, a chlorate salt, an acetate salt, a trifluoroacetate salt, a propionate salt, a butyrate salt, a pyridinium salt, a glycolate salt, a pyruvate salt, an oxalate salt, a malate salt, a maleate salt, a malonate salt, a succinate salt, a fumarate salt, a tartrate salt, a citrate salt, a benzoate salt, a cinnamate salt, a mandelate salt, a salicylic acid salt, a methane sulfonate salt, an ethane sulfonate salt, a p- toluen
- said peptide is part of a fusion protein.
- binding molecule that specifically recognizes a peptide comprising an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59, or said peptide comprising an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59 when bound to an MHC molecule.
- the binding molecule is typically a proteinaceous binding molecule or an aptamer.
- the binding molecule provided herein is an antibody, or a functional fragment thereof.
- the antibody is generally a soluble or a membrane-bound antibody.
- the binding molecule is a T cell receptor (TCR), or a functional fragment thereof.
- the TCR is generally a soluble TCR or a membrane-bound TCR.
- the antibody or functional fragment thereof may in some embodiments be a monoclonal antibody, a bi-specific antibody, a chimeric antibody, a human antibody and/or a humanized antibody or a fragment thereof.
- the binding molecule is an aptamer.
- the binding molecule may be a T cell engaging bispecific molecule, such as a bispecific molecule comprising a TCR domain and a T cell recruiting antibody domain.
- nucleic acid comprising a sequence encoding the peptide or the binding molecule provided herein, being a proteinaceous binding molecule.
- sequence is operably linked to a heterologous promoter sequence.
- an expression vector comprising the nucleic acid provided herein.
- a recombinant host cell comprising the peptide, the binding molecule and/or the nucleic acid provided herein.
- T lymphocytes that specifically recognize a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule.
- the method comprises (a) contacting in vitro T lymphocytes with antigen loaded human class I MHC molecules expressed on the surface of a suitable antigen-presenting cell or an artificial construct mimicking an antigen-presenting cell for a period of time sufficient to activate the T cells in an antigen specific manner, wherein said antigen is the peptide provided herein or (b) introducing in vitro into T lymphocytes a nucleic acid encoding a TCR that specifically recognizes a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule.
- T lymphocyte produced by the in vitro method for producing T lymphocytes provided herein, wherein said T lymphocyte specifically recognizes a cell which presents a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59.
- the T lymphocyte is an activated T lymphocyte.
- a pharmaceutical composition comprising one or more active ingredients as provided herein selected from the group consisting of a peptide, a pharmaceutically acceptable salt of the peptide, the binding molecule, the nucleic acid, the expression vector, the recombinant host cell, or the T lymphocyte, and a pharmaceutically acceptable carrier or a pharmaceutically acceptable excipient.
- the pharmaceutical composition provided herein is a vaccine.
- the pharmaceutical composition provided herein further comprises an adjuvant.
- a combination product comprising one or more of the peptide or salt thereof, the binding molecule, the nucleic acid, the expression vector or the recombinant host cell or the T lymphocyte as provided herein.
- the combination product comprises an adjuvant.
- the adjuvant of the pharmaceutical composition or of the combination product provided herein comprises IFN-alpha, IFN-beta or an interleukin, wherein the interleukin preferably is IL-1 , IL-2, IL-4, IL-7, IL-9, IL-10, IL-12, IL-13, IL-15, IL-17, IL-18, IL-21 , IL-23 or a combination thereof.
- the adjuvant of the pharmaceutical composition or of the combination product provided herein is IL-2, IL-12, IL-15, IL-18, IL-21 or a combination thereof.
- the pharmaceutical composition or the combination product provided herein comprises the T lymphocyte provided herein and an adjuvant selected from an interleukin.
- the interleukin is IL-2, IL-15 or IL-21 or a combination thereof.
- the binding molecule is a proteinaceous binding molecule.
- the method includes culturing the recombinant host cell provided herein and isolating the peptide or the proteinaceous binding molecule from the host cell and/or its culture medium.
- the peptide or the salt thereof is provided herein.
- the use is in the prevention, treatment and/or diagnosis of cancer.
- the cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, nonHodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer and another cancer that shows an over-presentation of a peptide provided herein.
- peptide or the salt thereof the binding molecule, the nucleic acid, the expression vector, the recombinant host cell, the T lymphocyte, the pharmaceutical composition or the combination product as disclosed herein in the manufacture of a medicament for the prevention, treatment and/or diagnosis of cancer.
- said cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer, and another cancer that shows an over-presentation of a peptide provided herein.
- the target cells present a peptide provided herein.
- the method comprises administering to the subject an effective number of T lymphocytes provided herein.
- the target cells are known or suspected of being cancer cells.
- the method includes contacting said cells with a plurality of T lymphocytes as provided herein.
- the cells are cells of a sample from a subject.
- the T lymphocyte provided herein is for use in the killing of target cells in a subject.
- the target cells present a peptide disclosed herein.
- the T lymphocyte provided herein is for use in the manufacture of a medicament for the killing of such target cells that present a peptide disclosed herein.
- the T lymphocyte is for use in the prevention or treatment of cancer.
- the use encompasses the killing of target cancer cells in a subject.
- the target cancer cells present a peptide disclosed herein.
- a method of treating a subject comprising administering to the subject an effective amount of the peptide, the binding molecule, the nucleic acid, the expression vector, the recombinant host cell, the T lymphocyte, the pharmaceutical composition or the combination product provided herein.
- said cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, nonHodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer or another cancer that shows an over-presentation of a peptide provided herein.
- kits includes (a) a container comprising the peptide or the salt thereof, the binding molecule, the nucleic acid, the expression vector, the recombinant host cell, the T lymphocyte, the pharmaceutical composition, and/or a combination product provided herein in solution or in lyophilized form.
- the kit includes (b) a further container containing a diluent or reconstituting solution for the lyophilized formulation.
- the kit includes (c) a further container containing an adjuvant.
- the kit includes both the container containing a diluent or reconstituting solution and a further container containing an adjuvant.
- the kit includes (d) an additional peptide that contains a sequence set forth in any one of SEQ ID NOs: 1 to 59 provided herein.
- the kit includes a container containing a diluent or reconstituting solution and an additional peptide that contains a sequence set forth in any one of SEQ ID NOs: 1 to 59.
- the kit includes the container containing a diluent or reconstituting solution, a further container containing an adjuvant and an additional peptide that contains a sequence set forth in any one of SEQ ID NOs: 1 to 59.
- the kit provided herein further comprises one or more of a buffer, a diluent, a filter, a needle, and/or a syringe.
- an in vitro method of diagnosing cancer in a sample comprising cells of an individual comprises (a) adding a binding molecule to a sample from the subject.
- the binding molecule is labeled.
- the label is a radionucleotide
- the binding molecule specifically binds to (i) a peptide comprising the amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 or (ii) said peptide comprising the amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule.
- the sample comprises cells of a tissue suspected to be a cancer tissue.
- an in vitro method of diagnosing cancer in a sample comprising cells of an individual comprises (a) identifying one or more peptides presented on cells comprised in a tumor sample from the individual; and (b) comparing the peptide(s) as identified in step (a) with a collection of peptides that have been pre-screened for immunogenicity and/or over-presentation in tumours as compared to normal tissues.
- the collection of peptides comprises one or more peptides comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, having a maximal length of 16 amino acids.
- the method also comprises (c) assessing whether a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 is a peptide comprised in a tumor sample from the individual.
- the sample comprises cells of a tissue suspected to be a cancer tissue.
- the cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer, or another cancer that shows an over-presentation of a peptide provided herein.
- the method comprises determining whether a peptide comprising the amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 is presented on cells in a sample from a subject suspected of having cancer.
- the sample comprises cancer cells from said subject.
- the method also comprises contacting in vitro a plurality of T lymphocytes from said subject with an antigen-presenting cell or an artificial antigen-presenting cell having antigen-loaded human class I MHC molecules on its surface.
- the antigen is a peptide comprising an amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59.
- the method includes activating said T lymphocytes in an antigen specific manner if they bind to said peptide.
- the method includes allowing proliferation of the obtained activated T lymphocytes.
- the method also includes isolating one or more activated T lymphocytees for manufacturing an individualized anti-cancer cellular therapeutic composition.
- manufacturing the individualized anti-cancer cellular therapeutic composition further comprises cloning the nucleic acid encoding the TCR expressed by one or more activated T lymphocytes into a suitable expression vector and introducing the vector into T lymphocytes.
- determining whether said peptide is presented on cells in a sample from the subject comprises adding a binding molecule to a sample from said subject. This binding molecule is (a) labeled and (b) specifically binds to said peptide or to said peptide when bound to an MHC molecule.
- the method includes (i) assessing whether cells of a cancer sample from the subject present a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59.
- the method also includes (ii) selecting the subject for treatment with a vaccine and/or an immunotherapeutic agent if the peptide that includes the sequence set forth in any one of SEQ ID NOs: 1 to 59 is presented on cells of the cancer sample.
- Said vaccine includes a peptide that includes a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a pharmaceutically acceptable salt thereof.
- the immunotherapeutic agent includes at least one of the binding molecule, the nucleic acid, the expression vector, the recombinant host cell or the T lymphocyte provided herein.
- the method of stratifying a subject further includes assessing whether the peptide that includes a sequence set forth in any one of SEQ ID NOs: 1 to 59 is bound by an MHC molecule.
- the anti-cancer vaccine includes a peptide that includes a sequence set forth in any one of SEQ ID NOs: 1 to 59.
- the compound-based and/or cellular therapeutic agent is based on a peptide that comprises a sequence set forth in any one of SEQ ID NOs: 1 to 59.
- the method includes (a) identifying one or more peptides presented on cells in a tumor sample from the individual; (b) comparing the peptide(s) as identified in (a) with a collection of peptides that have been pre-screened for immunogenicity and/or overpresentation in tumours as compared to normal tissues.
- the collection of peptides includes one or more peptides that include a sequence set forth in any one of SEQ ID NOs: 1 to 59, having a maximal length of 16 amino acids.
- the method also includes (c) selecting one or more peptides from the collection of peptides if it/they match(es) the peptide(s) identified in the subject.
- the selected peptide(s) is/are one or more peptide(s) comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, having a maximal length of 16 amino acids comprised in the collection of peptides.
- the method also includes (d) manufacturing and/or formulating the individualized vaccine or compoundbased and/or cellular therapeutic agent based on the selection in step (c).
- identifying said peptide(s) involves (a1 ) comparing expression data from the tumor sample to expression data from a sample of normal tissue corresponding to the tissue type of the tumor sample to identify proteins that are over-expressed or aberrantly expressed in the tumour sample. Identifying said peptide(s) also involves (a2) correlating the expression data with sequences of MHC ligands bound to MHC class I molecules in the tumor sample to identify MHC ligands derived from proteins over-expressed or aberrantly expressed by the tumor.
- the sequences of MHC ligands are identified by eluting bound peptides from MHC molecules isolated from the tumor sample and sequencing the eluted ligands; and/or the normal tissue corresponding to the tissue type of the tumor sample is obtained from the same individual; and/or the immunogenicity of the peptides included in the collection of peptides is determined by a method comprising an in vitro immunogenicity assay, subject immunomonitoring for individual HLA binding, MHC multimer staining, an ELISPOT assay and/or intracellular cytokine staining.
- a method as defined above further comprises identifying a mutation that is unique to the tumor sample relative to normal corresponding tissue from the individual, and selecting a peptide that correlates with the mutation for inclusion in the vaccine or for the generation of a cellular therapeutic, wherein the mutation is optionally identified by whole genome sequencing.
- the one or more peptides included in the collection of peptides is/are identified based on the following steps (aa) performing genome-wide messenger ribonucleic acid (mRNA) expression analysis by highly parallel methods, such as microarrays or sequencing-based expression profiling, comprising identifying genes that are over-expressed in a malignant tissue, compared to a normal tissue or tissues; (ab) selecting peptides encoded by selectively expressed or overexpressed genes as detected in step (aa), and (ac) determining an induction of an in vivo T-cell response by the peptides as selected comprising in vitro immunogenicity assays using human T cells from healthy donors or the individual; or
- identifying MHC ligands e.g. HLA ligands
- mRNA messenger ribonucleic acid
- highly parallel methods such as microarrays or sequencingbased expression profiling, comprising identifying genes that are over-expressed in a malignant tissue, compared with a normal tissue or tissues;
- comparing the identified MHC ligands to the gene expression data comprising selecting peptides encoded by selectively expressed or over-expressed genes as detected in step be;
- determining an induction of an in vivo T-cell response by the peptides as selected comprising an in vitro immunogenicity assay using human T cells from a healthy donor or the individual.
- a peptide consisting of an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59, or a pharmaceutically acceptable salt thereof.
- the peptide has an overall length of up to 13 amino acids, including of up to 9 amino acids.
- a peptide comprising, an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59, or a pharmaceutically acceptable salt thereof.
- the peptide has an overall length of up to 30 amino acids, including of up to 16 amino acids.
- the peptide defines an epitope and its sequence can be identified in the PRAME protein amino acid sequence.
- nucleic acid vaccine that comprises a region encoding a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
- the nucleic acid vaccine may comprise a concatemer that contains multiple copies of a nucleic acid sequence encoding a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
- a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59 may be used as an epitope in a protein vaccine. Accordingly, provided herein is furthermore a protein vaccine comprising a section consisting of a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59. Said protein vaccine may comprise several repetitions of a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
- the peptide as provided herein has the ability to bind to an MHC molecule class I and/or MHC molecule class II, such as human leukocyte antigens (HLA).
- MHC molecule class I and/or MHC molecule class II such as human leukocyte antigens (HLA).
- HLA human leukocyte antigens
- an elongated version of the peptide has the ability to bind to an MHC molecule class II.
- a respective peptide, when bound to the MHC is capable of being recognized by CD4 and/or CD8 T cells.
- a respective MHC class I molecule is an HLA allotype MHC molecule specified further below.
- the peptide consists of an amino acid sequence according to SEQ ID NO: 1 to SEQ ID NO: 59.
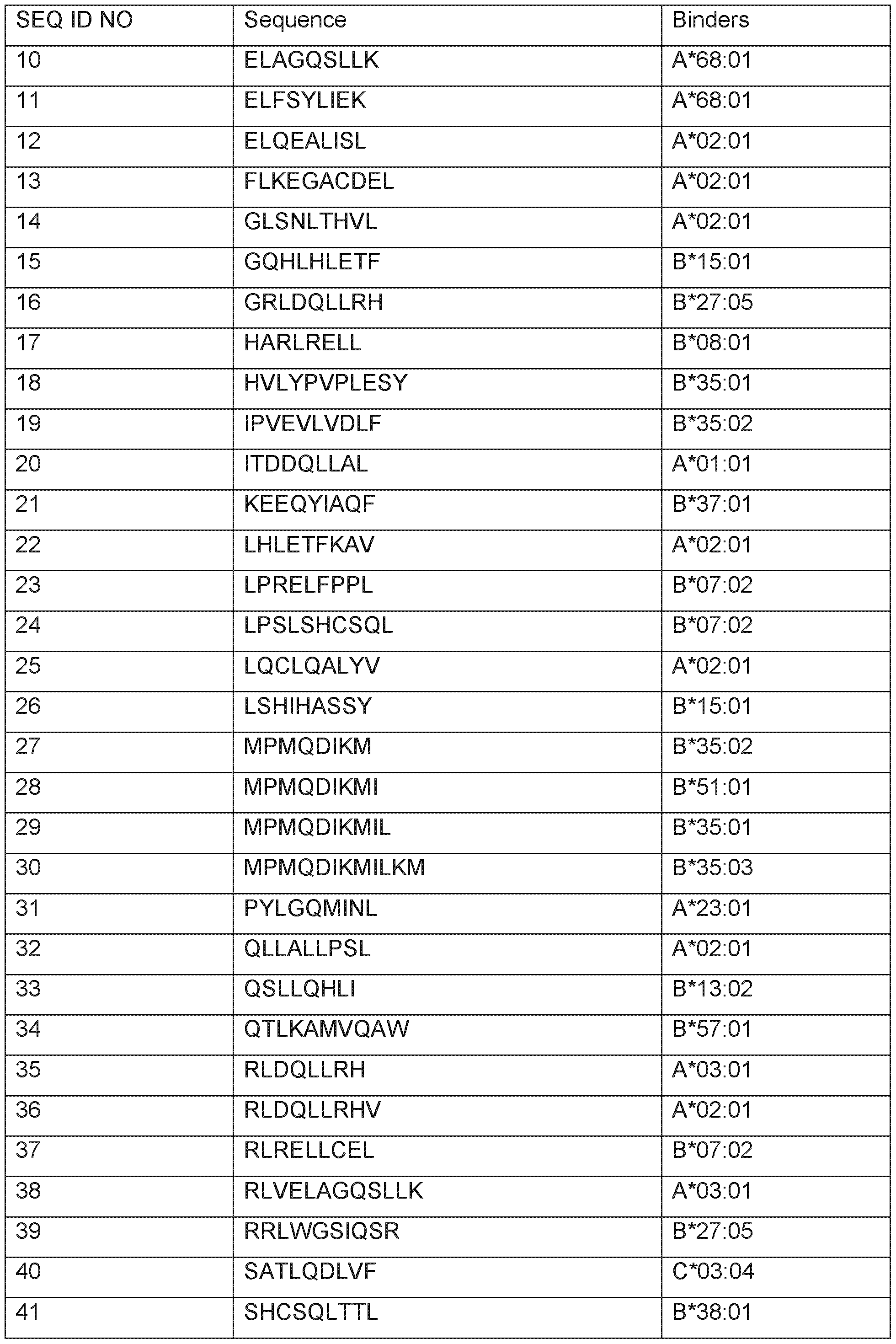
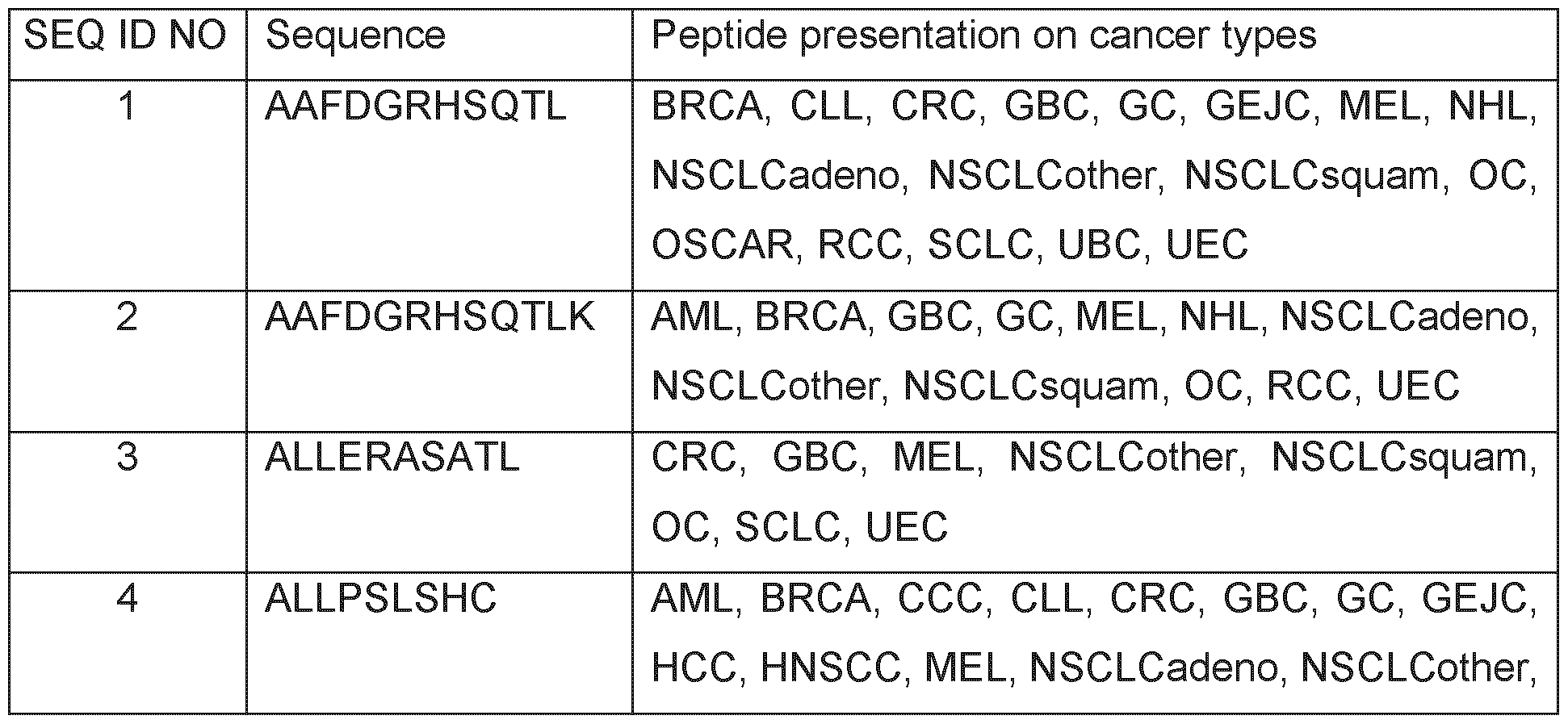
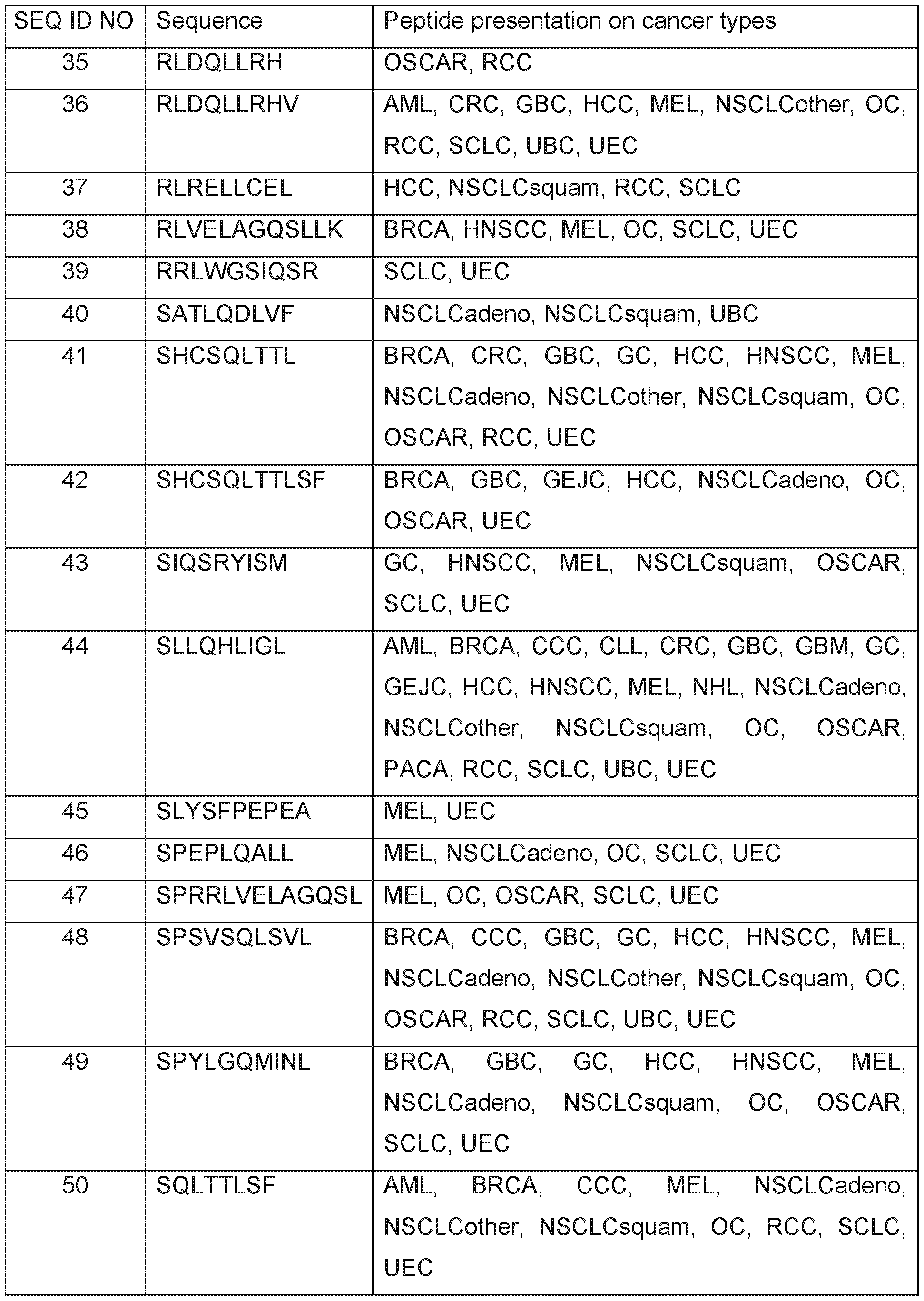
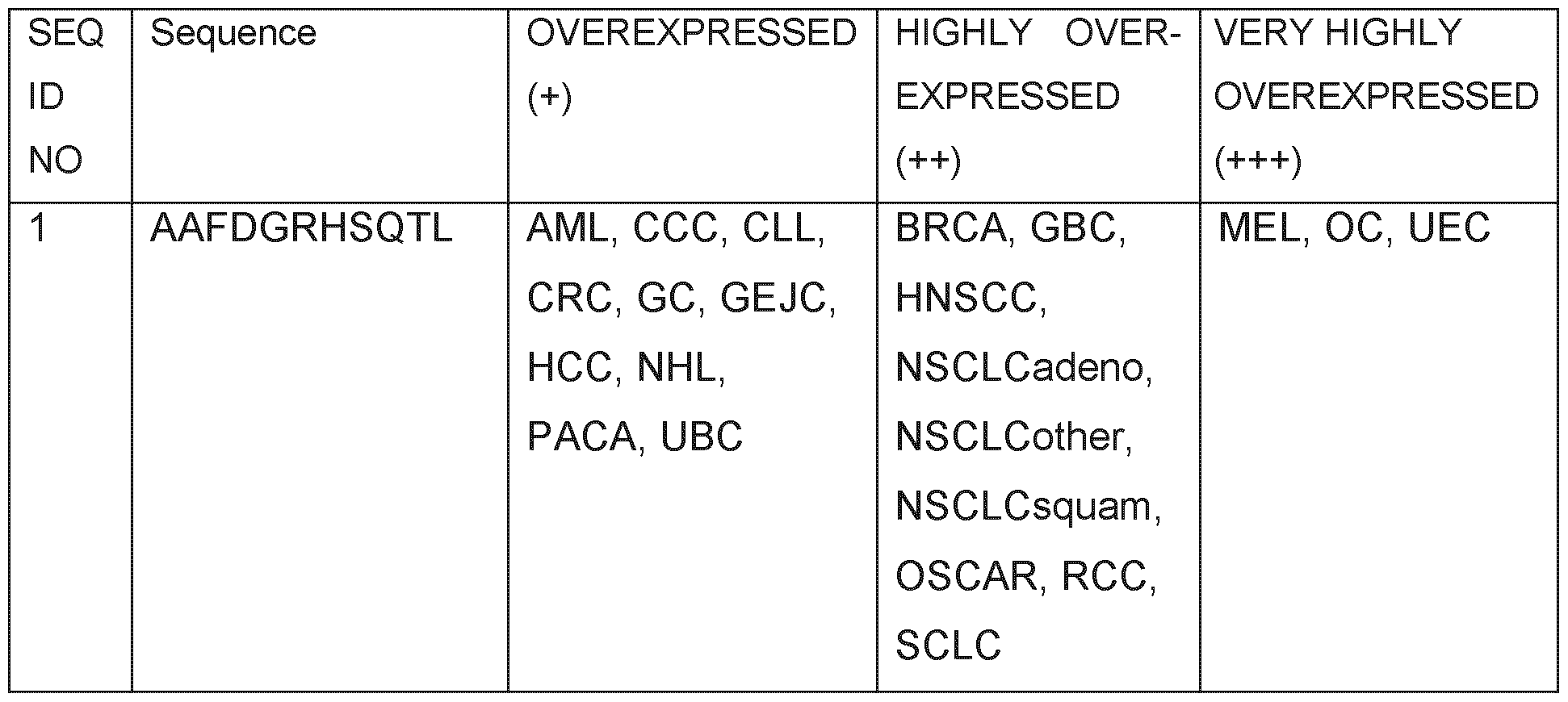
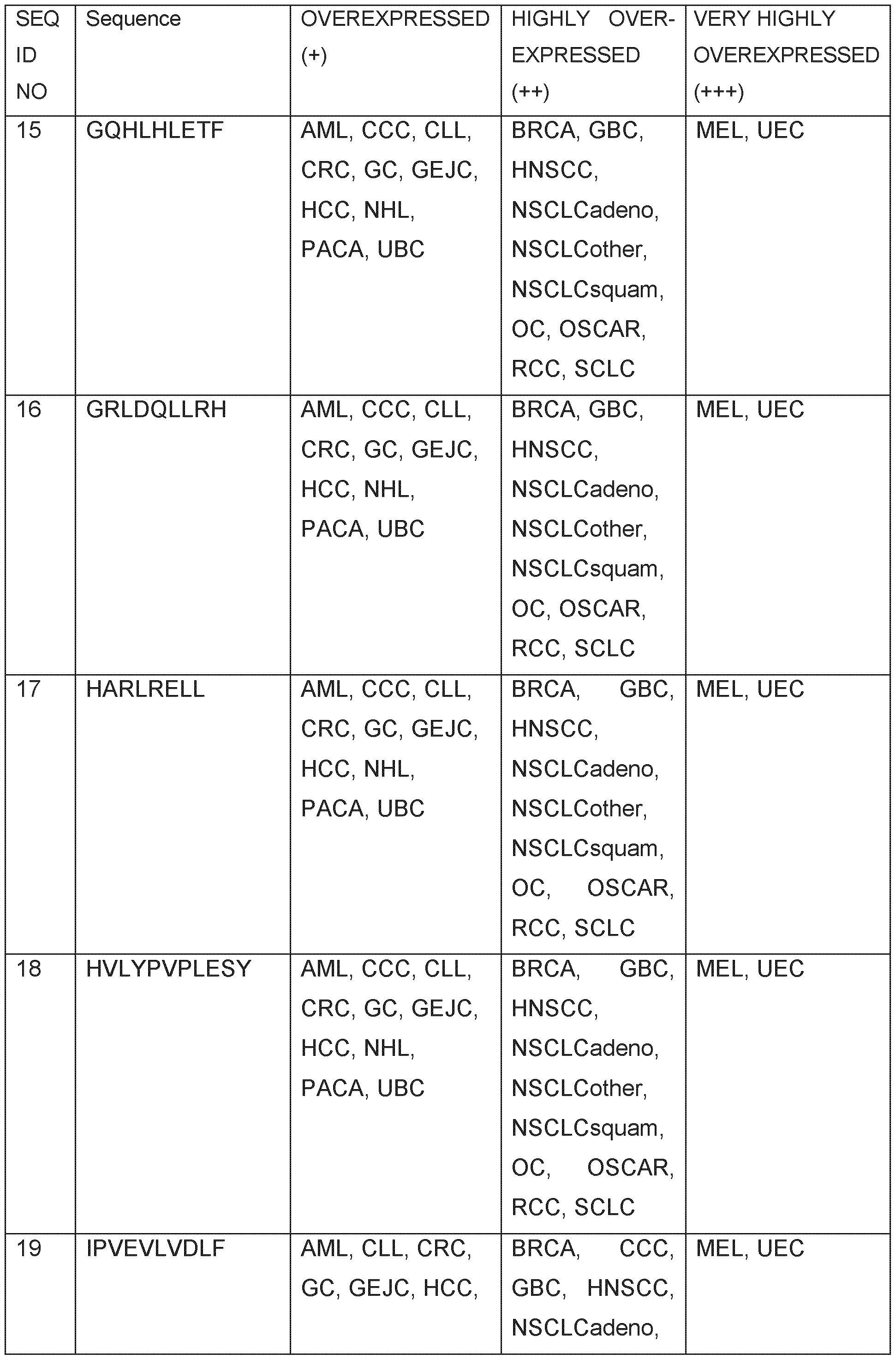
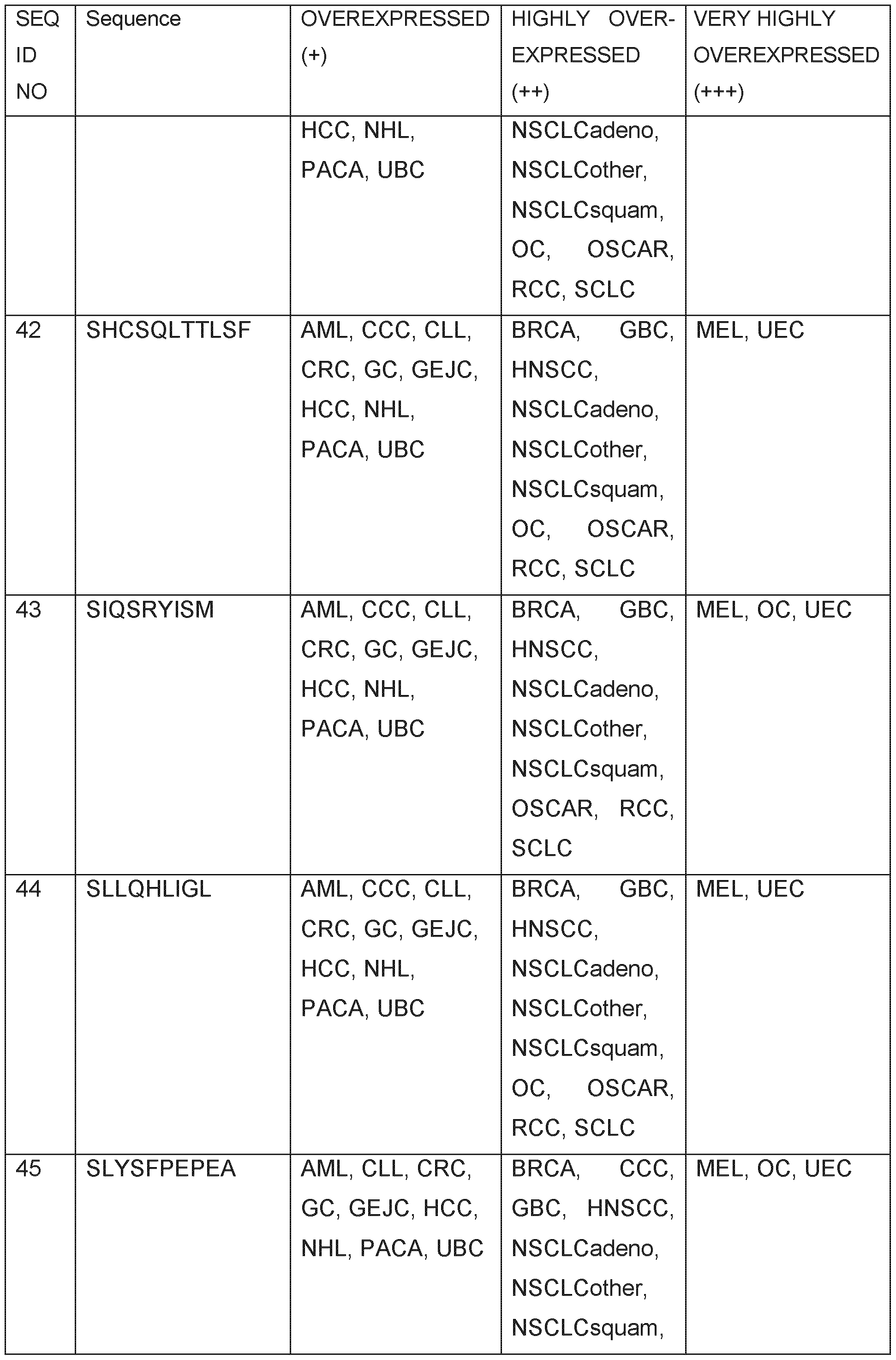
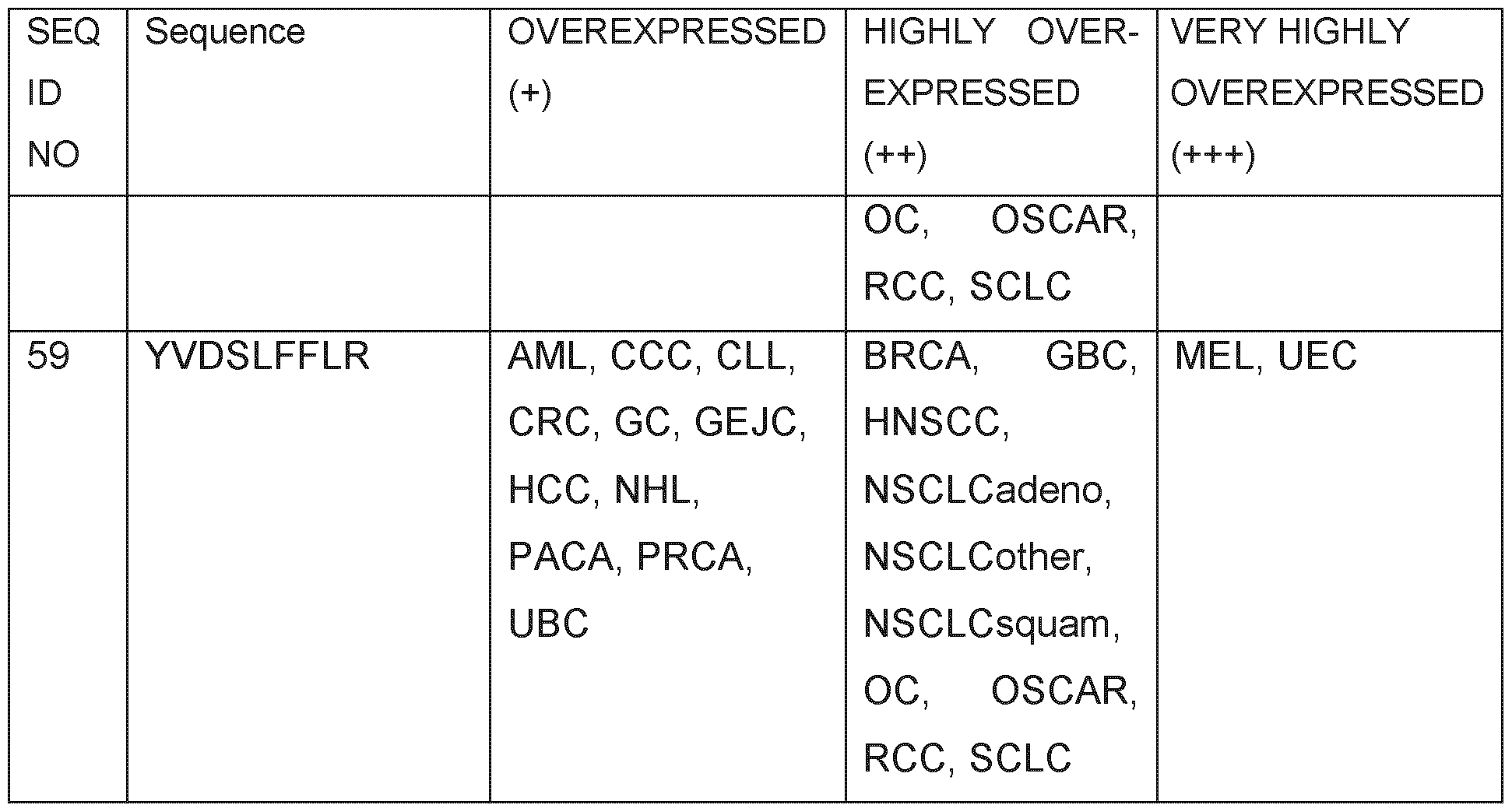
- Table 1 shows the peptides with their amino acid sequences disclosed herein and their respective SEQ ID NOs.
- Table 1 Peptides provided and their respective SEQ ID NO.
- a peptide provided herein has the ability to bind to components of the major histocompatibility complex (MHC), in the case of humans also known as human leukocyte antigens (HLAs).
- MHC major histocompatibility complex
- HLAs human leukocyte antigens
- the MHC is an MHC class I MHC molecule.
- a peptide as disclosed herein is a synthetic peptide.
- a peptide as disclosed herein is provided as a chemically synthesized peptide.
- the peptide may for example have been synthesized by means of solid phase synthesis.
- a peptide as disclosed herein is provided as a recombinant peptide.
- the peptide may for example have been secreted by a recombinant host cell expressing the peptide.
- the peptide may have been isolated from the media encompassing the cells.
- the cells can be sedimented, and the supernatant be collected for isolating the peptide.
- the peptide may in some embodiments have been expressed in a recombinant host cell without being secreted. In such embodiments the peptide may have been isolated from the cells.
- a peptide may for example be provided as a salt that has an inorganic counter ion.
- a peptide may be provided as a halogen salt such as a chloride.
- a peptide as disclosed herein may in some embodiments be provided as a salt that has an organic compound, such as a carboxylic acid containing compound, as a counter ion.
- a peptide may be provided as an acetate salt or a trifluoroacetate salt.
- a peptide as disclosed herein includes one or more non-peptide bonds. In some embodiments a peptide as disclosed herein includes an isopeptide bond.
- a peptide as disclosed herein is modified, such as chemically modified.
- the peptide may for example contain one or more modified side chains. It may for example include one or more modified functional groups.
- a peptide as disclosed herein contains one or more amino acid side chains modified by acylation or amidation.
- a peptide as disclosed herein contains one or more amino acid side chains modified by the formation of an ether or an ester.
- a peptide as disclosed herein contains one or more amino acid side chains modified by the formation of a thioether or a thioester.
- a peptide as disclosed herein contains one or more lysine side chains modified by pyridoxylation.
- a peptide as disclosed herein contains one or more amino acid side chains modified by reductive alkylation or by carboxymethylation. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by trinitrobenzylation of amino groups or amide modification of carboxyl groups. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by oxidation of cysteine to cysteic acid or it contains mixed disulphides with other thiol compounds.
- a peptide as disclosed herein contains one or more amino acid side chains modified in the form of an adduct of arginine with phenylglyoxal, 2,3-butanedione or in the form of a mercurial derivative. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by reaction with maleimide or by carbamoylation. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by an intra-molecular crosslink between lysine and a glutamic acid.
- a peptide as disclosed herein contains one or more amino acid side chains modified in the form of carbethoxylated histidine obtained with diethyl pyrocarbonate. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by 1 ,2-cyclohexanedione or by methionine thioether formation using iodoacetamide. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by bromoethylamine or by methionine oxidation with chloramine T.
- a peptide as disclosed herein contains one or more amino acid side chains modified by reaction with tyrosine tetranitromethane or N- acetyl imidazole. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by reaction with N-bromosuccinimide or modified with PEG. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by reaction with 2-hydroxy-5-nitrobenzyl bromide or 3- bromo-3-methyl-2-(2-nitrophenylmercapto)-3H-indole (BPNS-skatole).
- a peptide as disclosed herein is part of a fusion protein, for example fused to the N-terminal amino acids of the HLA-DR antigen-associated invariant chain (li) or fused to (or into the sequence of) an antibody, such as, for example, an antibody that is specific for dendritic cells.
- Binding Molecules in a further aspect there is provided a binding molecule that specifically binds to a peptide as provided herein, or that specifically binds to the respective peptide when in a complex with an MHC molecule.
- the binding molecule may be a proteinaceous binding molecule.
- the binding molecule may be an antibody or a functional fragment thereof.
- the binding molecule may also be a T cell receptor (TCR) or a functional fragment thereof.
- TCR T cell receptor
- the binding molecule may also be a proteinaceous binding molecule with antibody-like functions.
- a binding molecule as provided herein contains a detectable moiety such as a detectable marker.
- a respective detectable moiety may in some embodiments be or include a fluorescent or luminescent portion.
- a respective detectable moiety may in some embodiments be or include a radiolabel or an enzyme capable of producing a detectable product.
- a respective detectable moiety may in some embodiments be or include a magnetic resonance imaging or a computer tomography X- ray contrast agent.
- a binding molecule provided herein e.g. an antibody a TCR, or an aptamer, carries a further effector function such as an immune stimulating domain or toxin.
- a binding molecule provided herein is coupled to a detectable label.
- a binding molecule such as a proteinaceous binding molecule, provided herein is coupled to an active agent such as a toxin.
- a binding molecule provided herein is coupled to a protein kinase modifying moiety.
- an immunotoxin contains a binding molecule provided herein, e.g. an antibody, a TCR or an aptamer, which is coupled to a toxin.
- a binding molecule provided herein, e.g. an antibody, a TCR or an aptamer, which is coupled to a toxin.
- an immunotoxin may be defined by a monoclonal antibody, or a functional monoclonal antibody fragment, that specifically binds to a peptide disclosed herein, covalently bonded to a toxin.
- the respective toxin may for instance be a truncated bacterial toxin.
- a respective antibody or functional fragment thereof may be a soluble antibody or binding fragment thereof.
- a respective antibody or functional fragment thereof may be a membrane-bound antibody or binding fragment thereof.
- the antibody or functional fragment thereof may be a monoclonal antibody or binding fragment thereof.
- the antibody or functional fragment thereof may be a human or a humanized antibody or binding fragment thereof.
- the antibody or functional fragment thereof may be a bi-, tri- or multi-specific antibody or binding fragment thereof.
- the antibody or functional fragment thereof may be a chimeric antibody or binding fragment thereof.
- a respective antibody or fragment thereof may in some embodiments carry a further effector function.
- the antibody or fragment thereof may for example be fused or covalently linked to an immune stimulating domain such as a cytokine (e.g. an inflammatory cytokine, as illustrated in Table 3), or toxin.
- a cytokine e.g. an inflammatory cytokine, as illustrated in Table 3
- toxin e.g. toxin
- a respective antibody may be a monoclonal antibody and/or a bi-specific antibody or a functional fragment thereof.
- a respective antibody may be a chimeric antibody or a functional fragment thereof.
- a respective antibody may also be TCR like antibody.
- a respective antibody or functional fragment thereof may in some embodiments carry a further effector function.
- the antibody or functional fragment thereof may for example be fused or covalently linked to an immune stimulating domain such as a cytokine (e.g. an inflammatory cytokine, as illustrated in Table 3), or toxin.
- the antibody or functional fragment thereof may also be fused or covalently linked to an antibody recognizing a T cell structure, such as an anti CD3 antibody.
- a respective T cell receptor or functional fragment thereof may be soluble or membranebound.
- a respective T cell receptor or functional T cell receptor functional fragment may be a monoclonal T cell receptor or T cell receptor functional fragment.
- a respective T cell receptor or functional fragment thereof may in some embodiments carry a further effector function.
- the T cell receptor or functional fragment thereof may for example be fused or covalently linked to an immune stimulating domain such as a cytokine (e.g. an inflammatory cytokine, as illustrated in Table 3), or toxin.
- the T cell receptor or functional fragment thereof may also be fused or covalently linked to an antibody recognizing a T cell structure, such as an anti CD3 antibody.
- Such a functional TCR fragment provided herein is an antigen-binding TCR fragment, capable of binding to a peptide as disclosed herein.
- a functional TCR fragment is an extracellular TCR fragment.
- a functional TCR fragment is a single chain TCR (scTCR).
- a single chain TCR may include an alpha chain and a beta chain, linked by a linker sequence.
- a TCR provided herein is a dimeric TCR (dTCR).
- the TCR provided herein may be a T cell engaging bispecific molecule, such as a bispecific molecule comprising a TCR domain and a T cell recruiting antibody domain.
- the TCR domain may comprise the alpha and beta variable regions of a TCR capable of binding to a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
- the T cell recruiting antibody may bind to CD3 and/or to the alpha/beta TCR/CD3 complex.
- the method may be an in vitro method.
- the method may involve introducing in vitro a nucleic acid encoding a TCR that specifically recognizes a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule into T lymphocytes.
- the method may additionally or alternatively involve contacting in vitro a T lymphocyte, e.g. a CTL or Th cell, with antigen-loaded human class I or class II MHC molecules expressed on the surface of a suitable antigen-presenting cell for a period of time sufficient to activate the T lymphocyte in an antigen specific manner.
- the respective antigen is the peptide defined above.
- the method may furthermore include providing in vitro a T lymphocyte and providing antigen-loaded human class I MHC molecules that are expressed on the surface of a suitable antigen-presenting cell.
- the method may furthermore include isolating the activated T lymphocytes.
- the antigen is loaded onto class I or II MHC molecules expressed on the surface of a suitable antigen-presenting cell or artificial antigen-presenting cell by contacting a sufficient amount of the antigen with an antigen-presenting cell.
- the antigen-presenting cell contains an expression vector capable of expressing a peptide as disclosed herein.
- T lymphocyte that specifically recognizes a cell which presents a peptide as disclosed herein.
- the respective presentation may be an over-presentation.
- the over-presentation may be based on aberrant expression.
- the over-presentation is not based on an aberrant expression.
- the T lymphocyte is an activated T lymphocyte that specifically recognizes the peptide on the cell which presents the same, when the peptide is bound by MHC.
- the (activated) T lymphocyte may be a cytotoxic T lymphocyte.
- the T lymphocytes selectively recognize a cell which presents a peptide that contains an amino acid sequence as defined herein.
- the T lymphocytes may selectively recognize a cell which presents a peptide as provided herein.
- the respective presentation may be an over-presentation.
- the over-presentation may be based on aberrant expression.
- the over-presentation is not based on aberrant expression.
- the T lymphocyte is an activated T lymphocyte that specifically recognizes the peptide on the cell which presents the same, when the peptide is bound by MHC.
- the (activated) T lymphocytes may be an alpha beta T lymphocyte or a gamma delta T lymphocyte.
- the (activated) T lymphocyte may be a CD8+ T lymphocyte or CD4+ T lymphocyte.
- the (activated) T lymphocyte may be a cytotoxic T lymphocyte.
- T lymphocyte that includes a T-cell receptor that specifically binds to a peptide as disclosed herein, when presented by an MHC molecule.
- the T- cell receptor specifically binds to a peptide as disclosed herein, when presented by an MHC molecule on the surface of a cell.
- the T lymphocyte is an activated T lymphocyte that specifically recognizes the peptide on the cell which presents the same, when the peptide is bound by MHC.
- the (activated) T lymphocyte may be an alpha beta T lymphocyte or a gamma delta T lymphocyte.
- the (activated) T lymphocyte may be a CD8+ T lymphocyte or CD4+ T lymphocyte.
- the (activated) T lymphocyte may be a cytotoxic T lymphocyte (CTL).
- CTL cytotoxic T lymphocyte
- the T lymphocytes are activated T lymphocytes that specifically recognizes the peptide on the cell which presents the same, when the peptide is bound by MHC.
- the (activated) T lymphocytes may be alpha beta T lymphocytes or gamma delta T lymphocytes.
- the (activated) T lymphocytes may be CD8+ T lymphocytes or CD4+ T lymphocytes.
- the plurality of (activated) T lymphocytes define a population of T lymphocytes.
- the plurality of (activated) T lymphocytes is included in or defines a cell line, such as a human cell line.
- an in vitro method of screening a TCR that specifically binds to a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59 includes contacting in vitro a pool of T cells (e.g. derived from the blood of a human donor), with antigen-presenting cells that express on the surface MHC molecules that are loaded with a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
- the method further involves selecting activated T cells.
- the method may furthermore include isolating the activated T cells.
- the method may further include isolating the TCR from the activated T cells.
- the method includes labelling at least two different antigens with different markers (e.g. fluorescence markers):
- the first antigen is a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
- the second antigen is (i) a peptide that is similar to the first antigen, or (ii) several different peptides that are similar to the first antigen.
- T cells that bind to the first antigen but not or less to the second antigen are selected as T cells that specifically bind to a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
- 2D-color tetramer combinations may be used to stain T cells that bind to the first antigen, T cells that bind to the second antigen and T cells that bind to the first and the second antigen.
- a method is described, e.g. in WO 2020/245326, the content of which is incorporated by reference in its entirety.
- Cell sorting may be used to exclusively obtain those T cells that only bind the first antigen (but not or less the second antigen).
- the method may further comprise isolating the TCR from said T cells that only bind to the first antigen (but not or less the second antigen).
- a T lymphocyte expressing a TCR that is specifically binding to a peptide as disclosed herein, typically when in a complex with an MHC molecule.
- the T lymphocyte is a T cell line or a cell of a T cell line.
- the T lymphocyte is part of a T cell population.
- the T lymphocyte is in some embodiments an alpha beta T lymphocyte.
- the T lymphocyte is in some embodiments a CD8+ T lymphocyte.
- the T lymphocyte is in some embodiments a CD4+ T lymphocyte.
- the T lymphocyte is in some embodiments a gamma delta T lymphocyte.
- the T lymphocyte is in some embodiments genetically modified to express a TCR that specifically recognizes a peptide comprising an amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59, typically when in a complex with an MHC molecule.
- T lymphocytes expressing a T-cell receptor (TCR) that specifically binds to a peptide as disclosed herein when in a complex with an MHC molecule.
- the T lymphocytes are in some embodiments alpha beta T lymphocytes.
- the T lymphocytes are in some embodiments CD8+ T lymphocytes or CD4+ T lymphocytes.
- the T lymphocytes are in some embodiments gamma delta T lymphocytes.
- the plurality of T lymphocytes defines a population of T lymphocytes.
- the plurality of T lymphocytes is included in or defines a cell line, such as a human cell line.
- the nucleic acid encodes a T-cell receptor reactive with an MHC ligand.
- the cells of the plurality of T lymphocytes express a T-cell receptor that specifically binds to the MHC ligand.
- the MHC ligand is a peptide as disclosed herein.
- the plurality of T lymphocytes is transduced by a lentivirus or retrovirus that includes the nucleic acid encoding the T-cell receptor.
- the T lymphocytes are in some embodiments alpha beta T lymphocytes.
- the T lymphocytes are in some embodiments CD8+ T lymphocytes or CD4+ T lymphocytes.
- the T lymphocytes are in some embodiments gamma delta T lymphocytes.
- the plurality of T lymphocytes defines a population of T lymphocytes.
- the plurality of T lymphocytes is included in or defines a cell line, such as a human cell line.
- an activated alpha beta T lymphocyte Provided is also an activated gamma delta T lymphocyte. Provided is also an activated CD8+ T lymphocyte. Provided is also an activated CD4+ T lymphocyte. Provided is also an activated gamma delta T lymphocyte. Provided is also a cytotoxic T lymphocyte (CTL) or activated T helper lymphocyte (Th cells). This T lymphocyte is stimulated with the peptide disclosed herein.
- the activated CTL is a cell line or a cell of a cell line.
- the activated Th cell is a cell line.
- the method is typically an in vitro method.
- the respective T cell population expresses a T-cell receptor that specifically binds to an MHC ligand.
- the MHC ligand is a peptide as disclosed herein.
- the method includes cloning a nucleic acid that encodes a TCR-alpha and/or TCR-beta chain specifically binding to an MHC ligand into a gamma retrovirus or lentivirus expression vector.
- the ligand is a peptide as disclosed herein.
- the method furthermore includes generating a recombinant virus with antigen specificity and functional avidity for the MHC ligand.
- the method also includes transducing a target T- cell population with the recombinant virus that has been generated.
- the method furthermore includes expanding the T lymphocyte population transduced with the recombinant virus.
- the method includes administering an effective number of T lymphocytes, in particular activated T lymphocytes, to the subject.
- the activated T lymphocytes are in some embodiments produced as described herein.
- the respective target cells in the subject are cancer cells.
- the T lymphocytes are autologous to the subject.
- the T lymphocytes have been obtained from a donor other than the subject.
- the T lymphocytes have been obtained from a healthy donor.
- the T lymphocytes are allogeneic to the subject.
- the T lymphocytes have been derived from autologous tumor infiltrating lymphocytes or from peripheral blood mononuclear cells.
- the T lymphocytes are or have been expanded in vitro.
- the target cancer cells present or over-present a peptide comprising any amino acid sequence defined herein.
- the method includes contacting T lymphocytes, in particular activated T lymphocytes, with the target cancer cells.
- the activated T lymphocytes are in some embodiments produced as described herein.
- the respective target cancer cells have been obtained from a subject.
- the T lymphocytes have been obtained from the same subject.
- the T lymphocytes have been obtained from a donor other than the subject.
- the T lymphocytes have been obtained from a healthy donor.
- the T lymphocytes are allogeneic to the subject.
- the T lymphocytes have been derived from autologous tumor infiltrating lymphocytes or from peripheral blood mononuclear cells.
- the T lymphocytes are or have been expanded in vitro.
- the method includes allowing target cancer cells in a subject to be killed.
- the target cancer cells over-present and/or aberrantly express a peptide comprising any amino acid sequence defined herein.
- the method or use may include administering an effective number of T lymphocytes, in particular activated T lymphocytes, to the subject, produced as described herein.
- an autologous or allogeneic human alpha beta T lymphocyte or gamma delta T lymphocyte recombinantly transfected with a T-cell receptor as defined herein.
- the alpha beta T lymphocyte can be a CD8+ T lymphocyte or a CD4+ T lymphocyte, and can specifically be a cytotoxic T lymphocyte (CTL) or T helper cell (Th cell).
- CTL cytotoxic T lymphocyte
- Th cell T helper cell
- TCR that is identified and isolated from an activated T lymphocyte as disclosed herein.
- TCR TCR
- a functional fragment thereof that is prepared based on a peptide as disclosed herein, a nucleic acid encoding the peptide, an expression vector that contains such nucleic acid or a host cell as described herein.
- a medical use of a peptide as provided herein In this regard there is provided a peptide as described herein for use in the treatment of a disease and in medicine. Typically, the disease is cancer.
- nucleic acid or an expression vector as provided herein.
- a nucleic acid or an expression vector as described herein for use in the treatment of a disease and in medicine.
- the disease is cancer.
- any peptide as described as a medicament or in the manufacture of a medicament there is provided the nucleic acid provided herein or the expression vector provided herein as a medicament or in the manufacture of a medicament.
- the cell provided herein or the activated T lymphocyte as a medicament or in the manufacture of a medicament.
- the proteinaceous binding molecule e.g. the TCR or the antibody or other peptide and/or peptide-MHC binding molecules provided herein, as a medicament or in the manufacture of a medicament.
- the medicament is active against cancer.
- the medicament is a cellular therapeutic agent or a vaccine.
- the medicament is a protein based on a soluble TCR or antibody.
- the medicament is a vaccine, such as an anti-cancer vaccine.
- the medicament is a cellular therapeutic agent.
- the cancer is acute myeloid leukemia or chronic lymphocytic leukemia.
- the cancer is breast cancer or cholangiocellular carcinoma.
- the cancer is colorectal cancer or gallbladder cancer.
- the cancer is glioblastoma or gastric cancer.
- the cancer is gastro-esophageal junction cancer or hepatocellular carcinoma.
- the cancer is head and neck squamous cell carcinoma or melanoma.
- the cancer is non-Hodgkin lymphoma or non-small cell lung cancer.
- the cancer is ovarian cancer or esophageal cancer.
- the cancer is pancreatic cancer or prostate cancer.
- the cancer is renal cell carcinoma or small cell lung cancer.
- the cancer is urinary bladder carcinoma or uterine endometrial cancer.
- Figures 1 A through 1 G show the over-presentation of various peptides in different cancer cells or tissues compared to normal cells or tissues.
- Upper part Median MS signal intensities from technical replicate measurements are plotted as dots for single normal (grey dots, left part of figure) and tumor samples (black dots, right part of figure) of the respective HLA allotype on which the peptide was detected. Boxes display median, 25th and 75th percentile of normalized signal intensities, while whiskers extend to the lowest data point still within 1.5 interquartile range (IQR) of the lower quartile, and the highest data point still within 1.5 IQR of the upper quartile.
- Lower part The relative peptide detection frequency in every organ is shown as spine plot.
- Tissues (from left to right): Normal samples: adipose (adipose tissue); adrenal gl (adrenal gland); bile duct; bladder; bloodcells; bloodvess (blood vessels); bone marrow; brain; breast; esoph (esophagus); eye; gall bl (gallbladder); head&neck; heart; intest, la (large intestine); intest, sm (small intestine); kidney; liver; lung; lymph nodes; nerve cent (central nerve); nerve periph (peripheral nerve); ovary; pancreas; parathyr (parathyroid gland); perit (peritoneum); pituit (pituitary); placenta; pleura; prostate; skel.
- Tumor samples AML (acute myeloid leukemia); BRCA (breast cancer); CCC (cholangiocellular carcinoma); CLL (chronic lymphocytic leukemia); CRC (colorectal cancer); GBC (gallbladder cancer); GBM (glioblastoma); GC (gastric cancer); GEJC (gastro-esophageal junction cancer); HCC (hepatocellular carcinoma); HNSCC (head and neck squamous cell carcinoma); MEL (melanoma); NHL (non-Hodgkin lymphoma); NSCLCadeno (non-small cell lung cancer adenocarcinoma); NSCLCother (NSCLC samples that could not unambiguously be assigned to NSCLCadeno or NSCLCsquam); NSCLCsquam (squamous cell nonsmall
- Figure 1A Peptide: AAFDGRHSQTLK (SEQ ID NO: 2), Figure 1 B) Peptide: GQHLHLETF (SEQ ID NO: 15), Figure 1 C) Peptide: MPMQDIKMI (SEQ ID NO: 28), Figure 1 D) Peptide: MPMQDIKMIL (SEQ ID NO: 29), Figure 1 E) Peptide: SLLQHLIGL (SEQ ID NO: 44), Figure 1 F) Peptide: SPSVSQLSVL (SEQ ID NO: 48), Figure 1 G) Peptide: YLHARLREL (SEQ ID NO: 58).
- Figures 2A through 2G show exemplary expression profile of source genes of the present invention that are overexpressed in different cancer samples.
- Tumor black dots
- normal grey dots
- Box-and-whisker plots represent median value, 25th and 75th percentile (box) plus whiskers that extend to the lowest data point still within 1 .5 interquartile range (IQR) of the lower quartile and the highest data point still within 1 .5 IQR of the upper quartile.
- IQR interquartile range
- Tissues (from left to right): Normal samples: adipose (adipose tissue); adrenal gl (adrenal gland); bile duct; bladder; bloodcells; bloodvess (blood vessels); bone marrow; brain; breast; esoph (esophagus); eye; gall bl (gallbladder); head&neck; heart; intest, la (large intestine); intest, sm (small intestine); kidney; liver; lung; lymph nodes; nerve periph (peripheral nerve); ovary; pancreas; parathyr (parathyroid gland); perit (peritoneum); pituit (pituitary); placenta; pleura; prostate; skel.
- adipose adipose tissue
- adrenal gl adrenal gland
- bile duct bladder
- bloodcells bloodvess
- bone marrow brain
- brain breast
- esoph es
- Tumor samples AML (acute myeloid leukemia); BRCA (breast cancer); CCC (cholangiocellular carcinoma); CLL (chronic lymphocytic leukemia); CRC (colorectal cancer); GBC (gallbladder cancer); GBM (glioblastoma); GC (gastric cancer); GEJC (gastro-esophageal junction cancer); HCC (hepatocellular carcinoma); HNSCC (head and neck squamous cell carcinoma); MEL (melanoma); NHL (non-Hodgkin lymphoma); NSCLCadeno (non-small cell lung cancer adenocarcinoma); NSCLCother (NSCLC samples that could not unambiguously be assigned to NSCLCadeno or NSCLCsquam); NSCLCsquam (squamous cell non-
- Figure 3 shows the results of the IdentControl experiments for one exemplary peptide SLLQHLIGL (SEQ ID NO: 44).
- the peptide was confirmed by IdentControl comparing the fragmentations of stable isotope labeled (SIL) standards in data-dependent acquisition (DDA) mode. Identity was confirmed using an in-house determined spectral correlation threshold.
- SIL stable isotope labeled
- Figure 4 shows one exemplary results for a CoElution experiment for the peptide YLHARLREL (SEQ ID NO: 58).
- the peptide was confirmed by CoElution using stable isotope labeled (SIL) internal standard and targeted MS (sPRM or IS-PRM).
- SIL stable isotope labeled
- MS sPRM or IS-PRM
- Non overlapping MS2 isolation windows for the SIL-peptide and the natural peptide are used.
- Control experiments using non-HLA peptidome sample e.g. tryptic digest or 5% FA
- Peptide identity is confirmed based on objective, predefined criteria in expert manual review.
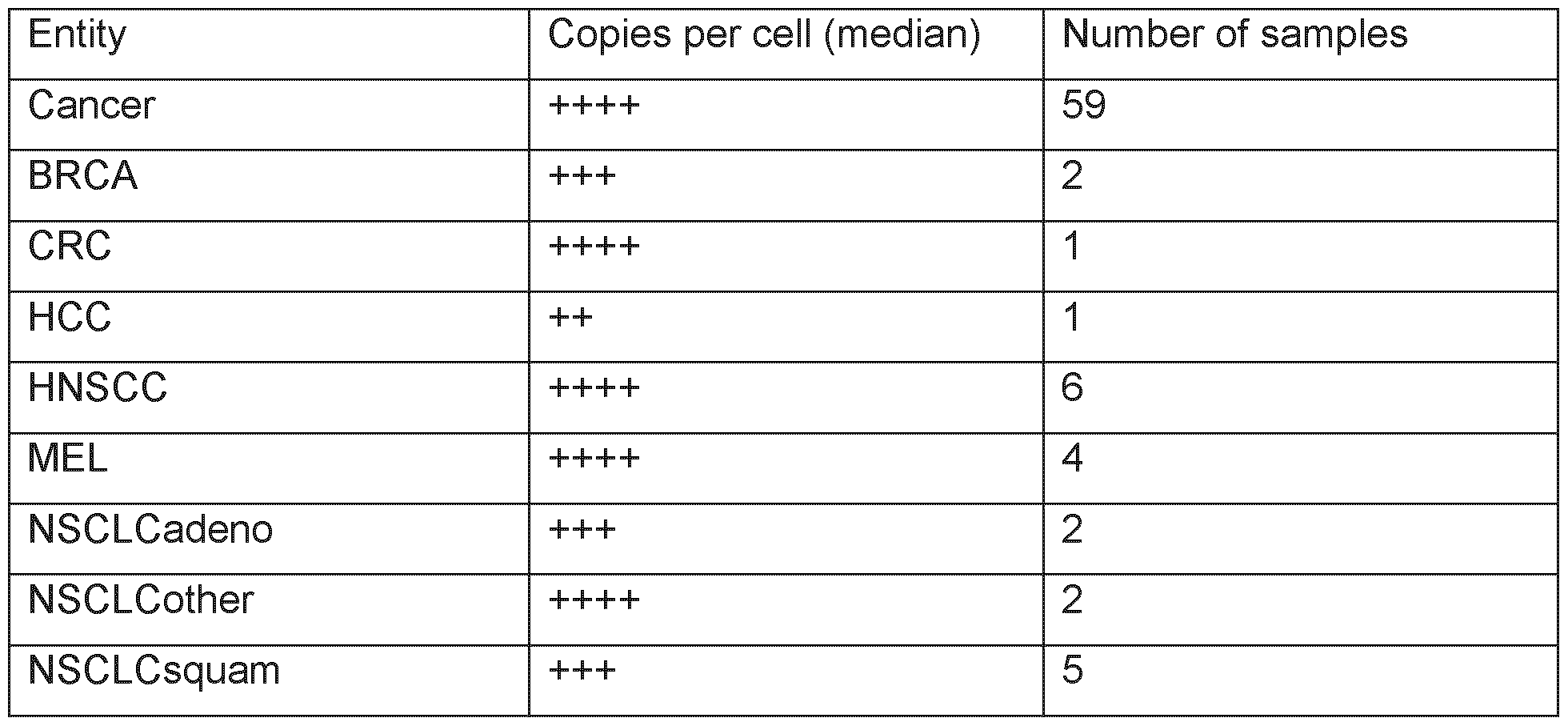
- Figures 5A through 5B shows absolute copies per cell in different tumor samples. Absolute copies per cell are plotted as black dots for tumor samples of the respective HLA allotype on which the peptide was detected. Boxes display median, 25th and 75th percentile of absolute copies per cell, while whiskers extend to the lowest data point still within 1.5 interquartile range (IQR) of the lower quartile, and the highest data point still within 1.5 IQR of the upper quartile.
- IQR interquartile range
- BRCA breast cancer
- CRC colonal cancer
- HCC hepatocellular carcinoma
- HNSCC head and neck squamous cell carcinoma
- MEL melanoma
- NSCLCadeno non-small cell lung cancer adenocarcinoma
- NSCLCother NSCLC samples that could not unambiguously be assigned to NSCLCadeno or NSCLCsquam
- NSCLCsquam squamous cell non-small cell lung cancer
- OC ovarian cancer
- OSCAR esophageal cancer
- PRCA prostate cancer
- RCC renal cell carcinoma
- SCLC small cell lung cancer
- UBC urinary bladder carcinoma
- UEC uterine endometrial cancer
- the present inventors identified peptides from the PRAME protein which are overpresented on several cancer cells.
- the over-presentation of these peptides can serve as the basis for a method of predicting a risk of occurrence of cancer and for a use and a method of diagnosing cancer.
- a cell over-presenting one or more of the respective peptides may also be a target in a method and a use in the prevention of cancer and/or the treatment of cancer.
- Peptides that represent T cell epitopes can be identified based on in silico epitope prediction or directly using mass spectrometry-based immunopeptidomics. A prediction based on algorithms does not necessarily lead to the successful identification of peptides that indeed exist in vivo. If a peptide is synthesized based on in silico epitope prediction, the synthesized peptide may turn out not to bind to MHC with high affinity and/or not to be recognized by T cells. This is due to the fact that the processes involved in the formation of peptides that are presented on the cell surface by MHC are complex and only partly understood today. Peptides identified using in silico prediction therefore need to be assessed with regard to their capability of binding to MHC and their capability of being recognized by CD4 and/or CD8 T cells.
- HLA peptide All peptides disclosed in the application were isolated from patient samples in the form of a complex with HLA molecules or MHC class I, also known as a pMHC complex and denoted in the Working Examples as “HLA peptide”.
- a HLA-specific antibody was used to recover HLA from patient samples.
- the antibody is conformation-specific and only binds to folded MHC class I complexes.
- MHC is only folded if binding to a peptide, so that no antibody binding and no isolation can occur if there is no peptide bound to the MHC.
- the peptide/MHC complexes were then immobilized on affinity chromatography material via an MHC- specific antibody. Peptides were then released from the MHC complex by acid treatment, and subsequently analyzed by mass spectrometry.
- PRAME Preferentially Expressed Antigen in Melanoma
- OIP4 Opa-interacting protein 4
- CT130 and MAPE is a protein and tumor antigen of the Cancer/Testis antigen group. It is a member of the PRAME family, which includes inter alia LRRC14, PRAME family members 1 , 2, 6, 25 or 33 (PRAMEF1 , PRAMEF2, PRAMEF6, PRAMEF25, PRAMEF33), and isoforms of the protein such as isoform “CRA_a” of UniProt accession number A0A024R1 E6, are known.
- PRAME has a length of 509 amino acids and a mass of 57,890 Da.
- PRAME has the NCBI accession number CAG30435, version 1 as of 2 February 2011 , and the UniProt accession number P78395, version 1 , release 2023JD3 of 28 June 2023, GenBank accession number CAG30435.1 , Gene ID 23532.
- PRAME is a component of a Cul2-RING (CRL2) E3 ubiquitin-protein ligase complex, which mediates ubiquitination of target proteins, leading to their degradation. The protein is expressed at a high level in a large proportion of tumors as well as several types of leukemia.
- PRAME is the best characterized member of the PRAME family of leucine-rich repeat (LRR) proteins. Mammalian genomes contain multiple members of the PRAME family whereas in other vertebrate genomes only one PRAME-like LRR protein was identified.
- PRAME is a cancer/testis antigen that is expressed at very low levels in normal adult tissues except testis but at high levels in a variety of cancer cells. Unless otherwise defined, all other scientific and technical terms used in the description, figures and claims have their ordinary meaning as commonly understood by one of ordinary skill in the art.
- a peptide may refer to one specific peptide or to a plurality of peptides, and terms such as “the method” are meant to reference similar steps and/or techniques known to those skilled in the art.
- the term “at least” preceding a series of elements is to be understood to refer to every element in the series.
- the terms “at least one” and “at least one of” include for example, one, two, three, four, or five or more elements. It is furthermore understood that slight variations above and below a stated range can be used to achieve substantially the same results as a value within the range. Also, unless indicated otherwise, the disclosure of ranges is intended as a continuous range including every value between the minimum and maximum values.
- Consisting of when reference is made to a peptide, shall mean a peptide provided herein with an amino acid sequence according to any of SEQ ID NO: 1 to SEQ ID NO: 59 that has the capacity of binding or binds to MHC molecules.
- the term “consisting of” excludes any element, component, integer step, or ingredient not specified in the context of the term. Where a method is for example defined as consisting of one or more steps or operations, it is meant that the method does not involve any other steps than those specified.
- peptide is used herein to designate a series of amino acid residues, connected one to the other typically by peptide bonds between the alpha-amino and carboxyl groups of the adjacent amino acids.
- peptide as used herein also includes “tumor associated peptide” or “TUMAP”.
- TUMAP tumor associated peptide
- tumor-associated peptide or “TUMAP”, also called “tumor antigenic peptide” in the art, is an antigenic peptide epitope that can be found presented by MHC on the surface of cells of a tumor.
- a tumor-associated peptide is recognized by a specific TCR on a T lymphocyte.
- a TUMAP as naturally presented on a target cell can be derived from one or more source proteins by degradation within that cell, herein also addressed as the source protein of the TUMAP.
- selection of peptides denotes a plurality of peptides or a plurality of peptide sequences.
- the collection of peptides may include 2, 3 or more peptides or peptide sequences.
- the collection of peptides may for example define a library of peptides.
- the collection of peptides may also be a database or included in a database.
- the peptides of the collection of peptides may have been pre-screened for immunogenicity and/or over-presentation in a particular tumor
- elongated peptide refers to a peptide that includes a defined amino acid sequence, and that in addition to this sequence contains further amino acids at the N- and/or C-terminus of the defined sequence.
- polypeptide designates a series of amino acid residues, connected one to the other typically by peptide bonds between the alpha-amino and carboxyl groups of the adjacent amino acids.
- polypeptide is meant to refer to molecules containing more than about 30 amino acid residues.
- a peptide, protein, nucleic acid molecule or polynucleotide coding for such a molecule is “immunogenic” (and thus is an “immunogen” within the present disclosure), if it is capable of inducing an immune response.
- immunogenicity is more specifically defined as the ability to induce a T cell response.
- an “immunogen” would be a molecule that is capable of inducing an immune response, and in the case of the present disclosure, a molecule capable of inducing a T cell response.
- the immunogen can be the peptide, the complex of the peptide with MHC, and/or protein that is used to raise a specific antibody or a TCR against it.
- the same peptide can be immunogenic in one person but not immunogenic in a different person, depending on the personal T cell receptor repertoires in each person which may or not contain T cell receptors able to recognize said peptide.
- immunogenicity of a peptide plays an important role when the immune response relies on the native pool of available T cell clones within a body, e.g. in the context of vaccination.
- immunogenicity of peptide can also be modulated using adjuvants which can evoke an immune response against peptide that would otherwise not be immunogenic, effectively turning a non-immunogenic peptide into an immunogenic peptide.
- a therapy involves binding molecules specific for a peptide being administered into the body, immunogenicity of said peptide plays a lesser role as the immune response does not rely on the naturally available T cell receptor repertoire present in the patient.
- a class I T cell “epitope” generally requires a short peptide that is bound to a class I MHC molecule (MHC class I alpha chain, beta-2-microglobulin, and peptide) that can be recognized by a T cell bearing a matching T cell receptor binding to the MHC-peptide complex with appropriate affinity.
- MHC class I alpha chain, beta-2-microglobulin, and peptide Peptides binding to MHC class I molecules are typically 8-13 amino acids in length, for example 9 amino acids in length.
- MHC major histocompatibility complex
- HLA human leukocyte antigen
- binding molecule refers to a molecule that specifically binds to an (e.g. antigenic) determinant.
- a “binding molecule” is generally one of a proteinaceous binding molecule and a nucleic acid molecule.
- a binding molecule is able to direct an entity to which it is coupled or attached (e.g. a (second) antigen binding moiety) to a target site, for example to a specific type of tumor cell or tumor stroma bearing the antigenic determinant (e.g. the complex of a peptide with MHC, according to the application at hand).
- a binding molecule is able to activate signaling through its target antigen, for example a T cell receptor complex antigen.
- binding molecules include, but are not limited to, antibodies and fragments thereof, binding proteins comprising at least one ankyrin repeat motif and single domain antigen binding (SDAB) molecules, aptamers, and (soluble) TCRs and fragments thereof.
- SDAB ankyrin repeat motif and single domain antigen binding
- a functional fragment as used herein is to be understood as a portion of a binding molecule that retains the biological function of the binding molecule from which the functional fragment is derived.
- a functional fragment of a binding molecule shall mean a portion of a binding molecule that specifically binds to the same antigenic determinant as the binding molecule said functional fragment is derived from.
- Such functional fragments of binding molecules are also referred to herein as binding fragments.
- peptide may for example be a peptide that is presented on the surface of a cell, including a peptide that is presented by MHC.
- an antibody with a binding affinity of ⁇ 100 nM, or ⁇ 50 nM to the peptide-MHC complex may be regarded as “specific” in the context of the present disclosure.
- binding means that the binding molecule, e.g. proteinaceous binding molecule, binds the peptide-MHC complex of interest better than other peptide-MHC complexes.
- Specific binding can be determined, for example, in accordance with a Western blot, an ELISA-, RIA-, ECL-, IRMA-test, FACS, IHC and a peptide scan.
- the antibody or fragment thereof, TCR or fragment thereof, or binding molecule can thereby form a complex with the respective peptide.
- Such binding may be exemplified by the specificity of a “lock-and- key-principle”.
- a peptide as disclosed herein may define the epitope to which a binding molecule such as a TCR specifically binds.
- the peptide is in some embodiments presented, and thereby fixed, by an MHC molecule.
- antibody or “antibodies” is used herein in a broad sense and includes both polyclonal and monoclonal antibodies.
- antibody fragment e.g. a CDR, Fv, Fab and or an Fc fragment
- polymer of such antibody molecule e.g. a CDR, Fv, Fab and or an Fc fragment
- humanized version of an antibody molecule e.g., a humanized version of an antibody molecule
- mAb monoclonal antibody
- mAb monoclonal antibody
- the monoclonal antibodies herein specifically include "chimeric" antibodies in which a portion of the heavy and/or light chain is identical with or homologous to corresponding sequences in antibodies derived from a particular species or belonging to a particular antibody class or subclass, while the remainder of the chain(s) is identical with or homologous to corresponding sequences in antibodies derived from another species or belonging to another antibody class or subclass, as well as fragments of such antibodies, so long as they exhibit the desired binding activity (e.g., specific binding of a peptide disclosed herein).
- antibody fragment shall refer to a fragment of such an antibody retaining its specific binding capacities, e.g.
- an IgG or IgM heavy chain (consisting of VH, CH1 , hinge, CH2 and CH3 regions)
- an IgG or IgM light chain (consisting of VL and CL regions), and/or
- antibody Also included in the term “antibody” are diabodies, camelid antibodies, nanobodies, domain antibodies, bivalent homodimers with two chains consisting of scFvs, IgAs (two IgG structures joined by a J chain and a secretory component), shark antibodies, antibodies consisting of new world primate framework plus non-new world primate CDR, dimerized constructs comprising CH3+VL+VH, and antibody conjugates (e.g. antibody or fragments or derivatives linked to a toxin, a radioisotope or a label).
- diabodies camelid antibodies, nanobodies, domain antibodies, bivalent homodimers with two chains consisting of scFvs, IgAs (two IgG structures joined by a J chain and a secretory component), shark antibodies, antibodies consisting of new world primate framework plus non-new world primate CDR, dimerized constructs comprising CH3+VL+VH, and antibody conjugates (e.g. antibody or fragments or
- T cell receptor refers to a heterodimeric molecule typically comprising an alpha polypeptide chain (alpha chain) and a beta polypeptide chain (beta chain), wherein the heterodimeric receptor is capable of binding to a peptide antigen presented by an MHC molecule.
- the term also includes a so-called gamma/delta TCR.
- TCR fragment refers to either a portion of a native TCR or a recombinant/engineered protein that contains TCR-derived complementaritydetermining region (CDR) sequences.
- CDR TCR-derived complementaritydetermining region
- a TCR fragment is an antigen-binding TCR fragment, which specifically binds to a peptide when bound by MHC proteins.
- binding to the respective peptide/MHC complex is defined by the CDR sequences, in particular CDR1 and CDR3, these sequences are included in an antigen-binding TCR fragment.
- CDR sequences in particular CDR1 and CDR3
- a variable domain comprising TCR-derived CDRs and antibody-derived framework regions may thus be considered an antigen-binding TCR fragment.
- an antigen-binding TCR fragment can contain an alpha chain variable domain (Va), or a portion thereof, and a beta chain variable domain (VP), or a portion thereof, each comprising three CDRs.
- an antigen-binding TCR fragment can contain a gamma chain variable domain (Vy), or a portion thereof, and a delta chain variable domain (V8), or a portion thereof, each comprising three CDRs.
- a TCR fragment may also be a single-chain TCR fragment, which contains only the Va and Vp domains, connected by a linker.
- immune response means a physiological process involving the activation and/or induction of an effector function of the body’s immune system in, by way of non-limiting examples, a T cell, B cell, natural killer (NK) cell, and/or an antigen- presenting cell (APC).
- An example of an immune response is any detectable antigenspecific activation and/or induction of a helper T cell or cytotoxic T cell activity or response, production of antibodies, antigen-presenting cell activity or infiltration, macrophage activity or infiltration, or neutrophil activity or infiltration.
- T cell response means the specific proliferation and activation of effector functions induced by a peptide in vitro or in vivo.
- effector functions may be lysis of peptide-pulsed, peptide-precursor pulsed or naturally peptide-presenting target cells, secretion of cytokines, such as Interferongamma, TNF-alpha, or IL-2 induced by peptide, secretion of effector molecules, such as granzymes or perforins induced by peptide, or degranulation.
- aberrantly expressed and “aberrantly express” it is meant that a polypeptide or protein is expressed at unusual high levels (overexpressed) in tumor tissue or tumor cells when compared to levels of expression in normal tissues or normal cells, or that the gene is silent in the tissue or cells from which the tumor is derived whereas in the tumor, it is expressed.
- polypeptide, protein or gene transcript is present at a level at least 1 .2-fold of that present in normal tissue or on normal cells; such as at least 2-fold.
- polypeptide, peptide or gene transcript is present at a level at least 5-fold or 10-fold the level present in normal tissue or on normal cells.
- over-presented and variants of the term such as “overpresented” or “overpresentation”, as used herein refers to the relative count of a peptide presented on a tissue or cell population addressed, when compared to an average count, i.e. the frequency with which the same peptide is presented on a healthy tissue or cell population of the same type. Furthermore, it can refer to the relative intensity of peptide presented on a tissue or cell population addressed, when compared to an average intensity, i.e. the amount of the same peptide with which it is presented on a healthy tissue or cell population of the same type.
- the term indicates that the peptide is presented by MHC proteins on the cell surface, both on the cells showing overpresentation and on the healthy reference. If a certain peptide is over-presented on cells of a certain cancer, a higher number of the peptide is bound to MHC proteins on the cell surface of the cells of this cancer than on the cell surface of corresponding tissue or cell population that is healthy.
- nucleic acid and “nucleic acid molecule” as used herein refer to any nucleic acid in any possible configuration, such as single stranded, double stranded or a combination thereof.
- the specific sequence unless the context indicates otherwise, refers to the single strand DNA of such sequence, the duplex of such sequence with its complement (double stranded DNA) and the complement of such sequence.
- nucleic acid sequence in some instances also referred to as a “nucleotide sequence”, refers to a heteropolymer of deoxyribonucleotides.
- a nucleic acid coding for (or encoding) a peptide is meant to include a reference to a nucleic acid sequence coding for the respective peptide, including artificial (man-made) start and stop codons compatible with the biological system by which the sequence is to be expressed.
- the nucleic acid sequence coding for a particular peptide or binding molecule may be naturally occurring or it may be synthetically constructed.
- a DNA segment encoding a peptide, polypeptide, and protein as provided herein is assembled from cDNA fragments and short oligonucleotide linkers, or from a series of oligonucleotides, to provide a synthetic gene that is capable of being expressed in a recombinant transcriptional unit comprising regulatory elements derived from a microbial or viral operon.
- the system may, for example, be a dendritic cell or another cell system useful for the production of TCRs.
- coding region refers to that portion of a gene which either naturally or normally codes for the expression product of that gene in its natural genomic environment, i.e. , the region coding in vivo for the native expression product of the gene.
- the coding region can be derived from a non-mutated (“normal”), mutated or altered gene, or can even be derived from a DNA sequence, or gene, wholly synthesized in the laboratory using methods well known to those of skill in the art of DNA synthesis.
- expression product means the polypeptide or protein that is the natural translation product of the gene and any nucleic acid sequence coding equivalents resulting from genetic code degeneracy and thus coding for the same amino acid(s).
- a peptide as disclosed herein may also be an expression product, when expressed from a recombinant nucleic acid construct encoding such peptide.
- DNA segment refers to a DNA polymer, in the form of a separate fragment or as a component of a larger DNA construct, which has been derived from DNA isolated at least once in a substantially pure form, i.e., free of contaminating endogenous materials and in a quantity or concentration enabling identification, manipulation, and recovery of the segment and its component nucleotide sequences by standard biochemical methods, for example, by using a cloning vector.
- Such segments are provided in the form of an open reading frame uninterrupted by internal non-translated sequences, or introns, which are typically present in eukaryotic genes. Sequences of non-translated DNA may be present downstream from the open reading frame, where the same do not interfere with manipulation or expression of the coding regions.
- vector refers to a vehicle into which a polynucleotide encoding a peptide or protein may be operably inserted.
- a vector contains a site that is adapted for the insertion of a polynucleotide, such as a multiple cloning site.
- a vector is a linear or circular nucleic acid molecule.
- expression vector refers to a vector that is cable of effecting the expression of a peptide or protein that is encoded by a nucleic acid sequence included in the vector.
- An expression vector contains particular sequences allowing the expression of such a nucleic acid sequence, to which the nucleic acid sequence can be operably coupled. Such a particular sequence is a promoter. Further examples of such sequences include a start codon, a polyadenylation signal, a termination signal, and a ribosome binding site.
- An expression vector may also contain one or more control sequences. An example of a control sequence is an enhancer.
- Various types of expression vectors are known, for example plasmid and viral-based expression vectors.
- promoter means a region of a nucleic acid, typically DNA, involved in binding of RNA polymerase to initiate transcription.
- recombinant or “heterologous” is used in this specification interchangeably to describe a nucleic acid molecule that, by virtue of its origin, manipulation, or both is not associated with all or a portion of the nucleic acid molecule with which it is associated in nature.
- a recombinant nucleic acid molecule includes a sequence which does not naturally occur in the respective wildtype organism or cell.
- a recombinant nucleic acid molecule is obtained by genetic engineering, usually constructed outside of a cell.
- a recombinant nucleic acid molecule is substantially identical and/or substantial complementary to at least a portion of the corresponding nucleic acid molecule occurring in nature.
- a recombinant nucleic acid molecule may be of any origin, such as genomic, cDNA, mammalian, bacterial, viral, semisynthetic or synthetic origin.
- the term "recombinant" as used herein with respect to a protein / peptide means a peptide produced by expression of a recombinant nucleic acid.
- isolated means that the material is removed from its original environment (e.g., the natural environment, if it is naturally occurring).
- a naturally- occurring nucleic acid molecule or protein present in a living animal is not isolated, but the same nucleic acid molecule or protein, separated from some or all of the coexisting materials in the natural system, is isolated.
- Such nucleic acid molecule could be part of a vector and/or such nucleic acid molecule or protein could be part of a composition, and still be isolated in that such vector or composition is not part of its natural environment.
- a peptide as disclosed herein may be isolated from natural sources as illustrated in the Working Examples. It may also be expressed in a host cell, and subsequently isolated from this host cell or from the media that includes the host cell. It may also be isolated from both the host cell and the media that includes the host cell.
- a “pharmaceutical composition” as used herein is a composition suitable for administration to a human being in a medical setting.
- a pharmaceutical composition is sterile and produced according to GMP guidelines.
- the pharmaceutical composition may for example be a solution or a solid.
- the pharmaceutical composition may for example be a suspension.
- Examples of a solid form of a composition include, but are not limited to a granule, a pellet, a micro-pellet, a sphere, a core, a coated core, a pill, a compressed tablet, a mini tablet, a layered tablet, a bead, a capsule, a powder or any form of a particle.
- the pharmaceutical composition may be a mixture that contains an agent as disclosed herein, e.g.
- a pharmaceutical composition may also be a mixture that includes a population of immune cells, such as human immune cells, and a protein such as albumin.
- the pharmaceutical composition may also be a mixture that contains a plurality of (activated) T lymphocytes as described herein. Said (activated) T lymphocytes may be alpha beta T lymphocytes or gamma delta T lymphocytes.
- the (activated) T lymphocytes may be CD8+ T lymphocytes or CD4+ T lymphocytes.
- the (activated) T lymphocytes may be cytotoxic T cells (CTLs) as disclosed herein.
- the pharmaceutical composition may or may not comprise one or more antigen-presenting cells and/or one or more MHC molecules.
- a “combination product” refers to a product that includes a plurality, i.e. two or more, individual pharmaceutical compositions.
- the pharmaceutical compositions may be provided for administration either together or separately.
- a combination product may include storage means for pharmaceutical preparations as individual or separate preparations.
- a combination product may also include means for dispensing a pharmaceutical preparation.
- an “active agent” is an entity, such as a pharmaceutical compound, that can be used to achieve a desired effect, e.g., a therapeutic effect.
- a therapeutic effect may be an effect in the treatment, including the prevention, of cancer.
- it is also referred to as a “therapeutic agent”.
- an active agent is intended to be used for administration to a subject.
- an active agent defines or contains an active ingredient, which can be used in a pharmacologically, including therapeutically, effective amount.
- An active agent may be included in a delivery system or in a kit. Examples of an active agent include a peptide as disclosed herein or a cell. Two further examples of an active agent are a proteinaceous binding molecule as disclosed herein or an adjuvant, such as an interleukin. It should be noted that as long as an active agent is referred to, this does not necessarily include its medical use.
- An “individualized pharmaceutical” shall mean a specifically tailored therapy product for one individual subject that will only be used for therapy in such individual subject, including an actively individualized cancer vaccine and/or an adoptive cellular therapy using autologous tissue from a subject, e.g. autologous patient tissue.
- a “personalized pharmaceutical” shall mean a specifically tailored therapy product for one individual person that will only be used for therapy in such individual person. As long as the subject is a human, “individualized” and “personalized” can be used interchangeably.
- a salt refers to an ionized form of a disclosed peptide where the peptide is provided with one or more counter ions, thereby providing an acid or base salt of the peptide.
- an acid salt is prepared from the free base (typically wherein the neutral form of the peptide has a neutral -NH2 group) involving reaction with a suitable acid.
- Suitable acids for preparing acid salts include both organic acid and inorganic acid, as explained in more detail below.
- pharmaceutically acceptable salt refers to any salt of a peptide provided herein which retains its biological properties and which is not toxic or otherwise undesirable for pharmaceutical use.
- a salt may be derived from a variety of organic or inorganic counter-ions well known in the art. Examples of a respective salt include, but are not limited to the chloride salt, the acetate salt or the trifluoroacetate salt, see also below.
- lipid nanoparticle a particle having at least one dimension on the order of nanometers (e.g., 1 -1 ,000 nm), which includes one or more lipids.
- cationic lipid refers to a lipid having a net positive charge. Cationic lipids bind a negatively charged nucleic acid molecule such as RNA by electrostatic interaction to the lipid matrix. Generally, cationic lipids possess a lipophilic moiety, such as a sterol, an acyl or diacyl chain, and the head group of the lipid typically carries the positive charge.
- a cationic lipid 1 ,2-di-O- octadecenyl-3-trimethylammonium propane (DOTMA) and dimethyldioctadecylammonium (DDAB).
- DOTMA dioleoyl-3-trimethylammonium propane
- DODAP dimethyldioleoyl-3- dimethylammonium-propane
- 1 ,2-diacyloxy-3-dimethylammonium propanes and 1 ,2-dialkyloxy-3-dimethylammonium propanes are further examples of a cationic lipid.
- a cationic lipid examples include dioctadecyldimethyl ammonium chloride (DODAC) and 2,3-dioleoyloxy-N-[2(spermine carboxamide)ethyl]-N,N-dimethyl-l- propanamium trifluoroacetate (DOSPA).
- DODAC dioctadecyldimethyl ammonium chloride
- DOSPA 2,3-dioleoyloxy-N-[2(spermine carboxamide)ethyl]-N,N-dimethyl-l- propanamium trifluoroacetate
- DMRIE 2,3-di(tetradecoxy)propyl-(2-hydroxyethyl)-dimethylazanium
- DMEPC dimyristoyl-sn-glycero-3-ethylphosphocholine
- Examples of a cationic lipid are also 1 ,2-dimyristoyl-3-trimethylammonium propane (DMTAP) and 1 ,2-dioleyloxypropyl- 3-dimethyl-hydroxyethyl ammonium bromide (DORIE).
- DMTAP 1,2-dimyristoyl-3-trimethylammonium propane
- DORIE 1,2-dioleyloxypropyl- 3-dimethyl-hydroxyethyl ammonium bromide
- neutral lipid refers to a lipid having a net charge of zero.
- Two illustrative examples of a neutral lipid are 1 ,2-di-(9Z-octadecenoyl)-sn-glycero-3- phosphoethanolamine (DOPE) and 1 ,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC).
- DOPE 1 ,2-dioleoyl-sn-glycero-3-phosphocholine
- DOPC 1,2-dioleoyl-sn-glycero-3-phosphocholine
- Two further examples of a neutral lipid are diacylphosphatidyl choline and diacylphosphatidyl ethanol amine.
- Two further examples of a neutral lipid are ceramide and sphingoemyelin.
- Yet two examples of a neutral lipid are cephalin and cholesterol.
- Another example of a neutral lipid is cerebroside.
- tissue as used herein, is intended to refer to a group of cells that have a similar structure and act together to perform a specific function.
- a tissue may be a cluster of cells.
- a tissue may contain extracellular matrix.
- normal tissue or “healthy tissue” are used herein, reference is made to cells of healthy tissue, including healthy peripheral blood mononuclear cells (PBMCs).
- PBMCs peripheral blood mononuclear cells
- Illustrative examples of respective tissue include, but are not limited to, healthy blood cells, brain, heart, liver, lung, adipose tissue, adrenal gland, bile duct, bladder, bone, bone marrow, esophagus, eye, gallbladder, head and neck, large intestine, small intestine, kidney, lymph node, central nerve, peripheral nerve, pancreas, parathyroid gland, peritoneum, pituitary, pleura, skeletal muscle, skin, spinal cord, spleen, stomach, thyroid, trachea, and ureter cells or other normal tissue cells such as breast, ovary, placenta, prostate, testis, thymus and uterus.
- PBMCs peripheral blood mononuclear cells
- normal tissue or “healthy tissue” are also used to address a sample from a healthy subject.
- tumor tissue typically cancer, including bone marrow, blood-forming cells and blood cells.
- body fluid is understood as an extracellular fluid found within an organism, typically fluid that can be found in a body cavity or hollow organ.
- Illustrative examples of a suitable body fluid include, but are not limited to, blood or cerebrospinal fluid.
- Two further examples of a suitable body fluid are pleural fluid or synovial fluid.
- a further illustrative example of a body fluid is ascitic fluid.
- healthy individual it is meant that the individual is generally in good health.
- a healthy individual generally has a competent immune system and is typically not suffering from any disease that can be readily tested for and detected.
- subject refers to a human or non-human animal, generally a mammal.
- a subject may be a mammalian species such as a rabbit, a mouse, a rat, a guinea pig, a hamster, a dog, a cat, a pig, a cow, a goat, a sheep, a horse, a monkey, an ape or a human.
- the methods, uses and compositions described in this document are generally applicable to both human and veterinary disease.
- the sample used to determine over-presentation and/or aberrant expression of a peptide on a target cell has been obtained from the subject.
- a subject is typically a living organism.
- a subject may also be a non-living organism. Where the subject is a living human who is receiving medical care for a disease or condition, it may also be addressed as a “patient”.
- stratifying and “stratification” as used herein indicate that a subject is assigned to a group, for example with regard to the suitability of the subject for a particular therapy.
- individuals suffering from cancer or being at risk of suffering from cancer may be stratified according to one or more peptides being overpresented on cancer tissue.
- T lymphocytes bind to short continuous amino acid sequences, see above. These sequences are often buried within the native structure of the protein. In the folded source protein these short amino acid sequences are thus typically not accessible (Murphy, 2022, p. 161 ). When presented as a small peptide on the cell surface, the sequence can only be recognized by a matching T lymphocyte if bound by MHC (Murphy, 2022, p. 161 ).
- Peptides presented on the cell surface can originate from any protein found within the cell (Gfeller, D, & Bassani-Sternberg, M, Front. Immunol. (2016), Sec. B Cell Biology, 9 doi: 10.3389/fimmu.2018.01716; Berg et al., 2015, p. 996). It can, however, not reliably be predicted which source protein will give rise to what sort of peptide(s).
- the present inventors have furthermore found that the peptides disclosed herein are typically not presented on cells of healthy tissue or presented in very small numbers. This means, for example, that in a healthy organism a peptide as provided herein is not presented to cells of the cellular immune response.
- Cell-mediated immunity evolved to cope with intracellular pathogens (Berg et al., 2023, p. 89; Berg et al., 2015, p. 995), and the same mechanisms also recognize cancer cells.
- Peptides not presented by healthy cells or present in very small numbers, but presented on the cell surface of cancer cells are recognized by T cells as non-self antigens. The cancer cell concerned is then eliminated by the T cells (Berg et al., 2015, p. 996).
- Such a peptide can be regarded as immunogenic.
- T cells carrying TCRs specific for non-self antigen peptides are allowed to mature, whereas T lymphocytes carrying TCRs specific for self-antigens are eliminated by clonal deletion or by a process called anergy, unless the respective T lymphocyte is an unaware cell. As a result, there is no immune response against self-antigens.
- TCRs The genetic architecture of TCRs resembles that of antibodies (Berg et al., 2015, p. 998). Combinations of J gene segments, D gene segments, and V gene segments for example lead to the formation of the variable region. As a result, at least 10 15 different specificities can arise from combinations of this repertoire of genes (Lythe, G, et al., Journal of Theoretical Biology (2016) 389, 214-224. Similar to the situation for antibodies, the combination of rearrangement of gene segments and RNA splicing leads to a diversity that ensures coverage of any potential antigenic epitope.
- a T cell that carries a TCR that is capable of binding a peptide that contains a sequence of one of SEQ ID NOs: 1 to 59, or consists of a sequence of one of SEQ ID NOs: 1 to 59 exists in an organism, e.g. in a blood sample. If blood from a plurality of donors is collected, it is furthermore highly likely that a T lymphocyte carrying a TCR specific for a peptide that contains a sequence of one of SEQ ID NOs: 1 to 59, or consists of a sequence of one of SEQ ID NOs: 1 to 59, can be identified.
- tumor cells are therefore generally eliminated by T lymphocytes that bind peptides that are presented on cancer cells, but not, or in very low numbers, on healthy cells. It is nevertheless possible that the cellular immune response fails.
- the initially formed cancer cells may not have been completely destroyed due to an initially low number of T cell clones specific against the particular peptide presented, and subsequently tumor infiltration may for instance be limited.
- An inhibiting tumor microenvironment may also have formed. In such cases a tumor has developed.
- a T cell-based therapy has therefore become a promising approach for an individual suffering from cancer.
- a peptide as disclosed herein has been isolated in a form bound to an MHC molecule from cancer cells and identified by way of mass spectrometric analysis. For this purpose, a tumor tissue sample or a tumor cell sample has been lysed. The peptide/MHC complex has then been isolated from the lysed sample by way of immunoprecipitation.
- Binding of a peptide to an MHC complex has been tested by methods known in the art. Binding of two potential binding partners can for example generally be tested by immobilizing one of the binding partners and/or by detectably labelling one of the binding partners. ELISA-based binding and competition techniques have for instance been developed for the detection of peptide/MHC binding. Immunoprecipitation, for example with a conformation dependent monoclonal antibody, or cell-based detection are additional options available in this regard.
- the antibody used for immunoprecipitation was conformation-specific.
- the antibody captured only folded MHC class I complexes.
- An antibody specific for the MHC molecule to which the peptide is bound has been used.
- peptide/MHC complexes bound to an antibody were washed to remove unbound peptides. MHC-bound peptides were then released from MHC.
- the discovery platform XPRESIDENT® v2.1 allows the identification and selection of relevant over-presented peptides which are potential targets for immunotherapy based on direct relative quantitation of HLA restricted peptide levels on cancer tissues in comparison to several different non-cancerous tissues and organs. See e.g. WO 2011/128448, U.S. 9791443 and U.S.
- a presentation profile was calculated showing the median sample presentation as well as replicate variation.
- the profile juxtaposes samples of the tumor entity of interest to a baseline of normal cell samples or tissue samples.
- Each of these profiles was then consolidated into an over-presentation score by calculating the p-value of a Linear Mixed-Effects Model (Pinheiro et al., 2015) adjusting for multiple testing by False Discovery Rate (Benjamini and Hochberg, 1995, the content of which is incorporated by reference in its entirety).
- MHC molecules specifically HLA molecules
- shock-frozen cell samples or tissue samples were purified and HLA associated peptides were isolated.
- the isolated peptides were separated and sequences were identified by online nano- electrospray-ionization (nanoESI) liquid chromatography-mass spectrometry (LC-MS) experiments.
- nanoESI nano- electrospray-ionization
- LC-MS liquid chromatography-mass spectrometry
- the resulting peptide sequences were verified by comparison of the fragmentation pattern of natural peptides (also referred to as TUMAPs) recorded from acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastroesophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer samples (N >750 samples) with the fragmentation patterns of corresponding synthetic reference peptides of identical sequences.
- TUMAPs fragmentation pattern of natural peptides
- the peptide, that is bound to the MHC is itself not recognized by the antibody, it relies on the structural integrity of the MHC complex to ensure binding to the antibodies used for immunopurification.
- the MHC class molecule is not stable without a peptide. If there is no peptide bound to the MHC class I peptide binding groove, MHC class I conformation is lost. As a result, the epitope recognized by the antibody (e.g. BB7.2) is no longer available and immunoprecipitation of MHC class I is unsuccessful. Therefore, isolation of peptides is not possible if the peptide is not bound to MHC class I binding groove and only MHC-class I - bound peptides can be detected using mass spectrometry.
- All peptides disclosed in the present application were identified and their binding to MHC was shown with this approach on acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, nonHodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer samples confirming their presentation on acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous
- the method assumes that LC-MS signal areas of a peptide correlate with its abundance in the sample. All quantitative signals of a peptide in various LC-MS experiments were normalized based on central tendency, averaged per sample and merged into a bar plot, called presentation profile.
- presentation profile consolidates different analysis methods like protein database search, spectral clustering, charge state deconvolution (decharging) and retention time alignment and normalization.
- RNA expression of the underlying gene was tested.
- mRNA data were obtained via RNASeq analyses of normal tissues and cancer tissues (see Example 2, Figures 2A-2G).
- An additional source of normal tissue data was a database of publicly available RNA expression data from around 3000 normal tissue samples (Lonsdale, 2013, the content of which is incorporated by reference in its entirety).
- a peptide as disclosed herein is derived from a protein the coding mRNA of which is highly expressed in cancer tissue, but very low or absent in healthy tissues.
- MHC Class I binding peptides are presented on virtually any nucleated cell. MHC Class I binding peptides are generated mainly following degradation of intracellular proteins by the ubiquitin-proteasome system of the respective cell.
- MHC ligandome is influenced by cancer-specific alterations affecting, among others, translational regulation, protein metabolism, proteasomal processing, cytosolic proteolysis, but also alterations in antigen processing. For example, the expression of chaperones is often altered in cancer cells.
- identifying peptides presented by MHC requires their isolation from cancer tissues. Identifying peptides presented by MHC only on cancer tissue furthermore requires the analysis of both healthy tissue and cancer tissue samples. Thousands of cancer tissue and healthy tissue samples need to be collected, prepared, measured, and the data analysed in a highly reproducible manner.
- sample preparation might introduce proteolytic fragments mimicking HLA ligands, which can be assessed by in silico methods. Chromatographic procedures pose the risk of peptide carryover between samples, which can be monitored by blank runs.
- electrospray ionization MS generates in-source fragments that can be identified computationally.
- the peptide sequencing is prone to false-positive identifications that can only be uncovered by experimental sequence validation through synthetic standards.
- peptides When identifying peptides for cancer therapy, potential peptides cannot simply be reduced to those peptides with low false discovery rate, since this would limit the sensitivity of identifying veritable peptides. Additionally, in other applications, the top- ranking peptide-spectrum match will usually be correct when identifying a peptide.
- An example for such other application is the commonly used approach for MS-based proteomic analysis where trypsin is used to digest proteins into tryptic peptides, i.e., peptides with either arginine or lysine at their C-terminus.
- a peptide found may actually not be a true MHC ligand since in-source fragmentation can occur.
- the mass spectrometry techniques of electrospray ionization may generate ions that might be selected for acquisition of MS/MS spectra that may be mistaken for MHC ligands.
- C-terminal source fragments may be falsely identified as MHC ligands. Eliminating such peptides from consideration requires close attention and experience.
- the parental peptides that the generated fragment are derived from are not detected upon fragmentation any longer and can lead to false negative results.
- Peptides not related to presentation by MHC can be generated as a result of proteolytic cleavage by endogenous proteases and peptidases. These enzymes originate from the sample analysed. This mechanism can lead to the false positive generation of peptides that are not presented by cells in vivo as well as missing identifications of relevant peptides due to degradation. Approaches for detection of such contaminations use a statistical analysis of the protein coverage as metric to exclude problematic proteins. This requires detailed analysis of large data sets and, again, close human attention going far beyond any routine experimentation.
- peptides comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 1 to SEQ ID NO: 59, or a pharmaceutically acceptable salt thereof.
- the peptides disclosed bind to inter alia HLA-A*01 , HLA-A*02, HLA-A*03, HLA-A*11 , HLA-A*23, HLA-A*68, HLA-B*07, HLA-B*08, HLA-B*13, HLA-B*14, HLA-B*15, HLA- B*18, HLA-B*27, HLA-B*35, HLA-B*37, HLA-B*38, HLA-B*40, HLA-B*44, HLA-B*51 , HLA-B*57, and HLA-C*03 allotype MHC molecules.
- said peptide thereof has a length according to the respective SEQ ID NO: 1 to SEQ ID NO: 59.
- the peptide consists of the amino acid sequence according to SEQ ID NO: 1 to SEQ ID NO: 59.
- Table 1 shows the amino acid sequences of the peptides disclosed herein and their respective SEQ ID NOs.
- a peptide disclosed herein can include up to four amino acids at the N- or the C-terminus in any combination between 4:0 and 0:4, relative to an amino acid sequence provided herein. That is, for example 1 or 2 amino acids can be added to either end. Also, amino acids can be added to either end. Combinations of possible elongations can be found in Table 2.
- the amino acids for the elongation/extension can be the peptides of the original sequence of the protein or any other amino acid(s).
- the elongation can be used to enhance the stability or solubility of the peptides.
- a peptide sequence used herein may be identical to a naturally occurring tumor- associated or tumor specific peptide sequence.
- the peptide is elongated on either or both ends by more than 4 amino acids, such as to a total length of up to 30 amino acids. This may lead to MHC class ii binding peptides. Binding to MHC class ii can be tested by methods known in the art.
- a peptide provided herein may have a maximal length of up to 30 amino acids, including a maximal length of up to 20 amino acids. In some embodiments a peptide provided herein may have a maximal length of up to 18, including of up to 15 amino acids. In some embodiments a peptide provided herein may have a maximal length of up to 14, including of up to 13 amino acids. In some embodiments a peptide provided herein may have a minimal length of 8, including of 9 amino acids. In some embodiments a peptide provided herein may have a minimal length of 10 amino acids.
- a peptide provided herein may have an overall length from 8 to 30 amino acids, including from 8 to 13 amino acids. In some embodiments a peptide provided herein may have an overall length from 10 to 16 amino acids, including from 10 to 12 amino acids.
- a peptide provided herein may have a length of 8 or 9 amino acids. In some embodiments a peptide provided herein may have a length of 10 or 11 amino acids. In some embodiments a peptide provided herein may have a length of 12 or 13 amino acids. In some embodiments a peptide provided herein, in particular, in case of an elongated class II binding peptide, may have a length of 14 or 15 amino acids. In some embodiments a peptide provided herein, in particular, in case of an elongated class II binding peptide, may have a length of 16 or 17 amino acids.
- a peptide provided herein, in particular, in case of an elongated class II binding peptide may have a length of 18 or 19 amino acids. In some embodiments a peptide provided herein, in particular, in case of an elongated class II binding peptide, may have a length of 20 or 21 amino acids. In some embodiments a peptide provided herein, in particular, in case of an elongated class II binding peptide, may have a length of 22 or 23 amino acids.
- a peptide provided herein in addition to the sequence according to any of SEQ ID NO: 1 to SEQ ID NO: 59 contains one or more further N- and/or C-term inally located stretches of amino acids that are not necessarily forming part of the peptide that functions as an epitope for MHC molecules.
- the peptide is part of a fusion protein which includes, for example, the 80 N-terminal amino acids of the HLA-DR antigen-associated invariant chain (p33, in the following “Ii”) as derived from the NCBI, GenBank Accession number X00497.
- the peptides provided herein can be fused to an antibody as described herein, or a functional part thereof, in particular, into a sequence of an antibody, so as to be specifically targeted by said antibody, or, for example, to or into an antibody that is specific for dendritic cells as described herein.
- a peptide as disclosed herein when bound by a member of the immune response or by a binding molecule can be taken to define an antigenic epitope (Murphy, 2022).
- the peptide may be modified further to improve stability and/or binding to MHC molecules in order to elicit a stronger immune response.
- Methods for such an optimization of a peptide sequence are well known in the art and include, for example, the introduction of reverse peptide bonds or non-peptide bonds.
- Non-peptide bond is a reverse peptide bond, in which amino acid residues are not joined by peptide (-CO-NH-) linkages but the peptide bond is reversed.
- retro-inverse peptidomimetics may be made using methods known in the art, for example such as those described by Meziere and colleagues (Meziere et al., 1997, incorporated herein by reference). This approach involves making pseudopeptides containing changes involving the backbone, and not the orientation of side chains. They show that for MHC binding and T helper cell responses, these pseudopeptides are useful. Retro-inverso peptides, which contain D-amino acids in the reverse sequence, are much more resistant to proteolysis (Meziere et al., 1997).
- the peptide as disclosed herein may in some embodiments include one or more non- peptide bonds.
- a non-peptide bond is, for example, -CH2-NH or -CH2S-.
- Two further examples of a non- peptide bond are -COCH2- or -CH(OH)CH2-.
- a further example of a non-peptide bond is -CH2SO-.
- a further example of a non-peptide bond is an N-methyl or an N-hydroxyl group contained in the bond between two amino acids.
- US 4,897,445 provides a method for the solid phase synthesis of non-peptide bonds (-CH2-NH) in polypeptide chains which involves polypeptides synthesized by standard procedures and the non-peptide bond synthesized by reacting an amino aldehyde and an amino acid in the presence of NaCNBHs.
- Peptides comprising the bonds described above may be synthesized with additional chemical groups present at their amino and/or carboxy termini, to enhance the stability, bioavailability, and/or affinity of the peptides.
- additional chemical groups such as carbobenzoxyl, dansyl, or t-butyloxycarbonyl groups may be added to the peptides' amino termini.
- an acetyl group or a 9-fluorenylmethoxy-carbonyl group may be placed at the peptides' amino termini.
- the hydrophobic group, t- butyloxycarbonyl, or an amido group may be added to the peptides' carboxy terminus.
- the peptides disclosed herein comprise no non-peptide bonds. In other words, the peptides disclosed herein in some embodiments comprise only peptide bonds.
- a peptide as provided herein may be provided in the form of a salt, in particular a pharmaceutically acceptable salt.
- the peptide may be provided in the form of a pharmaceutically acceptable salt. Additionally, the peptide may be delivered using a sustained-release system.
- a non-naturally occurring peptide has been synthetically produced (e.g. synthesized) as a pharmaceutically acceptable salt.
- Methods to synthetically produce peptides are well known in the art. See below.
- a salt of a peptide provided herein differs substantially from the respective peptide in its state in vivo, as the peptide is not in the form of a salt or associated with counterions in vivo.
- a salt may be provided to increase the shelf-life of a peptide provided herein.
- a salt may also be provided to alter the solubility of a peptide provided herein in an aqueous solvent, including saline or water.
- the salt form of the peptide mediates the solubility of the peptide, in particular in the context of pharmaceutical compositions comprising a peptide as disclosed herein.
- a sufficient and at least substantial solubility of the peptide(s) is required in order to efficiently provide the peptides to the subject to be treated.
- a salt as used herein is a pharmaceutically acceptable salt of the corresponding peptide.
- An acid salt may be prepared by contacting the peptide with a suitable acid.
- Suitable acids for preparing an acid salt include both an organic acid and an inorganic acid.
- a suitable organic acid may in some embodiments be a carboxylic acid.
- Two examples of a suitable organic acid are acetic acid and propionic acid.
- Further examples of a suitable organic acid are glycolic acid and pyruvic acid.
- Oxalic acid and malic acid are yet further examples of a suitable organic acid.
- Two further organic acids that may be used are malonic acid and succinic acid.
- Yet two further examples of a suitable organic acid are maleic acid and fumaric acid.
- Further examples of a suitable organic acid are tartaric acid and citric acid.
- Benzoic acid and cinnamic acid are further examples of a suitable organic acid.
- mandelic acid and salicylic acid are suitable organic acids.
- Yet two further suitable organic acids are methane sulfonic acid and an aryl sulfonic acid.
- methane sulfonic acid and an aryl sulfonic acid Two examples of an aryl sulfonic acid are benzenesulfonic acid and p-toluenesulfonic acid.
- An example of a suitable organic acid is also ethane sulfonic acid.
- a suitable inorganic acid is hydrochloric acid and hydrobromic acid Sulfuric acid and nitric acid are further examples of a suitable inorganic acid. Yet a further example of a suitable inorganic acid is phosphoric acid.
- preparation of a basic salt of an acid moiety which may be present on a peptide may be prepared using a pharmaceutically acceptable base.
- a pharmaceutically acceptable base may for instance be an inorganic base or an organic base.
- Two suitable inorganic bases are sodium hydroxide and potassium hydroxide.
- Yet two further suitable inorganic bases are ammonium hydroxide and calcium hydroxide.
- Two suitable organic bases are trimethylamine and triethylamine.
- a salt as provided herein may be an alkaline or an earth alkaline salt, such as a salt of the Hofmeister series.
- the salt may include PO4 3 or SCU 2 ’.
- Two further suitable anions are CHsCFteCOO and HOCHsCOO.
- Yet two further suitable anions are Cl’ and Br. NO 3 ’ and CIO4 are two further suitable anions.
- Two further suitable anions are I’ and SCN’.
- Two suitable cations are NH4 + and Rb + .
- Two further suitable cations are K + and Na + .
- Cs + and Li + are two further suitable cations.
- Two further suitable cations are Zn 2+ and Mg 2+ .
- Yet two further suitable cations are Ca 2+ and Mn 2+ .
- Cu 2+ and Ba 2+ are two suitable cations.
- Particularly salts are selected from (NH4)3PO4, (NH4)2HPO4, (NH4)H2PO4, (NH4)2SO4, NH4CH3COO, NH4CI, NH 4 Br, NH4NO3, NH4CIO4, NH 4 I, NH4SCN, Rb 3 PO 4 , Rb 2 HPO 4 , RbH 2 PO 4 , Rb 2 SO 4 , Rb 4 CH 3 COO, Rb 4 CI, Rb 4 Br, Rb 4 NO 3 , Rb 4 CIO 4 , Rb 4 l, Rb 4 SCN, K3PO4, K2HPO4, KH2PO4, K2SO4, KCH3COO, KCI, KBr, KNO3, KCIO4, KI, KSCN, Na 3 PO 4 , Na 2 HPO 4 , NaH 2 PO 4 , Na 2 SO 4 , NaCHsCOO, NaCI, NaBr, NaNOs, NaCICU, Nal, NaSCN, ZnCI 2 Cs
- the salt is ammonium acetate, MgCh, KH2PO4, Na2SO4, KCI, NaCI, and CaCh, such as, for example, the chloride or acetate (trifluoroacetate) salts (see e.g. Berge, S.M., et al., Journal of Pharmaceutical Science (1977) 66, 1 -19, the contents of which are incorporated by reference in their entirety).
- a pharmaceutically acceptable salt can be taken to be a derivative of a peptide as disclosed herein, wherein the peptide is modified by making an acid or base salt of the agent.
- acid salts are prepared from the free base (typically wherein the neutral form of the drug has a neutral -NH2 group) involving reaction with a suitable acid.
- suitable acids for preparing acid salts include both organic acids e.g.
- acetic acid trifluoro acetic acid, propionic acid, glycolic acid, pyruvic acid, oxalic acid, malic acid, malonic acid, succinic acid, maleic acid, fumaric acid, tartaric acid, citric acid, benzoic acid, cinnamic acid, mandelic acid, methane sulfonic acid, ethane sulfonic acid, p- toluenesulfonic acid, salicylic acid, and the like, as well as inorganic acids, e.g., hydrochloric acid, hydrobromic acid, sulfuric acid, nitric acid phosphoric acid and the like.
- inorganic acids e.g., hydrochloric acid, hydrobromic acid, sulfuric acid, nitric acid phosphoric acid and the like.
- a pharmaceutical composition contains one or more peptides as salts of acetic acid (acetates), trifluoro acetates or hydrochloric acid (chlorides).
- a peptide as disclosed herein has been chemically synthesized.
- peptides may be synthesized by the Fmoc-polyamide mode of solid-phase peptide synthesis as disclosed by Lukas et al. (Lukas et al., 1981 , the content of which is incorporated by reference in its entirety) and by references as cited therein.
- Temporary N-amino group protection is afforded by the 9- fluorenylmethyloxycarbonyl (Fmoc) group. Repetitive cleavage of this highly base-labile protecting group is done using 20% piperidine in N,N-dimethylformamide.
- Side-chain functionalities may be protected as their butyl ethers (in the case of serine, threonine and tyrosine), butyl esters (in the case of glutamic acid and aspartic acid), butyloxycarbonyl derivative (in the case of lysine and histidine), trityl derivative (in the case of cysteine) and 4-methoxy-2,3,6-trimethylbenzenesulphonyl derivative (in the case of arginine).
- glutamine or asparagine are C-terminal residues, use is made of the 4,4'- dimethoxybenzhydryl group for protection of the side chain amido functionalities.
- the solid-phase support is based on a polydimethyl-acrylamide polymer constituted from the three monomers dimethylacrylamide (backbone-monomer), bisacryloylethylene diamine (cross linker) and acryloylsarcosine methyl ester (functionalizing agent).
- the peptide-to- resin cleavable linked agent used is the acid-labile 4-hydroxymethyl-phenoxyacetic acid derivative. All amino acid derivatives are added as their preformed symmetrical anhydride derivatives with the exception of asparagine and glutamine, which are added using a reversed coupling procedure involving N,N-dicyclohexyl-carbodiimide and 1 - hydroxybenzotriazole.
- Trifluoroacetic acid may be removed by evaporation in vacuo, with subsequent trituration with diethyl ether affording the crude peptide.
- Any scavengers present can be removed by a simple extraction procedure which on lyophilization of the aqueous phase affords the crude peptide free of scavengers.
- Reagents for peptide synthesis are generally available from e.g., Calbiochem-Novabiochem (Nottingham, UK).
- Purification may involve any desired technique, such as re-crystallization, size exclusion chromatography, ion-exchange chromatography, hydrophobic interaction chromatography and (usually) reverse-phase high performance liquid chromatography using e.g., aceton itrile/water gradient separation.
- Analysis of synthesized peptides may be carried out using thin layer chromatography, electrophoresis, in particular capillary electrophoresis, solid phase extraction (CSPE), reverse-phase high performance liquid chromatography, amino-acid analysis after acid hydrolysis and by fast atom bombardment (FAB) mass spectrometric analysis, as well as MALDI and ESI-Q-TOF mass spectrometric analysis.
- electrophoresis in particular capillary electrophoresis
- CSPE solid phase extraction
- FAB fast atom bombardment
- MALDI and ESI-Q-TOF mass spectrometric analysis as well as MALDI and ESI-Q-TOF mass spectrometric analysis.
- the method includes culturing a host cell such as a recombinant host cell that expresses or contains the peptide.
- the method furthermore includes isolating the peptide, from the host cell and/or from the culture medium that contains the host cell.
- a cDNA encoding a peptide according as disclosed herein can be expressed in prokaryotic cells (e.g., bacteria) or eukaryotic cells (e.g., yeast, insect, or mammalian cells), after which the recombinant peptide can be purified.
- prokaryotic cells e.g., bacteria
- eukaryotic cells e.g., yeast, insect, or mammalian cells
- a method of treating a subject having cancer or of preventing a subject from suffering from cancer includes administering to the subject a peptide as provided herein.
- the method may include administering to a subject a pharmaceutically effective amount of the peptide.
- a method of treating a subject having cancer or of preventing a subject from suffering from cancer includes administering to the subject a binding molecule as provided herein.
- the method may include administering to a subject a pharmaceutically effective amount of the binding molecule.
- a method of treating a subject having cancer or of preventing a subject from suffering from cancer includes administering to the subject a nucleic acid or an expression vector as provided herein.
- the method may include administering to a subject a pharmaceutically effective amount of the nucleic acid or the expression vector.
- a method of treating a subject having cancer or of preventing a subject from suffering from cancer includes administering to the subject a recombinant host cell, including a plurality of recombinant host cells, as provided herein.
- the method may include administering to a subject a pharmaceutically effective amount of the host cell(s).
- a method of treating a subject having cancer or of preventing a subject from suffering from cancer includes administering to the subject an activated T lymphocyte, including a plurality of activated T lymphocytes, as provided herein.
- the method may include administering to a subject a pharmaceutically effective amount of the activated T lymphocyte(s).
- a method of treating a subject having cancer or of preventing a subject from suffering from cancer includes administering to the subject a pharmaceutical composition or a combination product as provided herein.
- the method may include administering to a subject a pharmaceutically effective amount of the pharmaceutical composition or the combination product.
- a method of treating a subject having cancer or of preventing a subject from suffering from cancer includes administering to the subject a composition that contains a population of activated T cells that kill cancer cells in the subject that over-present a peptide as described herein on their cell surface.
- the method may include administering to a subject a pharmaceutically effective amount of the population of activated T cells.
- Any peptide or pharmaceutically acceptable salt thereof as disclosed herein may be used in medicine.
- any binding molecule as disclosed herein may be used in medicine.
- Any nucleic acid and any expression vector that contains such a nucleic acid as disclosed herein may be used in medicine.
- any host cell and any T lymphocyte as disclosed herein may be used in medicine.
- Any cell line as disclosed herein may also be used in medicine.
- any pharmaceutical composition as disclosed herein may be used in medicine.
- Any kit or combination product as disclosed herein may also be used in medicine.
- a peptide as disclosed herein may for example be used in generating a binding molecule, such as a TCR, a TCR fragment, an antibody, an antibody fragment or an aptamer.
- the binding molecule may then be administered to an individual.
- the binding molecule may be coupled to a toxin, in particular a compound having cytotoxic effect as illustrated below.
- the binding molecule may for example be coupled to an RNA polymerase inhibitor or to an RNA-splicing inhibitor.
- the antibody may for example be coupled to a toxin via a cleavable linker.
- a peptide as disclosed herein may also be administered to a subject in order to induce or stimulate an immune response.
- a host cell may be used to produce a proteinaceous binding molecule or a peptide as disclosed herein.
- the proteinaceous binding molecule or the peptide may then be administered to a subject.
- a nucleic acid encoding a peptide as disclosed herein may be used for preventing occurrence of a cancer or for treating a cancer.
- the nucleic acid may be included in a vector, such as an expression vector.
- the nucleic acid, including the vector may be designed to enter cells and transcribe, translate and process a peptide as disclosed herein.
- the respective vector can in some embodiments be taken to define a recombinant viral or bacterial vaccine.
- a T lymphocyte or a T cell line may be directly administered to a subject.
- the T lymphocyte may for example be autologous to the subject.
- the T lymphocyte may also be allogenic.
- a T lymphocyte used may be obtained from a healthy donor.
- the T lymphocyte may be derived from a tumor infiltrating lymphocyte or from a peripheral blood mononuclear cell.
- the T lymphocyte may be recombinant in that it has been genetically engineered to express a TCR of a desired specificity or affinity to a peptide as disclosed herein.
- the method includes administering to a subject suffering from cancer an effective amount of a peptide or a salt thereof, a binding molecule, a nucleic acid, an expression vector, a host cell, a T lymphocyte as disclosed herein or any combination thereof.
- peptide or the salt thereof as disclosed herein for use in the diagnosis of cancer.
- nucleic acid, or the vector that contains the nucleic acid, as disclosed herein for use in the diagnosis of cancer are furthermore the host cell or the T lymphocyte as disclosed herein for use in the diagnosis of cancer.
- pharmaceutical composition or combination product as disclosed herein for use in the diagnosis of cancer.
- kit as disclosed herein for use in the diagnosis of cancer.
- a use in the diagnosis of cancer may also be carried out in the form of an in vitro method.
- Diagnosing cancer may also be carried out in the form of an in vivo method
- the method may be an in vitro method carried out on one or more samples that have been obtained from the respective subject.
- the method includes detecting whether a peptide as disclosed herein is presented on cells of the sample.
- a binding molecule as disclosed herein may be used, for example in labelled form.
- the method may include detecting the presence of the peptide, or the relative amount of the peptide, relative to a control.
- the control may be defined by one or more healthy samples of the same tissue type or cell type.
- the method may be an in vitro method carried out on one or more cancer samples that have been obtained from the respective subject.
- the method includes detecting whether a peptide as disclosed herein is presented on cells of the sample.
- the method may also include estimating or assessing whether a peptide as disclosed herein is presented on cells of the sample.
- a binding molecule as disclosed herein may be used.
- the method may include detecting the presence of the peptide, or the relative amount of the peptide, relative to a control.
- the control may be defined by one or more healthy samples of the same tissue type or cell type.
- the method includes administering to the subject a plurality of T lymphocytes as disclosed herein.
- the T lymphocytes may be activated T lymphocytes.
- T lymphocytes kill cancer cells in the subject that present a peptide as disclosed herein.
- the T lymphocytes may in some embodiments be autologous to the subject.
- the T lymphocytes may in some embodiments be allogenic.
- the T lymphocytes used may be obtained from a healthy donor.
- the T lymphocytes may be derived from tumor infiltrating lymphocytes or from peripheral blood mononuclear cells.
- the T lymphocytes may be recombinant in that they have been genetically engineered to express a TCR of a desired specificity or affinity to a peptide as disclosed herein.
- the cancer cells present a peptide as disclosed herein, bound by MHC.
- the method includes administering to the subject an effective number of T lymphocytes as disclosed herein.
- the T lymphocytes may be activated T lymphocytes.
- a method of killing cancer cells may also be carried out in the form of an in vitro method.
- a method of killing cancer cells may also be carried out in the form of an in vivo method.
- Provided is furthermore an activated T lymphocyte as disclosed herein for use in the killing of target cells in a subject.
- the target cells present a peptide that contains any amino acid sequence disclosed herein.
- the respective target cells are cancer cells.
- T lymphocyte as disclosed herein for use in the manufacture of a medicament for the killing of cancer cells in a subject.
- the cancer cells present a peptide as disclosed herein, bound by MHC.
- the T lymphocytes may be activated T lymphocytes.
- T lymphocyte as disclosed herein in the killing of target cells in a subject.
- the cancer cells present a peptide as disclosed herein, bound by MHC.
- the T lymphocytes may be activated T lymphocytes.
- the cancer may be acute myeloid leukemia or breast cancer.
- the cancer may be cholangiocellular carcinoma or chronic lymphocytic leukemia.
- the cancer may in some embodiments be colorectal cancer or gallbladder cancer.
- the cancer may be glioblastoma.
- the cancer may in some embodiments be gastric cancer or gastroesophageal junction cancer.
- the cancer may be hepatocellular carcinoma or head and neck squamous cell carcinoma.
- the cancer may be melanoma.
- the cancer may in some embodiments be non-Hodgkin lymphoma.
- the cancer may in some embodiments be non-small cell lung cancer or small cell lung cancer.
- the cancer may be ovarian cancer or uterine endometrial cancer.
- the cancer may in some embodiments be esophageal cancer.
- the cancer may in some embodiments be pancreatic cancer or prostate cancer.
- the cancer may be renal cell carcinoma or urinary bladder carcinoma.
- the cancer may also be another cancer that shows an over-presentation of a peptide as disclosed herein.
- a method of diagnosing cancer in a subject includes using a binding molecule as provided herein.
- a binding molecule is a proteinaceous binding molecule that specifically binds to a peptide as provided herein, or to the respective peptide when in complex with an MHC molecule, see above.
- the method may include administering the binding molecule to a subject or contacting the binding molecule with a sample that has been obtained from a subject.
- the method involves targeting cells, including cells as part of tissue, from or in the subject with a binding molecule as disclosed herein.
- the method includes providing a sample from the subject.
- the method furthermore includes detection of binding of the binding molecule to cells, for example cells in one or more tissue samples, of or in the subject.
- detecting binding of the binding molecule generally the presentation of a peptide as disclosed herein on cells originating from the subject is identified.
- the peptide and/or the TCR, antibody, aptamer or other binding molecule provided herein can be used in the diagnosis of cancer.
- the peptide or binding molecule can be used to verify a pathologist’s diagnosis of a cancer based on a sample, e.g. a biopsied sample.
- the cells originating from the subject may be cells of a tissue or of a body fluid obtained from the subject.
- a biopsy of a tumor or a cancerous body fluid sample is collected and subjected to immunoprecipitation of peptide MHC complexes, with subsequent analysis of the peptidome thus obtained by means of mass spectrometry.
- Respective methods are e.g. disclosed in Fritsche et al. (Fritsche, J, et al., Proteomics (2016) 18, 1700284), the content of which is incorporated herein by reference.
- a labelled binding molecule e.g. a T cell receptor or a TCR mimetic antibody, specific for the peptide MHC complex comprising a peptide as disclosed herein.
- a biopsy or sample of the cancer tissue is obtained, rated with a desired immunological method (e.g. sliced or homogenized in case of a solid tumor) and then incubated with the labelled binding molecule. See e.g. Hoydahl et al. (Hoydahl, LS, et al., Antibodies (Basel), (2019) 8, 2, 32, doi:10.3390/antib8020032) for methods, the content of which is incorporated herein by reference.
- an antibody, TCR, aptamer or other binding molecule may also be used in both in vitro and in vivo diagnostic assays.
- the binding molecule is labeled with a radionucleotide (such as 111 1n, "Tc, 14 C, 131 l, 3 H, 32 P or 35 S) so that the tumor can be localized using immunoscintiography.
- the peptides were generated from acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, nonHodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer cells and since it was determined that these peptides are not or at lower levels present in normal tissues, these peptides can be used to diagnose the presence of a cancer.
- a peptide as disclosed herein on cells of a cancer sample can assist a pathologist in diagnosis of cancer.
- Detection of certain peptides by means of antibodies, mass spectrometry or other methods known in the art can tell the pathologist that the tissue sample is malignant or inflamed or generally diseased, or can be used as a biomarker for acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial
- peptides on diseased tissue specimen can enable the decision about the benefit of therapies involving the immune system, especially if T-lymphocytes are known or expected to be involved in the mechanism of action.
- Loss of MHC expression is a well described mechanism by which infected or malignant cells escape immunosurveillance. Thus, presence of peptides shows that this mechanism is not exploited by the analyzed cells.
- an in vitro method of assessing the risk of a subject to have or to develop cancer includes adding a binding molecule, such as a proteinaceous binding molecule, to a sample that contains cells of from said subject.
- the binding molecule is labeled with a probe or a radionucleotide.
- the method furthermore includes detecting whether the binding molecule binds to cells in the sample from the subject.
- the binding molecule specifically binds to (i) a peptide that includes the amino acid sequence according to any one of SEQ ID NOs: 1 to 59 or (ii) said peptide that includes the amino acid sequence according to any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule.
- the detection of binding of the binding molecule to cells in the sample from the subject identifies the presentation of said peptide on said cells. The detection thereby indicates a risk of the subject to have or to develop cancer.
- a peptide provided herein may be used to monitor and analyze lymphocyte responses against those peptides such as T cell responses or antibody responses against the peptide or the peptide complexed to MHC molecules. These lymphocyte responses can be used as prognostic markers for decision on further therapy steps. These responses can also be used as surrogate response markers in immunotherapy approaches aiming to induce lymphocyte responses by different means, e.g. vaccination of protein, nucleic acids, autologous materials, adoptive transfer of lymphocytes. In gene therapy settings, lymphocyte responses against peptides can be considered in the assessment of side effects. Monitoring of lymphocyte responses might also be a valuable tool for follow-up examinations of transplantation therapies, e.g. for the detection of graft versus host and host versus graft diseases.
- the method includes using a binding molecule, such as a proteinaceous binding molecule, as provided herein.
- the method may include administering the binding molecule to a subject or contacting the binding molecule with a sample that has been obtained from a subject.
- the method involves targeting cells, including cells as part of tissue, from or in the subject with a binding molecule as disclosed herein.
- the method includes providing a sample from the subject.
- the method furthermore includes detection of binding of the binding molecule to cells, for example cells in one or more tissue samples, of or in the subject.
- targeting cells from or in the subject with the binding molecule is performed at certain, e.g. predetermined, time intervals. Samples from the subject may be provided that have been obtained at the corresponding time points.
- a peptide as disclosed herein is over-presented in cancer tissue compared to healthy tissues (see Table 6) and thus can be used for the production of antibodies and/or TCRs, such as soluble TCRs, as disclosed herein for use in the treatment of cancer, or for the production of vaccines against cancer.
- the peptides when complexed with the respective MHC can be used for the production of an antibody and/or a TCR, for example a soluble TCR for use in the treatment of cancer.
- Respective methods are well known to the person of skill in the art and can be found in the respective literature as well (see also below).
- peptides that are useful in treating cancers/tumors, such as acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastroesophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer that overpresent a peptide as disclosed herein.
- cancers/tumors such as acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastroesophageal junction cancer, hepat
- peptides were shown by MS to be naturally presented by MHC molecules on primary human acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer samples.
- MHC bound peptides can be recognized by the immune system, specifically T lymphocytes.
- T cells can destroy the cells presenting the recognized MHC-peptide complex, e.g. acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer cells presenting the derived peptides.
- acute myeloid leukemia breast cancer
- cholangiocellular carcinoma chronic lymphocytic leukemia
- colorectal cancer gallbladder cancer
- glioblastoma gas
- nucleic acid molecule may contain a sequence that encodes a peptide as disclosed herein.
- the sequence that encodes the peptide may be linked to a heterologous promoter sequence.
- the sequence that encodes the peptide may be operatively coupled to a heterologous promoter sequence.
- nucleic acid molecule contains an open reading frame (ORF) that contains a sequence coding for a peptide as disclosed herein.
- the nucleic acid molecule may in some embodiments contain a sequence that encodes a proteinaceous binding molecule as disclosed herein.
- the sequence that encodes the proteinaceous binding molecule may be linked to a heterologous promoter sequence.
- the sequence that encodes the proteinaceous binding molecule may be operatively coupled to a heterologous promoter sequence.
- a respective nucleic acid may for instance be a DNA molecule or a cDNA molecule.
- a respective nucleic acid may also be a PNA molecule or an RNA molecule.
- a respective nucleic acid may be a combination of one or more of a DNA molecule, a cDNA molecule, a PNA molecule and/or an RNA molecule.
- the nucleic acid molecule is an artificial nucleic acid molecule.
- the nucleic acid is included in a vector, such as an expression vector.
- the sequence encoding a peptide may be operably linked to one or more regulatory sequences such as a promoter.
- the sequence encoding a proteinaceous binding molecule may be operably linked to one or more regulatory sequences such as a promoter.
- the vector is in some embodiments an artificial vector.
- the nucleic acid may be complexed or associated with a lipid. In some embodiments the nucleic acid may be encapsulated in a carrier particle that contains a lipid.
- a nucleic acid molecule containing a sequence that encodes a peptide or variant as provided herein may be included in a vaccine composition.
- a respective vaccine composition may for example be used for eliciting or for enhancing an anti-tumor immune response.
- nucleic acid molecule provided herein contains a codon- optimized region. In some embodiments a nucleic acid molecule contains a sequence encoding a peptide or variant thereof or a proteinaceous binding molecule as disclosed herein that is codon-optimized. In some embodiments a nucleic acid molecule as disclosed herein contains an open reading frame that is codon-optimized.
- an expression vector in a further aspect there is provided an expression vector.
- the expression vector includes a nucleic acid as disclosed herein.
- a host cell such as a recombinant host cell, that includes a peptide as disclosed herein.
- a host cell such as a recombinant host cell, that includes a proteinaceous binding molecule, e.g. an antibody or fragment thereof or a T cell receptor or fragment thereof, as provided herein.
- a host cell that includes a nucleic acid as disclosed herein or an expression vector as disclosed herein.
- a respective host cell may be an antigen- presenting cell.
- the host cell is a dendritic cell.
- the host cell is a T cell or an NK cell.
- the host cell is a T lymphocyte or an NK cell expressing a heterologous proteinaceous binding molecule.
- the heterologous proteinaceous binding molecule is in some embodiments a heterologous TCR.
- the heterologous TCR specifically recognizes a peptide comprising an amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising an amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule.
- a pharmaceutical composition contains at least one active ingredient and a pharmaceutically acceptable carrier and/or a pharmaceutically acceptable excipient.
- the active ingredient may be a peptide or salt thereof as disclosed herein.
- the active ingredient may also be a binding molecule as disclosed herein, for instance an antibody, a functional fragment thereof, a T-cell receptor or a functional fragment thereof, or an aptamer.
- the active ingredient may also be a nucleic acid or an expression vector as disclosed herein.
- the active ingredient may also be a recombinant host cell or an T lymphocyte as disclosed herein.
- a respective pharmaceutical composition may be provided for therapeutic purposes, for diagnostic purposes, or for both.
- a pharmaceutical composition provided herein may be a pharmaceutical composition for inducing an immune response in a human or animal.
- the pharmaceutical composition is included in a vaccine. In some embodiments the pharmaceutical composition is a cellular therapeutic composition.
- such a pharmaceutical composition is an individualized, including a personalized, pharmaceutical composition for an individual subject.
- a pharmaceutical composition may contain one or more peptide(s) as disclosed herein either in the free form or in the form of a pharmaceutically acceptable salt (see also above).
- a pharmaceutical composition contains one or more peptides as salts of acetic acid (acetates), trifluoro acetates or hydrochloric acid (chlorides).
- the pharmaceutical composition may contain a stabilizing substance such as those disclosed in WO2023/044455.
- a respective stabilizing substance may for instance be dimethyl sulfoxide (DMSO) or diethyl sulfoxide.
- a respective stabilizing substance may also be dimethyl sulfone or N-Methyl-2-pyrrolidone.
- a supplemental RNA stabilizing substance included such as hexylene glycol or Na-hexametaphosphate.
- Further examples of a supplemental RNA stabilizing substance are trimetaphosphate or Na-Glycerol Phosphate.
- a pharmaceutical composition whether an individualized product or not, that includes an agent as disclosed herein may be manufactured in a manner that is itself known, e.g., by means of a mixing, dissolving, granulating, dragee-making, levigating, emulsifying, encapsulating, entrapping or lyophilizing process.
- the pharmaceutical composition contains a conjugated or labelled active ingredient. In some embodiments the pharmaceutical composition contains an adjuvant. In some embodiments the pharmaceutical composition contains an immunogenicity enhancing amount of an adjuvant. In some embodiments the adjuvant is an interleukin.
- the interleukin may for example be IL-2, IL-12, IL-15, IL-18, IL-21 or a combination thereof. In one embodiment the interleukin may be IL-2. In one embodiment the interleukin may be IL-12. In one embodiment the interleukin may be IL-15. In one embodiment the interleukin may be IL-18. In one embodiment the interleukin may be IL- 21. In some embodiments, the interleukin may be expressed by the T lymphocyte and may be membrane-bound or soluble.
- a pharmaceutical composition e.g. a vaccine composition, as disclosed herein may include one or more adjuvants.
- An adjuvant may also be included in a combination product.
- An adjuvant is a substance that non-specifically enhances or potentiates the immune response e.g. an immune response mediated by CD8-positive T cells and/or helper T (TH) cells to an antigen.
- Suitable adjuvants include, but are not limited to, 1018 ISS, aluminum salts, AMPLIVAX®, AS15, BCG, CP-870,893, CpG7909, CyaA, dSLIM, flagellin or TLR5 ligands derived from flagellin, FLT3 ligand, GM-CSF, IC30, IC31 , Imiquimod (ALDARA®), Resiquimod, ImuFact IMP321 , an Interleukin, Interferon-alpha or -beta, or pegylated derivatives thereof, IS Patch, ISS, ISCOMATRIX, ISCOMs, Juvlmmune®, LipoVac, MALP2, MF59, monophosphoryl lipid A, Montanide IMS 1312, Montanide ISA 206, Montanide ISA 50V, Montanide ISA-51 , water-in-oil and oil-in-water emulsions, OK- 432, OM-174,
- Adjuvants such as Freund's or GM-CSF may be used.
- An example of an interleukin is IL-2.
- a further example of an interleukin is IL-15.
- Yet another example of an interleukin is IL-18.
- a further illustrative example of an interleukin is IL-21.
- Suitable adjuvants further include cytokines.
- suitable cytokines are provided in Table 3 (Gabrilovich et al., 1996, the contents of which are incorporated by reference in their entirety).
- cytokines and immunological adjuvants may be used in vitro, such as for expansion or activation of T cells, or for ex vivo uses.
- Suitable adjuvants further include CpG oligonucleotides.
- useful adjuvants include, but are not limited to, a chemically modified CpG (e.g. CpR, Idera), a dsRNA analogue such as Poly(l:C) and a derivative thereof (e.g. AmpliGen®, Hiltonol®, poly(ICLC), poly(IC-R), poly(l:C12U), non-CpG bacterial DNA or RNA as well as a mimetic of the bacterial lipopeptide Pam3Cys-Ser-Ser, such as Pam3Cys-GDPKHPKSF (XS15). See Gouttefangeas and Rammensee 2018 or Rammensee et al. 2019, the contents of which are incorporated herein by reference in their entireties.
- a further example of a suitable adjuvant is an immunoactive small molecule or an antibody such as cyclophosphamide, sunitinib, an immune checkpoint inhibitor including ipilimumab, nivolumab, pembrolizumab, atezolizumab, avelumab, durvalumab, and cemiplimab, Bevacizumab®, Celebrex, NCX-4016, sildenafil, tadalafil, vardenafil, sorafenib, temozolomide, temsirolimus, XL-999, CP-547632, pazopanib, VEGF Trap, ZD2171 , AZD2171 , anti-CTLA4, another antibody targeting a key structure of the immune system (e.g.
- anti-CD40, anti-TGF-beta, anti-TNF-alpha receptor) and SC58175, which may act therapeutically and/or as an adjuvant may act therapeutically and/or as an adjuvant.
- concentration of an adjuvant and/or additive useful in the context of a method or use disclosed herein can readily be determined by the skilled artisan without undue experimentation.
- a suitable adjuvant examples include anti-CD40, imiquimod, Resiquimod, GM-CSF, cyclophosphamide, sunitinib, bevacizumab, interferon-alpha, a CpG oligonucleotide or a derivative thereof, Poly(l:C) or a derivative thereof, RNA, sildenafil, and a particulate formulation with poly(lactide co-glycolide) (PLG), a virosome, and/or an interleukin.
- a respective interleukin may for instance be one or more of IL1 , IL-2, IL-4, IL-7, IL-12, IL- 13, IL-15, IL-18, IL-21 , and IL-23.
- the pharmaceutical composition is used for parenteral administration, such as subcutaneous, intradermal, intramuscular or oral administration.
- the peptides, TCRs, antibodies or cells and optionally other molecules can be dissolved or suspended in a pharmaceutically acceptable carrier, for instance an aqueous carrier, in the pharmaceutical composition.
- a pharmaceutically acceptable carrier for instance an aqueous carrier
- An antibody or TCR as disclosed herein may be administered to a subject in a pharmaceutically acceptable carrier.
- a pharmaceutically acceptable carrier include saline, Ringer's solution and dextrose solution.
- the pH of the solution may for instance be from about 5 to about 8, such as from about 7 to about 7.5.
- Further carriers include sustained release preparations such as semipermeable matrices of solid hydrophobic polymers containing the antibody, which matrices are in the form of shaped articles, e.g., films, liposomes or microparticles. It will be apparent to those persons skilled in the art that certain carriers may be more preferable depending upon, for instance, the route of administration and concentration of antibody being administered.
- the pharmaceutical composition can contain excipients, such as buffers, binding agents, blasting agents, diluents, flavors, lubricants, emulsifier, vehicle etc. or a combination thereof.
- excipients such as buffers, binding agents, blasting agents, diluents, flavors, lubricants, emulsifier, vehicle etc. or a combination thereof.
- the peptides, TCRs, antibodies or T cells recognizing the peptide of the present disclosure in a complex with an MHC molecule can also be administered together with immune stimulating substances, such as cytokines shown in Table 3. These cytokines may be expressed by the T lymphocyte in soluble or membrane-bound form or may be administered separately.
- One or more cytokines may also be used in the activation and/or expansion of T cells, such as T cells recognizing the peptide of the present disclosure in a complex with an MHC molecule.
- composition can be used for a prevention, prophylaxis and/or therapy of adenomatous or cancerous diseases.
- Exemplary formulations can be for example found in EP2112253.
- a pharmaceutical composition e.g. a vaccine composition, as disclosed herein does not contain any adjuvant.
- a formulation as provided herein is one that is suitable for administration of the peptide or other agent by any acceptable route such as oral (enteral), nasal, ophthal, subcutaneous, intradermal, intramuscular, intravenous or transdermal.
- the administration is s.c., and in some embodiments i.d. administration by infusion pump.
- a peptide or a peptide sequence as provided herein may be included in a vaccine composition.
- a respective vaccine composition may for example be used for eliciting or for enhancing an anti-tumor immune response.
- a peptide sequence as provided herein may be used as a target for the development of a pharmaceutically / immunologically active compound and/or cell.
- the peptide, the salt thereof, the nucleic acid, a vector comprising the nucleic acid or the pharmaceutical composition comprising the peptide, the salt thereof, the nucleic acid or the vector disclosed herein is an immunotherapeutic such as a vaccine.
- a peptide as disclosed herein is useful for generating an immune response in a subject by which tumor cells can be destroyed.
- An immune response in a subject can be induced by direct administration of the described peptide or suitable precursor substance (e.g. an elongated peptide, protein, or nucleic acid encoding the peptide) to the subject, in some embodiments in combination with an agent enhancing the immunogenicity (i.e. an adjuvant).
- the immune response originating from such a therapeutic vaccination can be expected to be highly specific against tumor cells because a target peptide as provided herein is not presented on healthy tissues in comparable copy numbers, preventing the risk of undesired autoimmune reactions against normal cells in the subject.
- the vaccine includes at least one peptide having the amino acid sequence set forth SEQ ID NO: 1 to SEQ ID NO: 59.
- the vaccine includes at least one or more additional peptides, such as two to 50 or two to 25 peptides. In some embodiments two to 20 additional peptides are included. In some embodiments two, three, four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, fifteen, sixteen, seventeen or eighteen peptides may be included.
- the peptide(s) may be derived from one or more source proteins and may bind to an MHC class I molecule.
- a peptide provided herein may bind to a molecule of any human leukocyte antigen serotype.
- a peptide provided herein generally binds to at least one selected from the group consisting of HLA-A*01 :01 , HLA-A*02:01 , HLA-A*03:01 , HLA-A*24:02, HLA-B*07:02, HLA-B*08:01 and HLA- B*44:02, plus optionally other HLA allotypes.
- a peptide binds to HLA-A*01 :01.
- a peptide binds to HLA-A*02:01 or HLA-A*03:01.
- a peptide binds to HLA-A*24:02 or HLA-B*07:02. In some embodiments a peptide binds to HLA-B*08:01 or HLA-B*44:02. Due to similarities in binding pattern such as the relevant anchoring positions some peptides bind to more than one allele, such an overlap is most likely, but not limited to, HLA-A*01 binding peptides also binding to HLA-B*15, HLA-A*03 binding peptides also binding to HLA-A*11 , HLA-B*07 binding peptides also binding to HLA-B*35 and HLA-B*51.
- a peptide provided herein is combined with one or more peptides binding to another allele, a higher percentage of any population of individuals can be treated compared with addressing either MHC class I allele alone. For example, with the combination of an HLA- A*02:01 binding peptide with an HLA-A*24:02 binding peptide 60% of patients in any relevant population can be treated, whereas in most populations less than 50% of patients could be addressed by either allele alone.
- a cancer vaccine comprising a nucleic acid molecule encoding a recombinant protein or peptide.
- the recombinant protein or peptide contains an amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
- the cancer vaccine further comprises a nucleic acid molecule encoding a recombinant MHC molecule.
- the recombinant MHC molecule is generally capable of forming a complex with the recombinant peptide that contains the amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
- the nucleic acid molecule is an RNA molecule such as an mRNA molecule.
- the nucleic acid molecule encodes a recombinant peptide that consists of an amino acid sequence selected from selected from the group consisting of SEQ ID NOs: 1 to 59.
- the nucleic acid molecule includes a 5’UTR.
- the nucleic acid molecule includes a 3’UTR.
- the nucleic acid molecule includes both a 5’UTR and a 3’UTR.
- the nucleic acid molecule is an RNA that includes a poly(A) tail.
- a respective poly(A) tail has in some embodiments a length of 100 to 150 adenosine residues, including 120 adenosine residues.
- a pharmaceutical composition as provided herein may also include one or more nucleic acid molecules, such as one or more RNA molecules. At least one nucleic acid molecule included in such a composition may include an amino acid sequence encoding a peptide as disclosed herein.
- the composition contains mRNA with a capping degree of 70% or more, such that at least 70% or at least 80% of the mRNA species contain a Cap1 structure. In some embodiments at least 90% of the mRNA species contain a Cap1 structure.
- 5'-capping of a polynucleotide may be completed concomitantly during the in vitro transcription reaction using one or more of the following chemical RNA cap analogues to generate the 5'-guanosine cap structure according to manufacturer protocols: 3'-0-Me- m7G(5')ppp(5') G [the ARCA cap]; G(5')ppp(5')A; G(5')ppp(5')G; m7G(5')ppp(5')A; m7G(5')ppp(5')G (New England BioLabs, Ipswich, MA).
- 5'-capping of modified RNA may be completed post-transcriptionally using a Vaccinia Vims Capping Enzyme to generate the “Cap-0” structure: m7G(5')ppp(5')G (New England BioLabs, Ipswich, MA).
- Cap-1 structure may be generated using both Vaccinia Vims Capping Enzyme and a 2'-0 methyl-transferase to generate: m7G(5')ppp(5')G-2'-O-methyl.
- the Cap-2 structure may be generated from the Cap-1 structure followed by the 2'-O-methylation of the 5'- antepenultimate nucleotide using a 2'-O-methyl-transferase.
- the Cap-3 structure may be generated from the Cap-2 structure followed by the 2'-O-methylation of the 5'- preantepenultimate nucleotide using a 2'-O-methyl-transferase.
- Enzymes may be derived from a recombinant source.
- the 5’ end of a nucleic acid molecule includes a Cap structure that has the following general formula:
- Ri and R2 are independently hydroxy or methoxy.
- W ⁇ , X’ and Y ⁇ are independently oxygen, sulfur, selenium, or BH3.
- the 5’-cap structure is capable of increasing the stability of the RNA, increasing translation efficiency of the RNA, prolonging translation of the RNA, increasing total protein expression of the RNA, and/or increasing the immune response against an antigen or antigen peptide encoded by said RNA when compared to the same RNA without the 5’-cap structure.
- the one or more nucleic acids may be complexed or associated with one or more lipids or lipid-based carriers, thereby forming liposomes, lipid nanoparticles (LNP), lipoplexes, and/or nanoliposomes, which may encapsulate the one or more nucleic acids.
- the LNP may contain any lipid capable of forming a particle to which the one or more nucleic acid molecules are attached, or in which the one or more nucleic acid molecules are encapsulated.
- An LNP may contain one or more cationic lipids or one or more neutral lipids.
- An LNP may also contain one or more steroids or steroid analogues.
- An LNP may also contain one or more polymer conjugated lipids, such as a polyethyleneglycol-(PEG-) lipid.
- the LNP contains one or more cationic lipids, and one or more neutral or pegylated lipids.
- PEG-lipid Two illustrative examples of a PEG-lipid are 1 ,2 dimyristoyl-sn-glycerol, methoxy-polyethylene-glycol (PEG2000 DMG) or O-Methyl-O'-succinylpolyethylenglykol 2'000 (PEG2000-cDMA).
- a further example of a PEG-lipid is PEG-DPG (1 ,2-Dipalm itoyl- sn-glycerol, methoxypolyethylene glycol).
- An illustrative example of a non-cationic lipid is 1 ,2 distearoyl-sn-glycero-3-phosphocholine (DSPC).
- one or more nucleic acid molecules may be included in the composition, which include(s) an amino acid sequence encoding a cytokine or a portion thereof.
- the combination product includes a peptide or salt thereof as disclosed herein.
- the combination product may also include a proteinaceous binding molecule as disclosed herein, for instance an antibody, fragment thereof, or a T-cell receptor or fragment thereof.
- the combination product may also include a nucleic acid or an expression vector as disclosed herein.
- the combination product may also include a recombinant host cell or an activated T lymphocyte as disclosed herein.
- the combination product includes a first active agent that is a peptide or salt thereof as disclosed herein. In some embodiments the combination product includes a first active agent that is a proteinaceous binding molecule as disclosed herein. In some embodiments the combination product includes a first active agent that is a nucleic acid or an expression vector as disclosed herein. In some embodiments the combination product includes a first active agent that is a recombinant host cell or an activated T lymphocyte as disclosed herein.
- the combination product includes a second active agent that is an adjuvant.
- the adjuvant is provided in an immunogenicity enhancing amount.
- the adjuvant is an interleukin.
- the interleukin may for example be IL-2, IL-12, IL-15, IL-18, IL-21 or a combination thereof.
- the interleukin may be IL-2.
- the interleukin may be IL- 12.
- the interleukin may be IL-15.
- the interleukin may be IL-18.
- the interleukin may be IL-21.
- the combination product includes a T lymphocyte, such as a recombinant T lymphocyte, as a first active agent and IL-2 as the second active agent.
- the combination product is or includes an individualized product.
- the combination product is provided in the form of a kit.
- the kit may include a first pharmaceutical composition that contains a peptide or salt thereof as disclosed herein.
- the first pharmaceutical composition may also include a proteinaceous binding molecule as disclosed herein, for instance an antibody, fragment thereof, or a T- cell receptor or fragment thereof.
- the first pharmaceutical composition may also include a nucleic acid or an expression vector as disclosed herein.
- the first pharmaceutical composition may also include a recombinant host cell or an activated T lymphocyte as disclosed herein.
- the kit may include a first pharmaceutical composition that contains an adjuvant as described above.
- the peptide may be substantially pure or combined with an immune-stimulating adjuvant (see below) or used in combination with immune-stimulatory cytokines, or be administered with a suitable delivery system, for example liposomes.
- the peptide may also be conjugated to a suitable carrier such as keyhole limpet hemocyanin (KLH) or mannan (see WO 95/18145 and Longenecker et al., 1993, the contents of both of which are incorporated by reference in their entirety).
- KLH keyhole limpet hemocyanin
- mannan see WO 95/18145 and Longenecker et al., 1993, the contents of both of which are incorporated by reference in their entirety.
- the peptide may also be tagged, may be a fusion protein, or may be a hybrid molecule.
- kits contains a container that contains a component as disclosed herein for the pharmaceutical composition, in solution or in lyophilized form.
- the component as disclosed herein may be the peptide provided herein.
- the component as disclosed herein may also be the proteinaceous binding molecule provided herein.
- the component may also be the nucleic acid or the vector provided herein.
- the component may also be the recombinant host cell or the activated T lymphocyte provided herein.
- the component as disclosed herein may be provided in the form of a pharmaceutical composition.
- the component as disclosed herein may also be the pharmaceutical composition disclosed above.
- the kit provided herein may include a lyophilized formulation provided herein in a suitable container and instructions for its reconstitution and/or use.
- suitable containers include for example bottles, vials (e.g. dual chamber vials), syringes (such as dual chamber syringes) and test tubes.
- the container may be formed from a variety of materials such as glass or plastic.
- the kit and/or container contain/s instructions on or associated with the container that indicates directions for reconstitution and/or use.
- the label may indicate that the lyophilized formulation is to be reconstituted to peptide concentrations as described above.
- the label may further indicate that the formulation is useful or intended for subcutaneous administration.
- the kit may in some embodiments furthermore contain a further container.
- the further container may contain a diluent for the solution or for the lyophilized formulation.
- the further container may also contain a reconstituting solution for the lyophilized formulation.
- the further container may be a second container.
- the kit may in some embodiments contain a further container.
- This further container contains an adjuvant.
- the further container may be a second container or a third container.
- the kit may include at least one further peptide that includes a sequence selected from the group consisting of SEQ ID NO: 1 to SEQ ID NO: 59.
- the kit may include any further peptide as provided herein.
- the kit may furthermore include a buffer and/or a diluent.
- the kit may furthermore include a filter and/or a needle.
- the kit may in some embodiments furthermore contain a syringe.
- the container holding the formulation may be a multi-use vial, which allows for repeat administrations (e.g., from 2-6 administrations) of the reconstituted formulation.
- the kit may further comprise a second container comprising a suitable diluent (e.g., sodium bicarbonate solution).
- the kit may further include other materials desirable from a commercial and user standpoint, including other buffers, diluents, filters, needles, syringes, and package inserts with instructions for use.
- kits provided herein may have a single container that contains the formulation of a pharmaceutical composition provided herein with or without other molecules, pharmaceutical compositions or cells of these other compounds) or may have a distinct container for each component.
- a kit provided herein includes a pharmaceutical composition as provided herein packaged for use in combination with the co-administration of a second compound (such as an adjuvant (e.g. GM-CSF), a chemotherapeutic agent, a natural product, a hormone or antagonist, an anti-angiogenesis agent or inhibitor, an apoptosisinducing agent or a chelator) or a pharmaceutical composition thereof.
- a second compound such as an adjuvant (e.g. GM-CSF), a chemotherapeutic agent, a natural product, a hormone or antagonist, an anti-angiogenesis agent or inhibitor, an apoptosisinducing agent or a chelator
- a respective composition and second compound may be provided as separate components, typically packed for independent use.
- the kit may in such embodiments define a combination product.
- Molecules included in the kit may be pre-complexed.
- each molecule may be in a separate distinct container prior to administration to a subject.
- the molecules included in the kit may be provided in one or more liquid solutions, such as an aqueous solution, for example a sterile aqueous solution.
- the molecules included in the kit may also be provided as solids, which may be converted into liquids by addition of suitable solvents, which are in some embodiments provided in another distinct container.
- the container of a therapeutic kit may be a vial, test tube, flask, bottle, syringe, or any other means of enclosing a solid or liquid.
- the kit will contain a second vial or other container, which allows for separate dosing.
- the kit may also contain another container for a pharmaceutically acceptable liquid.
- a therapeutic kit contains an apparatus (e.g., one or more needles, syringes, eye droppers, pipette, etc.), which enables administration of an agent as provided herein that is a component of the present kit.
- a kit may in some embodiments be an example of a combination product, as provided herein.
- a combination product may be a combination of any peptide, nucleic acid, cell, binding molecule or other agent as provided herein with any other agent provided herein.
- a combination product may be for administration at independent points in time.
- the agents, e.g. a binding agent and an adjuvant, included in a combination may be administered concomitantly or subsequently.
- a further aspect provides a nucleic acid molecule (for example a polynucleotide) encoding a peptide of the disclosure.
- An aspect provides a nucleic acid molecule encoding a TCR or a TCR fragment specific for the peptide of the disclosure.
- Another aspect provides a nucleic acid molecule encoding an antibody or an antibody fragment specific for the peptide of the disclosure.
- the nucleic acid molecule contains a nucleic acid sequence that codes in 5’— >3’ direction for the peptide, TCR or TCR fragment.
- the polynucleotide may be, for example, DNA, cDNA, PNA, RNA or combinations thereof, either single- and/or double-stranded, or native or stabilized forms of polynucleotides, such as, for example, polynucleotides with a phosphorothioate backbone and it may or may not contain introns so long as it codes for the peptide.
- polynucleotides such as, for example, polynucleotides with a phosphorothioate backbone and it may or may not contain introns so long as it codes for the peptide.
- polynucleotide may be, for example, DNA, cDNA, PNA, RNA or combinations thereof, either single- and/or double-stranded, or native or stabilized forms of polynucleotides, such as, for example, polynucleotides with a phosphorothioate backbone and it may or may not contain introns so long as it codes
- nucleic acids examples include for instance DNA molecules, RNA molecules, analogues of the DNA or RNA generated using nucleotide analogues or using nucleic acid chemistry, locked nucleic acid molecules (LNA), protein nucleic acids molecules (PNA), alkylphosphonate and alkylphosphotriester nucleic acid molecules and tecto-RNA molecules (e.g., Liu, B., et al., J. Am. Chem. Soc. (2004) 126, 4076-4077).
- LNA has a modified RNA backbone with a methylene bridge between C4' and 02', providing the respective molecule with a higher duplex stability and nuclease resistance.
- Alkylphosphonate and alkylphosphotriester nucleic acid molecules can be viewed as a DNA or an RNA molecule, in which phosphate groups of the nucleic acid backbone are neutralized by exchanging the P-OH groups of the phosphate groups in the nucleic acid backbone to an alkyl and to an alkoxy group, respectively.
- DNA or RNA may be of genomic or synthetic origin and may be single or double stranded.
- Such nucleic acid can be e.g. mRNA, cRNA, synthetic RNA, genomic DNA, cDNA synthetic DNA, a copolymer of DNA and RNA, oligonucleotides, etc.
- a respective nucleic acid may furthermore contain non-natural nucleotide analogues and/or be linked to an affinity tag or a label.
- RNA may be selected from messenger RNA (mRNA), self-amplifying RNA (saRNA), siRNA or miRNA.
- nucleotide analogues and nucleoside analogues are known and can be used in nucleic acids provided herein and used in the methods disclosed herein.
- a nucleotide analogue is a nucleotide containing a modification at for instance the base, sugar, or phosphate moieties.
- a nucleoside analogue is accordingly a nucleoside containing a modification at the base or sugar.
- a substitution of 2'-OH residues of siRNA with 2'F, 2'0-Me or 2'H residues is known to improve the in vivo stability of the respective RNA.
- nucleoside analogue is 5-methyluridine, also called ribothymidine, which is the ribonucleoside counterpart to the deoxyribonucleoside thymidine, which lacks a hydroxyl group at the 2' position.
- a further example of a nucleoside analogue is pseudouridine, a C5-glycoside isomer of uridine, in which the uracil is bonded to ribose via a carbon-carbon instead of a nitrogen-carbon glycosidic bond.
- An additional methylation at the N1 position of pseudouridine removes the additional hydrogen bond donor and generates N1 -methyl-pseudouridine.
- nucleoside analogues are nucleosides in which the 4’-oxygen of furanose is replaced by another heteroatom or by carbon.
- Azanucleosides for instance have a nitrogen atom at this position, and thionucleosides have a sulfur atom instead of oxygen.
- a suitable nucleoside analogue are also 4'-th iourid ine and 5-methylcytosine.
- Further examples of a suitable nucleoside analogue are 2-thio-1 -methyl-1 -deaza-pseudouridine or 2-thio-1 - methyl-pseudouridine.
- Suitable examples of a nucleoside analogue are also 2-thio-5- aza-uridine or 2-thio-dihydropseudouridine.
- Examples of a suitable nucleoside analogue are also 2-thio-dihydrouridine or 2-thio-pseudouridine. Further examples of a suitable nucleoside analogue are 4-methoxy-2-thio-pseudouridine or 4-methoxy-pseudouridine. Yet further examples of a suitable nucleoside analogue are 4-thio-1 -methyl- pseudouridine or 4-thio-pseudouridine. Examples of a suitable nucleoside analogue are also 5-aza-uridine or dihydropseudouridine. Examples of a suitable nucleoside analogue are also 5-methyluridine or 5-methoxyuridine.
- nucleoside analogues examples include also 2-methoxyuridine or 2'-O-methyl uridine. Further examples of a suitable nucleoside analogue are 2-methoxy-4-thio-uridine or 1 -methyl-3-(3-amino-3- carboxypropyl)pseudouridine.
- Modifications at the base moiety may be a natural or a synthetic modification of A, C, G, and T/U, a different purine or pyrimidine base, such as uracil-5-yl, hypoxanthin-9-yl, and 2-aminoadenin-9-yl, as well as a non-purine or a non-pyrimidine nucleotide base.
- Pseudouridine can for example be formed using a pseudouridine synthase enzyme.
- Other nucleotide analogues serve as universal bases. Examples of universal bases include 3-nitropyrrole and 5-nitroindole. Universal bases are able to form a base pair with any other base. Base modifications often can be combined with for example a sugar modification, such as for instance 2'-O-methoxyethyl, e.g. to achieve unique properties such as increased duplex stability.
- a nucleic acid contains two or more encoding repeats (“concatemer”), separated by short nucleotide stretches (“spacers”).
- the nucleic acid for example an mRNA molecule, includes a 5' untranslated region (UTR) and/or a 3' UTR.
- a 3’-UTR in some embodiments contains or consists of a nucleic acid sequence derived from a 3’-UTR of a gene selected from PSMB3, ALB7, alpha-globin, CASP1 , COX6B1 , GNAS, NDUFA1 , and RPS9 or from a homolog, a fragment, or a variant of any one of these genes.
- a 5’-UTR contains or consists of a nucleic acid sequence derived from a 5’-UTR of a gene selected from HSD17B4, RPL32, ASAH1 , ATP5A1 , MP68, NDUFA4, NOSIP, RPL31 , SLC7A3, TUBB4B, and UBQLN2 or from a homolog, a fragment, or variant of any one of these genes.
- a 5’-UTR and/or a heterologous 3’ UTR is selected from UTR design a-1 (HSD17B4/PSMB3) or a-3 (SLC7A3/PSMB3).
- a 5’-UTR and/or a heterologous 3’ UTR is selected from UTR design e-2 (RPL31/RPS9), or i-3 (-/muag).
- the nucleic acid contains a coding sequence which is codon- optimized and/or in which the G/C content is increased and the uridine content is decreased compared to the respective naturally occurring coding sequence.
- the codonoptimization and/or the increase in the G/C content can be carried out in such a way that it does not change the sequence of the encoded amino acid sequence.
- Generating a G/C content optimized nucleic acid sequence may for instance be carried out using a method according to WO 2002/098443, the disclosure of which is incorporated herein in its entirety.
- nucleic acid may include or consist of adapting the codons in a coding sequence to human codon usage (herein referred to as “human codon usage adapted coding sequence”).
- Codons encoding the same amino acid occur at different frequencies in humans. Accordingly, the coding sequence of the nucleic acid may be modified such that the frequency of the codons encoding the same amino acid corresponds to the naturally occurring frequency of that codon according to the human codon usage.
- the wild type or reference coding sequence may be adapted in a way that the codon “GCC” is used with a frequency of 0.40, the codon “GOT” is used with a frequency of 0.28, the codon “GCA” is used with a frequency of 0.22 and the codon “GCG” is used with a frequency of 0.10.
- a corresponding procedure (as exemplified for alanine) may be applied to any, including each, amino acid encoded by the coding sequence of a nucleic acid. Where all codons for each amino acid are adapted in this manner, a sequence is obtained that is adapted to human codon usage.
- a still further aspect provides an expression vector comprising a nucleic acid sequence encoding a peptide as disclosed herein.
- an expression vector comprising a nucleic acid sequence encoding a proteinaceous binding molecule as disclosed herein.
- the nucleic acid sequence may in some embodiments encode an antibody or an antibody fragment as disclosed herein.
- the nucleic acid sequence may in some embodiments encode a TCR or a TCR fragment as disclosed herein.
- the nucleic acid molecule or polynucleotide used for active vaccination may be substantially pure or contained in a suitable vector or delivery system.
- the nucleic acid may be DNA, cDNA, PNA, RNA or a combination thereof. Methods for designing and introducing such a nucleic acid are well known in the art.
- Suitable vectors and delivery systems include viral DNA and/or RNA, such as systems based on adenovirus, vaccinia virus, retroviruses, herpes virus, adeno-associated virus or hybrids containing elements of more than one virus.
- Non-viral delivery systems include cationic lipids and cationic polymers and are well known in the art of DNA delivery. Physical delivery, such as via a “gene-gun” may also be used.
- the peptide or peptides encoded by the nucleic acid may be a fusion protein, for example with an epitope that stimulates T cells for the respective opposite CDR as noted above.
- a variety of methods have been developed to link polynucleotides, especially DNA, to vectors for example via complementary cohesive termini. For instance, complementary homopolymer tracts can be added to the DNA segment to be inserted to the vector DNA. The vector and DNA segment are then joined by hydrogen bonding between the complementary homopolymeric tails to form recombinant DNA molecules.
- Synthetic linkers containing one or more restriction sites provide an alternative method of joining the DNA segment to vectors.
- Synthetic linkers containing a variety of restriction endonuclease sites are commercially available from a number of sources including International Biotechnologies Inc. New Haven, CN, USA.
- a desirable method of modifying the DNA encoding a peptide as disclosed herein, or encoding the respective TCR or antibody, or TCR or antibody fragment employs the polymerase chain reaction as disclosed by Saiki et al. (Saiki et al., 1988, the contents of which are incorporated by reference in their entirety).
- This method may be used for introducing the DNA into a suitable vector, for example by engineering in suitable restriction sites, or it may be used to modify the DNA in other useful ways as is known in the art.
- Illustrative examples of viral vectors are pox- or adenovirus vectors.
- the DNA (or in the case of retroviral vectors, RNA) may then be expressed in a suitable host to produce a polypeptide comprising the peptide as disclosed herein.
- RNA Ribonucleic acid
- the DNA encoding a peptide of the disclosure may be used in accordance with known techniques, appropriately modified in view of the teachings contained herein, to construct an expression vector, which is then used to transform an appropriate host cell for the expression and production of the polypeptide of the invention.
- Such techniques include those disclosed for example in US 4,440,859, 4,530,901 , 4,582,800, 4,677,063, 4,678,751 , 4,704,362, 4,710,463, 4,757,006, 4,766,075, and 4,810,648, the contents each of which are incorporated by reference in their entirety.
- DNA (or in the case of retroviral vectors, RNA) encoding a polypeptide that encodes the peptide, TCR, TCR fragment, antibody or antibody fragment as disclosed herein may be joined to a wide variety of other DNA sequences for introduction into an appropriate host.
- the companion DNA will depend on the nature of the host, the manner of the introduction of the DNA into the host, and whether episomal maintenance or integration is desired.
- the DNA is inserted into an expression vector, such as a plasmid, in proper orientation and correct reading frame for expression.
- an expression vector such as a plasmid
- the DNA may be linked to the appropriate transcriptional and translational regulatory control nucleic acid sequences recognized by the desired host, although such controls are generally available in the expression vector.
- the vector is then introduced into the host through standard techniques. Generally, not all of the hosts will be transformed by the vector. Therefore, it will be necessary to select for transformed host cells.
- One selection technique involves incorporating a DNA sequence into the expression vector, with any necessary control elements, that codes for a selectable trait in the transformed cell, such as antibiotic resistance.
- the gene for such selectable trait can be on another vector, which is used to co-transform the desired host cell.
- Host cells that have been transformed by a recombinant DNA as disclosed herein are then cultured for a sufficient time and under appropriate conditions known to those skilled in the art in view of the teachings disclosed herein to permit the expression of the polypeptide or peptide, which can then be recovered.
- the system can be mammalian cells such as CHO cells available from the ATCC Cell Biology Collection.
- a typical mammalian cell vector plasmid for constitutive expression comprises the CMV or SV40 promoter with a suitable poly A tail and a resistance marker, such as neomycin.
- a suitable poly A tail and a resistance marker, such as neomycin.
- pSVL available from Pharmacia, Piscataway, NJ, USA.
- An example of an inducible mammalian expression vector is pMSG, also available from Pharmacia.
- Useful yeast plasmid vectors are pRS403-406 and pRS413-416 and are generally available from Stratagene Cloning Systems, La Jolla, CA 92037, USA.
- Plasmids pRS403, pRS404, pRS405 and pRS406 are Yeast Integrating plasmids (Yips) and incorporate the yeast selectable markers HIS3, TRP1 , LEU2 and URA3.
- Plasmids pRS413-416 are Yeast Centromere plasmids (Ycps).
- CMV promoter-based vectors (for example from Sigma-Aldrich) provide transient or stable expression, cytoplasmic expression or secretion, and N-terminal or C-terminal tagging in various combinations of FLAG, 3xFLAG, c-myc or MAT. These fusion proteins allow for detection, purification and analysis of recombinant protein. Dual-tagged fusions provide flexibility in detection.
- CMV human cytomegalovirus
- the strong human cytomegalovirus (CMV) promoter regulatory region drives constitutive protein expression levels as high as 1 mg/L in COS cells. For less potent cell lines, protein levels are typically ⁇ 0.1 mg/L.
- the presence of the SV40 replication origin will result in high levels of DNA replication in SV40 replication permissive COS cells.
- CMV vectors for example, can contain the pMB1 (derivative of pBR322) origin for replication in bacterial cells, the b-lactamase gene for ampicillin resistance selection in bacteria, hGH polyA, and the f1 origin.
- Vectors containing the pre-pro-trypsin leader (PPT) sequence can direct the secretion of FLAG fusion proteins into the culture medium for purification using ANTI-FLAG antibodies, resins, and plates.
- Other vectors and expression systems are well known in the art for use with a variety of host cells.
- two or more peptides as disclosed herein are encoded and thus expressed in a successive order (similar to “beads on a string” constructs).
- the peptides may be linked or fused together by stretches of linker amino acids, such as for example LLLLLL, or may be linked without any additional peptide(s) between them.
- linker amino acids such as for example LLLLLL
- These constructs can also be used for cancer therapy and may induce immune responses both involving MHC I and MHC II.
- the host cell can be either prokaryotic or eukaryotic.
- Bacterial cells may be prokaryotic host cells in some circumstances and typically are a strain of E. coli such as, for example, the E. coli strains DH5 available from Bethesda Research Laboratories Inc., Bethesda, MD, USA, and RR1 available from the American Type Culture Collection (ATCC) of Rockville, MD, USA (No ATCC 31343).
- suitable eukaryotic host cells include yeast, insect and mammalian cells.
- suitable eukaryotic host cells are vertebrate cells such as those from a mouse, rat, monkey or human fibroblastic and colon cell lines.
- Yeast host cells include YPH499, YPH500 and YPH501 , which are generally available from Stratagene Cloning Systems, La Jolla, CA 92037, USA.
- mammalian host cells include Chinese hamster ovary (CHO) cells available from the ATCC as CCL61 , NIH Swiss mouse embryo cells NIH/3T3 available from the ATCC as CRL 1658, monkey kidney- derived COS-1 cells available from the ATCC as CRL 1650 and 293 cells which are human embryonic kidney cells.
- suitable insect cells are Sf9 cells which can be transfected with baculovirus expression vectors. An overview regarding the choice of suitable host cells for expression can be found in the literature (Baibas and Lorence, 2004).
- Transformation of appropriate cell hosts with a DNA construct of the present disclosure is accomplished by well-known methods that typically depend on the type of vector used.
- transformation of prokaryotic host cells see for example Cohen et al. or Green and Sambrook (Cohen et al., 1972; Green and Sambrook, 2012). Transformation of yeast cells is described in Sherman et al. (Sherman et al. , 1986). The method of Beggs (Beggs, 1978) is also useful.
- reagents useful in transfecting such cells for example calcium phosphate and DEAE-dextran or liposome formulations, are available from Stratagene Cloning Systems, or Life Technologies Inc., Gaithersburg, MD 20877, USA. Electroporation is also useful for transforming and/or transfecting cells and is well known in the art for transforming yeast cell, bacterial cells, insect cells and vertebrate cells. The contents of each of these references is herein incorporated by reference in their entirety.
- Successfully transformed cells i.e., cells that contain a DNA construct as disclosed herein, can be identified by well-known techniques such as PCR.
- the presence of the protein in the supernatant can furthermore be detected using antibodies.
- host cells disclosed herein are useful in the preparation of a peptide disclosed herein, for example bacterial, yeast and insect cells. Such host cells may also be useful in expressing a TCR or an antibody. Other host cells may be useful in certain therapeutic methods. For example, antigen-presenting cells, such as dendritic cells, may be used to express a peptide disclosed herein such that they may be loaded into appropriate MHC molecules. Thus, provided is also a host cell that includes a nucleic acid molecule or an expression vector as disclosed herein.
- the host cell is an antigen-presenting cell, in particular a dendritic cell or antigen-presenting cell.
- APCs loaded with a recombinant fusion protein containing prostatic acid phosphatase (PAP) were approved by the U.S. Food and Drug Administration (FDA) on April 29, 2010, to treat asymptomatic or minimally symptomatic metastatic HRPC (Sipuleucel-T) (Rini et al., 2006; Small et al., 2006, the contents of which are incorporated by reference in their entirety).
- a method of producing a peptide comprising culturing a host cell and isolating the peptide from the host cell or its culture medium.
- a binding molecule is generally a proteinaceous binding molecule or a nucleic acid molecule.
- a proteinaceous binding molecule that specifically binds to a peptide as disclosed herein or to an MHC class I or ii molecule being complexed with a peptide as disclosed herein.
- a respective proteinaceous binding molecule is in some embodiments an antibody or a fragment thereof.
- a further example of a suitable proteinaceous binding molecule is a TCR or a fragment thereof.
- Yet another example of a suitable proteinaceous binding molecule is a chimeric antigen receptor (CAR).
- a CAR is a recombinant or synthetic molecule which combines antibody-based specificity for a certain antigen with a T cell receptor-activating intracellular domain thereby defining a chimeric protein that exhibits cellular immune activity to the respective antigen.
- the proteinaceous binding molecule is not a chimeric antigen receptor.
- the proteinaceous binding molecule is an antibody fragment.
- an antibody fragment are a diabody or a domain antibody (Holt, L.J., et al., Trends Biotechnol. (2003), 21 , 11 , 484-490).
- a proteinaceous binding molecule with antibody-like functions is a mutein based on a polypeptide of the lipocalin family (WO 03/029462, Beste et al., Proc. Natl. Acad. Sci. USA (1999) 96, 1898-1903).
- Lipocalins such as the bilin binding protein, the human neutrophil gelatinase-associated lipocalin, human Apolipoprotein D or glycodelin, possess natural ligand-binding sites that can be modified so that they bind to selected small protein regions known as haptens.
- a further example of a proteinaceous binding molecule is a so-called glubody (see e.g. international patent application WO 96/23879 or Napolitano, E.W., et al., Chemistry & Biology (1996) 3, 5, 359-367).
- Examples of further proteinaceous binding molecules are a protein based on the ankyrin scaffold (Mosavi, L.K., et al., Protein Science (2004) 13, 6, 1435-1448) or a protein based on the crystalline scaffold (e.g. international patent application WO 01/04144).
- Yet further examples of a proteinaceous binding molecule are AdNectins or tetranectins.
- a further example of a proteinaceous binding molecule is an avimer.
- Avimers contain so called A-domains that occur as strings of multiple domains in several cell surface receptors (Silverman, J., et al., Nature Biotechnology (2005) 23, 1556-1561 ).
- Adnectins derived from a domain of human fibronectin, contain three loops that can be engineered for antibody -like binding to targets (Gill, D.S. & Damle, N.K., Current Opinion in Biotechnology (2006) 17, 653-658).
- Tetranectins derived from the respective human homotrimeric protein, likewise contain loop regions in a C-type lectin domain that can be engineered for desired binding (ibid.).
- Peptoids which can act as protein ligands, are oligo(N-alkyl) glycines that differ from peptides in that the side chain is connected to the amide nitrogen rather than the a carbon atom. Peptoids are typically resistant to proteases and other modifying enzymes and can have a much higher cell permeability than peptides (see e.g. Kwon, Y.-U., and Kodadek, T., J. Am. Chem. Soc. (2007) 129, 1508-1509).
- a further illustrative example of a proteinaceous binding molecule is a soluble bispecific binding molecule, which contains a peptide/MHC recognition domain and a T cellengaging domain.
- the T cell-engaging domain is typically specific for the CD3 receptor.
- the peptide/MHC recognition domain may be a TCR domain or an antibody domain. Examples are the so-called immune-mobilizing monoclonal T cell receptors against cancer (ImmTAC) or the bispecific T cell engagers (BiTEs®).
- an antibody, a TCR or another proteinaceous binding molecule may have a binding specificity to a peptide that can be regarded as an epitope.
- a peptide disclosed herein can be used to generate and develop a specific antibody or antibody fragment against the peptide when in a complex with MHC (Dao et al., Sci Transl Med. (2013) 5, 176, 176ra33; Dao et al., Nat. Biotechnol. (2015) 33, 10, 1079-1086).
- the respective binding molecule can be used for therapy, targeting toxins or radioactive substances to the diseased tissue.
- Another use of such antibody or antibody fragment can be targeting radionuclides to the diseased tissue for imaging purposes such as PET. This use can help to detect small metastases or to determine the size and precise localization of diseased tissues.
- the antibody is a TCR-like antibody, also termed T-cell receptor mimic (TCRm) antibody.
- TCR-like antibody can be produced with the antibody generation methods described herein, and recognizes a peptide/MHC complex, for example a peptide/MHC complex on the surface of a tumor cell, in the same way as a TCR (see also He et al., 2019).
- a TCR-like antibody is thus an antibody which binds to a peptide as disclosed herein complexed with an MHC.
- a TCR-like antibody is a monoclonal antibody.
- TCR-like antibody is a soluble antibody.
- TCR- like antibodies are known to have effector functions including direct killing of cancer cells, complement-dependent cytotoxicity (CDC), antibody-dependent cellular cytotoxicity (ADCC) and antibody-dependent phagocytosis.
- CDC complement-dependent cytotoxicity
- ADCC antibody-dependent cellular cytotoxicity
- a TCR-like antibody typically has a higher affinity than a corresponding TCR.
- TCR mimic antibodies binding to MHC restricted peptide derived from PRAME are for example disclosed in Chang et al. (Chang et al., J Clin Invest (2017) 127, 7: 2705-2718), the content of which is incorporated herein by reference in its entirety. See, also, US 2018/0148503 (T cell receptor-like antibodies specific for a PRAME peptide) (Eureka Therapeutics Inc), the content of which is incorporated herein by reference in its entirety.
- a suitable antibody may in some embodiments also be a multispecific antibody that includes several antibody fragments.
- the antibodies are tested for their desired activity by known methods, in accordance with the purpose for which the antibodies are to be used (e.g., ELISA, immunohistochemistry, immunotherapy, etc. (Greenfield, 2014, the contents of which are incorporated by reference in their entirety)).
- the antibodies may be tested in ELISA assays or, Western blots, immunohistochemical staining of formalin-fixed cancers or frozen tissue sections. After their initial in vitro characterization, antibodies intended for therapeutic or in vivo diagnostic use are tested according to known clinical testing methods.
- a bispecific antibody includes an antibody capable of selectively binding two or more epitopes.
- Bispecific antibodies may be manufactured in a variety of ways (Holliger & Winter, 1993, the contents of which are incorporated by reference in their entirety), for instance, prepared chemically or from hybrid hybridomas, or may be any of the bispecific antibody fragments mentioned above.
- scFv dimers or diabodies may be used, rather than whole antibodies.
- Diabodies and scFv can be constructed without an Fc region, using only variable domains (usually including the variable domain components from both light and heavy chains of the source antibody), potentially reducing the effects of anti-idiotypic reaction.
- Other forms of bispecific antibodies include the single chain “Janusins” described by Traunecker and colleagues (Traunecker et al., 1991 , the content of which is incorporated by reference in its entirety).
- Bispecific antibodies generally include two different binding domains, with each binding domain specifically binding a different epitope either on two different antigens or on the same antigen. If a bispecific antibody is capable of selectively binding two different epitopes (a first epitope and a second epitope), the affinity of the first binding for the first epitope will generally be at least one to two or three or four orders of magnitude lower than the affinity of the first binding domain for the second epitope, and vice versa.
- the epitopes recognized by the bispecific antibody can be on the same or a different target (e.g., on the same or a different protein). Bispecific antibodies can be made for example by combining binding domains that recognize different epitopes of the same antigen.
- Bispecific antibodies can be prepared as full-length antibodies or antibody fragments (for example, F(ab’)2 bispecific antibodies). See e.g. WO 1996/016673; U.S. Patent No. 5,837,234; WO 1998/002463; U.S. Patent No. 5,821 ,337, the contents of which are incorporated by reference in their entirety.
- a bispecific antibody can have an extended half-life.
- half-life extension of a bispecific antibody can be achieved by: increasing the hydrodynamic volume of the antibody by coupling to inert polymers such as polyethylene glycol or other mimetic hydrophilic polymers; fusion or conjugation to large disordered peptides; or fusing or coupling the antibody to a ligand.
- inert polymers such as polyethylene glycol or other mimetic hydrophilic polymers
- fusion or conjugation to large disordered peptides or fusing or coupling the antibody to a ligand.
- bispecific antibodies are known in the art. Production of full-length bispecific antibodies is based on the co-expression of two antibody heavy chain-light chain pairs, where the two chains have different specificities. See e.g., WO 1993/008829 and Traunecker et al., 1991 , the contents of which are incorporated by reference in their entirety.
- Polyclonal antibodies are heterogeneous populations of antibody molecules derived from the sera of animals immunized with an antigen.
- Polyclonal antibodies which selectively bind a peptide according to SEQ ID NO: 1 to SEQ ID NO: 59, or a fragment thereof may be made by methods well-known in the art (see, e.g., Howard & Kaser, 2007, the contents of which are incorporated by reference in their entirety).
- Chimeric antibodies are molecules different portions of which are derived from different animal species, such as those having variable region derived from a murine antibody and a human antibody constant region, which are primarily used to reduce immunogenicity in application and to increase yields in production for example where murine monoclonal antibodies have higher yields from hybridomas but higher immunogenicity in humans, such that human murine chimeric monoclonal antibodies are used.
- Chimeric antibodies and methods for their production are known in the art (Cabilly et al., 1984; Morrison et al., 1984; Boulianne et al., 1984; European Patent Application 173494 (1986); WO 86/01533 (1986); European Patent Application 184187 (1986); Sahagan et al., 1986; Liu et al., 1987; Sun et al., 1987; Better et al., 1988; Harlow & Lane, 1998; U.S. Patent No. 5,624,659, the contents of which are incorporated by reference in their entirety).
- An antibody fragment whether attached to other sequences or not, can also include one or more insertions, deletions, substitutions, or other selected modifications of particular regions or specific amino acids residues, provided the activity of the fragment is not significantly altered or impaired compared to the non-modified antibody or antibody fragment.
- Such modification can provide for some additional property, such as to remove/add an amino acid capable of disulfide bonding, to increase its bio-longevity, to alter its secretory characteristics, etc.
- the antibody fragment must possess a bioactive property, such as binding activity, regulation of binding at the binding domain, etc.
- Functional or active regions of the antibody may be identified by mutagenesis of a specific region of the protein, followed by expression and testing of the expressed peptide. Such methods are readily apparent to a skilled practitioner in the art and can include site-specific mutagenesis of the nucleic acid encoding the antibody fragment.
- An antibody provided herein may in some embodiments be a humanized antibody or a human antibody.
- Humanized forms of non-human (e.g., murine) antibodies are chimeric antibodies, antibody chains or fragments thereof (such as Fv, Fab, Fab' or other antigenbinding subsequences of antibodies) which contain minimal sequence derived from a non-human antibody.
- Humanized antibodies include human antibody (recipient antibody) in which residues from a complementary determining region (CDR) of the recipient are replaced by residues from a CDR of a non-human species (donor antibody) such as mouse, rat or rabbit having the desired specificity, affinity and capacity.
- Fv framework (FR) residues of the human antibody are replaced by corresponding non-human residues.
- Humanized antibodies may also comprise residues which are found neither in the recipient antibody nor in the imported CDR or framework sequences.
- the humanized antibody will comprise substantially all of at least one, and typically two, variable domains, in which all or substantially all of the CDR regions correspond to those of a non-human antibody and all or substantially all of the FR regions are those of a human antibody consensus sequence.
- the humanized antibody typically contains a portion of an antibody constant region (Fc), for example that of a human antibody.
- antibody fragments comprising the epitope binding site (e.g., Fab, F(ab’)2, or other fragments) may be synthesized. “Fragments” or minimal antibodies may be designed utilizing recombinant antibody techniques. For instance, “Fv” antibodies for use in the present invention may be produced by synthesizing a fused variable light chain region and a variable heavy chain region. Combinations of antibodies are also of interest, e.g., diabodies, which comprise two distinct Fv specificities.
- Antigen-binding fragments of antibody include, but are not limited to, SMIPs (small molecule immunopharmaceuticals), camelbodies, nanobodies, and IgNAR.
- the antibody is binding with a binding affinity of ⁇ 100 nM, or ⁇ 50 nM to the complex, which is also regarded as “specific” in the context of the present invention. In some embodiments, the antibody is binding with a binding affinity of ⁇ 10 nM, such as ⁇ 1 nM to the complex. In some embodiments, the antibody is binding with a binding affinity of ⁇ 0.1 nM, for instance ⁇ 0.01 nM, to the complex.
- the present disclosure also relates to a method of producing an antibody specifically binding to a peptide as disclosed herein, as well as an antibody specifically binding to an MHC class I and/or class II molecule complexed with a peptide as provided herein.
- the antibody may in some embodiments be recombinant.
- a respective method may involve immunizing genetically engineered non-human mammal containing cells expressing the MHC class I and/or class II molecule with a soluble form of an MHC class I or II molecule complexed with a peptide as provided herein. The method also involves isolating mRNA molecules from antibody producing cells of the non-human mammal.
- the method may in some embodiments also include producing a phage display library displaying protein molecules encoded by the above-named mRNA molecules.
- the method may furthermore include isolating a phage from the phage display library.
- the phage may display the antibody specifically binding to said MHC class I or ii molecule being complexed with the respective HLA restricted antigen.
- a hybridoma cell In general, techniques for preparing monoclonal antibodies and hybridomas are well known in the art. Methods for the production of a hybridoma cell are disclosed in Kohler & Milstein (1975). Any animal such as a goat, a mouse or a rabbit that is known to produce antibodies can be immunized with the selected peptide, e.g. a peptide as provided herein. Methods for immunization are well known in the art. Such methods include subcutaneous or intraperitoneal injection of the peptide to elicit lymphocytes that produce or are capable of producing antibodies that will specifically bind to the immunizing agent. Typically, the immunized mammal is bled and the serum from each blood sample is analysed for particular antibodies using appropriate screening assays. Alternatively, the lymphocytes may be immunized in vitro.
- lymphocytes typically splenocytes
- an immortal cell line typically myeloma cells, such as SP2/0-Agl4 myeloma cells
- the immortal cell line such as a myeloma cell line is derived from the same mammalian species as the lymphocytes.
- Illustrative immortal cell lines are mouse myeloma cell lines that are sensitive to culture medium containing hypoxanthine, aminopterin and thymidine ("HAT medium").
- HAT-sensitive mouse myeloma cells are fused to mouse splenocytes using 1500 molecular weight polyethylene glycol ("PEG 1500"). Hybridoma cells resulting from the fusion may then be selected using HAT medium, which kills unfused and unproductively fused myeloma cells.
- PEG 1500 1500 molecular weight polyethylene glycol
- a method of producing the antibody may involve producing a phage display library displaying protein molecules encoded by the mRNA molecules.
- the method furthermore includes isolating a phage from the phage display library.
- the phage displays the above antibody specifically binding to the MHC class I molecule complexed with a peptide as provided herein.
- In vitro antibody libraries are, among others, disclosed in US6300064 by MorphoSys and US6248516 by MRC/Scripps/Stratagene. Phage Display techniques are for example disclosed in US5223409 by Dyax.
- One further method to generate antibodies is by means of a transgenic mammal platforms, as for example described in EP1480515A2 by TaconicArtemis.
- a method of producing an antibody as disclosed herein may further include humanizing the antibody.
- Methods for humanizing non-human antibodies are well known in the art.
- Humanized forms of the antibodies of the present invention may be generated using one of the procedures known in the art such as chimerization or CDR grafting.
- a humanized antibody has one or more amino acid residues introduced into it from a source which is non-human. These non-human amino acid residues are often referred to as "import" residues, which are typically taken from an “import” variable domain. Humanization can be essentially performed by substituting rodent CDRs or CDR sequences for the corresponding sequences of a human antibody. Accordingly, such "humanized” antibodies are chimeric antibodies (US 4,816,567, the contents of which are incorporated by reference in their entirety), wherein substantially less than an intact human variable domain has been substituted by the corresponding sequence from a non- human species. In practice, humanized antibodies are typically human antibodies in which some CDR residues and possibly some FR residues are substituted by residues from analogous sites in rodent antibodies.
- a method of producing an antibody may further include conjugating the antibody with a toxin. In some embodiments, a method of producing an antibody may further include conjugating the antibody with an immune stimulating domain.
- Methods for the production and/or selection of fully human mAbs are known in the art. These can involve the use of a transgenic animal which is immunized with the respective protein or peptide, or the use of a suitable display technique, like yeast display, phage display, B-cell display or ribosome display, where antibodies from a library are screened against human iRhom2 in a stationary phase.
- the monoclonal antibody may also be made by a recombinant DNA method, such as those described in US 4,816,567.
- DNA encoding a monoclonal antibody provided herein can be readily isolated and sequenced using conventional procedures (e.g., by using oligonucleotide probes that are capable of binding specifically to genes encoding the heavy and light chains of murine antibodies).
- In vitro methods are also suitable for preparing monovalent antibodies.
- Digestion of antibodies to produce fragments thereof, particularly Fab fragments can be accomplished using routine techniques known in the art. For instance, digestion can be performed using papain. Examples of papain digestion are described in WO 94/29348 and US 4,342,566, the contents of which are incorporated by reference in their entirety.
- Papain digestion of antibodies typically produces two identical antigen binding fragments, called Fab fragments, each with a single antigen binding site, and a residual Fc fragment. Pepsin treatment yields a F(ab')2 fragment and a pFc' fragment.
- An antibody of the present disclosure can be obtained through phage display, or ribosome display, or yeast display, or bacteria display, or Baculovirus display, or mammal cell display, or mRNA display. These methods are all conventional techniques in the art, the specific operations thereof can be seen in corresponding textbooks or operation manuals (Mondon et al., 2008; the content of which is hereby incorporated by reference in its entirety).
- phage display as an example, separate antibody genes may be inserted into the DNA of phages, so that the variable regions on the antibody molecules that can bind the antigens may be coupled to the capsid protein of the phage. After the phage infecting E. coli, single stranded DNA may be replicated in E.
- the phage may be reassembled and secreted into the culture medium, while the E. coli may not be lysed.
- the phage may be co-incubated with target antigens; and after the bound phages are isolated, amplification and purification may be then conducted so that great amounts of clones can be screened.
- the phage display technique can be found in the literature (Liu et al., 2004; the contents of which are hereby incorporated by reference in its entirety).
- the present disclosure may include methods for producing a monoclonal antibody using a phage display method.
- mRNA may be prepared from an animal, e.g., rabbits, rats, mice, guinea pigs, hamsters, goats, horses, chickens, sheep, and camelids (e.g., llamas), immunized by the method for immunizing an animal, whereupon cDNA may be prepared using the mRNA as a template, so that a single-chain antibody (scFv) gene encoding only an antibody variable region may be prepared.
- the gene may be cloned to a phagemid vector. E. coli, into which the phagemid vector is transduced, is infected with phage, so as to express the scFV antibody on the phage capsid.
- Screening of the scFv expressed in this way against an antigen protein or against a peptide-MHC complex may prepare a monoclonal scFV antibody specific to the antigen protein or the peptide-MHC complex.
- preparation of mRNA, preparation of cDNA, subcloning to phagemid or transduction to E. coli, phage infection, and screening of a monoclonal scFV antibody specific to an antigen protein or a peptide-MHC complex each may be performed by the known method.
- subcloning of a scFV gene to a phagemid vector containing two elements consisting of a gene fragment encoding a leader sequence (signal sequence) and a phage capsid protein iii and a replication origin of M13 and using of M13 phage as a phage can express a scFV antibody on the M13 phage.
- a phage obtained by screening may be infected to a specific bacterium and cultured, so that a monoclonal antibody specific to an antigen protein may also be collected in large quantities from the culture.
- an scFV antibody but also an antibody fragment having no constant region, such as a Fab antibody fragment, may be prepared.
- the present disclosure may include phage display libraries, in which the heavy and light chain variable regions of an antibody may be synthesized such that they include nearly all possible specificities.
- the present disclosure may include generation of phage display libraries containing phage other than M13.
- Other bacteriophages such as lambda phage, may also be useful in the method of the present disclosure.
- Lambda phage display libraries have been generated, which display peptides encoded by heterologous DNA on their surface (Sternberg et al., 1995; the content of which is hereby incorporated by reference in its entirety).
- the method of the present disclosure may be extended to include viruses other than bacteriophage, such as eukaryotic viruses.
- Eukaryotic viruses may be generated that encode genes suitable for delivery to a mammal and that encode and display an antibody capable of targeting a specific cell type or tissue into which the gene is to be delivered.
- retroviral vectors have been generated, which display functional antibody fragments (Russell et al., 1993; the content of which is hereby incorporated by reference in its entirety).
- a TCR including a soluble TCR.
- the present disclosure further relates to a TCR comprising an alpha chain and a beta chain (“alpha/beta TCRs”).
- a peptide disclosed herein that is capable of binding to a TCR and an antibody when presented by an MHC molecule.
- T-cell receptors bind to short protein fragments, such as the peptides disclosed herein, that are presented on the surface of most nucleated cells by MHC molecules.
- Each T cell expresses a unique TCR that is generated by randomly assorting genes. This ensures that the cellular immune response can respond to almost any infection and all sorts of cancer.
- a respective TCR may be a TCR that has been isolated from a given T cell clone.
- Single cell TCR alpha and beta chains can be sequenced using emulsion-based PCR techniques. Such technique may be combined with a PCR-based single cell barcoding strategy and next-generation sequencing (NGS) (De Simone et al., 2018).
- NGS next-generation sequencing
- MHC bound peptides can be recognized by the immune system, specifically T lymphocytes.
- T lymphocytes expressing a TCR that specifically binds a peptide can destroy the cells presenting the recognized MHC-peptide complex, e.g. acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer cells presenting the derived peptides.
- acute myeloid leukemia breast cancer
- cholangiocellular carcinoma chronic lymphocytic leukemia
- the present disclosure also relates to a fragment of a TCR provided herein that is capable of binding to a peptide antigen provided herein when presented by an MHC molecule.
- the term particularly relates to a soluble TCR fragment, for example a TCR lacking the transmembrane part and/or constant regions, a single chain TCR, and a fusion thereof, for example to an antibody.
- the generation of soluble TCR fragments, specifically binding to an antigen have been disclosed more than 30 years ago (Novotny, J, et al., Proc. Nati. Acad. Sci. USA (1991 ) 88, 19, 8646-8650).
- TCR fragment examples include, but are not limited to, a fragment of the a, p, 8 or the y chain and a combination of any of these.
- a TCR fragment may contain one or both of a Va-Ca or Vp-Cp or a portion thereof.
- a functional TCR fragment generally contains a variable domain such as V a , Vp, V y and/or V 8.
- the TCR fragment may in some embodiments be a single-chain TCR (scTCR), for example having an alpha chain variable region linked to a beta chain variable region.
- a TCR or TCR fragment may in some embodiments be humanized or a human TCR or TCR fragment.
- a provided TCR is a cloned TCR engineered into an autologous or allogeneic T cell.
- Provided is also a method of generating a T cell as disclosed herein.
- Provided is also a method of generating an NK cell or another cell bearing said TCR or cross-reacting with said TCR.
- the alpha and beta chains of alpha/beta TCR's, and the gamma and delta chains of gamma/delta TCRs, are generally regarded as each having two "domains", namely variable and constant domains.
- the variable domain consists of a concatenation of the variable region (V) and the joining region (J).
- the variable domain may also include a leader region (L).
- Beta and delta chains may also include a diversity region (D).
- the alpha and beta constant domains may also include C-terminal transmembrane (TM) domains that anchor the alpha and beta chains to the cell membrane.
- alpha/beta heterodimeric TCRs of the present disclosure may have a TRAC constant domain sequence and a TRBC1 or TRBC2 constant domain sequence, and the TRAC constant domain sequence and the TRBC1 or TRBC2 constant domain sequence of the TCR may be linked by the native disulfide bond between Cys4 of exon 2 of TRAC and Cys2 of exon 2 of TRBCI or TRBC2.
- a TCR of the present disclosure having at least one mutation in the alpha chain and/or having at least one mutation in the beta chain has modified glycosylation compared to the unmutated TCR.
- the present disclosure further relates to a method of identifying and isolating a TCR according to the present disclosure.
- the method includes incubating PBMCs from healthy donors negative for the respective HLA allotype with HLA-peptide monomers, incubating the PBMCs with tetramer-phycoerythrin (PE) and isolating the high avidity T cells by fluorescence activated cell sorting (FACS) Calibur analysis.
- PE tetramer-phycoerythrin
- the disclosure provides a method of producing a TCR as described herein, the method comprising culturing a host cell capable of expressing the TCR under conditions suitable to promote expression of the TCR.
- a soluble T cell receptor can be generated from a specific T cell clone, and its affinity can be increased by mutagenesis targeting the complementarity-determining regions.
- phage display can be used (US 2010/0113300, Liddy et al., 2012, the contents of which are incorporated by reference in their entirety).
- alpha and beta chains can be linked e.g.
- the T cell receptor can be linked to a toxin, a drug, a cytokine (see for example US 2013/0115191 , the content of which is incorporated by reference in its entirety), and a domain recruiting effector cells such as an anti-CD3 domain, etc., in order to execute particular functions on target cells. Moreover, it could be expressed in T cells used for adoptive transfer.
- the present disclosure further relates to a method of identifying and isolating a TCR according to the present disclosure, said method comprising obtaining a transgenic mouse with the entire human TCRap gene loci (1.1 and 0.7 Mb), whose T cells express a diverse human TCR repertoire that compensates for mouse TCR deficiency, immunizing the mouse with a peptide, incubating PBMCs obtained from the transgenic mice with tetramer-phycoerythrin (PE), and isolating the high avidity T cells by fluorescence activated cell sorting (FACS) Calibur analysis.
- a transgenic mouse with the entire human TCRap gene loci 1.1 and 0.7 Mb
- T cells express a diverse human TCR repertoire that compensates for mouse TCR deficiency
- immunizing the mouse with a peptide immunizing the mouse with a peptide
- nucleic acids encoding TCR-alpha and/or TCR-beta chains of the present disclosure are cloned into expression vectors, such as gamma retrovirus or lentivirus.
- the recombinant viruses are generated and then tested for functionality, such as antigen specificity and functional avidity.
- An aliquot of the final product is then used to transduce the target T cell population (generally purified from patient PBMCs), which is expanded before infusion into the subject / patient.
- TCR RNAs are synthesized by techniques known in the art e.g. in vitro transcription systems. The in vitro synthesized TCR RNAs are then introduced into primary CD8-positive T cells obtained from healthy donors by electroporation to re-express tumor specific TCR-alpha and/or TCR-beta chains.
- a nucleic acid encoding a TCR of the present disclosure may be operably linked to a strong promoter, such as a retroviral long terminal repeat (LTR), cytomegalovirus (CMV), murine stem cell virus (MSCV) U3, phosphoglycerate kinase (PGK), p-actin, ubiquitin, and a simian virus 40 (SV40)/CD43 composite promoter, elongation factor (EF)-1 a and the spleen focus-forming virus (SFFV) promoter.
- a strong promoter such as a retroviral long terminal repeat (LTR), cytomegalovirus (CMV), murine stem cell virus (MSCV) U3, phosphoglycerate kinase (PGK), p-actin, ubiquitin, and a simian virus 40 (SV40)/CD43 composite promoter, elongation factor (EF)-1 a and the spleen focus-forming virus (SFFV) promote
- Nucleic acids encoding TCRs of the present disclosure may be codon optimized to increase expression from a host cell. Redundancy in the genetic code allows some amino acids to be encoded by more than one codon, but certain codons are less “optimal” than others because of the relative availability of matching tRNAs as well as other factors (Gustafsson et al., 2004).
- TCR-alpha and TCR-beta gene sequences such that each amino acid is encoded by the optimal codon for mammalian gene expression, as well as eliminating mRNA instability motifs or cryptic splice sites, has been shown to significantly enhance TCR-alpha and TCR-beta gene expression (Scholten et al., 2006, the contents of these references are herein incorporated by reference in their entirety).
- mispairing between the introduced and endogenous TCR chains may result in the acquisition of specificities that pose a significant risk for autoimmunity.
- the formation of mixed TCR dimers may reduce the number of CD3 molecules available to form properly paired TCR complexes, and therefore can significantly decrease the functional avidity of the cells expressing the introduced TCR (Kuball et al., 2007, the contents of which are incorporated by reference in their entirety).
- the C-terminus domain of the introduced TCR chains of the present disclosure may be modified in order to promote interchain affinity, while decreasing the ability of the introduced chains to pair with the endogenous TCR.
- These strategies may include replacing the human TCR-alpha and TCR-beta C-terminus domains with their murine counterparts (murinized C-terminus domain); generating a second interchain disulfide bond in the C-terminus domain by introducing a second cysteine residue into both the TCR-alpha and TCR-beta chains of the introduced TCR (cysteine modification); swapping interacting residues in the TCR-alpha and TCR-beta chain C-terminus domains (“knob-in-hole”); and fusing the variable domains of the TCR-alpha and TCR-beta chains directly to CD3 ⁇ (CD3 ⁇ fusion) (Schmitt et al., 2009).
- a host cell is engineered to express a TCR of the present disclosure.
- the host cell is a human T cell or T cell progenitor.
- the T cell or T cell progenitor is obtained from a subject suffering from cancer.
- the T cell or T cell progenitor is obtained from a healthy donor.
- Host cells of the present disclosure can be allogeneic or autologous with respect to a subject to be treated.
- the host is a gamma/delta T cell transformed to express an alpha/beta TCR.
- a nucleic acid molecule as a binding molecule is typically an aptamer.
- An aptamer as provided herein may be an aptamer that specifically binds to a peptide as disclosed herein.
- An aptamer (see for example WO 2014/191359 and the literature as cited therein) is a short single-stranded nucleic acid molecule, which can fold into a defined three- dimensional structure and recognize a specific target structure. Aptamers have appeared to be suitable alternatives for developing targeted therapies. Aptamers have been shown to selectively bind to a variety of complex targets with high affinity and specificity.
- Aptamers recognizing cell surface located molecules have been identified within the past decades and provide means for developing diagnostic and therapeutic approaches. Since aptamers have been shown to possess almost no toxicity and immunogenicity, they are promising candidates for biomedical applications. Indeed aptamers, for example prostate-specific membrane-antigen recognizing aptamers, have been successfully employed for targeted therapies and shown to be functional in xenograft in vivo models. Furthermore, aptamers recognizing specific tumor cell lines have been identified.
- a DNA aptamer can be selected to reveal broad-spectrum recognition properties for various cancer cells, and particularly those derived from solid tumors, while non- tumorigenic and primary healthy cells are not recognized. If the identified aptamers recognize not only a specific tumor sub-type but rather interact with a series of tumors, this renders the aptamers applicable as so-called broad-spectrum diagnostics and therapeutics.
- Aptamers can be selected against complex targets such as cells and tissues and complexes of the peptides comprising, including consisting of, a sequence according to any of SEQ ID NOs: 1 to 59 disclosed herein with an MHC molecule. Selection may be done using the cell-SELEX (Systematic Evolution of Ligands by Exponential enrichment) technique.
- a binding molecule such as a TCR or an antibody, is coupled to a toxin.
- Peptides, binding molecules, such as antibodies or TCRs, and T lymphocytes as disclosed herein may be labeled for detection by any desired imaging method. Additionally, labels may be bi- or multi-functional and may be detectable by more than one method.
- the peptide, binding molecule, or T-lymphocyte may be directly or indirectly labeled. Attachment of a label to a binding molecule includes covalent attachment of the label, incorporation of the label into the binding molecule, and the covalent attachment of a chelating compound for binding of label, amongst others well recognized in the art. Methods for detection of labels include, but are not limited to, fluorescence, light, confocal and electron microscopy; magnetic resonance imaging and spectroscopy; fluoroscopy, computed tomography and positron emission tomography.
- Each binding molecule can include a labelling which provides that the bound binding molecule can be detected by determining the presence or absence of a signal provided by the label.
- the binding molecule can be labelled with a fluorescent dye or any other applicable cellular marker molecule.
- marker molecules are well known in the art.
- a fluorescence-labelling for example provided by a fluorescence dye, can provide a visualization of the bound aptamer by fluorescence or laser scanning microscopy or flow cytometry.
- a binding molecule as disclosed herein in particular, a proteinaceous binding molecule such as a TCR or an antibody or an TCR or antibody fragment, may have a detectable label.
- the binding molecule may in some embodiments be covalently coupled to a detectable label.
- a detectable label may be selected from the group consisting of a radionuclide, a fluorophore and biotin.
- a binding molecule of the present disclosure may be conjugated to a therapeutically or diagnostically active agent, such as a radionuclide, a chemotherapeutic agent, or a toxin.
- a respective label may be used in monitoring of a cancer therapy or in diagnosis of cancer.
- Immunohistochemical techniques are, for example, established in the art in diagnosis since decades.
- the disease tissue sample may be fresh or frozen or may be embedded in paraffin and fixed with a preservative such as formalin.
- the fixed or embedded section contains the sample are contacted with a labeled primary antibody and secondary antibody, wherein the antibody is used to detect the expression of the proteins in situ.
- Detectable labels of a peptide, binding molecule or T-lymphocyte may be for example fluorophores, enzymes, substrates, bioluminescent materials, radioactive materials, radioisotopes such as positron emitting radionucleotides, gold, gadolinium and other lanthanides, paramagnetic iron and chemiluminescent labels.
- Exemplary enzymes labels include, but are not limited to, horseradish peroxidase, acetylcholinesterase, alkaline phosphatase, b-galactosidase and luciferase.
- fluorophore include, but are not limited to, eosin, rhodamine, fluorescein, fluorescein isothiocyanate, umbelliferone, dichlorotriazinylamine, phycoerythrin and dansyl chloride.
- chemiluminescent labels include, but are not limited to, luminol.
- bioluminescent materials include, but are not limited to, luciferin and aequorin.
- radioactive materials include radionuclides.
- radionuclides are carbon-13 ( 13 C), nitrogen-15 ( 15 N), deuterium ( 2 H), as well as any combination thereof.
- Further examples of radionuclides include, but are not limited to, beta-emitters such as phosphorus-32 (32P), scandium-47 (47Sc), copper-67 (67Cu), gallium-67 (67Ga), yttrium-88 (88Y), yttrium-90 (90Y), iodine-125 (1251), iodine-131 (1311), samarium-153 (153Sm), lutetium-177 (177Lu), rhenium-186 (186Re), rhenium-188 (188Re), and alphaemitters such as astatine-211 (211 At), lead-212 (212Pb), bismuth-212 (212Bi), bismuth- 213 (213Bi), actinium-225 (225Ac) or any combination thereof.
- a peptide as disclosed herein is used to generate activated T lymphocytes in vitro.
- nucleic acids encoding binding molecules, such as T cells receptors as disclosed herein can be used to genetically engineer T lymphocytes in vitro.
- Such genetically engineered T lymphocytes express T cell receptors that specifically recognize a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule.
- the T lymphocytes obtained by either method may be used for administration to an individual.
- T cell-based therapies have been successfully developed in clinical trials.
- TIL therapy tumor-infiltrating lymphocyte therapy
- T lymphocytes within tumor tissue are often enriched in clones with specificity for peptides that are presented only on cells of the tumor, the efficiency of TIL therapy is based on the proliferation of such T lymphocytes.
- T lymphocytes are isolated from a subject’s tumor, stimulated and expanded in vitro, and administered back into the patient.
- T cells isolated from a donor peripheral blood cells are engineered to express TCRs of desired specificity and affinity to a peptide presented on cancer cells.
- the donor may be the subject of the treatment (autologous therapy) or different from the subject of the treatment (allogeneic therapy).
- peripheral blood mononuclear cells (PBMCs) are obtained from donors and/or from the subject, and T lymphocytes binding to one or more desired peptides identified as being presented on cells of the cancer to be treated are enriched and isolated by FAC-sorting.
- T cell clones are isolated and the TCRs sequenced, after T lymphocytes responsive to the desired peptide(s) have been isolated and expanded.
- T-cell therapy may rely on T lymphocytes obtained directly from the same individual, which is an autologous approach. T lymphocytes obtained from the same individual can be grown and selected in culture, and infused back into the individual for the treatment of the cancer concerned. In such an autologous setting, T lymphocytes obtained from the individual can be engineered to express a TCR transgene specific for a peptide disclosed herein. The recombinant TCR confers to the respective T lymphocytes the ability to recognize the peptide disclosed herein and kill tumor cells. The recombinant TCR may also be an affinity-maturated TCR. T-cell engineering also allows constructing a TCR with optimized affinity, selecting the most appropriate T-cell subset to be transduced and given to the individual, and to control over its in vivo efficacy and safety.
- T-cell therapy may also rely on T lymphocytes obtained from healthy donors other than the individual to be treated, which is an allogeneic approach.
- suitable lymphocytes such as gamma delta T lymphocytes
- T lymphocytes obtained from a donor may be engineered to express so called “suicide genes”, which renders expressing cells sensitivity to a prodrug.
- suicide genes which renders expressing cells sensitivity to a prodrug.
- Another aspect of the present disclosure involves an in vitro method for producing T lymphocytes.
- These methods may include the use of the peptides complexed with MHC (e.g. as presented on an antigen-presenting cell as described below) to generate a T cell receptor whose nucleic acid is cloned and is introduced into a host cell, for example a T lymphocyte.
- This engineered T lymphocyte can then be transferred to a subject for therapy of cancer.
- PBMCs are taken from a donor and the desired T lymphocyte subset for genetic engineering is enriched, e.g. by depleting monocytes or alpha beta T cells.
- the donor may or may not be the subject of the treatment.
- the T lymphocytes can then be activated, e.g. in the presence of anti-CD3 and anti- CD28 in case of alpha beta T cells for autologous T cell therapy, and subsequently be transduced with a nucleic acid or vector encoding the TCR.
- Suitable vectors include, e.g. lentiviral vectors.
- T lymphocytes may be cultured in the presence of suitable cytokines such as cytokine interleukin 7 (IL-7) and/or interleukin 15 (IL-15).
- suitable cytokines such as cytokine interleukin 7 (IL-7) and/or interleukin 15 (IL-15).
- IL-7 cytokine interleukin 7
- IL-15 interleukin 15
- T lymphocytes may be activated with zoledronate before being subjected to genetic engineering.
- An exemplary process is described in WO 2019104269 A1 , which is incorporated herein by reference in its entirety.
- the method may include contacting in vitro T lymphocytes with peptide loaded human MHC molecules expressed on the surface of a suitable antigen-presenting cell for a period of time sufficient to activate the T lymphocyte in an antigen specific manner, wherein the antigen is a peptide according to the invention.
- activated T lymphocytes can either be clonally expanded and reinfused into the subject, or be used to generate TCRs that are used to genetically engineer suitable T lymphocytes for immunotherapy as described above.
- a sufficient amount of the antigen is used with an antigen-presenting cell.
- the human peptide loading deficient cell line T2 is available from the American Type Culture Collection, 12301 Parklawn Drive, Rockville, Maryland 20852, USA under Catalogue No CRL 1992; the Drosophila cell line Schneider line 2 is available from the ATCC under Catalogue No CRL 19863; the mouse RMA-S cell line is described in Ljunggren et al. (Ljunggren and Karre, 1985, the contents of which are incorporated by reference in their entirety).
- the host cell expresses substantially no MHC class I molecules.
- the stimulator cell expresses a molecule important for providing a co-stimulatory signal for T cells such as any of B7.1 , B7.2, ICAM-1 and LFA3.
- the nucleic acid sequences of numerous MHC class I molecules and of the co-stimulator molecules are publicly available from the GenBank and EMBL databases.
- the T lymphocytes may be CD8-positive T lymphocytes or cytotoxic T lymphocytes (CTLs). These T cells may be further expanded and either directly reinfused into the patient, or used to obtain a suitable TCR that is introduced into a suitable host cell, typically a T lymphocyte (e.g., a CD8- positive T lymphocyte, a CD4-positive T lymphocyte or a gamma delta T lymphocyte).
- a T lymphocyte e.g., a CD8- positive T lymphocyte, a CD4-positive T lymphocyte or a gamma delta T lymphocyte.
- the cell may comprise an expression vector capable of expressing a peptide containing the sequence of one of SEQ ID NO: 1 to SEQ ID NO: 59.
- Walter et al. (Walter et al., 2003, the contents of which are incorporated by reference in their entirety) describe the in vitro priming of T cells by using artificial antigen-presenting cells (aAPCs), which is also a suitable way for generating T cells against the peptide of choice.
- aAPCs artificial antigen-presenting cells
- the inventors have generated aAPCs by the coupling of preformed MHC-peptide complexes to the surface of polystyrene particles (microbeads) by biotin:streptavidin biochemistry. This system permits the exact control of the MHC density on aAPCs, which allows to selectively elicit high- or low-avidity antigen specific T cell responses with high efficiency from blood samples.
- T cells may also be used in the preparation of T cells and a method is described in detail in WO 97/26328, incorporated herein by reference in its entirety.
- other cells may be used to present antigens such as CHO cells, baculovirus-infected insect cells, bacteria, yeast, and vaccinia- infected target cells.
- plant viruses may be used see for example Porta et al. (Porta et al., 1994, the content of which is incorporated by reference in its entirety) which describes the development of cowpea mosaic virus as a high-yielding system for the presentation of foreign peptides.
- T lymphocytes which are produced by the above method, will selectively recognize a cell that aberrantly expresses a peptide that comprises an amino acid sequence of SEQ ID NO: 1 to SEQ ID NO: 59.
- the T lymphocyte recognizes the target cell that presents a peptide as disclosed herein by interacting through its TCR with the MHC peptide complex (for example, binding).
- the T lymphocytes are useful in a method of killing target cells in a subject whose target cells aberrantly express a polypeptide or peptide that includes an amino acid sequence as provided herein wherein the subject is administered an effective number of the T lymphocytes.
- the T lymphocytes that are administered to the subject may be derived from the subject and optionally activated as described above (i.e. they are autologous T cells).
- the T lymphocytes are not from the subject but are from another individual. In typical embodiments the individual is a healthy individual.
- the target cells for the T lymphocytes may be cells of the tumor (which sometimes express MHC class II) and/or stromal cells surrounding the tumor (which sometimes also express MHC class II) (Dengjel et al., 2006).
- T lymphocytes provided herein may be used as active ingredients of a therapeutic composition.
- T lymphocytes may be obtained by methods known in the art, e.g., those described above.
- any molecule provided herein i.e. the peptide, nucleic acid, antibody, expression vector, cell, activated T cell, T cell receptor or the nucleic acid encoding it, is useful for the treatment of disorders, characterized by cells escaping an immune response. Therefore, any molecule provided herein may be used as medicament or in the manufacture of a medicament. The molecule may be used by itself or combined with other molecule(s) provided herein or (a) known molecule(s).
- any molecule provided herein may also be included in a pharmaceutical composition.
- the pharmaceutical composition provided herein may be included in a pharmaceutical combination, such as a combination product.
- Two pharmaceutical compositions that differ from each other in their active components, may define the combination product.
- the compositions of a combination product may be administered independently.
- the compositions of a combination product may be administered at different points of time.
- One of the compositions of a combination product may be administered in a time-delayed manner relative to another composition of the combination product.
- the compositions of a combination product may be administered concomitantly.
- a pharmaceutical composition as provided herein such as a therapeutic composition or a vaccine composition, is provided in the form of a solution.
- the components of the pharmaceutical composition need to be or are dissolved or suspended in a pharmaceutically acceptable liquid vehicle such as water, an organic solvent, a mixture of both or one or more pharmaceutically acceptable oils or fats.
- a composition such as a therapeutic composition or a vaccine composition as disclosed herein may be provided in lyophilized form or in freeze-dried form. The lyophilized form may when needed be solubilized in water, for example for injection, and gently mixed.
- an immunoadjuvant as described hereinabove may then be added. The mixture may then be gently mixed and charged into a suitable syringe.
- a pharmaceutical composition as disclosed herein may be an individualized pharmaceutical composition in that the composition has been specifically tailored for an individual.
- An individualized pharmaceutical composition has generally been verified to contain an active ingredient that is predicted to be effective in the particular individual.
- a pharmaceutical composition as disclosed herein may also be provided without being verified for any individual.
- the pharmaceutical composition may for example contain a peptide as disclosed herein and be administered as a vaccine on the basis of a risk assessment, without knowing whether there is an increased risk that the peptide will be or is presented on cancer cells of the individual.
- a peptide as disclosed herein may be used for risk stratification, for diagnosis of existing cancer, for monitoring for a risk level of cancer, including for a potential risk increase, and for predicting a future medical outcome.
- Determining presentation, including the presentation level, of a peptide as disclosed herein can be used to stratify a subject undergoing or about to undergo cancer treatment. Determining presentation, including the presentation level, of a peptide as disclosed herein can also be used to stratify a subject for vaccination.
- biomarkers for stratification of patients is a procedure well established in the art.
- This procedure includes or consists of linking one or more patient subpopulations, characterized by a certain feature, in the context of the present disclosure the over-presentation of a particular peptide by MHC on cancer cells, to a particular treatment.
- the general aim of stratification is to match patients with therapies that are more likely to be effective and safe.
- stratifying patients may in some embodiments include evaluation of patient history and physical assessment, combined with laboratory tests on the basis of a method as disclosed herein, and clinical observation.
- the method includes assessing whether cells of a cancer sample from the subject present a peptide as disclosed herein.
- the method also includes selecting the subject for treatment with a vaccine and/or an immunotherapeutic agent, if cells in the cancer sample over-present a peptide as disclosed herein, relative to one or more corresponding healthy tissue samples or cells.
- the vaccine includes a peptide that contains a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a pharmaceutically acceptable salt thereof.
- the immunotherapeutic agent in some embodiments includes the binding molecule as disclosed herein.
- a method of producing, e.g. formulating or synthesizing a pharmaceutical product, such as a therapeutic product may in some embodiments be a vaccine.
- the pharmaceutical product may in some embodiments be a compound-based product.
- the pharmaceutical product may in some embodiments be a cellular product.
- the individualized pharmaceutical product may be an anticancer vaccine or a compound-based and/or cellular therapeutical product.
- the individualized pharmaceutical product includes a peptide as provided herein.
- the collection of peptides may contain HLA-A*02, HLA-A*01 , HLA- A*03, HLA-A*24, HLA-B*07, HLA-B*08 and HLA-B*44 marker peptides.
- HLA-A*02, HLA-A*01 , HLA- A*03, HLA-A*24, HLA-B*07, HLA-B*08 and HLA-B*44 marker peptides are examples of the magnitude of T-cell immunity induced by peptides in a quantitative manner and hence allow important conclusion to be drawn on the capacity of the vaccine to elicit anti-tumor responses.
- Peptides for the collection of peptides may be identified by using an integrated functional omics approach combining gene expression analysis, mass spectrometry, and T cell immunology (XPresident ®). The approach assures that only peptides truly present on a high percentage of tumors but not or only minimally presented on normal tissue, are chosen for further analysis.
- colorectal cancer, glioblastoma, gastric cancer, hepatocellular carcinoma, and renal cell carcinoma samples from subjects, e.g. patients, were analyzed. Further tissues and bodily fluids from healthy donors were analyzed. Both types of samples were analyzed in a stepwise approach:
- HLA ligands from the malignant and healthy samples were identified by mass spectrometry
- mRNA expression analysis was used to identify genes over-expressed in the malignant tissue compared with a range of normal organs and tissues.
- HLA ligands were compared to gene expression data. Peptides overpresented or selectively presented on tumor tissue, preferably encoded by selectively expressed or over-expressed genes as detected in step 2 were considered suitable peptide candidates for a multi-peptide vaccine.
- the one or more peptides included in the individualized pharmaceutical product is/are identified by a method that involves: (a) identifying the peptides presented by a tumor sample from the individual subject; (b) comparing the peptides identified in (a) with a collection (e.g. a database) of peptides as described above; and (c) selecting one or more peptides from the collection of peptides (database) that correlates with a tumor-associated peptide identified in the subject.
- the peptide(s) presented by the tumor sample can be identified by: (a1 ) comparing expression data from the tumor sample to expression data from a sample of normal tissue corresponding to the tissue type of the tumor sample to identify proteins that are overexpressed or aberrantly expressed in the tumor sample; and (a2) correlating the expression data with sequences of MHC ligands bound to MHC class I and/or class II molecules in the tumor sample to identify MHC ligands derived from proteins overexpressed or aberrantly expressed by the tumor.
- the sequences of MHC ligands are identified by eluting bound peptides from MHC molecules isolated from the tumor sample, and sequencing the eluted ligands.
- the tumor sample and the normal tissue have been or are obtained from the same subject, (a) identifying TUMAPs presented by a tumor sample from the individual subject by the method as described above; (b) comparing the peptides identified in (a) with a warehouse of peptides that have been prescreened for immunogenicity and over presentation in tumors as compared to corresponding normal tissue; (c) selecting at least one peptide from the warehouse that correlates with a tumor-associated peptide identified in the subject; and (d) optionally, selecting at least one peptide identified de novo in (a) confirming its immunogenicity.
- the peptide(s) included in the individualized pharmaceutical product is/are identified by: (a) identifying one or more peptides presented by a tumor sample from the individual subject; and (b) selecting at least one peptide identified de novo in (a) and confirming its immunogenicity.
- the single peptide-DMSO solutions may be mixed in equal parts to achieve a solution containing all peptides to be included in the product with a concentration of ⁇ 2.5 mg/ml/peptide.
- the mixed solution may then for example be diluted 1 :3 with water for injection to achieve a concentration of 0.826 mg/ml/peptide in 33% DMSO.
- the diluted solution is typically filtered through a 0.22 pm sterile filter. A final bulk solution can thus be obtained.
- the method may also include selecting a peptide that contains the mutation for the generation of a cellular therapeutic or for the generation of a cellular therapeutic combination.
- a cancer vaccine for use in a method of inducing an immune response against cancer cells of a subject.
- the individualized cancer vaccine contains a nucleic acid molecule that encodes a recombinant protein or peptide.
- the recombinant protein or peptide contains an amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
- the method of inducing an immune response involves administering the vaccine to the subject. Expression of the recombinant protein or peptide in cells of the subject results in MHC class I presentation of a peptide having a sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
- the MHC class l-presented peptide having a sequence selected from the group consisting of SEQ ID NOs: 1 to 59 is capable of eliciting a CD8+ T cell response against cancer cells of the subject that present the peptide having a sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
- administering the vaccine is done intravenously or intraperitoneally. In some embodiments administering the vaccine is done intramuscularly or subcutaneously. In some embodiments administering the vaccine is done transdermally or intranodally.
- a method for producing an individualized anti-cancer vaccine for an individual subject contains a nucleic acid molecule that encodes a recombinant protein or peptide.
- the recombinant protein or peptide contains an amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
- the method includes identifying one or more peptides presented on cells in a tumor sample from the individual subject.
- the method furthermore includes comparing the identified peptide(s) with a collection or dataset of peptides that have been pre-screened for immunogenicity and/or over-presentation in tumor cells as compared to healthy cells, e.g. cells of normal tissues.
- the collection or dataset of peptides includes one or more peptides that contain a sequence selected from the group consisting of SEQ ID NO: 1 to SEQ ID NO: 59.
- the peptide(s) has/have a maximal length of up to 30 amino acids, including of up to 16 amino acids. In some embodiments, the peptide(s) has/have an overall length of up to 12 amino acids.
- the collection or dataset of peptides includes one or more peptides that consist of a sequence selected from the group consisting of SEQ ID NO: 1 to SEQ ID NO: 59.
- the method furthermore includes assessing whether the one or more peptide(s) presented on cells of the tumor sample is/are included in the collection or dataset of peptides.
- a nucleic acid molecule that encodes the recombinant protein or peptide is selected for inclusion into the individualized anti-cancer vaccine.
- the method in this case furthermore includes manufacturing and/or producing the individualized anti-cancer vaccine.
- the cancer vaccine is an individualized cancer vaccine.
- cancer cells, but not healthy cells, of the subject present a peptide containing an amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
- cancer cells, but not healthy cells, of the subject present a peptide consisting of an amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
- the immunogenicity of the peptides included in the collection of peptides is determined by a method that involves an in vitro immunogenicity assay and/or subject immunomonitoring for individual MHC binding. In some embodiments, the immunogenicity of the peptides included in the collection of peptides is determined by a method that involves MHC multimer staining and/or an ELISPOT assay. In some embodiments, the immunogenicity of the peptides included in the collection of peptide is determined by a method that involves intracellular cytokine staining
- mRNA expression analysis on a sample obtained from the subject by highly parallel methods, such as microarrays or sequencing-based expression profiling, comprising identifying genes that are overexpressed in a malignant tissue, compared to a normal tissue or tissues; - selecting peptides encoded by selectively expressed or over-expressed genes as detected in the genome-wide mRNA expression analysis, and
- determining an induction of an in vivo T-cell response by the peptides as selected comprising in vitro immunogenicity assays using human T lymphocytes from a healthy donor or from the subject.
- mRNA expression analysis by highly parallel methods, such as microarrays or sequencing-based expression profiling, comprising identifying genes that are over-expressed in a malignant tissue, compared with a normal tissue or tissues;
- BiolVT (Detroit, Ml, USA & Royston, Herts, UK); Bio-Options Inc. (Brea, CA, USA); BioServe (Beltsville, MD, USA); Capital BioScience Inc. (Rockville, MD, USA); Conversant Bio (Huntsville, AL, USA); Cureline Inc. (Brisbane, CA, USA); DxBiosamples (San Diego, CA, USA); Geneticist Inc. (Glendale, CA, USA); Indivumed GmbH (Hamburg, Germany); Kyoto Prefectural University of Medicine (KPUM) (Kyoto, Japan); Osaka City University (OCU) (Osaka, Japan); ProteoGenex Inc.
- KPUM Kyoto Prefectural University of Medicine
- OCU Osaka City University
- ProteoGenex Inc ProteoGenex Inc.
- HLA peptide pools from shock-frozen tissue samples were obtained by immune precipitation from solid tissues according to a slightly modified protocol (Falk et al. , 1991 ; Seeger et al., 1999) using the HLA-A*02 specific antibody BB7.2, the HLA-A, -B, -C specific antibody w6/32, the HLA-DR specific antibody L243 and the HLA-DP specific antibody B7/21 , CNBr-activated sepharose, acid treatment, and ultrafiltration.
- Table 4 shows the peptides and the HLA allotypes - from the group consisting of HLA- A*01 :01 , HLA-A*02:01 , HLA-A*03:01 , HLA-A*24:02, HLA-B*07:02, HLA-B*08:01 and HLA-B*44:02 - they bind to.
- HLA-A*01 binding peptides also binding to HLA-B*15
- HLA-A*03 binding peptides also binding to HLA-A*11
- HLA-B*07 binding peptides also binding to HLA-B*35 and HLA-B*51.
- HLA peptide pools as obtained were separated according to their hydrophobicity by reversed-phase chromatography (nanoAcquity UPLC system, Waters) and the eluting peptides were analyzed in LTQ velos and fusion hybrid mass spectrometers (ThermoElectron) equipped with an ESI source.
- Peptide pools were loaded directly onto the analytical fused-silica micro-capillary column (75 pm i.d. x 250 mm) packed with 1.7 pm C18 reversed-phase material (Waters) applying a flow rate of 400 nL per minute.
- each identified peptide can be associated with quantitative data allowing relative quantification between samples and tissues.
- all quantitative data acquired for peptide candidates was inspected manually to assure data consistency and to verify the accuracy of the automated analysis.
- a presentation profile was calculated showing the mean sample presentation as well as replicate variations.
- Cancer type AML (acute myeloid leukemia); BRCA (breast cancer); CCC (cholangiocellular carcinoma); CLL (chronic lymphocytic leukemia); CRC (colorectal cancer); GBC (gallbladder cancer); GBM (glioblastoma); GC (gastric cancer); GEJC (gastro-esophageal junction cancer); HCC (hepatocellular carcinoma); HNSCC (head and neck squamous cell carcinoma); MEL (melanoma); NHL (non-Hodgkin lymphoma); NSCLCadeno (non-small cell lung cancer adenocarcinoma); NSCLCother (NSCLC samples that could not unambiguously be assigned to NSCLCadeno or NSCLCsquam); NSCLCsquam (squamous
- RNA from healthy human tissues for RNASeq experiments was obtained from: Asterand (Detroit, Ml, USA & Royston, Herts, UK); Bio-Options Inc. (Brea, CA, USA); Geneticist Inc. (Glendale, CA, USA); ProteoGenex Inc. (Culver City, CA, USA); Tissue Solutions Ltd (Glasgow, UK).
- RNA from tumor tissues for RNASeq experiments was obtained from: Asterand (Detroit, Ml, USA & Royston, Herts, UK); BioCat GmbH (Heidelberg, Germany); BioServe (Beltsville, MD, USA); Geneticist Inc. (Glendale, CA, USA); Istituto Nazionale Tumori "Pascale” (Naples, Italy); ProteoGenex Inc. (Culver City, CA, USA); University Hospital Heidelberg (Heidelberg, Germany).
- RNA samples Quality and quantity of all RNA samples were assessed on an Agilent 2100 Bioanalyzer (Agilent, Waldbronn, Germany) using the RNA 6000 Pico LabChip Kit (Agilent).
- RNAseq next-generation sequencing
- GENEWIZ Germany GmbH Leipzig, Germany
- sequencing libraries were prepared from total RNA using the NEBNext® UltraTM II Directional RNA Library Prep Kit for Illumina according to the manufacturer’s instructions (New England Biolabs, Ipswich, MA, USA), which includes mRNA selection, RNA fragmentation, cDNA conversion and addition of sequencing adaptors.
- libraries were multiplexed and loaded onto the Illumina NovaSeq 6000 sequencer (Illumina Inc., San Diego, CA, USA) according to the manufacturer’s instructions, generating a minimum of 80 million 150 bp paired-end raw reads per sample.
- RNA reads supporting the peptide were counted and are shown as exemplary expression profiles of peptides of the present invention that are highly overexpressed or exclusively expressed in AML (acute myeloid leukemia); BRCA (breast cancer); CCC (cholangiocellular carcinoma); CLL (chronic lymphocytic leukemia); CRC (colorectal cancer); GBC (gallbladder cancer); GBM (glioblastoma); GC (gastric cancer); GEJC (gastro-esophageal junction cancer); HCC (hepatocellular carcinoma); HNSCC (head and neck squamous cell carcinoma); MEL (melanoma); NHL (non-Hodgkin lymphoma); NSCLCadeno (non-small cell lung cancer adenocarcinoma); NSCLCother (NSCLC samples that could not unambiguously be assigned to NSCLCadeno or NSCLCsquam); NSCLCsquam); NSCLCsquam
- Table 6 Expression scores. The table lists peptides that are very highly overexpressed in tumors compared to a panel of normal tissues (+++) or highly overexpressed in tumors compared to a panel of normal tissues (++) or overexpressed in tumors compared to a panel of normal tissues (+) according to the RNAseq experiments.
- the baseline for this score was calculated from measurements of the following relevant normal tissues: adipose tissue; adrenal gland; bile duct; bladder; bloodcells; blood vessels; bone marrow; brain; breast; esophagus; eye; gallbladder; nead&neck; heart; large intestine; small intestine; kidney; liver; lung; lymph nodes; peripheral nerve; ovary; pancreas; parathyroid gland; peritoneum; pituitary; placenta; pleura; prostate; skeletal muscle; skin; spinal cord; spleen; stomach; testis; thymus; thyroid; trachea; ureter; uterus.
- the arithmetic mean of all respective samples was used for the calculation.
- peptides were synthesized using standard and well-established solid phase peptide synthesis using the Fmoc- strategy. If necessary, stable isotope labeled (SIL-) amino acids were used to introduce a discriminating mass shift and allow for the use of these labeled peptides as internal standards (e.g. if a peptide was selected for identity confirmation in CoElution experiments). Identity and purity of each individual peptide were determined by mass spectrometry and analytical RP-HPLC. The peptides were obtained as white to off-white lyophilizes (trifluoro acetate salt) in purities of >50%. Tumor-associated peptides (TUMAPs) are usually administered as trifluoro-acetate salts or acetate salts, other saltforms are also possible.
- SIL- stable isotope labeled
- the initial validation of peptides was achieved by IdentControl via spectral comparison.
- synthetic peptides were used for validation of peptide identifications by acquisition of high-resolution reference MS2 spectra using matched fragmentation modes and collision energies as for acquisition of the natural spectra.
- Automated spectral comparison was performed using the sensitive metric of spectral correlation with a cutoff score determined to result in 90% sensitivity at ⁇ 1 % FDR based on a benchmark dataset comprising >10,000 manually validated spectra.
- Ambiguous identifications were further subjected to validation in CoElution experiments.
- a list of peptides validated by Identcontrol can be found in Table 7 and an exemplary validation via IdentControl for SEQ ID NO: 44 is shown in Figure 3.
- Table 7 Identcontrol Results.
- the spectral correlation indicates the similarity of the MS/MS spectra from the endogenous detected peptide compared to the synthetic peptide, the higher the value the more alike the spectra are.
- the peptide is validated when a threshold of 0.872 is met, or spectra are considered identical by manual review.
- peptides were subjected to CoElution experiments using SIL internal standard peptides. To this end, SIL peptides were spiked into HLA peptidome extracts from samples and subjected to liquid chromatography - targeted mass spectrometry (LC-MS) to confirm peptide identity based on spectral similarity as well as CoElution in the retention time dimension.
- LC-MS liquid chromatography - targeted mass spectrometry
- SIL-peptide amounts were adjusted to the peptide specific ionization factors (determined in calibration curves), if necessary.
- LC-MS was performed using pre-defined targeted MS2 scan events with non-overlapping isolation windows for SIL-peptide and natural peptide species to avoid co-fragmentation.
- control experiments were performed in a non-HLA peptide containing tryptic matrix, which had to confirm absence of any unlabeled signal.
- Peptide detection and validation by CoElution was determined by manual expert review based on multiple pre-defined objective criteria, including dot product (dotP) of SIL peptide compared to unlabeled peptide MS2 traces, the presence of the most intense transitions in multiple consecutive scans and aligned peak apexes.
- dotP dot product
- Table 8 A list of peptides validated by CoElution can be found in Table 8 and the CoElution result for SEQ ID NO 58 is shown in Figure 4.
- binders such as antibodies and/or TCRs
- selection criteria include, but are not restricted to, (i) exclusiveness of presentation and (ii) the density of peptide presented on the cell surface.
- the inventors analyzed absolute peptide copies per cell as described in WO 2016/107740. The quantitation of peptide copies per cell in solid tumor samples requires the absolute quantitation of the isolated peptide, the efficiency of the peptide isolation process, and the cell count of the tissue sample analyzed.
- a calibration curve was generated for SEQ ID NO: 44 and SEQ ID NO: 48, using two different isotope labeled peptide variants (one or two isotope-labeled amino acids are included during peptide synthesis). These isotope-labeled variants differ from the tumor-associated peptide only in their mass but show no difference in other physicochemical properties (Anderson et al., 2012).
- the doubly isotope-labeled peptide also called internal standard, was further spiked to each MS sample and all MS signals were normalized to the MS signal of the internal standard to level out potential technical variances between MS experiments.
- the calibration curves were prepared in at least three different matrices, i.e., HLA peptide eluates from natural samples similar to the routine MS samples, and each preparation was measured in duplicate MS runs.
- MS signals were normalized to the signal of the internal standard and a calibration curve was calculated by logistic regression.
- the respective samples were also spiked with the internal standard; the MS signals were normalized to the internal standard and quantified using the peptide calibration curve.
- peptide-MHC complexes were generated for all peptides selected for absolute quantitation.
- single-isotope-labelled versions of the peptides were used, i.e., one isotopelabelled amino acid was included in peptide synthesis.
- These complexes were spiked into the freshly prepared tissue lysates, i.e., at the earliest possible point of the peptide isolation procedure, and then captured like the natural peptide-MHC complexes in the following affinity purification. Measuring the recovery of the single-labelled peptides therefore allows conclusions regarding the efficiency of isolation of individual natural peptides.
- the efficiency of isolation was analyzed in a small set of samples and was comparable among these tissue samples. In contrast, the isolation efficiency differs between individual peptides. This suggests that the isolation efficiency, although determined in only a limited number of tissue samples, may be extrapolated to any other tissue preparation. However, it is necessary to analyze each peptide individually as the isolation efficiency may not be extrapolated from one peptide to others. Determination of the cell count in solid, frozen tissue
- the inventors applied DNA content analysis. This method is applicable to a wide range of samples of different origin and, most importantly, frozen samples (Alcoser et al. , 2011 ; Forsey and Chaudhuri, 2009; Silva et al. , 2013).
- a tissue sample is processed to a homogenous lysate, from which a small lysate aliquot is taken. The aliquot is divided in three parts, from which DNA is isolated (QiaAmp DNA Mini Kit, Qiagen, Hilden, Germany). The total DNA content from each DNA isolation is quantified using a fluorescence-based DNA quantitation assay (Qubit dsDNA HS Assay Kit, Life Technologies, Darmstadt, Germany) in at least two replicates.
- the standard curve is used to calculate the total cell content from the total DNA content from each DNA isolation.
- the mean total cell count of the tissue sample used for peptide isolation is then extrapolated considering the known volume of the lysate aliquots and the total lysate volume.
- Table 9A Copy cell numbers for SEQ ID NO: 44 in different cancer entities.
- Table 9B Copy cell numbers for SEQ ID NO: 48 in different cancer entities.
- Tables 9A and 9B list the results of absolute peptide quantitation in cancer samples.
- Baibas R and Lorence A “Methods in Molecular Biology Recombinant Gene Expression, Reviews and Protocols” (2004). Banchereau, J. et al., Cell 106 (2001 ): 271 -274.
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- General Health & Medical Sciences (AREA)
- Medicinal Chemistry (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Biophysics (AREA)
- Biochemistry (AREA)
- Genetics & Genomics (AREA)
- Gastroenterology & Hepatology (AREA)
- Molecular Biology (AREA)
- Zoology (AREA)
- Toxicology (AREA)
- Immunology (AREA)
- Oncology (AREA)
- Microbiology (AREA)
- Mycology (AREA)
- Pharmacology & Pharmacy (AREA)
- Epidemiology (AREA)
- Animal Behavior & Ethology (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
- Peptides Or Proteins (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
Abstract
Provided is a peptide comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 1 to SEQ ID NO: 59, or a pharmaceutically acceptable salt thereof.
Description
Peptides displayed by MHC for use in immunotherapy against different types of cancer
CROSS-REFERENCE TO RELATED APPLICATIONS
The present application claims the benefit of and the priority to an application for "Peptides displayed by MHC for use in immunotherapy against different types of cancer" filed on 18 August 2023 with the European Patent Office, and there duly assigned serial number EP 23 020 387.9. The content of said application filed on 18 August 2023 is incorporated herein by reference for all purposes, including an incorporation of any element or part of the description, claims or drawings not contained herein and referred to in Rule 20.5(a) of the PCT, pursuant to Rule 4.18 of the PCT.
REFERENCE TO SEQUENCE LISTING SUBMITTED AS A COMPLIANT XML 1.0 FORMAT FILE (.xml)
Pursuant to the EFS-Web legal framework and 37 CFR §§ 1.821 -825 (see MPEP § 2442.03(a)), Rule 30 EPC, and § 11 PatV, an electronic sequence listing compliant with WIPO standard ST.26 in the form of an XML 1.0 format file is submitted concurrently with the instant application, and the entire contents of the sequence listing are incorporated herein by reference. For the avoidance of doubt, if discrepancies exist between the sequences mentioned in the specification and the electronic sequence listing, the sequences in the specification shall be deemed to be the correct ones.
FIELD OF THE DISCLOSURE
Disclosed herein are peptides that are derived from the Preferentially Expressed Antigen in Melanoma (PRAME) protein, that can be used in immunotherapeutic methods. In particular, provided is an immunotherapy of cancer. The peptides are tumor-associated T cell peptide epitopes, that can serve as an active pharmaceutical ingredient of a vaccine composition that stimulates an anti-tumor immune response. The peptides disclosed herein can also be used to bind or stimulate T cells in vitro that may be transferred into a subject. Such a peptide bound to a molecule of the major histocompatibility complex (MHC), or a peptide as such, can also be the target of an antibody, a soluble T cell receptor, bispecific T cell engager, and another binding molecule, e.g. a proteinaceous binding molecule. Such binding molecules can be used in immunotherapeutic methods, wherein soluble binding molecules can be administered directly to a subject to be treated, whereas membrane-bound binding molecules can be expressed in suitable cells that can be used for immunotherapy.
Disclosed herein are peptide sequences derived from peptides from the PRAME protein which were isolated from complexes with MHC class I molecules of human tumor cells. Said peptide sequences can be used in vaccine compositions for eliciting anti-tumor immune responses, or as targets for the development of pharmaceutically / immunologically active compounds and cells.
BACKGROUND
The following discussion of the background is merely provided to aid the reader in understanding the invention and is not meant as an admission of prior art to the present invention.
According to the World Health Organization (WHO), cancer ranked among the four major non-communicable deadly diseases worldwide in 2012. For the same year, colorectal cancer, breast cancer and respiratory tract cancers were listed within the top 10 causes of death in high income countries.
Considering the severe side-effects and expenses associated with treating cancer, there is a need to identify factors that can be used in the treatment of cancer.
There is also a need to identify factors representing biomarkers for cancer, leading to better diagnosis of cancer, assessment of prognosis, and prediction of treatment success.
The presentation of peptides on almost any cell is to be distinguished from the presentation of peptides by specialized antigen-presenting cells, such as dendritic cells, via crosspresentation. In addition to peptides derived from proteins within dendritic cells, these cells are also capable of taking up extracellular proteins and presenting fragments thereof on their cell surface.
Origin of Peptides presented by MHC molecules
The present invention can be taken to generally be based on the identification of peptides that have been found on the extracellular surface of cancer cells, where they are bound to molecules of the major histocompatibility complex (MHC). This display of peptides is part of cell-mediated immunity evolved to cope with intracellular pathogens (Berg, J. M. et al., “Biochemistry”, 10th ed. (2023), Macmillan Learning, 89-91 ; Berg, J. M. et al., “Biochemistry”, 8th ed. (2015), W. H. Freeman and Company, 995-1005; Murphy, K. et al., “Janeway's Immunobiology”, 10th ed. (2022), WW Norton & Co Inc., 29-31 , 159-175, 218-246). Cell-mediated immunity is also able to recognize cancer cells.
Almost any vertebrate cell presents on its outer surface a vast collection of peptides derived from the degradation of proteins that are found in its cytoplasm. These peptides are continuously generated by degradation. A well-studied process in this regard involves proteasomes, and routing via the endoplasmic reticulum, where further trimming occurs (Berg et al. , 2015; Murphy, 2022). Proteasomes are not the only degradation source, and it is under discussion how significant the contribution of proteasomes to the complete MHC ligandome on the cell surface is. Only very few peptides thus generated are bound by newly synthesized MHC molecules, and transported to the cell surface (Berg et al., 2015; Murphy, 2022). Once presented on the cell surface in a complex with MHC proteins, the peptides can be contacted by T cell receptors on T lymphocytes. These short peptides often correspond to sequences that are buried within the native structure of the source protein and thus inaccessible. Only unfolding and fragmentation allows these sequences to be recognized by T-cell receptors.
MHC molecules possess a peptide-binding groove, within which a matching peptide can be bound. Different MHC variants have different amino acid residues at specific positions in the binding groove, which are called anchor residues (Murphy, 2022). A respective binding groove has two, in some cases three, pockets that undergo binding with amino acid side chains. Some of the bound peptides are too long to be encompassed in the cleft, which then typically protrude outside of the same (La Gruta, 2018).
The contact of MHC molecules with the T cell receptors mediates the detection of cancerspecific peptides and the activation of T killer cells. Activated killer T cells can destroy antigen-presenting cells by perforins and granzymes.
The T cell mediated immune response thus differs from the immunoglobulin mediated immune response. While antibodies can also recognize linear epitopes, they mainly recognize conformational epitopes exposed on the surfaces of proteins with tertiary structure. In contrast, CD8+ T cells recognize short linear peptide fragments, typically 8 to around 13 amino acids in length, which are presented in an extended conformation. MHC molecules can thus be taken to present a short linear epitope in a sterically fixed form (Murphy, 2022). The repertoire of T-cell receptor specificity may have both a genetic basis and a basis in selection in the thymus (La Gruta, 2018).
While the above explanations on the peptides presented by MHC molecules can be regarded as established knowledge, this knowledge does not extend to mechanisms that could allow determining what kind of peptide might be presented on the cell surface of a cell. As explained in more detail below, peptides presented by MHC molecules need to be identified and validated in a sophisticated approach.
Cancer Immunotherapy
Immunotherapy of cancer represents an option of specific targeting of cancer cells while minimizing side effects. Cancer immunotherapy makes use of the existence of tumor associated antigens.
T cell-based immunotherapy targets peptide epitopes derived from tumor-associated or tumor-specific proteins, which are presented by MHC molecules. The antigens that are recognized by the tumor-specific T lymphocytes, that is, the epitopes thereof, can be molecules derived from all protein classes, such as enzymes, receptors, transcription factors, etc. which are expressed and, as compared to unaltered cells of the same origin, usually up-regulated in cells of the respective tumor.
There are two classes of MHC molecules, MHC class I and MHC class ii. MHC class I molecules are composed of an alpha heavy chain and beta-2-microglobulin, MHC class ii molecules of an alpha and a beta chain. Their three-dimensional conformation results in a binding groove, which engages in non-covalent interactions with peptides.
MHC class I molecules can be found on most nucleated cells. They present peptides that result from proteolytic cleavage of predominantly endogenous proteins, defective ribosomal products (DRIPs) and larger peptides. Also, peptides derived from endosomal compartments or exogenous sources are also frequently found on MHC class I molecules. This non-classical way of class I presentation is referred to as crosspresentation in the literature (Brossart and Bevan, 1997; Rock et al., 1990). MHC class ii molecules can be found predominantly on professional antigen-presenting cells (APCs), and primarily present peptides of exogenous or transmembrane proteins that are taken up by APCs e.g. during endocytosis and are subsequently processed.
Complexes of peptide and MHC class I are recognized by CD8-positive T cells bearing the appropriate T cell receptor (TCR), whereas complexes of peptide and MHC class ii molecules are recognized by CD4-positive helper T cells bearing the appropriate TCR. It is well known that the TCR, the peptide and the MHC are thereby present in a stoichiometric amount of 1 : 1 : 1 .
For an MHC class I peptide to be able to trigger (elicit) a cellular immune response, it must first of all be bound to an MHC molecule. The binding is dependent on the allele of the MHC molecule and specific polymorphisms of the amino acid sequence of the peptide. MHC class I binding peptides are typically 8-13 amino acid residues in length and usually contain two conserved residues ("anchors") in their sequence that interact with the corresponding binding groove of the MHC molecule. In this way each MHC allele
has a “binding motif” determining which peptides can bind specifically to the binding groove.
In the MHC class I dependent immune reaction, peptides bound to MHC class I molecules expressed by cells, e.g., tumor cells, can be recognized by T cells bearing specific TCRs which in turn initiate the immune reaction.
For peptides to be recognized by T-lymphocytes as tumor-specific or associated antigenic epitopes, and to be used in a therapy, particular prerequisites must be fulfilled. The antigenic epitope should be over-presented on tumor cells in comparison to normal healthy cells. The expression level of the source protein of the peptide does not necessarily directly correlate with the presentation level of a peptide. It may be advantageous that a peptide is mainly displayed, or presented, on tumor cells and not, or in comparably small amounts, on normal healthy cells, for example cells of normal or healthy tissues. A peptide that is over-presented on the surface of a tumor cell can be derived from any protein found in the respective tumor cell.
In case of targeting peptide-MHC by specific TCRs (e.g., soluble TCRs) and antibodies or other binding molecules as disclosed herein, the therapeutic effect of the binder does not depend on the TCR repertoire of the person receiving said binding molecule. In these cases, the presentation is the determining factor.
SUMMARY
The scope of the invention is defined by the claims.
Specifically, provided herein is a peptide comprising an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59, or a pharmaceutically acceptable salt thereof, wherein the peptide has an overall length of up to 16 amino acids. In one aspect, the peptide, or the salt thereof provided herein has the ability to bind to an MHC class I molecule, and/or said peptide, when bound to said MHC, is capable of being recognized by CD8 T cells.
In another aspect, the peptide or salt thereof provided herein includes one or more nonpeptide bonds, wherein the one or more non-peptide bonds are optionally selected from -CH2-NH, -CH2S-, -CH2CH2-, -CH=CH-, -COCH2-, -CH(OH)CH2-, -CH2SO-, an N-methyl group or an N-hydroxyl group.
In one aspect, the pharmaceutically acceptable salt provided herein is one of a chloride salt, a bromide salt, an iodide salt, a thiocyanate salt, a phosphate salt, a phosphonium salt, a nitrate salt, a sulfate salt, a chlorate salt, an acetate salt, a trifluoroacetate salt, a propionate salt, a butyrate salt, a pyridinium salt, a glycolate salt, a pyruvate salt, an
oxalate salt, a malate salt, a maleate salt, a malonate salt, a succinate salt, a fumarate salt, a tartrate salt, a citrate salt, a benzoate salt, a cinnamate salt, a mandelate salt, a salicylic acid salt, a methane sulfonate salt, an ethane sulfonate salt, a p- toluenesulfonate salt, a salicylate salt, a sodium salt, a potassium salt, a barium salt, a zinc salt, a lithium salt, a rubidium salt, a cesium salt, a magnesium salt, a manganese salt, a copper salt, an ammonium salt, a calcium salt or a trimethylamine salt, wherein the salt is preferably selected from a chloride salt, an acetate salt, a trifluoroacetate salt or a combination thereof.
In another aspect, said peptide is part of a fusion protein.
Provided herein is also a binding molecule that specifically recognizes a peptide comprising an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59, or said peptide comprising an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59 when bound to an MHC molecule. The binding molecule is typically a proteinaceous binding molecule or an aptamer. In some embodiments, the binding molecule provided herein is an antibody, or a functional fragment thereof. The antibody is generally a soluble or a membrane-bound antibody. In some embodiments, the binding molecule is a T cell receptor (TCR), or a functional fragment thereof. The TCR is generally a soluble TCR or a membrane-bound TCR. The antibody or functional fragment thereof may in some embodiments be a monoclonal antibody, a bi-specific antibody, a chimeric antibody, a human antibody and/or a humanized antibody or a fragment thereof. In some embodiments, the binding molecule is an aptamer. In some embodiments, the binding molecule may be a T cell engaging bispecific molecule, such as a bispecific molecule comprising a TCR domain and a T cell recruiting antibody domain.
Provided herein is also a nucleic acid comprising a sequence encoding the peptide or the binding molecule provided herein, being a proteinaceous binding molecule. In some embodiments, the sequence is operably linked to a heterologous promoter sequence.
Provided herein is also an expression vector comprising the nucleic acid provided herein.
Provided herein is a recombinant host cell comprising the peptide, the binding molecule and/or the nucleic acid provided herein.
Provided herein is an in vitro method for producing T lymphocytes that specifically recognize a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule. The method comprises (a) contacting in vitro T lymphocytes with antigen loaded human class I MHC molecules expressed on the surface of a suitable
antigen-presenting cell or an artificial construct mimicking an antigen-presenting cell for a period of time sufficient to activate the T cells in an antigen specific manner, wherein said antigen is the peptide provided herein or (b) introducing in vitro into T lymphocytes a nucleic acid encoding a TCR that specifically recognizes a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule.
Provided herein is also a T lymphocyte, produced by the in vitro method for producing T lymphocytes provided herein, wherein said T lymphocyte specifically recognizes a cell which presents a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59. In some embodiments, the T lymphocyte is an activated T lymphocyte.
Provided herein is a pharmaceutical composition comprising one or more active ingredients as provided herein selected from the group consisting of a peptide, a pharmaceutically acceptable salt of the peptide, the binding molecule, the nucleic acid, the expression vector, the recombinant host cell, or the T lymphocyte, and a pharmaceutically acceptable carrier or a pharmaceutically acceptable excipient. In one aspect, the pharmaceutical composition provided herein is a vaccine. In another aspect, the pharmaceutical composition provided herein further comprises an adjuvant.
Provided herein is a combination product, comprising one or more of the peptide or salt thereof, the binding molecule, the nucleic acid, the expression vector or the recombinant host cell or the T lymphocyte as provided herein. In one aspect, the combination product comprises an adjuvant.
In one aspect, the adjuvant of the pharmaceutical composition or of the combination product provided herein, comprises IFN-alpha, IFN-beta or an interleukin, wherein the interleukin preferably is IL-1 , IL-2, IL-4, IL-7, IL-9, IL-10, IL-12, IL-13, IL-15, IL-17, IL-18, IL-21 , IL-23 or a combination thereof. In another aspect, the adjuvant of the pharmaceutical composition or of the combination product provided herein, is IL-2, IL-12, IL-15, IL-18, IL-21 or a combination thereof.
In another aspect, the pharmaceutical composition or the combination product provided herein comprises the T lymphocyte provided herein and an adjuvant selected from an interleukin. In some embodiments, the interleukin is IL-2, IL-15 or IL-21 or a combination thereof.
Provided herein is a method for producing the peptide or the binding molecule provided herein. The binding molecule is a proteinaceous binding molecule. The method includes culturing the recombinant host cell provided herein and isolating the peptide or the proteinaceous binding molecule from the host cell and/or its culture medium.
Provided herein is the peptide or the salt thereof, the binding molecule, the nucleic acid, the expression vector, the recombinant host cell, the T lymphocyte, the pharmaceutical composition or the combination product provided herein for use in medicine.
Provided herein is the peptide or the salt thereof, the binding molecule, the nucleic acid, the expression vector, the recombinant host cell, the T lymphocyte, the pharmaceutical composition or the combination product for use in medicine as provided herein. The use is in the prevention, treatment and/or diagnosis of cancer. In one aspect, the cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, nonHodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer and another cancer that shows an over-presentation of a peptide provided herein.
Provided herein is the use of the peptide or the salt thereof, the binding molecule, the nucleic acid, the expression vector, the recombinant host cell, the T lymphocyte, the pharmaceutical composition or the combination product as disclosed herein in the manufacture of a medicament for the prevention, treatment and/or diagnosis of cancer. In one aspect said cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer, and another cancer that shows an over-presentation of a peptide provided herein.
Provided herein is a method of killing target cells in a subject. The target cells present a peptide provided herein. The method comprises administering to the subject an effective number of T lymphocytes provided herein.
Provided herein is an in vitro method for killing target cells that present a peptide provided herein. The target cells are known or suspected of being cancer cells. The method includes contacting said cells with a plurality of T lymphocytes as provided herein. In some embodiments, the cells are cells of a sample from a subject.
In one aspect, the T lymphocyte provided herein is for use in the killing of target cells in a subject. The target cells present a peptide disclosed herein.
In one aspect, the T lymphocyte provided herein is for use in the manufacture of a medicament for the killing of such target cells that present a peptide disclosed herein.
In one aspect, the T lymphocyte is for use in the prevention or treatment of cancer. The use encompasses the killing of target cancer cells in a subject. The target cancer cells present a peptide disclosed herein.
Provided herein is a method of treating a subject (i) being diagnosed for, (ii) suffering from and/or (iii) being at risk of developing cancer. The method comprises administering to the subject an effective amount of the peptide, the binding molecule, the nucleic acid, the expression vector, the recombinant host cell, the T lymphocyte, the pharmaceutical composition or the combination product provided herein. In one aspect, said cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, nonHodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer or another cancer that shows an over-presentation of a peptide provided herein.
Provided herein is a kit. The kit includes (a) a container comprising the peptide or the salt thereof, the binding molecule, the nucleic acid, the expression vector, the recombinant host cell, the T lymphocyte, the pharmaceutical composition, and/or a combination product provided herein in solution or in lyophilized form. In some embodiments, the kit includes (b) a further container containing a diluent or reconstituting solution for the lyophilized formulation. In some embodiments, the kit includes (c) a further container containing an adjuvant. In some embodiments the kit includes both the container containing a diluent or reconstituting solution and a further container containing an adjuvant. In some embodiments, the kit includes (d) an additional peptide that contains a sequence set forth in any one of SEQ ID NOs: 1 to 59 provided herein. In some embodiments the kit includes a container containing a diluent or reconstituting solution and an additional peptide that contains a sequence set forth in any one of SEQ ID NOs: 1 to 59. In some embodiments, the kit includes the container containing a diluent or reconstituting solution, a further container containing an adjuvant and an additional peptide that contains a sequence set forth in any one of SEQ ID NOs: 1 to 59. In one aspect the kit provided herein further comprises one or more of a buffer, a diluent, a filter, a needle, and/or a syringe.
Provided herein is an in vitro method of diagnosing cancer in a sample comprising cells of an individual. The method comprises (a) adding a binding molecule to a sample from
the subject. The binding molecule is labeled. In some embodiments, the label is a radionucleotide, and (b) detecting whether the binding molecule binds to cells in the sample from the subject. The binding molecule specifically binds to (i) a peptide comprising the amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 or (ii) said peptide comprising the amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule. In some embodiments, the sample comprises cells of a tissue suspected to be a cancer tissue.
Provided herein is an in vitro method of diagnosing cancer in a sample comprising cells of an individual. The method comprises (a) identifying one or more peptides presented on cells comprised in a tumor sample from the individual; and (b) comparing the peptide(s) as identified in step (a) with a collection of peptides that have been pre-screened for immunogenicity and/or over-presentation in tumours as compared to normal tissues. The collection of peptides comprises one or more peptides comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, having a maximal length of 16 amino acids. The method also comprises (c) assessing whether a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 is a peptide comprised in a tumor sample from the individual. In some embodiments, the sample comprises cells of a tissue suspected to be a cancer tissue.
In one aspect of a method of diagnosing as provided herein, the cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer, or another cancer that shows an over-presentation of a peptide provided herein.
Provided herein is a method for producing an individualized anti-cancer cellular therapeutic composition for a subject. The method comprises determining whether a peptide comprising the amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 is presented on cells in a sample from a subject suspected of having cancer. In some embodiments, the sample comprises cancer cells from said subject. The method also comprises contacting in vitro a plurality of T lymphocytes from said subject with an antigen-presenting cell or an artificial antigen-presenting cell having antigen-loaded human class I MHC molecules on its surface. The antigen is a peptide comprising an amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59. Thereby the method includes activating said T lymphocytes in an antigen specific manner if they bind to said peptide.
In some embodiments, the method includes allowing proliferation of the obtained activated T lymphocytes. The method also includes isolating one or more activated T lymphocytees for manufacturing an individualized anti-cancer cellular therapeutic composition.
In one aspect of isolating one or more activated T lymphocytes, manufacturing the individualized anti-cancer cellular therapeutic composition further comprises cloning the nucleic acid encoding the TCR expressed by one or more activated T lymphocytes into a suitable expression vector and introducing the vector into T lymphocytes. In a further aspect of the method, determining whether said peptide is presented on cells in a sample from the subject comprises adding a binding molecule to a sample from said subject. This binding molecule is (a) labeled and (b) specifically binds to said peptide or to said peptide when bound to an MHC molecule.
Provided herein is also a method of stratifying a subject for individualized vaccination and/or immunotherapy against cancer. The method includes (i) assessing whether cells of a cancer sample from the subject present a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59. The method also includes (ii) selecting the subject for treatment with a vaccine and/or an immunotherapeutic agent if the peptide that includes the sequence set forth in any one of SEQ ID NOs: 1 to 59 is presented on cells of the cancer sample. Said vaccine includes a peptide that includes a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a pharmaceutically acceptable salt thereof. The immunotherapeutic agent includes at least one of the binding molecule, the nucleic acid, the expression vector, the recombinant host cell or the T lymphocyte provided herein. In one aspect, the method of stratifying a subject further includes assessing whether the peptide that includes a sequence set forth in any one of SEQ ID NOs: 1 to 59 is bound by an MHC molecule.
Provided herein is a method for producing an individualized anti-cancer vaccine or an individualized compound-based and/or cellular therapeutic agent for an individual. The anti-cancer vaccine includes a peptide that includes a sequence set forth in any one of SEQ ID NOs: 1 to 59. The compound-based and/or cellular therapeutic agent is based on a peptide that comprises a sequence set forth in any one of SEQ ID NOs: 1 to 59. The method includes (a) identifying one or more peptides presented on cells in a tumor sample from the individual; (b) comparing the peptide(s) as identified in (a) with a collection of peptides that have been pre-screened for immunogenicity and/or overpresentation in tumours as compared to normal tissues. The collection of peptides includes one or more peptides that include a sequence set forth in any one of SEQ ID NOs: 1 to 59, having a maximal length of 16 amino acids. The method also includes (c) selecting one or more peptides from the collection of peptides if it/they match(es) the peptide(s) identified in the subject. The selected peptide(s) is/are one or more peptide(s)
comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, having a maximal length of 16 amino acids comprised in the collection of peptides. The method also includes (d) manufacturing and/or formulating the individualized vaccine or compoundbased and/or cellular therapeutic agent based on the selection in step (c).
In one aspect of a method defined above, identifying said peptide(s) involves (a1 ) comparing expression data from the tumor sample to expression data from a sample of normal tissue corresponding to the tissue type of the tumor sample to identify proteins that are over-expressed or aberrantly expressed in the tumour sample. Identifying said peptide(s) also involves (a2) correlating the expression data with sequences of MHC ligands bound to MHC class I molecules in the tumor sample to identify MHC ligands derived from proteins over-expressed or aberrantly expressed by the tumor.
In another aspect of such a method, the sequences of MHC ligands are identified by eluting bound peptides from MHC molecules isolated from the tumor sample and sequencing the eluted ligands; and/or the normal tissue corresponding to the tissue type of the tumor sample is obtained from the same individual; and/or the immunogenicity of the peptides included in the collection of peptides is determined by a method comprising an in vitro immunogenicity assay, subject immunomonitoring for individual HLA binding, MHC multimer staining, an ELISPOT assay and/or intracellular cytokine staining.
In yet another aspect, a method as defined above further comprises identifying a mutation that is unique to the tumor sample relative to normal corresponding tissue from the individual, and selecting a peptide that correlates with the mutation for inclusion in the vaccine or for the generation of a cellular therapeutic, wherein the mutation is optionally identified by whole genome sequencing.
In another aspect of a method defined above, the one or more peptides included in the collection of peptides is/are identified based on the following steps (aa) performing genome-wide messenger ribonucleic acid (mRNA) expression analysis by highly parallel methods, such as microarrays or sequencing-based expression profiling, comprising identifying genes that are over-expressed in a malignant tissue, compared to a normal tissue or tissues; (ab) selecting peptides encoded by selectively expressed or overexpressed genes as detected in step (aa), and (ac) determining an induction of an in vivo T-cell response by the peptides as selected comprising in vitro immunogenicity assays using human T cells from healthy donors or the individual; or
(ba) identifying MHC ligands, e.g. HLA ligands, from the tumor sample using mass spectrometry; (bb) performing genome-wide messenger ribonucleic acid (mRNA) expression analysis by highly parallel methods, such as microarrays or sequencingbased expression profiling, comprising identifying genes that are over-expressed in a
malignant tissue, compared with a normal tissue or tissues; (be) comparing the identified MHC ligands to the gene expression data; (bd) selecting peptides encoded by selectively expressed or over-expressed genes as detected in step be; (be) re-detecting selected peptides from step bd on tumor tissue and detecting lack of or infrequent detection on healthy tissue and confirming the relevance of over-expression at the mRNA level; and (bf) determining an induction of an in vivo T-cell response by the peptides as selected comprising an in vitro immunogenicity assay using human T cells from a healthy donor or the individual.
Peptides
In a first aspect, there is provided a peptide consisting of an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59, or a pharmaceutically acceptable salt thereof. In some embodiments the peptide has an overall length of up to 13 amino acids, including of up to 9 amino acids. In a further aspect, there is provided a peptide comprising, an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59, or a pharmaceutically acceptable salt thereof. In that aspect the peptide has an overall length of up to 30 amino acids, including of up to 16 amino acids. The peptide defines an epitope and its sequence can be identified in the PRAME protein amino acid sequence.
Also provided herein is a nucleic acid vaccine that comprises a region encoding a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59. The nucleic acid vaccine may comprise a concatemer that contains multiple copies of a nucleic acid sequence encoding a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
As described herein, a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59 may be used as an epitope in a protein vaccine. Accordingly, provided herein is furthermore a protein vaccine comprising a section consisting of a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59. Said protein vaccine may comprise several repetitions of a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
The peptide as provided herein has the ability to bind to an MHC molecule class I and/or MHC molecule class II, such as human leukocyte antigens (HLA). In particular, an elongated version of the peptide has the ability to bind to an MHC molecule class II. A respective peptide, when bound to the MHC, is capable of being recognized by CD4 and/or CD8 T cells. In some embodiments, a respective MHC class I molecule is an HLA allotype MHC molecule specified further below.
In some embodiments, the peptide consists of an amino acid sequence according to SEQ ID NO: 1 to SEQ ID NO: 59.
The following table (Table 1 ) shows the peptides with their amino acid sequences disclosed herein and their respective SEQ ID NOs.
As explained below, another aspect of the disclosure relates to the use of the peptides disclosed herein, for example, for the combined treatment of a proliferative disease.
Generally, a peptide provided herein has the ability to bind to components of the major histocompatibility complex (MHC), in the case of humans also known as human leukocyte antigens (HLAs). In some embodiments, the MHC is an MHC class I MHC molecule.
Recombinant or Synthetic Peptide
In some embodiments a peptide as disclosed herein is a synthetic peptide. In some embodiments a peptide as disclosed herein is provided as a chemically synthesized peptide. The peptide may for example have been synthesized by means of solid phase synthesis.
In some embodiments a peptide as disclosed herein is provided as a recombinant peptide. The peptide may for example have been secreted by a recombinant host cell expressing the peptide. In such embodiments the peptide may have been isolated from the media encompassing the cells. In such embodiments the cells can be sedimented, and the supernatant be collected for isolating the peptide. The peptide may in some embodiments have been expressed in a recombinant host cell without being secreted. In such embodiments the peptide may have been isolated from the cells.
Salt of the Peptide
Provided is also a salt of a peptide as disclosed herein. A peptide may for example be provided as a salt that has an inorganic counter ion. As an example, a peptide may be provided as a halogen salt such as a chloride.
A peptide as disclosed herein may in some embodiments be provided as a salt that has an organic compound, such as a carboxylic acid containing compound, as a counter ion. As an example, a peptide may be provided as an acetate salt or a trifluoroacetate salt.
In some embodiments a respective peptide is provided as a combination of two or more of a chloride salt, an acetate salt and a trifluoroacetate salt.
Peptides with Modifications or Non-Peptide Bonds
In some embodiments a peptide as disclosed herein includes one or more non-peptide bonds. In some embodiments a peptide as disclosed herein includes an isopeptide bond.
In some embodiments a peptide as disclosed herein is modified, such as chemically modified. The peptide may for example contain one or more modified side chains. It may for example include one or more modified functional groups. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by
acylation or amidation. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by the formation of an ether or an ester. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by the formation of a thioether or a thioester. In some embodiments a peptide as disclosed herein contains one or more lysine side chains modified by pyridoxylation. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by reductive alkylation or by carboxymethylation. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by trinitrobenzylation of amino groups or amide modification of carboxyl groups. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by oxidation of cysteine to cysteic acid or it contains mixed disulphides with other thiol compounds. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified in the form of an adduct of arginine with phenylglyoxal, 2,3-butanedione or in the form of a mercurial derivative. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by reaction with maleimide or by carbamoylation. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by an intra-molecular crosslink between lysine and a glutamic acid. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified in the form of carbethoxylated histidine obtained with diethyl pyrocarbonate. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by 1 ,2-cyclohexanedione or by methionine thioether formation using iodoacetamide. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by bromoethylamine or by methionine oxidation with chloramine T. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by reaction with tyrosine tetranitromethane or N- acetyl imidazole. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by reaction with N-bromosuccinimide or modified with PEG. In some embodiments a peptide as disclosed herein contains one or more amino acid side chains modified by reaction with 2-hydroxy-5-nitrobenzyl bromide or 3- bromo-3-methyl-2-(2-nitrophenylmercapto)-3H-indole (BPNS-skatole).
Fusion Protein
In some embodiments a peptide as disclosed herein is part of a fusion protein, for example fused to the N-terminal amino acids of the HLA-DR antigen-associated invariant chain (li) or fused to (or into the sequence of) an antibody, such as, for example, an antibody that is specific for dendritic cells.
Binding Molecules
In a further aspect there is provided a binding molecule that specifically binds to a peptide as provided herein, or that specifically binds to the respective peptide when in a complex with an MHC molecule. The binding molecule may be a proteinaceous binding molecule. The binding molecule may be an antibody or a functional fragment thereof. The binding molecule may also be a T cell receptor (TCR) or a functional fragment thereof. The binding molecule may also be a proteinaceous binding molecule with antibody-like functions.
In some embodiments, a binding molecule as provided herein contains a detectable moiety such as a detectable marker. A respective detectable moiety may in some embodiments be or include a fluorescent or luminescent portion. A respective detectable moiety may in some embodiments be or include a radiolabel or an enzyme capable of producing a detectable product. A respective detectable moiety may in some embodiments be or include a magnetic resonance imaging or a computer tomography X- ray contrast agent.
In some embodiments, a binding molecule provided herein, e.g. an antibody a TCR, or an aptamer, carries a further effector function such as an immune stimulating domain or toxin. In some embodiments, a binding molecule provided herein is coupled to a detectable label. In some embodiments, a binding molecule such as a proteinaceous binding molecule, provided herein is coupled to an active agent such as a toxin. In some embodiments, a binding molecule provided herein is coupled to a protein kinase modifying moiety.
In a further aspect, there is provided an immunotoxin. The immunotoxin contains a binding molecule provided herein, e.g. an antibody, a TCR or an aptamer, which is coupled to a toxin. As an illustrative example, such an immunotoxin may be defined by a monoclonal antibody, or a functional monoclonal antibody fragment, that specifically binds to a peptide disclosed herein, covalently bonded to a toxin. The respective toxin may for instance be a truncated bacterial toxin.
Antibody or Functional Fragment Thereof
A respective antibody or functional fragment thereof may be a soluble antibody or binding fragment thereof. A respective antibody or functional fragment thereof may be a membrane-bound antibody or binding fragment thereof. In some embodiments the antibody or functional fragment thereof may be a monoclonal antibody or binding fragment thereof. In some embodiments the antibody or functional fragment thereof may be a human or a humanized antibody or binding fragment thereof. In some embodiments the antibody or functional fragment thereof may be a bi-, tri- or multi-specific antibody or
binding fragment thereof. In some embodiments the antibody or functional fragment thereof may be a chimeric antibody or binding fragment thereof.
A respective antibody or fragment thereof may in some embodiments carry a further effector function. The antibody or fragment thereof may for example be fused or covalently linked to an immune stimulating domain such as a cytokine (e.g. an inflammatory cytokine, as illustrated in Table 3), or toxin.
A respective antibody may be a monoclonal antibody and/or a bi-specific antibody or a functional fragment thereof. A respective antibody may be a chimeric antibody or a functional fragment thereof. A respective antibody may also be TCR like antibody. A respective antibody or functional fragment thereof may in some embodiments carry a further effector function. The antibody or functional fragment thereof may for example be fused or covalently linked to an immune stimulating domain such as a cytokine (e.g. an inflammatory cytokine, as illustrated in Table 3), or toxin. The antibody or functional fragment thereof may also be fused or covalently linked to an antibody recognizing a T cell structure, such as an anti CD3 antibody.
TCR or Functional Fragment Thereof
A respective T cell receptor or functional fragment thereof may be soluble or membranebound. In some embodiments a respective T cell receptor or functional T cell receptor functional fragment may be a monoclonal T cell receptor or T cell receptor functional fragment. A respective T cell receptor or functional fragment thereof may in some embodiments carry a further effector function. The T cell receptor or functional fragment thereof may for example be fused or covalently linked to an immune stimulating domain such as a cytokine (e.g. an inflammatory cytokine, as illustrated in Table 3), or toxin. The T cell receptor or functional fragment thereof may also be fused or covalently linked to an antibody recognizing a T cell structure, such as an anti CD3 antibody.
Such a functional TCR fragment provided herein is an antigen-binding TCR fragment, capable of binding to a peptide as disclosed herein. In some embodiments a functional TCR fragment is an extracellular TCR fragment. In some embodiments a functional TCR fragment is a single chain TCR (scTCR). A single chain TCR may include an alpha chain and a beta chain, linked by a linker sequence. In some embodiments a TCR provided herein is a dimeric TCR (dTCR).
In some embodiments a TCR provided herein is membrane bound. In some embodiments a TCR provided herein is a soluble TCR. A soluble TCR generally has no transmembrane or cytoplasmic domains. In some embodiments a soluble heterodimeric
TCR includes an introduced disulfide bond between residues of the respective constant domains, as described, for example, in WO 03/020763.
The TCR provided herein may be a T cell engaging bispecific molecule, such as a bispecific molecule comprising a TCR domain and a T cell recruiting antibody domain. The TCR domain may comprise the alpha and beta variable regions of a TCR capable of binding to a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59. The T cell recruiting antibody may bind to CD3 and/or to the alpha/beta TCR/CD3 complex.
Methods of producing T lymphocytes
Provided is also a method for producing T lymphocytes. The method may be an in vitro method. The method may involve introducing in vitro a nucleic acid encoding a TCR that specifically recognizes a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule into T lymphocytes.
The method may additionally or alternatively involve contacting in vitro a T lymphocyte, e.g. a CTL or Th cell, with antigen-loaded human class I or class II MHC molecules expressed on the surface of a suitable antigen-presenting cell for a period of time sufficient to activate the T lymphocyte in an antigen specific manner. The respective antigen is the peptide defined above. The method may furthermore include providing in vitro a T lymphocyte and providing antigen-loaded human class I MHC molecules that are expressed on the surface of a suitable antigen-presenting cell. The method may furthermore include isolating the activated T lymphocytes.
In some embodiments of the method for producing an activated T lymphocyte, the antigen is loaded onto class I or II MHC molecules expressed on the surface of a suitable antigen-presenting cell or artificial antigen-presenting cell by contacting a sufficient amount of the antigen with an antigen-presenting cell.
In some embodiments of the method for producing an activated T lymphocyte, the antigen-presenting cell contains an expression vector capable of expressing a peptide as disclosed herein.
Provided is furthermore a T lymphocyte that specifically recognizes a cell which presents a peptide as disclosed herein. The respective presentation may be an over-presentation. In some embodiments the over-presentation may be based on aberrant expression. In some embodiments the over-presentation is not based on an aberrant expression. In some embodiments, the T lymphocyte is an activated T lymphocyte that specifically
recognizes the peptide on the cell which presents the same, when the peptide is bound by MHC. The (activated) T lymphocyte may be a cytotoxic T lymphocyte.
Provided is also a T lymphocyte, produced by a method as disclosed herein. The T lymphocytes selectively recognize a cell which presents a peptide that contains an amino acid sequence as defined herein. The T lymphocytes may selectively recognize a cell which presents a peptide as provided herein. The respective presentation may be an over-presentation. In some embodiments the over-presentation may be based on aberrant expression. In some embodiments the over-presentation is not based on aberrant expression. In some embodiments, the T lymphocyte is an activated T lymphocyte that specifically recognizes the peptide on the cell which presents the same, when the peptide is bound by MHC. The (activated) T lymphocytes may be an alpha beta T lymphocyte or a gamma delta T lymphocyte. The (activated) T lymphocyte may be a CD8+ T lymphocyte or CD4+ T lymphocyte. The (activated) T lymphocyte may be a cytotoxic T lymphocyte.
Provided is also a T lymphocyte that includes a T-cell receptor that specifically binds to a peptide as disclosed herein, when presented by an MHC molecule. Generally, the T- cell receptor specifically binds to a peptide as disclosed herein, when presented by an MHC molecule on the surface of a cell. In some embodiments, the T lymphocyte is an activated T lymphocyte that specifically recognizes the peptide on the cell which presents the same, when the peptide is bound by MHC. The (activated) T lymphocyte may be an alpha beta T lymphocyte or a gamma delta T lymphocyte. The (activated) T lymphocyte may be a CD8+ T lymphocyte or CD4+ T lymphocyte. The (activated) T lymphocyte may be a cytotoxic T lymphocyte (CTL).
Provided is also a plurality of T lymphocytes stimulated in vitro with a peptide disclosed herein. In some embodiments, the T lymphocytes are activated T lymphocytes that specifically recognizes the peptide on the cell which presents the same, when the peptide is bound by MHC. The (activated) T lymphocytes may be alpha beta T lymphocytes or gamma delta T lymphocytes. The (activated) T lymphocytes may be CD8+ T lymphocytes or CD4+ T lymphocytes. In some embodiments the plurality of (activated) T lymphocytes define a population of T lymphocytes. In some embodiments the plurality of (activated) T lymphocytes is included in or defines a cell line, such as a human cell line.
There is furthermore provided an in vitro method of screening a TCR that specifically binds to a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59. The method includes contacting in vitro a pool of T cells (e.g. derived from the blood of a human donor), with antigen-presenting cells that express on the surface MHC molecules that are loaded with a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59. The
method further involves selecting activated T cells. The method may furthermore include isolating the activated T cells. The method may further include isolating the TCR from the activated T cells.
Also provided herein is a further in vitro method of screening a TCR that specifically binds to a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59. The method includes labelling at least two different antigens with different markers (e.g. fluorescence markers):
- The first antigen is a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
- The second antigen is (i) a peptide that is similar to the first antigen, or (ii) several different peptides that are similar to the first antigen.
T cells that bind to the first antigen but not or less to the second antigen are selected as T cells that specifically bind to a peptide according to any one of SEQ ID NO: 1 to SEQ ID NO: 59.
For example, 2D-color tetramer combinations may be used to stain T cells that bind to the first antigen, T cells that bind to the second antigen and T cells that bind to the first and the second antigen. Such a method is described, e.g. in WO 2020/245326, the content of which is incorporated by reference in its entirety. Cell sorting may be used to exclusively obtain those T cells that only bind the first antigen (but not or less the second antigen). The method may further comprise isolating the TCR from said T cells that only bind to the first antigen (but not or less the second antigen).
T Lymphocyte (ACT)
Furthermore, there is provided a T lymphocyte expressing a TCR that is specifically binding to a peptide as disclosed herein, typically when in a complex with an MHC molecule. In some embodiments the T lymphocyte is a T cell line or a cell of a T cell line. In some embodiments the T lymphocyte is part of a T cell population. The T lymphocyte is in some embodiments an alpha beta T lymphocyte. The T lymphocyte is in some embodiments a CD8+ T lymphocyte. The T lymphocyte is in some embodiments a CD4+ T lymphocyte. The T lymphocyte is in some embodiments a gamma delta T lymphocyte. The T lymphocyte is in some embodiments genetically modified to express a TCR that specifically recognizes a peptide comprising an amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59, typically when in a complex with an MHC molecule.
Provided is furthermore a plurality of T lymphocytes expressing a T-cell receptor (TCR) that specifically binds to a peptide as disclosed herein when in a complex with an MHC molecule. The T lymphocytes are in some embodiments alpha beta T lymphocytes. The
T lymphocytes are in some embodiments CD8+ T lymphocytes or CD4+ T lymphocytes. The T lymphocytes are in some embodiments gamma delta T lymphocytes. In some embodiments the plurality of T lymphocytes defines a population of T lymphocytes. In some embodiments the plurality of T lymphocytes is included in or defines a cell line, such as a human cell line.
Provided is also a plurality of T lymphocytes genetically modified with a nucleic acid. The nucleic acid encodes a T-cell receptor reactive with an MHC ligand. The cells of the plurality of T lymphocytes express a T-cell receptor that specifically binds to the MHC ligand. The MHC ligand is a peptide as disclosed herein. In some embodiments the plurality of T lymphocytes is transduced by a lentivirus or retrovirus that includes the nucleic acid encoding the T-cell receptor. The T lymphocytes are in some embodiments alpha beta T lymphocytes. The T lymphocytes are in some embodiments CD8+ T lymphocytes or CD4+ T lymphocytes. The T lymphocytes are in some embodiments gamma delta T lymphocytes. In some embodiments the plurality of T lymphocytes defines a population of T lymphocytes. In some embodiments the plurality of T lymphocytes is included in or defines a cell line, such as a human cell line.
Provided is also an activated alpha beta T lymphocyte. Provided is also an activated gamma delta T lymphocyte. Provided is also an activated CD8+ T lymphocyte. Provided is also an activated CD4+ T lymphocyte. Provided is also an activated gamma delta T lymphocyte. Provided is also a cytotoxic T lymphocyte (CTL) or activated T helper lymphocyte (Th cells). This T lymphocyte is stimulated with the peptide disclosed herein. In one embodiment the activated CTL is a cell line or a cell of a cell line. In one embodiment the activated Th cell is a cell line.
Provided is furthermore a method for providing a T cell population. The method is typically an in vitro method. The respective T cell population expresses a T-cell receptor that specifically binds to an MHC ligand. The MHC ligand is a peptide as disclosed herein. The method includes cloning a nucleic acid that encodes a TCR-alpha and/or TCR-beta chain specifically binding to an MHC ligand into a gamma retrovirus or lentivirus expression vector. As noted above, the ligand is a peptide as disclosed herein. The method furthermore includes generating a recombinant virus with antigen specificity and functional avidity for the MHC ligand. The method also includes transducing a target T- cell population with the recombinant virus that has been generated. The method furthermore includes expanding the T lymphocyte population transduced with the recombinant virus.
Provided is further a method of killing target cells in a subject whose target cells overpresent and/or aberrantly express a peptide comprising any amino acid sequence
defined herein. The method includes administering an effective number of T lymphocytes, in particular activated T lymphocytes, to the subject. The activated T lymphocytes are in some embodiments produced as described herein.
In some embodiments, the respective target cells in the subject are cancer cells. In some embodiments, the T lymphocytes are autologous to the subject. In one embodiment, the T lymphocytes have been obtained from a donor other than the subject. In some embodiments, the T lymphocytes have been obtained from a healthy donor. In some embodiments, the T lymphocytes are allogeneic to the subject. In some embodiments, the T lymphocytes have been derived from autologous tumor infiltrating lymphocytes or from peripheral blood mononuclear cells. In some embodiments, the T lymphocytes are or have been expanded in vitro.
Provided is also an in vitro method for killing target cancer cells. The target cancer cells present or over-present a peptide comprising any amino acid sequence defined herein. The method includes contacting T lymphocytes, in particular activated T lymphocytes, with the target cancer cells. The activated T lymphocytes are in some embodiments produced as described herein.
In some embodiments, the respective target cancer cells have been obtained from a subject. In some embodiments, the T lymphocytes have been obtained from the same subject. In one embodiment, the T lymphocytes have been obtained from a donor other than the subject. In some embodiments, the T lymphocytes have been obtained from a healthy donor. In some embodiments, the T lymphocytes are allogeneic to the subject. In some embodiments, the T lymphocytes have been derived from autologous tumor infiltrating lymphocytes or from peripheral blood mononuclear cells. In some embodiments, the T lymphocytes are or have been expanded in vitro.
Provided is further a method or a use of treating, including preventing, cancer. The method includes allowing target cancer cells in a subject to be killed. The target cancer cells over-present and/or aberrantly express a peptide comprising any amino acid sequence defined herein. The method or use may include administering an effective number of T lymphocytes, in particular activated T lymphocytes, to the subject, produced as described herein.
Provided is furthermore an autologous or allogeneic human alpha beta T lymphocyte or gamma delta T lymphocyte, recombinantly transfected with a T-cell receptor as defined herein. The alpha beta T lymphocyte can be a CD8+ T lymphocyte or a CD4+ T lymphocyte, and can specifically be a cytotoxic T lymphocyte (CTL) or T helper cell (Th cell).
Provided is also a TCR that is identified and isolated from an activated T lymphocyte as disclosed herein.
Provided is furthermore a TCR, or a functional fragment thereof, that is prepared based on a peptide as disclosed herein, a nucleic acid encoding the peptide, an expression vector that contains such nucleic acid or a host cell as described herein.
Production of a Medicament, First and Second Medical Uses
In a further aspect there is provided a medical use of a peptide as provided herein. In this regard there is provided a peptide as described herein for use in the treatment of a disease and in medicine. Typically, the disease is cancer.
In a further aspect there is provided a medical use of a nucleic acid or an expression vector as provided herein. In this regard there is provided a nucleic acid or an expression vector as described herein for use in the treatment of a disease and in medicine. Typically, the disease is cancer.
In a further aspect, there is provided the use of any peptide as described as a medicament or in the manufacture of a medicament. In a further aspect, there is provided the nucleic acid provided herein or the expression vector provided herein as a medicament or in the manufacture of a medicament. In a further aspect, there is provided the cell provided herein or the activated T lymphocyte as a medicament or in the manufacture of a medicament. In a further aspect, there is provided the proteinaceous binding molecule, e.g. the TCR or the antibody or other peptide and/or peptide-MHC binding molecules provided herein, as a medicament or in the manufacture of a medicament. In typical embodiments, the medicament is active against cancer. In some embodiments the medicament is a cellular therapeutic agent or a vaccine. In typical embodiments the medicament is a protein based on a soluble TCR or antibody.
Provided is furthermore a use as disclosed herein, wherein the medicament is a vaccine, such as an anti-cancer vaccine. Provided is also a use as disclosed herein, wherein the medicament is a cellular therapeutic agent.
In some embodiments the cancer is acute myeloid leukemia or chronic lymphocytic leukemia. In some embodiments the cancer is breast cancer or cholangiocellular carcinoma. In some embodiments the cancer is colorectal cancer or gallbladder cancer. In some embodiments the cancer is glioblastoma or gastric cancer. In some embodiments the cancer is gastro-esophageal junction cancer or hepatocellular carcinoma. In some embodiments the cancer is head and neck squamous cell carcinoma
or melanoma. In some embodiments the cancer is non-Hodgkin lymphoma or non-small cell lung cancer. In some embodiments the cancer is ovarian cancer or esophageal cancer. In some embodiments the cancer is pancreatic cancer or prostate cancer. In some embodiments the cancer is renal cell carcinoma or small cell lung cancer. In some embodiments the cancer is urinary bladder carcinoma or uterine endometrial cancer.
BRIEF DESCRIPTION OF THE FIGURES
Figures 1 A through 1 G show the over-presentation of various peptides in different cancer cells or tissues compared to normal cells or tissues. Upper part: Median MS signal intensities from technical replicate measurements are plotted as dots for single normal (grey dots, left part of figure) and tumor samples (black dots, right part of figure) of the respective HLA allotype on which the peptide was detected. Boxes display median, 25th and 75th percentile of normalized signal intensities, while whiskers extend to the lowest data point still within 1.5 interquartile range (IQR) of the lower quartile, and the highest data point still within 1.5 IQR of the upper quartile. Lower part: The relative peptide detection frequency in every organ is shown as spine plot. Numbers below the panel indicate number of samples on which the peptide was detected out of the total number of samples analyzed for each organ (N > 750 for HLA-A*02 positive normal samples, N > 210 for HLA-A*03 positive normal samples, N > 180 HLA-A*24 normal samples, and N > 210 HLA-B*44 normal samples) or tumor indication (N > 675 for HLA-A*02 positive cancer samples, N > 180 for HLA-A*03 positive cancer samples, N > 330 HLA-A*24 positive cancer samples and N > 210 HLA-B*44 positive cancer samples).
If the peptide has been detected on a sample but could not be quantified for technical reasons, the sample is included in this representation of detection frequency, but no dot is shown in the upper part of the figure. Tissues (from left to right): Normal samples: adipose (adipose tissue); adrenal gl (adrenal gland); bile duct; bladder; bloodcells; bloodvess (blood vessels); bone marrow; brain; breast; esoph (esophagus); eye; gall bl (gallbladder); head&neck; heart; intest, la (large intestine); intest, sm (small intestine); kidney; liver; lung; lymph nodes; nerve cent (central nerve); nerve periph (peripheral nerve); ovary; pancreas; parathyr (parathyroid gland); perit (peritoneum); pituit (pituitary); placenta; pleura; prostate; skel. mus (skeletal muscle); skin; spinal cord; spleen; stomach; testis; thymus; thyroid; trachea; ureter; uterus. Tumor samples: AML (acute myeloid leukemia); BRCA (breast cancer); CCC (cholangiocellular carcinoma); CLL (chronic lymphocytic leukemia); CRC (colorectal cancer); GBC (gallbladder cancer); GBM (glioblastoma); GC (gastric cancer); GEJC (gastro-esophageal junction cancer); HCC (hepatocellular carcinoma); HNSCC (head and neck squamous cell carcinoma); MEL (melanoma); NHL (non-Hodgkin lymphoma); NSCLCadeno (non-small cell lung cancer adenocarcinoma); NSCLCother (NSCLC samples that could not unambiguously
be assigned to NSCLCadeno or NSCLCsquam); NSCLCsquam (squamous cell nonsmall cell lung cancer); OC (ovarian cancer); OSCAR (esophageal cancer); PACA (pancreatic cancer); PRCA (prostate cancer); RCC (renal cell carcinoma); SCLC (small cell lung cancer); UBC (urinary bladder carcinoma); UEC (uterine endometrial cancer). Figure 1A) Peptide: AAFDGRHSQTLK (SEQ ID NO: 2), Figure 1 B) Peptide: GQHLHLETF (SEQ ID NO: 15), Figure 1 C) Peptide: MPMQDIKMI (SEQ ID NO: 28), Figure 1 D) Peptide: MPMQDIKMIL (SEQ ID NO: 29), Figure 1 E) Peptide: SLLQHLIGL (SEQ ID NO: 44), Figure 1 F) Peptide: SPSVSQLSVL (SEQ ID NO: 48), Figure 1 G) Peptide: YLHARLREL (SEQ ID NO: 58).
Figures 2A through 2G show exemplary expression profile of source genes of the present invention that are overexpressed in different cancer samples. Tumor (black dots) and normal (grey dots) samples are grouped according to organ of origin. Box-and-whisker plots represent median value, 25th and 75th percentile (box) plus whiskers that extend to the lowest data point still within 1 .5 interquartile range (IQR) of the lower quartile and the highest data point still within 1 .5 IQR of the upper quartile. Tissues (from left to right): Normal samples: adipose (adipose tissue); adrenal gl (adrenal gland); bile duct; bladder; bloodcells; bloodvess (blood vessels); bone marrow; brain; breast; esoph (esophagus); eye; gall bl (gallbladder); head&neck; heart; intest, la (large intestine); intest, sm (small intestine); kidney; liver; lung; lymph nodes; nerve periph (peripheral nerve); ovary; pancreas; parathyr (parathyroid gland); perit (peritoneum); pituit (pituitary); placenta; pleura; prostate; skel. mus (skeletal muscle); skin; spinal cord; spleen; stomach; testis; thymus; thyroid; trachea; ureter; uterus. Tumor samples: AML (acute myeloid leukemia); BRCA (breast cancer); CCC (cholangiocellular carcinoma); CLL (chronic lymphocytic leukemia); CRC (colorectal cancer); GBC (gallbladder cancer); GBM (glioblastoma); GC (gastric cancer); GEJC (gastro-esophageal junction cancer); HCC (hepatocellular carcinoma); HNSCC (head and neck squamous cell carcinoma); MEL (melanoma); NHL (non-Hodgkin lymphoma); NSCLCadeno (non-small cell lung cancer adenocarcinoma); NSCLCother (NSCLC samples that could not unambiguously be assigned to NSCLCadeno or NSCLCsquam); NSCLCsquam (squamous cell non-small cell lung cancer); OC (ovarian cancer); OSCAR (esophageal cancer); PACA (pancreatic cancer); PRCA (prostate cancer); RCC (renal cell carcinoma); SCLC (small cell lung cancer); UBC (urinary bladder carcinoma); UEC (uterine endometrial cancer). Figure 2A) Peptide: AAFDGRHSQTLK (SEQ ID NO: 2), Figure 2B) Peptide: GQHLHLETF (SEQ ID NO: 15), Figure 2C) Peptide: MPMQDIKMI (SEQ ID NO: 28), Figure 2D) Peptide: MPMQDIKMIL (SEQ ID NO: 29), Figure 2E) Peptide: SLLQHLIGL (SEQ ID NO: 44), Figure 2F) Peptide: SPSVSQLSVL (SEQ ID NO: 48), Figure 2G) Peptide: YLHARLREL (SEQ ID NO: 58).
Figure 3 shows the results of the IdentControl experiments for one exemplary peptide SLLQHLIGL (SEQ ID NO: 44). The peptide was confirmed by IdentControl comparing
the fragmentations of stable isotope labeled (SIL) standards in data-dependent acquisition (DDA) mode. Identity was confirmed using an in-house determined spectral correlation threshold.
Figure 4 shows one exemplary results for a CoElution experiment for the peptide YLHARLREL (SEQ ID NO: 58). The peptide was confirmed by CoElution using stable isotope labeled (SIL) internal standard and targeted MS (sPRM or IS-PRM). Non overlapping MS2 isolation windows for the SIL-peptide and the natural peptide are used. Control experiments using non-HLA peptidome sample (e.g. tryptic digest or 5% FA) as matrix are performed to confirm isotopic purity of the SIL internal standard. Peptide identity is confirmed based on objective, predefined criteria in expert manual review.
Figures 5A through 5B shows absolute copies per cell in different tumor samples. Absolute copies per cell are plotted as black dots for tumor samples of the respective HLA allotype on which the peptide was detected. Boxes display median, 25th and 75th percentile of absolute copies per cell, while whiskers extend to the lowest data point still within 1.5 interquartile range (IQR) of the lower quartile, and the highest data point still within 1.5 IQR of the upper quartile. BRCA (breast cancer); CRC (colorectal cancer); HCC (hepatocellular carcinoma); HNSCC (head and neck squamous cell carcinoma); MEL (melanoma); NSCLCadeno (non-small cell lung cancer adenocarcinoma); NSCLCother (NSCLC samples that could not unambiguously be assigned to NSCLCadeno or NSCLCsquam); NSCLCsquam (squamous cell non-small cell lung cancer); OC (ovarian cancer); OSCAR (esophageal cancer); PRCA (prostate cancer); RCC (renal cell carcinoma); SCLC (small cell lung cancer); UBC (urinary bladder carcinoma); UEC (uterine endometrial cancer). >1 - <25 copies = +, >25 copies = ++, >50 copies = +++, >75 copies = ++++. Figure 5A) Peptide: SLLQHLIGL (SEQ ID NO: 44), Figure 5B) Peptide: SPSVSQLSVL (SEQ ID NO: 48).
DETAILED DESCRIPTION
The present inventors identified peptides from the PRAME protein which are overpresented on several cancer cells. The over-presentation of these peptides can serve as the basis for a method of predicting a risk of occurrence of cancer and for a use and a method of diagnosing cancer. A cell over-presenting one or more of the respective peptides may also be a target in a method and a use in the prevention of cancer and/or the treatment of cancer.
Peptides that represent T cell epitopes can be identified based on in silico epitope prediction or directly using mass spectrometry-based immunopeptidomics. A prediction
based on algorithms does not necessarily lead to the successful identification of peptides that indeed exist in vivo. If a peptide is synthesized based on in silico epitope prediction, the synthesized peptide may turn out not to bind to MHC with high affinity and/or not to be recognized by T cells. This is due to the fact that the processes involved in the formation of peptides that are presented on the cell surface by MHC are complex and only partly understood today. Peptides identified using in silico prediction therefore need to be assessed with regard to their capability of binding to MHC and their capability of being recognized by CD4 and/or CD8 T cells.
The present inventors have therefore taken the approach of directly identifying peptides using mass spectrometry-based immunopeptidomics. All peptides disclosed in the application were isolated from patient samples in the form of a complex with HLA molecules or MHC class I, also known as a pMHC complex and denoted in the Working Examples as “HLA peptide”. A HLA-specific antibody was used to recover HLA from patient samples. The antibody is conformation-specific and only binds to folded MHC class I complexes. MHC is only folded if binding to a peptide, so that no antibody binding and no isolation can occur if there is no peptide bound to the MHC. The peptide/MHC complexes were then immobilized on affinity chromatography material via an MHC- specific antibody. Peptides were then released from the MHC complex by acid treatment, and subsequently analyzed by mass spectrometry.
PRAME (Preferentially Expressed Antigen in Melanoma), also known as Opa-interacting protein 4 (OIP4), CT130 and MAPE, is a protein and tumor antigen of the Cancer/Testis antigen group. It is a member of the PRAME family, which includes inter alia LRRC14, PRAME family members 1 , 2, 6, 25 or 33 (PRAMEF1 , PRAMEF2, PRAMEF6, PRAMEF25, PRAMEF33), and isoforms of the protein such as isoform “CRA_a” of UniProt accession number A0A024R1 E6, are known. PRAME has a length of 509 amino acids and a mass of 57,890 Da. PRAME has the NCBI accession number CAG30435, version 1 as of 2 February 2011 , and the UniProt accession number P78395, version 1 , release 2023JD3 of 28 June 2023, GenBank accession number CAG30435.1 , Gene ID 23532.
PRAME is a component of a Cul2-RING (CRL2) E3 ubiquitin-protein ligase complex, which mediates ubiquitination of target proteins, leading to their degradation. The protein is expressed at a high level in a large proportion of tumors as well as several types of leukemia. PRAME is the best characterized member of the PRAME family of leucine-rich repeat (LRR) proteins. Mammalian genomes contain multiple members of the PRAME family whereas in other vertebrate genomes only one PRAME-like LRR protein was identified. PRAME is a cancer/testis antigen that is expressed at very low levels in normal adult tissues except testis but at high levels in a variety of cancer cells.
Unless otherwise defined, all other scientific and technical terms used in the description, figures and claims have their ordinary meaning as commonly understood by one of ordinary skill in the art.
As used herein and except as noted otherwise all terms are defined as given below.
Definitions
The terms “comprising”, “including,” containing”, “having” and variations of the terms shall be read in an open-ended manner and without limitation. The word “comprise” and variations of the word, such as “comprising” and “comprises”, is not intended to exclude other elements, components, integers or steps. Where a method is for example defined as comprising one or more steps or operations, it is meant that the method involves each step as specified, but may involve other steps that are not expressly defined. Singular forms such as “a“, “an” or “the” include plural references unless the context clearly indicates otherwise. As an example, the use of “a peptide” may refer to one specific peptide or to a plurality of peptides, and terms such as “the method” are meant to reference similar steps and/or techniques known to those skilled in the art. Unless otherwise indicated, the term “at least” preceding a series of elements is to be understood to refer to every element in the series. The terms “at least one” and “at least one of” include for example, one, two, three, four, or five or more elements. It is furthermore understood that slight variations above and below a stated range can be used to achieve substantially the same results as a value within the range. Also, unless indicated otherwise, the disclosure of ranges is intended as a continuous range including every value between the minimum and maximum values.
“Consisting of”, when reference is made to a peptide, shall mean a peptide provided herein with an amino acid sequence according to any of SEQ ID NO: 1 to SEQ ID NO: 59 that has the capacity of binding or binds to MHC molecules. The term "consisting of" excludes any element, component, integer step, or ingredient not specified in the context of the term. Where a method is for example defined as consisting of one or more steps or operations, it is meant that the method does not involve any other steps than those specified.
The term “peptide” is used herein to designate a series of amino acid residues, connected one to the other typically by peptide bonds between the alpha-amino and carboxyl groups of the adjacent amino acids. The term “peptide” as used herein also includes “tumor associated peptide” or “TUMAP”.
By “non-peptide bond” is meant that the amide linkage -CO-NH- defining the peptide bond between two monomer units has been replaced by a different bond.
A “tumor-associated peptide” or “TUMAP”, also called “tumor antigenic peptide” in the art, is an antigenic peptide epitope that can be found presented by MHC on the surface of cells of a tumor. A tumor-associated peptide is recognized by a specific TCR on a T lymphocyte.
A TUMAP as naturally presented on a target cell can be derived from one or more source proteins by degradation within that cell, herein also addressed as the source protein of the TUMAP.
The term "collection of peptides” as used herein denotes a plurality of peptides or a plurality of peptide sequences. The collection of peptides may include 2, 3 or more peptides or peptide sequences. The collection of peptides may for example define a library of peptides. The collection of peptides may also be a database or included in a database. The peptides of the collection of peptides may have been pre-screened for immunogenicity and/or over-presentation in a particular tumor
The term "elongated peptide" refers to a peptide that includes a defined amino acid sequence, and that in addition to this sequence contains further amino acids at the N- and/or C-terminus of the defined sequence.
The term “polypeptide” designates a series of amino acid residues, connected one to the other typically by peptide bonds between the alpha-amino and carboxyl groups of the adjacent amino acids. In contrast to the term “peptide”, the term “polypeptide” is meant to refer to molecules containing more than about 30 amino acid residues.
A peptide, protein, nucleic acid molecule or polynucleotide coding for such a molecule is “immunogenic” (and thus is an “immunogen” within the present disclosure), if it is capable of inducing an immune response. In this disclosure, immunogenicity is more specifically defined as the ability to induce a T cell response. Thus, an “immunogen” would be a molecule that is capable of inducing an immune response, and in the case of the present disclosure, a molecule capable of inducing a T cell response. In another aspect, the immunogen can be the peptide, the complex of the peptide with MHC, and/or protein that is used to raise a specific antibody or a TCR against it.
The same peptide can be immunogenic in one person but not immunogenic in a different person, depending on the personal T cell receptor repertoires in each person which may or not contain T cell receptors able to recognize said peptide. As such, immunogenicity of a peptide plays an important role when the immune response relies on the native pool
of available T cell clones within a body, e.g. in the context of vaccination. In this case, immunogenicity of peptide can also be modulated using adjuvants which can evoke an immune response against peptide that would otherwise not be immunogenic, effectively turning a non-immunogenic peptide into an immunogenic peptide. If a therapy involves binding molecules specific for a peptide being administered into the body, immunogenicity of said peptide plays a lesser role as the immune response does not rely on the naturally available T cell receptor repertoire present in the patient.
A class I T cell “epitope” generally requires a short peptide that is bound to a class I MHC molecule (MHC class I alpha chain, beta-2-microglobulin, and peptide) that can be recognized by a T cell bearing a matching T cell receptor binding to the MHC-peptide complex with appropriate affinity. Peptides binding to MHC class I molecules are typically 8-13 amino acids in length, for example 9 amino acids in length.
The term “major histocompatibility complex”, abbreviated as MHC, refers to a group of proteins found on the surfaces of as good as any nucleated vertebrate cell. MHC proteins tightly bind peptides formed by protein degradation within the same cell, thereby presenting these epitopes to T lymphocytes. In humans, these proteins are called the “human leukocyte antigen” (HLA) proteins. As good as all MHC proteins are polymorphic, so that a subject has a particular combination of MHC alleles, known as the MHC haplotype.
As used herein, the term "binding molecule" refers to a molecule that specifically binds to an (e.g. antigenic) determinant. A "binding molecule" is generally one of a proteinaceous binding molecule and a nucleic acid molecule. In one embodiment, a binding molecule is able to direct an entity to which it is coupled or attached (e.g. a (second) antigen binding moiety) to a target site, for example to a specific type of tumor cell or tumor stroma bearing the antigenic determinant (e.g. the complex of a peptide with MHC, according to the application at hand). In another embodiment a binding molecule is able to activate signaling through its target antigen, for example a T cell receptor complex antigen. Examples of binding molecules include, but are not limited to, antibodies and fragments thereof, binding proteins comprising at least one ankyrin repeat motif and single domain antigen binding (SDAB) molecules, aptamers, and (soluble) TCRs and fragments thereof. To assess whether a molecule is a binding molecule with an affinity to a target that allows its binding thereto, binding assays can be performed.
The term “functional fragment” as used herein is to be understood as a portion of a binding molecule that retains the biological function of the binding molecule from which the functional fragment is derived. Specifically, a functional fragment of a binding molecule shall mean a portion of a binding molecule that specifically binds to the same
antigenic determinant as the binding molecule said functional fragment is derived from. Such functional fragments of binding molecules are also referred to herein as binding fragments.
The term “specific” and variant terms such as “specificity” or “specifically”, as used herein are understood to indicate that the binding molecule is directed against, binds to, recognizes or reacts with a peptide that has an indicated amino acid sequence, but at least essentially not with another peptide. The peptide may for example be a peptide that is presented on the surface of a cell, including a peptide that is presented by MHC. For example, an antibody with a binding affinity of <100 nM, or <50 nM to the peptide-MHC complex, may be regarded as “specific” in the context of the present disclosure.
In the context of binding to a peptide-MHC complex, “specific” binding means that the binding molecule, e.g. proteinaceous binding molecule, binds the peptide-MHC complex of interest better than other peptide-MHC complexes. “Specific binding” can be determined, for example, in accordance with a Western blot, an ELISA-, RIA-, ECL-, IRMA-test, FACS, IHC and a peptide scan. Generally, the antibody or fragment thereof, TCR or fragment thereof, or binding molecule can thereby form a complex with the respective peptide. Such binding may be exemplified by the specificity of a “lock-and- key-principle”. A peptide as disclosed herein may define the epitope to which a binding molecule such as a TCR specifically binds. The peptide is in some embodiments presented, and thereby fixed, by an MHC molecule.
The term “antibody” or “antibodies" is used herein in a broad sense and includes both polyclonal and monoclonal antibodies. In addition to intact or “full” antibody molecules, also included in the term ’’antibody" is an antibody fragment (e.g. a CDR, Fv, Fab and or an Fc fragment) or a polymer of such antibody molecule and a humanized version of an antibody molecule, as long as it exhibits any of the desired properties (e.g., specific binding of a peptide disclosed herein).
As used herein, the term “monoclonal antibody (mAb)” shall refer to an antibody composition having a homogenous antibody population, i.e. , a homogeneous population consisting of a whole antibody, or a fragment or derivative thereof retaining target binding capacities. The individual antibodies included in the population are identical except for possible naturally occurring mutations that may be present in minor amounts. The monoclonal antibodies herein specifically include "chimeric" antibodies in which a portion of the heavy and/or light chain is identical with or homologous to corresponding sequences in antibodies derived from a particular species or belonging to a particular antibody class or subclass, while the remainder of the chain(s) is identical with or
homologous to corresponding sequences in antibodies derived from another species or belonging to another antibody class or subclass, as well as fragments of such antibodies, so long as they exhibit the desired binding activity (e.g., specific binding of a peptide disclosed herein).
As used herein, the term “antibody fragment” shall refer to a fragment of such an antibody retaining its specific binding capacities, e.g.
■ a CDR (complementarity determining region)
■ a hypervariable region
■ a variable domain (Fv)
■ a single chain variable domain (scFv)
■ an IgG or IgM heavy chain (consisting of VH, CH1 , hinge, CH2 and CH3 regions)
■ an IgG or IgM light chain (consisting of VL and CL regions), and/or
■ a Fab and/or F(ab’)2.
Also included in the term “antibody” are diabodies, camelid antibodies, nanobodies, domain antibodies, bivalent homodimers with two chains consisting of scFvs, IgAs (two IgG structures joined by a J chain and a secretory component), shark antibodies, antibodies consisting of new world primate framework plus non-new world primate CDR, dimerized constructs comprising CH3+VL+VH, and antibody conjugates (e.g. antibody or fragments or derivatives linked to a toxin, a radioisotope or a label).
The term “T cell receptor” (abbreviated TCR) refers to a heterodimeric molecule typically comprising an alpha polypeptide chain (alpha chain) and a beta polypeptide chain (beta chain), wherein the heterodimeric receptor is capable of binding to a peptide antigen presented by an MHC molecule. The term also includes a so-called gamma/delta TCR.
As used herein, the term “TCR fragment” refers to either a portion of a native TCR or a recombinant/engineered protein that contains TCR-derived complementaritydetermining region (CDR) sequences. Generally, a TCR fragment is an antigen-binding TCR fragment, which specifically binds to a peptide when bound by MHC proteins. As binding to the respective peptide/MHC complex is defined by the CDR sequences, in particular CDR1 and CDR3, these sequences are included in an antigen-binding TCR fragment. The person skilled in the art is aware that the CDRs can be interspersed with framework regions, which may not be crucial for target antigen specificity. A variable domain comprising TCR-derived CDRs and antibody-derived framework regions may thus be considered an antigen-binding TCR fragment. As a further example, for a human ap TCR, an antigen-binding TCR fragment can contain an alpha chain variable domain (Va), or a portion thereof, and a beta chain variable domain (VP), or a portion thereof,
each comprising three CDRs. For a human y8 TCR, an antigen-binding TCR fragment can contain a gamma chain variable domain (Vy), or a portion thereof, and a delta chain variable domain (V8), or a portion thereof, each comprising three CDRs. A TCR fragment may also be a single-chain TCR fragment, which contains only the Va and Vp domains, connected by a linker.
The term “immune response,” as used herein, means a physiological process involving the activation and/or induction of an effector function of the body’s immune system in, by way of non-limiting examples, a T cell, B cell, natural killer (NK) cell, and/or an antigen- presenting cell (APC). An example of an immune response is any detectable antigenspecific activation and/or induction of a helper T cell or cytotoxic T cell activity or response, production of antibodies, antigen-presenting cell activity or infiltration, macrophage activity or infiltration, or neutrophil activity or infiltration.
The term “T cell response” means the specific proliferation and activation of effector functions induced by a peptide in vitro or in vivo. For MHC class I restricted cytotoxic T cells, effector functions may be lysis of peptide-pulsed, peptide-precursor pulsed or naturally peptide-presenting target cells, secretion of cytokines, such as Interferongamma, TNF-alpha, or IL-2 induced by peptide, secretion of effector molecules, such as granzymes or perforins induced by peptide, or degranulation.
By ’’aberrantly expressed" and “aberrantly express” it is meant that a polypeptide or protein is expressed at unusual high levels (overexpressed) in tumor tissue or tumor cells when compared to levels of expression in normal tissues or normal cells, or that the gene is silent in the tissue or cells from which the tumor is derived whereas in the tumor, it is expressed.
By ’’overexpressed" it is meant that the polypeptide, protein or gene transcript is present at a level at least 1 .2-fold of that present in normal tissue or on normal cells; such as at least 2-fold. In some embodiments the polypeptide, peptide or gene transcript is present at a level at least 5-fold or 10-fold the level present in normal tissue or on normal cells.
The term "over-presented” and variants of the term such as "overpresented” or "overpresentation”, as used herein refers to the relative count of a peptide presented on a tissue or cell population addressed, when compared to an average count, i.e. the frequency with which the same peptide is presented on a healthy tissue or cell population of the same type. Furthermore, it can refer to the relative intensity of peptide presented on a tissue or cell population addressed, when compared to an average intensity, i.e. the amount of the same peptide with which it is presented on a healthy tissue or cell
population of the same type. The term indicates that the peptide is presented by MHC proteins on the cell surface, both on the cells showing overpresentation and on the healthy reference. If a certain peptide is over-presented on cells of a certain cancer, a higher number of the peptide is bound to MHC proteins on the cell surface of the cells of this cancer than on the cell surface of corresponding tissue or cell population that is healthy.
The terms “nucleic acid” and “nucleic acid molecule” as used herein refer to any nucleic acid in any possible configuration, such as single stranded, double stranded or a combination thereof. Thus, for example for DNA, the specific sequence, unless the context indicates otherwise, refers to the single strand DNA of such sequence, the duplex of such sequence with its complement (double stranded DNA) and the complement of such sequence.
In some embodiments, the term “nucleic acid sequence”, in some instances also referred to as a “nucleotide sequence”, refers to a heteropolymer of deoxyribonucleotides.
As used herein the term “a nucleic acid coding for (or encoding) a peptide” is meant to include a reference to a nucleic acid sequence coding for the respective peptide, including artificial (man-made) start and stop codons compatible with the biological system by which the sequence is to be expressed. The nucleic acid sequence coding for a particular peptide or binding molecule may be naturally occurring or it may be synthetically constructed. Generally, a DNA segment encoding a peptide, polypeptide, and protein as provided herein, is assembled from cDNA fragments and short oligonucleotide linkers, or from a series of oligonucleotides, to provide a synthetic gene that is capable of being expressed in a recombinant transcriptional unit comprising regulatory elements derived from a microbial or viral operon. The system may, for example, be a dendritic cell or another cell system useful for the production of TCRs.
The term “coding region” refers to that portion of a gene which either naturally or normally codes for the expression product of that gene in its natural genomic environment, i.e. , the region coding in vivo for the native expression product of the gene. The coding region can be derived from a non-mutated (“normal”), mutated or altered gene, or can even be derived from a DNA sequence, or gene, wholly synthesized in the laboratory using methods well known to those of skill in the art of DNA synthesis.
The term “expression product” means the polypeptide or protein that is the natural translation product of the gene and any nucleic acid sequence coding equivalents resulting from genetic code degeneracy and thus coding for the same amino acid(s). In
the present specification, a peptide as disclosed herein may also be an expression product, when expressed from a recombinant nucleic acid construct encoding such peptide.
The term “DNA segment” refers to a DNA polymer, in the form of a separate fragment or as a component of a larger DNA construct, which has been derived from DNA isolated at least once in a substantially pure form, i.e., free of contaminating endogenous materials and in a quantity or concentration enabling identification, manipulation, and recovery of the segment and its component nucleotide sequences by standard biochemical methods, for example, by using a cloning vector. Such segments are provided in the form of an open reading frame uninterrupted by internal non-translated sequences, or introns, which are typically present in eukaryotic genes. Sequences of non-translated DNA may be present downstream from the open reading frame, where the same do not interfere with manipulation or expression of the coding regions.
The term “vector" refers to a vehicle into which a polynucleotide encoding a peptide or protein may be operably inserted. A vector contains a site that is adapted for the insertion of a polynucleotide, such as a multiple cloning site. Generally, a vector is a linear or circular nucleic acid molecule.
The term “expression vector" refers to a vector that is cable of effecting the expression of a peptide or protein that is encoded by a nucleic acid sequence included in the vector. An expression vector contains particular sequences allowing the expression of such a nucleic acid sequence, to which the nucleic acid sequence can be operably coupled. Such a particular sequence is a promoter. Further examples of such sequences include a start codon, a polyadenylation signal, a termination signal, and a ribosome binding site. An expression vector may also contain one or more control sequences. An example of a control sequence is an enhancer. Various types of expression vectors are known, for example plasmid and viral-based expression vectors.
The term “promoter” means a region of a nucleic acid, typically DNA, involved in binding of RNA polymerase to initiate transcription.
The word “recombinant” or “heterologous” is used in this specification interchangeably to describe a nucleic acid molecule that, by virtue of its origin, manipulation, or both is not associated with all or a portion of the nucleic acid molecule with which it is associated in nature. Generally, a recombinant nucleic acid molecule includes a sequence which does not naturally occur in the respective wildtype organism or cell. Typically, a recombinant nucleic acid molecule is obtained by genetic engineering, usually constructed outside of a cell. Generally, a recombinant nucleic acid molecule is substantially identical and/or
substantial complementary to at least a portion of the corresponding nucleic acid molecule occurring in nature. A recombinant nucleic acid molecule may be of any origin, such as genomic, cDNA, mammalian, bacterial, viral, semisynthetic or synthetic origin. The term "recombinant" as used herein with respect to a protein / peptide means a peptide produced by expression of a recombinant nucleic acid.
The term “isolated” means that the material is removed from its original environment (e.g., the natural environment, if it is naturally occurring). For example, a naturally- occurring nucleic acid molecule or protein present in a living animal is not isolated, but the same nucleic acid molecule or protein, separated from some or all of the coexisting materials in the natural system, is isolated. Such nucleic acid molecule could be part of a vector and/or such nucleic acid molecule or protein could be part of a composition, and still be isolated in that such vector or composition is not part of its natural environment. Similarly, a peptide as disclosed herein may be isolated from natural sources as illustrated in the Working Examples. It may also be expressed in a host cell, and subsequently isolated from this host cell or from the media that includes the host cell. It may also be isolated from both the host cell and the media that includes the host cell.
A “pharmaceutical composition” as used herein is a composition suitable for administration to a human being in a medical setting. In typical embodiments, a pharmaceutical composition is sterile and produced according to GMP guidelines. The pharmaceutical composition may for example be a solution or a solid. The pharmaceutical composition may for example be a suspension. Examples of a solid form of a composition include, but are not limited to a granule, a pellet, a micro-pellet, a sphere, a core, a coated core, a pill, a compressed tablet, a mini tablet, a layered tablet, a bead, a capsule, a powder or any form of a particle. The pharmaceutical composition may be a mixture that contains an agent as disclosed herein, e.g. a TCR or an antibody, and an adjuvant. A pharmaceutical composition may also be a mixture that includes a population of immune cells, such as human immune cells, and a protein such as albumin. The pharmaceutical composition may also be a mixture that contains a plurality of (activated) T lymphocytes as described herein. Said (activated) T lymphocytes may be alpha beta T lymphocytes or gamma delta T lymphocytes. The (activated) T lymphocytes may be CD8+ T lymphocytes or CD4+ T lymphocytes. The (activated) T lymphocytes may be cytotoxic T cells (CTLs) as disclosed herein. The pharmaceutical composition may or may not comprise one or more antigen-presenting cells and/or one or more MHC molecules.
A “combination product” refers to a product that includes a plurality, i.e. two or more, individual pharmaceutical compositions. The pharmaceutical compositions may be provided for administration either together or separately. A combination product may
include storage means for pharmaceutical preparations as individual or separate preparations. A combination product may also include means for dispensing a pharmaceutical preparation.
An “active agent” is an entity, such as a pharmaceutical compound, that can be used to achieve a desired effect, e.g., a therapeutic effect. A therapeutic effect may be an effect in the treatment, including the prevention, of cancer. Where the active agent is used to achieve a therapeutic effect, it is also referred to as a “therapeutic agent”. Generally, an active agent is intended to be used for administration to a subject. Typically, an active agent defines or contains an active ingredient, which can be used in a pharmacologically, including therapeutically, effective amount. An active agent may be included in a delivery system or in a kit. Examples of an active agent include a peptide as disclosed herein or a cell. Two further examples of an active agent are a proteinaceous binding molecule as disclosed herein or an adjuvant, such as an interleukin. It should be noted that as long as an active agent is referred to, this does not necessarily include its medical use.
An “individualized pharmaceutical” shall mean a specifically tailored therapy product for one individual subject that will only be used for therapy in such individual subject, including an actively individualized cancer vaccine and/or an adoptive cellular therapy using autologous tissue from a subject, e.g. autologous patient tissue. A “personalized pharmaceutical” shall mean a specifically tailored therapy product for one individual person that will only be used for therapy in such individual person. As long as the subject is a human, “individualized” and “personalized” can be used interchangeably.
The term "a salt" refers to an ionized form of a disclosed peptide where the peptide is provided with one or more counter ions, thereby providing an acid or base salt of the peptide. For example, an acid salt is prepared from the free base (typically wherein the neutral form of the peptide has a neutral -NH2 group) involving reaction with a suitable acid. Suitable acids for preparing acid salts include both organic acid and inorganic acid, as explained in more detail below.
The term “pharmaceutically acceptable salt” refers to any salt of a peptide provided herein which retains its biological properties and which is not toxic or otherwise undesirable for pharmaceutical use. Such a salt may be derived from a variety of organic or inorganic counter-ions well known in the art. Examples of a respective salt include, but are not limited to the chloride salt, the acetate salt or the trifluoroacetate salt, see also below.
By the term “lipid nanoparticle” is meant a particle having at least one dimension on the order of nanometers (e.g., 1 -1 ,000 nm), which includes one or more lipids.
The term "cationic lipid" as used herein refers to a lipid having a net positive charge. Cationic lipids bind a negatively charged nucleic acid molecule such as RNA by electrostatic interaction to the lipid matrix. Generally, cationic lipids possess a lipophilic moiety, such as a sterol, an acyl or diacyl chain, and the head group of the lipid typically carries the positive charge. Two illustrative examples of a cationic lipid are 1 ,2-di-O- octadecenyl-3-trimethylammonium propane (DOTMA) and dimethyldioctadecylammonium (DDAB). Two further examples of a cationic lipid are 1 ,2- dioleoyl-3-trimethylammonium propane (DOTAP) and 1 ,2-dioleoyl-3- dimethylammonium-propane (DODAP). 1 ,2-diacyloxy-3-dimethylammonium propanes and 1 ,2-dialkyloxy-3-dimethylammonium propanes are further examples of a cationic lipid. Further examples of a cationic lipid are are dioctadecyldimethyl ammonium chloride (DODAC) and 2,3-dioleoyloxy-N-[2(spermine carboxamide)ethyl]-N,N-dimethyl-l- propanamium trifluoroacetate (DOSPA). Two further illustrative examples of a cationic lipid are 2,3-di(tetradecoxy)propyl-(2-hydroxyethyl)-dimethylazanium (DMRIE) and 1 ,2- dimyristoyl-sn-glycero-3-ethylphosphocholine (DMEPC). Examples of a cationic lipid are also 1 ,2-dimyristoyl-3-trimethylammonium propane (DMTAP) and 1 ,2-dioleyloxypropyl- 3-dimethyl-hydroxyethyl ammonium bromide (DORIE).
The term "neutral lipid" refers to a lipid having a net charge of zero. Two illustrative examples of a neutral lipid are 1 ,2-di-(9Z-octadecenoyl)-sn-glycero-3- phosphoethanolamine (DOPE) and 1 ,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC). Two further examples of a neutral lipid are diacylphosphatidyl choline and diacylphosphatidyl ethanol amine. Two further examples of a neutral lipid are ceramide and sphingoemyelin. Yet two examples of a neutral lipid are cephalin and cholesterol. Another example of a neutral lipid is cerebroside.
The term “tissue” as used herein, is intended to refer to a group of cells that have a similar structure and act together to perform a specific function. A tissue may be a cluster of cells. A tissue may contain extracellular matrix.
Where the terms "normal tissue" or “healthy tissue” are used herein, reference is made to cells of healthy tissue, including healthy peripheral blood mononuclear cells (PBMCs). Illustrative examples of respective tissue include, but are not limited to, healthy blood cells, brain, heart, liver, lung, adipose tissue, adrenal gland, bile duct, bladder, bone, bone marrow, esophagus, eye, gallbladder, head and neck, large intestine, small intestine, kidney, lymph node, central nerve, peripheral nerve, pancreas, parathyroid gland, peritoneum, pituitary, pleura, skeletal muscle, skin, spinal cord, spleen, stomach, thyroid, trachea, and ureter cells or other normal tissue cells such as breast, ovary, placenta, prostate, testis, thymus and uterus. The terms "normal tissue" or “healthy tissue” are also used to address a sample from a healthy subject.
Where the term "tumor tissue" is used herein, reference is made to cells of a tumor, typically cancer, including bone marrow, blood-forming cells and blood cells. As examples, cells defining acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer are tumor tissue, but not cells of normal tissue (see Example 1). The term is also used to address a sample from a subject that has been identified as cancerous, including a sample of a hematologic neoplasm, such as chronic lymphocytic leukemia, chronic myeloid leukemia or acute myeloid leukemia.
As used herein, “body fluid” is understood as an extracellular fluid found within an organism, typically fluid that can be found in a body cavity or hollow organ. Illustrative examples of a suitable body fluid include, but are not limited to, blood or cerebrospinal fluid. Two further examples of a suitable body fluid are pleural fluid or synovial fluid. A further illustrative example of a body fluid is ascitic fluid.
By “healthy individual” it is meant that the individual is generally in good health. A healthy individual generally has a competent immune system and is typically not suffering from any disease that can be readily tested for and detected.
The term “subject” as used herein, also addressed as an individual, refers to a human or non-human animal, generally a mammal. A subject may be a mammalian species such as a rabbit, a mouse, a rat, a guinea pig, a hamster, a dog, a cat, a pig, a cow, a goat, a sheep, a horse, a monkey, an ape or a human. Thus, the methods, uses and compositions described in this document are generally applicable to both human and veterinary disease. As explained in more detail below, the sample used to determine over-presentation and/or aberrant expression of a peptide on a target cell has been obtained from the subject. A subject is typically a living organism. A subject may also be a non-living organism. Where the subject is a living human who is receiving medical care for a disease or condition, it may also be addressed as a “patient”.
The terms “stratifying” and “stratification” as used herein indicate that a subject is assigned to a group, for example with regard to the suitability of the subject for a particular therapy. As an illustrative example, individuals suffering from cancer or being at risk of suffering from cancer may be stratified according to one or more peptides being overpresented on cancer tissue.
Peptide identification and characterization
The present inventors have found that the peptides disclosed herein are presented on cells of cancer tissue. T lymphocytes bind to short continuous amino acid sequences, see above. These sequences are often buried within the native structure of the protein. In the folded source protein these short amino acid sequences are thus typically not accessible (Murphy, 2022, p. 161 ). When presented as a small peptide on the cell surface, the sequence can only be recognized by a matching T lymphocyte if bound by MHC (Murphy, 2022, p. 161 ).
Peptides presented on the cell surface can originate from any protein found within the cell (Gfeller, D, & Bassani-Sternberg, M, Front. Immunol. (2018), Sec. B Cell Biology, 9 doi: 10.3389/fimmu.2018.01716; Berg et al., 2015, p. 996). It can, however, not reliably be predicted which source protein will give rise to what sort of peptide(s).
The present inventors have furthermore found that the peptides disclosed herein are typically not presented on cells of healthy tissue or presented in very small numbers. This means, for example, that in a healthy organism a peptide as provided herein is not presented to cells of the cellular immune response. Cell-mediated immunity evolved to cope with intracellular pathogens (Berg et al., 2023, p. 89; Berg et al., 2015, p. 995), and the same mechanisms also recognize cancer cells. Peptides not presented by healthy cells or present in very small numbers, but presented on the cell surface of cancer cells are recognized by T cells as non-self antigens. The cancer cell concerned is then eliminated by the T cells (Berg et al., 2015, p. 996). Such a peptide can be regarded as immunogenic.
On the other hand, for a peptide that is predominantly presented on cells of healthy tissue, MHC restriction in T cell development of the respective organism typically leads to the peptide as being recognized as a self antigen. In this case the peptide cannot be immunogenic within the respective organism. This is due to what is known as the selftolerance of the immune system. Precursors of T cells undergo positive and negative selection in the thymus, which about 98 % of the precursors do not survive (Berg et al., 2015, p. 1004). T cells carrying TCRs specific for non-self antigen peptides are allowed to mature, whereas T lymphocytes carrying TCRs specific for self-antigens are eliminated by clonal deletion or by a process called anergy, unless the respective T lymphocyte is an ignorant cell. As a result, there is no immune response against self-antigens.
The genetic architecture of TCRs resembles that of antibodies (Berg et al., 2015, p. 998). Combinations of J gene segments, D gene segments, and V gene segments for example lead to the formation of the variable region. As a result, at least 1015 different specificities
can arise from combinations of this repertoire of genes (Lythe, G, et al., Journal of Theoretical Biology (2016) 389, 214-224. Similar to the situation for antibodies, the combination of rearrangement of gene segments and RNA splicing leads to a diversity that ensures coverage of any potential antigenic epitope. It is therefore most likely that a T cell that carries a TCR that is capable of binding a peptide that contains a sequence of one of SEQ ID NOs: 1 to 59, or consists of a sequence of one of SEQ ID NOs: 1 to 59 exists in an organism, e.g. in a blood sample. If blood from a plurality of donors is collected, it is furthermore highly likely that a T lymphocyte carrying a TCR specific for a peptide that contains a sequence of one of SEQ ID NOs: 1 to 59, or consists of a sequence of one of SEQ ID NOs: 1 to 59, can be identified.
Within an individual, tumor cells are therefore generally eliminated by T lymphocytes that bind peptides that are presented on cancer cells, but not, or in very low numbers, on healthy cells. It is nevertheless possible that the cellular immune response fails. The initially formed cancer cells may not have been completely destroyed due to an initially low number of T cell clones specific against the particular peptide presented, and subsequently tumor infiltration may for instance be limited. An inhibiting tumor microenvironment may also have formed. In such cases a tumor has developed. Inter alia a T cell-based therapy has therefore become a promising approach for an individual suffering from cancer.
A peptide as disclosed herein has been isolated in a form bound to an MHC molecule from cancer cells and identified by way of mass spectrometric analysis. For this purpose, a tumor tissue sample or a tumor cell sample has been lysed. The peptide/MHC complex has then been isolated from the lysed sample by way of immunoprecipitation.
Binding of a peptide to an MHC complex has been tested by methods known in the art. Binding of two potential binding partners can for example generally be tested by immobilizing one of the binding partners and/or by detectably labelling one of the binding partners. ELISA-based binding and competition techniques have for instance been developed for the detection of peptide/MHC binding. Immunoprecipitation, for example with a conformation dependent monoclonal antibody, or cell-based detection are additional options available in this regard.
The antibody used for immunoprecipitation was conformation-specific. The antibody captured only folded MHC class I complexes. An antibody specific for the MHC molecule to which the peptide is bound has been used. After immunoprecipitation, peptide/MHC complexes bound to an antibody were washed to remove unbound peptides. MHC-bound peptides were then released from MHC.
The discovery platform XPRESIDENT® v2.1 allows the identification and selection of relevant over-presented peptides which are potential targets for immunotherapy based on direct relative quantitation of HLA restricted peptide levels on cancer tissues in comparison to several different non-cancerous tissues and organs. See e.g. WO 2011/128448, U.S. 9791443 and U.S. 9791444, the contents of which are incorporated by reference in their entirety. This was achieved by the development of label-free differential quantitation using the acquired LC-MS data processed by a proprietary data analysis pipeline, combining algorithms for sequence identification, spectral clustering, ion counting, retention time alignment, charge state deconvolution and normalization.
In order to select over-presented peptides, a presentation profile was calculated showing the median sample presentation as well as replicate variation. The profile juxtaposes samples of the tumor entity of interest to a baseline of normal cell samples or tissue samples. Each of these profiles was then consolidated into an over-presentation score by calculating the p-value of a Linear Mixed-Effects Model (Pinheiro et al., 2015) adjusting for multiple testing by False Discovery Rate (Benjamini and Hochberg, 1995, the content of which is incorporated by reference in its entirety).
For the identification and relative quantitation of the disclosed MHC peptide ligands by mass spectrometry, MHC molecules, specifically HLA molecules, from shock-frozen cell samples or tissue samples were purified and HLA associated peptides were isolated. The isolated peptides were separated and sequences were identified by online nano- electrospray-ionization (nanoESI) liquid chromatography-mass spectrometry (LC-MS) experiments. The resulting peptide sequences were verified by comparison of the fragmentation pattern of natural peptides (also referred to as TUMAPs) recorded from acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastroesophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer samples (N >750 samples) with the fragmentation patterns of corresponding synthetic reference peptides of identical sequences. Since the peptides were directly identified as ligands of MHC molecules of primary tumors, these results provide direct evidence for the natural processing and presentation of the identified peptides on primary cancer tissue obtained from acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic
cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer patients (cf. Example 1 , Figures 1A-1 G).
Additional sequence information from public resources (Olexiouk et al., 2016; Subramanian et al., 2011 , the contents which are incorporated by reference in their entirety) were integrated into the XPRESIDENT® discovery pipeline to enable the identification of peptides from non-canonical origin. Presentation levels including error estimates for each peptide and sample were established. Peptides exclusively presented on tumor cells of tumor tissue or of a body fluid, and peptides over-presented on cancer cells versus corresponding non-cancer cells, including cells of a body fluid, or tumor versus non-cancerous tissues and organs have been identified. As the peptide, that is bound to the MHC, is itself not recognized by the antibody, it relies on the structural integrity of the MHC complex to ensure binding to the antibodies used for immunopurification. However, the MHC class molecule is not stable without a peptide. If there is no peptide bound to the MHC class I peptide binding groove, MHC class I conformation is lost. As a result, the epitope recognized by the antibody (e.g. BB7.2) is no longer available and immunoprecipitation of MHC class I is unsuccessful. Therefore, isolation of peptides is not possible if the peptide is not bound to MHC class I binding groove and only MHC-class I - bound peptides can be detected using mass spectrometry.
MHC-peptide complexes from acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer tissue samples were purified and peptides bound to MHC were released from the MHC binding cleft and analyzed by LC-MS (see Example 1 ). All peptides disclosed in the present application were identified and their binding to MHC was shown with this approach on acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, nonHodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer samples confirming their presentation on acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and
neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, uterine endometrial cancer, and combinations thereof.
Peptides identified on acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer and normal tissues were quantified using ioncounting of label-free LC-MS data. The method assumes that LC-MS signal areas of a peptide correlate with its abundance in the sample. All quantitative signals of a peptide in various LC-MS experiments were normalized based on central tendency, averaged per sample and merged into a bar plot, called presentation profile. The presentation profile consolidates different analysis methods like protein database search, spectral clustering, charge state deconvolution (decharging) and retention time alignment and normalization.
Besides over-presentation of the peptide, mRNA expression of the underlying gene was tested. mRNA data were obtained via RNASeq analyses of normal tissues and cancer tissues (see Example 2, Figures 2A-2G). An additional source of normal tissue data was a database of publicly available RNA expression data from around 3000 normal tissue samples (Lonsdale, 2013, the content of which is incorporated by reference in its entirety). In some embodiments, a peptide as disclosed herein is derived from a protein the coding mRNA of which is highly expressed in cancer tissue, but very low or absent in healthy tissues.
It should, however, be noted that there is no direct association between the expression level of a transcript or protein in a cell and the presentation of one or more peptides derived from the protein on the cell surface. Put differently, there is a very poor general correlation between the transcriptome or proteome and the MHC ligandome of a cell. Peptides presented on the surface of a cell are fragments of all proteins expressed in the respective cell. For this reason, attempts of identifying potential MHC ligand peptides solely on the basis of transcript or protein expression profile without further experimental validation are bound to fail. The method pursued by the present inventors is based on analyzing the peptides presented on the cell surface, and thus on the MHC ligandome.
In this regard, MHC Class I binding peptides are presented on virtually any nucleated cell. MHC Class I binding peptides are generated mainly following degradation of intracellular proteins by the ubiquitin-proteasome system of the respective cell. When a cancer cell is concerned, it should furthermore be noted that the MHC ligandome is influenced by cancer-specific alterations affecting, among others, translational regulation, protein metabolism, proteasomal processing, cytosolic proteolysis, but also alterations in antigen processing. For example, the expression of chaperones is often altered in cancer cells.
As mentioned above, identifying peptides presented by MHC requires their isolation from cancer tissues. Identifying peptides presented by MHC only on cancer tissue furthermore requires the analysis of both healthy tissue and cancer tissue samples. Thousands of cancer tissue and healthy tissue samples need to be collected, prepared, measured, and the data analysed in a highly reproducible manner.
Notably, when identifying peptides as presented only on the surface of cancer cells there also exist a couple of error sources that can lead to the identification of false positive results or missing/false negative identifications on healthy tissues, thereby requiring careful individual adaptions and validation of obtained results. For example, sample preparation might introduce proteolytic fragments mimicking HLA ligands, which can be assessed by in silico methods. Chromatographic procedures pose the risk of peptide carryover between samples, which can be monitored by blank runs. In addition, electrospray ionization MS generates in-source fragments that can be identified computationally. And finally, the peptide sequencing is prone to false-positive identifications that can only be uncovered by experimental sequence validation through synthetic standards.
While the techniques used for identifying peptides presented by MHC on the surface of cells may be generally known, their specific adjustments, specific combination, their individual application and analysis of the resulting data with respect to each and every individually identified over-presented peptide included in the present application by far exceeds what the person skilled in the art would consider routine experimentation. In fact, the present application shows the substantial effort the applicant and the present inventors have made to advance the knowledge in the present field.
Achieving sensible results while observing for the various pitfalls inherent to the system, as indicated above requires significant human intervention. If peptide identification and validation would be run in an automatic fashion or at least based on some kind of routine, such pitfalls would not be eliminated, and any results thus obtained would be meaningless:
(a) The number of possible MHC-ligands is virtually endless. Particularly, there are approximately 60 million theoretical class I peptides encoded by the proteome with a length between 8 and 12 amino acids. This dramatically increases the chances of falsely discovering MHC peptides, and requires additional attention to each potential peptide. When identifying peptides for cancer therapy, potential peptides cannot simply be reduced to those peptides with low false discovery rate, since this would limit the sensitivity of identifying veritable peptides. Additionally, in other applications, the top- ranking peptide-spectrum match will usually be correct when identifying a peptide. An example for such other application is the commonly used approach for MS-based proteomic analysis where trypsin is used to digest proteins into tryptic peptides, i.e., peptides with either arginine or lysine at their C-terminus. This limitation to only analyzing tryptic peptides drastically reduces the number of peptides within the analyte and hence reduces the complexity of the analysis by reducing the number of theoretically available peptide that might fit an experimentally derived peptide spectrum. However, this is not necessarily true for immunopeptidomics, which does not have the clearly defined peptide sequence features mentioned above and hence has a much larger search space than other applications, resulting in more mass ambiguities. Hence, analysing obtained data has to be performed in a more thorough manner than in other areas, requiring close human attention to each case.
(b) A peptide found may actually not be a true MHC ligand since in-source fragmentation can occur. In particular, the mass spectrometry techniques of electrospray ionization may generate ions that might be selected for acquisition of MS/MS spectra that may be mistaken for MHC ligands. Particularly, C-terminal source fragments may be falsely identified as MHC ligands. Eliminating such peptides from consideration requires close attention and experience. On the other hand, the parental peptides that the generated fragment are derived from, are not detected upon fragmentation any longer and can lead to false negative results.
(c) Due to the high sensitivity needed in immunopeptidomics, system- and application-specific sources of carryover need to be identified and clean-up protocols need to be designed, in order to avoid contamination of obtained results by previous samples, which can easily yield to false positive results. This issue is highly specific to the present research field and requires identifying and implementing such protocols in routine experimentation.
(d) Peptides not related to presentation by MHC can be generated as a result of proteolytic cleavage by endogenous proteases and peptidases. These enzymes originate from the sample analysed. This mechanism can lead to the false positive
generation of peptides that are not presented by cells in vivo as well as missing identifications of relevant peptides due to degradation. Approaches for detection of such contaminations use a statistical analysis of the protein coverage as metric to exclude problematic proteins. This requires detailed analysis of large data sets and, again, close human attention going far beyond any routine experimentation.
All these aspects need to be considered together when identifying peptides presented on the surface of certain cells. As a result, even the identification of a single peptide as an MHC ligand cannot be considered routine experimentation.
What is more, after identifying a peptide as being presented on cancer cells, it needs to be assessed whether the peptide is presented on cells of healthy tissue or not. As explained above, even the knowledge of a particular peptide that is being searched for, cannot help in its identification. Instead, in this case a careful screening of all peptides identified on healthy tissue is required, with the aim of detecting the presence of one particular peptide. Due to the inherent error sources of the techniques used, this renders analysis even more complicated than identifying a peptide presented on cancer tissue. That is, while identifying MHC ligands could be restricted based on the data to the most reliable results, since any identified MHC ligand could be useful, for the comparison whether a certain peptide is or is not presented on healthy tissue, no data can be omitted. Thereby, an additional level of complexity is introduced into the method required for identification of peptides suitable for cancer therapy, rendering this aspect of the identification even more complicated.
The approach followed by the present inventors thereby on one hand ensures that only peptides found in patient samples are identified, and on the other hand minimizes any risk that contaminations and errors falsify the results (Fritsche, J, et al., Mol Cell Proteomics (2021 ) 20, DOI: /10.1016/j.mcpro.2021 .100110).
Peptides
As described herein, provided are peptides comprising an amino acid sequence selected from the group consisting of SEQ ID NO: 1 to SEQ ID NO: 59, or a pharmaceutically acceptable salt thereof.
The peptides disclosed bind to inter alia HLA-A*01 , HLA-A*02, HLA-A*03, HLA-A*11 , HLA-A*23, HLA-A*68, HLA-B*07, HLA-B*08, HLA-B*13, HLA-B*14, HLA-B*15, HLA- B*18, HLA-B*27, HLA-B*35, HLA-B*37, HLA-B*38, HLA-B*40, HLA-B*44, HLA-B*51 , HLA-B*57, and HLA-C*03 allotype MHC molecules.
In one embodiment, said peptide thereof has a length according to the respective SEQ ID NO: 1 to SEQ ID NO: 59. In one embodiment, the peptide consists of the amino acid sequence according to SEQ ID NO: 1 to SEQ ID NO: 59. As disclosed herein, Table 1 shows the amino acid sequences of the peptides disclosed herein and their respective SEQ ID NOs.
A peptide disclosed herein can include up to four amino acids at the N- or the C-terminus in any combination between 4:0 and 0:4, relative to an amino acid sequence provided herein. That is, for example 1 or 2 amino acids can be added to either end. Also, amino acids can be added to either end. Combinations of possible elongations can be found in Table 2.
The amino acids for the elongation/extension can be the peptides of the original sequence of the protein or any other amino acid(s). The elongation can be used to enhance the stability or solubility of the peptides.
Thus, a peptide sequence used herein may be identical to a naturally occurring tumor- associated or tumor specific peptide sequence.
In some embodiments, the peptide is elongated on either or both ends by more than 4 amino acids, such as to a total length of up to 30 amino acids. This may lead to MHC
class ii binding peptides. Binding to MHC class ii can be tested by methods known in the art.
Accordingly, a peptide provided herein may have a maximal length of up to 30 amino acids, including a maximal length of up to 20 amino acids. In some embodiments a peptide provided herein may have a maximal length of up to 18, including of up to 15 amino acids. In some embodiments a peptide provided herein may have a maximal length of up to 14, including of up to 13 amino acids. In some embodiments a peptide provided herein may have a minimal length of 8, including of 9 amino acids. In some embodiments a peptide provided herein may have a minimal length of 10 amino acids.
In some embodiments a peptide provided herein may have an overall length from 8 to 30 amino acids, including from 8 to 13 amino acids. In some embodiments a peptide provided herein may have an overall length from 10 to 16 amino acids, including from 10 to 12 amino acids.
In some embodiments a peptide provided herein may have a length of 8 or 9 amino acids. In some embodiments a peptide provided herein may have a length of 10 or 11 amino acids. In some embodiments a peptide provided herein may have a length of 12 or 13 amino acids. In some embodiments a peptide provided herein, in particular, in case of an elongated class II binding peptide, may have a length of 14 or 15 amino acids. In some embodiments a peptide provided herein, in particular, in case of an elongated class II binding peptide, may have a length of 16 or 17 amino acids. In some embodiments a peptide provided herein, in particular, in case of an elongated class II binding peptide, may have a length of 18 or 19 amino acids. In some embodiments a peptide provided herein, in particular, in case of an elongated class II binding peptide, may have a length of 20 or 21 amino acids. In some embodiments a peptide provided herein, in particular, in case of an elongated class II binding peptide, may have a length of 22 or 23 amino acids.
As noted above, in some embodiments a peptide provided herein in addition to the sequence according to any of SEQ ID NO: 1 to SEQ ID NO: 59 contains one or more further N- and/or C-term inally located stretches of amino acids that are not necessarily forming part of the peptide that functions as an epitope for MHC molecules.
Nevertheless, these stretches can be important to provide an efficient introduction of the peptide provided herein into the cells. In one embodiment, the peptide is part of a fusion protein which includes, for example, the 80 N-terminal amino acids of the HLA-DR antigen-associated invariant chain (p33, in the following “Ii”) as derived from the NCBI, GenBank Accession number X00497. In other fusions, the peptides provided herein can
be fused to an antibody as described herein, or a functional part thereof, in particular, into a sequence of an antibody, so as to be specifically targeted by said antibody, or, for example, to or into an antibody that is specific for dendritic cells as described herein.
As already explained above, due to their length, a peptide as disclosed herein when bound by a member of the immune response or by a binding molecule, can be taken to define an antigenic epitope (Murphy, 2022).
Peptide modifications
In addition, the peptide may be modified further to improve stability and/or binding to MHC molecules in order to elicit a stronger immune response. Methods for such an optimization of a peptide sequence are well known in the art and include, for example, the introduction of reverse peptide bonds or non-peptide bonds.
An example of a non-peptide bond is a reverse peptide bond, in which amino acid residues are not joined by peptide (-CO-NH-) linkages but the peptide bond is reversed. Such retro-inverse peptidomimetics may be made using methods known in the art, for example such as those described by Meziere and colleagues (Meziere et al., 1997, incorporated herein by reference). This approach involves making pseudopeptides containing changes involving the backbone, and not the orientation of side chains. They show that for MHC binding and T helper cell responses, these pseudopeptides are useful. Retro-inverso peptides, which contain D-amino acids in the reverse sequence, are much more resistant to proteolysis (Meziere et al., 1997).
The peptide as disclosed herein may in some embodiments include one or more non- peptide bonds. A non-peptide bond is, for example, -CH2-NH or -CH2S-. A further example of a non-peptide bond is -CH2CH2- or -CH=CH- Two further examples of a non- peptide bond are -COCH2- or -CH(OH)CH2-. A further example of a non-peptide bond is -CH2SO-. A further example of a non-peptide bond is an N-methyl or an N-hydroxyl group contained in the bond between two amino acids. US 4,897,445 provides a method for the solid phase synthesis of non-peptide bonds (-CH2-NH) in polypeptide chains which involves polypeptides synthesized by standard procedures and the non-peptide bond synthesized by reacting an amino aldehyde and an amino acid in the presence of NaCNBHs.
Peptides comprising the bonds described above may be synthesized with additional chemical groups present at their amino and/or carboxy termini, to enhance the stability, bioavailability, and/or affinity of the peptides. For example, hydrophobic groups such as carbobenzoxyl, dansyl, or t-butyloxycarbonyl groups may be added to the peptides'
amino termini. Likewise, an acetyl group or a 9-fluorenylmethoxy-carbonyl group may be placed at the peptides' amino termini. Additionally, the hydrophobic group, t- butyloxycarbonyl, or an amido group may be added to the peptides' carboxy terminus.
In some embodiments, the peptides disclosed herein comprise no non-peptide bonds. In other words, the peptides disclosed herein in some embodiments comprise only peptide bonds.
Peptide salts
A peptide as provided herein may be provided in the form of a salt, in particular a pharmaceutically acceptable salt.
In embodiments where a peptide is to be included into a pharmaceutical composition that is of an overall hydrophobic nature, the peptide may be provided in the form of a pharmaceutically acceptable salt. Additionally, the peptide may be delivered using a sustained-release system.
In some embodiments a non-naturally occurring peptide has been synthetically produced (e.g. synthesized) as a pharmaceutically acceptable salt. Methods to synthetically produce peptides are well known in the art. See below.
It has to be noted that a salt of a peptide provided herein differs substantially from the respective peptide in its state in vivo, as the peptide is not in the form of a salt or associated with counterions in vivo. A salt may be provided to increase the shelf-life of a peptide provided herein.
A salt may also be provided to alter the solubility of a peptide provided herein in an aqueous solvent, including saline or water. The salt form of the peptide mediates the solubility of the peptide, in particular in the context of pharmaceutical compositions comprising a peptide as disclosed herein. A sufficient and at least substantial solubility of the peptide(s) is required in order to efficiently provide the peptides to the subject to be treated.
Typically, a salt as used herein is a pharmaceutically acceptable salt of the corresponding peptide.
An acid salt may be prepared by contacting the peptide with a suitable acid. Suitable acids for preparing an acid salt include both an organic acid and an inorganic acid. A suitable organic acid may in some embodiments be a carboxylic acid. Two examples of a suitable organic acid are acetic acid and propionic acid. Further examples of a suitable
organic acid are glycolic acid and pyruvic acid. Oxalic acid and malic acid are yet further examples of a suitable organic acid. Two further organic acids that may be used are malonic acid and succinic acid. Yet two further examples of a suitable organic acid are maleic acid and fumaric acid. Further examples of a suitable organic acid are tartaric acid and citric acid. Benzoic acid and cinnamic acid are further examples of a suitable organic acid. Furthermore, mandelic acid and salicylic acid are suitable organic acids. Yet two further suitable organic acids are methane sulfonic acid and an aryl sulfonic acid. Two examples of an aryl sulfonic acid are benzenesulfonic acid and p-toluenesulfonic acid. An example of a suitable organic acid is also ethane sulfonic acid.
Two examples of a suitable inorganic acid are hydrochloric acid and hydrobromic acid Sulfuric acid and nitric acid are further examples of a suitable inorganic acid. Yet a further example of a suitable inorganic acid is phosphoric acid. Conversely, preparation of a basic salt of an acid moiety which may be present on a peptide may be prepared using a pharmaceutically acceptable base. A pharmaceutically acceptable base may for instance be an inorganic base or an organic base. Two suitable inorganic bases are sodium hydroxide and potassium hydroxide. Yet two further suitable inorganic bases are ammonium hydroxide and calcium hydroxide. Two suitable organic bases are trimethylamine and triethylamine.
A salt as provided herein may be an alkaline or an earth alkaline salt, such as a salt of the Hofmeister series. As anions, the salt may include PO43 or SCU2’. Two further suitable anions are CHsCFteCOO and HOCHsCOO. Yet two further suitable anions are Cl’ and Br. NO3’ and CIO4 are two further suitable anions. Two further suitable anions are I’ and SCN’. Two suitable cations are NH4+ and Rb+. Two further suitable cations are K+ and Na+. Cs+ and Li+ are two further suitable cations. Two further suitable cations are Zn2+ and Mg2+. Yet two further suitable cations are Ca2+ and Mn2+. Also, Cu2+ and Ba2+ are two suitable cations.
Particularly salts are selected from (NH4)3PO4, (NH4)2HPO4, (NH4)H2PO4, (NH4)2SO4, NH4CH3COO, NH4CI, NH4Br, NH4NO3, NH4CIO4, NH4I, NH4SCN, Rb3PO4, Rb2HPO4, RbH2PO4, Rb2SO4, Rb4CH3COO, Rb4CI, Rb4Br, Rb4NO3, Rb4CIO4, Rb4l, Rb4SCN, K3PO4, K2HPO4, KH2PO4, K2SO4, KCH3COO, KCI, KBr, KNO3, KCIO4, KI, KSCN, Na3PO4, Na2HPO4, NaH2PO4, Na2SO4, NaCHsCOO, NaCI, NaBr, NaNOs, NaCICU, Nal, NaSCN, ZnCI2 Cs3PO4, CS2HPO4, CsH2PO4, Cs2SO4, CsCHsCOO, CsCI, CsBr, CsNOs, CsCIO4, Csl, CsSCN, U3PO4, Li2HPO4, LiH2PO4, U2SO4, LiCHsCOO, LiCI, LiBr, LiNOs, LiCICU, Lil, LiSCN, Cu2SO4, Mg3(PO4)2, Mg2HPO4, Mg(H2PO4)2, Mg2SO4, Mg(CH3COO)2, MgCI2, MgBr2, Mg(NO3)2, Mg(CIO4)2, Mgl2, Mg(SCN)2, MnCh, Ca3(PO4), Ca2HPO4, Ca(H2PO4)2, CaSO4, Ca(CH3COO)2, CaCI2, CaBr2, Ca(NO3)2, Ca(CIO4)2, Cal2, Ca(SCN)2, Ba3(PO4)2, Ba2HPO4, Ba(H2PO4)2, BaSO4, Ba(CH3COO)2, BaCh,
BaBr2, Ba(NO3)2, Ba(CIO4)2, Bah, and Ba(SCN)2. In some embodiments the salt is ammonium acetate, MgCh, KH2PO4, Na2SO4, KCI, NaCI, and CaCh, such as, for example, the chloride or acetate (trifluoroacetate) salts (see e.g. Berge, S.M., et al., Journal of Pharmaceutical Science (1977) 66, 1 -19, the contents of which are incorporated by reference in their entirety).
A pharmaceutically acceptable salt can be taken to be a derivative of a peptide as disclosed herein, wherein the peptide is modified by making an acid or base salt of the agent. For example, acid salts are prepared from the free base (typically wherein the neutral form of the drug has a neutral -NH2 group) involving reaction with a suitable acid. Suitable acids for preparing acid salts include both organic acids e.g. acetic acid, trifluoro acetic acid, propionic acid, glycolic acid, pyruvic acid, oxalic acid, malic acid, malonic acid, succinic acid, maleic acid, fumaric acid, tartaric acid, citric acid, benzoic acid, cinnamic acid, mandelic acid, methane sulfonic acid, ethane sulfonic acid, p- toluenesulfonic acid, salicylic acid, and the like, as well as inorganic acids, e.g., hydrochloric acid, hydrobromic acid, sulfuric acid, nitric acid phosphoric acid and the like. Conversely, preparation of basic salts of acid moieties which may be present on a peptide are prepared using a pharmaceutically acceptable base such as sodium hydroxide, potassium hydroxide, ammonium hydroxide, calcium hydroxide, or trimethylamine. In some embodiments, a pharmaceutical composition contains one or more peptides as salts of acetic acid (acetates), trifluoro acetates or hydrochloric acid (chlorides).
Chemical peptide synthesis
As noted above, in some embodiments a peptide as disclosed herein has been chemically synthesized. Generally, peptides may be synthesized by the Fmoc-polyamide mode of solid-phase peptide synthesis as disclosed by Lukas et al. (Lukas et al., 1981 , the content of which is incorporated by reference in its entirety) and by references as cited therein. Temporary N-amino group protection is afforded by the 9- fluorenylmethyloxycarbonyl (Fmoc) group. Repetitive cleavage of this highly base-labile protecting group is done using 20% piperidine in N,N-dimethylformamide. Side-chain functionalities may be protected as their butyl ethers (in the case of serine, threonine and tyrosine), butyl esters (in the case of glutamic acid and aspartic acid), butyloxycarbonyl derivative (in the case of lysine and histidine), trityl derivative (in the case of cysteine) and 4-methoxy-2,3,6-trimethylbenzenesulphonyl derivative (in the case of arginine). Where glutamine or asparagine are C-terminal residues, use is made of the 4,4'- dimethoxybenzhydryl group for protection of the side chain amido functionalities. The solid-phase support is based on a polydimethyl-acrylamide polymer constituted from the three monomers dimethylacrylamide (backbone-monomer), bisacryloylethylene diamine (cross linker) and acryloylsarcosine methyl ester (functionalizing agent). The peptide-to-
resin cleavable linked agent used is the acid-labile 4-hydroxymethyl-phenoxyacetic acid derivative. All amino acid derivatives are added as their preformed symmetrical anhydride derivatives with the exception of asparagine and glutamine, which are added using a reversed coupling procedure involving N,N-dicyclohexyl-carbodiimide and 1 - hydroxybenzotriazole. All coupling and deprotection reactions are monitored using ninhydrin, trinitrobenzene sulphonic acid or isotin test procedures. Upon completion of synthesis, peptides are cleaved from the resin support with concomitant removal of sidechain protecting groups by treatment with 95% trifluoroacetic acid containing a 50% scavenger mix. Scavengers commonly used include ethanedithiol, phenol, anisole and water, the exact choice depending on the constituent amino acids of the peptide being synthesized. Also, a combination of solid phase and solution phase methodologies for the synthesis of peptides is possible (see for example Bruckdorfer et al., 2004 and the references as cited therein, the content of which is incorporated by reference in its entirety).
Trifluoroacetic acid may be removed by evaporation in vacuo, with subsequent trituration with diethyl ether affording the crude peptide. Any scavengers present can be removed by a simple extraction procedure which on lyophilization of the aqueous phase affords the crude peptide free of scavengers. Reagents for peptide synthesis are generally available from e.g., Calbiochem-Novabiochem (Nottingham, UK).
Purification may involve any desired technique, such as re-crystallization, size exclusion chromatography, ion-exchange chromatography, hydrophobic interaction chromatography and (usually) reverse-phase high performance liquid chromatography using e.g., aceton itrile/water gradient separation.
Analysis of synthesized peptides may be carried out using thin layer chromatography, electrophoresis, in particular capillary electrophoresis, solid phase extraction (CSPE), reverse-phase high performance liquid chromatography, amino-acid analysis after acid hydrolysis and by fast atom bombardment (FAB) mass spectrometric analysis, as well as MALDI and ESI-Q-TOF mass spectrometric analysis.
Peptide Production by Expression
Provided is furthermore a method for producing the peptide as disclosed herein. The method includes culturing a host cell such as a recombinant host cell that expresses or contains the peptide. The method furthermore includes isolating the peptide, from the host cell and/or from the culture medium that contains the host cell.
In an aspect, a cDNA encoding a peptide according as disclosed herein, such as a peptide according to SEQ ID NO: 1 to SEQ ID NO: 59, can be expressed in prokaryotic
cells (e.g., bacteria) or eukaryotic cells (e.g., yeast, insect, or mammalian cells), after which the recombinant peptide can be purified.
Uses
Treatment and prevention
In a further aspect there is provided a method of treating a subject having cancer or of preventing a subject from suffering from cancer. The method includes administering to the subject a peptide as provided herein. The method may include administering to a subject a pharmaceutically effective amount of the peptide.
In a further aspect there is provided a method of treating a subject having cancer or of preventing a subject from suffering from cancer. The method includes administering to the subject a binding molecule as provided herein. The method may include administering to a subject a pharmaceutically effective amount of the binding molecule.
In a further aspect there is provided a method of treating a subject having cancer or of preventing a subject from suffering from cancer. The method includes administering to the subject a nucleic acid or an expression vector as provided herein. The method may include administering to a subject a pharmaceutically effective amount of the nucleic acid or the expression vector.
In a further aspect there is provided a method of treating a subject having cancer or of preventing a subject from suffering from cancer. The method includes administering to the subject a recombinant host cell, including a plurality of recombinant host cells, as provided herein. The method may include administering to a subject a pharmaceutically effective amount of the host cell(s).
In a further aspect there is provided a method of treating a subject having cancer or of preventing a subject from suffering from cancer. The method includes administering to the subject an activated T lymphocyte, including a plurality of activated T lymphocytes, as provided herein. The method may include administering to a subject a pharmaceutically effective amount of the activated T lymphocyte(s).
In a further aspect there is provided a method of treating a subject having cancer or of preventing a subject from suffering from cancer. The method includes administering to the subject a pharmaceutical composition or a combination product as provided herein. The method may include administering to a subject a pharmaceutically effective amount of the pharmaceutical composition or the combination product.
In a further aspect there is provided a method of treating a subject having cancer or of preventing a subject from suffering from cancer. The method includes administering to the subject a composition that contains a population of activated T cells that kill cancer cells in the subject that over-present a peptide as described herein on their cell surface. The method may include administering to a subject a pharmaceutically effective amount of the population of activated T cells.
Any peptide or pharmaceutically acceptable salt thereof as disclosed herein may be used in medicine. Furthermore, any binding molecule as disclosed herein may be used in medicine. Any nucleic acid and any expression vector that contains such a nucleic acid as disclosed herein may be used in medicine. Also, any host cell and any T lymphocyte as disclosed herein may be used in medicine. Any cell line as disclosed herein may also be used in medicine. Also, any pharmaceutical composition as disclosed herein may be used in medicine. Any kit or combination product as disclosed herein may also be used in medicine.
The use in medicine may be a use in treating and/or preventing cancer. A peptide as disclosed herein may for example be used in generating a binding molecule, such as a TCR, a TCR fragment, an antibody, an antibody fragment or an aptamer. The binding molecule may then be administered to an individual. In some embodiments the binding molecule may be coupled to a toxin, in particular a compound having cytotoxic effect as illustrated below. The binding molecule may for example be coupled to an RNA polymerase inhibitor or to an RNA-splicing inhibitor. The antibody may for example be coupled to a toxin via a cleavable linker. A peptide as disclosed herein may also be administered to a subject in order to induce or stimulate an immune response.
Within the context of a medical use or medical treatment, a host cell may be used to produce a proteinaceous binding molecule or a peptide as disclosed herein. The proteinaceous binding molecule or the peptide may then be administered to a subject.
In some embodiments a nucleic acid encoding a peptide as disclosed herein may be used for preventing occurrence of a cancer or for treating a cancer. The nucleic acid may be included in a vector, such as an expression vector. The nucleic acid, including the vector, may be designed to enter cells and transcribe, translate and process a peptide as disclosed herein. The respective vector can in some embodiments be taken to define a recombinant viral or bacterial vaccine.
A T lymphocyte or a T cell line may be directly administered to a subject. The T lymphocyte may for example be autologous to the subject. The T lymphocyte may also be allogenic. In some embodiments a T lymphocyte used may be obtained from a healthy
donor. The T lymphocyte may be derived from a tumor infiltrating lymphocyte or from a peripheral blood mononuclear cell. In some embodiments the T lymphocyte may be recombinant in that it has been genetically engineered to express a TCR of a desired specificity or affinity to a peptide as disclosed herein.
Provided is the peptide or the salt thereof as disclosed herein for use in the treatment and/or prevention of cancer. Provided is also the nucleic acid, or the vector that contains the nucleic acid, as disclosed herein for use in the treatment and/or prevention of cancer. Provided are furthermore the host cell or the T lymphocyte as disclosed herein for use in the treatment and/or prevention of cancer. Provided are furthermore the pharmaceutical composition or combination product as disclosed herein for use in the treatment and/or prevention of cancer. Provided is also the kit as disclosed herein for use in the treatment and/or prevention of cancer.
Provided is also the use of the peptide or the salt thereof as disclosed herein in the manufacture of a medicament for treating and/or preventing cancer. Provided is also the use of the nucleic acid, or the vector that contains the nucleic acid, as disclosed herein in the manufacture of a medicament for treating and/or preventing cancer. Provided are furthermore the use of the host cell or the T lymphocyte as disclosed herein in the manufacture of a medicament for treating and/or preventing cancer. Provided are furthermore the use of the pharmaceutical composition or combination product as disclosed herein in the manufacture of a medicament for treating and/or preventing cancer. Provided is also the use of the kit as disclosed herein in the manufacture of a medicament for treating and/or preventing cancer.
Provided is furthermore a method of treating and/or preventing cancer. The method includes administering to a subject suffering from cancer an effective amount of a peptide or a salt thereof, a binding molecule, a nucleic acid, an expression vector, a host cell, a T lymphocyte as disclosed herein or any combination thereof.
In a further aspect, provided is the peptide or the salt thereof as disclosed herein for use in the diagnosis of cancer. Provided is also the nucleic acid, or the vector that contains the nucleic acid, as disclosed herein for use in the diagnosis of cancer. Provided are furthermore the host cell or the T lymphocyte as disclosed herein for use in the diagnosis of cancer. Provided are furthermore the pharmaceutical composition or combination product as disclosed herein for use in the diagnosis of cancer. Provided is also the kit as disclosed herein for use in the diagnosis of cancer.
A use in the diagnosis of cancer may also be carried out in the form of an in vitro method. Diagnosing cancer may also be carried out in the form of an in vivo method
Provided is furthermore a method of monitoring the risk of occurrence of cancer in a subject. The method may be an in vitro method carried out on one or more samples that have been obtained from the respective subject. The method includes detecting whether a peptide as disclosed herein is presented on cells of the sample. A binding molecule as disclosed herein may be used, for example in labelled form. The method may include detecting the presence of the peptide, or the relative amount of the peptide, relative to a control. The control may be defined by one or more healthy samples of the same tissue type or cell type.
Provided is furthermore a method of monitoring the treatment of a cancer in a subject. The method may be an in vitro method carried out on one or more cancer samples that have been obtained from the respective subject. The method includes detecting whether a peptide as disclosed herein is presented on cells of the sample. The method may also include estimating or assessing whether a peptide as disclosed herein is presented on cells of the sample. A binding molecule as disclosed herein may be used. Again, the method may include detecting the presence of the peptide, or the relative amount of the peptide, relative to a control. The control may be defined by one or more healthy samples of the same tissue type or cell type.
Provided is also a method of eliciting an immune response in a subject that has cancer. The method includes administering to the subject a plurality of T lymphocytes as disclosed herein. The T lymphocytes may be activated T lymphocytes. T lymphocytes kill cancer cells in the subject that present a peptide as disclosed herein. The T lymphocytes may in some embodiments be autologous to the subject. The T lymphocytes may in some embodiments be allogenic. In some embodiments the T lymphocytes used may be obtained from a healthy donor. The T lymphocytes may be derived from tumor infiltrating lymphocytes or from peripheral blood mononuclear cells. In some embodiments the T lymphocytes may be recombinant in that they have been genetically engineered to express a TCR of a desired specificity or affinity to a peptide as disclosed herein.
Provided is furthermore a method of killing cancer cells in a subject. The cancer cells present a peptide as disclosed herein, bound by MHC. The method includes administering to the subject an effective number of T lymphocytes as disclosed herein. The T lymphocytes may be activated T lymphocytes.
A method of killing cancer cells may also be carried out in the form of an in vitro method. A method of killing cancer cells may also be carried out in the form of an in vivo method. Provided is furthermore an activated T lymphocyte as disclosed herein for use in the killing of target cells in a subject. The target cells present a peptide that contains any
amino acid sequence disclosed herein. As disclosed herein, the respective target cells are cancer cells.
In this regard, provided is a T lymphocyte as disclosed herein for use in the manufacture of a medicament for the killing of cancer cells in a subject. The cancer cells present a peptide as disclosed herein, bound by MHC. The T lymphocytes may be activated T lymphocytes.
Provided is also the use a T lymphocyte as disclosed herein in the killing of target cells in a subject. The cancer cells present a peptide as disclosed herein, bound by MHC. The T lymphocytes may be activated T lymphocytes.
In any of the uses and methods described above, the cancer may be acute myeloid leukemia or breast cancer. In some embodiments the cancer may be cholangiocellular carcinoma or chronic lymphocytic leukemia. The cancer may in some embodiments be colorectal cancer or gallbladder cancer. In some embodiments the cancer may be glioblastoma. The cancer may in some embodiments be gastric cancer or gastroesophageal junction cancer. In some embodiments the cancer may be hepatocellular carcinoma or head and neck squamous cell carcinoma. In some embodiments the cancer may be melanoma. The cancer may in some embodiments be non-Hodgkin lymphoma. The cancer may in some embodiments be non-small cell lung cancer or small cell lung cancer. In some embodiments the cancer may be ovarian cancer or uterine endometrial cancer. The cancer may in some embodiments be esophageal cancer. The cancer may in some embodiments be pancreatic cancer or prostate cancer. In some embodiments the cancer may be renal cell carcinoma or urinary bladder carcinoma. The cancer may also be another cancer that shows an over-presentation of a peptide as disclosed herein.
Diagnostics
In a further aspect there is provided a method of diagnosing cancer in a subject. The subject may be suspected of having cancer or be found to be at risk of developing cancer. The method includes using a binding molecule as provided herein. Generally, such a binding molecule is a proteinaceous binding molecule that specifically binds to a peptide as provided herein, or to the respective peptide when in complex with an MHC molecule, see above. The method may include administering the binding molecule to a subject or contacting the binding molecule with a sample that has been obtained from a subject. Typically, the method involves targeting cells, including cells as part of tissue, from or in the subject with a binding molecule as disclosed herein. In this regard in some embodiments the method includes providing a sample from the subject. The method furthermore includes detection of binding of the binding molecule to cells, for example
cells in one or more tissue samples, of or in the subject. By detecting binding of the binding molecule, generally the presentation of a peptide as disclosed herein on cells originating from the subject is identified.
The peptide and/or the TCR, antibody, aptamer or other binding molecule provided herein can be used in the diagnosis of cancer. The peptide or binding molecule can be used to verify a pathologist’s diagnosis of a cancer based on a sample, e.g. a biopsied sample.
The cells originating from the subject may be cells of a tissue or of a body fluid obtained from the subject. In some embodiments, a biopsy of a tumor or a cancerous body fluid sample is collected and subjected to immunoprecipitation of peptide MHC complexes, with subsequent analysis of the peptidome thus obtained by means of mass spectrometry. Respective methods are e.g. disclosed in Fritsche et al. (Fritsche, J, et al., Proteomics (2018) 18, 1700284), the content of which is incorporated herein by reference.
In some embodiments, a labelled binding molecule, e.g. a T cell receptor or a TCR mimetic antibody, specific for the peptide MHC complex comprising a peptide as disclosed herein. In one embodiment, a biopsy or sample of the cancer tissue is obtained, rated with a desired immunological method (e.g. sliced or homogenized in case of a solid tumor) and then incubated with the labelled binding molecule. See e.g. Hoydahl et al. (Hoydahl, LS, et al., Antibodies (Basel), (2019) 8, 2, 32, doi:10.3390/antib8020032) for methods, the content of which is incorporated herein by reference.
An antibody, TCR, aptamer or other binding molecule may also be used in both in vitro and in vivo diagnostic assays. Generally, the binding molecule is labeled with a radionucleotide (such as 1111n, "Tc, 14C, 131 l, 3H, 32P or 35S) so that the tumor can be localized using immunoscintiography.
Since the peptides were generated from acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, nonHodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer cells and since it was determined that these peptides are not or at lower levels present in normal tissues, these peptides can be used to diagnose the presence of a cancer.
The presence of a peptide as disclosed herein on cells of a cancer sample, e.g. on a tissue biopsy or in a blood sample, can assist a pathologist in diagnosis of cancer. Detection of certain peptides by means of antibodies, mass spectrometry or other methods known in the art can tell the pathologist that the tissue sample is malignant or inflamed or generally diseased, or can be used as a biomarker for acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer. The presence of groups of peptides can enable classification or sub-classification of diseased tissues.
The detection of peptides on diseased tissue specimen can enable the decision about the benefit of therapies involving the immune system, especially if T-lymphocytes are known or expected to be involved in the mechanism of action. Loss of MHC expression is a well described mechanism by which infected or malignant cells escape immunosurveillance. Thus, presence of peptides shows that this mechanism is not exploited by the analyzed cells.
In a related aspect there is provided an in vitro method of assessing the risk of a subject to have or to develop cancer. The method includes adding a binding molecule, such as a proteinaceous binding molecule, to a sample that contains cells of from said subject. The binding molecule is labeled with a probe or a radionucleotide. The method furthermore includes detecting whether the binding molecule binds to cells in the sample from the subject. The binding molecule specifically binds to (i) a peptide that includes the amino acid sequence according to any one of SEQ ID NOs: 1 to 59 or (ii) said peptide that includes the amino acid sequence according to any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule. The detection of binding of the binding molecule to cells in the sample from the subject identifies the presentation of said peptide on said cells. The detection thereby indicates a risk of the subject to have or to develop cancer.
Monitoring
A peptide provided herein may be used to monitor and analyze lymphocyte responses against those peptides such as T cell responses or antibody responses against the peptide or the peptide complexed to MHC molecules. These lymphocyte responses can be used as prognostic markers for decision on further therapy steps. These responses can also be used as surrogate response markers in immunotherapy approaches aiming to induce lymphocyte responses by different means, e.g. vaccination of protein, nucleic acids,
autologous materials, adoptive transfer of lymphocytes. In gene therapy settings, lymphocyte responses against peptides can be considered in the assessment of side effects. Monitoring of lymphocyte responses might also be a valuable tool for follow-up examinations of transplantation therapies, e.g. for the detection of graft versus host and host versus graft diseases.
Provided herein is a method of monitoring a cancer therapy in a subject. The method includes using a binding molecule, such as a proteinaceous binding molecule, as provided herein. The method may include administering the binding molecule to a subject or contacting the binding molecule with a sample that has been obtained from a subject. Typically, the method involves targeting cells, including cells as part of tissue, from or in the subject with a binding molecule as disclosed herein. In some embodiments the method includes providing a sample from the subject. The method furthermore includes detection of binding of the binding molecule to cells, for example cells in one or more tissue samples, of or in the subject.
According to a particular embodiment of the method of monitoring a cancer therapy, targeting cells from or in the subject with the binding molecule is performed at certain, e.g. predetermined, time intervals. Samples from the subject may be provided that have been obtained at the corresponding time points.
Explanations above on the method of diagnosing cancer in a subject also apply to the method of monitoring a cancer therapy in a subject.
Peptides
A peptide as disclosed herein is over-presented in cancer tissue compared to healthy tissues (see Table 6) and thus can be used for the production of antibodies and/or TCRs, such as soluble TCRs, as disclosed herein for use in the treatment of cancer, or for the production of vaccines against cancer. Specifically, the peptides when complexed with the respective MHC can be used for the production of an antibody and/or a TCR, for example a soluble TCR for use in the treatment of cancer. Respective methods are well known to the person of skill in the art and can be found in the respective literature as well (see also below).
Provided herein are peptides that are useful in treating cancers/tumors, such as acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastroesophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma,
small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer that overpresent a peptide as disclosed herein. These peptides were shown by MS to be naturally presented by MHC molecules on primary human acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer samples.
Some of the source genes/proteins (also designated “full-length proteins” or “underlying proteins”) from which the peptides are derived were shown to be highly overexpressed in cancer compared with normal tissues, demonstrating a high degree of tumor association of the source genes (see Example 2). However, as explained above, there is no general correlation between expression of the source protein and the presentation of peptides derived therefrom. Moreover, the peptides themselves are over-presented on tumor tissue relative to normal tissues (see Example 1 ).
MHC bound peptides can be recognized by the immune system, specifically T lymphocytes. T cells can destroy the cells presenting the recognized MHC-peptide complex, e.g. acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer cells presenting the derived peptides.
Nucleic Acid
In a further aspect, there is provided a nucleic acid molecule. The nucleic acid molecule may contain a sequence that encodes a peptide as disclosed herein. The sequence that encodes the peptide may be linked to a heterologous promoter sequence. The sequence that encodes the peptide may be operatively coupled to a heterologous promoter sequence. In some embodiments the nucleic acid molecule contains an open reading frame (ORF) that contains a sequence coding for a peptide as disclosed herein.
The nucleic acid molecule may in some embodiments contain a sequence that encodes a proteinaceous binding molecule as disclosed herein. The sequence that encodes the proteinaceous binding molecule may be linked to a heterologous promoter sequence.
The sequence that encodes the proteinaceous binding molecule may be operatively coupled to a heterologous promoter sequence.
A respective nucleic acid may for instance be a DNA molecule or a cDNA molecule. A respective nucleic acid may also be a PNA molecule or an RNA molecule. In some embodiments a respective nucleic acid may be a combination of one or more of a DNA molecule, a cDNA molecule, a PNA molecule and/or an RNA molecule. In some embodiments the nucleic acid molecule is an artificial nucleic acid molecule.
In some embodiments, the nucleic acid is included in a vector, such as an expression vector. In such embodiments the sequence encoding a peptide may be operably linked to one or more regulatory sequences such as a promoter. Likewise, the sequence encoding a proteinaceous binding molecule may be operably linked to one or more regulatory sequences such as a promoter. The vector is in some embodiments an artificial vector.
In some embodiments the nucleic acid may be complexed or associated with a lipid. In some embodiments the nucleic acid may be encapsulated in a carrier particle that contains a lipid.
In some embodiments a nucleic acid molecule containing a sequence that encodes a peptide or variant as provided herein, may be included in a vaccine composition. A respective vaccine composition may for example be used for eliciting or for enhancing an anti-tumor immune response.
In some embodiments a nucleic acid molecule provided herein contains a codon- optimized region. In some embodiments a nucleic acid molecule contains a sequence encoding a peptide or variant thereof or a proteinaceous binding molecule as disclosed herein that is codon-optimized. In some embodiments a nucleic acid molecule as disclosed herein contains an open reading frame that is codon-optimized.
Vector
In a further aspect there is provided an expression vector. The expression vector includes a nucleic acid as disclosed herein.
Host Cell
Provided is also a host cell, such as a recombinant host cell, that includes a peptide as disclosed herein. Provided is also a host cell, such as a recombinant host cell, that includes a proteinaceous binding molecule, e.g. an antibody or fragment thereof or a T cell receptor or fragment thereof, as provided herein.
Provided is also a host cell that includes a nucleic acid as disclosed herein or an expression vector as disclosed herein. A respective host cell may be an antigen- presenting cell. In some embodiments the host cell is a dendritic cell. In some embodiments the host cell is a T cell or an NK cell.
In some embodiments the host cell is a T lymphocyte or an NK cell expressing a heterologous proteinaceous binding molecule. The heterologous proteinaceous binding molecule is in some embodiments a heterologous TCR. The heterologous TCR specifically recognizes a peptide comprising an amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising an amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule.
Pharmaceutical Compositions
General
In a further aspect there is provided a pharmaceutical composition. The pharmaceutical composition contains at least one active ingredient and a pharmaceutically acceptable carrier and/or a pharmaceutically acceptable excipient. The active ingredient may be a peptide or salt thereof as disclosed herein. The active ingredient may also be a binding molecule as disclosed herein, for instance an antibody, a functional fragment thereof, a T-cell receptor or a functional fragment thereof, or an aptamer. The active ingredient may also be a nucleic acid or an expression vector as disclosed herein. The active ingredient may also be a recombinant host cell or an T lymphocyte as disclosed herein.
A respective pharmaceutical composition may be provided for therapeutic purposes, for diagnostic purposes, or for both. As an example, a pharmaceutical composition provided herein may be a pharmaceutical composition for inducing an immune response in a human or animal.
In some embodiments the pharmaceutical composition is included in a vaccine. In some embodiments the pharmaceutical composition is a cellular therapeutic composition.
In one embodiment, such a pharmaceutical composition is an individualized, including a personalized, pharmaceutical composition for an individual subject.
As noted above, a pharmaceutical composition may contain one or more peptide(s) as disclosed herein either in the free form or in the form of a pharmaceutically acceptable salt (see also above). In some embodiments, a pharmaceutical composition contains one or more peptides as salts of acetic acid (acetates), trifluoro acetates or hydrochloric acid (chlorides).
In some embodiments, for example where the pharmaceutical composition contains a nucleic acid in the form of RNA, it may contain a stabilizing substance such as those disclosed in WO2023/044455. A respective stabilizing substance may for instance be dimethyl sulfoxide (DMSO) or diethyl sulfoxide. A respective stabilizing substance may also be dimethyl sulfone or N-Methyl-2-pyrrolidone. There may also be a supplemental RNA stabilizing substance included such as hexylene glycol or Na-hexametaphosphate. Further examples of a supplemental RNA stabilizing substance are trimetaphosphate or Na-Glycerol Phosphate.
A pharmaceutical composition, whether an individualized product or not, that includes an agent as disclosed herein may be manufactured in a manner that is itself known, e.g., by means of a mixing, dissolving, granulating, dragee-making, levigating, emulsifying, encapsulating, entrapping or lyophilizing process.
Adjuvants
In some embodiments the pharmaceutical composition contains a conjugated or labelled active ingredient. In some embodiments the pharmaceutical composition contains an adjuvant. In some embodiments the pharmaceutical composition contains an immunogenicity enhancing amount of an adjuvant. In some embodiments the adjuvant is an interleukin. The interleukin may for example be IL-2, IL-12, IL-15, IL-18, IL-21 or a combination thereof. In one embodiment the interleukin may be IL-2. In one embodiment the interleukin may be IL-12. In one embodiment the interleukin may be IL-15. In one embodiment the interleukin may be IL-18. In one embodiment the interleukin may be IL- 21. In some embodiments, the interleukin may be expressed by the T lymphocyte and may be membrane-bound or soluble.
A pharmaceutical composition, e.g. a vaccine composition, as disclosed herein may include one or more adjuvants. An adjuvant may also be included in a combination product. An adjuvant is a substance that non-specifically enhances or potentiates the immune response e.g. an immune response mediated by CD8-positive T cells and/or helper T (TH) cells to an antigen.
Suitable adjuvants include, but are not limited to, 1018 ISS, aluminum salts, AMPLIVAX®, AS15, BCG, CP-870,893, CpG7909, CyaA, dSLIM, flagellin or TLR5 ligands derived from flagellin, FLT3 ligand, GM-CSF, IC30, IC31 , Imiquimod (ALDARA®), Resiquimod, ImuFact IMP321 , an Interleukin, Interferon-alpha or -beta, or pegylated derivatives thereof, IS Patch, ISS, ISCOMATRIX, ISCOMs, Juvlmmune®, LipoVac, MALP2, MF59, monophosphoryl lipid A, Montanide IMS 1312, Montanide ISA 206, Montanide ISA 50V, Montanide ISA-51 , water-in-oil and oil-in-water emulsions, OK- 432, OM-174, OM-197-MP-EC, ONTAK, OspA, PepTel® vector system, poly(lactid co-
glycolid) [PLG]-based and dextran microparticles, talactoferrin SRL172, Virosomes and other Virus-like particles, YF-17D, VEGF trap, R848, beta-glucan, Pam3Cys, Aquila's QS21 stimulon, which is derived from saponin, mycobacterial extracts and synthetic bacterial cell wall mimics, and other proprietary adjuvants such as Ribi's Detox, Quil, or Superfos. Adjuvants such as Freund's or GM-CSF may be used. An example of an interleukin is IL-2. A further example of an interleukin is IL-15. Yet another example of an interleukin is IL-18. A further illustrative example of an interleukin is IL-21. Several immunological adjuvants (e.g. MF59) specific for dendritic cells and their preparation have been described previously (Allison and Krummel, 1995, the contents of which are incorporated by reference in their entirety).
Suitable adjuvants further include cytokines. Examples of suitable cytokines are provided in Table 3 (Gabrilovich et al., 1996, the contents of which are incorporated by reference in their entirety). In an aspect, cytokines and immunological adjuvants may be used in vitro, such as for expansion or activation of T cells, or for ex vivo uses. Suitable adjuvants further include CpG oligonucleotides.
Other examples of useful adjuvants include, but are not limited to, a chemically modified CpG (e.g. CpR, Idera), a dsRNA analogue such as Poly(l:C) and a derivative thereof (e.g. AmpliGen®, Hiltonol®, poly(ICLC), poly(IC-R), poly(l:C12U), non-CpG bacterial DNA or RNA as well as a mimetic of the bacterial lipopeptide Pam3Cys-Ser-Ser, such as Pam3Cys-GDPKHPKSF (XS15). See Gouttefangeas and Rammensee 2018 or Rammensee et al. 2019, the contents of which are incorporated herein by reference in their entireties.
A further example of a suitable adjuvant is an immunoactive small molecule or an antibody such as cyclophosphamide, sunitinib, an immune checkpoint inhibitor including ipilimumab, nivolumab, pembrolizumab, atezolizumab, avelumab, durvalumab, and cemiplimab, Bevacizumab®, Celebrex, NCX-4016, sildenafil, tadalafil, vardenafil, sorafenib, temozolomide, temsirolimus, XL-999, CP-547632, pazopanib, VEGF Trap, ZD2171 , AZD2171 , anti-CTLA4, another antibody targeting a key structure of the immune system (e.g. anti-CD40, anti-TGF-beta, anti-TNF-alpha receptor) and SC58175, which may act therapeutically and/or as an adjuvant. The amount and concentration of an adjuvant and/or additive useful in the context of a method or use disclosed herein can readily be determined by the skilled artisan without undue experimentation.
Examples of a suitable adjuvant are anti-CD40, imiquimod, Resiquimod, GM-CSF, cyclophosphamide, sunitinib, bevacizumab, interferon-alpha, a CpG oligonucleotide or a derivative thereof, Poly(l:C) or a derivative thereof, RNA, sildenafil, and a particulate formulation with poly(lactide co-glycolide) (PLG), a virosome, and/or an interleukin. A
respective interleukin may for instance be one or more of IL1 , IL-2, IL-4, IL-7, IL-12, IL- 13, IL-15, IL-18, IL-21 , and IL-23.
Administration modes
The pharmaceutical composition is used for parenteral administration, such as subcutaneous, intradermal, intramuscular or oral administration.
Formulations
The peptides, TCRs, antibodies or cells and optionally other molecules can be dissolved or suspended in a pharmaceutically acceptable carrier, for instance an aqueous carrier, in the pharmaceutical composition.
An antibody or TCR as disclosed herein may be administered to a subject in a pharmaceutically acceptable carrier. Typically, an appropriate amount of a pharmaceutically acceptable salt is used in the formulation to render the formulation isotonic. Examples of the pharmaceutically acceptable carrier include saline, Ringer's solution and dextrose solution. The pH of the solution may for instance be from about 5 to about 8, such as from about 7 to about 7.5. Further carriers include sustained release preparations such as semipermeable matrices of solid hydrophobic polymers containing the antibody, which matrices are in the form of shaped articles, e.g., films, liposomes or microparticles. It will be apparent to those persons skilled in the art that certain carriers may be more preferable depending upon, for instance, the route of administration and concentration of antibody being administered.
In addition, the pharmaceutical composition can contain excipients, such as buffers, binding agents, blasting agents, diluents, flavors, lubricants, emulsifier, vehicle etc. or a combination thereof.
The peptides, TCRs, antibodies or T cells recognizing the peptide of the present disclosure in a complex with an MHC molecule can also be administered together with immune stimulating substances, such as cytokines shown in Table 3. These cytokines may be expressed by the T lymphocyte in soluble or membrane-bound form or may be administered separately.
One or more cytokines, e.g., IL-2, IL-7, IL-12, IL-15, IL-21 , IFN-a, and IFN-|3, may also be used in the activation and/or expansion of T cells, such as T cells recognizing the peptide of the present disclosure in a complex with an MHC molecule.
An extensive listing of excipients that can be used in such a composition, can be, for example, taken from A. Kibbe, Handbook of Pharmaceutical Excipients (Kibbe, 2000). Other examples of suitable pharmaceutical carriers are described in Remington’s Pharmaceutical Sciences (Gennaro, 1997; Banker and Rhodes, 2002, the contents of which are herein incorporated by reference in their entireties). The composition can be used for a prevention, prophylaxis and/or therapy of adenomatous or cancerous diseases. Exemplary formulations can be for example found in EP2112253.
In one embodiment, a pharmaceutical composition, e.g. a vaccine composition, as disclosed herein does not contain any adjuvant.
A formulation as provided herein is one that is suitable for administration of the peptide or other agent by any acceptable route such as oral (enteral), nasal, ophthal, subcutaneous, intradermal, intramuscular, intravenous or transdermal. In some embodiments, the administration is s.c., and in some embodiments i.d. administration by infusion pump.
Vaccines
In some embodiments a peptide or a peptide sequence as provided herein may be included in a vaccine composition. A respective vaccine composition may for example be used for eliciting or for enhancing an anti-tumor immune response. In some embodiments a peptide sequence as provided herein, may be used as a target for the development of a pharmaceutically / immunologically active compound and/or cell.
In some embodiments, the peptide, the salt thereof, the nucleic acid, a vector comprising the nucleic acid or the pharmaceutical composition comprising the peptide, the salt thereof, the nucleic acid or the vector disclosed herein is an immunotherapeutic such as a vaccine.
A peptide as disclosed herein is useful for generating an immune response in a subject by which tumor cells can be destroyed. An immune response in a subject can be induced
by direct administration of the described peptide or suitable precursor substance (e.g. an elongated peptide, protein, or nucleic acid encoding the peptide) to the subject, in some embodiments in combination with an agent enhancing the immunogenicity (i.e. an adjuvant). The immune response originating from such a therapeutic vaccination can be expected to be highly specific against tumor cells because a target peptide as provided herein is not presented on healthy tissues in comparable copy numbers, preventing the risk of undesired autoimmune reactions against normal cells in the subject.
In one aspect, the vaccine includes at least one peptide having the amino acid sequence set forth SEQ ID NO: 1 to SEQ ID NO: 59. In some embodiments, the vaccine includes at least one or more additional peptides, such as two to 50 or two to 25 peptides. In some embodiments two to 20 additional peptides are included. In some embodiments two, three, four, five, six, seven, eight, nine, ten, eleven, twelve, thirteen, fourteen, fifteen, sixteen, seventeen or eighteen peptides may be included. The peptide(s) may be derived from one or more source proteins and may bind to an MHC class I molecule.
A peptide provided herein, for instance when included into a vaccine described herein, may bind to a molecule of any human leukocyte antigen serotype. A peptide provided herein generally binds to at least one selected from the group consisting of HLA-A*01 :01 , HLA-A*02:01 , HLA-A*03:01 , HLA-A*24:02, HLA-B*07:02, HLA-B*08:01 and HLA- B*44:02, plus optionally other HLA allotypes. In some embodiments a peptide binds to HLA-A*01 :01. In some embodiments a peptide binds to HLA-A*02:01 or HLA-A*03:01. In some embodiments a peptide binds to HLA-A*24:02 or HLA-B*07:02. In some embodiments a peptide binds to HLA-B*08:01 or HLA-B*44:02. Due to similarities in binding pattern such as the relevant anchoring positions some peptides bind to more than one allele, such an overlap is most likely, but not limited to, HLA-A*01 binding peptides also binding to HLA-B*15, HLA-A*03 binding peptides also binding to HLA-A*11 , HLA-B*07 binding peptides also binding to HLA-B*35 and HLA-B*51.
If a peptide provided herein is combined with one or more peptides binding to another allele, a higher percentage of any population of individuals can be treated compared with addressing either MHC class I allele alone. For example, with the combination of an HLA- A*02:01 binding peptide with an HLA-A*24:02 binding peptide 60% of patients in any relevant population can be treated, whereas in most populations less than 50% of patients could be addressed by either allele alone.
It is important to realize that the immune response triggered by a vaccine provided herein attacks the cancer in different cell-stages and different stages of development. Furthermore, different cancer associated signaling pathways are attacked. This is an advantage over vaccines that address only one or few targets, which may cause the
tumor to easily adapt to the attack (tumor escape). Furthermore, not all individual tumors express the same pattern of antigens. Therefore, a combination of several tumor- associated peptides ensures that every single tumor bears at least some of the targets. The composition is designed in such a way that each tumor is expected to express several of the antigens and cover several independent pathways necessary for tumor growth and maintenance. Thus, the vaccine can easily be used “off-the-shelf” for a larger patient population. This means that a pre-selection of subjects / patients to be treated with the vaccine can be restricted to MHC typing, does not require any additional biomarker assessments for antigen expression, but it is still ensured that several targets are simultaneously attacked by the induced immune response, which is important for efficacy (Banchereau et al., 2001 ; Walter et al., 2012, the contents of which are incorporated by reference in their entirety). mRNA
In a further aspect there is provided a cancer vaccine comprising a nucleic acid molecule encoding a recombinant protein or peptide. The recombinant protein or peptide contains an amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59. In some embodiments the cancer vaccine further comprises a nucleic acid molecule encoding a recombinant MHC molecule. The recombinant MHC molecule is generally capable of forming a complex with the recombinant peptide that contains the amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
In some embodiments the nucleic acid molecule is an RNA molecule such as an mRNA molecule. In some embodiments the nucleic acid molecule encodes a recombinant peptide that consists of an amino acid sequence selected from selected from the group consisting of SEQ ID NOs: 1 to 59. In some embodiments the nucleic acid molecule includes a 5’UTR. In some embodiments the nucleic acid molecule includes a 3’UTR. In some embodiments the nucleic acid molecule includes both a 5’UTR and a 3’UTR.
In some embodiments the nucleic acid molecule is an RNA that includes a poly(A) tail. A respective poly(A) tail has in some embodiments a length of 100 to 150 adenosine residues, including 120 adenosine residues.
A pharmaceutical composition as provided herein may also include one or more nucleic acid molecules, such as one or more RNA molecules. At least one nucleic acid molecule included in such a composition may include an amino acid sequence encoding a peptide as disclosed herein.
In some embodiments, the composition contains mRNA with a capping degree of 70% or more, such that at least 70% or at least 80% of the mRNA species contain a Cap1
structure. In some embodiments at least 90% of the mRNA species contain a Cap1 structure.
5'-capping of a polynucleotide may be completed concomitantly during the in vitro transcription reaction using one or more of the following chemical RNA cap analogues to generate the 5'-guanosine cap structure according to manufacturer protocols: 3'-0-Me- m7G(5')ppp(5') G [the ARCA cap]; G(5')ppp(5')A; G(5')ppp(5')G; m7G(5')ppp(5')A; m7G(5')ppp(5')G (New England BioLabs, Ipswich, MA). 5'-capping of modified RNA may be completed post-transcriptionally using a Vaccinia Vims Capping Enzyme to generate the “Cap-0” structure: m7G(5')ppp(5')G (New England BioLabs, Ipswich, MA). Cap-1 structure may be generated using both Vaccinia Vims Capping Enzyme and a 2'-0 methyl-transferase to generate: m7G(5')ppp(5')G-2'-O-methyl. The Cap-2 structure may be generated from the Cap-1 structure followed by the 2'-O-methylation of the 5'- antepenultimate nucleotide using a 2'-O-methyl-transferase. The Cap-3 structure may be generated from the Cap-2 structure followed by the 2'-O-methylation of the 5'- preantepenultimate nucleotide using a 2'-O-methyl-transferase. Enzymes may be derived from a recombinant source.
As an illustrative example, in some embodiments the 5’ end of a nucleic acid molecule, e.g. RNA molecule, includes a Cap structure that has the following general formula:
In this formula Ri and R2 are independently hydroxy or methoxy. W~, X’ and Y~ are independently oxygen, sulfur, selenium, or BH3.
Further examples and explanations can be found in WO 2011/015347, which is incorporated herein by reference in its entirety.
Where the nucleic acid molecule is an RNA molecule, the 5’-cap structure is capable of increasing the stability of the RNA, increasing translation efficiency of the RNA, prolonging translation of the RNA, increasing total protein expression of the RNA, and/or increasing the immune response against an antigen or antigen peptide encoded by said RNA when compared to the same RNA without the 5’-cap structure.
The one or more nucleic acids may be complexed or associated with one or more lipids or lipid-based carriers, thereby forming liposomes, lipid nanoparticles (LNP), lipoplexes, and/or nanoliposomes, which may encapsulate the one or more nucleic acids. The LNP may contain any lipid capable of forming a particle to which the one or more nucleic acid molecules are attached, or in which the one or more nucleic acid molecules are encapsulated. An LNP may contain one or more cationic lipids or one or more neutral lipids. An LNP may also contain one or more steroids or steroid analogues. An LNP may also contain one or more polymer conjugated lipids, such as a polyethyleneglycol-(PEG-)
lipid. In one embodiment, the LNP contains one or more cationic lipids, and one or more neutral or pegylated lipids.
Two illustrative examples of a PEG-lipid are 1 ,2 dimyristoyl-sn-glycerol, methoxy-polyethylene-glycol (PEG2000 DMG) or O-Methyl-O'-succinylpolyethylenglykol 2'000 (PEG2000-cDMA). A further example of a PEG-lipid is PEG-DPG (1 ,2-Dipalm itoyl- sn-glycerol, methoxypolyethylene glycol). An illustrative example of a non-cationic lipid is 1 ,2 distearoyl-sn-glycero-3-phosphocholine (DSPC).
In some embodiments one or more nucleic acid molecules may be included in the composition, which include(s) an amino acid sequence encoding a cytokine or a portion thereof.
Combination Product
In some embodiments the combination product includes a peptide or salt thereof as disclosed herein. The combination product may also include a proteinaceous binding molecule as disclosed herein, for instance an antibody, fragment thereof, or a T-cell receptor or fragment thereof. The combination product may also include a nucleic acid or an expression vector as disclosed herein. The combination product may also include a recombinant host cell or an activated T lymphocyte as disclosed herein.
In some embodiments the combination product includes a first active agent that is a peptide or salt thereof as disclosed herein. In some embodiments the combination product includes a first active agent that is a proteinaceous binding molecule as disclosed herein. In some embodiments the combination product includes a first active agent that is a nucleic acid or an expression vector as disclosed herein. In some embodiments the combination product includes a first active agent that is a recombinant host cell or an activated T lymphocyte as disclosed herein.
In some embodiments the combination product includes a second active agent that is an adjuvant. In some embodiments the adjuvant is provided in an immunogenicity enhancing amount. In some embodiments the adjuvant is an interleukin. The interleukin may for example be IL-2, IL-12, IL-15, IL-18, IL-21 or a combination thereof. In one embodiment the interleukin may be IL-2. In one embodiment the interleukin may be IL- 12. In one embodiment the interleukin may be IL-15. In one embodiment the interleukin may be IL-18. In one embodiment the interleukin may be IL-21. In one embodiment the combination product includes a T lymphocyte, such as a recombinant T lymphocyte, as a first active agent and IL-2 as the second active agent.
In some embodiments, the combination product is or includes an individualized product. In some embodiments, the combination product is provided in the form of a kit. The kit may include a first pharmaceutical composition that contains a peptide or salt thereof as disclosed herein. The first pharmaceutical composition may also include a proteinaceous binding molecule as disclosed herein, for instance an antibody, fragment thereof, or a T- cell receptor or fragment thereof. The first pharmaceutical composition may also include a nucleic acid or an expression vector as disclosed herein. The first pharmaceutical composition may also include a recombinant host cell or an activated T lymphocyte as disclosed herein. The kit may include a first pharmaceutical composition that contains an adjuvant as described above.
The peptide may be substantially pure or combined with an immune-stimulating adjuvant (see below) or used in combination with immune-stimulatory cytokines, or be administered with a suitable delivery system, for example liposomes. The peptide may also be conjugated to a suitable carrier such as keyhole limpet hemocyanin (KLH) or mannan (see WO 95/18145 and Longenecker et al., 1993, the contents of both of which are incorporated by reference in their entirety). The peptide may also be tagged, may be a fusion protein, or may be a hybrid molecule.
Kits
Provided is furthermore a kit. A kit provided herein contains a container that contains a component as disclosed herein for the pharmaceutical composition, in solution or in lyophilized form. The component as disclosed herein may be the peptide provided herein. The component as disclosed herein may also be the proteinaceous binding molecule provided herein. The component may also be the nucleic acid or the vector provided herein. The component may also be the recombinant host cell or the activated T lymphocyte provided herein. The component as disclosed herein may be provided in the form of a pharmaceutical composition. The component as disclosed herein may also be the pharmaceutical composition disclosed above.
In one embodiment, the kit provided herein may include a lyophilized formulation provided herein in a suitable container and instructions for its reconstitution and/or use. Suitable containers include for example bottles, vials (e.g. dual chamber vials), syringes (such as dual chamber syringes) and test tubes. The container may be formed from a variety of materials such as glass or plastic. In some embodiments the kit and/or container contain/s instructions on or associated with the container that indicates directions for reconstitution and/or use. For example, the label may indicate that the lyophilized formulation is to be reconstituted to peptide concentrations as described
above. The label may further indicate that the formulation is useful or intended for subcutaneous administration.
The kit may in some embodiments furthermore contain a further container. The further container may contain a diluent for the solution or for the lyophilized formulation. The further container may also contain a reconstituting solution for the lyophilized formulation. The further container may be a second container.
The kit may in some embodiments contain a further container. This further container contains an adjuvant. The further container may be a second container or a third container.
In some embodiments the kit may include at least one further peptide that includes a sequence selected from the group consisting of SEQ ID NO: 1 to SEQ ID NO: 59. The kit may include any further peptide as provided herein. In some embodiments, the kit may furthermore include a buffer and/or a diluent. In some embodiments, the kit may furthermore include a filter and/or a needle. The kit may in some embodiments furthermore contain a syringe.
The container holding the formulation may be a multi-use vial, which allows for repeat administrations (e.g., from 2-6 administrations) of the reconstituted formulation. The kit may further comprise a second container comprising a suitable diluent (e.g., sodium bicarbonate solution).
Upon mixing of the diluent and the lyophilized formulation, the final peptide concentration in the reconstituted formulation is in some embodiments at least 0.15 mg/mL/peptide (=75 pg) and in some embodiments not more than 3 mg/mL/peptide (=1500 pg). The kit may further include other materials desirable from a commercial and user standpoint, including other buffers, diluents, filters, needles, syringes, and package inserts with instructions for use.
A kit provided herein may have a single container that contains the formulation of a pharmaceutical composition provided herein with or without other molecules, pharmaceutical compositions or cells of these other compounds) or may have a distinct container for each component.
In some embodiments, a kit provided herein includes a pharmaceutical composition as provided herein packaged for use in combination with the co-administration of a second compound (such as an adjuvant (e.g. GM-CSF), a chemotherapeutic agent, a natural
product, a hormone or antagonist, an anti-angiogenesis agent or inhibitor, an apoptosisinducing agent or a chelator) or a pharmaceutical composition thereof. In some embodiments a respective composition and second compound may be provided as separate components, typically packed for independent use. The kit may in such embodiments define a combination product.
Molecules included in the kit, such as the peptide or salt thereof, binding molecule, nucleic acid, expression vector, or a cell included in the kit may be pre-complexed. In some embodiments each molecule may be in a separate distinct container prior to administration to a subject. The molecules included in the kit may be provided in one or more liquid solutions, such as an aqueous solution, for example a sterile aqueous solution. The molecules included in the kit may also be provided as solids, which may be converted into liquids by addition of suitable solvents, which are in some embodiments provided in another distinct container.
The container of a therapeutic kit may be a vial, test tube, flask, bottle, syringe, or any other means of enclosing a solid or liquid. Usually, when there is more than one component, the kit will contain a second vial or other container, which allows for separate dosing. The kit may also contain another container for a pharmaceutically acceptable liquid. In some embodiments, a therapeutic kit contains an apparatus (e.g., one or more needles, syringes, eye droppers, pipette, etc.), which enables administration of an agent as provided herein that is a component of the present kit.
A kit may in some embodiments be an example of a combination product, as provided herein. A combination product may be a combination of any peptide, nucleic acid, cell, binding molecule or other agent as provided herein with any other agent provided herein. A combination product may be for administration at independent points in time. The agents, e.g. a binding agent and an adjuvant, included in a combination may be administered concomitantly or subsequently.
Nucleic acids
General
A further aspect provides a nucleic acid molecule (for example a polynucleotide) encoding a peptide of the disclosure. An aspect provides a nucleic acid molecule encoding a TCR or a TCR fragment specific for the peptide of the disclosure. Another aspect provides a nucleic acid molecule encoding an antibody or an antibody fragment specific for the peptide of the disclosure. Typically, the nucleic acid molecule contains a nucleic acid sequence that codes in 5’— >3’ direction for the peptide, TCR or TCR
fragment. The polynucleotide may be, for example, DNA, cDNA, PNA, RNA or combinations thereof, either single- and/or double-stranded, or native or stabilized forms of polynucleotides, such as, for example, polynucleotides with a phosphorothioate backbone and it may or may not contain introns so long as it codes for the peptide. Of course, only peptides that contain naturally occurring amino acid residues joined by naturally occurring peptide bonds are encodable by a polynucleotide.
Examples of nucleic acids include for instance DNA molecules, RNA molecules, analogues of the DNA or RNA generated using nucleotide analogues or using nucleic acid chemistry, locked nucleic acid molecules (LNA), protein nucleic acids molecules (PNA), alkylphosphonate and alkylphosphotriester nucleic acid molecules and tecto-RNA molecules (e.g., Liu, B., et al., J. Am. Chem. Soc. (2004) 126, 4076-4077). LNA has a modified RNA backbone with a methylene bridge between C4' and 02', providing the respective molecule with a higher duplex stability and nuclease resistance.
Alkylphosphonate and alkylphosphotriester nucleic acid molecules can be viewed as a DNA or an RNA molecule, in which phosphate groups of the nucleic acid backbone are neutralized by exchanging the P-OH groups of the phosphate groups in the nucleic acid backbone to an alkyl and to an alkoxy group, respectively. DNA or RNA may be of genomic or synthetic origin and may be single or double stranded. Such nucleic acid can be e.g. mRNA, cRNA, synthetic RNA, genomic DNA, cDNA synthetic DNA, a copolymer of DNA and RNA, oligonucleotides, etc. A respective nucleic acid may furthermore contain non-natural nucleotide analogues and/or be linked to an affinity tag or a label. RNA may be selected from messenger RNA (mRNA), self-amplifying RNA (saRNA), siRNA or miRNA.
Many nucleotide analogues and nucleoside analogues are known and can be used in nucleic acids provided herein and used in the methods disclosed herein. A nucleotide analogue is a nucleotide containing a modification at for instance the base, sugar, or phosphate moieties. A nucleoside analogue is accordingly a nucleoside containing a modification at the base or sugar. As an illustrative example, a substitution of 2'-OH residues of siRNA with 2'F, 2'0-Me or 2'H residues is known to improve the in vivo stability of the respective RNA. An example of a nucleoside analogue is 5-methyluridine, also called ribothymidine, which is the ribonucleoside counterpart to the deoxyribonucleoside thymidine, which lacks a hydroxyl group at the 2' position. A further example of a nucleoside analogue is pseudouridine, a C5-glycoside isomer of uridine, in which the uracil is bonded to ribose via a carbon-carbon instead of a nitrogen-carbon glycosidic bond. An additional methylation at the N1 position of pseudouridine removes the additional hydrogen bond donor and generates N1 -methyl-pseudouridine. Further
examples of nucleoside analogues are nucleosides in which the 4’-oxygen of furanose is replaced by another heteroatom or by carbon. Azanucleosides for instance have a nitrogen atom at this position, and thionucleosides have a sulfur atom instead of oxygen.
Yet further illustrative examples of a nucleoside analogue that may be included in a nucleic acid are N1 -ethylpseudouridine or 2 -thiouridine. Examples of a suitable nucleoside analogue are also 4'-th iourid ine and 5-methylcytosine. Further examples of a suitable nucleoside analogue are 2-thio-1 -methyl-1 -deaza-pseudouridine or 2-thio-1 - methyl-pseudouridine. Suitable examples of a nucleoside analogue are also 2-thio-5- aza-uridine or 2-thio-dihydropseudouridine. Examples of a suitable nucleoside analogue are also 2-thio-dihydrouridine or 2-thio-pseudouridine. Further examples of a suitable nucleoside analogue are 4-methoxy-2-thio-pseudouridine or 4-methoxy-pseudouridine. Yet further examples of a suitable nucleoside analogue are 4-thio-1 -methyl- pseudouridine or 4-thio-pseudouridine. Examples of a suitable nucleoside analogue are also 5-aza-uridine or dihydropseudouridine. Examples of a suitable nucleoside analogue are also 5-methyluridine or 5-methoxyuridine. Examples of a suitable nucleoside analogue are also 2-methoxyuridine or 2'-O-methyl uridine. Further examples of a suitable nucleoside analogue are 2-methoxy-4-thio-uridine or 1 -methyl-3-(3-amino-3- carboxypropyl)pseudouridine.
Modifications at the base moiety may be a natural or a synthetic modification of A, C, G, and T/U, a different purine or pyrimidine base, such as uracil-5-yl, hypoxanthin-9-yl, and 2-aminoadenin-9-yl, as well as a non-purine or a non-pyrimidine nucleotide base. Pseudouridine can for example be formed using a pseudouridine synthase enzyme. Other nucleotide analogues serve as universal bases. Examples of universal bases include 3-nitropyrrole and 5-nitroindole. Universal bases are able to form a base pair with any other base. Base modifications often can be combined with for example a sugar modification, such as for instance 2'-O-methoxyethyl, e.g. to achieve unique properties such as increased duplex stability.
In some embodiments, a nucleic acid contains two or more encoding repeats (“concatemer”), separated by short nucleotide stretches (“spacers”). In some embodiments, the nucleic acid, for example an mRNA molecule, includes a 5' untranslated region (UTR) and/or a 3' UTR. A 3’-UTR in some embodiments contains or consists of a nucleic acid sequence derived from a 3’-UTR of a gene selected from PSMB3, ALB7, alpha-globin, CASP1 , COX6B1 , GNAS, NDUFA1 , and RPS9 or from a homolog, a fragment, or a variant of any one of these genes. In some embodiments, a 5’-UTR contains or consists of a nucleic acid sequence derived from a 5’-UTR of a gene selected from HSD17B4, RPL32, ASAH1 , ATP5A1 , MP68, NDUFA4, NOSIP, RPL31 ,
SLC7A3, TUBB4B, and UBQLN2 or from a homolog, a fragment, or variant of any one of these genes. In some embodiments, a 5’-UTR and/or a heterologous 3’ UTR is selected from UTR design a-1 (HSD17B4/PSMB3) or a-3 (SLC7A3/PSMB3). In some embodiments, a 5’-UTR and/or a heterologous 3’ UTR is selected from UTR design e-2 (RPL31/RPS9), or i-3 (-/muag).
In some embodiments the nucleic acid contains a coding sequence which is codon- optimized and/or in which the G/C content is increased and the uridine content is decreased compared to the respective naturally occurring coding sequence. The codonoptimization and/or the increase in the G/C content can be carried out in such a way that it does not change the sequence of the encoded amino acid sequence. Generating a G/C content optimized nucleic acid sequence (RNA or DNA) may for instance be carried out using a method according to WO 2002/098443, the disclosure of which is incorporated herein in its entirety.
Where a nucleic acid is modified, this may include or consist of adapting the codons in a coding sequence to human codon usage (herein referred to as “human codon usage adapted coding sequence”).
Codons encoding the same amino acid occur at different frequencies in humans. Accordingly, the coding sequence of the nucleic acid may be modified such that the frequency of the codons encoding the same amino acid corresponds to the naturally occurring frequency of that codon according to the human codon usage. As an illustrative example, in the case of the amino acid alanine, the wild type or reference coding sequence may be adapted in a way that the codon “GCC” is used with a frequency of 0.40, the codon “GOT” is used with a frequency of 0.28, the codon “GCA” is used with a frequency of 0.22 and the codon “GCG” is used with a frequency of 0.10. A corresponding procedure (as exemplified for alanine) may be applied to any, including each, amino acid encoded by the coding sequence of a nucleic acid. Where all codons for each amino acid are adapted in this manner, a sequence is obtained that is adapted to human codon usage.
Vectors
A still further aspect provides an expression vector comprising a nucleic acid sequence encoding a peptide as disclosed herein. Provided is also an expression vector comprising a nucleic acid sequence encoding a proteinaceous binding molecule as disclosed herein. The nucleic acid sequence may in some embodiments encode an antibody or an antibody fragment as disclosed herein. The nucleic acid sequence may in some embodiments encode a TCR or a TCR fragment as disclosed herein.
The nucleic acid molecule or polynucleotide used for active vaccination may be substantially pure or contained in a suitable vector or delivery system. The nucleic acid may be DNA, cDNA, PNA, RNA or a combination thereof. Methods for designing and introducing such a nucleic acid are well known in the art. An overview is provided by Teufel et al. (Teufel et al., 2005, the contents of which are incorporated by reference in their entirety). Polynucleotide and nucleic acid vaccines are easy to prepare, but the mode of action of these vectors in inducing an immune response is not fully understood. Suitable vectors and delivery systems include viral DNA and/or RNA, such as systems based on adenovirus, vaccinia virus, retroviruses, herpes virus, adeno-associated virus or hybrids containing elements of more than one virus. Non-viral delivery systems include cationic lipids and cationic polymers and are well known in the art of DNA delivery. Physical delivery, such as via a “gene-gun” may also be used. The peptide or peptides encoded by the nucleic acid may be a fusion protein, for example with an epitope that stimulates T cells for the respective opposite CDR as noted above.
A variety of methods have been developed to link polynucleotides, especially DNA, to vectors for example via complementary cohesive termini. For instance, complementary homopolymer tracts can be added to the DNA segment to be inserted to the vector DNA. The vector and DNA segment are then joined by hydrogen bonding between the complementary homopolymeric tails to form recombinant DNA molecules.
Synthetic linkers containing one or more restriction sites provide an alternative method of joining the DNA segment to vectors. Synthetic linkers containing a variety of restriction endonuclease sites are commercially available from a number of sources including International Biotechnologies Inc. New Haven, CN, USA.
A desirable method of modifying the DNA encoding a peptide as disclosed herein, or encoding the respective TCR or antibody, or TCR or antibody fragment, employs the polymerase chain reaction as disclosed by Saiki et al. (Saiki et al., 1988, the contents of which are incorporated by reference in their entirety). This method may be used for introducing the DNA into a suitable vector, for example by engineering in suitable restriction sites, or it may be used to modify the DNA in other useful ways as is known in the art. Illustrative examples of viral vectors are pox- or adenovirus vectors.
In some embodiments the DNA (or in the case of retroviral vectors, RNA) may then be expressed in a suitable host to produce a polypeptide comprising the peptide as disclosed herein. Thus, the DNA encoding a peptide of the disclosure may be used in accordance with known techniques, appropriately modified in view of the teachings contained herein, to construct an expression vector, which is then used to transform an appropriate host cell for the expression and production of the polypeptide of the invention.
Such techniques include those disclosed for example in US 4,440,859, 4,530,901 , 4,582,800, 4,677,063, 4,678,751 , 4,704,362, 4,710,463, 4,757,006, 4,766,075, and 4,810,648, the contents each of which are incorporated by reference in their entirety.
The DNA (or in the case of retroviral vectors, RNA) encoding a polypeptide that encodes the peptide, TCR, TCR fragment, antibody or antibody fragment as disclosed herein may be joined to a wide variety of other DNA sequences for introduction into an appropriate host. The companion DNA will depend on the nature of the host, the manner of the introduction of the DNA into the host, and whether episomal maintenance or integration is desired.
Generally, the DNA is inserted into an expression vector, such as a plasmid, in proper orientation and correct reading frame for expression. If necessary, the DNA may be linked to the appropriate transcriptional and translational regulatory control nucleic acid sequences recognized by the desired host, although such controls are generally available in the expression vector. The vector is then introduced into the host through standard techniques. Generally, not all of the hosts will be transformed by the vector. Therefore, it will be necessary to select for transformed host cells. One selection technique involves incorporating a DNA sequence into the expression vector, with any necessary control elements, that codes for a selectable trait in the transformed cell, such as antibiotic resistance.
Alternatively, the gene for such selectable trait can be on another vector, which is used to co-transform the desired host cell.
Host cells that have been transformed by a recombinant DNA as disclosed herein are then cultured for a sufficient time and under appropriate conditions known to those skilled in the art in view of the teachings disclosed herein to permit the expression of the polypeptide or peptide, which can then be recovered.
Many expression systems are known, including bacteria (for example E. coli and Bacillus subtilis), yeasts (for example Saccharomyces cerevisiae), filamentous fungi (for example Aspergillus spec.), plant cells, animal cells and insect cells. In some embodiments, the system can be mammalian cells such as CHO cells available from the ATCC Cell Biology Collection.
A typical mammalian cell vector plasmid for constitutive expression comprises the CMV or SV40 promoter with a suitable poly A tail and a resistance marker, such as neomycin. One example is pSVL available from Pharmacia, Piscataway, NJ, USA. An example of
an inducible mammalian expression vector is pMSG, also available from Pharmacia. Useful yeast plasmid vectors are pRS403-406 and pRS413-416 and are generally available from Stratagene Cloning Systems, La Jolla, CA 92037, USA. Plasmids pRS403, pRS404, pRS405 and pRS406 are Yeast Integrating plasmids (Yips) and incorporate the yeast selectable markers HIS3, TRP1 , LEU2 and URA3. Plasmids pRS413-416 are Yeast Centromere plasmids (Ycps). CMV promoter-based vectors (for example from Sigma-Aldrich) provide transient or stable expression, cytoplasmic expression or secretion, and N-terminal or C-terminal tagging in various combinations of FLAG, 3xFLAG, c-myc or MAT. These fusion proteins allow for detection, purification and analysis of recombinant protein. Dual-tagged fusions provide flexibility in detection.
The strong human cytomegalovirus (CMV) promoter regulatory region drives constitutive protein expression levels as high as 1 mg/L in COS cells. For less potent cell lines, protein levels are typically ~0.1 mg/L. The presence of the SV40 replication origin will result in high levels of DNA replication in SV40 replication permissive COS cells. CMV vectors, for example, can contain the pMB1 (derivative of pBR322) origin for replication in bacterial cells, the b-lactamase gene for ampicillin resistance selection in bacteria, hGH polyA, and the f1 origin. Vectors containing the pre-pro-trypsin leader (PPT) sequence can direct the secretion of FLAG fusion proteins into the culture medium for purification using ANTI-FLAG antibodies, resins, and plates. Other vectors and expression systems are well known in the art for use with a variety of host cells.
In another embodiment two or more peptides as disclosed herein are encoded and thus expressed in a successive order (similar to “beads on a string” constructs). In doing so, the peptides may be linked or fused together by stretches of linker amino acids, such as for example LLLLLL, or may be linked without any additional peptide(s) between them. These constructs can also be used for cancer therapy and may induce immune responses both involving MHC I and MHC II.
Host cells
Provided is also a host cell transformed with a polynucleotide vector construct as disclosed herein. The host cell can be either prokaryotic or eukaryotic. Bacterial cells may be prokaryotic host cells in some circumstances and typically are a strain of E. coli such as, for example, the E. coli strains DH5 available from Bethesda Research Laboratories Inc., Bethesda, MD, USA, and RR1 available from the American Type Culture Collection (ATCC) of Rockville, MD, USA (No ATCC 31343). Examples of suitable eukaryotic host cells include yeast, insect and mammalian cells. Further examples of suitable eukaryotic host cells are vertebrate cells such as those from a mouse, rat, monkey or human fibroblastic and colon cell lines. Yeast host cells include
YPH499, YPH500 and YPH501 , which are generally available from Stratagene Cloning Systems, La Jolla, CA 92037, USA. In some embodiments mammalian host cells include Chinese hamster ovary (CHO) cells available from the ATCC as CCL61 , NIH Swiss mouse embryo cells NIH/3T3 available from the ATCC as CRL 1658, monkey kidney- derived COS-1 cells available from the ATCC as CRL 1650 and 293 cells which are human embryonic kidney cells. Examples of suitable insect cells are Sf9 cells which can be transfected with baculovirus expression vectors. An overview regarding the choice of suitable host cells for expression can be found in the literature (Baibas and Lorence, 2004).
Transformation of appropriate cell hosts with a DNA construct of the present disclosure is accomplished by well-known methods that typically depend on the type of vector used. With regard to transformation of prokaryotic host cells, see for example Cohen et al. or Green and Sambrook (Cohen et al., 1972; Green and Sambrook, 2012). Transformation of yeast cells is described in Sherman et al. (Sherman et al. , 1986). The method of Beggs (Beggs, 1978) is also useful. With regard to vertebrate cells, reagents useful in transfecting such cells, for example calcium phosphate and DEAE-dextran or liposome formulations, are available from Stratagene Cloning Systems, or Life Technologies Inc., Gaithersburg, MD 20877, USA. Electroporation is also useful for transforming and/or transfecting cells and is well known in the art for transforming yeast cell, bacterial cells, insect cells and vertebrate cells. The contents of each of these references is herein incorporated by reference in their entirety.
Successfully transformed cells, i.e., cells that contain a DNA construct as disclosed herein, can be identified by well-known techniques such as PCR. The presence of the protein in the supernatant can furthermore be detected using antibodies.
It will be appreciated that certain host cells disclosed herein are useful in the preparation of a peptide disclosed herein, for example bacterial, yeast and insect cells. Such host cells may also be useful in expressing a TCR or an antibody. Other host cells may be useful in certain therapeutic methods. For example, antigen-presenting cells, such as dendritic cells, may be used to express a peptide disclosed herein such that they may be loaded into appropriate MHC molecules. Thus, provided is also a host cell that includes a nucleic acid molecule or an expression vector as disclosed herein.
In one embodiment the host cell is an antigen-presenting cell, in particular a dendritic cell or antigen-presenting cell. APCs loaded with a recombinant fusion protein containing prostatic acid phosphatase (PAP) were approved by the U.S. Food and Drug Administration (FDA) on April 29, 2010, to treat asymptomatic or minimally symptomatic
metastatic HRPC (Sipuleucel-T) (Rini et al., 2006; Small et al., 2006, the contents of which are incorporated by reference in their entirety).
Provided is also a method of producing a peptide, the method comprising culturing a host cell and isolating the peptide from the host cell or its culture medium.
Binding molecule
As noted above, a binding molecule is generally a proteinaceous binding molecule or a nucleic acid molecule.
In one aspect there is provided a proteinaceous binding molecule that specifically binds to a peptide as disclosed herein or to an MHC class I or ii molecule being complexed with a peptide as disclosed herein.
A respective proteinaceous binding molecule is in some embodiments an antibody or a fragment thereof. A further example of a suitable proteinaceous binding molecule is a TCR or a fragment thereof. Yet another example of a suitable proteinaceous binding molecule is a chimeric antigen receptor (CAR). A CAR is a recombinant or synthetic molecule which combines antibody-based specificity for a certain antigen with a T cell receptor-activating intracellular domain thereby defining a chimeric protein that exhibits cellular immune activity to the respective antigen. In some embodiments the proteinaceous binding molecule is not a chimeric antigen receptor.
In some embodiments the proteinaceous binding molecule is an antibody fragment. Further examples of an antibody fragment are a diabody or a domain antibody (Holt, L.J., et al., Trends Biotechnol. (2003), 21 , 11 , 484-490).
An example of a proteinaceous binding molecule with antibody-like functions is a mutein based on a polypeptide of the lipocalin family (WO 03/029462, Beste et al., Proc. Natl. Acad. Sci. USA (1999) 96, 1898-1903). Lipocalins, such as the bilin binding protein, the human neutrophil gelatinase-associated lipocalin, human Apolipoprotein D or glycodelin, possess natural ligand-binding sites that can be modified so that they bind to selected small protein regions known as haptens.
A further example of a proteinaceous binding molecule is a so-called glubody (see e.g. international patent application WO 96/23879 or Napolitano, E.W., et al., Chemistry & Biology (1996) 3, 5, 359-367). Examples of further proteinaceous binding molecules are a protein based on the ankyrin scaffold (Mosavi, L.K., et al., Protein Science (2004) 13, 6, 1435-1448) or a protein based on the crystalline scaffold (e.g. international patent application WO 01/04144).
Yet further examples of a proteinaceous binding molecule are AdNectins or tetranectins. A further example of a proteinaceous binding molecule is an avimer. Avimers contain so called A-domains that occur as strings of multiple domains in several cell surface receptors (Silverman, J., et al., Nature Biotechnology (2005) 23, 1556-1561 ). Adnectins, derived from a domain of human fibronectin, contain three loops that can be engineered for antibody -like binding to targets (Gill, D.S. & Damle, N.K., Current Opinion in Biotechnology (2006) 17, 653-658). Tetranectins, derived from the respective human homotrimeric protein, likewise contain loop regions in a C-type lectin domain that can be engineered for desired binding (ibid.). Peptoids, which can act as protein ligands, are oligo(N-alkyl) glycines that differ from peptides in that the side chain is connected to the amide nitrogen rather than the a carbon atom. Peptoids are typically resistant to proteases and other modifying enzymes and can have a much higher cell permeability than peptides (see e.g. Kwon, Y.-U., and Kodadek, T., J. Am. Chem. Soc. (2007) 129, 1508-1509).
A further illustrative example of a proteinaceous binding molecule is a soluble bispecific binding molecule, which contains a peptide/MHC recognition domain and a T cellengaging domain. The T cell-engaging domain is typically specific for the CD3 receptor. The peptide/MHC recognition domain may be a TCR domain or an antibody domain. Examples are the so-called immune-mobilizing monoclonal T cell receptors against cancer (ImmTAC) or the bispecific T cell engagers (BiTEs®).
In some embodiments there is provided an antibody, a TCR or another proteinaceous binding molecule. The antibody, TCR or other proteinaceous binding molecule may have a binding specificity to a peptide that can be regarded as an epitope.
For further information on a polypeptide binding molecule see for example the background section of WO 2014/071978A1 and the references cited therein, the contents of which are incorporated by reference in their entirety.
Antibody
A peptide disclosed herein can be used to generate and develop a specific antibody or antibody fragment against the peptide when in a complex with MHC (Dao et al., Sci Transl Med. (2013) 5, 176, 176ra33; Dao et al., Nat. Biotechnol. (2015) 33, 10, 1079-1086). The respective binding molecule can be used for therapy, targeting toxins or radioactive substances to the diseased tissue. Another use of such antibody or antibody fragment can be targeting radionuclides to the diseased tissue for imaging purposes such as PET. This use can help to detect small metastases or to determine the size and precise localization of diseased tissues.
TCR-mimic antibody
In one embodiment, the antibody is a TCR-like antibody, also termed T-cell receptor mimic (TCRm) antibody. A TCR-like antibody can be produced with the antibody generation methods described herein, and recognizes a peptide/MHC complex, for example a peptide/MHC complex on the surface of a tumor cell, in the same way as a TCR (see also He et al., 2019). In the context of the present disclosure, a TCR-like antibody is thus an antibody which binds to a peptide as disclosed herein complexed with an MHC. Typically, a TCR-like antibody is a monoclonal antibody. Such MHC-restricted monoclonal antibodies that bind to a peptide when in an MHC complex can be generated using well-established techniques, including the generation of hybridoma and use of phage-display. In typical embodiments, a TCR-like antibody is a soluble antibody. TCR- like antibodies are known to have effector functions including direct killing of cancer cells, complement-dependent cytotoxicity (CDC), antibody-dependent cellular cytotoxicity (ADCC) and antibody-dependent phagocytosis. A TCR-like antibody typically has a higher affinity than a corresponding TCR. In order to increase the chances of obtaining antibodies that are capable of recognizing the composite peptide-major histocompatibility complex surface, Yang et al. (Yang, X, et al., Nature Biotechnology (2023) 41 , 932-943) have proposed re-engineering existing MHC-restricted TCR-mimic antibodies.
TCR mimic antibodies binding to MHC restricted peptide derived from PRAME are for example disclosed in Chang et al. (Chang et al., J Clin Invest (2017) 127, 7: 2705-2718), the content of which is incorporated herein by reference in its entirety. See, also, US 2018/0148503 (T cell receptor-like antibodies specific for a PRAME peptide) (Eureka Therapeutics Inc), the content of which is incorporated herein by reference in its entirety.
A suitable antibody may in some embodiments also be a multispecific antibody that includes several antibody fragments.
An antibody, a TCR or a proteinaceous binding molecule with antibody -like functions may be PEGylated or hyperglycosylated if desired.
In some embodiments an antibody, a TCR or a proteinaceous binding molecule as disclosed herein specifically binds to an MHC class I or II molecule being complexed with an MHC restricted antigen. The respective antibody may for example be a polyclonal antibody or a monoclonal antibody. The respective antibody may also be a bispecific antibody or a chimeric antibody. The respective antibody may also be a humanized antibody and/or an antibody fragment.
One of skill in the art will realize that the generation of two or more different sets of monoclonal or polyclonal antibodies maximizes the likelihood of obtaining an antibody
with the specificity and affinity required for its intended use (e.g., ELISA, immunohistochemistry, in vivo imaging, immunotoxin therapy). The antibodies are tested for their desired activity by known methods, in accordance with the purpose for which the antibodies are to be used (e.g., ELISA, immunohistochemistry, immunotherapy, etc. (Greenfield, 2014, the contents of which are incorporated by reference in their entirety)). For example, the antibodies may be tested in ELISA assays or, Western blots, immunohistochemical staining of formalin-fixed cancers or frozen tissue sections. After their initial in vitro characterization, antibodies intended for therapeutic or in vivo diagnostic use are tested according to known clinical testing methods.
Bispecific Antibody
In an aspect, a bispecific antibody includes an antibody capable of selectively binding two or more epitopes. Bispecific antibodies may be manufactured in a variety of ways (Holliger & Winter, 1993, the contents of which are incorporated by reference in their entirety), for instance, prepared chemically or from hybrid hybridomas, or may be any of the bispecific antibody fragments mentioned above. scFv dimers or diabodies may be used, rather than whole antibodies. Diabodies and scFv can be constructed without an Fc region, using only variable domains (usually including the variable domain components from both light and heavy chains of the source antibody), potentially reducing the effects of anti-idiotypic reaction. Other forms of bispecific antibodies include the single chain “Janusins” described by Traunecker and colleagues (Traunecker et al., 1991 , the content of which is incorporated by reference in its entirety).
Bispecific antibodies generally include two different binding domains, with each binding domain specifically binding a different epitope either on two different antigens or on the same antigen. If a bispecific antibody is capable of selectively binding two different epitopes (a first epitope and a second epitope), the affinity of the first binding for the first epitope will generally be at least one to two or three or four orders of magnitude lower than the affinity of the first binding domain for the second epitope, and vice versa. The epitopes recognized by the bispecific antibody can be on the same or a different target (e.g., on the same or a different protein). Bispecific antibodies can be made for example by combining binding domains that recognize different epitopes of the same antigen.
Some example bispecific antibodies have two heavy chains (each having three heavy chain CDRs, followed by (N-terminal to C-terminal) a CH1 domain, a hinge, a CH2 domain, and a CH3 domain), and two antibody light chains that confer antigen-binding specificity through association with each heavy chain. However, additional architectures are envisioned, including bispecific antibodies in which the light chain(s) associate with each heavy chain but do not (or minimally) contribute to antigen-binding specificity, or
that can bind one or more of the epitopes bound by the heavy chain antigen-binding regions, or that can associate with each heavy chain and enable binding of one or both of the heavy chains to one or both epitopes.
In particular embodiments, a bispecific antibody can include an antibody arm combined with an arm that binds to a triggering molecule on a leukocyte, such as a T cell receptor molecule (for example, CD3).
Bispecific antibodies can be prepared as full-length antibodies or antibody fragments (for example, F(ab’)2 bispecific antibodies). See e.g. WO 1996/016673; U.S. Patent No. 5,837,234; WO 1998/002463; U.S. Patent No. 5,821 ,337, the contents of which are incorporated by reference in their entirety.
A bispecific antibody can have an extended half-life. In particular embodiments, half-life extension of a bispecific antibody can be achieved by: increasing the hydrodynamic volume of the antibody by coupling to inert polymers such as polyethylene glycol or other mimetic hydrophilic polymers; fusion or conjugation to large disordered peptides; or fusing or coupling the antibody to a ligand. These alterations and a number of others are described elsewhere (U.S. Patent No. 7,083,784, U.S. Patent No. 7,670,600, U.S. Patent Application Publication No. 2010/0234575, and Zwolak et al.,2017, the contents of which are incorporated by reference in their entirety). Bispecific antibodies with extended halflives are described in for example U.S. Patent No. 8,921 ,528 and U.S. Patent Application Publication No. 2014/0308285, the contents of which are incorporated by reference in their entirety.
Methods for making bispecific antibodies are known in the art. Production of full-length bispecific antibodies is based on the co-expression of two antibody heavy chain-light chain pairs, where the two chains have different specificities. See e.g., WO 1993/008829 and Traunecker et al., 1991 , the contents of which are incorporated by reference in their entirety.
Polyclonal Antibody
Methods for making polyclonal antibodies are known in the art. Polyclonal antibodies are heterogeneous populations of antibody molecules derived from the sera of animals immunized with an antigen. Polyclonal antibodies which selectively bind a peptide according to SEQ ID NO: 1 to SEQ ID NO: 59, or a fragment thereof may be made by methods well-known in the art (see, e.g., Howard & Kaser, 2007, the contents of which are incorporated by reference in their entirety).
Chimeric Antibody
Chimeric antibodies are molecules different portions of which are derived from different animal species, such as those having variable region derived from a murine antibody and a human antibody constant region, which are primarily used to reduce immunogenicity in application and to increase yields in production for example where murine monoclonal antibodies have higher yields from hybridomas but higher immunogenicity in humans, such that human murine chimeric monoclonal antibodies are used. Chimeric antibodies and methods for their production are known in the art (Cabilly et al., 1984; Morrison et al., 1984; Boulianne et al., 1984; European Patent Application 173494 (1986); WO 86/01533 (1986); European Patent Application 184187 (1986); Sahagan et al., 1986; Liu et al., 1987; Sun et al., 1987; Better et al., 1988; Harlow & Lane, 1998; U.S. Patent No. 5,624,659, the contents of which are incorporated by reference in their entirety).
Antibody Fragments
An antibody fragment, whether attached to other sequences or not, can also include one or more insertions, deletions, substitutions, or other selected modifications of particular regions or specific amino acids residues, provided the activity of the fragment is not significantly altered or impaired compared to the non-modified antibody or antibody fragment. Such modification can provide for some additional property, such as to remove/add an amino acid capable of disulfide bonding, to increase its bio-longevity, to alter its secretory characteristics, etc. In any case, the antibody fragment must possess a bioactive property, such as binding activity, regulation of binding at the binding domain, etc. Functional or active regions of the antibody may be identified by mutagenesis of a specific region of the protein, followed by expression and testing of the expressed peptide. Such methods are readily apparent to a skilled practitioner in the art and can include site-specific mutagenesis of the nucleic acid encoding the antibody fragment.
Antibody production methods
An antibody provided herein may in some embodiments be a humanized antibody or a human antibody. Humanized forms of non-human (e.g., murine) antibodies are chimeric antibodies, antibody chains or fragments thereof (such as Fv, Fab, Fab' or other antigenbinding subsequences of antibodies) which contain minimal sequence derived from a non-human antibody. Humanized antibodies include human antibody (recipient antibody) in which residues from a complementary determining region (CDR) of the recipient are replaced by residues from a CDR of a non-human species (donor antibody) such as mouse, rat or rabbit having the desired specificity, affinity and capacity. In some instances, Fv framework (FR) residues of the human antibody are replaced by corresponding non-human residues. Humanized antibodies may also comprise residues which are found neither in the recipient antibody nor in the imported CDR or framework
sequences. In general, the humanized antibody will comprise substantially all of at least one, and typically two, variable domains, in which all or substantially all of the CDR regions correspond to those of a non-human antibody and all or substantially all of the FR regions are those of a human antibody consensus sequence. The humanized antibody typically contains a portion of an antibody constant region (Fc), for example that of a human antibody.
In addition to entire antibodies (or their recombinant counterparts), antibody fragments comprising the epitope binding site (e.g., Fab, F(ab’)2, or other fragments) may be synthesized. “Fragments” or minimal antibodies may be designed utilizing recombinant antibody techniques. For instance, “Fv” antibodies for use in the present invention may be produced by synthesizing a fused variable light chain region and a variable heavy chain region. Combinations of antibodies are also of interest, e.g., diabodies, which comprise two distinct Fv specificities. Antigen-binding fragments of antibody include, but are not limited to, SMIPs (small molecule immunopharmaceuticals), camelbodies, nanobodies, and IgNAR.
Respective methods for producing such antibodies and single chain MHC class I complexes, as well as other tools for the production of these antibodies are disclosed in WO 03/068201 , WO 2004/084798, WO 01/72768, WO 03/070752, and in publications (Cohen et al., 2003a; Cohen et al., 2003b; Denkberg et al., 2003, which for the purposes of the present invention are incorporated by reference in their entireties).
In some embodiments, the antibody is binding with a binding affinity of <100 nM, or <50 nM to the complex, which is also regarded as “specific” in the context of the present invention. In some embodiments, the antibody is binding with a binding affinity of <10 nM, such as <1 nM to the complex. In some embodiments, the antibody is binding with a binding affinity of <0.1 nM, for instance <0.01 nM, to the complex.
Methods of producing an antibody
The present disclosure also relates to a method of producing an antibody specifically binding to a peptide as disclosed herein, as well as an antibody specifically binding to an MHC class I and/or class II molecule complexed with a peptide as provided herein. The antibody may in some embodiments be recombinant.
A respective method may involve immunizing genetically engineered non-human mammal containing cells expressing the MHC class I and/or class II molecule with a soluble form of an MHC class I or II molecule complexed with a peptide as provided
herein. The method also involves isolating mRNA molecules from antibody producing cells of the non-human mammal.
The method may in some embodiments also include producing a phage display library displaying protein molecules encoded by the above-named mRNA molecules. The method may furthermore include isolating a phage from the phage display library. The phage may display the antibody specifically binding to said MHC class I or ii molecule being complexed with the respective HLA restricted antigen.
In general, techniques for preparing monoclonal antibodies and hybridomas are well known in the art. Methods for the production of a hybridoma cell are disclosed in Kohler & Milstein (1975). Any animal such as a goat, a mouse or a rabbit that is known to produce antibodies can be immunized with the selected peptide, e.g. a peptide as provided herein. Methods for immunization are well known in the art. Such methods include subcutaneous or intraperitoneal injection of the peptide to elicit lymphocytes that produce or are capable of producing antibodies that will specifically bind to the immunizing agent. Typically, the immunized mammal is bled and the serum from each blood sample is analysed for particular antibodies using appropriate screening assays. Alternatively, the lymphocytes may be immunized in vitro.
For a monoclonal antibody, lymphocytes, typically splenocytes, from the immunized animals are removed, fused with an immortal cell line, typically myeloma cells, such as SP2/0-Agl4 myeloma cells, and allowed to become monoclonal antibody producing hybridoma cells. Typically, the immortal cell line such as a myeloma cell line is derived from the same mammalian species as the lymphocytes. Illustrative immortal cell lines are mouse myeloma cell lines that are sensitive to culture medium containing hypoxanthine, aminopterin and thymidine ("HAT medium"). As an illustrative example, HAT-sensitive mouse myeloma cells are fused to mouse splenocytes using 1500 molecular weight polyethylene glycol ("PEG 1500"). Hybridoma cells resulting from the fusion may then be selected using HAT medium, which kills unfused and unproductively fused myeloma cells.
A method of producing the antibody may involve producing a phage display library displaying protein molecules encoded by the mRNA molecules. In such embodiments the method furthermore includes isolating a phage from the phage display library. The phage displays the above antibody specifically binding to the MHC class I molecule complexed with a peptide as provided herein.
In vitro antibody libraries are, among others, disclosed in US6300064 by MorphoSys and US6248516 by MRC/Scripps/Stratagene. Phage Display techniques are for example disclosed in US5223409 by Dyax.
One further method to generate antibodies is by means of a transgenic mammal platforms, as for example described in EP1480515A2 by TaconicArtemis.
In some embodiments, a method of producing an antibody as disclosed herein may further include humanizing the antibody. Methods for humanizing non-human antibodies are well known in the art. Humanized forms of the antibodies of the present invention may be generated using one of the procedures known in the art such as chimerization or CDR grafting.
Generally, a humanized antibody has one or more amino acid residues introduced into it from a source which is non-human. These non-human amino acid residues are often referred to as "import" residues, which are typically taken from an "import" variable domain. Humanization can be essentially performed by substituting rodent CDRs or CDR sequences for the corresponding sequences of a human antibody. Accordingly, such "humanized" antibodies are chimeric antibodies (US 4,816,567, the contents of which are incorporated by reference in their entirety), wherein substantially less than an intact human variable domain has been substituted by the corresponding sequence from a non- human species. In practice, humanized antibodies are typically human antibodies in which some CDR residues and possibly some FR residues are substituted by residues from analogous sites in rodent antibodies.
US6331415 by Genentech describes the production of chimeric antibodies, while US6548640 by Medical Research Council describes CDR grafting techniques and US5859205 by Celltech describes the production of humanised antibodies.
In some embodiments, a method of producing an antibody may further include conjugating the antibody with a toxin. In some embodiments, a method of producing an antibody may further include conjugating the antibody with an immune stimulating domain.
Methods for the production and/or selection of fully human mAbs are known in the art. These can involve the use of a transgenic animal which is immunized with the respective protein or peptide, or the use of a suitable display technique, like yeast display, phage display, B-cell display or ribosome display, where antibodies from a library are screened against human iRhom2 in a stationary phase.
The monoclonal antibody may also be made by a recombinant DNA method, such as those described in US 4,816,567. DNA encoding a monoclonal antibody provided herein can be readily isolated and sequenced using conventional procedures (e.g., by using oligonucleotide probes that are capable of binding specifically to genes encoding the heavy and light chains of murine antibodies).
In vitro methods are also suitable for preparing monovalent antibodies. Digestion of antibodies to produce fragments thereof, particularly Fab fragments, can be accomplished using routine techniques known in the art. For instance, digestion can be performed using papain. Examples of papain digestion are described in WO 94/29348 and US 4,342,566, the contents of which are incorporated by reference in their entirety. Papain digestion of antibodies typically produces two identical antigen binding fragments, called Fab fragments, each with a single antigen binding site, and a residual Fc fragment. Pepsin treatment yields a F(ab')2 fragment and a pFc' fragment.
Transgenic animals (e.g., mice) that are capable, upon immunization, of producing a full repertoire of human antibodies in the absence of endogenous antibody production can be employed. For example, it has been described that the homozygous deletion of the antibody heavy chain joining region gene in chimeric and germ-line mutant mice results in complete inhibition of endogenous antibody production. Transfer of the human germline antibody gene array in such germ-line mutant mice will result in the production of human antibodies upon antigen challenge. Human antibodies can also be produced in phage display libraries.
An antibody of the present disclosure can be obtained through phage display, or ribosome display, or yeast display, or bacteria display, or Baculovirus display, or mammal cell display, or mRNA display. These methods are all conventional techniques in the art, the specific operations thereof can be seen in corresponding textbooks or operation manuals (Mondon et al., 2008; the content of which is hereby incorporated by reference in its entirety). Using phage display as an example, separate antibody genes may be inserted into the DNA of phages, so that the variable regions on the antibody molecules that can bind the antigens may be coupled to the capsid protein of the phage. After the phage infecting E. coli, single stranded DNA may be replicated in E. coli, and the phage may be reassembled and secreted into the culture medium, while the E. coli may not be lysed. The phage may be co-incubated with target antigens; and after the bound phages are isolated, amplification and purification may be then conducted so that great amounts of clones can be screened. The phage display technique can be found in the literature (Liu et al., 2004; the contents of which are hereby incorporated by reference in its entirety).
In another aspect, the present disclosure may include methods for producing a monoclonal antibody using a phage display method. Specifically, mRNA may be prepared from an animal, e.g., rabbits, rats, mice, guinea pigs, hamsters, goats, horses, chickens, sheep, and camelids (e.g., llamas), immunized by the method for immunizing an animal, whereupon cDNA may be prepared using the mRNA as a template, so that a single-chain antibody (scFv) gene encoding only an antibody variable region may be prepared. The gene may be cloned to a phagemid vector. E. coli, into which the phagemid vector is transduced, is infected with phage, so as to express the scFV antibody on the phage capsid. Screening of the scFv expressed in this way against an antigen protein or against a peptide-MHC complex may prepare a monoclonal scFV antibody specific to the antigen protein or the peptide-MHC complex. Herein, preparation of mRNA, preparation of cDNA, subcloning to phagemid or transduction to E. coli, phage infection, and screening of a monoclonal scFV antibody specific to an antigen protein or a peptide-MHC complex each may be performed by the known method. For example, subcloning of a scFV gene to a phagemid vector containing two elements consisting of a gene fragment encoding a leader sequence (signal sequence) and a phage capsid protein iii and a replication origin of M13 and using of M13 phage as a phage can express a scFV antibody on the M13 phage. Further, a phage obtained by screening may be infected to a specific bacterium and cultured, so that a monoclonal antibody specific to an antigen protein may also be collected in large quantities from the culture. According to the method for producing a monoclonal antibody of the present disclosure, not only an scFV antibody but also an antibody fragment having no constant region, such as a Fab antibody fragment, may be prepared.
In another aspect, the present disclosure may include phage display libraries, in which the heavy and light chain variable regions of an antibody may be synthesized such that they include nearly all possible specificities.
In another aspect, the present disclosure may include generation of phage display libraries containing phage other than M13. Other bacteriophages, such as lambda phage, may also be useful in the method of the present disclosure. Lambda phage display libraries have been generated, which display peptides encoded by heterologous DNA on their surface (Sternberg et al., 1995; the content of which is hereby incorporated by reference in its entirety). Moreover, the method of the present disclosure may be extended to include viruses other than bacteriophage, such as eukaryotic viruses. Eukaryotic viruses may be generated that encode genes suitable for delivery to a mammal and that encode and display an antibody capable of targeting a specific cell type or tissue into which the gene is to be delivered. For example, retroviral vectors have been
generated, which display functional antibody fragments (Russell et al., 1993; the content of which is hereby incorporated by reference in its entirety).
T-cell receptors (TCRs)
Provided is furthermore a TCR, including a soluble TCR. The present disclosure further relates to a TCR comprising an alpha chain and a beta chain (“alpha/beta TCRs”). Also provided is a peptide disclosed herein that is capable of binding to a TCR and an antibody when presented by an MHC molecule.
As explained above, T-cell receptors (TCRs) bind to short protein fragments, such as the peptides disclosed herein, that are presented on the surface of most nucleated cells by MHC molecules. Each T cell expresses a unique TCR that is generated by randomly assorting genes. This ensures that the cellular immune response can respond to almost any infection and all sorts of cancer.
A respective TCR may be a TCR that has been isolated from a given T cell clone. Single cell TCR alpha and beta chains can be sequenced using emulsion-based PCR techniques. Such technique may be combined with a PCR-based single cell barcoding strategy and next-generation sequencing (NGS) (De Simone et al., 2018).
MHC bound peptides can be recognized by the immune system, specifically T lymphocytes. T lymphocytes expressing a TCR that specifically binds a peptide can destroy the cells presenting the recognized MHC-peptide complex, e.g. acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer cells presenting the derived peptides.
TCR fragments
The present disclosure also relates to a fragment of a TCR provided herein that is capable of binding to a peptide antigen provided herein when presented by an MHC molecule. The term particularly relates to a soluble TCR fragment, for example a TCR lacking the transmembrane part and/or constant regions, a single chain TCR, and a fusion thereof, for example to an antibody. The generation of soluble TCR fragments, specifically binding to an antigen, have been disclosed more than 30 years ago (Novotny, J, et al., Proc. Nati. Acad. Sci. USA (1991 ) 88, 19, 8646-8650).
Examples of a TCR fragment include, but are not limited to, a fragment of the a, p, 8 or the y chain and a combination of any of these. A TCR fragment may contain one or both of a Va-Ca or Vp-Cp or a portion thereof. As noted above, a functional TCR fragment generally contains a variable domain such as Va, Vp, V y and/or V 8. The TCR fragment may in some embodiments be a single-chain TCR (scTCR), for example having an alpha chain variable region linked to a beta chain variable region.
A TCR or TCR fragment may in some embodiments be humanized or a human TCR or TCR fragment.
In some embodiments a provided TCR is a cloned TCR engineered into an autologous or allogeneic T cell. Provided is also a method of generating a T cell as disclosed herein. Provided is also a method of generating an NK cell or another cell bearing said TCR or cross-reacting with said TCR.
The alpha and beta chains of alpha/beta TCR's, and the gamma and delta chains of gamma/delta TCRs, are generally regarded as each having two "domains", namely variable and constant domains. The variable domain consists of a concatenation of the variable region (V) and the joining region (J). The variable domain may also include a leader region (L). Beta and delta chains may also include a diversity region (D). The alpha and beta constant domains may also include C-terminal transmembrane (TM) domains that anchor the alpha and beta chains to the cell membrane.
With or without the introduced inter-chain bond mentioned above, alpha/beta heterodimeric TCRs of the present disclosure may have a TRAC constant domain sequence and a TRBC1 or TRBC2 constant domain sequence, and the TRAC constant domain sequence and the TRBC1 or TRBC2 constant domain sequence of the TCR may be linked by the native disulfide bond between Cys4 of exon 2 of TRAC and Cys2 of exon 2 of TRBCI or TRBC2.
In an embodiment, a TCR of the present disclosure having at least one mutation in the alpha chain and/or having at least one mutation in the beta chain has modified glycosylation compared to the unmutated TCR.
In an embodiment, a TCR comprising at least one mutation in the TCR alpha chain and/or TCR beta chain has a binding affinity for, and/or a binding half-life for, a peptide-MHC molecule complex, which is at least double that of a TCR comprising the unmutated TCR alpha chain and/or unmutated TCR beta chain. Affinity-enhancement of tumor specific TCRs, and its exploitation, relies on the existence of a window for optimal TCR affinities. The existence of such a window is based on observations that TCRs specific for e.g. MHC-restricted pathogens have Kd values that are generally about 10-fold lower when
compared to TCRs specific for e.g. MHC-restricted tumor-associated self-antigens. It is now known, although tumor antigens have the potential to be immunogenic, because tumors arise from the individual’s own cells only mutated proteins or proteins with altered translational processing will be seen as foreign by the immune system. Antigens that are upregulated or overexpressed (so called self-antigens) will not necessarily induce a functional immune response against the tumor: T cells expressing TCRs that are highly reactive to these antigens will have been negatively selected within the thymus in a process known as central tolerance, meaning that only T cells with low-affinity TCRs for self-antigens remain (Murphy et al., 2022, pp. 243-245, 333-341 Therefore, affinity of TCRs of the present disclosure to peptides can be enhanced by methods well known in the art.
The present disclosure further relates to a method of identifying and isolating a TCR according to the present disclosure. The method includes incubating PBMCs from healthy donors negative for the respective HLA allotype with HLA-peptide monomers, incubating the PBMCs with tetramer-phycoerythrin (PE) and isolating the high avidity T cells by fluorescence activated cell sorting (FACS) Calibur analysis.
Methods of producing a TCR
In one embodiment the disclosure provides a method of producing a TCR as described herein, the method comprising culturing a host cell capable of expressing the TCR under conditions suitable to promote expression of the TCR.
For example, a soluble T cell receptor can be generated from a specific T cell clone, and its affinity can be increased by mutagenesis targeting the complementarity-determining regions. For the purpose of T cell receptor selection, phage display can be used (US 2010/0113300, Liddy et al., 2012, the contents of which are incorporated by reference in their entirety). For the purpose of stabilization of T cell receptors during phage display and in case of practical use as drug, alpha and beta chains can be linked e.g. by nonnative disulfide bonds, other covalent bonds (single chain T cell receptor), or by dimerization domains (Boulter et al., 2003; Card et al., 2004; Willcox et al., 1999, the contents of which are incorporated by reference in their entirety). The T cell receptor can be linked to a toxin, a drug, a cytokine (see for example US 2013/0115191 , the content of which is incorporated by reference in its entirety), and a domain recruiting effector cells such as an anti-CD3 domain, etc., in order to execute particular functions on target cells. Moreover, it could be expressed in T cells used for adoptive transfer. Further information can be found in WO 2004/033685A1 and WO 2004/074322A1 . A combination of soluble TCRs is described in WO 2012/056407A1 . Further methods for the production are
disclosed in WO 2013/057586A1 , the content of which is incorporated by reference in its entirety.
The present disclosure further relates to a method of identifying and isolating a TCR according to the present disclosure, said method comprising obtaining a transgenic mouse with the entire human TCRap gene loci (1.1 and 0.7 Mb), whose T cells express a diverse human TCR repertoire that compensates for mouse TCR deficiency, immunizing the mouse with a peptide, incubating PBMCs obtained from the transgenic mice with tetramer-phycoerythrin (PE), and isolating the high avidity T cells by fluorescence activated cell sorting (FACS) Calibur analysis.
In one aspect, to obtain T cells expressing TCRs of the present disclosure, nucleic acids encoding TCR-alpha and/or TCR-beta chains of the present disclosure are cloned into expression vectors, such as gamma retrovirus or lentivirus. The recombinant viruses are generated and then tested for functionality, such as antigen specificity and functional avidity. An aliquot of the final product is then used to transduce the target T cell population (generally purified from patient PBMCs), which is expanded before infusion into the subject / patient.
In another aspect, to obtain T cells expressing TCRs of the present disclosure, TCR RNAs are synthesized by techniques known in the art e.g. in vitro transcription systems. The in vitro synthesized TCR RNAs are then introduced into primary CD8-positive T cells obtained from healthy donors by electroporation to re-express tumor specific TCR-alpha and/or TCR-beta chains.
To increase the expression, a nucleic acid encoding a TCR of the present disclosure may be operably linked to a strong promoter, such as a retroviral long terminal repeat (LTR), cytomegalovirus (CMV), murine stem cell virus (MSCV) U3, phosphoglycerate kinase (PGK), p-actin, ubiquitin, and a simian virus 40 (SV40)/CD43 composite promoter, elongation factor (EF)-1 a and the spleen focus-forming virus (SFFV) promoter. In one embodiment, the promoter is heterologous to the nucleic acid being expressed.
In addition to strong promoters, TCR expression cassettes of the present disclosure may contain additional elements that can enhance transgene expression, including a central polypurine tract (cPPT), which promotes the nuclear translocation of lentiviral constructs (Follenzi et al., 2000), and the woodchuck hepatitis virus post-transcriptional regulatory element (wPRE), which increases the level of transgene expression by increasing RNA stability (Zufferey et al., 1999, the contents of which are incorporated by reference in their entirety).
The alpha and beta chains of a TCR of the present disclosure may be encoded by nucleic acids located in separate vectors or may be encoded by polynucleotides located in the same vector.
Achieving high level TCR surface expression requires that both the TCR-alpha and TCR- beta chains of the introduced TCR be transcribed at high levels. To do so, the TCR-alpha and TCR-beta chains of the present disclosure may be cloned into bi-cistronic constructs in a single vector, which has been shown to be capable of overcoming this obstacle. The use of a viral internal ribosomal entry site (IRES) between the TCR-alpha and TCR-beta chains results in the coordinated expression of both chains, because the TCR-alpha and TCR-beta chains are generated from a single transcript that is broken into two proteins during translation, ensuring that an equal molar ratio of TCR-alpha and TCR-beta chains are produced (Schmitt et al., 2009, the contents of which are incorporated by reference in their entirety).
Nucleic acids encoding TCRs of the present disclosure may be codon optimized to increase expression from a host cell. Redundancy in the genetic code allows some amino acids to be encoded by more than one codon, but certain codons are less “optimal” than others because of the relative availability of matching tRNAs as well as other factors (Gustafsson et al., 2004). Modifying the TCR-alpha and TCR-beta gene sequences such that each amino acid is encoded by the optimal codon for mammalian gene expression, as well as eliminating mRNA instability motifs or cryptic splice sites, has been shown to significantly enhance TCR-alpha and TCR-beta gene expression (Scholten et al., 2006, the contents of these references are herein incorporated by reference in their entirety).
Furthermore, mispairing between the introduced and endogenous TCR chains may result in the acquisition of specificities that pose a significant risk for autoimmunity. For example, the formation of mixed TCR dimers may reduce the number of CD3 molecules available to form properly paired TCR complexes, and therefore can significantly decrease the functional avidity of the cells expressing the introduced TCR (Kuball et al., 2007, the contents of which are incorporated by reference in their entirety).
To reduce mispairing, the C-terminus domain of the introduced TCR chains of the present disclosure may be modified in order to promote interchain affinity, while decreasing the ability of the introduced chains to pair with the endogenous TCR. These strategies may include replacing the human TCR-alpha and TCR-beta C-terminus domains with their murine counterparts (murinized C-terminus domain); generating a second interchain disulfide bond in the C-terminus domain by introducing a second cysteine residue into both the TCR-alpha and TCR-beta chains of the introduced TCR (cysteine modification); swapping interacting residues in the TCR-alpha and TCR-beta chain C-terminus domains
(“knob-in-hole”); and fusing the variable domains of the TCR-alpha and TCR-beta chains directly to CD3< (CD3< fusion) (Schmitt et al., 2009).
In an embodiment, a host cell is engineered to express a TCR of the present disclosure. In some embodiments, the host cell is a human T cell or T cell progenitor. In some embodiments the T cell or T cell progenitor is obtained from a subject suffering from cancer. In other embodiments the T cell or T cell progenitor is obtained from a healthy donor. Host cells of the present disclosure can be allogeneic or autologous with respect to a subject to be treated. In one embodiment, the host is a gamma/delta T cell transformed to express an alpha/beta TCR.
It is a further aspect there is provided a method for producing a soluble T cell receptor recognizing a specific peptide-MHC complex.
Aptamers
A nucleic acid molecule as a binding molecule is typically an aptamer. An aptamer as provided herein may be an aptamer that specifically binds to a peptide as disclosed herein. An aptamer (see for example WO 2014/191359 and the literature as cited therein) is a short single-stranded nucleic acid molecule, which can fold into a defined three- dimensional structure and recognize a specific target structure. Aptamers have appeared to be suitable alternatives for developing targeted therapies. Aptamers have been shown to selectively bind to a variety of complex targets with high affinity and specificity.
Aptamers recognizing cell surface located molecules have been identified within the past decades and provide means for developing diagnostic and therapeutic approaches. Since aptamers have been shown to possess almost no toxicity and immunogenicity, they are promising candidates for biomedical applications. Indeed aptamers, for example prostate-specific membrane-antigen recognizing aptamers, have been successfully employed for targeted therapies and shown to be functional in xenograft in vivo models. Furthermore, aptamers recognizing specific tumor cell lines have been identified.
A DNA aptamer can be selected to reveal broad-spectrum recognition properties for various cancer cells, and particularly those derived from solid tumors, while non- tumorigenic and primary healthy cells are not recognized. If the identified aptamers recognize not only a specific tumor sub-type but rather interact with a series of tumors, this renders the aptamers applicable as so-called broad-spectrum diagnostics and therapeutics.
Further, investigation of cell-binding behavior with flow cytometry showed that the aptamers revealed very good apparent affinities that are within the nanomolar range.
Aptamers are useful for diagnostic and therapeutic purposes. Further, it could be shown that some of the aptamers are taken up by tumor cells and thus can function as molecular vehicles for the targeted delivery of anti-cancer agents such as siRNA into tumor cells.
Aptamers can be selected against complex targets such as cells and tissues and complexes of the peptides comprising, including consisting of, a sequence according to any of SEQ ID NOs: 1 to 59 disclosed herein with an MHC molecule. Selection may be done using the cell-SELEX (Systematic Evolution of Ligands by Exponential enrichment) technique.
Toxins
In some embodiments a binding molecule, such as a TCR or an antibody, is coupled to a toxin.
Detectable Labels
Peptides, binding molecules, such as antibodies or TCRs, and T lymphocytes as disclosed herein, may be labeled for detection by any desired imaging method. Additionally, labels may be bi- or multi-functional and may be detectable by more than one method. The peptide, binding molecule, or T-lymphocyte may be directly or indirectly labeled. Attachment of a label to a binding molecule includes covalent attachment of the label, incorporation of the label into the binding molecule, and the covalent attachment of a chelating compound for binding of label, amongst others well recognized in the art. Methods for detection of labels include, but are not limited to, fluorescence, light, confocal and electron microscopy; magnetic resonance imaging and spectroscopy; fluoroscopy, computed tomography and positron emission tomography.
Each binding molecule can include a labelling which provides that the bound binding molecule can be detected by determining the presence or absence of a signal provided by the label. For example, the binding molecule can be labelled with a fluorescent dye or any other applicable cellular marker molecule. Such marker molecules are well known in the art. For example, a fluorescence-labelling, for example provided by a fluorescence dye, can provide a visualization of the bound aptamer by fluorescence or laser scanning microscopy or flow cytometry.
A binding molecule as disclosed herein, in particular, a proteinaceous binding molecule such as a TCR or an antibody or an TCR or antibody fragment, may have a detectable label. The binding molecule may in some embodiments be covalently coupled to a detectable label. Such a detectable label may be selected from the group consisting of a radionuclide, a fluorophore and biotin. A binding molecule of the present disclosure may
be conjugated to a therapeutically or diagnostically active agent, such as a radionuclide, a chemotherapeutic agent, or a toxin.
In some embodiments a respective label may be used in monitoring of a cancer therapy or in diagnosis of cancer. Immunohistochemical techniques are, for example, established in the art in diagnosis since decades. For immunohistochemistry, the disease tissue sample may be fresh or frozen or may be embedded in paraffin and fixed with a preservative such as formalin. The fixed or embedded section contains the sample are contacted with a labeled primary antibody and secondary antibody, wherein the antibody is used to detect the expression of the proteins in situ.
Detectable labels of a peptide, binding molecule or T-lymphocyte may be for example fluorophores, enzymes, substrates, bioluminescent materials, radioactive materials, radioisotopes such as positron emitting radionucleotides, gold, gadolinium and other lanthanides, paramagnetic iron and chemiluminescent labels. Exemplary enzymes labels include, but are not limited to, horseradish peroxidase, acetylcholinesterase, alkaline phosphatase, b-galactosidase and luciferase. Exemplary fluorophore (fluorescent materials) include, but are not limited to, eosin, rhodamine, fluorescein, fluorescein isothiocyanate, umbelliferone, dichlorotriazinylamine, phycoerythrin and dansyl chloride. Exemplary chemiluminescent labels include, but are not limited to, luminol. Exemplary bioluminescent materials include, but are not limited to, luciferin and aequorin.
Exemplary radioactive materials include radionuclides. Examples of radionuclides are carbon-13 (13C), nitrogen-15 (15N), deuterium (2H), as well as any combination thereof. Further examples of radionuclides include, but are not limited to, beta-emitters such as phosphorus-32 (32P), scandium-47 (47Sc), copper-67 (67Cu), gallium-67 (67Ga), yttrium-88 (88Y), yttrium-90 (90Y), iodine-125 (1251), iodine-131 (1311), samarium-153 (153Sm), lutetium-177 (177Lu), rhenium-186 (186Re), rhenium-188 (188Re), and alphaemitters such as astatine-211 (211 At), lead-212 (212Pb), bismuth-212 (212Bi), bismuth- 213 (213Bi), actinium-225 (225Ac) or any combination thereof.
T cell therapy
In some embodiments, a peptide as disclosed herein is used to generate activated T lymphocytes in vitro. In some embodiments, nucleic acids encoding binding molecules, such as T cells receptors as disclosed herein can be used to genetically engineer T lymphocytes in vitro. Such genetically engineered T lymphocytes express T cell receptors that specifically recognize a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule. The T lymphocytes obtained by either method may be used for administration to an individual.
T cell-based therapies have been successfully developed in clinical trials. An approach used in the treatment of advanced solid tumors is tumor-infiltrating lymphocyte therapy (TIL therapy). While T lymphocytes within tumor tissue are often enriched in clones with specificity for peptides that are presented only on cells of the tumor, the efficiency of TIL therapy is based on the proliferation of such T lymphocytes. T lymphocytes are isolated from a subject’s tumor, stimulated and expanded in vitro, and administered back into the patient.
In adoptive cellular therapy (ACT), T cells isolated from a donor’s peripheral blood cells are engineered to express TCRs of desired specificity and affinity to a peptide presented on cancer cells. The donor may be the subject of the treatment (autologous therapy) or different from the subject of the treatment (allogeneic therapy). For this purpose, peripheral blood mononuclear cells (PBMCs) are obtained from donors and/or from the subject, and T lymphocytes binding to one or more desired peptides identified as being presented on cells of the cancer to be treated are enriched and isolated by FAC-sorting. T cell clones are isolated and the TCRs sequenced, after T lymphocytes responsive to the desired peptide(s) have been isolated and expanded. The respective TCRs can then, possibly after having been engineered, be expressed in T cells isolated from a donor or the subject. The number of T lymphocytes and thereby TCRs available that can be screened against a certain peptide has been found to be only limited by the number of donor PBMCs tested.
Both above approaches are successful because T cell clones against any non-self peptide can be identified in a subject and/or in donor PBMCs.
As explained above, if a peptide is presented on the surface of cells of healthy tissue, the peptide is recognized as a self-antigen due to MHC restriction in T lymphocyte development of the respective organism. Precursors of T lymphocytes undergo positive and negative selection in the thymus. T lymphocytes carrying TCRs that are specific for peptides presented on cells of the organism are eliminated by clonal deletion or by a process called anergy. The organism thus has a self-tolerance with regard to such peptides.
For a peptide not presented on healthy cells there is no clonal deletion in the thymus. The peptide is only found in an organism with cancer since the cancer concerned exists, and it is not found on any cell in a healthy organism. Among T lymphocytes isolated from a blood sample, for example of a healthy organism, there will typically be no T lymphocytes with a TCR that is specific for a peptide presented on the surface of healthy cells. However, T lymphocytes with a TCR that is specific for a peptide as disclosed herein can be expected to be found in a blood sample. It is estimated that in an average
adult human, there are around 4 x 1011 total circulating T cells and an estimated 1 O10 unique T cell clonotypes (Lythe, G, et al., J. Theor. Biol. (2016) 389, 214-224). Already in a millilitre of blood of a healthy donor there are typically around 200.000 to 800.000 CD8 T cells. A few millilitres thus already exceed the number of a million CD8 T cells. The skilled person can thus isolate a T lymphocyte of with a specificity for a peptide disclosed herein.
Peripheral blood T lymphocytes can be obtained and selected in vitro in order to obtain and express a T cell receptor (TCR) that is specific for the peptide comprising the sequence according to any one of SEQ ID NOs: 1 to 59. The selection can be achieved by coculturing the obtained T lymphocytes with antigen-presenting cells that express a peptide disclosed herein. Both in vivo and in vitro, T lymphocytes clonally expand upon recognition of the epitope peptide to which they bind. In vitro this leads to proliferation of the T lymphocytes specific for a peptide disclosed herein. Following in vitro selection and proliferation, a T cell clone can be produced that is specific for a peptide disclosed herein.
On this basis adoptive cell therapy (ACT) can be applied, which has become an established approach in cancer therapy. T-cell therapy may rely on T lymphocytes obtained directly from the same individual, which is an autologous approach. T lymphocytes obtained from the same individual can be grown and selected in culture, and infused back into the individual for the treatment of the cancer concerned. In such an autologous setting, T lymphocytes obtained from the individual can be engineered to express a TCR transgene specific for a peptide disclosed herein. The recombinant TCR confers to the respective T lymphocytes the ability to recognize the peptide disclosed herein and kill tumor cells. The recombinant TCR may also be an affinity-maturated TCR. T-cell engineering also allows constructing a TCR with optimized affinity, selecting the most appropriate T-cell subset to be transduced and given to the individual, and to control over its in vivo efficacy and safety.
A TCR transgene specific for a peptide disclosed herein may in some embodiments have been obtained from a TCR isolated from another species, e.g. a murine TCR. Chimeric HLA-A2 transgenic mice with a diverse T-cell repertoire have been used to allow the isolation of human TCRs against human tumor-associated antigens from antigennegative hosts.
T-cell therapy may also rely on T lymphocytes obtained from healthy donors other than the individual to be treated, which is an allogeneic approach. In the allogeneic setting, suitable lymphocytes, such as gamma delta T lymphocytes, may be genetically engineered to express a TCR that is selected, obtained and optionally affinity-maturated as described above. Alternatively or additionally, T lymphocytes obtained from a donor
may be engineered to express so called “suicide genes”, which renders expressing cells sensitivity to a prodrug. Thereby the T lymphocytes can be selectively eliminated in case of adverse events. This approach has been successfully used on transplanted patients. The most extensively tested suicide gene in clinical trials is the gene of the thymidine kinase from HSV (HSV-TK).
Accordingly, such a T lymphocyte and a TCR can be taken to define an additional embodiment of an immunotherapeutic use of a peptide as provided herein.
Method of Producing T lymphocytes
Another aspect of the present disclosure involves an in vitro method for producing T lymphocytes.
These methods may include the use of the peptides complexed with MHC (e.g. as presented on an antigen-presenting cell as described below) to generate a T cell receptor whose nucleic acid is cloned and is introduced into a host cell, for example a T lymphocyte. This engineered T lymphocyte can then be transferred to a subject for therapy of cancer. To this end, PBMCs are taken from a donor and the desired T lymphocyte subset for genetic engineering is enriched, e.g. by depleting monocytes or alpha beta T cells. The donor may or may not be the subject of the treatment.
The T lymphocytes can then be activated, e.g. in the presence of anti-CD3 and anti- CD28 in case of alpha beta T cells for autologous T cell therapy, and subsequently be transduced with a nucleic acid or vector encoding the TCR. Suitable vectors include, e.g. lentiviral vectors. Subsequently, T lymphocytes may be cultured in the presence of suitable cytokines such as cytokine interleukin 7 (IL-7) and/or interleukin 15 (IL-15). An exemplary process for genetically engineering T lymphocytes is described in WO 2019157298 A1 , which is incorporated herein by reference in its entirety. Other T lymphocyte subsets may require different handling. E.g., for gamma delta T cells in case of allogeneic therapy, T lymphocytes may be activated with zoledronate before being subjected to genetic engineering. An exemplary process is described in WO 2019104269 A1 , which is incorporated herein by reference in its entirety.
Additionally or alternatively, the method may include contacting in vitro T lymphocytes with peptide loaded human MHC molecules expressed on the surface of a suitable antigen-presenting cell for a period of time sufficient to activate the T lymphocyte in an antigen specific manner, wherein the antigen is a peptide according to the invention. Such activated T lymphocytes can either be clonally expanded and reinfused into the subject, or be used to generate TCRs that are used to genetically engineer suitable T lymphocytes for immunotherapy as described above.
Generally, a sufficient amount of the antigen is used with an antigen-presenting cell.
In some embodiments the mammalian antigen-presenting cell lacks or has a reduced level or function of the TAP peptide transporter. Suitable cells that lack the TAP peptide transporter include T2, RMA-S and Drosophila cells. TAP is the transporter associated with antigen processing.
The human peptide loading deficient cell line T2 is available from the American Type Culture Collection, 12301 Parklawn Drive, Rockville, Maryland 20852, USA under Catalogue No CRL 1992; the Drosophila cell line Schneider line 2 is available from the ATCC under Catalogue No CRL 19863; the mouse RMA-S cell line is described in Ljunggren et al. (Ljunggren and Karre, 1985, the contents of which are incorporated by reference in their entirety).
In typical embodiments, before transfection the host cell expresses substantially no MHC class I molecules. In some embodiments the stimulator cell expresses a molecule important for providing a co-stimulatory signal for T cells such as any of B7.1 , B7.2, ICAM-1 and LFA3. The nucleic acid sequences of numerous MHC class I molecules and of the co-stimulator molecules are publicly available from the GenBank and EMBL databases.
In case of an MHC class I epitope being used as an antigen; the T lymphocytes may be CD8-positive T lymphocytes or cytotoxic T lymphocytes (CTLs). These T cells may be further expanded and either directly reinfused into the patient, or used to obtain a suitable TCR that is introduced into a suitable host cell, typically a T lymphocyte (e.g., a CD8- positive T lymphocyte, a CD4-positive T lymphocyte or a gamma delta T lymphocyte).
If an antigen-presenting cell is transfected to express such an epitope, the cell may comprise an expression vector capable of expressing a peptide containing the sequence of one of SEQ ID NO: 1 to SEQ ID NO: 59.
A number of other methods may be used for generating T lymphocytes in vitro. For example, autologous tumor-infiltrating lymphocytes can be used in the generation of CTL. Piebanski et al. (Piebanski et al., 1995) made use of autologous peripheral blood lymphocytes (PLBs) in the preparation of T cells. Furthermore, the production of autologous T cells by pulsing dendritic cells with a peptide, or via infection with recombinant virus is possible. Also, B cells can be used in the production of autologous T cells. In addition, macrophages pulsed with a peptide, or infected with recombinant virus, may be used in the preparation of autologous T cells. S. Walter et al. (Walter et al., 2003, the contents of which are incorporated by reference in their entirety) describe the in vitro priming of T cells by using artificial antigen-presenting cells (aAPCs), which is
also a suitable way for generating T cells against the peptide of choice. The inventors have generated aAPCs by the coupling of preformed MHC-peptide complexes to the surface of polystyrene particles (microbeads) by biotin:streptavidin biochemistry. This system permits the exact control of the MHC density on aAPCs, which allows to selectively elicit high- or low-avidity antigen specific T cell responses with high efficiency from blood samples. Apart from MHC-peptide complexes, aAPCs should carry other proteins with co-stimulatory activity like anti-CD28 antibodies coupled to their surface. Furthermore, such aAPC-based systems often require the addition of appropriate soluble factors, e. g. cytokines, like interleukin-12.
Other cells may also be used in the preparation of T cells and a method is described in detail in WO 97/26328, incorporated herein by reference in its entirety. For example, in addition to Drosophila cells and T2 cells, other cells may be used to present antigens such as CHO cells, baculovirus-infected insect cells, bacteria, yeast, and vaccinia- infected target cells. In addition plant viruses may be used see for example Porta et al. (Porta et al., 1994, the content of which is incorporated by reference in its entirety) which describes the development of cowpea mosaic virus as a high-yielding system for the presentation of foreign peptides.
The T lymphocytes provided with the method described herein may be activated T lymphocytes. The T lymphocytes that are directed against the peptides provided herein are useful in therapy. Thus, a further aspect of the disclosure provides T cells obtainable by the foregoing methods disclosed herein.
T lymphocytes, which are produced by the above method, will selectively recognize a cell that aberrantly expresses a peptide that comprises an amino acid sequence of SEQ ID NO: 1 to SEQ ID NO: 59.
In some embodiments, the T lymphocyte recognizes the target cell that presents a peptide as disclosed herein by interacting through its TCR with the MHC peptide complex (for example, binding). The T lymphocytes are useful in a method of killing target cells in a subject whose target cells aberrantly express a polypeptide or peptide that includes an amino acid sequence as provided herein wherein the subject is administered an effective number of the T lymphocytes. The T lymphocytes that are administered to the subject may be derived from the subject and optionally activated as described above (i.e. they are autologous T cells). In some embodiments, the T lymphocytes are not from the subject but are from another individual. In typical embodiments the individual is a healthy individual.
In vivo, the target cells for the T lymphocytes, e.g. CD8-positive T lymphocytes, provided herein may be cells of the tumor (which sometimes express MHC class II) and/or stromal cells surrounding the tumor (which sometimes also express MHC class II) (Dengjel et al., 2006).
The T lymphocytes provided herein may be used as active ingredients of a therapeutic composition.
T lymphocytes may be obtained by methods known in the art, e.g., those described above.
Protocols for this so-called adoptive transfer of T lymphocytes are well known in the art. Several reviews can be found (Gattinoni et al., 2006; Morgan et al., 2006, the contents of which are incorporated by reference in their entirety).
Pharmaceutical composition
Any molecule provided herein, i.e. the peptide, nucleic acid, antibody, expression vector, cell, activated T cell, T cell receptor or the nucleic acid encoding it, is useful for the treatment of disorders, characterized by cells escaping an immune response. Therefore, any molecule provided herein may be used as medicament or in the manufacture of a medicament. The molecule may be used by itself or combined with other molecule(s) provided herein or (a) known molecule(s).
Any molecule provided herein may also be included in a pharmaceutical composition. In some embodiments, the pharmaceutical composition provided herein may be included in a pharmaceutical combination, such as a combination product. Two pharmaceutical compositions that differ from each other in their active components, may define the combination product. The compositions of a combination product may be administered independently. In some embodiments the compositions of a combination product may be administered at different points of time. One of the compositions of a combination product may be administered in a time-delayed manner relative to another composition of the combination product. In some embodiments the compositions of a combination product may be administered concomitantly.
In some embodiments a pharmaceutical composition as provided herein, such as a therapeutic composition or a vaccine composition, is provided in the form of a solution. In some embodiments the components of the pharmaceutical composition need to be or are dissolved or suspended in a pharmaceutically acceptable liquid vehicle such as water, an organic solvent, a mixture of both or one or more pharmaceutically acceptable oils or
fats. A composition such as a therapeutic composition or a vaccine composition as disclosed herein may be provided in lyophilized form or in freeze-dried form. The lyophilized form may when needed be solubilized in water, for example for injection, and gently mixed. In some embodiments, an immunoadjuvant as described hereinabove may then be added. The mixture may then be gently mixed and charged into a suitable syringe.
A pharmaceutical composition as disclosed herein may be an individualized pharmaceutical composition in that the composition has been specifically tailored for an individual. An individualized pharmaceutical composition has generally been verified to contain an active ingredient that is predicted to be effective in the particular individual. A pharmaceutical composition as disclosed herein may also be provided without being verified for any individual. The pharmaceutical composition may for example contain a peptide as disclosed herein and be administered as a vaccine on the basis of a risk assessment, without knowing whether there is an increased risk that the peptide will be or is presented on cancer cells of the individual.
Method of stratifying a subject
In some embodiments a peptide as disclosed herein may be used for risk stratification, for diagnosis of existing cancer, for monitoring for a risk level of cancer, including for a potential risk increase, and for predicting a future medical outcome.
Determining presentation, including the presentation level, of a peptide as disclosed herein can be used to stratify a subject undergoing or about to undergo cancer treatment. Determining presentation, including the presentation level, of a peptide as disclosed herein can also be used to stratify a subject for vaccination.
With regard to human individuals, the use of biomarkers for stratification of patients is a procedure well established in the art. This procedure includes or consists of linking one or more patient subpopulations, characterized by a certain feature, in the context of the present disclosure the over-presentation of a particular peptide by MHC on cancer cells, to a particular treatment. The general aim of stratification is to match patients with therapies that are more likely to be effective and safe. In a more general context stratifying patients may in some embodiments include evaluation of patient history and physical assessment, combined with laboratory tests on the basis of a method as disclosed herein, and clinical observation.
Provided herein is a method of stratifying a subject for individualized, including personalized, vaccination and/or immunotherapy against cancer. The method includes assessing whether cells of a cancer sample from the subject present a peptide as disclosed herein. The method also includes selecting the subject for treatment with a
vaccine and/or an immunotherapeutic agent, if cells in the cancer sample over-present a peptide as disclosed herein, relative to one or more corresponding healthy tissue samples or cells. The vaccine includes a peptide that contains a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a pharmaceutically acceptable salt thereof. The immunotherapeutic agent in some embodiments includes the binding molecule as disclosed herein. In some embodiments the immunotherapeutic agent includes a nucleic acid and/or an expression vector as disclosed herein. In some embodiments the immunotherapeutic agent includes a recombinant host cell as disclosed herein. In some embodiments the immunotherapeutic agent includes a T lymphocyte as disclosed herein, for example an activated T lymphocyte or a recombinant T lymphocyte.
Method to Produce an (Individualized) Pharmaceutical Product
In a further aspect there is provided a method of producing, e.g. formulating or synthesizing a pharmaceutical product, such as a therapeutic product. The pharmaceutical product may in some embodiments be a vaccine. The pharmaceutical product may in some embodiments be a compound-based product. The pharmaceutical product may in some embodiments be a cellular product.
Provided is furthermore a method for producing an individualized, including personalized, pharmaceutical product. The individualized pharmaceutical product may be an anticancer vaccine or a compound-based and/or cellular therapeutical product. Generally, the individualized pharmaceutical product includes a peptide as provided herein.
In typical embodiments, the method for producing an individualized pharmaceutical product for an individual includes manufacturing a pharmaceutical composition that includes at least one peptide selected from a collection of pre-screened peptides. The at least one peptide used in the pharmaceutical composition is selected for suitability in the individual. In one embodiment, the pharmaceutical composition is a vaccine. The method may also be adapted to produce T cell clones for down-stream applications, such as a TCR isolation, or a soluble antibody, and other treatment options.
As noted above, the collection of peptides (e.g., in the form of a database) is composed of tumor-associated peptides which were highly overexpressed in tumor tissue. The peptides may have been identified on cells of colorectal cancer, glioblastoma, gastric cancer, hepatocellular carcinoma, and subjects suffering from renal cell carcinoma with various HLA-A HLA-B and HLA-C alleles. The peptides may also have been identified on cells of prostate cancer, bladder cancer and brain cancer. The peptides may also have been identified on cells of breast cancer, esophageal cancer, liver cancer, lung cancer (NSCLC, SCLC) and/or ovarian cancer. The peptides may also have been identified on
cells of uterine cancer and/or bile duct cancer. In some embodiments the cells may have been identified on cells of pancreatic cancer and/or gallbladder cancer. The peptides may also have been identified on cells of melanoma, merkel cell carcinoma and/or leukemia (AML, CLL). It may contain MHC class I and MHC class ii peptides or elongated MHC class I peptides. In addition to the tumor associated peptides collected from several colorectal cancer, glioblastoma, gastric cancer, hepatocellular carcinoma, and renal cell carcinoma tissues, the collection of peptides may contain HLA-A*02, HLA-A*01 , HLA- A*03, HLA-A*24, HLA-B*07, HLA-B*08 and HLA-B*44 marker peptides. These peptides allow comparison of the magnitude of T-cell immunity induced by peptides in a quantitative manner and hence allow important conclusion to be drawn on the capacity of the vaccine to elicit anti-tumor responses. Secondly, they function as important positive control peptides derived from a “non-self” antigen in the case that any vaccine-induced T cell responses to peptides derived from “self” antigens in a subject are not observed. And thirdly, it may allow conclusions to be drawn, regarding the status of immunocompetence of the subject.
Peptides for the collection of peptides may be identified by using an integrated functional omics approach combining gene expression analysis, mass spectrometry, and T cell immunology (XPresident ®). The approach assures that only peptides truly present on a high percentage of tumors but not or only minimally presented on normal tissue, are chosen for further analysis. For initial peptide selection, colorectal cancer, glioblastoma, gastric cancer, hepatocellular carcinoma, and renal cell carcinoma samples from subjects, e.g. patients, were analyzed. Further tissues and bodily fluids from healthy donors were analyzed. Both types of samples were analyzed in a stepwise approach:
1. HLA ligands from the malignant and healthy samples were identified by mass spectrometry
2. Genome-wide messenger ribonucleic acid (mRNA) expression analysis was used to identify genes over-expressed in the malignant tissue compared with a range of normal organs and tissues.
3. Identified HLA ligands were compared to gene expression data. Peptides overpresented or selectively presented on tumor tissue, preferably encoded by selectively expressed or over-expressed genes as detected in step 2 were considered suitable peptide candidates for a multi-peptide vaccine.
4. Literature research was performed in order to identify additional evidence supporting the relevance of the identified peptides as TUMAPs.
5. The relevance of over-expression at the mRNA level was confirmed by redetection of selected peptides from step 3 on tumor tissue and lack of (or infrequent) detection on healthy tissues.
6. In order to assess, whether an induction of in vivo T-cell responses by the selected peptides may be feasible, in vitro immunogenicity assays were performed using human T cells from healthy donors as well as from subjects suffering from colorectal cancer, glioblastoma, gastric cancer, hepatocellular carcinoma, and renal cell carcinoma.
In some embodiments, the one or more peptides included in the individualized pharmaceutical product is/are identified by a method that involves: (a) identifying the peptides presented by a tumor sample from the individual subject; (b) comparing the peptides identified in (a) with a collection (e.g. a database) of peptides as described above; and (c) selecting one or more peptides from the collection of peptides (database) that correlates with a tumor-associated peptide identified in the subject. For example, the peptide(s) presented by the tumor sample can be identified by: (a1 ) comparing expression data from the tumor sample to expression data from a sample of normal tissue corresponding to the tissue type of the tumor sample to identify proteins that are overexpressed or aberrantly expressed in the tumor sample; and (a2) correlating the expression data with sequences of MHC ligands bound to MHC class I and/or class II molecules in the tumor sample to identify MHC ligands derived from proteins overexpressed or aberrantly expressed by the tumor. In some embodiments, the sequences of MHC ligands are identified by eluting bound peptides from MHC molecules isolated from the tumor sample, and sequencing the eluted ligands. In some embodiments, the tumor sample and the normal tissue have been or are obtained from the same subject, (a) identifying TUMAPs presented by a tumor sample from the individual subject by the method as described above; (b) comparing the peptides identified in (a) with a warehouse of peptides that have been prescreened for immunogenicity and over presentation in tumors as compared to corresponding normal tissue; (c) selecting at least one peptide from the warehouse that correlates with a tumor-associated peptide identified in the subject; and (d) optionally, selecting at least one peptide identified de novo in (a) confirming its immunogenicity.
In one exemplary embodiment, the peptide(s) included in the individualized pharmaceutical product, e.g. the vaccine, is/are identified by: (a) identifying one or more peptides presented by a tumor sample from the individual subject; and (b) selecting at least one peptide identified de novo in (a) and confirming its immunogenicity.
Once the peptide(s) for an individualized peptide-based pharmaceutical product, e.g. vaccine, is/are selected, the pharmaceutical product is produced. The pharmaceutical product is in some embodiments a liquid formulation consisting of the individual peptides dissolved in between 20-40% DMSO. In some embodiments the individual peptides are dissolved in about 30-35% DMSO, such as about 33% DMSO.
Each peptide to be included into a product may be dissolved in DMSO. The concentration of the single peptide solutions has to be chosen depending on the number of peptides to be included into the product. The single peptide-DMSO solutions may be mixed in equal parts to achieve a solution containing all peptides to be included in the product with a concentration of ~2.5 mg/ml/peptide. The mixed solution may then for example be diluted 1 :3 with water for injection to achieve a concentration of 0.826 mg/ml/peptide in 33% DMSO. The diluted solution is typically filtered through a 0.22 pm sterile filter. A final bulk solution can thus be obtained.
The final bulk solution may be filled into vials and stored at -20°C until use. One vial may contain 700 pL solution, containing 0.578 mg of each peptide. Of this, 500 pL (approx. 400 pg per peptide) may for example be applied for intradermal injection.
Furthermore, the method may also include selecting a peptide that contains the mutation for the generation of a cellular therapeutic or for the generation of a cellular therapeutic combination.
In a further aspect, there is provided a cancer vaccine for use in a method of inducing an immune response against cancer cells of a subject. The individualized cancer vaccine contains a nucleic acid molecule that encodes a recombinant protein or peptide. The recombinant protein or peptide contains an amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59. The method of inducing an immune response involves administering the vaccine to the subject. Expression of the recombinant protein or peptide in cells of the subject results in MHC class I presentation of a peptide having a sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
The MHC class l-presented peptide having a sequence selected from the group consisting of SEQ ID NOs: 1 to 59 is capable of eliciting a CD8+ T cell response against cancer cells of the subject that present the peptide having a sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
In some embodiments administering the vaccine is done intravenously or intraperitoneally. In some embodiments administering the vaccine is done intramuscularly or subcutaneously. In some embodiments administering the vaccine is done transdermally or intranodally.
In a further aspect there is provided a method for producing an individualized anti-cancer vaccine for an individual subject. The individualized anti-cancer vaccine contains a nucleic acid molecule that encodes a recombinant protein or peptide. The recombinant protein or peptide contains an amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59. The method includes identifying one or more peptides presented
on cells in a tumor sample from the individual subject. The method furthermore includes comparing the identified peptide(s) with a collection or dataset of peptides that have been pre-screened for immunogenicity and/or over-presentation in tumor cells as compared to healthy cells, e.g. cells of normal tissues. The collection or dataset of peptides includes one or more peptides that contain a sequence selected from the group consisting of SEQ ID NO: 1 to SEQ ID NO: 59. The peptide(s) has/have a maximal length of up to 30 amino acids, including of up to 16 amino acids. In some embodiments, the peptide(s) has/have an overall length of up to 12 amino acids. In one embodiment the collection or dataset of peptides includes one or more peptides that consist of a sequence selected from the group consisting of SEQ ID NO: 1 to SEQ ID NO: 59. The method furthermore includes assessing whether the one or more peptide(s) presented on cells of the tumor sample is/are included in the collection or dataset of peptides. If one or more of the peptide(s) is/are included in the collection or dataset of peptides, a nucleic acid molecule that encodes the recombinant protein or peptide is selected for inclusion into the individualized anti-cancer vaccine. The method in this case furthermore includes manufacturing and/or producing the individualized anti-cancer vaccine.
In some embodiments, the cancer vaccine is an individualized cancer vaccine. In such embodiments cancer cells, but not healthy cells, of the subject present a peptide containing an amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59. In some embodiments cancer cells, but not healthy cells, of the subject present a peptide consisting of an amino acid sequence selected from the group consisting of SEQ ID NOs: 1 to 59.
In some embodiments, the immunogenicity of the peptides included in the collection of peptides is determined by a method that involves an in vitro immunogenicity assay and/or subject immunomonitoring for individual MHC binding. In some embodiments, the immunogenicity of the peptides included in the collection of peptides is determined by a method that involves MHC multimer staining and/or an ELISPOT assay. In some embodiments, the immunogenicity of the peptides included in the collection of peptide is determined by a method that involves intracellular cytokine staining
In some embodiments of the method for producing an individualized anti-cancer vaccine for an individual subject the peptides included in the collection are identified based on the following steps:
- performing genome-wide messenger ribonucleic acid (mRNA) expression analysis on a sample obtained from the subject by highly parallel methods, such as microarrays or sequencing-based expression profiling, comprising identifying genes that are overexpressed in a malignant tissue, compared to a normal tissue or tissues;
- selecting peptides encoded by selectively expressed or over-expressed genes as detected in the genome-wide mRNA expression analysis, and
- determining an induction of an in vivo T-cell response by the peptides as selected comprising in vitro immunogenicity assays using human T lymphocytes from a healthy donor or from the subject.
In some embodiments of the method for producing an individualized anti-cancer vaccine for an individual subject the peptides included in the collection of peptides are identified based on the following steps:
- identifying MHC ligands from the tumor sample using mass spectrometry;
- performing genome-wide messenger ribonucleic acid (mRNA) expression analysis by highly parallel methods, such as microarrays or sequencing-based expression profiling, comprising identifying genes that are over-expressed in a malignant tissue, compared with a normal tissue or tissues;
- comparing the identified MHC ligands to the gene expression data;
- selecting peptides encoded by selectively expressed or over-expressed genes as detected in the comparison to the gene expression data;
- re-detecting selected peptides from the selecting step on tumor tissue and detecting lack of or infrequent detection on healthy tissue and confirming the relevance of overexpression at the mRNA level; and
- determining an induction of an in vivo T-cell response by the peptides as selected comprising an in vitro immunogenicity assay using human T lymphocytes from a healthy donor or the subject.
Safeguard - Proviso
The contents of the articles, patents, and patent applications, and all other documents and electronically available information mentioned or cited herein, are hereby incorporated by reference in their entirety to the same extent as if each individual publication was specifically and individually indicated to be incorporated by reference. In case of conflict, the present specification, including definitions, will control. Applicant reserves the right to physically incorporate into this application any and all materials and information from any such articles, patents, patent applications, or other physical and electronic documents. Nothing herein is to be construed as an admission that the present invention is not entitled to antedate such disclosure by virtue of prior invention. No admission is made that any reference constitutes prior art. The discussion of references states what their authors assert, and the applicant reserves the right to challenge the accuracy and pertinency of the cited documents.
The listing or discussion of a previously published document in this specification should not necessarily be taken as an acknowledgement that the document is part of the state of the art or is common general knowledge.
While the above detailed description has shown, described, and pointed out novel peptides and features as applied to various embodiments, it will be understood that various omissions, substitutions, and changes in the form and details of the uses, methods and compositions illustrated may be made by those skilled in the art without departing from principles described herein. The foregoing description is of the best mode presently contemplated. This description is in no way meant to be limiting, but rather should be taken as illustrative of the general principles described herein. The scope of the disclosure should be determined with reference to the claims.
The present invention will now be described in the following examples which describe preferred embodiments thereof, and with reference to the accompanying figures, nevertheless, without being limited thereto. For the purposes of the present invention, all references as cited herein are incorporated by reference in their entireties.
Further, note that experimental data and figures may only be disclosed herein for a selected set of peptides as claimed. Although for all peptides disclosed and claimed herein, complete data sets have been generated and can be made available upon request, applicant has decided to not incorporate herein all these complete date sets, because this would go beyond a manageable scope of this application text.
EXAMPLES
EXAMPLE 1
Identification and quantitation of tumor associated peptides presented on the cell surface
Tissue samples
Patients’ tissues were obtained from:
BiolVT (Detroit, Ml, USA & Royston, Herts, UK); Bio-Options Inc. (Brea, CA, USA); BioServe (Beltsville, MD, USA); Capital BioScience Inc. (Rockville, MD, USA); Conversant Bio (Huntsville, AL, USA); Cureline Inc. (Brisbane, CA, USA); DxBiosamples (San Diego, CA, USA); Geneticist Inc. (Glendale, CA, USA); Indivumed GmbH (Hamburg, Germany); Kyoto Prefectural University of Medicine (KPUM) (Kyoto, Japan); Osaka City University (OCU) (Osaka, Japan); ProteoGenex Inc. (Culver City, CA, USA); Tissue Solutions Ltd (Glasgow, UK); Universitat Bonn (Bonn, Germany); Asklepios Clinic St. Georg (Hamburg, Germany); Vai d'Hebron University Hospital (Barcelona, Spain);
Center for cancer immune therapy (CCIT), Herlev Hospital (Herlev, Denmark); Leiden University Medical Center (LUMC) (Leiden, Netherlands); Istituto Nazionale Tumori "Pascale", Molecular Biology and Viral Oncology Unit (Naples, Italy); Stanford Cancer Center (Palo Alto, CA, USA); University Hospital Geneva (Geneva, Switzerland); University Hospital Heidelberg (Heidelberg, Germany); University Hospital Munich (Munich, Germany); University Hospital Tuebingen (Tuebingen, Germany).
Written informed consents of all patients had been given before surgery or autopsy. Tissues were shock-frozen immediately after excision and stored until isolation of peptides/TUMAPs at -70°C or below.
Isolation of HLA peptides from tissue samples
HLA peptide pools from shock-frozen tissue samples were obtained by immune precipitation from solid tissues according to a slightly modified protocol (Falk et al. , 1991 ; Seeger et al., 1999) using the HLA-A*02 specific antibody BB7.2, the HLA-A, -B, -C specific antibody w6/32, the HLA-DR specific antibody L243 and the HLA-DP specific antibody B7/21 , CNBr-activated sepharose, acid treatment, and ultrafiltration.
Table 4 shows the peptides and the HLA allotypes - from the group consisting of HLA- A*01 :01 , HLA-A*02:01 , HLA-A*03:01 , HLA-A*24:02, HLA-B*07:02, HLA-B*08:01 and HLA-B*44:02 - they bind to. However, due to similarities in binding pattern such as the relevant anchoring positions some peptides bind to more than one allele, such an overlap is most likely, but not limited to, HLA-A*01 binding peptides also binding to HLA-B*15, HLA-A*03 binding peptides also binding to HLA-A*11 , HLA-B*07 binding peptides also binding to HLA-B*35 and HLA-B*51.
Mass spectrometry analyses
The HLA peptide pools as obtained were separated according to their hydrophobicity by reversed-phase chromatography (nanoAcquity UPLC system, Waters) and the eluting peptides were analyzed in LTQ velos and fusion hybrid mass spectrometers (ThermoElectron) equipped with an ESI source. Peptide pools were loaded directly onto the analytical fused-silica micro-capillary column (75 pm i.d. x 250 mm) packed with 1.7 pm C18 reversed-phase material (Waters) applying a flow rate of 400 nL per minute. Subsequently, the peptides were separated using a two-step 180 minute-binary gradient from 10% to 33% B at a flow rate of 300 nL per minute. The gradient was composed of solvent A (0.1 % formic acid in water) and solvent B (0.1 % formic acid in acetonitrile). A gold coated glass capillary (PicoTip, New Objective) was used for introduction into the nanoESI source. The LTQ-Orbitrap mass spectrometers were operated in the data- dependent mode using a TOP5 strategy. In brief, a scan cycle was initiated with a full scan of high mass accuracy in the orbitrap (R = 30000), which was followed by MS/MS
scans also in the orbitrap (R = 7500) on the 5 most abundant precursor ions with dynamic exclusion of previously selected ions. Tandem mass spectra were interpreted by SEQUEST at a fixed false discovery rate (q<0.05) and additional manual control.
Contaminating peptides were removed from the analysis as follows: Peptides not related to antigen presentation but arising from proteolytic cleavage via endogenous proteases were removed from subsequent analyses using statistical methods based on protein coverage ratio, peptide coverage ratio and HLA ligand binding propensity. Peptides arising from electrospray ionization within the mass spectrometer were detected by searching for peptide pairs where one peptide sequence is a substring of a longer peptide sequence and both peptides are coeluting from the chromatography. In cases where the identified peptide sequence was uncertain it was additionally validated by automatic and manual comparison of the generated natural peptide fragmentation pattern with the fragmentation pattern of a synthetic sequence-identical reference peptide. To avoid chromatographic carry-overs effects between different samples analyzed, monitored blank runs are run between analytical runs and further system suitability checks are performed prior to any analytical sample acquisition.
Label-free relative LC-MS quantitation was performed by ion counting i.e. , by extraction and analysis of LC-MS features (Mueller et al., 2007). The method assumes that the peptide’s LC-MS signal area correlates with its abundance in the sample. Extracted features were further processed by charge state deconvolution and retention time alignment (Mueller et al., 2008; Sturm et al., 2008). Finally, all LC-MS features were cross-referenced with the sequence identification results to combine quantitative data of different samples and tissues to peptide presentation profiles. The quantitative data were normalized in a two-tier fashion according to central tendency to account for variation within technical and biological replicates. Thus, each identified peptide can be associated with quantitative data allowing relative quantification between samples and tissues. In addition, all quantitative data acquired for peptide candidates was inspected manually to assure data consistency and to verify the accuracy of the automated analysis. For each peptide a presentation profile was calculated showing the mean sample presentation as well as replicate variations. The profiles juxtapose AML (acute myeloid leukemia); BRCA (breast cancer); CCC (cholangiocellular carcinoma); CLL (chronic lymphocytic leukemia); CRC (colorectal cancer); GBC (gallbladder cancer); GBM (glioblastoma); GC (gastric cancer); GEJC (gastro-esophageal junction cancer); HCC (hepatocellular carcinoma); HNSCC (head and neck squamous cell carcinoma); MEL (melanoma); NHL (non-Hodgkin lymphoma); NSCLCadeno (non-small cell lung cancer adenocarcinoma); NSCLCother (NSCLC samples that could not unambiguously be assigned to NSCLCadeno or NSCLCsquam); NSCLCsquam (squamous cell non-small cell lung
cancer); OC (ovarian cancer); OSCAR (esophageal cancer); PACA (pancreatic cancer); PRCA (prostate cancer); RCC (renal cell carcinoma); SCLC (small cell lung cancer); UBC (urinary bladder carcinoma); UEC (uterine endometrial cancer) samples to a baseline of normal tissue samples. Presentation profiles of exemplary over-presented peptides are shown in Figures 1A-1 G. The plots show only those identifications of peptides as dots which were made on tissue samples positive for the respective HLA allotype which were processed using HLA specific antibodies.
Peptide presentation on the various indications for all peptides (SEQ ID NO: 1 to SEQ ID NO: 59) are shown in Table 5. This table lists all indications on which the respective peptide was identified at least once, independent of the HLA typing of the sample or the antibody used to process said sample.
Table 5: Presentation on various cancer entities for peptides according to the invention, and thus the particular relevance of the peptides as mentioned for the diagnosis and/or treatment of the cancers as indicated. Cancer type: AML (acute myeloid leukemia); BRCA (breast cancer); CCC (cholangiocellular carcinoma); CLL (chronic lymphocytic leukemia); CRC (colorectal cancer); GBC (gallbladder cancer); GBM (glioblastoma); GC (gastric cancer); GEJC (gastro-esophageal junction cancer); HCC (hepatocellular carcinoma); HNSCC (head and neck squamous cell carcinoma); MEL (melanoma); NHL (non-Hodgkin lymphoma); NSCLCadeno (non-small cell lung cancer adenocarcinoma); NSCLCother (NSCLC samples that could not unambiguously be assigned to NSCLCadeno or NSCLCsquam); NSCLCsquam (squamous cell non-small cell lung cancer); OC (ovarian cancer); OSCAR (esophageal cancer); PACA (pancreatic cancer); PRCA (prostate cancer); RCC (renal cell carcinoma); SCLC (small cell lung cancer); UBC (urinary bladder carcinoma); UEC (uterine endometrial cancer).
EXAMPLE 2
Expression profiling of genes encoding the peptides of the invention
Over-presentation or specific presentation of a peptide on tumor cells compared to normal cells is sufficient for its usefulness in immunotherapy, and some peptides are tumor-specific despite their source protein occurring also in normal tissues. Still, mRNA expression profiling adds an additional level of safety in selection of peptide targets for immunotherapies. Especially for therapeutic options with high safety risks, such as
affinity-matured TCRs, the ideal target peptide will be derived from a protein that is unique to the tumor and not found on normal tissues.
RNA sources and preparation
Surgically removed tissue specimens were provided as indicated above (see Example 1 ) after written informed consent had been obtained from each patient. Tumor tissue specimens were snap-frozen immediately after surgery and later homogenized with mortar and pestle under liquid nitrogen. Total RNA was prepared from these samples using TRI Reagent (Ambion, Darmstadt, Germany) followed by a cleanup with RNeasy (QIAGEN, Hilden, Germany); both methods were performed according to the manufacturer's protocol.
Total RNA from healthy human tissues for RNASeq experiments was obtained from: Asterand (Detroit, Ml, USA & Royston, Herts, UK); Bio-Options Inc. (Brea, CA, USA); Geneticist Inc. (Glendale, CA, USA); ProteoGenex Inc. (Culver City, CA, USA); Tissue Solutions Ltd (Glasgow, UK).
Total RNA from tumor tissues for RNASeq experiments was obtained from: Asterand (Detroit, Ml, USA & Royston, Herts, UK); BioCat GmbH (Heidelberg, Germany); BioServe (Beltsville, MD, USA); Geneticist Inc. (Glendale, CA, USA); Istituto Nazionale Tumori "Pascale" (Naples, Italy); ProteoGenex Inc. (Culver City, CA, USA); University Hospital Heidelberg (Heidelberg, Germany).
Quality and quantity of all RNA samples were assessed on an Agilent 2100 Bioanalyzer (Agilent, Waldbronn, Germany) using the RNA 6000 Pico LabChip Kit (Agilent).
RNAseq experiments
Gene expression analysis of tumor and normal tissue RNA samples was performed by next-generation sequencing (RNAseq) by GENEWIZ Germany GmbH (Leipzig, Germany). Briefly, sequencing libraries were prepared from total RNA using the NEBNext® Ultra™ II Directional RNA Library Prep Kit for Illumina according to the manufacturer’s instructions (New England Biolabs, Ipswich, MA, USA), which includes mRNA selection, RNA fragmentation, cDNA conversion and addition of sequencing adaptors. For sequencing, libraries were multiplexed and loaded onto the Illumina NovaSeq 6000 sequencer (Illumina Inc., San Diego, CA, USA) according to the manufacturer’s instructions, generating a minimum of 80 million 150 bp paired-end raw reads per sample. After quality control, adapter trimming and mapping to the reference genome, RNA reads supporting the peptide were counted and are shown as exemplary expression profiles of peptides of the present invention that are highly overexpressed or
exclusively expressed in AML (acute myeloid leukemia); BRCA (breast cancer); CCC (cholangiocellular carcinoma); CLL (chronic lymphocytic leukemia); CRC (colorectal cancer); GBC (gallbladder cancer); GBM (glioblastoma); GC (gastric cancer); GEJC (gastro-esophageal junction cancer); HCC (hepatocellular carcinoma); HNSCC (head and neck squamous cell carcinoma); MEL (melanoma); NHL (non-Hodgkin lymphoma); NSCLCadeno (non-small cell lung cancer adenocarcinoma); NSCLCother (NSCLC samples that could not unambiguously be assigned to NSCLCadeno or NSCLCsquam); NSCLCsquam (squamous cell non-small cell lung cancer); OC (ovarian cancer); OSCAR (esophageal cancer); PACA (pancreatic cancer); PRCA (prostate cancer); RCC (renal cell carcinoma); SCLC (small cell lung cancer); UBC (urinary bladder carcinoma); UEC (uterine endometrial cancer) in Figures 2A-2G. Furthermore, expression scores from the RNAseq experiments are listed in Table 6.
Table 6: Expression scores. The table lists peptides that are very highly overexpressed in tumors compared to a panel of normal tissues (+++) or highly overexpressed in tumors compared to a panel of normal tissues (++) or overexpressed in tumors compared to a panel of normal tissues (+) according to the RNAseq experiments. The baseline for this score was calculated from measurements of the following relevant normal tissues: adipose tissue; adrenal gland; bile duct; bladder; bloodcells; blood vessels; bone marrow; brain; breast; esophagus; eye; gallbladder; nead&neck; heart; large intestine; small intestine; kidney; liver; lung; lymph nodes; peripheral nerve; ovary; pancreas; parathyroid gland; peritoneum; pituitary; placenta; pleura; prostate; skeletal muscle; skin; spinal cord; spleen; stomach; testis; thymus; thyroid; trachea; ureter; uterus. In case expression data for several samples of the same tissue type were available, the arithmetic mean of all respective samples was used for the calculation.
EXAMPLE 3
Validation of peptides by IdentControl and CoElution
In order to validate the peptides according to the invention, all peptides were synthesized using standard and well-established solid phase peptide synthesis using the Fmoc- strategy. If necessary, stable isotope labeled (SIL-) amino acids were used to introduce a discriminating mass shift and allow for the use of these labeled peptides as internal standards (e.g. if a peptide was selected for identity confirmation in CoElution experiments). Identity and purity of each individual peptide were determined by mass spectrometry and analytical RP-HPLC. The peptides were obtained as white to off-white lyophilizes (trifluoro acetate salt) in purities of >50%. Tumor-associated peptides (TUMAPs) are usually administered as trifluoro-acetate salts or acetate salts, other saltforms are also possible.
The initial validation of peptides was achieved by IdentControl via spectral comparison. For this, synthetic peptides were used for validation of peptide identifications by acquisition of high-resolution reference MS2 spectra using matched fragmentation modes and collision energies as for acquisition of the natural spectra. Automated spectral comparison was performed using the sensitive metric of spectral correlation with a cutoff score determined to result in 90% sensitivity at <1 % FDR based on a benchmark dataset comprising >10,000 manually validated spectra. Ambiguous identifications were further subjected to validation in CoElution experiments.
A list of peptides validated by Identcontrol can be found in Table 7 and an exemplary validation via IdentControl for SEQ ID NO: 44 is shown in Figure 3.
Table 7: Identcontrol Results. The spectral correlation indicates the similarity of the MS/MS spectra from the endogenous detected peptide compared to the synthetic peptide, the higher the value the more alike the spectra are. The peptide is validated when a threshold of 0.872 is met, or spectra are considered identical by manual review.
For further validation, peptides were subjected to CoElution experiments using SIL internal standard peptides. To this end, SIL peptides were spiked into HLA peptidome extracts from samples and subjected to liquid chromatography - targeted mass spectrometry (LC-MS) to confirm peptide identity based on spectral similarity as well as CoElution in the retention time dimension. Spiked SIL-peptide amounts were adjusted to the peptide specific ionization factors (determined in calibration curves), if necessary. LC-MS was performed using pre-defined targeted MS2 scan events with non-overlapping isolation windows for SIL-peptide and natural peptide species to avoid co-fragmentation. To confirm isotopic purity and absence of co-fragmentation of SIL- and natural peptide, control experiments were performed in a non-HLA peptide containing tryptic matrix, which had to confirm absence of any unlabeled signal. Peptide detection and validation by CoElution was determined by manual expert review based on multiple pre-defined objective criteria, including dot product (dotP) of SIL peptide compared to unlabeled peptide MS2 traces, the presence of the most intense transitions in multiple consecutive scans and aligned peak apexes. A list of peptides validated by CoElution can be found in Table 8 and the CoElution result for SEQ ID NO 58 is shown in Figure 4.
EXAMPLE 4:
Absolute quantitation of tumor-associated peptides presented on cell surface
The generation of binders, such as antibodies and/or TCRs, is a laborious process, which may be conducted only for a number of selected targets. In the case of tumor-associated and -specific peptides, selection criteria include, but are not restricted to, (i) exclusiveness of presentation and (ii) the density of peptide presented on the cell surface. In addition to the isolation and relative quantitation of peptides as described in the examples, the inventors analyzed absolute peptide copies per cell as described in WO 2016/107740. The quantitation of peptide copies per cell in solid tumor samples requires the absolute quantitation of the isolated peptide, the efficiency of the peptide isolation process, and the cell count of the tissue sample analyzed.
Peptide quantitation by nano LC-MS/MS
For an accurate quantitation of peptides by mass spectrometry, a calibration curve was generated for SEQ ID NO: 44 and SEQ ID NO: 48, using two different isotope labeled
peptide variants (one or two isotope-labeled amino acids are included during peptide synthesis). These isotope-labeled variants differ from the tumor-associated peptide only in their mass but show no difference in other physicochemical properties (Anderson et al., 2012). For the peptide calibration curve, a series of nano LC-MS/MS measurements was performed to determine the ration of MS/MS signals of titrated (singly isotopelabeled peptide) to constant (doubly isotope-labeled peptide) isotope-labeled peptides.
The doubly isotope-labeled peptide, also called internal standard, was further spiked to each MS sample and all MS signals were normalized to the MS signal of the internal standard to level out potential technical variances between MS experiments.
The calibration curves were prepared in at least three different matrices, i.e., HLA peptide eluates from natural samples similar to the routine MS samples, and each preparation was measured in duplicate MS runs. For evaluation, MS signals were normalized to the signal of the internal standard and a calibration curve was calculated by logistic regression.
For the quantitation of tumor-associated peptides from tissue samples, the respective samples were also spiked with the internal standard; the MS signals were normalized to the internal standard and quantified using the peptide calibration curve.
Efficiency of peptide-MHC isolation
As for any protein purification process, the isolation of proteins from tissue samples is associated with a certain loss of the protein of interest. To determine the efficiency of peptide isolation, peptide-MHC complexes were generated for all peptides selected for absolute quantitation. To be able to discriminate the spiked from the natural peptide-MHC complexes, single-isotope-labelled versions of the peptides were used, i.e., one isotopelabelled amino acid was included in peptide synthesis. These complexes were spiked into the freshly prepared tissue lysates, i.e., at the earliest possible point of the peptide isolation procedure, and then captured like the natural peptide-MHC complexes in the following affinity purification. Measuring the recovery of the single-labelled peptides therefore allows conclusions regarding the efficiency of isolation of individual natural peptides.
The efficiency of isolation was analyzed in a small set of samples and was comparable among these tissue samples. In contrast, the isolation efficiency differs between individual peptides. This suggests that the isolation efficiency, although determined in only a limited number of tissue samples, may be extrapolated to any other tissue preparation. However, it is necessary to analyze each peptide individually as the isolation efficiency may not be extrapolated from one peptide to others.
Determination of the cell count in solid, frozen tissue
In order to determine the cell count of the tissue samples subjected to absolute peptide quantitation, the inventors applied DNA content analysis. This method is applicable to a wide range of samples of different origin and, most importantly, frozen samples (Alcoser et al. , 2011 ; Forsey and Chaudhuri, 2009; Silva et al. , 2013). During the peptide isolation protocol, a tissue sample is processed to a homogenous lysate, from which a small lysate aliquot is taken. The aliquot is divided in three parts, from which DNA is isolated (QiaAmp DNA Mini Kit, Qiagen, Hilden, Germany). The total DNA content from each DNA isolation is quantified using a fluorescence-based DNA quantitation assay (Qubit dsDNA HS Assay Kit, Life Technologies, Darmstadt, Germany) in at least two replicates.
In order to calculate the cell number, a DNA standard curve from aliquots of isolated healthy blood cells from several donors, with a range of defined cell numbers, has been generated. The standard curve is used to calculate the total cell content from the total DNA content from each DNA isolation. The mean total cell count of the tissue sample used for peptide isolation is then extrapolated considering the known volume of the lysate aliquots and the total lysate volume.
Peptide copies per cell
With data of the aforementioned experiments, the inventors calculated the number of peptide copies per cell by dividing the total peptide amount by the total cell count of the sample, followed by division through isolation efficiency. Copy cell numbers for SEQ ID NO: 44 are shown in Table 9A and Figure 5A and for SEQ ID NO 48 in Table 9B and Figure 5B.
Tables 9A and 9B list the results of absolute peptide quantitation in cancer samples.
The number of samples, in which evaluable, high quality MS data are available, is indicated.
A more elaborate disclosure of the method to absolutely quantify the peptides is disclosed in international patent publication W02016107740A1 and US patent 10,545,154, the content of both of which is incorporated herein by reference.
REFERENCE LIST
Alcoser, S. Y. et al., BMC.Biotechnol. 11 (2011 ): 124.
Allison, J. P. et al., Science 270 (1995): 932-933.
Appay, V. et al., Eur.J Immunol. 36 (2006): 1805-1814.
Baibas R and Lorence A. “Methods in Molecular Biology Recombinant Gene Expression, Reviews and Protocols” (2004).
Banchereau, J. et al., Cell 106 (2001 ): 271 -274.
Banker, G. and Rhodes, C. “Modern Pharmaceutics”, CRC Press (2002).
Berg, J. M. et al, “Biochemistry”, 8th ed. (2015), W. H. Freeman and Company, 995-999.
Beggs, J. D. et al., Nature 275 (1978): 104-109.
Benjamini, Y. et al., Journal of the Royal Statistical Society. Series B (Methodological), Vol.57 (1995): 289-300.
Berge, S. M. et al., Journal of Pharmaceutical Science 66 (1977): 1 -19.
Better, M. et al., Science 240 (1988): 1041 -1043.
Boulianne, G. L. et al. Nature 312 (1984): 643-646.
Boulter, J. M. et al., Protein Eng 16 (2003): 707-711.
Brossart, P. et al., Blood 90 (1997): 1594-1599.
Bruckdorfer, T. et al., Curr.Pharm.Biotechnol. 5 (2004): 29-43.
Cabilly, S. et al. Proc Natl Acad Sci 81 (1984): 3273-3277.
Card, K. F. et al., Cancer Immunol. Immunother. 53 (2004): 345-357.
Cohen, C. J. et al., J Immunol. 170 (2003b): 4349-4361.
Cohen, C. J. et al., J Mol. Recognit. 16 (2003a): 324-332.
Cohen, S. N. et al., Proc.Natl.Acad.Sci. U.S.A. 69 (1972): 2110-2114.
Coligan, J. E. et al., Current Protocols in Protein Science (1995).
Colombetti, S. et al., J Immunol. 176 (2006): 2730-2738
Dayhoff, M. O. et al. „The Atlas of Protein Sequence and Structure” Natl Biomedical Research (1965).
Dengjel, J. et al., Clin Cancer Res 12 (2006): 4163-4170.
Denkberg, G. et al., J Immunol. 171 (2003): 2197-2207.
Denkberg, G, & Reiter, Y, Autoimmun. Rev. 5(4) (2006): 252-7.
De Simone, M. et al., Front. Immunol. 9 (2018) Article 1638; doi: 10.3389/fimmu.2018.01638.
Falk, K. et al., Nature 351 (1991 ): 290-296.
Follenzi, A. et al., Nat Genet. 25 (2000): 217-222.
Fong, L. et al., Proc.Natl.Acad.Sci.U.S.A 98 (2001 ): 8809-8814.
Forsey, R. W. et al., Biotechnol.Lett. 31 (2009): 819-823.
Gabrilovich, D. I. et al., Nat.Med 2 (1996): 1096-1103.
Gattinoni, L. et al., Nat.Rev.lmmunol. 6 (2006): 383-393.
Gennaro, A. “Remington’s: The Science and Practice of Pharmacy”, Lippincott Williams & Wilkins (1997).
Gilliland, D. G. et al., Proc Natl Acad Sci 77 (1980): 4539
Green, M. R. et al., Molecular Cloning, A Laboratory Manual 4th (2012).
Greenfield, E. A., Antibodies: A Laboratory Manual 2nd (2014).
Guo, H. H. et al., Proc Natl Acad Sci 101 (25): 9205-9210 (2004).
Gustafsson, C. et al., Trends Biotechnol. 22 (2004): 346-353.
Gouttefangeas, C., and Rammensee, H. G., Cancer Immunol Immunother 67 (2018): 1911 -18.
Harlow, E. & Lane, D. “Using Antibodies - A laboratory manual”, Cold Spring Harbor Laboratory, (1989).
He, E., et al., “TCR-like antibodies in cancer immunotherapy”, Journal of Hematology & Oncology 12 (2019): 99
Holliger, P. et al. Proc Natl Acad Sci, 90 (1993): 6444-6448.
Howard, G. C. & Kaser, M. R. “Making and Using antibodies: A practical handbook”, CRC
Press (2007)
Kibbe, A. H., Handbook of Pharmaceutical Excipients rd (2000).
Krieg, A. M., Nat. Rev. Drug Discov. 5 (2006): 471 -484.
Krolick, K. A. et al., Proc Nat’l Acad Sci 77 (1980): 5419.
Kuball, J. et al., Blood 109 (2007): 2331 -2338.
La Gruta, N. L. et al., Nat. Rev. Immunology 18(7) (2018): 467-478.
Liddy, N. et al., Nat.Med. 18 (2012): 980-987.
Liu, et al., Proc Natl Acad Sci 84 (1987):3439-3443
Ljunggren, H. G. et al., J Exp.Med 162 (1985): 1745-1759.
Longenecker, B. M. et al., Ann N.Y.Acad.Sci. 690 (1993): 276-291
Lonsdale, J., Nat.Genet. 45 (2013): 580-585
Lukas, T. J. et al., Proc.Natl.Acad.Sci. U.S.A. 78 (1981 ): 2791 -2795
Lundblad, R. L., Chemical Reagents for Protein Modification 3rd (2004)
Meziere, C. et al., J Immunol 159 (1997): 3230-3237.
Mondon R et al Front BioSci 13 (2008): 1117-1129, 1008.
Morgan, R. A. et al., Science 314 (2006): 126-129.
Morrison S. L. et al., Proc Natl Acad Sci (1984): 6851 -6855.
Mueller, L. N. et al., J Proteome.Res. 7 (2008): 51 -61.
Mueller, L. N. et al., Proteomics. 7 (2007): 3470-3480.
Murphy, K. et al.: “Janeway's Immunobiology”, 10th ed., W W Norton & Co Inc., (2022), 13-14, 159-173, 196-201 , 218-226, 369-372, 394-399 and 764-771.
Olexiouk, V. et al., Nucleic Acids Res 44 (2016): D324-D329.
Pinheiro, J. et al., nlme: Linear and Nonlinear Mixed Effects Models (CRAN.R- project.org/packe=nlme) (2015)
Piebanski, M. et al., Eur.J Immunol 25 (1995): 1783-1787.
Porta, C. et al., Virology 202 (1994): 949-955.
Rammensee, H. G. et al., Immunogenetics 50 (1999): 213-219
Rammensee, H. G. et al., Journal for Immunotherapy of Cancer 7 (2019): 307.
Rini, B. I. et al., Cancer 107 (2006): 67-74.
Rock, K. L. et al., Science 249 (1990): 918-921.
Russell, S. J. et al., Nucl Acids Res 21 (1993): 1081 -1085.
Saiki, R. K. et al., Science 239 (1988): 487-491
Schmitt, T. M. et al., Hum. Gene Then 20 (2009): 1240-1248.
Scholten, K. B. et al., Clin Immunol. 119 (2006): 135-145.
Seeger, F. H. et al., Immunogenetics 49 (1999): 571 -576.
Sherman, F. et al., Laboratory Course Manual for Methods in Yeast Genetics (1986).
Silva, L. P. et al., Anal.Chem. 85 (2013): 9536-9542.
Singh-Jasuja, H. et al., Cancer Immunol. Immunother. 53 (2004): 187-195.
Small, E. J. et al., J Clin Oncol. 24 (2006): 3089-3094.
Sternberg, N. et al. Proc Natl Acad Sci 92 (1995): 1609-1613.
Sturm, M. et al., BMC. Bioinformatics. 9 (2008): 163.
Subramanian, R. P. et al., Retrovirology. 8 (2011 ): 90.
Sun, L. K. et al., Proc Natl Acad Sci 84 (1987): 214-218.
Teufel, R. et al., Cell Mol. Life Sci. 62 (2005): 1755-1762.
Traunecker, A. et al., EMBO 10 (1991 ): 3655-3659.
Unverdorben, F. et al., Protein Engineering, Design & Selection 25 (2012): 81 -88
Walter, S. et al., J. Immunol. 171 (2003): 4974-4978.
Walter, S. et al., Nat Med. 18 (2012): 1254-1261.
Willcox, B. E. et al., Protein Sci. 8 (1999): 2418-2423.
Youle, R. J. et al., Proc Natl Acad Sci 77 (1980): 5483-5486.
Zaremba, S. et al., Cancer Res. 57 (1997): 4570-4577.
Zufferey, R. et al., J Virol. 73 (1999): 2886-2892.
Claims
1. A peptide comprising an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59, or a pharmaceutically acceptable salt thereof, wherein the peptide has an overall length of up to 16 amino acids.
2. The peptide, or the salt thereof, according to claim 1 , wherein
• said peptide has the ability to bind to an MHC class I molecule, and/or
• wherein said peptide, when bound to said MHC, is capable of being recognized by CD8 T cells.
3. The peptide or salt thereof according to claim 1 or 2, wherein said peptide
(i) includes one or more non-peptide bonds, wherein the one or more non-peptide bonds are optionally selected from -CH2-NH, -CH2S-, -CH2CH2-, -CH=CH-, -COCH2-, - CH(OH)CH2-, -CH2SO-, an N-methyl group or an N-hydroxyl group, and/or
(ii) wherein said pharmaceutically acceptable salt is one of a chloride salt, a bromide salt, an iodide salt, a thiocyanate salt, a phosphate salt, a phosphonium salt, a nitrate salt, a sulfate salt, a chlorate salt, an acetate salt, a trifluoroacetate salt, a propionate salt, a butyrate salt, a pyridinium salt, a glycolate salt, a pyruvate salt, an oxalate salt, a malate salt, a maleate salt, a malonate salt, a succinate salt, a fumarate salt, a tartrate salt, a citrate salt, a benzoate salt, a cinnamate salt, a mandelate salt, a salicylic acid salt, a methane sulfonate salt, an ethane sulfonate salt, a p-toluenesulfonate salt, a salicylate salt, a sodium salt, a potassium salt, a barium salt, a zinc salt, a lithium salt, a rubidium salt, a cesium salt, a magnesium salt, a manganese salt, a copper salt, an ammonium salt, a calcium salt or a trimethylamine salt, wherein the salt is preferably selected from a chloride salt, an acetate salt, a trifluoroacetate salt or a combination thereof.
4. The peptide or salt thereof according to any one of claims 1 to 3, wherein said peptide is part of a fusion protein.
5. A binding molecule that specifically recognizes a peptide comprising an amino acid sequence selected from the group consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59, or said peptide comprising an amino acid sequence selected from the group
consisting of any one of SEQ ID NO: 1 to SEQ ID NO: 59 when bound to an MHC molecule, wherein the binding molecule is preferably a proteinaceous binding molecule or an aptamer.
6. The binding molecule according to claim 5, being
(a) an antibody, or a functional fragment thereof, being a soluble or a membrane-bound antibody; or
(b) a T cell receptor (TCR), or a functional fragment thereof, being soluble or membranebound, wherein the antibody or functional fragment thereof optionally is a monoclonal antibody, a bi-specific antibody, a chimeric antibody, a human antibody and/or a humanized antibody or a fragment thereof; or
(c) an aptamer.
7. A nucleic acid comprising a sequence encoding the peptide according to claim 1 or 2 or the binding molecule according to claim 6, being a proteinaceous binding molecule, wherein the sequence is optionally operably linked to a heterologous promoter sequence.
8. An expression vector comprising the nucleic acid according to claim 7.
9. A recombinant host cell comprising the peptide according to claim 1 or 2, the binding molecule according to claim 6, and/or the nucleic acid according to claim 7.
10. An in vitro method for producing T lymphocytes that specifically recognize a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule, the method comprising (a) contacting in vitro T lymphocytes with antigen loaded human class I MHC molecules expressed on the surface of a suitable antigen- presenting cell or an artificial construct mimicking an antigen-presenting cell for a period of time sufficient to activate the T cells in an antigen specific manner, wherein said antigen is the peptide according to claim 1 or 2 or (b) introducing in vitro into T lymphocytes a nucleic acid encoding a TCR that specifically recognizes a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a peptide
comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule.
11. A T lymphocyte, produced by the method according to claim 10, that specifically recognizes a cell which presents a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, wherein the T lymphocyte is optionally an activated T lymphocyte.
12. A pharmaceutical composition comprising one or more active ingredients selected from the group consisting of
• the peptide according to any one of claims 1 to 4,
• a pharmaceutically acceptable salt of the peptide according to any one of claims 1 to 4,
• the binding molecule according to claim 5 or 6,
• the nucleic acid according to claim 7,
• the expression vector according to claim 8,
• the recombinant host cell according to claim 9,
• or the T lymphocyte according to claim 11 , and a pharmaceutically acceptable carrier or a pharmaceutically acceptable excipient.
13. The pharmaceutical composition according to claim 12, wherein the pharmaceutical composition is a vaccine.
14. The pharmaceutical composition according to claim 12 or 13, wherein the pharmaceutical composition further comprises an adjuvant.
15. A combination product, comprising one or more of the peptide or salt thereof according to any one of claims 1 to 4, the binding molecule according to claim 5 or 6, the nucleic acid according to claim 7, the expression vector according to claim 8 or the recombinant host cell according to claim 9 or the T lymphocyte according to claim 11 .
16. The combination product of claim 15, further comprising an adjuvant.
17. The pharmaceutical composition according to claim 14 or the combination product according to claim 15 or 16, wherein the adjuvant comprises IFN-alpha, IFN-beta or an interleukin, wherein the interleukin preferably is IL-1 , IL-2, IL-4, IL-7, IL-9, IL-10, IL-12, IL-13, IL-15, IL-17, IL-18, IL-21 , IL-23 or a combination thereof.
18. The pharmaceutical composition according to claim 14 or 17 or the combination product according to claims16 or 17, wherein the adjuvant is IL-2, IL-12, IL-15, IL-18, IL- 21 or a combination thereof.
19. The pharmaceutical composition according to any one of claims 14, 17 or 18, or the combination product according to any one of claims 16 to 18, comprising the T lymphocyte according to claim 11 and an adjuvant selected from an interleukin, preferably IL-2, IL-15 or IL-21 or a combination thereof.
20. A method for producing the peptide according to claim 1 or 2 or the binding molecule according to claim 5 or 6, being a proteinaceous binding molecule, the method comprising culturing the recombinant host cell according to claim 9, and isolating the peptide or the proteinaceous binding molecule from the host cell and/or its culture medium.
21. The peptide or the salt thereof according to any one of claims 1 to 4, the binding molecule according to claim 5 or 6, the nucleic acid according to claim 7, the expression vector according to claim 8, the recombinant host cell according to claim 9, the T lymphocyte according to claim 11 , the pharmaceutical composition according to any one of claims 12 to 14 and 17-19 or the combination product according to any one of claims 15 to 19 for use in medicine.
22. The peptide or the salt thereof, the binding molecule, the nucleic acid, the expression vector according, the recombinant host cell, the T lymphocyte, the pharmaceutical composition or the combination product for use according to claim 21 , being for use in the prevention, treatment and/or diagnosis of cancer.
23. The peptide, the binding molecule, the nucleic acid, the expression vector host cell, the T lymphocyte, the pharmaceutical composition or the combination product for use
according to claim 22, wherein said cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastroesophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer and another cancer that shows an over-presentation of a peptide according to claim 1 or 2.
24. Use of the peptide or the salt thereof according to any one of claims 1 to 4, the binding molecule according to claim 5 or 6, the nucleic acid according to claim 7, the expression vector according to claim 8, the recombinant host cell according to claim 9, the T lymphocyte according to claim 11 , the pharmaceutical composition according to any one of claims 12 to 14 and 17-19 or the combination product according to any one of claims 15 to 19 in the manufacture of a medicament for the prevention, treatment and/or diagnosis of cancer.
25. The use according to claim 24, wherein the cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer, and another cancer that shows an over-presentation of a peptide according to claim 1 or 2.
26. A method of killing target cells in a subject, wherein the target cells present a peptide according to claim 1 or 2, the method comprising administering to the subject an effective number of T lymphocytes according to claim 11 .
27. An in vitro method for killing target cells that present a peptide according to claim 1 or 2, wherein said cells are known or suspected of being cancer cells, the method
comprising contacting said cells with a plurality of T lymphocytes according to claim 11 , wherein said cells are optionally cells of a sample from a subject.
28. The T lymphocyte according to claim 11 for a) use in the killing of target cells in a subject, wherein the target cells present a peptide according to claim 1 or 2, or b) use in the manufacture of a medicament for the killing of such target cells, or c) use in the prevention or treatment of cancer, the use comprising the killing of target cancer cells in a subject, wherein the target cancer cells present a peptide according to claim 1 or 2.
29. A method of treating a subject
• being diagnosed for,
• suffering from and/or
• being at risk of developing cancer, the method comprising administering to the subject an effective amount of the peptide according to any one of claims 1 to 4, the binding molecule according to claim 5 or 6, the nucleic acid according to claim 7, the recombinant host cell according to claim 8, the T lymphocyte according to claim 11 , the pharmaceutical composition according to any one of claims 12 to 14 or 17 to 19 or the combination product according to any one of claims 15 to 19.
30. The method according to claim 29, wherein said cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer or another cancer that shows an over-presentation of a peptide according to claim 1 or 2.
31 . A kit comprising:
(a) a container comprising the peptide or the salt thereof according to any one of claims 1 to 4, the binding molecule according to claim 5 or 6, the nucleic acid according to claim
7, the expression vector according to claim 8, the recombinant host cell according to claim 9, the T lymphocyte according to claim 11 , the pharmaceutical composition according to any one of claims 12 to 14 and 17-19, and/or a combination product according to any one of claims claim 15 to 19 in solution or in lyophilized form;
(b) optionally, a further container containing a diluent or reconstituting solution for the lyophilized formulation; and/or
(c) optionally, a further container containing an adjuvant; and/or
(d) optionally, an additional peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59.
32. The kit according to claim 31 , further comprising one or more of a buffer, a diluent, a filter, a needle, and/or a syringe.
33. An in vitro method of diagnosing cancer in a sample comprising cells of an individual, the method comprising:
(a) adding a binding molecule to a sample from said subject, wherein the binding molecule is labeled, optionally wherein the label is a radionucleotide, and
(b) detecting whether the binding molecule binds to cells in the sample from the subject, wherein the binding molecule specifically binds to (i) a peptide comprising the amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 or (ii) said peptide comprising the amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 when bound to an MHC molecule, wherein said sample optionally comprises cells of a tissue suspected to be a cancer tissue.
34. An in vitro method of diagnosing cancer in a sample comprising cells of an individual, the method comprising:
(a) identifying one or more peptides presented on cells comprised in a tumor sample from the individual; and
(b) comparing the peptide(s) as identified in step (a) with a collection of peptides that have been pre-screened for immunogenicity and/or over-presentation in tumours as compared to normal tissues, wherein the collection of peptides comprises one or more
peptides comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, having a maximal length of 16 amino acids; and
(c) assessing whether a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 is a peptide comprised in a tumor sample from the individual, wherein said sample optionally comprises cells of a tissue suspected to be a cancer tissue.
35. The method according to claim 33 or 34, wherein said cancer is selected from the group consisting of acute myeloid leukemia, breast cancer, cholangiocellular carcinoma, chronic lymphocytic leukemia, colorectal cancer, gallbladder cancer, glioblastoma, gastric cancer, gastro-esophageal junction cancer, hepatocellular carcinoma, head and neck squamous cell carcinoma, melanoma, non-Hodgkin lymphoma, non-small cell lung cancer, ovarian cancer, esophageal cancer, pancreatic cancer, prostate cancer, renal cell carcinoma, small cell lung cancer, urinary bladder carcinoma, and uterine endometrial cancer, or another cancer that shows an over-presentation of a peptide according to claim 1 or 2.
36. A method for producing an individualized anti-cancer cellular therapeutic composition for a subject, the method comprising:
(a) determining whether a peptide comprising the amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59 is presented on cells in a sample from a subject suspected of having cancer, wherein said sample preferably comprises cancer cells from said subject;
(b) contacting in vitro a plurality of T lymphocytes from said subject with an antigen- presenting cell or an artificial antigen-presenting cell having antigen-loaded human class I MHC molecules on its surface, wherein said antigen is a peptide comprising an amino acid sequence set forth in any one of SEQ ID NOs: 1 to 59, thereby activating said T lymphocytes in an antigen specific manner if they bind to said peptide;
(c) optionally allowing proliferation of the obtained activated T lymphocytes; and
(d) isolating one or more activated T lymphocytes for manufacturing an individualized anti-cancer cellular therapeutic composition.
37. The method according to claim 36, wherein (d) manufacturing the individualized anticancer cellular therapeutic composition further comprises cloning the nucleic acid
encoding the TCR expressed by one or more activated T lymphocytes into a suitable expression vector and introducing the vector into T lymphocytes.
38. The method according to claim 36 or 37, wherein determining whether said peptide is presented on cells in a sample from the subject comprises adding a binding molecule to a sample from said subject, wherein the binding molecule is (a) labeled and (b) specifically binds to said peptide or to said peptide when bound to an MHC molecule.
39. A method of stratifying a subject for individualized vaccination and/or immunotherapy against cancer, the method comprising
(i) assessing whether cells of a cancer sample from the subject present a peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, and
(ii) selecting the subject for treatment with a vaccine and/or an immunotherapeutic agent, if said peptide comprising the sequence set forth in any one of SEQ ID NOs: 1 to 59 is presented on cells of the cancer sample, wherein said vaccine comprises a peptide that comprises a sequence set forth in any one of SEQ ID NOs: 1 to 59, or a pharmaceutically acceptable salt thereof, and wherein said immunotherapeutic agent comprises the binding molecule according to claim 5 or 6, the nucleic acid according to claim 7, the expression vector according to claim 8, the recombinant host cell according to claim 9 or the T lymphocyte according to claim 11.
40. The method according to claim 39, further comprising assessing whether the peptide comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59 is bound by an MHC molecule.
41 . A method for producing an individualized anti-cancer vaccine or an individualized compound-based and/or cellular therapeutic agent for an individual, wherein the anticancer vaccine comprises a peptide that comprises a sequence set forth in any one of SEQ ID NOs: 1 to 59, and the compound-based and/or cellular therapeutic agent is based on a peptide that comprises a sequence set forth in any one of SEQ ID NOs: 1 to 59, the method comprising:
(a) identifying one or more peptides presented on cells in a tumor sample from the individual;
(b) comparing the peptide(s) as identified in (a) with a collection of peptides that have been pre-screened for immunogenicity and/or over-presentation in tumours as compared to normal tissues, wherein the collection of peptides comprises one or more peptides comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, having a maximal length of 16 amino acids;
(c) selecting one or more peptides from the collection of peptides if it/they match(es) the peptide(s) identified in the subject, wherein the selected peptide(s) is/are one or more peptide(s) comprising a sequence set forth in any one of SEQ ID NOs: 1 to 59, having a maximal length of 16 amino acids comprised in the collection of peptides; and
(d) manufacturing and/or formulating the individualized vaccine or compound-based and/or cellular therapeutic agent based on the selection in step (c).
42. The method according to any one of claims 34, 35 and 41 , wherein identifying said peptide(s) comprises:
(a1 ) comparing expression data from the tumor sample to expression data from a sample of normal tissue corresponding to the tissue type of the tumor sample to identify proteins that are over-expressed or aberrantly expressed in the tumour sample; and
(a2) correlating the expression data with sequences of MHC ligands bound to MHC class I molecules in the tumor sample to identify MHC ligands derived from proteins overexpressed or aberrantly expressed by the tumor.
43. The method according to any one of claims 34, 35 and 41 , wherein the sequences of MHC ligands are identified by eluting bound peptides from MHC molecules isolated from the tumor sample and sequencing the eluted ligands; and/or wherein the normal tissue corresponding to the tissue type of the tumor sample is obtained from the same individual; and/or wherein the immunogenicity of the peptides included in the collection of peptides is determined by a method comprising an in vitro immunogenicity assay, subject immunomonitoring for individual MHC binding, MHC multimer staining, an ELISPOT assay and/or intracellular cytokine staining.
44. The method according to any one of claims 33 to 43, further comprising identifying a mutation that is unique to the tumor sample relative to normal corresponding tissue from
the individual, and selecting a peptide that correlates with the mutation for inclusion in the vaccine or for the generation of a cellular therapeutic, wherein the mutation is optionally identified by whole genome sequencing.
45. The method according to any one of claims 27 to 29 and 32 to 35, wherein the one or more peptides included in the collection of peptides is/are identified based on the following steps: aa. performing genome-wide messenger ribonucleic acid (mRNA) expression analysis by highly parallel methods, such as microarrays or sequencing-based expression profiling, comprising identifying genes that are over-expressed in a malignant tissue, compared to a normal tissue or tissues; ab. selecting peptides encoded by selectively expressed or over-expressed genes as detected in step aa, and ac. determining an induction of an in vivo T-cell response by the peptides as selected comprising in vitro immunogenicity assays using human T cells from healthy donors or the individual; or ba. identifying MHC ligands from the tumor sample using mass spectrometry; bb. performing genome-wide messenger ribonucleic acid (mRNA) expression analysis by highly parallel methods, such as microarrays or sequencing-based expression profiling, comprising identifying genes that are over-expressed in a malignant tissue, compared with a normal tissue or tissues; be. comparing the identified MHC ligands to the gene expression data; bd. selecting peptides encoded by selectively expressed or over-expressed genes as detected in step be; be. re-detecting selected peptides from step bd on tumor tissue and detecting lack of or infrequent detection on healthy tissue and confirming the relevance of over-expression at the mRNA level; and bf. determining an induction of an in vivo T-cell response by the peptides as selected comprising an in vitro immunogenicity assay using human T cells from a healthy donor or the individual.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP23020387.9 | 2023-08-18 | ||
| EP23020387 | 2023-08-18 |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| WO2025040598A2 true WO2025040598A2 (en) | 2025-02-27 |
| WO2025040598A3 WO2025040598A3 (en) | 2025-05-01 |
Family
ID=87760405
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/EP2024/073138 Pending WO2025040598A2 (en) | 2023-08-18 | 2024-08-16 | Peptides displayed by mhc for use in immunotherapy against different types of cancer |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025040598A2 (en) |
Citations (60)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4342566A (en) | 1980-02-22 | 1982-08-03 | Scripps Clinic & Research Foundation | Solid phase anti-C3 assay for detection of immune complexes |
| US4440859A (en) | 1977-05-27 | 1984-04-03 | The Regents Of The University Of California | Method for producing recombinant bacterial plasmids containing the coding sequences of higher organisms |
| US4530901A (en) | 1980-01-08 | 1985-07-23 | Biogen N.V. | Recombinant DNA molecules and their use in producing human interferon-like polypeptides |
| EP0173494A2 (en) | 1984-08-27 | 1986-03-05 | The Board Of Trustees Of The Leland Stanford Junior University | Chimeric receptors by DNA splicing and expression |
| WO1986001533A1 (en) | 1984-09-03 | 1986-03-13 | Celltech Limited | Production of chimeric antibodies |
| US4582800A (en) | 1982-07-12 | 1986-04-15 | Hoffmann-La Roche Inc. | Novel vectors and method for controlling interferon expression |
| EP0184187A2 (en) | 1984-12-04 | 1986-06-11 | Teijin Limited | Mouse-human chimaeric immunoglobulin heavy chain, and chimaeric DNA encoding it |
| US4677063A (en) | 1985-05-02 | 1987-06-30 | Cetus Corporation | Human tumor necrosis factor |
| US4678751A (en) | 1981-09-25 | 1987-07-07 | Genentech, Inc. | Hybrid human leukocyte interferons |
| US4704362A (en) | 1977-11-08 | 1987-11-03 | Genentech, Inc. | Recombinant cloning vehicle microbial polypeptide expression |
| US4710463A (en) | 1978-12-22 | 1987-12-01 | Biogen N.V. | Recombinant DNA molecules capable of expressing HBV core and surface antigens |
| US4757006A (en) | 1983-10-28 | 1988-07-12 | Genetics Institute, Inc. | Human factor VIII:C gene and recombinant methods for production |
| US4766075A (en) | 1982-07-14 | 1988-08-23 | Genentech, Inc. | Human tissue plasminogen activator |
| US4810648A (en) | 1986-01-08 | 1989-03-07 | Rhone Poulenc Agrochimie | Haloarylnitrile degrading gene, its use, and cells containing the gene |
| US4816567A (en) | 1983-04-08 | 1989-03-28 | Genentech, Inc. | Recombinant immunoglobin preparations |
| US4897445A (en) | 1986-06-27 | 1990-01-30 | The Administrators Of The Tulane Educational Fund | Method for synthesizing a peptide containing a non-peptide bond |
| WO1993008829A1 (en) | 1991-11-04 | 1993-05-13 | The Regents Of The University Of California | Compositions that mediate killing of hiv-infected cells |
| US5223409A (en) | 1988-09-02 | 1993-06-29 | Protein Engineering Corp. | Directed evolution of novel binding proteins |
| WO1994029348A2 (en) | 1993-06-03 | 1994-12-22 | Therapeutic Antibodies Inc. | Production of antibody fragments |
| WO1995018145A1 (en) | 1993-12-24 | 1995-07-06 | Ilexus Pty Ltd | Conjugates of human mucin and a carbohydrate polymer and their use in cancer treatment |
| WO1996016673A1 (en) | 1994-12-02 | 1996-06-06 | Chiron Corporation | Method of promoting an immune response with a bispecific antibody |
| WO1996023879A1 (en) | 1995-01-30 | 1996-08-08 | Terrapin Technologies, Inc. | Glubodies - multiplicities of proteins capable of binding a variety of small molecules |
| US5624659A (en) | 1993-03-19 | 1997-04-29 | Duke University | Method of treating brain tumors expressing tenascin |
| WO1997026328A1 (en) | 1996-01-17 | 1997-07-24 | Imperial College Innovations Limited | Immunotherapy using cytotoxic t lymphocytes (ctl) |
| WO1998002463A1 (en) | 1996-07-11 | 1998-01-22 | Medarex, Inc. | THERAPEUTIC MULTISPECIFIC COMPOUNDS COMPRISED OF ANTI-FCα RECEPTOR ANTIBODIES |
| US5821337A (en) | 1991-06-14 | 1998-10-13 | Genentech, Inc. | Immunoglobulin variants |
| US5837234A (en) | 1995-06-07 | 1998-11-17 | Cytotherapeutics, Inc. | Bioartificial organ containing cells encapsulated in a permselective polyether suflfone membrane |
| US5859205A (en) | 1989-12-21 | 1999-01-12 | Celltech Limited | Humanised antibodies |
| WO2001004144A2 (en) | 1999-07-13 | 2001-01-18 | Scil Proteins Gmbh | Fabrication of beta-pleated sheet proteins with specific binding properties |
| US6248516B1 (en) | 1988-11-11 | 2001-06-19 | Medical Research Council | Single domain ligands, receptors comprising said ligands methods for their production, and use of said ligands and receptors |
| WO2001072768A2 (en) | 2000-03-27 | 2001-10-04 | Technion Research And Development Foundation Ltd. | Single chain class i major histo-compatibility complexes, constructs encoding same and methods of generating same |
| US6300064B1 (en) | 1995-08-18 | 2001-10-09 | Morphosys Ag | Protein/(poly)peptide libraries |
| WO2002098443A2 (en) | 2001-06-05 | 2002-12-12 | Curevac Gmbh | Stabilised mrna with an increased g/c content and optimised codon for use in gene therapy |
| WO2003020763A2 (en) | 2001-08-31 | 2003-03-13 | Avidex Limited | Soluble t cell receptor |
| WO2003029462A1 (en) | 2001-09-27 | 2003-04-10 | Pieris Proteolab Ag | Muteins of human neutrophil gelatinase-associated lipocalin and related proteins |
| US6548640B1 (en) | 1986-03-27 | 2003-04-15 | Btg International Limited | Altered antibodies |
| WO2003068201A2 (en) | 2002-02-13 | 2003-08-21 | Technion Research & Development Foundation Ltd. | High affinity antibody having a tcr-like specificity and use in detection and treatment |
| WO2003070752A2 (en) | 2002-02-20 | 2003-08-28 | Dyax Corporation | Mhc-peptide complex binding ligands |
| WO2004033685A1 (en) | 2002-10-09 | 2004-04-22 | Avidex Ltd | Single chain recombinant t cell receptors |
| WO2004074322A1 (en) | 2003-02-22 | 2004-09-02 | Avidex Ltd | Modified soluble t cell receptor |
| WO2004084798A2 (en) | 2003-03-26 | 2004-10-07 | Technion Research & Development Foundation Ltd. | Antigen-presenting complex-binding compositions and uses thereof |
| EP1480515A2 (en) | 2002-03-05 | 2004-12-01 | ARTEMIS Pharmaceuticals GmbH | Inbred embryonic stem-cell derived mice |
| US7083784B2 (en) | 2000-12-12 | 2006-08-01 | Medimmune, Inc. | Molecules with extended half-lives, compositions and uses thereof |
| EP2112253A2 (en) | 2008-04-21 | 2009-10-28 | United Technologies Corporation | Method of restoring an article |
| US20100113300A1 (en) | 2002-11-09 | 2010-05-06 | Immunocore Limited | T cell receptor display |
| US20100234575A1 (en) | 2004-11-12 | 2010-09-16 | Xencor, Inc. | Fc variants with altered binding to fcrn |
| WO2011015347A1 (en) | 2009-08-05 | 2011-02-10 | Biontech Ag | Vaccine composition comprising 5'-cap modified rna |
| WO2011128448A1 (en) | 2010-04-16 | 2011-10-20 | Immatics Biotechnologies Gmbh | Method for differentially quantifying naturally processed hla-restricted peptides for cancer, autoimmune and infectious diseases immunotherapy development |
| WO2012056407A1 (en) | 2010-10-26 | 2012-05-03 | Technion Research & Development Foundation Ltd. | Antibodies which bind soluble t-cell receptor ligands |
| WO2013057586A1 (en) | 2011-10-19 | 2013-04-25 | Oslo Universitetssykehus Hf | Compositions and methods for producing soluble t - cell receptors |
| US20130115191A1 (en) | 2000-06-05 | 2013-05-09 | Altor Bioscience Corporation | T cell receptor fusions and conjugates and methods of use thereof |
| WO2014071978A1 (en) | 2012-11-08 | 2014-05-15 | Roche Diagnostics Gmbh | Nucleic acids encoding chimeric polypeptides for library screening |
| US20140308285A1 (en) | 2013-03-15 | 2014-10-16 | Amgen Inc. | Heterodimeric bispecific antibodies |
| WO2014191359A1 (en) | 2013-05-28 | 2014-12-04 | Rheinische Friedrich-Wilhelms Universität Bonn | Aptamers and use of the aptamers in the diagnosis and treatment of cancer |
| US8921528B2 (en) | 2004-06-01 | 2014-12-30 | Domantis Limited | Bispecific fusion antibodies with enhanced serum half-life |
| WO2016107740A1 (en) | 2014-12-30 | 2016-07-07 | Immatics Biotechnologies Gmbh | Method for the absolute quantification of naturally processed hla-restricted cancer peptides |
| WO2019104269A1 (en) | 2017-11-27 | 2019-05-31 | Immatics US, Inc. | Methods for activating, modifying and expanding gamma delta t cells for the treatment of cancer and related malignancies |
| WO2019157298A1 (en) | 2018-02-09 | 2019-08-15 | Immatics US, Inc. | Methods for manufacturing t cells |
| WO2020245326A1 (en) | 2019-06-06 | 2020-12-10 | Immatics Biotechnologies Gmbh | Sorting with counter selection using sequence similar peptides |
| WO2023044455A2 (en) | 2021-09-16 | 2023-03-23 | Team Medical Llc | Rna stabilization |
Family Cites Families (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| GB201521894D0 (en) * | 2015-12-11 | 2016-01-27 | Immatics Biotechnologies Gmbh | Novel peptides and combination of peptides for use in immunotherapy against various cancers |
| CN107266552B (en) * | 2016-03-30 | 2022-02-08 | 香雪生命科学技术(广东)有限公司 | Tumor antigen short peptide derived from PRAME |
| US10925947B2 (en) * | 2018-06-29 | 2021-02-23 | Immatics Biotechnologies Gmbh | A*03 restricted peptides for use in immunotherapy against cancers and related methods |
| EP4247402A4 (en) * | 2020-11-19 | 2025-03-26 | Board of Regents, The University of Texas System | Methods and compositions comprising mhc class i peptides |
-
2024
- 2024-08-16 WO PCT/EP2024/073138 patent/WO2025040598A2/en active Pending
Patent Citations (65)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4440859A (en) | 1977-05-27 | 1984-04-03 | The Regents Of The University Of California | Method for producing recombinant bacterial plasmids containing the coding sequences of higher organisms |
| US4704362A (en) | 1977-11-08 | 1987-11-03 | Genentech, Inc. | Recombinant cloning vehicle microbial polypeptide expression |
| US4710463A (en) | 1978-12-22 | 1987-12-01 | Biogen N.V. | Recombinant DNA molecules capable of expressing HBV core and surface antigens |
| US4530901A (en) | 1980-01-08 | 1985-07-23 | Biogen N.V. | Recombinant DNA molecules and their use in producing human interferon-like polypeptides |
| US4342566A (en) | 1980-02-22 | 1982-08-03 | Scripps Clinic & Research Foundation | Solid phase anti-C3 assay for detection of immune complexes |
| US4678751A (en) | 1981-09-25 | 1987-07-07 | Genentech, Inc. | Hybrid human leukocyte interferons |
| US4582800A (en) | 1982-07-12 | 1986-04-15 | Hoffmann-La Roche Inc. | Novel vectors and method for controlling interferon expression |
| US4766075A (en) | 1982-07-14 | 1988-08-23 | Genentech, Inc. | Human tissue plasminogen activator |
| US4816567A (en) | 1983-04-08 | 1989-03-28 | Genentech, Inc. | Recombinant immunoglobin preparations |
| US6331415B1 (en) | 1983-04-08 | 2001-12-18 | Genentech, Inc. | Methods of producing immunoglobulins, vectors and transformed host cells for use therein |
| US4757006A (en) | 1983-10-28 | 1988-07-12 | Genetics Institute, Inc. | Human factor VIII:C gene and recombinant methods for production |
| EP0173494A2 (en) | 1984-08-27 | 1986-03-05 | The Board Of Trustees Of The Leland Stanford Junior University | Chimeric receptors by DNA splicing and expression |
| WO1986001533A1 (en) | 1984-09-03 | 1986-03-13 | Celltech Limited | Production of chimeric antibodies |
| EP0184187A2 (en) | 1984-12-04 | 1986-06-11 | Teijin Limited | Mouse-human chimaeric immunoglobulin heavy chain, and chimaeric DNA encoding it |
| US4677063A (en) | 1985-05-02 | 1987-06-30 | Cetus Corporation | Human tumor necrosis factor |
| US4810648A (en) | 1986-01-08 | 1989-03-07 | Rhone Poulenc Agrochimie | Haloarylnitrile degrading gene, its use, and cells containing the gene |
| US6548640B1 (en) | 1986-03-27 | 2003-04-15 | Btg International Limited | Altered antibodies |
| US4897445A (en) | 1986-06-27 | 1990-01-30 | The Administrators Of The Tulane Educational Fund | Method for synthesizing a peptide containing a non-peptide bond |
| US5223409A (en) | 1988-09-02 | 1993-06-29 | Protein Engineering Corp. | Directed evolution of novel binding proteins |
| US6248516B1 (en) | 1988-11-11 | 2001-06-19 | Medical Research Council | Single domain ligands, receptors comprising said ligands methods for their production, and use of said ligands and receptors |
| US5859205A (en) | 1989-12-21 | 1999-01-12 | Celltech Limited | Humanised antibodies |
| US5821337A (en) | 1991-06-14 | 1998-10-13 | Genentech, Inc. | Immunoglobulin variants |
| WO1993008829A1 (en) | 1991-11-04 | 1993-05-13 | The Regents Of The University Of California | Compositions that mediate killing of hiv-infected cells |
| US5624659A (en) | 1993-03-19 | 1997-04-29 | Duke University | Method of treating brain tumors expressing tenascin |
| WO1994029348A2 (en) | 1993-06-03 | 1994-12-22 | Therapeutic Antibodies Inc. | Production of antibody fragments |
| WO1995018145A1 (en) | 1993-12-24 | 1995-07-06 | Ilexus Pty Ltd | Conjugates of human mucin and a carbohydrate polymer and their use in cancer treatment |
| WO1996016673A1 (en) | 1994-12-02 | 1996-06-06 | Chiron Corporation | Method of promoting an immune response with a bispecific antibody |
| WO1996023879A1 (en) | 1995-01-30 | 1996-08-08 | Terrapin Technologies, Inc. | Glubodies - multiplicities of proteins capable of binding a variety of small molecules |
| US5837234A (en) | 1995-06-07 | 1998-11-17 | Cytotherapeutics, Inc. | Bioartificial organ containing cells encapsulated in a permselective polyether suflfone membrane |
| US6300064B1 (en) | 1995-08-18 | 2001-10-09 | Morphosys Ag | Protein/(poly)peptide libraries |
| WO1997026328A1 (en) | 1996-01-17 | 1997-07-24 | Imperial College Innovations Limited | Immunotherapy using cytotoxic t lymphocytes (ctl) |
| WO1998002463A1 (en) | 1996-07-11 | 1998-01-22 | Medarex, Inc. | THERAPEUTIC MULTISPECIFIC COMPOUNDS COMPRISED OF ANTI-FCα RECEPTOR ANTIBODIES |
| WO2001004144A2 (en) | 1999-07-13 | 2001-01-18 | Scil Proteins Gmbh | Fabrication of beta-pleated sheet proteins with specific binding properties |
| WO2001072768A2 (en) | 2000-03-27 | 2001-10-04 | Technion Research And Development Foundation Ltd. | Single chain class i major histo-compatibility complexes, constructs encoding same and methods of generating same |
| US20130115191A1 (en) | 2000-06-05 | 2013-05-09 | Altor Bioscience Corporation | T cell receptor fusions and conjugates and methods of use thereof |
| US7670600B2 (en) | 2000-12-12 | 2010-03-02 | MedImmine, LLC | Molecules with extended half-lives, compositions and uses thereof |
| US7083784B2 (en) | 2000-12-12 | 2006-08-01 | Medimmune, Inc. | Molecules with extended half-lives, compositions and uses thereof |
| WO2002098443A2 (en) | 2001-06-05 | 2002-12-12 | Curevac Gmbh | Stabilised mrna with an increased g/c content and optimised codon for use in gene therapy |
| WO2003020763A2 (en) | 2001-08-31 | 2003-03-13 | Avidex Limited | Soluble t cell receptor |
| WO2003029462A1 (en) | 2001-09-27 | 2003-04-10 | Pieris Proteolab Ag | Muteins of human neutrophil gelatinase-associated lipocalin and related proteins |
| WO2003068201A2 (en) | 2002-02-13 | 2003-08-21 | Technion Research & Development Foundation Ltd. | High affinity antibody having a tcr-like specificity and use in detection and treatment |
| WO2003070752A2 (en) | 2002-02-20 | 2003-08-28 | Dyax Corporation | Mhc-peptide complex binding ligands |
| EP1480515A2 (en) | 2002-03-05 | 2004-12-01 | ARTEMIS Pharmaceuticals GmbH | Inbred embryonic stem-cell derived mice |
| WO2004033685A1 (en) | 2002-10-09 | 2004-04-22 | Avidex Ltd | Single chain recombinant t cell receptors |
| US20100113300A1 (en) | 2002-11-09 | 2010-05-06 | Immunocore Limited | T cell receptor display |
| WO2004074322A1 (en) | 2003-02-22 | 2004-09-02 | Avidex Ltd | Modified soluble t cell receptor |
| WO2004084798A2 (en) | 2003-03-26 | 2004-10-07 | Technion Research & Development Foundation Ltd. | Antigen-presenting complex-binding compositions and uses thereof |
| US8921528B2 (en) | 2004-06-01 | 2014-12-30 | Domantis Limited | Bispecific fusion antibodies with enhanced serum half-life |
| US20100234575A1 (en) | 2004-11-12 | 2010-09-16 | Xencor, Inc. | Fc variants with altered binding to fcrn |
| EP2112253A2 (en) | 2008-04-21 | 2009-10-28 | United Technologies Corporation | Method of restoring an article |
| WO2011015347A1 (en) | 2009-08-05 | 2011-02-10 | Biontech Ag | Vaccine composition comprising 5'-cap modified rna |
| WO2011128448A1 (en) | 2010-04-16 | 2011-10-20 | Immatics Biotechnologies Gmbh | Method for differentially quantifying naturally processed hla-restricted peptides for cancer, autoimmune and infectious diseases immunotherapy development |
| US9791443B2 (en) | 2010-04-16 | 2017-10-17 | Immatics Biotechnologies Gmbh | Method for differentially quantifying naturally processed HLA-restricted peptides for cancer, autoimmune and infectious diseases immunotherapy development |
| US9791444B2 (en) | 2010-04-16 | 2017-10-17 | Immatics Biotechnologies Gmbh | Method for differentially quantifying naturally processed HLA-restricted peptides for cancer, autoimmune and infectious diseases immunotherapy development |
| WO2012056407A1 (en) | 2010-10-26 | 2012-05-03 | Technion Research & Development Foundation Ltd. | Antibodies which bind soluble t-cell receptor ligands |
| WO2013057586A1 (en) | 2011-10-19 | 2013-04-25 | Oslo Universitetssykehus Hf | Compositions and methods for producing soluble t - cell receptors |
| WO2014071978A1 (en) | 2012-11-08 | 2014-05-15 | Roche Diagnostics Gmbh | Nucleic acids encoding chimeric polypeptides for library screening |
| US20140308285A1 (en) | 2013-03-15 | 2014-10-16 | Amgen Inc. | Heterodimeric bispecific antibodies |
| WO2014191359A1 (en) | 2013-05-28 | 2014-12-04 | Rheinische Friedrich-Wilhelms Universität Bonn | Aptamers and use of the aptamers in the diagnosis and treatment of cancer |
| WO2016107740A1 (en) | 2014-12-30 | 2016-07-07 | Immatics Biotechnologies Gmbh | Method for the absolute quantification of naturally processed hla-restricted cancer peptides |
| US10545154B2 (en) | 2014-12-30 | 2020-01-28 | Immatics Biotechnologies Gmbh | Method for the absolute quantification of naturally processed HLA-restricted cancer peptides |
| WO2019104269A1 (en) | 2017-11-27 | 2019-05-31 | Immatics US, Inc. | Methods for activating, modifying and expanding gamma delta t cells for the treatment of cancer and related malignancies |
| WO2019157298A1 (en) | 2018-02-09 | 2019-08-15 | Immatics US, Inc. | Methods for manufacturing t cells |
| WO2020245326A1 (en) | 2019-06-06 | 2020-12-10 | Immatics Biotechnologies Gmbh | Sorting with counter selection using sequence similar peptides |
| WO2023044455A2 (en) | 2021-09-16 | 2023-03-23 | Team Medical Llc | Rna stabilization |
Non-Patent Citations (107)
| Title |
|---|
| "GenBank", Database accession no. CAG30435.1 |
| "Gene", Database accession no. 23532 |
| "NCBI", Database accession no. CAG30435 |
| "UniProt", Database accession no. AOA024R1 E6 |
| ALCOSER, S. Y. ET AL., BMC.BIOTECHNOL, vol. 11, 2011, pages 124 |
| ALLISON, J. P ET AL., SCIENCE, vol. 270, 1995, pages 932 - 933 |
| APPAY, V. ET AL., EUR.J IMMUNOL., vol. 36, 2006, pages 1805 - 1814 |
| BALBÁS RLORENCE A, METHODS IN MOLECULAR BIOLOGY RECOMBINANT GENE EXPRESSION, REVIEWS AND PROTOCOLS, 2004 |
| BANCHEREAU, J ET AL., CELL, vol. 106, 2001, pages 271 - 274 |
| BANKER, GRHODES, C: "Modern Pharmaceutics", 2002, CRC PRESS |
| BEGGS, J. D. ET AL., NATURE, vol. 275, 1978, pages 104 - 109 |
| BENJAMINI, Y. ET AL., JOURNAL OF THE ROYAL STATISTICAL SOCIETY.SERIES B (METHODOLOGICAL, vol. 57, 1995, pages 289 - 300 |
| BERG ET AL., SEC. B CELL BIOLOGY, vol. 9, 2015, pages 996 - 1005 |
| BERG, J. M. ET AL.: "Biochemistry", MACMILLAN LEARNING, 2023, pages 89 - 91 |
| BERGE, S. M. ET AL., JOURNAL OF PHARMACEUTICAL SCIENCE, vol. 66, 1977, pages 1 - 19 |
| BESTE ET AL., PROC. NATL. ACAD. SCI. USA, vol. 96, 1999, pages 1898 - 1903 |
| BOULIANNE, G. L. ET AL., NATURE, vol. 312, 1984, pages 643 - 646 |
| BOULTER, J. M. ET AL., PROTEIN ENG, vol. 16, 2003, pages 707 - 711 |
| BROSSART, P. ET AL., BLOOD, vol. 90, 1997, pages 1594 - 1599 |
| BRUCKDORFER, T. ET AL., CURR.PHARM.BIOTECHNOL., vol. 5, 2004, pages 29 - 43 |
| CHANG ET AL., J CLIN INVEST, vol. 127, no. 7, 2017, pages 2705 - 2718 |
| COHEN, C. J. ET AL., J MOL.RECOGNIT, vol. 16, 2003, pages 324 - 332 |
| COHEN, S. N. ET AL., PROC.NATL.ACAD.SCI, vol. 69, 1972, pages 2110 - 2114 |
| COLIGAN, J. E. ET AL., CURRENT PROTOCOLS IN PROTEIN SCIENCE, 1995 |
| COLOMBETTI, S ET AL., J IMMUNOL., vol. 176, 2006, pages 2730 - 2738 |
| DAO ET AL., NAT. BIOTECHNOL., vol. 33, no. 10, 2015, pages 1079 - 1086 |
| DAO ET AL., SCI TRANSL MED, vol. 5, no. 176, 2013 |
| DAYHOFF, M. O. ET AL.: "The Atlas of Protein Sequence and Structure", NATL BIOMEDICAL RESEARCH, 1965 |
| DENGJEL, J. ET AL., CLIN CANCER RES, vol. 12, 2006, pages 4163 - 4170 |
| DENKBERG, G. ET AL., J IMMUNOL., vol. 171, 2003, pages 2197 - 2207 |
| DENKBERG, GREITER, Y, AUTOIMMUN. REV, vol. 5, no. 4, 2006, pages 252 - 7 |
| FALK, K. ET AL., NATURE, vol. 351, 1991, pages 290 - 296 |
| FOLLENZI, A. ET AL., NAT GENET, vol. 25, 2000, pages 217 - 222 |
| FONG, L. ET AL., PROC.NATL.ACAD.SCI.U.S.A, vol. 98, 2001, pages 8809 - 8814 |
| FORSEY, R. W. ET AL., BIOTECHNOL. LETT., vol. 31, 2009, pages 819 - 823 |
| FRITSCHE, J ET AL., MOL CELL PROTEOMICS, 2021 |
| FRITSCHE, J ET AL., PROTEOMICS, vol. 18, 2018, pages 1700284 |
| GABRILOVICH, D. I. ET AL., NAT.MED, vol. 2, 1996, pages 1096 - 1103 |
| GATTINONI, L. ET AL., NAT.REV.IMMUNOL., vol. 6, 2006, pages 383 - 393 |
| GFELLER, DBASSANI-STERNBERG, M, FRONT. IMMUNOL, vol. 9, 2018 |
| GILL, D.SDAMLE, N.K., CURRENT OPINION IN BIOTECHNOLOGY, vol. 17, 2006, pages 653 - 658 |
| GILLILAND, D. G. ET AL., PROC NATL ACAD SCI, vol. 77, 1980, pages 5483 - 5486 |
| GOUTTEFANGEAS, C.RAMMENSEE, H. G., CANCER IMMUNOL IMMUNOTHER, vol. 67, 2018, pages 1911 - 18 |
| GREEN, M. R. ET AL., MOLECULAR CLONING, A LABORATORY MANUAL, 2012 |
| GREENFIELD, E. A., ANTIBODIES: A LABORATORY MANUAL, 2014 |
| GUO, H. H. ET AL., PROC NATL ACAD SCI, vol. 101, no. 25, 2004, pages 9205 - 9210 |
| GUSTAFSSON, C. ET AL., TRENDS BIOTECHNOL, vol. 22, 2004, pages 346 - 353 |
| HARLOW, ELANE, D.: "Using Antibodies - A laboratory manual", 1989, COLD SPRING HARBOR LABORATORY |
| HE, E ET AL.: "TCR-like antibodies in cancer immunotherapy", JOURNAL OF HEMATOLOGY & ONCOLOGY, vol. 12, 2019, pages 99, XP055822672, DOI: 10.1186/s13045-019-0788-4 |
| HOLLIGER, P. ET AL., PROC NATL ACAD SCI, vol. 90, 1993, pages 6444 - 6448 |
| HOLT, L.J. ET AL., TRENDS BIOTECHNOL, vol. 21, no. 11, 2003, pages 484 - 490 |
| HOYDAHL, LS ET AL., ANTIBODIES, vol. 8, no. 2, 2019, pages 32 |
| KIBBE, A. H.: "Handbook of Pharmaceutical Excipients rd", 2000 |
| KRIEG, A. M., NAT.REV.DRUG DISCOV., vol. 5, 2006, pages 471 - 484 |
| KROLICK, K. A. ET AL., PROC NAT'L ACAD SCI, vol. 77, 1980, pages 5419 |
| KUBALL, J. ET AL., BLOOD, vol. 109, 2007, pages 2331 - 2338 |
| KWON, Y.-U.KODADEK, T., J. AM. CHEM. SOC., vol. 129, 2007, pages 1508 - 1509 |
| LA GRUTA, N. L. ET AL., NAT. REV. IMMUNOLOGY, vol. 18, no. 7, 2018, pages 467 - 478 |
| LIDDY, N. ET AL., NAT.MED., vol. 18, 2012, pages 980 - 987 |
| LIU, B. ET AL., J. AM. CHEM. SOC., vol. 126, 2004, pages 4076 - 4077 |
| LJUNGGREN, H. G. ET AL., J EXP.MED, vol. 162, 1985, pages 1745 - 1759 |
| LONGENECKER, B. M. ET AL., ANN N.Y.ACAD.SCI., vol. 690, 1993, pages 276 - 291 |
| LONSDALE, J., NAT. GENET., vol. 45, 2013, pages 580 - 585 |
| LUKAS, T. J ET AL., PROC.NATL.ACAD.SCI., vol. 78, 1981, pages 2791 - 2795 |
| LUNDBLAD, R. L., CHEMICAL REAGENTS FOR PROTEIN MODIFICATION, 2004 |
| LYTHE, G ET AL., J. THEOR. BIOL., vol. 389, 2016, pages 214 - 224 |
| LYTHE, G ET AL., JOURNAL OF THEORETICAL BIOLOGY, vol. 389, 2016, pages 214 - 224 |
| MEZIERE, C. ET AL., J IMMUNOL, vol. 159, 1997, pages 3230 - 3237 |
| MONDON R ET AL., FRONT BIOSCI, vol. 13, 2008, pages 1117 - 1129 |
| MORGAN, R. A. ET AL., SCIENCE, vol. 314, 2006, pages 126 - 129 |
| MORRISON S. L. ET AL., PROC NATL ACAD SCI, vol. 81, 1984, pages 6851 - 6855 |
| MOSAVI, L.K. ET AL., PROTEIN SCIENCE, vol. 13, no. 6, 2004, pages 1435 - 1448 |
| MUELLER, L. N. ET AL., J PROTEOME.RES., vol. 7, 2008, pages 51 - 61 |
| MUELLER, L. N. ET AL., PROTEOMICS, vol. 7, 2007, pages 3470 - 3480 |
| MURPHY, K. ET AL.: "Janeway's Immunobiology", 2022, W W NORTON & CO INC. |
| NAPOLITANO, E.W. ET AL., CHEMISTRY & BIOLOGY, vol. 3, no. 5, 1996, pages 359 - 367 |
| NOVOTNY, J ET AL., PROC. NATI. ACAD. SCI., vol. 88, no. 19, 1991, pages 8646 - 8650 |
| OLEXIOUK, V. ET AL., NUCLEIC ACIDS RES, vol. 44, 2016, pages D324 - D329 |
| PINHEIRO, J. ET AL., NLME: LINEAR AND NONLINEAR MIXED EFFECTS MODELS (CRAN.R-PROJECT.ORG/PACKE=NLME, 2015, pages 995 - 999 |
| PLEBANSKI, M. ET AL., EUR.J IMMUNOL, vol. 25, 1995, pages 1783 - 1787 |
| PORTA, C. ET AL., VIROLOGY, vol. 202, 1994, pages 949 - 955 |
| RAMMENSEE, H. G. ET AL., IMMUNOGENETICS, vol. 49, 1999, pages 571 - 576 |
| RAMMENSEE, H. G. ET AL., JOURNAL FOR IMMUNOTHERAPY OF CANCER, vol. 7, 2019, pages 307 |
| RINI, B. I. ET AL., CANCER, vol. 107, 2006, pages 67 - 74 |
| ROCK, K. L. ET AL., SCIENCE, vol. 249, 1990, pages 918 - 921 |
| RUSSELL, S. J. ET AL., NUCL ACIDS RES, vol. 21, 1993, pages 1081 - 1085 |
| SAIKI, R. K. ET AL., SCIENCE, vol. 239, 1988, pages 1041 - 1043 |
| SCHMITT, T. M. ET AL., HUM.GENE THER., vol. 20, 2009, pages 1240 - 1248 |
| SCHOLTEN, K. B. ET AL., CLIN IMMUNOL., vol. 119, 2006, pages 135 - 145 |
| SHERMAN, F. ET AL., LABORATORY COURSE MANUAL FOR METHODS IN YEAST GENETICS, 1986 |
| SILVA, L. P. ET AL., ANAL.CHEM., vol. 85, 2013, pages 9536 - 9542 |
| SILVERMAN, J. ET AL., NATURE BIOTECHNOLOGY, vol. 23, 2005, pages 1556 - 1561 |
| SINGH-JASUJA, H. ET AL., CANCER IMMUNOL.IMMUNOTHER, vol. 53, 2004, pages 187 - 195 |
| SMALL, E. J. ET AL., J CLIN ONCOL., vol. 24, 2006, pages 3089 - 3094 |
| STERNBERG, N. ET AL., PROC NATL ACAD SCI, vol. 92, 1995, pages 1609 - 1613 |
| STURM, M. ET AL., BMC.BIOINFORMATICS, vol. 9, 2008, pages 163 |
| SUBRAMANIAN, R. P. ET AL., RETROVIROLOGY, vol. 8, 2011, pages 90 |
| SUN, L. K. ET AL., PROC NATL ACAD SCI, vol. 84, 1987, pages 3439 - 3443 |
| TEUFEL, R. ET AL., CELL MOL.LIFE SCI., vol. 62, 2005, pages 1755 - 1762 |
| TRAUNECKER, A. ET AL., EMBO, vol. 10, 1991, pages 3655 - 3659 |
| UNVERDORBEN, F. ET AL., PROTEIN ENGINEERING, DESIGN & SELECTION, vol. 25, 2012, pages 81 - 88 |
| WALTER, S. ET AL., J.IMMUNOL., vol. 171, 2003, pages 4974 - 4978 |
| WALTER, S. ET AL., NAT MED, vol. 18, 2012, pages 1254 - 1261 |
| WILLCOX, B. E. ET AL., PROTEIN SCI, vol. 8, 1999, pages 2418 - 2423 |
| YANG, X ET AL., NATURE BIOTECHNOLOGY, vol. 41, 2023, pages 932 - 943 |
| ZAREMBA, S. ET AL., CANCER RES., vol. 57, 1997, pages 4570 - 4577 |
| ZUFFEREY, R. ET AL., J VIROL., vol. 73, 1999, pages 2886 - 2892 |
Also Published As
| Publication number | Publication date |
|---|---|
| WO2025040598A3 (en) | 2025-05-01 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US10106593B2 (en) | Peptides and combination of peptides for use in immunotherapy against various tumors | |
| US11407809B2 (en) | Peptides and combination of peptides for use in immunotherapy against various tumors | |
| US10954281B2 (en) | Immunotherapy with A*01 restricted peptides and combination of peptides against cancers and related methods | |
| WO2022152880A1 (en) | Peptides displayed by hla for use in immunotherapy against different types of cancers | |
| WO2025040598A2 (en) | Peptides displayed by mhc for use in immunotherapy against different types of cancer |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24782198 Country of ref document: EP Kind code of ref document: A2 |