WO2025039046A1 - Anti-rhesus d human monoclonal antibodies - Google Patents
Anti-rhesus d human monoclonal antibodies Download PDFInfo
- Publication number
- WO2025039046A1 WO2025039046A1 PCT/AU2024/050904 AU2024050904W WO2025039046A1 WO 2025039046 A1 WO2025039046 A1 WO 2025039046A1 AU 2024050904 W AU2024050904 W AU 2024050904W WO 2025039046 A1 WO2025039046 A1 WO 2025039046A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- sequence
- seq
- set forth
- cdr
- chain variable
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
- C07K16/34—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans against blood group antigens
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K16/00—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies
- C07K16/18—Immunoglobulins [IGs], e.g. monoclonal or polyclonal antibodies against material from animals or humans
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P43/00—Drugs for specific purposes, not provided for in groups A61P1/00-A61P41/00
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P7/00—Drugs for disorders of the blood or the extracellular fluid
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/505—Medicinal preparations containing antigens or antibodies comprising antibodies
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/20—Immunoglobulins specific features characterized by taxonomic origin
- C07K2317/21—Immunoglobulins specific features characterized by taxonomic origin from primates, e.g. man
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/40—Immunoglobulins specific features characterized by post-translational modification
- C07K2317/41—Glycosylation, sialylation, or fucosylation
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/50—Immunoglobulins specific features characterized by immunoglobulin fragments
- C07K2317/52—Constant or Fc region; Isotype
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/70—Immunoglobulins specific features characterized by effect upon binding to a cell or to an antigen
- C07K2317/73—Inducing cell death, e.g. apoptosis, necrosis or inhibition of cell proliferation
- C07K2317/732—Antibody-dependent cellular cytotoxicity [ADCC]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2317/00—Immunoglobulins specific features
- C07K2317/90—Immunoglobulins specific features characterized by (pharmaco)kinetic aspects or by stability of the immunoglobulin
- C07K2317/92—Affinity (KD), association rate (Ka), dissociation rate (Kd) or EC50 value
Definitions
- the present invention relates generally to recombinant antibodies which bind to RhD blood group antigens.
- the antibodies are useful for the treatment of the prevention of haemolytic disease of the newborn (HDN), treatment of idiopathic thrombocytopenic purpura (ITP), and to impede anti-RhD alloimmunisation in RhD negative patients who receive an RhD+ blood transfusion.
- Antibodies are naturally occurring molecules produced by the B-lymphocytes of the immune system to help protect against infection and disease.
- Rhesus D (RhD) is a type of blood group which is found on human red blood cells. Approximately 85% of people are RhD positive (RhD+) and 15% are RhD negative (RhD-), although these frequencies vary between populations.
- RhD- foetal red blood cells
- RBCs foetal red blood cells
- This ‘anti-D response’ increases with each pregnancy and can cause destruction of the baby’s red blood cells, known as haemolytic disease of the newborn (HDN). HDN can lead to miscarriage or stillbirth, and it has been a major cause of infant mortality throughout history, and it is still so in many parts of the world.
- anti-RhD serum i.e., serum containing anti-RhD antibodies
- RhD- women in order to rapidly remove the small population of foetal RhD+ blood cells that enter the maternal circulation, and thereby avoid sensitization of the mother’s immune system.
- HND is now rare in developed countries with an anti-RhD program in place.
- Anti-RhD plasma is derived from a small pool of altruistic blood donors who have been screened for the presence of anti-RhD antibodies. These donors are then periodically boosted with RhD+ RBCs to boost the immune system and maintain high levels of anti-RhD antibodies. Some donors have donated blood over 1,000 times for this purpose, but in many parts of the world, no such service exists. Additionally, the altruistic blood donors can become unwell after repeated immunization with ‘foreign’ RBCs. Furthermore, the donation process is time consuming and expensive, and it carries a small but appreciable risk of transferring communicable diseases. For example, in the UK, donor plasma was for many years sourced from North America to minimize any risk of transmission of variant Creutzfeldt-Jakob disease.
- the present inventors have surprisingly developed an alternative source of anti-D antibodies that is plasma and donor independent.
- the anti-D antibodies can be used for example as a prophylactic treatment and or treatment for potential sensitising events for Rh negative women who are pregnant or recently pregnant (up to for example 10 days post pregnancy cessation).
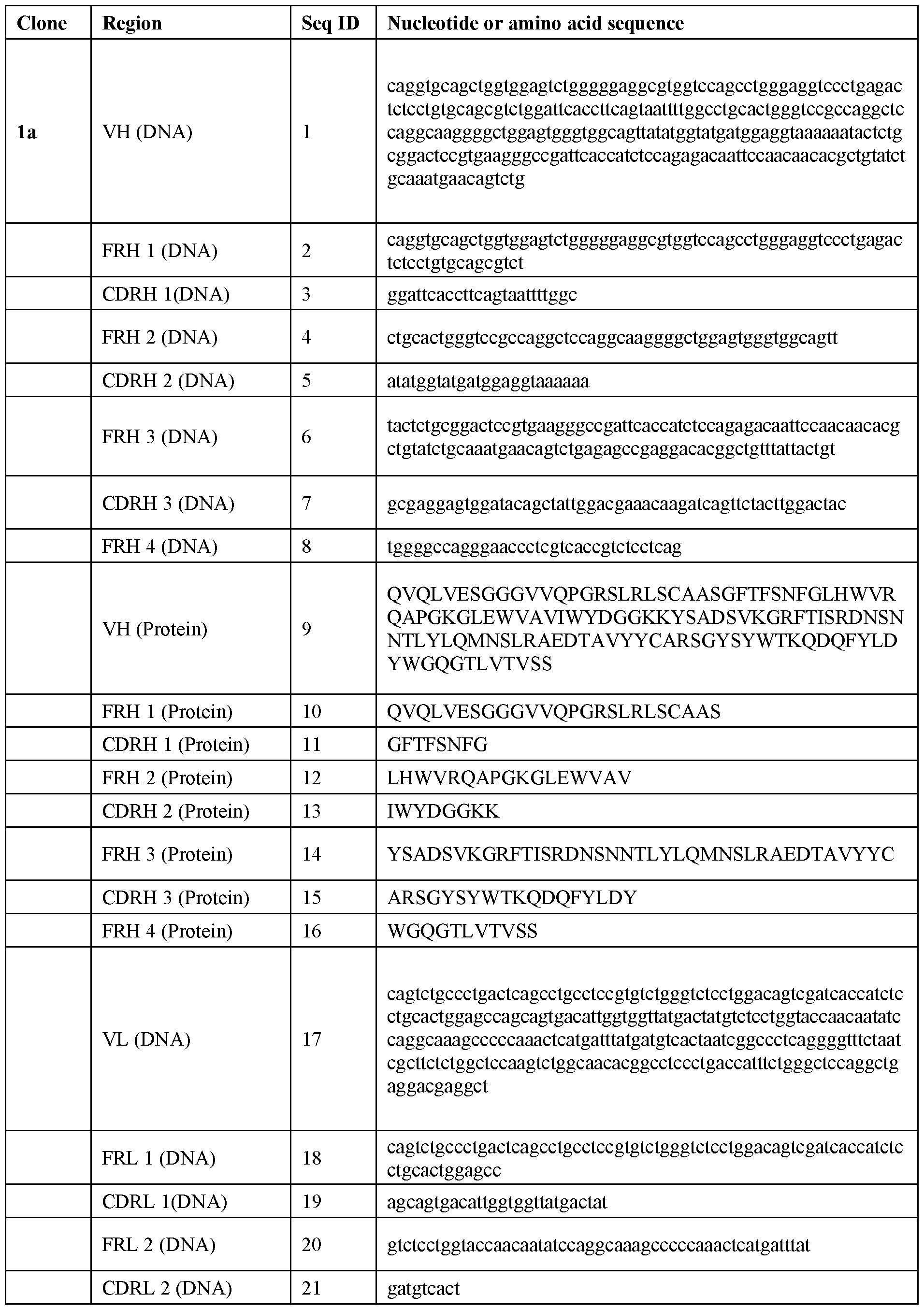
- an isolated or recombinant antibody or antigen binding fragment thereof capable of specifically binding to Rhesus D (RhD), wherein the antibody or antigen binding fragment thereof comprises a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in any one of SEQ ID NOs: 11, 43, 75, 107, 139, 171, 203, 235, 267, 299, 331, 363, 395, 427, 459, 491; a CDR H2 comprising a sequence as set forth in any one of SEQ ID NOs: 13, 45, 77, 109, 141, 173, 205, 237, 269, 301, 333, 365, 397, 429, 461, 493; and a CDR H3 comprising a sequence as set forth in any one of SEQ ID NOs: 15, 47, 79, 111, 143, 175, 207, 239, 271, 303, 3
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 11; a CDRH2 sequence as set forth in any one of SEQ ID NO: 13; and a CDR H3 sequence as set forth in of SEQ ID NO: 15.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 43; a CDR H2 sequence as set forth in SEQ ID NO: 45; and a CDR H3 sequence as set forth in SEQ ID NO: 47.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 75; a CDR H2 sequence as set forth in SEQ ID NO: 77; and a CDR H3 sequence as set forth in SEQ ID NO: 79.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 107; a CDR H2 sequence as set forth in SEQ ID NO: 109; and a CDR H3 sequence as set forth in SEQ ID NO: 111.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 139; a CDR H2 sequence as set forth in SEQ ID NO: 141; and a CDR H3 sequence as set forth SEQ ID NO: 143.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 171; a CDR H2 sequence as set forth in SEQ ID NO: 173; and a CDR H3 sequence as set forth in SEQ ID NO: 175
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 203; a CDR H2 sequence as set forth in SEQ ID NO: 205; and a CDR H3 sequence as set forth in SEQ ID NO: 207.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 235; a CDR H2 sequence as set forth in SEQ ID NO: 237; and a CDR H3 sequence as set forth in SEQ ID NO: 239.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 267; a CDR H2 sequence as set forth in SEQ ID NO: 269; and a CDR H3 sequence as set forth in SEQ ID NO: 271.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 299; a CDR H2 sequence as set forth in SEQ ID NO: 301; and a CDR H3 as set forth in SEQ ID NO: 303.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 331; a CDR H2 sequence as set forth in SEQ ID NO: 333; and a CDR H3 sequence as set forth in SEQ ID Nos: 335.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 363; a CDR H2 sequence as set forth in SEQ ID NO: 365; and a CDR H3 sequence as set forth in SEQ ID NO: 367.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 395; a CDR H2 sequence as set forth in SEQ ID NO: 397; and a CDR H3 sequence as set forth in SEQ ID NO: 399.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 427; a CDR H2 sequence as set forth in SEQ ID NO: 429; and a CDR H3 sequence as set forth in SEQ ID NO: 431.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 459; a CDR H2 sequence as set forth in SEQ ID NO: 461; and a CDR H3 sequence as set forth in SEQ ID NO: 463.
- the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 491; a CDR H2 sequence as set forth in SEQ ID NO: 493; and a CDR H3 sequence as set forth in SEQ ID NO: 495.
- the antibody or antigen binding fragment of the disclosure comprises a light chain variable region (VL) comprising a CDR LI sequence as set forth in any one of SEQ ID NOs: 27, 59, 91, 123, 155, 187, 219, 251, 283, 315, 347, 379, 411, 443, 475, 507; a CDR L2 sequence as set forth in any one of SEQ ID NOs: 29, 61, 93, 125, 157, 189, 221, 253, 285, 317, 349, 381, 413, 445, 477, 509; and a CDR L3 sequence as set forth in any one of SEQ ID NOs: 31, 63, 95, 127, 159, 191, 223, 255, 287, 319, 351, 383, 415, 447, 479, 511.
- VL light chain variable region
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 27; a CDR L2 sequence as set forth in SEQ ID NO: 29; and a CDR L3 sequence as set forth in SEQ ID NO: 31.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 59; a CDR L2 sequence as set forth in SEQ ID NO: 61; and a CDR L3 sequence as set forth in SEQ ID NO: 63.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 91; a CDR L2 sequence as set forth in SEQ ID NO: 93; and a CDR L3 sequence as set forth in SEQ ID NO: 95.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 123; a CDR L2 sequence as set forth in SEQ ID NO: 125; and a CDR L3 sequence as set forth in SEQ ID NO: 127.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 155; a CDR L2 sequence as set forth in SEQ ID NO: 157; and a CDR L3 sequence as set forth in SEQ ID NO: 159.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 187; a CDR L2 sequence as set forth in SEQ ID NO: 189; and a CDR L3 as set forth in SEQ ID NO: 191.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI as set forth in SEQ ID NO: 219; a CDR L2 sequence as set forth in SEQ ID NO: 221; and a CDR L3 sequence as set forth in SEQ ID NO: 223.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 251; a CDR L2 sequence as set forth in SEQ ID NO: 253; and a CDR L3 sequence as set forth in SEQ ID NO: 255.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 283; a CDR L2 sequence as set forth in SEQ ID NO: 285; and a CDR L3 sequence as set forth in SEQ ID NO: 287.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 315; a CDR L2 sequence as set forth in SEQ ID NO: 317; and a CDR L3 sequence as set forth in SEQ ID NO: 319.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 347; a CDR L2 sequence as set forth in SEQ ID NO: 349; and a CDR L3 sequence as set forth in SEQ ID NO: 351.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 379; a CDR L2 sequence as set forth in SEQ ID NO: 381; and a CDR L3 sequence as set forth in SEQ ID NO: 383.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 411, a CDR L2 sequence as set forth in SEQ ID NO: 413; and a CDR L3 sequence as set forth in SEQ ID NO: 415.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 443; a CDR L2 sequence as set forth in SEQ ID NO: 445; and a CDR L3 sequence as set forth in SEQ ID NO: 447.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 475; a CDR L2 sequence as set forth in SEQ ID NO: 477; and a CDR L3 sequence as set forth in SEQ ID NO: 479.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 507; a CDR L2 sequence as set forth in SEQ ID NO: 509; and a CDR L3 sequence as set forth in SEQ ID NO: 511.
- the antibody or antigen binding fragment of the disclosure comprises a VH comprising an amino acid sequence having at least 70% sequence identity to the sequence set forth in any one of SEQ ID NOs: 9, 41, 73, 105, 137, 169, 201, 233, 265, 297, 329, 361, 393, 425, 457, 489.
- the antibody or antigen binding fragment of the disclosure comprises a VL comprising an amino acid sequence having at least 70% sequence identity to the sequence set forth in any one of SEQ ID NOs: 25, 57, 89, 121, 153, 185, 217, 249, 281, 313, 345, 377, 409, 441, 473, 505.
- the disclosure provides an antibody or antigen binding fragment thereof capable of specifically binding to Rhesus D (RhD), wherein the antibody or antigen binding fragment thereof comprises: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 11; CDR H2 comprising a sequence as set forth in SEQ ID NO: 13; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 15; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 27; CDR L2 comprising a sequence as set forth in SEQ ID NO: 29; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 31; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 43; CDR H2 comprising a sequence as set forth in S
- the disclosure provides an antibody or antigen binding fragment thereof capable of specifically binding to Rhesus D (RhD), wherein the antibody or antigen binding fragment thereof comprises: a) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO:
- the antibody or antigen binding fragment is capable of specifically binding to RhD positive (RhD+), but not RhD negative (RhD-), RBCs.
- the antibody or antigen binding fragment is capable of agglutinating RhD+ red blood cells but not RhD- red blood cells.
- the antibody or antigen binding fragment is one of IgG subclass 1, IgG subclass 2, or IgG subclass 3.
- the antibody or antigen binding fragment is IgG subclass 1, IgG or IgG subclass 3.
- the antibody or antigen binding fragment is IgG subclass 1.
- the antibody or antigen binding fragment comprise a Fc region or part thereof.
- one or more of the antibodies or antigen binding fragments disclosed in the present application can be used in combination to provide a composition. Accordingly, the present disclosure provides a composition comprising one or more antibodies or antigen binding fragments as defined herein. In some embodiments, one or more of the antibodies or antigen binding fragments disclosed herein, can be used in combination with one more additional antibodies or antigen binding fragments capable of specifically binding to Rhesus D (RhD) that may optionally be identified using a method disclosed herein.
- RhD Rhesus D
- the present disclosure provides a composition comprising one or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein one or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 11; CDR H2 comprising a sequence as set forth in SEQ ID NO: 13; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 15; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 27; CDR L2 comprising a sequence as set forth in SEQ ID NO: 29; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 31; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set
- the disclosure provides a composition comprising one or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein one or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO:9; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 25; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 41; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 57; c) a heavy chain variable region (VH) comprising complementary determining regions (CDRs)
- the disclosure provides a composition comprising one or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein the one or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 75; CDR H2 comprising a sequence as set forth in SEQ ID NO: 77; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 79; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 91; CDR L2 comprising a sequence as set forth in SEQ ID NO: 93; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 95; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl
- the composition further comprises two or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD).
- the two or more antibodies or antigen binding fragments may be selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 75; CDR H2 comprising a sequence as set forth in SEQ ID NO: 77; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 79; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 91; CDR L2 comprising a sequence as set forth in SEQ ID NO: 93; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 95; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) H
- VH heavy chain variable
- the composition further comprises three or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD).
- the one or more antibodies or antigen binding fragments may be selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 75; CDR H2 comprising a sequence as set forth in SEQ ID NO: 77; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 79; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 91; CDR L2 comprising a sequence as set forth in SEQ ID NO: 93; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 95; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) H
- VH heavy chain variable
- the disclosure provides a composition comprising one or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein the one or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 89; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 185; c) a heavy chain variable region (VH) comprising complementary determining regions (VH)
- the disclosure provides a composition comprising a combination of two or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein the two or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 89; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 185; c) a heavy chain variable region (VH) comprising complementary
- the disclosure provides a composition comprising a combination of three or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein the three or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 89; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 185; c) a heavy chain variable region (VH) comprising complementary
- the composition comprises a combination of three or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD).
- the three or more antibodies or antigen binding fragments may be selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 89; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 137; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 153; c) a heavy chain variable region (VH) comprising complementary
- composition of any of the previous aspects further comprises a pharmaceutically acceptable carrier.
- the disclosure provides a nucleic acid molecule(s) encoding the antibody or antibody or antigen binding fragment according to the previous aspects.
- the disclosure provides a vector(s) comprising the nucleic acid molecule(s) of the disclosure.
- the vector(s) is an expression vector.
- the disclosure provides a host cell comprising the vector(s) of the disclosure.
- the host cell is a prokaryote or eukaryote.
- the host cell is selected from an Escherichia coli cell, a yeast cell, a mammalian cell, and other cells applicable to the preparation of an antibody or antigen binding fragment thereof.
- the mammalian cell is a CHO cell, HEK293 cell or COS cell.
- disclosure provides a production method of an antibody or antigen binding fragment thereof, wherein the host cell of the disclosure is cultured under conditions suitable for expression of a nucleic acid encoding the antibody or antigen binding fragment thereof of the disclosure, optionally wherein the antibody or antigen binding fragment thereof is separated, and optionally wherein the produced antibody or antigen-binding fragments thereof are collected.
- the disclosure provides a method of treating or preventing Rhesus alloimmunization of Rh-negative subjects, wherein the method comprises administering to a Rh-negative subject in need thereof an effective amount of an anti-Rd(D) antibody or antigen binding fragment of the disclosure, or a combination thereof; or the composition of the disclosure.
- the subject is pregnant.
- the method prevents haemolytic disease of the foetus and newborn.
- the disclosure provides for use of the antibody or antigen binding fragment of the disclosure, or a combination thereof, in the manufacture of a medicament for the treatment or prevention of haemolytic disease of the foetus and newborn.
- the disclosure provides for use of the antibody or antigen binding fragment of the disclosure, or a composition thereof, or a composition of the disclosure, for the treatment or prevention of Rhesus alloimmunization of Rh-negative subjects.
- the disclosure provides a method for profiling the most abundant antibodies that bind to the antigen (RhD) in a polyclonal (pAb) population, wherein the method comprises: (a) proteomic analysis of a RhD-specific IgG mixture from plasma, wherein plasma is provided from a donor; affinity purification of the total IgG mixture from the plasma; separation of the RhD-specific and non-RhD specific IgG subsets by incubation with either RhD+ or RhD- RBCs, respectively; independent fragmenting of the RhD-specific and non-RhD specific IgG subsets into short peptides of overlapping sequences using one or more proteases; analysis of the fragmented peptides by mass spectrometry to identify the sequence of each peptide; assembly of the short peptide sequences into longer peptide sequences that belong to individual antibodies, optionally using computational software; and excluding antibodies derived from the non-RhD specific subset from further analysis; and (b) BCR sequencing,
- a method for profiling the most abundant antibodies that bind to the antigen (RhD) in a polyclonal (pAb) population comprising: a) proteomic analysis of a RhD-specific IgG mixture from plasma to provide proteomic sequence information: b) BCR sequencing: c) combining the transcriptomics sequence information with the proteomic sequence information to further assemble and derive the complete amino acid sequences of the most abundant mAbs in the RhD specific IgG subset: d) further selecting from the most abundant mAbs a combination of mAbs each of which comprises a different genetic and/or amino acid sequence and each mAb containing a IgGl subclass heavy chain variable region (VH).
- VH IgGl subclass heavy chain variable region
- the blood is collected from RhD antigen donors.
- the donor plasma is separated from PBMCs.
- CD 19+ IgG+ B cells are separated from donor PBMCs.
- one or more of the amino acid sequences identified can be used to make recombinant antibodies or antigen binding fragments that can be used in combination with one or more of the antibodies or antigen binding fragments disclosed herein.
- FIGURE l is a schematic showing the discovery of anti-RhD antibodies using a combination of Next Generation Sequencing (NGS) and proteomics.
- NGS Next Generation Sequencing
- FIGURE 2 is a schematic showing the cloning of antibody variable genes and cloning into the expression vectors. The specificity of purified antibodies is evaluated in agglutination assays against both RhD+ and RhD- RBCs.
- FIGURE 3 is a schematic of the RhD antigen in the erythrocyte membrane.
- FIGURE 4 shows agglutination results (with bromelain treated or untreated RBCs).
- FIGURE 5 shows anti-RhD mAbs competition binding profiles.
- FIGURE 6 shows binding of anti-RhD mAbs to different Epitope RhD variants (A). Potential epitopes of anti-RhD mAbs according to the availability of epitopes in tested RhD variants (B).
- FIGURE 7 shows the inhibitory activity of RhD-pIgG-1 on binding of anti-RhD mAbs from FACS analysis.
- FIGURE 8 shows anti-RhD mAbs induced RBC clearance by THP-1 monocyte.
- A) The gating strategy used for analysis of in vitro phagocytosis assay in the presence of non-RhD mAb (Top panel) or anti-RhD mAb (Bottom panel).
- FIGURE 9 shows anti-RhD mAbs induced RBC clearance by ADCC.
- FIGURE 10 shows activation of NK cells by anti-RhD mAbs.
- FIGURE 11 shows the germline usage in anti-RhD heavy chains.
- FIGURE 12 shows the germline usage in anti-RhD light chains.
- FIGURE 13 is an image of the Alphafold modelling the interaction between anti-RhD mAbs with RhD antigen.
- the RhD antigen is depicted as a coiled structure (bottom molecule in each figure).
- the antibody heavy and light chains are represented as ribbon structures (top molecules in each figure).
- FIGURE 14 shows the prediction of the epitope interaction with the RhD antigen.
- FIGURE 15 shows a prediction of target sites of anti-RhD mAbs on extracellular RhD loops. The predicted binding sites by Alphafold is shown in rectangle for each mAb.
- Four mAbs interact with ASP350 (shown by black oval).
- FIGURE 16 shows RBC binding of modified anti-RhD mAbs.
- FIGURE 17 shows NK cell activation and RBC clearance through ADCC with FC modified anti-RhD mAbs.
- FIGURE 18 shows ADP activity results of Fc modified anti-RhD mAbs.
- ADP activity ADP score of antibody variants at various concentrations.
- B Area under curve (AUC).
- FIGURE 19 shows clearance of RhD+ RBCs mediated by anti-RhD antibody variants.
- ADCC activity of antibodies (Img/mL) against Bromelain-treated (A) or untreated (B) RhD+ RBCs is shown in the top panel.
- ADP activity of antibodies (lOOng/mL) against Bromelain-treated (C) or untreated (D) RhD+ RBCs is also shown in the bottom panel.
- FIGURE 20 shows clearance of RhD+ RBCs mediated by anti-RhD antibody pools.
- ADCC activity of antibodies (Img/mL) against Bromelain-treated (A) or untreated (B) RhD+ RBCs is shown in the top panel.
- ADP activity of antibody pools (lOOng/mL) against Bromelain- treated (C) or untreated (D) RhD+ RBCs is shown in the bottom panel.
- FIGURE 21 shows RBC binding of anti-RhD antibody fragments.
- composition of matter, group of steps or group of compositions of matter shall be taken to encompass one and a plurality (i.e. one or more) of those steps, compositions of matter, groups of steps or groups of compositions of matter.
- variable regions and parts thereof, antibodies and fragments thereof herein may be further clarified by the discussion in Kabat Sequences of Proteins of Immunological Interest, National Institutes of Health, Bethesda, Md. (1987 and 1991), Bork et al., J Mol. Biol. (1994) 242:309-320, 1994; Chothia and Lesk, J. Mol. Biol. (1987) 196:901-917; Chothia et al. Nature (1989) 342:877-883, and Al-Lazikani et al., J. Mol. Biol. (1997) 273:927-948.
- derived from shall be taken to indicate that a specified integer may be obtained from a particular source albeit not necessarily directly from that source.
- binding does not necessarily require exclusive binding or non-detectable binding of another antigen.
- the term “specifically binds” can be used interchangeably with “selectively binds” herein.
- reference herein to binding means specific binding, and each term shall be understood to provide explicit support for the other term. Methods for determining specific binding will be apparent to the skilled person. For example, a binding protein comprising the binding region of the disclosure is contacted with the component or a cell expressing same or a mutant form thereof or an alternative antigen. The binding to the component or mutant form or alternative antigen is then determined and a binding region that binds as set out above is considered to specifically bind to the component.
- recombinant shall be understood to mean the product of artificial genetic recombination. Accordingly, in the context of an antibody or antigen binding fragment thereof, this term does not encompass an antibody naturally occurring within a subject's body that is the product of natural recombination that occurs during B cell maturation. However, if such an antibody is isolated, it is to be considered an isolated protein comprising an antibody variable region. Similarly, if nucleic acid encoding the protein is isolated and expressed using recombinant means, the resulting protein is a recombinant protein. A recombinant protein also encompasses a protein expressed by artificial recombinant means when it is within a cell, tissue or subject, for example, in which it is expressed.
- polypeptide or "polypeptide chain” will be understood to mean a series of contiguous amino acids linked by peptide bonds.
- an "antibody” is generally considered to be a protein that comprises a variable region made up of a plurality of polypeptide chains, for example, a polypeptide comprising a light chain variable region (VL) and a polypeptide comprising a heavy chain variable region (VH).
- An antibody also generally comprises constant domains, some of which can be arranged into a constant region, which includes a constant fragment or fragment crystallizable (Fc), in the case of a heavy chain.
- Fc constant fragment or fragment crystallizable
- a light chain from mammals is either a K light chain or a light chain and a heavy chain from mammals is a a, 5, a, y, or p.
- Antibodies can be of any class (e.g., IgG, IgE, IgM, IgD, IgA, and IgY), or subclass (e.g., IgGl, IgG2, IgG3, IgG4, IgAl and IgA2) or subclass.
- the term "antibody" in the context of this invention encompasses human antibodies.
- antibody also includes variants, for example, variants missing an encoded C-terminal lysine residue, a deamidated variant and/or a glycosylated variant and/or a variant comprising a pyroglutamate, for example, at the N- terminus and/or a variant lacking a N-terminal residue, for example, a N-terminal glutamine in an antibody or V region and/or a variant comprising all or part of a secretion signal.
- Deamidated variants of encoded asparagine residues may result in isoaspartic, and aspartic acid isoforms being generated or even a succinamide involving an adjacent amino acid residue.
- Deamidated variants of encoded glutamine residues may result in glutamic acid.
- Compositions comprising a heterogeneous mixture of such sequences and variants are intended to be included when reference is made to a particular amino acid sequence.
- full-length antibody “intact antibody” or “whole antibody” are used interchangeably to refer to an antibody in its substantially intact form, as opposed to an antigen binding fragment of an antibody.
- whole antibodies include those with heavy and light chains including an Fc region.
- the constant domains may be wild-type sequence constant domains (e.g., human wild-type sequence constant domains) or amino acid sequence variants thereof).
- mAb monoclonal antibody
- mAb monoclonal antibody
- the term "monoclonal antibody” (mAb) as used herein refers to an antibody obtained from a population of substantially homogeneous antibodies, or to said population of antibodies.
- the individual antibodies comprising the population are essentially identical, except for possible naturally occurring mutations that may be present in minor amounts.
- Monoclonal antibodies are highly specific, being directed against a single antigenic site.
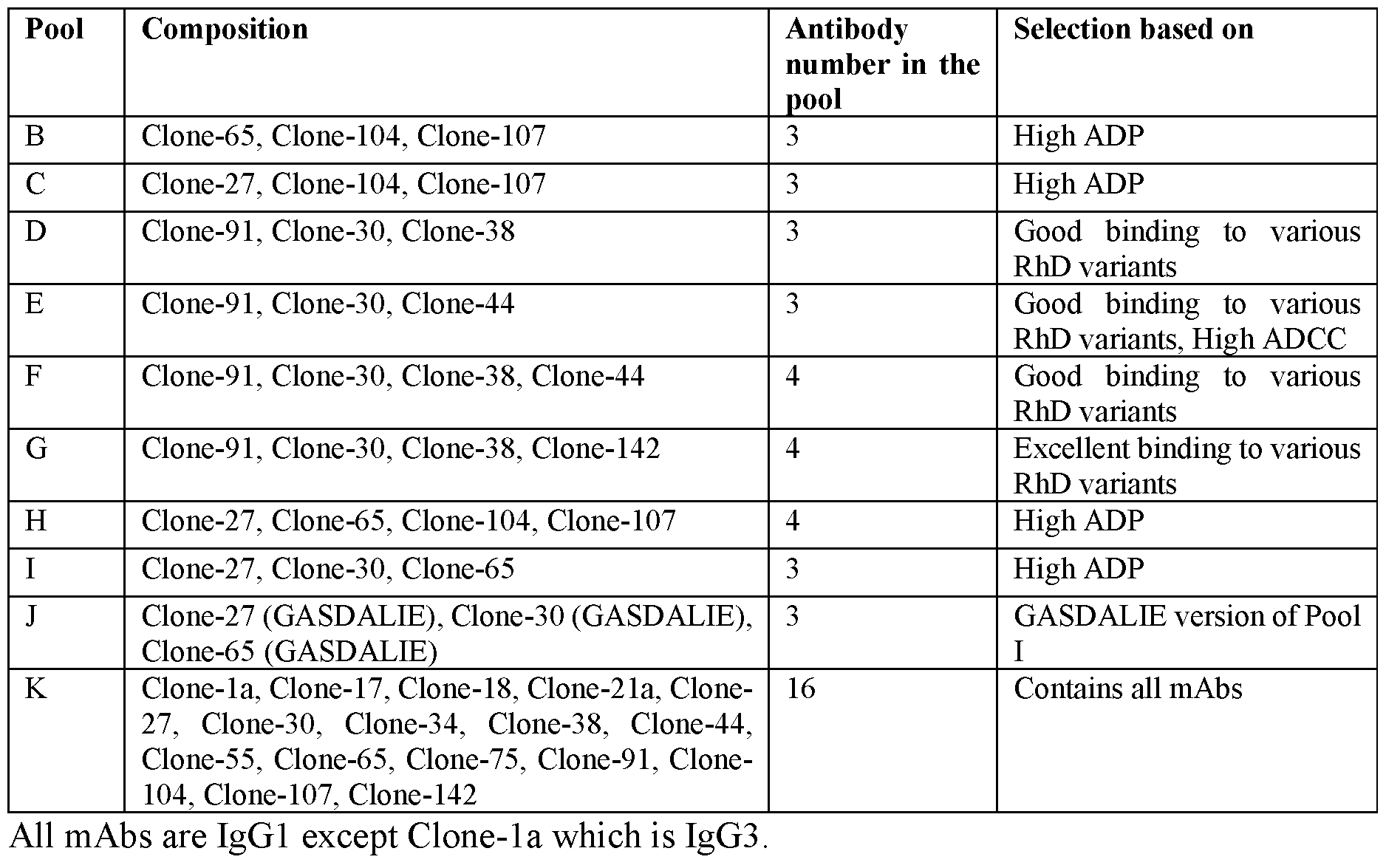
- the present disclosure provides for recombinant DNA expression of monoclonal antibodies. Two or more recombinant monoclonal antibodies (e.g., directed against different antigenic sites) can be pooled and used in methods of the disclosure. Specific combinations or pools can be chosen to provide a diverse range of recombinant antibodies each of which may bind to one or more different Rhesus D antigen epitopes,
- Human antibodies as used herein includes antibodies having the amino acid sequence of a human immunoglobulin and antibodies isolated from human immunoglobulin libraries.
- variable antibodies refers to structural chimeras with, for example, variable sequences derived from one species, and constant regions from another species or another isotype within the same species.
- variable region refers to the portions of the light and/or heavy chains of an antibody as defined herein that is capable of specifically binding to an antigen and, for example, includes amino acid sequences of complementarity determining regions (CDRs); i.e., CDR1, CDR2, and CDR3, and framework regions (FRs).
- CDRs complementarity determining regions
- FRs framework regions
- the variable region comprises three or four FRs (e.g., FRI, FR2, FR3 and optionally FR4) together with three CDRs.
- VH refers to the variable region of the heavy chain.
- VL refers to the variable region of the light chain.
- CDRs complementarity determining regions
- CDR1, CDR2, and CDR3 refers to the amino acid residues of an antibody variable region the presence of which contribute to specific antigen binding.
- Each variable region typically has three CDR regions identified as CDR1, CDR2 and CDR3.
- amino acid positions assigned to CDRs and FRs are defined according to Kabat Sequences of Proteins of Immunological Interest, National Institutes of Health, Bethesda, Md., 1987 and 1991.
- VH FRs and CDRs are positioned as follows: residues 1-30 (FRI), 31-35 (CDR1), 36-49 (FR2), 50-65 (CDR2), 66-94 (FR3), 95- 102 (CDR3) and 103- 113 (FR4).
- VL FRs and CDRs are positioned as follows: residues 1-23 (FRI), 24-34 (CDR1), 35-49 (FR2), 50-56 (CDR2), 57-88 (FR3), 89-97 (CDR3) and 98-107 (FR4).
- the present disclosure is not limited to FRs and CDRs as defined by the Kabat numbering system, but includes all numbering systems, including those discussed above.
- reference herein to a CDR (or a FR) is in respect of those regions according to the Kabat numbering system.
- FRs Framework regions
- the term “Fv” shall be taken to mean any protein, whether comprised of multiple polypeptides or a single polypeptide, in which a VH and a VL associate and form a complex having an antigen binding site, i.e., capable of specifically binding to an antigen.
- the VH and the VL which form the antigen binding site can be in a single polypeptide chain or in different polypeptide chains.
- an Fv of the disclosure (as well as any protein of the disclosure) may have multiple antigen binding sites which may or may not bind the same antigen. This term shall be understood to encompass fragments directly derived from an antibody as well as proteins corresponding to such a fragment produced using recombinant means.
- the VH is not linked to a heavy chain constant domain (CH), for example CH 1 and/or the VL is not linked to a light chain constant domain (CL).
- exemplary Fv containing polypeptides or proteins include a Fab fragment, a Fab’ fragment, a F(ab’) fragment, a scFv, a diabody, a triabody, a tetrabody or higher order complex, or any of the foregoing linked to a constant region or domain thereof, for example, one or both of a CH2or CH3 domain, for example, a minibody.
- a "Fab fragment” consists of a monovalent antigen-binding fragment of an immunoglobulin and can be produced by digestion of a whole antibody with the enzyme papain, to yield a fragment consisting of an intact light chain and a portion of a heavy chain or can be produced using recombinant means.
- a "Fab 1 fragment" of an antibody can be obtained by treating a whole antibody with pepsin, followed by reduction, to yield a molecule consisting of an intact light chain and a portion of a heavy chain comprising a VH and a single CH. Two Fab' fragments are obtained per antibody treated in this manner. A Fab’ fragment can also be produced by recombinant means.
- a “F(ab')2 fragment” consists of a dimer of two Fab' fragments held together by two disulfide bonds and is obtained by treating a whole antibody molecule with the enzyme pepsin, without subsequent reduction.
- a “Fab2” fragment is a recombinant fragment comprising two Fab fragments linked using, for example a leucine zipper or a CH3 domain.
- a “single chain Fv” or “scFv” is a recombinant molecule containing the variable region fragment (Fv) of an antibody in which the variable region of the light chain and the variable region of the heavy chain are covalently linked by a suitable, flexible polypeptide linker.
- constant region refers to a portion of heavy chain or light chain of an antibody other than the variable region.
- the constant region generally comprises a plurality of constant domains and a hinge region, for example, an IgG constant region comprises the following linked components, a CHI, a hinge, a CH2 and a CH3.
- a constant region comprises a Fc.
- a constant region In a light chain, a constant region generally comprises one constant domain (a CL1).
- fragment crystallizable or “Fc” or “Fc region” or “Fc portion” refers to a region of an antibody comprising at least one constant domain, and which is generally (though not necessarily) glycosylated and which is capable of binding to one or more Fc receptors and/or components of the complement cascade.
- the heavy chain constant region can be selected from any of the five isotypes: a, 5, a, y, or p.
- heavy chains of various subclasses are responsible for different effector functions and thus, by choosing the desired heavy chain constant region, proteins with desired effector function can be produced.
- Exemplary heavy chain constant regions are gamma 1 (IgGl), gamma 2 (IgG2) and gamma 3 (IgG3), or hybrids thereof.
- the terms “antigen-binding portion of an antibody”, “antigen-binding fragment”, “antigen-binding domain”, “antibody fragment”, or a “functional fragment of an antibody” are used interchangeably in the present disclosure to mean one or more fragments of an antibody that retain the ability to specifically bind to an antigen, (see generally, Holliger et al., Nature Biotech. (2005) 23 (9):1126-1129.
- Non-limiting examples of antibody fragments include (i) a Fab fragment, a monovalent fragment consisting of the VL , VH , CL and CHI domains; (ii) a F(ab')2 fragment, a bivalent fragment comprising two Fab fragments linked by a disulfide bridge at the hinge region; (iii) a Fd fragment consisting of the VH and CHI domains; (iv) a Fv fragment consisting of the VL and VH domains of a single arm of an antibody, (v) a dAb fragment (Ward et al., Nature (1989) 341 :544 546), which consists of a VH domain; and (vi) an isolated complementarity determining region (CDR).
- a Fab fragment a monovalent fragment consisting of the VL , VH , CL and CHI domains
- a F(ab')2 fragment a bivalent fragment comprising two Fab fragments linked by a disulfide bridge
- the two domains of the Fv fragment, VL and VH are coded for by separate genes, they can be joined, using recombinant methods, by a synthetic linker that enables them to be made as a single protein chain in which the VL and VH regions pair to form monovalent molecules (known as single chain Fv (scFv) (see, e.g., Bird et al., Science (1988) 242:423 426; and Huston et al., Proc. Natl. Acad. Sci. USA (1988) 85:5879 5883; and Osbourn et al., Nat. Biotechnol. (1998) 16:778).
- scFv single chain Fv
- single chain antibodies are also intended to be encompassed within the term “antigen-binding portion” of an antibody.
- Any VH and VL sequences of specific scFv can be linked to human immunoglobulin constant region cDNA or genomic sequences, to generate expression vectors encoding complete IgG molecules or other isotypes.
- VH and VL can also be used in the generation of Fab, Fv or other fragments of immunoglobulins using either protein chemistry or recombinant DNA technology.
- Other forms of single chain antibodies, such as diabodies are also encompassed.
- subject refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- patient refers to, humans.
- individual refers to, humans.
- individual refers to, humans.
- patient as used herein includes a living human that is receiving medical care or that should receive medical care due to a disease or condition. This includes subjects with no defined illness or observable symptoms of a disease or condition who are being investigated for signs of pathology.
- the term “subject” refers to Rh-negative subjects (e.g., a Rh- negative pregnant subject) at risk of or with symptoms of Rhesus alloimmunization.
- the “subject” refers to a fetus or newborn at risk of haemolytic disease owing to Rhesus alloimmunization of the “mother”.
- a subject "at risk” of developing a disease or condition or relapse thereof or relapsing may or may not have detectable symptoms and may or may not have displayed detectable symptoms prior to treatment according to the present disclosure.
- “At risk” denotes that a subject has one or more risk factors, which are measurable parameters that correlate with development of the disease or condition, as known in the art and/or described herein.
- the terms “treating”, “treat” or “treatment” include administering an antibody or antigen binding fragment of the disclosure to thereby reduce or eliminate at least one symptom of the disease or condition or to slow progression of the disease or condition.
- the anti-D antibodies can be used for example as a prophylactic treatment and or treatment for potential sensitising events for Rh negative women who are pregnant or recently pregnant (up to for example 10 days post pregnancy cessation).
- the term "preventing”, “prevent” or “prevention” includes providing prophylaxis with respect to occurrence or recurrence of the disease or condition.
- An individual may be predisposed to or at risk of developing the disease or disease relapse but has not yet been diagnosed with the disease or the relapse.
- an “effective amount” refers to at least an amount effective, at dosages and for periods of time necessary, to achieve the desired result.
- the desired result may be a therapeutic or prophylactic result.
- An effective amount can be provided in one or more administrations.
- the term “effective amount” means an amount necessary to effect treatment of the disease or condition.
- the term “effective amount” is meant as an amount necessary to deplete or eliminate RhD+ red blood cells but not RhD negative (RhD-) red blood cells. This can be referred to as RBC clearance in the art.
- RBC clearance may be mediated by antibody dependent cell mediated cytotoxicity (ADCC) and/or antibody-dependent cellular phagocytosis (ADCP).
- ADCC antibody dependent cell mediated cytotoxicity
- ADCP antibody-dependent cellular phagocytosis
- the effective amount may vary according to the disease or condition to be treated or factor to be altered and also according to the weight, age, racial background, sex, health and/or physical condition and other factors relevant to the subject being treated. Typically, the effective amount will fall within a relatively broad range (e.g.
- the effective amount can be administered in a single dose or in a dose repeated once or several times over a treatment period.
- the present invention relates antibodies to the RhD antigen of human red blood cells.
- the Rhesus blood group system is a major antigenic constituent of the human red blood cell membrane; of this group, the RhD antigen is of particular clinical importance in relation to isoimmune reactions.
- An Rh D- individual with anti-RhD who receives RhD+ blood is liable to suffer substantial red blood cell (RBc) destruction due to the Rh(D) phenotype incompatibility, and thus blood of donors must routinely be classified as RhD+ or RhD-.
- RBc red blood cell
- RhD antigen is also responsible for haemolytic disease of the newborn (HDN). This condition arises in newborn RhD+ infants of RhD- “mothers” previously sensitised to RhD antigen as a result of IgG anti-RhD antibodies crossing the placenta during pregnancy and causing foetal RBC destruction. Sensitization of the RhD- “mother” to RhD antigen often occurs during the birth of an earlier RhD+ “child” due to some foetal RBCs entering the maternal circulation and being recognised as foreign by the maternal immune system.
- the anti-RhD antibodies and antigen binding fragments of the disclosure are monoclonal antibodies and may be used singly or in combination.
- the antibodies may be genetically engineered to enhance specific functional activities.
- the antibodies and antigen binding fragments comprise all or a portion of a constant region of an antibody.
- the constant region is an isotype selected from: IgA (e.g., IgAl or IgA2), IgD, IgE, IgG (e.g., IgGl, IgG2, IgG3 or IgG4), and IgM.
- the antibody or antigen binding fragment is derived from germline genes including IGHV3-33, IGHV3-30, IGHV3-30 3, IGHV3-30 5, IGHV3-33, IGHV1-2, IGHV2-26, IGHV3-21, IGHV3-30, IGHV3-53, IGHV4-34, IGHV4-39, and IGHV4-59.
- the IGHD segment of the antibody or antigen binding fragment is derived from germline genes including IGHD 1-26, IGHD2-2, IGHD2-21, IGHD3-3, IGHD3-9, IGHD3-10, IGHD3-16, IGHD3-22, IGHD5-12, IGHD5-18, IGHD6-19, and IGHD6-6.
- the JH segment of the antibody or antigen binding fragment is derived from germline genes including IGHJ4, IGHJ6, IGHJ3, and IGHJ5.
- the light chain segment of the antibody or antigen binding fragment is derived from germline genes including IGLV1-47, IGLV2-14, IGLV2-23, IGLV7-43, IGLV1-51, IGKV1-39, and IGKV2-28.
- the anti-RhD antibodies of the disclosure may comprise a number of isotypes of the same antibody, or combinations of antibodies with different isotypes.
- the anti-RhD antibodies and antigen binding fragments of the disclosure may comprise, for example, one or more CDRs (e.g., CDR3), the variable region (or portions thereof), the constant region (or portions thereof), or combinations thereof.
- CDRs e.g., CDR3
- the antibody or antigen-binding fragment may comprise a heavy chain variable region and/or a light chain variable region comprising at least one CDR amino acid sequence (e.g., CDR3) or a VH or VL sequence as defined herein, or a sequence having an amino acid sequence that is at least50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95% or at least 99% identical thereto.
- CDR amino acid sequence e.g., CDR3
- VH or VL sequence as defined herein
- the sequences are aligned for optimal comparison purposes (e.g., gaps can be introduced in the sequence of a first amino acid or nucleic acid sequence for optimal alignment with a second amino acid or nucleic acid sequence).
- the amino acid residues or nucleotides at corresponding amino acid positions or nucleotide positions are then compared. When a position in the first sequence is occupied by the same amino acid residue or nucleotide as the corresponding position in the second sequence, then the molecules are identical at that position.
- the determination of percent identity between two sequences can also be accomplished using a mathematical algorithm.
- a mathematical algorithm utilized for the comparison of two sequences is the algorithm of Karlin and Altschul, Proc. Natl. Acad. Sci. USA (1990) 87:2264-2268, modified as in Karlin and Altschul, Proc. Natl. Acad. Sci. USA (1993) 90:5873-5877. Such an algorithm is incorporated into the NBLAST and XBLAST programs of Altschul et al., J. Mol. Biol. (1990) 215:403.
- Gapped BLAST can be utilized as described in Altschul et al., Nucleic Acids Res. (1997) 25:3389-3402.
- PSI-BLAST can be used to perform an iterated search which detects distant relationships between molecules (Id.).
- BLAST Gapped BLAST
- PSI- Blast programs the default parameters of the respective programs (e.g., of XBLAST and NBLAST) can be used (see, e.g., the NCBI website).
- Another preferred, non-limiting example of a mathematical algorithm utilized for the comparison of sequences is the algorithm of Myers and Miller, CABIOS (1988) 4:11-17. Such an algorithm is incorporated in the ALIGN program (version 2.0) which is part of the GCG sequence alignment software package.
- ALIGN program version 2.0
- a PAM 120 weight residue table, a gap length penalty of 12, and a gap penalty of 4 can be used.
- the antibody or antigen binding fragment of the disclosure may bind an epitope or could bind RhD sequence variants and partial D epitopes as described in (Vox Sang 1996;70: 123-131).
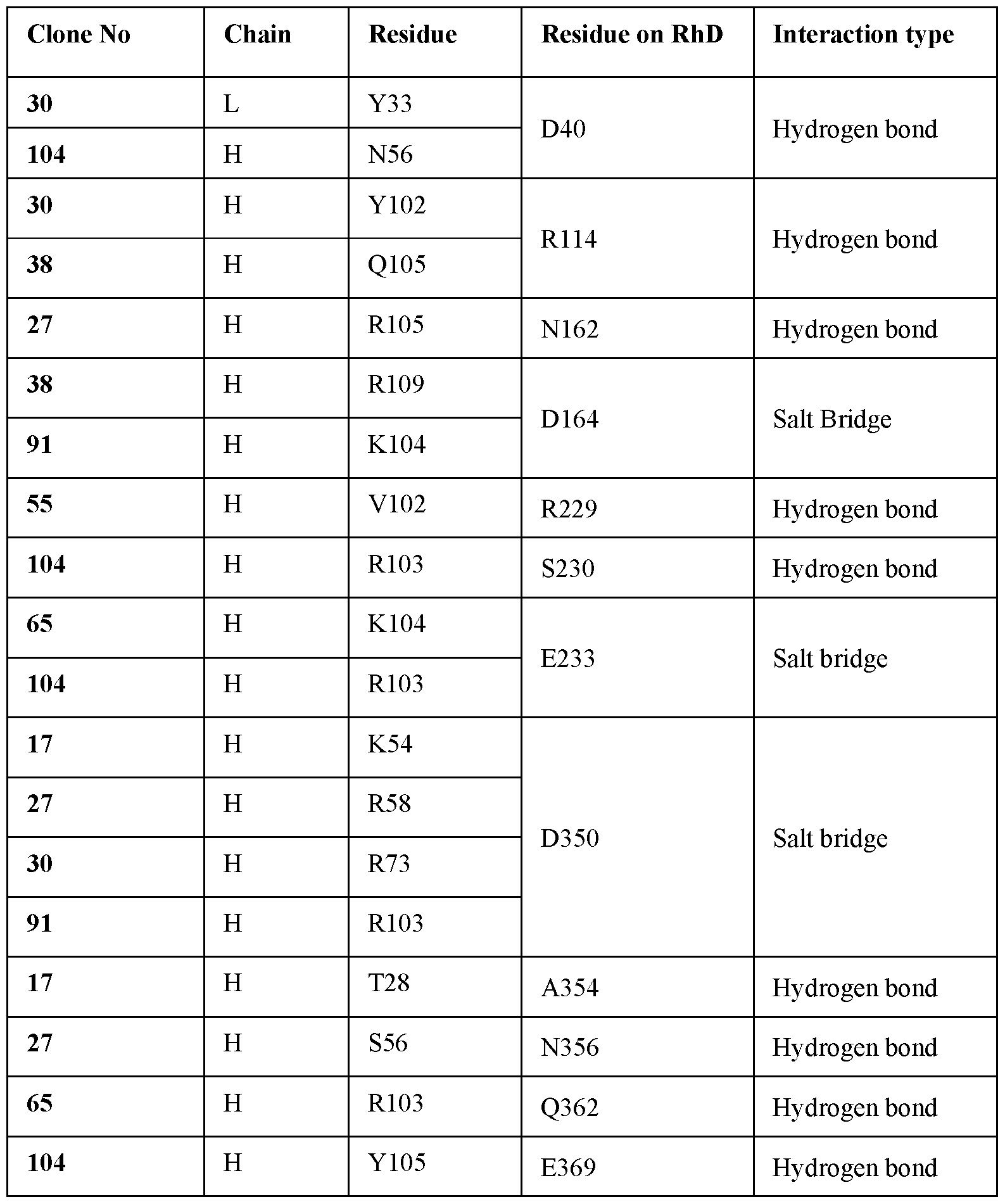
- the antibody or antigen binding fragment contacts residues on RhD including D40, R114, N162, D164, R229, S230, E233, D350, A354, N356, Q362, and/or E369.
- the CDRH3 loop plays a critical role in epitope recognition.
- the CDRH3 loop consists of 16 to 22 amino acids in length.
- antibodies or antigen binding fragments that interact with a greater number of residues on RhD have a greater capacity to inhibit binding of other anti- RhD antibodies or antigen binding fragments.
- the antibodies or binding fragments may be antibodies or fragments whose sequences have been modified to insert one or more amino acids into one or more of its hypervariable regions, for example as described in Jung and Pluckthun, Protein Engineering (1997) 10:9, 959-966; Yazaki et al., Protein Eng. Des Sei. (2004) 17(5):481-9; and US 2007/0280931.
- the antibody or antigen binding fragment comprises the constant region or a portion thereof such as a Fc.
- the constant region or a portion thereof may comprise one or more amino acid substitutions.
- the Fc may comprise one or more amino acid substitutions that alters binding to an Fc receptor and/or effector function.
- the Fc receptor is an Fey receptor. In one embodiment, the Fc receptor is a human Fc receptor. In one embodiment the Fc receptor is an activating Fc receptor. In a specific embodiment the Fc receptor is an activating human Fey receptor, more specifically human FcyRIIIa, FcyRI or FcyRIIa, most specifically human FcyRIIIa.
- the effector function is one or more selected from the group of complement dependent cytotoxicity (CDC), antibody-dependent cell-mediated cytotoxicity (ADCC), antibody-dependent cellular phagocytosis (ADCP), and secretion of cell death mediators such as cytokines or reactive oxygen species (ROS). In a particular embodiment, the effector function is ADCC.
- the Fc is modified by glycosylation, in particular fucosylation and/or galactosylation.
- glycosylation in particular fucosylation and/or galactosylation.
- the extent of glycosylation and type of glycosylation has been shown to influence antibody activity (Kumpel et al. Hum Antibodies Hybridomas. 1994;5(3-4): 143- 51.; Siberil et al Clin Immunol. 2006 Feb-Mar;118(2-3): 170-9.)
- the glycosylation pattern maybe modified by chemical or enzymatic modification. In some embodiments this modification may be carried out by Fc mutagenesis, glycoengineering and subclass switching.
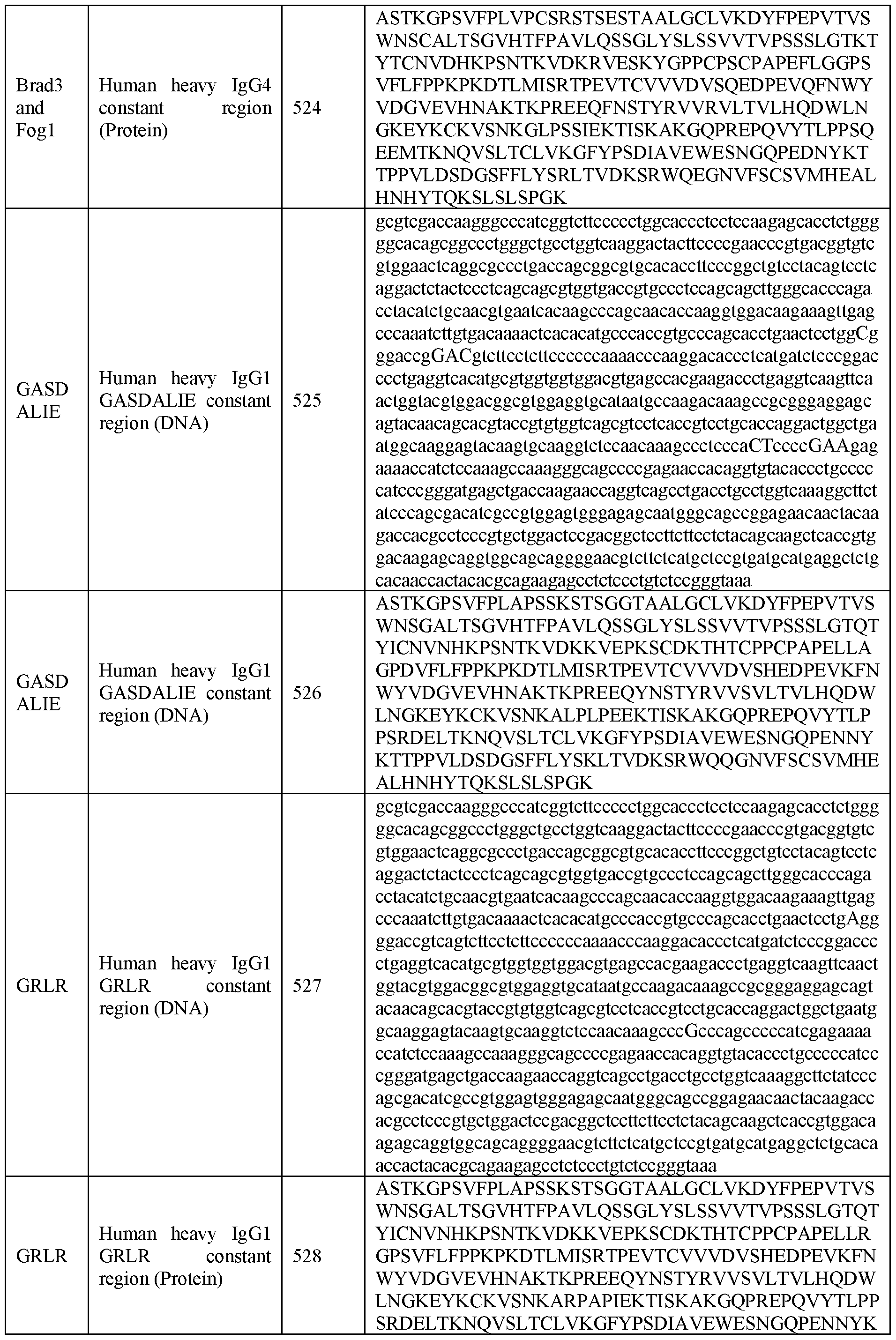
- modifications may be antibody Fc mutated variants that could include the mutations of IgGl-GASDALIE such as G236A, S239D, A330L and I332E (IgGl- GASDALIE antibody) and G236R and L328 (IgGl-GRLR antibody).
- Other modifications could be those referred to in Edwards et al. Enhancement of Antibody-Dependent Cellular Cytotoxicity and Phagocytosis in Anti-HIV-1 Human-Bovine Chimeric Broadly Neutralizing Antibodies. J Virol., 95(13, 2021 or Bournazos et al. Broadly neutralizing anti-HIV-1 antibodies require Fc effector functions for in vivo activity. Cell. 2014 Sep 11 ; 158(6): 1243- 1253.
- the present invention relates to antibody or antigen binding fragments capable of specifically binding to RhD positive (RhD+), but not RhD negative (RhD-), RBCs.
- the antibody or antigen binding fragment is capable of agglutinating RhD+ red blood cells but not RhD- red blood cells.
- the strength of the antibody or antigen binding fragment binding to RhD+ RBCs is correlated with its ability to induce RhD+ RBC agglutination.
- the antibody or antigen binding fragment may also induce RhD+ RBC natural killer (NK) cell-mediated haemolysis.
- NK natural killer
- Antibody or antigen binding fragment induced NK cell- mediated RhD+ RBC haemolysis may be unable to be supressed with caspase, cathepsin, RIPK1, RIPK2, or proteasome inhibitors.
- the antibody or antigen binding fragment may induce NK cell-mediated RhD+ RBC clearance through induction of antibody dependent cell-mediated cytotoxicity (ADCC), wherein the strength of antibody or antigen binding fragment binding to RhD+ RBCs is correlated with its ability to induce ADCC.
- ADCC antibody dependent cell-mediated cytotoxicity
- the antibody or antigen binding fragment may be capable of inducing RhD+ RBC NK cell-mediated haemolysis independent of cell-to-cell contact.
- the antibody or antigen binding fragment may also induce ROS production in the NK cells.
- incubation with the antibody or antigen binding fragment treated RhD+ RBCs may upregulate cytokine and chemokine production in NK cells, for example, one or more of RANTES, MIP-la, MIP-1B, IL-8, IL-9, IL-17, IL-12, IL-lb, IFN-y, TNF-a, CD69, NKP46, and NKG2D.
- the antibody or antigen binding fragment may be further capable of inducing monocyte-mediated clearance of RhD+ RBCs. Induction of monocyte-mediated clearance of RhD+ RBCs may occur through interactions between the antibody or antigen binding fragment and Fey receptors.
- the present disclosure encompasses nucleic acid molecules encoding immunoglobulin light and/or heavy chain variable regions for anti-RhD antibodies, vectors comprising such nucleic acids, and host cells capable of producing the anti-RhD antibodies of the disclosure.
- the nucleic acid molecules encode, and the host cells are capable of expressing the anti-RhD antibodies and antibody-binding fragments, as well as fusion proteins and chimeric antigen receptors containing them.
- An anti-RhD antibody of the disclosure can be prepared by recombinant expression of immunoglobulin light and heavy chain genes in a host cell.
- a host cell is transfected with one or more recombinant expression vectors carrying DNA fragments encoding the immunoglobulin light and heavy chains of the antibody such that the light and heavy chains are expressed in the host cell and, optionally, secreted into the medium in which the host cells are cultured, from which medium the antibodies can be recovered.
- Standard recombinant DNA methodologies are used to obtain antibody heavy and light chain genes, to incorporate these genes into recombinant expression vectors and to introduce the vectors into host cells, such as those described in Molecular Cloning; A Laboratory Manual, Second Edition (Sambrook, Fritsch and Maniatis (eds), Cold Spring Harbor, N. Y., 1989), Current Protocols in Molecular Biology (Ausubel, F. M. et al., eds., Greene Publishing Associates, 1989) and in U.S. Pat. No. 4,816,397.
- DNA fragments encoding the light and heavy chain variable regions are first obtained. These DNAs can be obtained by amplification and modification of germline DNA or cDNA encoding light and heavy chain variable sequences, for example using the polymerase chain reaction (PCR).
- PCR polymerase chain reaction
- Germline DNA sequences for human heavy and light chain variable region genes are known in the art (see, e.g., the "VBASE" human germline sequence database; see also Kabat et al., Sequences of Proteins of Immunological Interest, Fifth Edition, U.S. Department of Health and Human Services, NIH Publication No. 91-3242. 1991; Tomlinson et al., J. Mol. Biol. (1992) 22T: 116-198; and Cox et al., Eur. J. Immunol. (1994) 24:827-836).
- DNA fragments encoding anti-RhD antibody related VH and VL segments are obtained, these DNA fragments can be further manipulated by standard recombinant DNA techniques, for example to convert the variable region genes to full-length antibody chain genes, to Fab fragment genes or to a scFv gene.
- a VH- or VL - encoding DNA fragment is operatively linked to another DNA fragment encoding another protein, such as an antibody constant region or a flexible linker.
- the term “operatively linked,” as used in this context, is intended to mean that the two DNA fragments are joined such that the amino acid sequences encoded by the two DNA fragments remain in-frame.
- the isolated DNA encoding the VH region can be converted to a full-length heavy chain gene by operatively linking the VH-encoding DNA to another DNA molecule encoding heavy chain constant regions (CHI, CH2, CH3 and, optionally, CH4).
- CHI heavy chain constant regions
- the sequences of human heavy chain constant region genes are known in the art (see, e.g., Kabat et al., Sequences of Proteins of Immunological Interest, Fifth Edition, U.S. Department of Health and Human Services, NIH Publication No. 91-3242, 1991) and DNA fragments encompassing these regions can be obtained by standard PCR amplification.
- the isolated DNA encoding the VL region can be converted to a full-length light chain gene (as well as a Fab light chain gene) by operatively linking the VL-encoding DNA to another DNA molecule encoding the light chain constant region, CL.
- the sequences of human light chain constant region genes are known in the art (see, e.g., Kabat et al., Sequences of Proteins of Immunological Interest, Fifth Edition, U.S. Department of Health and Human Services, NTH Publication No. 91-3242, 1991) and DNA fragments encompassing these regions can be obtained by standard PCR amplification.
- the light chain constant region can be a K or constant region, but in certain embodiments is a K constant region.
- the VH- and VL-encoding DNA fragments can be operatively linked to another fragment encoding a flexible linker, for example, encoding the amino acid sequence (Gly4 ⁇ Ser)3 , such that the VH and VL sequences can be expressed as a contiguous single-chain protein, with the VH and VL regions joined by the flexible linker (see, e.g., Bird et al., Science (1988) 242:423-426; Huston et al., Proc. Natl. Acad. Sci. USA (1988) 85:5879- 5883; McCafferty et al., Nature (1990) 348:552-554).
- a flexible linker for example, encoding the amino acid sequence (Gly4 ⁇ Ser)3 , such that the VH and VL sequences can be expressed as a contiguous single-chain protein, with the VH and VL regions joined by the flexible linker (see, e.g., Bird et al
- the antibody genes are inserted into the expression vector by standard methods (e.g., ligation of complementary restriction sites on the antibody gene fragment and vector, or blunt end ligation if no restriction sites are present).
- the expression vector Prior to insertion of the anti-RhD antibody- related light or heavy chain sequences, the expression vector can already carry antibody constant region sequences.
- one approach to converting the anti-RhD monoclonal antibody related VH and VL sequences to full-length antibody genes is to insert them into expression vectors already encoding heavy chain constant and light chain constant regions, respectively, such that the VH segment is operatively linked to the CH segment(s) within the vector and the VL segment is operatively linked to the CL segment within the vector.
- the recombinant expression vector can encode a signal peptide that facilitates secretion of the antibody chain from a host cell.
- the antibody chain gene can be cloned into the vector such that the signal peptide is linked in-frame to the amino terminus of the antibody chain gene.
- the signal peptide can be an immunoglobulin signal peptide or a heterologous signal peptide (i.e., a signal peptide from a non-immunoglobulin protein).
- the recombinant expression vectors of the disclosure carry regulatory sequences that control the expression of the antibody chain genes in a host cell.
- the term “regulatory sequence” is intended to include promoters, enhancers and other expression control elements (e.g., polyadenylation signals) that control the transcription or translation of the antibody chain genes.
- Such regulatory sequences are described, for example, in Goeddel, Gene Expression Technology: Methods in Enzymology 185, Academic Press, San Diego, Calif., 1990. It will be appreciated by those skilled in the art that the design of the expression vector, including the selection of regulatory sequences may depend on such factors as the choice of the host cell to be transformed, the level of expression of protein desired, etc.
- Suitable regulatory sequences for mammalian host cell expression include viral elements that direct high levels of protein expression in mammalian cells, such as promoters and/or enhancers derived from cytomegalovirus (CMV) (such as the CMV promoter/enhancer), Simian Virus 40 (SV40) (such as the SV40 promoter/enhancer), adenovirus, (e.g., the adenovirus major late promoter (AdMLP)) and polyoma.
- CMV cytomegalovirus
- SV40 Simian Virus 40
- AdMLP adenovirus major late promoter
- the recombinant expression vectors of the disclosure can carry additional sequences, such as sequences that regulate replication of the vector in host cells (e.g., origins of replication) and selectable marker genes.
- the selectable marker gene facilitates selection of host cells into which the vector has been introduced (see, e.g., US 4,399,216, US 4,634,665 and US 5,179,017).
- the selectable marker gene confers resistance to drugs, such as G418, hygromycin or methotrexate, on a host cell into which the vector has been introduced.
- Suitable selectable marker genes include the dihydrofolate reductase (DHFR) gene (for use in DHFR- host cells with methotrexate selection/amplification) and the neo gene (for G418 selection).
- DHFR dihydrofolate reductase
- neo gene for G418 selection.
- the expression vector(s) encoding the heavy and light chains is transfected into a host cell by standard techniques.
- the various forms of the term “transfection” are intended to encompass a wide variety of techniques commonly used for the introduction of exogenous DNA into a prokaryotic or eukaryotic host cell, e.g., electroporation, lipofection, calcium-phosphate precipitation, DEAE— dextran transfection and the like.
- eukaryotic cells e.g., mammalian host cells
- expression of antibodies is performed in eukaryotic cells, e.g., mammalian host cells, of optimal secretion of a properly folded and immunologically active antibody.
- eukaryotic cells e.g., mammalian host cells
- Exemplary mammalian host cells for expressing the recombinant antibodies of the disclosure include Chinese Hamster Ovary (CHO cells) (including DHFR- CHO cells, described in Urlaub and Chasin, Proc. Natl. Acad. Sci. USA (1980) 77:4216-4220, used with a DHFR selectable marker, for example, as described in Kaufman and Sharp, Mol. Biol.
- FUT8 KO CHO cells (Yang et al, Frontiers Chem, 9 (2021)) have also been shown to produce completely non- fucosylated antibodies.
- Other ways to reduce fucosylation include manipulation of culture conditions (Konno et al Cytotechnology, 64(3) (2012)).
- Host cells can also be used to produce portions of intact antibodies, such as Fab fragments or scFv molecules. It is understood that variations on the above procedure are within the scope of the present disclosure. For example, it can be desirable to transfect a host cell with DNA encoding either the light chain or the heavy chain (but not both) of an anti- RhD antibody of this disclosure.
- an anti-RhD antibody of the disclosure can be purified by any method known in the art for purification of an immunoglobulin molecule, for example, by chromatography (e.g., hydroxyapatite chromatography, hydrophobic interaction chromatography, gel electrophoresis, dialysis, affinity chromatography (e.g., protein A affinity chromatography or protein G chromatography), size exclusion chromatography, ion exchange, centrifugation, differential solubility, or by any other standard technique for the purification of proteins.
- chromatography e.g., hydroxyapatite chromatography, hydrophobic interaction chromatography, gel electrophoresis, dialysis, affinity chromatography (e.g., protein A affinity chromatography or protein G chromatography), size exclusion chromatography, ion exchange, centrifugation, differential solubility, or by any other standard technique for the purification of proteins.
- affinity chromatography e.g., protein A affinity chromatography or protein G chromat
- a protease inhibitor such as PMSF may be included in any of the foregoing steps to inhibit proteolysis and antibiotics may be included to prevent the growth of adventitious contaminants.
- supernatants can be filtered and/or separated from cells expressing the protein, e.g., using continuous centrifugation.
- anti-RhD antibodies of the present disclosure and/or binding fragments can be fused to heterologous polypeptide sequences described herein or otherwise known in the art to facilitate purification.
- a poly-histidine tag for example, a hexahistidine tag, or an influenza virus hemagglutinin (HA) tag, or a Simian Virus 5 (V5) tag, or a FLAG tag, or a glutathione S-transferase (GST) tag.
- HA hexahistidine tag
- V5 Simian Virus 5
- FLAG tag or a glutathione S-transferase (GST) tag.
- GST glutathione S-transferase
- a protein comprising a hexa-his tag is purified by contacting a sample comprising the protein with nickel-nitrilotriacetic acid (Ni-NTA) that specifically binds a hexa-his tag immobilized on a solid or semi-solid support, washing the sample to remove unbound protein, and subsequently eluting the bound protein.
- Ni-NTA nickel-nitrilotriacetic acid
- a ligand or antibody that binds to a tag is used in an affinity purification method.
- the anti-RhD antibodies and fragments of the disclosure may be in the form of compositions comprising one or more of the anti-RhD antibodies or binding fragments and one or more carriers, excipients and/or diluents.
- the form of the composition e.g., dry powder, liquid formulation, etc.
- the excipients, diluents and/or carriers used will depend upon the intended uses of the antibody or fragment, for therapeutic uses, the mode of administration.
- the compositions consist of two or more anti-RhD antibodies or binding fragments.
- the compositions consist of three or more anti-RhD antibodies or binding fragments.
- an antibody or fragment or composition of the disclosure in a prophylactic manner for the prevention of the alloimmunization of a RhD- negative woman, immediately after the birth of a Rh-positive child, and to prevent, during subsequent pregnancies, haemolytic disease of the newborn; during abortions, extrauterine pregnancies in a situation of RhD incompatibility or in transplacental haemorrhages resulting from amniocentesis, chorionic biopsies, or traumatic obstetric manipulations in a situation of incompatibility for Rhesus D.
- donor blood is first separated into plasma and PBMC fractions by centrifugation.
- the total IgG content may be purified by incubation of the plasma fraction with, for example, Protein G agarose.
- RhD-specific antibodies may be further isolated by incubation of the total IgG pool with RhD-positive RBCs.
- Antibody-bound RBCs are preferably washed to remove non-specific binders, before eluting antibodies with, for example, EDTA/glycine acid, resulting in a pool of RhD-specific antibodies.
- a similar selection step can be performed by incubating the total IgG pool with RhD-negative RBCs, to isolate antibodies that bind to non-RhD antigens present on the RBCs.
- RhD-specific and non-RhD specific pools of antibodies may then be subjected to peptic digest and mass spectrometry to identify the antibody sequences.
- sequences identified in only the RhD-specific pool (considered to be RhD-specific binders) can be used for further analysis.
- B-lymphocytes may be analysed for relative antibody transcription levels.
- Live CD 19+ IgG+ cells may be enriched from donor PBMCs by fluorescence activated cell sorting (FACS), whereafter single-cell RNA sequencing can be performed.
- FACS fluorescence activated cell sorting
- BCRs B cell receptors
- Peripheral blood was obtained from 27 ‘super producers’ participating in the RhD program at Australian Red Cross Lifeblood (ARCL).
- Peripheral Blood Mononuclear cells (PBMC) were purified from the blood using standard techniques (see for example Panda and Ravindran, 2013, Bio-protocol 3(3): and Panda et al 2012, PLoS Pathog 8(5). RBC were lysed from the final cell pellet and subsequently, PBMCs were washed with cold PBS and frozen at -80 °C using heat-inactivated 90% foetal calf serum and 10% dimethylsulfoxide (DMSO).
- DMSO dimethylsulfoxide
- donor blood is first separated into plasma and PBMC fractions by centrifugation.
- the total immunoglobulin fraction is purified by affinity chromatography (using Protein G agarose).
- RhD-specific antibodies are further isolated by incubation of the total IgG pool with RhD-positive RBCs.
- Antibody-bound RBCs are washed to remove nonspecific binders, before eluting antibodies with EDTA/glycine acid, resulting in a pool of RhD-specific antibodies.
- the antibodies prepared as above were analysed by mass spectrometry-based proteomics. To obtain small peptides for further analysis samples were prepared by digesting with AspN, GluC and trypsin digestion and chymotrypsin as follows.
- Antibodies (2 pg) were incubated with 50 mM ammonium bicarbonate, 8M Urea and 1 mM DTT for 30 minutes and 37°C. Antibodies were alkylated using 10 mM iodoacetamide for 1 hour, and subsequently quenched with 10 mM DTT. Samples were diluted with 50 mM ammonium bicarbonate to reduce UREA to IM concentration. 100 ng trypsin, AspN or GluC was added for overnight digestion at 37°C. Peptides were then desalted using Cl 8 STAGE tips and collected into new tubes by centrifugation. The collected peptides were lyophilized to dryness using a CentriVap (Labconco), before reconstituting in 10 pl 0.1% FA/2% ACN ready for mass spectrometry analysis.
- Peptides (5 pl) were separated by reverse-phase chromatography on a Cis fused silica column packed into an emitter tip (lonOpticks), using a nano-flow HPLC (M-class, Waters).
- Raw data files were analysed by MaxQuant vl .6.17 software using the integrated Andromeda search engine.
- Experiment type was set as TIMS-DDA with no modification to default settings.
- Data was searched against an in-house generated heavy /light chain IgG .fasta database and a separate reverse decoy database using a strict trypsin, AspN, GluC or chymotrypsin specificity allowing up to 2 missed cleavages.
- the minimum required peptide length was set to 7 amino acids.
- Modifications Carbamidomethylation of Cys was set as a fixed modification, while N-acetylation of proteins and oxidation of Met were set as variable modifications.
- First search peptide tolerance was set at 10 ppm and main search set at 20 ppm (other settings left as default).
- Peptide-spectrum match and protein identifications were filtered using a target-decoy approach at an FDR of 1%.
- Label-free quantification (LFQ) quantification was selected, with a minimum ratio count of 2.
- Peptide-spectrum match scores and protein identifications were filtered using a target-decoy approach at an FDR of 1%.
- Missing values were imputed by using Missing Not At Random (MNAR) method. This was achieved by substituting ‘NAs’ with numbers that were drawn from a normal distribution with a mean that is left-shifted from the sample mean by 1.8 standard deviation with a width of 0.3.
- MNAR Missing Not At Random
- the R-package limma (v. 3.50.1) was used to perform the differential analysis to compare the differential protein abundance of protein sequences identified in RhdNeg vs RhdPos samples. Protein sequences were determined to be differentially expressed if the false discovery rate (FDR) was ⁇ 0.05 and fold change 2.
- FDR false discovery rate
- RhD specific antibody sequences were extracted and converted to a FASTA file and then subjected to analysis. The data sets were combined, and IGHM sequences were removed from the dataset. Heavy chains were assigned new names such as clone- 1, clone-2, etc., if the sequences were unique. In cases where cells had identical heavy-light pairs, only one cell was included in the analysis, and the repeated sequences from other cells were excluded. Thus, combining the transcriptomics sequence information with the proteomic sequence information to further assemble and derive the complete amino acid sequences of the most abundant mAbs in the RhD specific IgG subset proved very powerful and efficient.
- Example 2 The below method was used to identify antibodies with high affinity.
- the antibody genes identified in Experiment 1 were then synthesised and cloned into vectors expressing human antibody gamma, kappa, or lambda constant regions ( Figure 2). Each antibody variable gene was cloned into a DNA plasmid expressing its natural heavy and light constant region according to single cell NGS data. The paired antibody heavy and light expression DNA plasmids were co-transfected into Expi-CHO cells according to the manufacturer’s instructions (ThermoFisher Scientific). The antibodies were purified with protein G agarose and used for further characterization.
- RhD+ (ROr phenotype: Dce/dce) and RhD- (r’r phenotype: dCe/dce) O Blood group was collected and centrifuged at 200xg for 10 min. Plasma was discarded and 3mL DPBS (+2mM EDTA) was added to the cells. The tube was centrifuged at 200xg for 10 min and huffy coat and supernatant was discarded. The cells were washed two more times with DPBS (+2mM EDTA) and used in the experiments immediately.
- the cryovial was incubated at 37 °C for 2min, inverted gently and centrifuged at 1500xg for 5 min. The supernatant was discarded except the last 50pl layering RBC pellet. Packed RBCs were resuspended and 12% NaCl (25 pl, over 5min) was added. The cells were incubated at RT for 3 min to equilibrate. 237.5 pl of 1.6% NaCl was added (over 5 min) followed by an extra 0.5 mL 1.6% NaCl (also over 5min).
- RBCs with Bromelain removes surface glycans and is expected to improve antibody access which can enhance ADCC.
- the washed RBCs were resuspended in DPBS and treated with Bromelain as follows: 2% RBC in DPBS was mixed with 0.5% Bromelain (w/v) (1 :2 ratio) and incubated at 37 °C for 10 min. The cells were centrifuged at 1500xg for 5 min and washed with DPBS twice.
- Agglutination assay was performed to assess RhD binding of candidate antibodies. Briefly, 50mL Bromelain treated O blood group RhD+ or RhD- RBC (1% haematocrit), 25mL mAb and 25mL anti-IgG secondary antibody _AHG colour (BIO-RAD laboratories #804115) were added to each well of U-bottom 96-well plate and incubated 90 min at RT. Agglutination was then visualized by the naked eye.
- mAbs 50mL were incubated with 50mL Bromelain-treated O blood group RhD+ or RhD- RBC (200,000) in U-bottom 96-well plate at 37 °C for 30 min. The cells were washed with 200mL wash buffer (DPBS containing 0.5% BSA and 2mM EDTA) and centrifuged at 1500 xg for 2 minutes. The supernatant was discarded, and 1/1000 dilution of Fab goat antihuman IgG (H&L) Texas Red (Rockland antibodies and assays # 809-1902) was added to each well.
- wash buffer DPBS containing 0.5% BSA and 2mM EDTA
- the plate was incubated at room temperature for 30 min and centrifuged at 1500 xg for 2 minutes. The plate was washed once more with 200mL wash buffer and the cells were resuspended in 50mL wash buffer. The samples were acquired on an Fortessa instrument (BD) and analysed using FlowJo vlO.6.2. Commercial (non-enriched) polyclonal RhD-pIgG- 1 was used as a control in this experiment.
- BD Fortessa instrument
- Commercial (non-enriched) polyclonal RhD-pIgG- 1 was used as a control in this experiment.
- a competition RBC binding assay was conducted to evaluate whether the anti-RhD mAbs target the same or overlapping epitopes and to what extent they can compete with each other for binding.
- FITC-conjugated anti-RhD mAbs were evaluated the presence and absence of other anti-RhD mAbs. Briefly, 100,000 bromelain-treated RhD+ RBCs (50pL) were incubated with unconjugated mAbs (50pL) for 40 minutes at 37°C then centrifuged at 1500 xg for 1 minute to remove unbound antibodies. Supernatant discarded and RBCs were washed with DPBS. Afterwards, FITC-conjugated anti-RhD mAbs were added (lOOpL) and the cells were incubated at 37°C for 40 minutes. The cells were washed as above and resuspended in DPBS. The samples were acquired on an Fortessa instrument (BD) and analysed using FlowJo vl0.6. 2. Brad3 and Fogl mAbs served as positive controls for anti- RhD mAb binding.
- BD Fortessa instrument
- the monocytic THP-1 cell line was grown in RPMI culture medium + 10% FCS at 37 °C and 5% CO2 atmosphere. 10,000 cells were incubated with 1/40 dilutions of mouse anti-human CD64 (FcyRI)- PE Cy7 (BD Biosciences; Cat no. 561191), 1/10 dilutions of mouse anti-human CD32 (FcyRII)- PE (BD Biosciences; Cat no. 552884), and 1/10 dilutions of mouse anti-human CD16 (FcyRIII)- FITC (BD Biosciences; Cat no. 555406) for 30 min on ice to assess the expression of FcyRs. The cells were washed with DPBS by centrifugation at 500xg for 5 min. The samples were acquired on an Fortessa instrument (BD) and analysed using FlowJo vlO.6.2.
- BD Fortessa instrument
- monocytic THP-1 cells were labelled with 1 pM CellTraceTM Violet (CTV) (Invitrogen; Cat no.C34571) in DPBS at 37°C for 1 hours with gentle shaking.
- CTV CellTraceTM Violet
- the cells were washed with RPMI+ 10% FCS and centrifuged at 500xg 5 min.
- RPMI+ 10% FCS was added to the cells and incubated at 37°C for 15 min followed by washing and resuspending in RPMI+ 10% FCS (4 x 10 5 cells/ml).
- RhD+ RBCs Bromelain-treated RhD+ RBCs were resuspended in RPMI media containing 3% human AB serum and 7% FCS at a concentration of 80,000 cells per mL. Then 25mL RhD+ RBC (20,000 /well), 25mL purified antibody (lOOng/well or lug/mL in RPMI + 3% human AB serum + 7% FCS), and 50mL effector cells (600,000 monocyte-depleted PBMC/well) were incubated overnight at 37°C.
- Triton X-100 (ThermoFisher Scientific #28314) was added to the control well (No antibody control) and incubated for 5 minutes at 37°C to achieve the maximal haemoglobin release. The cells were then centrifuged at 1500xg for 2 minutes and the supernatants were collected. A 1/100 dilution of the supernatant was used to detect the haemoglobin in each well using Human HB (Hemoglobin) ELISA Kit (Elabscience #E- EL-H0415) according to the manufacturer's instructions.
- the percent specific RBC lysis was calculated using the formula: 100 * (experimental release - spontaneous release)/(maximum release - spontaneous release).
- Enriched RhD-pIgG-1, prepared according to Experiment 2 was used as a control in these assays.
- NK cells were enriched using human NK cell isolation kit (Miltenyi Biotec #130-092-657) according to the manufacturer’s instruction.
- 100,000 Bromelain-treated RhD+ RBCs (25 pL), with lOOng/mL antibody (25 pL) and 100,000 purified NK cells (50 pL) were incubated overnight at 37°C. Then the supernatant was harvested, and the presence of cytokines or chemokines was investigated using Bioplex kit (Bio-rad) according to the manufacturer’s instruction.
- Commercial (non-enriched) polyclonal RhD-pIgG-1 was used as a control in these assays.
- Micro-Agglutination assay 50mL Bromelain treated RBCs (1% haematocrit), 25mL antibody and 25mL anti-IgG secondary antibody _AHG colour (BIO-RAD laboratories #804115) were added to each well of U-bottom 96-well plate and incubated 90 min at RT. Agglutination was then visualized by the naked eye. The antibody concentration used in the assay was between 1 and 25 pg/ml for the various anti-RhD mAbs variants of this invention. Indirect antiglobulin test (IAT) assay was performed using Bio-Rad LISS/Coombs kit (BIO-RAD) according to the manufacturers’ instruction.
- IAT Indirect antiglobulin test
- FACS-based binding assay To evaluate the binding of the antibody (mAbs) to RhD antigen, a flow cytometry analysis was performed. In brief, antibody (50mL) was incubated with 50mL Bromelain-treated RBCs (200,000) in U-bottom 96-well plate at 37 °C for 30 min. The cells were washed with 200mL DPBS and centrifuged at 1500 xg for 2 minutes. The supernatant was discarded, and 1/1000 dilution of Fab goat anti-human IgG (H&L) Texas Red (Rockland antibodies and assays # 809-1902) was added to each well. The plate was incubated at room temperature for 30 min and centrifuged at 1500 xg for 2 minutes.
- H&L Fab goat anti-human IgG
- Texas Red Rockland antibodies and assays # 809-1902
- the plate was washed once more with 200mL wash buffer and the cells were resuspended in 50mL wash buffer.
- the samples were acquired on an Fortessa instrument (BD) and analysed using FlowJo vlO.6.2.
- Commercial (non-enriched) polyclonal RhD-pIgG-1 was used as a control in this assay.
- RhD-pIgG-1 A competition RBC binding assay was conducted to evaluate whether commercial polyclonal (non-enriched) RhD-pIgG-1 can suppress the binding of anti-RhD mAbs. This assay can demonstrate the relative binding affinities and potential epitope overlap between the polyclonal RhD-pIgG-1 and the anti-RhD mAbs.
- FITC-conjugated anti-RhD mAbs were evaluated the presence and absence of polyclonal IVIg (Pool of non-RhD antibodies) and RhD-pIgG-1 (derived from Australian donors). Briefly, 100,000 bromelain-treated RhD+ RBCs (50pL) were incubated with 5pg IVIg or RhD-pIgG-1 (50pL) for 30 minutes at 37°C then centrifuged at 1500 xg for 1 minute to remove unbound antibodies. Supernatant discarded and RBCs were washed with DPBS.
- RhD-pIgG consists of a pool of immunoglobulins (antibodies) with a higher titre of anti-RhD specific IgG compared to the titre found in normal plasma-derived IgG.
- Antibodies Commercial clinically used product is currently derived from the plasma of RhD- donors who have developed anti-RhD antibodies due to hemolytic disease of the newborn (HDN) or immunization against RhD+ RBCs. However, these preparations contain a mixture of both RhD-specific and non-RhD antibodies.
- RhD-pIgG-1 derived from Australian donors
- PBS PBS
- RhD+ RBCs RBCs
- the RBCs were then washed at least six times with PBS to remove non-specific binders.
- 16 mL of 50 mM glycine, 0.9% NaCl, 2 mM EDTA (pH 2.7) was added, the mixture was vortexed for 5 seconds, and incubated on ice for 2 minutes.
- the RBCs were immediately centrifuged at 2500xg for 5 minutes, and the supernatant was collected.
- the acidic pH was neutralized by adding 2.1 mL of 1 M Tris, 5.2% NaCl (pH 8), resulting in a pool of eluted RhD-specific antibodies.
- Anti-RhD monoclonal antibodies agglutinated RhD+ RBC.
- the agglutination endpoint concentration or agglutination titre indicates the lowest concentration of the antibody at which visible agglutination of RBCs occurs.
- a lower agglutination endpoint concentration suggests higher antibody affinity and potency, as it requires less antibody to achieve agglutination.
- RhD-pIgG-1 a polyclonal product containing anti-RhD antibodies derived from Australian blood donors and currently available on the market, agglutinated RhD+ RBCs while also exhibiting slight non-specific reactivity against RhD- RBCs.
- K kappa light chain
- L lambda light chain
- the anti-RhD mAbs displayed specific binding to RhD+ RBCs. Notably, Clone-30 mAb exhibited binding to over 75% of RhD+ RBCs, while seven mAbs (Clone 18, 27, 38, 55, 65, 91 and 104) showed binding to 51-75% of the cells.
- Four mAbs (Clone 21a, 34, 44 and 107) demonstrated moderate binding, interacting with 25-50% of the RBCs, and four other mAbs (Clone la, 17, 75 and 142) exhibited weak binding, attaching to less than 25% of the cells (Table 3).
- RhD mAbs including Brad3, Fogl, and commercial RhD-pIgG- 1, demonstrated specific binding to wild-type (WT) RhD variants (ROr, R1R1, and R2R2) but not to the rr variant (RhD- RBC).
- WT wild-type
- RhD variants ROr, R1R1, and R2R2
- RhD- RBC rr variant
- Testing Weak RhD type 2 from different donors revealed variable binding for Clone-17, Clone-75, and Clone-142. These mAbs also did not bind to Weak RhD type 1. Additionally, Clone-17, Clone-75, Clone-34, and Clone-55 did not bind to Weak RhD type 4. The other tested mAbs demonstrated satisfactory binding to the Weak RhD variants.
- preferred embodiments of this invention include selecting use of mAbs which have a predominance of IgGl subclass to enhance ADP activity
- the prozone-like effect observed at higher concentrations further underscores the potential for fine-tuning antibody formulations to achieve optimal therapeutic outcomes.
- the unique characteristics of these anti-RhD mAbs highlight their potential for development into effective therapeutic agents for conditions involving RhD+ RBCs.
- ADCC antibodydependent cellular cytotoxicity
- the mAbs of this invention show a comparable biological activity to other commercially available antibodies when evaluating cytokine and chemokine secretion by NK cells.
- the cytokine and chemokine secretion profile of the mAbs closely resembled that of commercial RhD-pIgG-1, indicating a similar pattern of immune response induced by both antibody preparations.
- cytokines and chemokines found to be upregulated were RANETS, MIP-la, MIP-1B, IL-8, IL-9, IL-17, IL-12, IL-lb, IFN-y, and TNF-a ( Figure 10A).
- These immune mediators play diverse roles in immune responses, including recruiting and activating immune cells to sites of inflammation.
- NK cells While their individual roles in inducing RBC death vary, the combined effect of these immune mediators may contribute to RBC clearance through phagocytosis and immune cell-mediated cytotoxicity. This enhanced immune response suggests that NK cells are activated upon encountering RBCs opsonised with anti- RhD antibodies, and highlights the complex interplay between these antibodies, NK cells, and RBCs in RhD-related immune-mediated haemolytic disorders.
- the nucleotide sequences of the antibody heavy and light variable genes were analysed using IMGT-V-Quest, which allowed for the identification and recognition of their respective germline gene counterparts.
- the mAbs of this invention displayed a CDRH3 size ranging from 15 to 22 amino acids (moderate-long length) (Table 4 and Table 5) and IGHV somatic hypermutation (SHM) levels ranging from 0.69% to 13.12% (Table 5).
- the CDRH3 size plays a critical role in antibody binding specificity and affinity, with longer CDRH3 regions often associated with increased antigen recognition and potential for enhanced binding to target epitopes.
- the IGHJ mutation displays relatively diverse levels, ranging from 0% to 20.83% (Table 5). Based on the findings, the level of SHM in the IGHV gene can be categorised as relatively low to moderate.
- the percentage of IGHV gene mutation indicates the extent to which the antibody sequences have undergone changes during the immune response, with higher percentages suggesting greater diversification. In this context, the observed mutation levels fall within a moderate range, implying that the antibodies have undergone some degree of mutation, but not to the extent seen in highly mutated antibodies.
- the CDRH3 size also plays a critical role in antibody binding specificity and affinity, with longer CDRH3 regions often associated with increased antigen recognition and potential for enhanced binding to target epitopes.
- the identified germline genes for the lambda light chains were IGLV1- 47, IGLV2-14, IGLV2-23, IGLV7-43, IGLV1-51, while for the kappa light chains, the genes were IGKV1-39 and IGKV2-28.
- the predominant germline genes for both light chain types were IGLV1-47 and IGKV1-39 ( Figure 12).
- the IGLV region showed a mutation range of 1.04% to 5.38%, while the J segment mutations ranged from 0% to 13.16% (Table 6).
- the isolated antibodies exhibited a higher SHM rate in IGHV and longer CDR compared to the light chain ( Figure 14A-B). Alignment of antibody heavy variable genes showed that the mAbs possess unique sequences in their CDRH3 regions.
- Antibody expression vectors containing heavy variable and constant (IgGl, IgGl- GASDALIE, IgGl-GRLR and IgG3) genes and light variable and constant (kappa) genes were co-transfected into Expi- Chinese Hamster Ovary (Expi-CHO) cells using an Expi-CHO expression system kit. Supernatants containing antibodies were harvested 4-5 days after transfection and filtered using 0.22 mm filters. The antibodies were purified using Protein G Agarose Fast Flow then buffer exchanged into PBS using Amicon 50 kDa spin membranes (Millipore) and sterilized.
- heavy chain expression plasmids IgGl and IgGl-GASDALIE
- light chain expression plasmids kappa
- the purified antibodies were run on NuPage 4-12% Bis-Tris Gel and either incubated with Goat horseradish peroxidase (HRP) conjugated anti-Human (H+L) or with biotinylated Lectin followed by Avidin HRP. Reduction of fucosylation was confirmed by western blot using IgGl-Afucosylated purified mAbs compared with IgGl mAbs.
- HRP horseradish peroxidase
- H+L horseradish peroxidase conjugated anti-Human
- Avidin HRP biotinylated Lectin followed by Avidin HRP. Reduction of fucosylation was confirmed by western blot using IgGl-Afucosylated purified mAbs compared with IgGl mAbs.
- RhD- O blood group RBC carrying RhD antigen was agglutinated in the presence of all Brad3 and Fogl variants but not VRC03 variants.
- RhD- O blood group RBC (r’r) was agglutinated with none of the antibodies, confirming the specificity of Brad3 and Fogl against RhD antigen.
- Agglutination is a very sensitive assay and the agglutination endpoint concentration of both Brad3 and Fogl variants was 5ng/mL showing that the agglutination activity was maintained in engineered anti-RhD mAbs.
- Figure 17A illustrates the method used to assess NK cell activation.
- the impact of these modifications on NK cell activation is shown in Figure 17B.
- Figure 17B Although there were some differences observed among the variants, there was no significant difference in the enhancement of NK cell activation between the GASDALIE Fc variant and the IgGl A fucosylated variant, or the variant with both modifications. Consistent results were obtained in the ADCC assay, indicating that these antibody modifications improved ADCC compared to WT variant and demonstrated activity comparable to enriched RhD-pIgG-1 ( Figure 17C).
- RhD-pIgG-2 2 mL (1500IU) of commercially available RhD-pIgG-2 was diluted with PBS and incubated with 8 mL of packed RhD+ RBCs (ROr variant). The RBCs were then washed at least six times with PBS to remove non-specific binders. To elute the RBC-bound antibodies, 16 mL of 50 mM glycine, 0.9% NaCl, 2 mM EDTA (pH 2.7) was added, the mixture was vortexed for 5 seconds and incubated on ice for 2 minutes.
- the RBCs were immediately centrifuged at 2500xg for 5 minutes, and the supernatant was collected and neutralized by adding 2.1 mL of 1 M Tris, 5.2% NaCl (pH 8). To remove potential RBC protein contaminants, further enrichment was achieved by affinity chromatography using Protein G agarose. This enriched sample was used as a benchmarking sample in the subsequent experiments.
- the final composition may preferably provide for a broader range of reactivity and functional diversity. These pools or combinations may more closely mirror the complex mixture of antibodies found in the commercial polyclonal RhD-pIgG products. Thus, combinations, or pools, of antibodies containing 3, 4 or 16 mAbs were generated (Table 8) and assessed in the agglutination assay (Table 9).
- non-RhD isotype control mAb was included and a positive ADCC/ ADP signal was defined as the signal above the background of these negative control samples.
- the background ADCC/ADP activity of the negative control mAb was subtracted from each experiment.
- the combinations, or pools, of mAbs exhibited ADCC activity against Bromelain-treated RBCs (Figure 20A) and ADP activity against both treated and untreated RBC ( Figure 20C- D) and the results was comparable to that of both commercially, enriched products, RhD- plgG-l and RhD-pIgG-2.
- a negative control pool (containing non-RhD isotype control mAbs: one IgGl mAb, one IgG2 mAb and one IgG3 mAb) was included and a positive ADCC signal was defined as a signal above the background of the negative control pool.
- the background ADCC activity of the negative control pool was subtracted from each experiment.
- Each bar represents the mean ADCC percentage with error bars indicating the standard error from the mean of two independent experiments.
- Statistical analysis was conducted to compare each antibody’s activity against RhD-pIgG-1 using one-way ANOVA followed by Dunnett’s multiplecomparison test (GraphPad Prism v9), ****: p ⁇ 0.0001.
- Table 9 The agglutination activity of mAb pools and engineered mAbs.
- Experiment 7 Investigating the minimal antibody fragments necessary for RBC binding
- RhD+ RBCs Bromelain-treated RhD+ RBCs were exposed to the anti-RhD mAb including Brad3_Full length, Brad3_F(ab’)2 and Brad3_Fab fragments. Then, the binding of antibody fragments was assessed in FACS.
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Medicinal Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- General Health & Medical Sciences (AREA)
- Immunology (AREA)
- Engineering & Computer Science (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Molecular Biology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Biophysics (AREA)
- Biochemistry (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Veterinary Medicine (AREA)
- Hematology (AREA)
- Genetics & Genomics (AREA)
- General Chemical & Material Sciences (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Pharmacology & Pharmacy (AREA)
- Animal Behavior & Ethology (AREA)
- Public Health (AREA)
- Diabetes (AREA)
- Peptides Or Proteins (AREA)
Abstract
The present invention provides for recombinant antibodies which bind to RhD blood group antigens as well as compositions comprises these recombinant antibodies and methods for selecting a combination of different recombinant antibodies derived from donor blood. The recombinant antibodies are useful for the treatment of the prevention of haemolytic disease of the newborn (HDN), treatment of idiopathic thrombocytopenic purpura (ITP), and to impede anti-RhD alloimmunisation in RhD negative patients who receive an RhD+ blood transfusion.
Description
Anti-Rhesus D human monoclonal antibodies
RELATED APPLICATION
The present application claims priority from Australian provisional patent application AU2023902692, filed August 23rd 2023, the entire contents of which is incorporated herein by reference.
Reference to Sequence Listing
The entire content of the electronic submission of the sequence listing is incorporated by reference in its entirety for all purposes.
FIELD OF INVENTION
The present invention relates generally to recombinant antibodies which bind to RhD blood group antigens. The antibodies are useful for the treatment of the prevention of haemolytic disease of the newborn (HDN), treatment of idiopathic thrombocytopenic purpura (ITP), and to impede anti-RhD alloimmunisation in RhD negative patients who receive an RhD+ blood transfusion.
BACKGROUND OF THE INVENTION
Antibodies are naturally occurring molecules produced by the B-lymphocytes of the immune system to help protect against infection and disease. Rhesus D (RhD) is a type of blood group which is found on human red blood cells. Approximately 85% of people are RhD positive (RhD+) and 15% are RhD negative (RhD-), although these frequencies vary between populations. When an RhD- mother is carrying an RhD+ infant, foetal red blood cells (RBCs) enter the maternal circulation, sensitizing the mother and provoking an antibody response to the RhD molecule. This ‘anti-D response’ increases with each pregnancy and can cause destruction of the baby’s red blood cells, known as haemolytic disease of the newborn (HDN). HDN can lead to miscarriage or stillbirth, and it has been a major cause of infant mortality throughout history, and it is still so in many parts of the world.
In the 1960s, it was suggested that anti-RhD serum (i.e., serum containing anti-RhD antibodies) might be given to RhD- women in order to rapidly remove the small population of foetal RhD+ blood cells that enter the maternal circulation, and thereby avoid sensitization of the mother’s immune system. This proved to be remarkably successful and since then the standard of care clinically has been to treat RhD- women preventatively (at 28 and 34 weeks
of pregnancy) with anti-RhD serum. As a result, HND is now rare in developed countries with an anti-RhD program in place.
Anti-RhD plasma is derived from a small pool of altruistic blood donors who have been screened for the presence of anti-RhD antibodies. These donors are then periodically boosted with RhD+ RBCs to boost the immune system and maintain high levels of anti-RhD antibodies. Some donors have donated blood over 1,000 times for this purpose, but in many parts of the world, no such service exists. Additionally, the altruistic blood donors can become unwell after repeated immunization with ‘foreign’ RBCs. Furthermore, the donation process is time consuming and expensive, and it carries a small but appreciable risk of transferring communicable diseases. For example, in the UK, donor plasma was for many years sourced from North America to minimize any risk of transmission of variant Creutzfeldt-Jakob disease.
The development of alternate, non-plasma-derived sources of anti-RhD antibodies is therefore desirable.
SUMMARY OF THE INVENTION
The present inventors have surprisingly developed an alternative source of anti-D antibodies that is plasma and donor independent. The anti-D antibodies can be used for example as a prophylactic treatment and or treatment for potential sensitising events for Rh negative women who are pregnant or recently pregnant (up to for example 10 days post pregnancy cessation).
Accordingly, in an aspect of the disclosure, there is provided an isolated or recombinant antibody or antigen binding fragment thereof capable of specifically binding to Rhesus D (RhD), wherein the antibody or antigen binding fragment thereof comprises a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in any one of SEQ ID NOs: 11, 43, 75, 107, 139, 171, 203, 235, 267, 299, 331, 363, 395, 427, 459, 491; a CDR H2 comprising a sequence as set forth in any one of SEQ ID NOs: 13, 45, 77, 109, 141, 173, 205, 237, 269, 301, 333, 365, 397, 429, 461, 493; and a CDR H3 comprising a sequence as set forth in any one of SEQ ID NOs: 15, 47, 79, 111, 143, 175, 207, 239, 271, 303, 335, 367, 399, 431, 463, 495.
In one embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 11; a CDRH2 sequence as set forth in any one of SEQ ID NO: 13; and a CDR H3 sequence as set forth in of SEQ ID NO: 15.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 43; a CDR H2 sequence as set forth in SEQ ID NO: 45; and a CDR H3 sequence as set forth in SEQ ID NO: 47.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 75; a CDR H2 sequence as set forth in SEQ ID NO: 77; and a CDR H3 sequence as set forth in SEQ ID NO: 79.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 107; a CDR H2 sequence as set forth in SEQ ID NO: 109; and a CDR H3 sequence as set forth in SEQ ID NO: 111.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 139; a CDR H2 sequence as set forth in SEQ ID NO: 141; and a CDR H3 sequence as set forth SEQ ID NO: 143.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 171; a CDR H2 sequence as set forth in SEQ ID NO: 173; and a CDR H3 sequence as set forth in SEQ ID NO: 175
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 203; a CDR H2 sequence as set forth in SEQ ID NO: 205; and a CDR H3 sequence as set forth in SEQ ID NO: 207.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 235; a CDR H2 sequence as set forth in SEQ ID NO: 237; and a CDR H3 sequence as set forth in SEQ ID NO: 239.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 267; a CDR H2 sequence as set forth in SEQ ID NO: 269; and a CDR H3 sequence as set forth in SEQ ID NO: 271.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 299; a CDR H2 sequence as set forth in SEQ ID NO: 301; and a CDR H3 as set forth in SEQ ID NO: 303.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 331; a CDR H2 sequence as set forth in SEQ ID NO: 333; and a CDR H3 sequence as set forth in SEQ ID Nos: 335.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 363; a CDR H2 sequence as set forth in SEQ ID NO: 365; and a CDR H3 sequence as set forth in SEQ ID NO: 367.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 395; a CDR H2 sequence as set forth in SEQ ID NO: 397; and a CDR H3 sequence as set forth in SEQ ID NO: 399.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 427; a CDR H2 sequence as set forth in SEQ ID NO: 429; and a CDR H3 sequence as set forth in SEQ ID NO: 431.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 459; a CDR H2 sequence as set forth in SEQ ID NO: 461; and a CDR H3 sequence as set forth in SEQ ID NO: 463.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a CDR Hl sequence as set forth in SEQ ID NO: 491; a CDR H2 sequence as set forth in SEQ ID NO: 493; and a CDR H3 sequence as set forth in SEQ ID NO: 495.
In one or a further embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a light chain variable region (VL) comprising a CDR LI sequence as set forth in any one of SEQ ID NOs: 27, 59, 91, 123, 155, 187, 219, 251, 283, 315, 347, 379, 411, 443, 475, 507; a CDR L2 sequence as set forth in any one of SEQ ID NOs: 29, 61, 93, 125, 157, 189, 221, 253, 285, 317, 349, 381, 413, 445, 477, 509; and a CDR L3 sequence as set forth in any one of SEQ ID NOs: 31, 63, 95, 127, 159, 191, 223, 255, 287, 319, 351, 383, 415, 447, 479, 511.
In one embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 27; a
CDR L2 sequence as set forth in SEQ ID NO: 29; and a CDR L3 sequence as set forth in SEQ ID NO: 31.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 59; a CDR L2 sequence as set forth in SEQ ID NO: 61; and a CDR L3 sequence as set forth in SEQ ID NO: 63.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 91; a CDR L2 sequence as set forth in SEQ ID NO: 93; and a CDR L3 sequence as set forth in SEQ ID NO: 95.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 123; a CDR L2 sequence as set forth in SEQ ID NO: 125; and a CDR L3 sequence as set forth in SEQ ID NO: 127.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 155; a CDR L2 sequence as set forth in SEQ ID NO: 157; and a CDR L3 sequence as set forth in SEQ ID NO: 159.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 187; a CDR L2 sequence as set forth in SEQ ID NO: 189; and a CDR L3 as set forth in SEQ ID NO: 191.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI as set forth in SEQ ID NO: 219; a CDR L2 sequence as set forth in SEQ ID NO: 221; and a CDR L3 sequence as set forth in SEQ ID NO: 223.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 251; a CDR L2 sequence as set forth in SEQ ID NO: 253; and a CDR L3 sequence as set forth in SEQ ID NO: 255.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 283; a CDR L2 sequence as set forth in SEQ ID NO: 285; and a CDR L3 sequence as set forth in SEQ ID NO: 287.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 315; a CDR L2 sequence as set forth in SEQ ID NO: 317; and a CDR L3 sequence as set forth in SEQ ID NO: 319.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 347; a CDR L2 sequence as set forth in SEQ ID NO: 349; and a CDR L3 sequence as set forth in SEQ ID NO: 351.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 379; a CDR L2 sequence as set forth in SEQ ID NO: 381; and a CDR L3 sequence as set forth in SEQ ID NO: 383.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 411, a CDR L2 sequence as set forth in SEQ ID NO: 413; and a CDR L3 sequence as set forth in SEQ ID NO: 415.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 443; a CDR L2 sequence as set forth in SEQ ID NO: 445; and a CDR L3 sequence as set forth in SEQ ID NO: 447.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 475; a CDR L2 sequence as set forth in SEQ ID NO: 477; and a CDR L3 sequence as set forth in SEQ ID NO: 479.
In another embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 507; a CDR L2 sequence as set forth in SEQ ID NO: 509; and a CDR L3 sequence as set forth in SEQ ID NO: 511.
In one or a further embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VH comprising an amino acid sequence having at least 70% sequence identity to the sequence set forth in any one of SEQ ID NOs: 9, 41, 73, 105, 137, 169, 201, 233, 265, 297, 329, 361, 393, 425, 457, 489.
In one or a further embodiment of the disclosure, the antibody or antigen binding fragment of the disclosure comprises a VL comprising an amino acid sequence having at least 70%
sequence identity to the sequence set forth in any one of SEQ ID NOs: 25, 57, 89, 121, 153, 185, 217, 249, 281, 313, 345, 377, 409, 441, 473, 505.
In another aspect, the disclosure provides an antibody or antigen binding fragment thereof capable of specifically binding to Rhesus D (RhD), wherein the antibody or antigen binding fragment thereof comprises: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 11; CDR H2 comprising a sequence as set forth in SEQ ID NO: 13; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 15; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 27; CDR L2 comprising a sequence as set forth in SEQ ID NO: 29; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 31; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 43; CDR H2 comprising a sequence as set forth in SEQ ID NO: 45; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 47; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 59, CDR L2 comprising a sequence as set forth in SEQ ID NO: 61, and CDR L3 comprising a sequence as set forth in SEQ ID NO: 63; c) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 75; CDR H2 comprising a sequence as set forth in SEQ ID NO: 77; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 79; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 91; CDR L2 comprising a sequence as set forth in SEQ ID NO: 93; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 95; d) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 107; CDR H2 comprising a sequence as set forth in SEQ ID NO: 109; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 111 ; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 123; CDR L2 comprising a sequence as set forth in SEQ ID NO: 125; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 127; e) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 139; CDR H2 comprising a sequence as set forth in SEQ ID NO: 141; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 143; and a light chain variable region (VL) comprising a CDR1 comprising a sequence as set forth in SEQ ID NO: 155, CDR L2 comprising a sequence as set forth in SEQ ID NO: 157; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 159; f) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 171; CDR H2 comprising a sequence as set forth in SEQ ID NO: 173; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 175; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 187; CDR L2 comprising a sequence as set forth in SEQ ID NO: 189; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 191;
g) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 203; CDR H2 comprising a sequence as set forth in SEQ ID NO: 205; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 207; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 219; CDR L2 comprising a sequence as set forth in SEQ ID NO: 221; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 223; h) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 235; CDR2 comprising a sequence as set forth in SEQ ID NO: 237; and CDR3 comprising a sequence as set forth in SEQ ID NO: 239; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 251; CDR L2 comprising a sequence as set forth in SEQ ID NO: 253; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 255; i) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 267; CDR H2 comprising a sequence as set forth in SEQ ID NO: 269; and CDR3 comprising a sequence as set forth in SEQ ID NO: 271; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 283; CDR L2 comprising a sequence as set forth in SEQ ID NO: 285; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 287; j) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 299; CDR H2 comprising a sequence as set forth in SEQ ID NO: 301; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 303; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 315; CDR L2 comprising a sequence as set forth in SEQ ID NO: 317; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 319; k) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 331; CDR H2 comprising a sequence as set forth in SEQ ID NO: 333; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 335; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 347; CDR L2 comprising a sequence as set forth in SEQ ID NO: 349; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 351; l) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 3631; CDR H2 comprising a sequence as set forth in SEQ ID NO: 365; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 367; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 379; CDR L2 comprising a sequence as set forth in SEQ ID NO: 381; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 383; m)a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 395; CDR H2 comprising a sequence as set forth in SEQ ID NO: 397; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 399; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 411; CDR L2 comprising a sequence as set forth in SEQ ID NO: 413; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 415;
n) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 427; CDR H2 comprising a sequence as set forth in SEQ ID NO: 429, and CDR H3 comprising a sequence as set forth in SEQ ID NO: 431 ; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 443; CDR L2 comprising a sequence as set forth in SEQ ID NO: 445; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 447; o) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 459; CDR H2 comprising a sequence as set forth in SEQ ID NO: 461; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 463; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 475; CDR L2 comprising a sequence as set forth in SEQ ID NO: 477; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 479; or p) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 491; CDR H2 comprising a sequence as set forth in SEQ ID NO: 493; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 495; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 507; CDR L2 comprising a sequence as set forth in SEQ ID NO: 509; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 511.
In another aspect, the disclosure provides an antibody or antigen binding fragment thereof capable of specifically binding to Rhesus D (RhD), wherein the antibody or antigen binding fragment thereof comprises: a) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO:
9; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 25; b) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 41; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 57; c) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 89; d) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 105; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 121; e) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 137; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 153;
f) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 185; g) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 201; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 217; h) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 233; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 249; i) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 265; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 281; j) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 297; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 313; k) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 329; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 345; l) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 361; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 377; m)a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 393; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 409; n) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 425; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 441; o) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 457; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 473; or p) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 489; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 505.
In some embodiments of any of the previous aspects of the disclosure, the antibody or antigen binding fragment is capable of specifically binding to RhD positive (RhD+), but not RhD negative (RhD-), RBCs. Advantageously, the antibody or antigen binding fragment is capable of agglutinating RhD+ red blood cells but not RhD- red blood cells.
In some or further embodiments of any of the previous aspects of the disclosure, the antibody or antigen binding fragment is one of IgG subclass 1, IgG subclass 2, or IgG subclass 3.
In some or further embodiments of any of the previous aspects of the disclosure, the antibody or antigen binding fragment is IgG subclass 1, IgG or IgG subclass 3.
In some or further embodiments of any of the previous aspects of the disclosure, the antibody or antigen binding fragment is IgG subclass 1.
In some or further embodiments of any of the previous aspects of the disclosure, the antibody or antigen binding fragment comprise a Fc region or part thereof.
Advantageously, one or more of the antibodies or antigen binding fragments disclosed in the present application can be used in combination to provide a composition. Accordingly, the present disclosure provides a composition comprising one or more antibodies or antigen binding fragments as defined herein. In some embodiments, one or more of the antibodies or antigen binding fragments disclosed herein, can be used in combination with one more additional antibodies or antigen binding fragments capable of specifically binding to Rhesus D (RhD) that may optionally be identified using a method disclosed herein.
Accordingly, in another aspect, the present disclosure provides a composition comprising one or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein one or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 11; CDR H2 comprising a sequence as set forth in SEQ ID NO: 13; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 15; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 27; CDR L2 comprising a sequence as set forth in SEQ ID NO: 29; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 31; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 43; CDR H2 comprising a sequence as set forth in SEQ ID NO: 45; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 47; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 59, CDR L2 comprising a sequence as set forth in SEQ ID NO: 61, and CDR L3 comprising a sequence as set forth in SEQ ID NO: 63; c) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 75; CDR H2 comprising
a sequence as set forth in SEQ ID NO: 77; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 79; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 91; CDR L2 comprising a sequence as set forth in SEQ ID NO: 93; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 95; d) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 107; CDR H2 comprising a sequence as set forth in SEQ ID NO: 109; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 111 ; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 123; CDR L2 comprising a sequence as set forth in SEQ ID NO: 125; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 127; e) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 139; CDR H2 comprising a sequence as set forth in SEQ ID NO: 141; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 143; and a light chain variable region (VL) comprising a CDR1 comprising a sequence as set forth in SEQ ID NO: 155, CDR L2 comprising a sequence as set forth in SEQ ID NO: 157; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 159; f) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 171; CDR H2 comprising a sequence as set forth in SEQ ID NO: 173; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 175; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 187; CDR L2 comprising a sequence as set forth in SEQ ID NO: 189; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 191; g) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 203; CDR H2 comprising a sequence as set forth in SEQ ID NO: 205; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 207; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 219; CDR L2 comprising a sequence as set forth in SEQ ID NO: 221; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 223; h) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 235; CDR2 comprising a sequence as set forth in SEQ ID NO: 237; and CDR3 comprising a sequence as set forth in SEQ ID NO: 239; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 251; CDR L2 comprising a sequence as set forth in SEQ ID NO: 253; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 255; i) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 267; CDR H2 comprising a sequence as set forth in SEQ ID NO: 269; and CDR3 comprising a sequence as set forth in SEQ ID NO: 271; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 283; CDR L2 comprising a sequence as set forth in SEQ ID NO: 285; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 287; j) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 299; CDR H2 comprising
a sequence as set forth in SEQ ID NO: 301; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 303; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 315; CDR L2 comprising a sequence as set forth in SEQ ID NO: 317; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 319; k) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 331; CDR H2 comprising a sequence as set forth in SEQ ID NO: 333; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 335; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 347; CDR L2 comprising a sequence as set forth in SEQ ID NO: 349; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 351; l) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 363; CDRH2 comprising a sequence as set forth in SEQ ID NO: 365; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 367; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 379; CDR L2 comprising a sequence as set forth in SEQ ID NO: 381; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 383; m)a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 395; CDR H2 comprising a sequence as set forth in SEQ ID NO: 397; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 399; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 411; CDR L2 comprising a sequence as set forth in SEQ ID NO: 413; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 415; n) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 427; CDR H2 comprising a sequence as set forth in SEQ ID NO: 429, and CDR H3 comprising a sequence as set forth in SEQ ID NO: 431 ; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 443; CDR L2 comprising a sequence as set forth in SEQ ID NO: 445; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 447; o) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 459; CDR H2 comprising a sequence as set forth in SEQ ID NO: 461; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 463; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 475; CDR L2 comprising a sequence as set forth in SEQ ID NO: 477; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 479; or p) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 491; CDR H2 comprising a sequence as set forth in SEQ ID NO: 493; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 495; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 507; CDR L2 comprising a sequence as set forth in SEQ ID NO: 509; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 511.
In another aspect, the disclosure provides a composition comprising one or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD),
wherein one or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO:9; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 25; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 41; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 57; c) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 89; d) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 105; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 121; e) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 137; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 153; f) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 185; g) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 201; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 217; h) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 233; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 249; i) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 265; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 281; j) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 297; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 313; k) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 329; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 345; l) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 361; and a light
chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 377; m)a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 393; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 409; n) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 425; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 441; o) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 457; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 473; or p) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 489; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 505.
In another aspect, the disclosure provides a composition comprising one or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein the one or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 75; CDR H2 comprising a sequence as set forth in SEQ ID NO: 77; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 79; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 91; CDR L2 comprising a sequence as set forth in SEQ ID NO: 93; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 95; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 171; CDR H2 comprising a sequence as set forth in SEQ ID NO: 173; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 175; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 187; CDR L2 comprising a sequence as set forth in SEQ ID NO: 189; and CDRL3 comprising a sequence as set forth in SEQ ID NO: 191; c) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 267; CDR H2 comprising a sequence as set forth in SEQ ID NO: 269; and CDR3 comprising a sequence as set forth in SEQ ID NO: 271; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 283; CDR L2 comprising a sequence as set forth in SEQ ID NO: 285; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 287; d) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO:299; CDR H2 comprising a sequence as set forth in SEQ ID NO: 301; and CDR3 comprising a sequence as set
forth in SEQ ID NO: 303; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 315; CDR L2 comprising a sequence as set forth in SEQ ID NO: 317; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 319; e) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 331; CDR H2 comprising a sequence as set forth in SEQ ID NO: 333; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 335; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 347; CDR L2 comprising a sequence as set forth in SEQ ID NO: 349; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 351; f) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 395; CDR H2 comprising a sequence as set forth in SEQ ID NO: 397; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 399; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 411; CDR L2 comprising a sequence as set forth in SEQ ID NO: 413; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 415; or g) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 427; CDR H2 comprising a sequence as set forth in SEQ ID NO: 429, and CDR H3 comprising a sequence as set forth in SEQ ID NO: 431 ; anda light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 443; CDR L2 comprising a sequence as set forth in SEQ ID NO: 445; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 447.
In one embodiment of the disclosure, the composition further comprises two or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD). For example, the two or more antibodies or antigen binding fragments may be selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 75; CDR H2 comprising a sequence as set forth in SEQ ID NO: 77; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 79; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 91; CDR L2 comprising a sequence as set forth in SEQ ID NO: 93; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 95; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 171; CDR H2 comprising a sequence as set forth in SEQ ID NO: 173; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 175; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 187; CDR L2 comprising a sequence as set forth in SEQ ID NO: 189; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 191;
c) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 267; CDR H2 comprising a sequence as set forth in SEQ ID NO: 269; and CDR3 comprising a sequence as set forth in SEQ ID NO: 271; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 283; CDR L2 comprising a sequence as set forth in SEQ ID NO: 285; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 287; d) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO:299; CDR H2 comprising a sequence as set forth in SEQ ID NO: 301; and CDR3 comprising a sequence as set forth in SEQ ID NO: 303; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 315; CDR L2 comprising a sequence as set forth in SEQ ID NO: 317; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 319; e) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 331; CDR H2 comprising a sequence as set forth in SEQ ID NO: 333; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 335; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 347; CDR L2 comprising a sequence as set forth in SEQ ID NO: 349; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 351; f) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 395; CDR H2 comprising a sequence as set forth in SEQ ID NO: 397; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 399; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 411; CDR L2 comprising a sequence as set forth in SEQ ID NO: 413; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 415; or g) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 427; CDR H2 comprising a sequence as set forth in SEQ ID NO: 429, and CDR H3 comprising a sequence as set forth in SEQ ID NO: 431; anda light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 443; CDR L2 comprising a sequence as set forth in SEQ ID NO: 445; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 447.
In one embodiment of the disclosure, the composition further comprises three or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD). For example, the one or more antibodies or antigen binding fragments may be selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 75; CDR H2 comprising a sequence as set forth in SEQ ID NO: 77; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 79; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 91; CDR
L2 comprising a sequence as set forth in SEQ ID NO: 93; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 95; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 171; CDR H2 comprising a sequence as set forth in SEQ ID NO: 173; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 175; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 187; CDR L2 comprising a sequence as set forth in SEQ ID NO: 189; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 191; c) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 267; CDR H2 comprising a sequence as set forth in SEQ ID NO: 269; and CDR3 comprising a sequence as set forth in SEQ ID NO: 271; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 283; CDR L2 comprising a sequence as set forth in SEQ ID NO: 285; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 287; d) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO:299; CDR H2 comprising a sequence as set forth in SEQ ID NO: 301; and CDR3 comprising a sequence as set forth in SEQ ID NO: 303; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 315; CDR L2 comprising a sequence as set forth in SEQ ID NO: 317; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 319; e) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 331; CDR H2 comprising a sequence as set forth in SEQ ID NO: 333; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 335; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 347; CDR L2 comprising a sequence as set forth in SEQ ID NO: 349; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 351; f) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 395; CDR H2 comprising a sequence as set forth in SEQ ID NO: 397; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 399; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 411; CDR L2 comprising a sequence as set forth in SEQ ID NO: 413; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 415; or g) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 427; CDR H2 comprising a sequence as set forth in SEQ ID NO: 429, and CDR H3 comprising a sequence as set forth in SEQ ID NO: 431; anda light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 443; CDR L2 comprising a sequence as set forth in SEQ ID NO: 445; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 447.
In another aspect, the disclosure provides a composition comprising one or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein the one or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 89; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 185; c) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 265; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 281; d) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 297; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 313; e) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 329; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 345; f) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 393; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 409; or g) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 425; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 441.
In another aspect, the disclosure provides a composition comprising a combination of two or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein the two or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 89; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 185;
c) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 265; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 281; d) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 297; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 313; e) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 329; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 345; f) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 393; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 409; or g) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 425; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 441
In another aspect, the disclosure provides a composition comprising a combination of three or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein the three or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 89; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 185; c) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 265; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 281; d) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 297; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 313; e) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 329; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 345; f) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 393; and a light
chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 409; or g) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 425; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 441
In one embodiment of the disclosure, the composition comprises a combination of three or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD). For example, the three or more antibodies or antigen binding fragments may be selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 89; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 137; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 153; c) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 185; d) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 233; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 249; e) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 265; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 281 f) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 329; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 345; g) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 393; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 409; h) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO425; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 444; or i) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO:489; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 505;
In some embodiments of the disclosure, the composition comprises a combination of three or more of the recombinant antibodies or antigen binding fragments selected to maximise either ADP activity or agglutination activity or both ADP activity and agglutination activity of the combination, compared to the ADP or agglutination of the individual antibodies or antigen binding fragments.
In some embodiments of the disclosure, the composition of any of the previous aspects further comprises a pharmaceutically acceptable carrier.
In another aspect, the disclosure provides a nucleic acid molecule(s) encoding the antibody or antibody or antigen binding fragment according to the previous aspects. In another aspect, the disclosure provides a vector(s) comprising the nucleic acid molecule(s) of the disclosure. In some embodiments of the disclosure, the vector(s) is an expression vector. In another aspect, the disclosure provides a host cell comprising the vector(s) of the disclosure. In some embodiments of the disclosure, the host cell is a prokaryote or eukaryote. In some embodiments of disclosure, the host cell is selected from an Escherichia coli cell, a yeast cell, a mammalian cell, and other cells applicable to the preparation of an antibody or antigen binding fragment thereof. In some embodiments of the disclosure, the mammalian cell is a CHO cell, HEK293 cell or COS cell.
In another aspect, disclosure provides a production method of an antibody or antigen binding fragment thereof, wherein the host cell of the disclosure is cultured under conditions suitable for expression of a nucleic acid encoding the antibody or antigen binding fragment thereof of the disclosure, optionally wherein the antibody or antigen binding fragment thereof is separated, and optionally wherein the produced antibody or antigen-binding fragments thereof are collected.
In another aspect, the disclosure provides a method of treating or preventing Rhesus alloimmunization of Rh-negative subjects, wherein the method comprises administering to a Rh-negative subject in need thereof an effective amount of an anti-Rd(D) antibody or antigen binding fragment of the disclosure, or a combination thereof; or the composition of the disclosure.
In some embodiments of the disclosure, the subject is pregnant.
In some embodiments of the disclosure, the method prevents haemolytic disease of the foetus and newborn.
In another aspect, the disclosure provides for use of the antibody or antigen binding fragment of the disclosure, or a combination thereof, in the manufacture of a medicament for the treatment or prevention of haemolytic disease of the foetus and newborn.
In another aspect, the disclosure provides for use of the antibody or antigen binding fragment of the disclosure, or a composition thereof, or a composition of the disclosure, for the treatment or prevention of Rhesus alloimmunization of Rh-negative subjects.
In another aspect, the disclosure provides a method for profiling the most abundant antibodies that bind to the antigen (RhD) in a polyclonal (pAb) population, wherein the method comprises: (a) proteomic analysis of a RhD-specific IgG mixture from plasma, wherein plasma is provided from a donor; affinity purification of the total IgG mixture from the plasma; separation of the RhD-specific and non-RhD specific IgG subsets by incubation with either RhD+ or RhD- RBCs, respectively; independent fragmenting of the RhD-specific and non-RhD specific IgG subsets into short peptides of overlapping sequences using one or more proteases; analysis of the fragmented peptides by mass spectrometry to identify the sequence of each peptide; assembly of the short peptide sequences into longer peptide sequences that belong to individual antibodies, optionally using computational software; and excluding antibodies derived from the non-RhD specific subset from further analysis; and (b) BCR sequencing, wherein the B cells provided are isolated from blood of the donor; extraction of RNA from the B cells; sequencing of the RNAs by Next Generation Sequencing (NGS) to provide transcriptomic sequence information; and combining the transcriptomics sequence information with the proteomic sequence information to further assemble and derive the complete amino acid sequences of the most abundant mAbs in the RhD specific IgG subset. In some preferred embodiments the method for profiling further comprises identification of most the abundant mAbs that were specifically enriched in both the LC and HC regions by ranking the sequences based on quantified fold changes of enrichment from the RhD specific IgG subset.
In another aspect of the disclosure provides for a method for profiling the most abundant antibodies that bind to the antigen (RhD) in a polyclonal (pAb) population, the method comprising: a) proteomic analysis of a RhD-specific IgG mixture from plasma to provide proteomic sequence information: b) BCR sequencing: c) combining the transcriptomics sequence information with the proteomic sequence information to further assemble and derive the complete amino acid sequences of the most abundant mAbs in the RhD specific IgG subset: d) further selecting from the most abundant mAbs a combination of mAbs each of which comprises a different genetic and/or amino acid sequence and each mAb containing a IgGl subclass heavy chain variable region (VH).
In some embodiments of the disclosure, the blood is collected from RhD antigen donors. In some embodiments of the disclosure, the donor plasma is separated from PBMCs. In further embodiments of the disclosure, CD 19+ IgG+ B cells are separated from donor PBMCs.
In some embodiments of the disclosure, one or more of the amino acid sequences identified can be used to make recombinant antibodies or antigen binding fragments that can be used in combination with one or more of the antibodies or antigen binding fragments disclosed herein.
BRIEF DESCRIPTION OF FIGURES
FIGURE l is a schematic showing the discovery of anti-RhD antibodies using a combination of Next Generation Sequencing (NGS) and proteomics.
FIGURE 2 is a schematic showing the cloning of antibody variable genes and cloning into the expression vectors. The specificity of purified antibodies is evaluated in agglutination assays against both RhD+ and RhD- RBCs.
FIGURE 3 is a schematic of the RhD antigen in the erythrocyte membrane.
FIGURE 4 shows agglutination results (with bromelain treated or untreated RBCs).
FIGURE 5 shows anti-RhD mAbs competition binding profiles. A) Competition RBC binding assay. The potency of each mAb is assessed based on its ability to inhibit RhD binding of other anti-RhD mAbs. The number of inhibited mAbs refers to the count of antibodies that their RhD binding is inhibited by at least 70% (B) or 50% (C) using the tested unconjugated mAb.
FIGURE 6 shows binding of anti-RhD mAbs to different Epitope RhD variants (A). Potential epitopes of anti-RhD mAbs according to the availability of epitopes in tested RhD variants (B).
FIGURE 7 shows the inhibitory activity of RhD-pIgG-1 on binding of anti-RhD mAbs from FACS analysis. A) The binding of each individual mAb to RhD+ RBCs in the absence and presence of other IgG products. B) Bar graph illustrates the percentage of binding inhibition mediated by other IgG products based on the reduction in the AUC of mAb binding in the absence of alternative IgG products.
FIGURE 8 shows anti-RhD mAbs induced RBC clearance by THP-1 monocyte. A) The gating strategy used for analysis of in vitro phagocytosis assay in the presence of non-RhD mAb (Top panel) or anti-RhD mAb (Bottom panel). B) Fey receptors on THP-1 monocytes, C) phagocytosis of anti-RhD mAbs:. F) Bar graphs depict the area under the curve (AUC) of phagocytosis scores across varying mAbs concentrations. E) subclass switching of clone 21a (from IgG2 to IgGl) and clone 104 (from IgG3 to IgGl).
FIGURE 9 shows anti-RhD mAbs induced RBC clearance by ADCC. A) Schematic of ADCC assay. B) ADCC of anti-RhD mAbs. C) subclass switching of clone 21a (from IgG2 to IgGl) and clone 104 (from IgG3 to IgGl)
FIGURE 10 shows activation of NK cells by anti-RhD mAbs. A) Detection of secreted cytokines by NK cells in the presence of anti-RhD antibodies using Bioplex kit. B) Assessment of NK cell activation markers using FACS in the presence of anti-RhD.
FIGURE 11 shows the germline usage in anti-RhD heavy chains.
FIGURE 12 shows the germline usage in anti-RhD light chains.
FIGURE 13 is an image of the Alphafold modelling the interaction between anti-RhD mAbs with RhD antigen. The RhD antigen is depicted as a coiled structure (bottom molecule in each figure). The antibody heavy and light chains are represented as ribbon structures (top molecules in each figure).
FIGURE 14 shows the prediction of the epitope interaction with the RhD antigen. A) The model of RhD antigen shown in dark grey with the epitopes of anti-RhD mAbs highlighted in pale grey, B) The predicted epitopes of anti-RhD mAbs on RhD antigen are shown in circles. FIGURE 15 shows a prediction of target sites of anti-RhD mAbs on extracellular RhD loops. The predicted binding sites by Alphafold is shown in rectangle for each mAb. Four mAbs interact with ASP350 (shown by black oval).
FIGURE 16 shows RBC binding of modified anti-RhD mAbs.
FIGURE 17 shows NK cell activation and RBC clearance through ADCC with FC modified anti-RhD mAbs. A) Gating strategy for evaluation of NK cell activation. RhD+ RBC was incubated with mAbs. B) The bar graphs show percentage of CD107+ cells, fFNy+ or total activated NK cells. C) ADCC assay results from experiments using monocyte-depleted PBMCs in the presence RhD+ RBCs opsonised with anti-RhD antibody.
FIGURE 18 shows ADP activity results of Fc modified anti-RhD mAbs. A) ADP activity (ADP score) of antibody variants at various concentrations. B) Area under curve (AUC).
FIGURE 19 shows clearance of RhD+ RBCs mediated by anti-RhD antibody variants. ADCC activity of antibodies (Img/mL) against Bromelain-treated (A) or untreated (B) RhD+ RBCs is shown in the top panel. ADP activity of antibodies (lOOng/mL) against Bromelain-treated (C) or untreated (D) RhD+ RBCs is also shown in the bottom panel.
FIGURE 20 shows clearance of RhD+ RBCs mediated by anti-RhD antibody pools. ADCC activity of antibodies (Img/mL) against Bromelain-treated (A) or untreated (B) RhD+ RBCs is shown in the top panel. ADP activity of antibody pools (lOOng/mL) against Bromelain- treated (C) or untreated (D) RhD+ RBCs is shown in the bottom panel.
FIGURE 21 shows RBC binding of anti-RhD antibody fragments.
KEY TO THE SEQUENCE LISTING
DETAILED DESCRIPTION OF THE INVENTION
Before describing the present invention in detail, it is to be understood that the terminology used herein is for the purpose of describing particular embodiments only and is not intended to be limiting.
A reference herein to a patent document or other matter which is given as prior art is not to be taken as admission that the document or matter was known or that the information it contains was part of the common general knowledge as at the priority date of any of the claims.
General Techniques and Definitions
Throughout this specification, unless specifically stated otherwise or the context requires otherwise, reference to a single step, composition of matter, group of steps or group of compositions of matter shall be taken to encompass one and a plurality (i.e. one or more) of those steps, compositions of matter, groups of steps or groups of compositions of matter.
Those skilled in the art will appreciate that the present disclosure is susceptible to variations and modifications other than those specifically described. It is to be understood that the disclosure includes all such variations and modifications. The disclosure also includes all of the steps, features, compositions and compounds referred to or indicated in this specification, individually or collectively, and any and all combinations or any two or more of said steps or features.
The present disclosure is not to be limited in scope by the specific examples described herein, which are intended for the purpose of exemplification only. Functionally equivalent products, compositions and methods are clearly within the scope of the present disclosure.
Any example of the present disclosure herein shall be taken to apply mutatis mutandis to any other example of the disclosure unless specifically stated otherwise. Stated another way, any specific example of the present disclosure may be combined with any other specific example of the disclosure (except where mutually exclusive).
Any example of the present disclosure disclosing a specific feature or group of features or method, or method steps will be taken to provide explicit support for disclaiming the specific feature or group of features or method or method steps.
Unless specifically defined otherwise, all technical and scientific terms used herein shall be taken to have the same meaning as commonly understood by one of ordinary skill in the art. In the present invention are standard procedures, well known to those skilled in the art. Such techniques are described and explained throughout the literature in sources such as, J. Perbal, A Practical Guide to Molecular Cloning, John Wiley and Sons (1984); J. Sambrook
et al., Molecular Cloning: A Laboratory Manual, Cold Spring Harbour Laboratory Press (1989); T. A. Brown (editor), Essential Molecular Biology: A Practical Approach, Volumes 1 and 2, IRL Press (1991); D.M. Glover and B.D. Hames (editors), DNA Cloning: A Practical Approach, Volumes 1-4, IRL Press (1995 and 1996); F.M. Ausubel et al., (editors), Current Protocols in Molecular Biology, Greene Pub. Associates and Wiley-Interscience (1988, including all updates until present); Ed Harlow and David Lane (editors) Antibodies: A Laboratory Manual, Cold Spring Harbour Laboratory, (1988); and J.E. Coligan et al., (editors) Current Protocols in Immunology, John Wiley & Sons (including all updates until present).
The description and definitions of variable regions and parts thereof, antibodies and fragments thereof herein may be further clarified by the discussion in Kabat Sequences of Proteins of Immunological Interest, National Institutes of Health, Bethesda, Md. (1987 and 1991), Bork et al., J Mol. Biol. (1994) 242:309-320, 1994; Chothia and Lesk, J. Mol. Biol. (1987) 196:901-917; Chothia et al. Nature (1989) 342:877-883, and Al-Lazikani et al., J. Mol. Biol. (1997) 273:927-948.
The term "and/or", e.g., "X and/or Y" shall be understood to mean either "X and Y" or "X or Y" and shall be taken to provide explicit support for both meanings or for either meaning.
As used herein, the singular forms “a”, “an” and “the” include plural referents unless the content clearly dictates otherwise. Thus, for example, reference to an antibody or antigen binding fragment, includes a combination of two or more such molecules.
Throughout this specification the word "comprise", or variations such as "comprises" or "comprising", will be understood to imply the inclusion of a stated element, integer or step, or group of elements, integers or steps, but not the exclusion of any other element, integer or step, or group of elements, integers or steps.
As used herein the term "derived from" shall be taken to indicate that a specified integer may be obtained from a particular source albeit not necessarily directly from that source.
As used herein, the term "specifically binds" shall be taken to mean that the binding interaction between the binding region on the antibody or binding fragment thereof is dependent on the presence of the antigenic determinant or epitope. The binding region preferentially binds or recognizes a specific antigenic determinant or epitope even when present in a mixture of other molecules or organisms. In one example, the binding region reacts or associates more frequently, more rapidly, with greater duration and/or with greater affinity with the specific component or cell expressing same than it does with alternative
antigens or cells. It is also understood by reading this definition that, for example, a binding region that specifically binds to a particular component may or may not specifically bind to a second antigen. As such, "specific binding" does not necessarily require exclusive binding or non-detectable binding of another antigen. The term "specifically binds" can be used interchangeably with "selectively binds" herein. Generally, reference herein to binding means specific binding, and each term shall be understood to provide explicit support for the other term. Methods for determining specific binding will be apparent to the skilled person. For example, a binding protein comprising the binding region of the disclosure is contacted with the component or a cell expressing same or a mutant form thereof or an alternative antigen. The binding to the component or mutant form or alternative antigen is then determined and a binding region that binds as set out above is considered to specifically bind to the component.
The term "recombinant" shall be understood to mean the product of artificial genetic recombination. Accordingly, in the context of an antibody or antigen binding fragment thereof, this term does not encompass an antibody naturally occurring within a subject's body that is the product of natural recombination that occurs during B cell maturation. However, if such an antibody is isolated, it is to be considered an isolated protein comprising an antibody variable region. Similarly, if nucleic acid encoding the protein is isolated and expressed using recombinant means, the resulting protein is a recombinant protein. A recombinant protein also encompasses a protein expressed by artificial recombinant means when it is within a cell, tissue or subject, for example, in which it is expressed.
The term "polypeptide" or "polypeptide chain" will be understood to mean a series of contiguous amino acids linked by peptide bonds.
The skilled person will be aware that an "antibody" is generally considered to be a protein that comprises a variable region made up of a plurality of polypeptide chains, for example, a polypeptide comprising a light chain variable region (VL) and a polypeptide comprising a heavy chain variable region (VH). An antibody also generally comprises constant domains, some of which can be arranged into a constant region, which includes a constant fragment or fragment crystallizable (Fc), in the case of a heavy chain. A VH and a VL interact to form an Fv comprising an antigen binding region that is capable of specifically binding to one or a few closely related antigens. Generally, a light chain from mammals is either a K light chain or a light chain and a heavy chain from mammals is a a, 5, a, y, or p. Antibodies can be of any class (e.g., IgG, IgE, IgM, IgD, IgA, and IgY), or subclass (e.g., IgGl, IgG2, IgG3, IgG4, IgAl and IgA2) or subclass. The term "antibody" in the context of this invention encompasses human antibodies. The term "antibody" also includes variants, for
example, variants missing an encoded C-terminal lysine residue, a deamidated variant and/or a glycosylated variant and/or a variant comprising a pyroglutamate, for example, at the N- terminus and/or a variant lacking a N-terminal residue, for example, a N-terminal glutamine in an antibody or V region and/or a variant comprising all or part of a secretion signal. Deamidated variants of encoded asparagine residues may result in isoaspartic, and aspartic acid isoforms being generated or even a succinamide involving an adjacent amino acid residue. Deamidated variants of encoded glutamine residues may result in glutamic acid. Compositions comprising a heterogeneous mixture of such sequences and variants are intended to be included when reference is made to a particular amino acid sequence.
The terms "full-length antibody", "intact antibody" or "whole antibody" are used interchangeably to refer to an antibody in its substantially intact form, as opposed to an antigen binding fragment of an antibody. Specifically, whole antibodies include those with heavy and light chains including an Fc region. The constant domains may be wild-type sequence constant domains (e.g., human wild-type sequence constant domains) or amino acid sequence variants thereof).
The term "monoclonal antibody" (mAb) as used herein refers to an antibody obtained from a population of substantially homogeneous antibodies, or to said population of antibodies. The individual antibodies comprising the population are essentially identical, except for possible naturally occurring mutations that may be present in minor amounts. Monoclonal antibodies are highly specific, being directed against a single antigenic site. The present disclosure provides for recombinant DNA expression of monoclonal antibodies. Two or more recombinant monoclonal antibodies (e.g., directed against different antigenic sites) can be pooled and used in methods of the disclosure. Specific combinations or pools can be chosen to provide a diverse range of recombinant antibodies each of which may bind to one or more different Rhesus D antigen epitopes,
“Human antibodies” as used herein includes antibodies having the amino acid sequence of a human immunoglobulin and antibodies isolated from human immunoglobulin libraries.
“Chimeric antibodies” as used herein refers to structural chimeras with, for example, variable sequences derived from one species, and constant regions from another species or another isotype within the same species. As used herein, “variable region" refers to the portions of the light and/or heavy chains of an antibody as defined herein that is capable of specifically binding to an antigen and, for example, includes amino acid sequences of complementarity determining regions (CDRs); i.e., CDR1, CDR2, and CDR3, and framework
regions (FRs). For example, the variable region comprises three or four FRs (e.g., FRI, FR2, FR3 and optionally FR4) together with three CDRs. VH refers to the variable region of the heavy chain. VL refers to the variable region of the light chain.
As used herein, the term "complementarity determining regions” (syn. CDRs; i.e., CDR1, CDR2, and CDR3) refers to the amino acid residues of an antibody variable region the presence of which contribute to specific antigen binding. Each variable region typically has three CDR regions identified as CDR1, CDR2 and CDR3. In one example, the amino acid positions assigned to CDRs and FRs are defined according to Kabat Sequences of Proteins of Immunological Interest, National Institutes of Health, Bethesda, Md., 1987 and 1991. For example, according to the numbering system of Kabat, VH FRs and CDRs are positioned as follows: residues 1-30 (FRI), 31-35 (CDR1), 36-49 (FR2), 50-65 (CDR2), 66-94 (FR3), 95- 102 (CDR3) and 103- 113 (FR4). According to the numbering system of Kabat, VL FRs and CDRs are positioned as follows: residues 1-23 (FRI), 24-34 (CDR1), 35-49 (FR2), 50-56 (CDR2), 57-88 (FR3), 89-97 (CDR3) and 98-107 (FR4). The present disclosure is not limited to FRs and CDRs as defined by the Kabat numbering system, but includes all numbering systems, including those discussed above. In one embodiment, reference herein to a CDR (or a FR) is in respect of those regions according to the Kabat numbering system.
"Framework regions" (FRs) are those variable region residues other than the CDR residues.
As used herein, the term “Fv” shall be taken to mean any protein, whether comprised of multiple polypeptides or a single polypeptide, in which a VH and a VL associate and form a complex having an antigen binding site, i.e., capable of specifically binding to an antigen. The VH and the VL which form the antigen binding site can be in a single polypeptide chain or in different polypeptide chains. Furthermore, an Fv of the disclosure (as well as any protein of the disclosure) may have multiple antigen binding sites which may or may not bind the same antigen. This term shall be understood to encompass fragments directly derived from an antibody as well as proteins corresponding to such a fragment produced using recombinant means. In some embodiments, the VH is not linked to a heavy chain constant domain (CH), for example CH 1 and/or the VL is not linked to a light chain constant domain (CL). Exemplary Fv containing polypeptides or proteins include a Fab fragment, a Fab’ fragment, a F(ab’) fragment, a scFv, a diabody, a triabody, a tetrabody or higher order complex, or any of the foregoing linked to a constant region or domain thereof, for example, one or both of a CH2or CH3 domain, for example, a minibody.
A "Fab fragment" consists of a monovalent antigen-binding fragment of an immunoglobulin and can be produced by digestion of a whole antibody with the enzyme papain, to yield a fragment consisting of an intact light chain and a portion of a heavy chain or can be produced using recombinant means.
A "Fab1 fragment" of an antibody can be obtained by treating a whole antibody with pepsin, followed by reduction, to yield a molecule consisting of an intact light chain and a portion of a heavy chain comprising a VH and a single CH. Two Fab' fragments are obtained per antibody treated in this manner. A Fab’ fragment can also be produced by recombinant means.
A "F(ab')2 fragment” consists of a dimer of two Fab' fragments held together by two disulfide bonds and is obtained by treating a whole antibody molecule with the enzyme pepsin, without subsequent reduction. A “Fab2” fragment is a recombinant fragment comprising two Fab fragments linked using, for example a leucine zipper or a CH3 domain.
A “single chain Fv” or “scFv” is a recombinant molecule containing the variable region fragment (Fv) of an antibody in which the variable region of the light chain and the variable region of the heavy chain are covalently linked by a suitable, flexible polypeptide linker.
The term "constant region" as used herein, refers to a portion of heavy chain or light chain of an antibody other than the variable region. In a heavy chain, the constant region generally comprises a plurality of constant domains and a hinge region, for example, an IgG constant region comprises the following linked components, a CHI, a hinge, a CH2 and a CH3. In a heavy chain, a constant region comprises a Fc. In a light chain, a constant region generally comprises one constant domain (a CL1).
The term "fragment crystallizable" or "Fc" or "Fc region" or "Fc portion" (which can be used interchangeably herein) refers to a region of an antibody comprising at least one constant domain, and which is generally (though not necessarily) glycosylated and which is capable of binding to one or more Fc receptors and/or components of the complement cascade. The heavy chain constant region can be selected from any of the five isotypes: a, 5, a, y, or p. Furthermore, heavy chains of various subclasses (such as the IgG subclasses of heavy chains) are responsible for different effector functions and thus, by choosing the desired heavy chain constant region, proteins with desired effector function can be produced. Exemplary heavy chain constant regions are gamma 1 (IgGl), gamma 2 (IgG2) and gamma 3 (IgG3), or hybrids thereof.
The terms “antigen-binding portion of an antibody”, “antigen-binding fragment”, “antigen-binding domain”, “antibody fragment”, or a “functional fragment of an antibody” are used interchangeably in the present disclosure to mean one or more fragments of an antibody that retain the ability to specifically bind to an antigen, (see generally, Holliger et al., Nature Biotech. (2005) 23 (9):1126-1129. Non-limiting examples of antibody fragments include (i) a Fab fragment, a monovalent fragment consisting of the VL , VH , CL and CHI domains; (ii) a F(ab')2 fragment, a bivalent fragment comprising two Fab fragments linked by a disulfide bridge at the hinge region; (iii) a Fd fragment consisting of the VH and CHI domains; (iv) a Fv fragment consisting of the VL and VH domains of a single arm of an antibody, (v) a dAb fragment (Ward et al., Nature (1989) 341 :544 546), which consists of a VH domain; and (vi) an isolated complementarity determining region (CDR). Furthermore, although the two domains of the Fv fragment, VL and VH , are coded for by separate genes, they can be joined, using recombinant methods, by a synthetic linker that enables them to be made as a single protein chain in which the VL and VH regions pair to form monovalent molecules (known as single chain Fv (scFv) (see, e.g., Bird et al., Science (1988) 242:423 426; and Huston et al., Proc. Natl. Acad. Sci. USA (1988) 85:5879 5883; and Osbourn et al., Nat. Biotechnol. (1998) 16:778). Such single chain antibodies are also intended to be encompassed within the term “antigen-binding portion” of an antibody. Any VH and VL sequences of specific scFv can be linked to human immunoglobulin constant region cDNA or genomic sequences, to generate expression vectors encoding complete IgG molecules or other isotypes. VH and VL can also be used in the generation of Fab, Fv or other fragments of immunoglobulins using either protein chemistry or recombinant DNA technology. Other forms of single chain antibodies, such as diabodies are also encompassed.
As used herein the term "subject" refers to, humans. The terms "subject," "patient," and "individual" are used interchangeably herein.
The term "patient" as used herein includes a living human that is receiving medical care or that should receive medical care due to a disease or condition. This includes subjects with no defined illness or observable symptoms of a disease or condition who are being investigated for signs of pathology.
In some examples, the term “subject” refers to Rh-negative subjects (e.g., a Rh- negative pregnant subject) at risk of or with symptoms of Rhesus alloimmunization. In other examples, the “subject” refers to a fetus or newborn at risk of haemolytic disease owing to Rhesus alloimmunization of the “mother”.
As used herein, a subject "at risk" of developing a disease or condition or relapse thereof or relapsing may or may not have detectable symptoms and may or may not have displayed detectable symptoms prior to treatment according to the present disclosure. "At risk" denotes that a subject has one or more risk factors, which are measurable parameters that correlate with development of the disease or condition, as known in the art and/or described herein.
As used herein, the terms "treating", "treat" or "treatment" include administering an antibody or antigen binding fragment of the disclosure to thereby reduce or eliminate at least one symptom of the disease or condition or to slow progression of the disease or condition. The anti-D antibodies can be used for example as a prophylactic treatment and or treatment for potential sensitising events for Rh negative women who are pregnant or recently pregnant (up to for example 10 days post pregnancy cessation).
As used herein, the term "preventing", "prevent" or "prevention" includes providing prophylaxis with respect to occurrence or recurrence of the disease or condition. An individual may be predisposed to or at risk of developing the disease or disease relapse but has not yet been diagnosed with the disease or the relapse.
An "effective amount" refers to at least an amount effective, at dosages and for periods of time necessary, to achieve the desired result. For example, the desired result may be a therapeutic or prophylactic result. An effective amount can be provided in one or more administrations. In some examples of the present disclosure, the term "effective amount" means an amount necessary to effect treatment of the disease or condition. In some examples of the present disclosure, the term "effective amount" is meant as an amount necessary to deplete or eliminate RhD+ red blood cells but not RhD negative (RhD-) red blood cells. This can be referred to as RBC clearance in the art. Although not wishing to be limited by theory, RBC clearance may be mediated by antibody dependent cell mediated cytotoxicity (ADCC) and/or antibody-dependent cellular phagocytosis (ADCP). See Cruz-Leal et al. Prevention of haemolytic disease of the foetus and newborn: what have we learned from animal models? Current Opinion in Hematology 24(6):p 536-543, 2017 for a general description of these methodologies as they apply to RBC clearance protocols. The effective amount may vary according to the disease or condition to be treated or factor to be altered and also according to the weight, age, racial background, sex, health and/or physical condition and other factors relevant to the subject being treated. Typically, the effective amount will fall within a relatively broad range (e.g. a "dosage" range) that can be determined through routine trial and experimentation by a medical practitioner. Accordingly, this term is not to be construed to
limit the disclosure to a specific quantity. The effective amount can be administered in a single dose or in a dose repeated once or several times over a treatment period.
Antibodies and Antigen Binding Fragments
The present invention relates antibodies to the RhD antigen of human red blood cells. The Rhesus blood group system is a major antigenic constituent of the human red blood cell membrane; of this group, the RhD antigen is of particular clinical importance in relation to isoimmune reactions. An Rh D- individual with anti-RhD who receives RhD+ blood is liable to suffer substantial red blood cell (RBc) destruction due to the Rh(D) phenotype incompatibility, and thus blood of donors must routinely be classified as RhD+ or RhD-.
The RhD antigen is also responsible for haemolytic disease of the newborn (HDN). This condition arises in newborn RhD+ infants of RhD- “mothers” previously sensitised to RhD antigen as a result of IgG anti-RhD antibodies crossing the placenta during pregnancy and causing foetal RBC destruction. Sensitization of the RhD- “mother” to RhD antigen often occurs during the birth of an earlier RhD+ “child” due to some foetal RBCs entering the maternal circulation and being recognised as foreign by the maternal immune system.
To reduce the incidence of HDN, it is routine practice to give polyclonal anti-RhD antibodies to RhD- “mothers” immediately after the birth of an RhD+ infant so that any RhD+ RBCs which may have entered the maternal circulation are rapidly removed (Mollison, P.L., Blood Transfusion is clinical Medicine, 7th Edn., Blackwell Scientific, Oxford, 1983; Laros Jr., R.K., Erythroblastosis Fe-talis. Blood Group Disorders in Pregnancy, 1986, Ch.7, p. 103).
The anti-RhD antibodies and antigen binding fragments of the disclosure are monoclonal antibodies and may be used singly or in combination. The antibodies may be genetically engineered to enhance specific functional activities. In various embodiments, the antibodies and antigen binding fragments comprise all or a portion of a constant region of an antibody. In some embodiments, the constant region is an isotype selected from: IgA (e.g., IgAl or IgA2), IgD, IgE, IgG (e.g., IgGl, IgG2, IgG3 or IgG4), and IgM.
In some embodiments, the antibody or antigen binding fragment is derived from germline genes including IGHV3-33, IGHV3-30, IGHV3-30 3, IGHV3-30 5, IGHV3-33, IGHV1-2, IGHV2-26, IGHV3-21, IGHV3-30, IGHV3-53, IGHV4-34, IGHV4-39, and IGHV4-59. In some embodiments, the IGHD segment of the antibody or antigen binding fragment is derived from germline genes including IGHD 1-26, IGHD2-2, IGHD2-21, IGHD3-3, IGHD3-9, IGHD3-10, IGHD3-16, IGHD3-22, IGHD5-12, IGHD5-18, IGHD6-19, and IGHD6-6. In some embodiments, the JH segment of the antibody or antigen binding fragment is derived from germline genes including IGHJ4, IGHJ6, IGHJ3, and IGHJ5. In
some embodiments, the light chain segment of the antibody or antigen binding fragment is derived from germline genes including IGLV1-47, IGLV2-14, IGLV2-23, IGLV7-43, IGLV1-51, IGKV1-39, and IGKV2-28.
In specific embodiments, the anti-RhD antibodies of the disclosure may comprise a number of isotypes of the same antibody, or combinations of antibodies with different isotypes.
The anti-RhD antibodies and antigen binding fragments of the disclosure may comprise, for example, one or more CDRs (e.g., CDR3), the variable region (or portions thereof), the constant region (or portions thereof), or combinations thereof.
The antibody or antigen-binding fragment may comprise a heavy chain variable region and/or a light chain variable region comprising at least one CDR amino acid sequence (e.g., CDR3) or a VH or VL sequence as defined herein, or a sequence having an amino acid sequence that is at least50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95% or at least 99% identical thereto.
To determine the percent identity of two amino acid sequences or of two nucleic acid sequences, the sequences are aligned for optimal comparison purposes (e.g., gaps can be introduced in the sequence of a first amino acid or nucleic acid sequence for optimal alignment with a second amino acid or nucleic acid sequence). The amino acid residues or nucleotides at corresponding amino acid positions or nucleotide positions are then compared. When a position in the first sequence is occupied by the same amino acid residue or nucleotide as the corresponding position in the second sequence, then the molecules are identical at that position. The percent identity between the two sequences is a function of the number of identical positions shared by the sequences (i.e., % identity = number of identical overlapping positions/total number of positions x 100%). In one embodiment, the two sequences are the same length.
The determination of percent identity between two sequences can also be accomplished using a mathematical algorithm. One, non-limiting example of a mathematical algorithm utilized for the comparison of two sequences is the algorithm of Karlin and Altschul, Proc. Natl. Acad. Sci. USA (1990) 87:2264-2268, modified as in Karlin and Altschul, Proc. Natl. Acad. Sci. USA (1993) 90:5873-5877. Such an algorithm is incorporated into the NBLAST and XBLAST programs of Altschul et al., J. Mol. Biol. (1990) 215:403. BLAST nucleotide searches can be performed with the NBLAST nucleotide program parameters set, for example, for score=100, wordlength=12 to obtain nucleotide sequences homologous to a nucleic acid molecule of the present invention. BLAST protein searches can
be performed with the XBLAST program parameters set, for example, to score-50, wordlength=3 to obtain amino acid sequences homologous to a protein molecule of the present invention. To obtain gapped alignments for comparison purposes, Gapped BLAST can be utilized as described in Altschul et al., Nucleic Acids Res. (1997) 25:3389-3402. Alternatively, PSI-BLAST can be used to perform an iterated search which detects distant relationships between molecules (Id.). When utilizing BLAST, Gapped BLAST, and PSI- Blast programs, the default parameters of the respective programs (e.g., of XBLAST and NBLAST) can be used (see, e.g., the NCBI website). Another preferred, non-limiting example of a mathematical algorithm utilized for the comparison of sequences is the algorithm of Myers and Miller, CABIOS (1988) 4:11-17. Such an algorithm is incorporated in the ALIGN program (version 2.0) which is part of the GCG sequence alignment software package. When utilizing the ALIGN program for comparing amino acid sequences, a PAM 120 weight residue table, a gap length penalty of 12, and a gap penalty of 4 can be used.
The antibody or antigen binding fragment of the disclosure may bind an epitope or could bind RhD sequence variants and partial D epitopes as described in (Vox Sang 1996;70: 123-131). In some embodiments, the antibody or antigen binding fragment contacts residues on RhD including D40, R114, N162, D164, R229, S230, E233, D350, A354, N356, Q362, and/or E369. In further embodiments, the CDRH3 loop plays a critical role in epitope recognition. In further embodiments, the CDRH3 loop consists of 16 to 22 amino acids in length. In further embodiments, antibodies or antigen binding fragments that interact with a greater number of residues on RhD have a greater capacity to inhibit binding of other anti- RhD antibodies or antigen binding fragments.
The antibodies or binding fragments may be antibodies or fragments whose sequences have been modified to insert one or more amino acids into one or more of its hypervariable regions, for example as described in Jung and Pluckthun, Protein Engineering (1997) 10:9, 959-966; Yazaki et al., Protein Eng. Des Sei. (2004) 17(5):481-9; and US 2007/0280931.
In some embodiments, the antibody or antigen binding fragment comprises the constant region or a portion thereof such as a Fc. The constant region or a portion thereof may comprise one or more amino acid substitutions. For example, the Fc may comprise one or more amino acid substitutions that alters binding to an Fc receptor and/or effector function.
In a particular embodiment the Fc receptor is an Fey receptor. In one embodiment, the Fc receptor is a human Fc receptor. In one embodiment the Fc receptor is an activating Fc receptor. In a specific embodiment the Fc receptor is an activating human Fey receptor, more specifically human FcyRIIIa, FcyRI or FcyRIIa, most specifically human FcyRIIIa. In
one embodiment the effector function is one or more selected from the group of complement dependent cytotoxicity (CDC), antibody-dependent cell-mediated cytotoxicity (ADCC), antibody-dependent cellular phagocytosis (ADCP), and secretion of cell death mediators such as cytokines or reactive oxygen species (ROS). In a particular embodiment, the effector function is ADCC.
In some embodiments the Fc is modified by glycosylation, in particular fucosylation and/or galactosylation. The extent of glycosylation and type of glycosylation has been shown to influence antibody activity (Kumpel et al. Hum Antibodies Hybridomas. 1994;5(3-4): 143- 51.; Siberil et al Clin Immunol. 2006 Feb-Mar;118(2-3): 170-9.) In some embodiments the glycosylation pattern maybe modified by chemical or enzymatic modification. In some embodiments this modification may be carried out by Fc mutagenesis, glycoengineering and subclass switching.
In some embodiments modifications may be antibody Fc mutated variants that could include the mutations of IgGl-GASDALIE such as G236A, S239D, A330L and I332E (IgGl- GASDALIE antibody) and G236R and L328 (IgGl-GRLR antibody). Other modifications could be those referred to in Edwards et al. Enhancement of Antibody-Dependent Cellular Cytotoxicity and Phagocytosis in Anti-HIV-1 Human-Bovine Chimeric Broadly Neutralizing Antibodies. J Virol., 95(13, 2021 or Bournazos et al. Broadly neutralizing anti-HIV-1 antibodies require Fc effector functions for in vivo activity. Cell. 2014 Sep 11 ; 158(6): 1243- 1253.
Antibody and Antigen Binding Fragment Activity
The present invention relates to antibody or antigen binding fragments capable of specifically binding to RhD positive (RhD+), but not RhD negative (RhD-), RBCs. Advantageously, the antibody or antigen binding fragment is capable of agglutinating RhD+ red blood cells but not RhD- red blood cells. In some embodiments, the strength of the antibody or antigen binding fragment binding to RhD+ RBCs is correlated with its ability to induce RhD+ RBC agglutination. The antibody or antigen binding fragment may be capable of one or more of the following: inducing phosphatidylserine exposure and Annexin V binding on RhD+ RBCs; inducing Reactive Oxygen Species (ROS) production specifically in RhD+ RBCs; increasing RhD+ RBC size; altering protein expression levels in RhD+ RBCs; and upregulating proteomic pathways associated with cellular stress and lipolysis in RhD+ RBCs.
The antibody or antigen binding fragment may also induce RhD+ RBC natural killer (NK) cell-mediated haemolysis. Antibody or antigen binding fragment induced NK cell-
mediated RhD+ RBC haemolysis may be unable to be supressed with caspase, cathepsin, RIPK1, RIPK2, or proteasome inhibitors. Further, although not wishing to be limited to theory, the antibody or antigen binding fragment may induce NK cell-mediated RhD+ RBC clearance through induction of antibody dependent cell-mediated cytotoxicity (ADCC), wherein the strength of antibody or antigen binding fragment binding to RhD+ RBCs is correlated with its ability to induce ADCC. The antibody or antigen binding fragment may be capable of inducing RhD+ RBC NK cell-mediated haemolysis independent of cell-to-cell contact. The antibody or antigen binding fragment may also induce ROS production in the NK cells. Further, incubation with the antibody or antigen binding fragment treated RhD+ RBCs may upregulate cytokine and chemokine production in NK cells, for example, one or more of RANTES, MIP-la, MIP-1B, IL-8, IL-9, IL-17, IL-12, IL-lb, IFN-y, TNF-a, CD69, NKP46, and NKG2D.
The antibody or antigen binding fragment may be further capable of inducing monocyte-mediated clearance of RhD+ RBCs. Induction of monocyte-mediated clearance of RhD+ RBCs may occur through interactions between the antibody or antigen binding fragment and Fey receptors.
Methods of Manufacture
Methods for generating antibodies are known in the art and/or described in Harlow and Lane (editors) Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory (1988). Nucleic Acids, Recombinant Vectors and Host Cells
The present disclosure encompasses nucleic acid molecules encoding immunoglobulin light and/or heavy chain variable regions for anti-RhD antibodies, vectors comprising such nucleic acids, and host cells capable of producing the anti-RhD antibodies of the disclosure. In certain aspects, the nucleic acid molecules encode, and the host cells are capable of expressing the anti-RhD antibodies and antibody-binding fragments, as well as fusion proteins and chimeric antigen receptors containing them.
An anti-RhD antibody of the disclosure can be prepared by recombinant expression of immunoglobulin light and heavy chain genes in a host cell. To express an antibody recombinantly, a host cell is transfected with one or more recombinant expression vectors carrying DNA fragments encoding the immunoglobulin light and heavy chains of the antibody such that the light and heavy chains are expressed in the host cell and, optionally, secreted into the medium in which the host cells are cultured, from which medium the antibodies can be recovered. Standard recombinant DNA methodologies are used to obtain antibody heavy and light chain genes, to incorporate these genes into recombinant expression vectors and to
introduce the vectors into host cells, such as those described in Molecular Cloning; A Laboratory Manual, Second Edition (Sambrook, Fritsch and Maniatis (eds), Cold Spring Harbor, N. Y., 1989), Current Protocols in Molecular Biology (Ausubel, F. M. et al., eds., Greene Publishing Associates, 1989) and in U.S. Pat. No. 4,816,397.
To generate nucleic acids encoding such anti-RhD antibodies, DNA fragments encoding the light and heavy chain variable regions are first obtained. These DNAs can be obtained by amplification and modification of germline DNA or cDNA encoding light and heavy chain variable sequences, for example using the polymerase chain reaction (PCR). Germline DNA sequences for human heavy and light chain variable region genes are known in the art (see, e.g., the "VBASE" human germline sequence database; see also Kabat et al., Sequences of Proteins of Immunological Interest, Fifth Edition, U.S. Department of Health and Human Services, NIH Publication No. 91-3242. 1991; Tomlinson et al., J. Mol. Biol. (1992) 22T: 116-198; and Cox et al., Eur. J. Immunol. (1994) 24:827-836).
Once DNA fragments encoding anti-RhD antibody related VH and VL segments are obtained, these DNA fragments can be further manipulated by standard recombinant DNA techniques, for example to convert the variable region genes to full-length antibody chain genes, to Fab fragment genes or to a scFv gene. In these manipulations, a VH- or VL - encoding DNA fragment is operatively linked to another DNA fragment encoding another protein, such as an antibody constant region or a flexible linker. The term “operatively linked,” as used in this context, is intended to mean that the two DNA fragments are joined such that the amino acid sequences encoded by the two DNA fragments remain in-frame.
The isolated DNA encoding the VH region can be converted to a full-length heavy chain gene by operatively linking the VH-encoding DNA to another DNA molecule encoding heavy chain constant regions (CHI, CH2, CH3 and, optionally, CH4). The sequences of human heavy chain constant region genes are known in the art (see, e.g., Kabat et al., Sequences of Proteins of Immunological Interest, Fifth Edition, U.S. Department of Health and Human Services, NIH Publication No. 91-3242, 1991) and DNA fragments encompassing these regions can be obtained by standard PCR amplification. The heavy chain constant region can be an IgGl, IgG2, IgG3, IgG4, IgA, IgE, IgM or IgD constant region, but in certain embodiments is an IgGl or IgG4 constant region. For a Fab fragment heavy chain gene, the VH-encoding DNA can be operatively linked to another DNA molecule encoding only the heavy chain CHI constant region.
The isolated DNA encoding the VL region can be converted to a full-length light chain gene (as well as a Fab light chain gene) by operatively linking the VL-encoding DNA to
another DNA molecule encoding the light chain constant region, CL. The sequences of human light chain constant region genes are known in the art (see, e.g., Kabat et al., Sequences of Proteins of Immunological Interest, Fifth Edition, U.S. Department of Health and Human Services, NTH Publication No. 91-3242, 1991) and DNA fragments encompassing these regions can be obtained by standard PCR amplification. The light chain constant region can be a K or constant region, but in certain embodiments is a K constant region.
To create an scFv gene, the VH- and VL-encoding DNA fragments can be operatively linked to another fragment encoding a flexible linker, for example, encoding the amino acid sequence (Gly4~Ser)3 , such that the VH and VL sequences can be expressed as a contiguous single-chain protein, with the VH and VL regions joined by the flexible linker (see, e.g., Bird et al., Science (1988) 242:423-426; Huston et al., Proc. Natl. Acad. Sci. USA (1988) 85:5879- 5883; McCafferty et al., Nature (1990) 348:552-554).
To express the anti-RhD antibodies of the disclosure, DNAs encoding partial or full- length light and heavy chains, obtained as described above, are inserted into expression vectors such that the genes are operatively linked to transcriptional and translational control sequences. In this context, the term "operatively linked" is intended to mean that an antibody gene is ligated into a vector such that transcriptional and translational control sequences within the vector serve their intended function of regulating the transcription and translation of the antibody gene. The expression vector and expression control sequences are chosen to be compatible with the expression host cell used. The antibody light chain gene and the antibody heavy chain gene can be inserted into separate vectors or, more typically, both genes are inserted into the same expression vector.
The antibody genes are inserted into the expression vector by standard methods (e.g., ligation of complementary restriction sites on the antibody gene fragment and vector, or blunt end ligation if no restriction sites are present). Prior to insertion of the anti-RhD antibody- related light or heavy chain sequences, the expression vector can already carry antibody constant region sequences. For example, one approach to converting the anti-RhD monoclonal antibody related VH and VL sequences to full-length antibody genes is to insert them into expression vectors already encoding heavy chain constant and light chain constant regions, respectively, such that the VH segment is operatively linked to the CH segment(s) within the vector and the VL segment is operatively linked to the CL segment within the vector. Additionally, or alternatively, the recombinant expression vector can encode a signal peptide that facilitates secretion of the antibody chain from a host cell. The antibody chain gene can be cloned into the vector such that the signal peptide is linked in-frame to the amino terminus
of the antibody chain gene. The signal peptide can be an immunoglobulin signal peptide or a heterologous signal peptide (i.e., a signal peptide from a non-immunoglobulin protein).
In addition to the antibody chain genes, the recombinant expression vectors of the disclosure carry regulatory sequences that control the expression of the antibody chain genes in a host cell. The term “regulatory sequence” is intended to include promoters, enhancers and other expression control elements (e.g., polyadenylation signals) that control the transcription or translation of the antibody chain genes. Such regulatory sequences are described, for example, in Goeddel, Gene Expression Technology: Methods in Enzymology 185, Academic Press, San Diego, Calif., 1990. It will be appreciated by those skilled in the art that the design of the expression vector, including the selection of regulatory sequences may depend on such factors as the choice of the host cell to be transformed, the level of expression of protein desired, etc. Suitable regulatory sequences for mammalian host cell expression include viral elements that direct high levels of protein expression in mammalian cells, such as promoters and/or enhancers derived from cytomegalovirus (CMV) (such as the CMV promoter/enhancer), Simian Virus 40 (SV40) (such as the SV40 promoter/enhancer), adenovirus, (e.g., the adenovirus major late promoter (AdMLP)) and polyoma. For further description of viral regulatory elements, and sequences thereof, see, for example, US 5,168,062, US 4,510,245, and US 4,968,615.
In addition to the antibody chain genes and regulatory sequences, the recombinant expression vectors of the disclosure can carry additional sequences, such as sequences that regulate replication of the vector in host cells (e.g., origins of replication) and selectable marker genes. The selectable marker gene facilitates selection of host cells into which the vector has been introduced (see, e.g., US 4,399,216, US 4,634,665 and US 5,179,017). For example, typically the selectable marker gene confers resistance to drugs, such as G418, hygromycin or methotrexate, on a host cell into which the vector has been introduced. Suitable selectable marker genes include the dihydrofolate reductase (DHFR) gene (for use in DHFR- host cells with methotrexate selection/amplification) and the neo gene (for G418 selection). For expression of the light and heavy chains, the expression vector(s) encoding the heavy and light chains is transfected into a host cell by standard techniques. The various forms of the term “transfection” are intended to encompass a wide variety of techniques commonly used for the introduction of exogenous DNA into a prokaryotic or eukaryotic host cell, e.g., electroporation, lipofection, calcium-phosphate precipitation, DEAE— dextran transfection and the like.
It is possible to express the antibodies of the disclosure in either prokaryotic or eukaryotic host cells. In certain embodiments, expression of antibodies is performed in eukaryotic cells, e.g., mammalian host cells, of optimal secretion of a properly folded and immunologically active antibody. Exemplary mammalian host cells for expressing the recombinant antibodies of the disclosure include Chinese Hamster Ovary (CHO cells) (including DHFR- CHO cells, described in Urlaub and Chasin, Proc. Natl. Acad. Sci. USA (1980) 77:4216-4220, used with a DHFR selectable marker, for example, as described in Kaufman and Sharp, Mol. Biol. (1982) 159:601-621), Expi-CHO, NSO myeloma cells, COS cells and SP2 cells. Other cells lines such as the rat hybridoma cell line, YB2/0 are also exemplary owing to their ability to generate low fucosylation of the antibody. (Teylaert et al, BMC Biotechnol 11(1) (2011)). In some embodiments the addition of supplements such as kifunensine or 2-F-peracetyl fucose, galactose or dexamethasone to the medium can also influence glycosylation (Ehret et al, Biotechol Bioeng 116(4) (2019)). FUT8 KO CHO cells (Yang et al, Frontiers Chem, 9 (2021)) have also been shown to produce completely non- fucosylated antibodies. Other ways to reduce fucosylation include manipulation of culture conditions (Konno et al Cytotechnology, 64(3) (2012)).
Host cells can also be used to produce portions of intact antibodies, such as Fab fragments or scFv molecules. It is understood that variations on the above procedure are within the scope of the present disclosure. For example, it can be desirable to transfect a host cell with DNA encoding either the light chain or the heavy chain (but not both) of an anti- RhD antibody of this disclosure.
Once an anti-RhD antibody of the disclosure has been produced by recombinant expression, it can be purified by any method known in the art for purification of an immunoglobulin molecule, for example, by chromatography (e.g., hydroxyapatite chromatography, hydrophobic interaction chromatography, gel electrophoresis, dialysis, affinity chromatography (e.g., protein A affinity chromatography or protein G chromatography), size exclusion chromatography, ion exchange, centrifugation, differential solubility, or by any other standard technique for the purification of proteins. Once isolated, the anti-RhD antibody can, if desired, be further purified, e.g., by high performance liquid chromatography, or by gel filtration chromatography.
Where a protein is secreted into culture medium, supernatants from such expression systems can be first concentrated using a commercially available protein concentration filter.
A protease inhibitor such as PMSF may be included in any of the foregoing steps to inhibit proteolysis and antibiotics may be included to prevent the growth of adventitious
contaminants. Alternatively, or additionally, supernatants can be filtered and/or separated from cells expressing the protein, e.g., using continuous centrifugation.
Further, the anti-RhD antibodies of the present disclosure and/or binding fragments can be fused to heterologous polypeptide sequences described herein or otherwise known in the art to facilitate purification. For example, a poly-histidine tag, for example, a hexahistidine tag, or an influenza virus hemagglutinin (HA) tag, or a Simian Virus 5 (V5) tag, or a FLAG tag, or a glutathione S-transferase (GST) tag. The resulting protein is then purified using methods known in the art, such as, affinity purification. For example, a protein comprising a hexa-his tag is purified by contacting a sample comprising the protein with nickel-nitrilotriacetic acid (Ni-NTA) that specifically binds a hexa-his tag immobilized on a solid or semi-solid support, washing the sample to remove unbound protein, and subsequently eluting the bound protein. Alternatively, or in addition, a ligand or antibody that binds to a tag is used in an affinity purification method.
Pharmaceutical Compositions
The anti-RhD antibodies and fragments of the disclosure may be in the form of compositions comprising one or more of the anti-RhD antibodies or binding fragments and one or more carriers, excipients and/or diluents. The form of the composition (e.g., dry powder, liquid formulation, etc.) and the excipients, diluents and/or carriers used will depend upon the intended uses of the antibody or fragment, for therapeutic uses, the mode of administration. In preferred embodiments, the compositions consist of two or more anti-RhD antibodies or binding fragments. In more preferred embodiments the compositions consist of three or more anti-RhD antibodies or binding fragments.
Methods of Use
The anti-RhD antibody or binding fragment or composition disclosed herein can be used in the treatment or prevention of neonatal haemolytic disease, idiopathic thrombocytopenic purpura (ITP), or RhD (-) individuals who have been transfused with RhD (+) blood.
In preferred embodiments, the anti-RhD antibody or binding fragment or composition of the disclosure is useful for the prevention of the Rh-negative alloimmunization of Rh- negative individuals. Without wishing to be bound by theory, the mechanism of action involves specific binding of antibodies or fragments thereof to RhD positive red cell D antigen, followed by removal of these red blood cells from the circulation, thought to occur primarily at the spleen. This clearance is associated with a dynamic mechanism of suppression of the primary immune response in individuals and thus prevents immunization.
Thus, the antibody or binding fragment or composition of the disclosure is particularly useful for the prevention of haemolytic disease in the newborn, by administration to a RhD negative “mother”. It is also possible to use an antibody or fragment or composition of the disclosure in a prophylactic manner for the prevention of the alloimmunization of a RhD- negative woman, immediately after the birth of a Rh-positive child, and to prevent, during subsequent pregnancies, haemolytic disease of the newborn; during abortions, extrauterine pregnancies in a situation of RhD incompatibility or in transplacental haemorrhages resulting from amniocentesis, chorionic biopsies, or traumatic obstetric manipulations in a situation of incompatibility for Rhesus D.
Profiling antibodies that bind to the antigen (RhD) in donor plasma
The present invention utilises a method of sequencing antibodies from the blood of donors who have been screened for anti-RhD antibody expression.
In some embodiments, donor blood is first separated into plasma and PBMC fractions by centrifugation. The total IgG content may be purified by incubation of the plasma fraction with, for example, Protein G agarose. RhD-specific antibodies may be further isolated by incubation of the total IgG pool with RhD-positive RBCs. Antibody-bound RBCs are preferably washed to remove non-specific binders, before eluting antibodies with, for example, EDTA/glycine acid, resulting in a pool of RhD-specific antibodies. A similar selection step can be performed by incubating the total IgG pool with RhD-negative RBCs, to isolate antibodies that bind to non-RhD antigens present on the RBCs. Both RhD-specific and non-RhD specific pools of antibodies may then be subjected to peptic digest and mass spectrometry to identify the antibody sequences. Advantageously, sequences identified in only the RhD-specific pool (considered to be RhD-specific binders) can be used for further analysis.
From the PBMC fraction of donor blood, B-lymphocytes may be analysed for relative antibody transcription levels. Live CD 19+ IgG+ cells may be enriched from donor PBMCs by fluorescence activated cell sorting (FACS), whereafter single-cell RNA sequencing can be performed. Such sequencing provides the benefit of pairing full-length heavy and light chain sequences in expressed B cell receptors (BCRs). Combining the resulting transcriptomic data with proteomic analysis of the RhD specific binders allows for identification of high frequency, linked heavy and light chains specific for RhD.
EXAMPLES
Herein is described the method for identifying and then preparing human monoclonal antibodies (mAbs) capable of activating effector cells such as NK cells and monocytic THP- 1 cells and resulting in RBC clearance through ADCC and phagocytosis, respectively. The resultant antibodies ae then tested for their biological function using a range of assays listed herein below and described in detail in the experimental sections. i. Agglutination assay (Plate-based) ii. RBC binding assay (FACS-based) iii. Competitive binding assay between mAbs iv. Epitope mapping v. Inhibition of anti-RhD mAb binding by a commercial hyperimmune RhD-pIgG-1 vi. Phagocytosis assay (FACS-based) vii. ADCC (FACS-based) viii. NK cell activation assay (FACS-based) ix. Cytokine secretion assay
1: Generation and characterization of anti-RhD antibodies
Methods for identification of RhD specific antibodies
Blood collection and processing
Peripheral blood was obtained from 27 ‘super producers’ participating in the RhD program at Australian Red Cross Lifeblood (ARCL). Peripheral Blood Mononuclear cells (PBMC) were purified from the blood using standard techniques (see for example Panda and Ravindran, 2013, Bio-protocol 3(3): and Panda et al 2012, PLoS Pathog 8(5). RBC were lysed from the final cell pellet and subsequently, PBMCs were washed with cold PBS and frozen at -80 °C using heat-inactivated 90% foetal calf serum and 10% dimethylsulfoxide (DMSO).
B cell sorting (antibody nucleotide sequencing).
Samples were divided into 4 groups (each group pool of 5-11 samples) and stained with BV605 anti -human IgG, APC-H7 CD 19 and SYTOX Green dead cell stain. Then, live CD19+ IgG+ cells were sorted and again each 2 samples were pooled (final n=2) fori OX Genomics gene sequencing (NGS).
Purification of anti-RhD antibodies from donors’ plasma.
In this present invention donor blood is first separated into plasma and PBMC fractions by centrifugation. The total immunoglobulin fraction is purified by affinity chromatography (using Protein G agarose). RhD-specific antibodies are further isolated by incubation of the total IgG pool with RhD-positive RBCs. Antibody-bound RBCs are washed to remove nonspecific binders, before eluting antibodies with EDTA/glycine acid, resulting in a pool of RhD-specific antibodies.
Proteomics was employed to determine differential protein abundances
The antibodies prepared as above were analysed by mass spectrometry-based proteomics. To obtain small peptides for further analysis samples were prepared by digesting with AspN, GluC and trypsin digestion and chymotrypsin as follows.
For AspN, GluC and trypsin digestion: Antibodies (2 pg) were incubated with 50 mM ammonium bicarbonate, 8M Urea and 1 mM DTT for 30 minutes and 37°C. Antibodies were alkylated using 10 mM iodoacetamide for 1 hour, and subsequently quenched with 10 mM DTT. Samples were diluted with 50 mM ammonium bicarbonate to reduce UREA to IM concentration. 100 ng trypsin, AspN or GluC was added for overnight digestion at 37°C. Peptides were then desalted using Cl 8 STAGE tips and collected into new tubes by centrifugation. The collected peptides were lyophilized to dryness using a CentriVap (Labconco), before reconstituting in 10 pl 0.1% FA/2% ACN ready for mass spectrometry analysis.
For chymotrypsin digestion: Antibodies (2 pg) were incubated with lOOmM Tris-HCl pH 7.6, 8M UREA and 1 mM DTT for 30 minutes and 37°C. Antibodies were alkylated using 10 mM iodoacetamide for 1 hour, and subsequently quenched with 10 mM DTT. Samples were diluted with lOOmM Tris-HCl to reduce UREA to IM concentration. 10 mM CaCh was added to each sample, and 100 ng chymotrypsin was added for overnight digestion at 37°C. Peptides were then desalted using C 18 STAGE tips and collected into new tubes by centrifugation. The collected peptides were lyophilized to dryness using a CentriVap (Labconco), before reconstituting in 10 pl 0.1% FA/2% ACN ready for mass spectrometry analysis.
Mass spectrometry analysis
Peptides (5 pl) were separated by reverse-phase chromatography on a Cis fused silica column packed into an emitter tip (lonOpticks), using a nano-flow HPLC (M-class, Waters).
Creating heavy and light chain IgG .fasta file
Using lOx Genomics the output of CellRanger (all contig annotations.j son) was used to create a .fasta file for subsequent MaxQuant searching. The .fasta file consisted of the barcodes, amino acid and nucleotide sequences that were identified as high confidence = TRUE and full_length= TRUE.
Raw data processing and analysis
Raw data files were analysed by MaxQuant vl .6.17 software using the integrated Andromeda search engine. Experiment type was set as TIMS-DDA with no modification to default settings. Data was searched against an in-house generated heavy /light chain IgG .fasta database and a separate reverse decoy database using a strict trypsin, AspN, GluC or chymotrypsin specificity allowing up to 2 missed cleavages. The minimum required peptide length was set to 7 amino acids. Modifications: Carbamidomethylation of Cys was set as a fixed modification, while N-acetylation of proteins and oxidation of Met were set as variable modifications. First search peptide tolerance was set at 10 ppm and main search set at 20 ppm (other settings left as default). Peptide-spectrum match and protein identifications were filtered using a target-decoy approach at an FDR of 1%. Label-free quantification (LFQ) quantification was selected, with a minimum ratio count of 2. Peptide-spectrum match scores and protein identifications were filtered using a target-decoy approach at an FDR of 1%.
Only proteins that were quantified in at least 50% of replicates in at least one condition were kept. Normalisation was performed by applying RUVIIIC[2] where the most invariant sequences were used as negative controls. The optimum k value was determined based on PCA, RLE and p-value distribution plots.
Missing values were imputed by using Missing Not At Random (MNAR) method. This was achieved by substituting ‘NAs’ with numbers that were drawn from a normal distribution with a mean that is left-shifted from the sample mean by 1.8 standard deviation with a width of 0.3.
Differential expression and enrichment analysis
The R-package limma (v. 3.50.1) was used to perform the differential analysis to compare the differential protein abundance of protein sequences identified in RhdNeg vs RhdPos samples. Protein sequences were determined to be differentially expressed if the false discovery rate (FDR) was < 0.05 and fold change 2.
Data refinement
Proteomics data for RhD specific antibodies was aligned with antibody sequences obtained from single cell sorted total IgG+ B cells.
Specifically, RhD specific antibody sequences were extracted and converted to a FASTA file and then subjected to analysis. The data sets were combined, and IGHM sequences were removed from the dataset. Heavy chains were assigned new names such as clone- 1, clone-2, etc., if the sequences were unique. In cases where cells had identical heavy-light pairs, only one cell was included in the analysis, and the repeated sequences from other cells were excluded. Thus, combining the transcriptomics sequence information with the proteomic sequence information to further assemble and derive the complete amino acid sequences of the most abundant mAbs in the RhD specific IgG subset proved very powerful and efficient.
This quantitative proteomics approach to identify the RhD peptide sequences that were specifically enriched from the RhD+ IgG compared with the RhD - IgG in both the LC and HC regions was a novel approach. Moreover, these sequences were ranked in order based on the adjusted p. value from the quantified fold changes of enrichment. This approach was different to merely sequencing the peptides. Without this quantitative proteomics comparison, it wouldn’t have been possible to identify the specific RhD sequences listed herein.
Example 2: The below method was used to identify antibodies with high affinity.
The antibody genes identified in Experiment 1 were then synthesised and cloned into vectors expressing human antibody gamma, kappa, or lambda constant regions (Figure 2). Each antibody variable gene was cloned into a DNA plasmid expressing its natural heavy and light constant region according to single cell NGS data. The paired antibody heavy and light expression DNA plasmids were co-transfected into Expi-CHO cells according to the manufacturer’s instructions (ThermoFisher Scientific). The antibodies were purified with protein G agarose and used for further characterization.
Characterization and analysis of the biological activity of the preferred mAbs (called specific clones throughout the specification) using the techniques herein described.
RBC preparation
RhD+ (ROr phenotype: Dce/dce) and RhD- (r’r phenotype: dCe/dce) O Blood group was collected and centrifuged at 200xg for 10 min. Plasma was discarded and 3mL DPBS (+2mM EDTA) was added to the cells. The tube was centrifuged at 200xg for 10 min and huffy coat and supernatant was discarded. The cells were washed two more times with DPBS (+2mM EDTA) and used in the experiments immediately.
To deglycerolize RBC, the cryovial was incubated at 37 °C for 2min, inverted gently and centrifuged at 1500xg for 5 min. The supernatant was discarded except the last 50pl layering RBC pellet. Packed RBCs were resuspended and 12% NaCl (25 pl, over 5min) was added. The cells were incubated at RT for 3 min to equilibrate. 237.5 pl of 1.6% NaCl was added (over 5 min) followed by an extra 0.5 mL 1.6% NaCl (also over 5min). The RBCs were transferred to sterile 15mL conical tubes and 1.25 mL isotonic wash (0.8% NaCl, 0.2% (l lmM) glucose, pH 7.4) was added over 5mins. Following addition of 4 mL isotonic wash (over 8 min), the cells were centrifuged at 1500 xg for 5min. The RBC pellet was resuspended in 5 mL buffer (3: 1 mix of 1.6% NaCl and the isotonic wash) and centrifuged at 1500 xg for 5 min. Washed RBC were finally resuspended in DPBS and used immediately or stored at 4 °C for a maximum of one week.
Treatment of RBCs with Bromelain removes surface glycans and is expected to improve antibody access which can enhance ADCC. For Bromelain treatment, the washed RBCs were resuspended in DPBS and treated with Bromelain as follows: 2% RBC in DPBS was mixed with 0.5% Bromelain (w/v) (1 :2 ratio) and incubated at 37 °C for 10 min. The cells were centrifuged at 1500xg for 5 min and washed with DPBS twice.
Agglutination assay
Agglutination assay was performed to assess RhD binding of candidate antibodies. Briefly, 50mL Bromelain treated O blood group RhD+ or RhD- RBC (1% haematocrit), 25mL mAb and 25mL anti-IgG secondary antibody _AHG colour (BIO-RAD laboratories #804115) were added to each well of U-bottom 96-well plate and incubated 90 min at RT. Agglutination was then visualized by the naked eye.
To further assess the agglutination activity of candidate mAbs on RBCs, both Bromelain- treated and untreated RBCs were incubated with the serial dilutions of mAbs in the presence anti-IgG secondary antibody _AHG colour. Th plate was incubated at 37 °C for 1 hour (using Bromelain treated RBC) or 2-3 hours (using untreated RBC). Agglutination was then visualized and scored by the naked eye. Commercial (non-enriched) polyclonal RhD-pIgG-1 was used as a control in this experiment.
RBC Binding (FACS-based)
To evaluate the binding of mAbs to RhD antigen, a flow cytometry analysis was performed. In brief, mAbs (50mL) were incubated with 50mL Bromelain-treated O blood group RhD+ or RhD- RBC (200,000) in U-bottom 96-well plate at 37 °C for 30 min. The cells were washed with 200mL wash buffer (DPBS containing 0.5% BSA and 2mM EDTA) and centrifuged at 1500 xg for 2 minutes. The supernatant was discarded, and 1/1000 dilution of Fab goat antihuman IgG (H&L) Texas Red (Rockland antibodies and assays # 809-1902) was added to each well. The plate was incubated at room temperature for 30 min and centrifuged at 1500 xg for 2 minutes. The plate was washed once more with 200mL wash buffer and the cells were resuspended in 50mL wash buffer. The samples were acquired on an Fortessa instrument (BD) and analysed using FlowJo vlO.6.2. Commercial (non-enriched) polyclonal RhD-pIgG- 1 was used as a control in this experiment.
Competitive binding assay between mAbs
A competition RBC binding assay was conducted to evaluate whether the anti-RhD mAbs target the same or overlapping epitopes and to what extent they can compete with each other for binding.
For this purpose, the binding of FITC-conjugated anti-RhD mAbs were evaluated the presence and absence of other anti-RhD mAbs. Briefly, 100,000 bromelain-treated RhD+ RBCs (50pL) were incubated with unconjugated mAbs (50pL) for 40 minutes at 37°C then centrifuged at 1500 xg for 1 minute to remove unbound antibodies. Supernatant discarded and RBCs were washed with DPBS. Afterwards, FITC-conjugated anti-RhD mAbs were added (lOOpL) and the cells were incubated at 37°C for 40 minutes. The cells were washed as above and resuspended in DPBS. The samples were acquired on an Fortessa instrument (BD) and analysed using FlowJo vl0.6. 2. Brad3 and Fogl mAbs served as positive controls for anti- RhD mAb binding.
FITC-conjugated mAbs were used at concentrations that achieved approximately 70% binding to RhD+ RBCs. Unconjugated mAbs were applied at concentrations that, where feasible, resulted in at least 90% inhibition in a self-competition assay with the corresponding FITC-conjugated mAb.
THP-1 phagocytosis assay (FACS-based)
The monocytic THP-1 cell line (ATCC) was grown in RPMI culture medium + 10% FCS at 37 °C and 5% CO2 atmosphere. 10,000 cells were incubated with 1/40 dilutions of mouse anti-human CD64 (FcyRI)- PE Cy7 (BD Biosciences; Cat no. 561191), 1/10 dilutions of
mouse anti-human CD32 (FcyRII)- PE (BD Biosciences; Cat no. 552884), and 1/10 dilutions of mouse anti-human CD16 (FcyRIII)- FITC (BD Biosciences; Cat no. 555406) for 30 min on ice to assess the expression of FcyRs. The cells were washed with DPBS by centrifugation at 500xg for 5 min. The samples were acquired on an Fortessa instrument (BD) and analysed using FlowJo vlO.6.2.
For the phagocytic assay, monocytic THP-1 cells were labelled with 1 pM CellTrace™ Violet (CTV) (Invitrogen; Cat no.C34571) in DPBS at 37°C for 1 hours with gentle shaking. The cells were washed with RPMI+ 10% FCS and centrifuged at 500xg 5 min. RPMI+ 10% FCS was added to the cells and incubated at 37°C for 15 min followed by washing and resuspending in RPMI+ 10% FCS (4 x 105 cells/ml). 2 x 107 bromelain-treated RBC was resuspended in ImL DPBS and IpL amine-reactive pHrodo™ Red, succinimidyl ester (ThermoFisher Scientific; Cat no. P36600) was added. The cells were incubated at 37°C for
1 hours (with gentle shaking), washed with RPMI+ 10% FCS and centrifuged at 1500xg 5 min. RBCs were resuspended in RPMI+ 10% FCS and incubated at 37°C for 15 min followed by washing and resuspending in RPMI+ 10% FCS (8 x 106cells/ml).
To examine phagocytosis, various concentrations of antibodies (25pL) were incubated with
2 x 105 pHrodo™ Red-labelled RhD+ (25pL) and 20,000 CTV THP-1 monocytes (50pL) for 16 hours at 37°C in a 96-well U-bottom plate. The plate was centrifuged at 500xg 5 min followed by addition of 100 pL cold RBC lysis buffer and incubation on ice for 5 min. The plate was washed with DPBS twice and the samples were acquired on an Fortessa instrument (BD) and analysed using FlowJo vlO.6.2. The FACS plot is shown in. The phagocytosis score was calculated using the formula: (% CTV+ /pHrodo red+ cells x mean fluorescent intensity)/! 000. The phagocytosis score of “no Ab” control was subtracted from all samples. Enriched RhD-pIgG-1 was used as a control in this experiment.
ADCC assay (FACS-based)
Bromelain-treated RhD+ RBCs were resuspended in RPMI media containing 3% human AB serum and 7% FCS at a concentration of 80,000 cells per mL. Then 25mL RhD+ RBC (20,000 /well), 25mL purified antibody (lOOng/well or lug/mL in RPMI + 3% human AB serum + 7% FCS), and 50mL effector cells (600,000 monocyte-depleted PBMC/well) were incubated overnight at 37°C. Subsequently, 2mL Triton X-100 (ThermoFisher Scientific #28314) was added to the control well (No antibody control) and incubated for 5 minutes at 37°C to achieve the maximal haemoglobin release. The cells were then centrifuged at 1500xg for 2 minutes and the supernatants were collected. A 1/100 dilution of the supernatant was used to detect the haemoglobin in each well using Human HB (Hemoglobin) ELISA Kit (Elabscience #E- EL-H0415) according to the manufacturer's instructions. The percent specific RBC lysis (% ADCC) was calculated using the formula: 100 * (experimental release - spontaneous release)/(maximum release - spontaneous release). Enriched RhD-pIgG-1, prepared according to Experiment 2, was used as a control in these assays.
NK cell activation (FACS-based)
To prepare the effector cells, frozen PBMCs were thawed at 37 °C then NK cells were enriched using human NK cell isolation kit (Miltenyi Biotec #130-092-657) according to the manufacturer’s instruction. 100,000 Bromelain-treated RhD+ RBCs (25 pL), with lOOng/mL antibody (25 pL) and 100,000 purified NK cells (50 pL) were incubated at 37 degrees for 4 hours then the cells were stained for surface or intracellular markers. Commercial (nonenriched) polyclonal RhD-pIgG-1 was used as a control in these assays.
Cytokine release assay
To prepare the effector cells, frozen PBMCs were thawed at 37 °C then NK cells were enriched using human NK cell isolation kit (Miltenyi Biotec #130-092-657) according to the manufacturer’s instruction. 100,000 Bromelain-treated RhD+ RBCs (25 pL), with lOOng/mL antibody (25 pL) and 100,000 purified NK cells (50 pL) were incubated overnight at 37°C. Then the supernatant was harvested, and the presence of cytokines or chemokines was investigated using Bioplex kit (Bio-rad) according to the manufacturer’s instruction. Commercial (non-enriched) polyclonal RhD-pIgG-1 was used as a control in these assays.
Epitope mapping
Epitope mapping was conducted using different assays.
Micro-Agglutination assay: 50mL Bromelain treated RBCs (1% haematocrit), 25mL antibody and 25mL anti-IgG secondary antibody _AHG colour (BIO-RAD laboratories #804115) were added to each well of U-bottom 96-well plate and incubated 90 min at RT. Agglutination was then visualized by the naked eye. The antibody concentration used in the assay was between 1 and 25 pg/ml for the various anti-RhD mAbs variants of this invention. Indirect antiglobulin test (IAT) assay was performed using Bio-Rad LISS/Coombs kit (BIO-RAD) according to the manufacturers’ instruction.
FACS-based binding assay: To evaluate the binding of the antibody (mAbs) to RhD antigen, a flow cytometry analysis was performed. In brief, antibody (50mL) was incubated with 50mL Bromelain-treated RBCs (200,000) in U-bottom 96-well plate at 37 °C for 30 min. The cells were washed with 200mL DPBS and centrifuged at 1500 xg for 2 minutes. The supernatant was discarded, and 1/1000 dilution of Fab goat anti-human IgG (H&L) Texas Red (Rockland antibodies and assays # 809-1902) was added to each well. The plate was incubated at room temperature for 30 min and centrifuged at 1500 xg for 2 minutes. The plate was washed once more with 200mL wash buffer and the cells were resuspended in 50mL wash buffer. The samples were acquired on an Fortessa instrument (BD) and analysed using FlowJo vlO.6.2. Commercial (non-enriched) polyclonal RhD-pIgG-1 was used as a control in this assay.
Inhibition of anti-RhD mAb binding by polyclonal RhD-pIgG-1
A competition RBC binding assay was conducted to evaluate whether commercial polyclonal (non-enriched) RhD-pIgG-1 can suppress the binding of anti-RhD mAbs. This assay can demonstrate the relative binding affinities and potential epitope overlap between the polyclonal RhD-pIgG-1 and the anti-RhD mAbs.
For this purpose, the binding of FITC-conjugated anti-RhD mAbs were evaluated the presence and absence of polyclonal IVIg (Pool of non-RhD antibodies) and RhD-pIgG-1 (derived from Australian donors). Briefly, 100,000 bromelain-treated RhD+ RBCs (50pL) were incubated with 5pg IVIg or RhD-pIgG-1 (50pL) for 30 minutes at 37°C then centrifuged at 1500 xg for 1 minute to remove unbound antibodies. Supernatant discarded and RBCs were washed with DPBS. Afterwards, serial dilutions of FITC-conjugated anti-RhD mAbs were added (lOOpL) and the cells were incubated at 37°C for 30 minutes. The cells were washed as above and resuspended in DPBS. The samples were acquired on an Fortessa instrument (BD) and analysed using FlowJo vl0.6. 2. Fogl mAb served as positive controls for anti-RhD mAb binding.
Enrichment of RhD-pIgGl
Commercial RhD-pIgG consists of a pool of immunoglobulins (antibodies) with a higher titre of anti-RhD specific IgG compared to the titre found in normal plasma-derived IgG. Commercial clinically used product is currently derived from the plasma of RhD- donors who have developed anti-RhD antibodies due to hemolytic disease of the newborn (HDN) or immunization against RhD+ RBCs. However, these preparations contain a mixture of both RhD-specific and non-RhD antibodies.
To directly compare the biological activities of the mAbs from this invention with a commercially available material, the inventors found it necessary to further increase the RhD titre of the commercial material, ensuring the RhD component was detectable in the assays described.
For this purpose, 1.5mL (625IU) of commercially available RhD-pIgG-1 (derived from Australian donors) was diluted with PBS and incubated with 8 mL of packed RhD+ RBCs (ROr variant). The RBCs were then washed at least six times with PBS to remove non-specific binders. To elute the RBC-bound antibodies, 16 mL of 50 mM glycine, 0.9% NaCl, 2 mM EDTA (pH 2.7) was added, the mixture was vortexed for 5 seconds, and incubated on ice for 2 minutes. The RBCs were immediately centrifuged at 2500xg for 5 minutes, and the supernatant was collected. The acidic pH was neutralized by adding 2.1 mL of 1 M Tris, 5.2% NaCl (pH 8), resulting in a pool of eluted RhD-specific antibodies.
To remove potential RBC protein contaminants, further enrichment was achieved by affinity chromatography using Protein G agarose followed by buffer exchange to PBS.
Results of Experiment 2
2.1 Anti-RhD monoclonal antibodies (mAb) agglutinated RhD+ RBC.
The results indicate that out of 180 candidate mAbs tested, 16 mAbs agglutinated RhD+ RBCs without showing any agglutination activity against RhD- RBCs(Table 2). The agglutination endpoint concentration or agglutination titre indicates the lowest concentration of the antibody at which visible agglutination of RBCs occurs. A lower agglutination endpoint concentration suggests higher antibody affinity and potency, as it requires less antibody to achieve agglutination.
Additionally, we observed no significant difference in the agglutination activity of most anti- RhD mAbs when tested on untreated and bromelain-treated RhD+ RBCs. These findings suggest that the binding of these mAbs is not influenced by RBC surface glycans and is
independent of glycan modifications (Figure 4). RhD-pIgG-1, a polyclonal product containing anti-RhD antibodies derived from Australian blood donors and currently available on the market, agglutinated RhD+ RBCs while also exhibiting slight non-specific reactivity against RhD- RBCs.
K: kappa light chain; L: lambda light chain.
2.2 Investigating binding of anti-RhD mAbs to RBCs.
The anti-RhD mAbs displayed specific binding to RhD+ RBCs. Notably, Clone-30 mAb exhibited binding to over 75% of RhD+ RBCs, while seven mAbs (Clone 18, 27, 38, 55, 65, 91 and 104) showed binding to 51-75% of the cells. Four mAbs (Clone 21a, 34, 44 and 107) demonstrated moderate binding, interacting with 25-50% of the RBCs, and four other mAbs (Clone la, 17, 75 and 142) exhibited weak binding, attaching to less than 25% of the cells (Table 3).
Table 3. RBC (%) binding of isolated mAbs
RBC binding of mAbs was assessed at a concentration of 1 pg/mL, except for Clone la-IgG3 (5 pg/mL), Clone 17 (5 pg/mL), Clone 21a-IgG2 (5 pg/mL), Clone 75 (25 pg/mL), Clone 142 (5 pg/mL), and the commercial RhD-pIgGl (25 pg/mL). Unless otherwise specified, all mAbs are IgGl . Pool K consists of 16 unique anti-RhD mAbs with different Fc isotypes. No binding is shown as The commercial RhD-pIgGl, although having a higher titre of RhD, is not specifically enriched for anti-RhD antibodies.
2.3 Competitive binding assay between mAbs
Among tested mAbs, 5 mAbs including (Clone-la, Clone -30, Clone -44, Clone -65, and Clone -104) exhibited significant suppression of RBC binding for all the other tested mAbs, with an inhibition rate above 70% (Figure 5A). Clone-38, Clone-55 and Clone-91 also showed broad suppressive capabilities, inhibiting the binding of all mAbs, but displayed slightly reduced suppression against 3-4 mAbs. On the other hand, the remaining mAbs, including Clone-18, Clone-21a, Clone-27, Clone-34, and Clone-107 exhibited partial suppression against other mAbs. While they were able to inhibit the binding of some mAbs, their suppression efficacy varied and was not as comprehensive as observed with the above listed 8 mAbs. Clone-17, Clone-75 and Clone-142 exhibited the lowest competition activity. Clone- 106, identified as a non-RhD mAb, did not show any suppression against anti-RhD mAbs, indicating the specificity of the assay. This comprehensive analysis provides valuable insights into the inhibitory potential of these mAbs and their cross-reactivity with one another.
Among the five mAbs capable of suppressing RBC binding for all 13 tested mAbs, Clone- 104 demonstrated the highest potency, achieving an inhibiting concentration of 0.5 pg/mL (Figure 5B-C). Following closely, Clone-30 exhibited significant inhibitory effects at a concentration of 2 pg/mL comparable to Brad3 control anti-RhD mAbs. For Clone-44, complete suppression required a concentration of 10 pg/mL, while Clone-la, Clone-55, and Clone-65 achieved full suppression at a higher concentration of 20 pg/mL. These findings highlight the varying potencies of the different mAbs in suppressing RBC binding, with Clone- 104 and Clone-30 demonstrating the most potent inhibitory effects at lower concentrations. Clone-142, Clone-17, and Clone-75 inhibited the binding of fewer than seven other mAbs, with less than 50% suppression observed. This suggests that the epitopes targeted by these mAbs may not overlap significantly with those recognised by the mAbs whose binding was not effectively suppressed.
2.4 Epitope mapping results
As shown in Figure 6A, all RhD mAbs, including Brad3, Fogl, and commercial RhD-pIgG- 1, demonstrated specific binding to wild-type (WT) RhD variants (ROr, R1R1, and R2R2) but not to the rr variant (RhD- RBC). Testing Weak RhD type 2 from different donors revealed variable binding for Clone-17, Clone-75, and Clone-142. These mAbs also did not bind to Weak RhD type 1. Additionally, Clone-17, Clone-75, Clone-34, and Clone-55 did not bind to
Weak RhD type 4. The other tested mAbs demonstrated satisfactory binding to the Weak RhD variants.
Most of the mAbs (clones) demonstrated reactivity against the DVII and DSC2 partial RhD variants, with only two mAbs (clones 27 and 142) exhibiting binding to the DVI variant. The mAbs displayed variable binding to other partial RhD variants, while a few mAbs (Clones 30, 38, 44, 91) consistently showed strong binding across partial RhD variants. As shown in Figure 6B, only Clone-27 and Clone-142 could potentially interact with epitopes 3.1 and 4.1. Clone- 142 was the only mAb to demonstrate potential reactivity with epitope 15.1, while other mAbs displayed variable results due to differing levels of reactivity with the tested RhD variants. Reactivity with epitopes 10.1 and 11.1 could not be determined due to the unavailability of RhD variants exposing these epitopes. Among the tested mAbs, Clone-30, Clone-38, Clone-44, Clone-55, Clone-91, and Clone-142 showed potential reactivity with all other listed epitopes, whereas the other mAbs produced variable results.
2.5 Inhibition of anti-RhD mAb binding by polyclonal RhD-pIgG-1
As shown in Figure 7A, the binding of all mAbs was suppressed by RhD-pIgG-1. Slight inhibition with IVIg was detected for Clone-la, Clone-21a, and Clone-142. Figure 7B shows that 44-79% of mAb binding is suppressed by RhD-pIgG-1, with the lowest inhibition for Clone-38 (44%) and the highest inhibition for Clone- la and Clone-27 (79%). The significant suppression of mAb binding by RhD-pIgG-1 suggests that these mAbs are targeting similar or overlapping epitopes on the RhD antigen. The differential inhibition observed among the mAbs indicates variability in epitope specificity and binding strength.
2.6 Assessing the activity of mAbs in mediating RBC clearance by THP-1 monocytes
Flow cytometric analysis was performed using the gating strategy in Figure 8A and it revealed that monocytic THP-1 cells expressed CD64 (FcyRI) and CD32 (FcyRII) receptors on their surface, but not CD 16 (FcyRIII) (Figure 8B). All tested anti-RhD mAbs, with the exception of Clone_21a_IgG2, were capable of mediating the clearance of RhD+ RBC by THP-1 monocytes. Most mAbs, along with RhD-pIgG-1, exhibited a prozone-like effect at concentrations exceeding 100 ng/mL (Figure 8C), indicating an excess of antibody relative to the antigen. The mAbs displayed varying degrees of ADP activity, with six of them (Clones 27, 55, 65, 75, 104, and 107) showing ADP activity comparable to RhD-pIgG-1 (Figure 8D). Notably, subclass switching from IgG2 to IgGl in Clone_21a enhanced ADP activity (Figure 8E).
These findings suggest that specific anti-RhD mAbs can effectively induce monocyte- mediated clearance of RhD+ RBCs, likely through interactions with Fey receptors and the subsequent activation of effector functions. Additionally, preferred embodiments of this invention include selecting use of mAbs which have a predominance of IgGl subclass to enhance ADP activity The prozone-like effect observed at higher concentrations further underscores the potential for fine-tuning antibody formulations to achieve optimal therapeutic outcomes. The unique characteristics of these anti-RhD mAbs highlight their potential for development into effective therapeutic agents for conditions involving RhD+ RBCs.
2.7 Assessing the activity of mAbs in mediating RBC clearance through antibodydependent cellular cytotoxicity (ADCC)
ADCC or RBC lysis was assessed based on a haemoglobin ELISA assay (Figure 9 A). While RhD-pIgG-1 demonstrated 48% ADCC activity, all tested mAbs exhibited less than 20% ADCC activity (Figure 9B). Among the IgGl mAbs, Clone 30 (19.9%), Clone 65 (16.1%), Clone 91 (17.6%), and Clone 107 (15.4%) showed the highest ADCC activity, suggesting that the targeted epitope may influence ADCC activity. Notably, Clone 104 (IgG3) exhibited 15.4% RBC lysis activity; however, switching to the IgGl subclass resulted in a reduction to 10.5% (Figure 9C). Furthermore, subclass switching from IgG2 to IgGl for Clone 21 enhanced ADCC activity by 2.5-fold (from 1.8% to 4.6%).
According to the literature, antibodies derived from plasma donors have different glycosylation patterns compared to normal antibodies, which improves their ADCC activity. This is potentially the case for RhD-pIgG-1. However, when producing the mAbs in CHO cells, the glycosylation differs from natural anti-RhD antibodies in donors, reducing their ADCC activity. This demonstrates the need for further antibody engineering to achieve comparable ADCC activity.
2.8 Investigating the mechanism of RBC clearance by NK cells
The mAbs of this invention show a comparable biological activity to other commercially available antibodies when evaluating cytokine and chemokine secretion by NK cells. The cytokine and chemokine secretion profile of the mAbs closely resembled that of commercial RhD-pIgG-1, indicating a similar pattern of immune response induced by both antibody preparations. Among the cytokines and chemokines found to be upregulated were RANETS, MIP-la, MIP-1B, IL-8, IL-9, IL-17, IL-12, IL-lb, IFN-y, and TNF-a (Figure 10A). These immune mediators play diverse roles in immune responses, including recruiting and activating immune cells to sites of inflammation. While their individual roles in inducing RBC death vary, the combined effect of these immune mediators may contribute to RBC clearance
through phagocytosis and immune cell-mediated cytotoxicity. This enhanced immune response suggests that NK cells are activated upon encountering RBCs opsonised with anti- RhD antibodies, and highlights the complex interplay between these antibodies, NK cells, and RBCs in RhD-related immune-mediated haemolytic disorders.
The upregulation of intracellular IFN-y and TNF-a, along with the increased expression of CD 107 on the surface of NK cells, was observed in response to anti-RhD mAbs mirroring the pattern seen with RhD-pIgG-1 (Figure 10B). Additionally, certain anti-RhD mAbs demonstrated enhanced expression of CD69, NKP46, and NKG2D on NK cell surfaces. These findings highlight the similarity in immune activation between anti-RhD mAbs, and the commercial RhD-pIgG-1, as evidenced by the shared upregulation of immune mediators and NK cell activation markers. The collective impact of these immune responses may contribute to RBC clearance in RhD-related haemolytic disorders.
2.9 Investigating the subtype and germline gene usage of mAbs.
The nucleotide sequences of the antibody heavy and light variable genes were analysed using IMGT-V-Quest, which allowed for the identification and recognition of their respective germline gene counterparts.
Among the 16 isolated anti-RhD mAbs, the majority (13) were classified as IgGl, with one antibody as IgG2 and two antibodies as IgG3, see Table 4. The primary germline gene utilized in these antibodies was IGHV3-33 (Figure 11, Table 5), in line with previous studies that reported VH3.33 superspecies (including IGHV3-30, IGHV3-30 3, IGHV3-30 5, and IGHV3-33) as the dominant gene segment for anti-RhD mAbs (McGowan et al., 2021; Andersen et al., 2007; Dohmen et al., 2006). Nonetheless, our isolated antibodies also exhibited usage of other germline genes such as IGHV1-2, IGHV2-26, IGHV3-21, IGHV3- 30, IGHV3-53, IGHV4-34, IGHV4-39, and IGHV4-59. In the IGHD segment, the germline genes IGHD 1-26, IGHD2-2, IGHD2-21, IGHD3-3, IGHD3-9, IGHD3-10, IGHD3-16, IGHD3-22, IGHD5-12, IGHD5-18, IGHD6-19, and IGHD6-6 were identified. Dominant germline genes in the JH domain were IGHJ4 and IGHJ6, while IGHJ3 and IGHJ5 were less prevalent (Table 5). Remarkably, antibody clones la and 91 shared identical germline genes (IGHV3-33, IGHD5-18, and IGHJ4), yet functional assays demonstrated distinct binding and activity against RhD+ RBCs.
The mAbs of this invention displayed a CDRH3 size ranging from 15 to 22 amino acids (moderate-long length) (Table 4 and Table 5) and IGHV somatic hypermutation (SHM) levels ranging from 0.69% to 13.12% (Table 5). The CDRH3 size plays a critical role in antibody
binding specificity and affinity, with longer CDRH3 regions often associated with increased antigen recognition and potential for enhanced binding to target epitopes.
The IGHJ mutation displays relatively diverse levels, ranging from 0% to 20.83% (Table 5). Based on the findings, the level of SHM in the IGHV gene can be categorised as relatively low to moderate. The percentage of IGHV gene mutation indicates the extent to which the antibody sequences have undergone changes during the immune response, with higher percentages suggesting greater diversification. In this context, the observed mutation levels fall within a moderate range, implying that the antibodies have undergone some degree of mutation, but not to the extent seen in highly mutated antibodies. The CDRH3 size also plays a critical role in antibody binding specificity and affinity, with longer CDRH3 regions often associated with increased antigen recognition and potential for enhanced binding to target epitopes.
Out of the 16 antibodies analyzed, 10 exhibited lambda light chains, while 6 displayed kappa light chains (Table 6). The identified germline genes for the lambda light chains were IGLV1- 47, IGLV2-14, IGLV2-23, IGLV7-43, IGLV1-51, while for the kappa light chains, the genes were IGKV1-39 and IGKV2-28. Notably, the predominant germline genes for both light chain types were IGLV1-47 and IGKV1-39 (Figure 12). In terms of mutations, the IGLV region showed a mutation range of 1.04% to 5.38%, while the J segment mutations ranged from 0% to 13.16% (Table 6).
The isolated antibodies exhibited a higher SHM rate in IGHV and longer CDR compared to the light chain (Figure 14A-B). Alignment of antibody heavy variable genes showed that the mAbs possess unique sequences in their CDRH3 regions.
Table 5. Germline gene usage of the heavy chain in anti-RhD mAbs
Table 6. Germline gene usage of the light chain in anti-RhD mAbs
2.10 Alphafold modelling.
Using Alphafold modelling, an Al system that predicts a protein 3D structure from its amino acid sequence and protein-protein interactions, it can be observed that the antibody heavy chains, specifically the CDR H3 regions, play a crucial role as the primary antibody paratopes (Figure 13). Additional analysis predicted that the antibodies formed hydrogen bonds and salt bridges with the external RhD loops (Table 7. Figure 14A shows the predicted interacting amino acids on RhD antigen while Figure 14B demonstrates the potential binding sites on the molecule. Asp350 was a common target site among modelled mAbs (Figure 15). Table 7. Prediction of the interactions between anti-RhD mAbs and RhD antigen.
Experiment 3: Production of single clones or a polycell bank.
With the high affinity clones now identified, the methods outlined in the invention could be adapted to produce single clones, from a single cell bank, or alternatively a polycell bank could be used, as described in Frandsen et al, Biotech Bioeng, 108 (9) 2011, or US20090017017.
Experiment 4: Fc modifications
Introducing point mutations into the antibody Fc region can improve ADCC and antibody dependent phagocytosis (ADP). Glycosylation, and more specifically fucosylation of antibody Fc regions can also enhance ADCC by improving the interaction of IgG with Fc gamma receptors (FcyRs) on NK cells.
In the following experiments the inventors used two anti-RhD antibodies described in the literature (Brad3 and Fogl) and undertook Fc mutagenesis, glycoengineering or both modifications to look to improve ADCC and ADP function.
4.1 Preparation of antibodies.
Heavy and light variable genes of Brad3 (GenBank: X64149.1 and X64162.1), Fogl (GenBank: X64150.1 and X64163.1) (both anti-RhD mAbs) and VRC03 (GenBank: GU980707.1 and GU980706.1) (non-RhD mAb) were codon optimized then synthesized by integrated DNA technologies. To construct IgGl variants, the synthesized heavy and light variable genes were digested with Agel/ Sall-HF and Agel BsiWi-HF. respectively and cloned into human gamma 1 constant region (heavy IgGl) and kappa constant region (light) expression vectors. IgG3 variants were constructed by synthesis of human gamma 3 (IgG3) constant gene (by Integrated DNA Technologies Inc) then digested with Sall-HF/BamHI-HF and substituted with IgGl constant gene in heavy expression vectors.
To make antibody Fc mutated variants (IgGl-GASDALIE and IgGl-GRLR), heavy gamma 1 gene (IgGl) was mutated to introduce following mutations: G236A, S239D, A330L and I332E (IgGl-GASDALIE antibody) and G236R and L328 (IgGl-GRLR antibody). An overlap extension PCR was used to mutate heavy Fc genes. Three separate initial PCR reactions were performed with unique oligos. The products of three PCR reactions were separately run on 2% agarose gel, extracted, and purified using the Nucleospin gel and PCR clean up kit. The purified PCR products were mixed in 1 : 1 : 1 ratio in an overlap extension PCR to form contiguous Fc sequences. The PCR was set up as mentioned before except that no oligos were added in the mix. The final PCR products were run on an agarose gel, extracted
and purified. The purified mutated Fc genes were digested with Sall-HF/BamHI-HF and substituted with IgGl gamma 1 constant gene in heavy expression vectors.
Antibody expression vectors containing heavy variable and constant (IgGl, IgGl- GASDALIE, IgGl-GRLR and IgG3) genes and light variable and constant (kappa) genes were co-transfected into Expi- Chinese Hamster Ovary (Expi-CHO) cells using an Expi-CHO expression system kit. Supernatants containing antibodies were harvested 4-5 days after transfection and filtered using 0.22 mm filters. The antibodies were purified using Protein G Agarose Fast Flow then buffer exchanged into PBS using Amicon 50 kDa spin membranes (Millipore) and sterilized.
To inhibit fucosylation and produce an IgGl-Afucosylated (IgGl-Afuc) antibody variant, heavy chain expression plasmids (IgGl and IgGl-GASDALIE) and light chain expression plasmids (kappa) were con-transfected into ExpiCHO cells in the presence of 50 pM Fucostatin. The antibodies were harvested and purified as above.
The purified antibodies were run on NuPage 4-12% Bis-Tris Gel and either incubated with Goat horseradish peroxidase (HRP) conjugated anti-Human (H+L) or with biotinylated Lectin followed by Avidin HRP. Reduction of fucosylation was confirmed by western blot using IgGl-Afucosylated purified mAbs compared with IgGl mAbs.
4.2 Anti-RhD antibody variants agglutinated RhD+ RBC.
O blood group RBC carrying RhD antigen (ROr) was agglutinated in the presence of all Brad3 and Fogl variants but not VRC03 variants. RhD- O blood group RBC (r’r) was agglutinated with none of the antibodies, confirming the specificity of Brad3 and Fogl against RhD antigen. Agglutination is a very sensitive assay and the agglutination endpoint concentration of both Brad3 and Fogl variants was 5ng/mL showing that the agglutination activity was maintained in engineered anti-RhD mAbs.
4.3 RBC binding was maintained in engineered antibodies.
To confirm that binding of Fc engineered antibodies to RhD antigen was maintained, a flow cytometry assay was performed. As shown in Figure 16 modification in the Fc region of anti- RhD antibodies did not affect RhD antigen binding. None of the antibody variants showed binding to RhD- RBC . VRC03 antibody was confirmed to not bind ROr RhD+ RBC.
4.4 Glycoengineering and Fc mutagenesis improved NK cell activation and RBC clearance through ADCC
Figure 17A illustrates the method used to assess NK cell activation. The impact of these modifications on NK cell activation is shown in Figure 17B. Although there were some differences observed among the variants, there was no significant difference in the
enhancement of NK cell activation between the GASDALIE Fc variant and the IgGl A fucosylated variant, or the variant with both modifications. Consistent results were obtained in the ADCC assay, indicating that these antibody modifications improved ADCC compared to WT variant and demonstrated activity comparable to enriched RhD-pIgG-1 (Figure 17C).
4.5. Fc modification maintained ADP activity in anti-RhD mAbs.
As shown in Figure 18 A, anti-RhD mAbs mediated phagocytosis of RhD+ RBCs but this activity was not obtained for control mAbs (non-RhD mAbs). The phagocytosis with monocytic THP-1 showed that Brad3 and Fogl mAb variants mediated RhD specific phagocytosis activity against RhD+ RBC (ROr phenotype) (Figure 18A) while RhD- RBC (r’r phenotype) failed to mediate any response (data not shown) in the presence of antibodies. The ADP activity was also maintained in Fc modificed mAbs (Figure 20D). The mAb variants demonstrated comparable ADP activity to enriched RhD-pIgG-1 (Figure 18B).
Experiment 5: Comparison of the biological activity of the mAbs generated from the method of the invention with a different commercial hyperimmune RhD IgG.
In previous experiments, the inventors used a commercially derived anti- RhD-pIgG derived from Australian donors. The next experiment repeated some of the previous ones with a different commercial product, this time from US donors. Use of differently sourced hyperimmune products, helps to demonstrate the robustness in the choice of the mAbs, since different ethnicities may give rise to subtly different antigen profiles and thus may give rise to slightly different antigen binding fragments that then may have some bearing on the efficacy and biological activity of selected mAbs.
Enrichment of anti-RhD antibodies from commercial anti-RhD-pIgG derived from U.S. donors (RhD-pIgG-2).
To directly compare the biological activities of the mAbs from this invention with this commercially available material, the inventors found, again, it necessary to further increase the RhD titre of the commercial material, ensuring the RhD component was detectable in the assays described.
For this purpose, 2 mL (1500IU) of commercially available RhD-pIgG-2 was diluted with PBS and incubated with 8 mL of packed RhD+ RBCs (ROr variant). The RBCs were then washed at least six times with PBS to remove non-specific binders. To elute the RBC-bound antibodies, 16 mL of 50 mM glycine, 0.9% NaCl, 2 mM EDTA (pH 2.7) was added, the
mixture was vortexed for 5 seconds and incubated on ice for 2 minutes. The RBCs were immediately centrifuged at 2500xg for 5 minutes, and the supernatant was collected and neutralized by adding 2.1 mL of 1 M Tris, 5.2% NaCl (pH 8). To remove potential RBC protein contaminants, further enrichment was achieved by affinity chromatography using Protein G agarose. This enriched sample was used as a benchmarking sample in the subsequent experiments.
Experiment 6: Generation of mAb pools. Different combinations of mAbs of this invention.
Different mAbs were pooled to provide different combinations to demonstrate the breadth of this invention. By pooling several different clones (variants) of mAbs of this invention, the final composition may preferably provide for a broader range of reactivity and functional diversity. These pools or combinations may more closely mirror the complex mixture of antibodies found in the commercial polyclonal RhD-pIgG products. Thus, combinations, or pools, of antibodies containing 3, 4 or 16 mAbs were generated (Table 8) and assessed in the agglutination assay (Table 9).
The Fc regions of Clone 27, Clone 30, and Clone 65 mAbs were engineered with the GASDALIE mutation, as detailed in Experiment 4. Pools of both wild-type and GASDALIE- mutated variants of these mAbs were subsequently evaluated in agglutination (Table 9), ADCC (Figure 19A-B), and ADP (Figure 19C-D) assays. GASDALIE engineering preserved the agglutination activity of the mAbs (Table 9). Both the individual GASDALIE mAbs and the GASDALIE pool (Pool J) demonstrated enhanced ADCC activity compared to wild-type mAbs and the wild-type pool (Pool I), respectively, particularly when normal (untreated) RBCs were used (Figure 19A-B). When Bromelain-treated RBCs were used, the ADCC activity of both the mAbs and pools was comparable to that of RhD-pIgGl (Figure 19A). However, the GASDALIE mAbs and GASDALIE pool (Pool J) exhibited higher ADCC activity than RhD-pIgGl when untreated RBCs were used (Figure 19B). The ADP activity was maintained in the GASDALIE variants compared to the wild-type formats, with the ADP activity of mAbs and pools being comparable to that of RhD-pIgGl (Figure 19C-D).
In Figure 19, non-RhD isotype control mAb was included and a positive ADCC/ ADP signal was defined as the signal above the background of these negative control samples. The background ADCC/ADP activity of the negative control mAb was subtracted from each experiment.
The combinations, or pools, of mAbs exhibited ADCC activity against Bromelain-treated RBCs (Figure 20A) and ADP activity against both treated and untreated RBC (Figure 20C- D) and the results was comparable to that of both commercially, enriched products, RhD- plgG-l and RhD-pIgG-2. While none of the pools demonstrated ADCC activity against untreated RhD+ RBCs, Pool J (which contains GASDALIE mAbs) showed superior activity, to that of RhD-pIgG-1 and RhD-pIgG-2 (Figure 20B). This enhanced activity of Pool J against untreated RhD+ RBCs highlights the efficacy of the GASDALIE-engineered mAbs, offering a potentially more effective therapeutic option in settings where untreated RBCs are targeted. The superior performance of Pool J underscores the innovation of this approach, establishing a novel and innovative method for enhancing ADCC activity in anti-RhD mAbs, which may have significant clinical and therapeutic applications.
In Figure 20 a negative control pool (containing non-RhD isotype control mAbs: one IgGl mAb, one IgG2 mAb and one IgG3 mAb) was included and a positive ADCC signal was defined as a signal above the background of the negative control pool. The background ADCC activity of the negative control pool was subtracted from each experiment. Each bar represents the mean ADCC percentage with error bars indicating the standard error from the mean of two independent experiments. Statistical analysis was conducted to compare each antibody’s activity against RhD-pIgG-1 using one-way ANOVA followed by Dunnett’s multiplecomparison test (GraphPad Prism v9), ****: p< 0.0001.
Table 9: The agglutination activity of mAb pools and engineered mAbs.
Experiment 7: Investigating the minimal antibody fragments necessary for RBC binding
Bromelain-treated RhD+ RBCs were exposed to the anti-RhD mAb including Brad3_Full length, Brad3_F(ab’)2 and Brad3_Fab fragments. Then, the binding of antibody fragments was assessed in FACS.
The findings, shown in Figure 21, revealed that Brad3 fragments, including Brad3_Full length, Brad3_F(ab’)2, and Brad3_Fab, exhibited binding capabilities to RhD+ RBCs. While the Full-length antibody and F(ab’)2 fragment showed similar binding activity, the Fab fragment exhibited slightly lower binding activity.
Claims
1. An isolated or recombinant antibody or antigen binding fragment thereof capable of specifically binding to Rhesus D (RhD), wherein the antibody or antigen binding fragment thereof comprises a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in any one of SEQ ID NOs: 11, 43, 75, 107, 139, 171, 203, 235, 267, 299, 331, 363, 395, 427, 459, 491; a CDR H2 comprising a sequence as set forth in any one of SEQ ID NOs: 13, 45, 77, 109, 141, 173, 205, 237, 269, 301, 333, 365, 397, 429, 461, 493; and a CDR H3 comprising a sequence as set forth in any one of SEQ ID NOs: 15, 47, 79, 111, 143, 175, 207, 239, 271, 303, 335, 367, 399, 431, 463, 495.
2. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 11; a CDR H2 sequence as set forth in any one of SEQ ID NO: 13; and a CDR H3 sequence as set forth in of SEQ ID NO: 15.
3. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR H 1 sequence as set forth in SEQ ID NO: 43; a CDR H2 sequence as set forth in SEQ ID NO: 45; and a CDR H3 sequence as set forth in SEQ ID NO: 47.
4. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl as set forth in SEQ ID NO: 75; a CDR H2 sequence as set forth in SEQ ID NO: 77; and a CDR H3 sequence as set forth in SEQ ID NO: 79.
5. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 107; a CDR H2 sequence as set forth in SEQ ID NO: 109; and a CDR H3 sequence as set forth in SEQ ID NO: 111.
6. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 139; a CDR H2 sequence as set forth in SEQ ID NO: 141; and a CDR H3 sequence as set forth SEQ ID NO: 143.
7. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 171; a CDR
H2 sequence as set forth in SEQ ID NO: 173; and a CDR H3 sequence as set forth in SEQ ID NO: 175.
8. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR H 1 sequence as set forth in SEQ ID NO: 203; a CDR H2 sequence as set forth in SEQ ID NO: 205; and a CDR H3 sequence as set forth in SEQ ID NO: 207.
9. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR H 1 sequence as set forth in SEQ ID NO: 235; a CDR H2 sequence as set forth in SEQ ID NO: 237; and a CDR H3 sequence as set forth in SEQ ID NO: 239.
10. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR H 1 sequence as set forth in SEQ ID NO: 267; a CDR H2 sequence as set forth in SEQ ID NO: 269; and a CDR H3 sequence as set forth in SEQ ID NO: 271.
11. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 299; a CDR H2 sequence as set forth in SEQ ID NO: 301; and a CDR H3 as set forth in SEQ ID NO: 303.
12. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 331; a CDR H2 sequence as set forth in SEQ ID NO: 333; and a CDR H3 sequence as set forth in SEQ ID NOs: 335.
13. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 363; a CDR H2 sequence as set forth in SEQ ID NO: 365; and a CDR H3 sequence as set forth in SEQ ID NO: 367.
14. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 395; a CDR H2 sequence as set forth in SEQ ID NO: 397; and a CDR H3 sequence as set forth in SEQ ID NO: 399.
15. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 427; a CDR H2 sequence as set forth in SEQ ID NO: 429; and a CDR H3 sequence as set forth in SEQ ID NO: 431.
16. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 459; a CDR H2 sequence as set forth in SEQ ID NO: 461; and a CDR H3 sequence as set forth in SEQ ID NO: 463.
17. The antibody or antigen binding fragment of claim 1, wherein the antibody or antigen binding fragment comprises a CDR Hl sequence as set forth in SEQ ID NO: 491; a CDR H2 sequence as set forth in SEQ ID NO: 493; and a CDR H3 sequence as set forth in SEQ ID NO: 495.
18. The antibody or antigen binding fragment of any one of claims 1 to 17, wherein the antibody or antigen binding fragment thereof comprises a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in any one of SEQ ID NOs: 27, 59, 91, 123, 155, 187, 219, 251, 283, 315, 347, 379, 411, 443, 475, 507; a CDR L2 comprising a sequence as set forth in any one of SEQ ID NOs: 29, 61, 93, 125, 157, 189, 221, 253, 285, 317, 349, 381, 413, 445, 477, 509; and a CDR L3 comprising a sequence as set forth in any one of SEQ ID NOs: 31, 63, 95, 127, 159, 191, 223, 255, 287, 319, 351, 383, 415, 447, 479, 511.
19. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 27; a CDR L2 sequence as set forth in SEQ ID NO: 29; and a CDR L3 sequence as set forth in SEQ ID NO: 31.
20. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 59; a CDR L2 sequence as set forth in SEQ ID NO: 61; and a CDR L3 sequence as set forth in SEQ ID NO: 63.
21. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 91; a CDR L2 sequence as set forth in SEQ ID NO: 93; and a CDR L3 sequence as set forth in SEQ ID NO: 95.
22. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 123; a CDR L2 sequence as set forth in SEQ ID NO: 125; and a CDR L3 sequence as set forth in SEQ ID NO: 127.
23. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as
set forth in SEQ ID NO: 155; a CDR L2 sequence as set forth in SEQ ID NO: 157; and a CDR L3 sequence as set forth in SEQ ID NO: 159.
24. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI as set forth in SEQ ID NO: 187; a CDR L2 sequence as set forth in SEQ ID NO: 189; and a CDR L3 as set forth in SEQ ID NO: 191.
25. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI as set forth in SEQ ID NO: 219; a CDR L2 sequence as set forth in SEQ ID NO: 221; and a CDR L3 sequence as set forth in SEQ ID NO: 223.
26. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 251 ; a CDR L2 sequence as set forth in SEQ ID NO: 253; and a CDR L3 sequence as set forth in SEQ ID NO: 255.
27. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 283; a CDR L2 sequence as set forth in SEQ ID NO: 285; and a CDR L3 sequence as set forth in SEQ ID NO: 287.
28. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 315; a CDR L2 sequence as set forth in SEQ ID NO: 317; and a CDR L3 sequence as set forth in SEQ ID NO: 319.
29. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 347; a CDR L2 sequence as set forth in SEQ ID NO: 349; and a CDR L3 sequence as set forth in SEQ ID NO: 351.
30. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 379; a CDR L2 sequence as set forth in SEQ ID NO: 381; and a CDR L3 sequence as set forth in SEQ ID NO: 383.
31. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 411, a CDR L2 sequence as set forth in SEQ ID NO: 413; and a CDR L3 sequence as set forth in SEQ ID NO: 415.
32. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 443; a CDR L2 sequence as set forth in SEQ ID NO: 445; and a CDR L3 sequence as set forth in SEQ ID NO: 447.
33. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 475; a CDR L2 sequence as set forth in SEQ ID NO: 477; and a CDR L3 sequence as set forth in SEQ ID NO: 479.
34. The antibody or antigen binding fragment of any one of claims 1 to 18, wherein the antibody or antigen binding fragment comprises a VL comprising a CDR LI sequence as set forth in SEQ ID NO: 507; a CDR L2 sequence as set forth in SEQ ID NO: 509; and a CDR L3 sequence as set forth in SEQ ID NO: 511.
35. The antibody or antigen binding fragment of any one of claims 1 to 34, wherein the VH comprises an amino acid sequence having at least 70% sequence identity to the sequence set forth in any one of SEQ ID NOs: 9, 41, 73, 105, 137, 169, 201, 233, 265, 297, 329, 361, 393, 425, 457, 489.
36. The antibody or antigen binding fragment of any one of claims 1 to 35, wherein the VL comprises an amino acid sequence having at least 70% sequence identity to the sequence set forth in any one of SEQ ID NOs: 25, 57, 89, 121, 153, 185, 217, 249, 281, 313, 345, 377, 409, 441, 473, 505.
37. An antibody or antigen binding fragment thereof capable of specifically binding to Rhesus D (RhD), wherein the antibody or antigen binding fragment thereof comprises: a) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 11; CDR H2 comprising a sequence as set forth in SEQ ID NO: 13; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 15; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 27; CDR L2 comprising a sequence as set forth in SEQ ID NO: 29; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 31; b) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 43; CDR H2 comprising a sequence as set forth in SEQ ID NO: 45; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 47; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 59, CDR L2 comprising a sequence
as set forth in SEQ ID NO: 61, and CDR L3 comprising a sequence as set forth in SEQ ID NO: 63; c) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 75; CDR H2 comprising a sequence as set forth in SEQ ID NO: 77; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 79; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 91; CDR L2 comprising a sequence as set forth in SEQ ID NO: 93; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 95; d) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 107; CDR H2 comprising a sequence as set forth in SEQ ID NO: 109; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 111; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 123; CDR L2 comprising a sequence as set forth in SEQ ID NO: 125; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 127; e) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 139; CDR H2 comprising a sequence as set forth in SEQ ID NO: 141; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 143; and a light chain variable region (VL) comprising a CDR1 comprising a sequence as set forth in SEQ ID NO: 155, CDR L2 comprising a sequence as set forth in SEQ ID NO: 157; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 159; f) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 171; CDR H2 comprising a sequence as set forth in SEQ ID NO: 173; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 175; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 187; CDR L2 comprising a sequence as set forth in SEQ ID NO: 189; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 191; g) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 203; CDR H2 comprising a sequence as set forth in SEQ ID NO: 205; and CDR H3 comprising a sequence as set
forth in SEQ ID NO: 207; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 219; CDR L2 comprising a sequence as set forth in SEQ ID NO: 221; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 223; h) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 235; CDR2 comprising a sequence as set forth in SEQ ID NO: 237; and CDR3 comprising a sequence as set forth in SEQ ID NO: 239; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 251; CDR L2 comprising a sequence as set forth in SEQ ID NO: 253; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 255; i) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 267; CDR H2 comprising a sequence as set forth in SEQ ID NO: 269; and CDR3 comprising a sequence as set forth in SEQ ID NO: 271; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 283; CDR L2 comprising a sequence as set forth in SEQ ID NO: 285; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 287; j) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 299; CDR H2 comprising a sequence as set forth in SEQ ID NO: 301; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 303; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 315; CDR L2 comprising a sequence as set forth in SEQ ID NO: 317; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 319; k) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 331; CDR H2 comprising a sequence as set forth in SEQ ID NO: 333; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 335; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 347; CDR L2 comprising a sequence as set forth in SEQ ID NO: 349; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 351;
l) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 363; CDR H2 comprising a sequence as set forth in SEQ ID NO: 365; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 367; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 379; CDR L2 comprising a sequence as set forth in SEQ ID NO: 381; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 383; m) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 395; CDR H2 comprising a sequence as set forth in SEQ ID NO: 397; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 399; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 411; CDR L2 comprising a sequence as set forth in SEQ ID NO: 413; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 415; n) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 427; CDR H2 comprising a sequence as set forth in SEQ ID NO: 429, and CDR H3 comprising a sequence as set forth in SEQ ID NO: 431; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 443; CDR L2 comprising a sequence as set forth in SEQ ID NO: 445; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 447; o) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 459; CDR H2 comprising a sequence as set forth in SEQ ID NO: 461; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 463; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 475; CDR L2 comprising a sequence as set forth in SEQ ID NO: 477; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 479; or p) a heavy chain variable region (VH) comprising a complementary determining region (CDR) Hl comprising a sequence as set forth in SEQ ID NO: 491; CDR H2 comprising a sequence as set forth in SEQ ID NO: 493; and CDR H3 comprising a sequence as set forth in SEQ ID NO: 495; and a light chain variable region (VL) comprising a CDR LI comprising a sequence as set forth in SEQ ID NO: 507; CDR L2 comprising a sequence
as set forth in SEQ ID NO: 509; and CDR L3 comprising a sequence as set forth in SEQ ID NO: 511.
38. An antibody or antigen binding fragment thereof capable of specifically binding to Rhesus D (RhD), wherein the antibody or antigen binding fragment thereof comprises: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO:9; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 25; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 41; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 57; c) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 89; d) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 105; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 121; e) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 137; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 153; f) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 185; g) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 201; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 217;
h) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 233; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 249; i) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 265; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 281; j) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 297; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 313; k) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 329; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 345; l) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 361; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 377; m) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 393; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 409; n) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 425; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 441; o) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 457; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 473; or
р) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 489; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 505.
39. The antibody or antigen binding fragment of claim 37 or claim 38, wherein the VH comprises an amino acid sequence having at least 70% sequence identity to the sequence set forth in any one of SEQ ID NOs: 9, 41, 73, 105, 137, 169, 201, 233, 265, 297, 329, 361, 393, 425, 457, 489.
40. The antibody or antigen binding fragment of any one of claims 37 to 39, wherein the VL comprises an amino acid sequence having at least 70% sequence identity to the sequence set forth in any one of SEQ ID NOs: 25, 57, 89, 121, 153, 185, 217, 249, 281, 313, 345, 377,409, 441, 473, 505.
41. An antibody or antigen binding fragment thereof capable of specifically binding to Rhesus D (RhD), wherein the antibody or antigen binding fragment thereof comprises: a) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 9; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 25; b) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 41; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 57; с) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 73; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 89; d) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 105; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 121; e) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 137; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 153; f) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 185;
g) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 201; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 217; h) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 233; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 249; i) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 265; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 281; j) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 297; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 313; k) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 329; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 345; l) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 361; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 377; m) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 393; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 409; n) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 425; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 441; o) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 457; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 473; or p) a heavy chain variable region (VH) comprising a sequence as set forth in SEQ ID NO: 489; and a light chain variable region (VL) comprising a sequence as set forth in SEQ ID NO: 505.
42. The antibody or antigen binding fragment of any one of claims 1 to 41, wherein the antibody or antigen binding fragment thereof is capable of depleting or eliminating RhD+ red blood cells but not RhD negative (RhD-) red blood cells
43. The antibody or binding fragment of any one of claims 1 to 42, wherein the antibody or binding fragment is of IgG subclass 1 and/or IgG subclass 3.
44. A composition comprising one or more antibodies or antigen binding fragments as defined in any one of claims 1 to 43.
45. A composition comprising a combination of three or more antibodies or antigen binding fragments thereof capable of specifically binding to Rhesus D (RhD), wherein the three or more antibodies or antigen binding fragments are selected from an antibody or antigen binding fragment thereof comprising: a) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 137; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 153; b) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 169; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 185; c) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 233; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 249; d) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 265; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 281 e) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 329; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 345; f) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO: 393; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 409; g) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO425; and a light chain
variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 444; or h) a heavy chain variable region (VH) comprising complementary determining regions (CDRs) Hl, H2, and H3 of the sequence as set forth in SEQ ID NO:489; and a light chain variable region (VL) comprising CDRs LI, L2, and L3 of the sequence as set forth in SEQ ID NO: 505;
46. The composition of claim 44 or 45, comprising a pharmaceutically acceptable carrier.
47. A method of treating or preventing Rhesus alloimmunization of Rh-negative subjects comprising administering to a Rh-negative subject in need thereof an effective amount of the anti-Rd(D) antibody or antigen binding fragment of any one of claims 1 to 46, or a combination thereof; or the composition of any one of claims 44 to 46.
48. The method of claim 47, wherein the subject is pregnant.
49. The method of claim 47, wherein the method prevents haemolytic disease of the fetus and newborn.
50. Use of the antibody or antigen binding fragment of any one of claims 1 to 45, or a combination thereof in the manufacture of a medicament for the treatment or prevention of haemolytic disease of the fetus and newborn.
51. The antibody or antigen binding fragment of any one of claims 1 to 45, or a combination thereof, or composition of claim 43 or claim 44 for treatment or prevention of treating or preventing Rhesus alloimmunization of Rh-negative subjects.
52. A method for profiling the most abundant antibodies that bind to the antigen (RhD) in a polyclonal (pAb) population, the method comprising: a) proteomic analysis of a RhD-specific IgG mixture from plasma to provide proteomic sequence information: i. providing plasma from a donor; ii. affinity purifying a total IgG mixture from the plasma; iii. separating RhD-specific and non-RhD specific IgG subsets by incubation with either RhD+ or RhD- RBCs, respectively; iv. independently fragmenting the RhD-specific and non-RhD specific IgG subsets into short peptides of overlapping sequences using one or more proteases; v. analysing the fragmented peptides by mass spectrometry to identify the sequence of each peptide;
vi. assembling the short peptides into longer peptides that belong to individual antibodies, optionally using computational software; excluding antibodies derived from the non-RhD specific subset from further analysis; b) BCR sequencing: i. providing B cells isolated from blood of the donor; ii. extracting RNA from the B cells; iii. sequencing the RNAs by NGS to provide transcriptomic sequence information; c) combining the transcriptomics sequence information with the proteomic sequence information to further assemble and derive the complete amino acid sequences of the most abundant mAbs in the RhD specific IgG subset.
53. The method of claim 52, wherein step (c ) further comprises identification of most the abundant mAbs that were specifically enriched in both the LC and HC regions by ranking the sequences based on quantified fold changes of enrichment from the RhD specific IgG subset.
54. A method for profiling the most abundant antibodies that bind to the antigen (RhD) in a polyclonal (pAb) population, the method comprising: a) proteomic analysis of a RhD-specific IgG mixture from plasma to provide proteomic sequence information: b) BCR sequencing: c) combining the transcriptomics sequence information with the proteomic sequence information to further assemble and derive the complete amino acid sequences of the most abundant mAbs in the RhD specific IgG subset: d) further selecting from the most abundant mAbs a combination of mAbs each of which comprises a different genetic and/or amino acid sequence and each mAb containing a IgGl subclass heavy chain variable region (VH).
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| AU2023902692A AU2023902692A0 (en) | 2023-08-23 | Anti-Rhesus D human monoclonal antibodies | |
| AU2023902692 | 2023-08-23 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025039046A1 true WO2025039046A1 (en) | 2025-02-27 |
Family
ID=94731150
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/AU2024/050904 Pending WO2025039046A1 (en) | 2023-08-23 | 2024-08-23 | Anti-rhesus d human monoclonal antibodies |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025039046A1 (en) |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1991007492A1 (en) * | 1989-11-13 | 1991-05-30 | Central Blood Laboratories Authority | Monoclonal antibodies |
| WO2006007850A1 (en) * | 2004-07-20 | 2006-01-26 | Symphogen A/S | Anti-rhesus d recombinant polyclonal antibody and methods of manufacture |
| WO2010079510A2 (en) * | 2008-12-31 | 2010-07-15 | Bharat Serums And Vaccines Ltd. | Anti-rhd monoclonal antibodies |
-
2024
- 2024-08-23 WO PCT/AU2024/050904 patent/WO2025039046A1/en active Pending
Patent Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO1991007492A1 (en) * | 1989-11-13 | 1991-05-30 | Central Blood Laboratories Authority | Monoclonal antibodies |
| WO2006007850A1 (en) * | 2004-07-20 | 2006-01-26 | Symphogen A/S | Anti-rhesus d recombinant polyclonal antibody and methods of manufacture |
| WO2010079510A2 (en) * | 2008-12-31 | 2010-07-15 | Bharat Serums And Vaccines Ltd. | Anti-rhd monoclonal antibodies |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP6718488B2 (en) | Modulation of complement-dependent cytotoxicity by modification of the C-terminus of antibody heavy chains | |
| JP6417442B2 (en) | Human tissue factor antibody and use thereof | |
| US10717785B2 (en) | Binding molecules that bind human complement factor C2 and uses thereof | |
| CN107660212A (en) | The monoclonal antibody of AntiCD3 McAb 03 | |
| RU2710532C2 (en) | NOVEL ANTI-HUMAN Igβ ANTIBODY | |
| EP2450381A1 (en) | Anti-RHD monoclonal antibodies | |
| CA2943857C (en) | Antibodies against hpa-1a | |
| EP4298125A1 (en) | Antibodies that bind cd123 and gamma-delta t cell receptors | |
| KR20210108972A (en) | Antiperiostin antibodies and uses thereof | |
| CN109963877B (en) | PCSK9 antibody, antigen-binding fragment thereof and medical application thereof | |
| US12180303B2 (en) | Anti-human MASP-2 antibody, preparation method therefor and application thereof | |
| WO2025039046A1 (en) | Anti-rhesus d human monoclonal antibodies | |
| EP4613767A1 (en) | Bispecific antibody for glypican-3 and use thereof | |
| CN119080933B (en) | Antibodies targeting CD16b and uses thereof | |
| RU2817146C2 (en) | Agent for inhibiting iron absorption by cells | |
| WO2024078381A1 (en) | Antibody or antigen-binding fragment thereof binding to human cd39, and preparation method therefor and use thereof | |
| TW201702263A (en) | Anti-CD20-/anti-BAFF bispecific antibodies | |
| WO2026020409A1 (en) | Upar antibody and use thereof | |
| TW202502827A (en) | Ige binding proteins and uses thereof | |
| CN116888150A (en) | anti-Plasmodium antibodies | |
| IL305346A (en) | Antibodies | |
| AU2014200308A1 (en) | Anti-RhD monoclonal antibodies | |
| WO2019212341A2 (en) | Improved anti-fibronectin eda antibodies | |
| TW201335184A (en) | Human tissue factor antibody and uses thereof |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24855149 Country of ref document: EP Kind code of ref document: A1 |