WO2025029872A1 - Magnetic beads for detergent removal - Google Patents
Magnetic beads for detergent removal Download PDFInfo
- Publication number
- WO2025029872A1 WO2025029872A1 PCT/US2024/040308 US2024040308W WO2025029872A1 WO 2025029872 A1 WO2025029872 A1 WO 2025029872A1 US 2024040308 W US2024040308 W US 2024040308W WO 2025029872 A1 WO2025029872 A1 WO 2025029872A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- beads
- hydrophobic
- hydrophilic
- magnetic
- ligands
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N33/00—Investigating or analysing materials by specific methods not covered by groups G01N1/00 - G01N31/00
- G01N33/48—Biological material, e.g. blood, urine; Haemocytometers
- G01N33/50—Chemical analysis of biological material, e.g. blood, urine; Testing involving biospecific ligand binding methods; Immunological testing
- G01N33/53—Immunoassay; Biospecific binding assay; Materials therefor
- G01N33/543—Immunoassay; Biospecific binding assay; Materials therefor with an insoluble carrier for immobilising immunochemicals
- G01N33/54313—Immunoassay; Biospecific binding assay; Materials therefor with an insoluble carrier for immobilising immunochemicals the carrier being characterised by its particulate form
- G01N33/54326—Magnetic particles
-
- B—PERFORMING OPERATIONS; TRANSPORTING
- B01—PHYSICAL OR CHEMICAL PROCESSES OR APPARATUS IN GENERAL
- B01J—CHEMICAL OR PHYSICAL PROCESSES, e.g. CATALYSIS OR COLLOID CHEMISTRY; THEIR RELEVANT APPARATUS
- B01J20/00—Solid sorbent compositions or filter aid compositions; Sorbents for chromatography; Processes for preparing, regenerating or reactivating thereof
- B01J20/28—Solid sorbent compositions or filter aid compositions; Sorbents for chromatography; Processes for preparing, regenerating or reactivating thereof characterised by their form or physical properties
- B01J20/28002—Solid sorbent compositions or filter aid compositions; Sorbents for chromatography; Processes for preparing, regenerating or reactivating thereof characterised by their form or physical properties characterised by their physical properties
- B01J20/28009—Magnetic properties
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2446/00—Magnetic particle immunoreagent carriers
- G01N2446/80—Magnetic particle immunoreagent carriers characterised by the agent used to coat the magnetic particles, e.g. lipids
- G01N2446/84—Polymer coating, e.g. gelatin
Definitions
- MAGNETIC BEADS FOR DETERGENT REMOVAL CROSS REFERENCE TO RELATED APPLICATION This application claims the benefit of the earlier filing date of U.S. provisional patent application No.63/517,029, filed August 1, 2023, which is incorporated herein by reference in its entirety.
- FIELD This disclosure concerns magnetic beads that contain ligands suitable for removing detergent(s) from mixtures, including mixtures that comprise biological molecules including polypeptides, proteins, RNA, and/or DNA, and methods for making the beads and using the beads.
- BACKGROUND Mass spectrometry (MS)-based proteomics has become the most comprehensive approach for protein identification, quantitation, interactions, modifications, and structural characterization.
- proteomics sample preparation is highly complex and variable, with many different protocols available.
- bottom-up proteomics sample preparation requires multiple steps including protein extraction, reduction and alkylation of cysteines, digestion of the proteins into peptides, cleanup, and concentration of the peptides for LC-MS analysis.
- the lack of standardization in proteomic sample preparation makes it difficult to accurately compare results from different laboratories and different protocols.
- SUMMARY Detergents including non-ionic and ionic detergents, are frequently used in mass spectrometry (mass spec; MS) sample preparation.
- the method comprises forming a suspension comprising a first plurality of magnetic beads and a first liquid that contains a first amount of a detergent; agitating the suspension; and removing the magnetic beads from the suspension to leave a solution comprising a second amount of the detergent that is less than the first amount of the detergent.
- the second amount of the detergent may be less than 0.1% of the second solution, such as less than 0.01% of the second solution, and may be substantially zero.
- Each magnetic bead may comprise a crosslinked polymer, a magnetic material, and a plurality of hydrophobic ligands formed from a reaction between azlactone moieties one or more hydrophobic molecules.
- the hydrophobic molecules may comprise a straight or branched chain C4-C20alkyl amine, for example, butylamine (C4), hexylamine (C6), octylamine (C8), decylamine (C10), dodecylamine (C12), tetradecylamine (C14), hexadecylamine (C16), octadecylamine (C18), eicosylamine (C20), or a combination thereof.
- C4-C20alkyl amine for example, butylamine (C4), hexylamine (C6), octylamine (C8), decylamine (C10), dodecylamine (C12), tetradecylamine (C14), hexadecylamine (C16), octadecylamine (C18), eicosylamine (C20), or a combination thereof.
- the straight or branched C4-C20 alkyl amine is selected from octylamine (C8), dodecylamine (C12), octadecylamine (C18), or a combination thereof.
- the hydrophobic molecule is octylamine and the hydrophobic ligand is .
- the magnetic bead may further comprise a hydrophilic ligand.
- the hydrophilic ligand may be formed by a reaction between azlactone moieties and one or more hydrophilic molecules.
- the one or more hydrophilic molecules may be selected from O-(2- aminopropyl)-O’-(2-methoxyethyl)polypropylene glycol, 4,7,10-Trioxa-1,13 tridecanediamine, ⁇ -cyclodextrin, dextran, PEG molecules ranging in MW from 2000 Da to 20,000 Da, or a combination thereof.
- the hydrophilic ligand is .
- the magnetic bead comprises as the hydrophilic ligand.
- the two types of ligand are present in a ratio of from 10% hydrophobic:90% hydrophilic to 90% hydrophobic:10% hydrophilic, such as from 20%:80% hydrophobic:hydrophilic to 40%:60% hydrophobic:hydrophilic.
- the suspension may further comprise a second plurality of magnetic beads where each magnetic bead in the second plurality of magnetic beads comprises a crosslinked polymer, a magnetic material, and a plurality of hydrophilic ligands formed from a reaction between azlactone moieties and one or more hydrophilic molecules.
- the first plurality of magnetic beads and the second plurality of magnetic beads may be each provided in amounts sufficient to provide a ratio of beads comprising hydrophobic ligands (referred to as hydrophobic beads) to beads comprising hydrophilic ligands (referred to as hydrophilic beads) of from 10% hydrophobic beads:90% hydrophilic beads to 90% hydrophobic beads:10% hydrophilic beads, such as from 20%:80% hydrophobic beads:hydrophilic beads to 40%:60% hydrophobic beads:hydrophilic beads.
- the detergent may be selected from polyethylene glycol sorbitan monooleate, t-Octylphenoxypolyethoxyethanol, octylphenoxypolyethoxyethanol (NP- 40) , n-Dodecyl- ⁇ -D-maltoside (DDM), sodium dodecyl sulfate (SDS), Glyco-diosgenin (GDN), Lauryl Maltose Neopentyl Glycol (LMNG), or a combination thereof.
- polyethylene glycol sorbitan monooleate t-Octylphenoxypolyethoxyethanol, octylphenoxypolyethoxyethanol (NP- 40) , n-Dodecyl- ⁇ -D-maltoside (DDM), sodium dodecyl sulfate (SDS), Glyco-diosgenin (GDN), Lauryl Maltose Neopentyl Glycol (LMNG), or a combination thereof
- a magnetic bead comprising a crosslinked polymer bead and a magnetic material contained within the polymer bead, and further comprising a plurality of ligands wherein the ligands are selected from: hydrophobic ligands formed from a reaction between azlactone moieties and one or more hydrophobic molecules; hydrophilic ligands formed from a reaction between azlactone moieties and one or more hydrophilic molecules; or a combination of hydrophobic ligands and hydrophilic ligands formed from reactions between azlactone moieties and one or more hydrophobic molecules and one or more hydrophilic molecules.
- the crosslinked polymer may be a crosslinked azlactone polymer, and the magnetic bead may be an azlactone bead.
- the one or more hydrophobic molecules may comprise a straight or branched chain C4-C20alkyl amine as disclosed herein.
- the hydrophilic ligand may be derived from O-(2-aminopropyl)-O’-(2- methoxyethyl)polypropylene glycol, 4,7,10-Trioxa-1,13 tridecanediamine, ⁇ - cyclodextrin, dextran, a PEG molecule having a molecular weight of from 2,000 to 20,000 Da, or a combination thereof.
- the magnetic material may comprise an iron source, such as iron oxide.
- the magnetic material is in the form of a particle, powder, flake, or cluster, and/or may be contained within an agarose bead which in turn may be contained within the crosslinked polymer.
- the magnetic bead may have a bead size of from 20 to 80 microns, such as from 30 to 60 microns. Also disclosed herein are aspects of a kit comprising the magnetic beads, and aspects of a method for making the magnetic beads.
- FIG.1 is a flow chart illustrating one aspect of a standard workflow using the disclosed magnetic beads.
- FIG.2 is a flowchart illustrating an SP3 workflow that includes a PAC workflow and an SP2 workflow.
- FIG.3 is a digital image illustrating one aspect of the disclosed magnetic beads.
- FIG.4 is a graph of differential volume versus size, illustrating the average size of the disclosed magnetic beads, and a table providing the average size and standard deviation.
- FIG.5 is a digital image illustrating one aspect of the disclosed magnetic beads synthesized using iron oxide clusters as the magnetic material.
- FIG.6 is a digital image illustrating one aspect of the disclosed magnetic beads synthesized using iron oxide particles as the magnetic material.
- FIG.7 is a digital image illustrating one aspect of the disclosed magnetic beads synthesized using Dynabeads TM as the magnetic material.
- FIGS.8A and 8B are a digital image of plates from various stages of an SP2 workflow (FIG.8A) and a table of results (FIG.8B), illustrating the surface absorption losses of different bead types.
- FIG.9 is a graph of peptide yield versus bead type, illustrating the peptide yields from different bead types, identified using a quantitative colorimetric peptide assay.
- FIG.10 is a graph of number of peptides versus bead types, illustrating the number of unique peptides isolated with different bead types and identified using nanoLC-MS analysis, 75 i.d.
- Alkyl refers to a saturated aliphatic hydrocarbyl group having from 1 to 25 (C 1-25 ) or more carbon atoms, more typically 1 to 10 (C 1-10 ) carbon atoms such as 1 to 6 (C1-6) carbon atoms or 1 to 4 (C1-4) carbon atoms.
- Carboxy refers to a -CO 2 H functional group.
- Ester refers to a -CO 2 R functional group, where R is an alkyl group, such as a C1-6 alkyl group.
- Nucleic acid refers to a polynucleotide molecule.
- the polynucleotide may be a naturally occurring polynucleotide or a synthetic polynucleotide.
- a nucleic acid may be a DNA, RNA or mixture of DNA and RNA nucleotides.
- a nucleic acid contains from 20 to 10,000 nucleotides or more, such as from 100, 200, 300, 400, 500, 600, 700, 800, 900, 1000, 2000, 3000, 4000, or 5000 nucleotides to 10,000 nucleotides.
- “Peptide” refers to a compound comprising amino acid residues connected by peptide bonds.
- Functional groups derived from an azlactone moiety may comprise moieties derived from a reaction between the azlactone moiety and a nucleophilic reactive group, such as, but not limited to, an amine-containing compound, thiol containing group, alcohol or water.
- Hydrolysis of azlactone or reaction of the azlactone group with an amine- containing compound can add functionality and/or alter the surface properties of the bead.
- hydrolysis of azlactone rings may produce a bead having a plurality of carboxylic acid functional groups on its surface.
- the azlactone group are reacted with an amine-containing compound to increase or decrease the hydrophobicity or hydrophilicity of the bead.
- an amine-containing compound that includes an additional amine reactive group may be reacted with the azlactone bead, where the reactive amine group can be converted to an epoxide, maleimide, or iodoacetyl group using standard methods known in the art.
- azlactone beads are reacted with a compound to expand the number of functional groups attached to the bead.
- Such an approach can be an effective approach for increasing the binding capacity of the bead for a particular target molecule.
- the beads may be reacted with an amine-containing compound (e.g., a polymer or dendrimer) that includes multiple reactive groups.
- Ligands for ion exchange including but not limited to PDA, PEI-25K, PEI-800 daltons.
- the amine-containing compound is an organic ligand suitable for removing detergent from a mixture, such as a solution or suspension resulting from cell lysis.
- Exemplary detergents that can be removed using the disclosed magnetic beads include, but are not limited to, Tween TM (polyethylene glycol sorbitan monooleate), such as Tween TM 80 (polysorbate 80) and Tween TM 20 (polysorbate 20); Triton TM X-100 (t-Octylphenoxypolyethoxyethanol, Polyethylene glycol tert- octylphenyl ether); NP-40 (octylphenoxypolyethoxyethanol), DDM (n-Dodecyl- ⁇ -D- maltoside), SDS (sodium dodecyl sulfate), GDN (Glyco-diosgenin), or LMNG (Lauryl Maltose Neopentyl Glycol).
- Tween TM polyethylene glycol sorbitan monooleate
- Tween TM 80 polysorbate 80
- Tween TM 20 polysorbate 20
- the organic ligand may be a hydrophobic amine, for example, an alkylamine, such as a straight or branched C4-C20 alkyl amines, such as a straight or branched chain C 6 -C 18 alkylamine, for example, butylamine (C4), hexylamine (C6), octylamine (C8), decylamine (C10), dodecylamine (C12), tetradecylamine (C14), hexadecylamine (C16), octadecylamine (C18), eicosylamine (C20), or a combination thereof; a hydrophilic amine, such as O-(2-aminopropyl)-O’-(2- methoxyethyl)polypropylene glycol (Methoxy Jeffamine; from 3 to 25, 4,7,10-Trioxa-1,13 tridecanediamine (C10PEG diamine), ⁇ -cycl
- an amine containing moiety is a protein such as an antibody, an antibody binding protein (e.g., Protein A, Protein A/G, Protein G, and Protein L), Streptavidin, Neutravidin, glutathione, enzyme, or other biomolecule containing amines.
- azlactone beads are conjugated to an amine containing biomolecule such as an alkali stable Protein A (asPA) and the final conjugate can then be used to purify IgG. It is desired that when the biomolecule is an enzyme, it maintains its enzymatic selectivity once coupled to the bead.
- enzymes include proteolytic enzymes (e.g., trypsin) and other types of enzymes known in the art.
- FIG.12 shows similar performance with azlactone beads (asPA-magUL) and magnetic agarose beads (asPA-magAg).
- the assay was performed with duplicate samples of each magnetic bead.
- an amine-containing compounds is or includes metal charged chelates (e.g., nickel-charged nitrilotriacetic acid, Ni-NTA).
- azlactone beads are conjugated to Ni-NTA and the final conjugate are used to purify polyhistidine-tagged proteins from a soluble protein extract.
- the polymer bead is an azlactone bead formed from a vinyl azlactone and a crosslinker.
- the vinyl azlactone may be 4,4-dimethyl-2-vinyloxazol- 5(4H)-one, 4,4-Dimethyl-2-vinyloxazol-5(4H)-one
- the cross linker maybe any crosslinker suitable to facilitate polymerization and form the bead.
- the crosslinker is a bisacrylamide, agarose, or a vinyl ether.
- the crosslinker is methylene bisacrylamide.
- the vinyl azlactone is 4,4-dimethyl-2-vinyloxazol-5(4H)- one and the crosslinker is a bisacrylamide, such as methylene bisacrylamide.
- the beads comprise a plurality of functional moieties on the surface of the bead. The functional moieties may be selected from
- the azlactone bead is reacted with an organic ligand, such as an organic amine or alcohol. In some aspects, the azlactone bead is reacted with a single species of organic ligand, such as one hydrophobic ligand or one hydrophilic ligand. But in other aspects, the azlactone bead is reacted with multiple organic ligands, such as 2, 3, 4 or more hydrophobic ligands, 2, 3, or 4 or more hydrophilic ligands, or a combination of one or more, such as 1, 2, 3 or more, hydrophobic ligands and one or more, such as 1, 2, 3 or more, hydrophilic ligands.
- the azlactone bead is reacted with one hydrophobic ligand and one hydrophilic ligand.
- the ratio of hydrophobic to hydrophilic ligands is from 1% hydrophobic:99% hydrophilic to 99% hydrophobic:1% hydrophilic by weight, such as 10% hydrophobic:90% hydrophilic to 90% hydrophobic:10% hydrophilic, 20% hydrophobic:80% hydrophilic to 80% hydrophobic:20% hydrophilic, 30% hydrophobic:70% hydrophilic to 70% hydrophobic:30% hydrophilic, 40% hydrophobic:60% hydrophilic to 60% hydrophobic:40% hydrophilic, or about 50%:50% hydrophobic to hydrophilic by weight.
- the ratio was from 20%:80% hydrophobic:hydrophilic to 40%:60% hydrophobic:hydrophilic by weight.
- Exemplary functional moieties comprising hydrophobic ligands include, but are not limited
- exemplary functional moieties comprising hydrophilic ligands include, but are not limited to, where m is from 40-450 (MW of from about 2,000 to 20,000 DA),
- a plurality of magnetic beads contains beads having a single type of ligand, for example, just beads with hydrophobic ligands, just beads having hydrophilic ligands, or just beads that each contain a hydrophobic and a hydrophilic ligand.
- a plurality of beads comprises a first portion of beads that comprise a hydrophobic ligand and a second portion of beads that comprise a hydrophilic ligand.
- the first portion and second portion of the beads are each provided in amounts sufficient to provide a ratio of beads comprising hydrophobic ligands (referred to as hydrophobic beads) to beads comprising hydrophilic ligands (referred to as hydrophilic beads) of from 10% hydrophobic beads:90% hydrophilic beads to 90% hydrophobic beads:10% hydrophilic beads. And in certain aspects, the ratio of hydrophobic beads to hydrophilic beads is from 20%:80% hydrophobic beads:hydrophilic beads to 40%:60% hydrophobic beads:hydrophilic beads.
- the magnetic material in the magnetic beads may be in any suitable form, such as, but not limited to, a particle, powder, flake, cluster, bead, or combination thereof.
- the magnetic material is or comprises magnetic particles.
- the magnetic material in the magnetic beads may comprise an iron source.
- the iron source may be any suitable iron source, such as an iron source that is added during the polymer bead formation reaction.
- the iron source may be in the form of iron-containing particles and/or may be an iron oxide, such as Fe 3 O 4 , Fe 2 O 3 , or a combination thereof.
- the beads may include magnetic particles as the iron source, for example Dynabeads TM (available from Thermo Fisher Scientific) and/or Sera-Mag TM Speedbeads (available from Cytiva Life Sciences) and/or other magnetic beads or particles.
- the magnetic material is, or comprise, Fe3O4 particles.
- the magnetic material is an activated iron oxide source, such as Fe 3 O 4 -NHS, to provide an addition source of reactivity during particle synthesis.
- the magnetic material is an activated magnetic material and may contain one or more functional groups, such as, for example, a carboxy group or ester group.
- the magnetic material is an activated iron source that comprises additional functional groups, such as, but not limited to, carboxy or ester groups.
- the magnetic material is contained within an agarose bead, and the agarose bead is contained within the crosslinked polymer to form the magnetic bead.
- the magnetic particles may have a size suitable for use in the disclosed beads, such as from 5 nm to 1,000 nm or more, such as from 5 nm to 200 nm or from 100 nm to 800 nm. In certain aspects, the magnetic particles have a size of from 15 nm to 100 nm, such as from 15 nm to 50 nm, or from 50 nm to 100 nm.
- the magnetic particles have a size of 100 nm to 800 nm, for example, iron clusters that may have an average particle size of from 100 nm to 800 nm, such as from 100 nm to 600 nm, or about 100 nm, 200 nm, 300 nm, 400 nm, 500 nm or 600 nm.
- the average particle size of the magnetic particles may be determined using a particle size analyzer.
- the magnetic particle is substantially spherical, but in other aspects, the particle is not substantially spherical.
- the magnetic particle is a substantially spherical iron cluster. In certain other aspects, the magnetic particle is not an iron cluster and is not substantially spherical.
- the magnetic beads may be coated with another material that provides a substantially spherical shape.
- the magnetic beads have an average size suitable for use in separation technology.
- the average particle size is determined by a particle analyzer (e.g. Beckman Coulter TM LS13320XR) or by scanning electron microscopy (SEM).
- the magnetic beads are substantially spherical and the average particle size is the average diameter of the beads.
- the magnetic beads have an average size of from 20 ⁇ m to 100 ⁇ m or more, such as from 20 ⁇ m to 80 ⁇ m, from 30 ⁇ m to 60 ⁇ m, from 30 ⁇ m to 50 ⁇ m, or from 40 ⁇ m to 80 ⁇ m.
- the magnetic beads may be porous or non-porous.
- the magnetic beads are porous and may comprise pores having a size of from 100 Angstroms to 200 Angstroms. And in some aspects, the pores extend from the surface of the particle into the interior, but do not pass completely though the particle. In some aspects, at least a portion, and may be substantially all, of the magnetic material is located in the pores.
- the magnetic beads may have a hydrated state and a non-hydrated state. Typically, the disclosed bead size is the bead size in the hydrated state, and the bead size of the magnetic beads in the hydrated state (the hydrated bead size) is larger than the non-hydrated bead.
- magnetic beads have the tendency to swell, from 3 times larger to 10 times larger or more, from 5 times larger to 8 times larger, or from 5 times to 6 times larger.
- nonmagnetic beads typically swell to about 8x larger when hydrated, compared to their non-hydrated state.
- other magnetic beads do not swell to a measurable extent.
- the swelling of the disclosed magnetic beads may allow the functional moieties on the bead surface, such as azlactone moieties, to be more accessible, and/or allow large biomolecules to bind to the functional moieties more easily because steric problems due to their size are reduced due to the swelling increasing the space between the functional moieties.
- the disclosed magnetic beads demonstrate a reduced bead loss during use in an automated process, for example, a Kingfisher apparatus, when compared with other magnetic beads.
- the bead loss may be due, at least in part, to surface adsorption onto the apparatus, for example, onto the tip plate and sample plate.
- a population of the disclosed magnetic beads exhibits a bead loss of less than about 5% by weight when used in an automated process.
- Magnetic bead synthesis in a suitable solvent such as an alkane solvent, for example, heptane or hexanes, or a chlorinated solvent, such as carbon tetrachloride, chloroform, or dichloromethane.
- the monomer may be an azlactone monomer, such as a vinyl azlactone, for example, 4,4- dimethyl-2-vinyloxazol-5(4H)-one.
- the polymeric stabilizer may be any polymeric stabilizer suitable for use in the polymerization reaction.
- polymeric stabilizers include, but are not limited to, copoly(isooctylacrylate/acrylic acid), copoly(isobutylmethacrylate/acrylic acid) or copoly(hexylacrylate/sodium acrylate).
- the magnetic material is added in a suitable form, such as, but not limited to, a particle, powder, flake, cluster, bead, or combination thereof.
- the magnetic material comprises an iron source, such as iron oxide.
- iron oxide particles are used.
- the first solution is prepared and maintained at a temperature of from 35 °C to 50 °C, such as from 40 °C to 45 °C.
- a second solution is prepared comprising a crosslinker in a suitable solvent.
- the crosslinker is any crosslinker suitable to facilitate bead formation. Exemplary crosslinkers include, but are not limited to, an acrylamide, bisacrylamide, agarose, or a vinyl ether. In certain aspects, the crosslinker is methylene bisacrylamide.
- the solvent may be any solvent suitable to facilitate the polymerization reaction, such as an alcohol (for example, methanol, ethanol, 2-propanol, 1-propanol, or a combination thereof), water, or a combination thereof.
- a radical initiator for example, sodium persulfate
- the second solution is prepared and maintained at a temperature of from 25 °C to 40 °C, such as from 30 °C to 35 °C.
- the first solution and the second solution are combined and the mixture is agitated such as by stirring, shaking, swirling, etc, at a temperature suitable to facilitate polymerization and bead formation, followed by addition of a polymerization initiator, such as tetramethylethylenediamine (TEMED).
- TEMED tetramethylethylenediamine
- the temperature is from 30 °C to 50 °C.
- the beads are filtered and washed with a suitable solvent, such as acetone.
- the beads then are resuspended in a suitable solvent, such as acetone, and agitated, such as by shaking, stirring, swirling, or sonication, with cooling, such as in an ice bath.
- the beads are then screened to select beads of a desired size range.
- the disclosed magnetic beads are suitable for use with a wide range of biomolecules, including proteins, polypeptides, peptides, DNA, RNA, and combinations thereof. Additionally, the disclosed magnetic beads are suitable for use with isotopically labeled biomolecules.
- the isotopic labelling may include labeling with deuterium, carbon-13, nitrogen-15, oxygen-18, or a combination thereof.
- Sample is washed twice with 200 ⁇ L of 100% HPLC-grade MeCN (or 85% MeCN, 10% EtOH, 5.0% water, 0.5% formic acid for improved salt removal, such as in TMT- or TMTpro-labeled samples).
- d Beads are collected and the supernatant is aspirated to waste.
- e Sample is eluted with 50 ⁇ L of 2-4% MeCN, 0.2%FA in water for 2 minutes with occasional vortex.
- f. Place on the magnetic rack for 2 minutes, and transfer supernatant to a clean tube.
- Samples are centrifuged at 10,000xg for 30 seconds.
- h. Sample is moved to a new tube without disturbing pellet. 13.
- Example 8 Comparison Experiment To compare the disclosed magnetic beads with other, commercially available beads, a comparative workflow was developed using a KingFisher TM Sample Purification System from Thermo Fisher Scientific TM . Table 2 provides the beads that were used for the comparison experiment. Table 2. Beads used in the comparison experiment A 25 ⁇ g HeLa lysate sample was used for each bead type. The comparative workflow was as follows: A. Protein Digestion (Off KingFisher) • Reduction/alkylation: 95 °C for 10 minutes, cooled to room temperature, and centrifuged for seconds. • Digestion: 37 °C for 1 hour with shaking at 1,000 rpm.
- Trypsin/Lys-C 0.67 ⁇ g/ ⁇ L (100 ⁇ g of Trypsin/Lys-C reconstituted in 150 ⁇ L of 0.1% acetic acid/10 mM CaCl2 for use, sample to trypsin ratio: 10:1) Table 3.
- Sample/reagent volume for protein digestion B. Peptide Cleanup (On KingFisher) • Bead conditioning: Load 20 ⁇ L of beads (5.0 mg) with 80 ⁇ L of Bead Wash/Binding Solution. Wash the beads with 200 ⁇ L of Bead Wash/Binding Solution. • Dilution: Transfer beads into the diluted sample. The total volume is 500 ⁇ L (>9% organic phase after dilution).
- Binding Mix the sample/beads at room temperature for 45 minutes at a medium rate.
- Wash Transfer the sample/beads into 500 ⁇ L of Wash Solution for peptide wash. Mix the plate at room temperature for 2 minutes at a medium rate.
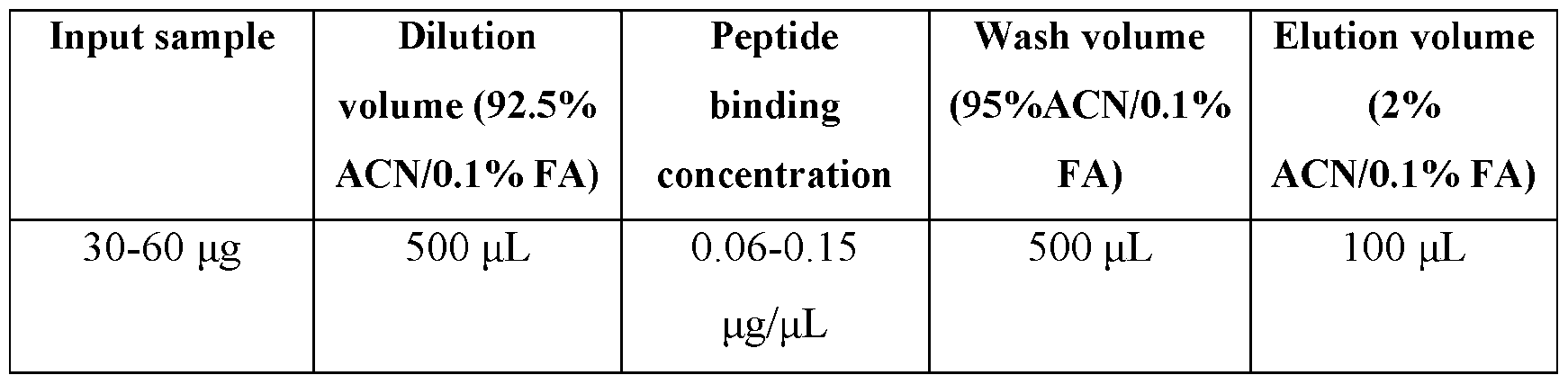
- Elution Transfer the sample/beads into 100 ⁇ L of Elution Solution for peptide elution. Mix the plate at room temperature for 5 minutes at a medium rate. Collect the output peptide samples for LC-MS analysis. Table 4. Peptide cleanup volumes Results from the experiment are provided in FIGS.8A, 8B, and 9-11.
- FIGS.8A and 8B demonstrate that the disclosed magnetic beads exhibit less surface absorption loss than the Sera-Mag TM carboxylate modified particles, MyOne TM , or silica beads in the SP2 workflow.
- the SeraMag TM , MyOne TM , and silica beads all demonstrated significant bead loss on the surfaces of the comb tip plate and sample plate (bead transfer/mix and binding steps).
- the disclosed magnetic beads did not exhibit significant bead loss.
- the data provided in FIGS.8A and 8B with respect to the disclosed magnetic beads refers to hydrolyzed beads, that is beads where the azlactone moieties on the surface of the beads have been hydrolyzed.
- FIG.9 provides data demonstrating that the peptide yield from the KingFisher SP2 workflow for the disclosed magnetic beads was substantially higher than the peptide yield for the Sera-Mag TM , MyOne TM Silane or silica beads, using a quantitative colorimetric peptide assay. No peptides were recovered using silica beads.
- FIG.10 demonstrates that using the disclosed magnetic beads, substantially more unique peptides were identified by nano-LCMS analysis, than were identified using the other bead types.
- FIG.11 demonstrates that substantially more protein groups were identified using the disclosed magnetic beads.
- Example 9 An experiment was designed to demonstrate the superior binding of the disclosed magnetic beads coupled to alkali stable Protein A (asPA), compared to a commercial magnetic bead that comprises asPA conjugates. Conjugate asPA to the disclosed magnetic beads Alkali stable Protein A was conjugated to the disclosed azlactone beads according to the following procedure: 1. Weigh out 77mg of the disclosed azlactone beads. 2.
- the amount of asPA bound to the disclosed azlactone magnetic beads is about 2-fold higher than the amount of asPA bound to the same amount of magnetic agarose beads.
- Example 10 Derivatization of Magnetic Beads with Ocylamine Ligands Various hydrophilic or hydrophobic molecules or proportional mixtures of hydrophilic and hydrophobic molecules were reacted to the amine-reactive, azlactone groups of magnetic beads to impart the support with detergent removal and peptide recovery properties.
- Scheme 1 shows the reaction of a representative hydrophobic ligand (octylamine, C8) to a magnetic bead bearing a terminal azlactone group.
- Scheme 1 A solution of ligand(s) in ethanol was prepared by dissolving 10mg/mL, 2mg/mL, or 40mg/mL of hydrophobic, hydrophilic or proportional mix of hydrophobic and hydrophilic ligand, respectively, in 8mL of ethanol with stirring. 250mg of magnetic base beads (swell volume : 1 gram of magnetic beads gives 6 mLs of beads) were added to the solution and allowed to react for 2 hours with end-over-end mixing. The reacted beads were then washed thoroughly twice with 1 bed volume of water, and then resuspended in water for later use.
- This general procedure was used to modify the magnetic beads with various types of hydrophobic ligands (i.e., octylamine (C8), dodecylamine (C12), octadecylamine (C18)) and hydrophilic ligands (i.e., methoxy Jeffamine (MJ), 4,7,10- Trioxa-1,13 tridecanediamine (C10PEG), B- cyclodextrin, dextran, and PEG molecules ranging in MW from 2000 Da to 20,000 Da), and combinations thereof.
- Table 5 lists different ligands that are reacted to the azlactone group of the magnetic beads.
- Example 11 Removal of Non-Ionic and Ionic Detergents by Azlactone Magnetic Beads Magnetic beads bearing azlactone groups were used to remove non-ionic and ionic detergents from biological samples (e.g., cell lysates and samples containing proteins and/or polypeptides. Examples of detergent removal workflows implementing various detection methods are provided below.
- Absorbance 280 protocol To determine removal of Triton TM X 100 and NP 40) 1. Pipette 200 ⁇ L of 25% slurry magnetic beads into Dynamag2 magnetic stand. 2. Remove supernate. 3. Pipette 100 ⁇ L of 1% detergent solution into bead sample. 4. Vortex detergent and beads until homogenous 5. Place solution in Dynamag2 magnetic stand and collect flow through. 6.
- Coomassie Protein Assay protocol To determine removal of detergents: 1. Pipette 200 ⁇ L of 25% slurry magnetic beads into Dynamag2 magnetic stand. 2. Remove supernate. 3. Pipette 100 ⁇ L of 1% detergent solution into bead sample. 4. Vortex detergent and beads until homogenous. 5. Place solution in Dynamag2 magnetic stand and collect flow through. 6. Pipette 10 ⁇ L of each sample and standard into a 96 well plate. 7. Pipette 300 ⁇ L of Coomassie Blue Stain Reagent into each occupied well. 8.
- Table 9 shows the % removal of detergents Triton TM X-100 and NP-40 at 1% concentration using hydrophilic ligands reacted to the beads.
- the hydrophilic ligands have poor capacity to trap detergents and as a result they were eliminated from further evaluation.
- Table 9 Detergent removal capacity of hydrophilic coated magnetic beads
- Example 14 Removal of Detergents from Beads Coupled to Blends of Hydrophobic and Hydrophilic Ligands
- Table 10 shows variations of octylamine ligand reacted beads blended with hydrophilic ligands such as O-(2- aminopropyl)-O’-(2-methoxyethyl)polypropylene glycol (Methoxy Jeffamine) and C10 PEG diamine in various blend ratios.
- Table 11 shows removal of three different non-ionic detergents 1% Triton TM X- 100, 1% NP-40 and 1% n-Dodecyl- ⁇ -D-maltoside (DDM). Beads with a mixture of hydrophobic and hydrophilic properties were found to remove a significant amount of detergent from the tested samples (e.g., 82%-100%). Table 11. Removal of Detergents from Beads with Hydrophobic and Hydrophilic Properties The samples also were evaluated for detergent removal from HeLa digests using the workflow described herein. Referring to Table 12, 40% or greater HeLa digest was recovered with C8 modified beads and its blends with hydrophilic ligands. Table 12.
- Table 17 provides the success rate of each bead compared to the untreated control sample.
- Table 17. Success Rate Example 18 MS analysis of detergent samples before and after bead processing Materials and Reagents PierceTM HeLa Protein Digest Standard (Thermo Scientific, Catalog number 88328) Instrument MS: Q Exactive HF mass spectrometer, Serial# SN05061L Method Dilute the detergent (n-Dodecyl- ⁇ -D-maltoside (DDM) and NP-40) samples before (control/reference sample) and after (samples) bead processing using 50% acetonitrile/0.1% formic acid with an appropriate ratio (e.g., 1:1000 – 1:100) for direct infusion MS analysis.
- DDM n-Dodecyl- ⁇ -D-maltoside
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Immunology (AREA)
- Life Sciences & Earth Sciences (AREA)
- Engineering & Computer Science (AREA)
- Analytical Chemistry (AREA)
- Urology & Nephrology (AREA)
- Molecular Biology (AREA)
- Biomedical Technology (AREA)
- Hematology (AREA)
- Cell Biology (AREA)
- Biotechnology (AREA)
- Microbiology (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Organic Chemistry (AREA)
- Food Science & Technology (AREA)
- Medicinal Chemistry (AREA)
- Physics & Mathematics (AREA)
- Biochemistry (AREA)
- General Health & Medical Sciences (AREA)
- General Physics & Mathematics (AREA)
- Pathology (AREA)
- Solid-Sorbent Or Filter-Aiding Compositions (AREA)
Abstract
Disclosed herein are aspects of magnetic beads that include functional groups suitable for removing detergent from liquids. The magnetic beads may comprise a vinyl azlactone-based crosslinked polymer and a magnetic material dispersed in the bead. The magnetic beads may comprise hydrophobic and/or hydrophilic ligands suitable to facilitate removal of the detergents from solutions or suspensions, while allowing for good recovery of any biomolecules of interest in the sample. Also disclosed are aspects of a method for making the beads, aspects of a method for using the beads, and aspects of a kit comprising the beads.
Description
MAGNETIC BEADS FOR DETERGENT REMOVAL CROSS REFERENCE TO RELATED APPLICATION This application claims the benefit of the earlier filing date of U.S. provisional patent application No.63/517,029, filed August 1, 2023, which is incorporated herein by reference in its entirety. FIELD This disclosure concerns magnetic beads that contain ligands suitable for removing detergent(s) from mixtures, including mixtures that comprise biological molecules including polypeptides, proteins, RNA, and/or DNA, and methods for making the beads and using the beads. BACKGROUND Mass spectrometry (MS)-based proteomics has become the most comprehensive approach for protein identification, quantitation, interactions, modifications, and structural characterization. It is also a challenging topic that requires expertise for sample preparation since a high-quality sample from a robust and reproducible sample preparation is critical for success. Proteomics sample preparation is highly complex and variable, with many different protocols available. For example, bottom-up proteomics sample preparation requires multiple steps including protein extraction, reduction and alkylation of cysteines, digestion of the proteins into peptides, cleanup, and concentration of the peptides for LC-MS analysis. The lack of standardization in proteomic sample preparation makes it difficult to accurately compare results from different laboratories and different protocols. SUMMARY Detergents, including non-ionic and ionic detergents, are frequently used in mass spectrometry (mass spec; MS) sample preparation. However, it is often necessary
to remove the detergents before injecting the sample into the MS machine. With new instruments becoming increasing sensitive, it is necessary to remove 90% or more, and may be 99%, 99.9%, or 99.99% or more of these detergents in order to observe the molecule(s) of interest, such as polypeptides, or proteins. Therefore, there is a need for new methods to remove detergents from samples, while allowing a good recovery, such as greater than 30%, greater than 40%, or greater than 50%, of the molecule(s) of interest, and without reacting with or otherwise changing the nature of the molecule(s) of interest. Disclosed herein is a method for removing detergent from a solution using magnetic beads. The method comprises forming a suspension comprising a first plurality of magnetic beads and a first liquid that contains a first amount of a detergent; agitating the suspension; and removing the magnetic beads from the suspension to leave a solution comprising a second amount of the detergent that is less than the first amount of the detergent. The second amount of the detergent may be less than 0.1% of the second solution, such as less than 0.01% of the second solution, and may be substantially zero. Each magnetic bead may comprise a crosslinked polymer, a magnetic material, and a plurality of hydrophobic ligands formed from a reaction between azlactone moieties
one or more hydrophobic molecules. The hydrophobic molecules may comprise a straight or branched chain C4-C20alkyl amine, for example, butylamine (C4), hexylamine (C6), octylamine (C8), decylamine (C10), dodecylamine (C12), tetradecylamine (C14), hexadecylamine (C16), octadecylamine (C18), eicosylamine (C20), or a combination thereof. In some aspects, the straight or branched C4-C20 alkyl amine is selected from octylamine (C8), dodecylamine (C12), octadecylamine (C18), or a combination thereof. In particular aspects, the hydrophobic molecule is octylamine and the hydrophobic ligand is
.
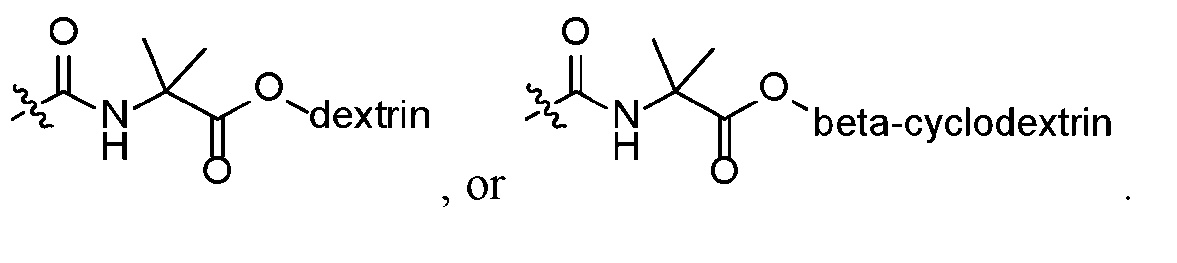
The magnetic bead may further comprise a hydrophilic ligand. The hydrophilic ligand may be formed by a reaction between azlactone moieties and one or more hydrophilic molecules. The one or more hydrophilic molecules may be selected from O-(2- aminopropyl)-O’-(2-methoxyethyl)polypropylene glycol, 4,7,10-Trioxa-1,13 tridecanediamine, β-cyclodextrin, dextran, PEG molecules ranging in MW from 2000 Da to 20,000 Da, or a combination thereof. In some aspects, the hydrophilic ligand is
. In a particular aspects, the magnetic bead comprises
as the hydrophilic ligand. In some aspects where the magnetic bead comprises a hydrophobic ligand and a hydrophilic ligand, the two types of ligand are present in a ratio of from 10% hydrophobic:90% hydrophilic to 90% hydrophobic:10% hydrophilic, such as from 20%:80% hydrophobic:hydrophilic to 40%:60% hydrophobic:hydrophilic. The suspension may further comprise a second plurality of magnetic beads where each magnetic bead in the second plurality of magnetic beads comprises a crosslinked polymer, a magnetic material, and a plurality of hydrophilic ligands formed from a reaction between azlactone moieties and one or more hydrophilic molecules. The first plurality of magnetic beads and the second plurality of magnetic beads may be
each provided in amounts sufficient to provide a ratio of beads comprising hydrophobic ligands (referred to as hydrophobic beads) to beads comprising hydrophilic ligands (referred to as hydrophilic beads) of from 10% hydrophobic beads:90% hydrophilic beads to 90% hydrophobic beads:10% hydrophilic beads, such as from 20%:80% hydrophobic beads:hydrophilic beads to 40%:60% hydrophobic beads:hydrophilic beads. In any aspects, the detergent may be selected from polyethylene glycol sorbitan monooleate, t-Octylphenoxypolyethoxyethanol, octylphenoxypolyethoxyethanol (NP- 40) , n-Dodecyl-β-D-maltoside (DDM), sodium dodecyl sulfate (SDS), Glyco-diosgenin (GDN), Lauryl Maltose Neopentyl Glycol (LMNG), or a combination thereof. Also disclosed herein are aspects of a magnetic bead comprising a crosslinked polymer bead and a magnetic material contained within the polymer bead, and further comprising a plurality of ligands wherein the ligands are selected from: hydrophobic ligands formed from a reaction between azlactone moieties and one or more hydrophobic molecules; hydrophilic ligands formed from a reaction between azlactone moieties and one or more hydrophilic molecules; or a combination of hydrophobic ligands and hydrophilic ligands formed from reactions between azlactone moieties and one or more hydrophobic molecules and one or more hydrophilic molecules. The crosslinked polymer may be a crosslinked azlactone polymer, and the magnetic bead may be an azlactone bead. The one or more hydrophobic molecules may comprise a straight or branched chain C4-C20alkyl amine as disclosed herein. And/or the hydrophilic ligand may be derived from O-(2-aminopropyl)-O’-(2- methoxyethyl)polypropylene glycol, 4,7,10-Trioxa-1,13 tridecanediamine, β- cyclodextrin, dextran, a PEG molecule having a molecular weight of from 2,000 to 20,000 Da, or a combination thereof. The magnetic material may comprise an iron source, such as iron oxide. In some aspects, the magnetic material is in the form of a particle, powder, flake, or cluster, and/or may be contained within an agarose bead which in turn may be contained within the crosslinked polymer.
In any aspects, the magnetic bead may have a bead size of from 20 to 80 microns, such as from 30 to 60 microns. Also disclosed herein are aspects of a kit comprising the magnetic beads, and aspects of a method for making the magnetic beads. The foregoing and other objects, features, and advantages of the disclosure will become more apparent from the following detailed description, which proceeds with reference to the accompanying figures. BRIEF DESCRIPTION OF THE DRAWINGS FIG.1 is a flow chart illustrating one aspect of a standard workflow using the disclosed magnetic beads. FIG.2 is a flowchart illustrating an SP3 workflow that includes a PAC workflow and an SP2 workflow. FIG.3 is a digital image illustrating one aspect of the disclosed magnetic beads. FIG.4 is a graph of differential volume versus size, illustrating the average size of the disclosed magnetic beads, and a table providing the average size and standard deviation. FIG.5 is a digital image illustrating one aspect of the disclosed magnetic beads synthesized using iron oxide clusters as the magnetic material. FIG.6 is a digital image illustrating one aspect of the disclosed magnetic beads synthesized using iron oxide particles as the magnetic material. FIG.7 is a digital image illustrating one aspect of the disclosed magnetic beads synthesized using DynabeadsTM as the magnetic material. FIGS.8A and 8B are a digital image of plates from various stages of an SP2 workflow (FIG.8A) and a table of results (FIG.8B), illustrating the surface absorption losses of different bead types. FIG.9 is a graph of peptide yield versus bead type, illustrating the peptide yields from different bead types, identified using a quantitative colorimetric peptide assay. FIG.10 is a graph of number of peptides versus bead types, illustrating the number of unique peptides isolated with different bead types and identified using
nanoLC-MS analysis, 75 i.d. column, 150 min gradient, QE plus mass spectrometry, PD 2.5 data. FIG.11 is a graph of protein group versus bead types, illustrating the number of different protein groups isolated with different bead types and identified using nanoLC- MS analysis, 75 i.d. column, 150 min gradient, QE plus mass spectrometry, PD 2.5 data. FIG.12 is a graph of bound antibody versus magnetic bead, illustrating the capacity of the disclosed magnetic beads conjugated to alkali stable Protein A (asPA) to bind rabbit IgG. FIG.13 is a graph of number versus peptides and peptide groups, illustrating the numbers of unique peptides and peptide groups identified from a HeLa protein digest after processing with an exemplary magnetic bead comprising octylamine at 10 mg/mL. FIG.14 is a graph of number versus peptides and peptide groups, illustrating the numbers of unique peptides and peptide groups identified from a HeLa protein digest after processing with an exemplary magnetic bead comprising octylamine at 20 mg/mL DETAILED DESCRIPTION I. Terms The following explanations of terms and methods are provided to better describe the present disclosure and to guide those of ordinary skill in the art in the practice of the present disclosure. The singular forms “a,” “an,” and “the” refer to one or more than one, unless the context clearly dictates otherwise. The term “or” refers to a single element of stated alternative elements or a combination of two or more elements, unless the context clearly indicates otherwise. As used herein, “comprises” means “includes.” Thus, “comprising A or B,” means “including A, B, or A and B,” without excluding additional elements. All references, including patents and patent applications cited herein, are incorporated by reference in their entirety, unless otherwise specified. Unless otherwise indicated, all numbers expressing quantities of components, molecular weights, percentages, temperatures, times, and so forth, as used in the specification or claims, are to be understood as being modified by the term “about.”
Accordingly, unless otherwise indicated, implicitly or explicitly, the numerical parameters set forth are approximations that may depend on the desired properties sought and/or limits of detection under standard test conditions/methods. When directly and explicitly distinguishing embodiments from discussed prior art, the embodiment numbers are not approximates unless the word “about” is expressly recited. Unless explained otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this disclosure pertains. Although methods and materials similar or equivalent to those described herein can be used in the practice or testing of the present disclosure, suitable methods and materials are described below. The materials, methods, and examples are illustrative only and not intended to be limiting. “Alkyl” refers to a saturated aliphatic hydrocarbyl group having from 1 to 25 (C1-25) or more carbon atoms, more typically 1 to 10 (C1-10) carbon atoms such as 1 to 6 (C1-6) carbon atoms or 1 to 4 (C1-4) carbon atoms. “Carboxy” refers to a -CO2H functional group. “Ester” refers to a -CO2R functional group, where R is an alkyl group, such as a C1-6 alkyl group. “Nucleic acid” refers to a polynucleotide molecule. The polynucleotide may be a naturally occurring polynucleotide or a synthetic polynucleotide. A nucleic acid may be a DNA, RNA or mixture of DNA and RNA nucleotides. Typically, a nucleic acid contains from 20 to 10,000 nucleotides or more, such as from 100, 200, 300, 400, 500, 600, 700, 800, 900, 1000, 2000, 3000, 4000, or 5000 nucleotides to 10,000 nucleotides. “Peptide” refers to a compound comprising amino acid residues connected by peptide bonds. Typically, a peptide compound has from 2 to about 50 amino acid residues. “Polypeptide” refers to a compound comprising amino acid residues connected by peptide bonds. In some aspects, a polypeptide has from about 50 amino acid residues to 2000 or more amino acid residues. The amino acids may include alpha-amino acids, and may comprise the L-optical isomer, the D-optical isomer, or combinations thereof.
“Protein” refers to a molecule or complex comprising one or more polypeptides having secondary, tertiary and/or quaternary structure. The secondary, tertiary and/or quaternary structure of a protein typically is stabilized using non-covalent bonds, such as hydrogen bonds, ionic bonds, hydrophobic interactions, and/or van der Walls interactions, and/or covalent bonds, for example, disulfide bonds, such as between the thiol groups of cysteine residues. II. Magnetic Beads Disclosed herein are aspects of a magnetic bead that comprises magnetic material dispersed in a polymer bead. In some aspects, the polymer bead comprises a crosslinked polymer that comprises one or more azlactone moieties
r one or more functional groups derived from an azlactone moiety. Functional groups derived from an azlactone moiety may comprise moieties derived from a reaction between the azlactone moiety and a nucleophilic reactive group, such as, but not limited to, an amine-containing compound, thiol containing group, alcohol or water. Hydrolysis of azlactone or reaction of the azlactone group with an amine- containing compound can add functionality and/or alter the surface properties of the bead. For example, hydrolysis of azlactone rings may produce a bead having a plurality of carboxylic acid functional groups on its surface. Alternatively, the azlactone group are reacted with an amine-containing compound to increase or decrease the hydrophobicity or hydrophilicity of the bead. In certain aspects, azlactone beads are reacted with an amine-containing compound that includes primary, secondary, or tertiary amine group(s) and can, optionally, further include one or more additional functional groups. In other aspects, azlactone beads are reacted with a thiol or alcohol containing compound resulting in thioamide and ester functionalized beads. In other aspects, the azlactone bead are reacted with an amine-containing compound to provide a linker that provides one or more additional functional groups for
further reaction with a ligand or biomolecule. For example, an amine-containing compound that includes an additional amine reactive group may be reacted with the azlactone bead, where the reactive amine group can be converted to an epoxide, maleimide, or iodoacetyl group using standard methods known in the art. In yet other aspects, azlactone beads are reacted with a compound to expand the number of functional groups attached to the bead. Such an approach can be an effective approach for increasing the binding capacity of the bead for a particular target molecule. For example, the beads may be reacted with an amine-containing compound (e.g., a polymer or dendrimer) that includes multiple reactive groups. In other aspects, the amine-containing compound are reacted with an azlactone group on the bead to provide a spacer to overcome steric hindrance upon ligand attachment or to provide a linker for subsequent reaction with a complementary reactive group using compounds and methods. Amine-containing compounds also include biomolecules (e.g., proteins, peptides, nucleic acids, nucleotides, and oligo- and polysaccharide), organic ligands, and large synthetic compounds, such as polymers (e.g., PEG) and dendrimers. Representative amine-containing organic ligands include, without limitation, alkylamines (e.g., straight or branched C4-C12 alkyl amines such as 1,5-diaminopentane, diethylamine, butylamine, and octylamine), arylamines (e.g., benzylamine and 4- aminobenzoic acid), aminocaproic acid, ethanolamine, 5-amino-2-methyl benzene sulfonic acid, aminoethyl-trimethylammonium chloride (AETMA), taurine, tris(hydroxymethyl)aminomethane (Tris), and the like. Ligands for ion exchange (cationic and anionic) including but not limited to PDA, PEI-25K, PEI-800 daltons. In certain aspects, the amine-containing compound is an organic ligand suitable for removing detergent from a mixture, such as a solution or suspension resulting from cell lysis. Exemplary detergents that can be removed using the disclosed magnetic beads include, but are not limited to, TweenTM (polyethylene glycol sorbitan monooleate), such as TweenTM 80 (polysorbate 80) and TweenTM 20 (polysorbate 20); TritonTM X-100 (t-Octylphenoxypolyethoxyethanol, Polyethylene glycol tert- octylphenyl ether); NP-40 (octylphenoxypolyethoxyethanol), DDM (n-Dodecyl-β-D-
maltoside), SDS (sodium dodecyl sulfate), GDN (Glyco-diosgenin), or LMNG (Lauryl Maltose Neopentyl Glycol). The organic ligand may be a hydrophobic amine, for example, an alkylamine, such as a straight or branched C4-C20 alkyl amines, such as a straight or branched chain C6-C18 alkylamine, for example, butylamine (C4), hexylamine (C6), octylamine (C8), decylamine (C10), dodecylamine (C12), tetradecylamine (C14), hexadecylamine (C16), octadecylamine (C18), eicosylamine (C20), or a combination thereof; a hydrophilic amine, such as O-(2-aminopropyl)-O’-(2- methoxyethyl)polypropylene glycol (Methoxy Jeffamine;
from 3 to 25, 4,7,10-Trioxa-1,13 tridecanediamine (C10PEG diamine), β-cyclodextrin, dextran, or PEG molecules ranging in MW from 2000 Da to 20,000 Da (as measured by proton NMR), or a combination thereof; or a combination thereof. In some aspects, an amine containing moiety is a protein such as an antibody, an antibody binding protein (e.g., Protein A, Protein A/G, Protein G, and Protein L), Streptavidin, Neutravidin, glutathione, enzyme, or other biomolecule containing amines. In one aspect, azlactone beads are conjugated to an amine containing biomolecule such as an alkali stable Protein A (asPA) and the final conjugate can then be used to purify IgG. It is desired that when the biomolecule is an enzyme, it maintains its enzymatic selectivity once coupled to the bead. Examples of enzymes include proteolytic enzymes (e.g., trypsin) and other types of enzymes known in the art. FIG.12 shows similar performance with azlactone beads (asPA-magUL) and magnetic agarose beads (asPA-magAg). The assay was performed with duplicate samples of each magnetic bead. In some aspects, an amine-containing compounds is or includes metal charged chelates (e.g., nickel-charged nitrilotriacetic acid, Ni-NTA). In one aspect, azlactone beads are conjugated to Ni-NTA and the final conjugate are used to purify polyhistidine-tagged proteins from a soluble protein extract.
In some aspects, the polymer bead is an azlactone bead formed from a vinyl azlactone and a crosslinker. The vinyl azlactone may be 4,4-dimethyl-2-vinyloxazol- 5(4H)-one,
4,4-Dimethyl-2-vinyloxazol-5(4H)-one The cross linker maybe any crosslinker suitable to facilitate polymerization and form the bead. In some aspects, the crosslinker is a bisacrylamide, agarose, or a vinyl ether. In certain aspects, the crosslinker is methylene bisacrylamide. In particular aspects, the vinyl azlactone is 4,4-dimethyl-2-vinyloxazol-5(4H)- one and the crosslinker is a bisacrylamide, such as methylene bisacrylamide. In some aspects, the beads comprise a plurality of functional moieties on the surface of the bead. The functional moieties may be selected from
.
In certain aspects, the azlactone bead is reacted with an organic ligand, such as an organic amine or alcohol. In some aspects, the azlactone bead is reacted with a single species of organic ligand, such as one hydrophobic ligand or one hydrophilic ligand. But in other aspects, the azlactone bead is reacted with multiple organic ligands, such as 2, 3, 4 or more hydrophobic ligands, 2, 3, or 4 or more hydrophilic ligands, or a combination of one or more, such as 1, 2, 3 or more, hydrophobic ligands and one or more, such as 1, 2, 3 or more, hydrophilic ligands. In certain aspects, the azlactone bead is reacted with one hydrophobic ligand and one hydrophilic ligand. In some aspects comprising a combination of hydrophobic and hydrophilic ligands, the ratio of hydrophobic to hydrophilic ligands is from 1% hydrophobic:99% hydrophilic to 99% hydrophobic:1% hydrophilic by weight, such as 10% hydrophobic:90% hydrophilic to 90% hydrophobic:10% hydrophilic, 20% hydrophobic:80% hydrophilic to 80% hydrophobic:20% hydrophilic, 30% hydrophobic:70% hydrophilic to 70% hydrophobic:30% hydrophilic, 40% hydrophobic:60% hydrophilic to 60% hydrophobic:40% hydrophilic, or about 50%:50% hydrophobic to hydrophilic by weight. In some aspects, the ratio was from 20%:80% hydrophobic:hydrophilic to 40%:60% hydrophobic:hydrophilic by weight. Exemplary functional moieties comprising hydrophobic ligands include, but are not limited to,
Exemplary functional moieties comprising hydrophilic ligands include, but are not limited to,
where m is from 40-450 (MW of from about 2,000 to 20,000 DA),
In some aspects, a plurality of magnetic beads contains beads having a single type of ligand, for example, just beads with hydrophobic ligands, just beads having
hydrophilic ligands, or just beads that each contain a hydrophobic and a hydrophilic ligand. However, in other aspects, a plurality of beads comprises a first portion of beads that comprise a hydrophobic ligand and a second portion of beads that comprise a hydrophilic ligand. In such aspects comprising a mixture of beads, the first portion and second portion of the beads are each provided in amounts sufficient to provide a ratio of beads comprising hydrophobic ligands (referred to as hydrophobic beads) to beads comprising hydrophilic ligands (referred to as hydrophilic beads) of from 10% hydrophobic beads:90% hydrophilic beads to 90% hydrophobic beads:10% hydrophilic beads. And in certain aspects, the ratio of hydrophobic beads to hydrophilic beads is from 20%:80% hydrophobic beads:hydrophilic beads to 40%:60% hydrophobic beads:hydrophilic beads. In any aspects, the magnetic material in the magnetic beads may be in any suitable form, such as, but not limited to, a particle, powder, flake, cluster, bead, or combination thereof. In some embodiments, the magnetic material is or comprises magnetic particles. In any aspects, the magnetic material in the magnetic beads may comprise an iron source. The iron source may be any suitable iron source, such as an iron source that is added during the polymer bead formation reaction. The iron source may be in the form of iron-containing particles and/or may be an iron oxide, such as Fe3O4, Fe2O3, or a combination thereof. Additionally, or alternatively, the beads may include magnetic particles as the iron source, for example DynabeadsTM (available from Thermo Fisher Scientific) and/or Sera-MagTM Speedbeads (available from Cytiva Life Sciences) and/or other magnetic beads or particles. In certain aspects, the magnetic material is, or comprise, Fe3O4 particles. In certain aspects, the magnetic material is an activated iron oxide source, such as Fe3O4-NHS, to provide an addition source of reactivity during particle synthesis. In some aspects, the magnetic material is an activated magnetic material and may contain one or more functional groups, such as, for example, a carboxy group or ester group. In certain aspects, the magnetic material is an activated iron source that
comprises additional functional groups, such as, but not limited to, carboxy or ester groups. In some aspects, the magnetic material is contained within an agarose bead, and the agarose bead is contained within the crosslinked polymer to form the magnetic bead. The magnetic particles may have a size suitable for use in the disclosed beads, such as from 5 nm to 1,000 nm or more, such as from 5 nm to 200 nm or from 100 nm to 800 nm. In certain aspects, the magnetic particles have a size of from 15 nm to 100 nm, such as from 15 nm to 50 nm, or from 50 nm to 100 nm. In certain other aspects, the magnetic particles have a size of 100 nm to 800 nm, for example, iron clusters that may have an average particle size of from 100 nm to 800 nm, such as from 100 nm to 600 nm, or about 100 nm, 200 nm, 300 nm, 400 nm, 500 nm or 600 nm. The average particle size of the magnetic particles may be determined using a particle size analyzer. In some aspects, the magnetic particle is substantially spherical, but in other aspects, the particle is not substantially spherical. In certain aspects, the magnetic particle is a substantially spherical iron cluster. In certain other aspects, the magnetic particle is not an iron cluster and is not substantially spherical. However, such magnetic particles may be coated with another material that provides a substantially spherical shape. The magnetic beads have an average size suitable for use in separation technology. The average particle size is determined by a particle analyzer (e.g. Beckman CoulterTM LS13320XR) or by scanning electron microscopy (SEM). In some aspects the magnetic beads are substantially spherical and the average particle size is the average diameter of the beads. In some aspects, the magnetic beads have an average size of from 20 μm to 100 μm or more, such as from 20 μm to 80 μm, from 30 μm to 60 μm, from 30 μm to 50 μm, or from 40 μm to 80 μm. The magnetic beads may be porous or non-porous. In some aspects, the magnetic beads are porous and may comprise pores having a size of from 100 Angstroms to 200 Angstroms. And in some aspects, the pores extend from the surface of the particle into the interior, but do not pass completely though the particle. In some
aspects, at least a portion, and may be substantially all, of the magnetic material is located in the pores. The magnetic beads may have a hydrated state and a non-hydrated state. Typically, the disclosed bead size is the bead size in the hydrated state, and the bead size of the magnetic beads in the hydrated state (the hydrated bead size) is larger than the non-hydrated bead. These magnetic beads have the tendency to swell, from 3 times larger to 10 times larger or more, from 5 times larger to 8 times larger, or from 5 times to 6 times larger. By way of comparison, nonmagnetic beads typically swell to about 8x larger when hydrated, compared to their non-hydrated state. And to the inventors’ knowledge, other magnetic beads do not swell to a measurable extent. The swelling of the disclosed magnetic beads may allow the functional moieties on the bead surface, such as azlactone moieties, to be more accessible, and/or allow large biomolecules to bind to the functional moieties more easily because steric problems due to their size are reduced due to the swelling increasing the space between the functional moieties. In some aspects, the disclosed magnetic beads demonstrate a reduced bead loss during use in an automated process, for example, a Kingfisher apparatus, when compared with other magnetic beads. The bead loss may be due, at least in part, to surface adsorption onto the apparatus, for example, onto the tip plate and sample plate. In some aspects, a population of the disclosed magnetic beads exhibits a bead loss of less than about 5% by weight when used in an automated process. In a comparative experiment, the disclosed magnetic beads demonstrated a bead loss of 4.7% by weight, where as Cytiva Sera-MagTM beads had a loss of 6.0% by weight, InvitrogenTM DynaGreenTM silica beads lost 11.5% by weight and DynabeadsTM MyOneTM Silane beads lost 6.8% by weight.
III. Synthesis of the Magnetic Beads A. Magnetic bead synthesis
in a suitable solvent, such as an alkane solvent, for example, heptane or hexanes, or a chlorinated solvent, such as carbon tetrachloride, chloroform, or dichloromethane. The monomer may be an azlactone monomer, such as a vinyl azlactone, for example, 4,4- dimethyl-2-vinyloxazol-5(4H)-one. The polymeric stabilizer may be any polymeric stabilizer suitable for use in the polymerization reaction. Example, polymeric stabilizers include, but are not limited to, copoly(isooctylacrylate/acrylic acid), copoly(isobutylmethacrylate/acrylic acid) or copoly(hexylacrylate/sodium acrylate). After agitation, the magnetic material is added in a suitable form, such as, but not limited to, a particle, powder, flake, cluster, bead, or combination thereof. In some aspects, the magnetic material comprises an iron source, such as iron oxide. In certain aspects, iron oxide particles are used. And in some aspects, the first solution is prepared and maintained at a temperature of from 35 °C to 50 °C, such as from 40 °C to 45 °C. A second solution is prepared comprising a crosslinker in a suitable solvent. The crosslinker is any crosslinker suitable to facilitate bead formation. Exemplary crosslinkers include, but are not limited to, an acrylamide, bisacrylamide, agarose, or a vinyl ether. In certain aspects, the crosslinker is methylene bisacrylamide. The solvent may be any solvent suitable to facilitate the polymerization reaction, such as an alcohol (for example, methanol, ethanol, 2-propanol, 1-propanol, or a combination thereof), water, or a combination thereof. In some aspects, a radical initiator, for example, sodium persulfate, is added to the second solution. And in some aspects, the second solution is prepared and maintained at a temperature of from 25 °C to 40 °C, such as from 30 °C to 35 °C.
The first solution and the second solution are combined and the mixture is agitated such as by stirring, shaking, swirling, etc, at a temperature suitable to facilitate polymerization and bead formation, followed by addition of a polymerization initiator, such as tetramethylethylenediamine (TEMED). In some aspects, the temperature is from 30 °C to 50 °C. After cooling to room temperature, the beads are filtered and washed with a suitable solvent, such as acetone. The beads then are resuspended in a suitable solvent, such as acetone, and agitated, such as by shaking, stirring, swirling, or sonication, with cooling, such as in an ice bath. The beads are then screened to select beads of a desired size range. Screening the beads may comprise using one or more sieves, such as two sieves, to select beads having the desired size or size range. In some aspects, a 63 μm sieve and a 25 μm sieve are used to select a population of magnetic beads having a size range of from 63-25 μm. The magnetic beads then are placed in a magnetic separate system to remove any non-magnetic residue. After filtering, the beads are dried, for example, under a vacuum and/or with warming at from 30 °C to 60 °C Before use, the beads are typically swelled in an aqueous liquid, such as water, a buffer solution, or an aqueous/organic solvent mixture. Additional information concerning synthesizing crosslinked beads can be found in United States Patent Nos.5,403,902 and 5,292,840, which are incorporated herein by reference in their entirety. B. Ligand attachment for detergent removal beads A solution of the desired ligand(s) is prepared in a suitable solvent, such as an alcohol (for example, methanol, ethanol or isopropanol). The ligand(s) are typically provided as an amine- or hydroxyl-containing molecule. The disclosed magnetic beads comprising azlactone functional groups on the surface are added to the solution and the mixture is allowed to react for a time period suitable to facilitate addition of the ligands to the bead. The time period may be from 1 hour or less to 6 hours or more, such as
from 1 hour to 4 hours, or from 1.5 hours to 2.5 hours. The mixture may be agitated during the reaction, such as by shaking, stirring, or end-over-end mixing. After the reaction time, the beads are separated from the reaction mixture, such as by magnetic separation or filtration, and then washed with water before being resuspended in water for use or storage. IV. Kit Comprising the Magnetic Beads Also disclosed herein are aspects of a kit comprising the disclosed magnetic beads. The components of the kit may depend on the intended use. In some aspects, the kit comprises the magnetic beads and a wash solution. In some aspects, the kit comprises, in addition to the magnetic beads, a lysis solution, universal nuclease, reduction solution, alkylation solution, enzyme reconstitution solution, Trypsin/Lys-C protease mix, bead wash/binding solution, wash solution, elution solution, or any combination thereof. In certain aspects, a kit comprises the magnetic beads, a lysis solution, a universal nuclease, a reduction solution, an alkylation solution, an enzyme reconstitution solution, and a Trypsin/Lys-C protease mix. In other aspects, a kit comprises the magnetic beads, a bead wash/binding solution, a wash solution, and an elution solution. V. Biomolecules and Reagents Successful mass spectrometry analysis of peptides requires high-quality input samples. Sample preparation prior to mass spectrometry (MS) often requires the introduction of chemicals which are incompatible with downstream MS analysis and must be removed from the sample prior to LC-MS analysis. However, complete cleanup of salts, detergents, and other chemical and non-chemical contaminants which simultaneously retains all peptide species and demonstrates reasonable recovery is very difficult, often requiring special equipment, highly complex processes, and/or large time commitments.
The disclosed magnetic beads may be used to simplify and accelerate sample cleanup while improving sample quality in terms of cleanliness and introducing amenability to automation. The beads also are compatible with plastics and solvents that are frequently used in biological assay. These features make the beads amenable to downstream applications prior to MS, and also other applications, such as chemical labeling, enrichment, and fractionation. The disclosed magnetic beads are suitable for use with a wide range of biomolecules, including proteins, polypeptides, peptides, DNA, RNA, and combinations thereof. Additionally, the disclosed magnetic beads are suitable for use with isotopically labeled biomolecules. The isotopic labelling may include labeling with deuterium, carbon-13, nitrogen-15, oxygen-18, or a combination thereof. Additionally, or alternatively, the biomolecules may be labelled with an isotopically-enriched mass tag, such as, but not limited to, Tandem Mass Tag (TMT and TMTpro) Reagents (available from Thermo Fisher ScientificTM); stable isotopically labeled peptides and/or proteins prepared using SILAC Metabolic Labeling Systems (available Thermo Fisher ScientificTM); rare-earth metals (e.g., lanthanide); fluorescent labels (e.g., dyes); tissues and/or cells grown with heavy amino acids; or a combination thereof. Disclosed beads comprising hydrophobic and/or hydrophilic ligands are useful for removing detergent from samples, for example, after lysing cells. The disclosed beads can remove 99% or more of the detergent from a sample, such as more than 99.5%, more than 99.9% or 99.99% or more of the detergent from the sample, while leaving at least 30% of the biomolecules, such as at least 40% or at least 50% of the biomolecules, for example, proteins, polypeptides, DNA molecules, RNA molecules etc., behind in the sample and substantially unchanged by the addition and removal of the magnetic beads. Such biomolecules can later be separated, if necessary, by using an alternative aspect of the magnetic beads disclosed herein that includes functional group(s) selected to bind to specific biomolecules. Such methods also are disclosed herein.
In some aspects, the disclosed magnetic beads are useful in multi- analyte analysis applications because of their distinct binding profiles to different types of biomolecules. For example, the disclosed magnetic beads typically bind DNA at low levels of organic solvent, such as about 30% organic, and are not affected by the presence of guanidine. In contrast, the disclosed beads bind to RNA at higher levels of organic solvent, such as about 90% organic. And the disclosed magnetic beads can bind to proteins in a solvent up to 45% organic in the presence of guanidine hydrochloride, or at about 80% without guanidine. These combinations of guanidine and organic presence can be utilized uniquely in the disclosed magnetic beads for sequential extraction of DNA, protein, and RNA from a single sample, with potential for much less cross-contamination of other analytes. Furthermore, the disclosed magnetic beads are compatible with a variety of organic solvents, including, but not limited to, alcohols such as methanol, ethanol, and isopropanol; acetonitrile; esters, such as ethyl acetate; and acids, such as trifluoroacetic acid and formic acid. VI. Applications 1. Detergent Removal In some aspects, a general method of using the disclosed magnetic beads to remove detergent from a sample comprises adding the magnetic beads comprising hydrophobic and/or hydrophilic ligands to the sample that contains the detergent. Typically after agitating the mixture, the magnetic beads are then separated from the sample, such as by using a magnetic stand, and washed to leave the biomolecules of interest in the sample solution for further separation, such as by other aspects of the magnetic beads disclosed herein, or analysis, such as by mass spectrometry. 2. Biomolecule removal In some aspects, a general method of using the magnetic beads disclosed herein to separate biomolecules comprises forming a first suspension comprising a plurality of the disclosed magnetic beads and a first liquid that contains a biomolecule of interest. After the biomolecule has bound to the magnetic beads, the beads are removed from the
first suspension and optionally washed to remove any unwanted or unbound material. A second suspension then is formed comprising the magnetic beads with the bound biomolecule and an elution solution. And the magnetic beads are then removed from the second suspension to leave a solution comprising the biomolecule of interest. The biomolecule may be any biomolecule suitable for binding to the disclosed magnetic beads. In some aspects, the biomolecule is a protein, peptide, polypeptide, DNA or RNA molecule. And in some aspects, the biomolecule is a composite molecule that comprises a combination of a protein, peptide, polypeptide, DNA, and/or RNA moiety. In any aspects, the biomolecule may be labeled. In some aspects, the label may be any label suitable for use in an analytical technique, such as mass spectrometry. Example labels include, but are not limited to, stable isotopes, such as deuterium, carbon-13, nitrogen-15, oxygen-18; isotopically-enriched mass tags, such as, but not limited to, Tandem Mass Tag (TMT and TMTpro) Reagents (available from Thermo Fisher ScientificTM); stable isotopically labeled peptides and/or proteins prepared using SILAC Metabolic Labeling Systems (available Thermo Fisher ScientificTM); rare-earth metals (e.g., lanthanide); fluorescent labels (e.g., dyes); tissues and/or cells grown with heavy amino acids; or any combination thereof. FIG.2 provide a flowchart of an SP3 workflow using the disclosed magnetic beads, that includes a PAC process and an SP2 process which are discussed below in more detail. A. Cleaning-up Peptides and/or Polypeptides In some aspects, the disclosed magnetic beads are used to clean up peptides and/or polypeptides. In some aspects, a mixture is formed comprising the disclosed beads and a sample containing the peptides and/or polypeptides to be cleaned-up. The sample may be a protein digest sample. Optionally, the magnetic beads may be washed before being combined with the sample, for example, to remove a storage buffer solution.
The mixture may further comprise a bead wash/binding solution. In some aspects, the bead wash/binding solution is 100% acetonitrile with 0.1% formic acid. The bead wash/binding solution may be added to the magnetic beads to form a bead slurry before the beads are added to the sample. The amount of the bead slurry that is added to the sample may vary depending on the amount of proteins in the sample. In some aspects, the total volume of the bead slurry for input proteins of 15-30 µg, 30-60 µg, or 60-150 µg should be 250 µL, 500 µL, or 1000 µL, respectively. In some aspects, the bead slurry has a bead concentration of 25%. In certain aspects, about 20 μL of a 25% bead slurry is used that provides 5 mg of beads per sample. In some aspects, from 3 mg to 10 mg of beads are used for 10-25 μg of proteins, such as about 5 mg of beads per 10-25 μg proteins. After combining the disclosed magnetic beads with the sample, the resulting mixture may be incubated at a temperature suitable to facilitate protein binding, such as from 20 °C to 30 °C or at about room temperature. The incubation is allowed to proceed for a time period suitable to facilitate protein binding, such as from 15 minutes to 180 minutes or more, or from 30 minutes to 60 minutes. The mixture may be agitated during incubation, such as by shaking, swirling, stirring, sonication, etc. After incubation, the magnetic beads are removed from the mixture and any unbound peptides or polypeptides by a suitable technique, such as using a magnetic stand. The magnetic beads may be washed to remove any remaining unbound material, solvents, etc. In some aspect s, the wash solution is from 90% to 99% organic in water with from 0.1% to 1% organic acid, and may be about 85% acetonitrile, 10% ethanol, 0.5% formic acid and 4.5% water. The magnetic beads then are treated with an elution solution to remove the bound peptides and/or polypeptides from the magnetic beads. In aspects, the elution solution is from 1% to 5% organic with from 0.05% to 0.25% acid in water, such as 2% acetonitrile/0.1% formic acid in water. The magnetic beads are incubated with the elution solution at a temperature and for a time period suitable to facilitate the peptides and/or polypeptides being eluted from the beads. Incubation may proceed at a
temperature of from 20 °C to 30 °C or at about room temperature, and/or for from 1 minute to 10 minutes or more, such as from 2 minute to 5 minutes, or about 3 minutes. After incubation, the magnetic beads are removed from the eluted peptide and/or polypeptide solution using a suitable technique, such as a magnetic stand. The eluted peptide and/or polypeptide solution then may be transferred to a suitable container for analysis, such as by mass spectrometry. In one aspect, a method for obtaining a peptide or polypeptide for LC-MS analysis comprises: forming a first suspension comprising a plurality of the disclosed magnetic beads and a first liquid that contains a protein digest sample comprising the peptide or polypeptide of interest; agitating the first suspension at room temperature for from 30 to 60 minutes; removing the magnetic beads from the first suspension and washing the magnetic beads using a second liquid; forming a second suspension comprising the magnetic beads and an elution solution and agitating the second suspension at room temperature for from 1 to 5 minutes; and separating the magnetic beads from the second suspension to leave a third liquid comprising the peptide or the polypeptide of interest. In some aspects, each magnetic bead in the plurality of magnetic beads comprises iron oxide particles distributed throughout an azlactone bead and a plurality
moieties located on the surface of the magnetic bead. B. Protein Aggregation Capture (PAC) Aspects of the disclosed magnetic beads are useful for protein aggregation capture (PAC) applications. In some aspects, a mixture is formed comprising the disclosed beads and a sample containing the proteins. Optionally, the magnetic beads may be washed before being combined with the sample, for example, to remove a
storage buffer solution. In some aspects, the washing solution may comprise from 70% to 95% by volume organic solvent, such as from 75% to 85% by volume organic solvent (for example, an alcohol such as 2-propanol, or acetonitrile), and also may comprise an organic acid, such as trifluoroacetic acid (TFA), in an amount of from greater than zero to 1% by volume, such as 0.25% to 1% by volume organic acid, such as TFA. In some aspects, the washed magnetic beads are reconstituted in a solution comprising from 30% to 95% organic solvent, such as from 75% to 85% organic solvent (for example, an alcohol such as 2-propanol, or acetonitrile), and an organic acid, such as TFA, in an amount of from greater than zero to 1%, such as 0.25% to 0.75% organic acid. In some aspects, about 80% 2-propanol with 0.5% TFA is used for the washing solution, reconstitution solution, or both. Organic solvent and optionally organic acid may be added to the protein sample to produce a final organic concentration of from 70% to 95% or more organic solvent in the protein sample, such as from 75% to 85% organic solvent, and optionally from greater than zero to 1% organic acid concentration, such as from 0.25% to 0.57% or about 0.5% organic acid concentration. In any aspects of the PAC method, unless otherwise specified, the organic solvent may be any suitable organic solvent, such as an alcohol (for example, 2- propanol, ethanol, methanol, 1-propanol), an ester solvent (for example, ethyl acetate), or acetonitrile, of a combination thereof. In any aspects of the PAC method, unless otherwise specified, the organic acid may be any suitable organic acid, such as trifluoroacetic acid (TFA) or formic acid (FA). In other aspects, the organic acid is replaced by an organic base, such as a trialkylamine, for example triethylamine. The disclosed magnetic beads are then added to the protein sample. In some aspects, the magnetic beads are added in an amount of from 5:1 w/w beads:protein to 15:1 w/w beads:protein, such as from 8:1 w/w beads:protein to 12:1 w/w beads:protein,
or about 10:1 w/w beads:protein. The magnetic beads may be added in a buffer at from 70% to 95% organic concentration, such as from 75% to 85% organic concentration. And/or the protein concentration may be from greater than zero to 5 mg/mL, such as from 0.5 mg/mL to 3 mg/mL, or from 1 mg/mL to 2 mg/mL. After combining the disclosed magnetic beads with the sample, the resulting mixture may be incubated at a temperature suitable to facilitate protein binding, such as from 20 °C to 30 °C or at about room temperature. The incubation is allowed to proceed for a time period suitable to facilitate protein binding, such as from 15 minutes to 180 minutes or more, or from 30 minutes to 60 minutes. The mixture may be agitated during incubation, such as by shaking, swirling, stirring, sonication, etc. After incubation, the magnetic beads are separated from the mixture. The separated beads are washed in a solution comprising from 65% to 100% organic solvent. The beads may be washed once, or more than once. In some aspects, the beads are washed with 100% organic solvent and followed by 65% to 95% organic solvent, such as 70% to 80% organic solvent. The organic solvent may be any organic solvent suitable for use in the PAC method. In certain aspects, the organic solvent may be acetonitrile or an alcohol. In particular aspects, the beads are washed with 100% acetonitrile followed by 70% alcohol, such as ethanol. After washing, the magnetic beads are exposed to a digestion mixture. In some aspects, from 10 μL to 25 μL digestion mixture is added for a 1-2 mg/mL protein sample. The digestion mixture may be any suitable digestion mixture. In some aspects, the digestion mixture comprises a buffer, such as a trialkyl ammonium buffer (for example, triethylammonium bicarbonate (TEAB)), an inorganic salt, such as calcium chloride, and one or more enzymes (for example, Trypsin and/or LysC). The enzyme or enzyme mixture may be present in an amount of from 1:7 to 1:30 w/w enzyme:protein, such as 1:10 to 1:25 w/w enzyme:protein. The magnetic beads are incubated in the digestion mixture for a time period suitable to facilitate digestion, such as from 1 hour to 5 hours or more, or from 2.4 hours to 3.5 hours. The mixture may be agitated during incubation, such as by shaking,
swirling, stirring, sonication, etc. The incubation may proceed at a temperature suitable to facilitate digestion, such as from 25 °C to 50 °C, from 30 °C to 40 °C, or from 35 °C to 40°C. After incubation, organic solvent is added to the mixture to achieve an organic concentration in the mixture of greater than 90%, such as greater than 92%, or greater than 92.5%. The magnetic beads may be separated and washed with a solution having a high organic concentration, such as an organic concentration of greater than 90%, greater than 92% or at least 95% organic. In some aspects, the washing solution comprises acetonitrile, an alcohol such as ethanol, and water, and may further comprise an organic acid such as formic acid. In one example, the washing solution comprises about 85% acetonitrile, about 10% ethanol, and about 5% water, and may further comprise 0.5% formic acid. After washing, the magnetic beads are eluted with an aqueous solution comprising from 1% to 5% organic solvent, such as from 2% to 4% organic solvent, and from 0.1% to 0.5% organic acid, such as from 0.2% to 0.3% organic acid. In some aspects, the organic solvent is acetonitrile and/or the organic acid is formic acid. After elution, the magnetic beads are removed from the mixture. In one aspect, a method for obtaining a protein comprises: forming a first suspension comprising the plurality of magnetic beads and a first liquid that contains the protein, wherein the first liquid is an aqueous solution comprising 0.5% trifluoroacetic acid, and 80% of an organic solvent selected from 2- propanol and acetonitrile; agitating the first suspension at room temperature for from 30 to 60 minutes; removing the magnetic beads from the first suspension and washing the magnetic beads with a second liquid comprising an organic solvent; incubating the magnetic beads with a digestion mixture comprising Trypsin and LysC for from 1 to 4 hours; washing the beads with a washing solution comprising water, an organic solvent and formic acid, to remove the digestion mixture;
forming a second suspension comprising the magnetic beads and an elution solution and agitating the second suspension at room temperature for from 1 to 5 minutes; and separating the magnetic beads from the second suspension to leave a third liquid comprising the protein. And in some aspects of the method, each magnetic bead comprises iron oxide particles distributed throughout an azlactone bead and a plurality
moieties located on the surface of the magnetic bead. C. Multi-Omic Sequential Aggregation Capture Aspects of the disclosed magnetic beads are useful for sequential aggregation capture applications for samples comprising combinations of nucleic acids and proteins. In some aspects, a mixture is formed comprising the disclosed beads and a sample containing the molecules of interest. The sample may be a cell lysate. In some aspects, from 10 mg to 20 mg of the disclosed magnetic beads are used per 1 x 106 cells of lysate, such as about 15 mg of the disclosed magnetic beads per 1 x 106 cells. Optionally, the magnetic beads may be washed before being combined with the sample, for example, to remove a storage buffer solution. Described below are aspects of a method for separating DNA, protein and RNA from a mixed sample. A person of ordinary skill in the art understands that the method steps can be used in any order, or omitted, depending on the sample provided and the nature of the desired molecules. a. DNA binding After the magnetic beads are added to the sample, organic solvent is added to produce an organic concentration in the sample of from 10% to 40%, such as from 20% to 35% or from 25% to 35%, or about 30% organic concentration. The organic solvent can be any organic solvent suitable for facilitate DNA binding. Unless otherwise specified, the organic solvent may be any suitable organic solvent, such as an alcohol
(for example, 2-propanol, ethanol, methanol, 1-propanol), and ester solvent (for example, an acetate ester, such as ethyl acetate), or acetonitrile, of a combination thereof. After combining the disclosed magnetic beads with the sample, the resulting mixture may be incubated at a temperature suitable to facilitate DNA binding, such as from 20 °C to 30 °C or at about room temperature. The incubation is allowed to proceed for a time period suitable to facilitate DNA binding, such as from 15 minutes to 180 minutes or more, or from 30 minutes to 60 minutes. The mixture may be agitated during incubation, such as by shaking, swirling, stirring, sonication, etc. After incubation, the magnetic beads with the DNA bound thereto are separated from the mixture by a suitable technique, such as a magnetic rack. The remaining lysate may be treated as necessary with additional magnetic beads to remove RNA and/or proteins as disclosed herein. The magnetic beads with the DNA bound thereto is treated with guanidine hydrochloride, a buffer, and a chelating agent, in an aqueous solvent comprising about from 20% to 40% organic solvent, such as from 25% to 35% organic solvent. The organic solvent may be an alcohol, such as 2-propanol, ethanol, methanol or 1-propanol. The resulting mixture is incubated for from 5 minutes to 30 minutes with agitation. After incubation, the beads are removed from the liquid, by a suitable technique, such as a magnetic rack, and are then treated with a second guanidine hydrochloride solution comprising a buffer, surfactant, and chelating agent. The buffer(s) may be Tris HCL, PBS, HEPES, triethylammonium bicarbonate (TEAB), or a combination thereof. The surfactant may be TweenTM, TritonTM X-100, SDS, or a combination thereof. And/or the chelating agent may be ethylenediaminetetraacetic acid (EDTA). After incubating, the supernatant is removed and proteinase K and/or RNase may be added to the supernatant if desired. The mixture is incubated at from 40 °C to 65°C, such as from 50 °C to 60 °C, for from 5 minutes to 30 minutes, with water added to the beads to prevent drying out. After incubation, the water is removed and the supernatant is returned to the beads.
The mixture is treated with a further portion of guanidine hydrochloride in a buffer solution containing EDTA and about 60% organic solvent, and the resulting mixture is incubated for from 30 minutes to 60 minutes at room temperature with agitation. After washing the beads with a solution comprising a buffer, such as a Tris hydrochloride buffer, and about 60% to 80% organic solvent, the DNA is eluted with an aqueous solution comprising a buffer such as Tris HCl and a chelating agent, such as EDTA. The magnetic beads are then removed by a suitable technique, and the DNA can be used or stored for future applications. b. Protein separation A lysate sample, such as the lysate sample from the DNA binding described above, is treated with fresh magnetic beads as disclosed herein, along with sufficient organic solvent to produce a total organic concentration of about 45%. Organic acid, such as TFA or formic acid, also may be added. The resultant mixture is incubated for from 30 minutes to 60 minutes or more with agitation to facilitate protein binding. After incubation the magnetic beads are removed by a suitable technique, such as magnetic rack. The RNA-containing supernatant may be used in an RNA-binding protocol described herein. The magnetic beads with the proteins bound thereto are washed one or more times, such as 1, 2, 3 or more times with from 70% to 100% organic solvent, for example, 70% ethanol and/or 100% acetonitrile. After washing the magnetic beads optionally may be reduced and/or alkylated, such as by treating with tris (2- carboxyethyl)phosphine (TCEP) and/or 2-chloroacetamide. The beads then are treated with universal nuclease followed by Trypsin/LysC. After digestion, the proteins are eluted as described herein with respect to the PAC method. c. RNA binding
Guanidine hydrochloride is added to a liquid sample containing RNA, such as the RNA-containing supernatant described herein, and the mixture is incubated at from 30 °C to 40 °C for from 1 minute to 20 minutes. A clean sample of the disclosed magnetic beads is added along with sufficient organic solvent such that the sample is at least 85% organic solvent, such as 90% organic solvent. Suitable organic solvents include, but are not limited to, acetonitrile or an alcohol, such as 2-propanol, 1- propanol, ethanol, methanol, or a combination thereof. The mixture is incubated at room temperature for a sufficient time to facilitate RNA binding to the magnetic beads, such as for 30 minutes to 60 minutes. After incubation, the magnetic beads are separated from the supernatant by a suitable technique, such as magnetic rack. The magnetic beads are treated with a mixture of buffer, guanidine hydrochloride, surfactant and a chelating agent. The buffer maybe PBS, Tris HCl, or a combination thereof. And/or the surfactant is 10% TweenTM, TritonTM X-100, or a combination thereof. And/or the chelating agent may be EDTA. After incubating, the supernatant is removed and proteinase K and/or RNase may be added to the supernatant if desired. The mixture is incubated at from 40 °C to 65°C, such as from 50 °C to 60 °C, for from 5 minutes to 30 minutes, with water added to the beads to prevent drying out. After incubation, the water is removed and the supernatant is returned to the beads. Organic solvent is added to the bead/sample mixture such that sample is 90% final organic concentration. Suitable organic solvents include, but are not limited to, acetonitrile or an alcohol, such as 2-propanol, 1-propanol, ethanol, methanol, or a combination thereof. The mixture is incubated with agitation at room temperature for a time sufficient to facilitate binding, such as from 30 minutes to 60 minutes. The magnetic beads are then washed with a mixture of from 60% to 80% organic solvent, such as from 65% to 75% organic solvent, a chelating agent, and a buffer. The chelating agent may be EDTA, and/or the buffer may be Tris HCl, PBS, or a combination thereof. Suitable organic solvents include, but are not limited to,
acetonitrile or an alcohol, such as 2-propanol, 1-propanol, ethanol, methanol, or a combination thereof. The RNA is then eluted with water and a chelating agent, such as EDTA, and the magnetic beads are separated from the RNA-containing mixture. VII. Examples Example 1 Preparation of Magnetic Beads A one-liter, Morton type round bottom flask (Flask 1) equipped with an overhead stirrer, nitrogen inlet and a thermocouple was charged with toluene (100-150 mL) and heated to 40 °C with a stirring rate of 200 rpm. A polymeric stabilizer was added (1 g/ml, 282 μl) after which the temperature increased to 45 °C. Heptane (400 mL) was slowly added to the reaction ensuring that the temperature remained above 40 °C at all times. In parallel, a 250-ml round bottom flask (Flask 2), equipped with a magnetic stirrer, nitrogen inlet and a thermocouple, was charged with methylene bisacrylamide (10.5 g, 68 mmol), isopropanol (72.5 mL) and product water (40 mL) and was heated to 30 °C. Vinyl dimethylazlactone (VDM or VDMA) (1.0 g, 7.2 mmol) was added to Flask 1 after the addition of heptane. The remaining steps were performed within 15 minutes. Stir rate was increased followed by the addition of iron oxide (3 g, 13 mmol) (Fe3O4). Sodium persulfate (0.435 g, 1.8 mmol), dissolved in 8 mL of DI water (which was purged with nitrogen for 1 hour prior to use) was added to Flask 2 and stirred for 1- 2 minutes. The magnetic stir bar was removed, and contents of Flask 2 were transferred to Flask 1. The final mixture was stirred for 5-10 minutes at 35 °C, after which time neat TEMED (0.435 mL, 3 mmol) was added to initiate polymerization. Within a minute, an exotherm of about 5-8 °C was observed followed by formation of gray/black beads. The reaction was stirred at 40 °C for 2 hours. Once the reaction had cooled down to room temperature, the beads were filtered and washed with acetone. The beads were resuspended in acetone and sonicated for 2-3 hours in an ice bath. The beads were then sieved using a 63 μm screen on top and 25
μm screen at the bottom with majority of the beads being collected on top of the 25 μm screen. The population of beads between 63-25 μm was isolated, transferred to a 1L Pyrex Bottle and placed in a magnet, such as a magnetic separation system from SepmagTM or equivalent, to remove any non-magnetic residue from the magnetic beads. The beads then were filtered using a Buchner funnel and transferred to a glass vial to be dried in a vacuum oven overnight at 35-40 °C. The magnetic beads were swelled in water and analyzed under a microscope. FIG.3 shows that the process produced round, spherical beads with iron oxide being distributed throughout the body of the beads. Particle size analysis was performed using a LS13-320 XRSW Particle Size Analyzer from Beckman Coulter (Indianapolis, IN). FIG.4 shows that the mean particle size was about 48 μm. FIGS.5-7 provide digital images showing beads made using different sources of magnetic material, including iron oxide clusters (FIG.5), iron oxide particles (FIG.6) and DynabeadsTM (FIG.7). FIGS.5-7 demonstrate that despite the different magnetic materials used, the magnetic beads were all substantially round and spherical, and had an average size of from 30 μm to 60 μm. Example 2 Preparation of Magnetic Beads A one-liter, Morton type round bottom flask (Flask 1) equipped with an overhead stirrer, nitrogen inlet and a thermocouple was charged with toluene (160 mL) and heated to 40 °C with a stirring rate of 200 rpm. A polymeric stabilizer was added (1 g/ml, 282 μl) after which the temperature increased to 45 °C. Heptane (400 mL) was slowly added to the reaction ensuring that the temperature remained above 40 °C at all times. In parallel, a 250-ml round bottom flask (Flask 2), equipped with a magnetic stirrer, nitrogen inlet and a thermocouple, was charged with methylene bisacrylamide (20 g), isopropanol (72.5 mL) and product water (40 mL) and was heated to 30 °C. Vinyl dimethylazlactone (VDM or VDMA) (1.0 g, 7.2 mmol) was added to Flask 1 after the addition of heptane. Stir rate was increased to 500 rpm. Iron source and sodium sulfate were added to flask 2 containing MBA and added to flask 1. The
final mixture was stirred for 5-10 minutes after which time neat TEMED was added to initiate polymerization. Within a minute, an exotherm of about 5-8 °C was observed followed by formation of gray/black beads. The reaction was stirred at 600 rpm and 40 °C for 2 hours. Once the reaction had cooled down to room temperature, magnetic beads were filtered and washed with acetone. The beads were sonicated, sieved and tested for microscopic analysis as well as particle size analysis using the procedure described in Example 1. Example 3 Preparation of Magnetic Beads A one-liter, Morton type round bottom flask (Flask 1) equipped with an overhead stirrer, nitrogen inlet and a thermocouple was charged with toluene (160 mL) and heated to 40 °C with a stirring rate of 400 rpm. A polymeric stabilizer was added (1 g/ml, 282 μl) after which the temperature increased to 45 °C. Heptane (200 mL) was slowly added to the reaction ensuring that the temperature remained above 40 °C at all times. In parallel, a 250-ml round bottom flask (Flask 2), equipped with a magnetic stirrer, nitrogen inlet and a thermocouple, was charged with methylene bisacrylamide (20 g), isopropanol (72.5 mL) and product water (40 mL) and was heated to 30 °C. Vinyl dimethylazlactone (VDM or VDMA) (1.0 g, 7.2 mmol) was added to Flask 1 after the addition of heptane. Stir rate was increased to 500 rpm after which DynabeadsTM (1g) and sodium sulfate were added to flask 2 containing MBA. The mixture was stirred for 2 minutes and added to flask 1. The final mixture was stirred for 5-10 minutes after which time neat TEMED was added to initiate polymerization. Within a minute, an exotherm of about 5-8 °C was observed followed by formation of gray/black beads. The reaction was stirred at 850 rpm and 40 °C for 2 hours. Once the reaction had cooled down to room temperature, magnetic beads were filtered and washed with acetone. The beads were sonicated, sieved and tested for microscopic analysis as well as particle size analysis using the procedure described in Example 1.
Example 4 Hydrolysis of Magnetic Beads
Magnetic beads were hydrolyzed according to the following procedure. Magnetic beads (0.50 g) were added to a 15 ml conical tube containing water (10 mL) and rotated overnight at room temperature. The beads were washed the next day with water using an appropriate magnetic stand. The hydrolyzed beads were then adjusted to a 25% slurry in water. Solutions used in Examples 5-8 are provided in Table 1 unless otherwise specified. Table 1. List of solutions
Example 5 Example Procedure (SP2) using the Disclosed Magnetic Beads Use 15-150 µg of protein per sample preparation. Rinse cultured cells or tissues 2-3 times with 1 × PBS to remove cell culture media or excess blood, respectively. Resuspend proteins, cells, or tissues in Lysis Solution without additional buffers. A. Extract protein, reduce, and alkylate 1. For cultured cells, add 50 µL of Lysis Buffer and 1 µL of Universal Nuclease to a minimum of 1 × 106 cells. Pipet up and down (with P200 tip) for 10-15 cycles until sample viscosity is reduced. Note: Lyse the cells in an appropriate volume of lysis buffer to have the final concentration of 2-5 µg/µl with a final volume of 7.5 µL, 15 µL, or 30 µL for input proteins of 15-30 µg, 30-60 µg, or 60-150 µg, respectively. Centrifugation of cultured cell lysates is typically not required after aspiration using a pipet. 2. For tissue samples, add 50 µL of Lysis Solution (containing 1 µL Universal Nuclease) per 5 mg of tissue and disrupt with tissue homogenizer until sample is homogenized. Centrifuge tissue lysates at 16,000 × g for 10 minutes. 3. For purified proteins, depleted and undepleted plasma, and serum samples, dilute samples directly in Lysis Solution to 2-5 µg/µL with a final volume of 7.5 µL, 15 µL, or 30 µL for input proteins of 10-25 µg, 25-50 µg, or 50-100 µg, respectively. Note: The concentration of plasma is around 60-70 µg/µL. For purified proteins and plasma samples, addition of Universal Nuclease is not required. 4. Determine the protein concentration of the supernatant using established methods such as the Pierce™ BCA Protein Assay Kit (Product No.23227) or Pierce™ Rapid Gold BCA Protein Assay Kit (Product No. A53226). Note: Dilution with Lysis Solution may be required for concentrated samples for BCA assay. 5. Transfer 15-150 µg of protein sample into a 1.5-mL low protein binding tube or a 96-well plate and adjust the final volume to 7.5 µL, 15 µL, or 30 µL for input proteins of 15-30 µg, 30-60 µg, or 60-150 µg, respectively with Lysis Solution.
6. Add 3.75 µL, 7.5 µL, or 15 µL of Reduction Solution for input proteins of 15-30 µg, 30-60 µg, or 60-150 µg, respectively to the sample and gently mix. 7. Add 3.75 µL, 7.5 µL, or 15 µL of Alkylation Solution for input proteins of 15- 30 µg, 30-60 µg, or 60-150 µg, respectively to the sample and gently mix. 8. Incubate the sample at 95 °C using a heat block for 10 minutes or 50 °C for 20- 30 minutes to reduce and alkylate the protein sample. 9. After incubation, allow the sample to cool to room temperature. 10. Briefly centrifuge the tube or plate before digestion. B. Digest protein 1. Add 150 µL of Enzyme Reconstitution Solution to 1 vial of Trypsin/Lys-C Protease Mix to prepare 0.67 µg/µL enzyme mix. 2. Add 3.75 µL, 7.5 µL, or 15 µL of the reconstituted enzyme solution for input proteins of 15-30 µg, 30-60 µg, or 60-150 µg, respectively to the reduced and alkylated protein sample solution. 3. Incubate with shaking at 37°C for 1-2 hours to digest the protein sample. Note: Optional labeling with TMT reagents can be performed at this point before peptide clean-up. C. Clean-up peptides 1. Transfer 20 µL of Magnetic Bead slurry to a microcentrifuge tube or a 96-well plate and remove the storage buffer using a magnetic stand. 2. Wash the resin by adding 200 µL of Bead Wash/Binding Solution, then vortex briefly and remove the buffer using a magnetic stand. 3. Add Bead Wash/Binding Solution to the resin and transfer the bead slurry to the protein digest sample. The total volume of the diluted sample for input proteins of 15- 30 µg, 30-60 µg, or 60-150 µg should be 250 µL, 500 µL, or 1000 µL, respectively. 4. Incubate the sample at room temperature for 30-60 minutes with mixing. 5. Collect the bead using a magnetic stand and remove the unbound peptide solution. 6. Wash the resin by adding 500 µL of Wash Solution, then vortex briefly and remove the buffer using a magnetic stand.
7. Add 30-100 µL of Elution Solution, then vortex briefly and incubate for 3 minutes with mixing at room temperature. 8. Transfer the eluted peptide solution using a magnetic stand to a new container for LC-MS analysis. Example 6 Example Procedure (PAC or Multi-analyte Aggregation Capture, with SP3 for protein) Note: Any amount of protein can be used as long as bead: protein ratios are scaled accordingly. Rinse cultured cells or tissues 2-3 times with 1 × PBS to remove cell culture media or excess blood, respectively. Resuspend proteins, cells, or tissues in a lysis solution, such as the Lysis Solution described herein. For protein extraction, many buffer components are allowable, including various detergents, salts, and chaotropic agents. For DNA extraction, chaotropic agents should be avoided. For RNA extraction, guanidine hydrochloride should be added to lysis buffer. Lysis and Reduction/Alkylation – General Notes and Getting Started: 1. For cultured cells, add lysis buffer to a minimum of 1 × 106 cells. Use enough lysis buffer for an anticipated concentration between 2-12 mg/mL protein. Let sit on ice for 15 minutes. Note: for 1 x 106 cells, assume 450 μg protein, 5 μg DNA, and 10 μg RNA and use 15 mg of the disclosed magnetic beads per each sample type. Note: For tissue samples, add lysis buffer to tissue and disrupt with tissue homogenizer until sample is homogenized. Note: For purposes of protein extraction only, add 1% (by volume) Universal nuclease to lysis buffer. Do not add nuclease or sonicate sample if the intention is to perform multi-omic sequential extractions of any combination of RNA, DNA, or protein. For this purpose, immediately move to “Bind DNA to Beads” section described in Example 5. 2. Centrifuge lysates at 16,000 × g for 15 minutes at 4 °C.
3. Determine the protein concentration of the supernatant using established methods such as the Pierce™ BCA Protein Assay Kit (Product No.23227) or Pierce™ Rapid Gold BCA Protein Assay Kit (Product No. A53226). 4. Proceed with reduction and alkylation, if preferred, to a final concentration of 10 mM TCEP and 20 mM 2-chloroacetamide (or equivalent protocol using dithiothreitol and iodoacetamide). TCEP and 2-chloroacetamide can be used together, but DTT and iodoacetamide must be used sequentially with DTT quench after incubation periods are complete. Note: Reduction and alkylation can be done before or after protein binding to beads. If reduction and alkylation are done after protein binding to beads, heat at 50 °C for 45 minutes. If reduction and alkylation are done prior to protein binding to beads, heat at 95 °C for 10 minutes. After incubation, allow the sample to cool to room temperature before adding beads or enzymes. Bind Protein to Beads (SP3 for protein ONLY) Note: This protocol is written with Reduction and Alkylation occurring prior to protein binding on bead at 95 °C for 10 minutes, with 5-minute cooling period prior to adding beads. Note: for protein only applications, 2-propanol and acetonitrile and inter- changeable. Note: Upstream of PAC workflow, lysis, reduction and alkylation chemistry is compatible. 1. Vortex the disclosed magnetic beads to ensure total suspension. 2. Remove supernatant from beads and wash in 500uL of 80% 2-propanol, 0.5% TFA. Remove wash. 3. Recon beads in 80% 2-propanol 0.5% TFA for [50 μg/μL]. 4. Add 99.5% 2-propanol 0.5% TFA to protein solution for a final organic concentration of 80% 2-propanol, 0.5% TFA. 5. Beads (10:1 bead: protein w/w) are added to protein in buffer for 80% organic concentration and 1-2mg/mL protein concentration.
6. Incubate 45 minutes room temp with sufficient agitation to keep beads in suspension, then remove the flow through. 7. Add 4x bead volume of 100% MeCN and vortex thoroughly. Remove wash solution. 8. Add 4x bead volume of 70% EtOH and vortex thoroughly. Remove wash solution. 9. The tubes are removed from the magnetic rack and 10-25uL digestion mixture is added to beads, comprising 50mM TEAB, 10mM CaCl2, and 1:10- 1:25 (enzyme: protein (w/w)) Trypsin and LysC. 10. Vortex briefly and incubate 3 hours in a Thermomixer at 37 °C, 1200 rpm. 11. After incubation, centrifuge briefly to collect any evaporated liquid. 12. Add 100% acetonitrile to sample and beads eluate for final organic concentration at least 92.5%. Note: from this point, SP2 is also compatible. 13. Sample vortexed 30 seconds. Total peptide concentration is at least 0.2 mg/mL. 14. Sample was incubated 10 minutes at room temperature. 15. Beads are collected and the supernatant is aspirated to waste. 16. Sample is washed twice with 4x bead volume of 85% acetonitrile, 10% ethanol, 5% water, 0.5% formic acid. 17. Beads are collected and the supernatant was aspirated to waste. 18. Sample is eluted with 2x bead volume of 2-4% MeCN, 0.2%FA in water for 2 minutes with vortexing (per 15 mg beads). 19. The tube is placed on the magnetic rack for 2 minutes, and the supernatant is transferred to a clean tube. Samples are centrifuged at 10,000xg for 30 seconds, and then the sample is moved to a new tube without disturbing pellet. 20. Quantitate with fluorometric or colorimetric peptide assay, or UV-Vis and/or move to downstream application. Example 7 Multi-OMIC Sequential Aggregation Capture
Bind DNA to Beads Note: this protocol is for sequential elution of DNA-Protein-RNA. Other sequences are possible with these beads. For extracting Protein-DNA only, start with 2% SDS, 100 mM TEAB, 1 mM EDTA, then reduce and alkylate with TCEP (10 mM) and 2-chloroacetamide (20mM), and bind protein at 80% 2-propanol, 0.5% TFA, take flow through and dilute with water to 30-40% 2-propanol for DNA binding. Presence of CH6ClN3 (guanidine hydrochloride) substantially alters the binding properties of each analyte on this magnetic bead and therefore organic concentrations are variable depending on when CH6ClN3 is introduced and the concentration thereof. 1. Dilute sample 1:1 with 0.2% SDS, 100 mM TEAB. Let sit on ice 15 minutes. 2. Wash 15 mg of the disclosed magnetic beads with 100% 2-propanol (per each 1 x 106 cells of lysate). 3. Add the required amount of beads to the lysates. 4. Add enough 2-propanol to the sample to achieve a total concentration of 30% organic. 5. Bind for 45 minutes at room temperature with sufficient agitation to keep beads in suspension. 6. Place on a magnetic rack and remove the lysate, which now contains RNA and protein, to “Bind Protein” section of this protocol. Move forward with DNA processing steps below, using only beads. 7. For each 15 mg of beads, add about 250 μL of 30% 2-propanol, 4.2 M CH6ClN3, 150 mM Tris HCl, 20 mM EDTA, remove from magnet and incubate 10 minutes at room temperature with sufficient agitation to keep beads in suspension. 8. Place on magnetic rack and discard supernatant to waste. 9. Add 100 μL of the following formulation per each 15 mg beads: 0.5x PBS, 28% CH6ClN3, 10% TweenTM, 0.5% Tris HCl, 0.5% EDTA, 0.5% TritonTM X-100. Invert several times to mix with beads and put on magnetic rack. 10. Remove supernatant to a new tube. Add proteinase K and RNase if desired, invert several times to mix, and incubate for 10 minutes at 55 °C, 400 rpm. Note: add water to beads during incubation so they do not dry out.
11. After incubation, remove water from beads and add sample back to beads. 12. Add a 1x volume of 60% 2-propanol, 2.4M CH6ClN3, 150 mM Tris HCl, 20 mM EDTA to bead/sample mixture and bind for 45 minutes at room temperature with sufficient agitation to keep beads in suspension. 13. Wash DNA-bound beads 2x with 200 μL 70% EtOH, 1 mM EDTA, and 25 mM Tris HCl (per 15 mg beads). 14. Elute in 50 μL 25 mM Tris-HCl, 0.1 mM EDTA (per 15 mg beads). 15. Place on magnetic rack, and transfer supernatant to new 1.5 mL tubes 16. Centrifuge 12,000 xg at 4 °C for 2 minutes. 17. Move to new tubes without beads. 18. Add EDTA to 1 mM and store at 4 °C until downstream application. Bind Protein to Beads (Sequential Method ONLY) 1. Retrieve flowthrough from “Bind DNA to Beads” or from “Lysis and Reduction/Alkylation”. 2. Wash a new set of 30 μL (15 mg) of the disclosed magnetic beads with 200uL 99.5% 2-propanol, 0.5% trifluoroacetic acid, then remove wash solution from beads. 3. Transfer samples to 15mg of the disclosed magnetic beads (per 1 x 106 cells). 4. Add enough 99% 2-Propanol, 1% TFA to a total concentration of 45% organic. 5. Bind for 45 minutes at room temperature with sufficient agitation to keep beads in suspension. 6. Place on magnetic rack and transfer RNA-containing supernatant to new 1.5 mL tubes. a. Add 0.1mM EDTA to RNA-containing supernatant and store at 4 °C for short term storage. 7. Wash with 200 μL 70% EtOH, vortex briefly (per 15 mg beads). 8. Wash with 200 μL 100% MeCN, vortex briefly (per 15 mg beads). 9. Resuspend bead-bound proteins in 40 μL 100mM TEAB, 10 mM CaCl2.
a. Reduce and alkylate with 10 μL total of TCEP and 2-chloroacetamide, if not done prior to bead binding. Final concentrations are 20 mM 2-chloroacetamide, 10 mM TCEP. Vortex sample and incubate at 50 °C for 45 minutes. b. If the samples have been previously reduced and alkylated, add 50uL 100 mM TEAB, 10mM CaCl2 to beads instead of 40 μL. 10. Add 0.6 μL universal nuclease. Pipette up and down repeatedly. 11. Take 1 vial of Trypsin/LysC from -20 °C and let thaw 5 minutes. a. Reconstitute vial in 50 μL 0.1% CH3COOH for 2.0 mg/mL. Mix gently and let sit 5 minutes. b. Add 1:10-1:25 of enzyme:protein (w/w). c. Incubate 2 hours in a Thermomixer at 37 °C, 1200rpm. Continue with the steps below for complete SP3 workflow of protein preparation. Alternatively, SP2 preparation is also compatible and can be used in place of this protocol, and still be SP3 (scaled for 15 mg beads) 12. Begin peptide cleanup by adding enough 2-propanol for >92.75% organic concentration. a. Sample is incubated 10 minutes at room temperature 1200 rpm. b. Beads are collected and the supernatant was aspirated to waste. c. Sample is washed twice with 200 μL of 100% HPLC-grade MeCN (or 85% MeCN, 10% EtOH, 5.0% water, 0.5% formic acid for improved salt removal, such as in TMT- or TMTpro-labeled samples). d. Beads are collected and the supernatant is aspirated to waste. e. Sample is eluted with 50 μL of 2-4% MeCN, 0.2%FA in water for 2 minutes with occasional vortex. f. Place on the magnetic rack for 2 minutes, and transfer supernatant to a clean tube. g. Samples are centrifuged at 10,000xg for 30 seconds. h. Sample is moved to a new tube without disturbing pellet.
13. Quantitate with fluorometric or colorimetric peptide assay, or UV-Vis and/or move to downstream application. Bind RNA to Beads 1. Retrieve RNA flowthru from 4 °C storage (45% 2-propanol with RNA). 2. Add 50 mg dry CH6ClN3 to 200 μL sample to reach 2.5M CH6ClN3 and incubate 10 minutes, 37 °C, 1200 rpm. 3. Wash 30 μL fresh magnetic beads disclosed herein with 200 μL 2-propanol. 4. Transfer 200 μL sample to 15 mg dry magnetic beads. 5. Add about 1200 μL 2-propanol such that sample is 90% final organic concentration. 6. Bind for 45 minutes at room temperature with sufficient agitation to keep beads in suspension. 7. Place on magnetic rack and discard supernatant. 8. Add 100 μL of the following formulation per each 15 mg beads: 0.5x PBS, 28% CH6ClN3, 10% TweenTM, 0.5% Tris HCl, 0.5% EDTA, 0.5% TritonTM X-100. Invert several times to mix with beads and put on magnetic rack. 9. Remove RNA-containing supernatant from beads and transfer to new 1.5 mL tube. a. Resuspend beads in 200 μL water. 10. Remove RNA-containing supernatant to a new tube. Add proteinase K and RNase if desired, invert several times to mix, and incubate for 10 minutes at 55 °C, 400 rpm. Note: add water to beads during incubation so they do not dry out. 11. After incubation, remove water from beads and add sample back to beads. 12. Add 2-propanol to bead/sample mixture such that sample is 90% final organic concentration and bind for 45 minutes at room temperature with sufficient agitation to keep beads in suspension. 13. Wash RNA-bound beads 2x with 400 μL 70% EtOH, 1 mM EDTA, and 25 mM Tris HCl (per 15mg bead, don’t vortex!).
14. Elute in 50 μL water, 0.1 mM EDTA (per 15mg bead). 15. Extract purified RNA from beads and place in new 1.5 mL tube and move to downstream application. Example 8 Comparison Experiment To compare the disclosed magnetic beads with other, commercially available beads, a comparative workflow was developed using a KingFisherTM Sample Purification System from Thermo Fisher ScientificTM. Table 2 provides the beads that were used for the comparison experiment. Table 2. Beads used in the comparison experiment
A 25 μg HeLa lysate sample was used for each bead type. The comparative workflow was as follows: A. Protein Digestion (Off KingFisher) • Reduction/alkylation: 95 °C for 10 minutes, cooled to room temperature, and centrifuged for seconds. • Digestion: 37 °C for 1 hour with shaking at 1,000 rpm. • Trypsin/Lys-C: 0.67 μg/μL (100 μg of Trypsin/Lys-C reconstituted in 150 μL of 0.1% acetic acid/10 mM CaCl2 for use, sample to trypsin ratio: 10:1)
Table 3. Sample/reagent volume for protein digestion
B. Peptide Cleanup (On KingFisher) • Bead conditioning: Load 20 μL of beads (5.0 mg) with 80 μL of Bead Wash/Binding Solution. Wash the beads with 200 μL of Bead Wash/Binding Solution. • Dilution: Transfer beads into the diluted sample. The total volume is 500 μL (>9% organic phase after dilution). • Binding: Mix the sample/beads at room temperature for 45 minutes at a medium rate. • Wash: Transfer the sample/beads into 500 μL of Wash Solution for peptide wash. Mix the plate at room temperature for 2 minutes at a medium rate. • Elution: Transfer the sample/beads into 100 μL of Elution Solution for peptide elution. Mix the plate at room temperature for 5 minutes at a medium rate. Collect the output peptide samples for LC-MS analysis. Table 4. Peptide cleanup volumes
Results from the experiment are provided in FIGS.8A, 8B, and 9-11. FIGS.8A and 8B demonstrate that the disclosed magnetic beads exhibit less surface absorption loss than the Sera-MagTM carboxylate modified particles, MyOneTM, or silica beads in the SP2 workflow. The SeraMagTM, MyOneTM, and silica beads all demonstrated significant bead loss on the surfaces of the comb tip plate and sample plate (bead
transfer/mix and binding steps). In contrast, the disclosed magnetic beads did not exhibit significant bead loss. The data provided in FIGS.8A and 8B with respect to the disclosed magnetic beads refers to hydrolyzed beads, that is beads where the azlactone moieties on the surface of the beads have been hydrolyzed. However, similar results were achieved using disclosed magnetic beads where the azlactone moieties were treated with aminobenzoic acid or aminoethyl-trimethylammonium chloride (AETMA) (data not shown). FIG.9 provides data demonstrating that the peptide yield from the KingFisher SP2 workflow for the disclosed magnetic beads was substantially higher than the peptide yield for the Sera-MagTM, MyOneTM Silane or silica beads, using a quantitative colorimetric peptide assay. No peptides were recovered using silica beads. FIG.10 demonstrates that using the disclosed magnetic beads, substantially more unique peptides were identified by nano-LCMS analysis, than were identified using the other bead types. And FIG.11 demonstrates that substantially more protein groups were identified using the disclosed magnetic beads. Example 9 An experiment was designed to demonstrate the superior binding of the disclosed magnetic beads coupled to alkali stable Protein A (asPA), compared to a commercial magnetic bead that comprises asPA conjugates. Conjugate asPA to the disclosed magnetic beads Alkali stable Protein A was conjugated to the disclosed azlactone beads according to the following procedure: 1. Weigh out 77mg of the disclosed azlactone beads. 2. Dilute asPA (20mg/mL in 250mM Sodium Acetate, pH 5.5, 0.5M NaCl, 0.05% TweenTM-20) into Coupling Buffer (0.1M Sodium Carbonate, 1.2M Sodium Citrate, pH 9) 1:3 (v/v); final concentration of components is 5mg/mL asPA, 0.075M Sodium Carbonate, 0.9M Sodium Carbonate, pH 9 3. Add 3.6mL of asPA in Coupling Buffer to azlactone beads.
4. Couple asPA to beads overnight at room temperature. 5. Wash asPA-beads with 2-3 mL water. 6. Block asPA-beads with 2-3 mL of 3M ethanolamine for 2 hours. 7. Wash asPA-beads with 2-3 mL water. 8. Wash asPA-beads twice with 2-3 mL 0.1M Glycine, pH 2 (Pierce IgG Elution Buffer, #21028) 9. Wash asPA-beads three times with 2-3 mL water. 10. Slurry asPA-beads to 25% using 20% ethanol. Measure the binding capacity of asPA-azlactone beads compared to asPA- magnetic agarose The procedure for measuring the binding capacity of asPA-azlactone beads compared to asPA-magnetic agarose (Pierce High Capacity Protein A MagBeads, alkali stable, #A53036, 25% slurry) was as follows: 1. Prepare 3mg/mL Rabbit IgG (RIgG) in PBS. 2. Add 2 x 50uL of 25% mag bead slurry (asPA-azlactone beads and asPA-magnetic agarose) to 1.5mL microcentrifuge tubes. 3. Wash beads twice with 500uL of PBS. 4. Add 500uL of 3mg/mL (RIgG) to beads and incubate for 1hr at room temperature with shaking/mixing. 5. Collect the flowthrough and save for analysis. 6. Create a RIgG standard curve from 0 to 3mg/mL by measuring absorbance at 280nm. 7. Measure absorbance of starting solution of RIgG and Flowthrough from each bead sample. 8. Calculate concentrations of start and flowthrough using standard curve. 9. Calculate RIgG bound to the mag beads by subtracting the RIgG in each Flowthrough from the starting RIgG. Extrapolate to amount RIgG bound per mL of settled beads.
Results The results clearly demonstrated that the asPA conjugated disclosed magnetic beads had about a 15% higher RIgG binding capacity that asPA-magentic agarose beads (FIG.12). Without being bound to a particular theory, this may be due, at least in part, to the disclosed azlactone magnetic beads being able to conjugate a larger amount of asPA than an amount asPA that the same amount of magnetic agarose beads can conjugate. And in some aspects, the amount of asPA bound to the disclosed azlactone magnetic beads is about 2-fold higher than the amount of asPA bound to the same amount of magnetic agarose beads. Example 10 Derivatization of Magnetic Beads with Ocylamine Ligands Various hydrophilic or hydrophobic molecules or proportional mixtures of hydrophilic and hydrophobic molecules were reacted to the amine-reactive, azlactone groups of magnetic beads to impart the support with detergent removal and peptide recovery properties. Scheme 1 shows the reaction of a representative hydrophobic ligand (octylamine, C8) to a magnetic bead bearing a terminal azlactone group.
Scheme 1 A solution of ligand(s) in ethanol was prepared by dissolving 10mg/mL, 2mg/mL, or 40mg/mL of hydrophobic, hydrophilic or proportional mix of hydrophobic and hydrophilic ligand, respectively, in 8mL of ethanol with stirring. 250mg of magnetic base beads (swell volume : 1 gram of magnetic beads gives 6 mLs of beads)
were added to the solution and allowed to react for 2 hours with end-over-end mixing. The reacted beads were then washed thoroughly twice with 1 bed volume of water, and then resuspended in water for later use. This general procedure was used to modify the magnetic beads with various types of hydrophobic ligands (i.e., octylamine (C8), dodecylamine (C12), octadecylamine (C18)) and hydrophilic ligands (i.e., methoxy Jeffamine (MJ), 4,7,10- Trioxa-1,13 tridecanediamine (C10PEG), B- cyclodextrin, dextran, and PEG molecules ranging in MW from 2000 Da to 20,000 Da), and combinations thereof. Table 5 lists different ligands that are reacted to the azlactone group of the magnetic beads. These ligands range from 4 carbon chain to 18 carbon chain across different concentrations immobilized on the bead. The hydrophobic and/or hydrophilic property that is imparted to the beads because of this reaction is also shown in the table. Table 5: Examples of Ligands for Attachment to Magnetic Beads
Example 11 Removal of Non-Ionic and Ionic Detergents by Azlactone Magnetic Beads Magnetic beads bearing azlactone groups were used to remove non-ionic and ionic detergents from biological samples (e.g., cell lysates and samples containing proteins and/or polypeptides. Examples of detergent removal workflows implementing various detection methods are provided below. Absorbance 280 protocol (To determine removal of TritonTM X 100 and NP 40) 1. Pipette 200 μL of 25% slurry magnetic beads into Dynamag2 magnetic stand. 2. Remove supernate. 3. Pipette 100 μL of 1% detergent solution into bead sample. 4. Vortex detergent and beads until homogenous 5. Place solution in Dynamag2 magnetic stand and collect flow through. 6. Pipette 4 μL of detergent flow through onto the NanoDrop One instrument to read the detergent absorbance at 280nm (include diluted standards). Coomassie Protein Assay protocol (To determine removal of detergents)
1. Pipette 200 μL of 25% slurry magnetic beads into Dynamag2 magnetic stand. 2. Remove supernate. 3. Pipette 100 μL of 1% detergent solution into bead sample. 4. Vortex detergent and beads until homogenous. 5. Place solution in Dynamag2 magnetic stand and collect flow through. 6. Pipette 10 μL of each sample and standard into a 96 well plate. 7. Pipette 300 μL of Coomassie Blue Stain Reagent into each occupied well. 8. Absorption measured on Multiskan GO instrument at 595 nm. CMC-535 Florescence Assay protocol (To determine removal of detergents) 1. Pipette 200 μL of 25% slurry magnetic beads into Dynamag2 magnetic stand. 2. Remove supernate. 3. Pipette 100 μL of 1% detergent solution into bead sample. 4. Vortex detergent and beads until homogenous 5. Place solution in Dynamag2 magnetic stand and collect flow through. 6. Pipette 50 μL of detergent flow through and 100uL of 1x fluorescent dye/ reagent 1 into a black 96 well plate (include diluted standards). 7. Shake plate for 2 minutes. 8. Pipette 50 μL of 1x Reagent 2 into each occupied well. 9. Shake plate for 2 minutes. 10. Read the fluorescence on the Varioskan instrument at 535 nm, using an excitation at 485 nm. HeLa Digest/Colorimetric Peptide Assay protocol (To determine recovery of peptides) 1. Pipette 200 μL of 25% slurry magnetic beads into Dynamag2 magnetic stand. 2. Remove supernate. 3. Pipette 100 μL of 1M Peptide digest assay solution into bead sample. 4. Vortex peptides and beads cautiously until homogenous 5. Place solution in Dynamag2 magnetic stand and collect flow through.
6. Pipette 20 μL of peptide flow through and appropriate standards into a clear- bottom 96 well plate. 7. Pipette 200 μL of colorimetric assay solution into each of the occupied wells. a) 5 mL of Colorimetric Assay Reagent A b) 4.8 mL of Colorimetric Assay Reagent B c) 200 μL of Colorimetric Assay Reagent C 8. Shake plate for 30 seconds. 9. Incubate plate for 15 minutes at 37 °C. 10. Read peptide absorbance on Multiskan GO instrument at 480 nm. Table 6 shows the removal of 1% NP-40 and 1% TritonTM X-100 using the beads prepared according to Example 10. Sample 4, 5, 6 and 7 (C8 and C12) removed 84% or more of the detergents from the sample. However, ligands with shorter (e.g., C4) or longer carbon chains (e.g., C18) removed substantially less detergent from the sample. Table 6. Removal of 1% NP-40 and 1% TritonTM X-100 Detergents
Example 12
Removal of Detergents from Cell Digests Magnetic beads were tested to determine their ability to remove ionic and non- ionic detergents from cell digest samples (Pierce™ HeLa Protein Digest Standard (Thermo Scientific, Cat. # 88328) and the recovery of proteins and peptides from the sample. Table 7 shows the recovery of peptides from a 1 mg/mL HeLa digest sample using the beads for detergent removal according to Example 10. Sample 4 (50 mg/mL C12 ligand) showed the lowest recovery of HeLa digest. Although the bead trapped detergents well, these beads also trapped 90% of the peptides. However, when the C12 ligand was blended with a hydrophilic ligand, the beads gave a moderate recovery of HeLa peptides at 37%. In comparison, C8 modified beads (Sample 6) and C8 blends with hydrophilic beads (Sample 7) resulted in 40% and 48% recovery, indicating that use of beads coupled to C8 ligand can be used for effective detergent removal from cell digest samples. Table 7. Recovery of peptides from a 1 mg/mL HeLa digest sample
Example 13 Removal of Detergents from Beads Coupled to Hydrophilic Ligands Different types of hydrophilic beads were coupled to the magnetic beads and evaluated using the workflows described herein. Table 4 shows different types of hydrophilic beads that are reacted to the magnetic beads. These beads are immobilized at various concentrations as show in in the table. Table 8. Magnetic Bead Samples with Hydrophilic Ligands
Table 9 shows the % removal of detergents TritonTM X-100 and NP-40 at 1% concentration using hydrophilic ligands reacted to the beads. The hydrophilic ligands have poor capacity to trap detergents and as a result they were eliminated from further evaluation. Table 9: Detergent removal capacity of hydrophilic coated magnetic beads
Example 14 Removal of Detergents from Beads Coupled to Blends of Hydrophobic and Hydrophilic Ligands Different blends of C8 and hydrophilic ligands beads were prepared and evaluated using the workflows described herein. Table 10 shows variations of octylamine ligand reacted beads blended with hydrophilic ligands such as O-(2- aminopropyl)-O’-(2-methoxyethyl)polypropylene glycol (Methoxy Jeffamine) and C10 PEG diamine in various blend ratios.
O-(2-aminopropyl)-O’-(2-methoxyethyl)polypropylene glycol where n is from 3 to 25 (Methoxy Jeffamine) Table 10. Octylamine and C8 blends with hydrophilic ligands immobilized magnetic beads
Table 11 shows removal of three different non-ionic detergents 1% TritonTM X- 100, 1% NP-40 and 1% n-Dodecyl-β-D-maltoside (DDM). Beads with a mixture of hydrophobic and hydrophilic properties were found to remove a significant amount of detergent from the tested samples (e.g., 82%-100%). Table 11. Removal of Detergents from Beads with Hydrophobic and Hydrophilic Properties
The samples also were evaluated for detergent removal from HeLa digests using the workflow described herein. Referring to Table 12, 40% or greater HeLa digest was recovered with C8 modified beads and its blends with hydrophilic ligands.
Table 12. Recovery of HeLa digest with C8 modified beads and its blends with hydrophilic ligands
Example 15 Removal of Detergents from Beads Coupled to Blends of Hydrophobic and Hydrophilic Ligands Hydrophobic C8 ligand at different concentrations ranging from 10 mg/mL to 30 mg/mL was immobilized on the magnetic beads and evaluated using the workflows described herein (Table 13). Table 13. Different concentrations of Octylamine (C8) immobilized on magnetic beads
Table 14 shows that octylamine immobilized on the beads tested at various concentrations yielded beads capable of removing at least twice the amount of detergent across all detergents tested as compared to the base beads with no ligand modification. For the non-ionic detergents, it can be advantageous to equilibrate the beads in low pH solution such as 0.1 N HCl to improve detergent removal efficiency.
Table 14. Detergent Removal with Octylamine Derivatized Magnetic Beads
The % recovery of three different cell lysates HeLa, HEK2 and Jurkat with the octylamine immobilized beads at three different concentrations of 20 mg/mL, 25 mg/mL and 30 mg/mL is shown in Table 15.45% or greater recovery is ideal for subsequent MS analysis of cell lysates, especially when evaluated using highly- sensitive instruments that are particularly sensitive to the presence of residual detergent. Table 15. Recovery of Cell Lysates
Example 16 Removal of Detergents from Beads Coupled to Hydrophobic Ligands Sample 22 was compared against a commercial bead (Pierce NHS-Activated Beads (Cat. # 88826)) (Sample 25) derivatized with C8 ligand at the same concentration
as used in the preparation of Sample 22. Table 16 compares the detergent removal capacity for each bead sample. Table 16. Comparison of magnetic beads with hydrophobic ligands
The exemplary disclosed beads comprising octylamine conjugated to azlactone groups on the magnetic beads surprisingly removed significantly more detergent than beads comprising octylamine conjugated to commercially available magnetic beads through NHS reactive groups. These data demonstrate the surprisingly superior results obtained using the disclosed magnetic beads compared to commercially available beads, even when the commercially available beads are conjugated to the same type of ligand as the disclosed beads. Example 17 Mass Spectrometry (MS) analysis of HeLa protein digest sample Materials and Reagents Pierce™ HeLa Protein Digest Standard (Thermo Scientific, Cat. # 88328) Instrument LC: DIONEX UltiMate 3000, Model# NCS-3500RS, Serial# 8100814 MS: Thermo Scientific™ Q Exactive™ Hybrid Quadrupole-Orbitrap™ Mass Spectrometer, Serial# SN03659L
Method Inject 500 ng of HeLa protein digest samples before (control, no addition of detergents) and after (samples, with/without addition of detergents TritonTM X-100 or NP-40) bead processing for LC-MS analysis using a 150-min gradient time. Beads used: Samples 21 (Octylamine 10 mg/mL) and 22 (Octylamine 10 mg/mL) from Example 15. Data analysis: Data was analyzed using Proteome Discoverer 2.2 software. Results For both beads (that is, both Samples 21 and 22), PEG contamination peaks were observed at the end of the nanoLC gradient/elution time, with no significant interference with the peptide separation and identification (data not shown). Both beads exhibited no significant bias regarding the MS chromatogram profile, indicating consistent peptide identification/coverage. Compared to the control sample (HeLa protein digest without processed with beads), both beads yielded comparable protein identification numbers and a <10% difference in identified peptides (FIGS.13 and 14). The data demonstrates that both beads provided comparable protein identification with only a small decrease in identification compared to the control sample that was not treated with a bead. And Table 17 provides the success rate of each bead compared to the untreated control sample. Table 17. Success Rate
Example 18 MS analysis of detergent samples before and after bead processing Materials and Reagents
Pierce™ HeLa Protein Digest Standard (Thermo Scientific, Catalog number 88328) Instrument MS: Q Exactive HF mass spectrometer, Serial# SN05061L Method Dilute the detergent (n-Dodecyl-β-D-maltoside (DDM) and NP-40) samples before (control/reference sample) and after (samples) bead processing using 50% acetonitrile/0.1% formic acid with an appropriate ratio (e.g., 1:1000 – 1:100) for direct infusion MS analysis. The relative detergent removal efficiency was calculated based on the percentage of MS signals between the testing samples and the control/reference samples, and removal efficiency is shown in Table 18. Table 18. Removal efficiency of magnetic beads
In view of the many possible aspects to which the principles of the disclosure may be applied, it should be recognized that the illustrated aspects are only preferred examples of the disclosure and should not be taken as limiting the scope of the disclosure. Rather, the scope of the disclosure is defined by the following claims. We therefore claim as our disclosure all that comes within the scope and spirit of these claims.
Claims
We claim: 1. A method, comprising: form a suspension comprising a first plurality of magnetic beads and a first solution that contains a first amount of a detergent; agitate the suspension; remove the magnetic beads from the suspension to leave a second solution comprising a second amount of a detergent that is less than the first amount of the detergent; wherein each magnetic bead in the first plurality of magnetic beads comprises a crosslinked polymer, a magnetic material, and a plurality of hydrophobic ligands formed from a reaction between azlactone moieties and one or more hydrophobic molecules.
2. The method of claim 1, wherein the one or more hydrophobic molecules comprise a straight or branched C4-C20 alkyl amine. 3. The method of claim 2, wherein the straight or branched C4-C20 alkyl amine is selected from butylamine (C4), hexylamine (C6), octylamine (C8), decylamine (C10), dodecylamine (C12), tetradecylamine (C14), hexadecylamine (C16), octadecylamine (C18), eicosylamine (C20), or a combination thereof. 4. The method of claim 3, wherein the straight or branched C4-C20 alkyl amine is selected from octylamine (C8), dodecylamine (C12), octadecylamine (C18), or a combination thereof. 5. The method of any one of claims 1-4, wherein the hydrophobic ligands are selected from
,
6. The method of any one of claims 1-5, wherein the hydrophobic ligand is
7. The method of any one of claims 1-6, wherein the magnetic beads in the first plurality of magnetic beads further comprise a hydrophilic ligand. 8. The method of claim 7, wherein the hydrophilic ligand is formed by a reaction between azlactone moieties and one or more hydrophilic molecules.
9. The method of any one of claims 7-8, wherein the beads comprise hydrophobic and hydrophilic ligands in a ratio of from 10% hydrophobic:90% hydrophilic to 90% hydrophobic:10% hydrophilic. 10. The method of claim 9, wherein the ratio of hydrophobic to hydrophilic ligands is from 20%:80% hydrophobic:hydrophilic to 40%:60% hydrophobic:hydrophilic. 12. The method of any one of claims 1-6, wherein the suspension further comprises a second plurality of magnetic beads where each magnetic bead in the second plurality of magnetic beads comprises a crosslinked polymer, a magnetic material, and a plurality of hydrophilic ligands formed from a reaction between azlactone moieties and one or more hydrophilic molecules. 13. The method of claim 12, wherein the first plurality of magnetic beads and the second plurality of magnetic beads are each provided in amounts sufficient to provide a ratio of beads comprising hydrophobic ligands (referred to as hydrophobic beads) to beads comprising hydrophilic ligands (referred to as hydrophilic beads) of from 10% hydrophobic beads:90% hydrophilic beads to 90% hydrophobic beads:10% hydrophilic beads. 14. The method of claim 13, wherein the ratio of hydrophobic beads to hydrophilic beads is from 20%:80% hydrophobic beads:hydrophilic beads to 40%:60% hydrophobic beads:hydrophilic beads. 15. The method of any one of claims 8 to 14, wherein the one or more hydrophilic molecules is selected from O-(2-aminopropyl)-O’-(2- methoxyethyl)polypropylene glycol, 4,7,10-Trioxa-1,13 tridecanediamine, β- cyclodextrin, dextran, PEG molecules ranging in MW from 2000 Da to 20,000 Da, or a combination thereof.
16. The method of any one of claims 7-15, wherein the hydrophilic ligands are selected from
where m is from 40-450 (MW of from about 2,000 to 20,000 DA),
17. The method of claim 16, wherein the hydrophilic ligand is
18. The method of any one of claims 7-16, wherein the hydrophobic ligand is
the hydrophilic ligand is
from 3 to 25,
. 19. The method of any one of claims 1-18, wherein the second amount of detergent is less than 0.1% of the second solution. 20. The method of claim 19, wherein the second amount of detergent is 0.01% or less of the second solution. 21. The method of any one of claims 1-20, wherein the detergent is selected from polyethylene glycol sorbitan monooleate, t-Octylphenoxypolyethoxyethanol, Octylphenoxypolyethoxyethanol (NP-40), n-Dodecyl-β-D-maltoside (DDM), sodium dodecyl sulfate (SDS), Glyco-diosgenin (GDN), Lauryl Maltose Neopentyl Glycol (LMNG), or a combination thereof. 22. A magnetic bead, comprising a crosslinked polymer bead and a magnetic material contained within the polymer bead, and further comprising a plurality of ligands formed wherein the ligands are selected from: hydrophobic ligands formed from a reaction between azlactone moieties and one or more hydrophobic molecules; hydrophilic ligands formed from a reaction between azlactone moieties and one or more hydrophilic molecules; or a combination of hydrophobic ligands and hydrophilic ligands formed from reactions between azlactone moieties and one or more hydrophobic molecules and one or more hydrophilic molecules.
23. The magnetic bead of claim 22, wherein the plurality of ligands are hydrophobic ligands formed from a reaction between azlactone moieties and one or more hydrophobic molecules. 24. The magnetic bead of claim 22, wherein the plurality of ligands are a combination of hydrophobic ligands and hydrophilic ligands formed from reactions between azlactone moieties and one or more hydrophobic molecules and one or more hydrophilic molecules. 25. The magnetic bead of any one of claims 22-24, wherein the magnetic material comprises an iron source. 26. The magnetic bead of claim 25, wherein the iron source comprises iron oxide. 27. The magnetic bead of any one of claims 22-26, wherein the magnetic material is contained within an agarose bead, and the agarose bead is contained within the crosslinked polymer. 28. The magnetic bead of any one of claims 22-27, wherein the magnetic material is in the form of a particle, powder, flake, or cluster. 29. The magnetic bead of any one of claims 22-28, wherein the crosslinked polymer is a crosslinked azlactone polymer, and the magnetic bead is an azlactone bead. 30. The magnetic bead of any one of claims 22-29, wherein the one or more hydrophobic molecules comprise a straight or branched chain C4-C20alkyl amine. 31. The magnetic bead of claim 30, wherein the straight or branched C4-C20 alkyl amine is selected from butylamine (C4), hexylamine (C6), octylamine (C8),
decylamine (C10), dodecylamine (C12), tetradecylamine (C14), hexadecylamine (C16), octadecylamine (C18), eicosylamine (C20), or a combination thereof. 32. The magnetic bead of claim 30, wherein the straight or branched C4-C20 alkyl amine is selected from octylamine (C8), dodecylamine (C12), octadecylamine (C18), or a combination thereof. 33. The magnetic bead of claim 32, wherein the hydrophobic ligand is selected from
34. The magnetic bead of any one of claims 22-33, wherein the hydrophobic
35. The magnetic bead of any one of claims 22 and 24-34, wherein the hydrophilic ligand is derived from O-(2-aminopropyl)-O’-(2- methoxyethyl)polypropylene glycol, 4,7,10-Trioxa-1,13 tridecanediamine, β- cyclodextrin, dextran, a PEG molecule having a molecular weight of from 2,000 to 20,000 Da, or a combination thereof. 36. The magnetic bead of any one of claims 22 and 24-35, wherein a ratio between the hydrophobic ligand and the hydrophilic ligand is from 10% hydrophobic:90% hydrophilic to 90% hydrophobic:10% hydrophilic by weight. 37. The magnetic bead of claim 36, wherein the ratio of hydrophobic to hydrophilic ligands is from 20%:80% hydrophobic:hydrophilic to 40%:60% hydrophobic:hydrophilic by weight. 38. The magnetic bead of any one of claims 22 and 24-37, wherein the magnetic bead comprises:
as the hydrophobic ligand; and
as the hydrophilic ligand. 39. The magnetic bead of any one of claims 22-38, wherein the bead has a bead size of from 20 to 80 microns.
40. The magnetic bead of claim 39, wherein the bead size is from 30 to 60 microns. 41. A mixture of magnetic beads according to any one of claims 22-40, the mixture comprising a first plurality of magnetic beads comprising hydrophobic ligands and a second plurality of magnetic beads comprising hydrophilic ligands. 42. The mixture of claim 41, wherein the first plurality of magnetic beads and the second plurality of magnetic beads are each provided in amounts sufficient to provide a ratio of beads comprising hydrophobic ligands (referred to as hydrophobic beads) to beads comprising hydrophilic ligands (referred to as hydrophilic beads) of from 10% hydrophobic beads:90% hydrophilic beads to 90% hydrophobic beads:10% hydrophilic beads. 43. The mixture of claim 42, wherein the ratio of hydrophobic beads to hydrophilic beads is from 20%:80% hydrophobic beads:hydrophilic beads to 40%:60% hydrophobic beads:hydrophilic beads. 44. A kit comprising a plurality of magnetic beads according to any one of claims 22-40 or a mixture of magnetic beads according to any one of clams 41-43. 45. A method for making a magnetic bead according to any one of claims 22-40, the method comprising: add a plurality magnetic bead comprising azlactone functional groups to a solution comprising molecule selected from hydrophobic molecules, hydrophilic molecules, or a combination of hydrophobic molecules and hydrophilic molecules; allow the magnetic beads to react with at least a portion of the molecules to form magnetic beads comprising ligands; and separate the magnetic beads comprising the ligands from the solution;
wherein when the molecules are hydrophobic molecules then the ligands are hydrophobic ligands, when the molecules are hydrophilic molecules then the ligands are hydrophilic ligands, and when the molecules comprise a combination of hydrophobic molecules and hydrophilic molecules then the ligands are a mixture of hydrophobic ligands and hydrophilic ligands. 46. The method of claim 45, wherein the molecules are hydrophobic molecules and the ligands are hydrophobic ligands. 47. The method of claim 45, wherein the molecules are hydrophilic molecules and the ligands are hydrophilic ligands. 48. The method of claim 45, wherein the molecules comprise a combination of hydrophobic molecules and hydrophilic molecules and the ligands are a mixture of hydrophobic ligands and hydrophilic ligands.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363517029P | 2023-08-01 | 2023-08-01 | |
| US63/517,029 | 2023-08-01 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025029872A1 true WO2025029872A1 (en) | 2025-02-06 |
Family
ID=92542682
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/040308 Pending WO2025029872A1 (en) | 2023-08-01 | 2024-07-31 | Magnetic beads for detergent removal |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2025029872A1 (en) |
Citations (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5292840A (en) | 1987-03-13 | 1994-03-08 | Minnesota Mining And Manufacturing Company | Polymeric supports |
| US5336742A (en) * | 1987-03-13 | 1994-08-09 | Minnesota Mining And Manufacturing Company | Polymeric supports |
| US5993935A (en) * | 1991-10-11 | 1999-11-30 | 3M Innovative Properties Company | Covalently reactive particles incorporated in a continous porous matrix |
| US8735455B2 (en) * | 2008-06-26 | 2014-05-27 | 3M Innovative Properties Company | Solid support with a grafted chain |
-
2024
- 2024-07-31 WO PCT/US2024/040308 patent/WO2025029872A1/en active Pending
Patent Citations (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US5292840A (en) | 1987-03-13 | 1994-03-08 | Minnesota Mining And Manufacturing Company | Polymeric supports |
| US5336742A (en) * | 1987-03-13 | 1994-08-09 | Minnesota Mining And Manufacturing Company | Polymeric supports |
| US5403902A (en) | 1987-03-13 | 1995-04-04 | Minnesota Mining And Manufacturing Company | Polymeric supports |
| US5993935A (en) * | 1991-10-11 | 1999-11-30 | 3M Innovative Properties Company | Covalently reactive particles incorporated in a continous porous matrix |
| US8735455B2 (en) * | 2008-06-26 | 2014-05-27 | 3M Innovative Properties Company | Solid support with a grafted chain |
Non-Patent Citations (4)
| Title |
|---|
| ANONYMOUS: "One-Step Detergent Removal kit", 27 November 2022 (2022-11-27), pages 1 - 3, XP093223103, Retrieved from the Internet <URL:https://bioclone.net/wp-content/uploads/pdf/instruction-manual/sample/One-Step%20Detergent%20Removal%20Kit.pdf> [retrieved on 20241111] * |
| ANONYMOUS: "Safety data sheet for product code BD101 and BD102 issued by Bioclone Inc.", 27 November 2022 (2022-11-27), pages 1 - 5, XP093223112, Retrieved from the Internet <URL:https://bioclone.net/wp-content/uploads/pdf/MSDS/SDS-Sample%20Preparation.pdf> [retrieved on 20241111] * |
| PRAI-IN YINGRAK ET AL: "Recyclable magnetic nanocluster crosslinked with poly(ethylene oxide)-block-poly(2-vinyl-4,4-dimethylazlactone) copolymer for adsorption with antibody", MATERIALS SCIENCE AND ENGINEERING C, ELSEVIER SCIENCE S.A, CH, vol. 67, 7 May 2016 (2016-05-07), pages 285 - 293, XP029579386, ISSN: 0928-4931, DOI: 10.1016/J.MSEC.2016.05.032 * |
| PRAY-IN YINGRAK ET AL: "Hydrophilic azlactone-functionalized magnetite nanoparticle for conjugation with folic acid", JOURNAL OF NANOPARTICLE RESEARCH 16, 22 March 2014 (2014-03-22), XP093217422, Retrieved from the Internet <URL:http://link.springer.com/article/10.1007/s11051-014-2357-7/fulltext.html> [retrieved on 20241023], DOI: 10.1007/s11051-014-2357-7 * |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| Filipović et al. | Affinity-based isolation of extracellular vesicles by means of single-domain antibodies bound to macroporous methacrylate-based copolymer | |
| US20180010169A1 (en) | Methods and reagents for selection of biological molecules | |
| Vergara-Barberan et al. | Recent advances in aptamer-based miniaturized extraction approaches in food analysis | |
| US10816546B2 (en) | Binding a target substance | |
| Li et al. | Efficient on‐chip proteolysis system based on functionalized magnetic silica microspheres | |
| CN100500837C (en) | Methods for separating molecules | |
| JP2005538929A (en) | Methods for isolating nucleic acids and proteins from a single sample | |
| EP3043905B1 (en) | New diagnostic assay using particles with magnetic properties | |
| JP7462972B2 (en) | Biomagnetic microspheres and methods of preparation and use thereof | |
| EP1774334B1 (en) | Use of magnetic material to direct isolation of compounds and fractionation of multipart samples | |
| Braun et al. | Peptides as biosorbents–promising tools for resource recovery | |
| US9983203B2 (en) | Method for protein analysis | |
| CN102421898A (en) | Nucleic acid purification method | |
| WO2003095646A1 (en) | Isolating nucleic acid | |
| WO2025029872A1 (en) | Magnetic beads for detergent removal | |
| WO2025029903A1 (en) | Functionalized magnetic beads and methods of preparation and use | |
| CN114907612A (en) | Magnetic microsphere and manufacturing method and application thereof | |
| AU2004279361A1 (en) | Methods and kits for purifying His-tagged proteins | |
| US20060030056A1 (en) | Use of magnetic material to fractionate samples | |
| EP4025911B1 (en) | Conjugates composed of membrane-targeting peptides for extracellular vesicles isolation, analysis and their integration thereof | |
| CN115449533A (en) | A low-abundance protein enrichment kit, preparation method and application | |
| JP2006327962A (en) | Method for separating target substance and molecular complex | |
| JP2001158800A (en) | Avidin particles | |
| CN117723748B (en) | Immunomagnetic bead for enhancing target protein signal for targeted detection, and preparation method and application thereof | |
| CN115494243B (en) | Method for quantitatively determining binding efficiency of streptavidin-biotin and application |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24761735 Country of ref document: EP Kind code of ref document: A1 |