WO2025019901A1 - Rna vaccines - Google Patents
Rna vaccines Download PDFInfo
- Publication number
- WO2025019901A1 WO2025019901A1 PCT/AU2024/050795 AU2024050795W WO2025019901A1 WO 2025019901 A1 WO2025019901 A1 WO 2025019901A1 AU 2024050795 W AU2024050795 W AU 2024050795W WO 2025019901 A1 WO2025019901 A1 WO 2025019901A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- rna
- amino acid
- sequence
- seq
- lipid
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Pending
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/195—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K39/02—Bacterial antigens
- A61K39/0216—Bacteriodetes, e.g. Bacteroides, Ornithobacter, Porphyromonas
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P1/00—Drugs for disorders of the alimentary tract or the digestive system
- A61P1/02—Stomatological preparations, e.g. drugs for caries, aphtae, periodontitis
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P31/00—Antiinfectives, i.e. antibiotics, antiseptics, chemotherapeutics
- A61P31/04—Antibacterial agents
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P37/00—Drugs for immunological or allergic disorders
- A61P37/02—Immunomodulators
- A61P37/04—Immunostimulants
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/62—DNA sequences coding for fusion proteins
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
- C12N9/48—Hydrolases (3) acting on peptide bonds (3.4)
- C12N9/50—Proteinases, e.g. Endopeptidases (3.4.21-3.4.25)
- C12N9/52—Proteinases, e.g. Endopeptidases (3.4.21-3.4.25) derived from bacteria or Archaea
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/51—Medicinal preparations containing antigens or antibodies comprising whole cells, viruses or DNA/RNA
- A61K2039/53—DNA (RNA) vaccination
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/55—Medicinal preparations containing antigens or antibodies characterised by the host/recipient, e.g. newborn with maternal antibodies
- A61K2039/552—Veterinary vaccine
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/555—Medicinal preparations containing antigens or antibodies characterised by a specific combination antigen/adjuvant
- A61K2039/55511—Organic adjuvants
- A61K2039/55555—Liposomes; Vesicles, e.g. nanoparticles; Spheres, e.g. nanospheres; Polymers
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/555—Medicinal preparations containing antigens or antibodies characterised by a specific combination antigen/adjuvant
- A61K2039/55511—Organic adjuvants
- A61K2039/55572—Lipopolysaccharides; Lipid A; Monophosphoryl lipid A
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K39/00—Medicinal preparations containing antigens or antibodies
- A61K2039/57—Medicinal preparations containing antigens or antibodies characterised by the type of response, e.g. Th1, Th2
- A61K2039/575—Medicinal preparations containing antigens or antibodies characterised by the type of response, e.g. Th1, Th2 humoral response
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K38/00—Medicinal preparations containing peptides
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12R—INDEXING SCHEME ASSOCIATED WITH SUBCLASSES C12C - C12Q, RELATING TO MICROORGANISMS
- C12R2001/00—Microorganisms ; Processes using microorganisms
- C12R2001/01—Bacteria or Actinomycetales ; using bacteria or Actinomycetales
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y304/00—Hydrolases acting on peptide bonds, i.e. peptidases (3.4)
- C12Y304/22—Cysteine endopeptidases (3.4.22)
- C12Y304/22037—Gingipain R (3.4.22.37)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y304/00—Hydrolases acting on peptide bonds, i.e. peptidases (3.4)
- C12Y304/22—Cysteine endopeptidases (3.4.22)
- C12Y304/22047—Gingipain K (3.4.22.47)
Definitions
- RNA vaccines Field of the invention relates to RNA-containing vaccine compositions for inducing an immune response to Porphyromonas gingivalis in a subject, and uses thereof.
- Related application [0002] This application claims priority from Australian provisional application AU 2023902379, the entire contents of which are hereby incorporated by reference.
- Background of the invention [0003] If dental plaque is left to accumulate around the tooth at the gingival (gum) margin this causes gingival inflammation (gingivitis). Chronic gingivitis can allow the emergence of a periodontal pathogen Porphyromonas gingivalis (P. gingivalis) at the base of a periodontal pocket to result in a chronic infection and the development of severe disease.
- P. gingivalis Periodontal pathogen Porphyromonas gingivalis
- Periodontitis This severe form of periodontal disease is called periodontitis and can lead to tooth loss in an approach by the immune system to eliminate the infection.
- Chronic periodontitis is an inflammatory disease of the supporting tissues of the teeth leading to resorption of alveolar bone and eventual tooth loss. The disease is a major public health problem in all societies and is estimated to affect up to 30% of the adult population with severe forms affecting 12-15% of the adult population.
- One in three adults have moderate to severe periodontitis. From epidemiological surveys, periodontitis has been linked to an increased risk of inflammatory diseases including cardiovascular diseases, certain cancers, preterm birth, rheumatoid arthritis and dementia. More recent research has linked chronic infection by P.
- the invention provides an RNA polynucleotide encoding a protein comprising or consisting of: - one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and/or - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. wherein the polynucleotide is capable of being translated in a mammalian cell. 1005272698
- the protein encoded by the RNA polynucleotide may further comprise: - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto.
- the protein encoded by the RNA may comprise the afore-mentioned domains in any order: for example: the one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P.
- gingivalis may be located N-terminally to the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, and/or the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis; or the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, may be located N-terminally to the one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P.
- ABSMs adhesin binding motifs
- gingivalis and/or the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis The domains may be directly joined in the context of the chimeric or fusion protein, or may be joined via a linker region, of one or more amino acid residues, as further defined herein.
- the protein may comprises or consist of, N to C terminus: active site (K) - ABMs (A); or active site (R) – ABMs (A); or active site (K) – DUF 2436 (D); or active site (R) – DUF2436 (D); or AMBs (A) – active site (K); or ABMs (A)-active site (R); or DUF2436 (D) – active site (K); or DUF2436 (D) – active site (R); or active site (R) – ABMs (A) - active site (K); or active site (R) – ABMs (A) - active site (R); or active site (K) – ABMs (A) - active site (R); or DUF2436 (D) –– ABMs (A); or DUF2436 (D) –– ABMs (A); or DUF2436 (D) –– ABMs (A) – active site (R) or (K).
- the RNA polynucleotide is in the form of a messenger RNA (mRNA) molecule.
- mRNA messenger RNA
- the RNA polynucleotide may be in any suitable format for being translated in a mammalian cell and enabling synthesis of the protein encoded by the RNA.
- the RNA polynucleotide may be composed entirely of ribose-containing nucleotides, or alternatively, may comprise a combination of ribose- containing nucleosides and of 2’-deoxyribose-containing nucleotides.
- the RNA polynucleotide may be a synthetic RNA molecule. 1005272698
- the RNA polynucleotide may be a circular RNA (circRNA) molecule.
- the RNA polynucleotide may be a complementary RNA (cRNA) molecule.
- the RNA polynucleotide made be a self-amplifying RA (saRNA) molecule or trans-amplifying (taRNA) molecule.
- saRNA self-amplifying RA
- taRNA trans-amplifying
- the RNA may further encode an N-terminal signal peptide for enabling secretion of the protein following translation thereof.
- the N-terminal signal peptide may comprise any amino acid sequence which enables the protein encoded by the RNA to be processed by ribosomes bound to the rough endoplasmic reticulum (ER) of a cell, and thereby results in threading of the protein into the ER. From the ER, the protein is capable of being transported to the plasma membrane and secreted from the mammalian cell.
- N-terminal secretion peptides are known to the skilled person and are further described herein.
- the RNA may further comprise a 5’ untranslated region (UTR) and a 3’ UTR.
- the RNA may also comprise a 5’ cap analog, such as 7mG(5′)ppp(5′)NlmpNp.
- the RNA may also comprise a polyadenine (polyA) tail.
- the poly(A) tail may be non-segmented or segmented with a short spacer element.
- the RNA may comprise a chemical modification. Examples of suitable chemical modification include a N1-methylpseudouridine modification or a N1-ethylpseudouridine modification or may comprise any chemical modification described herein.
- the polynucleotide has a uridine content of less than about 50%, less than about 45%, less than about 40%, less than about 35%, less than about 30%, less than about 25%, less than about 20% or less than about 15%. In preferred embodiments, the polynucleotide has a uridine content of between about 15% and about 35%, preferably between about 15% and about 25%. 1005272698
- the uridines in the polynucleotide are replaced with a chemical modification such as N1-methyl-pseudouridine.
- a chemical modification such as N1-methyl-pseudouridine.
- at least 25%, at least 30%, at least 35%, at least 40%, at least 45%, at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, or 100% of the uridine nucleosides are replaced with N1-methyl- pseudouridine.
- the RNA polynucleotide is in the form of a codon optimised RNA molecule, optionally depleted of uridine nucleosides.
- the codon optimisations comprises conversion of codons encoding serine to UCG.
- the protein encoded by the RNA polynucleotide is a chimeric or fusion protein comprising or consisting of: - one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P.
- the chimeric or fusion protein encoded by the RNA may comprise the afore-mentioned domains in any order: for example: the one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P.
- gingivalis may be located N-terminally to the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, may be located N-terminally to the one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P.
- ABSMs adhesin binding motifs
- the domains may be directly joined in the context of the chimeric or fusion protein, or may be joined via a linker region, of one or more amino acid residues, as further defined herein.
- the chimeric or fusion protein may comprises or consist of, N to C terminus: active site (K) - ABMs (A); or active site (R) – ABMs (A); or ABMs (A) – active site (K); or ABMs (A)-active site (R); or active site (R) – ABMs (A) - active site (K); or active site (R) – ABMs (A) - active site (R); or active site (K) – ABMs (A) - active site (R).) 1005272698
- amino acid sequences of an Arg- or Lys-gingipain of P. gingivalis are further described herein.
- the amino acid sequence of an active site of an Arg- or Lys- gingipain of P. gingivalis comprises the amino acid sequence of KAS or RAS (the Lysine or Arginine active site histidine sequence), ie a peptide including the active site histidine and surrounding area of the active site.
- gingivalis (also designated “R” herein), comprises the amino acid sequence of SEQ ID NO: 38, encoded by the RNA sequence as set forth in SEQ ID NO: 50, or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto.
- the amino acid sequence of an active site of a Lys- gingipain of P comprises the amino acid sequence of SEQ ID NO: 38, encoded by the RNA sequence as set forth in SEQ ID NO: 50, or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least
- gingivalis (also designated “K” herein), comprises the amino acid sequence of SEQ ID NO: 8, encoded by the RNA sequence as set forth in SEQ ID NO: 43, or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto.
- the amino acid sequence of the active site of an Arg- or Lys- gingipain does not comprise the entire catalytic domain of the gingipain.
- the chimeric or fusion protein encoded by the RNA polynucleotide comprises i) an amino acid sequence that comprises or consists of an amino acid sequence of the active site of an Arg-gingipain of P. gingivalis, or sequences that are at least 80% identical thereto; and ii) an amino acid sequence that comprises or consists of an amino acid sequence of the active site of a Lys-gingipain of P. gingivalis, or sequences that are at least 80% identical thereto.
- the chimeric or fusion protein encoded by the RNA polynucleotide comprises at least two amino acid sequences that comprise or consist of an amino acid sequence of the active site of an Arg- or Lys-gingipain of P. gingivalis, or sequences that 1005272698
- the at least two amino acid sequences may be located contiguously in the chimeric or fusion protein, or may be located in different locations within the chimeric or fusion protein.
- one of the at least two amino acid sequences may be located at the N terminus of the chimeric or fusion protein while the second of the at least two amino acid sequences may be located at the C-terminus of the chimeric or fusion protein.
- one of the at least two amino acid sequences may be located at the N or C terminus of the chimeric or fusion protein while the second of the at least two amino acid sequences may be located within the chimeric or fusion protein (ie not at either N or C termini).
- the at least two amino acid sequences may (both) be located at the N terminus of the chimeric or fusion protein or the at least two amino acid sequences may (both) be located at the C-terminus of the chimeric or fusion protein.
- the chimeric or fusion protein encoded by the RNA polynucleotide may further comprise: - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto.
- the amino acid sequence comprising the amino acid sequence of a DUF2436 domain is located between the amino acid sequence of the active site of the gingipain of P.
- the chimeric or fusion protein comprises, N to C terminus or C to N terminus: active site (K) or (R) – DUF domain (D) – ABMs (A); or active site (K) or (R) – DUF domain (D) – ABMs (A) – active site (K) or (R)).
- the amino acid sequence of a DUF2436 domain of an Arg or Lys gingipain of P is a DUF2436 domain of an Arg or Lys gingipain of P.
- gingivalis comprises or consists of the amino acid sequence of SEQ ID NO: 35 or 76, encoded by the RNA sequence as set forth in SEQ ID NO: 51, or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto. 1005272698
- a cysteine residue in the DUF2436 domain may be substituted to a serine or valine residue, preferably to a serine residue (such as shown in SEQ ID NO: 36).
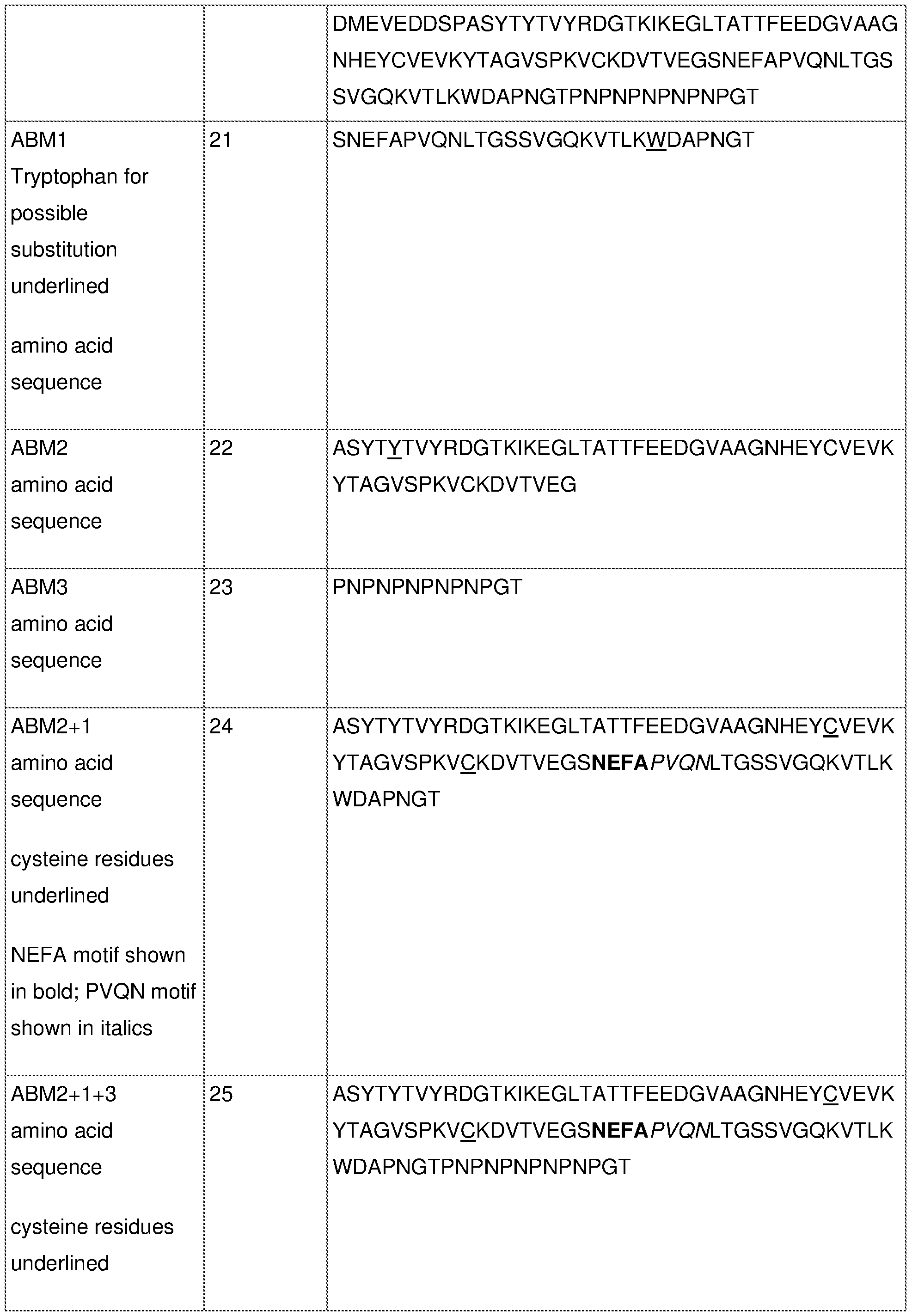
- the one or more adhesin binding motifs comprise or consist of the amino acid sequence of ABM2 and/or ABM1 (for example as set forth in SEQ ID NO: 22 and SEQ ID NO: 21, respectively, or comprising the amino acid sequence as set forth in SEQ ID NO: 24 (ABM2+1), or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto.
- Such amino acid sequences may be encoded by an RNA comprising the sequence set forth in SEQ ID NOs: 52, 53, 55, herein.
- the ABMs may further comprise ABM3 (eg SEQ ID NO: 73 or encoded by SEQ ID NO: 54).
- the ABMs may be provided in any order, but are preferably in the order ABM2 - ABM1 - ABM3.
- the one or more adhesin binding motifs may comprise or consist of the amino acid sequence of ABM2, ABM1 and ABM3 (for example as set forth in SEQ ID NO: 25, 72 or 73 and encoded by an RNA comprising the sequence of SEQ ID NO: 56 or 77 or 78).
- the one or more adhesin binding motifs may comprise one or more modifications selected from: a) one or more cysteine amino acid substitutions compared to the naturally occurring Arg- or Lys-gingipain sequences in corresponding regions; b) substitution of the proline and/or an asparagine residues in the sequence PxxN corresponding to, or at a position equivalent to, residues 6 to 9 of the sequence of SEQ ID NO: 21 (ABM1); c) substitution of the motif NxFA to SxYQ in the sequence, corresponding to, or at a position equivalent to residues 2 to 5 of the sequence of SEQ ID NO: 21 (ABM1); 1005272698
- the one or more cysteine amino acid substitutions may be a substitution to a serine residue or to a valine residue.
- the one or more cysteine substitutions may comprise one or more substitutions to a serine residue.
- only one cysteine residue may be substituted. In other embodiments, two or three cysteine residues may be substituted.
- the cysteine residues are substituted to a combination of valine and serine residues. In other embodiments, all substituted cysteine residues are substituted to serine or all substitute cysteine residues are substituted to valine.
- the motif PxxN eg PVQN, SEQ ID NO: 106
- the proline amino acid substitution is preferably a substitution to an alanine residue.
- the asparagine amino acid substitution may be a substitution to a proline residue or an alanine residue.
- the asparagine residue is substituted to a proline residue. In other embodiments, the asparagine residue is not substituted.
- the substitution is from PxxN to AxxP, (eg AVQP, SEQ ID NO: 107) (such as exemplified in the amino acid sequences of SEQ ID NOs: 30 to 32).
- the one or more adhesin binding motifs comprise the amino acid sequence as set forth in any one of SEQ ID NOs: 21 to 25, and comprising: a) one or more cysteine amino acid substitutions compared to the naturally occurring Arg- or Lys-gingipain sequences in corresponding regions, preferably substitution of all cysteine residues; and 1005272698
- the one or more adhesin binding motifs comprise or consist the amino acid sequence as set forth in any one of SEQ ID NOs: 26 to 34, or sequences at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto, provided that the sequences comprise the aforementioned substitutions of the cysteine and proline and asparagine residues.
- the RNA encodes a chimeric or fusion protein comprising or consisting of: [0055] a) - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P.
- ABSMs adhesin binding motifs
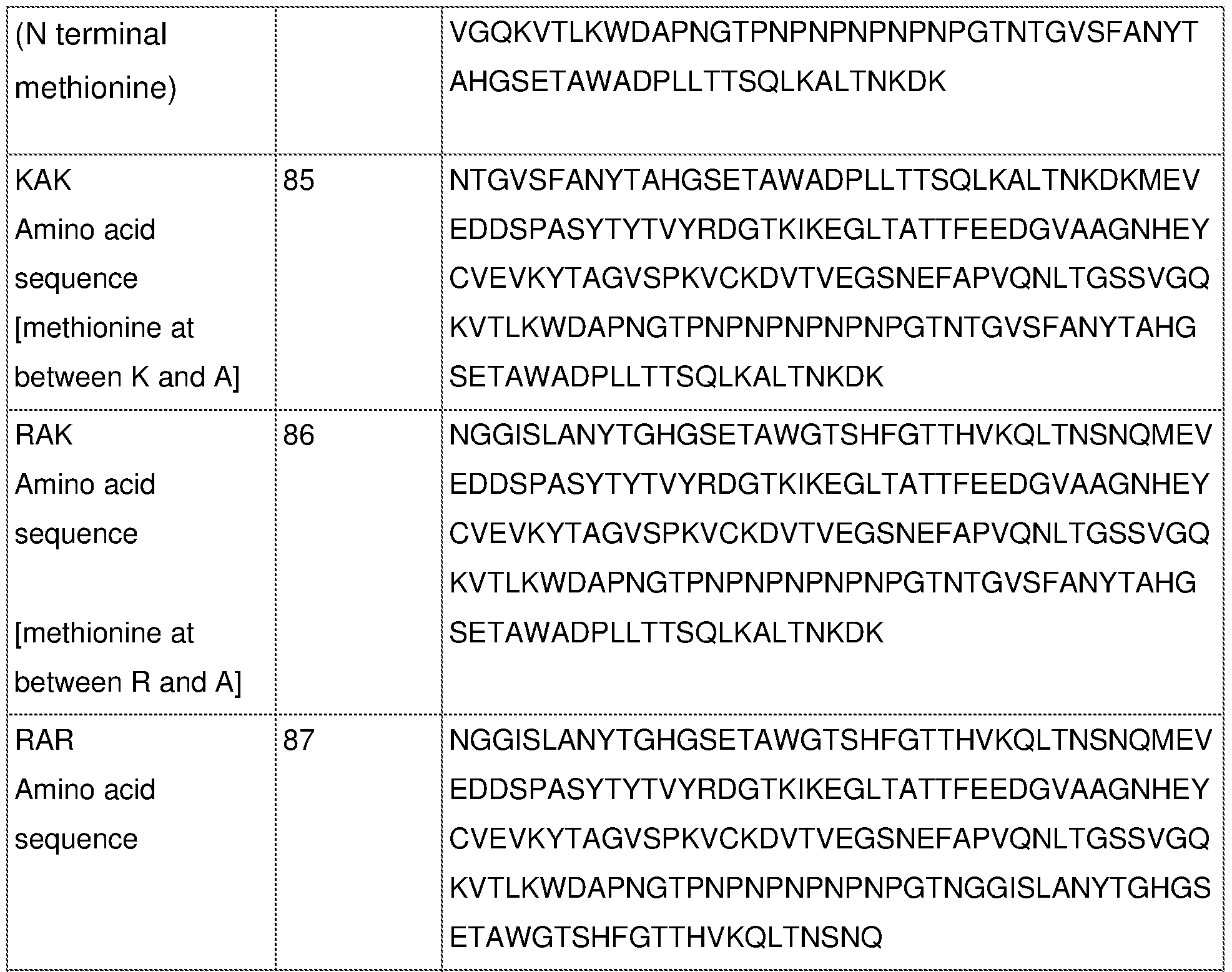
- the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 18, SEQ ID NO: 39 or any of SEQ ID NOs: 58 to 63; or SEQ ID NO: 81 to 87; or [0057] b) - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P.
- the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 4, 12, 16 or 20; - more preferably wherein the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 4; or [0058] c) - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P.
- ABSMs adhesin binding motifs
- the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 14; or [0059] d) - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P.
- the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 10 or SEQ ID NO: 108; or [0060] e) - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P.
- a linker region may be included between the amino acid sequence of a DUF2436 domain and the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain.
- ABSMs adhesin binding motifs
- the linker comprises or consists of the sequence EVEDDSP (SEQ ID NO: 109).
- the chimeric or fusion protein encoded by the RNA does not include a linker sequence between the DUF2436 domain and one or more adhesin domains.
- the amino acid sequence of an AMB2 domain as defined herein may further comprise at its N-terminus, the amino acid sequence of EVEDDSP (SEQ ID NO: 109), which is derived from the native P. gingivalis gingipain polyprotein sequence.
- EVEDDSP SEQ ID NO: 109
- the present disclosure provides basis for the generation of a chimeric or fusion protein from an RNA molecule, wherein the chimeric or fusion protein does not comprise an N terminal methionine residue.
- the RNA molecule comprises or consists of a nucleotide sequence encoding a protein comprising or consisting of the amino acid sequence of any one of: SEQ ID NO: 2, SEQ ID NO: 8 or SEQ ID NO: 38.
- the RNA molecule comprises or consists of a nucleotide sequence of any one of: a) SEQ ID NO: 48 or 57 b) SEQ ID NO: 45, 47 or 49 or SEQ ID NO: 41; c) SEQ ID NO: 46; d) SEQ ID NO: 44; e) SEQ ID NO: 42 1005272698
- the RNA comprises or consists of a nucleotide sequence of any one of: SEQ ID NO: 40, SEQ ID NO: 43 or SEQ ID NO: 50.
- the invention further provides for the use of any DNA polynucleotide described herein (and particularly any DNA polynucleotide comprising or consisting of a sequence exemplified in Table 1).
- the use of the DNA polynucleotide may be for obtaining an RNA polynucleotide of the invention.
- the present invention also provides a composition, including a pharmaceutical composition comprising an RNA as described herein.
- the composition comprises one or more pharmaceutically acceptable excipients.
- the RNA may comprise one or more agents for enabling delivery of the RNA to a mammalian cell, and thereby enabling translation of the RNA in the cell.
- the composition may comprise a combination of one or more of the RNA molecules described herein.
- the present invention further provides compositions comprising an RNA molecule as described herein, wherein the composition also comprises a lipid component.
- the RNA (e.g., RNA) vaccines of the disclosure can be formulated using one or more liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles [0070] In preferred embodiments, the RNA is formulated in a lipid nanoparticle.
- the RNA as described herein is the only polynucleotide species present in the composition or in the liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles.
- the polynucleotide as described herein is the only active ingredient present in the composition or in the liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles.
- the composition or liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles may comprise more than one RNA 1005272698
- composition or the liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles may comprise a single polynucleotide construct containing one or more RNA sequences as described herein (and thereby also encoding more than one chimeric or fusion protein amino acid sequence).
- the present invention contemplates the provision of compositions, liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles for delivering combinations of two or more of any of the RNA molecules described herein.
- the invention provides a lipid nanoparticle or other nanovehicle, such as nanopolymer, for delivery of the polynucleotide to a subject in need thereof.
- Lipid nanoparticles are well known in the art and are further described herein.
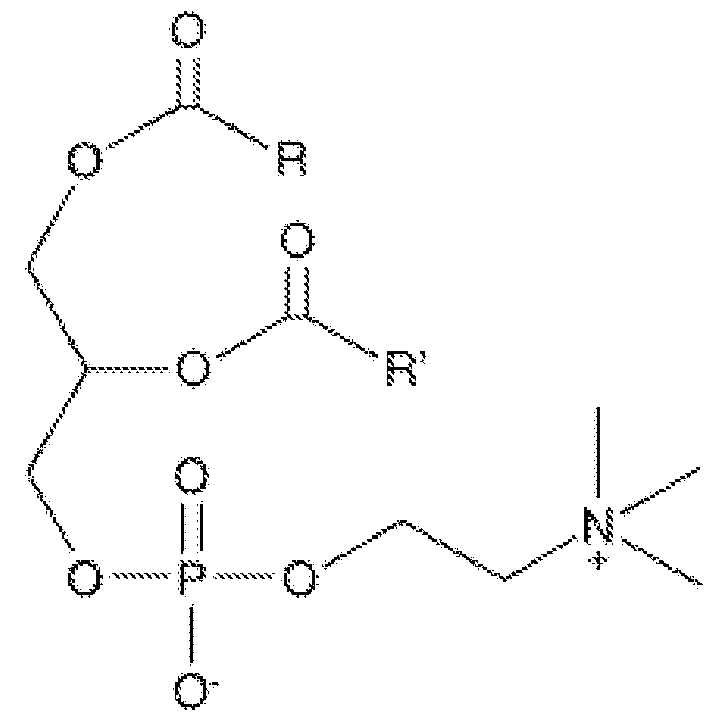
- the lipid nanoparticle comprises a cationic and/or ionisable lipid, a phospholipid, a PEG (or PEGylated) lipid, and a structural lipid.
- the lipid nanoparticle may comprise: - a cationic and/or ionisable lipid comprising from about 25 % to about 75 mol % of the total lipid present in the nanoparticle; - a sterol (structural lipid) comprising from about 5 mol % to about 60 mol % of the total lipid present in the nanoparticle; - a phospholipid comprising from about 5 mol % to about 50 mol % of the total lipid present in the nanoparticle; - a PEGylated lipid comprising from about 0.5 mol % to 20 mol % of the total lipid present in the nanoparticle.
- the lipid nanoparticle comprises: - an ionisable lipid in the form of [(4-hydroxybutyl)azanediyl]di(hexane-6,1-diyl) bis(2-hexyldecanoate) (ALC-0315), - a sterol in the form of cholesterol, 1005272698
- lipids are present in the lipid nanoparticle at molar lipid ratios (%) of 46.3 ALC-0315: 42.7 cholesterol : 9.4 DSPC : 1.6 ALC-0159, optionally in Tris/sucrose buffer (25 mM Tris pH 7.4, 8.8% sucrose w/v).
- the present invention also provides a method for producing a lipid nanoparticle comprising an RNA, encoding a protein, or a chimeric or fusion protein as described herein.
- the method comprises formulating any RNA molecule of the invention, with one or more lipids useful for producing a lipid nanoparticle.
- the lipid components comprise a phospholipid, a PEG lipid, and a structural lipid.
- the present invention also provides a nucleic acid construct or vector, comprising a polynucleotide as described herein.
- the vector may be any vector suitable for production of RNA from a DNA template.
- the vector may additionally comprise 3’UTR and 5’UTRs and polyadenine fragments.
- Examples of such vectors include: IVT RNA vector or similar vectors that comprise T7, T3 and SP6 signals for expression.
- the vector can be from a plasmid or produced through PCR or Phi29 DNA polymerase (e.g. GenomiPhiTM V2 DNA) or other bacterial constructs.
- the vector may be a self-amplifying RNA replicon, such as, but not limited to a self-amplifying RNA vector from an alphavirus, optionally Venezuelan Equine Encephalitis Virus (VEEV), bipartite VEEV, or variants thereof (including the TC83 mutated variant).
- VEEV Venezuelan Equine Encephalitis Virus
- capping of the polynucleotide may be performed using any commercially available capping reagent.
- Such reagents are known to the skilled person, such as the commercial capping reagent Cap1 from TriLink Biotechnologies Inc.
- Other capping reagents may be used, including but not limited to Cap 0 and Cap 2. 1005272698
- the present invention provides a method for eliciting an immune response to P. gingivalis in a subject in need thereof, the method comprising administering to the subject, a polynucleotide, vector, nanoparticle or composition described herein.
- the invention provides a method for eliciting an immune response to P. gingivalis in a subject in need thereof, the method comprising administering to the subject, a composition comprising: - an RNA as described herein, wherein said RNA is capable of being translated in a cell of a mammalian subject to produce the polypeptide encoded by the polynucleotide; - optionally, an agent for enabling delivery of the RNA into mammalian cells.
- the agent for delivery of the RNA into mammalian cells may be any suitable agent known to the skilled person for delivery of RNA. Such agents may include: cell penetrating peptides, lipid-based formulations.
- the invention provides a method for eliciting an immune response to P. gingivalis in a subject in need thereof, the method comprising administering to the subject, a nanoparticle composition comprising: - a lipid component; and - an RNA as described herein, wherein said RNA is capable of being translated in a cell of the subject to produce the polypeptide encoded by the polynucleotide.
- the invention also provides a method for producing a chimeric or fusion protein as described herein in a mammalian cell, the method comprising contacting the mammalian cell with a composition comprising: - an RNA as described herein, wherein said RNA is capable of being translated in the mammalian cell to produce the protein; - optionally, an agent for enabling delivery of the RNA into mammalian cells.
- the agent for delivery of the RNA into mammalian cells may be any suitable agent known to the skilled person for delivery of RNA. Such agents may include: cell penetrating peptides, lipid-based formulations. 1005272698
- the invention also provides a method for producing a chimeric or fusion protein as described herein in a mammalian cell, the method comprising contacting the mammalian cell with a nanoparticle composition, the composition comprising: - a lipid component; and - an RNA as described herein, wherein said RNA is capable of being translated in the mammalian cell to produce the protein.
- the lipid component comprises a cationic and/or ionisable lipid, a phospholipid, a PEG lipid, and a structural lipid.
- the invention also provides a method for delivering an RNA to a mammalian cell in a subject in need thereof, said method comprising administering to a subject in need thereof, a nanoparticle composition, the composition comprising: - a lipid component; and - an RNA comprising a polynucleotide sequence as described herein, wherein said RNA is capable of being translated in the mammalian cell to produce the chimeric or fusion protein described herein; wherein the administering comprises contacting said mammalian cell with the nanoparticle composition, thereby enabling delivery of the RNA to the mammalian cell.
- the lipid component comprises a cationic and/or ionisable lipid, a phospholipid, a PEG lipid, and a structural lipid.
- the ionisable lipid may be substituted or combined with an adjuvant lipidoid for enhancing RNA delivery.
- the present invention also provides the use of a polynucleotide, vector, or nanoparticle described herein, in the manufacture of a composition for eliciting an immune response to P. gingivalis in a subject.
- the present invention also provides the use of i) a lipid component as described herein, and ii) an RNA as described herein, in the manufacture of a composition for delivering the RNA to a mammalian cell in a subject in need thereof. 1005272698
- the present invention also provides a polynucleotide, vector, nanoparticle or composition as described herein, for use in eliciting an immune response to P. gingivalis in a subject.
- at least 80% identity should be taken to provide basis for “at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identity”.
- Antigen sequence mRNA sequence encoding protein antigen for expression in cell upon translation of mRNA; Poly(A) tail: polyadenosine tail for providing RNA stability and maximising translation.
- Figure 3 Expression and secretion by HeLa cells of mRNA constructs encoding chimeric proteins.
- m1 ⁇ N1-methylpseudouridine modified mRNA sequence.
- Figure 6 schematic of study protocol for determining in vivo efficacy of vaccines (therapeutic model).
- FIG. 8 Antibody responses in mice following alveolar bone loss experiments.
- F Heat-killed P. gingivalis IgG2a titres.
- m1 ⁇ N1- methylpseudouridine modified mRNA sequence.
- Figure 9 Expression and secretion of constructs encoding truncated antigens. A.
- KDAK-3S-AVQP mRNA encoding KDAK-3S-AVQP protein (SEQ ID NO: 6).
- m1 ⁇ N1-methylpseudouridine modified mRNA sequence.
- KA mRNA encoding KA protein (SEQ ID NO: 18).
- KD mRNA encoding KD protein (SEQ ID NO: 10).
- DA mRNA encoding DA protein (SEQ ID NO: 14).
- KDA21 mRNA encoding KDA ⁇ AMB3 protein (SEQ ID NO: 16).
- Figure 12 Immunogenicity of truncated antigens RDA and K.
- D Heat-killed
- Na ⁇ ve unvaccinated;
- K mRNA encoding K protein (SEQ ID NO: 8 ).
- KD mRNA encoding KD protein (SEQ ID NO: 10).
- Figure 13 schematic of study protocol for determining in vivo efficacy of vaccines in prophylaxis model.
- FIG. 16 Antibody responses in mice following alveolar bone loss experiments in the prophylaxis model.
- Table 2 exemplary RNA sequences of the invention Descriptor SEQ ID RNA Sequence No KDcAK1n 40 AAUACCGGAGUCAGCUUUGCAAACUAUACAGCGCAUGGAUCUG AGACCGCAUGGGCUGAUCCACUUCUGACUACUUCUACUGAA AGCACUCACUAAUAAGGACAAAUGGGGAGACAAUACGGGUUACC AGUUCUUGUUGGAUGCCGAUCACAAUACAUUCGGAAGUGUCAU UCCGGCAACCGGUCCUCUCUUUACCGGAACAGCUUCUUCCAAU CUUUACAGUGCGAACUUCGAGUAUUUGAUCCCGGCCAAUGCCG AUCCUGUUGUUACUACACAGAAUAUUAUCGUUACAGGACAGGG UGAAGUUGUAAUCCCCGGUGGUGUGUUUACGACUAUUGCAUUACG AACCCGGAACCUGCAUCCGGAAAGAUGUGGAUCGCAGGAUG 1005272698
- ABM3 54 CCGAAUCCGAAUCCAAAUCCGAAUCCGAAUCCGGGAACA ABM 2+1 55 GCAAGCUAUACCUACACGGUGUAUCGUGACGGCACGAAGAUCA AGGAAGGUCUGACAGCUACGACAUUCGAAGAAGACGGUGUAGC UGCAGGCAAUCAUGAGUAUUGCGUGGAAGUUAAGUACACAGCC GGCGUAUCUCCGAAGGUAUGUAAAGACGUUACGGUAGAAGGAU CCAAUGAAUUUGCUCCUGUACAGAACCUGACCGGUAGUUCAGU AGGUCAGAAAGUAACGCUUAAGUGGGAUGCACCUAAUGGUACC ABM 2+1+3 56 GCAAGCUAUACCUACACGGUGUAUCGUGACGGCACGAAGAUCA AGGAAGGUCUGACAGCUACGACAUUCGAAGAAGACGGUGUAGC UGCAGGCAAUCAUGAGUAUUGCGUGGAAGUUAAGUACACAGCC GGCGUAUCUCCGAAGGUAUGUAAAGACGUUACGGUAG
- RNA vaccine encoding KDcAK1n did not express well and therefore KDcAK1n was not a preferred candidate for use in an RNA vaccine.
- the inventors found that robust immune responses were obtained when providing an RNA encoding alternative chimeric proteins containing various domains derived from P. gingivalis Arg- or Lys-gingipain proteins. 1005272698
- the present invention is therefore concerned with the design of new RNA vaccines encoding protein antigens, or chimeric or fusion proteins comprising protein antigens, for use in inducing an immune response to P. gingivalis, and methods and uses comprising the same.
- Gingipains [0130] The pathogenicity of P. gingivalis is attributed to a number of surface-associated virulence factors that include cysteine proteinases (gingipains), fimbriae, haem-binding proteins, and outer membrane transport proteins amongst others. In particular, the extracellular Arg- and Lys-specific proteinases ‘gingipains’ (RgpA/B and Kgp) of P.
- gingivalis have been implicated as major virulence factors that are critical for colonisation, penetration into host tissue, dysregulation of the immune response, dysbiosis and disease.
- the gingipains in particular the Lys-specific proteinase Kgp are essential for P. gingivalis to induce alveolar bone resorption in the mouse periodontitis model.
- the gingipains have also been found in gingival tissue at sites of severe periodontitis at high concentrations proximal to the subgingival plaque and at lower concentrations at distal sites deeper into the gingival tissue.
- Lys-specific and Arg-specific proteinases have been shown to degrade a variety of host proteins in vitro, e.g., fibrinogen, fibronectin, and laminin.
- Plasma host defence and regulatory proteinase inhibitors ⁇ -trypsin, ⁇ 2- macroglobulin, anti-chymotrypsin, antithrombin III and antiplasmin are also degraded by Lys- and Arg- proteinases from P. gingivalis. This has led to the development of a cogent mechanism to explain the keystone role played by P. gingivalis in the development of chronic periodontitis.
- the RgpA, RgpB and Kgp genes all encode an N-terminal signal peptide of ⁇ 22 amino acids in length, an unusually long propeptide of ⁇ 200 amino acids in length, and a catalytic domain of ⁇ 480 amino acids.
- HA hemagglutinin-adhesin
- ABMs adhesin binding domains
- DUF2436 conserved Pfam Domain of Unknown Function
- CADs C-terminal adhesin domains or cleaved adhesin domains
- Kgp comprises (N terminus to C terminus): a catalytic domain, a first ABM (ABM1), DUF2436, a domain comprising ABM2, ABM1, ABM3, two CAD domains (termed K1 and K2), a further domain comprising ABM2 and ABM1, a further CAD domain (termed K3), ABM2, and a C-terminal domain.
- ABMs 1, 2 and 3 will be understood to generally refer to the ABMs found in the order ABM2, ABM1 and ABM3 in the sequence immediately C terminal to DUF2436 of Kgp, as depicted in Figure 1.
- the catalytic domains of RgpB and RgpA share a high-degree of sequence homology. However, RgpB lacks the HA domains and is located in a monomeric form on the outer membrane.
- HA domains have been alternatively described as C- terminal adhesin domains or cleaved adhesin domains (CADs) and some are DUF (“Domain of Unknown Function”) 2436 domains (conserved Pfam Domain of Unknown Function; IPR018832).
- DUF Domain of Unknown Function
- IPR018832 Pfam Domain of Unknown Function
- RgpA-Kgp proteinase-adhesin complex has been designated the RgpA-Kgp proteinase-adhesin complex (previously referred to as the PrtR-PrtK proteinase-adhesin complex).
- the complex is composed of a 45kDa Arg- specific calcium-stabilised cysteine proteinase and seven sequence-related adhesin domains, [0137]
- a Lys-gingipain catalytic domain may also be referred to as a KAS domain or PAS domain.
- an Arg-gingipain catalytic domain may also be referred to as a RAS domain or PAS domain.
- the catalytic domain of the Lys-gingipain or Arg-gingipains is located in the N-terminal ⁇ 480 amino acid region of the protein.
- the active site within the catalytic domain is typically located at amino acid residues 426-446 (for RgpA) and 432-453 (for Kgp).
- an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis will be understood to typically refer to the region of an Arg- or Lys-gingipain that is C- terminal to the catalytic or active site domain.
- the adhesin domain also referred to as 1005272698
- nucleic acids typically comprise a Domain of Unknown Function (DUF) domain (especially DUF 2436 conserved Pfam Domain of Unknown Function; IPR018832) and several adhesin binding motifs (ABM) domains and a cleaved adhesin domain (CAD).
- DUF Domain of Unknown Function
- ABSM adhesin binding motifs
- CAD cleaved adhesin domain
- Nucleic acids in its broadest sense, includes any compound and/or substance that comprise a polymer of nucleotides. These polymers are often referred to as polynucleotides.
- the polynucleotides of the invention are in the form of an RNA molecule, preferably an mRNA.
- RNA messenger RNA
- RNA refers to any polynucleotide which encodes a polypeptide of interest and which is capable of being translated to produce the encoded polypeptide of interest in vitro, in vivo, in situ or ex vivo.
- RNA messenger RNA
- any of the RNA polynucleotides encoded by a DNA identified by a particular sequence identification number may also comprise the corresponding RNA (e.g., RNA) sequence encoded by the DNA, where each “T” of the DNA sequence is substituted with “U.”
- the basic components of an RNA molecule include at least a coding region, a 5′UTR, a 3′UTR, a 5′ cap and a poly-A tail.
- Polynucleotides of the present disclosure may function as RNA but can be distinguished from wild-type RNA in their functional and/or structural design features, which serve to overcome existing problems of effective polypeptide expression using nucleic-acid based therapeutics.
- a “5′ untranslated region” refers to a region of an RNA that is directly upstream (i.e., 5′) from the start codon (i.e., the first codon of an RNA transcript translated by a ribosome) that does not encode a polypeptide.
- a “3′ untranslated region” (3′UTR) refers to a region of an RNA that is directly downstream (i.e., 3′) from the stop codon (i.e., the codon of an RNA transcript that signals a termination of translation) that does not encode a polypeptide.
- An “open reading frame” is a continuous stretch of DNA beginning with a start codon (e.g., methionine (ATG)), and ending with a stop codon (e.g., TAA, TAG or TGA) and encodes a polypeptide. 1005272698
- start codon e.g., methionine (ATG)
- stop codon e.g., TAA, TAG or TGA
- a “polyA tail” is a region of RNA (typically mRNA) that is downstream, e.g., directly downstream (i.e., 3′), from the 3′ UTR that contains multiple, sometimes consecutive adenosine monophosphates.
- a polyA tail may contain 10 to 300 adenosine monophosphates.
- a polyA tail may contain 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, 110, 120, 130, 140, 150, 160, 170, 180, 190, 200, 210, 220, 230, 240, 250, 260, 270, 280, 290 or 300 adenosine monophosphates.
- a polyA tail contains 50 to 250 adenosine monophosphates.
- a segmented polyA tail may be used (typically segments of consecutive adenosine monophosphates separated via a short spacer region between segments).
- the poly(A) tail functions to protect mRNA from enzymatic degradation, e.g., in the cytoplasm, and aids in transcription termination, export of the mRNA from the nucleus and translation.
- a polynucleotide includes 200 to 3,000 nucleotides.
- a polynucleotide may include 200 to 500, 200 to 1000, 200 to 1500, 200 to 3000, 500 to 1000, 500 to 1500, 500 to 2000, 500 to 3000, 1000 to 1500, 1000 to 2000, 1000 to 3000, 1500 to 3000, or 2000 to 3000 nucleotides.
- the present invention also contemplates the use of one or more structural and/or chemical modifications or alterations which impart useful properties to the polynucleotide including, in some embodiments, the lack of a substantial induction of the innate immune response of a cell into which the polynucleotide is introduced.
- modified RNA molecules of the present invention may also be termed “mmRNA.”
- mmRNA modified RNA molecules of the present invention
- a “structural” feature or modification is one in which two or more linked nucleotides are inserted, deleted, duplicated, inverted or randomized in a polynucleotide, primary construct or mRNA without significant chemical modification to the nucleotides themselves. Because chemical bonds will necessarily be broken and reformed to effect a structural modification, structural modifications are of a chemical nature and hence are chemical modifications. However, structural modifications will result in a different sequence of nucleotides. For example, the polynucleotide “ATCG” may be chemically modified to “AT-5meC-G”.
- RNA molecules of the invention may also comprise an 5’ terminal cap.
- the 5′ terminal cap is 7mG(5′)ppp(5′)NlmpNp although it will be 1005272698
- the RNA molecule comprises at least one chemical modification.
- chemical modification and “chemically modified” refer to modification with respect to adenosine (A), guanosine (G), uridine (U), thymidine (T) or cytidine (C) ribonucleosides or deoxyribnucleosides in at least one of their position, pattern, percent or population. Generally, these terms do not refer to the ribonucleotide modifications in naturally occurring 5′-terminal RNA cap moieties.
- RNA polynucleotides e.g., RNA polynucleotides, such as mRNA polynucleotides
- a particular region of a polynucleotide contains one, two or more (optionally different) nucleoside or nucleotide modifications.
- a modified RNA polynucleotide e.g., a modified RNA polynucleotide
- a modified RNA polynucleotide introduced to a cell or organism, exhibits reduced degradation in the cell or organism, respectively, relative to an unmodified polynucleotide.
- a modified RNA polynucleotide e.g., a modified RNA polynucleotide
- introduced into a cell or organism may exhibit reduced immunogenicity in the cell or organism, respectively (e.g., a reduced innate response).
- Modifications of polynucleotides include, without limitation, those described herein.
- Polynucleotides may comprise modifications that are naturally-occurring, non-naturally-occurring or the polynucleotide may comprise a combination of naturally-occurring and non-naturally- occurring modifications.
- Polynucleotides may include any useful modification, for example, of a sugar, a nucleobase, or an internucleoside linkage (e.g., to a linking phosphate, to a phosphodiester linkage or to the phosphodiester backbone).
- Polynucleotides e.g., RNA polynucleotides, such as mRNA polynucleotides
- RNA polynucleotides such as mRNA polynucleotides
- polynucleotides in some embodiments, comprise non-natural modified nucleotides that are introduced during synthesis or post-synthesis of the polynucleotides to achieve desired functions or properties.
- the modifications may be present on internucleotide linkages, purine or 1005272698
- the modification may be introduced with chemical synthesis or with a polymerase enzyme at the terminal of a chain or anywhere else in the chain. Any of the regions of a polynucleotide may be chemically modified.
- the present disclosure provides for modified nucleosides and nucleotides of a polynucleotide (e.g., RNA polynucleotides, such as mRNA polynucleotides).
- nucleoside refers to a compound containing a sugar molecule (e.g., a pentose or ribose) or a derivative thereof in combination with an organic base (e.g., a purine or pyrimidine) or a derivative thereof (also referred to herein as “nucleobase”).
- a nucleotide refers to a nucleoside, including a phosphate group. Modified nucleotides may by synthesized by any useful method, such as, for example, chemically, enzymatically, or recombinantly, to include one or more modified or non-natural nucleosides. Polynucleotides may comprise a region or regions of linked nucleosides.
- Modified nucleotide base pairing encompasses not only the standard adenosine-thymine, adenosine-uracil, or guanosine-cytosine base pairs, but also base pairs formed between nucleotides and/or modified nucleotides comprising non-standard or modified bases, wherein the arrangement of hydrogen bond donors and hydrogen bond acceptors permits hydrogen bonding between a non-standard base and a standard base or between two complementary non-standard base structures.
- Non-standard base pairing is the base pairing between the modified nucleotide inosine and adenine, cytosine or uracil. Any combination of base/sugar or linker may be incorporated into polynucleotides of the present disclosure.
- the at least one chemical modification may be selected from pseudouridine, N1-methylpseudouridine, N1-ethylpseudouridine, 2-thiouridine, 4′-thiouridine, 5- methylcytosine, 5-methyluridine, 2-thio-1-methyl-1-deaza-pseudouridine, 2-thio-1- methyl-pseudouridine, 2-thio-5-aza-uridine, 2-thio-dihydropseudouridine, 2-thio- dihydrouridine, 2-thio-pseudouridine, 4-methoxy-2-thio-pseudouridine, 4-methoxy- pseudouridine, 4-thio-1-methyl-pseudouridine, 4-thio-pseudouridine, 5-aza-uridine, dihydropseudouridine, 5-methoxyuridine and 2′-O-methyl uridine.
- the chemical modification is in the 5-position of the uracil. In some embodiments, the chemical modification is a N1-methylpseudouridine. In some embodiments, the chemical modification is a N1-ethylpseudouridine. In some embodiments, polynucleotides include 1005272698
- polynucleotides e.g., RNA polynucleotides, such as mRNA polynucleotides
- RNA polynucleotides are uniformly modified (e.g., fully modified, modified throughout the entire sequence) for a particular modification.
- a polynucleotide can be uniformly modified with 5-methyl-cytidine (m5C), meaning that all cytosine residues in the RNA sequence are replaced with 5-methyl-cytidine (m5C).
- nucleobases and nucleosides having a modified cytosine include N4-acetyl-cytidine (ac4C), 5-methyl-cytidine (m5C), 5-halo-cytidine (e.g., 5-iodo- cytidine), 5-hydroxymethyl-cytidine (hm5C), 1-methyl-pseudoisocytidine, 2-thio-cytidine (s2C), and 2-thio-5-methyl-cytidine.
- a modified nucleobase is a modified uridine.
- exemplary nucleobases and In some embodiments, a modified nucleobase is a modified cytosine.
- nucleosides having a modified uridine include 5-cyano uridine, and 4′-thio uridine.
- a modified nucleobase is a modified adenine.
- Exemplary nucleobases and nucleosides having a modified adenine include 7-deaza-adenine, 1- methyl-adenosine (m1A), 2-methyl-adenine (m2A), and N6-methyl-adenosine (m6A).
- a modified nucleobase is a modified guanine.
- exemplary nucleobases and nucleosides having a modified guanine include inosine (I), 1-methyl- inosine (m1I), wyosine (imG), methylwyosine (mimG), 7-deaza-guanosine, 7-cyano-7- deaza-guanosine (preQO), 7-aminomethyl-7-deaza-guanosine (preQ1), 7-methyl- guanosine (m7G), 1-methyl-guanosine (mlG), 8-oxo-guanosine, 7-methyl-8-oxo- guanosine.
- the polynucleotides of the present disclosure may be partially or fully modified along the entire length of the molecule.
- one or more or all or a given type of nucleotide e.g., purine or pyrimidine, or any one or more or all of A, G, U, C
- all nucleotides X in a polynucleotide of the present disclosure or in a given 1005272698
- X may any one of nucleotides A, G, U, C, or any one of the combinations A+G, A+U, A+C, G+U, G+C, U+C, A+G+U, A+G+C, G+U+C or A+G+C.
- the polynucleotide may contain from about 1% to about 100% modified nucleotides (either in relation to overall nucleotide content, or in relation to one or more types of nucleotide, i.e., any one or more of A, G, U or C) or any intervening percentage (e.g., from 1% to 20%, from 1% to 25%, from 1% to 50%, from 1% to 60%, from 1% to 70%, from 1% to 80%, from 1% to 90%, from 1% to 95%, from 10% to 20%, from 10% to 25%, from 10% to 50%, from 10% to 60%, from 10% to 70%, from 10% to 80%, from 10% to 90%, from 10% to 95%, from 10% to 100%, from 20% to 25%, from 20% to 50%, from 20% to 60%, from 20% to 70%, from 20% to 80%, from 20% to 90%, from 20% to 95%, from 20% to 100%, from 50% to 60%, from 50% to 70%, from 50% to 80%, from 50% to 90%, from 20% to 95%, from 20% to 100%, from
- the polynucleotides may contain at a minimum 1% and at maximum 100% modified nucleotides, or any intervening percentage, such as at least 5% modified nucleotides, at least 10% modified nucleotides, at least 25% modified nucleotides, at least 50% modified nucleotides, at least 80% modified nucleotides, or at least 90% modified nucleotides.
- the polynucleotides may contain a modified pyrimidine such as a modified uracil or cytosine.
- At least 5%, at least 10%, at least 25%, at least 50%, at least 80%, at least 90% or 100% of the uracil in the polynucleotide is replaced with a modified uracil (e.g., a 5-substituted uracil).
- the modified uracil can be replaced by a compound having a single unique structure, or can be replaced by a plurality of compounds having different structures (e.g., 2, 3, 4 or more unique structures).
- the modified nucleobase is a modified uracil.
- Exemplary nucleobases and nucleosides having a modified uracil include pseudouridine ( ⁇ ), pyridin- 1005272698
- the modified nucleobase is a modified cytosine.
- exemplary nucleobases and nucleosides having a modified cytosine include 5-aza- cytidine, 6-aza-cytidine, pseudoisocytidine, 3-methyl-cytidine, N4-acetyl-cytidine, 5- formyl-cytidine, N4-methyl-cytidine, 5-methyl-cytidine, 5-halo-cytidine (e.g., 5-iodo- cytidine), 5-hydroxymethyl-cytidine, 1-methyl-pseudoisocytidine, pyrrolo-cytidine, pyrrolo-pseudoisocytidine, 2-thio-cytidine, 2-thio-5-methyl-cytidine, 4-thio- pseudoisocytidine, 4-thio-1-methyl-pseudoisocytidine, 4-thio-1-methyl-pseudoi
- the modified nucleobase is a modified adenine.
- exemplary nucleobases and nucleosides having a modified adenine include 2-amino- purine, 2, 6-diaminopurine, 2-amino-6-halo-purine (e.g., 2-amino-6-chloro-purine), 6- halo-purine (e.g., 6-chloro-purine), 2-amino-6-methyl-purine, 8-azido-adenosine, 7- deaza-adenine, 7-deaza-8-aza-adenine, 7-deaza-2-amino-purine, 7-deaza-8-aza-2- amino-purine, 7-deaza-2,6-diaminopurine, 7-deaza-8-aza-2,6-diaminopurine, 1-methyl- adenosine, 2-methyl-adenine, N6-methyl-adenosine, 2-methylthio-
- the modified nucleobase is a modified guanine.
- exemplary nucleobases and nucleosides having a modified guanine include inosine, 1- methyl-inosine, wyosine, methylwyosine, 4-demethyl-wyosine, isowyosine (imG2), wybutosine, peroxywybutosine, hydroxywybutosine, undermodified hydroxywybutosine, 7-deaza-guanosine, queuosine, epoxyqueuosine, galactosyl-queuosine (galQ), mannosyl-queuosine, 7-cyano-7-deaza-guanosine, 7-aminomethyl-7-deaza-guanosine, archaeosine, 7-deaza-8-aza-guanosine, 6-thio-guanosine, 6-thio-7-deaza-guanosine,
- the RNA (e.g., RNA) vaccines comprise a 5′UTR element, an optionally codon optimized open reading frame, and a 3′UTR element, a poly(A) sequence and/or a polyadenylation signal wherein the RNA is not chemically modified.
- Polynucleotides of the present disclosure in some embodiments, are codon optimized. Codon optimization methods are known in the art and may be used as provided herein.
- Codon optimization may be used to match codon frequencies in target and host organisms to ensure proper folding; bias GC content to increase RNA stability or reduce secondary structures; minimize tandem repeat codons or base runs that may impair gene construction or expression; customize transcriptional and translational control regions; insert or remove protein trafficking sequences; remove/add post translation modification sites in encoded protein (e.g. glycosylation sites); add, remove or shuffle protein domains; insert or delete restriction sites; modify ribosome binding sites and RNA degradation sites; adjust translational rates to allow the various domains of the protein to fold properly; or to reduce or eliminate problem secondary structures within the polynucleotide.
- encoded protein e.g. glycosylation sites
- add, remove or shuffle protein domains add or delete restriction sites
- modify ribosome binding sites and RNA degradation sites adjust translational rates to allow the various domains of the protein to fold properly; or to reduce or eliminate problem secondary structures within the polynucleotide.
- Codon optimization tools, algorithms and services are known in the art—non-limiting examples include services from GeneArt (Life Technologies), DNA2.0 (Menlo Park Calif.) and/or proprietary methods.
- the open reading frame (ORF) sequence is optimized using optimization algorithms.
- a codon optimized sequence shares less than 95% sequence identity, less than 90% sequence identity, less than 85% sequence identity, less than 80% sequence identity, or less than 75% sequence identity to a naturally- occurring or wild-type sequence (e.g., a naturally-occurring or wild-type RNA sequence 1005272698
- a codon-optimized sequence shares between 65% and 85% (e.g., between about 67% and about 85%, or between about 67% and about 80%) sequence identity to a naturally-occurring sequence or a wild-type sequence (e.g., a naturally-occurring or wild-type RNA sequence encoding a polypeptide or protein of interest (e.g., an antigenic protein or polypeptide)).
- a codon- optimized sequence shares between 65% and 75%, or about 80% sequence identity to a naturally-occurring sequence or wild-type sequence (e.g., a naturally-occurring or wild- type RNA sequence encoding a polypeptide or protein of interest (e.g., an antigenic protein or polypeptide)).
- a codon-optimized RNA e.g., mRNA
- the G/C-content of nucleic acid molecules may influence the stability of the RNA.
- RNA having an increased amount of guanine (G) and/or cytosine (C) residues may be functionally more stable than nucleic acids containing a large amount of adenine (A) and thymine (T) or uracil (U) nucleotides.
- WO02/098443 discloses a pharmaceutical composition containing an RNA stabilized by sequence modifications in the translated region. Due to the degeneracy of the genetic code, the modifications work by substituting existing codons for those that promote greater RNA stability without changing the resulting amino acid. The approach is limited to coding regions of the RNA.
- Naturally-occurring eukaryotic RNA molecules have been found to contain stabilizing elements, including, but not limited to untranslated regions (UTR) at their 5′- end (5′UTR) and/or at their 3′-end (3′UTR), in addition to other structural features, such as a 5′-cap structure or a 3′-poly(A) tail. Both the 5′UTR and the 3′UTR are typically transcribed from the genomic DNA and are elements of the premature RNA. Characteristic structural features of mature RNA, such as the 5′-cap and the 3′-poly(A) tail are usually added to the transcribed (premature) RNA during RNA processing.
- stabilizing elements including, but not limited to untranslated regions (UTR) at their 5′- end (5′UTR) and/or at their 3′-end (3′UTR), in addition to other structural features, such as a 5′-cap structure or a 3′-poly(A) tail.
- the 3′-poly(A) tail is typically a stretch of adenine nucleotides added to the 3′-end of the transcribed RNA. It can comprise up to about 400 adenine nucleotides. In some embodiments the length of the 3′-poly(A) tail may be an essential element with respect to the stability of the individual RNA. 1005272698
- the RNA (e.g., mRNA) vaccine may include one or more stabilizing elements.
- Stabilizing elements may include for instance a histone stem-loop.
- a stem-loop binding protein (SLBP) a 32 kDa protein has been identified. It is associated with the histone stem-loop at the 3′-end of the histone messages in both the nucleus and the cytoplasm. Its expression level is regulated by the cell cycle; it peaks during the S- phase, when histone RNA levels are also elevated.
- the protein has been shown to be essential for efficient 3′-end processing of histone pre-RNA by the U7 snRNP.
- RNA binding domain of SLBP is conserved through metazoa and protozoa; its binding to the histone stem-loop depends on the structure of the loop.
- the minimum binding site includes at least three nucleotides 5′ and two nucleotides 3′ relative to the stem-loop.
- the RNA (e.g., mRNA) vaccines include a coding region, at least one histone stem-loop, and optionally, a poly(A) sequence or polyadenylation signal.
- the poly(A) sequence or polyadenylation signal generally should enhance the expression level of the encoded protein.
- the encoded protein in some embodiments, is not a histone protein, a reporter protein (e.g. Luciferase, GFP, EGFP, ⁇ -Galactosidase, EGFP), or a marker or selection protein (e.g. alpha-Globin, Galactokinase and Xanthine:guanine phosphoribosyl transferase (GPT)).
- a reporter protein e.g. Luciferase, GFP, EGFP, ⁇ -Galactosidase, EGFP
- a marker or selection protein e.g. alpha-Globin, Galactokinase and Xanthine:guanine phosphoribosyl transferase (GPT)
- the combination of a poly(A) sequence or polyadenylation signal and at least one histone stem-loop acts synergistically to increase the protein expression beyond the level observed with either of the individual elements. It has been found that the synergistic effect of the combination of poly(A) and at least one histone stem-loop does not depend on the order of the elements or the length of the poly(A) sequence.
- the RNA (e.g., mRNA) vaccine does not comprise a histone downstream element (HDE).
- Histone downstream element includes a purine-rich polynucleotide stretch of approximately 15 to 20 nucleotides 3′ of naturally occurring stem-loops, representing the binding site for the U7 snRNA, which is involved in processing of histone pre-RNA into mature histone RNA. Ideally, the inventive nucleic acid does not include an intron. 1005272698
- the RNA (e.g., mRNA) vaccine may or may not contain an enhancer and/or promoter sequence, which may be modified or unmodified or which may be activated or inactivated.
- the histone stem-loop is generally derived from histone genes, and includes an intramolecular base pairing of two neighbored partially or entirely reverse complementary sequences separated by a spacer, including (e.g., consisting of) a short sequence, which forms the loop of the structure.
- the unpaired loop region is typically unable to base pair with either of the stem loop elements. It occurs more often in RNA, as is a key component of many RNA secondary structures, but may be present in single-stranded DNA as well.
- the stem-loop structure generally depends on the length, number of mismatches or bulges, and base composition of the paired region. In some embodiments, wobble base pairing (non-Watson-Crick base pairing) may result. In some embodiments, the at least one histone stem-loop sequence comprises a length of 15 to 45 nucleotides.
- the RNA (e.g., mRNA) vaccine may have one or more AU-rich sequences removed. These sequences, sometimes referred to as AURES are destabilizing sequences found in the 3′UTR. The AURES may be removed from the RNA (e.g., mRNA) vaccines.
- the AURES may remain in the RNA (e.g., mRNA) vaccine.
- the RNA of the invention eg mRNA
- the RNA of the invention may comprise a ribosome skipping sequence, such as a 2A skipping sequence.
- a ribosome skipping sequence such as a 2A skipping sequence.
- an mRNA of the invention may encode two or more of the domains K, D, A (including A ⁇ AMB3) or R as defined elsewhere herein and also defined in Table 1, or may encode two or more of the proteins exemplified in Table 1 as being proteins that can be encoded by an RNA sequence of the invention.
- an mRNA of the invention could encode one or more of a KA chimeric protein, a DA chimeric protein, an RA chimeric protein, an AR chimeric protein, an AK chimeric protein, an AD chimeric protein, a KDA chimeric protein, an RDA chimeric protein, a DAR chimeric protein, a DAK chimeric protein or combinations thereof.
- 2A peptide sequence for use to introduce ribosome skipping include the T2A or T2A-like sequences derived from Thosea asigna virus and from Porcine teschovirus-12A. 1005272698
- polypeptide means a polymer of amino acid residues (natural or unnatural) linked together most often by peptide bonds.
- the term, as used herein, refers to proteins, polypeptides, and peptides of any size, structure, or function. In some instances the polypeptide encoded is smaller than about 50 amino acids and the polypeptide is then termed a peptide.
- polypeptides include gene products, naturally occurring polypeptides, synthetic polypeptides, homologs, orthologs, paralogs, fragments and other equivalents, variants, and analogs of the foregoing.
- a polypeptide may be a single molecule or may be a multi-molecular complex such as a dimer, trimer or tetramer. They may also comprise single chain or multichain polypeptides such as antibodies or insulin and may be associated or linked. Most commonly disulfide linkages are found in multichain polypeptides.
- polypeptide may also apply to amino acid polymers in which one or more amino acid residues are an artificial chemical analogue of a corresponding naturally occurring amino acid.
- a chimeric or fusion protein refers to a polypeptide that comprises amino acid sequences that are not arranged in the same spatial configuration as occurs in nature.
- the chimeric or fusion protein encoded by the polynucleotides of the invention comprises portions of an Arg or Lys- gingipain from P. gingivalis, which are in a different spatial arrangement to full length gingipain.
- the RNA preferably encodes chimeric or fusion proteins comprising various domains (as defined herein), derived from a P. gingivalis gingipain.
- the domains may be directly joined within the chimeric or fusion protein, or the chimeric or fusion protein may comprise linkers for joining the domains. 1005272698
- linker for joining amino acid sequences are well known to persons of skill in the art.
- the linker is non-immunogenic.
- the linker is comprised of amino acids, and may therefore be termed a peptide linker.
- a linker is usually a peptide having a length of up to 20 amino acids, although may be longer.
- the term “linked to” or “fused to” refers to a covalent bond, e.g., a peptide bond, formed between two moieties. Accordingly, in the context of the present invention the linker may have a length of 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 or 22 or more amino acids.
- the chimeric or fusion proteins encoded by the RNAs of the invention may comprise a linker between the amino acid sequence of a P. gingivalis gingipain active site, and the amino acid sequence of the adhesin domain of a P. gingivalis gingipain.
- linkers have the advantage that they can make it more likely that the different polypeptides of the fusion protein fold independently and behave as expected.
- Suitable linkers may be up to 50 amino acids in length, although less than 20, less than 15 or less than five amino acids is preferred.
- the linker may function to bring the domains into a closer spatial arrangement than normally observed in a P. gingivalis trypsin-like enzyme. Alternatively, it may space domains apart.
- linker for use in protein constructs, including those with minimal impact on solubility are known in the art.
- the linker may be any linker known in the art to the skilled person and may be a flexible linker (such as those comprising repeats of glycine and serine residues), a rigid linker (such as those comprising glutamic acid and lysine residues, flanking alanine repeats) and/or a cleavable linker (such as sequences that are susceptible by protease cleavage). Examples of such linkers are known to the skilled person and are described for example, in Chen et al., (2013) Advanced Drug Delivery Reviews, 65: 1357-1369.
- Useful linkers include glycine-serine (GlySer) linkers, which are well-known in the art and comprise glycine and serine units combined in various orders. Examples include, but are not limited to, (GS), (GSGGS)n (SEQ ID NO: 88), (GGGS)n (SEQ ID NO: 89) and (GGGGS)n (SEQ ID NO: 90), where n is an integer of at least one, typically an integer between 1 and about 10, for example, between 1 and about 8, between 1 and about 6, or between 1 and about 5.
- the peptide linker may include the amino acids glycine and serine in various lengths and combinations. In some aspects, the peptide linker can 1005272698
- the peptide linker can include the amino acid sequence GGGGS (a linker of 6 amino acids in length, SEQ ID NO: 90) or even longer.
- the linker may be a series of repeating glycine and serine residues (GS) of different lengths, i.e., (GS)n where n is any number from 1 to 15 or more.
- the linker may be (GS)3 (i.e., GSGSGS, SEQ ID NO: 91) or longer (GS)11 or longer. It will be appreciated that n can be any number including 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 or more. Fusion proteins having linkers of such length are included within the scope of the present invention.
- the linker may be a series of repeating glycine residues separated by serine residues.
- (GGGGS)3 i.e., the linker may comprise the amino acid sequence GGGGSGGGGSGGGGS, (G4S)3, SEQ ID NO: 92) and variations thereof.
- the peptide linker can include the amino acid sequence GGGGS (a linker of 6 amino acids in length) or even longer.
- the linker may a series of repeating glycine and serine residues (GS) of different lengths, i.e., (GS)n where n is any number from 1 to 15 or more.
- the linker may be (GS)3 (i.e., GSGSGS, SEQ ID NO: 91) or longer (GS)11 or longer. It will be appreciated that n can be any number including 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 or more.
- polypeptide variant refers to molecules which differ in their amino acid sequence from a native or reference sequence.
- the amino acid sequence variants may possess substitutions, deletions, and/or insertions at certain positions within the amino acid sequence, as compared to a native or reference sequence.
- variants will possess at least about 50% identity (homology) to a native or reference sequence, and preferably, they will be at least about 80%, more preferably at least about 90% identical (homologous) to a native or reference sequence.
- the present invention contemplates several types of compositions which encode polypeptides, including variants and derivatives. These include substitutional, insertional, deletion and covalent variants and derivatives.
- derivative is used 1005272698
- RNAs of the invention encoding polypeptides containing substitutions, insertions and/or additions, deletions and covalent modifications with respect to reference sequences, in particular the polypeptide sequences disclosed herein, are included within the scope of this invention.
- sequence tags or amino acids such as one or more lysines, can be added to the peptide sequences of the invention (e.g., at the N- terminal or C-terminal ends). Sequence tags can be used for peptide purification or localization.
- Lysines can be used to increase peptide solubility or to allow for biotinylation.
- amino acid residues located at the carboxy and amino terminal regions of the amino acid sequence of a peptide or protein may optionally be deleted providing for truncated sequences.
- Certain amino acids e.g., C-terminal or N-terminal residues
- “Substitutional variants” when referring to polypeptides are those that have at least one amino acid residue in a native or starting sequence removed and a different amino acid inserted in its place at the same position.
- substitutions may be single, where only one amino acid in the molecule has been substituted, or they may be multiple, where two or more (e.g., 3, 4 or 5) amino acids have been substituted in the same molecule.
- the substitutions may be conservative amino acid substitutions.
- conservative amino acid substitution refers to the substitution of an amino acid that is normally present in the sequence with a different amino acid of similar size, charge, or polarity. Examples of conservative substitutions include the substitution of a non-polar (hydrophobic) residue such as isoleucine, valine and leucine for another non-polar residue.
- examples of conservative substitutions include the substitution of one polar (hydrophilic) residue for another such as between arginine and lysine, between glutamine and asparagine, and between glycine and serine. Additionally, the substitution of a basic residue such as lysine, arginine or histidine for another, or the substitution of one acidic residue such as aspartic acid or glutamic acid for another acidic residue are additional examples of conservative substitutions. Examples of non-conservative substitutions include the substitution of a non-polar (hydrophobic) amino acid residue such as isoleucine, valine, leucine, alanine, 1005272698
- a polar (hydrophilic) residue such as cysteine, glutamine, glutamic acid or lysine and/or a polar residue for a non-polar residue.
- Amino acid deletions or insertions can also be made relative to the native sequence of the P. gingivalis protein. Thus, for example, amino acids which do not have a substantial effect on the activity of the polypeptide, or at least which do not eliminate such activity, can be deleted.
- domain refers to a motif of a polypeptide having one or more identifiable structural or functional characteristics or properties (e.g., binding capacity, serving as a site for protein-protein interactions).
- RNA molecule of the invention may make modifications to an RNA molecule of the invention so that it includes codons encoding additional amino acid residues derived from the naturally occurring domain sequences of the gingipain.
- an RNA of the invention encodes a chimeric protein (such as RA, KA, KDA, KDAK and the like), and wherein the sequences of K, R, D and A are as herein defined, it will be within the purview of the skilled person to include additional codons encoding additional amino acids at the N or C termini of each domain, for example, in order to further stabilise the encoded protein.
- the additional amino acids correspond to naturally occurring gingipain sequence.
- a methionine may be encoded at the N terminal region of the A domain.
- the RNA may encode 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10 or more additional amino acid residues.
- the D domain may comprise a C terminal methionine residue (or alternatively this residue may be omitted from the sequence of the D domain).
- Such extremity is not limited only to the first or final site of the polypeptide or polynucleotide but may include additional amino acids or nucleotides in the terminal regions.
- Polypeptide-based molecules may be characterized as having both an N- terminus (terminated by an amino acid with a free amino group (NH2)) and a C-terminus (terminated by an amino acid with a free carboxyl group (COOH)). Proteins are in some cases made up of multiple polypeptide chains brought together by disulfide bonds or by 1005272698
- protein fragments, functional protein domains, and homologous proteins are also considered to be within the scope of polypeptides of interest.
- any protein fragment meaning a polypeptide sequence at least one amino acid residue shorter than a reference polypeptide sequence but otherwise identical
- a reference protein having a length of 10, 20, 30, 40, 50, 60, 70, 80, 90, 100 or longer than 100 amino acids.
- any protein that includes a stretch of 20, 30, 40, 50, or 100 (contiguous) amino acids that are 40%, 50%, 60%, 70%, 80%, 90%, 95%, or 100% identical to any of the sequences described herein can be utilized in accordance with the disclosure.
- a polypeptide includes 2, 3, 4, 5, 6, 7, 8, 9, 10, or more mutations as shown in any of the sequences provided herein or referenced herein.
- any protein that includes a stretch of 20, 30, 40, 50, or 100 amino acids that are greater than 80%, 90%, 95%, or 100% identical to any of the sequences described herein, wherein the protein has a stretch of 5, 10, 15, 20, 25, or 30 amino acids that are less than 80%, 75%, 70%, 65% to 60% identical to any of the sequences described herein can be utilized in accordance with the disclosure.
- Polypeptide or polynucleotide molecules of the present disclosure may share a certain degree of sequence similarity or identity with recited SEQ ID NOs.
- identity refers to a relationship between the sequences of two or more polypeptides or polynucleotides, as determined by comparing the sequences.
- identity also means the degree of sequence relatedness between two sequences as determined by the number of matches between strings of two or more amino acid residues or nucleic acid residues. Identity measures the percent of identical matches between the smaller of two or more sequences with gap alignments (if any) addressed by a particular mathematical model or computer program (e.g., “algorithms”). Identity of related peptides can be readily calculated by known methods. “% identity” as it applies to polypeptide or polynucleotide sequences is defined as the percentage of residues (amino acid residues or nucleic acid residues) in the candidate amino acid or nucleic acid sequence that are identical with the residues in the amino acid sequence or nucleic acid 1005272698
- variants of a particular polynucleotide or polypeptide have at least 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% but less than 100% sequence identity to that particular reference polynucleotide or polypeptide as determined by sequence alignment programs and parameters described herein and known to those skilled in the art.
- Such tools for alignment include those of the BLAST suite (Stephen F. Altschul, et al. (1997).” Gapped BLAST and PSI-BLAST: a new generation of protein database search programs,” Nucleic Acids Res. 25:3389-3402).
- Another popular local alignment technique is based on the Smith-Waterman algorithm (Smith, T. F. & Waterman, M. S. (1981) “Identification of common molecular subsequences.” J. Mol. Biol. 147:195-197).
- a general global alignment technique based on dynamic programming is the Needleman-Wunsch algorithm (Needleman, S. B. & Wunsch, C. D. (1970) “A general method applicable to the search for similarities in the amino acid sequences of two proteins.” J.
- identity refers to the overall relatedness between polymeric molecules, for example, between polynucleotide molecules (e.g. DNA molecules and/or RNA molecules) and/or between polypeptide molecules.
- Calculation of the percent identity of two polynucleic acid sequences can be performed by aligning the two sequences for optimal comparison purposes (e.g., gaps can be introduced in one or both of a first and a second nucleic acid sequences for optimal alignment and non- identical sequences can be disregarded for comparison purposes).
- the length of a sequence aligned for comparison purposes is at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, or 100% of the length of the reference sequence.
- the nucleotides at corresponding nucleotide positions are then compared. When a position in the first sequence is occupied by the same nucleotide as the corresponding position in the second sequence, then the 1005272698
- the percent identity between the two sequences is a function of the number of identical positions shared by the sequences, taking into account the number of gaps, and the length of each gap, which needs to be introduced for optimal alignment of the two sequences.
- the comparison of sequences and determination of percent identity between two sequences can be accomplished using a mathematical algorithm.
- the percent identity between two nucleic acid sequences can be determined using methods such as those described in Computational Molecular Biology, Lesk, A. M., ed., Oxford University Press, New York, 1988; Biocomputing: Informatics and Genome Projects, Smith, D.
- the percent identity between two nucleic acid sequences can be determined using the algorithm of Meyers and Miller (CABIOS, 1989, 4:11-17), which has been incorporated into the ALIGN program (version 2.0) using a PAM 120 weight residue table, a gap length penalty of 12 and a gap penalty of 4.
- the percent identity between two nucleic acid sequences can, alternatively, be determined using the GAP program in the GCG software package using an NWSgapdna.CMP matrix. Methods commonly employed to determine percent identity between sequences include, but are not limited to those disclosed in Carillo, H., and Lipman, D., SIAM J Applied Math., 48:1073 (1988); incorporated herein by reference. Techniques for determining identity are codified in publicly available computer programs.
- Exemplary computer software to determine homology between two sequences include, but are not limited to, GCG program package, Devereux, J., et al., Nucleic Acids Research, 12(1), 387 (1984)), BLASTP, BLASTN, and FASTA Altschul, S. F. et al., J. Molec. Biol., 215, 403 (1990)).
- Signal peptides [0204]
- the polypeptides encoded by the polynucleotides of the invention typically comprise N-terminal signal peptides.
- Signal peptides comprising the N-terminal 15-60 amino acids of proteins, are typically needed for the translocation across the membrane on the secretory pathway and, thus, universally control the entry of most proteins both in eukaryotes and prokaryotes to the secretory pathway.
- Signal peptides generally include three regions: an N-terminal region of differing length, which usually comprises positively 1005272698
- the signal peptide of a nascent precursor protein directs the ribosome to the rough endoplasmic reticulum (ER) membrane and initiates the transport of the growing peptide chain across it for processing.
- ER processing produces mature proteins, wherein the signal peptide is cleaved from precursor proteins, typically by a ER- resident signal peptidase of the host cell, or they remain uncleaved and function as a membrane anchor.
- a signal peptide may also facilitate the targeting of the protein to the cell membrane. The signal peptide, however, is not responsible for the final destination of the mature protein.
- the N-terminal secretion signal peptide may comprise any amino acid sequence which enables the chimeric or fusion protein to be processed by ribosomes bound to the rough endoplasmic reticulum (ER) of a cell, and thereby results in threading of the chimeric or fusion protein into the ER.
- ER endoplasmic reticulum
- the N-terminal secretion signal peptide is any peptide that enables secretion of the encoded protein by the cell in which the RNA is expressed or translated.
- the signal peptide fused to the antigenic polypeptide is an artificial signal peptide.
- an artificial signal peptide fused to the antigenic polypeptide encoded by the RNA (e.g., mRNA) vaccine is obtained from an immunoglobulin protein, e.g., an IgE signal peptide or an IgG signal peptide.
- a signal peptide fused to the antigenic polypeptide encoded by a RNA (e.g., mRNA) vaccine is an Ig heavy chain epsilon-1 signal peptide (IgE HC SP) having the sequence of: MDWTWILFLVAAATRVHS (SEQ ID NO: 96).
- a signal peptide fused to the antigenic polypeptide encoded by the RNA (e.g., mRNA) vaccine is an IgGk chain V-III region HAH signal peptide (IgGk SP) having the sequence of METPAQLLFLLLLWLPDTTG (SEQ ID NO: 97).
- the signal peptide is selected from: Japanese encephalitis PRM signal sequence (MLGSNSGQRVVFTILLLLVAPAYS, SEQ ID NO: 98), VSVg 1005272698
- tPA tissue plasminogen activator
- MDAMKRGLCCVLLLCGAVFVSPS tissue plasminogen activator
- tPA tissue plasminogen activator
- tPA tissue plasminogen activator
- tPA tissue plasminogen activator
- VSAR MDAMKRGLCCVLLLCGAVFVSAR
- tPA VSP
- VSPS MDAMKRGLCCVLLLCGAVFVSP
- the amino acid sequence of the signal peptide comprises the sequence of SEAP (secreted embryonic alkaline phosphatase): MLLLLLLLGLRLQLSLG[A] (SEQ ID NO: 105), such that the expressed RNA product comprises the sequence MLLLLLLLGLRLQLSLG[A] (SEQ ID NO: 105) N terminal to the sequences defined herein including in Table 1.
- the signal peptide comprises the sequence as set forth in any of these examples, of a sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto.
- a signal peptide may have a length of 15-60 amino acids.
- a signal peptide may have a length of 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, or 60 amino acids.
- a signal peptide has a length of 20-60, 25-60, 30-60, 35-60, 40-60, 45-60, 50-60, 55-60, 15-55, 20-55, 25-55, 30-55, 35-55, 40-55, 45-55, 50-55, 15-50, 20-50, 25-50, 30-50, 35-50, 40-50, 45-50, 15- 45, 20-45, 25-45, 30-45, 35-45, 40-45, 15-40, 20-40, 25-40, 30-40, 35-40, 15-35, 20-35, 25-35, 30-35, 15-30, 20-30, 25-30, 15-25, 20-25, or 15-20 amino acids. 1005272698
- a signal peptide is typically cleaved from the nascent polypeptide at the cleavage junction during ER processing.
- the mature antigenic polypeptide produces by an RNA vaccine of the present disclosure typically does not comprise a signal peptide.
- the RNA of the invention may include sequence encoding an N terminal methionine, and/or other residues (such as alanine) for enabling expression, secretion and/or cleavage of the signal peptide.
- the RNA of the invention may encode one, two, three, four or more N terminal amino acids to the sequences defined herein in Table 1.
- an RNA encoding an amino acid sequence as set forth in any of SEQ ID NOs: 2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 38 or 39 may encode one or more additional N terminal amino acids, optionally an N terminal alanine residue and/or optionally an N terminal methionine residue.
- an RNA encoding an amino acid sequence of a D domain (eg in SEQ ID NOs: 35 and 36) may comprise the C terminal methionine residue (as shown in these sequence) or the C terminal methionine residue may be omitted from the D domain sequence.
- compositions and lipid nanoparticles contemplates the provision of a polynucleotide (preferably an RNA) encoding an a chimeric or fusion protein for inducing an immune response to P. gingivalis, preferably formulated in a lipid nanoparticle. Accordingly, the present invention also provides a lipid nanoparticle comprising a polynucleotide as described herein. It will be appreciated that in any embodiment, the nanoparticles of the invention may also be described as “vaccine” compositions or “immune stimulating” compositions. [0219] In some embodiments, the RNA of the invention is formulated in a lipid- polycation complex, referred to as a cationic lipid nanoparticle.
- the polycation may include a cationic peptide or a polypeptide such as, but not limited to, polylysine, polyornithine and/or polyarginine.
- the RNA may be 1005272698
- Lipid nanoparticle formulations typically comprise a lipid, in particular, an ionizable cationic lipid, and further comprise a non-cationic lipid, a sterol and a molecule capable of reducing particle aggregation, for example a PEG or PEG-modified lipid.
- a cationic lipid is an ionizable cationic lipid and the non- cationic lipid is a neutral lipid, and the sterol is a cholesterol.
- a cationic lipid is selected from the group consisting of 2,2-dilinoleyl-4-dimethylaminoethyl- [1,3]-dioxolane (DLin-KC2-DMA), dilinoleyl-methyl-4-dimethylaminobutyrate (DLin-MC3- DMA), di((Z)-non-2-en-1-yl) 9-((4-(dimethylamino)butanoyl)oxy)heptadecanedioate (L319), (12Z,15Z)—N,N-dimethyl-2-nonylhenicosa-12,15-dien-1-amine (L608), and N,N- dimethyl-1-[(1S,2R)-2-octylcyclopropyl]heptadecan-8-amine (L530).
- DLin-KC2-DMA 2,2-dilinoleyl-4-dimethylaminoethyl- [1,3]
- lipid nanoparticle formulations include 25-75% of a cationic lipid optionally selected from 2,2-dilinoleyl-4-dimethylaminoethyl-[1,3]-dioxolane (DLin-KC2-DMA), dilinoleyl-methyl-4-dimethylaminobutyrate (DLin-MC3-DMA), and di((Z)-non-2-en-1-yl) 9-((4-(dimethylamino)butanoyl)oxy)heptadecanedioate (L319), 0.5- 15% of the neutral lipid, 5-50% of the sterol, and 0.5-20% of the PEG or PEG-modified lipid on a molar basis.
- a cationic lipid optionally selected from 2,2-dilinoleyl-4-dimethylaminoethyl-[1,3]-dioxolane (DLin-KC2-DMA), dilinoleyl-methyl
- lipid nanoparticle formulations consists essentially of a lipid mixture in molar ratios of 20-70% cationic lipid: 5-45% neutral lipid: 20-55% cholesterol: 0.5-15% PEG-modified lipid. In some embodiments, lipid nanoparticle formulations consists essentially of a lipid mixture in a molar ratio of 20-60% cationic lipid: 5-25% neutral lipid: 25-55% cholesterol: 0.5-15% PEG-modified lipid.
- the molar lipid ratio is 50/10/38.5/1.5 (mol % cationic lipid/neutral lipid, e.g., DSPC/Chol/PEG-modified lipid, e.g., PEG-DMG, PEG-DSG or PEG-DPG), 57.2/7.1134.3/1.4 (mol % cationic lipid/neutral lipid, e.g., DPPC/Chol/PEG- modified lipid, e.g., PEG-cDMA), 40/15/40/5 (mol % cationic lipid/neutral lipid, e.g., DSPC/Chol/PEG-modified lipid, e.g., PEG-DMG), 50/10/35/4.5/0.5 (mol % cationic lipid/neutral lipid, e.g., DSPC/Chol/PEG-modified lipid, e.g., PEG-DSG), 50/10/35/5 (cationic lipid/neutral lipid
- a lipid nanoparticle formulation may be influenced by, but not limited to, the selection of the cationic lipid component, the degree of cationic lipid saturation, the nature of the PEGylation, ratio of all components and biophysical parameters such as size.
- the lipid nanoparticle formulation is composed of 57.1% cationic lipid, 7.1% dipalmitoylphosphatidylcholine, 34.3% cholesterol, and 1.4% PEG-c-DMA.
- the lipid nanoparticle comprises a molar ratio of about 20-60% cationic lipid, 0.5-15% PEG-modified lipid, 25-55% sterol, and 25% non-cationic lipid.
- the cationic lipid is an ionizable cationic lipid and the non- cationic lipid is a neutral lipid, and the sterol is a cholesterol.
- the cationic lipid is selected from 2,2-dilinoleyl-4-dimethylaminoethyl-[1,3]-dioxolane (DLin- KC2-DMA), dilinoleyl-methyl-4-dimethylaminobutyrate (DLin-MC3-DMA), and di((Z)-non- 2-en-1-yl) 9-((4-(dimethylamino)butanoyl)oxy)heptadecanedioate (L319).
- DLin- KC2-DMA 2,2-dilinoleyl-4-dimethylaminoethyl-[1,3]-dioxolane
- DLin-MC3-DMA dilinoleyl-methyl-4-dimethylaminobutyrate
- lipid nanoparticle formulations may comprise 35 to 45% cationic lipid, 40% to 50% cationic lipid, 50% to 60% cationic lipid and/or 55% to 65% cationic lipid.
- the ratio of lipid to RNA (e.g., mRNA) in lipid nanoparticles may be 5:1 to 20:1, 10:1 to 25:1, 15:1 to 30:1 and/or at least 30:1.
- the ratio of PEG in the lipid nanoparticle formulations may be increased or decreased and/or the carbon chain length of the PEG lipid may be modified from C14 to C18 to alter the pharmacokinetics and/or biodistribution of the lipid nanoparticle formulations.
- lipid nanoparticle formulations may contain 0.5% to 3.0%, 1.0% to 3.5%, 1.5% to 4.0%, 2.0% to 4.5%, 2.5% to 5.0% and/or 3.0% to 6.0% of the lipid molar ratio of PEG-c-DOMG (R-3-[( ⁇ -methoxy- poly(ethyleneglycol)2000)carbamoyl)]-1,2-dimyristyloxypropyl-3-amine) (also referred to herein as PEG-DOMG) as compared to the cationic lipid, DSPC and cholesterol.
- PEG-c-DOMG may be replaced with a PEG lipid such as, but not 1005272698
- the cationic lipid may be selected from any lipid known in the art such as, but not limited to, DLin-MC3-DMA, DLin-DMA, C12-200 and DLin-KC2- DMA.
- the amino alcohol cationic lipid may be a lipids described in and/or made by the methods described in U.S. Patent Publication No.
- the cationic lipid may be 2-amino- 3-[(9Z,12Z)-octadeca-9,12-dien-1-yloxy]-2- ⁇ [(9Z,2Z)-octadeca-9,12-dien-1-yloxy] methyl ⁇ propan-1-ol (Compound 1 in US20130150625); 2-amino-3-[(9Z)-octadec-9-en-1- yloxy]-2- ⁇ [(9Z)-octadec-9-en-1-yloxy]methyl ⁇ propan-1-ol (Compound 2 in US20130150625); 2-amino-3-[(9Z,12Z)-octadeca-9,12-dien-1-yloxy]-2- [(octyloxy)methyl]propan-1-ol (Compound 3 in US20130150625); and 2-(
- the cationic lipid may be any one of N,N-dioleyl-N,N-dimethylammonium chloride (DODAC), 1,2-dioeoyloxy-3-(dimethylamino)propane (DODAP), 1,2-dioleyloxy- N,N-dimethylaminopropane (DODMA), 1,2-distearyloxy-N,N-dimethylaminopropane (DSDMA), N-(1-(2,3-dioleyloxy)propyl)-N,N,N-trimethylammonium chloride (DOTMA), N,N-distearyl-N,N-dimethylammonium bromide (DDAB), N-(1-(2,3-dioleoyloxy)propyl)- N,N,N-trimethylammonium chloride (DOTAP), 3-(N—(N′,N′-dimethylaminoethane)- carbamoyl
- the cationic lipid may be of Formula I [0233] wherein R 1 and R 2 are independently selected and are H or C1-C3 alkyls, R 3 and R 4 are independently selected and are alkyl groups having from about 10 to about 20 carbon atoms, and at least one of R 3 and R 4 comprises at least two sites of unsaturation, preferably the cationic lipid of Formula I is 1,2-dilinoleyloxy-N,N-dimethylaminopropane (DLinDMA) or 1,2-dilinolenyloxy-N,N-dimethylaminopropane (DLenDMA).
- DLinDMA 1,2-dilinoleyloxy-N,N-dimethylaminopropane
- DLenDMA 1,2-dilinolenyloxy-N,N-dimethylaminopropane
- the cationic lipid may be of Formula II [0235] wherein R 1 and R 2 are independently selected and are H or C1-C3 alkyls, R 3 and R 4 are independently selected and are alkyl groups having from about 10 to about 20 carbon atoms, and at least one of R 3 and R 4 comprises at least two sites of unsaturation.
- the cationic lipid may be of Formula III [0237] wherein R 1 and R 2 are either the same or different and independently optionally substituted C12-C24 alkyl, optionally substituted C12-C24 alkenyl, optionally substituted C12- C24 alkynyl, or optionally substituted C12-C24 acyl; R 3 and R 4 are either the same or 1005272698
- R 3 and R 4 may join to form an optionally substituted heterocyclic ring of 4 to 6 carbon atoms and 1 or 2 heteroatoms chosen from nitrogen and oxygen;
- R 5 is either absent or hydrogen or C1-C6 alkyl to provide a quaternary amine;
- m, n, and p are either the same or different and independently either 0 or 1 with the proviso that m, n, and p are not simultaneously 0;
- q is 0, 1, 2, 3, or 4;
- Y and Z are either the same or different and independently O, S, or NH.
- the cationic lipid of Formula III may be 2,2-dilinoleyl-4-(2-dimethylaminoethyl)- [1,3]-dioxolane (DLin-K-C2-DMA; “XTC2”), 2,2-dilinoleyl-4-(3-dimethylaminopropyl)-[1,3]- dioxolane (DLin-K-C3-DMA), 2,2-dilinoleyl-4-(4-dimethylaminobutyl)-[1,3]-dioxolane (DLin-K-C4-DMA), 2,2-dilinoleyl-5-dimethylaminomethyl-[1,3]-dioxane (DLin-K6-DMA), 2,2-dilinoleyl-4-N-methylpepiazino-[1,3]-dioxolane (DLin-K-MPZ), 2,2-dilinoleyl-4- dimethylaminomethyl
- the phospholipid moiety may be selected from the group consisting of: phosphatidyl choline, phosphatidyl ethanolamine, phosphatidyl glycerol, phosphatidyl serine, phosphatidic acid, 2-lysophosphatidyl choline, and a sphingomyelin.
- the phospholipid may have a fatty acid moiety selected from the non-limiting group consisting of: lauric acid, myristic acid, myristoleic acid, palmitic acid, palmitoleic acid, stearic acid, oleic acid, linoleic acid, alpha-linolenic acid, erucic acid, phytanoic acid, arachidic acid, arachidonic acid, eicosapentaenoic acid, behenic acid, docosapentaenoic acid, and docosahexaenoic acid.
- lauric acid lauric acid
- myristic acid myristoleic acid
- palmitic acid palmitoleic acid
- stearic acid oleic acid
- linoleic acid alpha-linolenic acid
- erucic acid erucic acid
- phytanoic acid arachidic acid, arachidonic acid
- the phosopholipid may be lecithin, phosphatidylethanolamine, lysolecithin, lysophosphatidylethanolamine, phosphatidylserine, phosphatidylinositol, sphingomyelin, egg sphingomyelin (ESM), cephalin, cardiolipin, phosphatidic acid, cerebrosides, dicetylphosphate, distearoylphosphatidylcholine (DSPC), dioleoylphosphatidylcholine (DOPC), dipalmitoylphosphatidylcholine (DPPC), dioleoylphosphatidylglycerol (DOPG), dipalmitoylphosphatidylglycerol (DPPG), dioleoylphosphatidylethanolamine (DOPE), palmitoyloleoyl-phosphatidylcholine (POPC), palmitoyloleoyl-phosphatidy
- the phosopholipid may be distearoylphosphatidylcholine (DSPC).
- the phospholipid may comprise from about 5 mol % to about 20 mol %, from about 5 mol % to about 15 mol %, from about 5 mol % to about 10 mol %, from about 10 mol % to about 20 mol %, from about 15 mol % to about 20 mol % of the total lipid present in the particle. 1005272698
- the phospholipid may comprise from 5 mol % to 20 mol %, from 5 mol % to 15 mol %, from 5 mol % to 10 mol %, from 10 mol % to 20 mol %, or from 15 mol % to 20 mol % of the total lipid present in the particle.
- the structural lipid may be selected from the group consisting of cholesterol, fecosterol, sitosterol, ergosterol, campesterol, stigmasterol, brassicasterol, tomatidine, tomatine, ursolic acid, alpha-tocopherol, and mixtures thereof.
- the structural lipid may be cholesterol.
- the structural lipid includes cholesterol and a corticosteroid (such as prednisolone, dexamethasone, prednisone, and hydrocortisone), or a combination thereof. Further, the structural lipid may be squalene, squalene or combination thereof. [0248] The structural lipid may include lipids containing geranyl acetate, farnesyl acetate or geranyl-geranyl, or ether, ester, or other derivatives.
- a corticosteroid such as prednisolone, dexamethasone, prednisone, and hydrocortisone
- the structural lipid may be squalene, squalene or combination thereof.
- the structural lipid may include lipids containing geranyl acetate, farnesyl acetate or geranyl-geranyl, or ether, ester, or other derivatives.
- the structural lipid may comprise from about 30 mol % to about 50 mol %, from about 30 mol % to about 45 mol %, from about 30 mol % to about 40 mol %, from about 30 mol % to about 35 mol %, from about 35 mol % to about 50 mol %, from about 40 mol % to about 50 mol %, or from about 45 mol % to about 50 mol % of the total lipid present in the particle.
- Examples of lipid nanoparticle compositions and methods of making same are described, for example, in Semple et al. (2010) Nat. Biotechnol.28:172-176; Jayarama et al. (2012), Angew. Chem. Int.
- a lipid nanoparticle formulation includes 0.5% to 15% on a molar basis of the neutral lipid, e.g., 3 to 12%, 5 to 10% or 15%, 10%, or 7.5% on a molar basis.
- neutral lipids include, without limitation, DSPC, POPC, DPPC, DOPE and SM.
- the formulation includes 5% to 50% on a molar basis of the sterol (e.g., 15 to 45%, 20 to 40%, 40%, 38.5%, 35%, or 31% on a molar basis.
- a non-limiting example of a sterol is cholesterol.
- a lipid nanoparticle formulation includes 0.5% to 20% on a molar basis of the PEG or PEG- modified lipid (e.g., 0.5 to 10%, 0.5 to 5%, 1.5%, 0.5%, 1.5%, 3.5%, or 5% on a molar basis.
- a PEG or PEG modified lipid comprises a PEG molecule of 1005272698
- a PEG or PEG modified lipid comprises a PEG molecule of an average molecular weight of less than 2,000, for example around 1,500 Da, around 1,000 Da, or around 500 Da.
- PEG-modified lipids include PEG-distearoyl glycerol (PEG-DMG) (also referred herein as PEG-C14 or C14-PEG), PEG-cDMA (further discussed in Reyes et al. J. Controlled Release, 107, 276-287 (2005) the contents of which are herein incorporated by reference in their entirety).
- the PEGylated lipid may comprise about 0.05 mol %, about 0.1 mol %, about 0.15 mol %, about 0.2 mol %, about 0.25 mol %, about 0.3 mol %, about 0.35 mol %, about 0.4 mol %, about 0.45 mol %, about 0.5% mol, about 0.6% mol, about 0.7% mol, about 0.8% mol, about 1 % mol, about 1.2% mol, about 1,4 % mol, about 1.6% mol, about 1.8 % mol, or about 2 % mol or more of the total lipid present in the particle.
- the PEGylated lipid may be selected from the group consisting of PEG-modified phosphatidylethanolamines, PEG-modified phosphatidic acids, PEG-modified ceramides, PEG-modified dialkylamines, PEG-modified diacylglycerols, PEG-modified dialkylglycerols, and mixtures thereof.
- the PEGylated lipid may be selected from the group consisting of PEG-c- DOMG, PEG-DMG, PEG-DLPE, PEG-DMPE, PEG-DPPC, or a PEG-DSPE lipid.
- the PEGylated lipid may have a PEG component that has a molecular weight between about 100 Da and about 100,000 Dam between about 100 Da and about 100,000 Da, between about 1000 Da and 9,000 Da, between about 1000 Da and 8,000 Da, between about 1000 Da and 7,000 Da, between about 1000 Da and 6,000 Da, between about 1000 Da and 5,000 Da, between about 1000 Da and 4,000 Da, between about 1000 Da and 3,000 Da, or between about 1000 Da and 2,000 Da.
- the PEGylated lipid may be DSPE-PEG, wherein the PEG has a molecular weight of 2000 Da.
- the pharmaceutical compositions of the RNA (e.g., mRNA) vaccines may include at least one of the PEGylated lipids described in International Publication No. WO2012099755, the contents of which are herein incorporated by reference in their entirety. 1005272698
- the PEGylated lipid may be ALC-0159 (2-[(polyethylene glycol)-2000]-N,N- ditetradecylacetamide).
- ALC-0159 is a PEG/lipid conjugate (i.e. PEGylated lipid), specifically, it is the N,N-dimyristylamide of 2-hydroxyacetic acid, O-pegylated to a PEG chain mass of about 2 kilodaltons (corresponding to about 45-46 ethylene oxide units per molecule of N,N-dimyristyl hydroxyacetamide). It is a non-ionic surfactant by its nature.
- Non-limiting examples of lipid nanoparticle compositions and methods of making them are described, for example, in Semple et al. (2010) Nat. Biotechnol.28:172- 176; Jayarama et al. (2012), Angew. Chem. Int. Ed., 51: 8529-8533; and Maier et al. (2013) Molecular Therapy 21, 1570-1578 (the contents of each of which are incorporated herein by reference in their entirety).
- Relative amounts of the active ingredient, the pharmaceutically acceptable excipient, and/or any additional ingredients in a vaccine composition may vary, depending upon the identity, size, and/or condition of the subject being treated and further depending upon the route by which the composition is to be administered.
- the composition may comprise between 0.1% and 99% (w/w) of the active ingredient.
- the composition may comprise between 0.1% and 100%, e.g., between 0.5 and 50%, between 1-30%, between 5-80%, at least 80% (w/w) active ingredient.
- the RNA vaccine composition of the invention may comprise a polynucleotide described herein, formulated in a lipid nanoparticle comprising ALC—0315 ([(4-hydroxybutyl)azanediyl]di(hexane-6,1-diyl) bis(2-hexyldecanoate)) Cholesterol, DSPC and ALC-0159 (2-[(polyethylene glycol)-2000]-N,N- ditetradecylacetamide), the buffer Tris-sucrose and water for injection.
- ALC—0315 [(4-hydroxybutyl)azanediyl]di(hexane-6,1-diyl) bis(2-hexyldecanoate)
- Cholesterol, DSPC and ALC-0159 (2-[(polyethylene glycol)-2000]-N,N- ditetradecylacetamide), the buffer Tris-sucrose and water for injection.
- the composition comprises: 0.6 mg/mL of drug substance (e.g., polynucleotides encoding a chimeric or fusion protein described herein and comprising components of a P. gingivalis gingipain polyprotein complex), 8.58 mg/mL of ALC-0315, 3.99 mg/mL of cholesterol, 1.80 mg/mL of DSPC, 0.95 mg/mL of ALC-0159 (2-[(polyethylene glycol)-2000]-N,N-ditetradecylacetamide), 3.03 mg/mL of Tris (trishydroxymethyl)aminomethoane), 88 mg/mL of sucrose in water, with a typical volume for injection of 50 ⁇ L. 1005272698
- drug substance e.g., polynucleotides encoding a chimeric or fusion protein described herein and comprising components of a P. gingivalis gingipain polyprotein complex
- ALC-0315 3.99 mg/mL of cholesterol
- the RNA vaccine composition of the invention may comprise the four lipids DLin-MC3-DMA, Cholesterol, DSPC and DMG-PEG 2000 at a ratio of 50:39.8:10:0.2 (mol ratio).
- a nanoparticle e.g., a lipid nanoparticle
- a nanoparticle has a mean diameter of 10-500 nm, 20-400 nm, 30-300 nm, 40-200 nm.
- a nanoparticle e.g., a lipid nanoparticle
- the RNA (e.g., RNA) vaccines of the disclosure may be formulated in lipid nanoparticles having a diameter from about 10 to about 100 nm such as, but not limited to, about 10 to about 20 nm, about 10 to about 30 nm, about 10 to about 40 nm, about 10 to about 50 nm, about 10 to about 60 nm, about 10 to about 70 nm, about 10 to about 80 nm, about 10 to about 90 nm, about 20 to about 30 nm, about 20 to about 40 nm, about 20 to about 50 nm, about 20 to about 60 nm, about 20 to about 70 nm, about 20 to about 80 nm, about 20 to about 90 nm, about 20 to about 100 nm, about 30 to about 40 nm, about 30 to about 50 nm, about 30 to about 60 nm, about 30 to about 70 nm, about 30 to about 80 nm, about 30 to about 90 nm, about 20 to about 100 n
- the lipid nanoparticles may have a diameter from about 10 to 500 nm. [0267] In some embodiments, the lipid nanoparticle may have a diameter greater than 100 nm, greater than 150 nm, greater than 200 nm, greater than 250 nm, greater than 300 nm, greater than 350 nm, greater than 400 nm, greater than 450 nm, greater than 500 nm, greater than 550 nm, greater than 600 nm, greater than 650 nm, greater than 700 nm, greater than 750 nm, greater than 800 nm, greater than 850 nm, greater than 900 nm, greater than 950 nm or greater than 1000 nm. 1005272698
- the lipid nanoparticle may be a limit size lipid nanoparticle described in International Patent Publication No. WO2013059922, the contents of which are herein incorporated by reference in their entirety.
- the limit size lipid nanoparticle may comprise a lipid bilayer surrounding an aqueous core or a hydrophobic core; where the lipid bilayer may comprise a phospholipid such as, but not limited to, diacylphosphatidylcholine, a diacylphosphatidylethanolamine, a ceramide, a sphingomyelin, a dihydrosphingomyelin, a cephalin, a cerebroside, a C8-C20 fatty acid diacylphophatidylcholine, and 1-palmitoyl-2-oleoyl phosphatidylcholine (POPC).
- POPC 1-palmitoyl-2-oleoyl phosphatidylcholine
- the limit size lipid nanoparticle may comprise a polyethylene glycol-lipid such as, but not limited to, DLPE-PEG, DMPE-PEG, DPPC-PEG and DSPE-PEG.
- the RNA (e.g., RNA) vaccine may be associated with a cationic or polycationic compounds, including protamine, nucleoline, spermine or spermidine, or other cationic peptides or proteins, such as poly-L-lysine (PLL), polyarginine, basic polypeptides, cell penetrating peptides (CPPs), including HIV-binding peptides, HIV-1 Tat (HIV), Tat-derived peptides, Penetratin, VP derived or analog peptides, Pestivirus Erns, HSV, VP (Herpes simplex), MAP, KALA or protein transduction domains (PTDs), PpT620, prolin-rich peptides,
- PLL poly-L-lys
- PEI polyethyleneimine
- DOTMA [1-(2,3- sioleyloxy)propyl)]-N,N,N-trimethylammonium chloride
- DMRIE di-C14-amidine
- DOTIM DOTIM
- SAINT DC-Chol
- BGTC CTAP
- DOPC DODAP
- DOPE Dioleyl phosphatidylethanol- amine
- DOSPA DODAB
- DOIC DOMEPC
- DOGS Dioctadecylamidoglicylspermin
- DIMRI Dimyristooxypropyl dimethyl hydroxyethyl ammonium bromide
- DOTAP dioleoyloxy-3-(trimethylammonio)propane
- DC-6-14 O,O-ditetradecanoyl-N-.alpha.- trimethylammonioacetyl)diethanolamine chloride
- CLIP 1 rac-[(2,3- dioctadecyl)]-N,N,N-
- modified polyaminoacids such as beta-aminoacid- polymers or reversed polyamides, etc.
- modified polyethylenes such as PVP (poly(N- ethyl-4-vinylpyridinium bromide)), etc.
- modified acrylates such as pDMAEMA 1005272698
- RNA vaccine is not associated with a cationic or polycationic compounds.
- suitable lipid nanoparticle formulations are provided in US 10,702,600, the contents of which are hereby incorporated by reference.
- the lipid nanoparticles described herein may be made in a sterile environment.
- the nanoparticle formulations may comprise a phosphate conjugate.
- the phosphate conjugate may increase in vivo circulation times and/or increase the targeted delivery of the nanoparticle.
- the phosphate conjugates may include a compound of any one of the formulas described in International Application No. WO2013033438, the contents of which are herein incorporated by reference in its entirety.
- the nanoparticle formulation may comprise a polymer conjugate.
- the polymer conjugate may be a water soluble conjugate.
- the polymer conjugate may have a structure as described in U.S. Patent Application No. 20130059360, the contents of which are herein incorporated by reference in its entirety.
- polymer conjugates with the polynucleotides of the present disclosure may be made using the methods and/or segmented polymeric reagents described in U.S. Patent Application No.
- the polymer conjugate may have pendant side groups comprising ring moieties such as, but not limited to, the polymer conjugates described in U.S. Patent 1005272698
- the nanoparticle formulations may comprise a conjugate to enhance the delivery of nanoparticles of the present disclosure in a subject. Further, the conjugate may inhibit phagocytic clearance of the nanoparticles in a subject.
- the conjugate may be a “self” peptide designed from the human membrane protein CD47 (e.g., the “self” particles described by Rodriguez et al. (Science 2013339, 971-975), herein incorporated by reference in its entirety). As shown by Rodriguez et al., the self peptides delayed macrophage-mediated clearance of nanoparticles which enhanced delivery of the nanoparticles.
- the conjugate may be the membrane protein CD47 (e.g., see Rodriguez et al. Science 2013339, 971-975, herein incorporated by reference in its entirety).
- Rodriguez et al. showed that, similarly to “self” peptides, CD47 can increase the circulating particle ratio in a subject as compared to scrambled peptides and PEG coated nanoparticles.
- the RNA (e.g., RNA) vaccines of the present disclosure are formulated in nanoparticles which comprise a conjugate to enhance the delivery of the nanoparticles of the present disclosure in a subject.
- the conjugate may be the CD47 membrane or the conjugate may be derived from the CD47 membrane protein, such as the “self” peptide described previously.
- the nanoparticle may comprise PEG and a conjugate of CD47 or a derivative thereof.
- the nanoparticle may comprise both the “self” peptide described above and the membrane protein CD47.
- RNA e.g., RNA
- vaccine pharmaceutical compositions comprising the polynucleotides of the present disclosure and a conjugate that may have a degradable linkage.
- conjugates include an aromatic moiety comprising an ionizable hydrogen atom, a spacer moiety, and a water-soluble polymer.
- compositions comprising a conjugate with a degradable linkage and methods for delivering such pharmaceutical compositions are described in U.S. Patent Publication No. US20130184443, the contents of which are herein incorporated by reference in their entirety.
- the nanoparticle formulations may be a carbohydrate nanoparticle comprising a carbohydrate carrier and a RNA (e.g., RNA) vaccine.
- RNA e.g., RNA
- carbohydrate carrier may include, but is not limited to, an anhydride-modified phytoglycogen or glycogen-type material, phtoglycogen octenyl succinate, phytoglycogen beta-dextrin, anhydride-modified phytoglycogen beta-dextrin.
- anhydride-modified phytoglycogen or glycogen-type material phtoglycogen octenyl succinate
- phytoglycogen beta-dextrin anhydride-modified phytoglycogen beta-dextrin.
- Nanoparticle formulations of the present disclosure may be coated with a surfactant or polymer in order to improve the delivery of the particle.
- the nanoparticle may be coated with a hydrophilic coating such as, but not limited to, PEG coatings and/or coatings that have a neutral surface charge.
- the hydrophilic coatings may help to deliver nanoparticles with larger payloads such as, but not limited to, RNA (e.g., RNA) vaccines within the central nervous system.
- RNA e.g., RNA
- nanoparticles comprising a hydrophilic coating and methods of making such nanoparticles are described in U.S. Patent Publication No. US20130183244, the contents of which are herein incorporated by reference in their entirety.
- the lipid nanoparticles of the present disclosure may be hydrophilic polymer particles.
- the lipid nanoparticles of the present disclosure may be hydrophobic polymer particles.
- Lipid nanoparticle formulations may be improved by replacing the cationic lipid with a biodegradable cationic lipid which is known as a rapidly eliminated lipid nanoparticle (reLNP).
- Ionizable cationic lipids such as, but not limited to, DLinDMA, DLin- KC2-DMA, and DLin-MC3-DMA, have been shown to accumulate in plasma and tissues over time and may be a potential source of toxicity.
- the rapid metabolism of the rapidly eliminated lipids can improve the tolerability and therapeutic index of the lipid nanoparticles by an order of magnitude from a 1 mg/kg dose to a 10 mg/kg dose in rat.
- Inclusion of an enzymatically degraded ester linkage can improve the degradation and metabolism profile of the cationic component, while still maintaining the activity of the reLNP formulation.
- the ester linkage can be internally located within the lipid chain or it 1005272698
- an immune response may be elicited by delivering a lipid nanoparticle which may include a nanospecies, a polymer and an immunogen. (U.S. Publication No.20120189700 and International Publication No. WO2012099805; each of which is herein incorporated by reference in their entirety).
- the polymer may encapsulate the nanospecies or partially encapsulate the nanospecies.
- the immunogen may be a recombinant protein, a modified RNA and/or a polynucleotide described herein.
- the lipid nanoparticle may be formulated for use in a vaccine such as, but not limited to, against a pathogen.
- Lipid nanoparticles may be engineered to alter the surface properties of particles so the lipid nanoparticles may penetrate the mucosal barrier.
- Mucus is located on mucosal tissue such as, but not limited to, oral (e.g., the buccal and esophageal membranes and tonsil tissue), ophthalmic, gastrointestinal (e.g., stomach, small intestine, large intestine, colon, rectum), nasal, respiratory (e.g., nasal, pharyngeal, tracheal and bronchial membranes), genital (e.g., vaginal, cervical and urethral membranes). Nanoparticles larger than 10-200 nm which are preferred for higher drug encapsulation efficiency and the ability to provide the sustained delivery of a wide array of drugs have been thought to be too large to rapidly diffuse through mucosal barriers.
- oral e.g., the buccal and esophageal membranes and tonsil tissue
- ophthalmic e.g., gastrointestinal (e.g., stomach, small intestine, large intestine, colon, rectum)
- nasal, respiratory e.g.,
- Mucus is continuously secreted, shed, discarded or digested and recycled so most of the trapped particles may be removed from the mucosa tissue within seconds or within a few hours.
- Large polymeric nanoparticles 200 nm-500 nm in diameter
- PEG polyethylene glycol
- the transport of nanoparticles may be determined using rates of permeation and/or fluorescent microscopy techniques including, but not limited to, fluorescence recovery after photobleaching (FRAP) and high resolution multiple particle tracking (MPT).
- FRAP fluorescence recovery after photobleaching
- MPT high resolution multiple particle tracking
- compositions which can penetrate a mucosal barrier may be made as described in U.S. Pat. No.8,241,670 1005272698
- the lipid nanoparticle engineered to penetrate mucus may comprise a polymeric material (i.e. a polymeric core) and/or a polymer-vitamin conjugate and/or a tri-block co- polymer.
- the polymeric material may include, but is not limited to, polyamines, polyethers, polyamides, polyesters, polycarbamates, polyureas, polycarbonates, poly(styrenes), polyimides, polysulfones, polyurethanes, polyacetylenes, polyethylenes, polyethyeneimines, polyisocyanates, polyacrylates, polymethacrylates, polyacrylonitriles, and polyarylates.
- the polymeric material may be biodegradable and/or biocompatible. Non-limiting examples of biocompatible polymers are described in International Patent Publication No. WO2013116804, the contents of which are herein incorporated by reference in their entirety.
- the polymeric material may additionally be irradiated.
- the polymeric material may be gamma irradiated (see e.g., International App. No. WO201282165, herein incorporated by reference in its entirety).
- specific polymers include poly(caprolactone) (PCL), ethylene vinyl acetate polymer (EVA), poly(lactic acid) (PLA), poly(L-lactic acid) (PLLA), poly(glycolic acid) (PGA), poly(lactic acid-co-glycolic acid) (PLGA), poly(L-lactic acid-co- glycolic acid) (PLLGA), poly(D,L-lactide) (PDLA), poly(L-lactide) (PLLA), poly(D,L-lactide- co-caprolactone), poly(D,L-lactide-co-caprolactone-co-glycolide), poly(D,L-lactide-co- PEO-co-D,L-lactide), poly(D,L-lactide-co-P
- the lipid nanoparticle may be coated or associated with a co-polymer such as, but not limited to, a block co- polymer (such as a branched polyether-polyamide block copolymer described in International Publication No.
- WO2013012476 herein incorporated by reference in its entirety
- (poly(ethylene glycol))-(poly(propylene oxide))-(poly(ethylene glycol)) triblock copolymer see e.g., U.S. Publication 20120121718 and U.S. Publication 20100003337 and U.S. Pat. No. 8,263,665, the contents of each of which is herein incorporated by reference in their entirety).
- the co-polymer may be a polymer that is generally regarded as safe (GRAS) and the formation of the lipid nanoparticle may be in such a way that no new chemical entities are created.
- GRAS generally regarded as safe
- the lipid nanoparticle may comprise poloxamers coating PLGA nanoparticles without forming new chemical entities which are still able to rapidly penetrate human mucus (Yang et al. Angew. Chem. Int. Ed.201150:2597-2600; the contents of which are herein incorporated by reference in their entirety).
- a non-limiting scalable method to produce nanoparticles which can penetrate human mucus is described by Xu et al. (see, e.g., J Control Release 2013, 170(2):279-86; the contents of which are herein incorporated by reference in their entirety).
- the vitamin of the polymer-vitamin conjugate may be vitamin E.
- the vitamin portion of the conjugate may be substituted with other suitable components such as, but not limited to, vitamin A, vitamin E, other vitamins, cholesterol, a hydrophobic moiety, or a hydrophobic component of other surfactants (e.g., sterol chains, fatty acids, hydrocarbon chains and alkylene oxide chains).
- suitable components such as, but not limited to, vitamin A, vitamin E, other vitamins, cholesterol, a hydrophobic moiety, or a hydrophobic component of other surfactants (e.g., sterol chains, fatty acids, hydrocarbon chains and alkylene oxide chains).
- the lipid nanoparticle engineered to penetrate mucus may include surface altering agents such as, but not limited to, polynucleotides, anionic proteins (e.g., bovine serum albumin), surfactants (e.g., cationic surfactants such as for example dimethyldioctadecyl-ammonium bromide), sugars or sugar derivatives (e.g., cyclodextrin), nucleic acids, polymers (e.g., heparin, polyethylene glycol and poloxamer), mucolytic agents (e.g., N-acetylcysteine, mugwort, bromelain, papain, clerodendrum, acetylcysteine, bromhexine, carbocisteine, eprazinone, mesna, ambroxol, sobrerol, domiodol, letosteine, stepronin, tiopronin, gelsolin, thy
- the mucus penetrating lipid nanoparticles may comprise at least one polynucleotide described herein.
- the polynucleotide may be encapsulated in the lipid nanoparticle and/or disposed on the surface of the particle.
- the polynucleotide may be covalently coupled to the lipid nanoparticle.
- Formulations of mucus penetrating lipid nanoparticles may comprise a plurality of nanoparticles. Further, the formulations may contain particles which may interact with the mucus and alter the structural and/or adhesive properties of the surrounding mucus to decrease mucoadhesion, which may increase the delivery of the mucus penetrating lipid nanoparticles to the mucosal tissue.
- the mucus penetrating lipid nanoparticles may be a hypotonic formulation comprising a mucosal penetration enhancing coating.
- the formulation may be hypotonic for the epithelium to which it is being delivered.
- hypotonic formulations may be found in International Patent Publication No. WO2013110028, the contents of which are herein incorporated by reference in their entirety.
- the RNA (e.g., mRNA) vaccine formulation may comprise or be a hypotonic solution.
- the RNA (e.g., mRNA) vaccine is formulated as a lipoplex, such as, without limitation, the ATUPLEXTM system, the DACC system, the DBTC system and other siRNA-lipoplex technology from Silence Therapeutics (London, United Kingdom), STEMFECTTM from STEMGENT® (Cambridge, Mass.), and polyethylenimine (PEI) or protamine-based targeted and non-targeted delivery of nucleic 1005272698
- a lipoplex such as, without limitation, the ATUPLEXTM system, the DACC system, the DBTC system and other siRNA-lipoplex technology from Silence Therapeutics (London, United Kingdom), STEMFECTTM from STEMGENT® (Cambridge, Mass.), and polyethylenimine (PEI) or protamine-based targeted and non-targeted delivery of nucleic 1005272698
- the RNA (e.g., mRNA) vaccine is formulated as a solid lipid nanoparticle.
- a solid lipid nanoparticle may be spherical with an average diameter between 10 to 1000 nm.
- the SLN possess a solid lipid core matrix that can solubilize lipophilic molecules and may be stabilized with surfactants and/or emulsifiers.
- the lipid nanoparticle may be a self-assembly lipid-polymer nanoparticle (see Zhang et al., ACS Nano, 2008, 2 (8), pp 1696-1702; the contents of which are herein incorporated by reference in their entirety).
- the SLN may be the SLN described in International Patent Publication No. WO2013105101, the contents of which are herein incorporated by reference in their entirety.
- the SLN may be made by the methods or processes described in International Patent Publication No.
- the RNA (e.g., mRNA) vaccines of the present disclosure can be formulated for controlled release and/or targeted delivery.
- controlled release refers to a pharmaceutical composition or compound release profile that conforms to a particular pattern of release to effect a therapeutic outcome.
- the RNA (e.g., mRNA) vaccines may be encapsulated into a delivery agent described herein and/or known in the art for controlled release and/or targeted delivery.
- the term “encapsulate” means to enclose, surround or encase.
- encapsulation may be substantial, complete or partial.
- substantially encapsulated means that at least greater than 50, 60, 70, 80, 85, 90, 95, 96, 97, 98, 99, 99.9, 99.9 or greater than 99.999% of the pharmaceutical composition or compound of the disclosure may be enclosed, surrounded or encased within the delivery agent. “Partially encapsulated” means that less 1005272698
- encapsulation may be determined by measuring the escape or the activity of the pharmaceutical composition or compound of the disclosure using fluorescence and/or electron micrograph. For example, at least 1, 5, 10, 20, 30, 40, 50, 60, 70, 80, 85, 90, 95, 96, 97, 98, 99, 99.9, 99.99 or greater than 99.99% of the pharmaceutical composition or compound of the disclosure are encapsulated in the delivery agent.
- the controlled release formulation may include, but is not limited to, tri-block co-polymers.
- the formulation may include two different types of tri-block co-polymers (International Pub. No. WO2012131104 and WO2012131106, the contents of each of which are incorporated herein by reference in their entirety).
- the RNA (e.g., mRNA) vaccines may be encapsulated into a lipid nanoparticle or a rapidly eliminated lipid nanoparticle and the lipid nanoparticles or a rapidly eliminated lipid nanoparticle may then be encapsulated into a polymer, hydrogel and/or surgical sealant described herein and/or known in the art.
- the polymer, hydrogel or surgical sealant may be PLGA, ethylene vinyl acetate (EVAc), poloxamer, GELSITE® (Nanotherapeutics, Inc. Alachua, Fla.), HYLENEX® (Halozyme Therapeutics, San Diego Calif.), surgical sealants such as fibrinogen polymers (Ethicon Inc. Cornelia, Ga.), TISSELL® (Baxter International, Inc Deerfield, Ill.), PEG-based sealants, and COSEAL® (Baxter International, Inc Deerfield, Ill.).
- EVAc ethylene vinyl acetate
- poloxamer GELSITE®
- HYLENEX® HyLENEX®
- surgical sealants such as fibrinogen polymers (Ethicon Inc. Cornelia, Ga.), TISSELL® (Baxter International, Inc Deerfield, Ill.), PEG-based sealants, and COSEAL® (Baxter International, Inc Deerfield, Ill.).
- the lipid nanoparticle may be encapsulated into any polymer known in the art which may form a gel when injected into a subject.
- the lipid nanoparticle may be encapsulated into a polymer matrix which may be biodegradable.
- the RNA (e.g., mRNA) vaccine formulation for controlled release and/or targeted delivery may also include at least one controlled release coating.
- Controlled release coatings include, but are not limited to, OPADRY®, polyvinylpyrrolidone/vinyl acetate copolymer, polyvinylpyrrolidone, hydroxypropyl methylcellulose, hydroxypropyl cellulose, hydroxyethyl cellulose, EUDRAGIT RL®, 1005272698
- the RNA (e.g., mRNA) vaccine controlled release and/or targeted delivery formulation may comprise at least one degradable polyester which may contain polycationic side chains.
- Degradable polyesters include, but are not limited to, poly(serine ester), poly(L-lactide-co-L-lysine), poly(4-hydroxy-L-proline ester), and combinations thereof.
- the degradable polyesters may include a PEG conjugation to form a PEGylated polymer.
- the RNA (e.g., mRNA) vaccine controlled release and/or targeted delivery formulation comprising at least one polynucleotide may comprise at least one PEG and/or PEG related polymer derivatives as described in U.S. Pat. No. 8,404,222, the contents of which are incorporated herein by reference in their entirety.
- the RNA (e.g., mRNA) vaccine controlled release delivery formulation comprising at least one polynucleotide may be the controlled release polymer system described in US20130130348, the contents of which are incorporated herein by reference in their entirety.
- the RNA (e.g., mRNA) vaccines of the present disclosure may be encapsulated in a therapeutic nanoparticle, referred to herein as “therapeutic nanoparticle RNA (e.g., mRNA) vaccines.”
- Therapeutic nanoparticles may be formulated by methods described herein and known in the art such as, but not limited to, International Pub Nos. WO2010005740, WO2010030763, WO2010005721, WO2010005723, WO2012054923, U.S. Publication Nos.
- therapeutic polymer nanoparticles may be identified by the methods described in US Pub No. US20120140790, the contents of which are herein incorporated by reference in their entirety.
- the therapeutic nanoparticle RNA e.g., mRNA
- sustained release refers to a 1005272698
- the sustained release nanoparticle may comprise a polymer and a therapeutic agent such as, but not limited to, the polynucleotides of the present disclosure (see International Pub No.2010075072 and US Pub No. US20100216804, US20110217377 and US20120201859, the contents of each of which are incorporated herein by reference in their entirety).
- the sustained release formulation may comprise agents which permit persistent bioavailability such as, but not limited to, crystals, macromolecular gels and/or particulate suspensions (see U.S. Patent Publication No US20130150295, the contents of each of which are incorporated herein by reference in their entirety).
- the therapeutic nanoparticle RNA e.g., mRNA
- the therapeutic nanoparticles may include a corticosteroid (see International Pub. No. WO2011084518, the contents of which are incorporated herein by reference in their entirety).
- the therapeutic nanoparticles may be formulated in nanoparticles described in International Pub No. WO2008121949, WO2010005726, WO2010005725, WO2011084521 and US Pub No. US20100069426, US20120004293 and US20100104655, the contents of each of which are incorporated herein by reference in their entirety.
- the nanoparticles of the present disclosure may comprise a polymeric matrix.
- the nanoparticle may comprise two or more polymers such as, but not limited to, polyethylenes, polycarbonates, polyanhydrides, polyhydroxyacids, polypropylfumerates, polycaprolactones, polyamides, polyacetals, polyethers, polyesters, poly(orthoesters), polycyanoacrylates, polyvinyl alcohols, polyurethanes, polyphosphazenes, polyacrylates, polymethacrylates, polycyanoacrylates, polyureas, polystyrenes, polyamines, polylysine, poly(ethylene imine), poly(serine ester), poly(L-lactide-co-L-lysine), poly(4-hydroxy-L-proline ester) or combinations thereof.
- polymers such as, but not limited to, polyethylenes, polycarbonates, polyanhydrides, polyhydroxyacids, polypropylfumerates, polycaprolactones, polyamides, polyacetals, polyethers, polyesters, poly(
- the therapeutic nanoparticle comprises a diblock copolymer.
- the diblock copolymer may include PEG in combination with a polymer such as, but not limited to, polyethylenes, polycarbonates, polyanhydrides, polyhydroxyacids, polypropylfumerates, polycaprolactones, polyamides, 1005272698
- the diblock copolymer may be a high- X diblock copolymer such as those described in International Patent Publication No. WO2013120052, the contents of which are incorporated herein by reference in their entirety.
- the therapeutic nanoparticle comprises a PLGA-PEG block copolymer (see U.S. Publication No. US20120004293 and U.S. Pat. No.8,236,330, each of which is herein incorporated by reference in their entirety).
- the therapeutic nanoparticle is a stealth nanoparticle comprising a diblock copolymer of PEG and PLA or PEG and PLGA (see U.S. Pat. No. 8,246,968 and International Publication No. WO2012166923, the contents of each of which are herein incorporated by reference in their entirety).
- the therapeutic nanoparticle is a stealth nanoparticle or a target-specific stealth nanoparticle as described in U.S. Patent Publication No. US20130172406, the contents of which are herein incorporated by reference in their entirety.
- the therapeutic nanoparticle may comprise a multiblock copolymer (see e.g., U.S. Pat. Nos.8,263,665 and 8,287,910 and U.S. Patent Pub. No. US20130195987, the contents of each of which are herein incorporated by reference in their entirety).
- the lipid nanoparticle comprises the block copolymer PEG-PLGA-PEG (see e.g., the thermosensitive hydrogel (PEG-PLGA-PEG) was used as a TGF-beta1 gene delivery vehicle in Lee et al.
- Thermosensitive Hydrogel as a Tgf- ⁇ 1 Gene Delivery Vehicle Enhances Diabetic Wound Healing.
- RNA e.g., mRNA
- the therapeutic nanoparticle may comprise a multiblock copolymer (see e.g., U.S. Pat. Nos.8,263,665 and 8,287,910 and U.S. Patent Pub. No. US20130195987, the contents of each of which are herein incorporated by reference in their entirety).
- the block copolymers described herein may be included in a polyion complex comprising a non-polymeric micelle and the block copolymer. (see e.g., U.S.
- the therapeutic nanoparticle may comprise at least one acrylic polymer.
- Acrylic polymers include but are not limited to, acrylic acid, methacrylic acid, acrylic acid and methacrylic acid copolymers, methyl methacrylate copolymers, ethoxyethyl methacrylates, cyanoethyl methacrylate, amino alkyl methacrylate copolymer, poly(acrylic acid), poly(methacrylic acid), polycyanoacrylates and combinations thereof.
- the therapeutic nanoparticles may comprise at least one poly(vinyl ester) polymer.
- the poly(vinyl ester) polymer may be a copolymer such as a random copolymer.
- the random copolymer may have a structure such as those described in International Application No. WO2013032829 or U.S. Patent Publication No US20130121954, the contents of each of which are herein incorporated by reference in their entirety.
- the poly(vinyl ester) polymers may be conjugated to the polynucleotides described herein.
- the therapeutic nanoparticle may comprise at least one diblock copolymer.
- the diblock copolymer may be, but it not limited to, a poly(lactic) acid- poly(ethylene)glycol copolymer (see, e.g., International Patent Publication No. WO2013044219, the contents of which are herein incorporated by reference in their entirety).
- the therapeutic nanoparticle may be used to treat cancer (see International publication No. WO2013044219, the contents of which are herein incorporated by reference in their entirety). 1005272698
- the therapeutic nanoparticles may comprise at least one cationic polymer described herein and/or known in the art.
- the therapeutic nanoparticles may comprise at least one amine-containing polymer such as, but not limited to polylysine, polyethylene imine, poly(amidoamine) dendrimers, poly(beta-amino esters) (see, e.g., U.S. Pat. No. 8,287,849, the contents of which are herein incorporated by reference in their entirety) and combinations thereof.
- the nanoparticles described herein may comprise an amine cationic lipid such as those described in International Patent Application No.
- the cationic lipids may have an amino-amine or an amino- amide moiety.
- the therapeutic nanoparticles may comprise at least one degradable polyester which may contain polycationic side chains.
- Degradable polyesters include, but are not limited to, poly(serine ester), poly(L-lactide-co-L-lysine), poly(4-hydroxy-L-proline ester), and combinations thereof.
- the degradable polyesters may include a PEG conjugation to form a PEGylated polymer.
- the synthetic nanocarriers may be formulated for targeted release.
- the synthetic nanocarrier is formulated to release the polynucleotides at a specified pH and/or after a desired time interval.
- the synthetic nanoparticle may be formulated to release the RNA (e.g., mRNA) vaccines after 24 hours and/or at a pH of 4.5 (see International Publication Nos. WO2010138193 and WO2010138194 and US Pub Nos. US20110020388 and US20110027217, each of which is herein incorporated by reference in their entireties).
- the synthetic nanocarriers may be formulated for controlled and/or sustained release of the polynucleotides described herein.
- the synthetic nanocarriers for sustained release may be formulated by methods known in the art, described herein and/or as described in International Pub No. WO2010138192 and US Pub No.20100303850, each of which is herein incorporated by reference in their entirety. 1005272698
- the RNA (e.g., mRNA) vaccine may be formulated for controlled and/or sustained release wherein the formulation comprises at least one polymer that is a crystalline side chain (CYSC) polymer. CYSC polymers are described in U.S. Pat. No.8,399,007, herein incorporated by reference in its entirety.
- the synthetic nanocarrier may be formulated for use as a vaccine.
- the synthetic nanocarrier may encapsulate at least one polynucleotide which encode at least one antigen.
- the synthetic nanocarrier may include at least one antigen and an excipient for a vaccine dosage form (see International Publication No.
- a vaccine dosage form may include at least two synthetic nanocarriers with the same or different antigens and an excipient (see International Publication No. WO2011150249 and U.S. Publication No. US20110293701, the contents of each of which are herein incorporated by reference in their entirety).
- the vaccine dosage form may be selected by methods described herein, known in the art and/or described in International Publication No. WO2011150258 and U.S. Publication No. US20120027806, the contents of each of which are herein incorporated by reference in their entirety).
- the synthetic nanocarrier may comprise at least one polynucleotide which encodes at least one adjuvant.
- the adjuvant may comprise dimethyldioctadecylammonium-bromide, dimethyldioctadecylammonium-chloride, dimethyldioctadecylammonium-phosphate or dimethyldioctadecylammonium-acetate (DDA) and an apolar fraction or part of said apolar fraction of a total lipid extract of a mycobacterium (see, e.g., U.S. Pat. No.
- the synthetic nanocarrier may comprise at least one polynucleotide and an adjuvant.
- the synthetic nanocarrier comprising and adjuvant may be formulated by the methods described in International Publication No. WO2011150240 and U.S. Publication No. US20110293700, the contents of each of which are herein incorporated by reference in their entirety.
- the synthetic nanocarrier may encapsulate at least one polynucleotide that encodes a peptide, fragment or region from a virus.
- the synthetic nanocarrier may include, but is not limited to, any of the 1005272698
- the synthetic nanocarrier may be coupled to a polynucleotide which may be able to trigger a humoral and/or cytotoxic T lymphocyte (CTL) response (see, e.g., International Publication No. WO2013019669, the contents of which are herein incorporated by reference in their entirety).
- CTL cytotoxic T lymphocyte
- the RNA (e.g., mRNA) vaccine may be encapsulated in, linked to and/or associated with zwitterionic lipids.
- zwitterionic lipids Non-limiting examples of zwitterionic lipids and methods of using zwitterionic lipids are described in U.S. Patent Publication No. US20130216607, the contents of which are herein incorporated by reference in their entirety.
- the zwitterionic lipids may be used in the liposomes and lipid nanoparticles described herein.
- the RNA (e.g., mRNA) vaccine may be formulated in colloid nanocarriers as described in U.S. Patent Publication No.
- the nanoparticle may be optimized for oral administration.
- the nanoparticle may comprise at least one cationic biopolymer such as, but not limited to, chitosan or a derivative thereof.
- the nanoparticle may be formulated by the methods described in U.S. Publication No. 20120282343, the contents of which are herein incorporated by reference in their entirety.
- LNPs comprise the lipid KL52 (an amino-lipid disclosed in U.S. Application Publication No. 2012/0295832, the contents of which are herein incorporated by reference in their entirety.
- LNPs comprising KL52 may be administered intravenously and/or in one or more doses. In some embodiments, administration of LNPs comprising KL52 results in equal or improved mRNA and/or protein expression as compared to LNPs comprising MC3. 1005272698
- RNA (e.g., mRNA) vaccine may be delivered using smaller LNPs.
- Such particles may comprise a diameter from below 0.1 um up to 100 nm such as, but not limited to, less than 0.1 um, less than 1.0 um, less than 5 um, less than 10 um, less than 15 um, less than 20 um, less than 25 um, less than 30 um, less than 35 um, less than 40 um, less than 50 um, less than 55 um, less than 60 um, less than 65 um, less than 70 um, less than 75 um, less than 80 um, less than 85 um, less than 90 um, less than 95 um, less than 100 um, less than 125 um, less than 150 um, less than 175 um, less than 200 um, less than 225 um, less than 250 um, less than 275 um, less than 300 um, less than 325 um, less than 350 um, less than 375 um, less than 400 um, less than 425 um, less than 450 um, less than 475 um, less than 500 um,
- RNA (e.g., mRNA) vaccines may be delivered using smaller LNPs, which may comprise a diameter from about 1 nm to about 100 nm, from about 1 nm to about 10 nm, about 1 nm to about 20 nm, from about 1 nm to about 30 nm, from about 1 nm to about 40 nm, from about 1 nm to about 50 nm, from about 1 nm to about 60 nm, from about 1 nm to about 70 nm, from about 1 nm to about 80 nm, from about 1 nm to about 90 nm, from about 5 nm to about from 100 nm, from about 5 nm to about 10 nm, about 5 nm to about 20 nm, from about 5 nm to about 30 nm, from about 5 nm to about 40 nm, from about 5 nm to about 50 nm, from about 5 nm to about 60
- microfluidic mixers may include, but are not limited to, a slit interdigital micromixer including, but not limited to those manufactured by Microinnova (Allerheiligen bei Wildon, Austria) and/or a staggered herringbone micromixer (SHM) (Zhigaltsev, I. V. et al., Bottom-up design and synthesis of limit size lipid nanoparticle systems with aqueous and triglyceride cores using millisecond microfluidic mixing have been published (Langmuir. 2012. 28:3633-40; Belliveau, N. M.
- a slit interdigital micromixer including, but not limited to those manufactured by Microinnova (Allerheiligen bei Wildon, Austria) and/or a staggered herringbone micromixer (SHM) (Zhigaltsev, I. V. et al., Bottom-up design and synthesis of limit size lipid nanoparticle systems with aqueous and trigly
- methods of LNP generation comprising SHM, further comprise the mixing of at least two input streams wherein mixing occurs by microstructure-induced chaotic advection (MICA).
- MICA microstructure-induced chaotic advection
- fluid streams flow through channels present in a herringbone pattern causing rotational flow and folding the fluids around each other.
- This method may also comprise a surface for fluid mixing wherein the surface changes orientations during fluid cycling.
- Methods of generating LNPs using SHM include those disclosed in U.S. Application Publication Nos. 2004/0262223 and 2012/0276209, the contents of each of which are herein incorporated by reference in their entirety.
- the RNA (e.g., mRNA) vaccine of the present disclosure may be formulated in lipid nanoparticles created using a micromixer such as, but not limited to, a Slit Interdigital Microstructured Mixer (SIMM-V2) or a Standard Slit Interdigital Micro Mixer (SSIMM) or Caterpillar (CPMM) or Impinging-jet (IJMM) from the Institut fiir Mikrotechnik Mainz GmbH, Mainz Germany).
- a micromixer such as, but not limited to, a Slit Interdigital Microstructured Mixer (SIMM-V2) or a Standard Slit Interdigital Micro Mixer (SSIMM) or Caterpillar (CPMM) or Impinging-jet (IJMM) from the Institut fiir Mikrotechnik Mainz GmbH, Mainz Germany).
- the RNA (e.g., mRNA) vaccines of the present disclosure may be formulated in lipid nanoparticles created using microfluidic technology (see, e.g., Whitesides
- controlled microfluidic formulation includes a passive method for mixing streams of steady pressure-driven flows in micro channels at a low Reynolds number 1005272698
- RNA vaccines of the present disclosure may be formulated in lipid nanoparticles created using a micromixer chip such as, but not limited to, those from Harvard Apparatus (Holliston, Mass.) or Dolomite Microfluidics (Royston, UK).
- a micromixer chip can be used for rapid mixing of two or more fluid streams with a split and recombine mechanism.
- the RNA (e.g., mRNA) vaccines of the disclosure may be formulated for delivery using the drug encapsulating microspheres described in International Patent Publication No. WO2013063468 or U.S. Pat. No. 8,440,614, the contents of each of which are herein incorporated by reference in their entirety.
- the microspheres may comprise a compound of the formula (I), (II), (III), (IV), (V) or (VI) as described in International Patent Publication No. WO2013063468, the contents of which are herein incorporated by reference in their entirety.
- RNA e.g., mRNA
- a lipid-based formulation including any LNP disclosed herein may further comprise one or more adjuvants.
- an ionisable lipid present in the nanoparticle formulation may be substituted or combined with an adjuvant lipidoid for enhancing RNA delivery.
- the antibody titre produced by the mRNA vaccines of the invention is a neutralizing antibody titre.
- the neutralizing antibody titre is greater than a protein vaccine.
- the neutralizing antibody titre produced by the mRNA vaccines of the invention is greater than an adjuvanted protein vaccine.
- the neutralizing antibody titre produced by the mRNA vaccines of the invention is 1,000-10,000, 1,200-10,000, 1,400-10,000, 1,500-10,000, 1,000-5,000, 1,000-4,000, 1,800-10,000, 2000-10,000, 2,000-5,000, 2,000-3,000, 2,000- 1005272698
- the disclosure features a pharmaceutical composition comprising a nanoparticle composition according to the preceding embodiments and a pharmaceutically acceptable carrier.
- the pharmaceutically acceptable carrier may be as described herein, and may also include one or more agents for facilitating storage of the composition at low temperatures.
- the pharmaceutical composition may be refrigerated or frozen for storage and/or shipment (e.g., being stored at a temperature of 4° C or lower, such as a temperature between about ⁇ 150° C. and about 0° C.
- the pharmaceutical composition is a solution that is refrigerated for storage and/or shipment at, for example, about ⁇ 20° C, ⁇ 30° C, ⁇ 40° C, ⁇ 50° C, ⁇ 60° C, ⁇ 70° C, ⁇ 80° C, ⁇ 90° C, ⁇ 130° C or ⁇ 150° C.).
- the pharmaceutical composition is a solution that is refrigerated for storage and/or shipment at, for example, about ⁇ 20° C, ⁇ 30° C, ⁇ 40° C, ⁇ 50° C, ⁇ 60° C, ⁇ 70° C, or ⁇ 80° C.
- compositions described herein may further comprise one or more cryoprotectants or cryopreservatives.
- cryopreservative or cryoprotectant may comprise a sugar such as sucrose, glucose or related sugar-based cryoprotectant.
- liposomes and Lipoplexes, and Lipid Nanoparticles [0343]
- the RNA (e.g., mRNA) vaccines of the disclosure can be formulated using one or more liposomes, lipoplexes, or lipid nanoparticles.
- pharmaceutical compositions of RNA (e.g., mRNA) vaccines include liposomes.
- Liposomes are artificially-prepared vesicles which may primarily be composed of a lipid bilayer and may be used as a delivery vehicle for the administration of nutrients and pharmaceutical formulations.
- Liposomes can be of different sizes such as, but not limited to, a multilamellar vesicle (MLV) which may be hundreds of nanometers in diameter and may contain a series of concentric bilayers separated by narrow aqueous compartments, a small unicellular vesicle (SUV) which may be smaller than 50 nm in diameter, and a large unilamellar vesicle (LUV) which may be between 50 and 500 nm in diameter.
- MLV multilamellar vesicle
- SUV small unicellular vesicle
- LUV large unilamellar vesicle
- Liposome design may include, but is not limited to, opsonins or ligands in order to improve the attachment of liposomes to unhealthy tissue or to activate events such as, but not limited to, endocytosis.
- Liposomes may contain a low or a high pH in order to improve the delivery of the pharmaceutical formulations. 1005272698
- liposomes may depend on the physicochemical characteristics such as, but not limited to, the pharmaceutical formulation entrapped and the liposomal ingredients, the nature of the medium in which the lipid vesicles are dispersed, the effective concentration of the entrapped substance and its potential toxicity, any additional processes involved during the application and/or delivery of the vesicles, the optimization size, polydispersity and the shelf-life of the vesicles for the intended application, and the batch-to-batch reproducibility and possibility of large-scale production of safe and efficient liposomal products.
- compositions described herein may include, without limitation, liposomes such as those formed from 1,2-dioleyloxy-N,N- dimethylaminopropane (DODMA) liposomes, DiLa2 liposomes from Marina Biotech (Bothell, Wash.), 1,2-dilinoleyloxy-3-dimethylaminopropane (DLin-DMA), 2,2-dilinoleyl-4- (2-dimethylaminoethyl)-[1,3]-dioxolane (DLin-KC2-DMA), and MC3 (US20100324120; herein incorporated by reference in its entirety) and liposomes which may deliver small molecule drugs such as, but not limited to, DOXIL® from Janssen Biotech, Inc.
- DODMA 1,2-dioleyloxy-N,N- dimethylaminopropane
- DiLa2 liposomes from Marina Biotech (Bothell, Wash.
- DLin-DMA 1,2-dilino
- compositions described herein may include, without limitation, liposomes such as those formed from the synthesis of stabilized plasmid-lipid particles (SPLP) or stabilized nucleic acid lipid particle (SNALP) that have been previously described and shown to be suitable for oligonucleotide delivery in vitro and in vivo (see Wheeler et al. Gene Therapy.19996:271-281; Zhang et al. Gene Therapy.19996:1438-1447; Jeffs et al. Pharm Res.200522:362-372; Morrissey et al., Nat Biotechnol. 2005 2:1002-1007; Zimmermann et al., Nature.
- liposomes such as those formed from the synthesis of stabilized plasmid-lipid particles (SPLP) or stabilized nucleic acid lipid particle (SNALP) that have been previously described and shown to be suitable for oligonucleotide delivery in vitro and in vivo (see Wheeler et al. Gene Therapy.19996
- a liposome can contain, but is not limited to, 55% cholesterol, 20% disteroylphosphatidyl choline (DSPC), 10% PEG-S-DSG, and 15% 1,2-dioleyloxy-N,N-dimethylaminopropane (DODMA), as described by Jeffs et al.
- certain liposome formulations may contain, 1005272698
- liposome formulations may comprise from about 25.0% cholesterol to about 40.0% cholesterol, from about 30.0% cholesterol to about 45.0% cholesterol, from about 35.0% cholesterol to about 50.0% cholesterol and/or from about 48.5% cholesterol to about 60% cholesterol.
- formulations may comprise a percentage of cholesterol selected from the group consisting of 28.5%, 31.5%, 33.5%, 36.5%, 37.0%, 38.5%, 39.0% and 43.5%. In some embodiments, formulations may comprise from about 5.0% to about 10.0% DSPC and/or from about 7.0% to about 15.0% DSPC.
- the RNA (e.g., mRNA) vaccine pharmaceutical compositions may be formulated in liposomes such as, but not limited to, DiLa2 liposomes (Marina Biotech, Bothell, Wash.), SMARTICLES® (Marina Biotech, Bothell, Wash.), neutral DOPC (1,2-dioleoyl-sn-glycero-3-phosphocholine) based liposomes (e.g., siRNA delivery for ovarian cancer (Landen et al. Cancer Biology & Therapy 20065(12)1708- 1713); herein incorporated by reference in its entirety) and hyaluronan-coated liposomes (Quiet Therapeutics, Israel).
- liposomes such as, but not limited to, DiLa2 liposomes (Marina Biotech, Bothell, Wash.), SMARTICLES® (Marina Biotech, Bothell, Wash.), neutral DOPC (1,2-dioleoyl-sn-glycero-3-
- the cationic lipid may be a low molecular weight cationic lipid such as those described in U.S. Patent Application No.20130090372, the contents of which are herein incorporated by reference in their entirety.
- the RNA (e.g., mRNA) vaccines may be formulated in a lipid vesicle, which may have crosslinks between functionalized lipid bilayers.
- the RNA (e.g., mRNA) vaccines may be formulated in a lipid-polycation complex. The formation of the lipid-polycation complex may be accomplished by methods known in the art and/or as described in U.S. Pub. No.
- the polycation may include a cationic peptide or a polypeptide such as, but not limited to, polylysine, polyornithine and/or polyarginine.
- the RNA (e.g., mRNA) vaccines may be formulated in a lipid-polycation complex, which may further 1005272698
- 104 include a non-cationic lipid such as, but not limited to, cholesterol or dioleoyl phosphatidylethanolamine (DOPE).
- DOPE dioleoyl phosphatidylethanolamine
- suitable formulations for mRNA delivery are described in Guevara et al., (2020), Frontiers in Chemistry, 8:589959; Zhang et al., (2019), Frontiers in Immunology, 10:594; and Liu et al., (2022), Polymers, 14: 4195; incorporated by reference herein in their entirety.
- Subjects and methods of administration [0353]
- the present invention also provides uses of the polynucleotides and compositions of the invention for producing an antigen-specific immune response in a subject.
- Such methods typically comprise administering a polynucleotide of the invention, preferably formulated in a lipid nanoparticle as described herein, to a subject in need thereof.
- the invention further provides compositions comprising the polynucleotides (RNA) defined herein, and the use of such RNA in immunogenic or vaccine compositions in the treatment or prevention of P. gingivalis infection.
- RNA polynucleotides
- the term "vaccine composition” used herein is defined as a composition used to elicit an immune response against an antigen (immunogen) encoded by the RNA in the composition in order to protect or treat an organism against disease.
- the terms “immunostimulating composition”, “vaccine composition” and “immunogenic composition” may generally be used interchangeably.
- the present invention provides methods and compositions for treating or preventing infection or minimising the likelihood of infection with P. gingivalis, in an individual in need thereof, the methods comprising administering a vaccine composition of the invention.
- the present invention includes methods and compositions for preventing infection with P. gingivalis, minimising the likelihood of infection and/or reducing the severity and duration of P. gingivalis infection in a subject.
- the present invention also provides a method for obtaining an antibody directed to P. gingivalis, the method comprising administering a chimeric or fusion protein, 1005272698
- composition, vaccine or immune stimulating composition of the invention to a non-human animal, thereby generating antibodies directed to P. gingivalis in the animal.
- the method further comprises isolating the antibody from the animal (eg from the blood of the animal) or from an egg of the animal (eg in the case of generating IgY antibodies from chickens).
- the present invention also provides an antibody preparation comprising an antibody directed to P. gingivalis, wherein the antibody preparation is obtained by administering a composition, vaccine or immune stimulating composition of the invention, to a non-human animal, thereby generating antibodies directed to P. gingivalis in the animal, and isolating the antibodies from the animal or egg thereof.
- gingivalis may be used therapeutically to eliminate or reduce P. gingivalis infection or prophylactically, to prevent or reduce the severity of P. gingivalis infection.
- treatment or “treating” of a subject includes the application or administration of a composition of the invention to a subject (or application or administration of a compound of the invention to a cell or tissue from a subject) with the purpose of delaying, slowing, stabilizing, curing, regressing, healing, alleviating, relieving, altering, remedying, less worsening, ameliorating, improving, or affecting the disease or condition, the symptom of the disease or condition, or the risk of (or susceptibility to) the disease or condition.
- treating refers to any indication of success in the treatment or amelioration of an injury, pathology or condition, including any objective or subjective parameter such as abatement; remission; lessening of the rate of worsening; lessening severity of the disease; stabilization, diminishing of symptoms or making the injury, pathology or condition more tolerable to the subject; slowing in the rate of degeneration or decline; making the final point of degeneration less debilitating; or improving a subject's physical or mental well-being.
- preventing or “prevention” is intended to refer to at least the reduction of likelihood of the risk of (or susceptibility to) acquiring a disease or disorder (i.e., causing at least one of the clinical symptoms of the disease not to develop in a subject that may be exposed to or predisposed to the disease but does not yet experience or display symptoms of the disease).
- Biological and physiological parameters for identifying such subjects are provided herein and are also well known by physicians. 1005272698
- the vaccine compositions of the invention can be administered to subjects felt to be in greatest need thereof, for example in the context of human patients, to children or the elderly or individuals at risk of exposure to P. gingivalis.
- the vaccine compositions of the invention can also be administered to subjects suspected of having or diagnosed with having infection with P. gingivalis.
- the compositions and methods of the present invention extend equally to uses in both human and/or veterinary medicine, generation of diagnostic agents or the generation of other treatment reagents.
- the term “subject” shall be taken to mean any animal including humans, for example a mammal. Exemplary subjects include but are not limited to humans and non-human primates. For example, the subject may be a human.
- the subject may be a veterinary subject, such as a companion animal (cat, dog, guinea pig, and the like).
- a companion animal cat, dog, guinea pig, and the like.
- the terms “subject”, “individual” and “patient” may be used interchangeably.
- the skilled person will be familiar with methods for determining successful vaccination/immunisation with a chimeric or fusion protein or composition as described herein. For example, the skilled person will be familiar with methods for quantifying the antibodies generated following immunisation and/or for quantifying the extent of the humoural (Th2) response induced following immunisation or for quantifying the extent of a Th1 response generated.
- Th2 humoural
- the subject exhibits a seroconversion rate of at least 80% (e.g., at least 85%, at least 90%, or at least 95%) following the first dose or the second (booster) dose of the vaccine.
- Seroconversion is the time period during which a specific antibody develops and becomes detectable in the blood.
- antigens enter the blood, and the immune system begins to produce antibodies in response.
- the antigen itself may or may not be detectable, but antibodies are considered absent.
- antibodies are present but not yet detectable. Any time after seroconversion, the antibodies can be detected in the blood, indicating a prior or current infection. 1005272698
- a polynucleotide (e.g., mRNA) vaccine is administered to a subject by intradermal or intramuscular injection, subcutaneous, intravenous, or intranasal route, or any other suitable route for delivery of an RNA-based vaccine.
- RNA e.g.,mRNA
- methods of inducing an antigen specific immune response in a subject including administering to a subject a RNA (e.g.,mRNA) vaccine as described herein, in an effective amount to produce an antigen specific immune response in a subject.
- Antigen-specific immune responses in a subject may be determined, in some embodiments, by assaying for antibody titre following administration to the subject of any of the polynucleotide (e.g., mRNA) vaccines of the present disclosure.
- the anti-antigenic polypeptide antibody titre produced in the subject is increased by at least 1 log relative to a control. In some embodiments, the anti-antigenic polypeptide antibody titre produced in the subject is increased by 1-3 log relative to a control. [0372] In some embodiments, the anti-antigenic polypeptide antibody titre produced in a subject is increased at least 2 times relative to a control. In some embodiments, the anti-antigenic polypeptide titre produced in the subject is increased at least 5 times relative to a control.
- the anti-antigenic polypeptide antibody titre produced in the subject is increased at least 10 times relative to a control. In some embodiments, the anti-antigenic polypeptide antibody titre produced in the subject is increased 2-10 times relative to a control.
- the control is an anti-antigenic polypeptide antibody titre produced in a subject who has not been administered a RNA (e.g., mRNA) vaccine of the present disclosure. In some embodiments, the control is an anti-antigenic polypeptide antibody titre produced in a subject who has been administered a live attenuated or inactivated P. gingivalis vaccine (see, e.g., Ren J. et al. J of Gen.
- a polynucleotide (e.g., mRNA) vaccine of the present disclosure is administered to a subject in an effective amount (an amount effective to induce an immune response).
- the effective amount is a dose equivalent to an at least 2-fold, at least 4-fold, at least 10-fold, at least 100-fold, at least 1000-fold reduction in the standard 1005272698
- the anti-antigenic polypeptide antibody titre produced in the subject is equivalent to an anti-antigenic polypeptide antibody titre produced in a control subject administered the standard of care dose of a recombinant P. gingivalis protein vaccine, a purified P. gingivalis protein vaccine, a live attenuated P. gingivalis vaccine, an inactivated P. gingivalis vaccine.
- the effective amount is a dose equivalent to 2-1000-fold reduction in the standard of care dose of a recombinant P.
- the anti- antigenic polypeptide antibody titre produced in the subject is equivalent to an anti- antigenic polypeptide antibody titre produced in a control subject administered the standard of care dose of a recombinant P. gingivalis protein vaccine, a purified P. gingivalis protein vaccine, a live attenuated P. gingivalis vaccine, or an inactivated P. gingivalis vaccine.
- the polynucleotide (e.g., mRNA) vaccine is formulated in an effective amount to produce an antigen specific immune response in a subject.
- Dosages of the vaccine composition of the present invention can vary between wide limits, depending upon the age and condition of the individual to be treated, etc.
- the effective amount is a total dose of 25 ⁇ g to 1000 ⁇ g, or 50 ⁇ g to 1000 ⁇ g. In some embodiments, the effective amount is a total dose of 100 ⁇ g. In some embodiments, the effective amount is a dose of 25 ⁇ g administered to the subject a total of two times. In some embodiments, the effective amount is a dose of 100 ⁇ g administered to the subject a total of two times.
- the effective amount is a dose of 400 ⁇ g administered to the subject a total of two times. In some embodiments, the effective amount is a dose of 500 ⁇ g administered to the subject a total of two times.
- the efficacy (or effectiveness) of a polynucleotide (e.g., mRNA) vaccine is greater than 60%. In some embodiments, the efficacy (or 1005272698
- a polynucleotide (e.g., mRNA) vaccine is at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, or at least 90%.
- Vaccine efficacy may be assessed using standard analyses (see, e.g., Weinberg et al., J Infect Dis. 2010 Jun. 1; 201(11):1607-10). For example, vaccine efficacy may be measured by double-blind, randomized, clinical controlled trials.
- Vaccine effectiveness is proportional to vaccine efficacy (potency) but is also affected by how well target groups in the population are immunized, as well as by other non-vaccine-related factors that influence the ‘real-world’ outcomes of hospitalizations, ambulatory visits, or costs.
- a retrospective case control analysis may be used, in which the rates of vaccination among a set of infected cases and appropriate controls are compared.
- Vaccine efficacy may also be assessed by evidence of treatment of a P. gingivalis infection as herein defined.
- the evidence of treatment may comprise reduction in the severity or duration of a P. gingivalis infection in a subject, such as reduction in inflammation caused by infection. 1005272698
- the invention is a composition for or method of vaccinating a subject comprising administering to the subject a nucleic acid vaccine comprising one or more RNA polynucleotides having an open reading frame encoding a first antigenic polypeptide wherein a dosage of between 10 ⁇ g/kg and 400 ⁇ g /kg of the nucleic acid vaccine is administered to the subject.
- the dosage of the RNA polynucleotide is 1-5 ⁇ g, 5-10 ⁇ g, 10-15 ⁇ g, 15-20 ⁇ g, 10-25 ⁇ g, 20-25 ⁇ g, 20- 50 ⁇ g, 30-50 ⁇ g, 40-50 ⁇ g, 40-60 ⁇ g, 60-80 ⁇ g, 60-100 ⁇ g, 50-100 ⁇ g, 80-120 ⁇ g, 40-120 ⁇ g, 40-150 ⁇ g, 50-150 ⁇ g, 50-200 ⁇ g, 80-200 ⁇ g, 100-200 ⁇ g, 120-250 ⁇ g, 150-250 ⁇ g, 180-280 ⁇ g, 200-300 ⁇ g, 50-300 ⁇ g, 80-300 ⁇ g, 100-300 ⁇ g, 40-300 ⁇ g, 50-350 ⁇ g, 100- 350 ⁇ g, 200-350 ⁇ g, 300-350 ⁇ g, 320-400 ⁇ g, 40-380 ⁇ g, 40-100 ⁇ g, 100-
- the nucleic acid vaccine is administered to the subject by intradermal or intramuscular injection. In some embodiments, the nucleic acid vaccine is administered to the subject on day zero. In some embodiments, a second dose of the nucleic acid vaccine is administered to the subject on day twenty one. [0384] In some embodiments, a dosage of at least about 2 micrograms ( ⁇ g) or at least about 10 ⁇ g or at least about 20 ⁇ g or at least about 30 ⁇ g of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, a dosage of 100 micrograms ( ⁇ g) of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject.
- a dosage of 50 micrograms ( ⁇ g) of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, a dosage of 75 micrograms ( ⁇ g) of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, a dosage of 150 micrograms ( ⁇ g) of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, a dosage of 400 micrograms of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject.
- a dosage of 200 micrograms ( ⁇ g) of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject.
- the RNA polynucleotide accumulates at a 100 fold higher level in the local lymph node in comparison with the distal lymph node.
- 111 nucleic acid vaccine comprising (a) at least one RNA polynucleotide, said polynucleotide comprising at least one chemical modification or optionally no nucleotide modification and two or more codon-optimized open reading frames, said open reading frames encoding a set of reference antigenic polypeptides, and (b) optionally a pharmaceutically acceptable carrier or excipient.
- the vaccine is administered to the individual via a route selected from the group consisting of intramuscular administration, intradermal administration and subcutaneous administration.
- the administering step comprises contacting a muscle tissue of the subject with a device suitable for injection of the composition.
- the administering step comprises contacting a muscle tissue of the subject with a device suitable for injection of the composition in combination with electroporation.
- Embodiments of the invention provide methods of vaccinating a subject comprising administering to the subject a single dosage of between 25 ⁇ g/kg and 400 ⁇ g/kg of a nucleic acid vaccine comprising one or more RNA polynucleotides as described herein, in an effective amount to vaccinate the subject.
- the invention encompasses a method of treating an elderly subject age 60 years or older comprising administering to the subject a nucleic acid vaccine comprising one or more RNA polynucleotides having an open reading frame encoding an antigenic polypeptide as described herein in an effective amount to vaccinate the subject.
- the invention encompasses a method of treating a young subject age 17 years or younger comprising administering to the subject a nucleic acid vaccine comprising one or more RNA polynucleotides having an open reading frame encoding an antigenic polypeptide as described herein in an effective amount to vaccinate the subject.
- the invention encompasses a method of treating an adult subject comprising administering to the subject a nucleic acid vaccine comprising one or more RNA polynucleotides having an open reading frame encoding an antigenic polypeptide as described herein in an effective amount to vaccinate the subject.
- vaccines of the invention e.g., LNP-encapsulated mRNA vaccines
- antibody titre refers to the amount of antigen-specific antibody produced in a subject, e.g., a human subject.
- antibody titre is expressed as the inverse of the greatest dilution (in a serial dilution) that still gives a positive result.
- antibody titre is determined or measured by enzyme-linked immunosorbent assay (ELISA).
- ELISA enzyme-linked immunosorbent assay
- antibody titre is determined or measured by neutralization assay, e.g., by microneutralization assay.
- antibody titre measurement is expressed as a ratio, such as 1:40, 1:100, etc.
- an efficacious vaccine produces an antibody titre of greater than 1:40, greater that 1:100, greater than 1:400, greater than 1:1000, greater than 1:2000, greater than 1:3000, greater than 1:4000, greater than 1:500, greater than 1:6000, greater than 1:7500, greater than 1:10000.
- the antibody titre is produced or reached by 10 days following vaccination, by 20 days following vaccination, by 30 days following vaccination, by 40 days following vaccination, or by 50 or more days following vaccination.
- the titre is produced or reached following a single dose of vaccine administered to the subject.
- the titre is produced or reached following multiple doses, e.g., following a first and a second dose (e.g., a booster dose.)
- antigen-specific antibodies are measured in units of ⁇ g /ml or are measured in units of IU/L (International Units per liter) or mIU/ml (milli International Units per ml).
- an efficacious vaccine produces >0.5 ⁇ g/ml, >0.1 ⁇ g /ml, >0.2 ⁇ g /ml, >0.35 ⁇ g /ml, >0.5 ⁇ g /ml, >1 ⁇ g /ml, >2 ⁇ g /ml, >5 ⁇ g /ml or >10 ⁇ g /ml.
- an efficacious vaccine produces >10 mIU/ml, >20 mIU/ml, >50 mIU/ml, >100 mIU/ml, >200 mIU/ml, >500 mIU/ml or >1000 mIU/ml.
- the antibody level or concentration is produced or reached by 10 days following vaccination, by 20 days following vaccination, by 30 days following vaccination, by 40 days following vaccination, or by 50 or more days following vaccination.
- the level or concentration is produced or reached following a single dose of vaccine administered to the subject.
- the level or concentration is produced or reached following multiple doses, e.g., following a first and a second dose (e.g., a booster dose.)
- a first and a second dose e.g., a booster dose.
- antibody level or concentration is determined or measured by enzyme- linked immunosorbent assay (ELISA).
- ELISA enzyme- linked immunosorbent assay
- neutralization assay e.g., by microneutralization assay.
- Example 1 Materials and methods
- Production of mRNA-LNPs [0396] The mRNAs were in vitro transcribed using T7 in vitro transcription kit (NEB) according to manufacturer’s instructions, from linearised DNA templates encoding 5’ and 3’ UTRs, signal peptide, the candidate sequence and a 125 nucleotide poly(A) tail and were capped co-transcriptionally using Clean Cap mRNA capping technology (TriLink Biotech).
- N1-methyl pseudouridine N1- methyl pseudo-UTP, m1 ⁇
- DNA was removed using DNAse I (NEB), and double-stranded RNA (dsRNA) was removed using cellulose binding as previously described previously [Baiersdörfer et al. (2019) Mol Ther Nucleic Acids.
- LNPs lipid nanoparticles
- ALC-0315 Cholesterol, DSPC, and ALC-0159 at molar lipid ratios (%) of 46.3:42.7:9.4:1.6, in Tris/sucrose buffer (25 mM Tris pH 7.4, 8.8% sucrose w/v).
- Tris/sucrose buffer 25 mM Tris pH 7.4, 8.8% sucrose w/v.
- HeLa cells were cultured in culture media DMEM, high glucose, GlutaMAXTM Supplement, pyruvate (ThermoFisher, CAT#10569010) with 10% FBS at 37°C, 5% CO2, according to standard protocols.
- mRNA candidates (prior to LNP formulation), were formulated with Lipofectamine MessengerMAX (IThermoFisher) and used to transfect cells according to manufacturer’s instructions.
- mRNA Lipofectamine MessengerMAX
- mRNA candidates were formulated with Lipofectamine MessengerMAX (IThermoFisher) and used to transfect cells according to manufacturer’s instructions.
- 1.8 ⁇ g of an mRNA was used to transfect 35,000 cells/well on a six well plate.
- cells were pelleted by centrifuging at 14,000 g for 15 mins. Supernatants were transferred to a fresh tube and frozen at -20°C until Western Blotting.
- whole cell lysate collection cells were rinsed in each well with ⁇ 2 mL DPBS.
- the liquid was removed and 250 ⁇ L of RIPA lysis buffer with protease inhibitors (ThermoFisher) was added to each well, and gently swirled to mix for 10s. Using a cell scraper the lysates were transferred into tubes and centrifuged at 1005272698
- Proteins were transferred onto a PDV membrane (Life Technologies, cat# LC2002) that had been pre-activated in methanol for 30 s and washed in transfer buffer (10x Tris/Glycine Buffer for Western Blots and Native Gels (BioRad, cat# 1610734), using Trans-Blot® TurboTM Transfer System (BioRad) with the standard protocol: 25V and 1.0A for 30 min.
- PVDF membranes with proteins transferred onto them were blocked for 2 hours in blocking buffer (5% skim milk powder in PBS-T) on a rocker and probed with in-house primary antibodies for binding to KAS2 (1:2500 in blocking buffer) or KDAK-3S-AVQP (1:3500 in blocking buffer) overnight at 4°C on a rocker.
- blocking buffer 5% skim milk powder in PBS-T
- gingivalis strain; W50 (serotype C); was obtained from the culture collection of the Oral Health Cooperative Research Centre, The Melbourne Dental School, University of Melbourne, Australia.
- P. gingivalis W50 was grown on Horse Blood Agar (HBA) (20 g/L HBA; Oxoid Ltd., Hampshire, UK) supplemented with 10% v/v lysed horse blood (37°C) in an anaerobic N2 atmosphere containing 5% CO2 in a MK3 Anaerobic Workstation (Don Whitley Scientific Ltd., Sydney, Australia). Colonies were inoculated 1005272698
- starter culture comprised of 20 mL sterilised brain heart infusion (37 g/L BHI; Oxoid Ltd., Hamsphire, UK) medium supplemented with 5 mg/L hemin and 0.5 mg/L cysteine and incubated anaerobically (24 h, 37 °C). Absorbance of batch cultures were monitored at OD650nm using a spectrophotometer (model 295E, Perkin-Elmer, Germany). Bacterial cells were harvested during late exponential growth by centrifugation (7,000 g, 20 min, 4 °C). Bacterial purity was routinely confirmed by Gram stain [Slots (1982). In: Host- Parasite Interaction in Periodontal Disease, Genco, R.J.
- P. gingivalis W50 culture was harvested (6,500 g, 4 °C), washed once with phosphate buffered saline (PBS) (0.01 M Na2HPO4, 1.5 mM KH2PO4 and 0.15 M NaCl, pH 7.4) then pelleted by centrifugation (7,000 g, 20 min 4°C). Bacterial cells were resuspended in PBS and heated to 65°C for 15 minutes.
- PBS phosphate buffered saline
- mice Female BALB/c; 6-8 weeks old, 10 mice/group
- P. gingivalis consisting of four doses of P. gingivalis W50 [1 x 1010 viable P. gingivalis W50 cells suspended in 20 ⁇ L PG buffer (50 mM Tris-HCL, 150 mM NaCl, 10 mM MgSO4 and 14.3 mM Mercaptoethanol, pH 7.4) containing 2% w/v carboxymethylcellulose (CMC, Sigma, New South Wales, Australia)], given two days apart.
- the inocula were prepared anaerobically and then immediately applied to the gingival margin of the maxillary molar teeth.
- mice were immunised on day 19 after the first oral inoculation with either the protein vaccine control at 200 ⁇ g in 1005272698
- saline/alum Alhydrogel; 10mg/mL w/v aluminium hydroxide wet gel suspension; Invivogen
- mice receiving the mRNA vaccines were injected with either 30 ⁇ g or 3 ⁇ g of formulated mRNA intramuscularly in a total volume of 50 ⁇ L on the same days as the recombinant protein vaccine.
- the injection was performed into the right semitendinosis or semimembranosis thigh muscle using a 27G needle.
- mice received a second immunisation on day 40 via the subcutaneous route (for the alum- adjuvanted protein vaccine) or intramuscularly (for the mRNA vaccine candidates, left semitendinosis or semimembranosis thigh muscle).
- mice On Day 62, mice were bled by cardiac puncture and killed. Maxillae were removed and halved through the midline, with 10 halves used to determine alveolar bone loss. Sera were used to determine the antibody profile using ELISA.
- gingivalis-induced alveolar bone loss in mm2 was calculated by subtracting the total visible CEJ-ABC area of the uninoculated (N-C) group from the total visible CEJ-ABC area of each experimental group. Alveolar bone loss measurements were determined twice in a random and blinded protocol. Data are expressed as the mean +/- standard deviation in mm2 and were analysed using a one-way ANOVA or Kruskal-Wallis test. [0410] Determination of subclass antibody in sera using ELISA 1005272698
- ELISAs were performed to evaluate subclass antibody in sera as described in Pathirana et al. (2007). Infect Immun 75: 1436-1442) using a solution of either heat killed W50 cells (10 ⁇ g/mL), domain subunits or epitopes (1 ⁇ g/mL) in 0.1M PBS (pH 7.4) to coat wells (16 h, 4 oC) of flat-bottom polyvinyl microtiter plates (Microtiter; Dynatech Laboratories, McLean, VA, US).
- the inoculum begun on day 42 and was repeated three times a week for 3 weeks.
- the inocula were prepared anaerobically and then immediately applied to the gingival margin of the maxillary molar teeth.
- the number of viable bacteria in each inoculum was verified by CFU counts on blood agar. [0414]
- ELISPOT assay was performed using the BD® ELISPOT Kits for IL-4 and IFN ⁇ as per the manufacturer’s instructions. 3 x 10 5 cells were added to each well of ELSIPOT plates with or without restimulation with KDAK-3S-AVQP protein (10 ⁇ g/mL). Concanavalin A was used as a positive control mitogen (10 ⁇ g/mL). [0417] Plates were incubated at 37oC in an atmosphere of 5% CO2 in air for 48 hours in a humidified incubator.
- Example 2 Expression of mRNA constructs encoding chimeric proteins [0419] The inventors obtained mRNA constructs encoding chimeric proteins containing postulated antigenic domains from the Kgp gingipain protein.
- the mRNA constructs comprised the general architecture as shown in Figure 2 and the chimeric proteins (antigens) encoded by the mRNA expression constructs comprised the architectures designated KDcAK1n, KDAK and KDAK-3S-AVQP as shown in Figure 1 (amino acid sequences as set forth in SEQ ID NOs: 2, 4 and 6, respectively, mRNA sequences as set forth in SEQ ID NOs: 40, 41 and 42, respectively).
- the mRNAs were tested for in vitro antigen expression and secretion in 293T (not shown) or HeLa cells and using various secretion peptides including tPA-derived 1005272698
- FIG. 1 shows Western Blots demonstrating expression and secretion by HeLa cells of mRNA constructs encoding KDcAK1n, KDAK-3S-AVQP and KDAK. Various signal peptides were tested. The results using the SEAP secretion peptide are shown.
- the LNPs used in these experiments comprised: ALC-0315, cholesterol, distearoylphosphatidylcholine and ALC-0159 in mole percent (%) ratio of: 46.3: 42.7: 9.4: 1.6.
- mRNA constructs were generated using either native RNA sequence (unmodified) or N1-methyl-pseudouridine modified (M1 ⁇ ) sequence.
- M1 ⁇ N1-methyl-pseudouridine modified
- mice were intramuscularly immunised with mRNA-LNPs according to the schedule shown in Figure 4.
- sera were collected at Day 35 and tested using ELISA to determine serum antibody subclasses responses of the immunised mice.
- Antisera were used to probe against the absorbed antigens: KDAK-3S- AVQP (purified recombinant protein), heat killed P. gingivalis strain W50 (HKPg), Kgpcat (catalytic domain of the Kgp gingipain) and a biotinylated linear peptide corresponding to 1005272698
- K AS2 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDK SEQ ID NO: 8
- Antibody responses are expressed as the ELISA titre obtained minus triple the background level, with each titre representing the mean ⁇ s.d. of the 5 individual mice ( Figure 5). Both m1 ⁇ -KDAK and m1 ⁇ -KDAK-3S-AVQP induced strong responses to KDAK-3S-AVQP at both 30 ⁇ g and 3 ⁇ g doses, inducing strong total IgG, IgG1 and IgG2 titres.
- RNAs KDAK and KDAK-3S-AVQP produced strong responses at 30 ⁇ g but weaker at 3 ⁇ g (particularly KDAK-3S-AVQP).
- KDcAK1n and m1 ⁇ -Negative -RNA control produced no significant responses.
- m1 ⁇ -KDAK, m1 ⁇ -KDAK-3S-AVQP and KDAK at 30 ⁇ g provided strong IgG1 responses against HKPg and only m1 ⁇ -KDAK 30 ⁇ g induced significant IgG2 titres.
- m1 ⁇ -KDAK 30 ⁇ g, m1 ⁇ -KDAK-3S-AVQP 30 ⁇ g, KDAK 30 ⁇ g and m1 ⁇ -KDAK 3 ⁇ g induced significant IgG titres against Kgpcat and m1 ⁇ -KDAK 30 ⁇ g, m1 ⁇ -KDAK- 3S-AVQP 30 ⁇ g, KDAK 30 ⁇ g also induced significant IgG1.
- the efficacy of the protein vaccine was not recapitulated when using an mRNA vaccine encoding the sequence for the same protein, potentially due to poor expression and poor secretion of the KDcAK1n protein from mRNA constructs.
- the results also show that vaccines comprising 30 ⁇ g mRNA were generally more immunogenic than vaccines comprising 3 ⁇ g mRNA.
- modified (M1 ⁇ ) mRNA was more immunogenic than native (unmodified) mRNA.
- Example 4 determining in vivo efficacy of mRNA vaccines in periodontitis model
- Figure 6 shows a schematic of the vaccination protocol for determining in vivo efficacy of candidate mRNA vaccines (as assessed by protection from alveolar bone loss), and immunogenicity (as determined by antibody levels in sera). 1005272698
- FIG. 7 shows P. gingivalis induced bone loss (mm) in the different treatment groups.
- KDAK-3S-AVQP protein was used as a positive control and “infected” mice (no RNA or protein vaccination) were used as a negative control.
- mRNA vaccines were prepared and formulated as outlined above and as used in Example 3.
- the results show that vaccination with m1 ⁇ -KDAK 30 ⁇ g, m1 ⁇ -KDAK-3S- AVQP 30 ⁇ g mRNA vaccines provided for significant protection against P. gingivalis induced bone loss, similarly to the alum-adjuvanted polypeptide KDAK-3S-AVQP.
- KDAK 30 ⁇ g and KDAK-3S-AVQP provided partial protection
- Antisera were used to probe against the absorbed antigens: recombinant protein KDAK-3S-AVQP, heat killed P. gingivalis strain W50 (HKPg), Kgpcat and biotinylated linear peptides corresponding to sequences encoded by the tested mRNA vaccine candidates.
- Antibody responses are expressed as the ELISA titre obtained minus triple the background level, with each titre representing the mean ⁇ s.d. of the 10 individual mice ( Figure 8).
- K also referred to herein as KAS or KAS2 active site domain
- amino acid sequence SEQ ID NO: 8 - KD (amino acid sequence: SEQ ID NO: 10) - KA (amino acid sequence SEQ ID NO: 18) - DA (amino acid sequence SEQ ID NO: 14) - KDA (amino acid sequence SEQ ID NO: 12) - KDA ⁇ AMB3 (ie wherein the adhesin domain comprises ABMs 2 and 1 but not ABM3; amino acid sequence SEQ ID NO: 16) - RDA (replacing Arginine-dependent gingipain active site KAS, with active site from a Lysine-dependent gingipain RAS) (amino acid sequence: SEQ ID NO: 20) - positive control: KDAK-3S-AVQP.
- Antisera were used to probe against the absorbed antigens: recombinant protein KDAK-3S-AVQP heat killed P. gingivalis strain W50 (HKPg) and Kgpcat corresponding to sequences encoded by the tested mRNA vaccine candidates.
- Antibody responses are expressed as the ELISA titre obtained minus triple the background level, with each titre representing the mean ⁇ s.d. of the 10 individual mice. All tested constructs induced strong total IgG, IgG1 and IgG2 responses against KDAK- 3S-AVQP ( Figure 11).
- All tested constructs induced strong total IgG and IgG1 titres against HKPg.
- Example 7 further assessment of immunogenicity of truncated constructs
- a further series of immunogenicity assessments were performed using constructs: m1 ⁇ -K 30 ⁇ g and m1 ⁇ -RDA 30 ⁇ g.
- Mice were intramuscularly immunised with the mRNA-LNPs encoding the target antigens according to the schedule shown in in Figure 4. Serum antibody subclass responses of immunised mice were examined by ELISA. Antisera were used to probe 1005272698
- Antibody responses are expressed as the ELISA titre obtained minus triple the background level, with each titre representing the mean ⁇ s.d. of the 5 individual mice ( Figure 12). Only m1 ⁇ -RDA induced strong responses to KDAK-3S-AVQP inducing strong total IgG, IgG1 and IgG2 titres. It also induced significant total IgG primarily IgG1 but not IgG2, against HKPg and significant IgG1 titres against Kgpcat.
- Example 8 In vivo efficacy and immunogenicity in a prophylactic model [0465] mRNAs (formulated in LNPs) encoding KDAK-3S-AVQP were assessed in the prophylactic vaccination model (see Figure 13) and compared to protein vaccine. The mRNA was m1 ⁇ -modified and was administered at a dose of 30 ⁇ g mRNA. [0466] The results, shown in Figure 15, indicate that the mRNA vaccine provided robust protection against P.
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Genetics & Genomics (AREA)
- Organic Chemistry (AREA)
- Engineering & Computer Science (AREA)
- General Health & Medical Sciences (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Medicinal Chemistry (AREA)
- Zoology (AREA)
- Wood Science & Technology (AREA)
- Molecular Biology (AREA)
- Biomedical Technology (AREA)
- Biochemistry (AREA)
- General Engineering & Computer Science (AREA)
- Microbiology (AREA)
- Biotechnology (AREA)
- Animal Behavior & Ethology (AREA)
- Public Health (AREA)
- Pharmacology & Pharmacy (AREA)
- Veterinary Medicine (AREA)
- Immunology (AREA)
- Biophysics (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- General Chemical & Material Sciences (AREA)
- Mycology (AREA)
- Gastroenterology & Hepatology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Plant Pathology (AREA)
- Physics & Mathematics (AREA)
- Epidemiology (AREA)
- Communicable Diseases (AREA)
- Oncology (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
- Peptides Or Proteins (AREA)
- Pharmaceuticals Containing Other Organic And Inorganic Compounds (AREA)
Abstract
The present invention relates to RNA-containing vaccine compositions for inducing an immune response to Porphyromonas gingivalis in a subject, and uses thereof.
Description
1005272698
1 RNA vaccines Field of the invention [0001] The present invention relates to RNA-containing vaccine compositions for inducing an immune response to Porphyromonas gingivalis in a subject, and uses thereof. Related application [0002] This application claims priority from Australian provisional application AU 2023902379, the entire contents of which are hereby incorporated by reference. Background of the invention [0003] If dental plaque is left to accumulate around the tooth at the gingival (gum) margin this causes gingival inflammation (gingivitis). Chronic gingivitis can allow the emergence of a periodontal pathogen Porphyromonas gingivalis (P. gingivalis) at the base of a periodontal pocket to result in a chronic infection and the development of severe disease. This severe form of periodontal disease is called periodontitis and can lead to tooth loss in an approach by the immune system to eliminate the infection. [0004] Chronic periodontitis is an inflammatory disease of the supporting tissues of the teeth leading to resorption of alveolar bone and eventual tooth loss. The disease is a major public health problem in all societies and is estimated to affect up to 30% of the adult population with severe forms affecting 12-15% of the adult population. [0005] One in three adults have moderate to severe periodontitis. From epidemiological surveys, periodontitis has been linked to an increased risk of inflammatory diseases including cardiovascular diseases, certain cancers, preterm birth, rheumatoid arthritis and dementia. More recent research has linked chronic infection by P. gingivalis with dementia and rheumatoid arthritis. For example, in one study, 96% of Alzheimer’s disease (AD) brain samples showed the presence of P. ginigvalis. Another study shows that chronic oral infection of mice with P. gingivalis resulted in the brain plaques associated with AD in humans and that the P. gingivalis proteases could cleave amyloid precursor and tau proteins to form the plaques and tangles associated with AD.
1005272698
2 [0006] A number of virulence factors have been reported to contribute to the pathogenicity of P. gingivalis including; LPS, fimbriae, hemagglutinin, hemolysin and extracellular hydrolytic enzymes (especially the Arg-X and Lys-X specific proteinases), otherwise known as "P. gingivalis gingipains". [0007] The magnitude of the public health problem is such that there is a need for a vaccine that provides a strong protective response to P. gingivalis infection and means for providing same. [0008] One problem has been that it is not clear how to obtain a strong protective response to P. gingivalis infection where there are a plethora of virulence factors to select from. [0009] There is currently no commercially approved vaccine for use in preventing or reducing the incidence and/or severity of P. gingivalis infection or for treating P. gingivalis infection and disease in subjects. [0010] There is therefore a need for alternative and/or improved approaches for the design and manufacture of P. gingivalis vaccines, and alternative and/or improved vaccines produced from P. gingivalis. [0011] Reference to any prior art in the specification is not an acknowledgment or suggestion that this prior art forms part of the common general knowledge in any jurisdiction or that this prior art could reasonably be expected to be understood, regarded as relevant, and/or combined with other pieces of prior art by a skilled person in the art. Summary of the invention [0012] In a first aspect, the invention provides an RNA polynucleotide encoding a protein comprising or consisting of: - one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and/or - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. wherein the polynucleotide is capable of being translated in a mammalian cell.
1005272698
3 [0013] In any embodiment, the protein encoded by the RNA polynucleotide may further comprise: - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. [0014] Optionally, the protein encoded by the RNA may comprise the afore-mentioned domains in any order: for example: the one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P. gingivalis may be located N-terminally to the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, and/or the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis; or the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, may be located N-terminally to the one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P. gingivalis and/or the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, The domains may be directly joined in the context of the chimeric or fusion protein, or may be joined via a linker region, of one or more amino acid residues, as further defined herein. [0015] In other words, the protein may comprises or consist of, N to C terminus: active site (K) - ABMs (A); or active site (R) – ABMs (A); or active site (K) – DUF 2436 (D); or active site (R) – DUF2436 (D); or AMBs (A) – active site (K); or ABMs (A)-active site (R); or DUF2436 (D) – active site (K); or DUF2436 (D) – active site (R); or active site (R) – ABMs (A) - active site (K); or active site (R) – ABMs (A) - active site (R); or active site (K) – ABMs (A) - active site (R); or DUF2436 (D) –– ABMs (A); or DUF2436 (D) –– ABMs (A) – active site (R) or (K). [0016] In any embodiment herein, the RNA polynucleotide is in the form of a messenger RNA (mRNA) molecule. However, it will be appreciated that the RNA polynucleotide may be in any suitable format for being translated in a mammalian cell and enabling synthesis of the protein encoded by the RNA. [0017] In certain embodiments, the RNA polynucleotide may be composed entirely of ribose-containing nucleotides, or alternatively, may comprise a combination of ribose- containing nucleosides and of 2’-deoxyribose-containing nucleotides. [0018] In any embodiment, the RNA polynucleotide may be a synthetic RNA molecule.
1005272698
4 [0019] In any embodiment, the RNA polynucleotide may be a circular RNA (circRNA) molecule. [0020] In any embodiment, the RNA polynucleotide may be a complementary RNA (cRNA) molecule. [0021] In any embodiment, the RNA polynucleotide made be a self-amplifying RA (saRNA) molecule or trans-amplifying (taRNA) molecule. [0022] Further examples of various RNA molecule forms are described in Fang et al., (2022) Signal Transduction and Targeted Therapy, 7: article 94, incorporated herein by reference. [0023] In any embodiment herein, the RNA may further encode an N-terminal signal peptide for enabling secretion of the protein following translation thereof. The N-terminal signal peptide may comprise any amino acid sequence which enables the protein encoded by the RNA to be processed by ribosomes bound to the rough endoplasmic reticulum (ER) of a cell, and thereby results in threading of the protein into the ER. From the ER, the protein is capable of being transported to the plasma membrane and secreted from the mammalian cell. N-terminal secretion peptides are known to the skilled person and are further described herein. [0024] In preferred embodiments, the RNA may further comprise a 5’ untranslated region (UTR) and a 3’ UTR. The RNA may also comprise a 5’ cap analog, such as 7mG(5′)ppp(5′)NlmpNp. The RNA may also comprise a polyadenine (polyA) tail. The poly(A) tail may be non-segmented or segmented with a short spacer element. [0025] The RNA may comprise a chemical modification. Examples of suitable chemical modification include a N1-methylpseudouridine modification or a N1-ethylpseudouridine modification or may comprise any chemical modification described herein. [0026] Preferably, the polynucleotide has a uridine content of less than about 50%, less than about 45%, less than about 40%, less than about 35%, less than about 30%, less than about 25%, less than about 20% or less than about 15%. In preferred embodiments, the polynucleotide has a uridine content of between about 15% and about 35%, preferably between about 15% and about 25%.
1005272698
5 [0027] In preferred embodiments, the uridines in the polynucleotide are replaced with a chemical modification such as N1-methyl-pseudouridine. Preferably, at least 25%, at least 30%, at least 35%, at least 40%, at least 45%, at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, or 100% of the uridine nucleosides are replaced with N1-methyl- pseudouridine. [0028] In any embodiment of the invention, the RNA polynucleotide is in the form of a codon optimised RNA molecule, optionally depleted of uridine nucleosides. In any embodiment, the codon optimisations comprises conversion of codons encoding serine to UCG. [0029] In any embodiment, the protein encoded by the RNA polynucleotide is a chimeric or fusion protein comprising or consisting of: - one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. [0030] It will be appreciated that the chimeric or fusion protein encoded by the RNA may comprise the afore-mentioned domains in any order: for example: the one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P. gingivalis may be located N-terminally to the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, may be located N-terminally to the one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P. gingivalis, The domains may be directly joined in the context of the chimeric or fusion protein, or may be joined via a linker region, of one or more amino acid residues, as further defined herein. (In other words, the chimeric or fusion protein may comprises or consist of, N to C terminus: active site (K) - ABMs (A); or active site (R) – ABMs (A); or ABMs (A) – active site (K); or ABMs (A)-active site (R); or active site (R) – ABMs (A) - active site (K); or active site (R) – ABMs (A) - active site (R); or active site (K) – ABMs (A) - active site (R).)
1005272698
6 [0031] Exemplary amino acid sequences of an Arg- or Lys-gingipain of P. gingivalis, (and RNA sequences encoding the same) are further described herein. Preferably, the amino acid sequence of an active site of an Arg- or Lys- gingipain of P. gingivalis comprises the amino acid sequence of KAS or RAS (the Lysine or Arginine active site histidine sequence), ie a peptide including the active site histidine and surrounding area of the active site. [0032] In certain embodiments, the amino acid sequence of an active site of an Arg- gingipain of P. gingivalis, (also designated “R” herein), comprises the amino acid sequence of SEQ ID NO: 38, encoded by the RNA sequence as set forth in SEQ ID NO: 50, or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto. [0033] In certain embodiments, the amino acid sequence of an active site of a Lys- gingipain of P. gingivalis (also designated “K” herein), comprises the amino acid sequence of SEQ ID NO: 8, encoded by the RNA sequence as set forth in SEQ ID NO: 43, or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto. [0034] Preferably, the amino acid sequence of the active site of an Arg- or Lys- gingipain does not comprise the entire catalytic domain of the gingipain. [0035] In certain embodiments the chimeric or fusion protein encoded by the RNA polynucleotide comprises i) an amino acid sequence that comprises or consists of an amino acid sequence of the active site of an Arg-gingipain of P. gingivalis, or sequences that are at least 80% identical thereto; and ii) an amino acid sequence that comprises or consists of an amino acid sequence of the active site of a Lys-gingipain of P. gingivalis, or sequences that are at least 80% identical thereto. [0036] Optionally, the chimeric or fusion protein encoded by the RNA polynucleotide comprises at least two amino acid sequences that comprise or consist of an amino acid sequence of the active site of an Arg- or Lys-gingipain of P. gingivalis, or sequences that
1005272698
7 are at least 80% identical thereto. The at least two amino acid sequences may be located contiguously in the chimeric or fusion protein, or may be located in different locations within the chimeric or fusion protein. Optionally, one of the at least two amino acid sequences may be located at the N terminus of the chimeric or fusion protein while the second of the at least two amino acid sequences may be located at the C-terminus of the chimeric or fusion protein. Optionally, one of the at least two amino acid sequences may be located at the N or C terminus of the chimeric or fusion protein while the second of the at least two amino acid sequences may be located within the chimeric or fusion protein (ie not at either N or C termini). Optionally, the at least two amino acid sequences may (both) be located at the N terminus of the chimeric or fusion protein or the at least two amino acid sequences may (both) be located at the C-terminus of the chimeric or fusion protein. [0037] In any embodiment, the chimeric or fusion protein encoded by the RNA polynucleotide may further comprise: - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. [0038] Optionally, the amino acid sequence comprising the amino acid sequence of a DUF2436 domain is located between the amino acid sequence of the active site of the gingipain of P. gingivalis and the amino acid sequence of the one or more adhesin binding motifs (ABMs). (In other words, such that the chimeric or fusion protein comprises, N to C terminus or C to N terminus: active site (K) or (R) – DUF domain (D) – ABMs (A); or active site (K) or (R) – DUF domain (D) – ABMs (A) – active site (K) or (R)). [0039] In certain embodiments, the amino acid sequence of a DUF2436 domain of an Arg or Lys gingipain of P. gingivalis comprises or consists of the amino acid sequence of SEQ ID NO: 35 or 76, encoded by the RNA sequence as set forth in SEQ ID NO: 51, or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto.
1005272698
8 [0040] In any embodiment, a cysteine residue in the DUF2436 domain may be substituted to a serine or valine residue, preferably to a serine residue (such as shown in SEQ ID NO: 36). [0041] In any embodiment, the one or more adhesin binding motifs (ABMs) comprise or consist of the amino acid sequence of ABM2 and/or ABM1 (for example as set forth in SEQ ID NO: 22 and SEQ ID NO: 21, respectively, or comprising the amino acid sequence as set forth in SEQ ID NO: 24 (ABM2+1), or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto. Such amino acid sequences may be encoded by an RNA comprising the sequence set forth in SEQ ID NOs: 52, 53, 55, herein. [0042] The ABMs may further comprise ABM3 (eg SEQ ID NO: 73 or encoded by SEQ ID NO: 54). [0043] The ABMs may be provided in any order, but are preferably in the order ABM2 - ABM1 - ABM3. [0044] Optionally, the one or more adhesin binding motifs (ABMs) may comprise or consist of the amino acid sequence of ABM2, ABM1 and ABM3 (for example as set forth in SEQ ID NO: 25, 72 or 73 and encoded by an RNA comprising the sequence of SEQ ID NO: 56 or 77 or 78). [0045] In any embodiment, the one or more adhesin binding motifs may comprise one or more modifications selected from: a) one or more cysteine amino acid substitutions compared to the naturally occurring Arg- or Lys-gingipain sequences in corresponding regions; b) substitution of the proline and/or an asparagine residues in the sequence PxxN corresponding to, or at a position equivalent to, residues 6 to 9 of the sequence of SEQ ID NO: 21 (ABM1); c) substitution of the motif NxFA to SxYQ in the sequence, corresponding to, or at a position equivalent to residues 2 to 5 of the sequence of SEQ ID NO: 21 (ABM1);
1005272698
9 d) substitution of the second tyrosine residue, corresponding to or at a position equivalent to residues at position 5 of SEQ ID NO: 22 (ABM2), and of the tryptophan residue, corresponding to or at a position equivalent to residue at position 23 of SEQ ID NO: 21 (ABM1) to alanine residues. [0046] The one or more cysteine amino acid substitutions may be a substitution to a serine residue or to a valine residue. Preferably, the one or more cysteine substitutions may comprise one or more substitutions to a serine residue. [0047] In certain embodiments, only one cysteine residue may be substituted. In other embodiments, two or three cysteine residues may be substituted. In particularly preferred embodiments, the cysteine residues are substituted to a combination of valine and serine residues. In other embodiments, all substituted cysteine residues are substituted to serine or all substitute cysteine residues are substituted to valine. [0048] The motif PxxN (eg PVQN, SEQ ID NO: 106), corresponding to or at a position equivalent to residues 6 to 9 of SEQ ID NO: 21, may comprise a substitution of the proline and asparagine residues. [0049] The proline amino acid substitution is preferably a substitution to an alanine residue. [0050] The asparagine amino acid substitution may be a substitution to a proline residue or an alanine residue. Preferably the asparagine residue is substituted to a proline residue. In other embodiments, the asparagine residue is not substituted. [0051] Preferably, the substitution is from PxxN to AxxP, (eg AVQP, SEQ ID NO: 107) (such as exemplified in the amino acid sequences of SEQ ID NOs: 30 to 32). [0052] In certain further embodiments, the one or more adhesin binding motifs comprise the amino acid sequence as set forth in any one of SEQ ID NOs: 21 to 25, and comprising: a) one or more cysteine amino acid substitutions compared to the naturally occurring Arg- or Lys-gingipain sequences in corresponding regions, preferably substitution of all cysteine residues; and
1005272698
10 b) substitution of the motif PxxN corresponding to, or at a position equivalent to, residues 6 to 9 of the sequence of SEQ ID NO: 21 (ABM1), to AxxP. [0053] Accordingly, in such embodiments, the one or more adhesin binding motifs comprise or consist the amino acid sequence as set forth in any one of SEQ ID NOs: 26 to 34, or sequences at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto, provided that the sequences comprise the aforementioned substitutions of the cysteine and proline and asparagine residues. [0054] In particularly preferred embodiments of the invention, the RNA (preferably mRNA) encodes a chimeric or fusion protein comprising or consisting of: [0055] a) - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, [0056] wherein preferably the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 18, SEQ ID NO: 39 or any of SEQ ID NOs: 58 to 63; or SEQ ID NO: 81 to 87; or [0057] b) - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto,
1005272698
11 - wherein preferably the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 4, 12, 16 or 20; - more preferably wherein the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 4; or [0058] c) - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, - wherein preferably the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 14; or [0059] d) - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, - wherein preferably the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 10 or SEQ ID NO: 108; or [0060] e) - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto,
1005272698
12 - wherein preferably the chimeric or fusion protein comprises the amino acid sequence of SEQ ID NO: 6. [0061] In certain embodiments a linker region may be included between the amino acid sequence of a DUF2436 domain and the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain. The skilled person will be familiar with suitable linkers which may be used for joining any of the aforementioned domains. In certain embodiments, and as exemplified in the Tables herein, the linker comprises or consists of the sequence EVEDDSP (SEQ ID NO: 109). In certain embodiments, the chimeric or fusion protein encoded by the RNA does not include a linker sequence between the DUF2436 domain and one or more adhesin domains. [0062] In certain embodiments, the amino acid sequence of an AMB2 domain as defined herein (eg in SEQ ID NO: 22 or SEQ ID NO 24 to 34) may further comprise at its N-terminus, the amino acid sequence of EVEDDSP (SEQ ID NO: 109), which is derived from the native P. gingivalis gingipain polyprotein sequence. [0063] It will also be appreciated that in some circumstances, N-terminal methionine residues of polypeptides are cleaved following translation of the mRNA into protein. As such, the present disclosure provides basis for the generation of a chimeric or fusion protein from an RNA molecule, wherein the chimeric or fusion protein does not comprise an N terminal methionine residue. [0064] In other embodiments of the invention, the RNA molecule comprises or consists of a nucleotide sequence encoding a protein comprising or consisting of the amino acid sequence of any one of: SEQ ID NO: 2, SEQ ID NO: 8 or SEQ ID NO: 38. [0065] In particularly preferred embodiments of the invention, the RNA molecule comprises or consists of a nucleotide sequence of any one of: a) SEQ ID NO: 48 or 57 b) SEQ ID NO: 45, 47 or 49 or SEQ ID NO: 41; c) SEQ ID NO: 46; d) SEQ ID NO: 44; e) SEQ ID NO: 42
1005272698
13 or sequences at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto. [0066] In other embodiments of the invention, the RNA comprises or consists of a nucleotide sequence of any one of: SEQ ID NO: 40, SEQ ID NO: 43 or SEQ ID NO: 50. [0067] The invention further provides for the use of any DNA polynucleotide described herein (and particularly any DNA polynucleotide comprising or consisting of a sequence exemplified in Table 1). Optionally, the use of the DNA polynucleotide may be for obtaining an RNA polynucleotide of the invention. [0068] The present invention also provides a composition, including a pharmaceutical composition comprising an RNA as described herein. Preferably, the composition comprises one or more pharmaceutically acceptable excipients. Optionally, the RNA may comprise one or more agents for enabling delivery of the RNA to a mammalian cell, and thereby enabling translation of the RNA in the cell. In any embodiment, the composition may comprise a combination of one or more of the RNA molecules described herein. [0069] The present invention further provides compositions comprising an RNA molecule as described herein, wherein the composition also comprises a lipid component. The RNA (e.g., RNA) vaccines of the disclosure can be formulated using one or more liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles [0070] In preferred embodiments, the RNA is formulated in a lipid nanoparticle. [0071] In one embodiment, the RNA as described herein is the only polynucleotide species present in the composition or in the liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles. Preferably, the polynucleotide as described herein is the only active ingredient present in the composition or in the liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles. [0072] In further examples, the composition or liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles may comprise more than one RNA
1005272698
14 (polynucleotide) species, as described herein (eg thereby providing RNA molecules encoding more than one chimeric or fusion protein amino acid sequence). Further still, the composition or the liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles may comprise a single polynucleotide construct containing one or more RNA sequences as described herein (and thereby also encoding more than one chimeric or fusion protein amino acid sequence). Accordingly, the present invention contemplates the provision of compositions, liposomes, lipid vesicle, lipoplexes (such as a lipid-polycation complex), or lipid nanoparticles for delivering combinations of two or more of any of the RNA molecules described herein. [0073] In any embodiment, the invention provides a lipid nanoparticle or other nanovehicle, such as nanopolymer, for delivery of the polynucleotide to a subject in need thereof. [0074] Lipid nanoparticles are well known in the art and are further described herein. Preferably the lipid nanoparticle comprises a cationic and/or ionisable lipid, a phospholipid, a PEG (or PEGylated) lipid, and a structural lipid. [0075] In any embodiment, the lipid nanoparticle may comprise: - a cationic and/or ionisable lipid comprising from about 25 % to about 75 mol % of the total lipid present in the nanoparticle; - a sterol (structural lipid) comprising from about 5 mol % to about 60 mol % of the total lipid present in the nanoparticle; - a phospholipid comprising from about 5 mol % to about 50 mol % of the total lipid present in the nanoparticle; - a PEGylated lipid comprising from about 0.5 mol % to 20 mol % of the total lipid present in the nanoparticle. [0076] In non-limiting examples, the lipid nanoparticle comprises: - an ionisable lipid in the form of [(4-hydroxybutyl)azanediyl]di(hexane-6,1-diyl) bis(2-hexyldecanoate) (ALC-0315), - a sterol in the form of cholesterol,
1005272698
15 - a phospholipid in the form of distearoylphosphatidylcholine (DSPC), and - a PEGylated lipid in the form of 2-[(polyethylene glycol)-2000]-N,N- ditetradecylacetamide (ALC-0159). [0077] Preferably the lipids are present in the lipid nanoparticle at molar lipid ratios (%) of 46.3 ALC-0315: 42.7 cholesterol : 9.4 DSPC : 1.6 ALC-0159, optionally in Tris/sucrose buffer (25 mM Tris pH 7.4, 8.8% sucrose w/v). [0078] The present invention also provides a method for producing a lipid nanoparticle comprising an RNA, encoding a protein, or a chimeric or fusion protein as described herein. Preferably the method comprises formulating any RNA molecule of the invention, with one or more lipids useful for producing a lipid nanoparticle. Preferably the lipid components comprise a phospholipid, a PEG lipid, and a structural lipid. [0079] The present invention also provides a nucleic acid construct or vector, comprising a polynucleotide as described herein. [0080] The vector may be any vector suitable for production of RNA from a DNA template. The vector may additionally comprise 3’UTR and 5’UTRs and polyadenine fragments. [0081] Examples of such vectors include: IVT RNA vector or similar vectors that comprise T7, T3 and SP6 signals for expression. The vector can be from a plasmid or produced through PCR or Phi29 DNA polymerase (e.g. GenomiPhi™ V2 DNA) or other bacterial constructs. [0082] The vector may be a self-amplifying RNA replicon, such as, but not limited to a self-amplifying RNA vector from an alphavirus, optionally Venezuelan Equine Encephalitis Virus (VEEV), bipartite VEEV, or variants thereof (including the TC83 mutated variant). Examples of self-amplifying mRNA platforms are known to the skilled person, and are described for example in Maruggi et al., (2017), Vaccines 35: 361-368, incorporated herein by reference. [0083] 5’ capping of the polynucleotide may be performed using any commercially available capping reagent. Such reagents are known to the skilled person, such as the commercial capping reagent Cap1 from TriLink Biotechnologies Inc. Other capping reagents may be used, including but not limited to Cap 0 and Cap 2.
1005272698
16 [0084] The present invention provides a method for eliciting an immune response to P. gingivalis in a subject in need thereof, the method comprising administering to the subject, a polynucleotide, vector, nanoparticle or composition described herein. [0085] The invention provides a method for eliciting an immune response to P. gingivalis in a subject in need thereof, the method comprising administering to the subject, a composition comprising: - an RNA as described herein, wherein said RNA is capable of being translated in a cell of a mammalian subject to produce the polypeptide encoded by the polynucleotide; - optionally, an agent for enabling delivery of the RNA into mammalian cells. [0086] The agent for delivery of the RNA into mammalian cells may be any suitable agent known to the skilled person for delivery of RNA. Such agents may include: cell penetrating peptides, lipid-based formulations. [0087] The invention provides a method for eliciting an immune response to P. gingivalis in a subject in need thereof, the method comprising administering to the subject, a nanoparticle composition comprising: - a lipid component; and - an RNA as described herein, wherein said RNA is capable of being translated in a cell of the subject to produce the polypeptide encoded by the polynucleotide. [0088] The invention also provides a method for producing a chimeric or fusion protein as described herein in a mammalian cell, the method comprising contacting the mammalian cell with a composition comprising: - an RNA as described herein, wherein said RNA is capable of being translated in the mammalian cell to produce the protein; - optionally, an agent for enabling delivery of the RNA into mammalian cells. [0089] The agent for delivery of the RNA into mammalian cells may be any suitable agent known to the skilled person for delivery of RNA. Such agents may include: cell penetrating peptides, lipid-based formulations.
1005272698
17 [0090] The invention also provides a method for producing a chimeric or fusion protein as described herein in a mammalian cell, the method comprising contacting the mammalian cell with a nanoparticle composition, the composition comprising: - a lipid component; and - an RNA as described herein, wherein said RNA is capable of being translated in the mammalian cell to produce the protein. [0091] Preferably the lipid component comprises a cationic and/or ionisable lipid, a phospholipid, a PEG lipid, and a structural lipid. [0092] The invention also provides a method for delivering an RNA to a mammalian cell in a subject in need thereof, said method comprising administering to a subject in need thereof, a nanoparticle composition, the composition comprising: - a lipid component; and - an RNA comprising a polynucleotide sequence as described herein, wherein said RNA is capable of being translated in the mammalian cell to produce the chimeric or fusion protein described herein; wherein the administering comprises contacting said mammalian cell with the nanoparticle composition, thereby enabling delivery of the RNA to the mammalian cell. [0093] Preferably the lipid component comprises a cationic and/or ionisable lipid, a phospholipid, a PEG lipid, and a structural lipid. [0094] In any embodiment, the ionisable lipid may be substituted or combined with an adjuvant lipidoid for enhancing RNA delivery. [0095] The present invention also provides the use of a polynucleotide, vector, or nanoparticle described herein, in the manufacture of a composition for eliciting an immune response to P. gingivalis in a subject. [0096] The present invention also provides the use of i) a lipid component as described herein, and ii) an RNA as described herein, in the manufacture of a composition for delivering the RNA to a mammalian cell in a subject in need thereof.
1005272698
18 [0097] The present invention also provides a polynucleotide, vector, nanoparticle or composition as described herein, for use in eliciting an immune response to P. gingivalis in a subject. [0098] As used herein “at least 80% identity” should be taken to provide basis for “at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identity”. [0099] As used herein a sequence defined as having “at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identity” to a particular SEQ ID NO, may also be referred to as a “substitutional variant”. [0100] As used herein, except where the context requires otherwise, the term "comprise" and variations of the term, such as "comprising", "comprises" and "comprised", are not intended to exclude further additives, components, integers or steps. [0101] Further aspects of the present invention and further embodiments of the aspects described in the preceding paragraphs will become apparent from the following description, given by way of example and with reference to the accompanying drawings. Brief description of the drawings [0102] Figure 1: Schematic showing domain architecture of Kgp polyprotein and chimeric proteins derived therefrom. (KDcAK1n: K = Kgp active site, Dc = N terminally truncated DUF2436 domain, A = ABM2, ABM1 and ABM3, K1n = C terminally truncated CAD domain; KDAK: K = Kgp active site, D = DUF2436 domain, A = ABM2, ABM1 and ABM3; KDAK-3S-AVQP: K = Kgp active site, D = DUF2436 domain, A = ABM2, ABM1 and ABM3, 3S-AVQP = PVQN (SEQ ID NO: 106) to AVQP (SEQ ID NO: 107) substitution in ABM1 domain and substitution of the cysteine residues in the D and A domains to serine). [0103] Figure 2: Exemplary structure of mRNA. Cap = 5’ cap for maximising RNA stability; 5’UTR and 3’ UTR = 5’ and 3’ untranslated sequences; SP = signal peptide;
1005272698
19 Antigen sequence = mRNA sequence encoding protein antigen for expression in cell upon translation of mRNA; Poly(A) tail: polyadenosine tail for providing RNA stability and maximising translation. [0104] Figure 3: Expression and secretion by HeLa cells of mRNA constructs encoding chimeric proteins. A. HeLa supernatant. Lane 1 = SEAP-KDcAK1n, Lane 2 = KDcAK1n (no secretion peptide), Lane 3 = Mock, Lane 4 = Protein size marker. B. HeLa supernatant Lane 1: SEAP-KDAK, Lane 2 = SEAP-KDAK-3S-AVQP, Lane 3 = Mock, Lane 5 = Protein size marker. C. HeLa supernatant Lane 1 = SEAP-KDAK-3S-AVQP, Lane 2 = KDAK-3S-AVQP (no secretion peptide), Lane 3 = Mock, Lane 4 = Protein size marker. SEAP = SEAP secretion peptide from secreted embryonic alkaline phosphatase. Mock = negative control (mock transfection). [0105] Figure 4: Schematic of vaccination protocol for assessing immunogenicity of candidate vaccines. [0106] Figure 5: Immunogenicity of mRNA vaccines. A. KDAK-3S-AVQP total IgG titre. B. KDAK-3S-AVQP IgG1 titres. C. KDAK-3S-AVQP IgG2a titres. D. Heat-killed P. gingivalis total IgG titres. E. Heat-killed P. gingivalis IgG1 titres. F. Heat-killed P. gingivalis IgG2a titres. G. Total IgG Kgpcat titres. H. Kgpcat IgG titres. I. KAS2 titres. Naïve = unvaccinated; mψ1-Neg = mRNA encoding KDAK-3S-AVQP but not capped; Chimera = mRNA encoding KDcAK1n (SEQ ID NO: 2); KDAK = mRNA encoding KDAK protein (SEQ ID NO: 4); KDAK-3S-AVQP = RNA encoding KDAK-3S-AVQP protein (SEQ ID NO: 6). m1ψ = N1-methylpseudouridine modified mRNA sequence. Protein control: alum- adjuvanted protein KDAK-3S-AVQP at 200 µg. * = p ≤ 0.05, ** = p ≤ 0.01, *** = p ≤ 0.001, **** = p ≤ 0.0001. [0107] Figure 6: schematic of study protocol for determining in vivo efficacy of vaccines (therapeutic model). [0108] Figure 7: P. gingivalis induced alveolar bone loss (mm) following vaccination with different mRNA constructs. Naïve = uninfected. Infected = no vaccination control. Positive control: alum-adjuvanted protein KDAK-3S-AVQP at 200 µg. m1ψ = modified mRNA sequence. * = p ≤ 0.05, ** = p ≤ 0.01, *** = p ≤ 0.001, **** = p ≤ 0.0001.
1005272698
20 [0109] Figure 8: Antibody responses in mice following alveolar bone loss experiments. A. KDAK-3S-AVQP total IgG titres. B. KDAK-3S-AVQP IgG1 titres. C. KDAK-3S-AVQP IgG2a titres. D. Heat-killed P. gingivalis total IgG titres. E. Heat-killed P. gingivalis IgG1 titres. F. Heat-killed P. gingivalis IgG2a titres. G. Kgpcat total IgG titres. H. Kgpcat IgG1 titres. I. Kgpcat IgG2a titres. J. KAS2 titres. Naïve = unvaccinated; infected = no vaccine control; KDAK = mRNA encoding KDAK protein (SEQ ID NO: 4); KDAK-3S-AVQP = mRNA encoding KDAK-3S-AVQP protein (SEQ ID NO: 6). m1ψ = N1- methylpseudouridine modified mRNA sequence. Protein control: alum-adjuvanted protein KDAK-3S-AVQP at 200 µg. * = p ≤ 0.05, ** = p ≤ 0.01, *** = p ≤ 0.001, **** = p ≤ 0.0001. [0110] Figure 9: Expression and secretion of constructs encoding truncated antigens. A. Western blot of lysates at 24 hours after transfection. Left hand panel: using anti-KDAK-3S-AVQP antibody. Right hand panel: using anti-KAS antibody. B. Western blot of supernatants at 24 hours after transfection. Left hand panel: using anti-KAS antibody. Right hand panel: using anti-KDAK-3S-AVQP antibody. A and B: Lanes: L = ladder; 1 = K; 2 = KA; 3 = KD; 4 = DA; 5 = KDAΔABM3; 6 = KDA; 7 = RDA; 8 = KDA-3S- AVQP (positive control); 9 = mock transfection (negative control). All constructs included SEAP secretion peptide. C. Expression of K over time compared to KDAK-3S-AVQP: time-lapse of K expression at 6 hours, 24 hours and 48 hours after transfection. Lanes: L = ladder; 1 = K; 2 @ 6 hours = KDAK-3S-AVQP @ 6 hours; 3 = Mock transfection @ 6 hours; 4 = K; 5 = KDAK-3S-AVQP @ 24 hours; 6 = Mock transfection @ 24 hours; 7 = K @ 48 hours 8 = KDA-3S-AVQP @ 48 hours; 9 = mock transfection @ 48 hours. All constructs included SEAP secretion peptide. [0111] Figure 10: P. gingivalis induced alveolar bone loss (mm) following vaccination with different mRNA constructs. Naïve = uninfected. Infected = no vaccination control. Positive control: alum adjuvanted protein KDAK-3S-AVQP at 200 µg. mψ1 = modified mRNA sequence. * = p ≤ 0.05, ** = p ≤ 0.01, *** = p ≤ 0.001, **** = p ≤ 0.0001. [0112] Figure 11: Antibody responses in mice following alveolar bone loss experiments using mRNA vaccine constructs encoding truncated antigens. A. KDAK-3S-AVQP total IgG titres. B. KDAK-3S-AVQP IgG1 titres. C. KDAK-3S-AVQP IgG2a titres. D. Heat-killed P. gingivalis total IgG titres. E. Heat-killed P. gingivalis IgG1 titres. F. Heat-killed P. gingivalis IgG2a titres. G. Kgpcat total IgG titres. H. Kgpcat IgG1 titres. I. Kgpcat IgG2a titres. J. KAS2 titres. Naïve = unvaccinated; infected = no vaccine
1005272698
21 control; KDAK-3S-AVQP = mRNA encoding KDAK-3S-AVQP protein (SEQ ID NO: 6). K. Kgp-RgpA complex total IgG titres. L. Kgp-RgpA complex IgG1 titres. M. Kgp-RpgA complex IgG2a titres. m1ψ = N1-methylpseudouridine modified mRNA sequence. KA = mRNA encoding KA protein (SEQ ID NO: 18). KD = mRNA encoding KD protein (SEQ ID NO: 10). DA = mRNA encoding DA protein (SEQ ID NO: 14). KDA21 = mRNA encoding KDAΔAMB3 protein (SEQ ID NO: 16). Protein control: alum-adjuvanted protein KDAK-3S- AVQP at 200µg. * = p ≤ 0.05, ** = p ≤ 0.01, *** = p ≤ 0.001, **** = p ≤ 0.0001. [0113] Figure 12: Immunogenicity of truncated antigens RDA and K. A. KDAK-3S- AVQP total IgG titres. B. KDAK-3S-AVQP IgG1 titres. C. KDAK-3S-AVQP IgG2a titres. D. Heat-killed P. gingivalis total IgG titres. E. Heat-killed P. gingivalis IgG1 titres. F. Heat- killed P. gingivalis IgG2a titres. G. Kgpcat total IgG titres. H. Kgpcat IgG1 titres. I. Kgpcat IgG2a titres. J. KAS2 titres. K. RgpA-Kgp complex total IgG titres. L. RgpA-Kgp complex l IgG1 titres M. RgpA-Kgp complex IgG2a titres. Naïve = unvaccinated; K = mRNA encoding K protein (SEQ ID NO: 8 ). KD = mRNA encoding KD protein (SEQ ID NO: 10). RDA = mRNA encoding RDA protein (SEQ ID NO: 20). * = p ≤ 0.05, ** = p ≤ 0.01, *** = p ≤ 0.001, **** = p ≤ 0.0001. [0114] Figure 13: schematic of study protocol for determining in vivo efficacy of vaccines in prophylaxis model. [0115] Figure 14: Number of KDAK-3S-AVQP-specific interferon gamma secreting splenocytes, from a prophylactic vaccine murine model, as determined by ex vivo ELISpot.. Naïve = uninfected. Infected = no vaccination control. Positive control: alum-adjuvanted protein KDAK-3S-AVQP at 200 µg. m1ψ = modified mRNA sequence. * = p ≤ 0.05, ** = p ≤ 0.01, *** = p ≤ 0.001, **** = p ≤ 0.0001. [0116] Figure 15: P. gingivalis induced alveolar bone loss (mm) following prophylactic vaccination with different mRNA constructs prior to challenge with P. gingivalis. Naïve = uninfected. Infected = no vaccination control. Positive control: alum- adjuvanted protein KDAK-3S-AVQP at 200 µg. m1ψ = modified mRNA sequence. * = p ≤ 0.05, ** = p ≤ 0.01, *** = p ≤ 0.001, **** = p ≤ 0.0001. [0117] Figure 16: Antibody responses in mice following alveolar bone loss experiments in the prophylaxis model. A. total IgG titres. B. IgG1 titres. C. IgG2a titres. D. Heat-killed P. gingivalis total IgG titres. E. Heat-killed P. gingivalis IgG1 titres. F. Heat-
1005272698
22 killed P. gingivalis IgG2a titres. G. RgpA-Kgp complex total IgG titres. H. RgpA-Kgp complex IgG1 titres. I. RgpA-Kgp complex IgG2a titres. Naïve = unvaccinated; infected = no vaccine control; KDAK-3S-AVQP = mRNA encoding KDAK-3S-AVQP protein (SEQ ID NO: 6). m1ψ = N1-methylpseudouridine modified mRNA sequence. * = p ≤ 0.05, ** = p ≤ 0.01, *** = p ≤ 0.001, **** = p ≤ 0.0001. Sequence information [0118] Table 1: DNA and amino acid sequences Descriptor SEQ ID No Sequence KDcAK1n (DNA 1 AATACCGGAGTCAGCTTTGCAAACTATACAGCGCATGG sequence) ATCTGAGACCGCATGGGCTGATCCACTTCTGACTACTT CTCAACTGAAAGCACTCACTAATAAGGACAAATGGGGA GACAATACGGGTTACCAGTTCTTGTTGGATGCCGATCA CAATACATTCGGAAGTGTCATTCCGGCAACCGGTCCTC TCTTTACCGGAACAGCTTCTTCCAATCTTTACAGTGCGA ACTTCGAGTATTTGATCCCGGCCAATGCCGATCCTGTT GTTACTACACAGAATATTATCGTTACAGGACAGGGTGAA GTTGTAATCCCCGGTGGTGTTTACGACTATTGCATTACG AACCCGGAACCTGCATCCGGAAAGATGTGGATCGCAG GAGATGGAGGCAACCAGCCTGCACGTTATGACGATTTC ACATTCGAAGCAGGCAAGAAGTACACCTTCACGATGCG TCGCGCCGGAATGGGAGATGGAACTGATATGGAAGTC GAAGACGATTCACCTGCAAGCTATACCTACACGGTGTA TCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCTA CGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCAT GAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTATC TCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATCCA ATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTTCA GTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTAA TGGTACCCCGAATCCGAATCCAAATCCGAATCCGAATC CGGGAACAACACTTTCCGAATCATTCGAAAATGGTATTC CGGCATCTTGGAAGACGATCGATGCAGACGGTGACGG GCATGGCTGGAAACCTGGAAATGCTCCCGGAATCGCT GGCTACAATAGCAATGGTTGTGTATATTCAGAGTCATTC
1005272698
23 GGTCTTGGTGGTATAGGAGTTCTTACCCCTGACAACTA TCTGATAACACCGGCATTGGATTTGCCTAACGGAGGT KDcAK1n (amino 2 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKWGD acid sequence) NTGYQFLLDADHNTFGSVIPATGPLFTGTASSNLYSANFE YLIPANADPVVTTQNIIVTGQGEVVIPGGVYDYCITNPEPAS GKMWIAGDGGNQPARYDDFTFEAGKKYTFTMRRAGMGD GTDMEVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVA AGNHEYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLT GSSVGQKVTLKWDAPNGTPNPNPNPNPNPGTTLSESFEN GIPASWKTIDADGDGHGWKPGNAPGIAGYNSNGCVYSES FGLGGIGVLTPDNYLITPALDLPNGG KDAK (DNA 3 AATACCGGAGTCAGCTTTGCAAACTATACAGCGCATGG sequence) ATCTGAGACCGCATGGGCTGATCCACTTCTGACTACTT CTCAACTGAAAGCACTCACTAATAAGGACAAAGCAGAA GGTTCCCGTGAAGTAAAACGGATCGGAGACGGTCTTTT CGTTACGATCGAACCTGCAAACGATGTACGTGCCAACG AAGCCAAGGTTGTGCTTGCGGCAGACAACGTATGGGG AGACAATACGGGTTACCAGTTCTTGTTGGATGCCGATC ACAATACATTCGGAAGTGTCATTCCGGCAACCGGTCCT CTCTTTACCGGAACAGCTTCTTCCAATCTTTACAGTGCG AACTTCGAGTATTTGATCCCGGCCAATGCCGATCCTGT TGTTACTACACAGAATATTATCGTTACAGGACAGGGTGA AGTTGTAATCCCCGGTGGTGTTTACGACTATTGCATTAC GAACCCGGAACCTGCATCCGGAAAGATGTGGATCGCA GGAGATGGAGGCAACCAGCCTGCACGTTATGACGATTT CACATTCGAAGCAGGCAAGAAGTACACCTTCACGATGC GTCGCGCCGGAATGGGAGATGGAACTGATATGGAAGT CGAAGACGATTCACCTGCAAGCTATACCTACACGGTGT ATCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCT ACGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCA TGAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTAT CTCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATCC AATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTTCA GTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTAA TGGTACCCCGAATCCGAATCCAAATCCGAATCCGAATC CGGGAACAAATACCGGAGTCAGCTTTGCAAACTATACA GCGCATGGATCTGAGACCGCATGGGCTGATCCACTTCT
1005272698
24 GACTACTTCTCAACTGAAAGCACTCACTAATAAGGACAA A KDAK (amino acid 4 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKAEG sequence) SREVKRIGDGLFVTIEPANDVRANEAKVVLAADNVWGDNT GYQFLLDADHNTFGSVIPATGPLFTGTASSNLYSANFEYLI PANADPVVTTQNIIVTGQGEVVIPGGVYDYCITNPEPASGK MWIAGDGGNQPARYDDFTFEAGKKYTFTMRRAGMGDGT DMEVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAG NHEYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGS SVGQKVTLKWDAPNGTPNPNPNPNPNPGTNTGVSFANY TAHGSETAWADPLLTTSQLKALTNKDK KDAK-3S-AVQP 5 AATACCGGAGTCAGCTTTGCAAACTATACAGCGCATGG (DNA sequence) ATCTGAGACCGCATGGGCTGATCCACTTCTGACTACTT CTCAACTGAAAGCACTCACTAATAAGGACAAAGCAGAA GGTTCCCGTGAAGTAAAACGGATCGGAGACGGTCTTTT CGTTACGATCGAACCTGCAAACGATGTACGTGCCAACG AAGCCAAGGTTGTGCTTGCGGCAGACAACGTATGGGG AGACAATACGGGTTACCAGTTCTTGTTGGATGCCGATC ACAATACATTCGGAAGTGTCATTCCGGCAACCGGTCCT CTCTTTACCGGAACAGCTTCTTCCAATCTTTACAGTGCG AACTTCGAGTATTTGATCCCGGCCAATGCCGATCCTGT TGTTACTACACAGAATATTATCGTTACAGGACAGGGTGA AGTTGTAATCCCCGGTGGTGTTTACGACTATAGCATTAC GAACCCGGAACCTGCATCCGGAAAGATGTGGATCGCA GGAGATGGAGGCAACCAGCCTGCACGTTATGACGATTT CACATTCGAAGCAGGCAAGAAGTACACCTTCACGATGC GTCGCGCCGGAATGGGAGATGGAACTGATATGGAAGT CGAAGACGATTCACCTGCAAGCTATACCTACACGGTGT ATCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCT ACGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCA TGAGTATAGCGTGGAAGTTAAGTACACAGCCGGCGTAT CTCCGAAGGTAAGTAAAGACGTTACGGTAGAAGGATCC AATGAATTTGCTGCTGTACAGCCGCTGACCGGTAGTTC AGTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTA ATGGTACCCCGAATCCGAATCCAAATCCGAATCCGAAT CCGGGAACAAATACCGGAGTCAGCTTTGCAAACTATAC AGCGCATGGATCTGAGACCGCATGGGCTGATCCACTTC
1005272698
25 TGACTACTTCTCAACTGAAAGCACTCACTAATAAGGACA AA KDAK-3S-AVQP 6 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKAEG (amino acid SREVKRIGDGLFVTIEPANDVRANEAKVVLAADNVWGDNT sequence) GYQFLLDADHNTFGSVIPATGPLFTGTASSNLYSANFEYLI PANADPVVTTQNIIVTGQGEVVIPGGVYDYSITNPEPASGK MWIAGDGGNQPARYDDFTFEAGKKYTFTMRRAGMGDGT DMEVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAG NHEYSVEVKYTAGVSPKVSKDVTVEGSNEFAAVQPLTGS SVGQKVTLKWDAPNGTPNPNPNPNPNPGTNTGVSFANY TAHGSETAWADPLLTTSQLKALTNKDK K (DNA 7 AATACCGGAGTCAGCTTTGCAAACTATACAGCGCATGG sequence) ATCTGAGACCGCATGGGCTGATCCACTTCTGACTACTT CTCAACTGAAAGCACTCACTAATAAGGACAAA K (amino acid 8 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDK sequence) KD (DNA 9 AATACCGGAGTCAGCTTTGCAAACTATACAGCGCATGG sequence) ATCTGAGACCGCATGGGCTGATCCACTTCTGACTACTT CTCAACTGAAAGCACTCACTAATAAGGACAAAGCAGAA GGTTCCCGTGAAGTAAAACGGATCGGAGACGGTCTTTT CGTTACGATCGAACCTGCAAACGATGTACGTGCCAACG AAGCCAAGGTTGTGCTTGCGGCAGACAACGTATGGGG AGACAATACGGGTTACCAGTTCTTGTTGGATGCCGATC ACAATACATTCGGAAGTGTCATTCCGGCAACCGGTCCT CTCTTTACCGGAACAGCTTCTTCCAATCTTTACAGTGCG AACTTCGAGTATTTGATCCCGGCCAATGCCGATCCTGT TGTTACTACACAGAATATTATCGTTACAGGACAGGGTGA AGTTGTAATCCCCGGTGGTGTTTACGACTATTGCATTAC GAACCCGGAACCTGCATCCGGAAAGATGTGGATCGCA GGAGATGGAGGCAACCAGCCTGCACGTTATGACGATTT CACATTCGAAGCAGGCAAGAAGTACACCTTCACGATGC GTCGCGCCGGAATGGGAGATGGAACTGATATG KD (amino acid 10 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKAEG sequence) SREVKRIGDGLFVTIEPANDVRANEAKVVLAADNVWGDNT GYQFLLDADHNTFGSVIPATGPLFTGTASSNLYSANFEYLI PANADPVVTTQNIIVTGQGEVVIPGGVYDYCITNPEPASGK
1005272698
26 MWIAGDGGNQPARYDDFTFEAGKKYTFTMRRAGMGDGT DM KDA (DNA 11 AATACCGGAGTCAGCTTTGCAAACTATACAGCGCATGG sequence) ATCTGAGACCGCATGGGCTGATCCACTTCTGACTACTT CTCAACTGAAAGCACTCACTAATAAGGACAAAGCAGAA GGTTCCCGTGAAGTAAAACGGATCGGAGACGGTCTTTT CGTTACGATCGAACCTGCAAACGATGTACGTGCCAACG AAGCCAAGGTTGTGCTTGCGGCAGACAACGTATGGGG AGACAATACGGGTTACCAGTTCTTGTTGGATGCCGATC ACAATACATTCGGAAGTGTCATTCCGGCAACCGGTCCT CTCTTTACCGGAACAGCTTCTTCCAATCTTTACAGTGCG AACTTCGAGTATTTGATCCCGGCCAATGCCGATCCTGT TGTTACTACACAGAATATTATCGTTACAGGACAGGGTGA AGTTGTAATCCCCGGTGGTGTTTACGACTATTGCATTAC GAACCCGGAACCTGCATCCGGAAAGATGTGGATCGCA GGAGATGGAGGCAACCAGCCTGCACGTTATGACGATTT CACATTCGAAGCAGGCAAGAAGTACACCTTCACGATGC GTCGCGCCGGAATGGGAGATGGAACTGATATGGAAGT CGAAGACGATTCACCTGCAAGCTATACCTACACGGTGT ATCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCT ACGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCA TGAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTAT CTCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATCC AATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTTCA GTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTAA TGGTACCCCGAATCCGAATCCAAATCCGAATCCGAATC CGGGAACA KDA (amino acid 12 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKAEG sequence) SREVKRIGDGLFVTIEPANDVRANEAKVVLAADNVWGDNT GYQFLLDADHNTFGSVIPATGPLFTGTASSNLYSANFEYLI PANADPVVTTQNIIVTGQGEVVIPGGVYDYCITNPEPASGK MWIAGDGGNQPARYDDFTFEAGKKYTFTMRRAGMGDGT DMEVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAG NHEYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGS SVGQKVTLKWDAPNGTPNPNPNPNPNPGT
1005272698
27 DA (DNA 13 GCAGAAGGTTCCCGTGAAGTAAAACGGATCGGAGACG sequence) GTCTTTTCGTTACGATCGAACCTGCAAACGATGTACGT GCCAACGAAGCCAAGGTTGTGCTTGCGGCAGACAACG TATGGGGAGACAATACGGGTTACCAGTTCTTGTTGGAT GCCGATCACAATACATTCGGAAGTGTCATTCCGGCAAC CGGTCCTCTCTTTACCGGAACAGCTTCTTCCAATCTTTA CAGTGCGAACTTCGAGTATTTGATCCCGGCCAATGCCG ATCCTGTTGTTACTACACAGAATATTATCGTTACAGGAC AGGGTGAAGTTGTAATCCCCGGTGGTGTTTACGACTAT TGCATTACGAACCCGGAACCTGCATCCGGAAAGATGTG GATCGCAGGAGATGGAGGCAACCAGCCTGCACGTTAT GACGATTTCACATTCGAAGCAGGCAAGAAGTACACCTT CACGATGCGTCGCGCCGGAATGGGAGATGGAACTGAT ATGGAAGTCGAAGACGATTCACCTGCAAGCTATACCTA CACGGTGTATCGTGACGGCACGAAGATCAAGGAAGGT CTGACAGCTACGACATTCGAAGAAGACGGTGTAGCTGC AGGCAATCATGAGTATTGCGTGGAAGTTAAGTACACAG CCGGCGTATCTCCGAAGGTATGTAAAGACGTTACGGTA GAAGGATCCAATGAATTTGCTCCTGTACAGAACCTGAC CGGTAGTTCAGTAGGTCAGAAAGTAACGCTTAAGTGGG ATGCACCTAATGGTACCCCGAATCCGAATCCAAATCCG AATCCGAATCCGGGAACA DA (amino acid 14 AEGSREVKRIGDGLFVTIEPANDVRANEAKVVLAADNVWG sequence) DNTGYQFLLDADHNTFGSVIPATGPLFTGTASSNLYSANF EYLIPANADPVVTTQNIIVTGQGEVVIPGGVYDYCITNPEPA SGKMWIAGDGGNQPARYDDFTFEAGKKYTFTMRRAGMG DGTDMEVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGV AAGNHEYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNL TGSSVGQKVTLKWDAPNGTPNPNPNPNPNPGT KDAΔABM3 15 AATACCGGAGTCAGCTTTGCAAACTATACAGCGCATGG (DNA sequence) ATCTGAGACCGCATGGGCTGATCCACTTCTGACTACTT CTCAACTGAAAGCACTCACTAATAAGGACAAAGCAGAA GGTTCCCGTGAAGTAAAACGGATCGGAGACGGTCTTTT CGTTACGATCGAACCTGCAAACGATGTACGTGCCAACG AAGCCAAGGTTGTGCTTGCGGCAGACAACGTATGGGG AGACAATACGGGTTACCAGTTCTTGTTGGATGCCGATC ACAATACATTCGGAAGTGTCATTCCGGCAACCGGTCCT
1005272698
28 CTCTTTACCGGAACAGCTTCTTCCAATCTTTACAGTGCG AACTTCGAGTATTTGATCCCGGCCAATGCCGATCCTGT TGTTACTACACAGAATATTATCGTTACAGGACAGGGTGA AGTTGTAATCCCCGGTGGTGTTTACGACTATTGCATTAC GAACCCGGAACCTGCATCCGGAAAGATGTGGATCGCA GGAGATGGAGGCAACCAGCCTGCACGTTATGACGATTT CACATTCGAAGCAGGCAAGAAGTACACCTTCACGATGC GTCGCGCCGGAATGGGAGATGGAACTGATATGGAAGT CGAAGACGATTCACCTGCAAGCTATACCTACACGGTGT ATCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCT ACGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCA TGAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTAT CTCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATCC AATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTTCA GTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTAA TGGTACC KDAΔABM3 16 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKAEG (amino acid SREVKRIGDGLFVTIEPANDVRANEAKVVLAADNVWGDNT sequence) GYQFLLDADHNTFGSVIPATGPLFTGTASSNLYSANFEYLI PANADPVVTTQNIIVTGQGEVVIPGGVYDYCITNPEPASGK MWIAGDGGNQPARYDDFTFEAGKKYTFTMRRAGMGDGT DMEVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAG NHEYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGS SVGQKVTLKWDAPNGT KA (DNA 17 AATACCGGAGTCAGCTTTGCAAACTATACAGCGCATGG sequence) ATCTGAGACCGCATGGGCTGATCCACTTCTGACTACTT CTCAACTGAAAGCACTCACTAATAAGGACAAAGAAGTC GAAGACGATTCACCTGCAAGCTATACCTACACGGTGTA TCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCTA CGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCAT GAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTATC TCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATCCA ATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTTCA GTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTAA TGGTACCCCGAATCCGAATCCAAATCCGAATCCG AATCCGGGAACA
1005272698
29 KA (amino acid 18 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKEVE sequence) DDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNHEYC VEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSVGQK VTLKWDAPNGTPNPNPNPNPNPGT RDA 19 AACGGAGGAATCTCGTTGGCCAACTATACGGGCCACG (DNA sequence) GTAGCGAAACAGCTTGGGGTACGTCTCACTTCGGCACC ACTCATGTGAAGCAGCTTACCAACAGCAACCAGGCAGA AGGTTCCCGTGAAGTAAAACGGATCGGAGACGGTCTTT TCGTTACGATCGAACCTGCAAACGATGTACGTGCCAAC GAAGCCAAGGTTGTGCTTGCGGCAGACAACGTATGGG GAGACAATACGGGTTACCAGTTCTTGTTGGATGCCGAT CACAATACATTCGGAAGTGTCATTCCGGCAACCGGTCC TCTCTTTACCGGAACAGCTTCTTCCAATCTTTACAGTGC GAACTTCGAGTATTTGATCCCGGCCAATGCCGATCCTG TTGTTACTACACAGAATATTATCGTTACAGGACAGGGTG AAGTTGTAATCCCCGGTGGTGTTTACGACTATTGCATTA CGAACCCGGAACCTGCATCCGGAAAGATGTGGATCGC AGGAGATGGAGGCAACCAGCCTGCACGTTATGACGATT TCACATTCGAAGCAGGCAAGAAGTACACCTTCACGATG CGTCGCGCCGGAATGGGAGATGGAACTGATATGGAAG TCGAAGACGATTCACCTGCAAGCTATACCTACACGGTG TATCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGC TACGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATC ATGAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTA TCTCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATC CAATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTT CAGTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCT AATGGTACCCCGAATCCGAATCCAAATCCGAATCCGAA TCCGGGAACA RDA 20 NGGISLANYTGHGSETAWGTSHFGTTHVKQLTNSNQAEG (amino acid SREVKRIGDGLFVTIEPANDVRANEAKVVLAADNVWGDNT sequence) GYQFLLDADHNTFGSVIPATGPLFTGTASSNLYSANFEYLI PANADPVVTTQNIIVTGQGEVVIPGGVYDYCITNPEPASGK MWIAGDGGNQPARYDDFTFEAGKKYTFTMRRAGMGDGT
1005272698
33 R (Arg gingipain 37 AACGGAGGAATCTCGTTGGCCAACTATACGGGCCACG active site; DNA GTAGCGAAACAGCTTGGGGTACGTCTCACTTCGGCACC sequence) ACTCATGTGAAGCAGCTTACCAACAGCAACCAG R (Arg gingipain 38 NGGISLANYTGHGSETAWGTSHFGTTHVKQLTNSNQ active site; amino acid sequence) RA 39 NGGISLANYTGHGSETAWGTSHFGTTHVKQLTNSNQEVE Amino acid DDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNHEYC sequence VEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSVGQK VTLKWDAPNGTPNPNPNPNPNPGT AR 58 EVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNH Amino acid EYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSV sequence GQKVTLKWDAPNGTPNPNPNPNPNPGTNGGISLANYTGH GSETAWGTSHFGTTHVKQLTNSNQ AK 59 EVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNH amino acid EYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSV sequence GQKVTLKWDAPNGTPNPNPNPNPNPGTNTGVSFANYTA HGSETAWADPLLTTSQLKALTNKDK KAK 60 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKEVE Amino acid DDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNHEYC sequence VEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSVGQK VTLKWDAPNGTPNPNPNPNPNPGTNTGVSFANYTAHGS ETAWADPLLTTSQLKALTNKDK RAK 61 NGGISLANYTGHGSETAWGTSHFGTTHVKQLTNSNQEVE Amino acid DDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNHEYC sequence VEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSVGQK VTLKWDAPNGTPNPNPNPNPNPGTNTGVSFANYTAHGS ETAWADPLLTTSQLKALTNKDK RAR 62 NGGISLANYTGHGSETAWGTSHFGTTHVKQLTNSNQEVE Amino acid DDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNHEYC sequence VEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSVGQK VTLKWDAPNGTPNPNPNPNPNPGTNGGISLANYTGHGSE TAWGTSHFGTTHVKQLTNSNQ
1005272698
34 KAR 63 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKEVE (amino acid DDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNHEYC sequence) VEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSVGQK VTLKWDAPNGTPNPNPNPNPNPGTNGGISLANYTGHGSE TAWGTSHFGTTHVKQLTNSNQ RA 64 AACGGAGGAATCTCGTTGGCCAACTATACGGGCCACG (DNA sequence) GTAGCGAAACAGCTTGGGGTACGTCTCACTTCGGCACC ACTCATGTGAAGCAGCTTACCAACAGCAACCAGGAAGT CGAAGACGATTCACCTGCAAGCTATACCTACACGGTGT ATCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCT ACGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCA TGAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTAT CTCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATCC AATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTTCA GTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTAA TGGTACCCCGAATCCGAATCCAAATCCGAATCCGAATC CGGGAACA AR 65 GAAGTCGAAGACGATTCACCTGCAAGCTATACCTACAC (DNA sequence) GGTGTATCGTGACGGCACGAAGATCAAGGAAGGTCTG ACAGCTACGACATTCGAAGAAGACGGTGTAGCTGCAGG CAATCATGAGTATTGCGTGGAAGTTAAGTACACAGCCG GCGTATCTCCGAAGGTATGTAAAGACGTTACGGTAGAA GGATCCAATGAATTTGCTCCTGTACAGAACCTGACCGG TAGTTCAGTAGGTCAGAAAGTAACGCTTAAGTGGGATG CACCTAATGGTACCCCGAATCCGAATCCAAATCCGAAT CCGAATCCGGGAACAAACGGAGGAATCTCGTTGGCCA ACTATACGGGCCACGGTAGCGAAACAGCTTGGGGTAC GTCTCACTTCGGCACCACTCATGTGAAGCAGCTTACCA ACAGCAACCAG AK 66 GAAGTCGAAGACGATTCACCTGCAAGCTATACCTACAC (DNA sequence) GGTGTATCGTGACGGCACGAAGATCAAGGAAGGTCTG ACAGCTACGACATTCGAAGAAGACGGTGTAGCTGCAGG CAATCATGAGTATTGCGTGGAAGTTAAGTACACAGCCG GCGTATCTCCGAAGGTATGTAAAGACGTTACGGTAGAA GGATCCAATGAATTTGCTCCTGTACAGAACCTGACCGG TAGTTCAGTAGGTCAGAAAGTAACGCTTAAGTGGGATG
1005272698
35 CACCTAATGGTACCCCGAATCCGAATCCAAATCCGAAT CCGAATCCGGGAACAAATACCGGAGTCAGCTTTGCAAA CTATACAGCGCATGGATCTGAGACCGCATGGGCTGATC CACTTCTGACTACTTCTCAACTGAAAGCACTCACTAATA AGGACAAA KAK 67 AATACCGGAGTCAGCTTTGCAAACTATACAGCGCATGG (DNA sequence) ATCTGAGACCGCATGGGCTGATCCACTTCTGACTACTT CTCAACTGAAAGCACTCACTAATAAGGACAAAGAAGTC GAAGACGATTCACCTGCAAGCTATACCTACACGGTGTA TCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCTA CGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCAT GAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTATC TCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATCCA ATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTTCA GTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTAA TGGTACCCCGAATCCGAATCCAAATCCGAATCCGAATC CGGGAACAAATACCGGAGTCAGCTTTGCAAACTATACA GCGCATGGATCTGAGACCGCATGGGCTGATCCACTTCT GACTACTTCTCAACTGAAAGCACTCACTAATAAGGACAA A RAK 68 AACGGAGGAATCTCGTTGGCCAACTATACGGGCCACG (DNA sequence) GTAGCGAAACAGCTTGGGGTACGTCTCACTTCGGCACC ACTCATGTGAAGCAGCTTACCAACAGCAACCAGGAAGT CGAAGACGATTCACCTGCAAGCTATACCTACACGGTGT ATCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCT ACGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCA TGAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTAT CTCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATCC AATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTTCA GTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTAA TGGTACCCCGAATCCGAATCCAAATCCGAATCCGAATC CGGGAACAAATACCGGAGTCAGCTTTGCAAACTATACA GCGCATGGATCTGAGACCGCATGGGCTGATCCACTTCT GACTACTTCTCAACTGAAAGCACTCACTAATAAGGACAA A
1005272698
36 RAR 69 AACGGAGGAATCTCGTTGGCCAACTATACGGGCCACG (DNA sequence) GTAGCGAAACAGCTTGGGGTACGTCTCACTTCGGCACC ACTCATGTGAAGCAGCTTACCAACAGCAACCAGGAAGT CGAAGACGATTCACCTGCAAGCTATACCTACACGGTGT ATCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCT ACGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCA TGAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTAT CTCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATCC AATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTTCA GTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTAA TGGTACCCCGAATCCGAATCCAAATCCGAATCCGAATC CGGGAACAAACGGAGGAATCTCGTTGGCCAACTATACG GGCCACGGTAGCGAAACAGCTTGGGGTACGTCTCACTT CGGCACCACTCATGTGAAGCAGCTTACCAACAGCAACC AG KAR 70 AATACCGGAGTCAGCTTTGCAAACTATACAGCGCATGG (DNA sequence) ATCTGAGACCGCATGGGCTGATCCACTTCTGACTACTT CTCAACTGAAAGCACTCACTAATAAGGACAAAGAAGTC GAAGACGATTCACCTGCAAGCTATACCTACACGGTGTA TCGTGACGGCACGAAGATCAAGGAAGGTCTGACAGCTA CGACATTCGAAGAAGACGGTGTAGCTGCAGGCAATCAT GAGTATTGCGTGGAAGTTAAGTACACAGCCGGCGTATC TCCGAAGGTATGTAAAGACGTTACGGTAGAAGGATCCA ATGAATTTGCTCCTGTACAGAACCTGACCGGTAGTTCA GTAGGTCAGAAAGTAACGCTTAAGTGGGATGCACCTAA TGGTACCCCGAATCCGAATCCAAATCCGAATCCGAATC CGGGAACAAACGGAGGAATCTCGTTGGCCAACTATACG GGCCACGGTAGCGAAACAGCTTGGGGTACGTCTCACTT CGGCACCACTCATGTGAAGCAGCTTACCAACAGCAACC AG KAR 71 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKMEV (amino acid EDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNHEY sequence with CVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSVGQ Methionine at N KVTLKWDAPNGTPNPNPNPNPNPGTNGGISLANYTGHGS term of A domain) ETAWGTSHFGTTHVKQLTNSNQ
1005272698
37 A 72 EVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNH (amino acid EYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSV sequence) GQKVTLKWDAPNGTPNPNPNPNPNPGT A 73 MEVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGN (amino acid HEYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSS sequence with N VGQKVTLKWDAPNGTPNPNPNPNPNPGT terminal Met) A 74 GAAGTCGAAGACGATTCACCTGCAAGCTATACCTACAC (DNA sequence) GGTGTATCGTGACGGCACGAAGATCAAGGAAGGTCTG ACAGCTACGACATTCGAAGAAGACGGTGTAGCTGCAGG CAATCATGAGTATTGCGTGGAAGTTAAGTACACAGCCG GCGTATCTCCGAAGGTATGTAAAGACGTTACGGTAGAA GGATCCAATGAATTTGCTCCTGTACAGAACCTGACCGG TAGTTCAGTAGGTCAGAAAGTAACGCTTAAGTGGGATG CACCTAATGGTACCCCGAATCCGAATCCAAATCCGAAT CCGAATCCGGGAACA A 75 ATGGAAGTCGAAGACGATTCACCTGCAAGCTATACCTA (alternative DNA CACGGTGTATCGTGACGGCACGAAGATCAAGGAAGGT sequence with 5’ CTGACAGCTACGACATTCGAAGAAGACGGTGTAGCTGC codon encoding AGGCAATCATGAGTATTGCGTGGAAGTTAAGTACACAG methionine) CCGGCGTATCTCCGAAGGTATGTAAAGACGTTACGGTA GAAGGATCCAATGAATTTGCTCCTGTACAGAACCTGAC CGGTAGTTCAGTAGGTCAGAAAGTAACGCTTAAGTGGG ATGCACCTAATGGTACCCCGAATCCGAATCCAAATCCG AATCCGAATCCGGGAACA DUF2436 76 AEGSREVKRIGDGLFVTIEPANDVRANEAKVVLAADNVWG (alternative amino DNTGYQFLLDADHNTFGSVIPATGPLFTGTASSNLYSANF acid sequence EYLIPANADPVVTTQNIIVTGQGEVVIPGGVYDYCITNPEPA absent C terminal SGKMWIAGDGGNQPARYDDFTFEAGKKYTFTMRRAGMG methionine) DGTD AR 79 ATGGAAGTCGAAGACGATTCACCTGCAAGCTATACCTA (DNA sequence; CACGGTGTATCGTGACGGCACGAAGATCAAGGAAGGT 5’ codon encoding CTGACAGCTACGACATTCGAAGAAGACGGTGTAGCTGC methionine] AGGCAATCATGAGTATTGCGTGGAAGTTAAGTACACAG CCGGCGTATCTCCGAAGGTATGTAAAGACGTTACGGTA GAAGGATCCAATGAATTTGCTCCTGTACAGAACCTGAC
1005272698
38 CGGTAGTTCAGTAGGTCAGAAAGTAACGCTTAAGTGGG ATGCACCTAATGGTACCCCGAATCCGAATCCAAATCCG AATCCGAATCCGGGAACAAACGGAGGAATCTCGTTGGC CAACTATACGGGCCACGGTAGCGAAACAGCTTGGGGT ACGTCTCACTTCGGCACCACTCATGTGAAGCAGCTTAC CAACAGCAACCAG AK 80 ATGGAAGTCGAAGACGATTCACCTGCAAGCTATACCTA (DNA sequence; CACGGTGTATCGTGACGGCACGAAGATCAAGGAAGGT 5’ codon encoding CTGACAGCTACGACATTCGAAGAAGACGGTGTAGCTGC methionine] AGGCAATCATGAGTATTGCGTGGAAGTTAAGTACACAG CCGGCGTATCTCCGAAGGTATGTAAAGACGTTACGGTA GAAGGATCCAATGAATTTGCTCCTGTACAGAACCTGAC CGGTAGTTCAGTAGGTCAGAAAGTAACGCTTAAGTGGG ATGCACCTAATGGTACCCCGAATCCGAATCCAAATCCG AATCCGAATCCGGGAACAAATACCGGAGTCAGCTTTGC AAACTATACAGCGCATGGATCTGAGACCGCATGGGCTG ATCCACTTCTGACTACTTCTCAACTGAAAGCACTCACTA ATAAGGACAAA RA 81 NGGISLANYTGHGSETAWGTSHFGTTHVKQLTNSNQMEV Amino acid EDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNHEY sequence (Met at CVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSVGQ N term of A) KVTLKWDAPNGTPNPNPNPNPNPGT AR 82 MEVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGN Amino acid HEYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSS sequence VGQKVTLKWDAPNGTPNPNPNPNPNPGTNGGISLANYTG HGSETAWGTSHFGTTHVKQLTNSNQ [N terminal methionine] KA (amino acid 83 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDKMEV sequence; Met at EDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGNHEY N term of A) CVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSSVGQ KVTLKWDAPNGTPNPNPNPNPNPGT AK 84 MEVEDDSPASYTYTVYRDGTKIKEGLTATTFEEDGVAAGN HEYCVEVKYTAGVSPKVCKDVTVEGSNEFAPVQNLTGSS
1005272698
39
[0119] Table 2: exemplary RNA sequences of the invention Descriptor SEQ ID RNA Sequence No KDcAK1n 40 AAUACCGGAGUCAGCUUUGCAAACUAUACAGCGCAUGGAUCUG AGACCGCAUGGGCUGAUCCACUUCUGACUACUUCUCAACUGAA AGCACUCACUAAUAAGGACAAAUGGGGAGACAAUACGGGUUACC AGUUCUUGUUGGAUGCCGAUCACAAUACAUUCGGAAGUGUCAU UCCGGCAACCGGUCCUCUCUUUACCGGAACAGCUUCUUCCAAU CUUUACAGUGCGAACUUCGAGUAUUUGAUCCCGGCCAAUGCCG AUCCUGUUGUUACUACACAGAAUAUUAUCGUUACAGGACAGGG UGAAGUUGUAAUCCCCGGUGGUGUUUACGACUAUUGCAUUACG AACCCGGAACCUGCAUCCGGAAAGAUGUGGAUCGCAGGAGAUG
1005272698
40 GAGGCAACCAGCCUGCACGUUAUGACGAUUUCACAUUCGAAGC AGGCAAGAAGUACACCUUCACGAUGCGUCGCGCCGGAAUGGGA GAUGGAACUGAUAUGGAAGUCGAAGACGAUUCACCUGCAAGCU AUACCUACACGGUGUAUCGUGACGGCACGAAGAUCAAGGAAGG UCUGACAGCUACGACAUUCGAAGAAGACGGUGUAGCUGCAGGC AAUCAUGAGUAUUGCGUGGAAGUUAAGUACACAGCCGGCGUAU CUCCGAAGGUAUGUAAAGACGUUACGGUAGAAGGAUCCAAUGA AUUUGCUCCUGUACAGAACCUGACCGGUAGUUCAGUAGGUCAG AAAGUAACGCUUAAGUGGGAUGCACCUAAUGGUACCCCGAAUC CGAAUCCAAAUCCGAAUCCGAAUCCGGGAACAACACUUUCCGAA UCAUUCGAAAAUGGUAUUCCGGCAUCUUGGAAGACGAUCGAUG CAGACGGUGACGGGCAUGGCUGGAAACCUGGAAAUGCUCCCGG AAUCGCUGGCUACAAUAGCAAUGGUUGUGUAUAUUCAGAGUCA UUCGGUCUUGGUGGUAUAGGAGUUCUUACCCCUGACAACUAUC UGAUAACACCGGCAUUGGAUUUGCCUAACGGAGGU KDAK 41 AAUACCGGAGUCAGCUUUGCAAACUAUACAGCGCAUGGAUCUG AGACCGCAUGGGCUGAUCCACUUCUGACUACUUCUCAACUGAA AGCACUCACUAAUAAGGACAAAGCAGAAGGUUCCCGUGAAGUAA AACGGAUCGGAGACGGUCUUUUCGUUACGAUCGAACCUGCAAA CGAUGUACGUGCCAACGAAGCCAAGGUUGUGCUUGCGGCAGAC AACGUAUGGGGAGACAAUACGGGUUACCAGUUCUUGUUGGAUG CCGAUCACAAUACAUUCGGAAGUGUCAUUCCGGCAACCGGUCC UCUCUUUACCGGAACAGCUUCUUCCAAUCUUUACAGUGCGAAC UUCGAGUAUUUGAUCCCGGCCAAUGCCGAUCCUGUUGUUACUA CACAGAAUAUUAUCGUUACAGGACAGGGUGAAGUUGUAAUCCC CGGUGGUGUUUACGACUAUUGCAUUACGAACCCGGAACCUGCA UCCGGAAAGAUGUGGAUCGCAGGAGAUGGAGGCAACCAGCCUG CACGUUAUGACGAUUUCACAUUCGAAGCAGGCAAGAAGUACACC UUCACGAUGCGUCGCGCCGGAAUGGGAGAUGGAACUGAUAUGG AAGUCGAAGACGAUUCACCUGCAAGCUAUACCUACACGGUGUA UCGUGACGGCACGAAGAUCAAGGAAGGUCUGACAGCUACGACA UUCGAAGAAGACGGUGUAGCUGCAGGCAAUCAUGAGUAUUGCG UGGAAGUUAAGUACACAGCCGGCGUAUCUCCGAAGGUAUGUAA AGACGUUACGGUAGAAGGAUCCAAUGAAUUUGCUCCUGUACAG AACCUGACCGGUAGUUCAGUAGGUCAGAAAGUAACGCUUAAGU GGGAUGCACCUAAUGGUACCCCGAAUCCGAAUCCAAAUCCGAA UCCGAAUCCGGGAACAAAUACCGGAGUCAGCUUUGCAAACUAUA CAGCGCAUGGAUCUGAGACCGCAUGGGCUGAUCCACUUCUGAC UACUUCUCAACUGAAAGCACUCACUAAUAAGGACAAA KDAK-3S- 42 AAUACCGGAGUCAGCUUUGCAAACUAUACAGCGCAUGGAUCUG AVQP AGACCGCAUGGGCUGAUCCACUUCUGACUACUUCUCAACUGAA AGCACUCACUAAUAAGGACAAAGCAGAAGGUUCCCGUGAAGUAA AACGGAUCGGAGACGGUCUUUUCGUUACGAUCGAACCUGCAAA CGAUGUACGUGCCAACGAAGCCAAGGUUGUGCUUGCGGCAGAC AACGUAUGGGGAGACAAUACGGGUUACCAGUUCUUGUUGGAUG CCGAUCACAAUACAUUCGGAAGUGUCAUUCCGGCAACCGGUCC UCUCUUUACCGGAACAGCUUCUUCCAAUCUUUACAGUGCGAAC UUCGAGUAUUUGAUCCCGGCCAAUGCCGAUCCUGUUGUUACUA CACAGAAUAUUAUCGUUACAGGACAGGGUGAAGUUGUAAUCCC CGGUGGUGUUUACGACUAUAGCAUUACGAACCCGGAACCUGCA UCCGGAAAGAUGUGGAUCGCAGGAGAUGGAGGCAACCAGCCUG CACGUUAUGACGAUUUCACAUUCGAAGCAGGCAAGAAGUACACC
1005272698
41 UUCACGAUGCGUCGCGCCGGAAUGGGAGAUGGAACUGAUAUGG AAGUCGAAGACGAUUCACCUGCAAGCUAUACCUACACGGUGUA UCGUGACGGCACGAAGAUCAAGGAAGGUCUGACAGCUACGACA UUCGAAGAAGACGGUGUAGCUGCAGGCAAUCAUGAGUAUAGCG UGGAAGUUAAGUACACAGCCGGCGUAUCUCCGAAGGUAAGUAA AGACGUUACGGUAGAAGGAUCCAAUGAAUUUGCUGCUGUACAG CCGCUGACCGGUAGUUCAGUAGGUCAGAAAGUAACGCUUAAGU GGGAUGCACCUAAUGGUACCCCGAAUCCGAAUCCAAAUCCGAA UCCGAAUCCGGGAACAAAUACCGGAGUCAGCUUUGCAAACUAUA K 43 KD 44 KDA 45 DA 46
1005272698
42 CAAGGUUGUGCUUGCGGCAGACAACGUAUGGGGAGACAAUACG GGUUACCAGUUCUUGUUGGAUGCCGAUCACAAUACAUUCGGAA GUGUCAUUCCGGCAACCGGUCCUCUCUUUACCGGAACAGCUUC UUCCAAUCUUUACAGUGCGAACUUCGAGUAUUUGAUCCCGGCC AAUGCCGAUCCUGUUGUUACUACACAGAAUAUUAUCGUUACAG GACAGGGUGAAGUUGUAAUCCCCGGUGGUGUUUACGACUAUUG CAUUACGAACCCGGAACCUGCAUCCGGAAAGAUGUGGAUCGCA GGAGAUGGAGGCAACCAGCCUGCACGUUAUGACGAUUUCACAU UCGAAGCAGGCAAGAAGUACACCUUCACGAUGCGUCGCGCCGG AAUGGGAGAUGGAACUGAUAUGGAAGUCGAAGACGAUUCACCU GCAAGCUAUACCUACACGGUGUAUCGUGACGGCACGAAGAUCA AGGAAGGUCUGACAGCUACGACAUUCGAAGAAGACGGUGUAGC UGCAGGCAAUCAUGAGUAUUGCGUGGAAGUUAAGUACACAGCC GGCGUAUCUCCGAAGGUAUGUAAAGACGUUACGGUAGAAGGAU CCAAUGAAUUUGCUCCUGUACAGAACCUGACCGGUAGUUCAGU AGGUCAGAAAGUAACGCUUAAGUGGGAUGCACCUAAUGGUACC CCGAAUCCGAAUCCAAAUCCGAAUCCGAAUCCGGGAACA KDAΔABM3 47 AAUACCGGAGUCAGCUUUGCAAACUAUACAGCGCAUGGAUCUG AGACCGCAUGGGCUGAUCCACUUCUGACUACUUCUCAACUGAA AGCACUCACUAAUAAGGACAAAGCAGAAGGUUCCCGUGAAGUAA AACGGAUCGGAGACGGUCUUUUCGUUACGAUCGAACCUGCAAA CGAUGUACGUGCCAACGAAGCCAAGGUUGUGCUUGCGGCAGAC AACGUAUGGGGAGACAAUACGGGUUACCAGUUCUUGUUGGAUG CCGAUCACAAUACAUUCGGAAGUGUCAUUCCGGCAACCGGUCC UCUCUUUACCGGAACAGCUUCUUCCAAUCUUUACAGUGCGAAC UUCGAGUAUUUGAUCCCGGCCAAUGCCGAUCCUGUUGUUACUA CACAGAAUAUUAUCGUUACAGGACAGGGUGAAGUUGUAAUCCC CGGUGGUGUUUACGACUAUUGCAUUACGAACCCGGAACCUGCA UCCGGAAAGAUGUGGAUCGCAGGAGAUGGAGGCAACCAGCCUG CACGUUAUGACGAUUUCACAUUCGAAGCAGGCAAGAAGUACACC UUCACGAUGCGUCGCGCCGGAAUGGGAGAUGGAACUGAUAUGG AAGUCGAAGACGAUUCACCUGCAAGCUAUACCUACACGGUGUA UCGUGACGGCACGAAGAUCAAGGAAGGUCUGACAGCUACGACA UUCGAAGAAGACGGUGUAGCUGCAGGCAAUCAUGAGUAUUGCG UGGAAGUUAAGUACACAGCCGGCGUAUCUCCGAAGGUAUGUAA AGACGUUACGGUAGAAGGAUCCAAUGAAUUUGCUCCUGUACAG AACCUGACCGGUAGUUCAGUAGGUCAGAAAGUAACGCUUAAGU GGGAUGCACCUAAUGGUACC KA 48 AAUACCGGAGUCAGCUUUGCAAACUAUACAGCGCAUGGAUCUG AGACCGCAUGGGCUGAUCCACUUCUGACUACUUCUCAACUGAA AGCACUCACUAAUAAGGACAAAGAAGUCGAAGACGAUUCACCUG CAAGCUAUACCUACACGGUGUAUCGUGACGGCACGAAGAUCAA GGAAGGUCUGACAGCUACGACAUUCGAAGAAGACGGUGUAGCU GCAGGCAAUCAUGAGUAUUGCGUGGAAGUUAAGUACACAGCCG GCGUAUCUCCGAAGGUAUGUAAAGACGUUACGGUAGAAGGAUC CAAUGAAUUUGCUCCUGUACAGAACCUGACCGGUAGUUCAGUA
1005272698
43 GGUCAGAAAGUAACGCUUAAGUGGGAUGCACCUAAUGGUACCC CGAAUCCGAAUCCAAAUCCGAAUCCGAAUCCGGGAACA RDA 49 AACGGAGGAAUCUCGUUGGCCAACUAUACGGGCCACGGUAGCG AAACAGCUUGGGGUACGUCUCACUUCGGCACCACUCAUGUGAA GCAGCUUACCAACAGCAACCAGGCAGAAGGUUCCCGUGAAGUA AAACGGAUCGGAGACGGUCUUUUCGUUACGAUCGAACCUGCAA ACGAUGUACGUGCCAACGAAGCCAAGGUUGUGCUUGCGGCAGA CAACGUAUGGGGAGACAAUACGGGUUACCAGUUCUUGUUGGAU GCCGAUCACAAUACAUUCGGAAGUGUCAUUCCGGCAACCGGUC CUCUCUUUACCGGAACAGCUUCUUCCAAUCUUUACAGUGCGAA CUUCGAGUAUUUGAUCCCGGCCAAUGCCGAUCCUGUUGUUACU ACACAGAAUAUUAUCGUUACAGGACAGGGUGAAGUUGUAAUCC CCGGUGGUGUUUACGACUAUUGCAUUACGAACCCGGAACCUGC AUCCGGAAAGAUGUGGAUCGCAGGAGAUGGAGGCAACCAGCCU GCACGUUAUGACGAUUUCACAUUCGAAGCAGGCAAGAAGUACA CCUUCACGAUGCGUCGCGCCGGAAUGGGAGAUGGAACUGAUAU GGAAGUCGAAGACGAUUCACCUGCAAGCUAUACCUACACGGUG UAUCGUGACGGCACGAAGAUCAAGGAAGGUCUGACAGCUACGA CAUUCGAAGAAGACGGUGUAGCUGCAGGCAAUCAUGAGUAUUG CGUGGAAGUUAAGUACACAGCCGGCGUAUCUCCGAAGGUAUGU AAAGACGUUACGGUAGAAGGAUCCAAUGAAUUUGCUCCUGUAC AGAACCUGACCGGUAGUUCAGUAGGUCAGAAAGUAACGCUUAA GUGGGAUGCACCUAAUGGUACCCCGAAUCCGAAUCCAAAUCCG AAUCCGAAUCCGGGAACA R 50 AACGGAGGAAUCUCGUUGGCCAACUAUACGGGCCACGGUAGCG AAACAGCUUGGGGUACGUCUCACUUCGGCACCACUCAUGUGAA GCAGCUUACCAACAGCAACCAG DUF2436 51 GCAGAAGGUUCCCGUGAAGUAAAACGGAUCGGAGACGGUCUUU UCGUUACGAUCGAACCUGCAAACGAUGUACGUGCCAACGAAGC CAAGGUUGUGCUUGCGGCAGACAACGUAUGGGGAGACAAUACG GGUUACCAGUUCUUGUUGGAUGCCGAUCACAAUACAUUCGGAA GUGUCAUUCCGGCAACCGGUCCUCUCUUUACCGGAACAGCUUC UUCCAAUCUUUACAGUGCGAACUUCGAGUAUUUGAUCCCGGCC AAUGCCGAUCCUGUUGUUACUACACAGAAUAUUAUCGUUACAG GACAGGGUGAAGUUGUAAUCCCCGGUGGUGUUUACGACUAUUG CAUUACGAACCCGGAACCUGCAUCCGGAAAGAUGUGGAUCGCA GGAGAUGGAGGCAACCAGCCUGCACGUUAUGACGAUUUCACAU UCGAAGCAGGCAAGAAGUACACCUUCACGAUGCGUCGCGCCGG AAUGGGAGAUGGAACUGAUAUG ABM1 52 UCCAAUGAAUUUGCUCCUGUACAGAACCUGACCGGUAGUUCAG UAGGUCAGAAAGUAACGCUUAAGUGGGAUGCACCUAAUGGUAC C ABM2 53 GCAAGCUAUACCUACACGGUGUAUCGUGACGGCACGAAGAUCAA GGAAGGUCUGACAGCUACGACAUUCGAAGAAGACGGUGUAGCU GCAGGCAAUCAUGAGUAUUGCGUGGAAGUUAAGUACACAGCCG GCGUAUCUCCGAAGGUAUGUAAAGACGUUACGGUAGAAGGA
1005272698
44 ABM3 54 CCGAAUCCGAAUCCAAAUCCGAAUCCGAAUCCGGGAACA ABM 2+1 55 GCAAGCUAUACCUACACGGUGUAUCGUGACGGCACGAAGAUCA AGGAAGGUCUGACAGCUACGACAUUCGAAGAAGACGGUGUAGC UGCAGGCAAUCAUGAGUAUUGCGUGGAAGUUAAGUACACAGCC GGCGUAUCUCCGAAGGUAUGUAAAGACGUUACGGUAGAAGGAU CCAAUGAAUUUGCUCCUGUACAGAACCUGACCGGUAGUUCAGU AGGUCAGAAAGUAACGCUUAAGUGGGAUGCACCUAAUGGUACC ABM 2+1+3 56 GCAAGCUAUACCUACACGGUGUAUCGUGACGGCACGAAGAUCA AGGAAGGUCUGACAGCUACGACAUUCGAAGAAGACGGUGUAGC UGCAGGCAAUCAUGAGUAUUGCGUGGAAGUUAAGUACACAGCC GGCGUAUCUCCGAAGGUAUGUAAAGACGUUACGGUAGAAGGAU CCAAUGAAUUUGCUCCUGUACAGAACCUGACCGGUAGUUCAGU AGGUCAGAAAGUAACGCUUAAGUGGGAUGCACCUAAUGGUACC CCGAAUCCGAAUCCAAAUCCGAAUCCGAAUCCGGGAACA RA 57 AACGGAGGAAUCUCGUUGGCCAACUAUACGGGCCACGGUAGCG AAACAGCUUGGGGUACGUCUCACUUCGGCACCACUCAUGUGAA GCAGCUUACCAACAGCAACCAGGAAGUCGAAGACGAUUCACCU GCAAGCUAUACCUACACGGUGUAUCGUGACGGCACGAAGAUCA AGGAAGGUCUGACAGCUACGACAUUCGAAGAAGACGGUGUAGC UGCAGGCAAUCAUGAGUAUUGCGUGGAAGUUAAGUACACAGCC GGCGUAUCUCCGAAGGUAUGUAAAGACGUUACGGUAGAAGGAU CCAAUGAAUUUGCUCCUGUACAGAACCUGACCGGUAGUUCAGU AGGUCAGAAAGUAACGCUUAAGUGGGAUGCACCUAAUGGUACC CCGAAUCCGAAUCCAAAUCCGAAUCCGAAUCCGGGAACA A 77 GAAGUCGAAGACGAUUCACCUGCAAGCUAUACCUACACGGUGU AUCGUGACGGCACGAAGAUCAAGGAAGGUCUGACAGCUACGAC AUUCGAAGAAGACGGUGUAGCUGCAGGCAAUCAUGAGUAUUGC GUGGAAGUUAAGUACACAGCCGGCGUAUCUCCGAAGGUAUGUA AAGACGUUACGGUAGAAGGAUCCAAUGAAUUUGCUCCUGUACA GAACCUGACCGGUAGUUCAGUAGGUCAGAAAGUAACGCUUAAG UGGGAUGCACCUAAUGGUACCCCGAAUCCGAAUCCAAAUCCGA AUCCGAAUCCGGGAACA A (with 5’ 78 [AUG]GAAGUCGAAGACGAUUCACCUGCAAGCUAUACCUACACGG AUG UGUAUCGUGACGGCACGAAGAUCAAGGAAGGUCUGACAGCUAC codon) GACAUUCGAAGAAGACGGUGUAGCUGCAGGCAAUCAUGAGUAU UGCGUGGAAGUUAAGUACACAGCCGGCGUAUCUCCGAAGGUAU GUAAAGACGUUACGGUAGAAGGAUCCAAUGAAUUUGCUCCUGU ACAGAACCUGACCGGUAGUUCAGUAGGUCAGAAAGUAACGCUU AAGUGGGAUGCACCUAAUGGUACCCCGAAUCCGAAUCCAAAUC CGAAUCCGAAUCCGGGAACA Detailed description of the embodiments [0120] It will be understood that the invention disclosed and defined in this specification extends to all alternative combinations of two or more of the individual features mentioned or evident from the text or drawings. All of these different combinations constitute various alternative aspects of the invention.
1005272698
45 [0121] Reference will now be made in detail to certain embodiments of the invention. While the invention will be described in conjunction with the embodiments, it will be understood that the intention is not to limit the invention to those embodiments. On the contrary, the invention is intended to cover all alternatives, modifications, and equivalents, which may be included within the scope of the present invention as defined by the claims. [0122] One skilled in the art will recognize many methods and materials similar or equivalent to those described herein, which could be used in the practice of the present invention. The present invention is in no way limited to the methods and materials described. It will be understood that the invention disclosed and defined in this specification extends to all alternative combinations of two or more of the individual features mentioned or evident from the text or drawings. All of these different combinations constitute various alternative aspects of the invention. [0123] All of the patents and publications referred to herein are incorporated by reference in their entirety. [0124] For purposes of interpreting this specification, terms used in the singular will also include the plural and vice versa. [0125] In work leading to the present invention, the inventors investigated various chimeric or fusion proteins for use in inducing immune responses to P. gingivalis and methods for large-scale production of such chimeras for use as vaccine candidates. [0126] One such candidate fusion protein (termed KDcAK1n as further described herein), was found to elicit a robust immune response to P. gingivalis, but suffered from large-scale manufacturing and production issues. [0127] The inventors initially considered whether the manufacturing issues for this protein could be overcome when the protein was provided to subjects when encoded by an RNA vaccine. Surprisingly, the inventors found that an RNA vaccine encoding KDcAK1n did not express well and therefore KDcAK1n was not a preferred candidate for use in an RNA vaccine. [0128] Surprisingly, the inventors found that robust immune responses were obtained when providing an RNA encoding alternative chimeric proteins containing various domains derived from P. gingivalis Arg- or Lys-gingipain proteins.
1005272698
46 [0129] The present invention is therefore concerned with the design of new RNA vaccines encoding protein antigens, or chimeric or fusion proteins comprising protein antigens, for use in inducing an immune response to P. gingivalis, and methods and uses comprising the same. Gingipains [0130] The pathogenicity of P. gingivalis is attributed to a number of surface-associated virulence factors that include cysteine proteinases (gingipains), fimbriae, haem-binding proteins, and outer membrane transport proteins amongst others. In particular, the extracellular Arg- and Lys-specific proteinases ‘gingipains’ (RgpA/B and Kgp) of P. gingivalis have been implicated as major virulence factors that are critical for colonisation, penetration into host tissue, dysregulation of the immune response, dysbiosis and disease. [0131] The gingipains, in particular the Lys-specific proteinase Kgp are essential for P. gingivalis to induce alveolar bone resorption in the mouse periodontitis model. The gingipains have also been found in gingival tissue at sites of severe periodontitis at high concentrations proximal to the subgingival plaque and at lower concentrations at distal sites deeper into the gingival tissue. Lys-specific and Arg-specific proteinases have been shown to degrade a variety of host proteins in vitro, e.g., fibrinogen, fibronectin, and laminin. Plasma host defence and regulatory proteinase inhibitors α-trypsin, α2- macroglobulin, anti-chymotrypsin, antithrombin III and antiplasmin are also degraded by Lys- and Arg- proteinases from P. gingivalis. This has led to the development of a cogent mechanism to explain the keystone role played by P. gingivalis in the development of chronic periodontitis. [0132] The RgpA, RgpB and Kgp genes all encode an N-terminal signal peptide of ∼22 amino acids in length, an unusually long propeptide of ∼200 amino acids in length, and a catalytic domain of ∼480 amino acids. C-terminal to the catalytic domain is a large hemagglutinin-adhesin (HA) domain which is comprised of adhesin binding domains (ABMs, of which 5 distinct sequences have been described), a “Domain of Unknown Function” (termed DUF2436 which is defined as conserved Pfam Domain of Unknown Function; IPR018832) and C-terminal adhesin domains or cleaved adhesin domains (or CADs). The particular arrangement of the ABMs, DUF and CADs varies between naturally occurring Kgp and RgpA/B.
1005272698
47 [0133] The architecture of the domains in the Kgp polyprotein is illustrated in Figure 1. For example, Kgp comprises (N terminus to C terminus): a catalytic domain, a first ABM (ABM1), DUF2436, a domain comprising ABM2, ABM1, ABM3, two CAD domains (termed K1 and K2), a further domain comprising ABM2 and ABM1, a further CAD domain (termed K3), ABM2, and a C-terminal domain. [0134] As used herein, reference to ABMs 1, 2 and 3 will be understood to generally refer to the ABMs found in the order ABM2, ABM1 and ABM3 in the sequence immediately C terminal to DUF2436 of Kgp, as depicted in Figure 1. [0135] The catalytic domains of RgpB and RgpA share a high-degree of sequence homology. However, RgpB lacks the HA domains and is located in a monomeric form on the outer membrane. Some of the HA domains have been alternatively described as C- terminal adhesin domains or cleaved adhesin domains (CADs) and some are DUF (“Domain of Unknown Function”) 2436 domains (conserved Pfam Domain of Unknown Function; IPR018832). [0136] The RgpA and Kgp precursor proteins are cleaved into multiple domains that remain non-covalently associated forming large outer membrane protein complexes. In vivo, Arg- and Lys-specific proteinases are therefore found in a cell-associated complex of non-covalently associated proteinases and adhesins. One such complex has been designated the RgpA-Kgp proteinase-adhesin complex (previously referred to as the PrtR-PrtK proteinase-adhesin complex). The complex is composed of a 45kDa Arg- specific calcium-stabilised cysteine proteinase and seven sequence-related adhesin domains, [0137] As used herein a Lys-gingipain catalytic domain may also be referred to as a KAS domain or PAS domain. As used herein an Arg-gingipain catalytic domain may also be referred to as a RAS domain or PAS domain. Typically, the catalytic domain of the Lys-gingipain or Arg-gingipains is located in the N-terminal ~480 amino acid region of the protein. The active site within the catalytic domain is typically located at amino acid residues 426-446 (for RgpA) and 432-453 (for Kgp). [0138] As used herein an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis will be understood to typically refer to the region of an Arg- or Lys-gingipain that is C- terminal to the catalytic or active site domain. The adhesin domain (also referred to as
1005272698
48 the HA domain) typically comprise a Domain of Unknown Function (DUF) domain (especially DUF 2436 conserved Pfam Domain of Unknown Function; IPR018832) and several adhesin binding motifs (ABM) domains and a cleaved adhesin domain (CAD). Nucleic acids [0139] The term “nucleic acid,” in its broadest sense, includes any compound and/or substance that comprise a polymer of nucleotides. These polymers are often referred to as polynucleotides. Typically, the polynucleotides of the invention are in the form of an RNA molecule, preferably an mRNA. As used herein, the term “messenger RNA” (RNA) refers to any polynucleotide which encodes a polypeptide of interest and which is capable of being translated to produce the encoded polypeptide of interest in vitro, in vivo, in situ or ex vivo. The skilled artisan will appreciate that, except where otherwise noted, polynucleotide sequences set forth in the instant application will recite “T”s in a representative DNA sequence but where the sequence represents RNA (e.g., RNA), the “T”s would be substituted for “U”s. Thus, any of the RNA polynucleotides encoded by a DNA identified by a particular sequence identification number may also comprise the corresponding RNA (e.g., RNA) sequence encoded by the DNA, where each “T” of the DNA sequence is substituted with “U.” [0140] The basic components of an RNA molecule include at least a coding region, a 5′UTR, a 3′UTR, a 5′ cap and a poly-A tail. Polynucleotides of the present disclosure may function as RNA but can be distinguished from wild-type RNA in their functional and/or structural design features, which serve to overcome existing problems of effective polypeptide expression using nucleic-acid based therapeutics. [0141] A “5′ untranslated region” (5′UTR) refers to a region of an RNA that is directly upstream (i.e., 5′) from the start codon (i.e., the first codon of an RNA transcript translated by a ribosome) that does not encode a polypeptide. [0142] A “3′ untranslated region” (3′UTR) refers to a region of an RNA that is directly downstream (i.e., 3′) from the stop codon (i.e., the codon of an RNA transcript that signals a termination of translation) that does not encode a polypeptide. [0143] An “open reading frame” is a continuous stretch of DNA beginning with a start codon (e.g., methionine (ATG)), and ending with a stop codon (e.g., TAA, TAG or TGA) and encodes a polypeptide.
1005272698
49 [0144] A “polyA tail” is a region of RNA (typically mRNA) that is downstream, e.g., directly downstream (i.e., 3′), from the 3′ UTR that contains multiple, sometimes consecutive adenosine monophosphates. A polyA tail may contain 10 to 300 adenosine monophosphates. For example, a polyA tail may contain 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, 110, 120, 130, 140, 150, 160, 170, 180, 190, 200, 210, 220, 230, 240, 250, 260, 270, 280, 290 or 300 adenosine monophosphates. In some embodiments, a polyA tail contains 50 to 250 adenosine monophosphates. In some embodiments, a segmented polyA tail may be used (typically segments of consecutive adenosine monophosphates separated via a short spacer region between segments). In a relevant biological setting (e.g., in cells, in vivo) the poly(A) tail functions to protect mRNA from enzymatic degradation, e.g., in the cytoplasm, and aids in transcription termination, export of the mRNA from the nucleus and translation. [0145] In some embodiments, a polynucleotide includes 200 to 3,000 nucleotides. For example, a polynucleotide may include 200 to 500, 200 to 1000, 200 to 1500, 200 to 3000, 500 to 1000, 500 to 1500, 500 to 2000, 500 to 3000, 1000 to 1500, 1000 to 2000, 1000 to 3000, 1500 to 3000, or 2000 to 3000 nucleotides. [0146] The present invention also contemplates the use of one or more structural and/or chemical modifications or alterations which impart useful properties to the polynucleotide including, in some embodiments, the lack of a substantial induction of the innate immune response of a cell into which the polynucleotide is introduced. As such, modified RNA molecules of the present invention may also be termed “mmRNA.” As used herein, a “structural” feature or modification is one in which two or more linked nucleotides are inserted, deleted, duplicated, inverted or randomized in a polynucleotide, primary construct or mRNA without significant chemical modification to the nucleotides themselves. Because chemical bonds will necessarily be broken and reformed to effect a structural modification, structural modifications are of a chemical nature and hence are chemical modifications. However, structural modifications will result in a different sequence of nucleotides. For example, the polynucleotide “ATCG” may be chemically modified to “AT-5meC-G”. The same polynucleotide may be structurally modified from “ATCG” to “ATCCCG”. Here, the dinucleotide “CC” has been inserted, resulting in a structural modification to the polynucleotide. [0147] The RNA molecules of the invention may also comprise an 5’ terminal cap. In some embodiments, the 5′ terminal cap is 7mG(5′)ppp(5′)NlmpNp although it will be
1005272698
50 appreciated that any number of different 5’ terminal caps commonly used in the art may be employed. [0148] In some embodiments, the RNA molecule comprises at least one chemical modification. The terms “chemical modification” and “chemically modified” refer to modification with respect to adenosine (A), guanosine (G), uridine (U), thymidine (T) or cytidine (C) ribonucleosides or deoxyribnucleosides in at least one of their position, pattern, percent or population. Generally, these terms do not refer to the ribonucleotide modifications in naturally occurring 5′-terminal RNA cap moieties. With respect to a polypeptide, the term “modification” refers to a modification relative to the canonical set 20 amino acids. Polypeptides, as provided herein, are also considered “modified” of they contain amino acid substitutions, insertions or a combination of substitutions and insertions. [0149] Polynucleotides (e.g., RNA polynucleotides, such as mRNA polynucleotides), in some embodiments, comprise various (more than one) different modifications. In some embodiments, a particular region of a polynucleotide contains one, two or more (optionally different) nucleoside or nucleotide modifications. In some embodiments, a modified RNA polynucleotide (e.g., a modified RNA polynucleotide), introduced to a cell or organism, exhibits reduced degradation in the cell or organism, respectively, relative to an unmodified polynucleotide. In some embodiments, a modified RNA polynucleotide (e.g., a modified RNA polynucleotide), introduced into a cell or organism, may exhibit reduced immunogenicity in the cell or organism, respectively (e.g., a reduced innate response). [0150] Modifications of polynucleotides include, without limitation, those described herein. Polynucleotides (e.g., RNA polynucleotides, such as mRNA polynucleotides) may comprise modifications that are naturally-occurring, non-naturally-occurring or the polynucleotide may comprise a combination of naturally-occurring and non-naturally- occurring modifications. Polynucleotides may include any useful modification, for example, of a sugar, a nucleobase, or an internucleoside linkage (e.g., to a linking phosphate, to a phosphodiester linkage or to the phosphodiester backbone). [0151] Polynucleotides (e.g., RNA polynucleotides, such as mRNA polynucleotides), in some embodiments, comprise non-natural modified nucleotides that are introduced during synthesis or post-synthesis of the polynucleotides to achieve desired functions or properties. The modifications may be present on internucleotide linkages, purine or
1005272698
51 pyrimidine bases, or sugars. The modification may be introduced with chemical synthesis or with a polymerase enzyme at the terminal of a chain or anywhere else in the chain. Any of the regions of a polynucleotide may be chemically modified. [0152] The present disclosure provides for modified nucleosides and nucleotides of a polynucleotide (e.g., RNA polynucleotides, such as mRNA polynucleotides). A “nucleoside” refers to a compound containing a sugar molecule (e.g., a pentose or ribose) or a derivative thereof in combination with an organic base (e.g., a purine or pyrimidine) or a derivative thereof (also referred to herein as “nucleobase”). A nucleotide” refers to a nucleoside, including a phosphate group. Modified nucleotides may by synthesized by any useful method, such as, for example, chemically, enzymatically, or recombinantly, to include one or more modified or non-natural nucleosides. Polynucleotides may comprise a region or regions of linked nucleosides. Such regions may have variable backbone linkages. The linkages may be standard phosphdioester linkages, in which case the polynucleotides would comprise regions of nucleotides. [0153] Modified nucleotide base pairing encompasses not only the standard adenosine-thymine, adenosine-uracil, or guanosine-cytosine base pairs, but also base pairs formed between nucleotides and/or modified nucleotides comprising non-standard or modified bases, wherein the arrangement of hydrogen bond donors and hydrogen bond acceptors permits hydrogen bonding between a non-standard base and a standard base or between two complementary non-standard base structures. One example of such non-standard base pairing is the base pairing between the modified nucleotide inosine and adenine, cytosine or uracil. Any combination of base/sugar or linker may be incorporated into polynucleotides of the present disclosure. [0154] The at least one chemical modification may be selected from pseudouridine, N1-methylpseudouridine, N1-ethylpseudouridine, 2-thiouridine, 4′-thiouridine, 5- methylcytosine, 5-methyluridine, 2-thio-1-methyl-1-deaza-pseudouridine, 2-thio-1- methyl-pseudouridine, 2-thio-5-aza-uridine, 2-thio-dihydropseudouridine, 2-thio- dihydrouridine, 2-thio-pseudouridine, 4-methoxy-2-thio-pseudouridine, 4-methoxy- pseudouridine, 4-thio-1-methyl-pseudouridine, 4-thio-pseudouridine, 5-aza-uridine, dihydropseudouridine, 5-methoxyuridine and 2′-O-methyl uridine. In some embodiments, the chemical modification is in the 5-position of the uracil. In some embodiments, the chemical modification is a N1-methylpseudouridine. In some embodiments, the chemical modification is a N1-ethylpseudouridine. In some embodiments, polynucleotides include
1005272698
52 a combination of at least two (e.g., 2, 3, 4 or more) of the aforementioned modified nucleobases. [0155] In some embodiments, polynucleotides (e.g., RNA polynucleotides, such as mRNA polynucleotides) are uniformly modified (e.g., fully modified, modified throughout the entire sequence) for a particular modification. For example, a polynucleotide can be uniformly modified with 5-methyl-cytidine (m5C), meaning that all cytosine residues in the RNA sequence are replaced with 5-methyl-cytidine (m5C). Similarly, a polynucleotide can be uniformly modified for any type of nucleoside residue present in the sequence by replacement with a modified residue such as those set forth above. [0156] Exemplary nucleobases and nucleosides having a modified cytosine include N4-acetyl-cytidine (ac4C), 5-methyl-cytidine (m5C), 5-halo-cytidine (e.g., 5-iodo- cytidine), 5-hydroxymethyl-cytidine (hm5C), 1-methyl-pseudoisocytidine, 2-thio-cytidine (s2C), and 2-thio-5-methyl-cytidine. [0157] In some embodiments, a modified nucleobase is a modified uridine. Exemplary nucleobases and In some embodiments, a modified nucleobase is a modified cytosine. nucleosides having a modified uridine include 5-cyano uridine, and 4′-thio uridine. [0158] In some embodiments, a modified nucleobase is a modified adenine. Exemplary nucleobases and nucleosides having a modified adenine include 7-deaza-adenine, 1- methyl-adenosine (m1A), 2-methyl-adenine (m2A), and N6-methyl-adenosine (m6A). [0159] In some embodiments, a modified nucleobase is a modified guanine. Exemplary nucleobases and nucleosides having a modified guanine include inosine (I), 1-methyl- inosine (m1I), wyosine (imG), methylwyosine (mimG), 7-deaza-guanosine, 7-cyano-7- deaza-guanosine (preQO), 7-aminomethyl-7-deaza-guanosine (preQ1), 7-methyl- guanosine (m7G), 1-methyl-guanosine (mlG), 8-oxo-guanosine, 7-methyl-8-oxo- guanosine. [0160] The polynucleotides of the present disclosure may be partially or fully modified along the entire length of the molecule. For example, one or more or all or a given type of nucleotide (e.g., purine or pyrimidine, or any one or more or all of A, G, U, C) may be uniformly modified in a polynucleotide of the disclosure, or in a given predetermined sequence region thereof (e.g., in the mRNA including or excluding the polyA tail). In some embodiments, all nucleotides X in a polynucleotide of the present disclosure (or in a given
1005272698
53 sequence region thereof) are modified nucleotides, wherein X may any one of nucleotides A, G, U, C, or any one of the combinations A+G, A+U, A+C, G+U, G+C, U+C, A+G+U, A+G+C, G+U+C or A+G+C. [0161] The polynucleotide may contain from about 1% to about 100% modified nucleotides (either in relation to overall nucleotide content, or in relation to one or more types of nucleotide, i.e., any one or more of A, G, U or C) or any intervening percentage (e.g., from 1% to 20%, from 1% to 25%, from 1% to 50%, from 1% to 60%, from 1% to 70%, from 1% to 80%, from 1% to 90%, from 1% to 95%, from 10% to 20%, from 10% to 25%, from 10% to 50%, from 10% to 60%, from 10% to 70%, from 10% to 80%, from 10% to 90%, from 10% to 95%, from 10% to 100%, from 20% to 25%, from 20% to 50%, from 20% to 60%, from 20% to 70%, from 20% to 80%, from 20% to 90%, from 20% to 95%, from 20% to 100%, from 50% to 60%, from 50% to 70%, from 50% to 80%, from 50% to 90%, from 50% to 95%, from 50% to 100%, from 70% to 80%, from 70% to 90%, from 70% to 95%, from 70% to 100%, from 80% to 90%, from 80% to 95%, from 80% to 100%, from 90% to 95%, from 90% to 100%, and from 95% to 100%). Any remaining percentage is accounted for by the presence of unmodified A, G, U, or C. [0162] The polynucleotides may contain at a minimum 1% and at maximum 100% modified nucleotides, or any intervening percentage, such as at least 5% modified nucleotides, at least 10% modified nucleotides, at least 25% modified nucleotides, at least 50% modified nucleotides, at least 80% modified nucleotides, or at least 90% modified nucleotides. For example, the polynucleotides may contain a modified pyrimidine such as a modified uracil or cytosine. In some embodiments, at least 5%, at least 10%, at least 25%, at least 50%, at least 80%, at least 90% or 100% of the uracil in the polynucleotide is replaced with a modified uracil (e.g., a 5-substituted uracil). The modified uracil can be replaced by a compound having a single unique structure, or can be replaced by a plurality of compounds having different structures (e.g., 2, 3, 4 or more unique structures). n some embodiments, at least 5%, at least 10%, at least 25%, at least 50%, at least 80%, at least 90% or 100% of the cytosine in the polynucleotide is replaced with a modified cytosine (e.g., a 5-substituted cytosine). The modified cytosine can be replaced by a compound having a single unique structure, or can be replaced by a plurality of compounds having different structures (e.g., 2, 3, 4 or more unique structures). [0163] In some embodiments, the modified nucleobase is a modified uracil. Exemplary nucleobases and nucleosides having a modified uracil include pseudouridine (ψ), pyridin-
1005272698
54 4-one ribonucleoside, 5-aza-uridine, 6-aza-uridine, 2-thio-5-aza-uridine, 2-thio-uridine, 4- thio-uridine, 4-thio-pseudouridine, 2-thio-pseudouridine, 5-hydroxy-uridine, 5-aminoallyl- uridine, 5-halo-uridine (e.g., 5-iodo-uridineor 5-bromo-uridine), 3-methyl-uridine, 5- methoxy-uridine, uridine 5-oxyacetic acid, uridine 5-oxyacetic acid methyl ester, 5- carboxymethyl-uridine, 1-carboxymethyl-pseudouridine, 5-carboxyhydroxymethyl- uridine, 5-carboxyhydroxymethyl-uridine methyl ester, 5-methoxycarbonylmethyl-uridine, 5-methoxycarbonylmethyl-2-thio-uridine, 5-aminomethyl-2-thio-uridine, 5- methylaminomethyl-uridine, 5-methylaminomethyl-2-thio-uridine, 5-methylaminomethyl- 2-seleno-uridine, 5-carbamoylmethyl-uridine, 5-carboxymethylaminomethyl-uridine, 5- carboxymethylaminomethyl-2-thio-uridine, 5-propynyl-uridine, 1-propynyl-pseudouridine, 5-taurinomethyl-uridine, 1-taurinomethyl-pseudouridine, 5-taurinomethyl-2-thio-uridine, 1-taurinomethyl-4-thio-pseudouridine, 5-methyl-uridine (i.e., having the nucleobase deoxythymine), 1-methyl-pseudouridine, 5-methyl-2-thio-uridine, 1-methyl-4-thio- pseudouridine, 4-thio-1-methyl-pseudouridine, 3-methyl-pseudouridine, 2-thio-1-methyl- pseudouridine, 1-methyl-1-deaza-pseudouridine, 2-thio-1-methyl-1-deaza- pseudouridine, dihydrouridine (D), dihydropseudouridine, 5,6-dihydrouridine, 5-methyl- dihydrouridine, 2-thio-dihydrouridine, 2-thio-dihydropseudouridine, 2-methoxy-uridine, 2- methoxy-4-thio-uridine, 4-methoxy-pseudouridine, 4-methoxy-2-thio-pseudouridine, N1- methyl-pseudouridine, 3-(3-amino-3-carboxypropyl)uridine, 1-methyl-3-(3-amino-3- carboxypropyl)pseudouridine, 5-(isopentenylaminomethyl)uridine, 5- (isopentenylaminomethyl)-2-thio-uridine, α-thio-uridine, 2′-O-methyl-uridine, 5,2′-O- dimethyl-uridine, 2′-O-methyl-pseudouridine (Wm), 2-thio-2′-O-methyl-uridine, 5- methoxycarbonylmethyl-2′-O-methyl-uridine, 5-carbamoylmethyl-2′-O-methyl-uridine, 5- carboxymethylaminomethyl-2′-O-methyl-uridine, 3,2′-O-dimethyl-uridine, and 5- (isopentenylaminomethyl)-2′-O-methyl-uridine, 1-thio-uridine, deoxythymidine, 2′-F-ara- uridine, 2′-F-uridine, 2′-OH-ara-uridine, 5-(2-carbomethoxyvinyl) uridine, and 5-[3-(1-E- propenylamino)]uridine. [0164] In some embodiments, the modified nucleobase is a modified cytosine. Exemplary nucleobases and nucleosides having a modified cytosine include 5-aza- cytidine, 6-aza-cytidine, pseudoisocytidine, 3-methyl-cytidine, N4-acetyl-cytidine, 5- formyl-cytidine, N4-methyl-cytidine, 5-methyl-cytidine, 5-halo-cytidine (e.g., 5-iodo- cytidine), 5-hydroxymethyl-cytidine, 1-methyl-pseudoisocytidine, pyrrolo-cytidine, pyrrolo-pseudoisocytidine, 2-thio-cytidine, 2-thio-5-methyl-cytidine, 4-thio- pseudoisocytidine, 4-thio-1-methyl-pseudoisocytidine, 4-thio-1-methyl-1-deaza-
1005272698
55 pseudoisocytidine, 1-methyl-1-deaza-pseudoisocytidine, zebularine, 5-aza-zebularine, 5- methyl-zebularine, 5-aza-2-thio-zebularine, 2-thio-zebularine, 2-methoxy-cytidine, 2- methoxy-5-methyl-cytidine, 4-methoxy-pseudoisocytidine, 4-methoxy-1-methyl- pseudoisocytidine, lysidine, α-thio-cytidine, 2′-O-methyl-cytidine, 5,2′-O-dimethyl- cytidine, N4-acetyl-2′-O-methyl-cytidine, N4,2′-O-dimethyl-cytidine, 5-formyl-2′-O-methyl- cytidine, N4,N4,2′-O-trimethyl-cytidine, 1-thio-cytidine, 2′-F-ara-cytidine, 2′-F-cytidine, and 2′-OH-ara-cytidine. [0165] In some embodiments, the modified nucleobase is a modified adenine. Exemplary nucleobases and nucleosides having a modified adenine include 2-amino- purine, 2, 6-diaminopurine, 2-amino-6-halo-purine (e.g., 2-amino-6-chloro-purine), 6- halo-purine (e.g., 6-chloro-purine), 2-amino-6-methyl-purine, 8-azido-adenosine, 7- deaza-adenine, 7-deaza-8-aza-adenine, 7-deaza-2-amino-purine, 7-deaza-8-aza-2- amino-purine, 7-deaza-2,6-diaminopurine, 7-deaza-8-aza-2,6-diaminopurine, 1-methyl- adenosine, 2-methyl-adenine, N6-methyl-adenosine, 2-methylthio-N6-methyl-adenosine, N6-isopentenyl-adenosine, 2-methylthio-N6-isopentenyl-adenosine, N6-(cis- hydroxyisopentenyl)adenosine, 2-methylthio-N6-(cis-hydroxyisopentenyl)adenosine, N6- glycinylcarbamoyl-adenosine, N6-threonylcarbamoyl-adenosine, N6-methyl-N6- threonylcarbamoyl-adenosine, 2-methylthio-N6-threonylcarbamoyl-adenosine, N6,N6- dimethyl-adenosine, N6-hydroxynorvalylcarbamoyl-adenosine, 2-methylthio-N6- hydroxynorvalylcarbamoyl-adenosine, N6-acetyl-adenosine, 7-methyl-adenine, 2- methylthio-adenine, 2-methoxy-adenine, α-thio-adenosine, 2′-O-methyl-adenosine, N6,2′-O-dimethyl-adenosine, N6,N6,2′-O-trimethyl-adenosine, 1,2′-O-dimethyl- adenosine, 2′-O-ribosyladenosine (phosphate) (Ar(p)), 2-amino-N6-methyl-purine, 1-thio- adenosine, 8-azido-adenosine, 2′-F-ara-adenosine, 2′-F-adenosine, 2′-OH-ara- adenosine, and N6-(19-amino-pentaoxanonadecyl)-adenosine. [0166] In some embodiments, the modified nucleobase is a modified guanine. Exemplary nucleobases and nucleosides having a modified guanine include inosine, 1- methyl-inosine, wyosine, methylwyosine, 4-demethyl-wyosine, isowyosine (imG2), wybutosine, peroxywybutosine, hydroxywybutosine, undermodified hydroxywybutosine, 7-deaza-guanosine, queuosine, epoxyqueuosine, galactosyl-queuosine (galQ), mannosyl-queuosine, 7-cyano-7-deaza-guanosine, 7-aminomethyl-7-deaza-guanosine, archaeosine, 7-deaza-8-aza-guanosine, 6-thio-guanosine, 6-thio-7-deaza-guanosine, 6- thio-7-deaza-8-aza-guanosine, 7-methyl-guanosine, 6-thio-7-methyl-guanosine, 7-
1005272698
56 methyl-inosine, 6-methoxy-guanosine, 1-methyl-guanosine, N2-methyl-guanosine, N2,N2-dimethyl-guanosine, N2,7-dimethyl-guanosine, N2, N2,7-dimethyl-guanosine, 8- oxo-guanosine, 7-methyl-8-oxo-guanosine, 1-methyl-6-thio-guanosine, N2-methyl-6- thio-guanosine, N2,N2-dimethyl-6-thio-guanosine, α-thio-guanosine, 2′-O-methyl- guanosine, N2-methyl-2′-O-methyl-guanosine, N2,N2-dimethyl-2′-O-methyl-guanosine, 1-methyl-2′-O-methyl-guanosine, N2,7-dimethyl-2′-O-methyl-guanosine, 2′-O-methyl- inosine, 1,2′-O-dimethyl-inosine, 2′-O-ribosylguanosine (phosphate) (Gr(p)), 1-thio- guanosine, 06-methyl-guanosine, 2′-F-ara-guanosine, and 2′-F-guanosine. [0167] In some embodiments, the RNA (e.g., RNA) vaccines comprise a 5′UTR element, an optionally codon optimized open reading frame, and a 3′UTR element, a poly(A) sequence and/or a polyadenylation signal wherein the RNA is not chemically modified. [0168] Polynucleotides of the present disclosure, in some embodiments, are codon optimized. Codon optimization methods are known in the art and may be used as provided herein. Codon optimization, in some embodiments, may be used to match codon frequencies in target and host organisms to ensure proper folding; bias GC content to increase RNA stability or reduce secondary structures; minimize tandem repeat codons or base runs that may impair gene construction or expression; customize transcriptional and translational control regions; insert or remove protein trafficking sequences; remove/add post translation modification sites in encoded protein (e.g. glycosylation sites); add, remove or shuffle protein domains; insert or delete restriction sites; modify ribosome binding sites and RNA degradation sites; adjust translational rates to allow the various domains of the protein to fold properly; or to reduce or eliminate problem secondary structures within the polynucleotide. Codon optimization tools, algorithms and services are known in the art—non-limiting examples include services from GeneArt (Life Technologies), DNA2.0 (Menlo Park Calif.) and/or proprietary methods. In some embodiments, the open reading frame (ORF) sequence is optimized using optimization algorithms. [0169] In some embodiments, a codon optimized sequence shares less than 95% sequence identity, less than 90% sequence identity, less than 85% sequence identity, less than 80% sequence identity, or less than 75% sequence identity to a naturally- occurring or wild-type sequence (e.g., a naturally-occurring or wild-type RNA sequence
1005272698
57 encoding a polypeptide or protein of interest (e.g., an antigenic protein or antigenic polypeptide)). [0170] In some embodiments, a codon-optimized sequence shares between 65% and 85% (e.g., between about 67% and about 85%, or between about 67% and about 80%) sequence identity to a naturally-occurring sequence or a wild-type sequence (e.g., a naturally-occurring or wild-type RNA sequence encoding a polypeptide or protein of interest (e.g., an antigenic protein or polypeptide)). In some embodiments, a codon- optimized sequence shares between 65% and 75%, or about 80% sequence identity to a naturally-occurring sequence or wild-type sequence (e.g., a naturally-occurring or wild- type RNA sequence encoding a polypeptide or protein of interest (e.g., an antigenic protein or polypeptide)). [0171] In some embodiments a codon-optimized RNA (e.g., mRNA) may, for instance, be one in which the levels of G/C are enhanced. The G/C-content of nucleic acid molecules may influence the stability of the RNA. RNA having an increased amount of guanine (G) and/or cytosine (C) residues may be functionally more stable than nucleic acids containing a large amount of adenine (A) and thymine (T) or uracil (U) nucleotides. WO02/098443 discloses a pharmaceutical composition containing an RNA stabilized by sequence modifications in the translated region. Due to the degeneracy of the genetic code, the modifications work by substituting existing codons for those that promote greater RNA stability without changing the resulting amino acid. The approach is limited to coding regions of the RNA. [0172] Naturally-occurring eukaryotic RNA molecules have been found to contain stabilizing elements, including, but not limited to untranslated regions (UTR) at their 5′- end (5′UTR) and/or at their 3′-end (3′UTR), in addition to other structural features, such as a 5′-cap structure or a 3′-poly(A) tail. Both the 5′UTR and the 3′UTR are typically transcribed from the genomic DNA and are elements of the premature RNA. Characteristic structural features of mature RNA, such as the 5′-cap and the 3′-poly(A) tail are usually added to the transcribed (premature) RNA during RNA processing. The 3′-poly(A) tail is typically a stretch of adenine nucleotides added to the 3′-end of the transcribed RNA. It can comprise up to about 400 adenine nucleotides. In some embodiments the length of the 3′-poly(A) tail may be an essential element with respect to the stability of the individual RNA.
1005272698
58 [0173] In some embodiments the RNA (e.g., mRNA) vaccine may include one or more stabilizing elements. Stabilizing elements may include for instance a histone stem-loop. A stem-loop binding protein (SLBP), a 32 kDa protein has been identified. It is associated with the histone stem-loop at the 3′-end of the histone messages in both the nucleus and the cytoplasm. Its expression level is regulated by the cell cycle; it peaks during the S- phase, when histone RNA levels are also elevated. The protein has been shown to be essential for efficient 3′-end processing of histone pre-RNA by the U7 snRNP. SLBP continues to be associated with the stem-loop after processing, and then stimulates the translation of mature histone RNAs into histone proteins in the cytoplasm. The RNA binding domain of SLBP is conserved through metazoa and protozoa; its binding to the histone stem-loop depends on the structure of the loop. The minimum binding site includes at least three nucleotides 5′ and two nucleotides 3′ relative to the stem-loop. [0174] In some embodiments, the RNA (e.g., mRNA) vaccines include a coding region, at least one histone stem-loop, and optionally, a poly(A) sequence or polyadenylation signal. The poly(A) sequence or polyadenylation signal generally should enhance the expression level of the encoded protein. The encoded protein, in some embodiments, is not a histone protein, a reporter protein (e.g. Luciferase, GFP, EGFP, β-Galactosidase, EGFP), or a marker or selection protein (e.g. alpha-Globin, Galactokinase and Xanthine:guanine phosphoribosyl transferase (GPT)). [0175] In some embodiments, the combination of a poly(A) sequence or polyadenylation signal and at least one histone stem-loop, even though both represent alternative mechanisms in nature, acts synergistically to increase the protein expression beyond the level observed with either of the individual elements. It has been found that the synergistic effect of the combination of poly(A) and at least one histone stem-loop does not depend on the order of the elements or the length of the poly(A) sequence. [0176] In some embodiments, the RNA (e.g., mRNA) vaccine does not comprise a histone downstream element (HDE). “Histone downstream element” (HDE) includes a purine-rich polynucleotide stretch of approximately 15 to 20 nucleotides 3′ of naturally occurring stem-loops, representing the binding site for the U7 snRNA, which is involved in processing of histone pre-RNA into mature histone RNA. Ideally, the inventive nucleic acid does not include an intron.
1005272698
59 [0177] In some embodiments, the RNA (e.g., mRNA) vaccine may or may not contain an enhancer and/or promoter sequence, which may be modified or unmodified or which may be activated or inactivated. In some embodiments, the histone stem-loop is generally derived from histone genes, and includes an intramolecular base pairing of two neighbored partially or entirely reverse complementary sequences separated by a spacer, including (e.g., consisting of) a short sequence, which forms the loop of the structure. The unpaired loop region is typically unable to base pair with either of the stem loop elements. It occurs more often in RNA, as is a key component of many RNA secondary structures, but may be present in single-stranded DNA as well. Stability of the stem-loop structure generally depends on the length, number of mismatches or bulges, and base composition of the paired region. In some embodiments, wobble base pairing (non-Watson-Crick base pairing) may result. In some embodiments, the at least one histone stem-loop sequence comprises a length of 15 to 45 nucleotides. [0178] In other embodiments the RNA (e.g., mRNA) vaccine may have one or more AU-rich sequences removed. These sequences, sometimes referred to as AURES are destabilizing sequences found in the 3′UTR. The AURES may be removed from the RNA (e.g., mRNA) vaccines. Alternatively the AURES may remain in the RNA (e.g., mRNA) vaccine. [0179] In still further embodiments, the RNA of the invention (eg mRNA) may comprise a ribosome skipping sequence, such as a 2A skipping sequence. The use of such sequences in RNA coding sequences are known to the skilled person and may enable the expression of multiple proteins or peptides from a single mRNA. Accordingly, in any embodiment, an mRNA of the invention may encode two or more of the domains K, D, A (including AΔAMB3) or R as defined elsewhere herein and also defined in Table 1, or may encode two or more of the proteins exemplified in Table 1 as being proteins that can be encoded by an RNA sequence of the invention. In certain non-limiting examples, an mRNA of the invention could encode one or more of a KA chimeric protein, a DA chimeric protein, an RA chimeric protein, an AR chimeric protein, an AK chimeric protein, an AD chimeric protein, a KDA chimeric protein, an RDA chimeric protein, a DAR chimeric protein, a DAK chimeric protein or combinations thereof. [0180] Non-limiting examples of 2A peptide sequence for use to introduce ribosome skipping include the T2A or T2A-like sequences derived from Thosea asigna virus and from Porcine teschovirus-12A.
1005272698
60 Polypeptides [0181] It will be appreciated that the polynucleotides of the invention encode a chimeric or fusion protein. The protein encoded by the RNA molecules of the invention may also be termed an “antigenic polypeptide” or simply “antigen”. [0182] As used herein, “polypeptide” means a polymer of amino acid residues (natural or unnatural) linked together most often by peptide bonds. The term, as used herein, refers to proteins, polypeptides, and peptides of any size, structure, or function. In some instances the polypeptide encoded is smaller than about 50 amino acids and the polypeptide is then termed a peptide. If the polypeptide is a peptide, it will be at least about 2, 3, 4, or at least 5 amino acid residues long. Thus, polypeptides include gene products, naturally occurring polypeptides, synthetic polypeptides, homologs, orthologs, paralogs, fragments and other equivalents, variants, and analogs of the foregoing. A polypeptide may be a single molecule or may be a multi-molecular complex such as a dimer, trimer or tetramer. They may also comprise single chain or multichain polypeptides such as antibodies or insulin and may be associated or linked. Most commonly disulfide linkages are found in multichain polypeptides. The term polypeptide may also apply to amino acid polymers in which one or more amino acid residues are an artificial chemical analogue of a corresponding naturally occurring amino acid. [0183] As used herein, a chimeric or fusion protein refers to a polypeptide that comprises amino acid sequences that are not arranged in the same spatial configuration as occurs in nature. For example, and in the context of the present invention, the chimeric or fusion protein encoded by the polynucleotides of the invention, comprises portions of an Arg or Lys- gingipain from P. gingivalis, which are in a different spatial arrangement to full length gingipain. Linkers [0184] In the context of the present invention, the RNA preferably encodes chimeric or fusion proteins comprising various domains (as defined herein), derived from a P. gingivalis gingipain. The domains may be directly joined within the chimeric or fusion protein, or the chimeric or fusion protein may comprise linkers for joining the domains.
1005272698
61 [0185] Suitable linkers for joining amino acid sequences are well known to persons of skill in the art. Preferably, the linker is non-immunogenic. Typically, the linker is comprised of amino acids, and may therefore be termed a peptide linker. [0186] A linker is usually a peptide having a length of up to 20 amino acids, although may be longer. The term “linked to” or “fused to” refers to a covalent bond, e.g., a peptide bond, formed between two moieties. Accordingly, in the context of the present invention the linker may have a length of 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21 or 22 or more amino acids. For example, the chimeric or fusion proteins encoded by the RNAs of the invention, may comprise a linker between the amino acid sequence of a P. gingivalis gingipain active site, and the amino acid sequence of the adhesin domain of a P. gingivalis gingipain. Such linkers have the advantage that they can make it more likely that the different polypeptides of the fusion protein fold independently and behave as expected. Suitable linkers may be up to 50 amino acids in length, although less than 20, less than 15 or less than five amino acids is preferred. The linker may function to bring the domains into a closer spatial arrangement than normally observed in a P. gingivalis trypsin-like enzyme. Alternatively, it may space domains apart. [0187] Suitable linkers for use in protein constructs, including those with minimal impact on solubility are known in the art. The linker may be any linker known in the art to the skilled person and may be a flexible linker (such as those comprising repeats of glycine and serine residues), a rigid linker (such as those comprising glutamic acid and lysine residues, flanking alanine repeats) and/or a cleavable linker (such as sequences that are susceptible by protease cleavage). Examples of such linkers are known to the skilled person and are described for example, in Chen et al., (2013) Advanced Drug Delivery Reviews, 65: 1357-1369. [0188] Useful linkers include glycine-serine (GlySer) linkers, which are well-known in the art and comprise glycine and serine units combined in various orders. Examples include, but are not limited to, (GS), (GSGGS)n (SEQ ID NO: 88), (GGGS)n (SEQ ID NO: 89) and (GGGGS)n (SEQ ID NO: 90), where n is an integer of at least one, typically an integer between 1 and about 10, for example, between 1 and about 8, between 1 and about 6, or between 1 and about 5. [0189] In some embodiments, the peptide linker may include the amino acids glycine and serine in various lengths and combinations. In some aspects, the peptide linker can
1005272698
62 include the sequence Gly-Gly-Ser (GGS), Gly-Gly-Gly-Ser (GGGS, SEQ ID NO: 89) or Gly-Gly-Gly-Gly-Ser (GGGGS, SEQ ID NO: 90) and variations or repeats thereof. In some aspects, the peptide linker can include the amino acid sequence GGGGS (a linker of 6 amino acids in length, SEQ ID NO: 90) or even longer. The linker may be a series of repeating glycine and serine residues (GS) of different lengths, i.e., (GS)n where n is any number from 1 to 15 or more. For example, the linker may be (GS)3 (i.e., GSGSGS, SEQ ID NO: 91) or longer (GS)11 or longer. It will be appreciated that n can be any number including 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 or more. Fusion proteins having linkers of such length are included within the scope of the present invention. Similarly, the linker may be a series of repeating glycine residues separated by serine residues. For example (GGGGS)3 (i.e., the linker may comprise the amino acid sequence GGGGSGGGGSGGGGS, (G4S)3, SEQ ID NO: 92) and variations thereof. [0190] In one embodiment, the peptide linker can include the amino acid sequence GGGGS (a linker of 6 amino acids in length) or even longer. The linker may a series of repeating glycine and serine residues (GS) of different lengths, i.e., (GS)n where n is any number from 1 to 15 or more. For example, the linker may be (GS)3 (i.e., GSGSGS, SEQ ID NO: 91) or longer (GS)11 or longer. It will be appreciated that n can be any number including 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11 or more. [0191] Other useful linkers include DSSG (SEQ ID NO: 93), DSSGAS (SEQ ID NO: 94), KLDSSG (SEQ ID NO: 95) and variations thereof. Examples of other suitable linkers are described in Chen et al., (2013) Advanced Drug Delivery Reviews, 65: 1357-1369. [0192] The term “polypeptide variant” refers to molecules which differ in their amino acid sequence from a native or reference sequence. The amino acid sequence variants may possess substitutions, deletions, and/or insertions at certain positions within the amino acid sequence, as compared to a native or reference sequence. Ordinarily, variants will possess at least about 50% identity (homology) to a native or reference sequence, and preferably, they will be at least about 80%, more preferably at least about 90% identical (homologous) to a native or reference sequence. [0193] The present invention contemplates several types of compositions which encode polypeptides, including variants and derivatives. These include substitutional, insertional, deletion and covalent variants and derivatives. The term “derivative” is used
1005272698
63 synonymously with the term “variant” but generally refers to a molecule that has been modified and/or changed in any way relative to a reference molecule or starting molecule. [0194] As such, RNAs of the invention encoding polypeptides containing substitutions, insertions and/or additions, deletions and covalent modifications with respect to reference sequences, in particular the polypeptide sequences disclosed herein, are included within the scope of this invention. For example, sequence tags or amino acids, such as one or more lysines, can be added to the peptide sequences of the invention (e.g., at the N- terminal or C-terminal ends). Sequence tags can be used for peptide purification or localization. Lysines can be used to increase peptide solubility or to allow for biotinylation. Alternatively, amino acid residues located at the carboxy and amino terminal regions of the amino acid sequence of a peptide or protein may optionally be deleted providing for truncated sequences. Certain amino acids (e.g., C-terminal or N-terminal residues) may alternatively be deleted depending on the use of the sequence, as for example, expression of the sequence as part of a larger sequence which is soluble, or linked to a solid support. [0195] “Substitutional variants” when referring to polypeptides are those that have at least one amino acid residue in a native or starting sequence removed and a different amino acid inserted in its place at the same position. Substitutions may be single, where only one amino acid in the molecule has been substituted, or they may be multiple, where two or more (e.g., 3, 4 or 5) amino acids have been substituted in the same molecule. In certain embodiments, the substitutions may be conservative amino acid substitutions. [0196] As used herein the term “conservative amino acid substitution” refers to the substitution of an amino acid that is normally present in the sequence with a different amino acid of similar size, charge, or polarity. Examples of conservative substitutions include the substitution of a non-polar (hydrophobic) residue such as isoleucine, valine and leucine for another non-polar residue. Likewise, examples of conservative substitutions include the substitution of one polar (hydrophilic) residue for another such as between arginine and lysine, between glutamine and asparagine, and between glycine and serine. Additionally, the substitution of a basic residue such as lysine, arginine or histidine for another, or the substitution of one acidic residue such as aspartic acid or glutamic acid for another acidic residue are additional examples of conservative substitutions. Examples of non-conservative substitutions include the substitution of a non-polar (hydrophobic) amino acid residue such as isoleucine, valine, leucine, alanine,
1005272698
64 methionine for a polar (hydrophilic) residue such as cysteine, glutamine, glutamic acid or lysine and/or a polar residue for a non-polar residue. [0197] Amino acid deletions or insertions can also be made relative to the native sequence of the P. gingivalis protein. Thus, for example, amino acids which do not have a substantial effect on the activity of the polypeptide, or at least which do not eliminate such activity, can be deleted. [0198] As used herein when referring to polypeptides the term “domain” refers to a motif of a polypeptide having one or more identifiable structural or functional characteristics or properties (e.g., binding capacity, serving as a site for protein-protein interactions). [0199] In any embodiment herein, the skilled person may make modifications to an RNA molecule of the invention so that it includes codons encoding additional amino acid residues derived from the naturally occurring domain sequences of the gingipain. For example, where an RNA of the invention encodes a chimeric protein (such as RA, KA, KDA, KDAK and the like), and wherein the sequences of K, R, D and A are as herein defined, it will be within the purview of the skilled person to include additional codons encoding additional amino acids at the N or C termini of each domain, for example, in order to further stabilise the encoded protein. Typically, the additional amino acids correspond to naturally occurring gingipain sequence. In one non-limiting example, a methionine may be encoded at the N terminal region of the A domain. It will be appreciated that the RNA may encode 1, 2, 3, 4, 5, 6, 7, 8, 9 or 10 or more additional amino acid residues. In another example, the D domain may comprise a C terminal methionine residue (or alternatively this residue may be omitted from the sequence of the D domain). [0200] As used herein the terms “termini” or “terminus” when referring to polypeptides or polynucleotides refers to an extremity of a polypeptide or polynucleotide respectively. Such extremity is not limited only to the first or final site of the polypeptide or polynucleotide but may include additional amino acids or nucleotides in the terminal regions. Polypeptide-based molecules may be characterized as having both an N- terminus (terminated by an amino acid with a free amino group (NH2)) and a C-terminus (terminated by an amino acid with a free carboxyl group (COOH)). Proteins are in some cases made up of multiple polypeptide chains brought together by disulfide bonds or by
1005272698
65 non-covalent forces (multimers, oligomers). These proteins have multiple N- and C- termini. Alternatively, the termini of the polypeptides may be modified such that they begin or end, as the case may be, with a non-polypeptide based moiety such as an organic conjugate. [0201] As recognized by those skilled in the art, protein fragments, functional protein domains, and homologous proteins are also considered to be within the scope of polypeptides of interest. For example, provided herein is any protein fragment (meaning a polypeptide sequence at least one amino acid residue shorter than a reference polypeptide sequence but otherwise identical) of a reference protein having a length of 10, 20, 30, 40, 50, 60, 70, 80, 90, 100 or longer than 100 amino acids. In another example, any protein that includes a stretch of 20, 30, 40, 50, or 100 (contiguous) amino acids that are 40%, 50%, 60%, 70%, 80%, 90%, 95%, or 100% identical to any of the sequences described herein can be utilized in accordance with the disclosure. In some embodiments, a polypeptide includes 2, 3, 4, 5, 6, 7, 8, 9, 10, or more mutations as shown in any of the sequences provided herein or referenced herein. In another example, any protein that includes a stretch of 20, 30, 40, 50, or 100 amino acids that are greater than 80%, 90%, 95%, or 100% identical to any of the sequences described herein, wherein the protein has a stretch of 5, 10, 15, 20, 25, or 30 amino acids that are less than 80%, 75%, 70%, 65% to 60% identical to any of the sequences described herein can be utilized in accordance with the disclosure. [0202] Polypeptide or polynucleotide molecules of the present disclosure may share a certain degree of sequence similarity or identity with recited SEQ ID NOs. The term “identity,” as known in the art, refers to a relationship between the sequences of two or more polypeptides or polynucleotides, as determined by comparing the sequences. In the art, identity also means the degree of sequence relatedness between two sequences as determined by the number of matches between strings of two or more amino acid residues or nucleic acid residues. Identity measures the percent of identical matches between the smaller of two or more sequences with gap alignments (if any) addressed by a particular mathematical model or computer program (e.g., “algorithms”). Identity of related peptides can be readily calculated by known methods. “% identity” as it applies to polypeptide or polynucleotide sequences is defined as the percentage of residues (amino acid residues or nucleic acid residues) in the candidate amino acid or nucleic acid sequence that are identical with the residues in the amino acid sequence or nucleic acid
1005272698
66 sequence of a second sequence after aligning the sequences and introducing gaps, if necessary, to achieve the maximum percent identity. Methods and computer programs for the alignment are well known in the art. Identity depends on a calculation of percent identity but may differ in value due to gaps and penalties introduced in the calculation. Generally, variants of a particular polynucleotide or polypeptide have at least 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99% but less than 100% sequence identity to that particular reference polynucleotide or polypeptide as determined by sequence alignment programs and parameters described herein and known to those skilled in the art. Such tools for alignment include those of the BLAST suite (Stephen F. Altschul, et al. (1997).” Gapped BLAST and PSI-BLAST: a new generation of protein database search programs,” Nucleic Acids Res. 25:3389-3402). Another popular local alignment technique is based on the Smith-Waterman algorithm (Smith, T. F. & Waterman, M. S. (1981) “Identification of common molecular subsequences.” J. Mol. Biol. 147:195-197). A general global alignment technique based on dynamic programming is the Needleman-Wunsch algorithm (Needleman, S. B. & Wunsch, C. D. (1970) “A general method applicable to the search for similarities in the amino acid sequences of two proteins.” J. Mol. Biol.48:443- 453). More recently, a Fast Optimal Global Sequence Alignment Algorithm (FOGSAA) was developed that purportedly produces global alignment of nucleotide and protein sequences faster than other optimal global alignment methods, including the Needleman- Wunsch algorithm. Other tools are described herein, specifically in the definition of “identity” below. [0203] The term “identity” refers to the overall relatedness between polymeric molecules, for example, between polynucleotide molecules (e.g. DNA molecules and/or RNA molecules) and/or between polypeptide molecules. Calculation of the percent identity of two polynucleic acid sequences, for example, can be performed by aligning the two sequences for optimal comparison purposes (e.g., gaps can be introduced in one or both of a first and a second nucleic acid sequences for optimal alignment and non- identical sequences can be disregarded for comparison purposes). In certain embodiments, the length of a sequence aligned for comparison purposes is at least 30%, at least 40%, at least 50%, at least 60%, at least 70%, at least 80%, at least 90%, at least 95%, or 100% of the length of the reference sequence. The nucleotides at corresponding nucleotide positions are then compared. When a position in the first sequence is occupied by the same nucleotide as the corresponding position in the second sequence, then the
1005272698
67 molecules are identical at that position. The percent identity between the two sequences is a function of the number of identical positions shared by the sequences, taking into account the number of gaps, and the length of each gap, which needs to be introduced for optimal alignment of the two sequences. The comparison of sequences and determination of percent identity between two sequences can be accomplished using a mathematical algorithm. For example, the percent identity between two nucleic acid sequences can be determined using methods such as those described in Computational Molecular Biology, Lesk, A. M., ed., Oxford University Press, New York, 1988; Biocomputing: Informatics and Genome Projects, Smith, D. W., ed., Academic Press, New York, 1993; Sequence Analysis in Molecular Biology, von Heinje, G., Academic Press, 1987; Computer Analysis of Sequence Data, Part I, Griffin, A. M., and Griffin, H. G., eds., Humana Press, New Jersey, 1994; and Sequence Analysis Primer, Gribskov, M. and Devereux, J., eds., M Stockton Press, New York, 1991; each of which is incorporated herein by reference. For example, the percent identity between two nucleic acid sequences can be determined using the algorithm of Meyers and Miller (CABIOS, 1989, 4:11-17), which has been incorporated into the ALIGN program (version 2.0) using a PAM 120 weight residue table, a gap length penalty of 12 and a gap penalty of 4. The percent identity between two nucleic acid sequences can, alternatively, be determined using the GAP program in the GCG software package using an NWSgapdna.CMP matrix. Methods commonly employed to determine percent identity between sequences include, but are not limited to those disclosed in Carillo, H., and Lipman, D., SIAM J Applied Math., 48:1073 (1988); incorporated herein by reference. Techniques for determining identity are codified in publicly available computer programs. Exemplary computer software to determine homology between two sequences include, but are not limited to, GCG program package, Devereux, J., et al., Nucleic Acids Research, 12(1), 387 (1984)), BLASTP, BLASTN, and FASTA Altschul, S. F. et al., J. Molec. Biol., 215, 403 (1990)). Signal peptides [0204] The polypeptides encoded by the polynucleotides of the invention typically comprise N-terminal signal peptides. Signal peptides, comprising the N-terminal 15-60 amino acids of proteins, are typically needed for the translocation across the membrane on the secretory pathway and, thus, universally control the entry of most proteins both in eukaryotes and prokaryotes to the secretory pathway. Signal peptides generally include three regions: an N-terminal region of differing length, which usually comprises positively
1005272698
68 charged amino acids; a hydrophobic region; and a short carboxy-terminal peptide region. In eukaryotes, the signal peptide of a nascent precursor protein (pre-protein) directs the ribosome to the rough endoplasmic reticulum (ER) membrane and initiates the transport of the growing peptide chain across it for processing. ER processing produces mature proteins, wherein the signal peptide is cleaved from precursor proteins, typically by a ER- resident signal peptidase of the host cell, or they remain uncleaved and function as a membrane anchor. A signal peptide may also facilitate the targeting of the protein to the cell membrane. The signal peptide, however, is not responsible for the final destination of the mature protein. Secretory proteins devoid of additional address tags in their sequence are by default secreted to the external environment. During recent years, a more advanced view of signal peptides has evolved, showing that the functions and immunodominance of certain signal peptides are much more versatile than previously anticipated. [0205] In any embodiment, the N-terminal secretion signal peptide may comprise any amino acid sequence which enables the chimeric or fusion protein to be processed by ribosomes bound to the rough endoplasmic reticulum (ER) of a cell, and thereby results in threading of the chimeric or fusion protein into the ER. [0206] Preferably the N-terminal secretion signal peptide is any peptide that enables secretion of the encoded protein by the cell in which the RNA is expressed or translated. [0207] In some embodiments, the signal peptide fused to the antigenic polypeptide is an artificial signal peptide. In some embodiments, an artificial signal peptide fused to the antigenic polypeptide encoded by the RNA (e.g., mRNA) vaccine is obtained from an immunoglobulin protein, e.g., an IgE signal peptide or an IgG signal peptide. In some embodiments, a signal peptide fused to the antigenic polypeptide encoded by a RNA (e.g., mRNA) vaccine is an Ig heavy chain epsilon-1 signal peptide (IgE HC SP) having the sequence of: MDWTWILFLVAAATRVHS (SEQ ID NO: 96). [0208] In some embodiments, a signal peptide fused to the antigenic polypeptide encoded by the RNA (e.g., mRNA) vaccine is an IgGk chain V-III region HAH signal peptide (IgGk SP) having the sequence of METPAQLLFLLLLWLPDTTG (SEQ ID NO: 97). In some embodiments, the signal peptide is selected from: Japanese encephalitis PRM signal sequence (MLGSNSGQRVVFTILLLLVAPAYS, SEQ ID NO: 98), VSVg
1005272698
69 protein signal sequence (MKCLLYLAFLFIGVNCA, SEQ ID NO: 99) and Japanese encephalitis JEV signal sequence (MWLVSLAIVTACAGA, SEQ ID NO: 100). [0209] Further examples of suitable signal peptides include sequences derived from tPA (tissue plasminogen activator): MDAMKRGLCCVLLLCGAVFVSPS (SEQ ID NO: 101) variants, such as: tPA (VSA): MDAMKRGLCCVLLLCGAVFVSA (SEQ ID NO: 102), tPA (VSAR): MDAMKRGLCCVLLLCGAVFVSAR (SEQ ID NO: 103), tPA (VSP): MDAMKRGLCCVLLLCGAVFVSP (SEQ ID NO: 104), tPA (VSPS): MDAMKRGLCCVLLLCGAVFVSPS (SEQ ID NO: 101). [0210] In certain embodiments, the amino acid sequence of the signal peptide comprises the sequence of SEAP (secreted embryonic alkaline phosphatase): MLLLLLLLGLRLQLSLG[A] (SEQ ID NO: 105), such that the expressed RNA product comprises the sequence MLLLLLLLGLRLQLSLG[A] (SEQ ID NO: 105) N terminal to the sequences defined herein including in Table 1. [0211] In preferred embodiments the invention, the signal peptide comprises the sequence as set forth in any of these examples, of a sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto. [0212] The examples disclosed herein are not meant to be limiting and any signal peptide that is known in the art to facilitate targeting of a protein to ER for processing and/or targeting of a protein to the cell membrane may be used in accordance with the present disclosure. [0213] A signal peptide may have a length of 15-60 amino acids. For example, a signal peptide may have a length of 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, or 60 amino acids. In some embodiments, a signal peptide has a length of 20-60, 25-60, 30-60, 35-60, 40-60, 45-60, 50-60, 55-60, 15-55, 20-55, 25-55, 30-55, 35-55, 40-55, 45-55, 50-55, 15-50, 20-50, 25-50, 30-50, 35-50, 40-50, 45-50, 15- 45, 20-45, 25-45, 30-45, 35-45, 40-45, 15-40, 20-40, 25-40, 30-40, 35-40, 15-35, 20-35, 25-35, 30-35, 15-30, 20-30, 25-30, 15-25, 20-25, or 15-20 amino acids.
1005272698
70 [0214] A signal peptide is typically cleaved from the nascent polypeptide at the cleavage junction during ER processing. The mature antigenic polypeptide produces by an RNA vaccine of the present disclosure typically does not comprise a signal peptide. [0215] It will be well within the purview of the skilled person to design a polypeptide encoded by an RNA of the invention, to facilitate expression and translation thereof in vivo. For example, in certain non-limiting examples, the RNA of the invention may include sequence encoding an N terminal methionine, and/or other residues (such as alanine) for enabling expression, secretion and/or cleavage of the signal peptide. [0216] Accordingly, in any embodiment, the RNA of the invention may encode one, two, three, four or more N terminal amino acids to the sequences defined herein in Table 1. For example, an RNA encoding an amino acid sequence as set forth in any of SEQ ID NOs: 2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 38 or 39 may encode one or more additional N terminal amino acids, optionally an N terminal alanine residue and/or optionally an N terminal methionine residue. [0217] In further examples, an RNA encoding an amino acid sequence of a D domain (eg in SEQ ID NOs: 35 and 36) may comprise the C terminal methionine residue (as shown in these sequence) or the C terminal methionine residue may be omitted from the D domain sequence. Compositions and lipid nanoparticles [0218] The present invention contemplates the provision of a polynucleotide (preferably an RNA) encoding an a chimeric or fusion protein for inducing an immune response to P. gingivalis, preferably formulated in a lipid nanoparticle. Accordingly, the present invention also provides a lipid nanoparticle comprising a polynucleotide as described herein. It will be appreciated that in any embodiment, the nanoparticles of the invention may also be described as “vaccine” compositions or “immune stimulating” compositions. [0219] In some embodiments, the RNA of the invention is formulated in a lipid- polycation complex, referred to as a cationic lipid nanoparticle. As a non-limiting example, the polycation may include a cationic peptide or a polypeptide such as, but not limited to, polylysine, polyornithine and/or polyarginine. In some embodiments, the RNA may be
1005272698
71 formulated in a lipid nanoparticle that includes a non-cationic lipid such as, but not limited to, cholesterol or dioleoyl phosphatidylethanolamine (DOPE). [0220] Lipid nanoparticle formulations typically comprise a lipid, in particular, an ionizable cationic lipid, and further comprise a non-cationic lipid, a sterol and a molecule capable of reducing particle aggregation, for example a PEG or PEG-modified lipid. [0221] In some embodiments, a cationic lipid is an ionizable cationic lipid and the non- cationic lipid is a neutral lipid, and the sterol is a cholesterol. In some embodiments, a cationic lipid is selected from the group consisting of 2,2-dilinoleyl-4-dimethylaminoethyl- [1,3]-dioxolane (DLin-KC2-DMA), dilinoleyl-methyl-4-dimethylaminobutyrate (DLin-MC3- DMA), di((Z)-non-2-en-1-yl) 9-((4-(dimethylamino)butanoyl)oxy)heptadecanedioate (L319), (12Z,15Z)—N,N-dimethyl-2-nonylhenicosa-12,15-dien-1-amine (L608), and N,N- dimethyl-1-[(1S,2R)-2-octylcyclopropyl]heptadecan-8-amine (L530). [0222] In some embodiments, lipid nanoparticle formulations include 25-75% of a cationic lipid optionally selected from 2,2-dilinoleyl-4-dimethylaminoethyl-[1,3]-dioxolane (DLin-KC2-DMA), dilinoleyl-methyl-4-dimethylaminobutyrate (DLin-MC3-DMA), and di((Z)-non-2-en-1-yl) 9-((4-(dimethylamino)butanoyl)oxy)heptadecanedioate (L319), 0.5- 15% of the neutral lipid, 5-50% of the sterol, and 0.5-20% of the PEG or PEG-modified lipid on a molar basis. [0223] In some embodiments, lipid nanoparticle formulations consists essentially of a lipid mixture in molar ratios of 20-70% cationic lipid: 5-45% neutral lipid: 20-55% cholesterol: 0.5-15% PEG-modified lipid. In some embodiments, lipid nanoparticle formulations consists essentially of a lipid mixture in a molar ratio of 20-60% cationic lipid: 5-25% neutral lipid: 25-55% cholesterol: 0.5-15% PEG-modified lipid. [0224] In some embodiments, the molar lipid ratio is 50/10/38.5/1.5 (mol % cationic lipid/neutral lipid, e.g., DSPC/Chol/PEG-modified lipid, e.g., PEG-DMG, PEG-DSG or PEG-DPG), 57.2/7.1134.3/1.4 (mol % cationic lipid/neutral lipid, e.g., DPPC/Chol/PEG- modified lipid, e.g., PEG-cDMA), 40/15/40/5 (mol % cationic lipid/neutral lipid, e.g., DSPC/Chol/PEG-modified lipid, e.g., PEG-DMG), 50/10/35/4.5/0.5 (mol % cationic lipid/neutral lipid, e.g., DSPC/Chol/PEG-modified lipid, e.g., PEG-DSG), 50/10/35/5 (cationic lipid/neutral lipid, e.g., DSPC/Chol/PEG-modified lipid, e.g., PEG-DMG), 40/10/40/10 (mol % cationic lipid/neutral lipid, e.g., DSPC/Chol/PEG-modified lipid, e.g.,
1005272698
72 PEG-DMG or PEG-cDMA), 35/15/40/10 (mol % cationic lipid/neutral lipid, e.g., DSPC/Chol/PEG-modified lipid, e.g., PEG-DMG or PEG-cDMA) or 52/13/30/5 (mol % cationic lipid/neutral lipid, e.g., DSPC/Chol/PEG-modified lipid, e.g., PEG-DMG or PEG- cDMA). [0225] A lipid nanoparticle formulation may be influenced by, but not limited to, the selection of the cationic lipid component, the degree of cationic lipid saturation, the nature of the PEGylation, ratio of all components and biophysical parameters such as size. In one example by Semple et al. (Nature Biotech.201028:172-176), the lipid nanoparticle formulation is composed of 57.1% cationic lipid, 7.1% dipalmitoylphosphatidylcholine, 34.3% cholesterol, and 1.4% PEG-c-DMA. [0226] In some embodiments, the lipid nanoparticle comprises a molar ratio of about 20-60% cationic lipid, 0.5-15% PEG-modified lipid, 25-55% sterol, and 25% non-cationic lipid. In some embodiments, the cationic lipid is an ionizable cationic lipid and the non- cationic lipid is a neutral lipid, and the sterol is a cholesterol. In some embodiments, the cationic lipid is selected from 2,2-dilinoleyl-4-dimethylaminoethyl-[1,3]-dioxolane (DLin- KC2-DMA), dilinoleyl-methyl-4-dimethylaminobutyrate (DLin-MC3-DMA), and di((Z)-non- 2-en-1-yl) 9-((4-(dimethylamino)butanoyl)oxy)heptadecanedioate (L319). [0227] In some embodiments, lipid nanoparticle formulations may comprise 35 to 45% cationic lipid, 40% to 50% cationic lipid, 50% to 60% cationic lipid and/or 55% to 65% cationic lipid. [0228] In some embodiments, the ratio of lipid to RNA (e.g., mRNA) in lipid nanoparticles may be 5:1 to 20:1, 10:1 to 25:1, 15:1 to 30:1 and/or at least 30:1. [0229] In some embodiments, the ratio of PEG in the lipid nanoparticle formulations may be increased or decreased and/or the carbon chain length of the PEG lipid may be modified from C14 to C18 to alter the pharmacokinetics and/or biodistribution of the lipid nanoparticle formulations. As a non-limiting example, lipid nanoparticle formulations may contain 0.5% to 3.0%, 1.0% to 3.5%, 1.5% to 4.0%, 2.0% to 4.5%, 2.5% to 5.0% and/or 3.0% to 6.0% of the lipid molar ratio of PEG-c-DOMG (R-3-[(ω-methoxy- poly(ethyleneglycol)2000)carbamoyl)]-1,2-dimyristyloxypropyl-3-amine) (also referred to herein as PEG-DOMG) as compared to the cationic lipid, DSPC and cholesterol. In some embodiments, the PEG-c-DOMG may be replaced with a PEG lipid such as, but not
1005272698
73 limited to, PEG-DSG (1,2-Distearoyl-sn-glycerol, methoxypolyethylene glycol), PEG- DMG (1,2-Dimyristoyl-sn-glycerol) and/or PEG-DPG (1,2-Dipalmitoyl-sn-glycerol, methoxypolyethylene glycol). The cationic lipid may be selected from any lipid known in the art such as, but not limited to, DLin-MC3-DMA, DLin-DMA, C12-200 and DLin-KC2- DMA. [0230] The amino alcohol cationic lipid may be a lipids described in and/or made by the methods described in U.S. Patent Publication No. US20130150625, herein incorporated by reference in its entirety. As a non-limiting example, the cationic lipid may be 2-amino- 3-[(9Z,12Z)-octadeca-9,12-dien-1-yloxy]-2-{[(9Z,2Z)-octadeca-9,12-dien-1-yloxy] methyl}propan-1-ol (Compound 1 in US20130150625); 2-amino-3-[(9Z)-octadec-9-en-1- yloxy]-2-{[(9Z)-octadec-9-en-1-yloxy]methyl}propan-1-ol (Compound 2 in US20130150625); 2-amino-3-[(9Z,12Z)-octadeca-9,12-dien-1-yloxy]-2- [(octyloxy)methyl]propan-1-ol (Compound 3 in US20130150625); and 2-(dimethylamino)- 3-[(9Z,12Z)-octadeca-9,12-dien-1-yloxy]-2-{[(9Z, 12Z)-octadeca-9,12-dien-1- yloxy]methyl}propan-1-ol (Compound 4 in US20130150625); or any pharmaceutically acceptable salt or stereoisomer thereof. [0231] The cationic lipid may be any one of N,N-dioleyl-N,N-dimethylammonium chloride (DODAC), 1,2-dioeoyloxy-3-(dimethylamino)propane (DODAP), 1,2-dioleyloxy- N,N-dimethylaminopropane (DODMA), 1,2-distearyloxy-N,N-dimethylaminopropane (DSDMA), N-(1-(2,3-dioleyloxy)propyl)-N,N,N-trimethylammonium chloride (DOTMA), N,N-distearyl-N,N-dimethylammonium bromide (DDAB), N-(1-(2,3-dioleoyloxy)propyl)- N,N,N-trimethylammonium chloride (DOTAP), 3-(N—(N′,N′-dimethylaminoethane)- carbamoyl)cholesterol (DC-Chol), N-(1,2-dimyristyloxyprop-3-yl)-N,N-dimethyl-N- hydroxyethyl ammonium bromide (DMRIE), 2,3-dioleyloxy-N-[2(spermine- carboxamido)ethyl]-N,N-dimethyl-1-propanaminiumtrifluoroacetate (DOSPA), dioctadecylamidoglycyl spermine (DOGS), 3-dimethylamino-2-(cholest-5-en-3-beta- oxybutan-4-oxy)-1-(cis,cis-9,12-octadecadienoxy)propane (CLinDMA), 2-[5′-(cholest-5- en-3.beta.-oxy)-3′-oxapentoxy)-3-dimethy-1-(cis,cis-9′,1-2′-octadecadienoxy)propane (CpLinDMA), N,N-dimethyl-3,4-dioleyloxybenzylamine (DMOBA), 1,2-N,N′- dioleylcarbamyl-3-dimethylaminopropane (DOcarbDAP), 1,2-N,N′-Dilinoleylcarbamyl-3- dimethylaminopropane (DLincarbDAP), 1,2-Dilinoleoylcarbamyl-3- dimethylaminopropane (DLinCDAP), 4-Hydroxybutyl)azanediyl)bis(hexane-6,1-diyl) bis(2-hexyldecanoate) (ALC-0315), 8-[(2-hydroxyethyl)[6-oxo-6-
1005272698
74 (undecyloxy)hexyl]amino]-octanoic acid,1-octylnonyl ester (SM-102) and mixtures thereof. [0232] The cationic lipid may be of Formula I
[0233] wherein R1 and R2 are independently selected and are H or C1-C3 alkyls, R3 and R4 are independently selected and are alkyl groups having from about 10 to about 20 carbon atoms, and at least one of R3 and R4 comprises at least two sites of unsaturation, preferably the cationic lipid of Formula I is 1,2-dilinoleyloxy-N,N-dimethylaminopropane (DLinDMA) or 1,2-dilinolenyloxy-N,N-dimethylaminopropane (DLenDMA). [0234] The cationic lipid may be of Formula II
[0235] wherein R1 and R2 are independently selected and are H or C1-C3 alkyls, R3 and R4 are independently selected and are alkyl groups having from about 10 to about 20 carbon atoms, and at least one of R3 and R4 comprises at least two sites of unsaturation. [0236] The cationic lipid may be of Formula III
[0237] wherein R1 and R2 are either the same or different and independently optionally substituted C12-C24 alkyl, optionally substituted C12-C24 alkenyl, optionally substituted C12- C24 alkynyl, or optionally substituted C12-C24 acyl; R3 and R4 are either the same or
1005272698
75 different and independently optionally substituted C1-C6 alkyl, optionally substituted C1- C6 alkenyl, or optionally substituted C1-C5 alkynyl or R3 and R4 may join to form an optionally substituted heterocyclic ring of 4 to 6 carbon atoms and 1 or 2 heteroatoms chosen from nitrogen and oxygen; R5 is either absent or hydrogen or C1-C6 alkyl to provide a quaternary amine; m, n, and p are either the same or different and independently either 0 or 1 with the proviso that m, n, and p are not simultaneously 0; q is 0, 1, 2, 3, or 4; and Y and Z are either the same or different and independently O, S, or NH. [0238] The cationic lipid of Formula III may be 2,2-dilinoleyl-4-(2-dimethylaminoethyl)- [1,3]-dioxolane (DLin-K-C2-DMA; “XTC2”), 2,2-dilinoleyl-4-(3-dimethylaminopropyl)-[1,3]- dioxolane (DLin-K-C3-DMA), 2,2-dilinoleyl-4-(4-dimethylaminobutyl)-[1,3]-dioxolane (DLin-K-C4-DMA), 2,2-dilinoleyl-5-dimethylaminomethyl-[1,3]-dioxane (DLin-K6-DMA), 2,2-dilinoleyl-4-N-methylpepiazino-[1,3]-dioxolane (DLin-K-MPZ), 2,2-dilinoleyl-4- dimethylaminomethyl-[1,3]-dioxolane (DLin-K-DMA), 1,2-dilinoleylcarbamoyloxy-3- dimethylaminopropane (DLin-C-DAP), 1,2-dilinoleyoxy-3- (dimethylamino)acetoxypropane (DLin-DAC), 1,2-dilinoleyoxy-3-morpholinopropane (DLin-MA), 1,2-dilinoleoyl-3-dimethylaminopropane (DLinDAP), 1,2-dilinoleylthio-3- dimethylaminopropane (DLin-S-DMA), 1-linoleoyl-2-linoleyloxy-3-dimethylaminopropane (DLin-2-DMAP), 1,2-dilinoleyloxy-3-trimethylaminopropane chloride salt (DLin-TMA.C1), 1,2-dilinoleoyl-3-trimethylaminopropane chloride salt (DLin-TAP.C1), 1,2-dilinoleyloxy-3- (N-methylpiperazino)propane (DLin-MPZ), 3-(N,N-dilinoleylamino)-1,2-propanediol (DLinAP), 3-(N,N-dioleylamino)-1,2-propanedio (DOAP), 1,2-dilinoleyloxo-3-(2-N,N- dimethylamino)ethoxypropane (DLin-EG-DMA), or mixtures thereof. [0239] The phospholipid may be according to Formula (IV):
1005272698
76 in which represents a phospholipid moiety and R and R’ represent fatty acid moieties with or without unsaturation that may be the same or different. [0240] The phospholipid moiety may be selected from the group consisting of: phosphatidyl choline, phosphatidyl ethanolamine, phosphatidyl glycerol, phosphatidyl serine, phosphatidic acid, 2-lysophosphatidyl choline, and a sphingomyelin. [0241] The phospholipid may have a fatty acid moiety selected from the non-limiting group consisting of: lauric acid, myristic acid, myristoleic acid, palmitic acid, palmitoleic acid, stearic acid, oleic acid, linoleic acid, alpha-linolenic acid, erucic acid, phytanoic acid, arachidic acid, arachidonic acid, eicosapentaenoic acid, behenic acid, docosapentaenoic acid, and docosahexaenoic acid. [0242] The phosopholipid may be lecithin, phosphatidylethanolamine, lysolecithin, lysophosphatidylethanolamine, phosphatidylserine, phosphatidylinositol, sphingomyelin, egg sphingomyelin (ESM), cephalin, cardiolipin, phosphatidic acid, cerebrosides, dicetylphosphate, distearoylphosphatidylcholine (DSPC), dioleoylphosphatidylcholine (DOPC), dipalmitoylphosphatidylcholine (DPPC), dioleoylphosphatidylglycerol (DOPG), dipalmitoylphosphatidylglycerol (DPPG), dioleoylphosphatidylethanolamine (DOPE), palmitoyloleoyl-phosphatidylcholine (POPC), palmitoyloleoyl-phosphatidylethanolamine (POPE), palmitoyloleyol-phosphatidylglycerol (POPG), dioleoylphosphatidylethanolamine 4-(N-maleimidomethyl)-cyclohexane-1-carboxylate (DOPE-mal), dipalmitoyl-phosphatidylethanolamine (DPPE), dimyristoyl- phosphatidylethanolamine (DMPE), distearoyl-phosphatidylethanolamine (DSPE), monomethyl-phosphatidylethanolamine, dimethyl-phosphatidylethanolamine, dielaidoyl- phosphatidylethanolamine (DEPE), stearoyloleoyl-phosphatidylethanolamine (SOPE), lysophosphatidylcholine, dilinoleoylphosphatidylcholine, and mixtures thereof. [0243] The phosopholipid may be distearoylphosphatidylcholine (DSPC). [0244] The phospholipid may comprise from about 5 mol % to about 20 mol %, from about 5 mol % to about 15 mol %, from about 5 mol % to about 10 mol %, from about 10 mol % to about 20 mol %, from about 15 mol % to about 20 mol % of the total lipid present in the particle.
1005272698
77 [0245] The phospholipid may comprise from 5 mol % to 20 mol %, from 5 mol % to 15 mol %, from 5 mol % to 10 mol %, from 10 mol % to 20 mol %, or from 15 mol % to 20 mol % of the total lipid present in the particle. [0246] The structural lipid may be selected from the group consisting of cholesterol, fecosterol, sitosterol, ergosterol, campesterol, stigmasterol, brassicasterol, tomatidine, tomatine, ursolic acid, alpha-tocopherol, and mixtures thereof. [0247] The structural lipid may be cholesterol. In some embodiments, the structural lipid includes cholesterol and a corticosteroid (such as prednisolone, dexamethasone, prednisone, and hydrocortisone), or a combination thereof. Further, the structural lipid may be squalene, squalene or combination thereof. [0248] The structural lipid may include lipids containing geranyl acetate, farnesyl acetate or geranyl-geranyl, or ether, ester, or other derivatives. [0249] The structural lipid may comprise from about 30 mol % to about 50 mol %, from about 30 mol % to about 45 mol %, from about 30 mol % to about 40 mol %, from about 30 mol % to about 35 mol %, from about 35 mol % to about 50 mol %, from about 40 mol % to about 50 mol %, or from about 45 mol % to about 50 mol % of the total lipid present in the particle. [0250] Examples of lipid nanoparticle compositions and methods of making same are described, for example, in Semple et al. (2010) Nat. Biotechnol.28:172-176; Jayarama et al. (2012), Angew. Chem. Int. Ed., 51: 8529-8533; and Maier et al. (2013) Molecular Therapy 21, 1570-1578 (the contents of each of which are incorporated herein by reference in their entirety). [0251] In some embodiments, a lipid nanoparticle formulation includes 0.5% to 15% on a molar basis of the neutral lipid, e.g., 3 to 12%, 5 to 10% or 15%, 10%, or 7.5% on a molar basis. Examples of neutral lipids include, without limitation, DSPC, POPC, DPPC, DOPE and SM. In some embodiments, the formulation includes 5% to 50% on a molar basis of the sterol (e.g., 15 to 45%, 20 to 40%, 40%, 38.5%, 35%, or 31% on a molar basis. A non-limiting example of a sterol is cholesterol. In some embodiments, a lipid nanoparticle formulation includes 0.5% to 20% on a molar basis of the PEG or PEG- modified lipid (e.g., 0.5 to 10%, 0.5 to 5%, 1.5%, 0.5%, 1.5%, 3.5%, or 5% on a molar basis. In some embodiments, a PEG or PEG modified lipid comprises a PEG molecule of
1005272698
78 an average molecular weight of 2,000 Da. In some embodiments, a PEG or PEG modified lipid comprises a PEG molecule of an average molecular weight of less than 2,000, for example around 1,500 Da, around 1,000 Da, or around 500 Da. Non-limiting examples of PEG-modified lipids include PEG-distearoyl glycerol (PEG-DMG) (also referred herein as PEG-C14 or C14-PEG), PEG-cDMA (further discussed in Reyes et al. J. Controlled Release, 107, 276-287 (2005) the contents of which are herein incorporated by reference in their entirety). [0252] In any embodiment, the PEGylated lipid may comprise about 0.05 mol %, about 0.1 mol %, about 0.15 mol %, about 0.2 mol %, about 0.25 mol %, about 0.3 mol %, about 0.35 mol %, about 0.4 mol %, about 0.45 mol %, about 0.5% mol, about 0.6% mol, about 0.7% mol, about 0.8% mol, about 1 % mol, about 1.2% mol, about 1,4 % mol, about 1.6% mol, about 1.8 % mol, or about 2 % mol or more of the total lipid present in the particle. [0253] The PEGylated lipid may be selected from the group consisting of PEG-modified phosphatidylethanolamines, PEG-modified phosphatidic acids, PEG-modified ceramides, PEG-modified dialkylamines, PEG-modified diacylglycerols, PEG-modified dialkylglycerols, and mixtures thereof. [0254] The PEGylated lipid may be selected from the group consisting of PEG-c- DOMG, PEG-DMG, PEG-DLPE, PEG-DMPE, PEG-DPPC, or a PEG-DSPE lipid. [0255] The PEGylated lipid may have a PEG component that has a molecular weight between about 100 Da and about 100,000 Dam between about 100 Da and about 100,000 Da, between about 1000 Da and 9,000 Da, between about 1000 Da and 8,000 Da, between about 1000 Da and 7,000 Da, between about 1000 Da and 6,000 Da, between about 1000 Da and 5,000 Da, between about 1000 Da and 4,000 Da, between about 1000 Da and 3,000 Da, or between about 1000 Da and 2,000 Da. [0256] The PEGylated lipid may be DSPE-PEG, wherein the PEG has a molecular weight of 2000 Da. [0257] In some embodiments, the pharmaceutical compositions of the RNA (e.g., mRNA) vaccines may include at least one of the PEGylated lipids described in International Publication No. WO2012099755, the contents of which are herein incorporated by reference in their entirety.
1005272698
79 [0258] The PEGylated lipid may be ALC-0159 (2-[(polyethylene glycol)-2000]-N,N- ditetradecylacetamide). ALC-0159 is a PEG/lipid conjugate (i.e. PEGylated lipid), specifically, it is the N,N-dimyristylamide of 2-hydroxyacetic acid, O-pegylated to a PEG chain mass of about 2 kilodaltons (corresponding to about 45-46 ethylene oxide units per molecule of N,N-dimyristyl hydroxyacetamide). It is a non-ionic surfactant by its nature. [0259] Non-limiting examples of lipid nanoparticle compositions and methods of making them are described, for example, in Semple et al. (2010) Nat. Biotechnol.28:172- 176; Jayarama et al. (2012), Angew. Chem. Int. Ed., 51: 8529-8533; and Maier et al. (2013) Molecular Therapy 21, 1570-1578 (the contents of each of which are incorporated herein by reference in their entirety). [0260] Relative amounts of the active ingredient, the pharmaceutically acceptable excipient, and/or any additional ingredients in a vaccine composition may vary, depending upon the identity, size, and/or condition of the subject being treated and further depending upon the route by which the composition is to be administered. For example, the composition may comprise between 0.1% and 99% (w/w) of the active ingredient. By way of example, the composition may comprise between 0.1% and 100%, e.g., between 0.5 and 50%, between 1-30%, between 5-80%, at least 80% (w/w) active ingredient. [0261] In some embodiments, the RNA vaccine composition of the invention may comprise a polynucleotide described herein, formulated in a lipid nanoparticle comprising ALC—0315 ([(4-hydroxybutyl)azanediyl]di(hexane-6,1-diyl) bis(2-hexyldecanoate)) Cholesterol, DSPC and ALC-0159 (2-[(polyethylene glycol)-2000]-N,N- ditetradecylacetamide), the buffer Tris-sucrose and water for injection. [0262] As a non-limiting example, the composition comprises: 0.6 mg/mL of drug substance (e.g., polynucleotides encoding a chimeric or fusion protein described herein and comprising components of a P. gingivalis gingipain polyprotein complex), 8.58 mg/mL of ALC-0315, 3.99 mg/mL of cholesterol, 1.80 mg/mL of DSPC, 0.95 mg/mL of ALC-0159 (2-[(polyethylene glycol)-2000]-N,N-ditetradecylacetamide), 3.03 mg/mL of Tris (trishydroxymethyl)aminomethoane), 88 mg/mL of sucrose in water, with a typical volume for injection of 50 µL.
1005272698
80 [0263] In alternative embodiments, the RNA vaccine composition of the invention may comprise the four lipids DLin-MC3-DMA, Cholesterol, DSPC and DMG-PEG 2000 at a ratio of 50:39.8:10:0.2 (mol ratio). [0264] In some embodiments, a nanoparticle (e.g., a lipid nanoparticle) has a mean diameter of 10-500 nm, 20-400 nm, 30-300 nm, 40-200 nm. In some embodiments, a nanoparticle (e.g., a lipid nanoparticle) has a mean diameter of 50-150 nm, 50-200 nm, 80-100 nm or 80-200 nm. [0265] In some embodiments, the RNA (e.g., RNA) vaccines of the disclosure may be formulated in lipid nanoparticles having a diameter from about 10 to about 100 nm such as, but not limited to, about 10 to about 20 nm, about 10 to about 30 nm, about 10 to about 40 nm, about 10 to about 50 nm, about 10 to about 60 nm, about 10 to about 70 nm, about 10 to about 80 nm, about 10 to about 90 nm, about 20 to about 30 nm, about 20 to about 40 nm, about 20 to about 50 nm, about 20 to about 60 nm, about 20 to about 70 nm, about 20 to about 80 nm, about 20 to about 90 nm, about 20 to about 100 nm, about 30 to about 40 nm, about 30 to about 50 nm, about 30 to about 60 nm, about 30 to about 70 nm, about 30 to about 80 nm, about 30 to about 90 nm, about 30 to about 100 nm, about 40 to about 50 nm, about 40 to about 60 nm, about 40 to about 70 nm, about 40 to about 80 nm, about 40 to about 90 nm, about 40 to about 100 nm, about 50 to about 60 nm, about 50 to about 70 nm about 50 to about 80 nm, about 50 to about 90 nm, about 50 to about 100 nm, about 60 to about 70 nm, about 60 to about 80 nm, about 60 to about 90 nm, about 60 to about 100 nm, about 70 to about 80 nm, about 70 to about 90 nm, about 70 to about 100 nm, about 80 to about 90 nm, about 80 to about 100 nm and/or about 90 to about 100 nm. [0266] In some embodiments, the lipid nanoparticles may have a diameter from about 10 to 500 nm. [0267] In some embodiments, the lipid nanoparticle may have a diameter greater than 100 nm, greater than 150 nm, greater than 200 nm, greater than 250 nm, greater than 300 nm, greater than 350 nm, greater than 400 nm, greater than 450 nm, greater than 500 nm, greater than 550 nm, greater than 600 nm, greater than 650 nm, greater than 700 nm, greater than 750 nm, greater than 800 nm, greater than 850 nm, greater than 900 nm, greater than 950 nm or greater than 1000 nm.
1005272698
81 [0268] In some embodiments, the lipid nanoparticle may be a limit size lipid nanoparticle described in International Patent Publication No. WO2013059922, the contents of which are herein incorporated by reference in their entirety. The limit size lipid nanoparticle may comprise a lipid bilayer surrounding an aqueous core or a hydrophobic core; where the lipid bilayer may comprise a phospholipid such as, but not limited to, diacylphosphatidylcholine, a diacylphosphatidylethanolamine, a ceramide, a sphingomyelin, a dihydrosphingomyelin, a cephalin, a cerebroside, a C8-C20 fatty acid diacylphophatidylcholine, and 1-palmitoyl-2-oleoyl phosphatidylcholine (POPC). In some embodiments, the limit size lipid nanoparticle may comprise a polyethylene glycol-lipid such as, but not limited to, DLPE-PEG, DMPE-PEG, DPPC-PEG and DSPE-PEG. [0269] In some embodiments the RNA (e.g., RNA) vaccine may be associated with a cationic or polycationic compounds, including protamine, nucleoline, spermine or spermidine, or other cationic peptides or proteins, such as poly-L-lysine (PLL), polyarginine, basic polypeptides, cell penetrating peptides (CPPs), including HIV-binding peptides, HIV-1 Tat (HIV), Tat-derived peptides, Penetratin, VP derived or analog peptides, Pestivirus Erns, HSV, VP (Herpes simplex), MAP, KALA or protein transduction domains (PTDs), PpT620, prolin-rich peptides, arginine-rich peptides, lysine-rich peptides, MPG-peptide(s), Pep-1, L-oligomers, Calcitonin peptide(s), Antennapedia- derived peptides (particularly from Drosophila antennapedia), pAntp, pIsl, FGF, Lactoferrin, Transportan, Buforin-2, Bac715-24, SynB, SynB(1), pVEC, hCT-derived peptides, SAP, histones, cationic polysaccharides, for example chitosan, polybrene, cationic polymers, e.g. polyethyleneimine (PEI), cationic lipids, e.g. DOTMA: [1-(2,3- sioleyloxy)propyl)]-N,N,N-trimethylammonium chloride, DMRIE, di-C14-amidine, DOTIM, SAINT, DC-Chol, BGTC, CTAP, DOPC, DODAP, DOPE: Dioleyl phosphatidylethanol- amine, DOSPA, DODAB, DOIC, DMEPC, DOGS: Dioctadecylamidoglicylspermin, DIMRI: Dimyristooxypropyl dimethyl hydroxyethyl ammonium bromide, DOTAP: dioleoyloxy-3-(trimethylammonio)propane, DC-6-14: O,O-ditetradecanoyl-N-.alpha.- trimethylammonioacetyl)diethanolamine chloride, CLIP 1: rac-[(2,3- dioctadecyloxypropyl)(2-hydroxyethyl)]-dimethylammonium chloride, CLIP6: rac-[2(2,3- dihexadecyloxypropyloxymethyloxy)ethyl]-trimethylammonium, CLIP9: rac-[2(2,3- dihexadecyloxypropyloxysuccinyloxy)ethyl]-trimethylammonium, oligofectamine, or cationic or polycationic polymers, e.g. modified polyaminoacids, such as beta-aminoacid- polymers or reversed polyamides, etc., modified polyethylenes, such as PVP (poly(N- ethyl-4-vinylpyridinium bromide)), etc., modified acrylates, such as pDMAEMA
1005272698
82 (poly(dimethylaminoethyl methylacrylate)), etc., modified amidoamines such as pAMAM (poly(amidoamine)), etc., modified polybetaminoester (PBAE), such as diamine end modified 1,4 butanediol diacrylate-co-5-amino-1-pentanol polymers, etc., dendrimers, such as polypropylamine dendrimers or pAMAM based dendrimers, etc., polyimine(s), such as PEI: poly(ethyleneimine), poly(propyleneimine), etc., polyallylamine, sugar backbone based polymers, such as cyclodextrin based polymers, dextran based polymers, chitosan, etc., silan backbone based polymers, such as PMOXA-PDMS copolymers, etc., blockpolymers consisting of a combination of one or more cationic blocks (e.g. selected from a cationic polymer as mentioned above) and of one or more hydrophilic or hydrophobic blocks (e.g. polyethyleneglycole), etc. [0270] In other embodiments the RNA (e.g., RNA) vaccine is not associated with a cationic or polycationic compounds. [0271] Other examples of suitable lipid nanoparticle formulations are provided in US 10,702,600, the contents of which are hereby incorporated by reference. [0272] The lipid nanoparticles described herein may be made in a sterile environment. [0273] The nanoparticle formulations may comprise a phosphate conjugate. The phosphate conjugate may increase in vivo circulation times and/or increase the targeted delivery of the nanoparticle. As a non-limiting example, the phosphate conjugates may include a compound of any one of the formulas described in International Application No. WO2013033438, the contents of which are herein incorporated by reference in its entirety. [0274] The nanoparticle formulation may comprise a polymer conjugate. The polymer conjugate may be a water soluble conjugate. The polymer conjugate may have a structure as described in U.S. Patent Application No. 20130059360, the contents of which are herein incorporated by reference in its entirety. In some embodiments, polymer conjugates with the polynucleotides of the present disclosure may be made using the methods and/or segmented polymeric reagents described in U.S. Patent Application No. 20130072709, the contents of which are herein incorporated by reference in its entirety. In some embodiments, the polymer conjugate may have pendant side groups comprising ring moieties such as, but not limited to, the polymer conjugates described in U.S. Patent
1005272698
83 Publication No. US20130196948, the contents which are herein incorporated by reference in its entirety. [0275] The nanoparticle formulations may comprise a conjugate to enhance the delivery of nanoparticles of the present disclosure in a subject. Further, the conjugate may inhibit phagocytic clearance of the nanoparticles in a subject. In one embodiment, the conjugate may be a “self” peptide designed from the human membrane protein CD47 (e.g., the “self” particles described by Rodriguez et al. (Science 2013339, 971-975), herein incorporated by reference in its entirety). As shown by Rodriguez et al., the self peptides delayed macrophage-mediated clearance of nanoparticles which enhanced delivery of the nanoparticles. In another embodiment, the conjugate may be the membrane protein CD47 (e.g., see Rodriguez et al. Science 2013339, 971-975, herein incorporated by reference in its entirety). Rodriguez et al. showed that, similarly to “self” peptides, CD47 can increase the circulating particle ratio in a subject as compared to scrambled peptides and PEG coated nanoparticles. [0276] In some embodiments, the RNA (e.g., RNA) vaccines of the present disclosure are formulated in nanoparticles which comprise a conjugate to enhance the delivery of the nanoparticles of the present disclosure in a subject. The conjugate may be the CD47 membrane or the conjugate may be derived from the CD47 membrane protein, such as the “self” peptide described previously. In some embodiments, the nanoparticle may comprise PEG and a conjugate of CD47 or a derivative thereof. In some embodiments, the nanoparticle may comprise both the “self” peptide described above and the membrane protein CD47. [0277] In some embodiments, RNA (e.g., RNA) vaccine pharmaceutical compositions comprising the polynucleotides of the present disclosure and a conjugate that may have a degradable linkage. Non-limiting examples of conjugates include an aromatic moiety comprising an ionizable hydrogen atom, a spacer moiety, and a water-soluble polymer. As a non-limiting example, pharmaceutical compositions comprising a conjugate with a degradable linkage and methods for delivering such pharmaceutical compositions are described in U.S. Patent Publication No. US20130184443, the contents of which are herein incorporated by reference in their entirety. [0278] The nanoparticle formulations may be a carbohydrate nanoparticle comprising a carbohydrate carrier and a RNA (e.g., RNA) vaccine. As a non-limiting example, the
1005272698
84 carbohydrate carrier may include, but is not limited to, an anhydride-modified phytoglycogen or glycogen-type material, phtoglycogen octenyl succinate, phytoglycogen beta-dextrin, anhydride-modified phytoglycogen beta-dextrin. (See e.g., International Publication No. WO2012109121; the contents of which are herein incorporated by reference in their entirety). [0279] Nanoparticle formulations of the present disclosure may be coated with a surfactant or polymer in order to improve the delivery of the particle. In some embodiments, the nanoparticle may be coated with a hydrophilic coating such as, but not limited to, PEG coatings and/or coatings that have a neutral surface charge. The hydrophilic coatings may help to deliver nanoparticles with larger payloads such as, but not limited to, RNA (e.g., RNA) vaccines within the central nervous system. As a non- limiting example nanoparticles comprising a hydrophilic coating and methods of making such nanoparticles are described in U.S. Patent Publication No. US20130183244, the contents of which are herein incorporated by reference in their entirety. [0280] In some embodiments, the lipid nanoparticles of the present disclosure may be hydrophilic polymer particles. Non-limiting examples of hydrophilic polymer particles and methods of making hydrophilic polymer particles are described in U.S. Patent Publication No. US20130210991, the contents of which are herein incorporated by reference in their entirety. [0281] In some embodiments, the lipid nanoparticles of the present disclosure may be hydrophobic polymer particles. [0282] Lipid nanoparticle formulations may be improved by replacing the cationic lipid with a biodegradable cationic lipid which is known as a rapidly eliminated lipid nanoparticle (reLNP). Ionizable cationic lipids, such as, but not limited to, DLinDMA, DLin- KC2-DMA, and DLin-MC3-DMA, have been shown to accumulate in plasma and tissues over time and may be a potential source of toxicity. The rapid metabolism of the rapidly eliminated lipids can improve the tolerability and therapeutic index of the lipid nanoparticles by an order of magnitude from a 1 mg/kg dose to a 10 mg/kg dose in rat. Inclusion of an enzymatically degraded ester linkage can improve the degradation and metabolism profile of the cationic component, while still maintaining the activity of the reLNP formulation. The ester linkage can be internally located within the lipid chain or it
1005272698
85 may be terminally located at the terminal end of the lipid chain. The internal ester linkage may replace any carbon in the lipid chain. [0283] In some embodiments, the internal ester linkage may be located on either side of the saturated carbon. [0284] In some embodiments, an immune response may be elicited by delivering a lipid nanoparticle which may include a nanospecies, a polymer and an immunogen. (U.S. Publication No.20120189700 and International Publication No. WO2012099805; each of which is herein incorporated by reference in their entirety). The polymer may encapsulate the nanospecies or partially encapsulate the nanospecies. The immunogen may be a recombinant protein, a modified RNA and/or a polynucleotide described herein. In some embodiments, the lipid nanoparticle may be formulated for use in a vaccine such as, but not limited to, against a pathogen. [0285] Lipid nanoparticles may be engineered to alter the surface properties of particles so the lipid nanoparticles may penetrate the mucosal barrier. Mucus is located on mucosal tissue such as, but not limited to, oral (e.g., the buccal and esophageal membranes and tonsil tissue), ophthalmic, gastrointestinal (e.g., stomach, small intestine, large intestine, colon, rectum), nasal, respiratory (e.g., nasal, pharyngeal, tracheal and bronchial membranes), genital (e.g., vaginal, cervical and urethral membranes). Nanoparticles larger than 10-200 nm which are preferred for higher drug encapsulation efficiency and the ability to provide the sustained delivery of a wide array of drugs have been thought to be too large to rapidly diffuse through mucosal barriers. Mucus is continuously secreted, shed, discarded or digested and recycled so most of the trapped particles may be removed from the mucosa tissue within seconds or within a few hours. Large polymeric nanoparticles (200 nm-500 nm in diameter) which have been coated densely with a low molecular weight polyethylene glycol (PEG) diffused through mucus only 4 to 6-fold lower than the same particles diffusing in water (Lai et al. PNAS 2007 104(5):1482-487; Lai et al. Adv Drug Deliv Rev.200961(2): 158-171; each of which is herein incorporated by reference in their entirety). The transport of nanoparticles may be determined using rates of permeation and/or fluorescent microscopy techniques including, but not limited to, fluorescence recovery after photobleaching (FRAP) and high resolution multiple particle tracking (MPT). As a non-limiting example, compositions which can penetrate a mucosal barrier may be made as described in U.S. Pat. No.8,241,670
1005272698
86 or International Patent Publication No. WO2013110028, the contents of each of which are herein incorporated by reference in its entirety. [0286] The lipid nanoparticle engineered to penetrate mucus may comprise a polymeric material (i.e. a polymeric core) and/or a polymer-vitamin conjugate and/or a tri-block co- polymer. The polymeric material may include, but is not limited to, polyamines, polyethers, polyamides, polyesters, polycarbamates, polyureas, polycarbonates, poly(styrenes), polyimides, polysulfones, polyurethanes, polyacetylenes, polyethylenes, polyethyeneimines, polyisocyanates, polyacrylates, polymethacrylates, polyacrylonitriles, and polyarylates. The polymeric material may be biodegradable and/or biocompatible. Non-limiting examples of biocompatible polymers are described in International Patent Publication No. WO2013116804, the contents of which are herein incorporated by reference in their entirety. The polymeric material may additionally be irradiated. As a non-limiting example, the polymeric material may be gamma irradiated (see e.g., International App. No. WO201282165, herein incorporated by reference in its entirety). Non-limiting examples of specific polymers include poly(caprolactone) (PCL), ethylene vinyl acetate polymer (EVA), poly(lactic acid) (PLA), poly(L-lactic acid) (PLLA), poly(glycolic acid) (PGA), poly(lactic acid-co-glycolic acid) (PLGA), poly(L-lactic acid-co- glycolic acid) (PLLGA), poly(D,L-lactide) (PDLA), poly(L-lactide) (PLLA), poly(D,L-lactide- co-caprolactone), poly(D,L-lactide-co-caprolactone-co-glycolide), poly(D,L-lactide-co- PEO-co-D,L-lactide), poly(D,L-lactide-co-PPO-co-D,L-lactide), polyalkyl cyanoacralate, polyurethane, poly-L-lysine (PLL), hydroxypropyl methacrylate (HPMA), polyethyleneglycol, poly-L-glutamic acid, poly(hydroxy acids), polyanhydrides, polyorthoesters, poly(ester amides), polyamides, poly(ester ethers), polycarbonates, polyalkylenes such as polyethylene and polypropylene, polyalkylene glycols such as poly(ethylene glycol) (PEG), polyalkylene oxides (PEO), polyalkylene terephthalates such as poly(ethylene terephthalate), polyvinyl alcohols (PVA), polyvinyl ethers, polyvinyl esters such as poly(vinyl acetate), polyvinyl halides such as poly(vinyl chloride) (PVC), polyvinylpyrrolidone, polysiloxanes, polystyrene (PS), polyurethanes, derivatized celluloses such as alkyl celluloses, hydroxyalkyl celluloses, cellulose ethers, cellulose esters, nitro celluloses, hydroxypropylcellulose, carboxymethylcellulose, polymers of acrylic acids, such as poly(methyl(meth)acrylate) (PMMA), poly(ethyl(meth)acrylate), poly(butyl(meth)acrylate), poly(isobutyl(meth)acrylate), poly(hexyl(meth)acrylate), poly(isodecyl(meth)acrylate), poly(lauryl(meth)acrylate), poly(phenyl(meth)acrylate), poly(methyl acrylate), poly(isopropyl acrylate), poly(isobutyl acrylate), poly(octadecyl
1005272698
87 acrylate) and copolymers and mixtures thereof, polydioxanone and its copolymers, polyhydroxyalkanoates, polypropylene fumarate, polyoxymethylene, poloxamers, poly(ortho)esters, poly(butyric acid), poly(valeric acid), poly(lactide-co-caprolactone), PEG-PLGA-PEG and trimethylene carbonate, polyvinylpyrrolidone. The lipid nanoparticle may be coated or associated with a co-polymer such as, but not limited to, a block co- polymer (such as a branched polyether-polyamide block copolymer described in International Publication No. WO2013012476, herein incorporated by reference in its entirety), and (poly(ethylene glycol))-(poly(propylene oxide))-(poly(ethylene glycol)) triblock copolymer (see e.g., U.S. Publication 20120121718 and U.S. Publication 20100003337 and U.S. Pat. No. 8,263,665, the contents of each of which is herein incorporated by reference in their entirety). The co-polymer may be a polymer that is generally regarded as safe (GRAS) and the formation of the lipid nanoparticle may be in such a way that no new chemical entities are created. For example, the lipid nanoparticle may comprise poloxamers coating PLGA nanoparticles without forming new chemical entities which are still able to rapidly penetrate human mucus (Yang et al. Angew. Chem. Int. Ed.201150:2597-2600; the contents of which are herein incorporated by reference in their entirety). A non-limiting scalable method to produce nanoparticles which can penetrate human mucus is described by Xu et al. (see, e.g., J Control Release 2013, 170(2):279-86; the contents of which are herein incorporated by reference in their entirety). [0287] The vitamin of the polymer-vitamin conjugate may be vitamin E. The vitamin portion of the conjugate may be substituted with other suitable components such as, but not limited to, vitamin A, vitamin E, other vitamins, cholesterol, a hydrophobic moiety, or a hydrophobic component of other surfactants (e.g., sterol chains, fatty acids, hydrocarbon chains and alkylene oxide chains). [0288] The lipid nanoparticle engineered to penetrate mucus may include surface altering agents such as, but not limited to, polynucleotides, anionic proteins (e.g., bovine serum albumin), surfactants (e.g., cationic surfactants such as for example dimethyldioctadecyl-ammonium bromide), sugars or sugar derivatives (e.g., cyclodextrin), nucleic acids, polymers (e.g., heparin, polyethylene glycol and poloxamer), mucolytic agents (e.g., N-acetylcysteine, mugwort, bromelain, papain, clerodendrum, acetylcysteine, bromhexine, carbocisteine, eprazinone, mesna, ambroxol, sobrerol, domiodol, letosteine, stepronin, tiopronin, gelsolin, thymosin 34 dornase alfa, neltenexine,
1005272698
88 erdosteine) and various DNases including rhDNase. The surface altering agent may be embedded or enmeshed in the particle's surface or disposed (e.g., by coating, adsorption, covalent linkage, or other process) on the surface of the lipid nanoparticle. (see e.g., U.S. Publication 20100215580 and U.S. Publication 20080166414 and US20130164343; the contents of each of which are herein incorporated by reference in their entirety). [0289] In some embodiments, the mucus penetrating lipid nanoparticles may comprise at least one polynucleotide described herein. The polynucleotide may be encapsulated in the lipid nanoparticle and/or disposed on the surface of the particle. The polynucleotide may be covalently coupled to the lipid nanoparticle. Formulations of mucus penetrating lipid nanoparticles may comprise a plurality of nanoparticles. Further, the formulations may contain particles which may interact with the mucus and alter the structural and/or adhesive properties of the surrounding mucus to decrease mucoadhesion, which may increase the delivery of the mucus penetrating lipid nanoparticles to the mucosal tissue. [0290] In some embodiments, the mucus penetrating lipid nanoparticles may be a hypotonic formulation comprising a mucosal penetration enhancing coating. The formulation may be hypotonic for the epithelium to which it is being delivered. Non-limiting examples of hypotonic formulations may be found in International Patent Publication No. WO2013110028, the contents of which are herein incorporated by reference in their entirety. [0291] In some embodiments, in order to enhance the delivery through the mucosal barrier the RNA (e.g., mRNA) vaccine formulation may comprise or be a hypotonic solution. [0292] Hypotonic solutions were found to increase the rate at which mucoinert particles such as, but not limited to, mucus-penetrating particles, were able to reach the vaginal epithelial surface (see e.g., Ensign et al. Biomaterials 201334(28):6922-9, the contents of which are herein incorporated by reference in their entirety). [0293] In some embodiments, the RNA (e.g., mRNA) vaccine is formulated as a lipoplex, such as, without limitation, the ATUPLEX™ system, the DACC system, the DBTC system and other siRNA-lipoplex technology from Silence Therapeutics (London, United Kingdom), STEMFECT™ from STEMGENT® (Cambridge, Mass.), and polyethylenimine (PEI) or protamine-based targeted and non-targeted delivery of nucleic
1005272698
89 acids acids (Aleku et al. Cancer Res. 200868:9788-9798; Strumberg et al. Int J Clin Pharmacol Ther 201250:76-78; Santel et al., Gene Ther 200613:1222-1234; Santel et al., Gene Ther 200613:1360-1370; Gutbier et al., Pulm Pharmacol. Ther.201023:334- 344; Kaufmann et al. Microvasc Res 201080:286-293Weide et al. J Immunother.2009 32:498-507; Weide et al. J Immunother. 200831:180-188; Pascolo Expert Opin. Biol. Ther.4:1285-1294; Fotin-Mleczek et al., 2011 J. Immunother.34:1-15; Song et al., Nature Biotechnol.2005, 23:709-717; Peer et al., Proc Natl Acad Sci USA.20076; 104:4095- 4100; deFougerolles Hum Gene Ther.200819:125-132, the contents of each of which are incorporated herein by reference in their entirety). [0294] In some embodiments, the RNA (e.g., mRNA) vaccine is formulated as a solid lipid nanoparticle. A solid lipid nanoparticle (SLN) may be spherical with an average diameter between 10 to 1000 nm. SLN possess a solid lipid core matrix that can solubilize lipophilic molecules and may be stabilized with surfactants and/or emulsifiers. In some embodiments, the lipid nanoparticle may be a self-assembly lipid-polymer nanoparticle (see Zhang et al., ACS Nano, 2008, 2 (8), pp 1696-1702; the contents of which are herein incorporated by reference in their entirety). As a non-limiting example, the SLN may be the SLN described in International Patent Publication No. WO2013105101, the contents of which are herein incorporated by reference in their entirety. As another non-limiting example, the SLN may be made by the methods or processes described in International Patent Publication No. WO2013105101, the contents of which are herein incorporated by reference in their entirety. [0295] In some embodiments, the RNA (e.g., mRNA) vaccines of the present disclosure can be formulated for controlled release and/or targeted delivery. As used herein, “controlled release” refers to a pharmaceutical composition or compound release profile that conforms to a particular pattern of release to effect a therapeutic outcome. In some embodiments, the RNA (e.g., mRNA) vaccines may be encapsulated into a delivery agent described herein and/or known in the art for controlled release and/or targeted delivery. As used herein, the term “encapsulate” means to enclose, surround or encase. As it relates to the formulation of the compounds of the disclosure, encapsulation may be substantial, complete or partial. The term “substantially encapsulated” means that at least greater than 50, 60, 70, 80, 85, 90, 95, 96, 97, 98, 99, 99.9, 99.9 or greater than 99.999% of the pharmaceutical composition or compound of the disclosure may be enclosed, surrounded or encased within the delivery agent. “Partially encapsulated” means that less
1005272698
90 than 10, 10, 20, 30, 4050 or less of the pharmaceutical composition or compound of the disclosure may be enclosed, surrounded or encased within the delivery agent. Advantageously, encapsulation may be determined by measuring the escape or the activity of the pharmaceutical composition or compound of the disclosure using fluorescence and/or electron micrograph. For example, at least 1, 5, 10, 20, 30, 40, 50, 60, 70, 80, 85, 90, 95, 96, 97, 98, 99, 99.9, 99.99 or greater than 99.99% of the pharmaceutical composition or compound of the disclosure are encapsulated in the delivery agent. [0296] In some embodiments, the controlled release formulation may include, but is not limited to, tri-block co-polymers. As a non-limiting example, the formulation may include two different types of tri-block co-polymers (International Pub. No. WO2012131104 and WO2012131106, the contents of each of which are incorporated herein by reference in their entirety). [0297] In some embodiments, the RNA (e.g., mRNA) vaccines may be encapsulated into a lipid nanoparticle or a rapidly eliminated lipid nanoparticle and the lipid nanoparticles or a rapidly eliminated lipid nanoparticle may then be encapsulated into a polymer, hydrogel and/or surgical sealant described herein and/or known in the art. As a non-limiting example, the polymer, hydrogel or surgical sealant may be PLGA, ethylene vinyl acetate (EVAc), poloxamer, GELSITE® (Nanotherapeutics, Inc. Alachua, Fla.), HYLENEX® (Halozyme Therapeutics, San Diego Calif.), surgical sealants such as fibrinogen polymers (Ethicon Inc. Cornelia, Ga.), TISSELL® (Baxter International, Inc Deerfield, Ill.), PEG-based sealants, and COSEAL® (Baxter International, Inc Deerfield, Ill.). [0298] In some embodiments, the lipid nanoparticle may be encapsulated into any polymer known in the art which may form a gel when injected into a subject. As another non-limiting example, the lipid nanoparticle may be encapsulated into a polymer matrix which may be biodegradable. [0299] In some embodiments, the RNA (e.g., mRNA) vaccine formulation for controlled release and/or targeted delivery may also include at least one controlled release coating. Controlled release coatings include, but are not limited to, OPADRY®, polyvinylpyrrolidone/vinyl acetate copolymer, polyvinylpyrrolidone, hydroxypropyl methylcellulose, hydroxypropyl cellulose, hydroxyethyl cellulose, EUDRAGIT RL®,
1005272698
91 EUDRAGIT RS® and cellulose derivatives such as ethylcellulose aqueous dispersions (AQUACOAT® and SURELEASE®). [0300] In some embodiments, the RNA (e.g., mRNA) vaccine controlled release and/or targeted delivery formulation may comprise at least one degradable polyester which may contain polycationic side chains. Degradeable polyesters include, but are not limited to, poly(serine ester), poly(L-lactide-co-L-lysine), poly(4-hydroxy-L-proline ester), and combinations thereof. In some embodiments, the degradable polyesters may include a PEG conjugation to form a PEGylated polymer. [0301] In some embodiments, the RNA (e.g., mRNA) vaccine controlled release and/or targeted delivery formulation comprising at least one polynucleotide may comprise at least one PEG and/or PEG related polymer derivatives as described in U.S. Pat. No. 8,404,222, the contents of which are incorporated herein by reference in their entirety. [0302] In some embodiments, the RNA (e.g., mRNA) vaccine controlled release delivery formulation comprising at least one polynucleotide may be the controlled release polymer system described in US20130130348, the contents of which are incorporated herein by reference in their entirety. [0303] In some embodiments, the RNA (e.g., mRNA) vaccines of the present disclosure may be encapsulated in a therapeutic nanoparticle, referred to herein as “therapeutic nanoparticle RNA (e.g., mRNA) vaccines.” Therapeutic nanoparticles may be formulated by methods described herein and known in the art such as, but not limited to, International Pub Nos. WO2010005740, WO2010030763, WO2010005721, WO2010005723, WO2012054923, U.S. Publication Nos. US20110262491, US20100104645, US20100087337, US20100068285, US20110274759, US20100068286, US20120288541, US20130123351 and US20130230567 and U.S. Pat. Nos.8,206,747, 8,293,276, 8,318,208 and 8,318,211; the contents of each of which are herein incorporated by reference in their entirety. In some embodiments, therapeutic polymer nanoparticles may be identified by the methods described in US Pub No. US20120140790, the contents of which are herein incorporated by reference in their entirety. [0304] In some embodiments, the therapeutic nanoparticle RNA (e.g., mRNA) vaccine may be formulated for sustained release. As used herein, “sustained release” refers to a
1005272698
92 pharmaceutical composition or compound that conforms to a release rate over a specific period of time. The period of time may include, but is not limited to, hours, days, weeks, months and years. As a non-limiting example, the sustained release nanoparticle may comprise a polymer and a therapeutic agent such as, but not limited to, the polynucleotides of the present disclosure (see International Pub No.2010075072 and US Pub No. US20100216804, US20110217377 and US20120201859, the contents of each of which are incorporated herein by reference in their entirety). In another non-limiting example, the sustained release formulation may comprise agents which permit persistent bioavailability such as, but not limited to, crystals, macromolecular gels and/or particulate suspensions (see U.S. Patent Publication No US20130150295, the contents of each of which are incorporated herein by reference in their entirety). [0305] In some embodiments, the therapeutic nanoparticle RNA (e.g., mRNA) vaccines may be formulated to be target specific. As a non-limiting example, the therapeutic nanoparticles may include a corticosteroid (see International Pub. No. WO2011084518, the contents of which are incorporated herein by reference in their entirety). As a non- limiting example, the therapeutic nanoparticles may be formulated in nanoparticles described in International Pub No. WO2008121949, WO2010005726, WO2010005725, WO2011084521 and US Pub No. US20100069426, US20120004293 and US20100104655, the contents of each of which are incorporated herein by reference in their entirety. [0306] In some embodiments, the nanoparticles of the present disclosure may comprise a polymeric matrix. As a non-limiting example, the nanoparticle may comprise two or more polymers such as, but not limited to, polyethylenes, polycarbonates, polyanhydrides, polyhydroxyacids, polypropylfumerates, polycaprolactones, polyamides, polyacetals, polyethers, polyesters, poly(orthoesters), polycyanoacrylates, polyvinyl alcohols, polyurethanes, polyphosphazenes, polyacrylates, polymethacrylates, polycyanoacrylates, polyureas, polystyrenes, polyamines, polylysine, poly(ethylene imine), poly(serine ester), poly(L-lactide-co-L-lysine), poly(4-hydroxy-L-proline ester) or combinations thereof. [0307] In some embodiments, the therapeutic nanoparticle comprises a diblock copolymer. In some embodiments, the diblock copolymer may include PEG in combination with a polymer such as, but not limited to, polyethylenes, polycarbonates, polyanhydrides, polyhydroxyacids, polypropylfumerates, polycaprolactones, polyamides,
1005272698
93 polyacetals, polyethers, polyesters, poly(orthoesters), polycyanoacrylates, polyvinyl alcohols, polyurethanes, polyphosphazenes, polyacrylates, polymethacrylates, polycyanoacrylates, polyureas, polystyrenes, polyamines, polylysine, poly(ethylene imine), poly(serine ester), poly(L-lactide-co-L-lysine), poly(4-hydroxy-L-proline ester) or combinations thereof. In yet another embodiment, the diblock copolymer may be a high- X diblock copolymer such as those described in International Patent Publication No. WO2013120052, the contents of which are incorporated herein by reference in their entirety. [0308] As a non-limiting example the therapeutic nanoparticle comprises a PLGA-PEG block copolymer (see U.S. Publication No. US20120004293 and U.S. Pat. No.8,236,330, each of which is herein incorporated by reference in their entirety). In another non-limiting example, the therapeutic nanoparticle is a stealth nanoparticle comprising a diblock copolymer of PEG and PLA or PEG and PLGA (see U.S. Pat. No. 8,246,968 and International Publication No. WO2012166923, the contents of each of which are herein incorporated by reference in their entirety). In yet another non-limiting example, the therapeutic nanoparticle is a stealth nanoparticle or a target-specific stealth nanoparticle as described in U.S. Patent Publication No. US20130172406, the contents of which are herein incorporated by reference in their entirety. [0309] In some embodiments, the therapeutic nanoparticle may comprise a multiblock copolymer (see e.g., U.S. Pat. Nos.8,263,665 and 8,287,910 and U.S. Patent Pub. No. US20130195987, the contents of each of which are herein incorporated by reference in their entirety). [0310] In yet another non-limiting example, the lipid nanoparticle comprises the block copolymer PEG-PLGA-PEG (see e.g., the thermosensitive hydrogel (PEG-PLGA-PEG) was used as a TGF-beta1 gene delivery vehicle in Lee et al. Thermosensitive Hydrogel as a Tgf-β1 Gene Delivery Vehicle Enhances Diabetic Wound Healing. Pharmaceutical Research, 200320(12): 1995-2000; as a controlled gene delivery system in Li et al. Controlled Gene Delivery System Based on Thermosensitive Biodegradable Hydrogel. Pharmaceutical Research 200320(6):884-888; and Chang et al., Non-ionic amphiphilic biodegradable PEG-PLGA-PEG copolymer enhances gene delivery efficiency in rat skeletal muscle. J Controlled Release.2007118:245-253, the contents of each of which are herein incorporated by reference in their entirety). The RNA (e.g., mRNA) vaccines
1005272698
94 of the present disclosure may be formulated in lipid nanoparticles comprising the PEG- PLGA-PEG block copolymer. [0311] In some embodiments, the therapeutic nanoparticle may comprise a multiblock copolymer (see e.g., U.S. Pat. Nos.8,263,665 and 8,287,910 and U.S. Patent Pub. No. US20130195987, the contents of each of which are herein incorporated by reference in their entirety). [0312] In some embodiments, the block copolymers described herein may be included in a polyion complex comprising a non-polymeric micelle and the block copolymer. (see e.g., U.S. Publication No.20120076836, the contents of which are herein incorporated by reference in their entirety). [0313] In some embodiments, the therapeutic nanoparticle may comprise at least one acrylic polymer. Acrylic polymers include but are not limited to, acrylic acid, methacrylic acid, acrylic acid and methacrylic acid copolymers, methyl methacrylate copolymers, ethoxyethyl methacrylates, cyanoethyl methacrylate, amino alkyl methacrylate copolymer, poly(acrylic acid), poly(methacrylic acid), polycyanoacrylates and combinations thereof. [0314] In some embodiments, the therapeutic nanoparticles may comprise at least one poly(vinyl ester) polymer. The poly(vinyl ester) polymer may be a copolymer such as a random copolymer. As a non-limiting example, the random copolymer may have a structure such as those described in International Application No. WO2013032829 or U.S. Patent Publication No US20130121954, the contents of each of which are herein incorporated by reference in their entirety. In some embodiments, the poly(vinyl ester) polymers may be conjugated to the polynucleotides described herein. [0315] In some embodiments, the therapeutic nanoparticle may comprise at least one diblock copolymer. The diblock copolymer may be, but it not limited to, a poly(lactic) acid- poly(ethylene)glycol copolymer (see, e.g., International Patent Publication No. WO2013044219, the contents of which are herein incorporated by reference in their entirety). [0316] As a non-limiting example, the therapeutic nanoparticle may be used to treat cancer (see International publication No. WO2013044219, the contents of which are herein incorporated by reference in their entirety).
1005272698
95 [0317] In some embodiments, the therapeutic nanoparticles may comprise at least one cationic polymer described herein and/or known in the art. [0318] In some embodiments, the therapeutic nanoparticles may comprise at least one amine-containing polymer such as, but not limited to polylysine, polyethylene imine, poly(amidoamine) dendrimers, poly(beta-amino esters) (see, e.g., U.S. Pat. No. 8,287,849, the contents of which are herein incorporated by reference in their entirety) and combinations thereof. [0319] In some embodiments, the nanoparticles described herein may comprise an amine cationic lipid such as those described in International Patent Application No. WO2013059496, the contents of which are herein incorporated by reference in their entirety. In some embodiments, the cationic lipids may have an amino-amine or an amino- amide moiety. [0320] In some embodiments, the therapeutic nanoparticles may comprise at least one degradable polyester which may contain polycationic side chains. Degradeable polyesters include, but are not limited to, poly(serine ester), poly(L-lactide-co-L-lysine), poly(4-hydroxy-L-proline ester), and combinations thereof. In some embodiments, the degradable polyesters may include a PEG conjugation to form a PEGylated polymer. [0321] In some embodiments, the synthetic nanocarriers may be formulated for targeted release. In some embodiments, the synthetic nanocarrier is formulated to release the polynucleotides at a specified pH and/or after a desired time interval. As a non-limiting example, the synthetic nanoparticle may be formulated to release the RNA (e.g., mRNA) vaccines after 24 hours and/or at a pH of 4.5 (see International Publication Nos. WO2010138193 and WO2010138194 and US Pub Nos. US20110020388 and US20110027217, each of which is herein incorporated by reference in their entireties). [0322] In some embodiments, the synthetic nanocarriers may be formulated for controlled and/or sustained release of the polynucleotides described herein. As a non- limiting example, the synthetic nanocarriers for sustained release may be formulated by methods known in the art, described herein and/or as described in International Pub No. WO2010138192 and US Pub No.20100303850, each of which is herein incorporated by reference in their entirety.
1005272698
96 [0323] In some embodiments, the RNA (e.g., mRNA) vaccine may be formulated for controlled and/or sustained release wherein the formulation comprises at least one polymer that is a crystalline side chain (CYSC) polymer. CYSC polymers are described in U.S. Pat. No.8,399,007, herein incorporated by reference in its entirety. [0324] In some embodiments, the synthetic nanocarrier may be formulated for use as a vaccine. In some embodiments, the synthetic nanocarrier may encapsulate at least one polynucleotide which encode at least one antigen. As a non-limiting example, the synthetic nanocarrier may include at least one antigen and an excipient for a vaccine dosage form (see International Publication No. WO2011150264 and U.S. Publication No. US20110293723, the contents of each of which are herein incorporated by reference in their entirety). As another non-limiting example, a vaccine dosage form may include at least two synthetic nanocarriers with the same or different antigens and an excipient (see International Publication No. WO2011150249 and U.S. Publication No. US20110293701, the contents of each of which are herein incorporated by reference in their entirety). The vaccine dosage form may be selected by methods described herein, known in the art and/or described in International Publication No. WO2011150258 and U.S. Publication No. US20120027806, the contents of each of which are herein incorporated by reference in their entirety). [0325] In some embodiments, the synthetic nanocarrier may comprise at least one polynucleotide which encodes at least one adjuvant. As non-limiting example, the adjuvant may comprise dimethyldioctadecylammonium-bromide, dimethyldioctadecylammonium-chloride, dimethyldioctadecylammonium-phosphate or dimethyldioctadecylammonium-acetate (DDA) and an apolar fraction or part of said apolar fraction of a total lipid extract of a mycobacterium (see, e.g., U.S. Pat. No. 8,241,610, the content of which is herein incorporated by reference in its entirety). In some embodiments, the synthetic nanocarrier may comprise at least one polynucleotide and an adjuvant. As a non-limiting example, the synthetic nanocarrier comprising and adjuvant may be formulated by the methods described in International Publication No. WO2011150240 and U.S. Publication No. US20110293700, the contents of each of which are herein incorporated by reference in their entirety. [0326] In some embodiments, the synthetic nanocarrier may encapsulate at least one polynucleotide that encodes a peptide, fragment or region from a virus. As a non-limiting example, the synthetic nanocarrier may include, but is not limited to, any of the
1005272698
97 nanocarriers described in International Publication No. WO2012024621, WO201202629, WO2012024632 and U.S. Publication No. US20120064110, US20120058153 and US20120058154, the contents of each of which are herein incorporated by reference in their entirety. [0327] In some embodiments, the synthetic nanocarrier may be coupled to a polynucleotide which may be able to trigger a humoral and/or cytotoxic T lymphocyte (CTL) response (see, e.g., International Publication No. WO2013019669, the contents of which are herein incorporated by reference in their entirety). [0328] In some embodiments, the RNA (e.g., mRNA) vaccine may be encapsulated in, linked to and/or associated with zwitterionic lipids. Non-limiting examples of zwitterionic lipids and methods of using zwitterionic lipids are described in U.S. Patent Publication No. US20130216607, the contents of which are herein incorporated by reference in their entirety. [0329] In some embodiment, the zwitterionic lipids may be used in the liposomes and lipid nanoparticles described herein. [0330] In some embodiments, the RNA (e.g., mRNA) vaccine may be formulated in colloid nanocarriers as described in U.S. Patent Publication No. US20130197100, the contents of which are herein incorporated by reference in their entirety. [0331] In some embodiments, the nanoparticle may be optimized for oral administration. The nanoparticle may comprise at least one cationic biopolymer such as, but not limited to, chitosan or a derivative thereof. As a non-limiting example, the nanoparticle may be formulated by the methods described in U.S. Publication No. 20120282343, the contents of which are herein incorporated by reference in their entirety. [0332] In some embodiments, LNPs comprise the lipid KL52 (an amino-lipid disclosed in U.S. Application Publication No. 2012/0295832, the contents of which are herein incorporated by reference in their entirety. Activity and/or safety (as measured by examining one or more of ALT/AST, white blood cell count and cytokine induction, for example) of LNP administration may be improved by incorporation of such lipids. LNPs comprising KL52 may be administered intravenously and/or in one or more doses. In some embodiments, administration of LNPs comprising KL52 results in equal or improved mRNA and/or protein expression as compared to LNPs comprising MC3.
1005272698
98 [0333] In some embodiments, RNA (e.g., mRNA) vaccine may be delivered using smaller LNPs. Such particles may comprise a diameter from below 0.1 um up to 100 nm such as, but not limited to, less than 0.1 um, less than 1.0 um, less than 5 um, less than 10 um, less than 15 um, less than 20 um, less than 25 um, less than 30 um, less than 35 um, less than 40 um, less than 50 um, less than 55 um, less than 60 um, less than 65 um, less than 70 um, less than 75 um, less than 80 um, less than 85 um, less than 90 um, less than 95 um, less than 100 um, less than 125 um, less than 150 um, less than 175 um, less than 200 um, less than 225 um, less than 250 um, less than 275 um, less than 300 um, less than 325 um, less than 350 um, less than 375 um, less than 400 um, less than 425 um, less than 450 um, less than 475 um, less than 500 um, less than 525 um, less than 550 um, less than 575 um, less than 600 um, less than 625 um, less than 650 um, less than 675 um, less than 700 um, less than 725 um, less than 750 um, less than 775 um, less than 800 um, less than 825 um, less than 850 um, less than 875 um, less than 900 um, less than 925 um, less than 950 um, less than 975 um, or less than 1000 um. [0334] In some embodiments, RNA (e.g., mRNA) vaccines may be delivered using smaller LNPs, which may comprise a diameter from about 1 nm to about 100 nm, from about 1 nm to about 10 nm, about 1 nm to about 20 nm, from about 1 nm to about 30 nm, from about 1 nm to about 40 nm, from about 1 nm to about 50 nm, from about 1 nm to about 60 nm, from about 1 nm to about 70 nm, from about 1 nm to about 80 nm, from about 1 nm to about 90 nm, from about 5 nm to about from 100 nm, from about 5 nm to about 10 nm, about 5 nm to about 20 nm, from about 5 nm to about 30 nm, from about 5 nm to about 40 nm, from about 5 nm to about 50 nm, from about 5 nm to about 60 nm, from about 5 nm to about 70 nm, from about 5 nm to about 80 nm, from about 5 nm to about 90 nm, about 10 to about 50 nm, from about 20 to about 50 nm, from about 30 to about 50 nm, from about 40 to about 50 nm, from about 20 to about 60 nm, from about 30 to about 60 nm, from about 40 to about 60 nm, from about 20 to about 70 nm, from about 30 to about 70 nm, from about 40 to about 70 nm, from about 50 to about 70 nm, from about 60 to about 70 nm, from about 20 to about 80 nm, from about 30 to about 80 nm, from about 40 to about 80 nm, from about 50 to about 80 nm, from about 60 to about 80 nm, from about 20 to about 90 nm, from about 30 to about 90 nm, from about 40 to about 90 nm, from about 50 to about 90 nm, from about 60 to about 90 nm and/or from about 70 to about 90 nm.
1005272698
99 [0335] In some embodiments, such LNPs are synthesized using methods comprising microfluidic mixers. Examples of microfluidic mixers may include, but are not limited to, a slit interdigital micromixer including, but not limited to those manufactured by Microinnova (Allerheiligen bei Wildon, Austria) and/or a staggered herringbone micromixer (SHM) (Zhigaltsev, I. V. et al., Bottom-up design and synthesis of limit size lipid nanoparticle systems with aqueous and triglyceride cores using millisecond microfluidic mixing have been published (Langmuir. 2012. 28:3633-40; Belliveau, N. M. et al., Microfluidic synthesis of highly potent limit-size lipid nanoparticles for in vivo delivery of siRNA. Molecular Therapy-Nucleic Acids.2012.1:e37; Chen, D. et al., Rapid discovery of potent siRNA-containing lipid nanoparticles enabled by controlled microfluidic formulation. J Am Chem Soc.2012.134(16):6948-51, the contents of each of which are herein incorporated by reference in their entirety). In some embodiments, methods of LNP generation comprising SHM, further comprise the mixing of at least two input streams wherein mixing occurs by microstructure-induced chaotic advection (MICA). According to this method, fluid streams flow through channels present in a herringbone pattern causing rotational flow and folding the fluids around each other. This method may also comprise a surface for fluid mixing wherein the surface changes orientations during fluid cycling. Methods of generating LNPs using SHM include those disclosed in U.S. Application Publication Nos. 2004/0262223 and 2012/0276209, the contents of each of which are herein incorporated by reference in their entirety. [0336] In some embodiments, the RNA (e.g., mRNA) vaccine of the present disclosure may be formulated in lipid nanoparticles created using a micromixer such as, but not limited to, a Slit Interdigital Microstructured Mixer (SIMM-V2) or a Standard Slit Interdigital Micro Mixer (SSIMM) or Caterpillar (CPMM) or Impinging-jet (IJMM) from the Institut fiir Mikrotechnik Mainz GmbH, Mainz Germany). [0337] In some embodiments, the RNA (e.g., mRNA) vaccines of the present disclosure may be formulated in lipid nanoparticles created using microfluidic technology (see, e.g., Whitesides, George M. The Origins and the Future of Microfluidics. Nature, 2006442: 368-373; and Abraham et al. Chaotic Mixer for Microchannels. Science, 2002 295: 647-651; each of which is herein incorporated by reference in its entirety). As a non- limiting example, controlled microfluidic formulation includes a passive method for mixing streams of steady pressure-driven flows in micro channels at a low Reynolds number
1005272698
100 (see, e.g., Abraham et al. Chaotic Mixer for Microchannels. Science, 2002295: 647-651, the contents of which are herein incorporated by reference in their entirety). [0338] In some embodiments, the RNA (e.g., mRNA) vaccines of the present disclosure may be formulated in lipid nanoparticles created using a micromixer chip such as, but not limited to, those from Harvard Apparatus (Holliston, Mass.) or Dolomite Microfluidics (Royston, UK). A micromixer chip can be used for rapid mixing of two or more fluid streams with a split and recombine mechanism. [0339] In some embodiments, the RNA (e.g., mRNA) vaccines of the disclosure may be formulated for delivery using the drug encapsulating microspheres described in International Patent Publication No. WO2013063468 or U.S. Pat. No. 8,440,614, the contents of each of which are herein incorporated by reference in their entirety. The microspheres may comprise a compound of the formula (I), (II), (III), (IV), (V) or (VI) as described in International Patent Publication No. WO2013063468, the contents of which are herein incorporated by reference in their entirety. In some embodiments, the amino acid, peptide, polypeptide, lipids (APPL) are useful in delivering the RNA (e.g., mRNA) vaccines of the disclosure to cells (see International Patent Publication No. WO2013063468, the contents of which are herein incorporated by reference in their entirety). [0340] In any embodiment, a lipid-based formulation including any LNP disclosed herein may further comprise one or more adjuvants. For example, in any embodiment, an ionisable lipid present in the nanoparticle formulation may be substituted or combined with an adjuvant lipidoid for enhancing RNA delivery. Examples of such approaches are described in the prior art, such as in Han et al., (2023) Nature Nanotechnology, https://doi.org/10.1038/s41565-023-01404-4, and Salleh et al., (2022), Peer J, 10:e13083; incorporated herein by reference. [0341] In some embodiments, the antibody titre produced by the mRNA vaccines of the invention is a neutralizing antibody titre. In some embodiments the neutralizing antibody titre is greater than a protein vaccine. In other embodiments the neutralizing antibody titre produced by the mRNA vaccines of the invention is greater than an adjuvanted protein vaccine. In yet other embodiments the neutralizing antibody titre produced by the mRNA vaccines of the invention is 1,000-10,000, 1,200-10,000, 1,400-10,000, 1,500-10,000, 1,000-5,000, 1,000-4,000, 1,800-10,000, 2000-10,000, 2,000-5,000, 2,000-3,000, 2,000-
1005272698
101 4,000, 3,000-5,000, 3,000-4,000, or 2,000-2,500. A neutralization titre is typically expressed as the highest serum dilution required to achieve a 50% reduction in the number of plaques. [0342] In some embodiments, the disclosure features a pharmaceutical composition comprising a nanoparticle composition according to the preceding embodiments and a pharmaceutically acceptable carrier. The pharmaceutically acceptable carrier may be as described herein, and may also include one or more agents for facilitating storage of the composition at low temperatures. For example, the pharmaceutical composition may be refrigerated or frozen for storage and/or shipment (e.g., being stored at a temperature of 4° C or lower, such as a temperature between about −150° C. and about 0° C. or between about −80° C and about −20° C (e.g., about −5° C, −10° C, −15° C, −20° C, −25° C, −30° C, −40° C, −50° C, −60° C, −70° C, −80° C, −90° C, −130° C or −150° C.). For example, the pharmaceutical composition is a solution that is refrigerated for storage and/or shipment at, for example, about −20° C, −30° C, −40° C, −50° C, −60° C, −70° C, or −80° C. Accordingly, it will be appreciated that the compositions described herein may further comprise one or more cryoprotectants or cryopreservatives. Optionally the cryopreservative or cryoprotectant may comprise a sugar such as sucrose, glucose or related sugar-based cryoprotectant. Liposomes and Lipoplexes, and Lipid Nanoparticles [0343] The RNA (e.g., mRNA) vaccines of the disclosure can be formulated using one or more liposomes, lipoplexes, or lipid nanoparticles. In some embodiments, pharmaceutical compositions of RNA (e.g., mRNA) vaccines include liposomes. Liposomes are artificially-prepared vesicles which may primarily be composed of a lipid bilayer and may be used as a delivery vehicle for the administration of nutrients and pharmaceutical formulations. Liposomes can be of different sizes such as, but not limited to, a multilamellar vesicle (MLV) which may be hundreds of nanometers in diameter and may contain a series of concentric bilayers separated by narrow aqueous compartments, a small unicellular vesicle (SUV) which may be smaller than 50 nm in diameter, and a large unilamellar vesicle (LUV) which may be between 50 and 500 nm in diameter. Liposome design may include, but is not limited to, opsonins or ligands in order to improve the attachment of liposomes to unhealthy tissue or to activate events such as, but not limited to, endocytosis. Liposomes may contain a low or a high pH in order to improve the delivery of the pharmaceutical formulations.
1005272698
102 [0344] The formation of liposomes may depend on the physicochemical characteristics such as, but not limited to, the pharmaceutical formulation entrapped and the liposomal ingredients, the nature of the medium in which the lipid vesicles are dispersed, the effective concentration of the entrapped substance and its potential toxicity, any additional processes involved during the application and/or delivery of the vesicles, the optimization size, polydispersity and the shelf-life of the vesicles for the intended application, and the batch-to-batch reproducibility and possibility of large-scale production of safe and efficient liposomal products. [0345] In some embodiments, pharmaceutical compositions described herein may include, without limitation, liposomes such as those formed from 1,2-dioleyloxy-N,N- dimethylaminopropane (DODMA) liposomes, DiLa2 liposomes from Marina Biotech (Bothell, Wash.), 1,2-dilinoleyloxy-3-dimethylaminopropane (DLin-DMA), 2,2-dilinoleyl-4- (2-dimethylaminoethyl)-[1,3]-dioxolane (DLin-KC2-DMA), and MC3 (US20100324120; herein incorporated by reference in its entirety) and liposomes which may deliver small molecule drugs such as, but not limited to, DOXIL® from Janssen Biotech, Inc. (Horsham, Pa.). [0346] In some embodiments, pharmaceutical compositions described herein may include, without limitation, liposomes such as those formed from the synthesis of stabilized plasmid-lipid particles (SPLP) or stabilized nucleic acid lipid particle (SNALP) that have been previously described and shown to be suitable for oligonucleotide delivery in vitro and in vivo (see Wheeler et al. Gene Therapy.19996:271-281; Zhang et al. Gene Therapy.19996:1438-1447; Jeffs et al. Pharm Res.200522:362-372; Morrissey et al., Nat Biotechnol. 2005 2:1002-1007; Zimmermann et al., Nature. 2006 441:111-114; Heyes et al. J Contr Rel.2005107:276-287; Semple et al. Nature Biotech.201028:172- 176; Judge et al. J Clin Invest.2009119:661-673; deFougerolles Hum Gene Ther.2008 19:125-132; U.S. Patent Publication No US20130122104; all of which are incorporated herein in their entireties). The original manufacture method by Wheeler et al. was a detergent dialysis method, which was later improved by Jeffs et al. and is referred to as the spontaneous vesicle formation method. The liposome formulations are composed of 3 to 4 lipid components in addition to the polynucleotide. As an example a liposome can contain, but is not limited to, 55% cholesterol, 20% disteroylphosphatidyl choline (DSPC), 10% PEG-S-DSG, and 15% 1,2-dioleyloxy-N,N-dimethylaminopropane (DODMA), as described by Jeffs et al. As another example, certain liposome formulations may contain,
1005272698
103 but are not limited to, 48% cholesterol, 20% DSPC, 2% PEG-c-DMA, and 30% cationic lipid, where the cationic lipid can be 1,2-distearloxy-N,N-dimethylaminopropane (DSDMA), DODMA, DLin-DMA, or 1,2-dilinolenyloxy-3-dimethylaminopropane (DLenDMA), as described by Heyes et al. [0347] In some embodiments, liposome formulations may comprise from about 25.0% cholesterol to about 40.0% cholesterol, from about 30.0% cholesterol to about 45.0% cholesterol, from about 35.0% cholesterol to about 50.0% cholesterol and/or from about 48.5% cholesterol to about 60% cholesterol. In some embodiments, formulations may comprise a percentage of cholesterol selected from the group consisting of 28.5%, 31.5%, 33.5%, 36.5%, 37.0%, 38.5%, 39.0% and 43.5%. In some embodiments, formulations may comprise from about 5.0% to about 10.0% DSPC and/or from about 7.0% to about 15.0% DSPC. [0348] In some embodiments, the RNA (e.g., mRNA) vaccine pharmaceutical compositions may be formulated in liposomes such as, but not limited to, DiLa2 liposomes (Marina Biotech, Bothell, Wash.), SMARTICLES® (Marina Biotech, Bothell, Wash.), neutral DOPC (1,2-dioleoyl-sn-glycero-3-phosphocholine) based liposomes (e.g., siRNA delivery for ovarian cancer (Landen et al. Cancer Biology & Therapy 20065(12)1708- 1713); herein incorporated by reference in its entirety) and hyaluronan-coated liposomes (Quiet Therapeutics, Israel). [0349] In some embodiments, the cationic lipid may be a low molecular weight cationic lipid such as those described in U.S. Patent Application No.20130090372, the contents of which are herein incorporated by reference in their entirety. [0350] In some embodiments, the RNA (e.g., mRNA) vaccines may be formulated in a lipid vesicle, which may have crosslinks between functionalized lipid bilayers. [0351] In some embodiments, the RNA (e.g., mRNA) vaccines may be formulated in a lipid-polycation complex. The formation of the lipid-polycation complex may be accomplished by methods known in the art and/or as described in U.S. Pub. No. 20120178702, herein incorporated by reference in its entirety. As a non-limiting example, the polycation may include a cationic peptide or a polypeptide such as, but not limited to, polylysine, polyornithine and/or polyarginine. In some embodiments, the RNA (e.g., mRNA) vaccines may be formulated in a lipid-polycation complex, which may further
1005272698
104 include a non-cationic lipid such as, but not limited to, cholesterol or dioleoyl phosphatidylethanolamine (DOPE). [0352] Further examples of suitable formulations for mRNA delivery are described in Guevara et al., (2020), Frontiers in Chemistry, 8:589959; Zhang et al., (2019), Frontiers in Immunology, 10:594; and Liu et al., (2022), Polymers, 14: 4195; incorporated by reference herein in their entirety. Subjects and methods of administration [0353] The present invention also provides uses of the polynucleotides and compositions of the invention for producing an antigen-specific immune response in a subject. Such methods typically comprise administering a polynucleotide of the invention, preferably formulated in a lipid nanoparticle as described herein, to a subject in need thereof. [0354] Accordingly, the invention further provides compositions comprising the polynucleotides (RNA) defined herein, and the use of such RNA in immunogenic or vaccine compositions in the treatment or prevention of P. gingivalis infection. [0355] The term "vaccine composition" used herein is defined as a composition used to elicit an immune response against an antigen (immunogen) encoded by the RNA in the composition in order to protect or treat an organism against disease. [0356] As used herein, the terms “immunostimulating composition”, “vaccine composition” and “immunogenic composition” may generally be used interchangeably. [0357] The present invention provides methods and compositions for treating or preventing infection or minimising the likelihood of infection with P. gingivalis, in an individual in need thereof, the methods comprising administering a vaccine composition of the invention. [0358] As such, the present invention includes methods and compositions for preventing infection with P. gingivalis, minimising the likelihood of infection and/or reducing the severity and duration of P. gingivalis infection in a subject. [0359] The present invention also provides a method for obtaining an antibody directed to P. gingivalis, the method comprising administering a chimeric or fusion protein,
1005272698
105 composition, vaccine or immune stimulating composition of the invention, to a non-human animal, thereby generating antibodies directed to P. gingivalis in the animal. Preferably the method further comprises isolating the antibody from the animal (eg from the blood of the animal) or from an egg of the animal (eg in the case of generating IgY antibodies from chickens). [0360] The present invention also provides an antibody preparation comprising an antibody directed to P. gingivalis, wherein the antibody preparation is obtained by administering a composition, vaccine or immune stimulating composition of the invention, to a non-human animal, thereby generating antibodies directed to P. gingivalis in the animal, and isolating the antibodies from the animal or egg thereof. [0361] The antibody directed to P. gingivalis may be used therapeutically to eliminate or reduce P. gingivalis infection or prophylactically, to prevent or reduce the severity of P. gingivalis infection. [0362] As used herein, the terms "treatment" or "treating" of a subject includes the application or administration of a composition of the invention to a subject (or application or administration of a compound of the invention to a cell or tissue from a subject) with the purpose of delaying, slowing, stabilizing, curing, regressing, healing, alleviating, relieving, altering, remedying, less worsening, ameliorating, improving, or affecting the disease or condition, the symptom of the disease or condition, or the risk of (or susceptibility to) the disease or condition. The term "treating" refers to any indication of success in the treatment or amelioration of an injury, pathology or condition, including any objective or subjective parameter such as abatement; remission; lessening of the rate of worsening; lessening severity of the disease; stabilization, diminishing of symptoms or making the injury, pathology or condition more tolerable to the subject; slowing in the rate of degeneration or decline; making the final point of degeneration less debilitating; or improving a subject's physical or mental well-being. [0363] As used herein, "preventing" or "prevention" is intended to refer to at least the reduction of likelihood of the risk of (or susceptibility to) acquiring a disease or disorder (i.e., causing at least one of the clinical symptoms of the disease not to develop in a subject that may be exposed to or predisposed to the disease but does not yet experience or display symptoms of the disease). Biological and physiological parameters for identifying such subjects are provided herein and are also well known by physicians.
1005272698
106 [0364] The vaccine compositions of the invention can be administered to subjects felt to be in greatest need thereof, for example in the context of human patients, to children or the elderly or individuals at risk of exposure to P. gingivalis. The vaccine compositions of the invention can also be administered to subjects suspected of having or diagnosed with having infection with P. gingivalis. [0365] The compositions and methods of the present invention extend equally to uses in both human and/or veterinary medicine, generation of diagnostic agents or the generation of other treatment reagents. [0366] As used herein, the term “subject” shall be taken to mean any animal including humans, for example a mammal. Exemplary subjects include but are not limited to humans and non-human primates. For example, the subject may be a human. In further examples, the subject may be a veterinary subject, such as a companion animal (cat, dog, guinea pig, and the like). [0367] As used herein, the terms “subject”, “individual” and “patient” may be used interchangeably. [0368] The skilled person will be familiar with methods for determining successful vaccination/immunisation with a chimeric or fusion protein or composition as described herein. For example, the skilled person will be familiar with methods for quantifying the antibodies generated following immunisation and/or for quantifying the extent of the humoural (Th2) response induced following immunisation or for quantifying the extent of a Th1 response generated. [0369] In some embodiments, following administration of a polynucleotide or composition of the invention, the subject exhibits a seroconversion rate of at least 80% (e.g., at least 85%, at least 90%, or at least 95%) following the first dose or the second (booster) dose of the vaccine. Seroconversion is the time period during which a specific antibody develops and becomes detectable in the blood. During an infection or immunization, antigens enter the blood, and the immune system begins to produce antibodies in response. Before seroconversion, the antigen itself may or may not be detectable, but antibodies are considered absent. During seroconversion, antibodies are present but not yet detectable. Any time after seroconversion, the antibodies can be detected in the blood, indicating a prior or current infection.
1005272698
107 [0370] In some embodiments, a polynucleotide (e.g., mRNA) vaccine is administered to a subject by intradermal or intramuscular injection, subcutaneous, intravenous, or intranasal route, or any other suitable route for delivery of an RNA-based vaccine. [0371] In some embodiments of the present disclosure, there is provided methods of inducing an antigen specific immune response in a subject, including administering to a subject a RNA (e.g.,mRNA) vaccine as described herein, in an effective amount to produce an antigen specific immune response in a subject. Antigen-specific immune responses in a subject may be determined, in some embodiments, by assaying for antibody titre following administration to the subject of any of the polynucleotide (e.g., mRNA) vaccines of the present disclosure. In some embodiments, the anti-antigenic polypeptide antibody titre produced in the subject is increased by at least 1 log relative to a control. In some embodiments, the anti-antigenic polypeptide antibody titre produced in the subject is increased by 1-3 log relative to a control. [0372] In some embodiments, the anti-antigenic polypeptide antibody titre produced in a subject is increased at least 2 times relative to a control. In some embodiments, the anti-antigenic polypeptide titre produced in the subject is increased at least 5 times relative to a control. In some embodiments, the anti-antigenic polypeptide antibody titre produced in the subject is increased at least 10 times relative to a control. In some embodiments, the anti-antigenic polypeptide antibody titre produced in the subject is increased 2-10 times relative to a control. [0373] In some embodiments, the control is an anti-antigenic polypeptide antibody titre produced in a subject who has not been administered a RNA (e.g., mRNA) vaccine of the present disclosure. In some embodiments, the control is an anti-antigenic polypeptide antibody titre produced in a subject who has been administered a live attenuated or inactivated P. gingivalis vaccine (see, e.g., Ren J. et al. J of Gen. Virol.2015; 96: 1515- 1520), or wherein the control is an anti-antigenic polypeptide antibody titre produced in a subject who has been administered a recombinant or purified P. gingivalis protein vaccine. [0374] A polynucleotide (e.g., mRNA) vaccine of the present disclosure is administered to a subject in an effective amount (an amount effective to induce an immune response). In some embodiments, the effective amount is a dose equivalent to an at least 2-fold, at least 4-fold, at least 10-fold, at least 100-fold, at least 1000-fold reduction in the standard
1005272698
108 of care dose of a recombinant P. gingivalis protein vaccine, wherein the anti-antigenic polypeptide antibody titre produced in the subject is equivalent to an anti-antigenic polypeptide antibody titre produced in a control subject administered the standard of care dose of a recombinant P. gingivalis protein vaccine, a purified P. gingivalis protein vaccine, a live attenuated P. gingivalis vaccine, an inactivated P. gingivalis vaccine. In some embodiments, the effective amount is a dose equivalent to 2-1000-fold reduction in the standard of care dose of a recombinant P. gingivalis protein vaccine, wherein the anti- antigenic polypeptide antibody titre produced in the subject is equivalent to an anti- antigenic polypeptide antibody titre produced in a control subject administered the standard of care dose of a recombinant P. gingivalis protein vaccine, a purified P. gingivalis protein vaccine, a live attenuated P. gingivalis vaccine, or an inactivated P. gingivalis vaccine. [0375] In some embodiments, the polynucleotide (e.g., mRNA) vaccine is formulated in an effective amount to produce an antigen specific immune response in a subject. [0376] Dosages of the vaccine composition of the present invention can vary between wide limits, depending upon the age and condition of the individual to be treated, etc. and a physician will ultimately determine appropriate dosages to be used. [0377] This dosage can be repeated as often as appropriate. For example, an initial dose of the vaccine may be administered and then a booster administered at a later date. [0378] In some embodiments, the effective amount is a total dose of 25 µg to 1000 µg, or 50 µg to 1000 µg. In some embodiments, the effective amount is a total dose of 100 µg. In some embodiments, the effective amount is a dose of 25 µg administered to the subject a total of two times. In some embodiments, the effective amount is a dose of 100 µg administered to the subject a total of two times. In some embodiments, the effective amount is a dose of 400 µg administered to the subject a total of two times. In some embodiments, the effective amount is a dose of 500 µg administered to the subject a total of two times. [0379] In some embodiments, the efficacy (or effectiveness) of a polynucleotide (e.g., mRNA) vaccine is greater than 60%. In some embodiments, the efficacy (or
1005272698
109 effectiveness) of a polynucleotide (e.g., mRNA) vaccine is at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, or at least 90%. [0380] Vaccine efficacy may be assessed using standard analyses (see, e.g., Weinberg et al., J Infect Dis. 2010 Jun. 1; 201(11):1607-10). For example, vaccine efficacy may be measured by double-blind, randomized, clinical controlled trials. Vaccine efficacy may be expressed as a proportionate reduction in disease attack rate (AR) between the unvaccinated (ARU) and vaccinated (ARV) study cohorts and can be calculated from the relative risk (RR) of disease among the vaccinated group with use of the following formulas: Efficacy = (ARU−ARV)/ARU × 100; and Efficacy = (1−RR) × 100. [0381] Similarly, vaccine effectiveness may be assessed using standard analyses (see, e.g., Weinberg et al., J Infect Dis.2010 Jun.1; 201(11):1607-10). Vaccine effectiveness is an assessment of how a vaccine (which may have already proven to have high vaccine efficacy) reduces disease in a population. This measure can assess the net balance of benefits and adverse effects of a vaccination program, not just the vaccine itself, under natural field conditions rather than in a controlled clinical trial. Vaccine effectiveness is proportional to vaccine efficacy (potency) but is also affected by how well target groups in the population are immunized, as well as by other non-vaccine-related factors that influence the ‘real-world’ outcomes of hospitalizations, ambulatory visits, or costs. For example, a retrospective case control analysis may be used, in which the rates of vaccination among a set of infected cases and appropriate controls are compared. Vaccine effectiveness may be expressed as a rate difference, with use of the odds ratio (OR) for developing infection despite vaccination: Effectiveness = (1−OR) × 100. [0382] Vaccine efficacy may also be assessed by evidence of treatment of a P. gingivalis infection as herein defined. The evidence of treatment may comprise reduction in the severity or duration of a P. gingivalis infection in a subject, such as reduction in inflammation caused by infection.
1005272698
110 [0383] In other embodiments the invention is a composition for or method of vaccinating a subject comprising administering to the subject a nucleic acid vaccine comprising one or more RNA polynucleotides having an open reading frame encoding a first antigenic polypeptide wherein a dosage of between 10 µg/kg and 400 µg /kg of the nucleic acid vaccine is administered to the subject. In some embodiments the dosage of the RNA polynucleotide is 1-5 µg, 5-10 µg, 10-15 µg, 15-20 µg, 10-25 µg, 20-25 µg, 20- 50 µg, 30-50 µg, 40-50 µg, 40-60 µg, 60-80 µg, 60-100 µg, 50-100 µg, 80-120 µg, 40-120 µg, 40-150 µg, 50-150 µg, 50-200 µg, 80-200 µg, 100-200 µg, 120-250 µg, 150-250 µg, 180-280 µg, 200-300 µg, 50-300 µg, 80-300 µg, 100-300 µg, 40-300 µg, 50-350 µg, 100- 350 µg, 200-350 µg, 300-350 µg, 320-400 µg, 40-380 µg, 40-100 µg, 100-400 µg, 200- 400 µg, or 300-400 µg per dose. In some embodiments, the nucleic acid vaccine is administered to the subject by intradermal or intramuscular injection. In some embodiments, the nucleic acid vaccine is administered to the subject on day zero. In some embodiments, a second dose of the nucleic acid vaccine is administered to the subject on day twenty one. [0384] In some embodiments, a dosage of at least about 2 micrograms (µg) or at least about 10 µg or at least about 20 µg or at least about 30 µg of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, a dosage of 100 micrograms (µg) of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, a dosage of 50 micrograms (µg) of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, a dosage of 75 micrograms (µg) of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, a dosage of 150 micrograms (µg) of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, a dosage of 400 micrograms of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, a dosage of 200 micrograms (µg) of the RNA polynucleotide is included in the nucleic acid vaccine administered to the subject. In some embodiments, the RNA polynucleotide accumulates at a 100 fold higher level in the local lymph node in comparison with the distal lymph node. [0385] Embodiments of the invention provide methods of creating, maintaining or restoring antigenic memory to a P. gingivalis in an individual or population of individuals comprising administering to said individual or population an antigenic memory booster
1005272698
111 nucleic acid vaccine comprising (a) at least one RNA polynucleotide, said polynucleotide comprising at least one chemical modification or optionally no nucleotide modification and two or more codon-optimized open reading frames, said open reading frames encoding a set of reference antigenic polypeptides, and (b) optionally a pharmaceutically acceptable carrier or excipient. In some embodiments, the vaccine is administered to the individual via a route selected from the group consisting of intramuscular administration, intradermal administration and subcutaneous administration. In some embodiments, the administering step comprises contacting a muscle tissue of the subject with a device suitable for injection of the composition. In some embodiments, the administering step comprises contacting a muscle tissue of the subject with a device suitable for injection of the composition in combination with electroporation. [0386] Embodiments of the invention provide methods of vaccinating a subject comprising administering to the subject a single dosage of between 25 µg/kg and 400 µg/kg of a nucleic acid vaccine comprising one or more RNA polynucleotides as described herein, in an effective amount to vaccinate the subject. [0387] In other embodiments the invention encompasses a method of treating an elderly subject age 60 years or older comprising administering to the subject a nucleic acid vaccine comprising one or more RNA polynucleotides having an open reading frame encoding an antigenic polypeptide as described herein in an effective amount to vaccinate the subject. [0388] In other embodiments the invention encompasses a method of treating a young subject age 17 years or younger comprising administering to the subject a nucleic acid vaccine comprising one or more RNA polynucleotides having an open reading frame encoding an antigenic polypeptide as described herein in an effective amount to vaccinate the subject. [0389] In other embodiments the invention encompasses a method of treating an adult subject comprising administering to the subject a nucleic acid vaccine comprising one or more RNA polynucleotides having an open reading frame encoding an antigenic polypeptide as described herein in an effective amount to vaccinate the subject. [0390] In preferred embodiments, vaccines of the invention (e.g., LNP-encapsulated mRNA vaccines) produce prophylactically- and/or therapeutically-efficacious levels,
1005272698
112 concentrations and/or titres of antigen-specific antibodies in the blood or serum of a vaccinated subject. [0391] As defined herein, the term antibody titre refers to the amount of antigen-specific antibody produced in a subject, e.g., a human subject. In exemplary embodiments, antibody titre is expressed as the inverse of the greatest dilution (in a serial dilution) that still gives a positive result. [0392] In exemplary embodiments, antibody titre is determined or measured by enzyme-linked immunosorbent assay (ELISA). In exemplary embodiments, antibody titre is determined or measured by neutralization assay, e.g., by microneutralization assay. In certain embodiments, antibody titre measurement is expressed as a ratio, such as 1:40, 1:100, etc. In exemplary embodiments of the invention, an efficacious vaccine produces an antibody titre of greater than 1:40, greater that 1:100, greater than 1:400, greater than 1:1000, greater than 1:2000, greater than 1:3000, greater than 1:4000, greater than 1:500, greater than 1:6000, greater than 1:7500, greater than 1:10000. In exemplary embodiments, the antibody titre is produced or reached by 10 days following vaccination, by 20 days following vaccination, by 30 days following vaccination, by 40 days following vaccination, or by 50 or more days following vaccination. In exemplary embodiments, the titre is produced or reached following a single dose of vaccine administered to the subject. In other embodiments, the titre is produced or reached following multiple doses, e.g., following a first and a second dose (e.g., a booster dose.) In exemplary embodiments of the invention, antigen-specific antibodies are measured in units of µg /ml or are measured in units of IU/L (International Units per liter) or mIU/ml (milli International Units per ml). In exemplary embodiments of the invention, an efficacious vaccine produces >0.5 µg/ml, >0.1 µg /ml, >0.2 µg /ml, >0.35 µg /ml, >0.5 µg /ml, >1 µg /ml, >2 µg /ml, >5 µg /ml or >10 µg /ml. In exemplary embodiments of the invention, an efficacious vaccine produces >10 mIU/ml, >20 mIU/ml, >50 mIU/ml, >100 mIU/ml, >200 mIU/ml, >500 mIU/ml or >1000 mIU/ml. In exemplary embodiments, the antibody level or concentration is produced or reached by 10 days following vaccination, by 20 days following vaccination, by 30 days following vaccination, by 40 days following vaccination, or by 50 or more days following vaccination. In exemplary embodiments, the level or concentration is produced or reached following a single dose of vaccine administered to the subject. In other embodiments, the level or concentration is produced or reached following multiple doses, e.g., following a first and a second dose (e.g., a booster dose.) In exemplary
1005272698
113 embodiments, antibody level or concentration is determined or measured by enzyme- linked immunosorbent assay (ELISA). [0393] In exemplary embodiments, antibody level or concentration is determined or measured by neutralization assay, e.g., by microneutralization assay. Examples [0394] Example 1: Materials and methods [0395] Production of mRNA-LNPs [0396] The mRNAs were in vitro transcribed using T7 in vitro transcription kit (NEB) according to manufacturer’s instructions, from linearised DNA templates encoding 5’ and 3’ UTRs, signal peptide, the candidate sequence and a 125 nucleotide poly(A) tail and were capped co-transcriptionally using Clean Cap mRNA capping technology (TriLink Biotech). Where clearly stated, the UTP was replaced by N1-methyl pseudouridine (N1- methyl pseudo-UTP, m1Ψ) during mRNA production. DNA was removed using DNAse I (NEB), and double-stranded RNA (dsRNA) was removed using cellulose binding as previously described previously [Baiersdörfer et al. (2019) Mol Ther Nucleic Acids. Apr 15;15:26-35] and formulated in lipid nanoparticles (LNPs) of the following lipid composition: ALC-0315, Cholesterol, DSPC, and ALC-0159 at molar lipid ratios (%) of 46.3:42.7:9.4:1.6, in Tris/sucrose buffer (25 mM Tris pH 7.4, 8.8% sucrose w/v). [0397] Expression of mRNA vaccine candidates in cell culture in vitro [0398] HeLa cells were cultured in culture media DMEM, high glucose, GlutaMAX™ Supplement, pyruvate (ThermoFisher, CAT#10569010) with 10% FBS at 37°C, 5% CO2, according to standard protocols. mRNA candidates (prior to LNP formulation), were formulated with Lipofectamine MessengerMAX (IThermoFisher) and used to transfect cells according to manufacturer’s instructions.1.8 µg of an mRNA was used to transfect 35,000 cells/well on a six well plate. For supernatant collection, cells were pelleted by centrifuging at 14,000 g for 15 mins. Supernatants were transferred to a fresh tube and frozen at -20°C until Western Blotting. For whole cell lysate collection, cells were rinsed in each well with ~2 mL DPBS. The liquid was removed and 250 µL of RIPA lysis buffer with protease inhibitors (ThermoFisher) was added to each well, and gently swirled to mix for 10s. Using a cell scraper the lysates were transferred into tubes and centrifuged at
1005272698
114 4°C at 16,000 g for 15 mins to pellet debris. The clarified lysates were collected into new tubes and stored at -20°C until Western Blotting. [0399] Western Blot [0400] Samples were mixed with 4X Laemmli sample buffer (10%v/v) supplemented with 2-mercaptoethanol and 100mM DTT, centrifuged at 13,000 g speed for 30 seconds and incubated at room temperature for 20 min, then denatured at 95°C for 10 min. Samples were loaded onto Any kD™ Mini-PROTEAN® TGX™ Precast Protein Gels (BioRad, ca# 4569036) and proteins were separated by SDS-PAGE with Tris/Glycine/SDS running buffer (BioRad, cat# 1610732) at 150V for 45 minutes. Precision Plus Protein™ WesternC™ Blotting Standard was used as a protein marker (BioRad, cat# 161-0376). Proteins were transferred onto a PDV membrane (Life Technologies, cat# LC2002) that had been pre-activated in methanol for 30 s and washed in transfer buffer (10x Tris/Glycine Buffer for Western Blots and Native Gels (BioRad, cat# 1610734), using Trans-Blot® Turbo™ Transfer System (BioRad) with the standard protocol: 25V and 1.0A for 30 min. [0401] PVDF membranes with proteins transferred onto them were blocked for 2 hours in blocking buffer (5% skim milk powder in PBS-T) on a rocker and probed with in-house primary antibodies for binding to KAS2 (1:2500 in blocking buffer) or KDAK-3S-AVQP (1:3500 in blocking buffer) overnight at 4°C on a rocker. Membranes were washed 5 times with PBS-T and incubated with goat anti mouse HRP (1:2500 in blocking buffer, supplemented with Precision Protein StrepTactin-HRP Conjugate BioRad, cat# 1610381 at 1:10,000) at room temperature for an hour on a rocker, washed 5 times with PBS-T, developed using Clarity™ Western ECL Substrate (BioRad, cat# 170-5060) and imaged using ChemiDoc™ Touch Imaging System using chemiluminescence detection. [0402] Culture of bacteria for mouse model of periodontitis [0403] P. gingivalis strain; W50 (serotype C); was obtained from the culture collection of the Oral Health Cooperative Research Centre, The Melbourne Dental School, University of Melbourne, Australia. P. gingivalis W50 was grown on Horse Blood Agar (HBA) (20 g/L HBA; Oxoid Ltd., Hampshire, UK) supplemented with 10% v/v lysed horse blood (37°C) in an anaerobic N2 atmosphere containing 5% CO2 in a MK3 Anaerobic Workstation (Don Whitley Scientific Ltd., Adelaide, Australia). Colonies were inoculated
1005272698
115 into starter culture comprised of 20 mL sterilised brain heart infusion (37 g/L BHI; Oxoid Ltd., Hamsphire, UK) medium supplemented with 5 mg/L hemin and 0.5 mg/L cysteine and incubated anaerobically (24 h, 37 °C). Absorbance of batch cultures were monitored at OD650nm using a spectrophotometer (model 295E, Perkin-Elmer, Germany). Bacterial cells were harvested during late exponential growth by centrifugation (7,000 g, 20 min, 4 °C). Bacterial purity was routinely confirmed by Gram stain [Slots (1982). In: Host- Parasite Interaction in Periodontal Disease, Genco, R.J. and Merganhagan, S.E. (eds). Washington D.C.: American Society for Microbiology. pp.27-45.]. [0404] Preparation of heat-killed bacteria [0405] P. gingivalis W50 culture was harvested (6,500 g, 4 °C), washed once with phosphate buffered saline (PBS) (0.01 M Na2HPO4, 1.5 mM KH2PO4 and 0.15 M NaCl, pH 7.4) then pelleted by centrifugation (7,000 g, 20 min 4°C). Bacterial cells were resuspended in PBS and heated to 65°C for 15 minutes. The suspension was centrifuged (7,000 g, 20 min 4ºC) and resuspended in sterile PBS and this was repeated once. After the second wash, the supernatant was discarded and the cell pellet was resuspended in sterile PBS to obtain a cell density of 2 x 1010 cells/mL, and protein concentration determined using Biorad Protein Assay Dye Reagent Concentrate (Life Science, NSW, Australia). [0406] Mouse periodontitis model [0407] The mouse periodontitis experiments were performed as described previously by O’Brien-Simpson et al. (2005 J Immunol 175: 3980-3989). Mice (female BALB/c; 6-8 weeks old, 10 mice/group), from Day 0 were intra-orally inoculated with P. gingivalis consisting of four doses of P. gingivalis W50 [1 x 1010 viable P. gingivalis W50 cells suspended in 20 µL PG buffer (50 mM Tris-HCL, 150 mM NaCl, 10 mM MgSO4 and 14.3 mM Mercaptoethanol, pH 7.4) containing 2% w/v carboxymethylcellulose (CMC, Sigma, New South Wales, Australia)], given two days apart. The inocula were prepared anaerobically and then immediately applied to the gingival margin of the maxillary molar teeth. The number of viable bacteria in each inoculum was verified by flow cytometry and CFU counts on blood agar. Groups of animals consisted of: P. gingivalis W50 orally inoculated (infected control), a non-bacterial inoculated control, and immunised groups. For the therapeutic vaccination periodontitis model (Figure 6) mice were immunised on day 19 after the first oral inoculation with either the protein vaccine control at 200 μg in
1005272698
116 saline/alum (Alhydrogel; 10mg/mL w/v aluminium hydroxide wet gel suspension; Invivogen) via the intraperitoneal route or with mRNAs formulated in LNPs in Tris/sucrose buffer (25 mM Tris, 8.7% sucrose). Mice receiving the mRNA vaccines were injected with either 30 µg or 3 µg of formulated mRNA intramuscularly in a total volume of 50 µL on the same days as the recombinant protein vaccine. The injection was performed into the right semitendinosis or semimembranosis thigh muscle using a 27G needle. Mice received a second immunisation on day 40 via the subcutaneous route (for the alum- adjuvanted protein vaccine) or intramuscularly (for the mRNA vaccine candidates, left semitendinosis or semimembranosis thigh muscle). On Day 62, mice were bled by cardiac puncture and killed. Maxillae were removed and halved through the midline, with 10 halves used to determine alveolar bone loss. Sera were used to determine the antibody profile using ELISA. [0408] Measurement of alveolar bone loss in mouse maxillae [0409] Maxillae to be examined for bone loss were boiled (1 min) in deionised water, mechanically defleshed, and immersed in 2% w/v potassium hydroxide (16 h, 25°C). Maxillae were washed twice with deionised water (25 °C), dried (1 h, 37 °C) and stained with 0.5% w/v aqueous methylene blue. Digital images of the buccal side of the maxillae were captured with an Olympus DP12 digital camera mounted on a dissecting microscope, using OLYSIA BioReport software version 3.2 (Olympus Australia Pty Ltd, New South Wales, Australia) to assess horizontal bone loss. Maxillae were oriented so that the buccal and lingual molar cusps were superimposed. Images were captured with a micrometre in frame, so that measurements could be standardised for each image. Horizontal bone loss was defined as the loss occurring in a horizontal plane, perpendicular to the alveolar bone crest that resulted in a reduction of the crest height. The visible area from the cemento-enamel junction (CEJ) to the alveolar bone crest (ABC) for each molar was measured using OLYSIA BioReport software version 3.2 imaging software to give the total visible CEJ-ABC area in mm2. P. gingivalis-induced alveolar bone loss in mm2 was calculated by subtracting the total visible CEJ-ABC area of the uninoculated (N-C) group from the total visible CEJ-ABC area of each experimental group. Alveolar bone loss measurements were determined twice in a random and blinded protocol. Data are expressed as the mean +/- standard deviation in mm2 and were analysed using a one-way ANOVA or Kruskal-Wallis test. [0410] Determination of subclass antibody in sera using ELISA
1005272698
117 [0411] ELISAs were performed to evaluate subclass antibody in sera as described in Pathirana et al. (2007). Infect Immun 75: 1436-1442) using a solution of either heat killed W50 cells (10 μg/mL), domain subunits or epitopes (1 μg/mL) in 0.1M PBS (pH 7.4) to coat wells (16 h, 4 ºC) of flat-bottom polyvinyl microtiter plates (Microtiter; Dynatech Laboratories, McLean, VA, US). In these experiments the following antibody dilutions were used; a dilution of 1/4000 dilution of goat anti-mouse; IgG (M8642), IgG1 (M8770), IgG2a (M4434) antibodies (Sigma, New South Wales, Australia). A 1/4000 dilution of a horseradish peroxidase-conjugated swine anti-goat IgG antibody (M5420; Sigma, New South Wales, Australia) was used to develop ELISA experiment. For the epitope ELISAs biotinylated peptides were bound to pre-blocked streptavidin coated flat bottom plates (Pierce; Thermo-Fisher) at 10 μg/mL. Following incubation with sera, the ELISA was developed with 1/4000 goat anti-mouse IgG and 1/4000 horseradish peroxidase- conjugated swine anti-goat IgG antibody. In all ELISA experiments washing was performed between steps using three washes of 220µL of PBS-Tween 20 (0.1% v/v). All optical density measurements were conducted on a Wallac VICTOR31420 Multilabel counter (Perkin Elmer) at 405nm and data were analysed using a one-way ANOVA or Kruskal-Wallis test. [0412] Prophylactic vaccination model [0413] Mice (female C57BL6) were intra-orally inoculated with P. gingivalis [1 x 1010 viable P. gingivalis W50 cells per dose suspended in 20 µL PG buffer [50 mM Tris-HCL, 150 mM NaCl, 10 mM MgSO4 and 14.3 mM mercaptoethanol, pH 7.4 containing 2% (w/v) carboxymethylcellulose]. The inoculum begun on day 42 and was repeated three times a week for 3 weeks. The inocula were prepared anaerobically and then immediately applied to the gingival margin of the maxillary molar teeth. The number of viable bacteria in each inoculum was verified by CFU counts on blood agar. [0414] Groups of animals consisted of: P. gingivalis W50 orally inoculated (infected control); a non-bacterial inoculated control (naive) and vaccinated group. On Day 84, mice were killed and bled by cardiac puncture. Maxillae were removed and halved through the midline, with 10 halves used to determine alveolar bone loss. Sera were used to determine the antibody profile using ELISA. Spleens were removed from mice and stored in prepared in Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 10% (v/v) heat-inactivated FBS (56 °C, 30 min), 2 mM L-glutamine, 2 mM sodium pyruvate, 100 IU/mL penicillin/Strep prior to tissue disruption for the ELISPOT assay.
1005272698
118 [0415] Collection of splenocytes and ELISPOT assay [0416] Mice (female C57BL6) were intra-orally inoculated with P. gingivalis [1 x 1010 viable P. gingivalis W50 cells per dose suspended in 20 µL PG buffer [50 mM Tris-HCL, 150 mM NaCl, 10 mM MgSO4. Single cell suspensions of splenocytes were prepared from spleens using the GentleMACS system according to the manufacturer’s instructions (Miltenyi Biotec) and were then treated with red blood cell lysis buffer for 5 minutes at RT (Sigma-Aldrich Pty. Ltd., New South Wales, Australia) and then washed twice in Dulbecco’s PBS with centrifugation at 800 g, 5 min. Cell suspensions were then filtered through 0.45 µM filters and ready for analysis. ELISPOT assay was performed using the BD® ELISPOT Kits for IL-4 and IFNγ as per the manufacturer’s instructions. 3 x 105 cells were added to each well of ELSIPOT plates with or without restimulation with KDAK-3S-AVQP protein (10 µg/mL). Concanavalin A was used as a positive control mitogen (10 µg/mL). [0417] Plates were incubated at 37ºC in an atmosphere of 5% CO2 in air for 48 hours in a humidified incubator. The plates were then washed and developed as per the manufacturer’s instructions and spots were allowed to develop for 20 – 30 minutes, before stopping the reaction by washing with water. The spots were counted using EliSpot Reader Lite (version 2.9. Autoimmun Diagnostika GmbH, Ebinger Strasse 4, Strassberg, Germany). Data are expressed as spot forming cells per million (SFC/million) and statistically analysed using one-way ANOVA and Dunnett’s 3T test (Graphpad Prism). [0418] Example 2: Expression of mRNA constructs encoding chimeric proteins [0419] The inventors obtained mRNA constructs encoding chimeric proteins containing postulated antigenic domains from the Kgp gingipain protein. The mRNA constructs comprised the general architecture as shown in Figure 2 and the chimeric proteins (antigens) encoded by the mRNA expression constructs comprised the architectures designated KDcAK1n, KDAK and KDAK-3S-AVQP as shown in Figure 1 (amino acid sequences as set forth in SEQ ID NOs: 2, 4 and 6, respectively, mRNA sequences as set forth in SEQ ID NOs: 40, 41 and 42, respectively). [0420] The mRNAs were tested for in vitro antigen expression and secretion in 293T (not shown) or HeLa cells and using various secretion peptides including tPA-derived
1005272698
119 signal peptides and SEAP signal peptide. Cells were transfected with the mRNA constructs using lipofectants as outlined in Example 1. After 48 hours, supernatants were collected and subjected to Western Blot analysis to test for secreted polypeptide. [0421] Figure 3 shows Western Blots demonstrating expression and secretion by HeLa cells of mRNA constructs encoding KDcAK1n, KDAK-3S-AVQP and KDAK. Various signal peptides were tested. The results using the SEAP secretion peptide are shown. [0422] The results indicate that constructs encoding KDcAK1n did not express well, whereas constructs encoding KDAK and KDAK-3S-AVQP expressed abundant amounts of protein which was secreted to the supernatant of the HeLa cells. [0423] Example 3: determining immunogenicity of candidate mRNA vaccines [0424] Figure 4 shows a schematic of the vaccination protocol used to assess the immunogenicity of candidate vaccines. [0425] Briefly, mRNA constructs as described above and encoding chimeras KDcAK1n, KDAK and KDAK-3S-AVQP, were formulated into lipid nanoparticles (LNP) using standard techniques. The LNPs used in these experiments comprised: ALC-0315, cholesterol, distearoylphosphatidylcholine and ALC-0159 in mole percent (%) ratio of: 46.3: 42.7: 9.4: 1.6. [0426] For each antigen, mRNA constructs were generated using either native RNA sequence (unmodified) or N1-methyl-pseudouridine modified (M1ψ) sequence. [0427] Two doses of mRNA vaccine were tested: 30 µg or 3 µg of mRNA formulated in LNP. The positive control used in experiments was the protein KDAK-3S-AVQP adjuvanted with alum. [0428] Mice were intramuscularly immunised with mRNA-LNPs according to the schedule shown in Figure 4. [0429] As outlined in Example 1, following immunisation, sera were collected at Day 35 and tested using ELISA to determine serum antibody subclasses responses of the immunised mice. Antisera were used to probe against the absorbed antigens: KDAK-3S- AVQP (purified recombinant protein), heat killed P. gingivalis strain W50 (HKPg), Kgpcat (catalytic domain of the Kgp gingipain) and a biotinylated linear peptide corresponding to
1005272698
120 sequence encoded by the tested RNA vaccine candidates. The sequence of the biotinylated KAS2 peptide is shown below: KAS2 NTGVSFANYTAHGSETAWADPLLTTSQLKALTNKDK (SEQ ID NO: 8) [0430] Antibody responses are expressed as the ELISA titre obtained minus triple the background level, with each titre representing the mean ± s.d. of the 5 individual mice (Figure 5). Both m1Ψ-KDAK and m1Ψ-KDAK-3S-AVQP induced strong responses to KDAK-3S-AVQP at both 30 µg and 3 µg doses, inducing strong total IgG, IgG1 and IgG2 titres. Native RNAs KDAK and KDAK-3S-AVQP produced strong responses at 30 µg but weaker at 3 µg (particularly KDAK-3S-AVQP). KDcAK1n and m1Ψ-Negative -RNA control produced no significant responses. [0431] m1Ψ-KDAK, m1Ψ-KDAK-3S-AVQP and KDAK at 30 µg provided strong IgG1 responses against HKPg and only m1Ψ-KDAK 30 µg induced significant IgG2 titres. [0432] m1Ψ-KDAK 30 µg, m1Ψ-KDAK-3S-AVQP 30 µg, KDAK 30 µg and m1Ψ-KDAK 3 µg induced significant IgG titres against Kgpcat and m1Ψ-KDAK 30 µg, m1Ψ-KDAK- 3S-AVQP 30 µg, KDAK 30 µg also induced significant IgG1. [0433] This was a surprising finding given that the prior art describes the utility of the protein KDcAK1n in generating robust immune responses to P. gingivalis antigens including to heat killed P. gingivalis, and recombinant Kgpcat protein. In.the present case, the efficacy of the protein vaccine was not recapitulated when using an mRNA vaccine encoding the sequence for the same protein, potentially due to poor expression and poor secretion of the KDcAK1n protein from mRNA constructs. [0434] The results also show that vaccines comprising 30 µg mRNA were generally more immunogenic than vaccines comprising 3 µg mRNA. Similarly, modified (M1ψ) mRNA was more immunogenic than native (unmodified) mRNA. [0435] Example 4: determining in vivo efficacy of mRNA vaccines in periodontitis model [0436] Figure 6 shows a schematic of the vaccination protocol for determining in vivo efficacy of candidate mRNA vaccines (as assessed by protection from alveolar bone loss), and immunogenicity (as determined by antibody levels in sera).
1005272698
121 [0437] Figure 7 shows P. gingivalis induced bone loss (mm) in the different treatment groups. KDAK-3S-AVQP protein was used as a positive control and “infected” mice (no RNA or protein vaccination) were used as a negative control. [0438] mRNA vaccines were prepared and formulated as outlined above and as used in Example 3. [0439] The results show that vaccination with m1Ψ-KDAK 30 µg, m1Ψ-KDAK-3S- AVQP 30 µg mRNA vaccines provided for significant protection against P. gingivalis induced bone loss, similarly to the alum-adjuvanted polypeptide KDAK-3S-AVQP. [0440] KDAK 30 µg and KDAK-3S-AVQP provided partial protection [0441] Antisera were used to probe against the absorbed antigens: recombinant protein KDAK-3S-AVQP, heat killed P. gingivalis strain W50 (HKPg), Kgpcat and biotinylated linear peptides corresponding to sequences encoded by the tested mRNA vaccine candidates. Antibody responses are expressed as the ELISA titre obtained minus triple the background level, with each titre representing the mean ± s.d. of the 10 individual mice (Figure 8). [0442] All tested constructs: m1Ψ-KDAK, m1Ψ-KDAK-3S-AVQP, KDAK and KDAK- 3S-AVQP provided strong total IgG, IgG1 and IgG2a responses against recombinant protein KDAK-3S-AVQP. m1Ψ-KDAK, m1Ψ-KDAK-3S-AVQP and KDAK induced strong total IgG and IgG1 titres against HKPg Only m1Ψ-KDAK induced weak IgG2a responses. [0443] All constructs induced IgG (primarily IgG1) responses against Kgpcat, but only m1ψ-KDAK, m1ψ-KDAK-3S-AVQP and KDAK were statistically significant. [0444] All constructs induced antibodies against the KAS2 peptide. [0445] Example 5: expression and secretion of truncated antigens encoded by alternative mRNA constructs [0446] Various truncated mRNA constructs were assessed, each expressing truncated variations of the protein antigens encoded by the mRNAs tested in the above examples. All mRNAs were m1ψ-modified. [0447] The mRNA constructs tested encoded:
1005272698
122 - K (also referred to herein as KAS or KAS2 active site domain; amino acid sequence SEQ ID NO: 8) - KD (amino acid sequence: SEQ ID NO: 10) - KA (amino acid sequence SEQ ID NO: 18) - DA (amino acid sequence SEQ ID NO: 14) - KDA (amino acid sequence SEQ ID NO: 12) - KDAΔAMB3 (ie wherein the adhesin domain comprises ABMs 2 and 1 but not ABM3; amino acid sequence SEQ ID NO: 16) - RDA (replacing Arginine-dependent gingipain active site KAS, with active site from a Lysine-dependent gingipain RAS) (amino acid sequence: SEQ ID NO: 20) - positive control: KDAK-3S-AVQP. [0448] The mRNAs encoding the target antigens were tested for in vitro expression and secretion in HeLa cells by Western Blot. Figure 9 shows the results of these studies. SEAP signal peptide was encoded at the N-terminus of each antigen. [0449] All constructs expressed well except the construct encoding K (m1Ψ-K) which was poorly expressed (potentially due to protein degradation following expression). The m1Ψ-K construct was initially not detected in either the lysate or the supernatant in the 24 hour transfected samples (Figure 9A). An additional time course analysis of 6 – 48 hours, plus an increased loading (x 3 volume) of m1Ψ-K samples revealed that it was poorly expressed and rapidly degraded (Figure 9B). All other constructs: m1Ψ-KD, m1Ψ-KA, m1Ψ-DA, m1Ψ-KDA, m1Ψ-KDA21 showed good expression and secretion (Figure 9A). [0450] The RDA antigen which was predominantly found in the whole cell lysate rather than supernatant (suggesting poor secretion). [0451] Example 6: in vivo efficacy and immunogenicity of truncated constructs [0452] mRNAs (formulated in LNPs) encoding KDA, KDAΔABM3, DA, KA and KD were assessed in this next study and compared to KDAK-3S-AVQP mRNA and protein vaccines. All mRNAs were m1ψ-modified and were administered at a dose of 30 µg mRNA.
1005272698
123 [0453] The results, shown in Figure 10, indicate that the mRNA vaccine encoding KA, DA, KDAΔABM3 and KDA provided robust protection against P. gingivalis-induced alveolar bone loss. [0454] Vaccines containing m1Ψ-KDAK-3S-AVQP, m1Ψ-KDA21 protected against bone loss in the animal model, similarly to the alum-adjuvanted recombinant protein KDAK-3S-AVQP. m1Ψ-DA and m1Ψ-KA provided partial protection (Figure 10), suggesting the importance of A and K domains for protection. [0455] Antisera were used to probe against the absorbed antigens: recombinant protein KDAK-3S-AVQP heat killed P. gingivalis strain W50 (HKPg) and Kgpcat corresponding to sequences encoded by the tested mRNA vaccine candidates. [0456] Antibody responses are expressed as the ELISA titre obtained minus triple the background level, with each titre representing the mean ± s.d. of the 10 individual mice. All tested constructs induced strong total IgG, IgG1 and IgG2 responses against KDAK- 3S-AVQP (Figure 11). [0457] All tested constructs induced strong total IgG and IgG1 titres against HKPg. Only m1Ψ-DA and m1Ψ-KDA21 induced IgG2a responses. [0458] All constructs except m1Ψ-DA (as it doesn’t contain the K domain that is part of the Kgpcat) induced IgG (primarily IgG1) responses against Kgpcat, but the response of m1Ψ-KDAK-3S-AVQP was weaker and did not show statistical significance for IgG1. Only m1Ψ-KDA21 and, m1Ψ-KDAK induced IgG2 titres. [0459] All constructs except m1Ψ-KDA21 (as it doesn’t contain the KAS2 epitope) induced antibodies against the KAS2 peptide. [0460] Example 7: further assessment of immunogenicity of truncated constructs [0461] A further series of immunogenicity assessments were performed using constructs: m1Ψ-K 30 µg and m1Ψ-RDA 30 µg. [0462] Mice were intramuscularly immunised with the mRNA-LNPs encoding the target antigens according to the schedule shown in in Figure 4. Serum antibody subclass responses of immunised mice were examined by ELISA. Antisera were used to probe
1005272698
124 against the absorbed antigens: KDAK-3S-AVQP polypeptide, heat killed P. gingivalis strain W50 (HKPg), and Kgpcat. [0463] Antibody responses are expressed as the ELISA titre obtained minus triple the background level, with each titre representing the mean ± s.d. of the 5 individual mice (Figure 12). Only m1Ψ-RDA induced strong responses to KDAK-3S-AVQP inducing strong total IgG, IgG1 and IgG2 titres. It also induced significant total IgG primarily IgG1 but not IgG2, against HKPg and significant IgG1 titres against Kgpcat. It also induced antibody titres against the KAS2 and ABM3 peptides. m1Ψ-K showed no immunogenicity, likely due to poor expression/stability as described above. [0464] Example 8: In vivo efficacy and immunogenicity in a prophylactic model [0465] mRNAs (formulated in LNPs) encoding KDAK-3S-AVQP were assessed in the prophylactic vaccination model (see Figure 13) and compared to protein vaccine. The mRNA was m1ψ-modified and was administered at a dose of 30 µg mRNA. [0466] The results, shown in Figure 15, indicate that the mRNA vaccine provided robust protection against P. gingivalis-induced alveolar bone loss in the prophylactic model of vaccination [0467] The mRNA vaccine containing m1Ψ-KDAK-3S-AVQP protected against bone loss in the animal model, similarly to the alum-adjuvanted recombinant protein KDAK- 3S-AVQP (Figure 15). [0468] Antisera were used to probe against the absorbed antigens: recombinant protein KDAK-3S-AVQP, heat killed P. gingivalis strain W50 (HKPg) and RgpA-Kgp complex. [0469] Antibody responses are expressed as the ELISA titre obtained minus triple the background level, with each titre representing the mean ± s.d. of the 10 individual mice. The tested mRNA construct induced strong total IgG, IgG1 and IgG2 responses against KDAK-3S-AVQP, HK-Pg and RgpA-Kgp complex (Figure 16). [0470] Ex vivo restimulation of recovered splenocytes with KDAK-3S-AVQP protein, in an ELISpot assay, demonstrated the presence of KDAK-3S-AVQP-specific interferon gamma secreting CD4+ T cells (Figure 14).
1005272698
125 [0471] It will be understood that the invention disclosed and defined in this specification extends to all alternative combinations of two or more of the individual features mentioned or evident from the text or drawings. All of these different combinations constitute various alternative aspects of the invention.
Claims
1005272698
126 CLAIMS 1. An RNA polynucleotide encoding a protein comprising or consisting of: - one or more amino acid sequences of an active site of an Arg- or Lys- gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and/or - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. wherein the polynucleotide is capable of being translated in a mammalian cell. 2. The RNA of claim 1, wherein the protein encoded by the RNA comprises or consists of: - one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. 3. The RNA of claim 1 or 2 wherein the protein encoded by the RNA polynucleotide further comprises: - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. 4. The RNA of any one of the preceding claims, wherein the protein encoded by the RNA polynucleotide is a chimeric or fusion protein comprising or consisting of: - one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto.
1005272698
127 5. The RNA of any one of the preceding claims, wherein the RNA encodes an amino acid sequence of an active site of an Arg-gingipain of P. gingivalis comprising the amino acid sequence of SEQ ID NO: 38 (eg encoded by the RNA sequence as set forth in SEQ ID NO: 50), or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto. 6. The RNA of any one of the preceding claims, wherein the RNA encodes an amino acid sequence of an active site of a Lys-gingipain of P. gingivalis comprising the amino acid sequence of SEQ ID NO: 8, (eg encoded by the RNA sequence as set forth in SEQ ID NO: 43), or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto. 7. The RNA of any one of the preceding claims, wherein the RNA encodes a chimeric or fusion protein that comprises: i) an amino acid sequence that comprises or consists of an amino acid sequence of the active site of an Arg-gingipain of P. gingivalis, or sequences that are at least 80% identical thereto; and ii) an amino acid sequence that comprises or consists of an amino acid sequence of the active site of a Lys-gingipain of P. gingivalis, or sequences that are at least 80% identical thereto. 8. The RNA of any one of the preceding claims wherein the RNA encodes a chimeric or fusion protein that comprises at least two amino acid sequences that comprise or consist of an amino acid sequence of the active site of an Arg- or Lys- gingipain of P. gingivalis, or sequences that are at least 80% identical thereto. 9. The RNA of claim 8, wherein the at least two amino acid sequences are located contiguously in the chimeric or fusion protein.
1005272698
128 10. The RNA of claim 8, wherein one of the at least two amino acid sequences is located at the N terminus of the chimeric or fusion protein and the second of the at least two amino acid sequences is located at the C-terminus of the chimeric or fusion protein. 11. The RNA of claim 8, wherein one of the at least two amino acid sequences is located at the N or C terminus of the chimeric or fusion protein and the second of the at least two amino acid sequences is located within the chimeric or fusion protein. 12. The RNA of claim 8, wherein the at least two amino acid sequences are (both) located at the N terminus of the chimeric or fusion protein or the at least two amino acid sequences are (both) located at the C-terminus of the chimeric or fusion protein. 13. The RNA of any one of claims 4 to 12, wherein the chimeric or fusion protein encoded by the RNA polynucleotide further comprises: - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. 14. The RNA of claim 13, wherein the RNA encodes: - one or more amino acid sequences of an active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and wherein the amino acid sequence of a DUF2436 domain is located between an amino acid sequence of an active site of the gingipain of P. gingivalis and the amino acid sequence of the or more adhesin binding motifs (ABMs). 15. The RNA of claim 13 or 14, wherein RNA encodes an amino acid sequence of a DUF2436 domain of an Arg or Lys gingipain of P. gingivalis comprising or consisting of the amino acid sequence of SEQ ID NO: 35, (eg encoded by the RNA sequence as set forth in SEQ ID NO: 50), or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto.
1005272698
129 16. The RNA of any one of claims 13 to 15, wherein the RNA encodes a substitution of a cysteine residue in the DUF2436 domain, such as a substitution to a serine or valine residue. 17. The RNA of claim 16, wherein the RNA encodes an amino acid sequence of a DUF2436 domain comprising a cysteine to serine substitution (such as shown in SEQ ID NO: 36). 18. The RNA of any one of the preceding claims wherein the RNA encodes one or more adhesin binding motifs (ABMs) comprising or consisting of the amino acid sequence of ABM2 and ABM1. 19. The RNA of claim 18, wherein the RNA encodes the amino acid sequence of a polypeptide comprising ABM2 and ABM1 as set forth in SEQ ID NO: 22 and SEQ ID NO: 21, respectively, or comprising the amino acid sequence as set forth in SEQ ID NO: 24 (ABM2+1), or a sequence at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto. 20. The RNA of claim 19, wherein the RNA comprises the nucleotide sequence as set forth in any one of SEQ ID NOs: 52 to 55. 21. The RNA of any one of claims 18 to 20, wherein the RNA encodes one or more adhesin binding motifs (ABMs) comprising or consisting of the amino acid sequence of ABM2, ABM1 and ABM3. 22. The RNA of claim 21, wherein the RNA encodes the amino acid sequence as set forth in SEQ ID NO: 25 (eg encoded by an RNA comprising the sequence of SEQ ID NO: 56). 23. The RNA of any one of claims 18 to 22, wherein the RNA encodes one or more adhesin binding motifs comprising one or more modifications selected from: a) one or more cysteine amino acid substitutions compared to the naturally occurring Arg- or Lys-gingipain sequences in corresponding regions;
1005272698
130 b) substitution of the proline and/or an asparagine residues in the sequence PxxN corresponding to, or at a position equivalent to, residues 6 to 9 of the sequence of SEQ ID NO: 21 (ABM1); c) substitution of the motif NxFA to SxYQ in the sequence, corresponding to, or at a position equivalent to residues 2 to 5 of the sequence of SEQ ID NO: 21 (ABM1); d) substitution of the second tyrosine residue, corresponding to or at a position equivalent to residues at position 5 of SEQ ID NO: 22 (ABM2), and of the tryptophan residue, corresponding to or at a position equivalent to residue at position 23 of SEQ ID NO: 21 (ABM1) to alanine residues. 24. The RNA of claim 23, wherein the RNA encodes one or more adhesin binding motifs comprising a substitution of one or more cysteine residues to a serine residue or to a valine residue. 25. The RNA of claim 24, wherein the RNA encodes one or more adhesin binding motifs that comprise substitution of all cysteine residues to serine or valine residues. 26. The RNA of any one of claims 23 to 25, wherein the RNA encodes one or more adhesin binding motifs that comprise a proline and/or asparagine substitution of the motif PxxN (eg PVQN, SEQ ID NO: 106), corresponding to or at a position equivalent to residues 6 to 9 of SEQ ID NO: 21. 27. The RNA of claim 26, wherein the proline amino acid substitution is a substitution to an alanine residue. 28. The RNA of claim 26 or 27, wherein the asparagine amino acid substitution is a substitution to a proline residue or an alanine residue. 29. The RNA of any one of claims 23 to 28, wherein the RNA encodes one or more adhesin binding motifs comprising a substitution from PxxN to AxxP, (eg AVQP, SEQ ID NO: 107) in the region encoding the ABM1 (such as exemplified in the amino acid sequences of SEQ ID NOs: 30 to 32). 30. The RNA of any one of claims 23 to 29, wherein the RNA encodes one or more adhesin binding motifs that comprise the amino acid sequence as set forth in any one of SEQ ID NOs: 21 to 25, and comprising:
1005272698
131 a) one or more cysteine amino acid substitutions compared to the naturally occurring Arg- or Lys-gingipain sequences in corresponding regions, preferably substitution of all cysteine residues; and b) substitution of the motif PxxN corresponding to, or at a position equivalent to, residues 6 to 9 of the sequence of SEQ ID NO: 21 (ABM1), to AxxP. 31. The RNA of claim 30, wherein the RNA encodes one or more adhesin binding motifs comprising or consisting of the amino acid sequence as set forth in any one of SEQ ID NOs: 26 to 34, or sequences at least 80%, at least 81%, at least 82%, at least 83%, at least 84%, at least 85%, at least 86%, at least 87%, at least 88%, at least 98%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98% or at least 99% identical thereto, provided that the sequences comprise the aforementioned substitutions of the cysteine and proline and asparagine residues. 32. An RNA polynucleotide encoding a chimeric or fusion protein comprising or consisting of: - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, 33. The RNA of claim 32, wherein the RNA encodes a chimeric or fusion protein comprising or consisting of the amino acid sequence of SEQ ID NO: 18 or 83 or SEQ ID NO: 39 or 81, or SEQ ID NO: 58 to 63, or SEQ ID NO: 82 to 87. 34. An RNA polynucleotide encoding a chimeric or fusion protein comprising or consisting of: - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, and
1005272698
132 - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. 35. The RNA of claim 34, wherein the RNA encodes a chimeric or fusion protein comprising or consisting of the amino acid sequence of the amino acid sequence of SEQ ID NO: 12, 16 or 20. 36. The RNA of claim 34, wherein the RNA encodes a chimeric or fusion protein comprising or consisting of the amino acid sequence of SEQ ID NO: 4. 37. An RNA polynucleotide encoding a chimeric or fusion protein comprising or consisting of: - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. 38. The RNA of claim 37, wherein the RNA encodes a chimeric or fusion protein comprising or consisting of the amino acid sequence of SEQ ID NO: 14. 39. An RNA polynucleotide encoding a chimeric or fusion protein comprising or consisting of: - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; and - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. 40. The RNA of claim 39, wherein the RNA encodes a chimeric or fusion protein comprising or consisting of the amino acid sequence of SEQ ID NO: 10 or 108. 41. An RNA polynucleotide encoding a chimeric or fusion protein comprising or consisting of:
1005272698
133 - one or more amino acid sequences of active site of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto; - the amino acid sequence of a DUF2436 domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto, and - the amino acid sequence of one or more adhesin binding motifs (ABMs) of an adhesin domain of an Arg- or Lys-gingipain of P. gingivalis, or a sequence that is at least 80% identical thereto. 42. The RNA of claim 41, wherein the RNA encodes a chimeric or fusion protein comprising or consisting of the amino acid sequence of SEQ ID NO: 6. 43. An RNA polynucleotide comprising or consisting of a nucleotide sequence encoding a protein comprising or consisting of the amino acid sequence of any one of: SEQ ID NO: 2, SEQ ID NO: 3 or SEQ ID NO: 38. 44. An RNA comprising or consisting of a nucleotide sequence of any one of: a) SEQ ID NO: 48, 57, b) SEQ ID NO: 45, 47 or 49 or SEQ ID NO: 41; c) SEQ ID NO: 46; d) SEQ ID NO: 44; e) SEQ ID NO: 42. 45. An RNA comprising or consisting of a nucleotide sequence of any one of: SEQ ID NO: 40, SEQ ID NO: 43 or SEQ ID NO: 50. 46. The RNA of any one of the preceding claims, wherein the RNA is an mRNA. 47. The RNA of any one of the preceding claims wherein the RNA further encodes an N-terminal signal peptide for enabling secretion of the protein following translation thereof. 48. The RNA of any one of the preceding claims, wherein the RNA further comprises a 5’ untranslated region (UTR) and/or a 3’ UTR.
1005272698
134 49. The RNA of any one of the preceding claims, wherein the RNA also comprises a 5’ cap analog, such as 7mG(5′)ppp(5′)NlmpNp. 50. The RNA of any one of the preceding claims wherein the RNA also comprises a polyadenine (polyA) tail. 50. The RNA of any one of the preceding claims, wherein the RNA comprises a chemical modification, preferably wherein the chemical modification is a 1- methylpseudouridine modification or a 1-ethylpseudouridine modification. 52. The RNA of any one of the preceding claims wherein the RNA has a uridine content of less than about 50%, less than about 45%, less than about 40%, less than about 35%, less than about 30%, less than about 25%, less than about 20% or less than about 15%. 53. The RNA of any one of the preceding claims, wherein the uridines in the RNA are replaced with a chemical modification such as N-methyl-pseudouridine. 54. The RNA of claim 53, wherein at least 25%, at least 30%, at least 35%, at least 40%, at least 45%, at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 95%, or 100% of the uridine nucleosides are replaced with N-methyl-pseudouridine. 55. The RNA of any one of the preceding claims wherein the RNA is in the form of a codon optimised RNA molecule. 56. A composition comprising an RNA of any one of the preceding claims. 57. A lipid nanoparticle composition comprising an RNA of any one of claims 1 to 55. 58. The lipid nanoparticle composition of claim 57, comprising: - a cationic and/or ionisable lipid comprising from about 25 % to about 75 mol % of the total lipid present in the nanoparticle; - a sterol (structural lipid) comprising from about 5 mol % to about 60 mol % of the total lipid present in the nanoparticle;
1005272698
135 - a phospholipid comprising from about 5 mol % to about 50 mol % of the total lipid present in the nanoparticle; - a PEGylated lipid comprising from about 0.5 mol % to 20 mol % of the total lipid present in the nanoparticle. 59. The lipid nanoparticle composition of claim 57 or 58, comprising: - an ionisable lipid in the form of [(4-hydroxybutyl)azanediyl]di(hexane-6,1-diyl) bis(2-hexyldecanoate) (ALC-0315), - a sterol in the form of cholesterol, - a phospholipid in the form of distearoylphosphatidylcholine (DSPC), and - a PEGylated lipid in the form of 2-[(polyethylene glycol)-2000]-N,N- ditetradecylacetamide (ALC-0159). 60. The lipid nanoparticle composition of claim 59, wherein the lipids are present in the lipid nanoparticle at molar lipid ratios (%) of 46.3 ALC-0315: 42.7 cholesterol : 9.4 DSPC : 1.6 ALC-0159, optionally in Tris/sucrose buffer (25 mM Tris pH 7.4, 8.8% sucrose w/v). 61. A method for producing a lipid nanoparticle comprising an RNA of any one of claims 1 to 55, wherein preferably the method comprises formulating any RNA molecule of any one of claims 1 to 55, with one or more lipids useful for producing a lipid nanoparticle. 62. The method of claim 61, wherein the lipid components comprise a phospholipid, a PEG lipid, and a structural lipid. 63. A nucleic acid construct or vector, comprising an RNA polynucleotide of any one of claims 1 to 55. 64. A method for eliciting an immune response P. gingivalis in a subject in need thereof, the method comprising administering to the subject, an RNA of any one of claims 1 to 55 or a nanoparticle or composition of any one of claims 56 to 60.
1005272698
136 65. A method for eliciting an immune response to P. gingivalis in a subject in need thereof, the method comprising administering to the subject, a nanoparticle composition comprising: - an RNA of any one of claims 1 to 55, - an agent for enabling delivery of the RNA to a cell of the subject; - wherein said RNA is capable of being translated in a cell of the subject to produce the polypeptide encoded by the polynucleotide. 66. A method for producing a chimeric or fusion protein in a mammalian cell, the method comprising contacting the mammalian cell with a nanoparticle composition, the composition comprising: - an RNA of any one of claims 1 to 55, - an agent for enabling delivery of the RNA to a cell of the subject; - wherein said RNA is capable of being translated in the mammalian cell to produce the protein. 67. A method for delivering an RNA to a mammalian cell in a subject in need thereof, said method comprising administering to a subject in need thereof, a nanoparticle composition, the composition comprising: - an RNA comprising a polynucleotide sequence of any one of claims 1 to 55, - an agent for enabling delivery of the RNA to a cell of the subject; - wherein said RNA is capable of being translated in the mammalian cell to produce the chimeric or fusion protein described herein; wherein the administering comprises contacting said mammalian cell with the nanoparticle composition, thereby enabling delivery of the RNA to the mammalian cell. 68. The method of any one of claims 65 to 67, wherein the agent for enabling delivery of the RNA to a cell of the subject is a lipid.
1005272698
137 69. The method of claim 68, wherein the lipid comprises a cationic and/or ionisable lipid, a phospholipid, a PEG lipid, and a structural lipid. 70. Use of an RNA of any one of claims 1 to 55, or a vector, or nanoparticle comprising the same, in the manufacture of a composition for eliciting an immune response to P. gingivalis in a subject. 71. Use of i) a lipid component, preferably comprising a cationic and/or ionisable lipid, a phospholipid, a PEG lipid, and a structural lipid., and ii) an RNA of any one of claims 1 to 55, in the manufacture of a composition for delivering the RNA to a mammalian cell in a subject in need thereof.
Priority Applications (1)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| AU2024301059A AU2024301059A1 (en) | 2023-07-26 | 2024-07-26 | Rna vaccines |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| AU2023902379A AU2023902379A0 (en) | 2023-07-26 | RNA vaccines | |
| AU2023902379 | 2023-07-26 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2025019901A1 true WO2025019901A1 (en) | 2025-01-30 |
Family
ID=94373846
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/AU2024/050795 Pending WO2025019901A1 (en) | 2023-07-26 | 2024-07-26 | Rna vaccines |
Country Status (2)
| Country | Link |
|---|---|
| AU (1) | AU2024301059A1 (en) |
| WO (1) | WO2025019901A1 (en) |
Citations (8)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2001047961A1 (en) * | 1999-12-24 | 2001-07-05 | Csl Limited | P. gingivalis antigenic composition |
| WO2010022463A1 (en) * | 2008-08-29 | 2010-03-04 | Oral Health Australia Pty Ltd | Prevention, treatment and diagnosis of p.gingivalis infection |
| WO2014183168A1 (en) * | 2013-05-14 | 2014-11-20 | Oral Health Australia Pty Ltd | Inhibitory propeptides |
| WO2016191389A2 (en) * | 2015-05-27 | 2016-12-01 | Merial, Inc. | Compositions containing antimicrobial igy antibodies, for treatment and prevention of disorders and diseases caused by oral health compromising (ohc) microorganisms |
| WO2021156267A1 (en) * | 2020-02-04 | 2021-08-12 | Curevac Ag | Coronavirus vaccine |
| WO2023137522A1 (en) * | 2022-01-20 | 2023-07-27 | Denteric Pty Ltd | Chimeric polypeptides |
| WO2023137521A1 (en) * | 2022-01-20 | 2023-07-27 | Denteric Pty Ltd | Chimeric polypeptides |
| WO2023137519A1 (en) * | 2022-01-20 | 2023-07-27 | Denteric Pty Ltd | Chimeric polypeptides |
-
2024
- 2024-07-26 AU AU2024301059A patent/AU2024301059A1/en active Pending
- 2024-07-26 WO PCT/AU2024/050795 patent/WO2025019901A1/en active Pending
Patent Citations (8)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2001047961A1 (en) * | 1999-12-24 | 2001-07-05 | Csl Limited | P. gingivalis antigenic composition |
| WO2010022463A1 (en) * | 2008-08-29 | 2010-03-04 | Oral Health Australia Pty Ltd | Prevention, treatment and diagnosis of p.gingivalis infection |
| WO2014183168A1 (en) * | 2013-05-14 | 2014-11-20 | Oral Health Australia Pty Ltd | Inhibitory propeptides |
| WO2016191389A2 (en) * | 2015-05-27 | 2016-12-01 | Merial, Inc. | Compositions containing antimicrobial igy antibodies, for treatment and prevention of disorders and diseases caused by oral health compromising (ohc) microorganisms |
| WO2021156267A1 (en) * | 2020-02-04 | 2021-08-12 | Curevac Ag | Coronavirus vaccine |
| WO2023137522A1 (en) * | 2022-01-20 | 2023-07-27 | Denteric Pty Ltd | Chimeric polypeptides |
| WO2023137521A1 (en) * | 2022-01-20 | 2023-07-27 | Denteric Pty Ltd | Chimeric polypeptides |
| WO2023137519A1 (en) * | 2022-01-20 | 2023-07-27 | Denteric Pty Ltd | Chimeric polypeptides |
Non-Patent Citations (1)
| Title |
|---|
| ZHENG D ET AL.: "Enhancing specific-antibody production to the ragB vaccine with GITRL that expand Tfh, IFN-y(+) T cells and attenuates Porphyromonas gingivalis infection in mice", PLOS ONE, vol. 8, no. 4, 2013, pages 59604, XP093029848, DOI: 10.1371/journal.pone.0059604 * |
Also Published As
| Publication number | Publication date |
|---|---|
| AU2024301059A1 (en) | 2026-01-08 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US12208288B2 (en) | Betacoronavirus RNA vaccines | |
| US10695419B2 (en) | Human cytomegalovirus vaccine | |
| US10383937B2 (en) | Human cytomegalovirus RNA vaccines | |
| US20200038499A1 (en) | Rna bacterial vaccines | |
| WO2018170245A1 (en) | Broad spectrum influenza virus vaccine | |
| WO2017015457A1 (en) | Ebola vaccine | |
| US20250099572A1 (en) | Vaccine compositions | |
| WO2023111907A1 (en) | Polynucleotide compositions and uses thereof | |
| WO2025019901A1 (en) | Rna vaccines | |
| WO2025019903A1 (en) | Rna vaccines for use in animal health | |
| CA3146392A1 (en) | Compositions and methods for the prevention and/or treatment of covid-19 | |
| HK40076103A (en) | Metapneumovirus mrna vaccines | |
| HK40038642A (en) | Respiratory virus vaccines |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24844114 Country of ref document: EP Kind code of ref document: A1 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: AU2024301059 Country of ref document: AU |
|
| ENP | Entry into the national phase |
Ref document number: 2024301059 Country of ref document: AU Date of ref document: 20240726 Kind code of ref document: A |